The full dataset viewer is not available (click to read why). Only showing a preview of the rows.
Error code: JobManagerCrashedError
Need help to make the dataset viewer work? Make sure to review how to configure the dataset viewer, and open a discussion for direct support.
name int64 | portal_run_id int64 | split string |
|---|---|---|
3,434 | 3,434 | test |
3,466 | 3,466 | train |
3,473 | 3,473 | train |
3,474 | 3,474 | train |
3,476 | 3,476 | train |
3,484 | 3,484 | train |
3,523 | 3,523 | train |
3,494 | 3,494 | train |
3,503 | 3,503 | train |
4,635 | 4,635 | train |
4,636 | 4,636 | train |
4,627 | 4,627 | test |
4,641 | 4,641 | train |
4,643 | 4,643 | train |
6,973 | 6,973 | train |
6,951 | 6,951 | test |
6,980 | 6,980 | train |
6,985 | 6,985 | train |
7,004 | 7,004 | train |
8,047 | 8,047 | train |
8,049 | 8,049 | train |
8,017 | 8,017 | train |
8,067 | 8,067 | train |
8,063 | 8,063 | test |
8,070 | 8,070 | train |
8,074 | 8,074 | train |
8,034 | 8,034 | train |
8,038 | 8,038 | train |
8,057 | 8,057 | train |
8,078 | 8,078 | train |
8,079 | 8,079 | train |
7,981 | 7,981 | test |
7,996 | 7,996 | train |
8,004 | 8,004 | train |
8,417 | 8,417 | train |
8,418 | 8,418 | train |
8,421 | 8,421 | train |
8,379 | 8,379 | test |
8,389 | 8,389 | train |
8,403 | 8,403 | train |
8,411 | 8,411 | train |
8,397 | 8,397 | train |
8,425 | 8,425 | train |
8,448 | 8,448 | train |
8,432 | 8,432 | test |
8,436 | 8,436 | train |
8,455 | 8,455 | train |
8,499 | 8,499 | train |
8,494 | 8,494 | train |
8,507 | 8,507 | train |
8,513 | 8,513 | train |
10,264 | 10,264 | train |
10,265 | 10,265 | train |
10,266 | 10,266 | test |
10,269 | 10,269 | train |
10,270 | 10,270 | train |
10,252 | 10,252 | test |
10,258 | 10,258 | train |
10,259 | 10,259 | train |
10,280 | 10,280 | train |
10,285 | 10,285 | train |
11,792 | 11,792 | train |
11,788 | 11,788 | train |
11,790 | 11,790 | train |
11,803 | 11,803 | train |
11,776 | 11,776 | test |
11,782 | 11,782 | test |
11,796 | 11,796 | train |
11,799 | 11,799 | train |
11,804 | 11,804 | train |
11,811 | 11,811 | train |
13,208 | 13,208 | train |
13,184 | 13,184 | test |
13,180 | 13,180 | train |
13,181 | 13,181 | train |
13,185 | 13,185 | train |
13,186 | 13,186 | train |
13,195 | 13,195 | train |
13,196 | 13,196 | train |
13,190 | 13,190 | train |
Dataset Card for POPSICLE
POPSICLE (Particle/Object Picking & Segmentation In CryoET Learning & Evaluation) is a benchmark suite for cryo-electron tomography (cryoET) covering both dense voxel-wise segmentation of cellular structures and sparse localization of macromolecular complexes. The suite is built directly on the CryoET Data Portal: each sub-benchmark in this repository is a thin mlcroissant manifest plus CSV indices over data that already lives on the portal. Tools that speak the copick/Croissant interface stream the raw data straight from the portal.
Quickstart
Load and visualize each sub-benchmark in one click — demos.ipynb lives
at the root of this repo.
Dataset Details
Dataset Description
POPSICLE bundles four sub-benchmarks chosen to span the main design axes of cryoET evaluation: eukaryotic and prokaryotic systems, controlled and fully in situ imaging settings, and both dense voxel-wise segmentation and sparse macromolecular localization. Each sub-benchmark inherits its splits, annotations, and licensing from the underlying CryoET Data Portal deposition; this repository contributes the unified manifest layer (splits, canonical class names, ground-truth filtering, evaluation protocol) so models can be trained and compared under one roof.
| Sub-benchmark | Task | Organism / sample | # train | # test |
|---|---|---|---|---|
| Phantom | Localization | Lysate | 7 | 485 |
| Yeast | Segmentation | S. pombe | 16 | 4 |
| Bacterial | Segmentation | Prokaryote (8 genera) | 68 | 12 |
| MotorBench | Localization | V. cholerae (test) + multi-genus prokaryotes (train) | 1,559 | 843 |
- Curated by: Authors of the POPSICLE benchmark paper (NeurIPS 2026, under review).
- Shared by: Biohub Dynamic Structural Biology / CryoET Data Portal community.
- License: CC0-1.0 (Public Domain Dedication) — covers both the manifests + CSVs in this repo and the underlying tomograms and annotations on the CryoET Data Portal depositions linked above. No attribution requirement; we still appreciate citation of the POPSICLE paper and the benchmark publication.
Dataset Sources
- Repository: https://huggingface.co/datasets/biohub/popsicle
- Paper: POPSICLE (NeurIPS 2026 submission).
- Underlying datasets:
- DS-10440 (Phantom train),
- DS-10445 (Phantom public test),
- DS-10446 (Phantom private test),
- CZCDP-10350 (Bacterial),
- CZCDP-10351 (Yeast; underlying tomograms in DS-10000 / DS-10001),
- CZCDP-10332/CZCDP-10347 (MotorBench train/test).
- Data Portal: https://cryoetdataportal.czscience.com (Ermel et al., Nature Methods 21:2200–2202, 2024).
- Tooling: copick reads each Croissant manifest and resolves data URLs against the portal.
Uses
Direct Use
- Training and evaluating 3D segmentation networks (nnU-Net, MedNeXt, SwinUNETR, Octopi, …) on cellular cryoET tomograms.
- Training and evaluating particle-picking / localization methods (DeepFinder, DeepETPicker, Octopi, segmentation-as-localization approaches, …) on multi-class macromolecular detection problems.
- Benchmarking foundation / promptable / multi-task models that must generalize across both segmentation and localization regimes — model rankings on POPSICLE differ substantially across tasks, exposing inductive-bias trade-offs that single-task evaluations miss.
Out-of-Scope Use
- POPSICLE is not a structure-determination dataset: per-particle picks are coordinates suitable for downstream subtomogram averaging, but no aligned 3D maps or refined orientations are released here.
- The Phantom training split is intentionally tiny (7 tomograms, reflecting realistic annotation budgets). It should not be used to estimate large-model scaling behavior.
- The benchmark is designed for evaluation; using the test/val splits to tune hyperparameters defeats its purpose.
Dataset Structure
Each sub-benchmark lives in its own subdirectory and is self-contained:
popsicle/
phantom/
Croissant/
metadata.json # Croissant 1.1 JSON-LD
runs.csv # name, portal_run_id, split
voxel_spacings.csv # run, voxel_size
tomograms.csv # run, voxel_size, tomo_type, url, ...
features.csv
picks.csv # run, user_id, session_id, object_name, url, sha256, ...
meshes.csv
segmentations.csv # run, voxel_size, user_id, session_id, name, is_multilabel, url, ...
objects.csv
bacterial/ # segmentation across 8 bacterial genera
yeast/ # segmentation in *S. pombe*
motorbench/ # flagellar-motor localization
Phantom — multi-class macromolecular localization
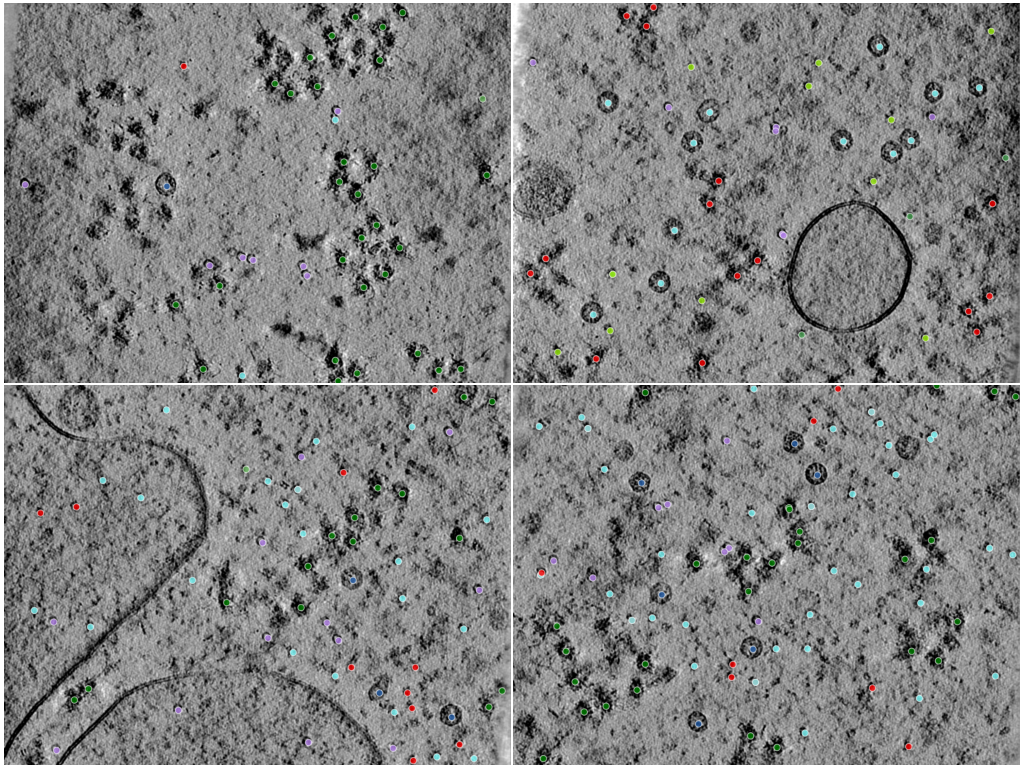
Phantom is built from an experimentally acquired lysate sample enriched for lysosomal components, with additional purified targets introduced to control object diversity and class balance (Peck et al. 2025). The six particle classes span over an order of magnitude in molecular weight (~268–4300 kDa) and have visibly distinct shapes, encouraging detection methods that generalize across particle scale and morphology.
The dataset is the basis of the CZII — CryoET Object Identification Kaggle challenge; splits and the F4 evaluation protocol below mirror that challenge.
| Class | Approx. MW | Portal aliases |
|---|---|---|
apo-ferritin |
~480 kDa | ferritin-complex |
beta-amylase |
~268 kDa | — |
beta-galactosidase |
~470 kDa | — |
ribosome |
~3300 kDa | cytosolic-ribosome |
thyroglobulin |
~660 kDa | — |
virus-like-particle |
~4300 kDa | virus-like-capsid |
Splits
| Split | CDP dataset | # tomograms | Notes |
|---|---|---|---|
train |
DS-10440 | 7 | Public training set |
val |
DS-10445 | 121 | CZII Kaggle "public test" set |
test |
DS-10446 | 364 | CZII Kaggle "private test" set |
Picks are restricted to the original ground-truth annotations. Community-derived submissions hosted on the same CDP datasets, including Kaggle challenge entries, are intentionally excluded so the benchmark scores against the canonical reference labels.
Loading with copick
(click to expand)
import copick
import numpy as np
import matplotlib.pyplot as plt
root = copick.from_croissant(
"https://huggingface.co/datasets/biohub/popsicle/resolve/main/phantom/Croissant/metadata.json",
overlay_root="/tmp/popsicle-overlay",
static_fs_args={"anon": True}, # public portal bucket
)
# Loop over every split and every run
for split, run_names in root.splits.items():
print(f"{split}: {len(run_names)} runs")
for run in root.get_runs_in_split(split):
for pick_set in run.picks:
print(f" {run.name} {pick_set.pickable_object_name}: {len(pick_set.points)} pts")
# Visualize: midplane slice of one tomogram with picks overlaid as scatter
run = root.get_runs_in_split("train")[0]
vs = run.voxel_spacings[0] # 10 Å for Phantom
tomo = vs.tomograms[0] # any tomo_type at this voxel spacing
arr = tomo.numpy() # (Z, Y, X) — streams from portal S3
z_mid = arr.shape[0] // 2
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(arr[z_mid], cmap="gray")
for pick_set in run.picks:
# CopickPoint locations are in physical units (Å); convert to voxel indices.
pts = np.array(
[(p.location.x, p.location.y, p.location.z) for p in pick_set.points]
) / vs.voxel_size
near = pts[np.abs(pts[:, 2] - z_mid) < 5] # within ±5 voxels of the slice
ax.scatter(near[:, 0], near[:, 1], s=24, label=pick_set.pickable_object_name)
ax.legend(loc="upper right", fontsize=8)
ax.set_title(f"{run.name} z={z_mid}")
plt.show()
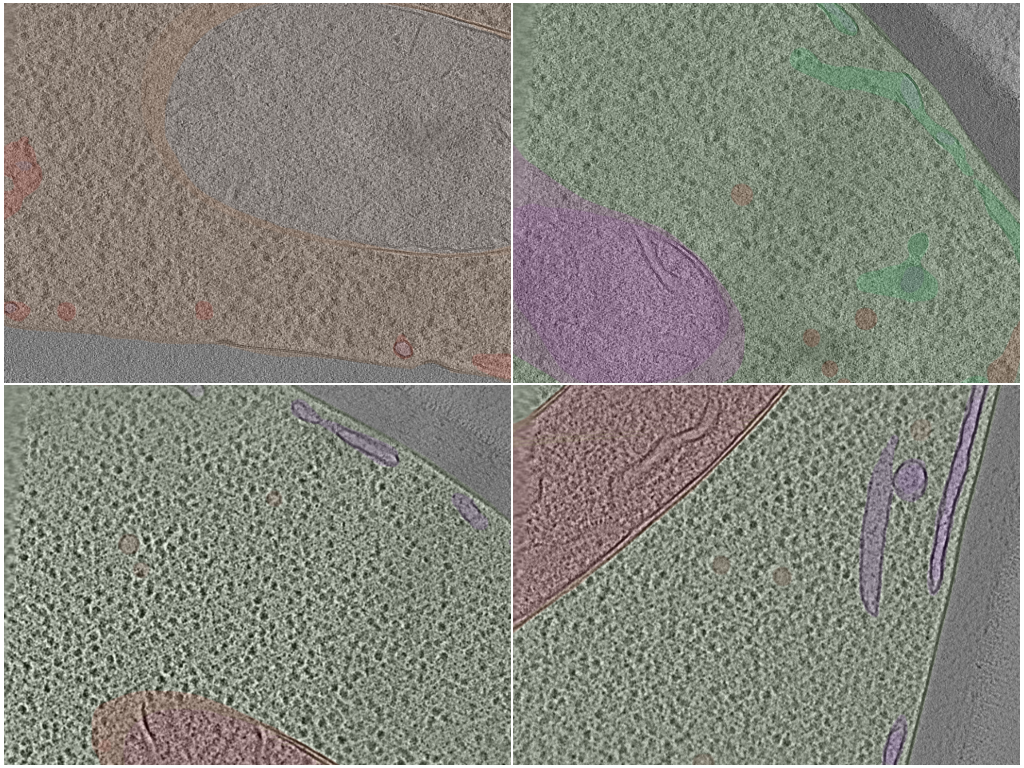
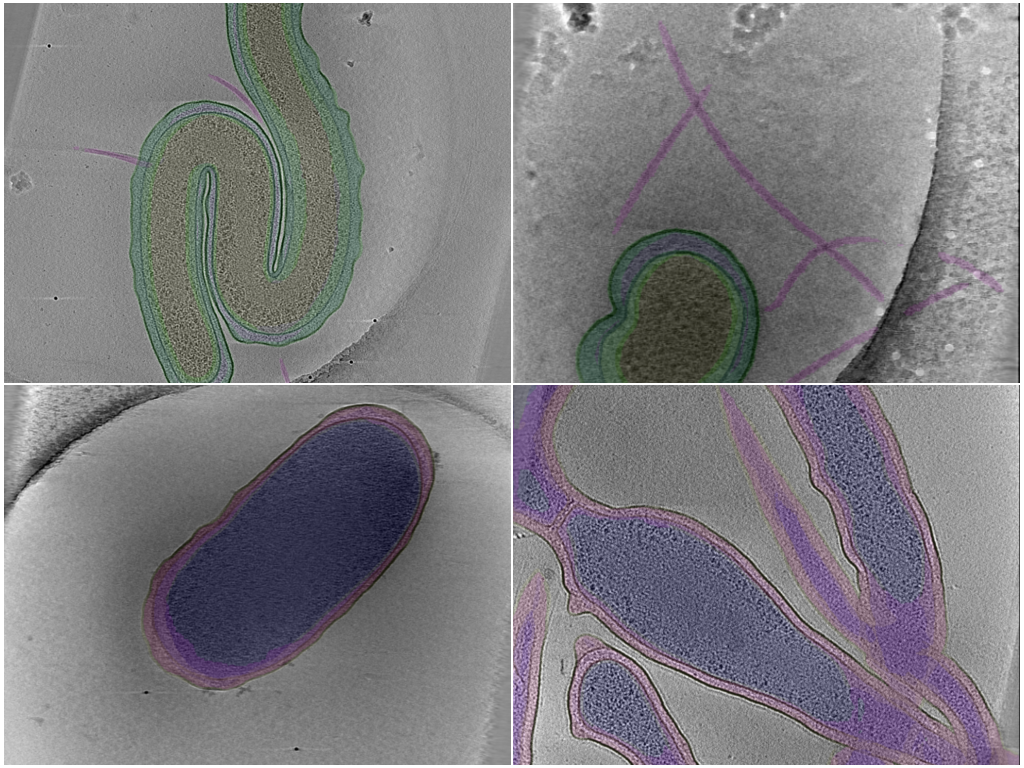
Bacterial — multi-class compartment segmentation
Dense voxel-wise segmentation of cellular compartments in in situ bacterial cryoET tomograms. 80 tomograms total, drawn from 13 underlying portal datasets that span 8 bacterial genera (bdellovibrio, coxiella, hylemonella, hyphomonas, legionella, pseudomonas, salmonella, vibrio), each annotated with up to five compartment classes. Annotations are sourced from the POPSICLE Bacterial Segmentation Dataset deposition (CZCDP-10350).
| Class | Portal object_name |
|---|---|
cytosole |
cytoplasm |
flagellum |
bacterial-type flagellum |
inclusion |
dense body |
intermembrane-space |
periplasmic space |
membrane |
membrane |
Splits
| Split | # tomograms | Notes |
|---|---|---|
train |
68 | 8 genera, ≤19 tomograms each |
test |
12 | held out from training |
Class incidence (number of runs containing each class) is non-uniform —
cytosole, intermembrane-space, and membrane are present in every
run; flagellum and inclusion appear in subsets:
| Class | Train | Test |
|---|---|---|
cytosole |
68 | 12 |
membrane |
68 | 12 |
intermembrane-space |
68 | 12 |
flagellum |
34 | 8 |
inclusion |
29 | 6 |
Segmentations are scoped to deposition CZCDP-10350 so that other
annotation collections living on the same underlying tomograms do not
leak into the benchmark.
Loading with copick
(click to expand)
import copick
import numpy as np
import matplotlib.pyplot as plt
root = copick.from_croissant(
"https://huggingface.co/datasets/biohub/popsicle/resolve/main/bacterial/Croissant/metadata.json",
overlay_root="/tmp/popsicle-overlay",
static_fs_args={"anon": True}, # public portal bucket
)
# Loop over every split and every run
for split, run_names in root.splits.items():
print(f"{split}: {len(run_names)} runs")
for run in root.get_runs_in_split(split):
seg_names = sorted({s.name for s in run.segmentations})
print(f" {run.name}: {seg_names}")
# Visualize: midplane slice of one tomogram with segmentation masks overlaid
run = root.get_runs_in_split("train")[0]
vs = run.voxel_spacings[0]
tomo = vs.tomograms[0]
arr = tomo.numpy() # (Z, Y, X) — streams from portal S3
z_mid = arr.shape[0] // 2
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(arr[z_mid], cmap="gray")
cmap = plt.get_cmap("tab10")
for i, seg in enumerate(run.segmentations):
mask = seg.numpy()[z_mid] # (Y, X) — same shape as the tomo slice
layer = np.ma.masked_where(mask == 0, np.full_like(mask, i, dtype=np.int8))
ax.imshow(layer, cmap=cmap, vmin=0, vmax=10, alpha=0.4, interpolation="none")
# one-line legend hack: scatter a single point off-image with the class color
ax.scatter([], [], color=cmap(i), label=seg.name)
ax.legend(loc="upper right", fontsize=8)
ax.set_title(f"{run.name} z={z_mid}")
plt.show()
Yeast — multi-class organelle segmentation
Dense voxel-wise segmentation of organelles in cryoET tomograms of Schizosaccharomyces pombe (fission yeast) — a low-data, high-variance eukaryotic counterpart to the well-sampled bacterial setting. 20 tomograms from the two CryoET Data Portal S. pombe datasets (DS-10000, DS-10001), annotated with up to six organelle classes per run. Annotations are sourced from the POPSICLE Yeast Segmentation Dataset deposition (CZCDP-10351).
| Class | Portal object_name |
|---|---|
cytoplasm |
cytoplasm |
nucleus |
nucleus |
nuclear-envelope |
nuclear envelope |
vesicle |
vesicle |
membrane-tubule |
membrane-enclosed lumen |
mitochondrion |
mitochondrion |
Splits
| Split | # tomograms | Notes |
|---|---|---|
train |
16 | All 16 from the user-supplied list |
test |
4 | Held out from training |
Class distribution is uneven — large structures like cytoplasm are
present in nearly every run, while small organelles (vesicle,
membrane-tubule) are sparse and mitochondrion / nucleus are present
in only a subset:
| Class | Total runs |
|---|---|
cytoplasm |
19 |
membrane-tubule |
18 |
vesicle |
18 |
mitochondrion |
9 |
nuclear-envelope |
8 |
nucleus |
8 |
This unevenness — together with the small total tomogram count — makes yeast a challenging low-data, class-imbalanced regime, complementary to the better-sampled bacterial benchmark (paper §4, §6, Table 2).
Loading with copick
(click to expand)
import copick
import numpy as np
import matplotlib.pyplot as plt
root = copick.from_croissant(
"https://huggingface.co/datasets/biohub/popsicle/resolve/main/yeast/Croissant/metadata.json",
overlay_root="/tmp/popsicle-overlay",
static_fs_args={"anon": True}, # public portal bucket
)
# Loop over every split and every run
for split, run_names in root.splits.items():
print(f"{split}: {len(run_names)} runs")
for run in root.get_runs_in_split(split):
seg_names = sorted({s.name for s in run.segmentations})
print(f" {run.name}: {seg_names}")
# Visualize: midplane slice of one tomogram with organelle segmentations overlaid
run = root.get_runs_in_split("train")[0]
vs = run.voxel_spacings[0]
tomo = vs.tomograms[0]
arr = tomo.numpy() # (Z, Y, X) — streams from portal S3
z_mid = arr.shape[0] // 2
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(arr[z_mid], cmap="gray")
cmap = plt.get_cmap("tab10")
for i, seg in enumerate(run.segmentations):
mask = seg.numpy()[z_mid]
layer = np.ma.masked_where(mask == 0, np.full_like(mask, i, dtype=np.int8))
ax.imshow(layer, cmap=cmap, vmin=0, vmax=10, alpha=0.4, interpolation="none")
ax.scatter([], [], color=cmap(i), label=seg.name)
ax.legend(loc="upper right", fontsize=8)
ax.set_title(f"{run.name} z={z_mid}")
plt.show()
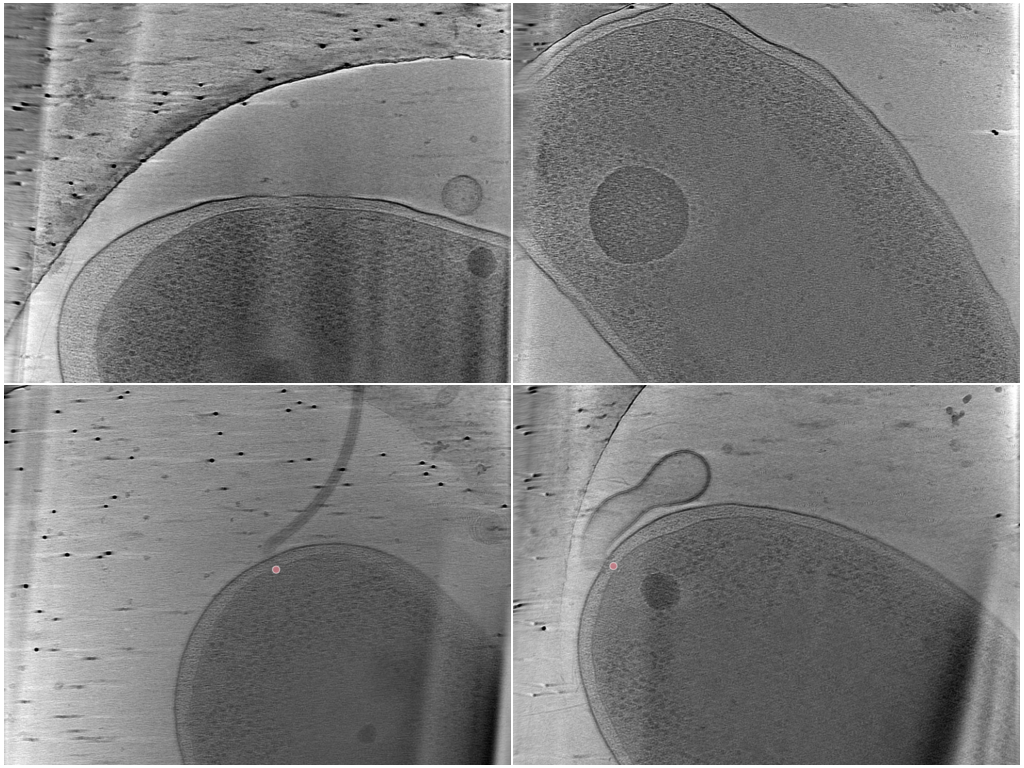
MotorBench — single-class flagellar motor localization
Sparse 3D point localization of bacterial flagellar motors in whole-cell cryoET tomograms. The benchmark is derived from the BYU — Locating Bacterial Flagellar Motors 2025 Kaggle challenge (CZCDP-10332 train, CZCDP-10347 test). The training side is annotation-only on the portal — its 1,559 motor picks are scattered across 92 host datasets spanning many bacterial and archaeal genera, contributed by the BYU competition authors plus the first-place (Brenden Artley) and MIC-DKFZ (Isensee et al.) follow-up releases. The held-out test side is a complete deposition: 5 Vibrio cholerae datasets (DS-10485…10489) authored by Owens et al. 2025.
| Class | Portal object_name |
|---|---|
flagellar-motor |
bacterial-type-flagellum-motor |
Splits
| Split | CDP source(s) | # tomograms | # motor picks | Notes |
|---|---|---|---|---|
train |
CZCDP-10332 → 92 host datasets | 1,559 | 1,559 | One or more motor picks per training run |
test |
CZCDP-10347 (DS-10485…10489) | 843 | 275 | Includes pickless negative samples |
The train slice was filtered down from the 3,528 host-dataset runs to only those that carry a CZCDP-10332 motor annotation — runs without a motor pick belong to unrelated experiments and are not part of the benchmark. The test slice keeps all 843 V. cholerae runs (motor and no-motor) because negative samples are an explicit part of the held-out evaluation per Owens et al. 2025.
Loading with copick
(click to expand)
import copick
import numpy as np
import matplotlib.pyplot as plt
root = copick.from_croissant(
"https://huggingface.co/datasets/biohub/popsicle/resolve/main/motorbench/Croissant/metadata.json",
overlay_root="/tmp/popsicle-overlay",
static_fs_args={"anon": True}, # public portal bucket
)
# Loop over every split and every run; report runs that carry motor picks
for split, run_names in root.splits.items():
runs_with = [r for r in root.get_runs_in_split(split) if r.picks]
print(f"{split}: {len(run_names)} runs ({len(runs_with)} with motor picks)")
# Visualize: midplane slice of one *V. cholerae* test tomogram with motor
# centers overlaid as crosses.
run = root.get_run("33914")
vs = run.voxel_spacings[0]
tomo = vs.tomograms[0]
arr = tomo.numpy() # (Z, Y, X) — streams from portal S3
z_mid = arr.shape[0] // 2
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(arr[z_mid], cmap="gray")
for pick_set in run.picks:
pts = np.array(
[(p.location.x, p.location.y, p.location.z) for p in pick_set.points]
) / vs.voxel_size
near = pts[np.abs(pts[:, 2] - z_mid) < 5]
ax.scatter(near[:, 0], near[:, 1], marker="x", s=80, c="red",
label=pick_set.pickable_object_name)
ax.legend(loc="upper right", fontsize=8)
ax.set_title(f"{run.name} z={z_mid}")
plt.show()
Dataset Creation
Curation Rationale
Existing cryoET evaluations are typically small, task-specific, and assembled in isolation, which makes cross-method comparison unreliable (paper §1, §2). POPSICLE provides one consistent evaluation surface that spans:
- Both task regimes: dense voxel-wise segmentation of cellular structures and sparse localization of macromolecular complexes.
- Both biological regimes: eukaryotic S. pombe and prokaryotic bacterial cells, plus a controlled lysate phantom.
- Heterogeneous data scales: from 7-tomogram low-supervision regimes to 1,000+ tomogram well-sampled regimes, exposing whether model rankings are stable across data abundance.
Because the underlying CryoET Data Portal is a continuously growing, ML-ready resource, POPSICLE is designed to evolve as new datasets and annotations become available rather than being a static challenge release (paper §1, §7).
Source Data
Data Collection and Processing
All source tomograms were acquired by tilt-series cryoET on vitrified biological samples and reconstructed via standard pipelines described on the CryoET Data Portal (paper §C). Biohub DSB uses AreTomo3 for motion correction, tilt-series alignment, CTF correction, and weighted back-projection, with optional self-supervised denoising via DenoisET. Phantom tomograms are reconstructed by the original Peck et al. 2025 pipeline and ingested into the portal as DS-10440/10445/10446.
Who are the source data producers?
Provenance differs per sub-benchmark and is summarized below. Authoritative credits live on each portal deposition page linked under Dataset Sources.
| Sub-benchmark | Tomogram source | Annotation source |
|---|---|---|
| Phantom | Acquired and reconstructed by Biohub DSB. | Created by Biohub DSB (Peck et al. 2025). |
| Bacterial | Contributed by external labs (8 bacterial genera). | Created by the POPSICLE authors. |
| Yeast | Acquired by external labs (de Teresa-Trueba et al.). | Curated/refined by Biohub DSB from prior releases. |
| MotorBench | Contributed by external labs (BYU + collaborators). | Contributed by the BYU Kaggle community . |
In short: POPSICLE produces some new annotations (Bacterial), curates others (Yeast), and indexes the rest (Phantom, MotorBench) — none of the underlying tomograms are re-acquired by this project.
Annotations
Annotation process
- Phantom: a sample of recombinantly expressed and purified macromolecules was prepared in lysate-like conditions and imaged by cryoET; Biohub DSB annotators identified the 3D centers of all six target classes per tomogram (Peck et al. 2025). Annotations are stored on the portal as point picks.
- Bacterial: POPSICLE authors generated dense voxel-wise segmentations across five compartment classes (cytosole, flagellum, inclusion, intermembrane-space, membrane) on 80 cellular tomograms from 8 bacterial genera, deposited as CZCDP-10350.
- Yeast: organelle segmentations originally produced by de Teresa-Trueba et al. 2023; curated and standardized by Biohub DSB for inclusion in POPSICLE.
- MotorBench: bacterial flagellar motor centers picked by the BYU Kaggle competition contributors (training corpus CZCDP-10332, expanded by the first-place team and MIC-DKFZ); the held-out test set (CZCDP-10347, V. cholerae) was authored by Owens et al. 2025.
Who are the annotators?
- Phantom: Ariana Peck and Biohub DSB co-authors of Peck et al. 2025 (point picks restricted in this release to exclude community Kaggle submissions hosted on the same datasets).
- Bacterial: POPSICLE author team (CZCDP-10350 contributors).
- Yeast: de Teresa-Trueba et al. 2023, with curatorial refinement by Biohub DSB.
- MotorBench: BYU competition + first-place team (Brenden Artley) + MIC-DKFZ (Fabian Isensee et al.) for the training corpus; Owens et al. 2025 (V. cholerae) for the held-out test set.
Personal and Sensitive Information
None. The dataset contains 3D microscopy images of in vitro and cellular biological samples; no human-subject data is present.
Bias, Risks, and Limitations
- Annotation completeness. Even expert ground truth for cryoET data may miss particles that are heavily distorted, edge-of-field, or occluded. Phantom's evaluation protocol uses a recall-weighted F4 precisely because false negatives are more costly than false positives in downstream subtomogram averaging (paper §3.2).
- Class imbalance. The Phantom β-amylase class (~268 kDa) is the smallest, lowest-contrast target; its reference labels carry lower confidence and the class is excluded from the aggregate F̄₄ score in the official evaluation protocol (paper §B.6, following the CZII Kaggle challenge). β-amylase is preserved in the dataset and reported per-class so methods can still attempt it.
- Imaging artifacts. All cryoET data carries the missing-wedge artifact and anisotropic resolution; performance is sensitive to both biological variation and reconstruction choices (paper §D).
- Living dataset. POPSICLE is built on a continuously expanding data resource. Future revisions will add sub-benchmarks and may refine annotations; pin the dataset revision when reproducing results.
Recommendations
- Train per sub-benchmark using the official
train(and, where applicable,val) splits; reservetestfor held-out evaluation. - Report per-class scores in addition to aggregates — Phantom rankings shift substantially when β-amylase weighting changes (paper §6, Table 4).
- When comparing to prior in-field work or Kaggle reference points, follow the original challenge protocol exactly (recall-weighted Fβ, class weights, β-amylase exclusion).
Citation
BibTeX:
@inproceedings{popsicle2026,
title = {POPSICLE: Benchmark Datasets for Segmentation and Localization in CryoET},
author = {Anonymous},
booktitle = {Advances in Neural Information Processing Systems},
year = {2026},
note = {Under review}
}
The Phantom sub-benchmark inherits its tomograms and reference picks from:
@article{peck2025phantom,
title = {A realistic phantom dataset for benchmarking cryo-{ET} data annotation},
author = {Peck, Ariana and Yu, Yue and Schwartz, Jonathan and Cheng, Anchi
and Ermel, Utz Heinrich and Hutchings, Joshua and Kandel, Saugat
and Kimanius, Dari and Montabana, Elizabeth A. and Serwas, Daniel
and Siems, Hannah and Wang, Feng and Zhao, Zhuowen and Zheng, Shawn
and Haury, Matthias and Agard, David A. and Potter, Clinton S.
and Carragher, Bridget and Harrington, Kyle and Paraan, Mohammadreza},
journal = {Nature Methods},
volume = {22},
pages = {1819--1823},
year = {2025},
doi = {10.1038/s41592-025-02800-5}
}
The MotorBench held-out test set is sourced from:
@article{owens2025motorbench,
title = {MotorBench: A cryo-electron tomography dataset of bacterial flagellar motors for testing detection algorithms},
author = {Owens, C. Braxton and Webb, Rachel and Hart, T. J. and Ward, Matthew M.
and Darley, Andrew J. and Maggi, Stefano and Morse, Bryan S.
and Jensen, Grant J. and Reade, Walter C. and Kaplan, Mohammed
and Hart, Gus L. W.},
journal = {bioRxiv},
year = {2025},
doi = {10.1101/2025.04.23.650258}
}
The Yeast sub-benchmark re-curates the upstream organelle segmentations introduced by DeePiCt:
@article{deteresa2023deepict,
title = {Convolutional networks for supervised mining of molecular patterns within cellular context},
author = {de Teresa-Trueba, Irene and Goetz, Sara K. and Mattausch, Alexander
and Stojanovska, Frosina and Zimmerli, Christian E. and Toro-Nahuelpan, Mauricio
and Cheng, Dorothy W. C. and Tollervey, Fergus and Pape, Constantin
and Beck, Martin and Diz-Mu{\~n}oz, Alba and Kreshuk, Anna
and Mahamid, Julia and Zaugg, Judith B.},
journal = {Nature Methods},
volume = {20},
pages = {284--294},
year = {2023},
doi = {10.1038/s41592-022-01746-2}
}
CryoET Data Portal:
@article{ermel2024dataportal,
title = {A data portal for providing standardized annotations for cryo-electron tomography},
author = {Ermel, Utz and others},
journal = {Nature Methods},
volume = {21},
number = {12},
pages = {2200--2202},
year = {2024}
}
copick toolkit:
@article{ermel2026copick,
title = {copick: An open dataset interface and toolkit for collaborative annotation and analysis of cryo-electron tomography data},
author = {Ermel, Utz Heinrich and Schwartz, Jonathan and Zhao, Zhuowen and Ji, Daniel
and Peck, Ariana and Yu, Yue and Paraan, Mohammadreza and Carragher, Bridget
and Frangakis, Achilleas S. and Harrington, Kyle I. S.},
journal = {Protein Science},
volume = {35},
number = {5},
pages = {e70578},
year = {2026},
doi = {10.1002/pro.70578}
}
References
Works cited inline in this dataset card. Author-year ordering.
- POPSICLE 2026 — POPSICLE: Benchmark Datasets for Segmentation and Localization in CryoET. NeurIPS 2026 (under review). (Anonymous during review; this dataset card.)
- de Teresa-Trueba et al. 2023 — Convolutional networks for supervised mining of molecular patterns within cellular context. Nature Methods 20, 284–294 (2023). doi:10.1038/s41592-022-01746-2
- Ermel et al. 2024 — A data portal for providing standardized annotations for cryo-electron tomography. Nature Methods 21, 2200–2202 (2024). doi:10.1038/s41592-024-02475-4
- Ermel et al. 2026 — copick: An open dataset interface and toolkit for collaborative annotation and analysis of cryo-electron tomography data. Protein Science 35, e70578 (2026). doi:10.1002/pro.70578
- Owens et al. 2025 — MotorBench: A cryo-electron tomography dataset of bacterial flagellar motors for testing detection algorithms. bioRxiv (2025). doi:10.1101/2025.04.23.650258
- Peck et al. 2025 — A realistic phantom dataset for benchmarking cryo-ET data annotation. Nature Methods 22, 1819–1823 (2025). doi:10.1038/s41592-025-02800-5
- Peck et al. 2025 (AreTomo3) — AreTomoLive: Automated reconstruction of comprehensively-corrected and denoised cryo-electron tomograms in real-time and at high throughput. bioRxiv (2025). doi:10.1101/2025.03.11.642690
Glossary
- Tomogram — a 3D reconstruction of a vitrified biological sample computed from a tilt series of 2D projections.
- Pick — a 3D coordinate identifying the center of a target macromolecule within a tomogram.
- Segmentation — a voxel-wise label volume assigning each voxel to one of a fixed set of classes (membrane, organelle, cytoplasm, …).
- Run (CryoET Data Portal) — all data acquired and derived from
imaging a single location in a sample: the original tilt series,
motion-corrected frames, one or more reconstructed tomograms (often at
multiple voxel spacings), and any annotations attached to those
tomograms. A run belongs to exactly one dataset and is identified by
a unique numeric Run ID. Copick exposes this as
run.name(a string cast of the Run ID for portal-backed projects). - Dataset (CryoET Data Portal, "DS-XXXXX") — a collection of runs that share a common sample and acquisition context (organism, strain, preparation, microscope session). Datasets carry their own metadata (sample type, growth conditions, instrument, contributing lab) and belong to exactly one deposition. The CDP "DS-10440" notation refers to a Dataset row.
- Deposition (CryoET Data Portal, "CZCDP-XXXXX") — a higher-level grouping under which one or more datasets and/or annotations are contributed to the portal as a single submission, typically tied to a publication or a community release (e.g. CZCDP-10350 for the POPSICLE Bacterial deposition). A deposition can be full (it carries new datasets + their tomograms + annotations) or annotation-only (it carries new annotations attached to runs that already exist in datasets contributed earlier under different depositions — used by POPSICLE for the Bacterial annotations and by the BYU MotorBench train deposition).
- Missing wedge — the cone of unmeasured Fourier-space data caused by the limited tilt range during cryoET acquisition; produces anisotropic resolution and characteristic elongation artifacts.
- F4 / F2 score — recall-weighted Fβ scores used in localization evaluation; β=4 (Phantom) and β=2 (MotorBench) emphasize recall over precision because missed targets are more costly downstream than filterable false positives.
More Information
- copick documentation: https://github.com/copick/copick
- CryoET Data Portal: https://cryoetdataportal.czscience.com
Dataset Card Authors
POPSICLE benchmark authors (NeurIPS 2026 submission, anonymous during review).
Dataset Card Contact
Open an issue at the dataset repository.
- Downloads last month
- 22