branch_name stringclasses 149 values | text stringlengths 23 89.3M | directory_id stringlengths 40 40 | languages listlengths 1 19 | num_files int64 1 11.8k | repo_language stringclasses 38 values | repo_name stringlengths 6 114 | revision_id stringlengths 40 40 | snapshot_id stringlengths 40 40 |
|---|---|---|---|---|---|---|---|---|
refs/heads/master | <file_sep>/* ①book.h文件的完整内容 */
#ifndef _BOOK /*条件编译,防止重复包含的错误*/
#define _BOOK
#include <string.h>
#define NUM 20 /*定义图书常量,此处可以根据实际需要修改常量值*/
struct Student /*学生记录的数据域*/
{
long num; /*学号 */
char name[20]; /*姓名 */
};
struct Book
{
char title[20]; /*书名 */
long number[20]; /*索引号 */
char state[4]; /*借阅状态 */
int kc; /*库存*/
};
typedef struct Book Book;
#define sizeBook sizeof(Book) /*一条图书记录所需要的内存空间大小*/
int readBook(Book book[],int n); /*读入书书籍名称*/
void printB00k( Book *Book , int n); /*输出所有书籍记录的值*/
void sortBook(Book book[],int n,int condition); /*选择法从大到小排序,按condition所规定的条件*/
int searchBook(Book book[],int n,Book b,int condition,int f[]) ; /*根据条件找数组中与b相等的各元素*/
int addBook(Book book[],int n,Book b); /*向数组中增加一条图书信息*/
int deleteBook(Book book[],int n,Book b); /*从数组中删除一条图书信息*/
#endif<file_sep>#include<stdio.h>
#include<stdlib.h>
#include"file.h"
#include"book.h"
void printHead( ) /*打印图书信息的表头*/
{
printf("%ld%s%ld%s%d\n","索引号","书名","学号","姓名","库存");
}
void menu( ) /*顶层菜单函数*/
{
printf("******** 1. 显示基本信息 ********\n");
printf("******** 2. 图书信息管理 ********\n");
printf("******** 3. 借阅人信息管理 ********\n");
printf("******** 4. 借阅数据统计********\n");
printf("******** 5. 根据条件查询 ********\n");
printf("******** 0. 退出 ********\n");
}
void menuBase( ) /*2、基本信息管理菜单函数*/
{
printf("%%%%%%%% 1. 插入图书记录 %%%%%%%%\n");
printf("%%%%%%%% 2. 删除图书记录 %%%%%%%%\n");
printf("%%%%%%%% 3. 修改图书记录 %%%%%%%%\n");
printf("%%%%%%%% 0. 返回上层菜单 %%%%%%%%\n");
}
void peopleBase() /*3、借阅人信息管理菜单函数*/
{
printf("@@@@@@@@@ 1. 插入借阅人信息记录 @@@@@@@@\n");
printf("@@@@@@@@@ 2. 删除借阅人信息记录 @@@@@@@@\n");
printf("@@@@@@@@@ 3. 修改借阅人信息记录 @@@@@@@@\n");
printf("@@@@@@@@@ 0. 返回上层菜单 @@@@@@@@\n");
}
void menuCount( ) /*4、统计菜单函数*/
{
printf("&&&&&&&& 1. 求借阅次数最高 &&&&&&&&\n");
printf("&&&&&&&& 1. 求借阅次数最低 &&&&&&&&\n");
printf("&&&&&&&& 0. 返回上层菜单 &&&&&&&&\n");
}
void menuSearch() /*5、根据条件查询菜单函数*/
{
printf("######## 1. 按书名查询 ########\n");
printf("######## 2. 按索引号查询 ########\n");
printf("######## 3. 借阅人信息查询 ########\n");
printf("######## 0. 返回上层菜单 ########\n");
}
int baseManage(Book book[],int n) /*该函数完成基本图书信息管理*/
/*按索引号进行插入删除修改,索引号不能重复*/
{
int choice,t,find[NUM];
Book b;
do
{
menuBase( ); /*显示对应的二级菜单*/
printf("choose one operation you want to do:\n");
scanf("%d",&choice); /*读入选项*/
switch(choice)
{
case 1: readBook(&b,1); /*读入一条待插入的图书记录*/
n=addBook(book,n,b); /*调用函数插入图书记录*/
break;
case 2: printf("Input the number deleted\n");
scanf("%ld",&b.number); /*读入一个待删除的图书索引号*/
n=deleteBook(book,n,b); /*调用函数删除指定索引号的图书记录*/
break;
case 3: printf("Input the number modified\n");
scanf("%ld",&b.number); /*读入一个待修改的图书索引号*/
t=searchBook(book,n,b,1,find) ; /*调用函数查找指定索引号的图书记录*/
if (t) /*如果该索引号的记录存在*/
{
readBook(&b,1); /*读入一条完整的图书记录信息*/
book[find[0]]=b; /*将刚读入的记录赋值给需要修改的数组记录*/
}
else /*如果该索引号的记录不存在*/
printf("this book is not in,can not be modified.\n"); /*输出提示信息*/
break;
case 0: break;
}
}while(choice);
return n; /*返回当前操作结束后的实际记录条数*/
}
int baseManage1(Student student[],int n) /*该函数完成借阅人基本信息管理*/
/*按学号进行插入删除修改, 学号不能重复*/
{
int choice,t,find[NUM];
Student s;
do
{
peopleBase( ); /*显示对应的二级菜单*/
printf("choose one operation you want to do:\n");
scanf("%d",&choice); /*读入选项*/
switch(choice)
{
case 1: readStudent(&s,1); /*读入一条待插入的借阅人记录*/
n=insertStudent(student,n,s); /*调用函数插入借阅人记录*/
break;
case 2: printf("Input the student number deleted\n");
scanf("%ld",&s.num); /*读入一个待删除的学号*/
n=deleteStudent(student,n,s); /*调用函数删除指定学号的借阅人记录*/
break;
case 3: printf("Input the student number modified\n");
scanf("%ld",&s.num); /*读入一个待修改的学号记录*/
t=searchStudent(student,n,s,1,find) ; /*调用函数查找指定学号的借阅人记录*/
if (t) /*如果该学号的记录存在*/
{
readStudent(&s,1); /*读入一条完整的借阅人记录信息*/
student[find[0]]=s; /*将刚读入的记录赋值给需要修改的数组记录*/
}
else /*如果该学好的借阅人记录不存在*/
printf("this people is not in,can not be modified.\n"); /*输出提示信息*/
break;
case 0: break;
}
}while(choice);
return n; /*返回当前操作结束后的实际记录条数*/
}
void printBookTimes(char *b,double m[NUM][2],int k) /*打印借阅次数通用函数,被countManage 调用
/*形式参数k代表输出不同的内容*/
{
int i;
printf(b);
for(i=0;i<NUM;i++)
printf("%d",m[i][k]);
printf("\n");
}
void countManage(Book book[],int n) /*该函数完成借阅次数的统计功能*/
{
int choice;
double mark[NUM][2];
do
{
menuCount( ); /*显示对应的二级菜单*/
printf("choose one operation you want to do:\n");
scanf("%d",&choice);
switch(choice)
{
case 1: printBookTimes("10本书的最高借阅次数是:\n",mark,0); /*输出最高次数*/
break;
case 2: printBookTimes("·10本书的最低借阅次数是:\n",mark,1); /*输出最低次数*/
break;
case 0: break;
}
}while (choice);
}
void searchManage(Book book[],int n) /*该函数完成根据条件查询功能*/
{
int i,choice,findnum,f[NUM];
Book b;
Student s;
do
{
menuSearch( ); /*显示对应的二级菜单*/
printf("choose one operation you want to do:\n");
scanf("%d",&choice);
switch(choice)
{
case 1: printf("Input a book\'s title will be searched:\n");
scanf("%ld",&b.title); /*输入待查询图书的索引号*/
break;
case 2: printf("Input a book\'s number will be searched:\n");
scanf("%s",&b.number); /*输入待查询图书的书名*/
break;
case 3: printf(" search the informatiom of the readers");
scanf("%ld",&s.num); /*输入待查询借阅人的学号*/
case 0: break;
}
if (choice>=1&&choice<=2)
{
findnum=searchBook(book,n,b,choice,f); /*查找的符合条件元素的下标存于f数组中*/
if (findnum) /*如果查找成功*/
{
printHead( ); /*打印表头*/
for (i=0;i<findnum;i++) /*循环控制f数组的下标*/
printBook(&book[f[i]],1); /*每次输出一条记录*/
}
else
printf("this record does not exist!\n"); /*如果查找不到元素,则输出提示信息*/
}
}while (choice);
}
int runMain(Book book[],Student student[],int n,int choice) /*主控模块,对应于一级菜单其下各功能选择执行*/
{
switch(choice)
{
case 1: printHead( ); /* 1. 显示基本信息*/
sortBook(book,n,1); /*按借阅次数由小到大的顺序排序记录*/
printBook(book,n); /*按借阅次数由小到大的顺序输出所有记录*/
break;
case 2: n=baseManage(book,n); /* 2. 图书基本信息管理*/
break;
case 3: n=baseManage1(student,n); /* 3. 借阅人信息管理*/
break;
case 4: countManage(book,n); /* 4. 借阅次数统计*/
break;
case 5: searchManage(book,n); /* 5. 根据条件查询*/
break;
case 0: break;
}
return n;
}
int main( )
{
Book book[NUM];
Student student[STU]; /*定义实参一维数组存储图书记录*/
int choice,n,m;
n=readFile(book); /*首先读取文件,记录条数返回赋值给n*/
m=readstudentFile(student);
if (!n) /*如果原来的文件为空*/
{
n=createFile(book); /*则首先要建立文件,从键盘上读入一系列记录存于文件*/
}
else if(!m)
{
m=createstudentfile(student);
}
do
{
menu(); /*显示主菜单*/
printf("Please input your choice: ");
scanf("%d",&choice);
if (choice>=0&&choice<=5)
n=runMain(book,student,n,choice); /*通过调用此函数进行一级功能项的选择执行*/
else
printf("error input,please input your choice again!\n");
} while (choice);
sortBook(book,n,1); /*存入文件前借阅次数由小到大排序*/
saveFile(book,n); /*将结果存入文件*/
return 0;
}
<file_sep> /*③ file.h文件的内容如下:*/
#include <stdio.h>
#include <stdlib.h>
#include "book.h"
int createFile(Book book[]) /*建立初始的数据文件*/
{
FILE *fp;
int n;
if((fp=fopen("d:\\book.dat", "wb")) == NULL) /*指定好文件名,以写入方式打开*/
{
printf("can not open book file !\n"); /*若打开失败,输出提示信息*/
exit(0); /*然后退出*/
}
printf("input book's information:\n");
n=readBook(book,NUM); /*调用book.h中的函数读数据*/
fwrite(book,sizeBook,n,fp); /*将刚才读入的所有记录一次性写入文件*/
fclose(fp);
return n;
}
int readFile(Book book[ ] ) /*将文件中的内容读出置于结构体数组book中*/
{
FILE *fp;
int i=0;
if((fp=fopen("d:\\book.dat", "rb")) == NULL) /*以读的方式打开指定文件*/
{
printf("bookfile does not exist,create it first:\n"); /*如果打开失败输出提示信息*/
return 0; /*然后返回0*/
}
fread(&book[i],sizeBook,1,fp); /*读出第一条记录*/
while(!feof(fp)) /*文件未结束时循环*/
{
i++;
fread(&book[i],sizeBook,1,fp); /*再读出下一条记录*/
}
fclose(fp); /*关闭文件*/
return i; /*返回记录条数*/
}
void saveFile(Book book[],int n) /*将结构体数组的内容写入文件*/
{
FILE *fp;
if((fp=fopen("d:\\book.dat", "wb")) == NULL) /*以写的方式打开指定文件*/
{
printf("can not open book file !\n"); /*如果打开失败,输出提示信息*/
exit(0); /*然后退出*/
}
fwrite(book,sizeBook,n,fp);
fclose(fp); /*关闭文件*/
}
int createstudentfile(Student student[])
{
FILE *fq;
int m;
if((fq=fopen("d:\\student.dat","wb"))==NULL)
{
printf("can not open student file !\n");
exit(0);
}
printf("input borrower's information:\n"); /*关闭文件*/
m=readStudent(student,STU);
fwrite(student,sizeStudent,m,fq);
fclose(fq);
return m;
}
int readstudentFile(Student student[ ] ) /*将文件中的内容读出置于结构体数组book中*/
{
FILE *fq;
int j=0;
if((fq=fopen("d:\\student.dat", "rb")) == NULL) /*以读的方式打开指定文件*/
{
printf("studentfile does not exist,create it first:\n"); /*如果打开失败输出提示信息*/
return 0; /*然后返回0*/
}
fread(&student[j],sizeStudent,1,fq); /*读出第一条记录*/
while(!feof(fq)) /*文件未结束时循环*/
{
j++;
fread(&student[j],sizeStudent,1,fq); /*再读出下一条记录*/
}
fclose(fq); /*关闭文件*/
return j; /*返回记录条数*/
}
void savestudentFile(Student student[],int n) /*将结构体数组的内容写入文件*/
{
FILE *fq;
if((fq=fopen("d:\\student.dat", "wb")) == NULL) /*以写的方式打开指定文件*/
{
printf("can not open student file !\n"); /*如果打开失败,输出提示信息*/
exit(0); /*然后退出*/
}
fwrite(student,sizeStudent,n,fq);
fclose(fq); /*关闭文件*/
}<file_sep>#ifndef _BOOK /*条件编译,防止重复包含的错误*/
#define _BOOK
#include <string.h>
#define NUM 3
#define STU 3
struct Book
{
char title[20]; /*书名 */
long number; /*索引号 */
int time; /*借阅次数 */
int kc; /*该图书总数量*/
};
typedef struct Book Book; /*定义图书常量,此处可以根据实际需要修改常量值*/
struct Student /*学生记录的数据域*/
{
long num; /*学号 */
char name[20]; /*姓名 */
Book message;
};typedef struct Student Student;
#define sizeBook sizeof(Book) /*一条图书记录所需要的内存空间大小*/
#define sizeStudent sizeof(Student) /*一个借阅人记录所需要的内存空间大小*/
int readBook(Book book[],int n); /*读入书书籍名称*/
void printBook( Book *Book , int n); /*输出所有书籍记录的值*/
int equal(Book b1,Book b2,int condition);/*比较图书记录是否相同*/
int larger(Book b1,Book b2,int condition); /*根据condition条件比较两本图书索引号的大小*/
void sortBook(Book book[],int n,int condition); /*选择法从小到大排序,按condition所规定的条件*/
int searchBook(Book book[],int n,Book b,int condition,int f[]) ; /*根据条件找数组中与b相等的各元素*/
int addBook(Book book[],int n,Book b); /*向数组中增加一条图书信息*/
int deleteBook(Book book[],int n,Book b); /*从数组中删除一条图书信息*/
int readStudent(Student student[], int n); /*读入借阅人记录值,借阅人为0或读满规定条数记录时停止*/
void printStudent( Student *student, int n); /*输出所有借阅人记录的值*/
int equal1(Student s1,Student s2,int condition);/*比较学生记录是否相同*/
void sortStudent(Student stu[],int n,int condition); /*选择法排序,按condition条件由小到大排序*/
int searchStudent(Student stu[],int n,Student s,int condition,int f[ ]); /*在stu数组中依condition条件查找*/
int insertStudent(Student student[],int n,Student s); /*向数组中增加一条学生信息*/
int deleteStudent(Student student[],int n,Student s);
#endif | 71d9eeb3952ac7aafd44d297d0c57dffde6b325d | [
"C",
"C++"
] | 4 | C | B15090806/B150908 | ab1b31b18ccdaea75881c1d2175474e6e7ed248b | fe7dd24b47b21ec2532763514e11e81ab1ae7ea7 |
refs/heads/master | <repo_name>ciena/ZeroTouchProvisioning<file_sep>/main.go
package main
import (
"bytes"
"flag"
"fmt"
"github.com/tmc/scp"
"golang.org/x/crypto/ssh"
"log"
"os"
"time"
)
const (
//onosIP = "10.0.0.1"
TIMEOUT = 3
)
func main() {
ip := flag.String("ip", "", "IP address of the switch")
host := flag.String("hostname", "", "Hoatname of the switch")
dpid := flag.String("dpid", "", "DPID of the switch")
user := flag.String("user", "root", "Username for the switch login")
password := flag.String("password", "onl", "Password for the switch login")
onosIP := flag.String("onosip", "10.1.0.1", "ONOS controller IP")
flag.Parse()
var buf bytes.Buffer
logger := log.New(&buf, "AUTOCONFIG: ", log.Ltime)
logger.Println("logger initialized")
config := &ssh.ClientConfig{
User: *user,
Auth: []ssh.AuthMethod{
ssh.Password(*password),
},
}
client, err := ssh.Dial("tcp", *ip+":22", config)
if err != nil {
panic("Failed to dial: " + err.Error())
}
cmd1 := "Working on... Hostname: " + *host + " with DPID: " + *dpid + " IP: " + *ip
fmt.Println(cmd1)
scpCmd := "scp"
cmdRC := "echo dpkg -i --force-overwrite /mnt/flash2/ofdpa-i.12.1.1_12.1.1+accton1.7-1_amd64.deb > /etc/rc.local"
hostnameString := fmt.Sprintf("hostname %s", *host)
cmdRChost := "echo " + hostnameString + " >> /etc/rc.local"
cmdRCexit := "echo exit 0 >> /etc/rc.local"
connect := "brcm-indigo-ofdpa-ofagent --dpid=" + *dpid + " --controller=" + *onosIP
cmds := []string{"test -e /etc/.configured && echo 'found' || echo 'notFound'",
"test -e /etc/.connected && echo 'connected' || echo 'notConnected'",
"persist /etc/network/interfaces",
"savepersist",
scpCmd,
"service ofdpa stop",
"dpkg -i --force-overwrite /mnt/flash2/ofdpa-i.12.1.1_12.1.1+accton1.7-1_amd64.deb",
"service ofdpa restart",
"persist /etc/accton/ofdpa.conf",
"savepersist",
cmdRC,
cmdRChost,
cmdRCexit,
"persist /etc/rc.local",
"savepersist",
connect,
"touch /etc/.configured",
"persist /etc/.configured",
"savepersist",
}
for cmdNumber, cmd := range cmds {
session, err := client.NewSession()
if err != nil {
panic("Failed to create session: " + err.Error())
}
defer session.Close()
var b bytes.Buffer
session.Stdout = &b
if cmd == scpCmd {
src := "ofdpa-i.12.1.1_12.1.1+accton1.7-1_amd64.deb"
dst := "/mnt/flash2/" + src
err = scp.CopyPath(src, dst, session)
if _, err := os.Stat(src); os.IsNotExist(err) {
fmt.Printf("no such file or directory: %s", src)
panic(err)
} else {
fmt.Println("SCP Success")
continue
}
}
fmt.Println(" RUNNING: " + cmd)
if cmd == "savepersist" {
session.Run(cmd) //savepersist returns error even if it succeeds (ONL bug)
} else if cmd == connect {
go func() {
time.Sleep(TIMEOUT * time.Millisecond)
timeout <- true
}()
go func() {
fmt.Println(" RUNNING: " + connect)
session.Run(cmd)
}()
time.Sleep(2 * time.Second)
} else {
if err := session.Run(cmd); err != nil {
fmt.Println("Failed to run cmd: " + cmd + " ERROR: " + err.Error())
}
}
rpl := b.String()
if cmdNumber == 0 {
fmt.Println(rpl[:5])
if rpl[:5] == "found" {
fmt.Println("Switch is already configured!")
}
}
if cmdNumber == 1 {
fmt.Println(rpl[:9])
if rpl[:9] == "connected" {
fmt.Println("Switch is already CONNECTED!")
break
} else {
fmt.Println("Switch is configured but not connected to ONOS, connecting now...")
go func() {
fmt.Println(" RUNNING: " + connect)
session.Run(connect)
}()
time.Sleep(2 * time.Second)
connd := "touch /etc/.connected"
session, err := client.NewSession()
if err != nil {
panic("Failed to create session: " + err.Error())
}
defer session.Close()
fmt.Println(" RUNNING: " + connd)
if err := session.Run(connd); err != nil {
fmt.Println("Failed to run cmd: " + connd + " ERROR: " + err.Error())
}
}
break
}
}
}
<file_sep>/Makefile
build:
GOOS=linux GOARCH=amd64 go build -o switchGo
image:
sudo docker build -t cord/fabricdhcpharvester .
run1:
sudo docker-compose -f harvest-compose-1.yml up
run2:
sudo docker-compose -f harvest-compose-2.yml up
runquiet1:
sudo docker-compose -f harvest-compose-1.yml up -d
runquiet2:
sudo docker-compose -f harvest-compose-2.yml up -d<file_sep>/Dockerfile
FROM iron/base
RUN apk update && apk upgrade \
&& apk add python \
&& rm -rf /var/cache/apk/*
ADD dhcpharvester.py /dhcpharvester.py
ADD switchGo /switchGo
ADD main.go /main.go
ADD ofdpa-i.12.1.1_12.1.1+accton1.7-1_amd64.deb /ofdpa-i.12.1.1_12.1.1+accton1.7-1_amd64.deb
ENTRYPOINT [ "python", "/dhcpharvester.py" ]
<file_sep>/README.md
# Zero Touch Provisioning (ZTP) for Whitebox fabric
- It uses DHCP information to see if a switch bootsup and matches it's config based on the MAC in the DHCP request (using DHCP harvest)
- Then it calls the Go program wich configures the switch
- Configuration includes, copying the appropriate OFDPA image, installing ofdpa, restarting ofdpa, hostname, make sure the configuration is persisted across reboots, etc. | f1ea56528cc76499a1b9432584f73db9daf45b2a | [
"Makefile",
"Go",
"Dockerfile",
"Markdown"
] | 4 | Go | ciena/ZeroTouchProvisioning | a262fc3630a17ff415ce06d1ff82af7550a20804 | da75719b39a6cb8b0175a4d7cacdf17f143e5658 |
refs/heads/master | <file_sep># RNCryptor
RNCryptor加密解密Python代码
#### **Dependencies**
/assets/pycrypto-2.6.1.tar.gz
```bash
tar -zxvf /assets/pycrypto-2.6.1.tar.gz
sudo python setup.py install
```
#### **用法**
```bash
python RNCryptor.py
```
```python
cryptor = RNCryptor()
#加密
cryptor.encrypt(Data,password)
#解密
cryptor.decrypt(Data,password)
```

<file_sep>#!/usr/bin/env python
# coding: utf-8
from __future__ import print_function
import hashlib
import hmac
import sys
import base64
from Crypto.Cipher import AES
from Crypto.Protocol import KDF
from Crypto import Random
PY2 = sys.version_info[0] == 2
PY3 = sys.version_info[0] == 3
if PY2:
def to_bytes(s):
if isinstance(s, str):
return s
if isinstance(s, unicode):
return s.encode('utf-8')
to_str = to_bytes
def bchr(s):
return chr(s)
def bord(s):
return ord(s)
elif PY3:
def to_bytes(s):
if isinstance(s, bytes):
return s
if isinstance(s, str):
return s.encode('utf-8')
def to_str(s):
if isinstance(s, bytes):
return s.decode('utf-8')
if isinstance(s, str):
return s
def bchr(s):
return bytes([s])
def bord(s):
return s
class RNCryptor(object):
"""Cryptor for RNCryptor"""
AES_BLOCK_SIZE = AES.block_size
AES_MODE = AES.MODE_CBC
SALT_SIZE = 8
def pre_decrypt_data(self, data):
""" Change this function for handling data before decryption. """
data = to_bytes(data)
return data
def post_decrypt_data(self, data):
""" Removes useless symbols which appear over padding for AES (PKCS#7). """
data = data[:-bord(data[-1])]
return to_str(data)
def decrypt(self, data, password):
data = self.pre_decrypt_data(data)
password = to_<PASSWORD>(password)
n = len(data)
version = data[0]
options = data[1]
encryption_salt = data[2:10]
hmac_salt = data[10:18]
iv = data[18:34]
cipher_text = data[34:n - 32]
hmac = data[n - 32:]
encryption_key = self._pbkdf2(password, encryption_salt)
hmac_key = self._pbkdf2(password, hmac_salt)
if self._hmac(hmac_key, data[:n - 32]) != hmac:
raise Exception("Bad data")
decrypted_data = self._aes_decrypt(encryption_key, iv, cipher_text)
return self.post_decrypt_data(decrypted_data)
def pre_encrypt_data(self, data):
""" Does padding for the data for AES (PKCS#7). """
data = to_bytes(data)
rem = self.AES_BLOCK_SIZE - len(data) % self.AES_BLOCK_SIZE
return data + bchr(rem) * rem
def post_encrypt_data(self, data):
""" Change this function for handling data after encryption. """
return data
def encrypt(self, data, password):
data = self.pre_encrypt_data(data)
password = <PASSWORD>(password)
encryption_salt = self.encryption_salt
encryption_key = self._pbkdf2(password, encryption_salt)
hmac_salt = self.hmac_salt
hmac_key = self._pbkdf2(password, hmac_salt)
iv = self.iv
cipher_text = self._aes_encrypt(encryption_key, iv, data)
version = b'\x03'
options = b'\x01'
new_data = b''.join([version, options, encryption_salt, hmac_salt, iv, cipher_text])
encrypted_data = new_data + self._hmac(hmac_key, new_data)
return self.post_encrypt_data(encrypted_data)
@property
def encryption_salt(self):
return Random.new().read(self.SALT_SIZE)
@property
def hmac_salt(self):
return Random.new().read(self.SALT_SIZE)
@property
def iv(self):
return Random.new().read(self.AES_BLOCK_SIZE)
def _aes_encrypt(self, key, iv, text):
return AES.new(key, self.AES_MODE, iv).encrypt(text)
def _aes_decrypt(self, key, iv, text):
return AES.new(key, self.AES_MODE, iv).decrypt(text)
def _hmac(self, key, data):
return hmac.new(key, data, hashlib.sha256).digest()
def _pbkdf2(self, password, salt, iterations=10000, key_length=32):
return KDF.PBKDF2(password, salt, dkLen=key_length, count=iterations,
prf=lambda p, s: hmac.new(p, s, hashlib.sha1).digest())
def main():
from time import time
cryptor = RNCryptor()
###################password,text#########################
#F17T7AN3HG71
password = "<PASSWORD>($@^"
########encrypt###########
yourimail_password = "<PASSWORD>"
encrypt_password = cryptor.encrypt(yourimail_password,password)
en_yourimail_password = base64.b64encode(encrypt_password)
#######decrypt##########
yourimail_password_base64encode = "<KEY>
base64decodetext = base64.b64decode(yourimail_password_base64encode)
de_yourimail_password = cryptor.decrypt(base64decodetext,password)
#####################################################################################################################################################
#encrypt
print("\n\n\n\n密文:\n",en_yourimail_password)
print("\n\n\n");
#=======================================================
#decrypt
print("明文:\n",de_yourimail_password)
print("\n\n\n\n\n")
#=======================================================
if __name__ == '__main__':
main()
| d8886da4e3098fb83fd93885d69621c798535c4b | [
"Markdown",
"Python"
] | 2 | Markdown | Vxer-Lee/RNCryptor | 53ff159a47300ddaa6d4a5be7948a056710992ba | efe9d22f0767dc3d5649dcd4b0c070192ab928c8 |
refs/heads/main | <file_sep>const colors = ['#DE6E51', '#E8A339', '#68B253'];
const sampleData = [
{
"name": "Feed",
"unit": "Kilograms per hector (kg/ha)",
"values": [
{
"timestamp": "2021-01-15T10:00:00.000",
"value": 0.5
},
{
"timestamp": "2021-01-16T10:00:00.000",
"value": 1.5
},
{
"timestamp": "2021-01-17T10:00:00.000",
"value": 1.4
},
{
"timestamp": "2021-01-18T10:00:00.000",
"value": 1.3
},
{
"timestamp": "2021-01-19T10:00:00.000",
"value": 1.2
},
{
"timestamp": "2021-01-20T10:00:00.000",
"value": 2.5
},
{
"timestamp": "2021-01-21T10:00:00.000",
"value": 2.0
}
]
},
{
"name": "Biomass",
"unit": "Kilograms per hector (kg/ha)",
"values": [
{
"timestamp": "2021-01-15T10:00:00.000",
"value": 10
},
{
"timestamp": "2021-01-16T10:00:00.000",
"value": 11
},
{
"timestamp": "2021-01-17T10:00:00.000",
"value": 12
},
{
"timestamp": "2021-01-18T10:00:00.000",
"value": 12
},
{
"timestamp": "2021-01-19T10:00:00.000",
"value": 13.5
},
{
"timestamp": "2021-01-20T10:00:00.000",
"value": 9
},
{
"timestamp": "2021-01-21T10:00:00.000",
"value": 8
}
]
},
{
"name": "Ammonia",
"unit": "Kilograms per hector (kg/ha)",
"values": [
{
"timestamp": "2021-01-15T10:00:00.000",
"value": 5.1
},
{
"timestamp": "2021-01-16T10:00:00.000",
"value": 7
},
{
"timestamp": "2021-01-17T10:00:00.000",
"value": 7
},
{
"timestamp": "2021-01-18T10:00:00.000",
"value": 6
},
{
"timestamp": "2021-01-19T10:00:00.000",
"value": 5
},
{
"timestamp": "2021-01-20T10:00:00.000",
"value": 4
},
{
"timestamp": "2021-01-21T10:00:00.000",
"value": 3.1
}
]
}
];
// set the dimensions and margins of the graph
const margin = { top: 40, right: 80, bottom: 60, left: 50 },
width = 960 - margin.left - margin.right,
height = 280 - margin.top - margin.bottom;
const parseDate = d3.timeParse("%m/%d/%Y");
const formatDate = d3.timeFormat("%m %d, %Y");
const formatMonth = d3.timeFormat("%b");
const x = d3.scaleTime().range([0, width]);
const y = d3.scaleLinear().range([height, 0]);
// append the svg object to the body of the page
const svg = d3
.select("#root")
.append("svg")
.attr(
"viewBox",
`0 0 ${width + margin.left + margin.right} ${height + margin.top + margin.bottom}`)
.append("g")
.attr("transform", "translate(" + margin.left + "," + margin.top + ")");
/* x axis */
svg
.append("g")
.attr("class", "x axis")
.attr("transform", "translate(0," + height + ")")
.call(d3.axisBottom(x).ticks(3).tickFormat(formatDate));
/* y axis */
svg.append("g").attr("class", "y axis").call(d3.axisLeft(y));
/* y axis lable */
svg
.append("text")
.attr("y", -20)
.attr("x", -20)
.attr("dy", "1em")
.style("text-anchor", "middle")
.text("kg/ha");
appendData(sampleData);
function appendData(sampleData) {
/* combine all values for domain */
const allValues = sampleData.map(d => d.values).flat();
sampleData.forEach((line, index) => {
// line.values = line.values.reverse();
line.values.forEach((d) => {
d.timestamp = new Date(d.timestamp);
d.value = Number(d.value);
});
x.domain(
d3.extent(allValues, (d) => d.timestamp)
);
y.domain([
0,
d3.max(allValues, (d) => d.value),
]);
/* animate axis */
svg
.select(".x.axis")
.transition()
.duration(750)
.call(d3.axisBottom(x).ticks(3).tickFormat(formatDate));
svg
.select(".y.axis")
.transition()
.duration(750)
.call(d3.axisLeft(y));
/* The data lines */
const linePath =
svg.append("path")
.data([line.values])
.attr("class", `line-${index}`)
.attr("d", d3
.line()
.x((d) => x(d.timestamp))
.y((d) => y(d.value))
.curve(d3.curveCardinal))
/* data point circles */
line.values.forEach(d => {
svg.selectAll()
.data([d])
.enter()
.append("circle")
.attr("fill", "white")
.attr("stroke", colors[index])
.attr("cx", (d) => x(d.timestamp))
.attr("cy", (d) => y(d.value))
.attr("r", 3)
})
const pathLength = linePath.node().getTotalLength();
linePath
.attr("stroke-dasharray", pathLength)
.attr("stroke-dashoffset", pathLength)
.attr("stroke-width", 3)
.transition()
.duration(1000)
.attr("stroke-width", 0)
.attr("stroke-dashoffset", 0);
/* dynamic legend text */
svg
.append("text")
.attr("class", "title")
.attr("x", (width / 2) + 100 * (index - 1))
.attr("y", 0 - margin.top / 2)
.attr("text-anchor", "left")
.text(line.name);
/* dynamic legend dots */
svg.append("circle")
.attr("fill", colors[index])
.attr("stroke", "none")
.attr("cx", ((width / 2) + 100 * (index - 1)) - 8)
.attr("cy", -4 - margin.top / 2)
.attr("r", 4)
const focus = svg
.append("g")
.attr("class", "focus")
.style("display", "none");
focus
.append("line")
.attr("class", "x")
.style("stroke-dasharray", "3,3")
.style("opacity", 0.5)
.attr("y1", 0)
.attr("y2", height);
focus
.append("line")
.attr("class", "y")
.style("stroke-dasharray", "3,3")
.style("opacity", 0.5)
.attr("x1", width)
.attr("x2", width);
focus
.append("circle")
.attr("class", "y")
.style("fill", "none")
.attr("r", 4);
focus.append("text").attr("class", "y1").attr("dx", 8).attr("dy", "-.3em");
focus.append("text").attr("class", "y2").attr("dx", 8).attr("dy", "-.3em");
focus.append("text").attr("class", "y3").attr("dx", 8).attr("dy", "1em");
focus.append("text").attr("class", "y4").attr("dx", 8).attr("dy", "1em");
function mouseMove(event) {
const bisect = d3.bisector((d) => d.timestamp).left;
const x0 = x.invert(d3.pointer(event, this)[0]);
const i = bisect(line.values, x0, 1);
const d0 = line.values[i - 1];
const d1 = line.values[i];
const d = x0 - d0.timestamp > d1.timestamp - x0 ? d1 : d0;
focus
.select("circle.y")
.attr("transform", "translate(" + x(d.timestamp) + "," + y(d.value) + ")");
focus
.select("text.y1")
.attr("transform", "translate(" + x(d.timestamp) + "," + y(d.value) + ")")
.text(`${line.name} ${d.value} ${line.unit}`);
/* focus
.select("text.y2")
.attr("transform", "translate(" + x(d.timestamp) + "," + y(d.value) + ")")
.text(d.value); */
focus
.select("text.y3")
.attr("transform", "translate(" + x(d.timestamp) + "," + y(d.value) + ")")
.text(formatDate(d.timestamp));
focus
.select("text.y4")
.attr("transform", "translate(" + x(d.timestamp) + "," + y(d.value) + ")")
.text(formatDate(d.timestamp));
focus
.select(".x")
.attr("transform", "translate(" + x(d.timestamp) + "," + y(d.value) + ")")
.attr("y2", height - y(d.value));
focus
.select(".y")
.attr("transform", "translate(" + width * -1 + "," + y(d.value) + ")")
.attr("x2", width + width);
}
svg
.append("rect")
.attr("width", width)
.attr("height", height)
.style("fill", "none")
.style("pointer-events", "all")
.on("mouseover", () => {
focus.style("display", null);
})
.on("mouseout", () => {
focus.style("display", "none");
})
.on("touchmove mousemove", mouseMove);
});
}
<file_sep># d3-line-chart
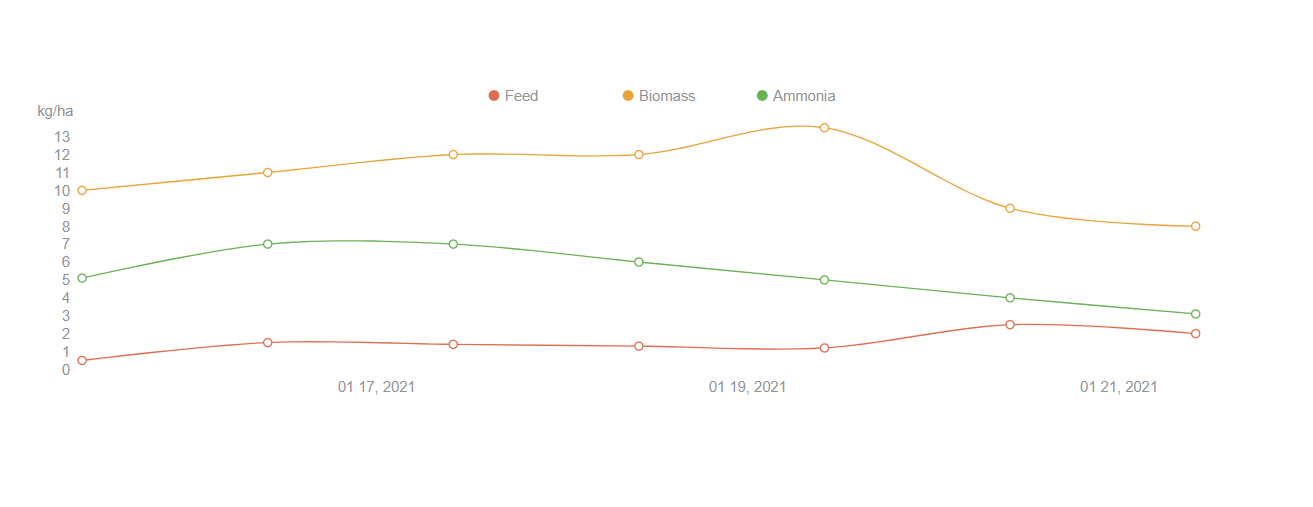
***This is an in-progress peek*** <br/>
<p>The chart is setup to handle a non-specified number of lines. I could set the color designation to loop back around, if there were ever more lines than colors. I used a color picker to find the colors and wrote a color array to dynamically color each line as it is looped. Typically, if an API endpoint is not available to set up a microservice, I would run a Json server, to access the initial data from a local json file, like the file I included, but did not use, yet. This data is a hard-coded const. Often my first step for a rough-in.</p>
<p>For interactivity, when you hover over data points, the info for the nearest focussed point should display. Currently, only the last line in the loop works, and I have that in my list of todos, below. </p>
***With more time, I would:*** <br/>
<ul>
<li>refine style, including adding subtle y grid dashed lines, etc. (I could see the very subtle horizontal grid in the assignment doc)</li>
<li>figure out how to have the data lines begin and end before and after the first and last data points</li>
<li>setup promised data (I chose not to use d3.csv, because the provided data sample was basically, already Json. I could have formatted the JSON into csv and posted it to a remote location to grab as a promise.)</li>
<li>make mouse move work for all lines</li>
<li>move the chart into React (The example I demoed, durring the first interview, was a d3.js bar chart reusable component with a live API search in React)</li>
<li>figure out what's going on with the y axis initial format, before the animation</li>
</ul>
| 1e74e259fb9dd3bd84443645366dbca06af15be9 | [
"JavaScript",
"Markdown"
] | 2 | JavaScript | JeffACDev/shrimpy-d3 | 6c086ae77ea29d89a1a31292b9c00a8947626231 | cdc03ee410bddf726c301163255df8cbc9f4c1dd |
refs/heads/main | <file_sep>
# solver.py
def solve(gr):
find = find_empty(gr)
if not find:
return True
else:
row, col = find
for i in range(1,10):
if valid(gr, i, (row, col)):
gr[row][col] = i
if solve(gr):
return True
gr[row][col] = 0
return False
def valid(gr, num, pos):
# Checking row
for i in range(len(gr[0])):
if gr[pos[0]][i] == num and pos[1] != i:
return False
# Checking column
for i in range(len(gr)):
if gr[i][pos[1]] == num and pos[0] != i:
return False
# Checking 3x3 subgrid
gr_x = pos[1] // 3
gr_y = pos[0] // 3
for i in range(gr_y*3, gr_y*3 + 3):
for j in range(gr_x * 3, gr_x*3 + 3):
if gr[i][j] == num and (i,j) != pos:
return False
return True
def print_grid(gr):
for i in range(len(gr)):
if i % 3 == 0 and i != 0:
print("- - - - - - - - - - - -")
for j in range(len(gr[0])):
if j % 3 == 0 and j != 0:
print(" | ", end="")
print(gr[i][j], end = ' ')
print()
def find_empty(gr):
for i in range(len(gr)):
for j in range(len(gr[0])):
if gr[i][j] == 0:
return (i, j) # row, col
return None<file_sep># Sudoku-solver-py
## Designed a sudoku solver in python using pygame library.
* I have used backtracking algorithm to solve the sudoku puzzle.
For GUI I have used pygame library of python.
To input your number click on the blank box and type in the number and press enter:
- If your value is correct then it will take it otherwise a cross will be shown at bottom left corner.
- To clear your choice provided it is wrong press : 'DEL'
If you want the computer to solve it - Simply press 'SPACE'
THANK YOU FOR VISITING THIS REPO!!
| e019fe2e2875978aeca2e015d1d2f81fc0811ab2 | [
"Markdown",
"Python"
] | 2 | Python | shubham25121999/Sudoku-solver-py | 26e4e49ea5c597683ef8c057c028fcf29fcdf8c2 | 72aa15a3ae387e7bf930fec2dd5f38c86afb7326 |
refs/heads/master | <repo_name>eduardoacv2/StoreApp-EL-ICOM5016<file_sep>/appjs/account.js
function Account(uId, uFirstName, uLastName, uBillingAddress, uCreditCard, aType, uBuyHistory, uSellHistory, uPInventory, uPassword)
{
this.uId = "123";
this.uFirstName = fName;
this.uLastName = lName;
this.uBillingAddress = address;
this.uCreditCard = cardNumber;
this.aType = accountType;
this.uBuyHistory = productList1;
this.uSellHistory = productList2;
this.uPInventory = productList3;
this.uPassword = <PASSWORD>;
this.toJSON = toJSON;
}
<file_sep>/appjs/app.js
$(document).on('pagebeforeshow', "#product", function( event, ui ) {
console.log("John");
$.ajax({
url : "http://localhost:3412/StoreApp/product",
contentType: "application/json",
success : function(data, textStatus, jqXHR){
var productList = data.product;
var len = productList.length;
var list = $("#product-list");
list.empty();
var product;
for (var i=0; i < len; ++i){
product = productList[i];
list.append("<li><a onclick=GetProduct(" + product.pName+ ")>" +
"<h2>" + product.pType1 + " " + product.pType2 + "</h2>" +
"<p><strong> Seller: " + product.pSeller + "</strong></p>" +
"<p>" + product.pCondition + "</p>" +
"<p class=\"ui-li-aside\">" + accounting.formatMoney(product.pBidPrice) + "</p>" +
"</a></li>");
}
list.listview("refresh");
},
error: function(data, textStatus, jqXHR){
console.log("textStatus: " + textStatus);
alert("Data not found!");
}
});
});
$(document).on('pagebeforeshow', "#product-view", function( event, ui ) {
$("#upd-type").val(currentProduct.pType1);
$("#upd-typeOfType").val(currentProduct.pType2);
$("#upd-name").val(currentProduct.pName);
$("#upd-price").val(currentProduct.pPrice);
$("#upd-condition").val(currentProduct.pCondition);
$("#upd-startingBid").val(currentProduct.pStartingBid);
$("#upd-bidPrice").val(currentProduct.pBidPrice);
$("#upd-seller").val(currentProduct.pSeller);
$("#upd-buyer").val(currentProduct.pBuyer);
$("#upd-image").val(currentProduct.pImage);
});
////////////////////////////////////////////////////////////////////////////////////////////////
/// Functions Called Directly from Buttons ///////////////////////
function ConverToJSON(formData){
var result = {};
$.each(formData,
function(i, o){
result[o.name] = o.value;
});
return result;
}
function SaveProduct(){
$.mobile.loading("show");
var form = $("#product-form");
var formData = form.serializeArray();
console.log("form Data: " + formData);
var newProduct = ConverToJSON(formData);
console.log("New Car: " + JSON.stringify(newProduct));
var newProductJSON = JSON.stringify(newProduct);
$.ajax({
url : "http://localhost:3412/StoreApp/product",
method: 'post',
data : newProductJSON,
contentType: "application/json",
dataType:"json",
success : function(data, textStatus, jqXHR){
$.mobile.loading("hide");
$.mobile.navigate("#product");
},
error: function(data, textStatus, jqXHR){
console.log("textStatus: " + textStatus);
$.mobile.loading("hide");
alert("Data could not be added!");
}
});
}
var currentPro = {};
function GetProduct(id){
$.mobile.loading("show");
$.ajax({
url : "http://localhost:3412/StoreApp/product/" + id,
method: 'get',
contentType: "application/json",
dataType:"json",
success : function(data, textStatus, jqXHR){
currentPro = data.product;
$.mobile.loading("hide");
$.mobile.navigate("#product-view");
},
error: function(data, textStatus, jqXHR){
console.log("textStatus: " + textStatus);
$.mobile.loading("hide");
if (data.status == 404){
alert("Product not found.");
}
else {
alter("Internal Server Error.");
}
}
});
}
function UpdateProduct(){
$.mobile.loading("show");
var form = $("#product-view-form");
var formData = form.serializeArray();
console.log("form Data: " + formData);
var updPro = ConverToJSON(formData);
updPro.id = currentPro.id;
console.log("Updated Product: " + JSON.stringify(updPro));
var updProJSON = JSON.stringify(updPro);
$.ajax({
url : "http://localhost:3412/StoreApp/product/" + updPro.id,
method: 'put',
data : updProJSON,
contentType: "application/json",
dataType:"json",
success : function(data, textStatus, jqXHR){
$.mobile.loading("hide");
$.mobile.navigate("#product");
},
error: function(data, textStatus, jqXHR){
console.log("textStatus: " + textStatus);
$.mobile.loading("hide");
if (data.status == 404){
alert("Data could not be updated!");
}
else {
alert("Internal Error.");
}
}
});
}
function DeleteProduct(){
$.mobile.loading("show");
var id = currentPro.id;
$.ajax({
url : "http://localhost:3412/StoreApp/product/" + id,
method: 'delete',
contentType: "application/json",
dataType:"json",
success : function(data, textStatus, jqXHR){
$.mobile.loading("hide");
$.mobile.navigate("#product");
},
error: function(data, textStatus, jqXHR){
console.log("textStatus: " + textStatus);
$.mobile.loading("hide");
if (data.status == 404){
alert("Product not found.");
}
else {
alter("Internal Server Error.");
}
}
});
}<file_sep>/appjs/product.js
function Product(pId, pName, pType1, pType2, pBidPrice, pPrice, pStartingBid, pCondition, pImage, pSeller, pBuyer)
{
this.pId = "0";
this.pName = name;
this.pType1 = type;
this.pType2 = typeOfType;
this.pBidPrice = bidPrice;
this.pPrice = price;
this.pStartingBid = startingBid;
this.pCondition = condition;
this.pImage = image;
this.pSeller = seller;
this.pBuyer = buyer;
this.toJSON = toJSON;
}
| 3c3b17422cfd9ab200caaecd2304876ea7648dc8 | [
"JavaScript"
] | 3 | JavaScript | eduardoacv2/StoreApp-EL-ICOM5016 | 97beddc119fe83b6107545d71859eb9e6adbd2ef | d002af41659536480e0843973442a226de1d9230 |
refs/heads/main | <repo_name>Gideon-Zozingao/express-sessions-and-cookies<file_sep>/README.md
# express-sessions-and-cookies | c3c0d430d0130cb0f31a6aa5764f9bf78ad2c0b5 | [
"Markdown"
] | 1 | Markdown | Gideon-Zozingao/express-sessions-and-cookies | cad4265f3dcdcc9ad52c585bb36d70c300b5ca14 | 4460bfbd9308779bc59ce91a4571b808d64e9c94 |
refs/heads/main | <file_sep># :oncoming_bus: Real Time Bus Tracker :oncoming_bus:
On this project, the goal was to make an API connection to gather the bus geolocation every 20 seconds and tracks its route.
# :bomb: How to run
1. Download the repository files
2. Get an API token at MapBox's website and replace it on the mapanimation.js
3. Open the index.html with your favorite browser
# :dart: Future Improvements:
- Calculate and show an ETA (Estimated Time of Arrival), based on historic information
# :registered: Licence Information:
Copyright 2021 <NAME>
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
<file_sep>const colors = ['#4d0ff1', '#6e0723', '#65202e', '#844aef', '#11a94c', '#5af68e', '#838452', '#1ac8cb', '#4e42e8', '#5d6eeb', '#0d3155', '#60d4d0', '#7ecdf3', '#d00cdd', '#af534b', '#fd1269', '#101ff8', '#9aa72c', '#cc5446', '#1a3ab7']
mapboxgl.accessToken = '<KEY>';
let map = new mapboxgl.Map({
container: 'map',
style: 'mapbox://styles/mapbox/streets-v11',
center: [-71.104081, 42.365554],
zoom: 14,
});
map.resize();
var busMark = [];
async function run() {
const locations = await getBusLocation();
console.log(location);
location.forEach(bus, i) => {
var marker = new mapboxgl.marker({'color': colors[i] })
.setLngLat([bus.attributes.longitude, bus.attributes.latitude])
.setPopup(new mapboxgl.Popup({offset: 25, closeOnClick: false, closeButton: false}).setHTML(`<h3>Bus ID <br>${bus.attributes.label}</h3>`))
.addTo(map)
.toogglePopup();
busesMarkers.push(marker);
});
function deleteMark() {
if (busMark!==null) {
for (var i = busMark.length - 1; i >=0; i--) {
busMark[i].remove();
}
}
}
locations.forEach((marker, i) => {
let popUp = document.getElementsByClassName('mapboxgl-popup-content');
popUp[i].style.background = colors[i];
});
setTimeout(deleteMark, 8000);
setTimeout(run, 20000);
}
async function getBusLocation() {
const url = 'https://api-v3.mbta.com/vehicles?filter[route]=1&include=trip';
const response = await fetch(url);
const json = await response.json();
return json.data;
}
map.on('load', function() {
run();
}); | 41ea3281db549d3c2add7b65d22278aedeb6aa9d | [
"Markdown",
"JavaScript"
] | 2 | Markdown | ejhenriques/Real_Time_Bus_Tracker | 38b094bd6852613c528425537d69f40462eec6d8 | 041722a438b978b4ba7ecca8aaf9e208da79111c |
refs/heads/master | <file_sep>import numpy as np
import cv2
from matplotlib import pyplot as plt
# 1.載入原始圖像並灰值化,原始影像為1000x2000 ,3 channel #灰:1 #RGB:3
img = cv2.imread('digits.png')
print("img shape=", img.shape)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)#RGB變成GRAY
print("gray shape=", gray.shape)
# 2.切成每一小塊20x20 pixel
# 先將gray 1000x2000 [rowsxcols] pixel , 將row=1000/50 :意為50列20pixel單位
# 再將產生的cols =2000/100 =>意為100行20pixel單位
cells = [np.hsplit(row, 100) for row in np.vsplit(gray, 50)]
# 3.將cells為一50x100 的list轉成array (50,100,20,20)
x = np.array(cells)
print("x shape=", x.shape)
# 4.把20x20 pixel 展平成一行400 pixel
# 將cells array X 轉成5000x400 後並分成兩半 train's data and test's data
train = x[:, :50].reshape(-1, 400).astype(np.float32) # Size = (2500,400)
test = x[:, 50:100].reshape(-1, 400).astype(np.float32) # Size = (2500,400)
print("train shape=", train.shape)
print("test shape=", test.shape)
# 5.Create labels for train and test data
k = np.arange(10)
train_labels = np.repeat(k, 250)[:, np.newaxis]
test_labels = train_labels.copy()
print("train_labels.shape=", train_labels.shape)
print("test_labels.shape=", test_labels.shape)
# 6. Initiate kNN, train the data, then test it with test data for k=1
knn = cv2.ml.KNearest_create()
knn.train(train, cv2.ml.ROW_SAMPLE, train_labels)
ret, result, neighbours, dist = knn.findNearest(test, k=1)
# 7. Now we check the accuracy of classification
# For that, compare the result with test_labels and check which are wrong
matches = result == test_labels
correct = np.count_nonzero(matches)
accuracy = correct * 100.0 / result.size
print(accuracy)
# 8.save the data
np.savez('knn_data.npz', train=train, train_labels=train_labels)
##-------------------------------------------------------------------------
##=========================================================================
##======Predict testing====================================================
# A.Now re-load the data
with np.load('knn_data.npz') as data:
print(data.files)
train = data['train']
train_labels = data['train_labels']
# B.輸入自己手寫的image data 必須是20x20 pixel
Input_Numer = [0] * 10
img_num = [0] * 10
img_res = [0] * 10
testData_r = [0] * 10
result = [0] * 10
result_str = [0] * 10
Input_Numer[0] = "0.png"
Input_Numer[1] = "1.png"
Input_Numer[2] = "2.png"
Input_Numer[3] = "3.png"
Input_Numer[4] = "4.png"
Input_Numer[5] = "5.png"
Input_Numer[6] = "6.png"
Input_Numer[7] = "7.png"
Input_Numer[8] = "8.png"
Input_Numer[9] = "9.png"
font = cv2.FONT_HERSHEY_SIMPLEX
# C.Predicting
for i in range(10): # input 10 number
img_num[i] = cv2.imread(Input_Numer[i], 0)
testData_r[i] = img_num[i][:, :].reshape(-1, 400).astype(np.float32) # Size = (1,400)
ret, result[i], neighbours, dist = knn.findNearest(testData_r[i], k=5)
# 產生white screen以顯示預測結果的白底
img_res[i] = np.zeros((64, 64, 3), np.uint8)
img_res[i][:, :] = [255, 255, 255]
# 將結果轉成字串以便顯示在圖上
print("result[i][0][0] =", result[i][0][0].astype(np.int32)) # change type from float32 to int32
result_str[i] = str(result[i][0][0].astype(np.int32))
if result[i][0][0].astype(np.int32) == i:
cv2.putText(img_res[i], result_str[i], (15, 52), font, 2, (0, 255, 0), 3, cv2.LINE_AA)
else:
cv2.putText(img_res[i], result_str[i], (15, 52), font, 2, (255, 0, 0), 3, cv2.LINE_AA)
# ===顯示輸入與預測結果圖======
Input_Numer_name = ['Input 0', 'Input 1', 'Input 2', 'Input 3', 'Input 4', \
'Input 5', 'Input 6', 'Input 7', 'Input8', 'Input9']
predict_Numer_name = ['predict 0', 'predict 1', 'predict 2', 'predict 3', 'predict 4', \
'predict 5', 'predict6 ', 'predict 7', 'predict 8', 'predict 9']
for i in range(10):
plt.subplot(2, 10, i + 1), plt.imshow(img_num[i], cmap='gray')
plt.title(Input_Numer_name[i]), plt.xticks([]), plt.yticks([])
plt.subplot(2, 10, i + 11), plt.imshow(img_res[i], cmap='gray')
plt.title(predict_Numer_name[i]), plt.xticks([]), plt.yticks([])
plt.show()
| 950bee852e439323a5683c2a6648c158dbaefb14 | [
"Python"
] | 1 | Python | luyihsien/knn-project | 6370415f6795eeaa6cfbbba08f796e4c758cbee7 | 1f03fa2cd39e49a9c8e07946ff2e4f6476d6d770 |
refs/heads/master | <repo_name>JaarenDSacharow/BurgerBuilder<file_sep>/src/containers/BurgerBuilder/BurgerBuilder.js
import React, {Component} from 'react';
import Aux from '../../hoc/Aux/Aux';
import Burger from '../../components/Burger/Burger';
import BuildControls from '../../components/Burger/BuildControls/BuildControls';
import Modal from '../../components/UI/Modal/Modal';
import OrderSummary from '../../components/Burger/OrderSummary/OrderSummary';
import Spinner from '../../components/UI/Spinner/Spinner';
//HOC to wrap this
import withErrorHandler from '../../hoc/WithErrorHandler/WithErrorHandler';
// our own axios instance
import axios from '../../axios-orders-instance';
//global prices to use for calculations
const INGREDIENT_PRICES = {
salad : 0.2,
cheese: 0.3,
meat: 1.0,
bacon: 0.3
}
//the main stateful class component (container)
// keep state and handlers in containers.
class BurgerBuilder extends Component {
// constructor(props) {
// super(props);
// this.state = {};
// }
state = {
ingredients: null, //<-- this is stored in firebase
totalPrice: 5, //base price
purchaseable: false, //for enabling/disabling the order now button
purchasing: false, //for determining if we are in the modal or not
loading: false, //for checking whether or not to show the spinner
error: false // in case the ingredients can't be loaded, we can get rid of the spinner.
}
componentDidMount(){
axios.get('/ingredients.json') //<-- it's firebase, don't forget the JSON extension or you'll get a CORS error
.then((response) =>{
this.setState({
ingredients: response.data
})
}).catch((error)=>{
this.setState({
error: true //this is the last line of defense to display an error to the user
})
})
}
updatePurchaseableState = (updatedIngredients) => {
// const ingredients = {
// ...this.state.ingredients
// };
//take the key and return the value in a new array,
//which we then reduce to get the s
const sum = Object.keys(updatedIngredients)
.map((ingredientKey)=>{
return updatedIngredients[ingredientKey]
})
.reduce((sum, el) =>{
return sum + el
}, 0);
this.setState({purchaseable: sum > 0});
}
purchasingHandler = () =>{
this.setState({
purchasing: true
})
}
purchaseCancelHandler = () => {
this.setState({
purchasing: false
})
}
//here's where we hit the firebase instance
purchaseContinueHandler = () => {
// alert('You Continued!');
//first set loading to true
this.setState({
loading : true
})
//this is a firebase specific thing for real time database
// firbase realtime database creates nodes with data based
// on this request
// you target the base URL node with a .json extension
//it will store data beneath that node
const order = {
ingredients: this.state.ingredients,
price: this.state.totalPrice, //recalc the price on the server in a real app
customer: {
name: '<NAME>',
address : {
street: '123 test street',
city: 'test city',
zip: '123456',
country: 'US'
},
email: "<EMAIL>"
},
deilveryMethod: 'fastest'
}
axios.post('/orders.json', order )
.then((response) =>{
//when you get the response, set loading to false to hide spinner
// and also set purchasing to false to have the modal leave
this.setState({
loading: false,
purchasing: false
})
console.log(response);
}).catch((error) => {
this.setState({
loading: false,
purchasing: false
})
})
}
addIngredientHandler = (type) => {
const oldCount = this.state.ingredients[type];
const newCount = oldCount + 1;
//update state immutably by making a copy with the spread operator
// since we're dealing with an entire object
const updatedIngredients = {
...this.state.ingredients
};
updatedIngredients[type] = newCount;
//now update the price based on the constant above
const oldPrice = this.state.totalPrice;
const priceAdd = INGREDIENT_PRICES[type]; //reference a key, not a property :/
const newPrice = oldPrice + priceAdd;
this.setState({
ingredients: updatedIngredients,
totalPrice : newPrice
});
this.updatePurchaseableState(updatedIngredients);
}
removeIngredientHandler = (type) => {
const oldCount = this.state.ingredients[type];
// to prevent sending a -1 array key to the burger component
// do nothing to update the amount of ingredients if a user
//tries to remove an ingredient whose count is 0
if (oldCount <= 0) {
return;
}
const newCount = oldCount - 1;
//update state immutably by making a copy with the spread operator
// since we're dealing with an entire object
const updatedIngredients = {
...this.state.ingredients
};
updatedIngredients[type] = newCount;
//now update the price based on the constant above
const oldPrice = this.state.totalPrice;
// to prevent sending a deduction when there are no ingredients
let priceToRemove = 0;
if (oldCount !==0) {
priceToRemove = INGREDIENT_PRICES[type]; //reference a key, not a property :/
}
const newPrice = oldPrice - priceToRemove;
this.setState({
ingredients: updatedIngredients,
totalPrice : newPrice
});
this.updatePurchaseableState(updatedIngredients);
}
render(){ //required lifecycle method
//let's add a check here to disable the "LESS" button of a given Build Control
//if there are no ingredients to take away
const disabledButtonInfo = {
...this.state.ingredients
}
for (let key in disabledButtonInfo ){
disabledButtonInfo[key] = disabledButtonInfo[key] <= 0
}
//here we have a check for loading state, replacing the order summary with a spinner
// but now that we're retriving ingredients from firebase, we need to also check for
//this.state.ingredients
let orderSummary = null;
if(this.state.ingredients){
orderSummary = <OrderSummary
ingredients={this.state.ingredients}
price={this.state.totalPrice}
cancel={this.purchaseCancelHandler}
continue={this.purchaseContinueHandler}
/>
}
if (this.state.loading) {
orderSummary = <Spinner />
}
//here we check to see if the burger's ingredients have been populated into state
//if not, show a spinner.
// if you don't have these checks, the app will break because the state isn't
//ready yet, as it relies on an external call
let burger = !this.state.error ? <Spinner /> : <p>Ingredients can't be loaded</p>
if(this.state.ingredients){
burger =
<Aux>
<Burger ingredients={this.state.ingredients} />
<BuildControls
ingredientAdded={this.addIngredientHandler}
ingredientRemoved={this.removeIngredientHandler}
disabled={disabledButtonInfo}
price={this.state.totalPrice}
purchaseable={!this.state.purchaseable}
ordered={this.purchasingHandler}
/>
</Aux>
}
return(
<Aux>
<Modal
show={this.state.purchasing}
modalClosed={this.purchaseCancelHandler}>
{orderSummary}
</Modal>
{burger}
</Aux>
)
}
}
//wrap this component with the HOC to display a global error modal
export default withErrorHandler(BurgerBuilder, axios);<file_sep>/src/components/Burger/OrderSummary/OrderSummary.js
import React from 'react';
import './OrderSummary.css';
import Aux from '../../../hoc/Aux/Aux';
import Button from '../../UI/Button/Button';
const OrderSummary = (props) => {
//create an array with the object's keys, again
const ingredientSummary = Object.keys(props.ingredients)
.map((ingredientKey, index) => {
return(
//now use that key and the corresponding value for it for display
<li key={ingredientKey}><span style={{textTransform:'capitalize'}}>{ingredientKey}</span>: {props.ingredients[ingredientKey]}</li>
)
});
return(
<Aux>
<h3>Your Order</h3>
<p>Delicious burger with the follwing ingredients</p>
<ul>
{ingredientSummary}
</ul>
<p>Total Price: ${props.price.toFixed(2)}</p>
<p>Continue to Checkout?</p>
<Button clicked={props.cancel} btnType={"Danger"}>Cancel</Button>
<Button clicked={props.continue} btnType={"Success"}>Continue</Button>
</Aux>
)
}
export default OrderSummary;<file_sep>/src/components/Burger/Burger.js
import React from 'react';
import './Burger.css';
import BurgerIngredient from './BurgerIngredient/BurgerIngredient';
import Aux from '../../hoc/Aux/Aux';
const Burger = (props) => {
//transformed ingredients take the key value pairs
//from the props and returns an array based on the value
// of the ingredients and the number (key : value)
//Object.keys returns an array based on the object keys
let transFormedIngredients = Object.keys(props.ingredients)
.map((ingredientKey) => {
console.log(ingredientKey);
//props.ingredients[ingredientsKey] returns the value from that key
console.log(props.ingredients[ingredientKey]);
//now we use the Array method to create a new array of a length that matches the key above
//spread operator creates a new array based on the current key
return[...Array(props.ingredients[ingredientKey])]
//and finally we map that array with a blank arg and index
//and render BurgerIngredient components in a list
.map((_, index) => {
return <BurgerIngredient key={ingredientKey + index} type={ingredientKey}/>
})
})
//after all of this, call reduce on the array to flatten it so we can see
//if there are any ingredients for the purposes of displaying a message
// reduce returns a value based on the elements of the array
// in this case we take each subarray and concat it to the main array
.reduce((arr,el) =>{
return arr.concat(el);
}, []);
console.log(transFormedIngredients.length);
if(transFormedIngredients.length === 0) {
transFormedIngredients = <p>Please begin adding ingredients.</p>;
}
return(
<Aux>
<div className="Burger">
<BurgerIngredient type="bread-top"/>
{transFormedIngredients}
<BurgerIngredient type="bread-bottom"/>
</div>
</Aux>
);
}
export default Burger;<file_sep>/src/hoc/Layout/Layout.js
import React, {Component} from 'react';
import Aux from '../../hoc/Aux/Aux';
import './Layout.css';
import Toolbar from '../../components/Navigation/Toolbar/Toolbar';
import SideDrawer from '../../components/Navigation/SideDrawer/SideDrawer';
//this component is merely a wrapper that returns props.children (see how it's used in APP)
// this is so we can have a universal layout for all child components
// it also includes all navigation and default layouts
//I changed it to a container so I can handle the
//opening and closing of the sidedrawer component
class Layout extends Component {
state = {
showSideDrawer: false
}
sideDrawerClosedHandler = () => {
this.setState({
showSideDrawer: false
})
}
sideDrawerToggle= () => {
this.setState({
showSideDrawer: !this.state.showSideDrawer
})
}
render() {
return(
<Aux>
<Toolbar toggleSideDrawer={this.sideDrawerToggle} />
<SideDrawer
open={this.state.showSideDrawer}
clicked={this.sideDrawerClosedHandler} />
<main className="Content">
{this.props.children}
</main>
</Aux>
);
}
}
export default Layout;<file_sep>/src/components/UI/Modal/Modal.js
import React, {Component} from 'react';
import './Modal.css';
import Aux from '../../../hoc/Aux/Aux';
import Backdrop from '../Backdrop/Backdrop'
class Modal extends Component {
//without shouldComponentUpdate, you'll trigger a rerender
//EVERYTIME state changes in the build control component.
// this lifecycle method improves performance by only rerendering
//if the right props change, in this case, we only want the modal
//to trigger a renender if it is visible.
//NOTE: because this component has children, you have to check for that too
// if you want to display the spinner in the ORDERSUMMARY child component.
shouldComponentUpdate(nextProps, nextState) {
return nextProps.show !== this.props.show || nextProps.children !== this.props.children;
}
componentDidUpdate() {
console.log('[Modal] will update');
}
render(){
return(
<Aux>
<Backdrop show={this.props.show} clicked={this.props.modalClosed}/>
<div style={{
//inline styles to display based on the show prop
transform: this.props.show ? 'translateY(0)' : 'translateY(-100vh)',
opacity: this.props.show ? '1' : '0'
}}
className="Modal"
>
{this.props.children}
</div>
</Aux>
)
}
}
export default Modal; | 4b232894468459ece3f685749997ba6103855980 | [
"JavaScript"
] | 5 | JavaScript | JaarenDSacharow/BurgerBuilder | 3db50112c25508a03373a180d3896cb4e7f1509b | a111dc291d3157f4e5abae74740f32a6cf4be7e2 |
refs/heads/master | <repo_name>jane-great/ChainBook<file_sep>/README.md
## Build Setup
``` bash
# install dependencies
npm install
# serve with hot reload at localhost:8080
npm run dev
# build for production with minification
npm run build
# build for production and view the bundle analyzer report
npm run build --report
# run unit tests
npm run unit
# run all tests
npm test
```
# mongodb install
从mongodb官网现在并安装mongodb
https://www.mongodb.com/download-center?jmp=tutorials&_ga=2.99390234.503106171.1527646565-1887027323.1527385485#atlas
启动mongodb
https://docs.mongodb.com/tutorials/install-mongodb-on-windows/
导入db文件夹中的db初始化文件
<file_sep>/front/src/apis/base.js
export default function (request) {
return {
getCommon() {
return request({ url: 'src/mock/base/test.json' }).then(({ data }) => data);
},
getForText() {
return Promise.resolve({});
}
};
}
<file_sep>/smartcontracts/test/Transaction.test.js
var BuyAndSell = artifacts.require("./BuyAndSell.sol");
var RentAndLease = artifacts.require("./RentAndLease.sol");
var Transaction = artifacts.require("./Transaction.sol");
var BookOwnerShip = artifacts.require("./BookOwnerShip.sol");
contract("Transaction", function(accounts) {
var buys;
var rents;
var book;
var trans;
BookOwnerShip.deployed().then(function(ins) {
book = ins;
})
Transaction.deployed().then(function(ins) {
trans = ins;
})
BuyAndSell.deployed().then(function(ins) {
buys = ins;
})
RentAndLease.deployed().then(function(ins) {
rents = ins;
})
it("初始化成功", async () => {
let addr = await trans.ceoAddress();
assert.equal(addr, accounts[0], "ceo地址设置正确");
await buys.addAddressToWhitelist(trans.address);
await rents.addAddressToWhitelist(trans.address);
})
it("只有ceo可以设置合约地址", async () => {
await trans.setBuyAndSell(buys.address,{from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "error message must contain revert");
});
let buyAddr = await trans.buyAndSell();
assert.equal(buyAddr, 0, "买卖合约地址尚未设置");
await trans.setBuyAndSell(buys.address);
let buyAddr2 = await trans.buyAndSell();
assert.equal(buyAddr2, buys.address, "买卖合约地址设置正确");
await trans.setBuyAndSell(rents.address,{from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "error message must contain revert");
});
let rentAddr = await trans.rentAndLease();
assert.equal(rentAddr, 0, "租赁合约地址尚未设置");
await trans.setRentAndLease(rents.address);
let rentAddr2 = await trans.rentAndLease();
assert.equal(rentAddr2, rents.address, "租赁合约地址设置正确");
})
it("只有ceo可以设置交易费用", async () => {
// 不是ceo不能设置交易价格
await trans.setBuyFees(1,{from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "error message must contain revert");
})
let buyfees = await trans.getBuyFees();
assert.equal(buyfees, 0, "买卖比例为初始值");
// ceo 可以设置交易价格
await trans.setBuyFees(100);
let buyfees2 = await trans.getBuyFees();
assert.equal(buyfees2, 100, "买卖比例为100");
// 不是ceo,不能设置租赁价格
await trans.setLeaseFees(1,{from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "error message must contain revert");
})
let rentfees = await trans.getLeaseFees();
assert.equal(rentfees, 0, "租赁比例为0");
// ceo,可以设置租赁价格
await trans.setLeaseFees(100);
let rentfees2 = await trans.getLeaseFees();
assert.equal(rentfees2, 100, "租赁比例为100");
})
it("出售书籍", async () => {
// 不是所有者,不允许出售书籍
await book.approve(trans.address,0, {from:accounts[0]});
await trans.sell(book.address, 0, 100, {from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "非所有者不能出售图书");
})
let owner = await book.ownerOf(0);
assert(owner, accounts[0], "图书所有者不变");
// 图书所有者,允许出售图书
await trans.sell(book.address, 0, 100, {from:accounts[0]}); // 出售图书
let owner3 = await book.ownerOf(0);
assert.equal(owner3, trans.address, "图书所有权转移到合约");
await book.transfer(accounts[0], 0).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "出售书籍后,原所有者无法转移图书所有权");
})
let owner2 = await book.ownerOf(0);
assert.equal(owner2, trans.address, "图书所有权不变");
let allowRead = await book.allowToRead(accounts[0], 0);
assert.equal(allowRead, false, "出售书籍后无法阅读图书");
})
it("修改出售价格", async () => {
// 不是所有者,不允许修改价格
await trans.setSellPrice(book.address, 0, 1, {from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "不是所有者不允许修改价格");
})
let sellinfo = await trans.getSellInfo(book.address, 0);
assert.equal(sellinfo[0], accounts[0], "售卖人不变");
assert.equal(sellinfo[1].toNumber(), 100, "售卖价格不变");
// 图书所有者,允许修改价格
await trans.setSellPrice(book.address, 0 , 1000, {from:accounts[0]});
let info = await trans.getSellInfo(book.address, 0);
assert.equal(info[0], accounts[0], "售卖人不变");
assert.equal(info[1].toNumber(), 1000, "售卖价格变为1000");
let owner = await book.ownerOf(0);
assert.equal(owner, trans.address, "图书所有权在合约");
})
it("取消出售", async () => {
// 不是所有者,不允许出售
await trans.cancelSell(book.address, 0, {from:accounts[1]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "不是所有者不允许取消出售");
})
let addr = await book.ownerOf(0);
assert.equal(addr, trans.address, "图书所有权不变");
// 图书所有者,出售成功
await trans.cancelSell(book.address,0);
let addr2 = await book.ownerOf(0);
assert.equal(addr2, accounts[0], "图书所有者变为accounts[0]");
let approveAddr = await book.getApproved(0);
assert.equal(approveAddr, 0, "图书授权取消");
let allowRead = await book.allowToRead(accounts[0], 0);
assert.equal(allowRead, true, "accounts[0]允许阅读图书");
})
it("购买图书", async () => {
// 重新出售图书
await book.approve(trans.address, 0);
await trans.sell(book.address, 0, 999);
// 金额不够,购买失败
await trans.buy(book.address, 0, {from:accounts[1], value: 10}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "金额不够,购买失败");
})
let owner = await book.ownerOf(0);
assert.equal(owner, trans.address, "图书所有权不变");
// 金额足够,购买成功
await trans.buy(book.address, 0, {from:accounts[2], value: 1000});
let owner2 = await book.ownerOf(0);
assert.equal(owner2, accounts[2], "accounts[2]拥有图书0");
let allowRead = await book.allowToRead(accounts[2], 0);
assert.equal(allowRead, true, "accounts[2]允许阅读图书");
})
it("出租图书", async () => {
await book.approve(trans.address, 0, {from:accounts[2]});
// 不是所有者,不能出租图书
await trans.rent(book.address, 0 , 10, 3600, {from:accounts[0]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "不是所有者不允许出租");
})
// 所有者允许出租图书
await trans.rent(book.address, 0, 10000, 3600, {from:accounts[2]});
let owner = await book.ownerOf(0);
assert.equal(owner, accounts[2], "accounts[2]拥有图书所有权");
let allowRead = await book.allowToRead(accounts[2], 0 );
assert.equal(allowRead, true, "accounts[2]允许阅读图书");
})
it("修改出售信息", async () => {
// 不是所有者,不允许修改出售信息
await trans.setRentInfo(book.address, 0, 1, 7200, {from: accounts[0]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "不是所有者修改出售信息");
})
let info = await trans.getRentInfo(book.address, 0);
assert.equal(info[0], accounts[2], "所有者不变");
assert.equal(info[1].toNumber(), 10000, "价格不变");
assert.equal(info[3].toNumber(), 3600, "出售时间不变");
// 图书所有者允许修改出售信息
await trans.setRentInfo(book.address, 0, 20000, 7200, {from:accounts[2]});
let res = await trans.getRentInfo(book.address, 0);
assert.equal(res[0], accounts[2], "所有者不变");
assert.equal(res[1].toNumber(), 20000, "价格变为20000");
assert.equal(res[3].toNumber(), 7200, "出售时间变为7200");
let owner = await book.ownerOf(0);
assert.equal(owner, accounts[2], "图书所有者还是accounts[2]");
})
it("取消出租", async () => {
// 不是所有者,不允许取消出租
await trans.cancelRent(book.address, 0, {from:accounts[0]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "不是所有者不允许取消");
})
let res = await book.isRent(0);
assert.equal(res, true, "图书正在出租");
// 图书所有者运行取消出租
await trans.cancelRent(book.address, 0, {from:accounts[2]});
let res2 = await book.isRent(0);
assert.equal(res2, false, "图书已经取消出租");
let owner = await book.ownerOf(0);
assert.equal(owner, accounts[2], "图书所有者不变");
let allowRead = await book.allowToRead(accounts[2], 0);
assert.equal(allowRead, true, "账户2允许阅读图书");
})
it("租赁图书", async () => {
// 重新出售图书
await book.approve(trans.address, 0, {from:accounts[2]});
await trans.rent(book.address, 0, web3.toWei(1,'ether'), 2, {from:accounts[2]});
// 金额不够,租赁失败
await trans.lease(book.address, 0, {from:accounts[1], value:1000}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "金额不够,租赁失败");
})
let allowRead = await book.allowToRead(accounts[1], 0);
assert.equal(allowRead, false, "不允许阅读");
let leaser = await book.tokenIdToLeaser(0);
assert.equal(leaser, 0, "无人租赁");
// 金钱足够,租赁成功
await trans.lease(book.address, 0, {from:accounts[1], value:web3.toWei(3,'ether')});
let allowRead2 = await book.allowToRead(accounts[1],0);
assert.equal(allowRead2, true, "允许账户1阅读");
let allowRead3 = await book.allowToRead(accounts[2],0);
assert.equal(allowRead3, false, "不允许账户2阅读");
let owner = await book.ownerOf(0);
assert.equal(owner, accounts[2], "图书所有权不转移");
let leaser2 = await book.tokenIdToLeaser(0);
assert.equal(leaser2, accounts[1], "账户1为租赁人");
let islease = await book.isLease(0);
assert.equal(islease, true, "书籍正在租赁");
await trans.cancelRent(book.address, 0, {from:accounts[2]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "租赁时不允许取消租赁");
})
await book.rentCancel(0,{from:accounts[2]}).then(assert.fail).catch(function(error) {
assert(error.message.indexOf('revert') >= 0, "租赁时不允许取消租赁");
})
})
it("提款成功", async () => {
let balance02 = web3.fromWei(web3.eth.getBalance(accounts[2]).toNumber(),'ether');
await trans.withdraw({from:accounts[2]});
let balance12 = web3.fromWei(web3.eth.getBalance(accounts[2]).toNumber(),'ether');
assert.equal(balance12 > balance02 + web3.toWei(0.5,'ether'), true, "转账成功");
let balance01 = web3.fromWei(web3.eth.getBalance(accounts[1]).toNumber(),'ether');
await trans.withdraw({from:accounts[1]});
let balance11 = web3.fromWei(web3.eth.getBalance(accounts[1]).toNumber(),'ether');
assert.equal(balance11 > balance01 + web3.toWei(0.5,'ether'), true, "转账成功");
})
})
<file_sep>/server/utils/logger.js
var log4js = require('log4js');
var logger = log4js.getLogger();
logger.level = 'debug';
//debug
EVENT.ENTER_FUNCTION = "enter function";
log4js.configure({
appenders: [
{
type: 'console',
layout: {
type: 'pattern',
pattern: "[%d{ISO8601}][%p][%x{pid}][%c] >>>%m",
tokens: {
"pid": function () {
return process.pid;
}
}
}
},
{
type: 'file',
filename: config.log.file,//文件的配置路径
//日志文件大小10m
maxLogSize: config.log.maxLogSize,
backups: config.log.backups,
layout: {
type: 'pattern',
pattern: "[%d{ISO8601}][%p][%x{pid}][%c] >>>%m",
tokens: {
"pid": function () {
return process.pid;
}
}
}
}
],
replaceConsole: true
});
function Logger(filepath) {
var category;
if (filepath == undefined || filepath.replace == undefined) {
category = 'common';
} else {
category = filepath.replace(/.*([\/\\]server)+/, "");
}
this._logger = log4js.getLogger(category);
this._logger.setLevel(config.log.level);
}
Logger.prototype.debug = function (eventName, msg, json) {
var _self = this;
_self._logger.debug(splice(eventName, msg, json));
}
Logger.prototype.warn = function (eventName, msg, json) {
var _self = this;
_self._logger.warn(splice(eventName, msg, json));
}
Logger.prototype.info = function (eventName, msg, json) {
var _self = this;
_self._logger.info(splice(eventName, msg, json));
}
Logger.prototype.error = function (eventName, msg, json) {
var _self = this;
_self._logger.error(splice(eventName, msg, json));
}
Logger.prototype.isDebugEnabled = function () {
var _self = this;
return _self._logger.isDebugEnabled();
}
Logger.prototype.isInfoEnabled = function () {
var _self = this;
return _self._logger.isInfoEnabled();
}
Logger.prototype.isWarnEnabled = function () {
var _self = this;
return _self._logger.isWarnEnabled();
}
function splice(eventName, msg, json) {
var c = "[" + eventName + "]";
if (json) {
//TODO:判断json的类型
c += " json=" + JSON.stringify(json);
}
if (msg) {
c += " msg=" + msg;
}
return c;
}
module.exports = exports = function (filepath) {
return new Logger(filepath);
};
exports.EVENT = EVENT;
<file_sep>/front/src/vuex/actions/base.js
import { GET_LOGIN_USERNAME } from '../mutation-types';
// initial state
const state = {
username: ''
};
// getters
const getters = {
};
// actions
const actions = {
getUsername({ commit, username }) {
commit(GET_LOGIN_USERNAME, username);
}
};
// mutations
const mutations = {
[GET_LOGIN_USERNAME](state, username) {
state.username = username;
},
};
export default {
namespaced: true,
state,
getters,
actions,
mutations
};
<file_sep>/server/controller/user.js
const log4js = require("log4js");
const logger = log4js.getLogger("controller/user");
const encrypt = require('../utils/encrypt');
const thunder = require("../utils/thunder");
const resourceContract = require("../dao/resourceContract");
const transactionContract = require("../dao/transactionContract");
const userDao = require("../dao/user");
const resourceInfoDao = require("../dao/resourceInfo");
const config = require("../config");
//预出售的状态
const preSellStatus = 1;
const preRentStatus = 1;
var userCount = 0;
/**
* 登录
* @param req
* @param res
* @param next
*/
exports.login = function(req, res, next) {
res.send({status:1,msg:"login success"});
}
/**
* 登出操作
* @param req
* @param res
* @param next
*/
exports.logout = async function(req, res, next) {
let user = req.session.passport.user;
if (user) {
logger.info("enter local logout", { "user": user });
try{
await userDao.deleteSessionById(req.session.id);
req.logout();
logger.info("success to logout from local", { "username": user.userName });
res.send({status:1,msg:"local logout success"});
}catch (e) {
logger.error("local logout fail",{ "username": user.userName },e);
res.send({status:0,msg:"local logout failed"});
}
} else {
logger.warn("local logout failed,no user in req.session.passport object ,maybe already expired.", {
"session": req.session.id,
"user": user
});
res.send({status:0,msg:"local logout failed"});
}
};
/**
* // 注册用户,且注册用户的链克口袋 post:方法,用户只需要输入用户名,手机号,邮箱
* //密码在传输的时候使用MD5加密,在存入进数据库的时候再次经过随机数加MD5加密保存
* @param req
* @param res
* @param next
*/
exports.register = async function(req, res, next) {
logger.info("register user");
let random = encrypt.getRandom();
//TODO 临时加的控制demo的注册
if(userCount >=9){
logger.error("user account limit",userCount);
res.send({status:0,msg:"注册失败"});
return;
}
let user = {
userName:req.body.userName,
pwd:encrypt.getMD5(req.body.pwd,random),
email:req.body.email,
mobile:req.body.mobile,
randomNum:random,
account:config.server.userAccount[userCount++] //TODO 对接钱包注册地址,获得真实的地址。
}
//TODO 校验注册的基本内容
try{
//1、先向迅雷注册账号 TODO 测试
/*let registerData = await thunder.register(user.email);
user.account = registerData.service_id;*/
//2、保存账号信息至数据库
await userDao.add(user);
res.send({status:1,msg:"恭喜注册成功"});
} catch (e) {
logger.error("register user fail.",user);
res.send({status:0,msg:"注册失败"});
}
};
/**
* 获取当前登录的用户信息
* @param req
* @param res
* @param next
*/
exports.getCurrentUserInfo = function(req, res, next) {
var user = req.session.passport.user;
res.send({ "success": true, "data": user });
};
exports.getUserInfoByAccount = async function(req,res,next){
let account = req.param("account");
if(account == undefined || account == "" || account == null){
res.send({status:0,msg:"account 必传"});
return;
}
try{
let userInfo = await userDao.findUserInfoByAccount(account);
res.send({status:1,msg:"success",data:userInfo});
}catch (e) {
logger.error("getUserInfoByAccount fail",{account:account},e);
res.send({status:0,msg:"getUserInfoByAccount fail"});
}
}
/**
* 获取当前用户已购买的资源列表
* @param req
* @param res
* @param next
*/
exports.getPurchasedResourcesByUser = async function(req, res, next) {
logger.info("get purchased file");
//1、先拿到当前用户信息,判断用户是否是登录状态
var user = req.session.passport.user;
//2、从数据库中获取当前已购买的资源列表
let list = await userDao.getPurchasedResourcesByUserId(user._id);
res.send({status:1,msg:"success",data:list});
};
/**
* 获取当前用户已租赁的资源列表
* @param req
* @param res
* @param next
*/
exports.getRentResourcesByUser = async function(req, res, next) {
logger.info("get rent file");
//1、先拿到当前用户信息,判断用户是否是登录状态
var user = req.session.passport.user;
//2、从数据库中获取当前已购买的资源列表
let list = await userDao.getRentResourcesByUserId(user._id);
res.send({status:1,msg:"success",data:list});
};
/**
* 获取当前用户提交的版权记录
* @param req
* @param res
* @param next
*/
exports.getCopyRightsByUser = async function(req, res, next) {
logger.info("get rent file");
//1、先拿到当前用户信息,判断用户是否是登录状态
var user = req.session.passport.user;
//2、从数据库中获取当前版权的资源列表
let list = await userDao.getCopyrightsByUserId(user._id);
res.send({status:1,msg:"success",data:list});
};
/**
* 出售一个资源
* @param req
* @param res
* @param next
*/
exports.sell = async function(req, res, next) {
let tokenId = req.body.tokenId;
let resourceId = req.body.resourceId;
let sellPrice = req.body.sellPrice;
let user = req.session.passport.user;
logger.info("selling file.", {
tokenId: tokenId,
resourceId:resourceId
});
try {
//todo 校验
//1、先查看这个资源个体的交易信息
let purchasedResourceDoc = await userDao.findOneUserPurchasedResourceByResourceIdAndTokenId(user._id,resourceId,tokenId);
let sellResource = purchasedResourceDoc.purchasedResources[0];
//校验这个资源是否可以售卖
if(sellResource.sellStatus>0){
logger.warn("sell resource fail.",{
user:user,
sellResource:sellResource
});
res.send({ status: 0, msg: "资源出售失败,该资源可能已出售、或正在出售中" });
return;
}
//如果这资源已出租也不能售卖
if(sellResource.rentOutStatus>1){
logger.warn("rent out resource fail.",{
user:user,
sellResource:sellResource
});
res.send({ status: 0, msg: "资源出售失败,该资源已出租成功,请出租时间结束后再出售" });
return;
}
let resourceInfo = await resourceInfoDao.findById(resourceId);
//2.1先判断拿到tokenId和合约地址,是否属于当前用个人账户,并且判断是否属于二手交易的首次交易,如果是则创建交易合约,如果不是就获取交易合约地址合约,将交易合约处于挂起状态,让资源处于售卖状态
resourceContract.approve(resourceInfo.resourceAddress,user.account,transactionContract.address(),tokenId);
//2.2 交易hash
let transactionAddressTmp = transactionContract.sell(user.account,resourceInfo.resourceAddress,tokenId,sellPrice);
//3、合约创建部署成功触发事件,更新登记交易合约的地址,还有售卖状态至1
let transactionAddress = transactionAddressTmp == null?sellResource.transactionAddress:transactionAddressTmp;
var sellResourceObj = {
tokenId:tokenId,
ownerAccount:user.account,
ownerId:user._id,
sellPrice:sellPrice,
transactionAddress:transactionAddress
}
await resourceInfoDao.addSellResourceById(resourceId,sellResourceObj)
let updateObj = await userDao.modifySellStatusAndTransactionAddress(user._id, tokenId, preSellStatus,transactionAddress,sellPrice);
if (updateObj !== undefined ) {
res.send({ status: 1, msg: "the resource sell success." });
} else {
res.send({ status: 0, msg: "资源出售失败,请稍后重试." });
}
} catch (err) {
logger.error("sell resource fail.",{
tokenId: tokenId,
user:user
},err);
res.send({ status: 0, msg: "资源出售失败,请稍后重试" });
}
};
/**
* 出租一个资源
* @param req
* @param res
* @param next
*/
exports.rentOut = async function(req, res, next) {
let tokenId = req.body.tokenId;
let resourceId = req.body.resourceId;
let rentPrice = req.body.rentPrice;
let rentTime = req.body.rentTime;
let user = req.session.passport.user;
logger.info("rentOut file.", {
tokenId: tokenId,
resourceId:resourceId,
rentPrice:rentPrice,
rentTime:rentTime,
user:user
});
try {
//todo 校验
//1、先查看这个资源个体的交易信息
let purchasedResourceDoc = await userDao.findOneUserPurchasedResourceByResourceIdAndTokenId(user._id,resourceId,tokenId);
let rentOutResource = purchasedResourceDoc.purchasedResources[0];
//校验这个资源是否可以售卖
if(rentOutResource.sellStatus>0){
logger.warn("sell resource fail.",{
user:user,
rentOutResource:rentOutResource
});
res.send({ status: 0, msg: "资源出租失败,该资源可能已出售、或正在出售中" });
return;
}
//如果这资源已出租也不能售卖
if(rentOutResource.rentOutStatus>0){
logger.warn("rent out resource fail.",{
user:user,
rentOutResource:rentOutResource
});
res.send({ status: 0, msg: "资源出租失败,该资源已出租成功,请出租时间结束后再出售" });
return;
}
let resourceInfo = await resourceInfoDao.findById(resourceId);
//2、先判断拿到tokenId和合约地址,是否属于当前用个人账户,并且判断是否属于二手交易的首次交易,如果是则创建交易合约,如果不是就获取交易合约地址合约,将交易合约处于挂起状态,让资源处于售卖状态
resourceContract.approve(resourceInfo.resourceAddress,user.account,transactionContract.address(),tokenId);
let transactionAddressTmp = transactionContract.rent(user.account,resourceInfo.resourceAddress,tokenId,rentPrice,rentTime);
//let transactionAddressTmp = transactionContract.rentOut(resourceInfo.resourceAddress,tokenId,rentOutResource.transactionAddress);
let transactionAddress = transactionAddressTmp == null?rentOutResource.transactionAddress:transactionAddressTmp;
//3、合约创建部署成功触发事件,更新登记交易合约的地址,还有售卖状态至1
var rentOutResourceObj = {
tokenId:tokenId,
ownerId:user._id,
ownerAccount:user.account,
rentPrice:rentPrice,
rentTime:rentTime,
transactionAddress:transactionAddress
}
await resourceInfoDao.addRentOutResourceById(resourceId,rentOutResourceObj)
let updateObj = await userDao.modifyRentStatusAndTransactionAddress(user._id, tokenId, preRentStatus,rentPrice,transactionAddress);
if (updateObj !== undefined ) {
res.send({ status: 1, msg: "the resource rent out success." });
} else {
res.send({ status: 0, msg: "资源出租失败,请稍后重试" });
}
} catch (err) {
logger.error("sell resource fail.",{
tokenId: tokenId,
user:user
},err);
res.send({ status: 0, msg: "资源出租失败,请稍后重试" });
}
}
<file_sep>/server/route.js
var express = require('express');
var router = express.Router();
var user = require('./controller/user');
var resourceCopyright = require('./controller/resourceCopyright');
var resourceInfo = require('./controller/resourceInfo');
var helloWorld = require('./controller/helloWorld');
var config = require('./config');
var passport = require("passport");
var auth = require("./controller/auth");
var fs = require("fs");
var URL = "/" + config.appName;
//和合约交互的例子
router.get(URL+'/helloWorld',auth.isAuthenticated,helloWorld.sayHello);
//用户登录、注册、及获取当前用户
router.post(URL+"/user/register", user.register);
router.post(URL+"/user/localLogin",passport.authenticate('local',{
session:true,
failureFlash: true
}),user.login);
router.post(URL+"/user/localLogout",auth.isAuthenticated,user.logout);
router.get(URL+"/user/renderUser",auth.isAuthenticated, user.getCurrentUserInfo);
router.get(URL+"/user/getUserInfoByAccount",auth.isAuthenticated, user.getUserInfoByAccount);
//用户个人管理
router.get(URL+"/user/getCopyRightsByUser",auth.isAuthenticated,user.getCopyRightsByUser);
router.get(URL+"/user/getPurchasedResourcesByUser",auth.isAuthenticated,user.getPurchasedResourcesByUser);
router.get(URL+"/user/getRentResourcesByUser",auth.isAuthenticated,user.getRentResourcesByUser);
router.post(URL+"/user/purchasedResources/sell",auth.isAuthenticated,user.sell);
router.post(URL+"/user/purchasedResource/rentOut",auth.isAuthenticated,user.rentOut);
//版权
router.post(URL+"/copyright/apply",auth.isAuthenticated,resourceCopyright.applyCopyright);
router.post(URL+"/copyright/audit",auth.isAuthenticated,resourceCopyright.auditCopyright);
router.post(URL+"/copyright/upload/sample",resourceCopyright.uploadSample);
router.get(URL+"/copyright/getResourceCopyrightDetailById",resourceCopyright.getResourceCopyrightDetailById);
//资源信息
router.post(URL+"/resource/publish",auth.isAuthenticated,resourceInfo.publishResource);
router.post(URL+"/resource/upload/coverImg",resourceInfo.uploadCoverImg);
//测试上传用
router.get(URL+'/form',function(req, res, next) {
fs.readFile('dist/testUploadImage.html', {encoding: 'utf8'},function(err,data) {
res.send(data);
});
});
router.get(URL+"/resource/getResourceDetailById",resourceInfo.getResourceDetailById);
router.post(URL+"/resource/getResourceListByPage",resourceInfo.getResourceListByPage);
router.post(URL+"/resource/getPurchasedResourceListByPage",resourceInfo.getPurchasedResourceListByPage);
router.post(URL+"/resource/getTenantableResourceListByPage",resourceInfo.getTenantableResourceListByPage);
router.get(URL+"/resource/getPurchasedResourceOwnerListById",resourceInfo.getPurchasedResourceOwnerListById);
router.get(URL+"/resource/getTenantableResourceOwnerListById",resourceInfo.getTenantableResourceOwnerListById);
router.post(URL+"/resource/buyFromAuthor",auth.isAuthenticated,resourceInfo.buyFromAuthor);
router.post(URL+"/resource/buy",auth.isAuthenticated,resourceInfo.buy);
router.post(URL+"/resource/rent",auth.isAuthenticated,resourceInfo.rent);
module.exports = router;
<file_sep>/front/src/config/user/data.js
import { ListType } from 'src/config/user/enum';
export const getTableHeader = (listType) => {
switch (listType) {
case ListType.CopyRight: {
return [
{ field: 'copyrightId', name: '版权ID', width: 120 },
{ field: 'workName', name: '版权名称', width: 120 },
{ field: 'copyrightAddress', name: '版权合约地址', width: 120 },
{ field: 'resourceAddress', name: '资源合约地址', width: 120 },
{ field: 'resourcesIpfsHash', name: '资源ipfsHash', hidden: true, width: 120 },
{ field: 'resourcesIpfsDHash', name: '资源ipfsDHash', hidden: true, width: 120 },
{ field: 'localUrl', name: '本地URL', hidden: true, width: 120 }
];
}
case ListType.Purchase: {
return [
{ field: 'resourceId', name: '资源ID', width: 120 },
{ field: 'resourceName', name: '名称', width: 120 },
{ field: 'type', name: '类型', width: 40 },
{ field: 'tokenId', name: 'tokenId', width: 120, hidden: true },
{ field: '__sellStatus', name: '出售状态', width: 60 },
{ field: 'sellPrice', name: '出售价格(元)', width: 60 },
{ field: '__rentOutStatus', name: '租赁状态', width: 60 },
{ field: 'rentPrice', name: '租赁价格(元)', width: 60 }
];
}
case ListType.Rent: {
return [
{ field: 'resourceId', name: '资源ID', width: 120 },
{ field: 'resourceName', name: '名称', width: 120 },
{ field: 'type', name: '类型', width: 40 },
{ field: 'tokenId', name: 'tokenId', width: 120 },
{ field: 'rentTime', name: '出售时间(天)', width: 80 }
];
}
default:
return [];
}
};
export const getCopyRightApplyInitData = () => ({
workName: '',
workCategory: '',
localUrl: [],
authors: [{
authorName: '',
identityType: '身份证',
identityNum: ''
}],
workProperty: '',
rights: [],
belong: ''
});
export const getCopyRightPublishInitData = () => ({
copyrightId: '',
resourceName: '',
total: '',
coverImage: '',
desc: '',
price: ''
});
<file_sep>/server/config.js
/**
*
* App配置文件读取
*/
function getRootPath() {
var breakIndex = __dirname.lastIndexOf('server');
if (breakIndex > 0) {
return __dirname.substring(0, breakIndex - 1);
}
return __dirname;
}
//应用主目录
const path = getRootPath();
//配置文件,优先从环境变量读取
const configPath = process.env.DR_CONFIG ? process.env.DR_CONFIG : path + '/chainbook_config.json';
const config = require('config.json')(configPath);
config.appName = 'ChainBook';
config.log = {
level: "INFO",
file: "chainbook.log",
maxLogSize: 1024 * 1024 * 10,//日志文件最大为10m
backups: 5
};
/*//合约相关
config.contract = {
url:config.contract.url
};
//迅雷相关配置
config.thunder = {
baseURL:config.thunder.baseURL
};*/
module.exports = config;
<file_sep>/front/src/config/user/converter.js
import { ListType } from 'src/config/user/enum';
export const getApiToRow = (listType, raw) => {
switch (listType) {
case ListType.CopyRight:
return raw.reduce((arr, cur) => arr.concat(cur.copyrights), []);
case ListType.Purchase:
return raw.reduce((arr, cur) => arr.concat(cur.purchasedResources), []).map(book =>
Object.assign(book, {
__sellStatus: book.sellStatus ? '已出售' : '未出售',
__rentOutStatus: book.rentOutStatus ? '已出租' : '未出租'
}));
// return [{
// resourceId: '5b0f597905373eafe9ceed62',
// resourceName: '以太坊白皮书',
// type: 'book',
// tokenId: 'book_contracts_token_id',
// sellStatus: 1,
// sellPrice: '20.00',
// rentOutStatus: 0,
// rentPrice: '2.00'
// }];
case ListType.Rent:
// return [{
// resourceId: '5b0f597905373eafe9ceed62',
// resourceName: '以太坊白皮书',
// type: 'book',
// tokenId: 'book_contracts_token_id',
// rentTime: 30
// }];
return raw.reduce((arr, cur) => arr.concat(cur.rentResources), []);
default:
return [];
}
};
export const getApiToRes = () => [];
export const getResToApi = (original) => {
const data = JSON.parse(JSON.stringify(original));
return Object.assign(data, {
authors: data.authors,
rights: data.rights,
localUrl: data.localUrl.map(sample => sample.url).join(',')
});
};
<file_sep>/front/src/router/index.js
import Vue from 'vue';
import Router from 'vue-router';
import Index from 'src/views/Index';
import SecondHand from 'src/views/SecondHand';
import Rent from 'src/views/Rent';
import FirstResource from 'src/views/FirstResource';
import User from 'src/views/User';
Vue.use(Router);
export const routerMap = [
{
path: '/',
name: 'Index',
component: Index,
title: '首页'
},
{
path: '/FirstResource',
name: 'FirstResource',
component: FirstResource,
title: '首发资源'
},
{
path: '/SecondHand',
name: 'SecondHand',
component: SecondHand,
title: '二手市场'
},
{
path: '/Rent',
name: 'Rent',
component: Rent,
title: '租赁市场'
},
{
path: '/User',
name: 'User',
component: User,
title: '个人中心',
visible: false
}
];
const router = new Router({
routes: routerMap
});
router.beforeEach((to, from, next) => {
const matchPath = routerMap.filter(item => item.path === to.path);
if (matchPath) {
document.title = `链书吧-${matchPath[0].title}`;
} else {
window.alert('不存在路径');
}
next();
});
export default router;
<file_sep>/server/model/user.js
var mongoose = require("mongoose");
var userSchema = new mongoose.Schema({
userName: String,
pwd: String,
account:String,
email: String,
mobile: String,
randomNum: String,
copyrights:[new mongoose.Schema({
_id: false,
copyrightId:String,
workName: String,
resourcesIpfsHash:String,
resourcesIpfsDHash:String,
localUrl:String,
copyrightAddress:String,
resourceAddress:String,
resourceId:String,
})],
purchasedResources:[new mongoose.Schema({
_id: false,
resourceId:String,
resourceName: String,
transactionAddress:String,
type:String,
tokenId:String,
sellStatus:Number,
sellPrice:String,
rentOutStatus:Number,
rentPrice:String
})],
rentResources:[new mongoose.Schema({
_id: false,
resourceId:String,
resourceName:String,
transactionAddress:String,
type:String,
tokenId:String,
rentTime:Number
})],
createDate: {type: Date, default: Date.now},
createBy: String,
updateDate: {type: Date, default: Date.now},
updateBy: String
}, {versionKey: false});
var User = mongoose.model("user", userSchema, "user");
module.exports = User;
<file_sep>/front/src/apis/copyright.js
export default function (request) {
return {
// 获取资源版权信息详情
getResourceCopyrightDetailById(id) {
return request({
url: '/copyright/getResourceCopyrightDetailById',
method: 'get',
params: { id }
}).then(data => data);
},
/**
* 申请资源版权信息
* @param {*} data { workName, workCategory, samplePath, authors, workProperty, rights, belong }
*/
apply(data) {
return request({
url: '/copyright/apply',
method: 'post',
data
}).then(data => data);
},
// 上传样本
sample(file) {
return request({
url: '/copyright/upload/sample',
method: 'post',
data: file
}).then(data => data);
},
// 手动审核
audit(copyrightId) {
return request({
url: '/copyright/audit',
method: 'post',
data: { copyrightId }
}).then(data => data);
}
// // 发行审核通过的版权信息
// publish(data) {
// return request({
// url: '/resource/publish',
// method: 'post',
// data
// }).then(data => data);
// }
};
}
<file_sep>/server/dao/resourceContract.js
const fs = require('fs');
const config = require('../config');
const log4js = require('log4js');
const logger = log4js.getLogger('dao/resourceContract');
const copyrightDao = require('./copyrightContract');
const Web3 = require('web3');
const solc = require('solc');
let web3;
if (typeof web3 !== 'undefined') {
web3 = new Web3(web3.currentProvider);
} else {
// set the provider you want from Web3.providers
web3 = new Web3(new Web3.providers.HttpProvider(config.contract.url));
}
const BookOwnerShip_SOURCE = config.server.bookOwnerShip_source;
const basePath = config.server.contract_path;
const chainbookAddress = config.server.address;
const chainbookGas = config.server.gas;
const limit = config.server.bookOwnership_limit;
const ResourceContractDao = class dao {
constructor(){
this._compile();
}
//编译合约
_compile(){
let output;
try{
let bookOwnerShip= fs.readFileSync(BookOwnerShip_SOURCE,'utf-8');
var input = {
'BookOwnerShip.sol': bookOwnerShip
}
function findImports (path) {
let dependentContract = fs.readFileSync(basePath+path,'utf-8');
if (dependentContract !== undefined && dependentContract !== null)
return { contents: dependentContract }
else
return { error: 'File not found' }
}
output = solc.compile({ sources: input },1,findImports)
for (var contractName in output.contracts){
logger.info(contractName + ': ' + output.contracts[contractName]);
if(contractName.includes("BookOwnerShip.sol")){
dao.bookOwnerShipCompiled = output.contracts[contractName];
dao.bookOwnerShipContactWeb3 = web3.eth.contract(JSON.parse(dao.bookOwnerShipCompiled.interface));
}
}
}catch (e) {
logger.error("compile contract fail",e);
//如果捕捉到异常就直接抛出去,就可以启动时就知道合约编译异常
throw e;
}
//如果编译后依然为空的话,说明合约有问题,直接启动报错
if(dao.bookOwnerShipCompiled == undefined || dao.bookOwnerShipContactWeb3 == undefined){
logger.error("compile BookOwnerShip.sol contract fail",output);
throw new Error("compile BookOwnerShip.sol error");
}
}
/**
* TODO 私有的方法
* 部署合约,这个合约由平台部署,所属地址是平台的
* @param _deployContract
* @returns {Promise<any>}
*/
_deployContract(userObj,resourceObj) {
//TODO 判空
return new Promise((resolve, reject) => {
//获取合约的代码,部署时传递的就是合约编译后的二进制码,
let deployCode = dao.bookOwnerShipCompiled.bytecode;
dao.bookOwnerShipContactWeb3.new(copyrightDao.getCopyrightContractAddress(),parseInt(resourceObj.copyrightAddress),parseInt(resourceObj.price),resourceObj.total,limit,{
data: deployCode,
from: userObj.account,
gas:6000000
}, function(err, contract) {
if (!err) {
// 注意:这个回调会触发两次,一次是合约的交易哈希属性完成,另一次是在某个地址上完成部署
// 通过判断是否有地址,来确认是第一次调用,还是第二次调用。
if (!contract.address) {
logger.info("contract deploy transaction hash: " + contract.transactionHash) //部署合约的交易哈希值
} else {
// 合约发布成功后,才能调用后续的方法
logger.info("contract deploy address: " + contract.address) // 合约的部署地址
resolve(contract);
}
}else {
reject(err);
}
});
});
}
async publishResource(userObj,resourceInfoObj){
//使用transaction方式调用,写入到区块链上,sendTransaction 方式调用
logger.info("enter resources contract",{
userObj:userObj,
resourceInfoObj:resourceInfoObj
});
let contract = await this._deployContract(userObj,resourceInfoObj);
//返回一个资源合约地址
return contract.address;
}
buyFromAuthor(resourceAddress,userObj,price){
return new Promise((resolve, reject) => {
if(resourceAddress == undefined){
reject(new Error("resourceAddress not null"));
}
//已有bookOwnership地址先获取实例
let contractInstance = dao.bookOwnerShipContactWeb3.at(resourceAddress);
var sendTransactionId = contractInstance.buyFromAuthor.sendTransaction({from:userObj.account, value:price,gas:chainbookGas});
//设置注册成功的监听事件
let buyFromAuthorEvent = contractInstance.CreatBook();
// 监听事件,监听到事件后会执行回调函数
buyFromAuthorEvent.watch(function(err, result) {
if (err) {
reject(e);
}else{
if(sendTransactionId === result.transactionHash){
logger.info("_BuyFromAuthor event",{
result:result,
_tokenId:result.args._tokenId.toString()
});
buyFromAuthorEvent.stopWatching();
resolve(result.args._tokenId.toString());
}
}
});
});
}
// 允许第三方平台交易图书
approve(resourceAddress,sender, _to, _tokenId) {
let contractInstance = dao.bookOwnerShipContactWeb3.at(resourceAddress);
contractInstance.approve.sendTransaction(_to, _tokenId, {from:sender});
}
}
module.exports = new ResourceContractDao();
<file_sep>/front/src/config/user/enum.js
export const ListType = {
CopyRight: 'CopyRight', // 登记版权
Purchase: 'Purchase', // 已购买
Rent: 'Rent' // 已租赁
};
export const Operation = {
Create: 1, // 新增版权登记
Update: 2, // 更新版权登记
Remove: 3, // 删除版权登记
Publish: 4, // 发行资源
Preview: 5, // 预览发行资源
Sell: 6, // 出售资源
Rent: 7, // 租赁资源
};
<file_sep>/server/app.js
var createError = require('http-errors');
var express = require('express');
var fs = require('fs');
var path = require('path');
var cookieParser = require('cookie-parser');
var log4js = require('log4js');
var mainRouter = require('./route');
var config = require('./config');
var mongoose = require("mongoose");
var passport = require('passport');
var LocalStrategy = require("passport-local").Strategy;
//持久化session
var session = require("express-session");
var MongoStore = require('connect-mongo')(session);
var flash = require('connect-flash');
var userDao = require("./dao/user");
const resolve = file=>path.resolve(__dirname, file);
//日志配置,TODO 挪开
log4js.configure({
appenders: {
out: { type: 'console' },
chainbook: { type: 'dateFile', filename: 'chainbook.log', alwaysIncludePattern:true },
},
categories: {
default: { appenders: ['out','chainbook'], level: 'INFO'}
}
});
var logger = log4js.getLogger();
var app = express();
app.use(express.json());
app.use(express.urlencoded({ extended: false }));
app.use(cookieParser());
app.use(session({
secret: 'chainbook',
resave: false,
saveUninitialized: true,
cookie: { secure: false },
store: new MongoStore({ mongooseConnection: mongoose.connection }) //session存储位置
}));
app.use(passport.initialize());
app.use(passport.session());
app.use(flash());
passport.use(new LocalStrategy({
usernameField: 'userName',
passwordField: 'pwd'
},function(userName,password,done) {
//校验用户名密码是否正确
userDao.verifyUser(userName,password).then(function(result){
if(result){
logger.info("passport verify success:",{
userName: userName
});
done(null,result);
}else{
logger.warn("passport verify fail.",{
userName: userName,
password: <PASSWORD>
});
done(null,false,{status:0,msg:"Invalid username or password"});
}
}).catch(err =>{
logger.error("passport verify fail.",{
userName: userName,
password: <PASSWORD>
},err);
done(err,null);
});
}));
passport.serializeUser(function (user,done){
done(null,user);
});
passport.deserializeUser(function (id, done) {
userDao.findUserInfoById(id).then(user => {
done(null, user);
}).catch( err =>{
logger.error("passport deserialize user fail",err);
done(null, false);
});
});
// 访问静态资源
app.use(express.static(path.join(__dirname, '../dist')));
// 路由的配置需要放置在前面,且使用根目录路由
app.use('/', mainRouter);
app.get("/" + config.appName+'/*', function (req, res, next) {
if(req.originalUrl.indexOf('/user') ==0 && req.originalUrl.indexOf('/copyright') == 0 && req.originalUrl.indexOf('/resource') ==0) {
const html = fs.readFileSync(resolve('../dist/index.html'), 'utf-8');
res.send(html);
}else{
next();
}
});
// catch 404 and forward to error handler
app.use(function(req, res, next) {
next(createError(404));
});
// error handler
app.use(function(err, req, res, next) {
// set locals, only providing error in development
res.locals.message = err.message;
res.locals.error = req.app.get('env') === 'development' ? err : {};
logger.error(err);
// render the error page
res.status(err.status || 500);
res.send(err);
});
//mongoose config
var dbOption = {
db: {
native_parser: true
},
server: {poolSize: config.db.poolSize},
user: config.db.userName,
pass: config.db.password
}
mongoose.connect(config.db.uri, dbOption);
mongoose.connection.on('connected', function () {
logger.info("DB connected", "success to connect DB", {"url:": config.db.uri});
});
mongoose.connection.on('error', function (err) {
logger.error("DB connected error", 'DB connected failed', {"url": config.db.uri, "err:": err});
});
mongoose.connection.on('disconnected', function () {
logger.info('DB disconnected', "success to close DB connection", {"url": config.db.uri});
});
process.on('SIGINT', function () {
mongoose.connection.close(function () {
logger.info('DB disconnected', "disconnected while app stopped", {"url": config.db.uri});
process.exit(0);
});
});
module.exports = app;
<file_sep>/server/dao/resourceCopyright.js
var util = require("util");
var ResourceCopyright = require("../model/resourceCopyright");
var SuperDao = require("./super");
var ObjectUtil = require("../utils/objectUtils");
var log4js = require('log4js');
var logger = log4js.getLogger('dao/user');
var ResourceCopyrightDao = function () {
SuperDao.call(this);
this.model = ResourceCopyright;
};
util.inherits(ResourceCopyrightDao, SuperDao);
/**
* 查找版权的审核和发行状态
* @param id
* @returns {*}
*/
ResourceCopyrightDao.prototype.findStatusById = function(id) {
try{
ObjectUtil.notNullAssert(id);
return new Promise((resolve, reject) => {
ResourceCopyright.find({_id:id},{auditStatus:1,publishStatus:1},function(err,obj){
if(err){
reject(err);
}else{
resolve(obj);
}
});
});
}catch(err){
logger.error("findStatusById error.id:{}",id,err)
return new Promise((reject) =>{
reject(err);
});
}
}
/**
* 修改版权的审核状态
* @param id
* @param auditStatus 0:未审核,1:审核成功,-1:审核失败
* @returns {*}
*/
ResourceCopyrightDao.prototype.modifyAuditStatus = function(id,auditStatus){
try{
ObjectUtil.notNullAssert(id);
ObjectUtil.notNullAssert(auditStatus);
return new Promise((resolve, reject) => {
ResourceCopyright.update({'_id':id},
{$set:{ 'auditStatus': auditStatus }},function(err,updateObj){
if(err){
reject(err);
}else{
resolve(updateObj);
}
}
);
});
}catch(err){
logger.error("findStatusById error.id:{}",id,err)
return new Promise((reject) =>{
reject(err);
});
}
}
/**
* 改版权的发行状态
* @param id
* @param publishStatus 0:未发行,1:已发行
* @returns {*}
*/
ResourceCopyrightDao.prototype.modifyPublishStatus = function(id,publishStatus,resourceAddress,resourceId){
try{
ObjectUtil.notNullAssert(id);
ObjectUtil.notNullAssert(publishStatus);
ObjectUtil.notNullAssert(resourceAddress);
return new Promise((resolve, reject) => {
ResourceCopyright.update({'_id':id},
{$set:{ 'publishStatus': publishStatus,'resourceAddress':resourceAddress,'resourceId':resourceId }},function(err,updateObj){
if(err){
reject(err);
}else{
resolve(updateObj);
}
}
);
});
}catch(err){
logger.error("modifyPublishStatus error.",{
id:id,
publishStatus:publishStatus
},err)
return new Promise((reject) =>{
reject(err);
});
}
}
ResourceCopyrightDao.prototype.updateResourceCopyrightInfo = function(id, copyrightAddress, resourceHash, resourceDhash, auditStatus){
try{
ObjectUtil.notNullAssert(id);
ObjectUtil.notNullAssert(copyrightAddress);
ObjectUtil.notNullAssert(resourceHash);
ObjectUtil.notNullAssert(resourceDhash);
return new Promise((resolve, reject) => {
ResourceCopyright.update({'_id':id},
{$set:{
'auditStatus': auditStatus,
'copyrightAddress':copyrightAddress,
'resourceHash':resourceHash,
'resourceDhash':resourceDhash
}},function(err,updateObj){
if(err){
reject(err);
}else{
resolve(updateObj);
}
}
);
});
}catch(err){
logger.error("updateResourceCopyrightInfo error",{
id:id,
auditStatus: auditStatus,
copyrightAddress:copyrightAddress,
resourceHash:resourceHash,
resourceDhash:resourceDhash
},err)
return new Promise((reject) =>{
reject(err);
});
}
}
/*初始化*/
module.exports = new ResourceCopyrightDao();
<file_sep>/server/model/session.js
var mongoose = require("mongoose");
var sessionsSchema = new mongoose.Schema({
_id:String,
session:String,
expires:{type: Date}
}, {versionKey: false});
var Sessions = mongoose.model("sessions", sessionsSchema, "sessions");
module.exports = Sessions;
<file_sep>/smartcontracts/migrations/4_BuyAndSell.js
var BuyAndSell = artifacts.require("./BuyAndSell.sol");
module.exports = function(deployer) {
deployer.deploy(BuyAndSell);
};
<file_sep>/server/controller/helloWorld.js
var express = require('express');
var fs = require('fs');
var Web3 = require('web3');
var web3 = new Web3(new Web3.providers.HttpProvider('http://localhost:7545'));
exports.sayHello = function(req, res, next) {
fs.readFile('./smartcontracts/build/contracts/helloWorld.json',function(err,data){
var helloWorldContract = new web3.eth.Contract(JSON.parse(data).abi,'0xc132c1e1a347883c5170a332eb2e61a024f354a8', {
from: '0x1234567890123456789012345678901234567891',
gasPrice: '20000000000'
});
console.log(helloWorldContract);
helloWorldContract.methods.sayHello().call().then(function(str) {
res.send(str+"2018");
});
});
};
<file_sep>/smartcontracts/migrations/3_BookOwnership.js
var BookOwnerShip = artifacts.require("./BookOwnerShip.sol");
var BookCopyrightCreate = artifacts.require("./BookCopyrightCreate.sol");
module.exports = function(deployer) {
var copyrightAddress = BookCopyrightCreate.address;
deployer.deploy(BookOwnerShip,copyrightAddress, 0, 0, 3, 1);
};
<file_sep>/server/dao/transactionContract.js
const fs = require('fs');
const config = require('../config');
const log4js = require('log4js');
const logger = log4js.getLogger('dao/transactionContract');
const Web3 = require('web3');
const solc = require('solc');
let web3;
if (typeof web3 !== 'undefined') {
web3 = new Web3(web3.currentProvider);
} else {
// set the provider you want from Web3.providers
web3 = new Web3(new Web3.providers.HttpProvider(config.contract.url));
}
const chainbookAddress = config.server.address;
const chainbookGas = config.server.gas;
const TransactionContractDao = class dao {
constructor() {
this._init();
}
async _init(){
logger.info("hiiiiii");
//编译buyAndSell
let buyAndSell = fs.readFileSync(config.server.buyAndSell_source,'utf-8');
dao.buyAndSellContract = this._compile({ 'BuyAndSell.sol': buyAndSell});
//编译rentAndLease
let rentAndLease = fs.readFileSync(config.server.rentAndLease_source,'utf-8');
dao.rentAndLeaseContract = this._compile({ 'RentAndLease.sol': rentAndLease});
//编译transaction
let transaction = fs.readFileSync(config.server.transaction_source,'utf-8');
dao.transactionContract = this._compile({ 'Transtaction.sol': transaction});
//部署buyAndSell
let buyContract = await this._deployContract(dao.buyAndSellContract);
dao.buyAndSellContract.address = buyContract.address;
dao.buyAndSellContract.contractInstance = dao.buyAndSellContract.contractWeb3.at(buyContract.address);
//部署rentAndLease
let rentContract = await this._deployContract(dao.rentAndLeaseContract);
dao.rentAndLeaseContract.address = rentContract.address;
dao.rentAndLeaseContract.contractInstance = dao.rentAndLeaseContract.contractWeb3.at(rentContract.address);
//部署transaction
let transtractContract = await this._deployContract(dao.transactionContract);
dao.transactionContract.address = transtractContract.address;
dao.transactionContract.contractInstance = dao.transactionContract.contractWeb3.at(transtractContract.address);
//初始化合约的之间的关系
dao.transactionContract.contractInstance.setBuyAndSell(dao.buyAndSellContract.address,{from:chainbookAddress});
dao.transactionContract.contractInstance.setRentAndLease(dao.rentAndLeaseContract.address,{from:chainbookAddress});
dao.buyAndSellContract.contractInstance.addAddressToWhitelist(dao.transactionContract.address,{from:chainbookAddress})
dao.rentAndLeaseContract.contractInstance.addAddressToWhitelist(dao.transactionContract.address,{from:chainbookAddress});
}
//编译合约
_compile(input){
let output;
let compileInfo;
try{
function findImports (path) {
let dependentContract = fs.readFileSync(config.server.contract_path+path,'utf-8');
if (dependentContract !== undefined && dependentContract !== null)
return { contents: dependentContract }
else
return { error: 'File not found' }
}
output = solc.compile({ sources: input },1,findImports);
for (var contractName in output.contracts){
logger.info(contractName + ': ' + output.contracts[contractName]);
for(let key in input){
if(contractName.includes(key)){
compileInfo = {
compiled:output.contracts[contractName],
contractWeb3:web3.eth.contract(JSON.parse(output.contracts[contractName].interface))
};
}
}
}
}catch (e) {
logger.error("compile contract fail",e);
//如果捕捉到异常就直接抛出去,就可以启动时就知道合约编译异常
throw e;
}
//如果编译后依然为空的话,说明合约有问题,直接启动报错
if(compileInfo == undefined ||compileInfo.compiled == undefined || compileInfo.contractWeb3 == undefined ){
logger.error("compile contract fail",{
input:input,
output:output
});
throw new Error("compile error,"+input);
}else{
return compileInfo;
}
}
/**
* TODO 私有的方法
* 部署合约,这个合约由平台部署,所属地址是平台的
* @param _deployContract
* @returns {Promise<any>}
*/
_deployContract(contractInfo) {
return new Promise((resolve, reject) => {
//获取合约的代码,部署时传递的就是合约编译后的二进制码,
let deployCode = contractInfo.compiled.bytecode;
//获取部署该合约预估的费用
let gasEstimate = web3.eth.estimateGas({data:deployCode});
contractInfo.contractWeb3.new({
data: deployCode,
from: chainbookAddress,
gas:gasEstimate
}, function(err, contract) {
if (!err) {
// 注意:这个回调会触发两次,一次是合约的交易哈希属性完成,另一次是在某个地址上完成部署
// 通过判断是否有地址,来确认是第一次调用,还是第二次调用。
if (!contract.address) {
logger.info("contract deploy transaction hash: ",{
transactionHash:contract.transactionHash,
contractInfo:contractInfo
}); //部署合约的交易哈希值
} else {
// 合约发布成功后,才能调用后续的方法
logger.info("contract deploy address: ",{
transactionHash:contract.transactionHash,
address:contract.address,
contractInfo:contractInfo
});
contractInfo.address = contract.address;
contractInfo.contractInstance = contractInfo.contractWeb3.at(contract.address);
resolve(contract);
}
}else {
reject(err);
}
});
});
}
address() {
return dao.transactionContract.address;
}
setBuyAndSell(sender, _addr){
dao.transactionContract.contractInstance.setBuyAndSell.sendTransaction(_addr, {from:sender,gas:6000000});
}
setRentAndLease(sender, _addr){
dao.transactionContract.contractInstance.setRentAndLease.sendTransaction(_addr,{from:sender,gas:6000000});
}
setNewAddress(sender, _addr) {
dao.transactionContract.contractInstance.setNewAddress.sendTransaction(_addr, {from:sender,gas:6000000});
}
setBuyFees(sender, _price){
dao.transactionContract.contractInstance.setBuyFees.sendTransaction(_price, {from:sender,gas:6000000});
}
setLeaseFess(sender, _price) {
dao.transactionContract.contractInstance.setLeaseFess.sendTransaction(_price, {from:sender,gas:6000000});
}
sell(sender, _contract, _tokenId, _price) {
return dao.transactionContract.contractInstance.sell.sendTransaction(_contract, _tokenId, _price, {from:sender,gas:6000000});
}
rent(sender, _contract, _tokenId, _price, _rentTime) {
return dao.transactionContract.contractInstance.rent.sendTransaction(_contract, _tokenId, _price, _rentTime, {from:sender,gas:6000000});
}
buy(sender, val, _contract, _tokenId) {
return dao.transactionContract.contractInstance.buy.sendTransaction(_contract, _tokenId, {from:sender, value:val,gas:6000000});
}
lease(sender, val, _contract, _tokenId) {
return dao.transactionContract.contractInstance.lease.sendTransaction(_contract, _tokenId, {from:sender, value:val,gas:6000000});
}
cancelSell(sender, _contract, _tokenId) {
dao.transactionContract.contractInstance.cancelSell.sendTransaction(_contract, _tokenId, {from:sender,gas:6000000});
}
cancelRent(sender, _contract, _tokenId) {
dao.transactionContract.contractInstance.cancelRent.sendTransaction(_contract, _tokenId, {from:sender,gas:6000000});
}
setSellPrice(sender, _contract, _tokenId, _price) {
dao.transactionContract.contractInstance.setSellPrice.sendTransaction(_contract, _tokenId, _price, {from:sender,gas:6000000});
}
setRentInfo(sender, _contract, _tokenId, _price, _rentTime) {
dao.transactionContract.contractInstance.setRentInfo.sendTransaction(_contract, _tokenId, _price, _rentTime, {from:sender,gas:6000000});
}
getSellInfo( _contract, _tokenId) {
return dao.transactionContract.contractInstance.getSellInfo.call(_contract, _tokenId);
}
getRentInfo( _contract, _tokenId) {
return dao.transactionContract.contractInstance.getRentInfo.call(_contract, _tokenId);
}
pauseContract(sender) {
dao.transactionContract.contractInstance.pauseContract.sendTransaction({from:sender,gas:6000000});
}
unpauseContract(sender) {
dao.transactionContract.contractInstance.pauseContract.sendTransaction({from:sender,gas:6000000});
}
}
module.exports = new TransactionContractDao();
<file_sep>/server/utils/objectUtils.js
/**
* Created by jane.zhang on 2016/1/4.
*/
var util = require("util");
var isEmptyObject = function (obj) {
if (util.isObject(obj)) {
for (var name in obj) {
if (obj[name]) return false;
}
return true;
}
return false;
}
var toRegex = function (obj) {
var option = obj;
for (item in obj) {
if (obj[item] != undefined) {
option[item] = new RegExp(obj[item]);
}
}
return option;
}
/**
* 数组去重。
* @param array
* @returns {Array}
*/
var uniqueArray = function (array) {
var n = [];//临时数组
for (var i = 0; i < array.length; i++) {
if (n.indexOf(array[i]) == -1) n.push(array[i]);
}
return n;
}
/**
* 参数为空断言
* @param param
* @returns {Error}
*/
var notNullAssert = function(param) {
if(param == null || param == undefined ){
throw new Error("param is not null or undefined");
}
}
var notEmptyObjectAssert = function(obj) {
if(obj == undefined || obj == null || util.isEmpty(obj)){
throw new Error("obj is empty");
}
}
exports.isEmptyObject = isEmptyObject;
exports.toRegex = toRegex;
exports.uniqueArray = uniqueArray;
exports.notNullAssert = notNullAssert;
exports.notEmptyObjectAssert = notEmptyObjectAssert;
<file_sep>/server/model/resourceInfo.js
var mongoose = require("mongoose");
var resourceInfoSchema = new mongoose.Schema({
resourceName:String,
desc:String,
total:Number,
coverImage:String,
price:String,
copyrightId:String,
copyrightAddress:String,
resourceAddress: String,
authorAccount:String,
hasSellOut:Number,
sellResources:[ new mongoose.Schema({
_id: false,
tokenId:String,
ownerId:String,
ownerAccount:String,
sellPrice:String,
transactionAddress:String,
})],
tenantableResources:[new mongoose.Schema({
_id: false,
tokenId:String,
ownerId:String,
ownerAccount:String,
rentPrice:String,
rentTime:Number,
transactionAddress:String
})],
createDate: {type: Date, default: Date.now},
createBy: String,
updateDate: {type: Date, default: Date.now},
updateBy: String
}, {versionKey: false});
var ResourceInfo = mongoose.model("resourceInfo", resourceInfoSchema, "resourceInfo");
module.exports = ResourceInfo;
<file_sep>/front/src/apis/resource.js
/* eslint-disable */
export default function (request) {
return {
// 获取所有首发资源
getResourceListByPage({ page, pageSize, lastId }) {
return request({
url: '/resource/getResourceListByPage',
method: 'post',
data: {
page, pageSize, lastId
}
}).then(({ data }) => data);
},
// 获取所有二手买卖资源列表
getPurchasedResourceListByPage(page, pageSize, lastId) {
return request({
url: '/resource/getPurchasedResourceListByPage',
method: 'post',
data: {
page, pageSize, lastId
}
}).then(({ data }) => data);
},
// 获取所有可租赁资源列表
getTenantableResourceListByPage(page, pageSize, lastId) {
return request({
url: '/resource/getTenantableResourceListByPage',
method: 'post',
data: {
page, pageSize, lastId
}
}).then(({ data }) => data);
},
// 根据id获取当前资源二手买卖的owners信息
getPurchasedResourceOwnerListById(id) {
return request({
url: '/resource/getPurchasedResourceOwnerListById',
method: 'get',
params: { id }
}).then(({ data }) => {
if (Array.isArray(data) && data.length > 0){
return data[0].sellResources[0];
}
return {};
});
},
// 根据id获取当前租赁者买卖的owners信息
getTenantableResourceOwnerListById(id) {
return request({
url: '/resource/getTenantableResourceOwnerListById',
method: 'get',
params: { id }
}).then(({ data }) => {
if (Array.isArray(data) && data.length > 0){
return data[0].tenantableResources[0];
}
return {};
});
},
// 获取资源信息详情
getResourceDetailById(id) {
return request({
url: '/resource/getResourceDetailById',
method: 'get',
params: { id }
}).then(({ data }) => data);
},
// 发行审核通过的版权
publish(data) {
return request({
url: '/resource/publish',
method: 'post',
data
}).then(({ data }) => data);
},
// 上传封面图片
coverImg(data) {
return request({
url: '/resource/upload/coverImg',
method: 'post',
data
}).then(({ data }) => data);
},
// 购买首发资源
buyFromAuthor(resourceId) {
return request({
url: '/resource/buyFromAuthor',
method: 'post',
data: {
resourceId
}
}).then(({ data }) => data);
},
// 购买二手资源
buy({ tokenId, resourceId }) {
return request({
url: '/resource/buy',
method: 'post',
data: {
tokenId,
resourceId
}
}).then(({ data }) => data);
},
// 租赁
rent({ tokenId, resourceId }) {
return request({
url: '/resource/rent',
method: 'post',
data: {
tokenId,
resourceId
}
}).then(({ data }) => data);
}
};
}
<file_sep>/front/src/vuex/mutation-types.js
export const PUSH_LOADING = 'PUSH_LOADING';
export const POP_LOADING = 'POP_LOADING';
// base
export const GET_LOGIN_USERNAME = 'GET_LOGIN_USERNAME';
<file_sep>/server/utils/thunder.js
/**
* 迅雷api交互模块
*/
const axios = require('axios')
const config = require('../config');
const log4js = require('log4js');
const logger = log4js.getLogger("utils/thunder");
//创建一个axios实例
const instance = axios.create({
baseURL: config.thunder.baseURL,
timeout: 1000,
headers: {'X-Custom-Header': 'foobar'}
});
const register = function(email) {
return instance.post('/api/linktest/regist',{
email:email
});
}
exports.register = register;
<file_sep>/server/dao/copyrightContract.js
const fs = require('fs');
const config = require('../config');
const log4js = require('log4js');
const logger = log4js.getLogger('dao/copyrightContract');
const Web3 = require('web3');
const solc = require('solc');
let web3;
if (typeof web3 !== 'undefined') {
web3 = new Web3(web3.currentProvider);
} else {
// set the provider you want from Web3.providers
web3 = new Web3(new Web3.providers.HttpProvider(config.contract.url));
}
const BookCopyrightCreate_SOURCE = config.server.bookCopyrightCreate_source;
const basePath = config.server.contract_path;
const chainbookAddress = config.server.address;
const chainbookGas = config.server.gas;
const CopyrightContractDao = class dao {
constructor(){
this._compile();
this._deployContract();
}
//编译合约
_compile(){
let output;
try{
let bookCopyrightCreate = fs.readFileSync(BookCopyrightCreate_SOURCE,'utf-8');
var input = {
'BookCopyrightCreate.sol': bookCopyrightCreate
}
function findImports (path) {
let dependentContract = fs.readFileSync(basePath+path,'utf-8');
if (dependentContract !== undefined && dependentContract !== null)
return { contents: dependentContract }
else
return { error: 'File not found' }
}
output = solc.compile({ sources: input },1,findImports)
for (var contractName in output.contracts){
logger.info(contractName + ': ' + output.contracts[contractName]);
if(contractName.includes("BookCopyrightCreate.sol")){
dao.bookCopyrightCreateCompiled = output.contracts[contractName];
dao.bookCopyrightCreateContactWeb3 = web3.eth.contract(JSON.parse(dao.bookCopyrightCreateCompiled.interface));
}
}
}catch (e) {
logger.error("compile contract fail",e);
//如果捕捉到异常就直接抛出去,就可以启动时就知道合约编译异常
throw e;
}
//如果编译后依然为空的话,说明合约有问题,直接启动报错
if(dao.bookCopyrightCreateCompiled == undefined || dao.bookCopyrightCreateContactWeb3 == undefined){
logger.error("compile BookCopyrightCreate.sol contract fail",output);
throw new Error("compile BookCopyrightCreate.sol error");
}
}
/**
* TODO 私有的方法
* 部署合约,这个合约由平台部署,所属地址是平台的
* @param _deployContract
* @returns {Promise<any>}
*/
_deployContract() {
return new Promise((resolve, reject) => {
//获取合约的代码,部署时传递的就是合约编译后的二进制码,
let deployCode = dao.bookCopyrightCreateCompiled.bytecode;
//获取部署该合约预估的费用
let gasEstimate = web3.eth.estimateGas({data:deployCode});
dao.bookCopyrightCreateContactWeb3.new({
data: deployCode,
from: chainbookAddress,
gas:gasEstimate
}, function(err, contract) {
if (!err) {
// 注意:这个回调会触发两次,一次是合约的交易哈希属性完成,另一次是在某个地址上完成部署
// 通过判断是否有地址,来确认是第一次调用,还是第二次调用。
if (!contract.address) {
logger.info("contract deploy transaction hash: " + contract.transactionHash) //部署合约的交易哈希值
} else {
// 合约发布成功后,才能调用后续的方法
dao.bookCopyrightAddress = contract.address;//合约的部署后的地址
dao.contractInstance = dao.bookCopyrightCreateContactWeb3.at(dao.bookCopyrightAddress);
logger.info("contract deploy address: " + contract.address) // 合约的部署地址
resolve(contract)
}
}else {
reject(err);
}
});
});
}
registerCopyright(userObj,copyrightObj) {
return new Promise((resolve, reject) => {
//获取一个已有地址的合约实例
try{
//调用这个合约的
let transactionId = dao.contractInstance.registerCopyright(
copyrightObj.workName,
copyrightObj.authors[0].authorName,
copyrightObj.account,
copyrightObj.resourceHash,{
from: chainbookAddress,
gas:chainbookGas //由平台提供gas
});
logger.info("registerCopyright transactionId:",transactionId);
//设置注册成功的监听事件
let registerCopyrightEvent = dao.contractInstance._RegisterCopyright();
// 监听事件,监听到事件后会执行回调函数
registerCopyrightEvent.watch(function(err, result) {
if (err) {
reject(e);
}else{
if(transactionId === result.transactionHash){
logger.info("_RegisterCopyright event",{
result:result,
_newBookCopyrightId:result.args._newBookCopyrightId.toString()
});
registerCopyrightEvent.stopWatching();
resolve(result.args._newBookCopyrightId.toString());
}
}
});
}catch (e) {
reject(e);
}
});
}
getCopyRightsByCopyrightAddress(copyrightAddress) {
return new Promise((resolve, reject) => {
try{
resolve(dao.contractInstance.getCopyright.call(copyrightAddress));
}catch (e) {
reject(e);
}
});
}
//返回版权合约得地址
getCopyrightContractAddress(){
return dao.bookCopyrightAddress;
}
}
module.exports = new CopyrightContractDao();
<file_sep>/smartcontracts/migrations/2_BookCopyrightCreat.js
var BookCopyrightCreate = artifacts.require("./BookCopyrightCreate.sol");
module.exports = function(deployer) {
var account = web3.eth.accounts[0];
deployer.deploy(BookCopyrightCreate).then(function(instance) {
instance.registerCopyright("chainbook","chainbook",account,"chainbook");
})
};
<file_sep>/server/model/resourceCopyright.js
var mongoose = require("mongoose");
var ResourceCopyrightSchema = new mongoose.Schema({
workName:String,
workCategory:String,
copyrightAddress:String,
resourceId:String,
resourceAddress:String,
resourceHash:String,
resourceDHash:String,
account:String,
localUrl:String,
authors:[new mongoose.Schema({
_id: false,
authorName:String,
identityType:String,
identityNum:Number
})],
workProperty:String,
rights:[String],
belong:String,
auditStatus:Number,
publishStatus:Number,
createDate: {type: Date, default: Date.now},
createBy: String,
updateDate: {type: Date, default: Date.now},
updateBy: String
}, {versionKey: false});
var ResourceCopyright = mongoose.model("resourceCopyright", ResourceCopyrightSchema, "resourceCopyright");
module.exports = ResourceCopyright;
<file_sep>/server/controller/resourceCopyright.js
const log4js = require('log4js');
const logger = log4js.getLogger('controller/resourceCopyright');
const userDao = require("../dao/user");
const resourceCopyrightDao = require("../dao/resourceCopyright");
const copyrightContractDao = require("../dao/copyrightContract");
const objectUtils = require("../utils/objectUtils");
const encrypt = require("../utils/encrypt");
const localUpload = require("../component/localUpload");
const ipfsResourcesDao = require("../component/ipfsResources");
const NO_AUDIT = 0;
const NO_PUBLISH = 0;
/**
* 申请版权
* @param req
* @param res
* @param next
*/
exports.applyCopyright = async function(req,res,next){
let user = req.session.passport.user;
let copyright = {
workName: req.body.workName,
workCategory: req.body.workCategory,
localUrl:req.body.localUrl,
account:user.account,
authors:req.body.authors,
workProperty:req.body.workProperty,
rights:req.body.rights,
belong:req.body.belong,
auditStatus:NO_AUDIT,
publishStatus:NO_PUBLISH
}
try{
//TODO 需要校验是否重复提交内容
//TODO 校验数据的合法性,表单校验
objectUtils.isEmptyObject(copyright);
objectUtils.isEmptyObject(copyright.authors);
objectUtils.isEmptyObject(copyright.rights);
//先将申请的版权信息登记进数据库
let savedObj = await resourceCopyrightDao.add(copyright);
//将数据保存至user
let copyrightObj = userDao.buildEmptyCopyright();
copyrightObj.localUrl = savedObj.localUrl;
copyrightObj.copyrightId = savedObj._id;
copyrightObj.workName = savedObj.workName;
await userDao.addCopyRightByUser(user._id,copyrightObj);
logger.info("apply copyright success",copyright);
res.send({status:1,msg:"申请版权信息保存成功"});
}catch (e) {
logger.error("save copyright fail",{ copyright: copyright},e);
res.send({status:0,msg:"申请版权信息保存失败"});
}
}
/** TODO
* 上传样本至服务器
* @param req
* @param res
* @param next
*/
exports.uploadSample = function(req,res,next){
let upload = localUpload.fileUpload.single("sample");
upload(req,res,function(err) {
if(err){
logger.error("upload fail",err);
res.send({status:0,msg:'上传样本失败'});
return;
}
var file = req.file;
if(!file){
res.send({status:0,msg:'上传样本失败'});
return;
}
logger.info('file info',{
mimetype:file.mimetype,
originalname:file.originalname,
size:file.size,
path:file.path });
res.send({status:1,msg:'success',data:{path:file.path.replace(/\\/g,"/")}});
});
}
exports.auditCopyright = async function(req,res,next){
let user = req.session.passport.user;
let copyrightId = req.body.copyrightId;
let isSuccess = await registerCopyright(user,copyrightId);
if(isSuccess){
res.send({status:1,msg:"审核版权成功"});
}else{
res.send({status:0,msg:"审核版权失败"});
}
}
/**
* 登记版权至合约,将被登记合约的定时任务调用 TODO
* @param req
* @param res
* @param next
*/
let registerCopyright = async function(userObj,copyrightId){
try{
//1、根据userAccount捞取基本用户信息
let account = userObj.account;
let copyrightObj = await resourceCopyrightDao.findOne({_id:copyrightId,account:account});
if(copyrightObj.auditStatus > 0){
logger.warn("该版权审核通过或者失败",{
userObj:userObj,
copyrightObj:copyrightObj
});
return false;
}
logger.info("register copyright",{
user:userObj,
copyrightId:copyrightId,
copyrightObj:copyrightObj
});
//2、将本地的最新样本上传至ipfs,并获得hash值
let resourceHash = await ipfsResourcesDao.upload(copyrightObj.localUrl,userObj);
let resourceDHash = encrypt.getMD5(resourceHash,"");
//3、调用合约的登记版权的方法,登记合约,保存版权的信息和版权的hash值,并返回合约地址
copyrightObj.resourceHash = resourceHash;
copyrightObj.resourceDHash = resourceDHash;
let copyrightAddress = await copyrightContractDao.registerCopyright(userObj,copyrightObj);
//4、登记hash,dhash,合约地址,审核状态等至resourceCopyright和user中
await resourceCopyrightDao.updateResourceCopyrightInfo(copyrightObj._id,copyrightAddress,resourceHash,resourceDHash,1);
//5、登记到用户的版权信息下
await userDao.modifyCopyrightAuditInfo(userObj._id,copyrightObj._id,resourceHash,resourceDHash,copyrightAddress);
return true;
}catch (e) {
logger.error("registerCopyright fail.",{
user:userObj,
copyrightId:copyrightId
},e);
return false;
}
}
/**
* 根据id获取资源版权细节
* @param req
* @param res
* @param next
*/
exports.getResourceCopyrightDetailById = async function(req, res, next) {
let id = req.param('id');
logger.info("getResourceCopyrightDetailById",{id:id});
try{
objectUtils.notNullAssert(id);
let resourceCopyright = await resourceCopyrightDao.findById(id);
res.send({status:1,msg:"success",data:resourceCopyright});
}catch (e) {
logger.error("get resource copyright detail fail.",id,e);
res.send({status:0,msg:"get resource copyright detail fail."});
}
}
<file_sep>/front/src/config/resource/enum.js
export const ListType = {
FirstResource: 'FirstResource', // 首发资源
SecondHand: 'SecondHand', // 二手市场
Rent: 'Rent' // 租赁市场
};
export const Operation = {
};
<file_sep>/front/.eslintrc.js
// https://eslint.org/docs/user-guide/configuring
module.exports = {
root: true,
parser: 'babel-eslint',
parserOptions: {
sourceType: 'module'
},
env: {
browser: true,
},
extends: 'airbnb-base',
// required to lint *.vue files
plugins: [
'html'
],
// check if imports actually resolve
'settings': {
'import/resolver': {
'webpack': {
'config': 'build/webpack.base.conf.js'
}
}
},
// add your custom rules here
'rules': {
// don't require .vue extension when importing
'no-lone-blocks': 0,
'global-require': 0,
'import/no-unresolved': 0,
'newline-per-chained-call': [
0, {
'ignoreChainWithDepth': true
}
],
'no-restricted-syntax': [
2,
'ForInStatement',
'LabeledStatement',
'WithStatement',
],
// allow debugger during development
'no-debugger': process.env.NODE_ENV === 'production' ? 2 : 0,
'no-console': 0,
'comma-dangle': 0,
'no-unused-vars': 1,
'no-param-reassign': 0,
'no-shadow': 0,
'no-trailing-spaces': 0,
'no-underscore-dangle': 0,
'max-len': [2, 100, 2, {
'ignoreUrls': true,
'ignoreComments': true,
'ignoreStrings': true,
'ignoreTemplateLiterals': true
}],
'import/extensions': ['error', 'never', { // TODO: 从always改到了never
'js': 'never',
'vue': 'never'
}],
// allow optionalDependencies
'import/no-extraneous-dependencies': ['error', {
'optionalDependencies': ['test/unit/index.js']
}],
// allow debugger during development
'no-debugger': process.env.NODE_ENV === 'production' ? 2 : 0,
'prefer-promise-reject-errors': ['off', {
'allowEmptyReject': true
}],
'object-curly-newline': 0
}
}
| 52eec035f0791c854a584ce20bea748dd21b2d9c | [
"Markdown",
"JavaScript"
] | 33 | Markdown | jane-great/ChainBook | 08982919a022bab116aeaa2c06ad78e174cb776f | c667ef319b86487a60ee9f0fb337b3e6dbdaed67 |
refs/heads/master | <file_sep># -*- coding: utf-8 -*-
# Generated by Django 1.10.3 on 2017-01-11 11:28
from __future__ import unicode_literals
import django.core.validators
from django.db import migrations, models
import django.utils.timezone
class Migration(migrations.Migration):
initial = True
dependencies = [
]
operations = [
migrations.CreateModel(
name='Mattress',
fields=[
('id', models.AutoField(auto_created=True, primary_key=True, serialize=False, verbose_name='ID')),
('name', models.CharField(max_length=200)),
('width', models.IntegerField(default=90, validators=[django.core.validators.MinValueValidator(70), django.core.validators.MaxValueValidator(200)])),
('height', models.IntegerField(default=200, validators=[django.core.validators.MinValueValidator(180), django.core.validators.MaxValueValidator(220)])),
('thickness', models.IntegerField(default=20, validators=[django.core.validators.MinValueValidator(3), django.core.validators.MaxValueValidator(50)])),
('description', models.TextField()),
('publish_date', models.DateField(default=django.utils.timezone.now)),
('price', models.DecimalField(decimal_places=2, max_digits=8)),
('stock', models.IntegerField(default=0)),
],
),
]
<file_sep>from django.contrib import admin
from store.models import Mattress
# Register your models here.
class MattressAdmin(admin.ModelAdmin):
list_display = ('name', 'price', 'stock', 'width', 'height')
admin.site.register(Mattress, MattressAdmin)
<file_sep>from django.db import models
from datetime import date
import sys
from django.utils import timezone
from django.core.validators import MinValueValidator, MaxValueValidator
# Create your models here.
class Mattress(models.Model):
name = models.CharField(max_length=200)
width = models.IntegerField(validators=[MinValueValidator(70),MaxValueValidator(200)], default=90)
height = models.IntegerField(validators=[MinValueValidator(180),MaxValueValidator(220)], default=200)
thickness = models.IntegerField(validators=[MinValueValidator(3),MaxValueValidator(50)], default=20)
description = models.TextField()
publish_date = models.DateField(default=timezone.now)
price = models.DecimalField(max_digits=8, decimal_places=2)
stock = models.IntegerField(default=0)<file_sep>from django.db import models
from django.utils import timezone
class Article(models.Model):
title = models.CharField(max_length=200)
author = models.CharField(max_length=200)
summary = models.TextField()
main_text = models.TextField()
publish_date = models.DateField(default=timezone.now)
def publish(self):
self.published_date = timezone.now()
self.save()
def __str__(self):
return self.title | 1334aa2be1b6f1a452f8c4dacdcce0e2df77ab51 | [
"Python"
] | 4 | Python | khortur/michalbiernacki | 3912786d75e804762b4f13e6cb18c005abcbecac | 1fc8da61fd4a4cd2af07ea5bc6b176803eec0c81 |
refs/heads/master | <file_sep>var video = document.querySelector('#video');
var cPallete = document.querySelector("#doit");
video.addEventListener('click', takeimage, true);
cPallete.addEventListener('click', takeimage, true);
//---------------
//normalize window.URL
window.URL || (window.URL = window.webkitURL || window.msURL || window.oURL);
//normalize navigator.getUserMedia
navigator.getUserMedia || (navigator.getUserMedia = navigator.webkitGetUserMedia || navigator.mozGetUserMedia || navigator.msGetUserMedia);
//detect if {video: true} or "video" style options
//by creating an iframe and blowing it up
//style jacked from @kangax
var optionStyle = (function(win){
//only test if there's something to test
if (!navigator.getUserMedia) return;
var el = document.createElement('iframe'),
root = document.body || document.documentElement,
string = true, object = true, nop = function(){};
root.appendChild(el);
var f = win.frames[win.frames.length-1];
f.navigator.getUserMedia || (f.navigator.getUserMedia = f.navigator.webkitGetUserMedia || f.navigator.mozGetuserMedia || f.navigator.msGetUserMedia);
try { //try it with spec syntax
f.navigator.getUserMedia({video: true}, nop);
} catch (e) {
object = false;
try { //try it with old spec string syntax
f.navigator.getUserMedia("video", nop);
} catch (e) { //neither is supported
string = false;
}
} finally { //clean up
root.removeChild(el);
el = null;
}
return {string: string, object: object}
})(window),
//normalize the options object to a string
//if that's the only thing supported
norm = function(opts){ // has to be {video: false, audio: true}. caveat emptor.
var stringOptions = [];
if (optionStyle.string && !optionStyle.object) {
//pluck the "true"s
for (var o in opts) {
if (opts[o] == true) {
stringOptions.push(o);
}
}
return stringOptions.join(" ");
} else {
//really, this will blow up if you pass it {video: true, rofl: "copter"}. so don't.
return opts;
}
},
hollaback = function(stream) {
video.src = (window.URL && window.URL.createObjectURL) ? window.URL.createObjectURL(stream) : stream;
};
if (navigator.getUserMedia) {
navigator.getUserMedia(norm({video: true, audio: false}), hollaback, not_supported);
}
else
{alert('Browser does not support getUserMedia!');}
// ----------------
function not_supported(){
var vid_c = document.querySelector("#video_container");
vid_c.innerHTML = "It seems this browser does not support <code>navigator.getUserMedia()<\/code>, please use a browser which does in order to see this demo in action.";
cPallete.className = "hide";
}
function v_success(stream){
video.src = stream;
}
function v_error(error){
console.log("Error! Error code is:"+error.code);
}
function takeimage(){
var canvas = document.querySelector('#mycanvas');
var ctx = canvas.getContext('2d');
var cw = canvas.width;
var ch = canvas.height;
var pixelCount = cw*ch;
ctx.drawImage(video, 0, 0, cw, ch);
var pixels = ctx.getImageData(0, 0, cw, ch).data;
//otherColors(pixels, pixelCount);
dominantColor(pixels, pixelCount);
}
function dominantColor(pixels, pixelCount) {
var pixelArray = [];
for (var i = 0; i < pixelCount; i++) {
// If pixel is mostly opaque and not white
if(pixels[i*4+3] >= 125){
if(!(pixels[i*4] > 250 && pixels[i*4+1] > 250 && pixels[i*4+2] > 250)){
pixelArray.push( [pixels[i*4], pixels[i*4+1], pixels[i*4+2]]);
}
}
}
// Send array to quantize function which clusters values
// using median cut algorithm
var cmap = MMCQ.quantize(pixelArray, 16);
var newPalette = cmap.palette();
var colorArray = {"r": newPalette[0][0], "g": newPalette[0][1], "b": newPalette[0][2]};
var colors = document.querySelector("#colors");
var thediv = document.createElement('div');
thediv.className = 'othercolors';
var colors = ['#aaffee','#cc44cc','#00cc55','#0000aa'];
var rgb = ['+colorArray.r+','+colorArray.g+','+colorArray.b+'];
var best = 768;
var canvasColor = colors[0];
for (i=0; i< colors.length; i++){
t = 0;
for (j=0; j<3; j++){
t += Math.abs(rgb[j] - parseInt(colors[i].substring(j*2+1,j*2+3), 16));
}
if (t < best){
best = t;
canvasColor = colors[i];
}
}
thediv.setAttribute('style', "background-color:rgb("+colorArray.r+","+colorArray.g+","+colorArray.b+");");
colorlist.appendChild(thediv);
}
<file_sep><!DOCTYPE html>
<html>
<head>
<title>l a p s e n s e</title>
<meta http-equiv="content-type" content="text/html; charset=utf-8" />
<meta name="viewport" content="width=device-width, user-scalable=no, minimum-scale=1.0, maximum-scale=1.0">
<meta name="mobile-web-app-capable" content="yes">
<meta name="apple-mobile-web-app-capable" content="yes" />
<link rel="manifest" href="/manifest.json">
<script src="bower_components/webcomponentsjs/webcomponents.min.js"></script>
<script type="text/javascript" src="js/haptic/haptic.js"></script>
<script src="https://code.jquery.com/jquery-2.1.4.min.js"></script>
<script src="https://code.jquery.com/ui/1.11.4/jquery-ui.min.js"></script>
<link rel="import" href="bower_components/core-toolbar/core-toolbar.html">
<link rel="import" href="bower_components/core-menu/core-menu.html">
<link rel="import" href="bower_components/core-item/core-item.html">
<link rel="import" href="bower_components/core-header-panel/core-header-panel.html">
<link rel="import" href="bower_components/core-drawer-panel/core-drawer-panel.html">
<link rel="import" href="bower_components/core-icons/core-icons.html">
<link rel="import" href="bower_components/paper-icon-button/paper-icon-button.html">
<link rel="import" href="bower_components/font-roboto/roboto.html">
<link rel="import" href="bower_components/google-map/google-map.html">
<link rel="import" href="bower_components/geo-location/geo-location.html">
<link rel="import" href="bower_components/core-selector/core-selector.html">
<link rel="import" href="bower_components/core-image/core-image.html">
<link rel="import" href="bower_components/paper-item/paper-item.html">
<link rel="import" href="bower_components/paper-input/paper-input.html">
<link rel="import" href="bower_components/paper-fab/paper-fab.html">
<link rel="stylesheet" type="text/css" href="css/style.css">
</head>
<body fullbleed unresolved>
<geo-location watchpos></geo-location>
<!-- sidemenu + frontal layer -->
<script>
document.addEventListener('polymer-ready', function() {
var navicon = document.getElementById('navicon');
var drawerPanel = document.getElementById('drawerPanel');
navicon.addEventListener('click', function() {
drawerPanel.togglePanel();
});
});
</script>
<?php
$zoomMap = 14;
if(isset($_POST['zoomMap']) && $_POST['zoomMap']>14){
//$zoomMap = $_POST['zoomMap'];
//$_POST['zoomMap']=14;
//echo $_POST['zoomMap'];
}
?>
<div id="coreApp">
<core-drawer-panel id="drawerPanel" drawerWidth="300px">
<!-- sidemenu -->
<core-header-panel mode="waterfall-tall" drawer style="background-color: #eaeaea; ">
<?php include('slidemenu.php'); ?>
</core-header-panel>
<!-- frontal layer -->
<core-header-panel main mode="seamed" id='mainPanel'>
<core-toolbar id="mainheader" style="color:white" >
<paper-icon-button id="navicon" icon="menu"></paper-icon-button>
<form method='post' action="index.php" id='form'>
<input type="text" value="17" class="hidden" name='zoomMap'>
<paper-icon-button id="cancel" icon="clear" class="hidden" onclick="document.getElementById('form').submit();"></paper-icon-button>
</form>
<paper-input-decorator class="custom" label="#yourhastag"><input id="i1" is="core-input" maxlength="20" type="search">
<paper-char-counter class="counter" target="i1"></paper-char-counter>
</paper-input-decorator>
<paper-icon-button id="refresh" icon="refresh" onclick="window.open('index.php','_parent');"></paper-icon-button>
<paper-icon-button id="accept" icon="done" class="hidden"></paper-icon-button>
</core-toolbar>
<!-- threejs random geometry -->
<div id="container"></div>
<script src="js/threejs/three.min.js"></script>
<script src="js/threejs/geometries/ConvexGeometry.js"></script>
<script src="js/threejs/libs/OrbitControls.js"></script>
<script src="js/threejs/Detector.js"></script>
<script src="js/threejs/lapse_color.js"></script>
<!-- capture button -->
<div class="capture">
<paper-fab id="fab1" icon="polymer"></paper-fab>
<paper-icon-button id="iconList" icon="menu"></paper-icon-button>
</div>
</core-header-panel>
</core-drawer-panel>
<!-- map -->
<div class="map" >
<template is="auto-binding">
<geo-location latitude="{{lat}}" longitude="{{lng}}" watchpos highaccuracy></geo-location>
<google-map latitude="{{lat}}" longitude="{{lng}}" minZoom="<?php echo $zoomMap ?>" maxZoom="<?php echo $zoomMap ?>" disableDefaultUI showCenterMarker fit></google-map>
</template>
</div>
</div>
<!-- List of old captures-->
<div id='listLateral'>
<div id='lateral'>
<div id="listHeader">
<paper-icon-button id="back" icon="arrow-back"></paper-icon-button>
</div>
<div id='theList'>
<?php include('list.php'); ?>
</div>
</div>
</div>
<script src="js/captureButtonProcess.js"></script>
</body><file_sep>$(window).load(function() {
$("#fab1").click(function(){
shortAndSharp(1);
$("#fab1").addClass('hidden');
$("#iconList").addClass('hidden');
$("#navicon").addClass('hidden');
$("#refresh").addClass('hidden');
$("#accept").removeClass('hidden');
$("#cancel").removeClass('hidden');
if ( ! Detector.webgl ) Detector.addGetWebGLMessage();
// prepare three.js
var container;
var camera, controls, scene, renderer;
var def_col = ran_col();
var points = init(def_col);
creatObject3D(points,def_col);
render();
});
$('#accept').click(function(){
window.open('index.php','_parent');
});
});
jQuery(document).ready(function () {
jQuery("#iconList").click(function () {
jQuery("#drawerPanel").animate({"width": ["0%", 'easeOutExpo']}, {
duration: 700
});
$('#mainheader').addClass('hidden');
$('.capture').addClass('hidden');
jQuery("#listLateral").animate({"marginLeft": ["-1px", 'easeOutExpo']}, {
duration: 700
});
jQuery(".map").animate({"opacity": ["0", 'easeOutSine']}, {
duration: 700
});
});
jQuery("#myList").click(function () {
jQuery("#drawerPanel").animate({"width": ["0%", 'easeOutExpo']}, {
duration: 700
});
var drawerPanel = document.getElementById('drawerPanel');
drawerPanel.togglePanel();
$('#mainheader').addClass('hidden');
$('.capture').addClass('hidden');
jQuery("#listLateral").animate({"marginLeft": ["-1px", 'easeOutExpo']}, {
duration: 700
});
jQuery(".map").animate({"opacity": ["0", 'easeOutSine']}, {
duration: 700
});
});
jQuery("#back").click(function () {
jQuery("#drawerPanel").animate({"width": ["100%", 'easeOutExpo']}, {
duration: 700
});
jQuery(".map").animate({"opacity": ["1", 'easeOutSine']}, {
duration: 700
});
$('#mainheader').removeClass('hidden');
$('.capture').removeClass('hidden');
jQuery("#listLateral").animate({"marginLeft": ["100%", 'easeOutExpo']}, {
duration: 700
});
});
});
/*
function showList(){
//set margin for the whole container with a jquery UI animation
$("#coreApp").animate({"marginRight": ["100%", 'easeOutExpo']}, {
duration: 700
});
$('#coreApp').addClass('hidden');
$("#listLateral").removeClass('hidden');
}
function goBack(){
$('#coreApp').removeClass('hidden');
$("#listLateral").addClass('hidden');
}
*/<file_sep>
// random background color
function ran_col() {
var color = '#';
var letters = ['ef4437','e71f63','8f3e97','65499d','4554a4','478fcc','38a4dc','09bcd3', '009688','4cae4e','8bc248','cddc37','feea39','fdc010','f8971c','f0592b','607d8b', '795548'];
color += letters[Math.floor(Math.random() * letters.length)];
return color;
}
function animate() {
requestAnimationFrame(animate);
controls.update();
if(dragging && !dragging2){ controls.inertiaFunction(); }
}
function init(def_col) {
// color frontal layout
var meta = document.createElement('meta');
meta.name = "theme-color";
meta.content = def_col;
document.getElementsByTagName('head')[0].appendChild(meta);
document.getElementById("mainheader", "fab1").style.backgroundColor = def_col;
// create scene
camera = new THREE.PerspectiveCamera( 60, window.innerWidth / window.innerHeight, 1, 1000 );
camera.position.z = 900;
controls = new THREE.OrbitControls( camera );
controls.damping = 0.2;
controls.addEventListener( 'change', render );
points = [];
for ( var i = 3; i < 100; i ++ ) {
points.push( randomPointInSphere( 100 ) ); // scale
}
return points;
}
function creatObject3D(points,def_col){
scene = new THREE.Scene();
scene.fog = new THREE.FogExp2( def_col, 0.0009 );
bg = document.body.style;
// random convex
materials = [
new THREE.MeshLambertMaterial( { color: def_col, shading: THREE.FlatShading } ) // material random color
];
var light, geometry, material, mesh;
object = THREE.SceneUtils.createMultiMaterialObject( new THREE.ConvexGeometry( points ), materials );
scene.add( object );
// lights
light = new THREE.DirectionalLight( def_col, 0.5 );
light.position.set( 1, 1, 1 );
scene.add( light );
light = new THREE.DirectionalLight( def_col, 0.5 );
light.position.set( -1, -1, -1 );
scene.add( light );
// renderer
renderer = new THREE.WebGLRenderer( { alpha: true, antialias: true } );
renderer.setClearColor( 0x000000, 0 );
renderer.setPixelRatio( window.devicePixelRatio );
renderer.setSize( window.innerWidth, window.innerHeight );
container = document.getElementById( 'container' );
container.appendChild( renderer.domElement );
//
window.addEventListener( 'resize', onWindowResize, false );
animate();
}
function onWindowResize() {
camera.aspect = window.innerWidth / window.innerHeight;
camera.updateProjectionMatrix();
renderer.setSize( window.innerWidth, window.innerHeight );
render();
}
function randomPointInSphere( radius ) {
return new THREE.Vector3(
( Math.random() - 0.5 ) * Math.floor(Math.random() * Math.floor(Math.random() * 4) + 1) * radius,
( Math.random() - 0.5 ) * Math.floor(Math.random() * Math.floor(Math.random() * 4) + 1) * radius,
( Math.random() - 0.5 ) * Math.floor(Math.random() * Math.floor(Math.random() * 4) + 1) * radius
);
}
function render() {
renderer.render( scene, camera );
stats.update();
} | 17ddc8f2ceb19161876405026eec6b7d084784e5 | [
"JavaScript",
"PHP"
] | 4 | JavaScript | serg1mm/lapse | c1f1772dee3cf05072f51077714c75279b2fb30a | 53cfb60c7ac04538019a1c6cf861d68508bb66cf |
refs/heads/master | <repo_name>kdbanman/oneDim<file_sep>/ruleDoc.py
cond1 = '111'
cond2 = '110'
cond3 = '101'
cond4 = '100'
cond5 = '011'
cond6 = '010'
cond7 = '001'
cond8 = '000'
responses = []
for i in range(0,256):
responses.append(0)
responses[i] = bin(i)[2:]
prepend = []
for j in range(0,8-len(responses[i])):
prepend.append('0')
prestring = ''
for zero in prepend:
prestring = prestring + zero
responses[i] = prestring + responses[i]
rules = {}
i = 0
for response in responses:
rules[i] = {}
rules[i][cond1] = response[0]
rules[i][cond2] = response[1]
rules[i][cond3] = response[2]
rules[i][cond4] = response[3]
rules[i][cond5] = response[4]
rules[i][cond6] = response[5]
rules[i][cond7] = response[6]
rules[i][cond8] = response[7]
i += 1
doc = open('ruleDoc.xml','w')
tabLvl = 0
doc.write('<rules>\n')
for k,v in rules.items():
doc.write('\t<rule>\n')
doc.write('\t\t<name>' + str(k) + '</name>\n')
for cond,resp in v.items():
doc.write('\t\t<condition>\n')
doc.write('\t\t\t<locality>\n')
doc.write('\t\t\t\t<left>' + cond[0] + '</left>\n')
doc.write('\t\t\t\t<self>' + cond[1] + '</self>\n')
doc.write('\t\t\t\t<right>' + cond[2] + '</right>\n')
doc.write('\t\t\t</locality>\n')
doc.write('\t\t\t<response>' + resp + '</response>\n')
doc.write('\t\t</condition>\n')
doc.write('\t</rule>\n')
doc.write('</rules>')
doc.close()
<file_sep>/1d.py
# render subset of environment. view moved by mouse position, user-selectable width
# if positive state an negative state try and occupy the same cell; it dies, all live/dead cells in direct contact die, or just surrounding cells die?
import pygame, sys
from pygame.locals import *
import random
def render(env, gen, len):
x = 0
top = (gen*3)%(len)
for cell in env:
if cell == 1:
color = liveColor
else:
color = whiteColor
left = 3*x
pygame.draw.rect(windowSurf,color, (left, top, 3, 3))
x = x + 1
pygame.display.update()
def iterate(env):
newEnv = []
i = 0
while i < len(env):
local = (env[(i-1)%len(env)],env[i],env[(i+1)%len(env)])
if local in liveSet:
state = 1
else:
state = 0
newEnv.append(state)
i = i + 1
return newEnv
size = int(input('environment size: '))
lifespan = int(input('lifespan: '))
#until we have a controllable render subset, render width needs to be 3*size to be meaningful
window = int(input('render width: '))
antiState = input('antistate? (1 for yes, 0 for no) ')
if antiState == 1:
antiState = True
renderMode = int(input('mode: '))
if renderMode == 3:
prob = int(input('probability: '))
if renderMode == 4:
width = size
while width >= size or width <= 0:
width = int(input('block width: '))
window = window - window%3
length = 798
pygame.init()
windowSurf = pygame.display.set_mode((window,length))
pygame.display.set_caption('1D Automata, Bitch')
whiteColor = pygame.Color(255,255,255)
liveColor = pygame.Color(0,0,0)
liveSet = set( ((0,0,1),(1,1,0),(0,1,1),(0,1,0),(1,0,1)) )
environment = []
i = 0
if renderMode == 1:
while i < size:
if i == int(size/2):
environment.append(1)
else:
environment.append(0)
i = i + 1
if renderMode == 2:
while i < size:
if i >= int(2*size/5) and i <= int(3*size/5):
environment.append(1)
else:
environment.append(0)
i = i + 1
if renderMode == 3:
while i < size:
if random.randint(0,100) < prob:
environment.append(1)
else:
environment.append(0)
i = i + 1
if renderMode == 4:
while i < size:
if i%2 == 0 and i >= int((size - width)/2) and i <= int((size + width)/2):
environment.append(1)
else:
environment.append(0)
i = i + 1
if antiState:
antiEnv = []
i = 0
while i < size:
if i == 0:
antiEnv.append(1)
else:
antiEnv.append(0)
i = i + 1
generation= 0
while generation < lifespan:
render(environment, generation, length)
environment = iterate(environment)
if antiState == True:
antiEnv = iterate(antiEnv)
i = 0
while i < len(environment):
if antiEnv[i] == 1 and environment[i] == 1:
antiEnv[i] = 0
environment [i] = 0
i = i + 1
generation = generation + 1
| 7bfa091fafaa1974c4a5360377096961cd8cd1ec | [
"Python"
] | 2 | Python | kdbanman/oneDim | 8ff37dc56d7ede799a6fce5665d057b517a38ee5 | 4b9b52adc652018c493d160eb678ae46e23d5e94 |
refs/heads/master | <file_sep>//Description:
//This time no story, no theory. The examples below show you how to write function accum:
//Examples:
//accum("abcd"); // "A-Bb-Ccc-Dddd"
//accum("RqaEzty"); // "R-Qq-Aaa-Eeee-Zzzzz-Tttttt-Yyyyyyy"
//accum("cwAt"); // "C-Ww-Aaa-Tttt"
//The parameter of accum is a string which includes only letters from a..z and A..Z.
function accum(s) {
let output = "";
let dash = "-";
for ( let i = 0; i < s.length; i++) {
output = output + s[i].toUpperCase();
console.log("output before second for: ", output);
for (let j = 0; j < i; j++) {
output = output + s[i].toLowerCase();
}
output = output + dash;
console.log("output after second for: ", output);
}
return output.slice(0, output.length-1);
}
| 9517e4976fdb019c7f7f3f2f70208d378d95b550 | [
"JavaScript"
] | 1 | JavaScript | myraha/codewars1 | bbfbdd9ea6a58938aabd6c6b98f5aeb18b7912f4 | 919d3aeee2890521eb8f21657700075e48a57492 |
refs/heads/main | <file_sep>package basic;
import util.Util;
/**
* In a sorted array, use binary search to check the existence of given value
*/
public class Code004BinarySearchExist {
public static void main(String[] args) {
final int testSum = 1_000_000;
int[] arr, arr1;
int size = 100, range = 30;
int value;
long start = Util.getSysTime();
System.out.println("test start ...");
for (int i = 0; i < testSum; i++) {
arr = Util.generateRandomArray(size, range);
value = Util.generateRandomValue(range);
if (arr == null) {
throw new RuntimeException("invalid array not allowed");
}
arr1 = Util.copyArray(arr);
mergeSort(arr1);
if (searchValue(arr, value) != binarySearchValue(arr1, value)) {
System.out.println("test fail ...");
Util.printArray(arr);
Util.printArray(arr1);
return;
}
}
long end = Util.getSysTime();
System.out.println("test pass ... time taken : " + (end - start) / 1000f);
}
private static boolean searchValue(int[] arr, int value) {
if (arr == null || arr.length == 0) {
return false;
}
for (int i : arr) {
if (i == value) {
return true;
}
}
return false;
}
private static boolean binarySearchValue(int[] arr, int value) {
if (arr == null || arr.length == 0) {
return false;
}
int mid;
int left = 0;
int right = arr.length - 1;
while (left < right) {
mid = left + ((right - left) >> 1);
if (arr[mid] == value) {
return true;
} else if (arr[mid] < value) {
left = mid + 1;
} else {
right = mid - 1;
}
}
return arr[left] == value;
}
private static void mergeSort(int[] arr) {
if (arr == null || arr.length <= 1) {
return;
}
arrSort(arr, 0, arr.length - 1);
}
private static void arrSort(int[] arr, int left, int right) {
if (left == right) {
return;
}
int mid = left + ((right - left) >> 1);
arrSort(arr, left, mid);
arrSort(arr, mid + 1, right);
merge(arr, left, mid, right);
}
private static void merge(int[] arr, int left, int mid, int right) {
int[] help = new int[right - left + 1];
int i = 0;
int p1 = left;
int p2 = mid + 1;
while (p1 <= mid && p2 <= right) {
help[i++] = arr[p1] <= arr[p2] ? arr[p1++] : arr[p2++];
}
while (p1 <= mid) {
help[i++] = arr[p1++];
}
while (p2 <= right) {
help[i++] = arr[p2++];
}
int len = help.length;
for (i = 0; i < len; i++) {
arr[left + i] = help[i];
}
}
}
<file_sep>write algorithm in Java from basic to advance
| 9c7ced14f26d68218ba3a934c70737637da60126 | [
"Markdown",
"Java"
] | 2 | Java | nickchen-dev/algoexpert | a009acc6601f17e4cd6feb8670522e9a708ac460 | e5390649345925cc6a55f3ee37ec2100b7cfbf1d |
refs/heads/master | <repo_name>timeconsumer/elevate<file_sep>/models/maincontroller.py
from events import Events
from models.elevator import Elevator
class MainController:
def __init__(self, elevators: list):
self.elevators = elevators
self.available_elevators = elevators
def add_elevator(self, elevator: Elevator):
self.elevators.append(elevator)
def handle_dispatch(self, requesting_floor):
print('Handling dispatch')
target_elevators = []
best_candidate: Elevator = None
for elevator in self.elevators:
if elevator.is_en_route_to_floor(requesting_floor):
target_elevators.append(elevator)
for elevator in target_elevators:
if best_candidate is None:
best_candidate = elevator
elif elevator.distance_to_floor(requesting_floor) < best_candidate.distance_to_floor(requesting_floor):
best_candidate = elevator
if best_candidate is None:
best_candidate = self.available_elevators[0]
best_candidate.add_floor_to_queue(requesting_floor)
self.available_elevators.remove(best_candidate)
print(self.available_elevators)
print(best_candidate.queued_floors)
<file_sep>/models/elevator.py
import decimal
import math
class Elevator:
def __init__(self, speed: decimal, capacity: int, home_floor: int):
"""
Creates an instance of an elevator
:param speed: The speed in feet / second of the elevator
:param capacity: The capacity in persons of the elevator
:param home_floor: The floor which this elevator calls home
"""
self.home_floor = home_floor
self.current_floor = home_floor
self.target_floor = home_floor
self.speed = speed
self.capacity = capacity
self.is_moving = False
self.last_update = 0 # TODO figure out updates
self.queued_floors = []
def is_home(self):
return self.current_floor == self.home_floor
def moving_direction(self):
if not self.is_moving:
return 'not moving'
elif self.current_floor < self.target_floor:
return 'up'
else:
return 'down'
def pick_up(self, requesting_floor: int):
self.is_moving = True
self.target_floor = requesting_floor
self.queued_floors.append(requesting_floor)
def is_en_route_to_floor(self, requesting_floor):
if requesting_floor > self.current_floor:
if self.moving_direction() == 'up':
return self.target_floor >= requesting_floor
else:
return False
if requesting_floor < self.current_floor:
if self.moving_direction() == 'down':
return self.target_floor <= requesting_floor
else:
return False
def distance_to_floor(self, requesting_floor):
return math.fabs(self.current_floor - requesting_floor)
def add_floor_to_queue(self, requesting_floor):
if requesting_floor not in self.queued_floors:
self.queued_floors.append(requesting_floor)
def update(self, ticks):
# TODO update positioning with this
return
<file_sep>/main.py
from models.elevator import Elevator
from models.maincontroller import MainController
def run():
elevator1 = Elevator(5, 5, 1)
controller = MainController([elevator1])
controller.handle_dispatch(5)
if __name__ == '__main__':
run()
| da64724948e15bcd2fa31435927e2314403c0273 | [
"Python"
] | 3 | Python | timeconsumer/elevate | e8d62f59111623a4f123401cbc6e4e18644a7237 | 1216d85927b1a042cdd81ea4b83a6801195b1d11 |
refs/heads/master | <repo_name>mmarzio67/tt4ep<file_sep>/handler.go
package main
import (
"encoding/json"
"fmt"
"net/http"
"strconv"
"time"
"github.com/gin-gonic/gin"
_ "github.com/lib/pq"
)
func rectime(w http.ResponseWriter, r *http.Request) {
queryValues := r.URL.Query()
ac, _ := strconv.Atoi(queryValues.Get("a"))
pj, _ := strconv.Atoi(queryValues.Get("p"))
ta, _ := strconv.Atoi(queryValues.Get("t"))
fmt.Printf("registration of %s done!\n", queryValues.Get("a"))
rt := time.Now()
track := Trackt{UserId: 1, ActionId: ac, ProjectId: pj, TaskId: ta, CreatedAt: rt}
track.Create()
et := tpl.ExecuteTemplate(w, "tmpl.html", track)
if et != nil {
http.Error(w, et.Error(), http.StatusInternalServerError)
return
}
}
func postime(w http.ResponseWriter, r *http.Request) {
fmt.Println("method:", r.Method) //get request method
if r.Method == "GET" {
e1 := tpl.ExecuteTemplate(w, "rectime.html", d)
if e1 != nil {
http.Error(w, e1.Error(), http.StatusInternalServerError)
return
}
} else {
// logic part of log in
pj, _ := strconv.Atoi(r.FormValue("project"))
tk, _ := strconv.Atoi(r.FormValue("task"))
at, _ := strconv.Atoi(r.FormValue("action"))
ds := r.FormValue("description")
us := 1
rt := time.Now()
fmt.Printf("Action: %d\n", at)
fmt.Printf("Project: %d\n", pj)
fmt.Printf("Task: %d\n", tk)
fmt.Printf("Description: %s\n", ds)
fmt.Printf("record time: %s\n", rt)
track := Trackt{UserId: us, ActionId: at, ProjectId: pj, TaskId: tk, Descr: ds, CreatedAt: rt}
track.Create()
//redirect to listrecs
http.Redirect(w, r, "/sel", http.StatusSeeOther)
}
}
func listRecs(w http.ResponseWriter, r *http.Request) {
//Allow CORS here By * or specific origin
w.Header().Set("Access-Control-Allow-Origin", "*")
w.Header().Set("Access-Control-Allow-Headers", "Content-Type")
//uid, _ := strconv.Atoi(queryValues.Get("u"))
uid := 1 //user=1, me
ts, _ := SelectRecs(uid)
json.NewEncoder(w).Encode(ts)
fmt.Printf("returned array of trackt=%d\n", ts)
}
func listUsers(w http.ResponseWriter, r *http.Request) {
//Allow CORS here By * or specific origin
w.Header().Set("Access-Control-Allow-Origin", "*")
w.Header().Set("Access-Control-Allow-Headers", "Content-Type")
uid := 1 //user=1, me
ns, _ := SelectUsers(uid)
fmt.Printf("returned array of trackt=%d\n", ns)
}
func getUsers(c *gin.Context) {
}
func returnAllArticles(w http.ResponseWriter, r *http.Request) {
//Allow CORS here By * or specific origin
w.Header().Set("Access-Control-Allow-Origin", "*")
w.Header().Set("Access-Control-Allow-Headers", "Content-Type")
articles := Articles{
Article{Title: "Hello", Desc: "Article Description", Content: "Article Content"},
Article{Title: "Hello 2", Desc: "Article Description", Content: "Article Content"},
}
fmt.Println("Endpoint Hit: returnAllArticles")
json.NewEncoder(w).Encode(articles)
}
<file_sep>/ui/src/components/TodoItem.js
import "./TodoItem.css";
import React from "react";
const TodoItem = ({ todo, onTodoSelect }) => {
// above: instead of props (witch contains the video object from the API)
// I just destructure and take only the video portion ({video})
return (
<div onClick={() => onTodoSelect(todo)} className="todo-item item">
<div className="content">
<div className="header">Timestamp: {todo.timestamp}</div>
<div className="header">Description: {todo.descr}</div>
<div className="header">Action: {todo.action}</div>
<div className="header">Project: {todo.project}</div>
</div>
</div>
);
};
export default TodoItem;
<file_sep>/ui/src/componentsVideo/VideoItem.js
import "./VideoItem.css";
import React from "react";
const VideoItem = ({ video, onVideoSelect }) => {
// above: instead of props (witch contains the video object from the API)
// I just destructure and take only the video portion ({video})
return (
<div onClick={() => onVideoSelect(video)} className="video-item item">
<img
alt={video.snippet.title}
className="ui image"
src={video.snippet.thumbnails.medium.url}
/>
<div className="content">
<div className="header">{video.snippet.title}</div>
</div>
</div>
);
};
export default VideoItem;
<file_sep>/main.go
package main
import (
"database/sql"
"html/template"
"net/http"
"time"
_ "github.com/lib/pq"
)
var err error
var db *sql.DB
var tpl *template.Template
var dbUsers = map[string]user{} // user ID, user
var dbSessions = map[string]session{} // session ID, session
var dbSessionsCleaned time.Time
var d DropdownItem
const sessionLength int = 30
// connect to the Db
func init() {
tpl = template.Must(template.ParseGlob("templates/*"))
dbSessionsCleaned = time.Now()
//connect to the database and manage the errors
connStr := "user=trackt dbname=trackt password=<PASSWORD> sslmode=disable"
db, err = sql.Open("postgres", connStr)
if err != nil {
panic(err)
}
projects, _ := pdProject()
tasks, _ := pdTask()
actions, _ := createDdl()
d = DropdownItem{
P: projects,
T: tasks,
A: actions,
}
}
func main() {
server := http.Server{
Addr: "127.0.0.1:10000",
}
// application logic handlers
http.HandleFunc("/rec", rectime)
http.HandleFunc("/sel", listRecs)
http.HandleFunc("/entry", postime)
http.HandleFunc("/users", listUsers)
http.HandleFunc("/articles", returnAllArticles)
// session logic handlers
http.HandleFunc("/", index)
http.HandleFunc("/bar", bar)
http.HandleFunc("/signup", signup)
http.HandleFunc("/login", login)
http.HandleFunc("/logout", logout)
http.Handle("/favicon.ico", http.NotFoundHandler())
server.ListenAndServe()
}
<file_sep>/database.go
package main
import (
"fmt"
"log"
"time"
_ "github.com/lib/pq"
)
type Trackt struct {
Id int `json:"id"`
UserId int `json:"userid"`
ActionId int `json:"actionid"`
ProjectId int `json:"projectid"`
TaskId int `json:"taskid"`
Descr string `json:"descr"`
CreatedAt time.Time `json:"createdat"`
}
type TrackRen struct {
Project string `json:"project"`
Task string `json:"task"`
Action string `json:"action"`
User string `json:"user"`
Descr string `json:"descr"`
Timestamp time.Time `json:"timestamp"`
}
type User struct {
Id int `json:"id"`
Name string `json:"name"`
Surname string `json:"surname"`
Function string `json:"function"`
}
type Project struct {
Id int
Name string `sql:"not null"`
Description string
}
type Task struct {
Id int
Name string `sql:"not null"`
Description string
}
type Action struct {
Id int
Name string
}
type user struct {
UserName string
Password []byte
First string
Last string
Role string
}
type session struct {
un string
lastActivity time.Time
}
type DropdownItem struct {
A []actions
P []project
T []task
}
type actions = map[string]interface{}
type project = map[string]interface{}
type task = map[string]interface{}
var actionslice []actions
var projectslice []project
var taskslice []task
// Article - Our struct for all articles :::: this is an example
type Article struct {
Id int `json:"id"`
Title string `json:"title"`
Desc string `json:"desc"`
Content string `json:"content"`
}
type Articles []Article
func createDdl() ([]map[string]interface{}, error) {
statement := `SELECT * FROM actions ORDER BY Name ASC`
rows, _ := db.Query(statement)
cols, _ := rows.Columns()
for rows.Next() {
// Create a slice of interface{}'s to represent each column,
// and a second slice to contain pointers to each item in the columns slice.
columns := make([]interface{}, len(cols))
columnPointers := make([]interface{}, len(cols))
for i, _ := range columns {
columnPointers[i] = &columns[i]
}
// Scan the result into the column pointers...
if err := rows.Scan(columnPointers...); err != nil {
}
// Create our map, and retrieve the value for each column from the pointers slice,
// storing it in the map with the name of the column as the key.
m := make(map[string]interface{})
for i, colName := range cols {
val := columnPointers[i].(*interface{})
m[colName] = *val
}
// Outputs: map[columnName:value columnName2:value2 columnName3:value3 ...]
fmt.Println(m)
actionslice = append(actionslice, m)
}
return actionslice, nil
}
func pdProject() ([]map[string]interface{}, error) {
statement := `SELECT * FROM project ORDER BY Name ASC`
rows, _ := db.Query(statement)
cols, _ := rows.Columns()
for rows.Next() {
// Create a slice of interface{}'s to represent each column,
// and a second slice to contain pointers to each item in the columns slice.
columns := make([]interface{}, len(cols))
columnPointers := make([]interface{}, len(cols))
for i, _ := range columns {
columnPointers[i] = &columns[i]
}
// Scan the result into the column pointers...
if err := rows.Scan(columnPointers...); err != nil {
}
// Create our map, and retrieve the value for each column from the pointers slice,
// storing it in the map with the name of the column as the key.
m := make(map[string]interface{})
for i, colName := range cols {
val := columnPointers[i].(*interface{})
m[colName] = *val
}
// Outputs: map[columnName:value columnName2:value2 columnName3:value3 ...]
fmt.Println(m)
projectslice = append(projectslice, m)
}
return projectslice, nil
}
func pdTask() ([]map[string]interface{}, error) {
statement := `SELECT * FROM task ORDER BY Name ASC`
rows, _ := db.Query(statement)
cols, _ := rows.Columns()
for rows.Next() {
// Create a slice of interface{}'s to represent each column,
// and a second slice to contain pointers to each item in the columns slice.
columns := make([]interface{}, len(cols))
columnPointers := make([]interface{}, len(cols))
for i, _ := range columns {
columnPointers[i] = &columns[i]
}
// Scan the result into the column pointers...
if err := rows.Scan(columnPointers...); err != nil {
}
// Create our map, and retrieve the value for each column from the pointers slice,
// storing it in the map with the name of the column as the key.
m := make(map[string]interface{})
for i, colName := range cols {
val := columnPointers[i].(*interface{})
m[colName] = *val
}
// Outputs: map[columnName:value columnName2:value2 columnName3:value3 ...]
fmt.Println(m)
taskslice = append(taskslice, m)
}
return taskslice, nil
}
// Create a new tracktime record
func (track *Trackt) Create() (err error) {
err = db.QueryRow("insert into trackt (usid, project_id, action_id, task_id, description, createdat) values ($1, $2, $3,$4,$5, $6) returning id", track.UserId, track.ProjectId, track.ActionId, track.TaskId, track.Descr, track.CreatedAt).Scan(&track.Id)
if err != nil {
log.Fatalf("Unable to connect to the DB: %v", err)
}
return
}
func SelectRecs(uid int) ([]*TrackRen, error) {
statement := `SELECT actions.name, project.name, task.name, users.name, trackt.description, trackt.createdat
FROM actions, project, task, users, trackt
WHERE actions.id=trackt.action_id
AND project.id=trackt.project_id
AND task.id= trackt.task_id
AND users.id= $1`
rows, err := db.Query(statement, uid)
if err != nil {
log.Fatal(err)
}
defer rows.Close()
ts := make([]*TrackRen, 0)
for rows.Next() {
t := new(TrackRen)
err := rows.Scan(&t.Action, &t.Project, &t.Task, &t.User, &t.Descr, &t.Timestamp)
if err != nil {
log.Fatal(err)
}
ts = append(ts, t)
}
for _, t := range ts {
fmt.Printf("%s, %s, %s, %s,%s\n", t.Action, t.Project, t.Task, t.User, t.Descr, t.Timestamp)
}
return ts, nil
}
func SelectUsers(uid int) ([]*User, error) {
statement := `SELECT name FROM users WHERE id=$1`
rows, err := db.Query(statement, uid)
if err != nil {
log.Fatal(err)
}
defer rows.Close()
ns := make([]*User, 0)
for rows.Next() {
n := new(User)
err := rows.Scan(&n.Name)
if err != nil {
log.Fatal(err)
}
ns = append(ns, n)
}
fmt.Println(ns)
return ns, nil
}
<file_sep>/ui/src/backup/App.js
import React, { Component } from "react";
import logo from "./logo.svg";
import "./App.css";
class App extends Component {
constructor(props) {
super(props);
this.state = {
articles: []
};
}
componentDidMount() {
let myHeaders = new Headers();
let myInit = {
method: "GET",
headers: myHeaders,
mode: "cors",
cache: "default"
};
fetch("http://localhost:10000/sel", myInit)
.then(res => res.json()) //response type
.then(data => {
console.log(data); //log the data
this.setState({ articles: data });
});
}
render() {
return (
<div className="App">
<header className="App-header">
<img src={logo} className="App-logo" alt="logo" />
<h1 className="App-title">Fetch Example</h1>
</header>
{this.state.articles.map(article => {
return (
<ul key={article.descr}>
<li>{article.project}</li>
<li>{article.task}</li>
</ul>
);
})}
</div>
);
}
}
export default App;
<file_sep>/db/setup.sql
drop table trackt cascade;
drop table users;
drop table project;
drop table task;
drop table actions;
create table users (
id serial primary key,
name varchar(32),
surname varchar(32),
func varchar(32)
);
create table project (
id serial primary key,
name varchar(32),
description varchar(64)
);
create table task (
id serial primary key,
name varchar(32),
description varchar(64)
);
create table actions (
id serial primary key,
name varchar(16)
);
create table trackt (
id serial primary key,
usid integer references users(id),
action_id integer references actions(id),
project_id integer references project(id),
task_id integer references task(id),
createdAt timestamp
); | ce833b0904ab671c7253c18ab47a9ff71207eada | [
"JavaScript",
"SQL",
"Go"
] | 7 | Go | mmarzio67/tt4ep | 03eef7bd1629002b39237a39b14941e1cda78a81 | c1495764498354ccd57c06df60cbfb9a42a42f98 |
refs/heads/master | <repo_name>angus-ai/angus-gateway<file_sep>/README.rst
=====================
Angus Gateway Service
=====================
Configuration
=============
* PORT (default 8080): listen port
* SERVICE_FILE (default /etc/angus-gateway/services.json): the service
directory
Mono service option
+++++++++++++++++++
* SERVICE_DIR (default None): service directory in json format
* SERVICE_NAME (default None): the service name
* SERVICE_VERSION (default 1): the service version
* SERVICE_URL (default None): the service url
Releases
========
Angus Gateway Service - Release Note 1.0.0
++++++++++++++++++++++++++++++++++++++++++
Features added
--------------
* Add blob delete action
Fixes
------
* Do not try to convert to json when it is a binary resource.
Angus Gateway Service - Release Note 0.0.2
++++++++++++++++++++++++++++++++++++++++++
Features added
--------------
* Microservice architecture with docker (Dockerfile)
* Define a service name, version and url for atomic run
Update
------
* Default service file location
* Move to default port 80
Angus Gateway Service - Release Note 0.0.1
++++++++++++++++++++++++++++++++++++++++++
Features added
--------------
* Primitive (in-memory) blob storage
* HTPASSWD access management (angus-access)
* Service repository
<file_sep>/Dockerfile
FROM 773153459320.dkr.ecr.eu-west-1.amazonaws.com/angus.box/genericservice
#
# Service
#
COPY angus /angus
#
# Entrypoint
#
COPY docker-entrypoint.sh /
RUN chmod +x /docker-entrypoint.sh
<file_sep>/docker-entrypoint.sh
#!/bin/bash
python /angus/services/gateway.py
<file_sep>/angus/services/gateway.py
#!/usr/bin/env python
# -*- coding: utf-8 -*-
# Licensed to the Apache Software Foundation (ASF) under one
# or more contributor license agreements. See the NOTICE file
# distributed with this work for additional information
# regarding copyright ownership. The ASF licenses this file
# to you under the Apache License, Version 2.0 (the
# "License"); you may not use this file except in compliance
# with the License. You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing,
# software distributed under the License is distributed on an
# "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY
# KIND, either express or implied. See the License for the
# specific language governing permissions and limitations
# under the License.
""" This is the gateway for all services.
Right now, it manages blob storage and service repository.
"""
import collections
import json
import logging
import os
import uuid
import tornado.ioloop
import tornado.web
import angus.analytics
LOGGER = logging.getLogger(__name__)
__updated__ = "2018-07-18"
__author__ = "<NAME>"
__copyright__ = "Copyright 2015-2018, Angus.ai"
__credits__ = ["<NAME>", "<NAME>"]
__status__ = "Production"
class Storage(object):
""" Storage class for blobs
"""
def __init__(self, size=10):
self.size = size
self.inner = collections.OrderedDict()
def store(self, key, content, meta=None):
""" Store a new blob
"""
self.inner[key] = (content, meta)
if len(self.inner) > self.size:
self.inner.popitem(last=False)
def remove(self, key):
del self.inner[key]
def get(self, key):
""" Get back a blob
"""
return self.inner.get(key)
def iteritems(self):
""" Iterator over content
"""
return self.inner.iteritems()
class Services(tornado.web.RequestHandler):
""" Handler that present all server services
"""
def initialize(self, service_map):
"""
List all avaialble service based on a service map:
{ (service_name, service_version) => service_url }
"""
tornado.web.RequestHandler.initialize(self)
self.service_map = service_map
@angus.analytics.report
def get(self):
response = dict()
for service in self.service_map.keys():
response[service] = {
'url': "/services/%s" % (service),
}
response = dict(services=response)
self.write(json.dumps(response))
class Service(tornado.web.RequestHandler):
""" Handler for service
"""
def initialize(self, service_map):
"""
Create a handler on a dictionary versions
{ (service_name, service_version) => service_url }
"""
tornado.web.RequestHandler.initialize(self)
self.service_map = service_map
@angus.analytics.report
def get(self, service):
if service not in self.service_map:
self.set_status(404, "Unknown service %s" % (service))
return
response = dict()
for version, href in self.service_map[service].iteritems():
response[version] = dict(url=href)
response = dict(versions=response)
self.write(json.dumps(response))
class Versions(tornado.web.RequestHandler):
""" Handler for service versions
"""
def initialize(self, service_map):
"""
Create a handler on a dictionary versions
{ (service_name, service_version) => service_url }
"""
tornado.web.RequestHandler.initialize(self)
self.service_map = service_map
@angus.analytics.report
def get(self, service, version):
# get a description
response = {
'description': 'Not available',
}
self.write(json.dumps(response))
class BlobStorage(tornado.web.RequestHandler):
""" Handler for blob creation
"""
def initialize(self, storage):
self.storage = storage
@angus.analytics.report
def post(self):
status = 201
self.set_status(status, "Blob created")
data = self.request.body_arguments['meta'][0]
data = json.loads(data)
file_url = data['content']
content = self.request.files[file_url][0]['body']
uid = unicode(uuid.uuid1())
user = angus.framework.extract_user(self)
self.storage.store(uid, content, dict(owner=user))
public_url = "%s://%s" % (self.request.protocol, self.request.host)
response = {
'status': status,
'url': "%s/blobs/%s" % (public_url, uid),
}
self.write(json.dumps(response))
class Blob(tornado.web.RequestHandler):
""" Handler for a blob
"""
def initialize(self, storage):
self.storage = storage
def delete(self, uid):
try:
self.storage.remove(uid)
except KeyError:
self.set_status(404, "Blob resource %s not found" % (uid))
@angus.analytics.report
def get(self, uid):
user = angus.framework.extract_user(self)
res = self.storage.get(uid)
if res is not None:
content, meta = res
if meta['owner'] == user:
self.write(content)
self.set_status(200)
self.flush()
else:
self.set_status(403, "Not owner")
else:
self.set_status(404, "Blob resource %s not found" % (uid))
def find_services():
""" Retrive service description
"""
service_dir = os.environ.get('SERVICE_DIR', None)
service_file = os.environ.get('SERVICE_FILE', '/etc/angus-gateway/services.json')
service_name = os.environ.get('SERVICE_NAME', None)
service_url = os.environ.get('SERVICE_URL', None)
service_version = os.environ.get('SERVICE_VERSION', 1)
if service_dir is not None:
services = json.loads(service_dir)
elif service_name is not None and service_url is not None:
services = dict()
services[service_name] = dict()
services[service_name][service_version] = service_url
elif service_file is not None and os.path.isfile(service_file):
with open(service_file, 'r') as sfile:
services = json.loads(sfile.read())
else:
services = dict() # No available service
return services
def main():
""" Run the service
"""
logging.basicConfig(level=logging.INFO)
home = os.path.dirname(os.path.dirname(os.path.realpath(__file__)))
port = os.environ.get('PORT', 8080)
services = find_services()
msg = "Available services:\n" + json.dumps(services, indent=4)
LOGGER.info(msg)
storage = Storage(size=300)
application = tornado.web.Application([
(r"/services/(?P<service>.*)/(?P<version>.*)",
Versions,
dict(service_map=services)),
(r"/services/(?P<service>.*)", Service, dict(service_map=services)),
(r"/services", Services, dict(service_map=services)),
(r"/blobs/(.*)", Blob, dict(storage=storage)),
(r"/blobs", BlobStorage, dict(storage=storage)),
(r"/(.*)",
tornado.web.StaticFileHandler,
{'path': "%s/static/public/" % (home)}),
])
application.listen(port, xheaders=True)
tornado.ioloop.IOLoop.instance().start()
if __name__ == "__main__":
main()
| c810cc6fe7385f4f42741b7599f63d1c7ac9f38f | [
"Python",
"Dockerfile",
"reStructuredText",
"Shell"
] | 4 | reStructuredText | angus-ai/angus-gateway | 94a1067c776df239921cfb5aeff126d1a0ce6423 | 3f183dbf68c8944ec301e3d4d19adf6b758890b0 |
refs/heads/master | <repo_name>intoyuniot/CGQ01<file_sep>/src/CGQ01.cpp
/*
******************************************************************************
This library is free software; you can redistribute it and/or
modify it under the terms of the GNU Lesser General Public
License as published by the Free Software Foundation, either
version 3 of the License, or (at your option) any later version.
This library is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU
Lesser General Public License for more details.
You should have received a copy of the GNU Lesser General Public
License along with this library; if not, see <http://www.gnu.org/licenses/>.
This library is CGQ01 formaldehyde sensor module
******************************************************************************
*/
#include "CGQ01.h"
CGQ01::CGQ01()
{
}
char CGQ01::begin()
{
Serial.begin(9600);
while(!Serial)
{
}
// 校零协议
Serial.write(0XA5);
Serial.write(0X5A);
Serial.write(0X02);
Serial.write(0X85);
Serial.write(0XAA);
counter = 0;
sign = 0;
}
double CGQ01::getConcentrationPPM()
{
unsigned char i=0,sum=0;
Serial.flush();
Serial.write(0XA5); //获取甲醛浓度信息数据,读取协议模式
Serial.write(0X5A);
Serial.write(0X02);
Serial.write(0X80);
Serial.write(0XAA);
delay(50);
while (Serial.available())
{
Re_buf[counter]=(unsigned char)Serial.read();
if(counter==1 && Re_buf[0]!=0xA5 && Re_buf[1]!=0x5A)
{
counter = 0;
return -1; //检查帧头
}
if(counter==8&&Re_buf[8]!=0xAA)
{
counter = 0;
return -1; //检查帧尾
}
counter++;
if(counter==9) //接收到数据
{
counter=0; //重新赋值,准备下一帧数据的接收
sign=1;
}
}
if(sign)
{
sign=0;
ppm = (float)(Re_buf[6]<<8|Re_buf[7])/100; // 甲醛 ppm
return ppm;
}
return -1;
}
double CGQ01::getConcentrationMg()
{
unsigned char i=0,sum=0;
Serial.flush();
Serial.write(0XA5); //获取甲醛浓度信息数据,读取协议模式
Serial.write(0X5A);
Serial.write(0X02);
Serial.write(0X80);
Serial.write(0XAA);
delay(50);
while (Serial.available())
{
Re_buf[counter]=(unsigned char)Serial.read();
if(counter==1 && Re_buf[0]!=0xA5 && Re_buf[1]!=0x5A)
{
counter = 0;
return -1; //检查帧头
}
if(counter==8&&Re_buf[8]!=0xAA)
{
counter = 0;
return -1; //检查帧尾
}
counter++;
if(counter==9) //接收到数据
{
counter=0; //重新赋值,准备下一帧数据的接收
sign=1;
}
}
if(sign)
{
sign=0;
mg = (float)(Re_buf[4]<<8|Re_buf[5])/100; // 甲醛 mg
return ppm;
}
return -1;
}
<file_sep>/examples/CGQ01_read/CGQ01_read.ino
// 本程序使用CGQ01甲醛模块,获取甲醛浓度,并重串口输出数据
// 本程序使用的是Serial
// 连接说明:
// CGQ01 Atom
// 1.VCC +5.0V
// 2.GND GND
// 3.RX A3(TX)
// 4.TX A2(RX)
/*
测量范围:0-5ppm
输出信号:2000nA/pp
甲醛模块上电后需1分钟预热时间,1分钟后测的数据为有效数据。
*/
#include <CGQ01.h>
CGQ01 cgq01PPM; // 定义传感器变量
void setup()
{
Serial.begin(115200);
cgq01PPM.begin(); // 初始化
}
void loop()
{
double ppm = cgq01PPM.getConcentrationPPM(); // 获取甲醛传感器的甲醛浓度
if (ppm != -1)
{
Serial.print(ppm); // 串口输出甲醛浓度数据
}
delay(2000); // 延迟2s
}
<file_sep>/library.properties
name=CGQ01
version=1.0.0
author=robin <<EMAIL>>
maintainer=robin <<EMAIL>>
sentence=CGQ01甲醛传感器库
paragraph=Like this project? Please star it on GitHub!
category=hcho sensor
url=www.intoyun.com
architectures=*
| b71d7fa27c41d87a20da1a2e390b09dea3da85af | [
"C++",
"INI"
] | 3 | C++ | intoyuniot/CGQ01 | 564006ae1e436ed361b1471b07a566cec07ce413 | 832650acab1b545a00b8f65da48ae9513aa4858d |
refs/heads/master | <repo_name>josephxwf/Bookstore-Flask<file_sep>/coreapp/__init__.py
# __init__.py
from flask import Flask
app = Flask('coreapp')
import views
<file_sep>/main.py
# used to local test your code by command python main.py
from flask import Flask
from coreapp import app
app.debug = True
if __name__ == '__main__':
app.run()<file_sep>/README.md
# Bookstore-Flask
Simple online bookstore that was developed with Flask.
## Prerequisites
* Pyhton 2.7
## Installation
* Install pip
* `easy_install pip`
* Install virtual environment
* ` pip install --user virtualenv`
* Create Virtualenv
* `virtualenv ENV`
* Activtate Virtualenv
* `source /path/to/ENV/bin/activate`
* install flask
* `pip install Flask`
## Development set up and run the app
1. Method 1 to run flask app
* cd to root directory
* `python main.py`
* Now head over to http://127.0.0.1:5000/, and you should see the online bookstore
2. Method 2 to run flask app
* cd to root directory
* ` export FLASK_APP=my_application`
* `export FLASK_ENV=development`
* `flask run`
## Issues
1. oserror: [errno 98] address already in use flask:
* `ps -fA | grep python`
* You will get a pid number by naming of your flask number. Now copy the pid number from second column of your project row of terminal output.
* `kill -9 pid` The terminal will restart and then run the command below
* `flask run`
## Authors
* **<NAME>** - *init work* - [josephxwf](https://github.com/josephxwf)
<file_sep>/coreapp/views.py
from flask import render_template
from coreapp import app
from flask import url_for # used for redirect to another page see home.html
import re
from bs4 import BeautifulSoup
#from sortedcontainers import SortedDict
def get_chap_links(page):
soup = BeautifulSoup(page)
links = [str(link.get('name')) for link in soup.find_all('a') if not link.get('href')]# last if condition filters only links that not begin with href
return links
def parse_wizofoz(bookname):
page = render_template(bookname) # render back to web page
links = get_chap_links(page) # use the function above to extract link in the wizofoz.html page
sections = []
# get each section from wizofoz.html and put into sections dictionary.
for ind in range(len(links)):
section = {}
start = links[ind]
if ind < len(links) - 1:
end = links[ind + 1]
# pattern for getting each section by extracting content between two sections
patt = ('<a name="{}"(.*)' + '<a name="{}"').format(start,end)
match = re.search(patt, page, re.MULTILINE | re.DOTALL)
else:
patt = '<a name="{}"(.*)'.format(start)
match = re.search(patt, page, re.MULTILINE | re.DOTALL)
if match:
soup = BeautifulSoup(match.group(1))
plist = [p.contents[0] for p in soup.find_all('p') if p.contents]
#section['title'] = soup.find('h3').contents[0]
section['title'] = start
section['order'] = ind # order and label each dictionary
section['plist'] = plist
sections.append(section)
return links, sections
@app.route('/')
def home():
data = {'image_url1':'/static/wonderfulWizard.jpg','image_url2':'/static/lostPrincess.jpg'}
#data['anchor'] = SortedDict(parse_wizofoz()[1])
data['anchor'] = parse_wizofoz('wizofoz.html')[1]
#page = render_template('wizofoz.html')
#data1 = {'links':get_chap_links(page), 'y':parse_wizofoz()}
return render_template('home.html', **data)
#return render_template('home.html', **data1)
@app.route('/ebook1')
def ebook1():
data = {'title':'The Wonderful Wizard of Oz','image_url':'/static/wonderfulWizard.jpg'}
#data['anchor'] = SortedDict(parse_wizofoz()[1])
data['anchor'] = parse_wizofoz('wizofoz.html')[1]
data['source']= 'wizofoz.html'
#page = render_template('wizofoz.html')
#data1 = {'links':get_chap_links(page), 'y':parse_wizofoz()}
return render_template('ebook1.html', **data)
@app.route('/ebook2')
def ebook2():
data = {'title':'The Lost Princess of Oz','image_url':'/static/lostPrincess.jpg'}
#data['anchor'] = SortedDict(parse_wizofoz()[1])
data['anchor'] = parse_wizofoz('lostprincess.html')[1]
#page = render_template('wizofoz.html')
#data1 = {'links':get_chap_links(page), 'y':parse_wizofoz()}
data['source']= 'lostprincess.html'
return render_template('ebook2.html', **data)
# dreirect depends on the section num and the name of book in html .
@app.route('/<int:num><source>') # source identify the book name, num indentify the chapter
def section(num,source):
data = {'title':'The Wonderful Wizard of Oz','image_url':'/static/title_img.jpg'}
data['anchor'] = parse_wizofoz(source)[1]
data['dic'] = parse_wizofoz(source)[1][num]#get the specific dictionary that contains that chapter label by num
data['source']= source
return render_template('section.html',**data)
| 9b04f7ab63f40c1cfa62638edbaad83d4a46f5bd | [
"Markdown",
"Python"
] | 4 | Python | josephxwf/Bookstore-Flask | 08bdc03ef6710cba3c303bb244d93517b873e673 | 2be7d790acc98410d75d00299d7797033a9987e3 |
refs/heads/master | <repo_name>saranvananvasu/ReactDemo<file_sep>/src/Login.js
import React from 'react'
import Button from '@material-ui/core/Button'
import TextField from '@material-ui/core/TextField'
import classNames from 'classnames';
import {Router,Route,Link, Redirect, withRouter } from 'react-router-dom'
class Login extends React.Component {
constructor(props) {
super(props);
this.handleLogin = this.handleLogin.bind(this);
this.handleSignUp = this.handleSignUp.bind(this);
this.onusernameChange = this.onusernameChange.bind(this);
this.onpasswordChange = this.onpasswordChange.bind(this);
this.state = {
username: '',
password: '',
loggedin: 0,
}
}
handleLogin() {
this.setState({username: 'sddd'});
this.setState({loggedin: 1});
}
handleSignUp() {
}
onusernameChange (event) {
this.setState({username: event.target.value});
console.log(this.state.username);
}
onpasswordChange (event) {
this.setState({password: event.target.value});
console.log(this.state.password);
}
render () {
if (this.state.loggedin === 1) return (
<Redirect to={{pathname: "/SignUp",}}/>
);
return (
<div class='center'>
<TextField id='usn'
label = 'Username'
margin="normal"
onChange = {this.onusernameChange}
/>
<br/><br/>
<TextField
id='usn'
label = 'Password'
onChange = {this.onpasswordChange}
/>
<br/><br/>
<Button onClick={this.handleLogin}>Login</Button>
<Button component={Link} to={"/Signup"}>Sign up</Button>
<label> {this.state.username} </label>
<label> {this.state.password} </label>
</div>
);
}
}
export default Login; | e16ad5c39b84be23ff62b4d10b6d28f31745c6da | [
"JavaScript"
] | 1 | JavaScript | saranvananvasu/ReactDemo | 7aa2bd66880bb76bf2e0f8b1bdbf3bce21111ae7 | 2c9be55c104bcce46d14ee24c576467e59b7c5d7 |
refs/heads/main | <repo_name>CadmiumC4/mal-linguist<file_sep>/baskasey.rs
use std;
| 7cf960696747aee050ea68380d38589812bdc30d | [
"Rust"
] | 1 | Rust | CadmiumC4/mal-linguist | 3dfc3c16afa14fa2532997f534a5ef7cfc6a54c0 | f0de99961572844215124943b0317adb8253642e |
refs/heads/master | <repo_name>YousufQadri/Campus-Recruitment-System-react-client<file_sep>/src/Components/Dashboard/CompanyDashboard.js
import React, { Component } from "react";
import { getJWT } from "../../helpers/jwt";
import axios from "axios";
import JobsView from "../Display/JobsView";
import CompanyProfileView from "../Display/CompanyProfileView";
import StudentsView from "../Display/StudentsView";
import JobApplicantView from "../Display/JobAppicantView";
import CreateModal from "../CreateModal";
import { connect } from "react-redux";
import { loadUserCompany } from "../../store/Actions/authActions";
class CompanyDashboard extends Component {
state = {
students: [],
companyJobs: [],
applicants: [],
profile: [],
selectedMenu: "Profile",
isLoading: false
};
componentDidMount() {
this.props.loadUserCompany(this.props.history);
const jwt = getJWT();
if (jwt) {
this.setState({ isLoading: true });
// Fetch companies API
axios
.get("http://localhost:5000/api/v1/company/get-profile/", {
headers: {
"x-auth-token": `${jwt}`
}
})
.then(res => {
console.log("Company: ", res.data.company[0]);
this.setState({ profile: res.data.company[0] });
})
.catch(error => console.log("Error: ", error.response.data));
this.fetchData();
this.setState({ isLoading: false });
}
}
fetchData = () => {
const jwt = getJWT();
if (jwt) {
axios
.get("http://localhost:5000/api/v1/company/get-data/", {
headers: {
"x-auth-token": `${jwt}`
}
})
.then(res => {
console.log("Companies: ", res.data);
this.setState({
students: res.data.allStudents,
companyJobs: res.data.companyJobs,
applicants: res.data.applicants
});
})
.catch(error => console.log("Error: ", error.response.data));
}
};
render() {
if (this.state.isLoading) {
return <p>Loading...</p>;
}
return (
<React.Fragment>
<h1 className="mt-5 font-weight-bold">Company Dashboard</h1>
<div className="container mt-5">
<div className="row">
<div className="col-sm-12 col-md-3">
<ul className="list-group">
{/* Options Menu */}
<h3 className="text-center list-group-item list-group-item-secondary">
Menu
</h3>
<li
className="list-group-item d-flex justify-content-between list-group-item-action active`"
onClick={() => this.setState({ selectedMenu: "Profile" })}
>
Company Profile
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "Students" });
}}
// this.setState({ selectedMenu: "Companies" })
>
Students
<span className="badge badge-primary badge-pill">
{this.state.students.length}
</span>
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "Jobs" });
}}
>
Current Jobs
<span className="badge badge-primary badge-pill">
{this.state.companyJobs.length}
</span>
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "Applicants" });
}}
>
Applicants
<span className="badge badge-primary badge-pill">
{this.state.applicants.length}
</span>
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "CreateJob" });
}}
>
Create Job Post
</li>
</ul>
</div>
{/* Loader or content */}
{/* {!this.state.companies || !this.state.jobs ? (
<div>Loading...</div>
) : ( */}
<div className="col-sm-12 col-md-9">
<div className="row">
{this.state.selectedMenu === "Profile" && (
<CompanyProfileView profile={this.state.profile} />
)}
{this.state.selectedMenu === "Students" &&
this.state.students.map(student => (
<StudentsView key={student._id} student={student} />
))}
{this.state.selectedMenu === "Jobs" &&
this.state.companyJobs.map(job => (
<JobsView key={job._id} job={job} applyModal={false} />
))}
{this.state.selectedMenu === "Applicants" &&
(this.state.applicants.length === 1 ? (
<JobApplicantView
key={this.state.applicants._id}
applicant={this.state.applicants[0]}
applyModal={false}
/>
) : (
this.state.applicants.map(applicant => (
<JobApplicantView
key={applicant._id}
applicant={applicant}
/>
))
))}
{this.state.selectedMenu === "CreateJob" && (
<CreateModal profile={this.state.profile} />
)}
</div>
</div>
</div>
</div>
</React.Fragment>
);
}
}
const mapStateToProps = state => ({
auth: state.auth
});
export default connect(
mapStateToProps,
{ loadUserCompany }
)(CompanyDashboard);
<file_sep>/src/Components/Register.js
import React, { Component } from "react";
import axios from "axios";
import {
TabContent,
TabPane,
Nav,
NavItem,
NavLink,
Form,
FormGroup,
Label,
Input,
Button,
Alert
} from "reactstrap";
import Swal from "sweetalert2";
class Register extends Component {
state = {
studentName: "",
companyName: "",
email: "",
password: "",
description: "",
contactNo: undefined,
website: "",
qualification: "",
cgpa: undefined,
message: "",
flag: false,
activeTab: 1
};
// componentDidUpdate() {
// setTimeout(() => this.setState({ message: "" }), 10000);
// }
onChange = e => {
this.setState({
[e.target.name]: e.target.value
});
};
toggle = tab => {
if (this.state.activeTab !== tab)
this.setState({
activeTab: tab,
studentName: "",
companyName: "",
email: "",
password: "",
description: "",
contactNo: undefined,
website: "",
qualification: "",
cgpa: undefined
});
};
studentFormSubmit = e => {
e.preventDefault();
let { studentName, email, password, qualification, cgpa } = this.state;
if (studentName && email && password && qualification && cgpa) {
axios
.post("http://localhost:5000/api/v1/student/register", {
studentName,
email,
password,
qualification,
cgpa
})
.then(res => {
console.log(res.data);
this.setState({
message: res.data.message,
flag: true,
studentName: "",
email: "",
password: "",
qualification: "",
cgpa: undefined
});
Swal.fire({
icon: "success",
title: "Registered successfully!",
text: res.data.message
});
this.props.history.push("/");
})
.catch(error => {
this.setState({ message: error.response.data.message, flag: false });
console.log("Error: ", error.response.data.message);
});
} else {
console.log("Fill all fields");
}
};
companyFormSubmit = e => {
e.preventDefault();
let {
companyName,
email,
password,
description,
contactNo,
website
} = this.state;
if (
companyName &&
email &&
password &&
description &&
contactNo &&
website
) {
axios
.post("http://localhost:5000/api/v1/company/register", {
companyName,
email,
password,
description,
contactNo,
website
})
.then(res => {
console.log(res.data);
this.setState({
message: res.data.message,
flag: true,
companyName: "",
email: "",
password: "",
description: "",
contactNo: undefined,
website: ""
});
Swal.fire({
icon: "success",
title: "Registered successfully!",
text: res.data.message
});
this.props.history.push("/");
})
.catch(error => {
this.setState({ message: error.response.data.message, flag: false });
console.log("Error: ", error.response.data.message);
});
} else {
console.log("Fill all fields");
}
};
render() {
return (
<div className="container">
<h1 className="my-5">Registration Form</h1>
{this.state.message ? (
<Alert
className="mb-4"
color={this.state.flag ? "success" : "danger"}
>
{this.state.message}
</Alert>
) : null}
<Nav tabs fill className="mb-3">
<NavItem>
<NavLink
onClick={() => {
this.toggle("1");
}}
>
Student
</NavLink>
</NavItem>
<NavItem>
<NavLink
onClick={() => {
this.toggle("2");
}}
>
Company
</NavLink>
</NavItem>
</Nav>
<TabContent activeTab={this.state.activeTab}>
<TabPane tabId="1">
<Form onSubmit={e => this.studentFormSubmit(e)}>
<h4>Student Registration</h4>
<FormGroup>
<Label>Full Name</Label>
<Input
type="text"
name="studentName"
placeholder="Enter your full name"
value={this.state.studentName}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Email</Label>
<Input
type="email"
name="email"
placeholder="Enter your email"
value={this.state.email}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Password</Label>
<Input
type="<PASSWORD>"
name="password"
placeholder="Enter your password"
value={this.state.password}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Qualification</Label>
<Input
type="text"
name="qualification"
placeholder="Enter your qualification"
value={this.state.qualification}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>CGPA</Label>
<Input
type="number"
name="cgpa"
placeholder="Enter your cgpa"
value={this.state.cgpa}
onChange={this.onChange}
required
/>
</FormGroup>
<Button
type="submit"
color="primary"
block
onClick={this.studentFormSubmit}
>
Submit
</Button>
</Form>
</TabPane>
<TabPane tabId="2">
<Form onSubmit={this.companyFormSubmit}>
<h4>Company Registration</h4>
<FormGroup>
<Label>Company Name</Label>
<Input
type="text"
name="companyName"
placeholder="Enter Company name"
value={this.state.companyName}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Email</Label>
<Input
type="email"
name="email"
placeholder="Enter your email"
value={this.state.email}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Password</Label>
<Input
type="<PASSWORD>"
name="password"
placeholder="Enter your password"
value={this.state.password}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Description</Label>
<Input
type="text"
name="description"
placeholder="Enter Company description"
value={this.state.description}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Contact Number</Label>
<Input
type="number"
name="contactNo"
placeholder="Enter Contact no."
value={this.state.contactNo}
onChange={this.onChange}
required
/>
</FormGroup>
<FormGroup>
<Label>Website</Label>
<Input
type="text"
name="website"
placeholder="Enter Website URL "
value={this.state.website}
onChange={this.onChange}
required
/>
</FormGroup>
<Button
type="submit"
color="primary"
block
onClick={this.companyFormSubmit}
>
Submit
</Button>
</Form>
</TabPane>
</TabContent>
</div>
);
}
}
export default Register;
<file_sep>/src/Components/CreateModal.js
import React from "react";
import { getJWT } from "../helpers/jwt";
import axios from "axios";
import Swal from "sweetalert2";
class CreateModal extends React.Component {
state = {
jobTitle: "",
description: ""
};
onChange = e => {
this.setState({
[e.target.name]: e.target.value
});
};
createJob = e => {
e.preventDefault();
const jwt = getJWT();
let data = {
jobTitle: this.state.jobTitle,
description: this.state.description
};
if (jwt) {
axios
.post(`http://localhost:5000/api/v1/job/create-job/`, data, {
headers: {
"x-auth-token": `${jwt}`
}
})
.then(res => {
console.log("new job: ", res.data);
Swal.fire({
icon: "success",
title: "Created successfull",
text: res.data.message
});
this.setState({ title: "", description: "" });
})
.catch(error => {
console.log("Error: ", error.response.data);
Swal.fire({
icon: "error",
title: "Oops... Something went wrong",
text: error.response.data.message
});
this.setState({ title: "", description: "" });
});
} else {
console.log("no token found");
}
};
render() {
return (
<div>
<form
style={{ width: "100%", marginLeft: "100px" }}
onSubmit={this.createJob}
>
<div className="form-group">
<label>Job Title</label>
<input
type="text"
className="form-control"
name="jobTitle"
value={this.state.jobTitle}
onChange={this.onChange}
placeholder="Enter job title"
required
/>
</div>
<div className="form-group">
<label>Description</label>
<input
type="text"
className="form-control"
name="description"
value={this.state.description}
onChange={this.onChange}
placeholder="Enter job description"
required
/>
</div>
<button
type="submit"
className="btn btn-primary"
onClick={this.createJob}
>
Submit
</button>
</form>
</div>
);
}
}
export default CreateModal;
<file_sep>/src/Components/Display/CompanyProfileView.js
import React from "react";
const CompanyProfileView = ({ profile }) => {
return (
<div className="col-sm-6 col-md-12 col-lg-12">
<div className="card mb-3 border-primary">
<div className="card-header">
<h3>{profile.companyName}</h3>
</div>
<div className="card-body">
<h5 className="card-title">
<strong>Description:</strong> {profile.description}
</h5>
<p className="card-text">{/* <strong>CGPA:</strong> {profile} */}</p>
<div className="card-text">
<h4>
<strong>Contact info:</strong>
</h4>
<div className="row">
<div className="col-sm-4">
<strong>Email:</strong> {profile.email}
</div>
<div className="col-sm-4">
<strong>Contact:</strong> {profile.contactNo}
</div>
<div className="col-sm-4">
<strong>Website:</strong> {profile.website}
</div>
</div>
<br />
</div>
</div>
</div>
</div>
);
};
export default CompanyProfileView;
<file_sep>/src/Components/Display/AppliedJobsView.js
import React from "react";
import ApplyModal from "../ApplyModal";
const AppliedJobsView = ({ job }) => {
console.log("Job", job);
return (
<div className="col-sm-6 col-md-4 col-lg-4">
<div
className="card border-info mb-3 text-info"
style={{ maxWidth: "18rem" }}
>
<div className="card-header">
<h4>Title: {job.jobId.jobTitle}</h4>
</div>
<div className="card-body text-dark">
<h5 className="card-title">
<strong>Company: {job.companyId.companyName}</strong>
{/* {job.companyId.companyName} */}
</h5>
<h5 className="card-text">
<strong>Job description: </strong>
{job.jobId.description}
</h5>
</div>
</div>
</div>
);
};
export default AppliedJobsView;
<file_sep>/src/Components/Display/StudentsView.js
import React from "react";
import user from "../../assets/img/user.png";
const StudentsView = ({ student }) => {
return (
<div className="col-sm-12 col-md-6 col-lg-4">
<div className="card bg-light mb-3 border-info">
<div className="card-header">
<img src={user} alt="" width={190} />
<h3>{student.studentName}</h3>
</div>
<div className="card-body">
<h5 className="card-title">
<strong>Qualification:</strong> {student.qualification}
</h5>
<p className="card-text">
<strong>CGPA:</strong> {student.cgpa}
</p>
<p className="card-text">
<strong>Email:</strong> {student.email}
</p>
</div>
</div>
</div>
);
};
export default StudentsView;
<file_sep>/src/Components/Display/StudentProfileView.js
import React from "react";
import user from "../../assets/img/user.png";
const StudentProfileView = ({ profile }) => {
return (
<div className="col-sm-6 col-md-12 col-lg-12">
<div className="card mb-3 border-primary">
<div className="card-header">
<img src={user} alt="" width={170} />
<h3>{profile.studentName}</h3>
</div>
<div className="card-body">
<h5 className="card-title">
<strong>Qualification:</strong> {profile.qualification}
</h5>
<p className="card-text">
<strong>CGPA:</strong> {profile.cgpa}
</p>
<p className="card-text">
<strong>Email:</strong> {profile.email}
</p>
</div>
</div>
</div>
);
};
export default StudentProfileView;
<file_sep>/src/store/Actions/authActions.js
import axios from "axios";
import {
CLEAR_ERRORS,
USER_LOADING,
USER_LOADED,
LOGIN_SUCCESS,
LOGIN_FAIL,
LOGOUT_SUCCESS,
REGISTER_SUCCESS,
REGISTER_FAIL,
AUTH_ERROR
} from "./types";
import { returnErrors } from "./errorActions";
// Check student token & load user
export const loadUserStudent = () => (dispatch, getState) => {
// User loading
dispatch({ type: USER_LOADING });
const token = getState().auth.token;
axios
.get("http://localhost:5000/api/v1/student/auth", {
headers: {
"x-auth-token": token
}
})
.then(res =>
dispatch({
type: USER_LOADED,
payload: res.data
})
)
.catch(err => {
dispatch(returnErrors(err.response.data, err.response.status));
dispatch({
type: AUTH_ERROR
});
});
};
// Check company token & load user
export const loadUserCompany = () => (dispatch, getState) => {
// User loading
dispatch({ type: USER_LOADING });
const token = getState().auth.token;
axios
.get("http://localhost:5000/api/v1/company/auth", {
headers: {
"x-auth-token": token
}
})
.then(res =>
dispatch({
type: USER_LOADED,
payload: res.data
})
)
.catch(err => {
dispatch(returnErrors(err.response.data, err.response.status));
dispatch({
type: AUTH_ERROR
});
});
};
// Check admin token & load user
export const loadUserAdmin = () => (dispatch, getState) => {
// User loading
dispatch({ type: USER_LOADING });
const token = getState().auth.token;
axios
.get("http://localhost:5000/api/v1/admin/auth", {
headers: {
"x-auth-token": token
}
})
.then(res =>
dispatch({
type: USER_LOADED,
payload: res.data
})
)
.catch(err => {
dispatch(returnErrors(err.response.data, err.response.status));
dispatch({
type: AUTH_ERROR
});
});
};
// Student Register
// export const studentRegister = (data, history) => dispatch => {
// dispatch({ type: USER_LOADING });
// // Request body
// // const body = { name, email, password, qualification, cgpa };
// axios
// .post("http://localhost:5000/api/v1/student/register", data)
// .then(res => {
// console.log(res.data);
// // dispatch({ type: CLEAR_ERRORS });
// dispatch({ type: REGISTER_SUCCESS });
// history.push("/");
// })
// .catch(error => {
// dispatch(
// returnErrors(error.response.data.message, error.response.data.success)
// );
// dispatch({ type: REGISTER_FAIL });
// console.log("Error: ", error.response);
// });
// // axios
// // .post("/api/users", body, config)
// // .then(res =>
// // dispatch({
// // type: REGISTER_SUCCESS,
// // payload: res.data
// // })
// // )
// // .catch(err => {
// // dispatch();
// // // returnErrors(err.response.data, err.response.status, "REGISTER_FAIL")
// // dispatch({
// // type: REGISTER_FAIL
// // });
// // });
// };
// Student Login
export const studentLogin = ({ email, password }, history) => dispatch => {
dispatch({ type: USER_LOADING });
axios
.post("http://localhost:5000/api/v1/student/login", {
email,
password
})
.then(res => {
console.log("response from action: ", res.data);
dispatch({ type: CLEAR_ERRORS });
dispatch({ type: LOGIN_SUCCESS, payload: res.data });
history.push("/student-dashboard");
})
.catch(error => {
dispatch(
returnErrors(error.response.data.message, error.response.data.success)
);
dispatch({ type: LOGIN_FAIL });
console.log("Error: ", error.response);
});
};
// Company Login
export const companyLogin = ({ email, password }, history) => dispatch => {
dispatch({ type: USER_LOADING });
axios
.post("http://localhost:5000/api/v1/company/login", {
email,
password
})
.then(res => {
console.log("response from action: ", res.data);
dispatch({ type: CLEAR_ERRORS });
dispatch({ type: LOGIN_SUCCESS, payload: res.data });
history.push("/Company-dashboard");
})
.catch(error => {
dispatch(
returnErrors(error.response.data.message, error.response.data.success)
);
dispatch({ type: LOGIN_FAIL });
console.log("Error: ", error.response);
});
};
// Admin Login
export const adminLogin = ({ email, password }, history) => dispatch => {
dispatch({ type: USER_LOADING });
axios
.post("http://localhost:5000/api/v1/admin/login", {
email,
password
})
.then(res => {
console.log("response from action: ", res.data);
dispatch({ type: CLEAR_ERRORS });
dispatch({ type: LOGIN_SUCCESS, payload: res.data });
history.push("/admin-dashboard");
})
.catch(error => {
dispatch(
returnErrors(error.response.data.message, error.response.data.success)
);
dispatch({ type: LOGIN_FAIL });
console.log("Error: ", error.response);
});
};
// Logout User
export const logout = () => {
return {
type: LOGOUT_SUCCESS
};
};
// // Setup config/headers and token
// export const tokenConfig = getState => {
// // Get token from localstorage
// const token = getState().auth.token;
// // Headers
// const config = {
// headers: {
// "Content-type": "application/json"
// }
// };
// // If token, add to headers
// if (token) {
// config.headers["x-auth-token"] = token;
// }
// return config;
// };
<file_sep>/src/Components/Display/JobsView.js
import React from "react";
import ApplyModal from "../ApplyModal";
const JobsView = ({ job, applyModal }) => {
return (
<div className="col-sm-6 col-md-4 col-lg-4">
<div className="card bg-light mb-3" style={{ maxWidth: "18rem" }}>
<div className="card-header">
<h3>
<strong>{job.jobTitle}</strong>
</h3>
</div>
<div className="card-body">
<h5 className="card-title">
<strong>Company: </strong> {job.companyId.companyName}
</h5>
<p className="card-text">
{" "}
<strong>Description: </strong>
{job.description}
</p>
{applyModal ? <ApplyModal label="Apply now" job={job} /> : null}
</div>
</div>
</div>
);
};
export default JobsView;
<file_sep>/src/Components/Dashboard/StudentDashboard.js
import React, { Component } from "react";
import { getJWT } from "../../helpers/jwt";
import axios from "axios";
import CompaniesView from "../Display/CompaniesView";
import JobsView from "../Display/JobsView";
import StudentProfileView from "../Display/StudentProfileView";
import AppliedJobsView from "../Display/AppliedJobsView";
import { connect } from "react-redux";
import { loadUserStudent } from "../../store/Actions/authActions";
import { Spinner } from "reactstrap";
class StudentDashboard extends Component {
state = {
companies: [],
jobs: [],
appliedJobs: [],
profile: [],
selectedMenu: "Profile",
isLoading: false
};
componentDidMount() {
this.props.loadUserStudent(this.props.history);
const jwt = getJWT();
if (jwt) {
this.setState({ isLoading: true });
// Fetch companies API
axios
.get("http://localhost:5000/api/v1/student/get-profile/", {
headers: {
"x-auth-token": `${jwt}`
}
})
.then(res => {
console.log("Student: ", res.data.student[0]);
this.setState({ profile: res.data.student[0] });
})
.catch(error => console.log("Error: ", error.response.data));
this.fetchData();
this.setState({ isLoading: false });
}
}
fetchData = () => {
const jwt = getJWT();
if (jwt) {
axios
.get("http://localhost:5000/api/v1/student/get-data/", {
headers: {
"x-auth-token": `${jwt}`
}
})
.then(res => {
console.log("Companies: ", res.data);
this.setState({
companies: res.data.companies,
jobs: res.data.allJobs,
appliedJobs: res.data.appliedJobs
});
})
.catch(error => console.log("Error: ", error.response.data));
}
};
render() {
if (this.state.isLoading) {
return <Spinner style={{ width: "3rem", height: "3rem" }} />;
}
return (
<React.Fragment>
<h1 className="mt-5 font-weight-bold">Student Portal</h1>
<div className="container mt-5">
<div className="row">
<div className="col-sm-12 col-md-3">
<ul className="list-group">
{/* Options Menu */}
<h3 className="text-center list-group-item list-group-item-secondary">
Menu
</h3>
<li
className="list-group-item d-flex justify-content-between list-group-item-action active`"
onClick={() => this.setState({ selectedMenu: "Profile" })}
>
Profile
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "Companies" });
}}
// this.setState({ selectedMenu: "Companies" })
>
Companies
<span className="badge badge-primary badge-pill">
{this.state.companies.length}
</span>
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "Jobs" });
}}
>
Jobs
<span className="badge badge-primary badge-pill">
{this.state.jobs.length}
</span>
</li>
<li
className="list-group-item d-flex justify-content-between align-items-center list-group-item-action"
onClick={() => {
this.fetchData();
this.setState({ selectedMenu: "AppliedJobs" });
}}
>
Applied Jobs
<span className="badge badge-primary badge-pill">
{this.state.appliedJobs.length}
</span>
</li>
</ul>
</div>
{/* Loader or content */}
{!this.state.companies || !this.state.jobs ? (
<Spinner style={{ width: "3rem", height: "3rem" }} />
) : (
<div className="col-sm-12 col-md-9">
<div className="row">
{this.state.selectedMenu === "Profile" && (
<StudentProfileView profile={this.state.profile} />
)}
{this.state.selectedMenu === "Companies" &&
this.state.companies.map(company => (
<CompaniesView key={company._id} company={company} />
))}
{this.state.selectedMenu === "Jobs" &&
this.state.jobs.map(job => (
<JobsView key={job._id} job={job} applyModal={true} />
))}
{this.state.selectedMenu === "AppliedJobs" &&
(this.state.appliedJobs.length === 1 ? (
<AppliedJobsView
key={this.state.appliedJobs._id}
job={this.state.appliedJobs[0]}
/>
) : (
this.state.appliedJobs.map(apjob => (
<AppliedJobsView key={apjob._id} job={apjob} />
))
))}
</div>
</div>
)}
</div>
</div>
</React.Fragment>
);
}
}
const mapStateToProps = state => ({
auth: state.auth
});
export default connect(
mapStateToProps,
{ loadUserStudent }
)(StudentDashboard);
<file_sep>/src/Components/Header.js
import React from "react";
import { Link } from "react-router-dom";
import { withRouter } from "react-router-dom";
import { connect } from "react-redux";
import { logout } from "../store/Actions/authActions";
const Header = props => {
const logUserOut = () => {
// localStorage.removeItem("jwt");
// console.log(props);
props.logout();
props.history.push("/");
};
return (
<nav className="navbar navbar-expand-lg navbar-dark bg-dark">
<div className="container">
<h1 className="navbar-brand">Campus Recruitment System</h1>
<form className="nav navbar-nav navbar-right form-inline my-2 my-lg-0">
{!props.auth.isAuthenticated ||
props.auth.isAuthenticated === null ? (
<div>
<Link
to="/register"
className="btn btn-outline-success my-2 my-sm-0 mr-3"
>
{" "}
Sign Up
</Link>
<Link to="/" className="btn btn-outline-success my-2 my-sm-0">
{" "}
Log In
</Link>
</div>
) : (
<Link
to="/"
className="btn btn-outline-success my-2 my-sm-0"
onClick={() => logUserOut()}
>
{" "}
Log out
</Link>
)}
</form>
</div>
</nav>
);
};
const mapStateToProps = state => ({
auth: state.auth
});
export default connect(
mapStateToProps,
{ logout }
)(withRouter(Header));
<file_sep>/src/Components/Display/CompaniesView.js
import React from "react";
const CompaniesView = ({ company }) => {
console.log(company);
return (
<div className="col-sm-12 col-md-6 col-lg-6">
<div className="card bg-light mb-3 border-info">
<div className="card-header">
<h3>{company.companyName}</h3>
</div>
<div className="card-body">
<h5 className="card-title">
<strong>Description: </strong>
{company.description}
</h5>
<p className="card-text">Website: {company.website}</p>
</div>
</div>
</div>
);
};
export default CompaniesView;
<file_sep>/src/Components/Display/JobAppicantView.js
import React from "react";
import moment from "moment";
const JobApplicantView = ({ applicant }) => {
console.log(applicant);
return (
// {jobs.map(job =>
// console.log(job)
// })}
// <JobCollapse title={applicant.jobId.jobTitle} />
<div className="col-sm-6">
<div className="list-group mb-4">
<div className="list-group-item list-group-item-action flex-column align-items-start">
<div className="d-flex w-100 justify-content-between">
<h4 className="mb-1">
<strong>Job:</strong> {applicant.jobId.jobTitle}
</h4>
<small>{moment(applicant.createdAt).fromNow()}</small>
</div>
<hr />
<div className="flex">
<p className="mb-1">
<strong>Student Name: </strong> {applicant.studentId.studentName}
</p>
<p className="mb-1">
<strong>Qualification: </strong>
{applicant.studentId.qualification}
</p>
</div>
</div>
</div>
</div>
);
};
export default JobApplicantView;
| d7dc601285810d826a816518980eb4691dbe89b8 | [
"JavaScript"
] | 13 | JavaScript | YousufQadri/Campus-Recruitment-System-react-client | 905bd3aa28e1096105d8dc15ed4ee5562fa171b1 | 555acd3c62827ab69dd8b583e9986da6ce98a3c1 |
refs/heads/master | <file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
#include <stdio.h>
int main() {
int a, b, max;
printf("Enter first number: ");
scanf("%d", &a);
printf("Enter second number: ");
scanf("%d", &b);
if (a > b) {
max = a;
}
else {
max = b;
}
printf("Maximum is %d\n", max);
return 0;
}<file_sep>#
# Generated - do not edit!
#
# NOCDDL
#
CND_BASEDIR=`pwd`
CND_BUILDDIR=build
CND_DISTDIR=dist
# Debug configuration
CND_PLATFORM_Debug=GNU-Linux
CND_ARTIFACT_DIR_Debug=dist/Debug/GNU-Linux
CND_ARTIFACT_NAME_Debug=initializing_vars
CND_ARTIFACT_PATH_Debug=dist/Debug/GNU-Linux/initializing_vars
CND_PACKAGE_DIR_Debug=dist/Debug/GNU-Linux/package
CND_PACKAGE_NAME_Debug=initializingvars.tar
CND_PACKAGE_PATH_Debug=dist/Debug/GNU-Linux/package/initializingvars.tar
# Release configuration
CND_PLATFORM_Release=GNU-Linux
CND_ARTIFACT_DIR_Release=dist/Release/GNU-Linux
CND_ARTIFACT_NAME_Release=initializing_vars
CND_ARTIFACT_PATH_Release=dist/Release/GNU-Linux/initializing_vars
CND_PACKAGE_DIR_Release=dist/Release/GNU-Linux/package
CND_PACKAGE_NAME_Release=initializingvars.tar
CND_PACKAGE_PATH_Release=dist/Release/GNU-Linux/package/initializingvars.tar
#
# include compiler specific variables
#
# dmake command
ROOT:sh = test -f nbproject/private/Makefile-variables.mk || \
(mkdir -p nbproject/private && touch nbproject/private/Makefile-variables.mk)
#
# gmake command
.PHONY: $(shell test -f nbproject/private/Makefile-variables.mk || (mkdir -p nbproject/private && touch nbproject/private/Makefile-variables.mk))
#
include nbproject/private/Makefile-variables.mk
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
/*
* File: main.c
* Author: gwalters
*
* Created on May 28, 2018, 12:38 PM
*/
#include <stdio.h>
#include <stdlib.h>
/*
*
*/
int main(int argc, char** argv) {
int aVar, x;
printf("Enter an integer: ");
scanf("%d", &aVar);
x = aVar + 1;
printf("The value is: %d and %d\n", aVar, x);
return 0;
}
<file_sep>// A simple Hello, world program
// Note: The file name must be the same as the
// class name, with a .java extension
package examples;
public class HelloStars
{
public static void main(String[] args)
{
System.out.println("***************************************");
System.out.println("* \t\tHello Gavin!\t\t*");
System.out.println("***************************************");
}
}
<file_sep>package com.gavinwalters;
public class Main {
public static void main(String[] args) {
int newScore = calculateScore(500);
System.out.println("New score is " + newScore);
calculateScore(1);
calculateScore();
calcFeetAndInchesToCentimeters(157);
}
public static int calculateScore(String playerName, int score) {
System.out.println("Player " + playerName + " scored " + score + " points");
return score * 1000;
}
public static int calculateScore(int score) {
System.out.println("Unkown player scored " + score + " points");
return score * 1000;
}
public static int calculateScore() {
System.out.println("No player name, no player score.");
return 0;
}
public static double calcFeetAndInchesToCentimeters(int feet, int inches) {
if ((feet < 0) || ((inches < 0) || (inches > 12))) {
System.out.println("Invalid range");
return -1;
}
double calc = (((feet * 12) + inches) * 2.54);
System.out.println(feet + " ft " + inches + " inches = " + calc + " cm");
return calc;
}
public static double calcFeetAndInchesToCentimeters(int inches) {
int feet;
if (inches >= 0) {
if (inches >= 12) {
feet = inches / 12;
inches = inches - ((inches / 12) * 12);
return calcFeetAndInchesToCentimeters(feet, inches);
}
return (inches * 2.54);
}
System.out.println("Invalid range");
return -1;
}
}
<file_sep>#
# Generated - do not edit!
#
# NOCDDL
#
CND_BASEDIR=`pwd`
CND_BUILDDIR=build
CND_DISTDIR=dist
# Debug configuration
CND_PLATFORM_Debug=GNU-Linux
CND_ARTIFACT_DIR_Debug=dist/Debug/GNU-Linux
CND_ARTIFACT_NAME_Debug=income
CND_ARTIFACT_PATH_Debug=dist/Debug/GNU-Linux/income
CND_PACKAGE_DIR_Debug=dist/Debug/GNU-Linux/package
CND_PACKAGE_NAME_Debug=income.tar
CND_PACKAGE_PATH_Debug=dist/Debug/GNU-Linux/package/income.tar
# Release configuration
CND_PLATFORM_Release=GNU-Linux
CND_ARTIFACT_DIR_Release=dist/Release/GNU-Linux
CND_ARTIFACT_NAME_Release=income
CND_ARTIFACT_PATH_Release=dist/Release/GNU-Linux/income
CND_PACKAGE_DIR_Release=dist/Release/GNU-Linux/package
CND_PACKAGE_NAME_Release=income.tar
CND_PACKAGE_PATH_Release=dist/Release/GNU-Linux/package/income.tar
#
# include compiler specific variables
#
# dmake command
ROOT:sh = test -f nbproject/private/Makefile-variables.mk || \
(mkdir -p nbproject/private && touch nbproject/private/Makefile-variables.mk)
#
# gmake command
.PHONY: $(shell test -f nbproject/private/Makefile-variables.mk || (mkdir -p nbproject/private && touch nbproject/private/Makefile-variables.mk))
#
include nbproject/private/Makefile-variables.mk
<file_sep>package examples;
public class ThreeVariables
{
public static void main (String[] args)
{
int deptNum = 0;
float salary = 0;
int jobClass = 0;
java.util.Scanner scanner = new java.util.Scanner(System.in);
System.out.print("Enter your department number: ");
deptNum = scanner.nextInt();
System.out.print("Enter your salary amount: ");
salary = scanner.nextFloat();
System.out.print("Enter you job class number: ");
jobClass = scanner.nextInt();
System.out.printf("<Department Number: " + deptNum + " > <Salary: " +
salary + " > <Job Class "+
"Number: " + jobClass + " >\n");
System.out.printf("<Department Number: " + deptNum + " > \t<Salary: " +
salary + " >\t<Job Class "+
"Number: " + jobClass + " >\n");
System.out.printf("<Department Number: " + deptNum + " > \n<Salary: " +
salary + " >\n<Job Class "+
"Number: " + jobClass + " >\n");
}
}
<file_sep>import java.io.PrintWriter;
import java.util.Scanner;
//Add additional import statements as needed
public class WritingToDocument
{
//Declare appropriate variables
public static void main(String[] args)
throws FileNotFoundException
{
//Create and associate the stream objects
Scanner inFile =
new Scanner(new FileReader("prog.dat"));
PrintWriter outFile = new PrintWriter("prog.out");
//Code for data manipulation
//Close file
inFile.close();
outFile.close();
}
}
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
/*
* File: main.c
* Author: gwalters
*
* Created on June 3, 2018, 1:06 AM
*/
#include <stdio.h>
#include <stdlib.h>
/*
*
*/
int main(int argc, char** argv) {
int mood;
printf("Please enter the number of neighbours around Pupeta: ");
scanf("%d", &mood);
if (mood < 0 || mood > 8) {
printf("There is no way Pupeta can have any neighbhors NOT between the"
" numbers 0 and 8...you liar!");
}
else if (mood == 0) {
printf("Sad and will force you to listen to his story, will make you late.");
}
else if (mood == 1) {
printf("He will ask you to deliver half of the pizza to his only neighbor, you may be late.");
}
else if (mood >= 2 && mood <= 5) {
printf("Will have a great mood and will be singing song. May crack joke as well. You will get"
"good tips, definetely.");
}
else if (mood == 7) {
printf("He will be making great drink for his neighbors and will invite you to taste. But will not give you tip.");
}
else if (mood == 6 || mood == 8) {
printf("Too angry and furious, deliver the pizza and leave ASAP.");
}
return (EXIT_SUCCESS);
}
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
package javamasterclass;
/**
*
* @author gvnwa
*/
public class JavaMasterclass {
/**
* @param args the command line arguments
*/
public static void main(String[] args) {
// TODO code application logic here
// for (int i = 2; i < 9; i++) {
// System.out.println("10,000 at " + i + "% interest = " + String.format("%.2f", calculateInterest(10000.0, i)));
// }
for (int i = 8; i > 1; i--) {
System.out.println("10,000 at " + i + "% interest = " + String.format("%.2f", calculateInterest(10000.0, i)));
}
}
public static double calculateInterest(double amount, double interestRate) {
return (amount * (interestRate/100));
}
}
<file_sep>import java.io.*;
import java.util.*;
public class LoopWithBugsData1
{
public static void main(String[] args)
throws FileNotFoundException
{
int i;
int j;
int sum;
int num;
Scanner infile =
new Scanner(new FileReader("Ch5_LoopWithBugsData.txt"));
{
sum = 0;
for (j = 1; j <=4; j++)
{
num = infile.nextInt();
System.out.print(num + " ");
sum = sum + num;
}
System.out.println("sum = " + sum);
}
}
}
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
/*
* File: main.c
* Author: gwalters
*
* Created on June 3, 2018, 12:35 PM
*/
#include <stdio.h>
#include <stdlib.h>
/*
*
*/
int main(int argc, char** argv) {
double income, tax;
printf("Enter your income: ");
scanf("%lf", &income);
if (income < 10000) {
tax = 0.0;
}
else if (income >= 10000 && income < 20000) {
tax = income * .1;
}
else if (income >= 20000 && income < 50000) {
tax = income * .2;
}
else {
tax = income * .25;
tax = tax + tax * .1;
}
printf("The payable tax is $%0.2lf\n", tax);
return (EXIT_SUCCESS);
}
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
#include <stdio.h>
int main(int argc, char** argv) {
int age;
printf("Enter your age: ");
scanf("%d", &age);
if (age >= 13 && age <= 19) {
printf("Yup! You are a teenager!");
}
else {
printf("No, you are not a teenager!");
}
return 0;
}
<file_sep>// A simple Hello, world program
// Note: The file name must be the same as the
// class name, with a .java extension
package examples;
public class HelloTabs
{
public static void main(String[] args)
{
System.out.println("\t \t Hello Gavin!");
}
}
<file_sep>// A simple Hello, world program
// Note: The file name must be the same as the
// class name, with a .java extension
package examples;
public class TwoPrints
{
public static void main(String[] args)
{
System.out.println("Hello");
System.out.println("Gavin!");
}
}
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
/*
* File: main.c
* Author: gwalters
*
* Created on June 3, 2018, 1:01 AM
*/
#include <stdio.h>
#include <stdlib.h>
/*
*
*/
int main(int argc, char** argv) {
int a, b, c, max;
printf("Enter 3 numbers: ");
scanf("%d %d %d", &a, &b, &c);
if (a > b && a > c) {
max = a;
}
else if (b > c) {
max = b;
}
else {
max = c;
}
printf("Maximum is %d\n", max);
return (EXIT_SUCCESS);
}
<file_sep>/*
* To change this license header, choose License Headers in Project Properties.
* To change this template file, choose Tools | Templates
* and open the template in the editor.
*/
/*
* File: main.c
* Author: gwalters
*
* Created on May 28, 2018, 8:10 PM
*/
#include <stdio.h>
#include <stdlib.h>
/*
*
*/
int main(int argc, char** argv) {
int var = 45;
printf("var = %d\n", var);
printf("var = %o\n", var);
printf("var = %x\n\n", var);
int var1 = 056;
printf("var = %d\n", var1);
printf("var = %o\n", var1);
printf("var = %x\n\n", var1);
int var2 = 0xa;
printf("var = %d\n", var2);
printf("var = %o\n", var2);
printf("var = %x\n", var2);
return 0;
}
<file_sep>/*
* Lab 2.15: PaintEstimator
*
* IT-145 Foundations of App Development
* Southern New Hampshire University
*/
//package paintestimator;
import java.util.Scanner;
import java.lang.Math;
/**
*
* @author <EMAIL>
*/
public class PaintEstimator {
public static void main(String[] args) {
Scanner scnr = new Scanner(System.in);
double wallHeight = 0.0;
double wallWidth = 0.0;
double wallArea = 0.0;
double gallonsPaintNeeded = 0.0;
int cansNeeded = 0;
final double squareFeetPerGallons = 350.0;
final double gallonsPerCan = 1.0;
System.out.println("Enter wall height (feet): ");
wallHeight = scnr.nextDouble();
// Prompt user to input wall's width
System.out.println("Enter wall width (feet): ");
wallHeight = scnr.nextDouble();
// Calculate and output wall area
wallArea = wallHeight * wallWidth;
System.out.println("Wall area: square feet");
// Calculate and output the amount of paint in gallons needed to paint the wall
gallonsPaintNeeded = wallArea/squareFeetPerGallons;
System.out.println("Paint needed: " + gallonsPaintNeeded + " gallons");
// Calculate and output the number of 1 gallon cans needed to paint the wall, rounded up to nearest integer
cansNeeded = (int) (gallonsPaintNeeded / gallonsPerCan); //Hint: this line is missing two operations
System.out.println("Cans needed: " + cansNeeded + " can(s)");
}
}
<file_sep>//Just a basic hello world program
public class helloWolrdAgain
{
public static void main(String[] args)
{
System.out.println("Hello muthafucking world for the "
+ "billionth time!");
}
}
<file_sep>package examples;
public class AgeInitial
{
public static void main (String[] args)
{
int age = 28;
String firstName = "Gavin";
char middleInitial = 'G';
String lastName = "Davis";
System.out.printf("%s %c %s : Age: %d", firstName, middleInitial, lastName, age);
}
}
<file_sep>/*
*************************************************************************
*
* File: nondescendingOrder.java
* Date: 05/03/2016
*
* Author: <NAME>
*
*************************************************************************
*/
//Write a program that prompts the user to input three numbers. This program should then output the numbers in nondescending order.
//Submit your compiled Java code for this assignment, compress the .java file into a single .zip file.
//For additional details, refer to the Programming Problems Rubric in the Assignment Guidelines and Rubrics folder.
//1)Prompt user for 1st, 2nd, and 3rd number.
//2)Output the selected numbers in nondescending order.
//import library to use nice user interface for
//ux/ui
import javax.swing.JOptionPane;
//name program
public class nondescendingOrder
{
//declare main method to start program
public static void main(String[] args)
{
//declare variables to store user inputs
//and display user outputs after
//processing
String input;
String output = "";
String intro = "This program "
+ "will ask you for three "
+ "numbers that will subsequently "
+ "be ordered in nondescending order.";
//declare variable that will be used to store
//user inputs
int firstNum, secondNum, thirdNum;
//used to hold number for a kind of bubble sort
//holds the number if the first if conditonal statement
//is true. Used to move number where it needs to be in
//the order.
int hold;
//introduce user to progam and request
//inputs
JOptionPane.showMessageDialog(null,intro, "Welcome!", JOptionPane.INFORMATION_MESSAGE);
input = JOptionPane.showInputDialog
("Enter the first number: ");
//process inputs into integers
firstNum = Integer.parseInt(input);
input = JOptionPane.showInputDialog
("Enter the second number: ");
secondNum = Integer.parseInt(input);
input = JOptionPane.showInputDialog
("Enter the final number: ");
thirdNum = Integer.parseInt(input);
//Use relational operators to assess
//user inputs and rearrange them
//accordingly
if (thirdNum < firstNum)
{
hold = thirdNum;
if (secondNum < hold)
output = String.format("%d, %d, %d",secondNum, hold, firstNum); //format strings to input into JOptionPane
else
output = String.format("%d, %d, %d", hold, secondNum, firstNum);//if thirdNum is less than one but seconNum is not less
//than hold of thirdNum, then thirdNum stays to replace
//the first num. All following statements follow similar
//pattern
}
else if (secondNum < firstNum)
{
hold = secondNum;
if (thirdNum < hold)
output = String.format("%d, %d, %d",thirdNum, hold, firstNum);
else
output = String.format("%d, %d, %d",hold, firstNum, thirdNum);
}
else if (thirdNum < secondNum)
{
hold = thirdNum;
if (firstNum < hold)
output = String.format("%d, %d, %d",firstNum, hold, secondNum);
else
output = String.format("%d, %d, %d",firstNum, hold, secondNum);
}
else if (secondNum < thirdNum)
{
hold = secondNum;
if (firstNum < hold)
output = String.format("%d, %d, %d",firstNum, hold, thirdNum);
else
output = String.format("%d, %d, %d",firstNum, hold, thirdNum);
}
else if (firstNum < thirdNum)
{
hold = firstNum;
if (secondNum < hold)
output = String.format("%d, %d, %d",secondNum, hold, thirdNum);
else
output = String.format("%d, %d, %d",hold, secondNum, thirdNum);
}
else if (firstNum < secondNum)
{
hold = firstNum;
if (thirdNum < hold)
output = String.format("%d, %d, %d",thirdNum, hold, secondNum);
else
output = String.format("%d, %d, %d",firstNum, secondNum, thirdNum);
}
//show dialog window with nondescending order results store in output
JOptionPane.showMessageDialog(null, output,
"Nondescending Order",
JOptionPane.INFORMATION_MESSAGE);
}
}
| f39884b15e673dd4b62df662364006cc0805f444 | [
"Java",
"C",
"Makefile"
] | 21 | C | gvnwlt/java-projects | 644a65003b54b2852b5802834e4d24b09d29b6c9 | 3960c40e5cf02cf5b8f847d598aa4a9a1dcdcf69 |
refs/heads/main | <repo_name>vladkulikov/NCPalindrome<file_sep>/IPalindrome.java
package com.company;
public interface IPalindrome {
boolean isPalindromev1(int x);
boolean isPalindromev2(int x);
boolean isPalindromev3(int x);
}
<file_sep>/PalindromeTest.java
package com.company;
import org.junit.Assert;
import org.junit.jupiter.api.Assertions;
import static org.junit.jupiter.api.Assertions.*;
class PalindromeTest {
//проверка нечетного количества цифр, палиндром
@org.junit.jupiter.api.Test
void isPalindromev1Test1() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev1(121);
assertTrue(actual);
}
//проверка четного количества цифр, палиндром
@org.junit.jupiter.api.Test
void isPalindromev1Test2() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev1(1221);
assertTrue(actual);
}
//проверка не палиндрома
@org.junit.jupiter.api.Test
void isPalindromev1Test3() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev1(1212);
assertFalse(actual);
}
//проверка не палиндрома
@org.junit.jupiter.api.Test
void isPalindromev1Test4() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev1(-12121);
assertTrue(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev2Test1() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev2(121);
assertTrue(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev2Test2() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev2(1221);
assertTrue(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev2Test3() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev2(1212);
assertFalse(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev2Test4() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev1(-12121);
assertTrue(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev3Test1() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev2(121);
assertTrue(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev3Test2() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev2(1221);
assertTrue(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev3Test3() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev2(1212);
assertFalse(actual);
}
@org.junit.jupiter.api.Test
void isPalindromev3Test4() {
Palindrome palindrome = new Palindrome();
boolean actual = palindrome.isPalindromev1(-12121);
assertTrue(actual);
}
} | 38798ca95ec68ef2f4cb7cacf49a6b05a7e58ccd | [
"Java"
] | 2 | Java | vladkulikov/NCPalindrome | 3be8880c1a8723c9b959a07d4cde027ea108c3c9 | 8a42d57c5bd074dd957a8c21662c60ed13478526 |
refs/heads/master | <file_sep>using Npgsql;
using System;
using System.Collections.Generic;
using System.ComponentModel;
using System.Data;
using System.Drawing;
using System.Linq;
using System.Text;
using System.Threading.Tasks;
using System.Windows.Forms;
namespace OtelOtomasyon
{
public partial class Form1 : Form
{
public void DataGridGuncelle()
{
try
{
connect.Open();
}
catch (Exception ex)
{
MessageBox.Show(ex.Message);
}
dataSet.Reset();
string sql = "select \"Insan\".\"Id\" as \"ID\",\"Insan\".\"Adi\" as \"AD\",\"Insan\".\"Soyadi\" as \"SOYAD\"," +
"\"Insan\".\"TelNo\" as \"TEL NO\",\"Insan\".\"TcNo\" as \"TC\",\"Insan\".\"Adres\" as \"ADRES\"," +
"\"Insan\".\"Cinsiyet\" as \"CİNSİYET\"," + "\"Insan\".\"DogumTarihi\" as \"DOĞUM TARİHİ\"," +
"\"Musteriler\".\"MedeniDurum\" as \"MEDENİ DURUM\",\"Musteriler\".\"SirketAdi\" as \"ŞİRKET ADI\",\"Musteriler\".\"Aciklama\" as \"AÇIKLAMA\" " +
"from \"Insan\" inner join \"Musteriler\"on \"Insan\".\"Id\" = \"Musteriler\".\"Id\"";
NpgsqlDataAdapter add = new NpgsqlDataAdapter(sql, connect);
add.Fill(dataSet);
dataGridView1.DataSource = dataSet.Tables[0];
connect.Close();
}
public void Temizle()
{
txtAd.Clear();
txtSoyad.Clear();
txtTelNo.Clear();
txtTc.Clear();
txtAdres.Clear();
txtDogumTarihi.Clear();
txtSirketAdi.Clear();
txtAciklama.Clear();
rdbErkek.Checked = false;
rdbKadin.Checked = false;
rdbEvli.Checked = false;
rdbBekar.Checked = false;
}
NpgsqlConnection connect = new NpgsqlConnection("Server=localhost;Port=5432;Database=Otel;User Id=postgres;Password=<PASSWORD>");
DataSet dataSet = new DataSet();
public Form1()
{
InitializeComponent();
}
private void Form1_Load(object sender, EventArgs e)
{
txtDogumTarihi.Text = "gg/aa/yyyy";
txtDogumTarihi.ForeColor = Color.Gray;
txtId.Enabled = false;
DataGridGuncelle();
}
private void btnBul_Click(object sender, EventArgs e)
{
Temizle();
if (txtId.Text == "")
{
MessageBox.Show("Lütfen Id Giriniz.");
}
else
{
connect.Open();
int id = Convert.ToInt32(txtId.Text);
string sql= "select \"Insan\".\"Id\" as \"ID\",\"Insan\".\"Adi\" as \"AD\",\"Insan\".\"Soyadi\" as \"SOYAD\"," +
"\"Insan\".\"TelNo\" as \"TEL NO\",\"Insan\".\"TcNo\" as \"TC\",\"Insan\".\"Adres\" as \"ADRES\"," +
"\"Insan\".\"Cinsiyet\" as \"CİNSİYET\"," + "\"Insan\".\"DogumTarihi\" as \"DOĞUM TARİHİ\"," +
"\"Musteriler\".\"MedeniDurum\" as \"MEDENİ DURUM\",\"Musteriler\".\"SirketAdi\" as \"ŞİRKET ADI\",\"Musteriler\".\"Aciklama\" as \"AÇIKLAMA\" " +
"from \"Insan\" inner join \"Musteriler\"on \"Insan\".\"Id\" = \"Musteriler\".\"Id\"" +
"where \"Insan\".\"Id\"="+id+"";
NpgsqlCommand command = new NpgsqlCommand(sql, connect);
NpgsqlDataReader read = command.ExecuteReader();
if (read.HasRows == false)
{
MessageBox.Show("Aranan Kayıt Bulunamadı");
txtId.Clear();
}
while (read.Read()) {
txtAd.Text = read[1].ToString();
txtSoyad.Text = read[2].ToString();
txtTelNo.Text = read[3].ToString();
txtTc.Text = read[4].ToString();
txtAdres.Text = read[5].ToString();
txtDogumTarihi.Text = read[7].ToString();
txtSirketAdi.Text = read[9].ToString();
txtAciklama.Text = read[10].ToString();
if (read[6].ToString() == "1")
{
rdbErkek.Checked = true;
}
else
{
rdbKadin.Checked = true;
}
if (read[8].ToString() == "1")
rdbEvli.Checked = true;
else
rdbBekar.Checked = true;
}
connect.Close();
txtId.Enabled = false;
}
}
private void btnTemizle_Click(object sender, EventArgs e)
{
Temizle();
txtId.Clear();
}
private void btnBul_MouseEnter(object sender, EventArgs e)
{
txtId.Enabled = true;
}
private void txtId_MouseLeave(object sender, EventArgs e)
{
txtId.Enabled = false;
}
private void btnEkle_Click(object sender, EventArgs e)
{
if (txtId.Text != "")
{
MessageBox.Show("Id alanı otomatik belirlenir. Lütfen boş bırakınız.");
}
else
{
if(txtAd.Text=="" || txtDogumTarihi.Text=="" || txtSoyad.Text=="" || txtTc.Text=="" || txtTelNo.Text == "")
{
MessageBox.Show("Adres,Şirket Adı ve Açıklama Haricindeki bütün alanları doldurmak zorunludur!");
}
else
{
string ad = txtAd.Text;
string soyad = txtSoyad.Text;
string telno = txtTelNo.Text;
string tc = txtTc.Text;
string adres = txtAdres.Text;
string dogumtarihi = txtDogumTarihi.Text;
string sirketadi = txtSirketAdi.Text;
string aciklama = txtAciklama.Text;
int cinsiyet = 0;
if (rdbErkek.Checked == true)
cinsiyet = 1;
int medeni = 0;
if (rdbEvli.Checked == true)
medeni = 1;
string sqlInsan= "INSERT INTO \"Insan\" ( \"Adi\", \"Soyadi\", \"TelNo\", \"TcNo\", \"Adres\", \"Cinsiyet\", \"DogumTarihi\") " +
"VALUES(\'"+ad+ "\',\'" + soyad+ "\',\'" + telno+ "\',\'" + tc+ "\',\'" + adres+ "\',\'" + cinsiyet+ "\',\'" + dogumtarihi+ "\')";
string sqlMusteri = "INSERT INTO \"Musteriler\" ( \"Id\", \"MedeniDurum\", \"SirketAdi\", \"Aciklama\") " +
"VALUES(\"sonInsanSayisi\"(),\'" + medeni + "\',\'" + sirketadi + "\',\'" + aciklama + "\')";
connect.Open();
NpgsqlCommand cmd = new NpgsqlCommand(sqlInsan, connect);
cmd.ExecuteNonQuery();
cmd.CommandText = sqlMusteri;
cmd.ExecuteNonQuery();
connect.Close();
DataGridGuncelle();
}
}
}
private void txtDogumTarihi_MouseEnter(object sender, EventArgs e)
{
//txtDogumTarihi.Clear();
txtDogumTarihi.ForeColor = Color.Black;
}
private void txtDogumTarihi_MouseLeave(object sender, EventArgs e)
{
//txtDogumTarihi.Text = "gg/aa/yyyy";
//txtDogumTarihi.ForeColor = Color.Gray;
}
private void btnSil_MouseEnter(object sender, EventArgs e)
{
txtId.Enabled = true;
}
private void btnSil_Click(object sender, EventArgs e)
{
if (txtId.Text == "")
{
MessageBox.Show("Lütfen Id Giriniz.");
}
else
{
int id = Convert.ToInt32(txtId.Text);
string sql = "DELETE FROM \"Insan\" WHERE \"Id\" =" + id + "";
connect.Open();
NpgsqlCommand cmd = new NpgsqlCommand(sql, connect);
cmd.ExecuteNonQuery();
connect.Close();
DataGridGuncelle();
}
}
private void btnGuncelle_Click(object sender, EventArgs e)
{
if (txtId.Text == "")
{
MessageBox.Show("Önce Bul işlemi yapınız!");
}
else
{
int id = Convert.ToInt32(txtId.Text);
string ad = txtAd.Text;
string soyad = txtSoyad.Text;
string telno = txtTelNo.Text;
string tc = txtTc.Text;
string adres = txtAdres.Text;
string dogumtarihi = txtDogumTarihi.Text;
string sirketadi = txtSirketAdi.Text;
string aciklama = txtAciklama.Text;
int cinsiyet = 0;
if (rdbErkek.Checked == true)
cinsiyet = 1;
int medeni = 0;
if (rdbEvli.Checked == true)
medeni = 1;
string sqlInsan = "UPDATE \"Insan\" SET \"Adi\" = \'" + ad + "\',\"Soyadi\" =\'" + soyad + "\' ," +
"\"TelNo\" = \'" + telno + "\',\"TcNo\" = \'" + tc + "\',\"Adres\" =\'" + adres + "\' ," +
"\"Cinsiyet\" =\'" + cinsiyet + "\' ,\"DogumTarihi\" =\'" + dogumtarihi + "\' WHERE \"Id\"=" + id + " ";
string sqlMusteri = "UPDATE \"Musteriler\" SET \"MedeniDurum\" =\'" + medeni + "\' ,\"SirketAdi\" =" +
"\'" + sirketadi + "\' ,\"Aciklama\" =\'" + aciklama + "\' WHERE \"Id\" =" + id + " ";
connect.Open();
NpgsqlCommand cmd = new NpgsqlCommand(sqlInsan, connect);
cmd.ExecuteNonQuery();
cmd.CommandText = sqlMusteri;
cmd.ExecuteNonQuery();
connect.Close();
DataGridGuncelle();
Temizle();
txtId.Clear();
}
}
}
}
| 27f8e19cb6b3bb551d59d407cc746182b36f34b1 | [
"C#"
] | 1 | C# | emreharman/VeriTaban-YonetimSistemleri | 02115df1b05c4e9d4526f2c09731260c550876f8 | c68de2c11339f678a34bde170e00849b1bfe23dc |
refs/heads/master | <repo_name>SHRISHRIKANT/Python<file_sep>/beta.py
def run(): # defining a function
print('Hello, Python.')
print("Hi, Github")
run() # calling a function
| b5f0820a15259afb56d03082bb24a61be15c90cc | [
"Python"
] | 1 | Python | SHRISHRIKANT/Python | a54d164f8e3f45e5b5d691e172a588c4eecc8d7a | 33499751186edc7dda6ddfb47447078ce53d488a |
refs/heads/master | <repo_name>guer0157/nodejs<file_sep>/mongo-db/deleteDoc.js
const mongoose=require('mongoose');
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
let courseSchema= new mongoose.Schema({
author:String,
tags:[String],
name:String,
isPublished:Boolean,
price:Number,
date:{type:Date, default:d=Date.now}
})
let Course=mongoose.model('Course', courseSchema)
async function deleteCourse(id){
// let courses=await Course.deleteOne({_id:id})
//findByIdAndremove will return the docuement that was removed.
let courses= await Course.findByIdAndRemove({_id:id})
console.log(courses)
}
deleteCourse('5c3a8a8a7fea8b037503e4cf')<file_sep>/mongo-db/updateDocs.js
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
.then(()=>console.log('Connected to mongoDB'))
.catch(err=>console.error('Could not connecto to mngod', err))
const courseSchema=new mongoose.Schema({
id:String,
name: String,
author: String,
tags: [String],
date: {type:Date, default:Date.now},
isPublished: Boolean,
price:Number
})
const Course=mongoose.model('Course', courseSchema)
async function updateCourse(id){
// console.log(id)
//Approad: QueryFirst
//findByid()
//Modify its properties
//save()
//find the element by id
const courses=await Course.findById(id)
//check if there is a course with that id
if (!courses){ return}
//set the values of the properties you want to modify
courses.isPublished=false;
courses.author='Another Author';
//await the result fo the async func and save it.
const result= await courses.save()
console.log(courses)
}
updateCourse("5c3a8a8a7fea8b037503e4cf")<file_sep>/mongo-db/exercise2.js
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises', {useNewUrlParser:true})
.then(()=>console.log("connected to Mongo DB"))
.catch(err=>console.log(err))
const courseSchema=mongoose.Schema({
author:String,
tags:[String],
name:String,
price:Number,
date:{type:Date, default:Date.now},
isPublished:Boolean
})
const Course = mongoose.model('Course', courseSchema)
async function getCourses(){
return await Course
.find({isPublished:true, tags:{$in:['frontend', 'backend']}})
.sort({price:-1})
.select({name:1, author:1})
}
async function showCourses(){
const course= await getCourses()
console.log(course)
}
showCourses()
<file_sep>/mongo-db/customValidator.js
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
.then(()=>console.log('Connected to mongoDB'))
.catch(err=>console.error('Could not connecto to mngod', err))
const courseSchema=new mongoose.Schema({
name: {
type: String,
required:true,
minlength:5,
maxlength:255,
},
category:{
type:String,
required:true,
enum:['web', 'mobile', 'network']
},
author: String,
tags: {type:Array,
//to set a custom validator set the value of the property you want to validate to an object set the first key
//in the object to 'type' and set its value to the required datatype
//then set the second key to 'validate' and set the vlaue to an object with the key 'validator' with a value of function(){}
//where you can perform the logic you need to validate
//the below coe check if the property 'tags' is set to an array and if lengh of the array is more than 0
validate:{
validator: function(v){
return v&&v.lenght>0;
},
message:'Course should have atleast one tag'
}
},
date: {type:Date, default:Date.now},
isPublished: Boolean,
price:{ type:Number, required:function(){ return this.isPublished} }
})
const Course=mongoose.model('Course', courseSchema)
async function addCourse(){
const newCourse= await Course({
tags:null,
category:'web',
name:'Xamarin for Beginners',
author:'Silvia',
isPublished:true,
price:15
})
await newCourse.save().catch(err=>console.log("Error",err.message))
}
addCourse()<file_sep>/mongo-db/exercises.js
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
const courseSchema= mongoose.Schema({
tags:[String],
date:{type:Date,default:Date.now},
name:String,
author:String,
isPublished:Boolean,
price:Number
})
const Course = mongoose.model('Course', courseSchema)
async function getCourses(){
return await Course
.find({isPublished:true})
.sort({price:-1})
.select({name:1,author:1, price:1})
}
async function run(){
const courses = await getCourses()
console.log(courses);
}
run()<file_sep>/mongo-db/validation.js
const mongoose=require('mongoose')
//set the connection to the mongobd
//mongoose.connect("mongodb://localhost:27017/YourDB", { useNewUrlParser: true });
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
.then(()=>console.log('Connected to mongoDB'))
.catch(err=>console.error('Could not connecto to mngod', err))
const courseSchema=new mongoose.Schema({
//validate required fields by passing an object as the value of the key {type:String, required:true} setting the 'type' property to
//the data type ex: 'String' and adding an additional property called 'required' with the value set to true
name: {
//String validators: minlength,maxlength,match,enum
type: String,
required:true,
minlength:5,
maxlength:255,
// match:/patter/
},
category:{
type:String,
required:true,
//'enum' may be use when only a certain set of predefined values is valid
//we can set those values inside an array an set the array to the value of the 'enum' property
enum:['web', 'mobile', 'network']
},
author: String,
tags: [String],
//number and date validators: min, max
date: {type:Date, default:Date.now},
isPublished: Boolean,
//We can pass function as th value of the required property qwhen we want to set conditional validation
//for example is 'price' is only required when 'isPublished' is set to 'true'
price:{ type:Number, required:function(){ return this.isPublished} }
})
const Course=mongoose.model('Course', courseSchema)
async function addCourse(){
const newCourse= await Course({
tags:['frontend', 'Xamarin'],
category:'0',
name:'Xamarin for Beginners',
author:'Silvia',
isPublished:true,
price:15
})
// const result=
await newCourse.save().catch(err=>console.log("Error",err.message))
// console.log(result)
}
addCourse()<file_sep>/mongo-db/updateDocs2.js
//To update multiple object use the update method
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises', {useNewUrlParser:true})
const courseSchema= new mongoose.Schema({
tags:[String],
author:String,
name:String,
isPublished:Boolean,
date:{type:Date, default:Date.now},
price:Number
})
const Course=mongoose.model('Course', courseSchema)
async function updateCourse(id){
//you may use the .updateMany(({parameter}), {updateObject}) and pass it the parameter you're looking for and one of the mongo db update methods.
//example: Course.updateMany({isPublished:false}) will return all the courses where the
//'isPublished' property is false and you can then update all of them at once.
//*google mongo db update operators */
//Use updateOne(), updateMany(), or bulkWrite(), findByIdAndUpdate() return the document that was updated.
//the third argument is oprtional and it specifies if you want the method to return the new docuement after it is updated.
const courses= await Course.findByIdAndUpdate({_id:id},{$set:{author:'Cesar',isPublished:true}},{new:true})
console.log(courses)
}
updateCourse('5c3a8a8a7fea8b037503e4cf')<file_sep>/RESTful/middleware/logger.js
//call app.use to install a middleware function in the request processing pipeline
//function takes req, res, and "next". "next" is a reference to the next middleware function in the pipeline "the next function to run"
let log = (req, res, next)=>{
//perform code here
console.log('logging');
//then call the "next" function to pass control to the next request processing function. which will terminate the process and
//send reponse back to client
next()
}
module.exports = log<file_sep>/mongo-db/connect.js
const mongoose=require('mongoose')
//set the connection to the mongobd
//mongoose.connect("mongodb://localhost:27017/YourDB", { useNewUrlParser: true });
mongoose.connect('mongodb://localhost:27017/mongo-exercises/', { useNewUrlParser: true })
.then(()=>console.log('Connected to mongoDB'))
.catch(err=>console.error('Could not connecto to mngod', err))
<file_sep>/mongo-db/asyncValidation.js
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
.then(()=>console.log('Connected to mongoDB'))
.catch(err=>console.error('Could not connecto to mngod', err))
const courseSchema=new mongoose.Schema({
name: {
type: String,
required:true,
minlength:5,
maxlength:255,
},
category:{
type:String,
required:true,
enum:['web', 'mobile', 'network']
},
author: String,
tags: {type:Array,
validate:{
//we can set async functions as validators when we need to grab info from a different http server in order to
//validate to use async validation set the add the key 'isAsync' and set it to 'true'
//then in the validator function pass in the argument that is being validfated and a callback function
isAsync:true,
validator: function(v, callback){
setTimeout(()=>{
const result=v&&v.lenght>0;
callback(result);
},4000)
},
message:'Course should have atleast one tag'
}
},
date: {type:Date, default:Date.now},
isPublished: Boolean,
price:{ type:Number, required:function(){ return this.isPublished} }
})
const Course=mongoose.model('Course', courseSchema)
async function addCourse(){
const newCourse= await Course({
tags:null,
category:'web',
name:'Xamarin for Beginners',
author:'Silvia',
isPublished:true,
price:15
})
await newCourse.save().catch(err=>console.log("Error",err.message))
}
addCourse()<file_sep>/RESTful/routes/home.js
const express=require('express');
const router=express.Router()
const courses=[{id:1,name:"course1"},{id:2,name:"course2"},{id:3,name:"course3"}];
router.get('/',(req, res)=>{
/*
request argument has many options
*/
res.send(courses);
})
module.exports=router<file_sep>/RESTful/routes/courses.js
const express = require("express");
const router = express.Router();
const courses = [
{ id: 1, name: "course1" },
{ id: 2, name: "course2" },
{ id: 3, name: "course3" }
];
router.get("/", (req, res) => {
/*
request argument has many options
*/
res.send(courses);
});
//EXAMPLE 3
//Getting specific items with id
//request has a params property that hold all the parameters in the querystring
router.get("/:id", (req, res) => {
//below:
//courses.find will loop through the array and return true when the param passed matches
//an 'id' in the courses array.
//req.params return a strin so we use the parseInt method to make it an Int value
const course = courses.find(item => item.id === parseInt(req.params.id));
//catch any error if id is not found. set the status of the response and also a message
//can be appended.
if (!course)
return res.status(404).send("The course with the given id was not found.");
res.send(course);
});
//EXAMPLE 4
//it is possible to have multiple parameters in one route
//access an object with all parameters passed using "req.params" or
//access a specific param from the object by using "req.params" and the name of the param "req.params.year"
router.get("//:year/:month", (req, res) => {
res.send(req.params);
});
//EXAMPLE 5
//We can also read queryString parameter wich are identified by after a "?" in the url
//for ex: user sortBy=name to sort them by name
//to read queryString parameters use ".query" instead of params
router.get("/:year/:month", (req, res) => {
res.send(req.query);
});
/*
////// /// /// ////////////
// // // // // // //
// // // // /// //
// // // // //
// // ///// //
ADD "joi" package to add validation to your api.
*/
//ALWAYS VALIDATE THE INPUT!!!
router.post("/api/courses", (req, res) => {
//WITH JOI FIRST WEE NEED TO DEFINE A SCHEMA which mean which properties we will have.
/************Start here code was refactored to use a recyclable fucntion to validate *****************
const schema={
//define the properies expected
//call the joi variable "Joi" then set the data type ".string" then the min length if applies ".min(3)"
//and if required
name:Joi.string().min(3).required()
}
//to validate call the variable where joi is stored "Joi" then call the "validate" method
//takes 2 arguments the body of the request and the schema if body matches schema request is good
const joiResult=Joi.validate(req.body, schema)
//store the params passed in an object
//NOTE: id will be automatically created when dealing with real db
//use req.body to get the body of the request plus the property you want to read ex:"req.body.name"
if(joiResult.error){
console.log(joiResult.error.details[0].message)
// 400 codes are used for bad requests.
//SEND send the response back to the client
res.status(400).send(joiResult.error.details)
return;
}***********************Ends Here******************************************************/
const { error } = validateCourse(req.body);
if (error) return res.status(400).send(error.details[0].message);
console.log(error);
const newCourse = {
id: courses.length + 1,
name: req.body.name
};
//push object to the array.
//in read dv you would send the data to the db here.
courses.push(newCourse);
//when posting it is good practice to return the body of the object
res.send(newCourse);
});
/*
////// // // ////////////
// // // // //
// // // // //
// // // //
// // //
*/
//put method need the specific route for the course to change
router.put("/:id", (req, res) => {
//Look for the course
const course = courses.find(item => item.id === parseInt(req.params.id));
console.log("This is course", course);
//if courses does not exists return 404
if (!course)
return res.status(404).send("The course was not found, could no update.");
//if it exists validate data by passing to a recyclable function to validate
const { error } = validateCourse(req.body); //use object destruction to get the error only.
if (error) {
res.status(400).send(error.details[0].message);
return;
}
//Update the code
course.name = req.body.name;
res.send(course);
//return the updated course
});
/*
////// ///// //
// // // //
// // //// //
// // // //
///// ///// //////
*/
router.delete("/:id", (req, res) => {
const course = courses.find(item => item.id === parseInt(req.params.id));
if (!course) return res.status(404).send("The course could not be found");
//to delete find the index of the course that's stored in the course constant
const index = courses.indexOf(course);
console.log("This is index", index);
//splice deletes 1 item from the courses array at the passed index
courses.splice(index, 1);
res.send(course);
});
function validateCourse(course) {
const schema = {
name: Joi.string()
.min(3)
.required()
};
return Joi.validate(course, schema);
}
module.exports = router;
<file_sep>/RESTful/app.js
/*
install use nodemon <filename> to automatically listen on the port
install joi to add easy validation
install helmet to set up http headers
install debug to quickly debug you application
install morgan to log http request (NOTE: morgan is not necesary and logging each request will delay the
request processing time. try not to use in production.)
Create 'config' folder and require it to set the seetings of the app in different build modes
*/
//require express and joi from node_modules
const config=require('config')
const express= require('express')
const Joi = require('joi')
const logger=require('./middleware/logger')
const helmet=require('helmet');
const morgan=require('morgan');
const authenticate = require('./authenticator')
//Moved all routes with '/api/courses/' to separate file
//in order to use them we need to require them from the courses file and the tell express to use then
//see line 46 app.use('/api/courses', courses)
const courses=require('./routes/courses')
const home=require('./routes/home')
//use debug you need to store it in a constant and pass in a name for the name space where these
//log messages will be stored. you will also need to create a environment variable with that name
const debug=require('debug')('app:debug');
// create an instance of the express class and store in variable
const app = express()
/*These two approaches get the type of environment the project is set to (production, development, etc..)
NOTE: app.get will return a default value of development 'NODE_ENV' is undefined.
console.log(`NODE_ENV: ${process.env.NODE_ENV}`)
console.log(`app: ${app.get('env')}`);
*/
// enable parsing of json objects. this feature is disabled by default
//if there is a json object in the request this will add the json object to the "req" argument and the json
//can be accessed as "req.body"
//app.use tells express to use these pieces of middleware passed to the function ".use()"
app.use(express.json())
//this is used if you have a url encoded request "thorough urlquesryStrings."
//extended prop needs to be turned to true to allow arrays and object in the body of the request
// app.use(express.urlencoded({extended:true}))
app.use(express.static('public'))
app.use(logger)
app.use(authenticate)
app.use(helmet());
//in order to use the other module. we need to tell express to use this route and pass it two arguments
//the route used '/api/courses' and the constant where the route is stored 'course'
//for any route that is using that end point ex:'api/courses' use this router ex: courses
app.use('/api/courses',courses)
app.use('/',home)
//CONFIGURATION*********************************************
debug('Application Name: ',config.get('name'))
debug('Mail Server: ',config.get('mail.host'))
debug('Mail Password: ',config.get('mail.password'))
//tiny is the format of the log you want to see. 'tiny' is the shortest format
//you can use the value returned from "NODE_ENV" to make a conditional
if(app.get('env')==='development'){
app.use(morgan('tiny'));
debug('Morgan Enabled')
}
// const courses=[{id:1,name:"course1"},{id:2,name:"course2"},{id:3,name:"course3"}];
/*
Add a listener to listen for the request
listen takes 2 parameters:
First: a port to listen
Second is optional: a function that will be called when the listener is triggered
*/
/* Avoid hardcoding the value of the port use an
enviroment variable to set the port number*/
//use the 'process' object with the property "env" for for enviroment variable
//then add the name of the enviroment variable "PORT" you need to make this evironment variable using
//the following command on the terminal
// export <Environment Variable Name>=<Number Value>
const port = process.env.PORT || 3000;
app.listen(port,()=>{
console.log(`Listening on port ${port}`)
})
<file_sep>/mongo-db/exercise3.js
const mongoose=require('mongoose')
mongoose.connect('mongodb://localhost:27017/mongo-exercises',{useNewUrlParser:true})
const courseSchema= new mongoose.Schema({
author:String,
price:Number,
date:{type:Date, default:Date.now},
isPublished:Boolean,
name:String,
tags:[String]
})
const Course=mongoose.model('Course', courseSchema)
async function getCourses(){
return await Course
.find({isPublished:true})
.or([{name:/.*by.*/}, {price:{$gte:15}}])
}
async function showCourses(){
const courses=await getCourses()
console.log(courses)
}
showCourses() | 45da02a281f596dfe36b853f56340c97205f6ef4 | [
"JavaScript"
] | 14 | JavaScript | guer0157/nodejs | ac98a3aea16c5a450ac5807f755890072e40e444 | 49024cae198dfa73ba508888de8df8737cadeb6c |
refs/heads/main | <repo_name>bharat123-anki/directory_user_interview<file_sep>/application/controllers/User.php
<?php
defined('BASEPATH') or exit('No direct script access allowed');
require("App.php");
class User extends CI_Controller
{
/**
* Index Page for this controller.
*
* Maps to the following URL
* http://example.com/index.php/welcome
* - or -
* http://example.com/index.php/welcome/index
* - or -
* Since this controller is set as the default controller in
* config/routes.php, it's displayed at http://example.com/
*
* So any other public methods not prefixed with an underscore will
* map to /index.php/welcome/<method_name>
* @see https://codeigniter.com/user_guide/general/urls.html
*/
public function __construct()
{
parent::__construct();
$this->load->model('DirectoryInfo_model');
}
public function index()
{
$this->load->view('add_user');
}
public function add($value = '')
{
$response = array('status' => 500, 'msg' => 'Some Internal Error');
$required = ['name', 'email', 'password'];
$all_good = 1;
foreach ($required as $key => $val) {
if (empty($_POST[$val])) {
$response = ['status' => 201, 'field' => $val, 'msg' => 'Field Is Required'];
$all_good = 0;
echo json_encode($response);
exit;
}
}
if ($all_good) {
$name = $this->input->post('name');
$email = $this->input->post('email');
$password = $this->input->post('password');
$dataStatus = 'New';
if (isset($id) && !empty($id)) {
$dataStatus = 'Update';
} else {
$this->db->insert('users', array('name' => $name, 'email' => $email, 'password' => md5($password)));
}
if ($this->db->affected_rows() > 0) {
$disp_msg = "Data Added Sucessfully You Will Redirect To Login Page Shortly";
if ($dataStatus != "New") {
$disp_msg = "Data Updated Sucessfully";
}
$response = array('status' => 200, 'msg' => $disp_msg);
}
}
echo json_encode($response);
}
}
<file_sep>/application/views/login/login_footer.php
<!-- /.login-box -->
<script src="<?php echo base_url() ?>plugins/jquery/jquery.min.js"></script>
<!-- jQuery UI 1.11.4 -->
<script src="<?php echo base_url() ?>plugins/jquery-ui/jquery-ui.min.js"></script>
<!-- jQuery -->
<!-- Bootstrap 4 -->
<!-- AdminLTE App -->
</body>
</html><file_sep>/application/views/user/user.php
<?php $this->load->view('template/header.php'); ?>
<body class="hold-transition skin-blue sidebar-mini">
<div class="wrapper">
<?php $this->load->view('template/navigation.php') ?>
<?php $this->load->view('template/sidebar.php'); ?>
<!-- Left side column. contains the logo and sidebar -->
<!-- Content Wrapper. Contains page content -->
<div class="content-wrapper">
<!-- Content Header (Page header) -->
<div class="content-header">
<div class="container-fluid">
<div class="row mb-2">
<div class="col-sm-6">
<h1 class="m-0 text-dark">Directory</h1>
</div><!-- /.col -->
<div class="col-sm-6">
<ol class="breadcrumb float-sm-right">
<li class="breadcrumb-item"><a href="javascript:void(0)" class="btn btn-primary add_candidate_form">Add Directory</a></li>
</ol>
</div><!-- /.col -->
</div><!-- /.row -->
</div><!-- /.container-fluid -->
</div>
<!-- Main content -->
<section class="content">
<!-- Info boxes -->
<div class="card">
<div class="card-header">
<h3 class="card-title">Search Form</h3>
<!-- /.card-tools -->
</div>
<!-- /.card-header -->
<div class="card-body">
<div id="search_form_dash"> </div>
</div>
<!-- /.card-body -->
<!-- /.card-footer -->
</div>
<div class="card">
<div class="card-header">
<h3 class="card-title">Directory Listings</h3>
<div class="card-tools">
<!-- Buttons, labels, and many other things can be placed here! -->
<!-- Here is a label for example -->
</div>
<!-- /.card-tools -->
</div>
<!-- /.card-header -->
<div class="card-body">
<div class="table-responsive-sm table-responsive-md table-responsive">
<!-- begin: Datatable -->
<table class="table table-striped- table-bordered table-hover table-checkable directory_data" id="directory_data">
<thead>
<tr>
<th scope="col">#</th>
<th scope="col">First Name</th>
<th scope="col">Middle Name</th>
<th scope="col">Last Name</th>
<th scope="col">Email</th>
<th scope="col">Mobile</th>
<th scope="col">Landline</th>
<th scope="col">Image</th>
<th scope="col">Notes</th>
<th scope="col">Created</th>
<th scope="col">Action</th>
</tr>
</thead>
</table>
<!--end: Datatable -->
</div>
<!--
<div id="directory_data">
</div> -->
</div>
<!-- /.card-body -->
<div class="card-footer">
</div>
<!-- /.card-footer -->
</div>
<!-- /.row -->
</section>
<div class="container-fluid">
<!-- Small boxes (Stat box) -->
<div id="addDirectoryModal" class="modal fade" tabindex="-1">
<div class="modal-dialog modal-lg">
<div class="modal-content">
<div class="modal-header">
<h5 class="modal-title"> User Info</h5>
<button type="button" class="close" data-dismiss="modal">×</button>
</div>
<div class="modal-body">
<div id="modalData"></div>
</div>
<div class="modal-footer">
<button type="button" class="btn btn-secondary" data-dismiss="modal">Cancel</button>
</div>
</div>
</div>
</div>
</div><!-- /.container-fluid -->
<!-- /.content -->
</div>
<!-- /.content-wrapper -->
<!-- Control Sidebar -->
<!-- /.control-sidebar -->
<!-- Add the sidebar's background. This div must be placed
immediately after the control sidebar -->
<div class="control-sidebar-bg"></div>
</div>
<!-- ./wrapper -->
<?php $this->load->view('template/footer.php'); ?>
</body>
</html>
<script src="<?php echo base_url() ?>assets/js/custom.js"></script>
<script type="text/javascript">
$(document).ready(function() {
getSerchFormOnUser()
$('#directory_data').dataTable({
searching: false,
processing: true,
serverSide: true,
ajax: 'getAllDirectoryInfoDatatable',
columnDefs: [{
"render": function(data, type, row) {
// console.log(row)
// return 'bharat';
return '<a href="javascript:void(0)" class="candidateEditdata" data-id="' + data + '"><i class="fa fa-edit"></i> </a>|<a href="javascript:void(0)" class="candidatedeletedata" data-id="' + data + '"><i class="fa fa-trash"></i></a>|<a href="javascript:void(0)" class="candidateviewDirectory" data-id="' + data + '"><i class="fa fa-eye"></i></a>'
},
"targets": 10,
},
{
"render": function(data, type, row) {
// if(empty(data))
if (data == "" || data == "undefined")
return '<p style="white-space: nowrap;">No Image Found</p>';
var path = '<?php echo base_url('uploads/user_images'); ?>';
data = '<a href="' + path + '/' + data + '" target="_blank" >Image</a>';
// return 'bharat';
return data
},
"targets": 7,
},
],
})
})
$('body').on('submit', '#searchDistributorForm', function(e) {
e.preventDefault();
var table = $('#directory_data').dataTable();
var passData = "?" + $('#searchDistributorForm').serialize();
table.fnReloadAjax("getAllDirectoryInfoDatatable" + passData);
// getAllDirectoryInfo(formData)
})
</script><file_sep>/application/views/user/search_form_dash.php
<form action="" method="POST" id="searchDistributorForm">
<div class="row">
<div class="form-group col-md-4">
<label for="exampleInputEmail1">Name</label>
<select name="first_name" class="form-control f_name" id="f_name">
<option value="">--Select Any One--</option>
<?php foreach (array_unique($first_name) as $key => $value) { ?>
<option value="<?php echo $value ?>"><?php echo $value; ?></option>
<?php } ?>
</select>
</div>
<div class="form-group col-md-4">
<label for="exampleInputEmail1">Telephone No</label>
<select name="mobile_no" class="form-control mobile_no" id="mobile_no">
<option value="">--Select Any One--</option>
<?php foreach (array_unique($mobile_no) as $key => $value) { ?>
<option value="<?php echo $value ?>"><?php echo $value; ?></option>
<?php } ?>
</select>
</div>
<div class="form-group col-md-4">
<label for="exampleInputEmail1"> </label>
<label for="exampleInputEmail1"> </label>
<label for="exampleInputEmail1"> </label>
<input type="submit" class=" btn btn-primary" />
</div>
</div>
</form>
<script type="text/javascript">
$(document).ready(function() {
$('.mobile_no, .f_name').select2();
})
</script><file_sep>/application/views/user/user_add_modal.php
<div class="main_notify"></div>
<form action="" method="POST" id="candidate_add_form" enctype="multipart/form-data">
<input type="hidden" name="id" value="<?php echo isset($directory_data['id']) ? $directory_data['id'] : ''; ?>">
<div class="row">
<div class="col-md-6">
<div class="form-group">
<label for="exampleInputEmail1">First Name</label>
<input type="text" class="form-control" id="first_name" aria-describedby="emailHelp" name="first_name" value="<?php echo isset($directory_data['first_name']) ? $directory_data['first_name'] : ''; ?>" placeholder="Enter First name">
</div>
</div>
<div class="col-md-6">
<div class="form-group">
<label for="exampleInputEmail1">Middle Name</label>
<input type="text" class="form-control" id="middle_name" aria-describedby="emailHelp" name="middle_name" value="<?php echo isset($directory_data['middle_name']) ? $directory_data['middle_name'] : ''; ?>" placeholder="Enter Middle name">
</div>
</div>
</div>
<div class="row">
<div class="col-md-6">
<div class="form-group">
<label for="exampleInputEmail1">Last Name</label>
<input type="text" class="form-control" id="last_name" aria-describedby="emailHelp" name="last_name" value="<?php echo isset($directory_data['last_name']) ? $directory_data['last_name'] : ''; ?>" placeholder="Enter Last name">
</div>
</div>
<div class="col-md-6">
<div class="form-group">
<label for="exampleInputEmail1">Email</label>
<input type="text" class="form-control" id="email" aria-describedby="emailHelp" name="email" value="<?php echo isset($directory_data['email']) ? $directory_data['email'] : ''; ?>" placeholder="Enter Last name">
</div>
</div>
</div>
<div class="row">
<div class="col-md-6">
<div class="form-group">
<label for="exampleInputEmail1">Mobile No</label>
<input type="text" class="form-control" id="mobile_no" aria-describedby="emailHelp" name="mobile_no" value="<?php echo isset($directory_data['mobile_no']) ? $directory_data['mobile_no'] : ''; ?>" placeholder="Enter Last name">
</div>
</div>
<div class="col-md-6">
<div class="form-group">
<label for="exampleInputEmail1">Landline No</label>
<input type="text" class="form-control" id="landline_no" aria-describedby="emailHelp" name="landline_no" value="<?php echo isset($directory_data['landline_no']) ? $directory_data['landline_no'] : ''; ?>" placeholder="Enter Last name">
</div>
</div>
</div>
<div class="form-group">
<label for="exampleInputEmail1">Notes</label>
<textarea name="notes" class="form-control"><?php echo isset($directory_data['notes']) ? $directory_data['notes'] : ''; ?></textarea>
</div>
<div class="form-group">
<label for="exampleInputEmail1">Candidate Image</label>
<input type="file" name="user_image" class="form-control">
</div>
<input type="submit" class="btn btn-primary"></input>
</div>
</form><file_sep>/application/controllers/DirectoryInfo.php
<?php
defined('BASEPATH') or exit('No direct script access allowed');
require("App.php");
class DirectoryInfo extends APP_Controller
{
/**
* Index Page for this controller.
*
* Maps to the following URL
* http://example.com/index.php/welcome
* - or -
* http://example.com/index.php/welcome/index
* - or -
* Since this controller is set as the default controller in
* config/routes.php, it's displayed at http://example.com/
*
* So any other public methods not prefixed with an underscore will
* map to /index.php/welcome/<method_name>
* @see https://codeigniter.com/user_guide/general/urls.html
*/
public function __construct()
{
parent::__construct();
$this->load->model('DirectoryInfo_model');
}
public function index()
{
$this->load->view('user/user');
}
public function getDirectoryAddModal()
{
if ($this->input->is_ajax_request()) {
$id = $this->input->post('id');
$directory_data = [];
if (!empty($id)) {
$directory_data = $this->DirectoryInfo_model->getDirectoryDataById($id);
}
$data['directory_data'] = $directory_data;
$this->load->view('user/user_add_modal', $data);
}
}
public function directoryAdd()
{
$response = array('status' => 500, 'msg' => 'Some Internal Error');
$required = ['first_name', 'last_name'];
$proceed = 1;
// print_r($_POST);
foreach ($required as $key => $val) {
if (empty($_POST[$val])) {
$data = ['field' => $val, 'msg' => 'Field Is Required'];
$response['status'] = 201;
$response['msg'] = "No Fields";
$response['err'][] = $data;
$proceed = 0;
}
}
if ($proceed) {
$id = $this->input->post('id');
$fname = $this->input->post('first_name');
$middle_name = $this->input->post('middle_name');
$last_name = $this->input->post('last_name');
$email = $this->input->post('email');
$mobile_no = $this->input->post('mobile_no');
$landline_no = $this->input->post('landline_no');
$notes = $this->input->post('notes');
$session_id = ($this->session->userdata('logged_session')['id']);
if (!empty($email)) {
if (!filter_var($email, FILTER_VALIDATE_EMAIL)) {
$response['status'] = 201;
$response['err'][] = ['field' => 'email', 'msg' => 'Please Enter Valid Email'];
echo json_encode($response);
exit;
}
}
if (!empty($mobile_no)) {
if (!preg_match('/^[0-9]{10}+$/', $mobile_no)) {
$response['status'] = 201;
$response['err'][] = ['field' => 'mobile_no', 'msg' => 'Please Enter Valid Mobile No'];
echo json_encode($response);
exit;
}
}
if (!empty($landline_no)) {
if ($this->validate_phone_number($landline_no) === false) {
$response['status'] = 201;
$response['err'][] = ['field' => 'landline_no', 'msg' => 'Please Enter Valid Landline No'];
echo json_encode($response);
exit;
}
}
$type = "New";
if (isset($id) && !empty($id)) {
$type = "Update";
}
// print_r($type);
$image_path = "";
if (!empty($_FILES['user_image']['name'])) {
$config['upload_path'] = './uploads/user_images/';
$config['allowed_types'] = 'jpg|png|jpeg';
$config['max_size'] = 1500;
$config['max_width'] = 2000;
$config['max_height'] = 2000;
$this->load->library('upload', $config);
if (!$this->upload->do_upload('user_image')) {
$response = ['status' => 203, 'field' => 'user_image', 'msg' => $this->upload->display_errors()];
echo json_encode($response);
exit;
} else {
$path = $this->upload->data('file_name');
$image_path = $path;
}
}
// exit;
if ($type == "New") {
$this->db->insert('directory_info', array('first_name' => $fname, 'user_id' => $session_id, 'middle_name' => $middle_name, 'last_name' => $last_name, 'email' => $email, 'mobile_no' => $mobile_no, 'landline_no' => $landline_no, 'user_image_path' => $image_path, 'notes' => $notes, 'created_by' => $session_id));
} else {
// for image manipulation
$existing_user = $this->DirectoryInfo_model->getDirectoryDataById($id);
$image_path_existing = ($existing_user['user_image_path']);
if (empty($image_path)) {
$image_path_to_update = $image_path_existing;
} else {
$image_path_to_update = $image_path;
if (!empty($image_path_existing)) {
$path = FCPATH . 'uploads/user_images/';
$file_name = $path . $image_path_existing;
unlink($file_name);
}
}
$this->db->where('id', $id);
$this->db->update('directory_info', array('first_name' => $fname, 'middle_name' => $middle_name, 'last_name' => $last_name, 'email' => $email, 'mobile_no' => $mobile_no, 'landline_no' => $landline_no, 'user_image_path' => $image_path_to_update, 'notes' => $notes, 'updated_by' => $session_id));
}
// print_r($this->db->last_query());
$disp_msg = "Data Added Sucessfully";
if ($type != "New") {
$disp_msg = "Data Updated Sucessfully";
}
if ($this->db->affected_rows() > 0) {
$response = array('status' => 200, 'msg' => $disp_msg);
} else {
$response = array('status' => 200, 'msg' => $disp_msg);
}
}
echo json_encode($response);
}
public function validate_phone_number($landline = '')
{
if (!preg_match("/^[0-9]{11}/", $landline)) {
return false;
} else {
return true;
}
}
public function deleteCandidateData($value = '')
{
$response = array('status' => 500, 'msg' => 'Some Internal Error');
$id = $this->input->post('id');
if (!empty($id)) {
$session_id = ($this->session->userdata('logged_session')['id']);
$this->db->where('id', $id);
$this->db->update(
'directory_info',
array('is_deleted' => 1, 'updated_by' => $session_id),
);
$response = array('status' => 200, 'msg' => 'Data Deleted Sucessfully');
}
echo json_encode($response);
}
public function getViewDirectory()
{
if ($this->input->is_ajax_request()) {
$id = $this->input->post('id');
$directory_data = [];
if (!empty($id)) {
// $increasecount = $this->DirectoryInfo_model->increaseviewCount($id);
$directory_data = $this->DirectoryInfo_model->getDirectoryDataById($id);
}
$data['directory_data'] = $directory_data;
$this->load->view('user/user_view_modal', $data);
}
}
public function getAllDirectoryInfoDatatable()
{
$request = $_REQUEST;
// echo "<pre>";
// print_r($request);
$start = $request['start'] ? (int) $request['start'] : (int) 0;
$length = $request['length'] ? (int) $request['length'] : (int) 0;
$searchFilter = array(
'first_name' => array('type' => 'text', 'value' => isset($request['first_name']) ? $request['first_name'] : ''),
'mobile_no' => array('type' => 'text', 'value' => isset($request['mobile_no']) ? $request['mobile_no'] : ''),
);
// print_r($searchFilter);
// print_r($_REQUEST);
$order = [];
$orderBy = $request['order'] ? $request['order'] : array();
if (!empty($orderBy)) {
$columns = array('', 'first_name', 'middle_name', 'last_name', 'email', 'mobile_no', 'landline_no', '', '', 'created_at', '');
$orderBy = $orderBy[0];
$columnIndex = $orderBy['column'];
if (array_key_exists($columnIndex, $columns)) {
if (!empty($columns[$columnIndex])) {
$column = $columns[$columnIndex];
$order['order_by'] = $column;
$order['order_type'] = $orderBy['dir'];
} else {
$order['order_by'] = 'created_at';
$order['order_type'] = 'Asc';
}
} else {
$order['order_by'] = 'created_at';
$order['order_type'] = 'Asc';
}
}
$user_id = ($this->session->userdata('logged_session')['id']);
$rows = $this->DirectoryInfo_model->getAllDirecetoryInfoData($user_id, $searchFilter, array($start, $length), $order);
// print_r($rows);
$noOfRecords = count($rows);
$data = array();
if (count($rows)) {
$count = 1;
foreach ($rows as $row) {
$created_at = date("d/m/Y", strtotime($row['created_at']));;
$data[] = array(
$count++,
$row['first_name'],
$row['middle_name'],
$row['last_name'],
$row['email'],
$row['mobile_no'],
$row['landline_no'],
$row['user_image_path'],
$row['notes'],
$created_at,
$row['id'],
);
}
}
$directory_info = array(
'draw' => (int) $request['draw'] ? (int) $request['draw'] : (int) 0,
'recordsTotal' => ($noOfRecords),
'recordsFiltered' => ($noOfRecords),
'data' => $data,
);
$directory_info = json_encode($directory_info);
echo ($directory_info);
// return ($directory_info);
// listing view page open kar
}
public function getSearchViewDash()
{
$user_id = ($this->session->userdata('logged_session')['id']);
$result = $this->DirectoryInfo_model->getSearchFormData($user_id);
$first_name = array_column($result, 'first_name');
$mobile_no = array_column($result, 'mobile_no');
$this->load->view('user/search_form_dash', array('first_name' => $first_name, 'mobile_no' => $mobile_no));
}
}
<file_sep>/application/controllers/Welcome.php
<?php
defined('BASEPATH') or exit('No direct script access allowed');
class Welcome extends CI_Controller
{
/**
* Index Page for this controller.
*
* Maps to the following URL
* http://example.com/index.php/welcome
* - or -
* http://example.com/index.php/welcome/index
* - or -
* Since this controller is set as the default controller in
* config/routes.php, it's displayed at http://example.com/
*
* So any other public methods not prefixed with an underscore will
* map to /index.php/welcome/<method_name>
* @see https://codeigniter.com/user_guide/general/urls.html
*/
public function __construct()
{
parent::__construct();
}
public function index()
{
$this->load->view('login/login');
}
public function checkUserInfo()
{
if ($this->input->is_ajax_request()) {
$response = array('status' => 500, 'msg' => 'Some Internal Error');
$email = $this->input->post('email');
$password = $this->input->post('<PASSWORD>');
$required = ['email', 'password'];
// print_r($_POST);
$proceed = 1;
foreach ($required as $key => $val) {
if (empty($_POST[$val])) {
$data = ['field' => $val, 'msg' => 'Field Is Required'];
$response['status'] = 201;
$response['msg'] = "No Fields";
$response['err'][] = $data;
$proceed = 0;
// exit;
}
}
if ($proceed) {
if (!filter_var($email, FILTER_VALIDATE_EMAIL)) {
$response = array('status' => 202, 'msg' => 'Invalid Email');
} else {
$this->load->model('User_model');
$return_query = $this->User_model->checkEmailAndPassword($email, $password);
if (empty($return_query)) {
$response = array('status' => 203, 'msg' => 'Invalid User');
} else {
$this->session->set_userdata('logged_session', $return_query);
$response = array('status' => 200, 'msg' => 'Success You Will Redirect Shortly', 'data' => $return_query);
}
}
}
echo json_encode($response);
}
}
public function logout($value = '')
{
$this->session->unset_userdata('logged_session');
$this->load->view('login/login');
}
}
<file_sep>/directory_interview.sql
-- phpMyAdmin SQL Dump
-- version 4.8.4
-- https://www.phpmyadmin.net/
--
-- Host: 127.0.0.1
-- Generation Time: Jun 01, 2021 at 02:32 PM
-- Server version: 10.1.37-MariaDB
-- PHP Version: 7.3.1
SET SQL_MODE = "NO_AUTO_VALUE_ON_ZERO";
SET AUTOCOMMIT = 0;
START TRANSACTION;
SET time_zone = "+00:00";
/*!40101 SET @OLD_CHARACTER_SET_CLIENT=@@CHARACTER_SET_CLIENT */;
/*!40101 SET @OLD_CHARACTER_SET_RESULTS=@@CHARACTER_SET_RESULTS */;
/*!40101 SET @OLD_COLLATION_CONNECTION=@@COLLATION_CONNECTION */;
/*!40101 SET NAMES utf8mb4 */;
--
-- Database: `directory_interview`
--
-- --------------------------------------------------------
--
-- Table structure for table `directory_info`
--
CREATE TABLE `directory_info` (
`id` int(11) NOT NULL,
`user_id` int(11) NOT NULL,
`first_name` varchar(50) NOT NULL,
`middle_name` varchar(50) DEFAULT NULL,
`last_name` varchar(50) DEFAULT NULL,
`user_image_path` text,
`email` varchar(60) DEFAULT NULL,
`mobile_no` bigint(20) DEFAULT NULL,
`landline_no` varchar(20) DEFAULT NULL,
`notes` text,
`view_count` longtext,
`is_deleted` int(11) NOT NULL DEFAULT '0',
`created_by` int(11) DEFAULT NULL,
`created_at` datetime DEFAULT CURRENT_TIMESTAMP,
`updated_by` int(11) DEFAULT NULL,
`updated_at` datetime DEFAULT '0000-00-00 00:00:00' ON UPDATE CURRENT_TIMESTAMP
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
--
-- Dumping data for table `directory_info`
--
INSERT INTO `directory_info` (`id`, `user_id`, `first_name`, `middle_name`, `last_name`, `user_image_path`, `email`, `mobile_no`, `landline_no`, `notes`, `view_count`, `is_deleted`, `created_by`, `created_at`, `updated_by`, `updated_at`) VALUES
(3, 1, 'Bharat', 'JAGDAMBA', 'Yadav', '', '<EMAIL>', 909090909, NULL, NULL, '', 1, 1, '2021-04-29 18:17:37', 1, '2021-05-28 17:37:54'),
(4, 2, 'Ankita', 'Bharat', 'YAdav', '', NULL, NULL, NULL, NULL, '', 0, 1, '2021-04-29 18:17:42', 1, '2021-05-28 17:37:54'),
(5, 2, 'Pajav', 'Singh', 'Ture', '\r\n', '<EMAIL>', 9090909090, NULL, NULL, '', 0, 1, '2021-04-29 18:18:18', NULL, '2021-05-28 17:37:54'),
(6, 1, 'milind', 'Ganatra', 'Pravin', 'Screenshot_(21).png', '<EMAIL>', 8879715444, '', NULL, '', 1, 1, '2021-04-29 18:21:34', 1, '2021-06-01 15:55:36'),
(7, 2, 'Oiyush', 'Salunkhe', 'qwety', '', '<EMAIL>', NULL, NULL, NULL, '', 1, 1, '2021-04-29 18:22:11', 1, '2021-05-28 17:37:54'),
(8, 2, 'No image no', '3', '3', '', NULL, NULL, NULL, NULL, '', 1, 1, '2021-04-29 23:51:51', 1, '2021-05-28 17:37:54'),
(10, 1, 'Mirchi', 'Rajkamal', 'Yadav', 'Screenshot_(32).png', '<EMAIL>', 9090909090, '4567', NULL, '', 0, 1, '2021-05-28 15:34:08', 1, '2021-05-28 17:37:54'),
(11, 1, 'Darshini', 'Chauhan', 'Bharta', 'Screenshot_(29).png', '', 0, '', '', '', 1, 1, '2021-05-28 15:37:09', 1, '2021-06-01 15:53:46'),
(12, 1, 'darshini', 'rajkamal', 'yadav', 'download1.jpg', '<EMAIL>', 9099098977, '', 'hello', '', 0, 1, '2021-05-11 15:49:55', 1, '2021-06-01 16:27:29'),
(13, 5, 'aatish', 'harishf', 'aat', 'Screenshot_(24).png', '<EMAIL>', 9099098903, '', '', '', 0, 5, '2021-05-27 16:15:54', 5, '2021-06-01 17:42:06'),
(14, 5, 'checklead qqqqqqqqqqqqqqqqqqqqq', 'lead', 'leading upd', 'Screenshot_(39).png', '<EMAIL>', 1232123234, '4545545', NULL, '', 0, 5, '2021-05-28 16:16:26', 5, '2021-05-28 17:37:54'),
(15, 5, 'checklead qqqqqqqqqqqqqqqqqqqqq', 'lead', 'leading', '', '<EMAIL>', 8989767878, '', NULL, '', 1, 5, '2021-05-26 16:18:41', 5, '2021-06-01 17:09:11'),
(16, 5, 'charmi', 'pravin', 'ganatra', '', '<EMAIL>', 8097172307, '', NULL, '', 0, 5, '2021-05-30 16:50:44', NULL, '2021-06-01 17:05:59'),
(17, 5, 'ajinkya', 'rahane', 'shah', '', '<EMAIL>', 9099098903, '', 'hhh', '', 1, 5, '2021-05-28 16:54:49', 5, '2021-06-01 17:08:52'),
(18, 5, 'charmi.s', 'pravin', 'ganatra', '', '<EMAIL>', 8097172307, '123456', NULL, '', 0, 5, '2021-05-28 16:57:34', 5, '2021-05-28 17:37:54'),
(19, 6, 'mayuri', 'bhaarat', 'Yadav', 'Screenshot_(32)2.png', '<EMAIL>', 9999995454, '123456789', 'Hello <NAME>', NULL, 0, 6, '2021-05-28 20:44:29', 6, '2021-05-28 20:45:01'),
(20, 6, 'vishaka', 'smitesh', 'Yadav', '', '<EMAIL>', 1234567890, 'hhhh', 'yuiop[', NULL, 0, 6, '2021-05-28 21:07:08', NULL, '0000-00-00 00:00:00'),
(21, 1, 'genuine', 'test', 'user', '', '', 0, '', '', NULL, 1, 1, '2021-06-01 11:26:36', 1, '2021-06-01 16:09:17'),
(22, 1, 'genuines', 'user', 'info', 'download.jpg', '<EMAIL>', 1233213456, '', '', NULL, 1, 1, '2021-06-01 11:29:24', 1, '2021-06-01 15:57:44'),
(23, 9, 'tanvi', 'mayur', 'Deshpand', 'Screenshot_(27)1.png', '<EMAIL>', 9090909090, '312345686789', 'ff', NULL, 0, 9, '2021-06-01 16:51:49', 9, '2021-06-01 17:03:25'),
(24, 10, 'tarun', 'taru', 'taaa', 'Screenshot_(30).png', '<EMAIL>', 1234567890, '12345678908', 'hello', NULL, 0, 10, '2021-06-01 17:50:28', 10, '2021-06-01 17:52:54'),
(25, 10, 'checklead qqqqqqqqqqqqqqqqqqqqq', 'lead', 'leading', 'Screenshot_(14)1.png', '<EMAIL>', 1234567890, '', '', NULL, 0, 10, '2021-06-01 17:51:23', 10, '2021-06-01 17:52:36'),
(26, 10, 'laaaaaa', 'qqqqqqq', 'leading', '', '<EMAIL>', 9123456789, '', '', NULL, 0, 10, '2021-06-01 17:56:36', 10, '2021-06-01 17:56:46');
-- --------------------------------------------------------
--
-- Table structure for table `users`
--
CREATE TABLE `users` (
`id` int(11) NOT NULL,
`name` varchar(100) NOT NULL,
`email` varchar(100) NOT NULL,
`password` varchar(200) NOT NULL,
`created_by` int(11) NOT NULL,
`created_at` datetime NOT NULL DEFAULT CURRENT_TIMESTAMP,
`updated_by` int(11) NOT NULL,
`updated_at` datetime NOT NULL DEFAULT '0000-00-00 00:00:00' ON UPDATE CURRENT_TIMESTAMP
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
--
-- Dumping data for table `users`
--
INSERT INTO `users` (`id`, `name`, `email`, `password`, `created_by`, `created_at`, `updated_by`, `updated_at`) VALUES
(1, 'Bharat', '<EMAIL>', '<PASSWORD>', 1, '2021-04-28 15:58:41', 0, '2021-04-29 11:24:01'),
(2, 'testing', '<EMAIL>', '<PASSWORD>', 0, '2021-04-30 18:15:40', 0, '0000-00-00 00:00:00'),
(5, 'bhar', '<EMAIL>', '<PASSWORD>', 0, '2021-05-01 16:53:42', 0, '2021-06-01 17:05:03'),
(6, 'Mayuri', '<EMAIL>', '<PASSWORD>', 0, '2021-05-28 20:39:10', 0, '0000-00-00 00:00:00'),
(9, 'newuser', '<EMAIL>', '<PASSWORD>5ca4', 0, '2021-06-01 16:46:08', 0, '2021-06-01 16:46:41'),
(10, 'tanya', '<EMAIL>', 'd37eaa547940fdd713097006308bf6c9', 0, '2021-06-01 17:49:36', 0, '0000-00-00 00:00:00');
--
-- Indexes for dumped tables
--
--
-- Indexes for table `directory_info`
--
ALTER TABLE `directory_info`
ADD PRIMARY KEY (`id`);
--
-- Indexes for table `users`
--
ALTER TABLE `users`
ADD PRIMARY KEY (`id`);
--
-- AUTO_INCREMENT for dumped tables
--
--
-- AUTO_INCREMENT for table `directory_info`
--
ALTER TABLE `directory_info`
MODIFY `id` int(11) NOT NULL AUTO_INCREMENT, AUTO_INCREMENT=27;
--
-- AUTO_INCREMENT for table `users`
--
ALTER TABLE `users`
MODIFY `id` int(11) NOT NULL AUTO_INCREMENT, AUTO_INCREMENT=11;
COMMIT;
/*!40101 SET CHARACTER_SET_CLIENT=@OLD_CHARACTER_SET_CLIENT */;
/*!40101 SET CHARACTER_SET_RESULTS=@OLD_CHARACTER_SET_RESULTS */;
/*!40101 SET COLLATION_CONNECTION=@OLD_COLLATION_CONNECTION */;
<file_sep>/readme.txt
1)The Entry POint of project is :-http://localhost/directory_user_interview/index.php/Welcome/
i)username:<EMAIL>
ii)password:<PASSWORD>
<file_sep>/application/views/login/login_password.php
<?php $this->load->view('login_header.php'); ?>
<div class="card">
<div class="card-body login-card-body">
<span>
<p style="text-align: right;"><a href="<?php echo base_url() ?>">Back</a></p>
</span>
<p class="login-box-msg">Sign in to start your session</p>
<!-- <form action="<?php echo base_url() ?>index.php/Welcome/checkEmailPassword" method="post"> -->
<p> Email: <?php echo $this->session->tempdata('user_email'); ?> </p>
<form action="" id="passwordCheck" method="POST">
<div style="text-align: left; color:red" class="main_notify"></div>
<label>Password</label>
<div class="input-group mb-3">
<input type="<PASSWORD>" name="password" class="form-control" placeholder="<PASSWORD>">
<div class="input-group-append">
<div class="input-group-text">
<span class="fas fa-envelope"></span>
</div>
</div>
</div>
<!-- /.col -->
<div class="col-4">
<input type="submit" class="btn btn-primary btn-block">
</div>
<!-- /.col -->
</div>
</form>
<!-- /.login-card-body -->
</div>
</div>
<?php $this->load->view('login_footer.php'); ?>
<script type="text/javascript">
$('body').on('submit', '#passwordCheck', function(e) {
e.preventDefault();
var formName = '#passwordCheck';
$(formName).find('.main_notify').html("");
var formdata = $(formName).serialize();
$.ajax({
type: 'POST',
url: '<?php echo base_url() ?>index.php/Welcome/checkEmailPassword',
data: formdata,
success: function(resp) {
var obj = JSON.parse(resp)
if (obj.status == 200) {
$('.main_notify').html(obj.msg);
setTimeout(function() {
window.location.href = '<?php echo base_url('index.php/Dashboard/index') ?>';
}, 1500);
} else if (obj.status == 201) {
$('.main_notify').html(obj.msg);
} else if (obj.status == 404) {
$('.main_notify').html(obj.msg);
setTimeout(function() {
window.location.href = '<?php echo base_url('index.php/') ?>';
}, 1500);
} else if (obj.status == 203) {
$('.main_notify').html(obj.msg);
} else {
alert(obj.msg);
}
}
})
})
</script><file_sep>/application/views/login/login.php
<?php $this->load->view('login/login_header.php'); ?>
<div class="card">
<div class="card-body login-card-body">
<p style="color: red"><?php echo $this->session->flashdata('email'); ?></p>
<p class="login-box-msg">Sign in to start your session</p>
<form action="" id="loginSubmit" method="post">
<div class="main_notify"></div>
<label>Email</label>
<div class="form-group ">
<input type="email" name="email" id="email" class="form-control" placeholder="Email" autocomplete="off">
</div>
<label>Password</label>
<div class="form-group ">
<input type="<PASSWORD>" name="password" id="password" class="form-control" placeholder="Email" autocomplete="off">
</div>
<!-- /.col -->
<div class="row">
<div class="col-md-4">
<input type="submit" class="btn btn-primary btn-block"></input>
</div>
<div class="col-md-4">
</div>
<div class="col-md-4">
<a href="<?php echo base_url() ?>/index.php/User">Register</a>
</div>
</div>
<!-- /.col -->
</form>
</div>
<!-- /.login-card-body -->
</div>
</div>
<?php $this->load->view('login/login_footer.php'); ?>
<script src="<?php echo base_url() ?>assets/js/custom.js"></script><file_sep>/application/models/DirectoryInfo_model.php
<?php
class DirectoryInfo_model extends CI_Model
{
public function getAllDirecetoryInfoData($user_id = '', $condition = array(), $limit = array(), $order = array())
{
$limit = (array_reverse($limit));
$this->db->select('di.*,DATE(created_at) AS c_day');
$this->db->from('directory_info di');
$this->db->where('di.is_deleted', 0);
if (!empty($condition['first_name']['value'])) {
$first_name = $condition['first_name']['value'];
$this->db->where('di.first_name', $first_name);
}
// print_r($condition);
if (!empty($condition['mobile_no']['value'])) {
$mobile_no = $condition['mobile_no']['value'];
$this->db->where('di.mobile_no', $mobile_no);
}
$this->db->where('di.user_id', $user_id);
if (!empty($limit)) {
$this->db->limit($limit[0], $limit[1]);
}
if (!empty($order) && isset($order['order_by']) && isset($order['order_type'])) {
$this->db->order_by($order['order_by'], $order['order_type']);
}
// print_r($this->db->last_query());
// exit;
$query = $this->db->get()->result_array();
return ($query);
}
public function getDirectoryDataById($id)
{
$sql = "SELECT di.* FROM directory_info di where di.id='" . $id . "' ";
$query = $this->db->query($sql);
return ($query->row_array());
}
public function increaseviewCount($id)
{
$date = date("Y-m-d", strtotime("now"));
$this->db->select('dcl.*');
$this->db->from('directory_count_logs dcl');
$this->db->where('dcl.directory_id', $id);
$this->db->where('dcl.date', $date);
$query = $this->db->get()->result_array();
if (empty($query)) {
$ans = $this->db->insert('directory_count_logs', array('directory_id' => $id, 'date' => $date, 'count' => 1));
} else {
$query = "UPDATE directory_count_logs SET count = count + 1 WHERE directory_id = '" . $id . "' AND date = '" . $date . "' ";
$ans = $this->db->query($query);
}
if ($this->db->affected_rows() > 0) {
return true;
}
}
public function getSearchFormData($id = '')
{
$this->db->select('di.*');
$this->db->from('directory_info di');
$this->db->where('di.is_deleted', 0);
$this->db->where('di.user_id', $id);
$this->db->order_by('di.id', 'DESC');
$ans = $this->db->get()->result_array();
return $ans;
}
}
| 7ed425562b19e65826bd2f853a9a7755382d5690 | [
"SQL",
"Text",
"PHP"
] | 12 | PHP | bharat123-anki/directory_user_interview | 1c6d78f88da5cc38ee489a0144d5b70d50d2da2d | 50601b5eaa1954054c90056fd45c2d9a56e8b4a9 |
refs/heads/master | <file_sep># Movie Data
## Updated Jan. 3, 2019 ##
### This has been archived because nothing will change except the data. If you would keep up with new stats, send me a message and I will send you the updated Excel. If you would like to make any changes to the code, also feel free to send a message, and I may unarchive it.
This is a collection of movies that I have watched over the course of my life. With over 300 movies analyzed, the following were collected:
- Directors
- Ratings
- Genre
- Years
- Domestic Gross
- Runtime
- Franchise
- Studio
The average runtime is about 119 minutes, with the shortest being 101 Dalmatians (79 min) and the longest being Gone with the Wind (238 min).
The average domestic gross is $151,485,429, with the lowest being Ip Man 2 ($205,675) and the highest being Star Wars the Force Awakens ($936,662,225).
The top five counts for Directors, Ratings, Genre, Years, Franchises, and Studios are listed below. To see the full list, take a look at the Excel file.
| Directors | Count |
| ------------- | ------------- |
| <NAME> | 12 |
| <NAME> | 7 |
| <NAME> | 7 |
| <NAME> | 6 |
| <NAME> | 6 |
| Rating | Count |
| ------------- | ------------- |
| R | 167 |
| PG-13 | 127 |
| PG | 74 |
| G | 19 |
| UR (Unrated) | 3 |
| Genre | Count |
| ------------- | ------------- |
| Action | 118 |
| Drama | 44 |
| Animation | 43 |
| Comedy | 36 |
| Crime | 30 |
| Years | Count |
| ------------- | ------------- |
| 2014 | 17 |
| 2016 | 16 |
| 2007 | 14 |
| 2006 | 14 |
| 2004 | 14 |
| Franchise | Count |
| ------------- | ------------- |
| Marvel Cinematic Universe | 18 |
| Harry Potter | 9 |
| Star Wars | 9 |
| X-Men | 7 |
| DC Extended Universe | 5 |
* Note: The actual top 3 for the franchise category are as follows:
| Franchise | Count |
| ------------- | ------------- |
| - (not a part of a franchise) | 98 |
| (adaptation) | 76 |
| (biography) | 19 |
They were not included as they are not a franchise
| Studio | Count |
| ------------- | ------------- |
| <NAME>. | 71 |
| Disney | 57 |
| Fox | 45 |
| Sony | 41 |
| Universal | 38 |<file_sep>using System;
using System.Collections.Generic;
using System.Linq;
using System.Runtime.InteropServices;
using Excel = Microsoft.Office.Interop.Excel;
namespace movieData
{
class Program
{
static void Main(string[] args)
{
Excel.Application xlApp = new Excel.Application();
Excel.Workbook movieWorkbook = xlApp.Workbooks.Open(@"C:\Users\Class2018\source\repos\movieData\movieData\Movie Data.xlsx");
Excel._Worksheet xlWorksheet = movieWorkbook.Sheets[1];
Excel.Range titleCol = xlWorksheet.Columns[1];
Excel.Range yearCol = xlWorksheet.Columns[2];
Excel.Range genreCol = xlWorksheet.Columns[3];
Excel.Range dirCol = xlWorksheet.Columns[4];
Excel.Range runCol = xlWorksheet.Columns[5];
Excel.Range grossCol = xlWorksheet.Columns[6];
Excel.Range ratingCol = xlWorksheet.Columns[7];
Excel.Range franchiseCol = xlWorksheet.Columns[8];
Excel.Range studioCol = xlWorksheet.Columns[9];
System.Array listVals = (System.Array)titleCol.Cells.Value;
string[] allTitles = listVals.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals1 = (System.Array)yearCol.Cells.Value;
string[] allYears = listVals1.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals2 = (System.Array)genreCol.Cells.Value;
string[] allGenre = listVals2.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals3 = (System.Array)dirCol.Cells.Value;
string[] allDir = listVals3.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals4 = (System.Array)runCol.Cells.Value;
string[] allRun = listVals4.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals5 = (System.Array)grossCol.Cells.Value;
string[] allGross = listVals5.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals6 = (System.Array)ratingCol.Cells.Value;
string[] allRating = listVals6.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals7 = (System.Array)franchiseCol.Cells.Value;
string[] allFran = listVals7.OfType<object>().Select(o => o.ToString()).ToArray();
System.Array listVals8 = (System.Array)studioCol.Cells.Value;
string[] allStudio = listVals8.OfType<object>().Select(o => o.ToString()).ToArray();
Stats s = new Stats();
Dictionary<string, int> DictYears = s.AddYears(allYears);
Dictionary<string, int> DictGenres = s.AddGenres(allGenre);
Dictionary<string, int> DictDirs = s.AddDirectors(allDir);
Dictionary<string, int> DictRating = s.AddGenres(allRating);
Dictionary<string, int> DictFran = s.AddGenres(allFran);
Dictionary<string, int> DictStudio = s.AddGenres(allStudio);
s.SetGrossStats(allGross, allTitles);
s.SetRunStats(allRun, allTitles);
Console.WriteLine("Time Average: {0} min", s.GetRunAvg());
Console.WriteLine("Shortest: {0} at {1} min", s.GetShortestFilm(), s.GetMinRun());
Console.WriteLine("Longest: {0} at {1} min", s.GetLongestFilm(), s.GetMaxRun());
Console.WriteLine("Gross Average: ${0}", s.GetGrossAvg());
Console.WriteLine("Lowest: {0} at ${1}", s.GetLowestFilm(), s.GetMinGross());
Console.WriteLine("Highest: {0} at ${1}", s.GetHighestFilm(), s.GetMaxGross());
//write to excel sheets
Excel._Worksheet xlWorksheet2 = movieWorkbook.Sheets[2];
xlWorksheet2.Cells[1, 1] = "Directors";
xlWorksheet2.Cells[1, 2] = "Count";
int a = 1;
foreach (var directors in DictDirs)
{
a++;
xlWorksheet2.Cells[a, 1] = directors.Key;
xlWorksheet2.Cells[a, 2] = directors.Value;
}
Excel._Worksheet xlWorksheet3 = movieWorkbook.Sheets[3];
xlWorksheet3.Cells[1, 1] = "Years";
xlWorksheet3.Cells[1, 2] = "Count";
int b = 1;
foreach (var years in DictYears)
{
b++;
xlWorksheet3.Cells[b, 1] = years.Key;
xlWorksheet3.Cells[b, 2] = years.Value;
}
Excel._Worksheet xlWorksheet4 = movieWorkbook.Sheets[4];
xlWorksheet4.Cells[1, 1] = "Genre";
xlWorksheet4.Cells[1, 2] = "Count";
int c = 1;
foreach (var genres in DictGenres)
{
c++;
xlWorksheet4.Cells[c, 1] = genres.Key;
xlWorksheet4.Cells[c, 2] = genres.Value;
}
Excel._Worksheet xlWorksheet5 = movieWorkbook.Sheets[5];
xlWorksheet5.Cells[1, 1] = "Rating";
xlWorksheet5.Cells[1, 2] = "Count";
int d = 1;
foreach (var ratings in DictRating)
{
d++;
xlWorksheet5.Cells[d, 1] = ratings.Key;
xlWorksheet5.Cells[d, 2] = ratings.Value;
}
Excel._Worksheet xlWorksheet6 = movieWorkbook.Sheets[6];
xlWorksheet6.Cells[1, 1] = "Franchise";
xlWorksheet6.Cells[1, 2] = "Count";
int e = 1;
foreach (var franchises in DictFran)
{
e++;
xlWorksheet6.Cells[e, 1] = franchises.Key;
xlWorksheet6.Cells[e, 2] = franchises.Value;
}
Excel._Worksheet xlWorksheet7 = movieWorkbook.Sheets[7];
xlWorksheet7.Cells[1, 1] = "Studio";
xlWorksheet7.Cells[1, 2] = "Count";
int x = 1;
foreach (var studio in DictStudio)
{
x++;
xlWorksheet7.Cells[x, 1] = studio.Key;
xlWorksheet7.Cells[x, 2] = studio.Value;
}
Console.WriteLine("Done!");
Marshal.ReleaseComObject(yearCol);
Marshal.ReleaseComObject(genreCol);
Marshal.ReleaseComObject(dirCol);
Marshal.ReleaseComObject(runCol);
Marshal.ReleaseComObject(grossCol);
Marshal.ReleaseComObject(ratingCol);
Marshal.ReleaseComObject(franchiseCol);
Marshal.ReleaseComObject(studioCol);
Marshal.ReleaseComObject(xlWorksheet);
Marshal.ReleaseComObject(xlWorksheet2);
Marshal.ReleaseComObject(xlWorksheet3);
Marshal.ReleaseComObject(xlWorksheet4);
Marshal.ReleaseComObject(xlWorksheet5);
Marshal.ReleaseComObject(xlWorksheet6);
Marshal.ReleaseComObject(xlWorksheet7);
movieWorkbook.Save();
movieWorkbook.Close(0);
Marshal.ReleaseComObject(movieWorkbook);
xlApp.Quit();
Marshal.ReleaseComObject(xlApp);
}
}
}<file_sep>using System;
using System.Collections.Generic;
using System.Linq;
using System.Text;
using System.Threading.Tasks;
using System.Numerics;
namespace movieData
{
class Stats
{
private BigInteger GrossAvg { get; set; }
private int MaxGross { get; set; }
private string HighestFilm { get; set; }
private string LowestFilm { get; set; }
private int MinGross { get; set; }
private BigInteger RunAvg { get; set; }
private string LongestFilm { get; set; }
private string ShortestFilm { get; set; }
private int MaxRun { get; set; }
private int MinRun { get; set; }
public Dictionary<string, int> AddDirectors(string[] arr)
{
Dictionary<string, int> directors = new Dictionary<string, int>();
for (int i = 1; i < arr.Length; i++)
{
if (directors.ContainsKey(arr[i]))
{
directors[arr[i]] = directors[arr[i]] += 1;
}
else
{
directors.Add(arr[i], 1);
}
}
return directors;
}
public Dictionary<string, int> AddGenres(string[] arr)
{
Dictionary<string, int> genre = new Dictionary<string, int>();
for (int i = 1; i < arr.Length; i++)
{
if (genre.ContainsKey(arr[i]))
{
genre[arr[i]] = genre[arr[i]] += 1;
}
else
{
genre.Add(arr[i], 1);
}
}
return genre;
}
public Dictionary<string, int> AddYears(string[] arr)
{
Dictionary<string, int> years = new Dictionary<string, int>();
for (int i = 1; i < arr.Length; i++)
{
if (years.ContainsKey(arr[i]))
{
years[arr[i]] = years[arr[i]] += 1;
}
else
{
years.Add(arr[i], 1);
}
}
return years;
}
public Dictionary<string, int> AddRatings(string[] arr)
{
Dictionary<string, int> ratings = new Dictionary<string, int>();
for (int i = 1; i < arr.Length; i++)
{
if (ratings.ContainsKey(arr[i]))
{
ratings[arr[i]] = ratings[arr[i]] += 1;
}
else
{
ratings.Add(arr[i], 1);
}
}
return ratings;
}
public void SetGrossStats(string[] arr, string[] arr1)
{
BigInteger sum = 0;
int NON_NA = 0;
MinGross = Int32.MaxValue;
for (int i = 1; i < arr.Length; i++)
{
if (arr[i] != "N/A")
{
int ValToString = System.Convert.ToInt32(arr[i]);
sum += ValToString;
NON_NA++;
if (ValToString > MaxGross)
{
MaxGross = ValToString;
HighestFilm = arr1[i];
}
else if (ValToString < MinGross)
{
MinGross = ValToString;
LowestFilm = arr1[i];
}
}
}
GrossAvg = sum / NON_NA;
}
public BigInteger GetGrossAvg()
{
return GrossAvg;
}
public int GetMaxGross()
{
return MaxGross;
}
public int GetMinGross()
{
return MinGross;
}
public string GetHighestFilm()
{
return HighestFilm;
}
public string GetLowestFilm()
{
return LowestFilm;
}
public void SetRunStats(string[] arr, string[] arr1)
{
int sum = 0;
MinRun = Int32.MaxValue;
for (int i = 1; i < arr.Length; i++)
{
int ValToString = System.Convert.ToInt32(arr[i]);
sum += ValToString;
if (ValToString > MaxRun)
{
MaxRun = ValToString;
LongestFilm = arr1[i];
}
else if (ValToString < MinRun)
{
MinRun = ValToString;
ShortestFilm = arr1[i];
}
}
RunAvg = sum / arr.Length;
}
public BigInteger GetRunAvg()
{
return RunAvg;
}
public int GetMaxRun()
{
return MaxRun;
}
public int GetMinRun()
{
return MinRun;
}
public string GetLongestFilm()
{
return LongestFilm;
}
public string GetShortestFilm()
{
return ShortestFilm;
}
}
} | a45673c93c66d9b23816841238541826bd104f60 | [
"Markdown",
"C#"
] | 3 | Markdown | rzhou10/Movie-Data | b4a9a4e1dad17e0176de8424b4d59d338a55157a | 43c990623e1310c1085cab127b22da30dfa49c87 |
refs/heads/master | <repo_name>aholston/user_dash<file_sep>/README.md
This is one of the first full stack assignments I did when I was learning Python. It needs to be debugged. Admin and User models being seperate are affecting the functionality. I am saving it as a side project to practice debugging when I have more free time.
<file_sep>/apps/userapp/views.py
from django.shortcuts import render, redirect, HttpResponse
from .models import *
from django.contrib import messages
import bcrypt
# Create your views here.
def index(request):
return render(request, 'userapp/index.html')
def signin(request):
return render(request, 'userapp/signin.html')
def register(request):
return render(request, 'userapp/register.html')
def reginfo(request):
errors = User.objects.validate(request.POST)
if len(errors):
for tag, error in errors.iteritems():
messages.error(request, error, extra_tags=tag)
return redirect('/register')
else:
pwHash = bcrypt.hashpw(request.POST['password'].encode(), bcrypt.gensalt())
if len(User.objects.all()) < 1 and len(Admin.objects.all()) < 1:
newUser = Admin.objects.create(first_name=request.POST['first_name'], last_name=request.POST['last_name'], email=request.POST['email'], password=<PASSWORD>)
return redirect('/')
else:
newUser = Admin.objects.create(first_name=request.POST['first_name'], last_name=request.POST['last_name'], email=request.POST['email'], password=<PASSWORD>)
return redirect('/')
| 13c57b5ac707fe210da9fa5f471eae0221c0fd31 | [
"Markdown",
"Python"
] | 2 | Markdown | aholston/user_dash | 7b6185bdd8829644802b5c28d99b157275c4a9b4 | 7c57179e67f2c97db5d4c678ba068cc1983721b3 |
refs/heads/master | <file_sep>require 'spec_helper'
describe "skills/show" do
before(:each) do
@skill = assign(:skill, stub_model(Skill,
:level => 1,
:one_click_freight => false,
:one_click_mail => false,
:one_click_passengers => false,
:passengers => 2,
:storage => 3,
:local_trains => 4,
:international_trains => 5,
:depot_trains => 6,
:route_planners => 7,
:maglev => 8,
:stations => 9
))
end
it "renders attributes in <p>" do
render
# Run the generator again with the --webrat flag if you want to use webrat matchers
rendered.should match(/1/)
rendered.should match(/false/)
rendered.should match(/false/)
rendered.should match(/false/)
rendered.should match(/2/)
rendered.should match(/3/)
rendered.should match(/4/)
rendered.should match(/5/)
rendered.should match(/6/)
rendered.should match(/7/)
rendered.should match(/8/)
rendered.should match(/9/)
end
end
<file_sep>describe ApplicationHelper do
let(:base_title) { "TrainStation Admin App" }
describe "base_title" do
it "should be" do
base_title().should == base_title
end
end
describe "full_title" do
it "should include the page name" do
full_title("foo").should =~ /foo/
end
it "should include the base name" do
full_title("foo").should =~ /^TrainStation Admin App/
end
it "should not include a bar for the home page" do
full_title("").should_not =~ /\|/
end
end
end<file_sep># == Schema Information
#
# Table name: materials
#
# id :integer not null, primary key
# wood :integer default(0), not null
# nails :integer default(0), not null
# bricks :integer default(0), not null
# glass :integer default(0), not null
# steel :integer default(0), not null
# gravel :integer default(0), not null
# fuel :integer default(0), not null
# uranium :integer default(0), not null
# cement :integer default(0), not null
# rubber :integer default(0), not null
# carbon :integer default(0), not null
# titanium :integer default(0), not null
# marble :integer default(0), not null
# wire :integer default(0), not null
# plastics :integer default(0), not null
# silicon :integer default(0), not null
# user_id :integer not null
# created_at :datetime not null
# updated_at :datetime not null
#
class Material < ActiveRecord::Base
attr_accessible :bricks, :carbon, :cement, :fuel, :glass, :gravel,
:marble, :nails, :plastics, :rubber, :silicon, :steel,
:titanium, :uranium, :wire, :wood
attr_protected :user_id
belongs_to :user, inverse_of: :material
validates :bricks, presence: true, numericality: { only_integer: true }
validates :carbon, presence: true, numericality: { only_integer: true }
validates :cement, presence: true, numericality: { only_integer: true }
validates :fuel, presence: true, numericality: { only_integer: true }
validates :glass, presence: true, numericality: { only_integer: true }
validates :gravel, presence: true, numericality: { only_integer: true }
validates :marble, presence: true, numericality: { only_integer: true }
validates :nails, presence: true, numericality: { only_integer: true }
validates :plastics, presence: true, numericality: { only_integer: true }
validates :rubber, presence: true, numericality: { only_integer: true }
validates :silicon, presence: true, numericality: { only_integer: true }
validates :steel, presence: true, numericality: { only_integer: true }
validates :titanium, presence: true, numericality: { only_integer: true }
validates :uranium, presence: true, numericality: { only_integer: true }
validates :wire, presence: true, numericality: { only_integer: true }
validates :wood, presence: true, numericality: { only_integer: true }
validates :user_id, presence: true, numericality: { only_integer: true }
end
<file_sep>require 'spec_helper'
describe "materials/show" do
before(:each) do
@material = assign(:material, stub_model(Material,
:wood => "",
:nails => "",
:bricks => "",
:glass => "",
:steel => "",
:gravel => "",
:fuel => "",
:uranium => "",
:cement => "",
:rubber => "",
:carbon => "",
:titanium => "",
:marble => "",
:wire => "",
:plastics => "",
:silicon => 1
))
end
it "renders attributes in <p>" do
render
# Run the generator again with the --webrat flag if you want to use webrat matchers
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(//)
rendered.should match(/1/)
end
end
<file_sep>require 'spec_helper'
describe "skills/new" do
before(:each) do
assign(:skill, stub_model(Skill,
:level => 1,
:one_click_freight => false,
:one_click_mail => false,
:one_click_passengers => false,
:passengers => 1,
:storage => 1,
:local_trains => 1,
:international_trains => 1,
:depot_trains => 1,
:route_planners => 1,
:maglev => 1,
:stations => 1
).as_new_record)
end
it "renders new skill form" do
render
# Run the generator again with the --webrat flag if you want to use webrat matchers
assert_select "form", :action => skills_path, :method => "post" do
assert_select "input#skill_level", :name => "skill[level]"
assert_select "input#skill_one_click_freight", :name => "skill[one_click_freight]"
assert_select "input#skill_one_click_mail", :name => "skill[one_click_mail]"
assert_select "input#skill_one_click_passengers", :name => "skill[one_click_passengers]"
assert_select "input#skill_passengers", :name => "skill[passengers]"
assert_select "input#skill_storage", :name => "skill[storage]"
assert_select "input#skill_local_trains", :name => "skill[local_trains]"
assert_select "input#skill_international_trains", :name => "skill[international_trains]"
assert_select "input#skill_depot_trains", :name => "skill[depot_trains]"
assert_select "input#skill_route_planners", :name => "skill[route_planners]"
assert_select "input#skill_maglev", :name => "skill[maglev]"
assert_select "input#skill_stations", :name => "skill[stations]"
end
end
end
<file_sep>module Requests
class User_pages
end
end
<file_sep>class CreateMaterials < ActiveRecord::Migration
def change
default_settings = {
default: 0,
null: false,
}
create_table :materials do |t|
t.integer :wood, default_settings
t.integer :nails, default_settings
t.integer :bricks, default_settings
t.integer :glass, default_settings
t.integer :steel, default_settings
t.integer :gravel, default_settings
t.integer :fuel, default_settings
t.integer :uranium, default_settings
t.integer :cement, default_settings
t.integer :rubber, default_settings
t.integer :carbon, default_settings
t.integer :titanium, default_settings
t.integer :marble, default_settings
t.integer :wire, default_settings
t.integer :plastics, default_settings
t.integer :silicon, default_settings
t.integer :user_id, null: false
t.timestamps
end
add_index :materials, :user_id
end
end
<file_sep>class CreateSkills < ActiveRecord::Migration
def change
create_table :skills do |t|
t.integer :level, default: 0, null: false
t.integer :passengers, default: 0, null: false
t.integer :storage, default: 0, null: false
t.integer :local_trains, default: 0, null: false
t.integer :international_trains, default: 0, null: false
t.integer :depot_trains, default: 0, null: false
t.integer :route_planners, default: 0, null: false
t.integer :maglev, default: 0, null: false
t.integer :stations, default: 0, null: false
t.boolean :one_click_freight, default: false, null: false
t.boolean :one_click_mail, default: false, null: false
t.boolean :one_click_passengers, default: false, null: false
t.integer :user_id, null: false
t.timestamps
end
add_index :skills, [:level, :user_id]
end
end
<file_sep># == Schema Information
#
# Table name: materials
#
# id :integer not null, primary key
# wood :integer default(0), not null
# nails :integer default(0), not null
# bricks :integer default(0), not null
# glass :integer default(0), not null
# steel :integer default(0), not null
# gravel :integer default(0), not null
# fuel :integer default(0), not null
# uranium :integer default(0), not null
# cement :integer default(0), not null
# rubber :integer default(0), not null
# carbon :integer default(0), not null
# titanium :integer default(0), not null
# marble :integer default(0), not null
# wire :integer default(0), not null
# plastics :integer default(0), not null
# silicon :integer default(0), not null
# user_id :integer not null
# created_at :datetime not null
# updated_at :datetime not null
#
require 'spec_helper'
describe Material do
let(:user) { FactoryGirl.create(:user) }
# Note to myself:
# has_one uses <class>.build_<second_class> to reate relationship
# has_many uses <class>.build.<second_class> to create relationship
before { @material = user.build_material(
wood: 1,
nails: 1,
bricks: 1,
glass: 1,
steel: 1,
gravel: 1,
fuel: 1,
uranium: 1,
cement: 1,
rubber: 1,
carbon: 1,
titanium: 1,
marble: 1,
wire: 1,
plastics: 1,
silicon: 1,
) }
subject { @material }
it { should respond_to(:wood) }
it { should respond_to(:user) }
pending "add some more examples to (or delete) #{__FILE__}"
end
<file_sep># Facebook TrainStation Administration (ts_admin)
[](https://gemnasium.com/ivdmeer/ts_admin)
This is an application for the Facebook TrainStation game.
With this application you can share your game progress with a group of friends.
Features:
* Share your skill level
* Share your wishlist
* Share your stock of materials
* Share your required materias
This appliction is a proof-of-concept and based on:
[*Ruby on Rails Tutorial - Learn Rails by Example*](http://railstutorial.org/)
by [<NAME>](http://michaelhartl.com/).
<file_sep>require 'spec_helper'
describe "materials/index" do
before(:each) do
assign(:materials, [
stub_model(Material,
:wood => "",
:nails => "",
:bricks => "",
:glass => "",
:steel => "",
:gravel => "",
:fuel => "",
:uranium => "",
:cement => "",
:rubber => "",
:carbon => "",
:titanium => "",
:marble => "",
:wire => "",
:plastics => "",
:silicon => 1
),
stub_model(Material,
:wood => "",
:nails => "",
:bricks => "",
:glass => "",
:steel => "",
:gravel => "",
:fuel => "",
:uranium => "",
:cement => "",
:rubber => "",
:carbon => "",
:titanium => "",
:marble => "",
:wire => "",
:plastics => "",
:silicon => 1
)
])
end
it "renders a list of materials" do
render
# Run the generator again with the --webrat flag if you want to use webrat matchers
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => "".to_s, :count => 2
assert_select "tr>td", :text => 1.to_s, :count => 2
end
end
<file_sep>module ApplicationHelper
# Check application_controler for helper methods that do not
# include html methods, this way the view and the controllers
# benefit from this helper.
def display_boolean_icon(value, options = { })
tag('i', class: ((value) ? 'icon-ok' : 'icon-remove'), title: value)
end
end<file_sep># Read about factories at https://github.com/thoughtbot/factory_girl
FactoryGirl.define do
factory :material do
wood 1
nails 1
bricks 1
glass 1
steel 1
gravel 1
fuel 1
uranium 1
cement 1
rubber 1
carbon 1
titanium 1
marble 1
wire 1
plastics 1
silicon 1
end
end<file_sep>require 'spec_helper'
describe "materials/edit" do
before(:each) do
@material = assign(:material, stub_model(Material,
:wood => "",
:nails => "",
:bricks => "",
:glass => "",
:steel => "",
:gravel => "",
:fuel => "",
:uranium => "",
:cement => "",
:rubber => "",
:carbon => "",
:titanium => "",
:marble => "",
:wire => "",
:plastics => "",
:silicon => 1
))
end
it "renders the edit material form" do
render
# Run the generator again with the --webrat flag if you want to use webrat matchers
assert_select "form", :action => materials_path(@material), :method => "post" do
assert_select "input#material_wood", :name => "material[wood]"
assert_select "input#material_nails", :name => "material[nails]"
assert_select "input#material_bricks", :name => "material[bricks]"
assert_select "input#material_glass", :name => "material[glass]"
assert_select "input#material_steel", :name => "material[steel]"
assert_select "input#material_gravel", :name => "material[gravel]"
assert_select "input#material_fuel", :name => "material[fuel]"
assert_select "input#material_uranium", :name => "material[uranium]"
assert_select "input#material_cement", :name => "material[cement]"
assert_select "input#material_rubber", :name => "material[rubber]"
assert_select "input#material_carbon", :name => "material[carbon]"
assert_select "input#material_titanium", :name => "material[titanium]"
assert_select "input#material_marble", :name => "material[marble]"
assert_select "input#material_wire", :name => "material[wire]"
assert_select "input#material_plastics", :name => "material[plastics]"
assert_select "input#material_silicon", :name => "material[silicon]"
end
end
end
<file_sep># Read about factories at https://github.com/thoughtbot/factory_girl
FactoryGirl.define do
factory :skill do
level 1
one_click_freight false
one_click_mail false
one_click_passengers false
passengers 1
storage 1
local_trains 1
international_trains 1
depot_trains 1
route_planners 1
maglev 1
stations 1
end
end
<file_sep># == Schema Information
#
# Table name: skills
#
# id :integer not null, primary key
# level :integer default(0), not null
# passengers :integer default(0), not null
# storage :integer default(0), not null
# local_trains :integer default(0), not null
# international_trains :integer default(0), not null
# depot_trains :integer default(0), not null
# route_planners :integer default(0), not null
# maglev :integer default(0), not null
# stations :integer default(0), not null
# one_click_freight :boolean default(FALSE), not null
# one_click_mail :boolean default(FALSE), not null
# one_click_passengers :boolean default(FALSE), not null
# user_id :integer not null
# created_at :datetime not null
# updated_at :datetime not null
#
require 'spec_helper'
describe Skill do
let(:user) { FactoryGirl.create(:user) }
# Note to myself:
# has_one uses <class>.build_<second_class> to create relationship
# has_many uses <class>.build.<second_class> to create relationship
before { @skill = user.build_skill(
depot_trains: 0,
international_trains: 0,
level: 0,
local_trains: 0,
maglev: 0,
one_click_freight: :false,
one_click_mail: :false,
one_click_passengers: :false,
passengers: 0,
route_planners: 0,
stations: 0,
storage: 0,
) }
subject { @skill }
it { should respond_to(:depot_trains) }
it { should respond_to(:international_trains) }
it { should respond_to(:level) }
it { should respond_to(:local_trains) }
it { should respond_to(:maglev) }
it { should respond_to(:one_click_freight) }
it { should respond_to(:one_click_mail) }
it { should respond_to(:one_click_passengers) }
it { should respond_to(:passengers) }
it { should respond_to(:route_planners) }
it { should respond_to(:stations) }
it { should respond_to(:storage) }
it { should respond_to(:user_id) }
it { should respond_to(:user) }
its(:user) { should == user }
it { should be_valid }
describe "accessible attributes" do
it "should not allow access to user_id" do
expect do
Skill.new(user_id: user.id)
end.should raise_error(ActiveModel::MassAssignmentSecurity::Error)
end
end
describe "when depot_trains is not present" do
before { @skill.depot_trains = nil }
it { should_not be_valid }
end
describe "when international_trains is not present" do
before { @skill.international_trains = nil }
it { should_not be_valid }
end
describe "when level is not present" do
before { @skill.level = nil }
it { should_not be_valid }
end
describe "when local_trains is not present" do
before { @skill.local_trains = nil }
it { should_not be_valid }
end
describe "when maglev is not present" do
before { @skill.maglev = nil }
it { should_not be_valid }
end
describe "when one_click_freight is not present" do
before { @skill.one_click_freight = nil }
it { should_not be_valid }
end
describe "when one_click_freight is not present" do
before { @skill.one_click_freight = nil }
it { should_not be_valid }
end
describe "when one_click_mail is not present" do
before { @skill.one_click_mail = nil }
it { should_not be_valid }
end
describe "when one_click_passengers is not present" do
before { @skill.one_click_passengers = nil }
it { should_not be_valid }
end
describe "when passengers is not present" do
before { @skill.passengers = nil }
it { should_not be_valid }
end
describe "when route_planners is not present" do
before { @skill.route_planners = nil }
it { should_not be_valid }
end
describe "when stations is not present" do
before { @skill.stations = nil }
it { should_not be_valid }
end
describe "when storage is not present" do
before { @skill.storage = nil }
it { should_not be_valid }
end
describe "when user_id is not present" do
before { @skill.user_id = nil }
it { should_not be_valid }
end
describe "with one_click_freight attribute set to 'true'" do
before { @skill.toggle!(:one_click_freight) }
it { should be_one_click_freight }
end
describe "with one_click_mail attribute set to 'true'" do
before { @skill.toggle!(:one_click_mail) }
it { should be_one_click_mail }
end
describe "with one_click_passengers attribute set to 'true'" do
before { @skill.toggle!(:one_click_passengers) }
it { should be_one_click_passengers }
end
end
<file_sep># == Schema Information
#
# Table name: skills
#
# id :integer not null, primary key
# level :integer default(0), not null
# passengers :integer default(0), not null
# storage :integer default(0), not null
# local_trains :integer default(0), not null
# international_trains :integer default(0), not null
# depot_trains :integer default(0), not null
# route_planners :integer default(0), not null
# maglev :integer default(0), not null
# stations :integer default(0), not null
# one_click_freight :boolean default(FALSE), not null
# one_click_mail :boolean default(FALSE), not null
# one_click_passengers :boolean default(FALSE), not null
# user_id :integer not null
# created_at :datetime not null
# updated_at :datetime not null
#
class Skill < ActiveRecord::Base
attr_accessible :depot_trains, :international_trains, :level,
:local_trains, :maglev, :one_click_freight,
:one_click_mail, :one_click_passengers, :passengers,
:route_planners, :stations, :storage
attr_protected :user_id
belongs_to :user, inverse_of: :skill
validates :level, presence: true
validates :passengers, presence: true
validates :storage, presence: true
validates :local_trains, presence: true
validates :international_trains, presence: true
validates :depot_trains, presence: true
validates :route_planners, presence: true
validates :maglev, presence: true
validates :stations, presence: true
# TODO investigate more on validating boolean values. It doesn't work as expected.
# see http://ar.rubyonrails.org/classes/ActiveRecord/Validations/ClassMethods.html
# If you want to validate the presence of a boolean field
# (where the real values are true and false), you will want to use
# validates_inclusion_of :field_name, :in => [true, false]
# This is due to the way Object#blank? handles boolean values.
# false.blank? # => true
#
validates :one_click_freight, inclusion: { in: [true, false] }
validates :one_click_mail, inclusion: { in: [true, false] }
validates :one_click_passengers, inclusion: { in: [true, false] }
validates :user_id, presence: true
default_scope order: 'skills.level DESC'
end
| 617a3e5f46206b664ce9f0dfeb3638448369258e | [
"Markdown",
"Ruby"
] | 17 | Ruby | ivdmeer/ts_admin | 2d7c2934e636e7790ee9f7819ca09ee36ebe68e8 | 6dbdabd48d671a3cda01d4abbbd4b002b124cbfc |
refs/heads/master | <repo_name>huhai0403/react-TodoList<file_sep>/README.md
# react-TodoList
************************************
#####began in 2015.11.1
#####upload in 2016.3.16
*************************************
`React v0.14.3` `Sortable 1.4.2` `open by firefox` `code by webstorm` `not used webpack or gulp ...`
**********************************
It is very original and fall behind because I contact JavaScript first time. Last year,I could only code html and css, was a complete beginner(of cours,as it is also!QAQ)
*************************************
<file_sep>/todopage/main.js
/*** Created by Administrator on 2015/11/10 0010.*/
/*nav*/
/*HeadPic*/
var HeadPic = React.createClass({
render:function(){
return (<div id="headpic">
<a href="#"><img src="img/1.gif" /></a>
</div>)
}
});
var items = ['任务', '便签', '项目', '地点', '标签'];
var active = { color:"#fff", background:"#000"};
var i = 0 ;
var n ;
var views ;
var NavList = React.createClass({
getInitialState:function(){
return {data: [],
NumLink: 0,
NumTop: 0
}
},
navLinkClick:function(index){
/*alert(i);*/
i = index;
this.setState({
NumTop : index,
NumLink : index
});
},
render:function(){
var that = this;
return(<div className="nav">
<ul >
{items.map(function(item,index){
var className ;
that.state.NumLink == index ? className = 'active' : className = '';
return (<li key={index} className = {className} onClick={that.navLinkClick.bind(that,index)} ><a href="#">{item}</a></li>)
})}
</ul>
</div>
)
}
});
/*top*/
var myDate = new Date();
/*time*/
var TopBox = React.createClass({
getInitialState:function(){
return{ ThisYears : myDate.getFullYear(),
ThisMouth : myDate.getMonth() + 1,
ThisDays : myDate.getDate(),
ThisHours : myDate.getHours(),
ThisMinutes :myDate.getMinutes()
}
},
addDayClick:function(){
this.setState({ThisDays:this.state.ThisDays + 1})
},
minDayClick:function(){
this.setState({ThisDays:this.state.ThisDays - 1})
},
render:function(){
return(
<div className="top clear">
<span className="time">{this.state.ThisYears+'年'+this.state.ThisMouth+'月'+this.state.ThisDays+"日 "+this.state.ThisHours+':'+this.state.ThisMinutes}</span>
<div className="checktime">
<a href="#" onClick={this.minDayClick}>←</a>
<a href="#" onClick={this.addDayClick}>→</a>
</div>
</div>
)
}
});
/*main*/
/*任务*/
var MainBoxTask = React.createClass({
render:function(){
return(
<div className="box clear">
<FirstBlock />
<SecondBlock />
<ThirdBlock />
<FourthBlock />
</div>
)
}
});
/*便签*/
var MainBoxNote = React.createClass({
render:function(){
return(
<div className="box clear">
<h1>便签</h1>
</div>
)
}
});
/*项目*/
var MainBoxPro = React.createClass({
render:function(){
return(
<div className="box clear">
<h1>项目</h1>
</div>
)
}
});
/*地点*/
var MainBoxPlace = React.createClass({
render:function(){
return(
<div className="box clear">
<h1>地点</h1>
</div>
)
}
});
/*标签*/
var MainBoxTag = React.createClass({
render:function(){
return(
<div className="box clear">
<p>标签</p>
</div>
)
}
});
/*block*/
var EventEach = React.createClass({
getInitialState: function () {
return {eventDisplayed: true,
eventShowed:true,
eventChange:true
}
},
eventDisClick:function(){
this.setState({
eventDisplayed: false
});
},
eventChangeClick: function() {
this.setState({eventChange: !this.state.eventChange});
},
render:function(){
var styleObj={
display:this.state.eventDisplayed ? "block": "none"
};
return(
<li style={styleObj}>
<p className="caption clear"><input type="text" defaultvalue="" disabled={this.state.eventChange} /><em>{document.lastModified.substring(0,10)}</em><button title="修改" onClick={this.eventChangeClick} className="change"><img src="img/2.gif" /></button><button title="删除" className="change" onClick={this.eventDisClick} ><img src="img/3.gif" /></button></p>
<textarea className="describe" disabled={this.state.eventChange} ></textarea>
</li>
)
}
});
var eventFirstList = [<EventEach />], eventSecondList = [] , eventThirdList = [] , eventFourthList = [];
var BlockTitleTip = ['很重要-很紧急','重要-不紧急','不重要-紧急','不重要-不紧急']
var FirstBlock = React.createClass({
BuildEvent:function(){
n = i;
/*alert(n);*/
this.setState({eventFirstList:eventFirstList.push(<EventEach />)});
},
render:function(){
return(
<div className="fir_block br mr_1">
<div className="block">
<div className="title first clear">
<span>{BlockTitleTip[0]}</span>
<a href="#" onClick={this.BuildEvent}>+</a>
</div>
<div className="oh">
<ul className="list">
{eventFirstList.map(function(listeach,Maxnum){
return(<div key={Maxnum}>{listeach}</div>)
})}
</ul>
</div>
</div>
</div>
)
}
});
var SecondBlock = React.createClass({
BuildEvent:function(){
this.setState({eventSecondList:eventSecondList.push(<EventEach />)});
},
render:function(){
return(
<div className="sec_block">
<div className="block">
<div className="title second clear">
<span>{BlockTitleTip[1]}</span>
<a href="#" onClick={this.BuildEvent}>+</a>
</div>
<div className="oh">
<ul className="list">
{eventSecondList.map(function(listeach,Maxnum){
return(<div key={Maxnum}>{listeach}</div>)
})}
</ul>
</div>
</div>
</div>
)
}
});
var ThirdBlock = React.createClass({
BuildEvent:function(){
this.setState({eventThirdList:eventThirdList.push(<EventEach />)});
},
render:function(){
return(
<div className="thi_block br bt mt_1 mr_1">
<div className="block">
<div className="title third clear">
<span>{BlockTitleTip[2]}</span>
<a href="#" onClick={this.BuildEvent}>+</a>
</div>
<div className="oh">
<ul className="list">
{eventThirdList.map(function(listeach,Maxnum){
return(<div key={Maxnum}>{listeach}</div>)
})}
</ul>
</div>
</div>
</div>
)
}
});
var FourthBlock = React.createClass({
BuildEvent:function(){
this.setState({eventFourthList:eventFourthList.push(<EventEach />)});
},
render:function(){
return(
<div className="fou_block bt mt_1">
<div className="block">
<div className="title fourth clear">
<span>{BlockTitleTip[3]}</span>
<a href="#" onClick={this.BuildEvent}>+</a>
</div>
<div className="oh">
<ul className="list">
{eventFourthList.map(function(listeach,Maxnum){
return(<div key={Maxnum}>{listeach}</div>)
})}
</ul>
</div>
</div>
</div>
)
}
});
/*side*/
/*var SideBox = React.createClass({
render:function(){
return(
<div className="side">
<HeadPic />
<NavList />
</div>
)
}
});*/
var MainBoxViews = [<MainBoxTask />,<MainBoxNote/>,<MainBoxPro/>,<MainBoxPlace/>,<MainBoxTag/>];
var ContentBox = React.createClass({
render:function(){
/*{alert(i)}*/
return(
<div className="content">
<div className="wrap">
<TopBox />
{MainBoxViews[i]}
</div>
</div>
)
}
});
var MainPage = React.createClass({
render:function(){
return (<div>
<div className="side">
<HeadPic />
<NavList />
</div>
<ContentBox />
</div>)
}
});
ReactDOM.render(
<MainPage />,
document.getElementById('main')
);
| 4e8be8eac72857888e0eea7e64e9c770fcec2219 | [
"Markdown",
"JavaScript"
] | 2 | Markdown | huhai0403/react-TodoList | c715cd3ffb2b6e1528d4a7188fa04367d4e8ada0 | 25542a23d51872ae5cdefedb1688275e673ed3db |
refs/heads/master | <repo_name>remipichon/jmpresseditor2.js<file_sep>/js/kernel_composant.js
/*
* Classes :
* Slide
* Element
* Text
* Image
* Méthode :
* Object.size
* selectSlide(callback, param1, composant) return slide's matricule
* getSlideMother(matricule) return slide's matricule
* Global:
* container
*
*/
//pour connaitre la taille d'un objet
Object.size = function(obj) {
var size = 0, key;
for (key in obj) {
if (obj.hasOwnProperty(key))
size++;
}
return size;
};
/******************************************************/
/******* definition des clases ********/
globalCpt = 0;
function initContainer() {
container = {metadata: {
type: 'free', //free, tree
name: 'Unnamed'
}, slide: [],
getSlide: function(matricule) {
for (var i in container.slide) {
if (container.slide[i].matricule === matricule)
return container.slide[i];
}
//console.log('error : getSlide : matricule \'' + matricule + '\' doesn\'t exist as a slide');
return;
}
};
watch(container.metadata, 'name', function(attr, action, newVal, oldVal) {
//mise à jour du DOM
$('#slideshowNameFree').html(newVal);
});
}
function initJmpress() {
$('#slideArea').children().remove();
//il semblerait que Jmpress ait besoin d'au moins une slide dans slideArea pour pouvoir looper
$('#slideArea').append('<div id="home" class="hidden step slide overview " data-scale ="5" data-x="1000" data-z ="1000" style="display:block"></div>');
$('#slideArea').removeClass();
$('#slideArea').jmpress({
//mouse: {clickSelects: false},
// keyboard: {use: false},
// keyboard: {
// 112: '' //doesn't work although doc shows me this way
// },
viewPort: {
height: 400,
width: 3200,
maxScale: 1
}
});
// $('#slideArea').jmpress({
// viewPort: {
// height: 400,
// width: 3000,
// maxScale: 1
// }
// });
globalConfig = {
heightSlide: parseInt($('#home').css('height')),
widthSlide: parseInt($('#home').css('width'))
};
$('#slideArea').jmpress('deinit', $('#home'));
$('#home').removeClass('slide'); //sinon la slide prend de la place dans le DOM alors que Jmpress ne la connait pas
transform3D = new Transform3D();
}
/* classe slide
* matricule
* type
* pos
* rotate
* properties
* scale
* hierarchy
* watches
* show
* affiche coord et nb element
*show('element')
* affiche la liste des composants avec leur détals (héritage du swow pour ajouter des infos propres à la fille
*
* destroy
*/
Slide = Class.extend({
init: function(params) {
if (typeof params !== 'undefined') {
//if matricule is set, check if unique
if (typeof params.matricule !== 'undefined') {
if (typeof findObjectOfComposant(params.matricule) !== 'undefined') { //le matricule existe déjà !
console.log('Error : construct Slide : matricule ' + params.matricule + ' already set in container');
delete this;
return;
}
}
}
//default value
var matricule = 'slide' + globalCpt++;
this.matricule = matricule;
//check if unique matrcule already exists
while (typeof findObjectOfComposant(this.matricule) !== 'undefined') { //le matricule existe déjà !
var matricule = 'slide' + globalCpt++;
this.matricule = matricule;
}
this.type = 'slide';
this.pos = {
x: 0,
y: 0,
z: 0
};
this.rotate = {
x: 0,
y: 0,
z: 0
};
//je ne peux mettre des wacth que sur des objets, ce que sont les slides sur le papier.
//en pratique, simpleJsInheritance transforme l'objet en une fonction dotée de prototype.
//seul les objet au sein de la slide peuvent avoir des watch.
this.properties = {
scale: 1,
hierarchy: '0'
};
this.element = {};
//prise en compte des parametres utilisteurs
if (typeof params !== 'undefined') {
for (var param in params) {
if (typeof params[param] === 'object') {
for (var paramNested in params[param]) {
this[param][paramNested] = params[param][paramNested];
}
} else {
this[param] = params[param];
}
}
}
//definition des watch qui permettent d'agir sur le DOM lorsqu'on agit sur les objets des slides
watch(this.pos, function(attr, action, newVal, oldVal) {
//mise à jour du DOM
var $slide = $('#' + matricule);
$('#slideArea').jmpress('deinit', $slide);
var attribut = 'data-' + attr;
$slide.attr(attribut, newVal);
$('#slideArea').jmpress('init', $slide);
});
watch(this.rotate, function(attr, action, newVal, oldVal) {
//mise à jour du DOM
var $slide = $('#' + matricule);
$('#slideArea').jmpress('deinit', $slide);
var attribut = 'data-rotate-' + attr;
$slide.attr(attribut, newVal);
$('#slideArea').jmpress('init', $slide);
});
watch(this.properties, 'scale', function(attr, action, newVal, oldVal) {
//mise à jour du DOM
var $slide = $('#' + matricule);
$('#slideArea').jmpress('deinit', $slide);
var attribut = 'data-scale';
$slide.attr(attribut, newVal);
$('#slideArea').jmpress('init', $slide);
});
watch(this.properties, 'hierarchy', function(attr, action, newVal, oldVal) {
//mise à jour du DOM
var $slide = $('#' + matricule);
// $('#slideArea').jmpress('deinit', $slide);
var attribut = 'hierarchy';
$slide.attr(attribut, newVal);
// $('#slideArea').jmpress('init', $slide);
});
//ajout de la slide à l'espace de stockage
container.slide.push(this);
//create node via mustach
var template = $('#templateSlide').html();
var html = Mustache.to_html(template, this);
$('#slideArea >').append(html);
var $newSlide = $('#slideArea >').children().last();
$('#slideArea').jmpress('init', $newSlide);
handlerComposant($newSlide);
//ajout à la timeline
var idSlide = this.matricule;
var $slideButton = $('<li matricule=' + idSlide + '><span>' + ((this.type === 'overview') ? 'Overview' : idSlide) + '</span> <a class="cross" href="#">x</a></li>');
$('#sortable').append($slideButton);
$('#sortable').sortable({
start: function(event, ui) {
ui.item.startPos = ui.item.index();
},
stop: function(event, ui) {
var newIndex = ui.item.index(); //nouvelle place
var matriculeSorted = ui.item.attr('matricule'); //pour la slide qui vient prendre la place de
var slide = container.getSlide(matriculeSorted); //+ 2 car il y a pour le moment deux slides dans le DOM dès le départ
var slideAfter = $($('#slideArea>').children()[newIndex + 0 + 1]).attr('matricule'); //cette slide
if (newIndex > ui.item.startPos) { //maintenant
slide.reOrder(slideAfter, false);
}
else {
slide.reOrder(slideAfter, true);
}
// if (ui.item.startPos > newIndex)
// {
// $('.slide').eq(newIndex).before($('#' + $idSlideSorted + ''));
// }
// else
// {
// $('.slide').eq(newIndex).after($('#' + $idSlideSorted + ''));
// }
// $(".slide").each(function(index) {
// var idSlide = $(this).attr('id');
// //pressjson.slide[idSlide].index = index; // MaJ de l'index des slide
// });
////console.log(newIndex, slide.matricule, slideAfter);
},
axis: "y"
})
.disableSelection();
$("#sortable").on("sortupdate", function(event, ui) {
//////console.log(event, ui);
});
},
reOrder: function(slideAfter, isBefore) {
//deplace la slide avant slideAfter
if (isBefore == false) {
$('#' + this.matricule).insertAfter($('#' + slideAfter));
}
else {
$('#' + this.matricule).insertBefore($('#' + slideAfter));
}
//deplace le slide dans le container en s'appuyant sur le DOM
var newContainer = {
metadata: container.metadata,
slide: {}
};
$('#slideArea>').children().each(function() {
if ($(this).attr('id') !== 'profondeur') {
newContainer.slide[$(this).attr('matricule')] = container.slide[$(this).attr('matricule')];
}
});
container = newContainer;
},
show: function(i) {
if (typeof i === 'undefined') {
console.log('{ matricule:', this.matricule, ', pos:{x:', this.pos.x, ', y:', this.pos.y, 'z:', this.pos.z, '}, rotate:{x:', this.rotate.x, ',y:', this.rotate.y, 'z:', this.rotate.z, '}, scale:{scale:', this.properties.scale, '}, nb elements :', Object.size(this.element), '}');
}
else if (i === 'element') {
console.log('liste des elements');
for (var el in this.element) {
console.log(this.element[el].show());
}
} else {
return '{matricule: ' + this.matricule + ', pos:{x: ' + this.pos.x + ', y: ' + this.pos.y + ', z:' + this.pos.z + '}, rotate:{x: ' + this.rotate.x + ', y: ' + this.rotate.y + ', z: ' + this.rotate.z + '}, properties:{scale: ' + this.properties.scale + '}, nb elements :' + Object.size(this.element) + '}';
}
},
destroy: function() {
//Jmpress l'oublie
$('#slideArea').jmpress('deinit', $('#' + this.matricule));
//disparition du dom
$('#' + this.matricule).remove();
//nettoyage du container
delete container.getSlide(this.matricule);
}
});
/* Ebauche de tentative de gestion par la Class Element d'un mauvais matricule
* entrainant le demande de selection d'une slide de la part d'user.
*
* @param {type} slide (matricule)
* @param {type} matricule (du composant)
* @param {type} composant (instance of Element)
* @returns {undefined}
*/ /*
function addComposantToSlide(slide, matricule, composant) {
container.slide[slide].element[matricule] = composant;
composant.DOM(slide);
}*/
/* Interface Element : doit être instancié
* matricule
* pos
* rotate
* watches
* show
* destroy
*
* minimum pour fonctionner :
* slide : matricule d'une slide de destination (erreur gérée)
*
*/
Element = Class.extend({
init: function(slide, params, matricule) {
if (typeof params !== 'undefined') {
//if matricule is set, check if unique
if (typeof params.matricule !== 'undefined') {
if (typeof findObjectOfComposant(params.matricule) !== 'undefined') { //le matricule existe déjà !
console.log('Error : construct Element : matricule ' + params.matricule + ' already set in container');
delete this;
return;
}
}
}
//ce sont les filles qui se chargent de params !
//default values
this.pos = {
x: 300, //connecter à la moitié de la largeur de la slide type
y: 300, //idem pour la hauteur
z: 0
};
this.rotate = {
x: 0,
y: 0,
z: 0
};
this.properties = {
};
//definition des watch qui permettent d'agir sur le DOM lorsqu'on agit sur les objets des slides
watch(this.pos, function(attr, action, newVal, oldVal) {
//mise à jour du DOM
var $element = $('#' + matricule);
var attribut;
switch (attr) {
case 'x':
attribut = 'left';
break;
case 'y' :
attribut = 'top';
break;
default :
return;
}
$element.css(attribut, newVal);
});
watch(this.rotate, function(attr, action, newVal, oldVal) {
return;
});
//gestion de l'erreur de matricule
if (container.getSlide(slide) === undefined) {
//console.log('Error : Le matricule de la slide cible n\'existe pas ', slide);
return 0;
} else if (container.getSlide(slide).type !== 'slide') {
//console.log('Error : element constructor : insert elementin slide : Le composant cible doit être une slide ', slide);
return 0;
} else {
container.getSlide(slide).element[matricule] = this;
return 1;
}
},
show: function(i) {
if (typeof i === 'undefined') {
console.log('{ matricule:', this.matricule, ', pos:{x:', this.pos.x, ', y:', this.pos.y, 'z:', this.pos.z, '}, rotate:{x:', this.rotate.x, ',y:', this.rotate.y, 'z:', this.rotate.z, '} }');
}
else {
return '{ matricule: ' + this.matricule + ', pos:{x: ' + this.pos.x + ', y: ' + this.pos.y + ' z: ' + this.pos.z + '}, rotate:{x:' + this.rotate.x + ',y:' + this.rotate.y + 'z:' + this.rotate.z + '} }';
}
},
destroy: function() {
$('#' + this.matricule).remove();
delete container.getSlide(this.matricule);
}
/* Important
* Ne pas oublier d'appeler une fonction 'initHtml' à la fin du //constructeur 'init'
* 'initHtml se charge d'écrire l'objet dans le DOM à partir d'un template Mustache propre à la fille
*/
});
/* Class Texte
* properties :
* hierarchy
* content
* changement matricule
*/
Text = Element.extend({
init: function(slide, params) {
// Appelle du //constructeur de la mere
// Il écrit la totalité des objets de params dans les attributs de la mere
// Il prend en compte les attributs qui ne sont pas dans la mère, il faut alors
// définir des watches au besoin.
if (typeof params === 'undefined') {
params = {};
}
if (typeof slide === 'undefined') {
slide = 'null';
}
//le constructeur mère a besoin du matricule pour renseigner le container.
//ainsi, si params contient un matricule, il faut lui passer en priorité :
if (typeof params !== 'undefined')
if (typeof params.matricule !== 'undefined')
var matricule = params.matricule;
else {
var matricule = 'texteelement' + globalCpt++;
//check if unique matrcule already exists
while (typeof findObjectOfComposant(this.matricule) !== 'undefined') { //le matricule existe déjà !
var matricule = 'texteelement' + globalCpt++;
this.matricule = matricule;
}
}
this.matricule = matricule;
if (!this._super(slide, params, matricule)) {
return 0;
}
//pour mustache
this.texte = 'true';
//attribut propre aux textes
this.properties = {
hierarchy: 'bodyText',
content: 'Type text here'
};
//prise en compte des parametres utilisteurs
if (typeof params !== 'undefined') {
for (var param in params) {
if (typeof params[param] === 'object') {
for (var paramNested in params[param]) {
this[param][paramNested] = params[param][paramNested];
}
} else {
this[param] = params[param];
}
}
}
//definition des watch qui permettent d'agir sur le DOM lorsqu'on agit sur les objets des slides
watch(this.properties, function(attr, action, newVal, oldVal) {
//mise à jour du DOM
var $element = $('#' + matricule);
if (attr === 'content') {
//redondant si le texte est édité via contenteditable
$element.children().html(newVal);
} else if (attr === 'hierarchy') {
$element.children().removeClass().addClass(newVal);
}
});
//ajout dans le DOM
this.DOM(slide);
},
//ajout dans le DOM
//cette fonction est appelé deux fois huhu
DOM: function(slide) {
this.show();
var template = $('#templateElementCK').html();
var html = Mustache.to_html(template, this);
////console.log('html', html);
$('#' + slide).append(html);
var newEl = $('#' + this.matricule);
newEl.children('span').html(this.properties.content);
handlerComposant(newEl);
/* si pos.y = 'center', il faut centrer le texte en Y, le HTMl ne permet pas de faire cela
Le mustache insere le html, ensuite je passe derrière pour centrer le texte en y en fn de sa taille */
if (this.pos.y === 'center') {
bidule = $('#' + this.matricule);
var heightTxt = parseInt($('#' + this.matricule).css('height'));
var posTxt = globalConfig.heightSlide / 2 - heightTxt / 2;
$('#' + this.matricule).css('top', posTxt);
this.pos.y = posTxt;
}
/* si pos.y = noCollision, il faut s'arranger pour mettre le texte sous son précédent (utile lors de la création automatique)*/
if (this.pos.y === 'noCollision') {
var posPrev = parseInt($('#' + this.matricule).prev().css('top'));
var heightPrev = parseInt($('#' + this.matricule).prev().css('height'));
var posTxt = posPrev + heightPrev + 20;
$('#' + this.matricule).css('top', posTxt);
this.pos.y = posTxt;
}
},
show: function(i) {
if (typeof i === 'undefined') {
var str = '';
str = this._super(24);
str += ' properties: {content: ' + this.properties.content + ' ,hierarchy: ' + this.hierarchy + ' }';
return str;
}
else {
}
}
});
//les images sont en pixel
Image = Element.extend({
init: function(slide, params) {
if (typeof params === 'undefined') {
params = {};
}
if (typeof slide === 'undefined') {
slide = 'null';
}
//le constructeur mère a besoin du matricule pour renseigner le container.
//ainsi, si params contient un matricule, il faut lui passer en priorité :
if (typeof params !== 'undefined')
if (typeof params.matricule !== 'undefined')
var matricule = params.matricule;
else {
var matricule = 'imageelement' + globalCpt++;
//check if unique matricule already exists
while (typeof findObjectOfComposant(this.matricule) !== 'undefined') { //le matricule existe déjà !
var matricule = 'imageelement' + globalCpt++;
this.matricule = matricule;
}
}
this.matricule = matricule;
this._super(slide, params, matricule);
//pour mustache
this.image = true;
//attributs
this.properties = {
};
this.size = {
height: 100,
width: 100
};
this.source = 'images/bleu_twitter.png';
//prise en compte des parametres utilisteurs
if (typeof params !== 'undefined') {
for (var param in params) {
if (typeof params[param] === 'object') {
truc = params[param];
for (var paramNested in params[param]) {
this[param][paramNested] = params[param][paramNested];
}
} else {
this[param] = params[param];
}
}
}
//ajout dans le DOM
var template = $('#templateImage').html();
var test = {src: 'opuet'};
var html = Mustache.to_html(template, this);
$('#' + slide).append(html);
handlerComposant($('#' + this.matricule));
}
});
/* trouver le matricule de la slide contenant le composant
* Si le composant est déjà une slide, retourne le matricule de la slide
* input : matricule d'un composant
* output : matricule de la slide le contenant
*/
function getSlideMother(matricule) {
if (typeof container.getSlide(matricule) === 'undefined') { //si le matricule n'est pas celui d'une slide
//$.each(container.slide, function(rien, slide) { //parcours des slides
for (var mat in container.slide) {
var slide = container.slide[mat];
if (typeof slide.element[matricule] === 'undefined') {
////console.log('pas dans la slide ', slide);
} else { //si le matricule est un element de la slide, on return le matricule de sa mere
return slide.matricule;
}
}//});
} else { //si le matricule est celui d'une slide
return matricule;
}
return 'Error : matricule doesn\'t existe';
}
/* trouve l'object en fonction du matricule (que ce soit une slide ou un composant)
*
* @type object composant
*/
function findObjectOfComposant(matricule){
//console.log('info : findObjectOfComposant effectue un test avec getSlide');
if (typeof container.getSlide(matricule) === 'undefined') { //si le matricule n'est pas celui d'une slide
for( var i in container.slide) { //parcours des slides
// //console.log('debug findobjofcptmr',slide,slide.element,matricule,typeof slide.element[matricule]);
if (typeof container.slide[i].element[matricule] === 'undefined') {
// //console.log('pas dans la slide ', slide);
} else {
return container.slide[i].element[matricule]; //si le matricule est un element de la slide, on return l'object complet
}
};
} else { //si le matricule est celui d'une slide
return container.getSlide(matricule);
}
//console.log('Error : findObjectOfComposant : matricule '+matricule+' doesn\'t existe');
return;
} <file_sep>/README.md
jmpresseditor2.js
=================
Second version of the impress.js editor
Editeur Jmpress nouvelle version, mode d'emploi
Lancer le fichier HTMLpage.html pour accéder à l'éditeur 100% Javascript dont seulement 3000 lignes très aérées et documentées sont nécessaires (hors bibliothèques Mustache, Jquery, SimpleInheritance, Watch)
En terme de vocabulaire, une présentation est composée de composants répartis en deux famille; les slides et les éléments. Les éléments n'existent que dans les slides et peuvent être du texte ou bien une image. Il faut au moins une slide pour ajouter des éléments.
Avec le clavier
Créer des éléments
Pour peu que le curseur ne survole aucuns composants (ni slides, ni éléments), il est possible de créer des composants en appuyant sur les lettres présentées ci-dessus à savoir :
• j : slide
• k : texte de titre 1 (gros)
• l : texte de titre 2
• m : texte de titre 1 (petit)
• ù : texte de corps de texte
• * : Image
L'ajout de slide se fait au centre de la présentation, l'ajout d'élément se fait au milieu de la slide. Les textes peuvent se superposer, pas les images.
L'ajout d'un élément (texte divers ou image) demande à l'user de sélectionner une slide en cliquant sur l'une d'elle. L'ajout d'une image demande la source de l'image qui peut être un fichier dans le dossier Image à la racine ou bien une URL d'une image sur internet.
Déplacer un composant
Lorsque le curseur survole un composant (une slide ou du texte ou une image) vous pouvez déplacer le composant en utilisant les touches comme mentionnées ci-dessus à savoir :
• z : Y vers le haut
• x : Y vers le bas
• q : X vers la droite
• s : X cers la gauche
• a : Z en profondeur (n'existe que pour les slides)
• x : Z en hauteur (n'existe que pour les slides)
Les slides peuvent également subir une rotation selon le même principe. Les lettres (X,Y,Z) mentionnées ci-dessus correspondent aux axes d'un tétraèdre régulier dont le plan (0,X,Y) est celui de l'écran au début de la présentation. Lorsque la caméra aura tournée le plan de l'écran ne sera plus le même.
Se déplacer dans la présentation
Lorsque le curseur ne survole aucunes slides ni aucuns composants les mêmes touches que pour le déplacement et la rotation des slides permettent de se déplacer dans la présentation et de la faire tourner.
Pour le moment la rotation se fait avec comme centre de rotation le centre de la présentation. C'est contre-intuitif et il est facile de se perdre dans le monde des slides. Afin de constater l'effet de la rotation le mieux est de se mettre proche du centre du monde des slides c’est-à-dire de sorte à bien voir la slide 'profondeur' (utiliser les flèches pour passer d'une slide à l'autre).
Editer le texte
Un simple click sur un élément texte permet de l'éditer.
<file_sep>/js/kernel_transform3d.js
/*
* To change this template, choose Tools | Templates
* and open the template in the editor.
*/
/*
* Fonction Jquery permettant d'effectuer un déplacement animé et smooth via la transform3D
* from http://cameronspear.com/blog/animating-translate3d-with-jquery/
* Great thanks to Cameronspear 2013 !
*/
(function($) {
var delay = 0;
$.fn.translate3d = function(translations, speed, easing, complete) {
var opt = $.speed(speed, easing, complete);
opt.easing = opt.easing || 'ease';
translations = $.extend({x: 0, y: 0, z: 0}, translations);
return this.each(function() {
var $this = $(this);
/* tentative avec mes outils */
var dico = getTrans3D($this);
if (typeof dico.translate3d === 'undefined') {
//console.log($this, 'n\'a pas de transform !');
return;
}
dico.translate3d[0] = translations.x;
dico.translate3d[1] = translations.y;
dico.translate3d[2] = translations.z;
$this.css({
transitionDuration: opt.duration + 'ms',
transitionTimingFunction: opt.easing,
transform: "translate(" + dico.translate[0] + "%, " + dico.translate[1] + "%) scaleX(" + dico.scaleX + ") scaleY(" + dico.scaleY + ") scaleZ(" + dico.scaleZ + ") translate3d(0px,0px,0px) scaleX(1) scaleY(1) scaleZ(1) rotateZ(" + dico.rotateZ + "deg) rotateY(" + dico.rotateY + "deg) rotateX(" + dico.rotateX + "deg) translate3d(" + dico.translate3d[0] + "px," + dico.translate3d[1] + "px, " + dico.translate3d[2] + "px)"
});
setTimeout(function() {
$this.css({
transitionDuration: '0s',
transitionTimingFunction: 'ease'
});
opt.complete();
}, opt.duration + (delay || 0));
});
};
})(jQuery);
/* pour modifier le css transform de la mere*/
/*
*
* @returns un objet avec les attributs du transform en key/value
rotateX:
rotateY:
rotateZ:
scaleX:
scaleY:
scaleZ:
translate: Object
0: -50
1: -50
__proto__: Object
translate3d: Object
0:
1:
2:
*/
function getTrans3D($node) {
////console.log('getTrans3D',$node);
var prefix = (pfx('transform'));
if (typeof $node[0] === 'undefined')
return;
var trans = $node[0].style['' + prefix + ''].match(/.+?\(.+?\)/g);
var dico = {};
for (var el in trans) {
var ele = trans[el];
var key = ele.match(/.+?\(/g).join("").match(/[a-zA-Z0-9]/g).join("");
var value = ele.match(/\(.+\)/g)[0].split(",");
if (value.length <= 1) {
value = parseFloat(value[0].match(/-[0-9]+|[0-9]+/g)[0]);
dico[key] = value;
} else {
dico[key] = {};
for (val in value) {
var vale = parseFloat(value[val].match(/-[0-9]+|[0-9]+/g)[0]);
dico[key][val] = vale;
}
}
}
return dico;
}
/*
* prend en parametre un object contenant TOUS les attributs du transform
* @param {type} dico
*
*/
function setTrans3D(dico, $node) {
if (typeof $node === 'undefined') {
$node = $("#slideArea>div");
}
//var transform = "translate(" + dico.translate[0] + "%, " + dico.translate[1] + "%) scaleX(" + dico.scaleX + ") scaleY(" + dico.scaleY + ") scaleZ(" + dico.scaleZ + ") rotateX(" + dico.rotateX + "deg) rotateY(" + dico.rotateY + "deg) rotateZ(" + dico.rotateZ + "deg) translate3d(" + dico.translate3d[0] + "px," + dico.translate3d[1] + "px, " + dico.translate3d[2] + "px)";
var transform = "translate(" + dico.translate[0] + "%, " + dico.translate[1] + "%) scaleX(" + dico.scaleX + ") scaleY(" + dico.scaleY + ") scaleZ(" + dico.scaleZ + ") translate3d(0px,0px,0px) scaleX(1) scaleY(1) scaleZ(1) rotateZ(" + dico.rotateZ + "deg) rotateY(" + dico.rotateY + "deg) rotateX(" + dico.rotateX + "deg) translate3d(" + dico.translate3d[0] + "px," + dico.translate3d[1] + "px, " + dico.translate3d[2] + "px)";
$node.css({'transform': transform});
}
/* ======================================================================================
* petits utilitaires
* from jmpress.js
* used to get the right vendor prefix
* ====================================================================================== */
/**
* Set supported prefixes
*
* @access protected
* @return Function to get prefixed property
*/
pfx = (function() {
var style = document.createElement('dummy').style,
prefixes = 'Webkit Moz O ms Khtml'.split(' '),
memory = {};
return function(prop) {
if (typeof memory[ prop ] === "undefined") {
var ucProp = prop.charAt(0).toUpperCase() + prop.substr(1),
props = (prop + ' ' + prefixes.join(ucProp + ' ') + ucProp).split(' ');
memory[ prop ] = null;
for (var i in props) {
if (style[ props[i] ] !== undefined) {
memory[ prop ] = props[i];
break;
}
}
}
return memory[ prop ];
};
}());
Transform3D = Class.extend({
init: function() {
var transform = getTrans3D($('#slideArea >'));
this.pos = {
x: 0, //transform.translate3d[0],
y: 0, //transform.translate3d[1],
z: 0//transform.translate3d[2]
};
this.rotate = {
x: 0, //transform.translate3d[0],
y: 0, //transform.translate3d[1],
z: 0//transform.translate3d[2]
};
watch(this.pos, function(attr, action, newVal, oldVal) {
WatchJS.noMore = true; //prevent invoking watcher in this scope
this.x = 0;
this.y = 0;
this.z = 0;
// var transform = 'translate(-50%, -50%), scaleX(1), scaleY(1), scaleZ(1), rotateZ(0deg), rotateY(0deg), rotateX(0deg), \n\
// translate3d(' + this.x + 'px, ' + this.y + 'px, ' + this.z + 'px)';
// //console.log('transform :', transform);
// $("#slideArea>div").css({'transform': transform});
// $("#slideArea").css({'transform': transform});
//magouille qui fera fonctionner
var dico = getTrans3D($('#slideArea >'));
var i;
switch (attr) {
case 'x':
i = 0;
break;
case 'y':
i = 1;
break;
case 'z':
i = 2;
break;
}
dico.translate3d[i] += newVal;
setTrans3D(dico);
//console.log(dico);
});
watch(this.rotate, function(attr, action, newVal, oldVal) {
WatchJS.noMore = true; //prevent invoking watcher in this scope
this.x = 0;
this.y = 0;
this.z = 0;
// var transform = 'translate(-50%, -50%), scaleX(1), scaleY(1), scaleZ(1), rotateZ(0deg), rotateY(0deg), rotateX(0deg), \n\
// translate3d(' + this.x + 'px, ' + this.y + 'px, ' + this.z + 'px)';
// //console.log('transform :', transform);
// $("#slideArea>div").css({'transform': transform});
// $("#slideArea").css({'transform': transform});
//magouille qui fera fonctionner
var dico = getTrans3D($('#slideArea >'));
var i;
switch (attr) {
case 'x':
i = 'X';
break;
case 'y':
i = 'Y';
break;
case 'z':
i = 'Z';
break;
}
dico['rotate' + i] += newVal;
setTrans3D(dico);
//console.log(dico);
});
}
});<file_sep>/js/auto_create.js
/*
*
* Calcul des positions d'une présentation à partir d'une liste comprenant le plan et les contenus
*
* Mode présentation à plat avec changement de niveau à chaque changement de partie
* Les contents sont au meme niveau que leur partie, vers le bas
*
*
*
*/
function initAutomatic() {
initContainer();
container.metadata.type = 'tree';
container.metadata.name = $('#slideshowNameTree').html();
// var config = {
// cranX: 1800,
// cranY: 1000,
// cranZ: -1000,
// liveX0: 10000,
// liveY0: 10000,
// liveZ0: 10000,
// endX0: -10000,
// endY0: -10000,
// endZ0: -10000
// };
var config = {
cranX: globalConfig.widthSlide * 1.5,
cranY: globalConfig.heightSlide * 1.5,
cranZ: -1000,
liveX0: -1500,
liveY0: 0,
liveZ0: 0,
endX0: 0,
endY0: -1500,
endZ0: 0
};
// goCK(config);
goNormalize();
// //console.log('de');
goDepth(config);
// //console.log('pos');
goPosition(config);
// //console.log('posEnd');
goPositionEnd(config);
// //console.log('jmpre');
goJmpress(config);
// //console.log('dyna');
dynamic(config);
alertify.success('slideShow '+container.metadata.name+' created </br> use SPACE to navigate </br> click \'tree\' to edit again ');
}
function goNormalize() {
$('#tree ol').each(function() {
if ($(this).children().length === 0) { //si l'ol a été ajouté pour 'rien'
$(this).remove();
}
});
$('#tree .textarea').each(function() {
// var content = $(this).val();
var content = $(this).html();
content = '<span class=\'textarea\'>' + content + '</span>';
$(this).parent().attr('type', 'body');
$(this).parent().append(content);
$(this).remove();
});
$('#tree li').each(function() {
if (typeof $(this).attr('type') === 'undefined')
$(this).attr('type', '');
});
}
/*
* Determination des positions des slides pendant la présentation, les coordonnées sont stockées dans la liste
* Chaque li représente une slide
*
*/
function goPosition(config) {
var cranX = config.cranX;
var cranY = config.cranY;
var cranZ = config.cranZ;
//////calcul des positions
$('#tree').attr('number', '');
$('#tree').prepend("<span style='display:none'>Jmpress Editor -</span>");
//premiers niveaux
$('#tree li').each(function() {
if ($(this).attr('depth') === '1') {
var x = config.liveX0 + parseInt($(this).index()) * cranX;
var y = config.liveY0;
if ($(this).index() === 0) { //initialisation de la positio de la toute première slide
var z = config.liveZ0;
} else {
var z = parseInt($(this).prev().attr('data-z')) + (parseInt(maxDepth($(this).prev(), 0))) * cranZ;
}
$(this).attr('data-x', x).attr('data-y', y).attr('data-z', z).attr('data-rotate-x', '-45');
$(this).attr('type', 'title');
var indice = parseFloat($(this).index()) + 1;
$(this).attr('number', indice);
}
});
//les autres niveaux (ne pas factoriser avec le traitement des premiers niveaux car les autres niveaux ont besoin des premiers niveaux pour s'appuyer
$('#tree li').each(function() {
if ($(this).attr('depth') !== '1' && $(this).attr('type') !== 'body') {//&& $(this).attr('nbChild') !== '0') {
if ($(this).index() === 0) { //si première fille
var x = $(this).parent().parent().attr('data-x');
} else {
var x = parseInt($($(this).prev()).attr('data-x')) + cranX; //position de sa grande soeur
}
var y = parseInt($(this).parent().parent().attr('data-y')) + cranY; //pour atteindre la li qui la stocke
var z = parseInt($(this).parent().parent().attr('data-z')) + parseInt($(this).index()) * cranZ;
$(this).attr('type', 'title');
// } else if ($(this).attr('depth') !== '1' && $(this).attr('nbChild') === '0') { //si pas d'enfants, c'est du contenu, slides horizontales
} else if ($(this).attr('depth') !== '1' && $(this).attr('type') === 'body') { //slide horizontales
var x = parseInt($(this).parent().parent().attr('data-x'));//+ $(this).index() * cranX / 2;
var y = parseInt($(this).parent().parent().attr('data-y')) + parseInt($(this).index() + 1) * cranY; //pour atteindre la li qui la stocke
var z = parseInt($(this).parent().parent().attr('data-z'));//+ parseInt($(this).index()) * cranZ;
$(this).attr('type', 'body');
}
$(this).attr('data-x', x).attr('data-y', y).attr('data-z', z);
var indice = parseFloat($(this).index()) + 1;
var parentNumber = $(this).parent().parent().attr('number');
$(this).attr('number', (parentNumber === '') ? indice : parentNumber + "." + indice);
});
}
/*
* Calcul des positions des slides pour l'état final
* Les coordonnées sont stockées dans la liste ol, chaque li correspond à une slide
*/
function goPositionEnd(config) {
var cranX = config.cranX;
var cranY = config.cranY;
var cranZ = config.cranZ;
//archivage de la coord Y de la plus haute et de la plus basse slide pour l'overview final
var upperY = config.endY0;
var lowerY;
//premiers niveaux
$('#tree li').each(function() {
//console.log('tree li', $(this));
if ($(this).attr('depth') === '1') {
var x = config.endX0;
var z = config.endZ0;
if ($(this).index() === 0) { //initialisation de la positio de la toute première slide
var y = upperY;
} else {
var y = parseInt($(this).prev().attr('data-end-y')) + (parseInt(maxDepth($(this).prev(), 0))) * cranY * 1.2;
var y = parseInt(getLastChild($(this).prev()).attr('data-end-y')) + cranY;
}
$(this).attr('data-end-x', x).attr('data-end-y', y).attr('data-end-z', z);
$(this).attr('type', 'title');
var indice = parseFloat($(this).index()) + 1;
$(this).attr('number', indice);
lowerY = y;
}
// });
//les autres niveaux
// $('#tree li').each(function() {
if ($(this).attr('depth') !== '1' && $(this).attr('nbChild') !== '0') {
var x = config.endX0 + (parseInt($(this).attr('depth')) - 1) * cranX;
var z = config.endZ0;
if ($(this).index() === 0) { //si première fille
var y = parseInt($(this).parent().parent().attr('data-end-y')) + ($(this).index() + 1) * cranY * 0.5;
} else {
// var y = parseInt($($(this).prev()).attr('data-end-y')) + (maxDepth($(this).prev(), 0) - parseInt($(this).attr('depth'))) * cranY ;//* 1.2; //position de sa grande soeur//depth-1 pour virer le content
var y = parseInt(getLastChild($(this).prev()).attr('data-end-y')) + cranY;
}
$(this).attr('data-end-x', x).attr('data-end-y', y).attr('data-end-z', z);
$(this).attr('type', 'title');
var indice = parseInt($(this).index()) + 1;
$(this).attr('number', $(this).parent().parent().attr('number') + "." + indice);
} // sinon c'est du contenu, on ne s'en occupe pas
});
//ajout de la slide de questions
var li = "<li class='questions' uppery = '" + upperY + "' lowery = '" + lowerY + "' >Any questions ?</li>";
$('#tree').children('ol').append(li);
//lowerY a été instancié pour la dernière fois par l'element le plus 'bas' dans la lsite ul li
}
/*
* A partir de la liste qui a été parsée pour ajouter les coordonnées, créer les slides et leurs contenus
* Ajoute également les slides qui ne font pas partie de la présentation, à savoir les overviews, les slides d'accueils et de concluion (any questions ?)
*/
function goJmpress(config) {
initJmpress();
//creation des slides jmpress
$('#tree li').each(function() {
if (typeof $(this).attr('uppery') !== 'undefined') {
var upperY = parseInt($(this).attr('uppery'));
var lowerY = parseInt($(this).attr('lowery'));
new Slide({
auto: true,
properties: {
scale: Math.abs((upperY - lowerY)) * 4 / 3 / 1000
},
matricule: 'end',
pos: {
x: config.endX0 + 10000,
y: (upperY + lowerY) / 2,
z: config.endZ0
},
type: 'overview'
}
);
var slide = new Slide({
auto: true,
matricule: 'questions',
properties: {
//hierarchy: '' + $(this).attr('number'),
scale: 10
},
pos: {
x: config.endX0 + 15000,
y: Math.abs(upperY - lowerY) / 2 + upperY,
z: config.endZ0
}
}
);
new Text('questions', {
//matricule: 'questionstexte',
properties: {
content: '<em>Merci de votre attention</em> <br/> Avez vous des questions ?',
hierarchy: 'H1Text'
},
pos: {
x: 0,
y: 'center'//globalConfig.heightSlide/2
}
});
return;
}
//si besoin ajout de l'overview
if ($(this).attr('type') === 'title') {
//s'il y au moins une petite soeur
////console.log($(this).index() , parseInt($(this).attr('siblings')) -1);
if ($(this).index() < parseInt($(this).attr('siblings')) - 1) {
// //console.log($($(this).siblings()[parseInt($(this).attr('siblings')) - 2]),$(this).siblings());
new Slide({
type: 'overview',
pos: {
//long bug à trouver ! Il faut -2 dans les siblings car le siblings ne compatabilise pas lui meme, logique !
x: (parseInt($(this).attr('data-x')) + parseInt($($(this).siblings()[parseInt($(this).attr('siblings')) - 2]).attr('data-x'))) / 2,
y: $(this).attr('data-y'),
z: $(this).attr('data-z')
},
properties: {
scale: $(this).attr('siblings')
}
});
}
}
//ajout de la slideet de son texte
var numberArray = $(this).attr('number').split('.');
number = numberArray.join('-');
//console.log(number);
var slide = new Slide({
matricule: number,
style: $(this).attr('type'),
pos: {
x: $(this).attr('data-x'),
y: $(this).attr('data-y'),
z: $(this).attr('data-z')
},
properties: {
hierarchy: '' + $(this).attr('number'),
scale: 1
}
});
if ($(this).children('.textarea').length !== 0) {
//rappel de la partie
var upHierarchy = $(this).parent('ol').siblings('span');
// //console.log('debug : goJmpress',upHierarchy);
new Text(slide.matricule, {
auto: true, //il ne faut que le treeMakerFromContainer en tienne compte
properties: {
content: upHierarchy.html(),
hierarchy: 'H3Text'
},
pos: {
x: 0,
y: globalConfig.heightSlide * 0.05
}
});
/*** gestion spéciale pour le contenu en code à mettre sous hilight (wrapper dans un pre code avec un id puis init via hilight.js **/
/*
if ($(this).children('.textarea').html().search('<?php') !== -1) {
//remplacement des balises php par PHP
var test = $('.textarea').html().replace('<?php', 'PHP');
test = test.replace('?>', 'PHPF');
//suppression des sauts de lignes
test = test.replace(/(\r\n|\n|\r)/gm, "");
//recupération du contenu entre PHP et PHPF
test = test.match(/PHP(.*?)PHP/i);
//recupération que ce qui nous interesse
var code = '';
test = test[0].match(/<p>(.*?)<\/p>/g);
$(test).each(function(indice, txt) {
////console.log(indice,txt,test[indice]);
code += test[indice].match(/<p>(.*?)<\/p>/i)[1] + '\n';
});
code = "<pre id='HAHA' class='php'><code class='php'>"+code+"</code></pre>";
truc = code;
// $(this).children('.textarea').append(code);
} else {
truc = $(this).children('.textarea').html();
}*/
//contenu
new Text(slide.matricule, {
properties: {
hierarchy: 'bodyText',
content: $(this).children('.textarea').html()
},
pos: {
x: 0,
y: 'noCollision'//globalConfig.heightSlide/2
}
});
} else if ($(this).children('.liTitle').length !== 0) {
new Text(slide.matricule, {
properties: {
hierarchy: 'H1Text',
content: numberArray.join('.') + ' | ' + $(this).children('.liTitle').html()
},
pos: {
x: 0,
y: 'center'
}
});
}
//ecriture du matricule de la slide dans la liste
$(this).attr('matricule', slide.matricule);
$(this).attr('id', 'li_' + slide.matricule);
});
} //fin goJmpress
<file_sep>/js/gui_mouse.js
/*
* Contient les élements communs à slideShowEditor et GUIEditor à savoir les outils permettant à l'Editor en lui même de fonctionner
* Contient :
* class objectEvent
* listener joystick
* appel directement Model avec un objEvt
* orthogonalProjection
*
*/
/**
* jQuery mousehold plugin - fires an event while the mouse is clicked down.
* Additionally, the function, when executed, is passed a single
* argument representing the count of times the event has been fired during
* this session of the mouse hold.
*
* @author <NAME> (<EMAIL>)
* @date 2006-12-15
* @example $("img").mousehold(200, function(i){ })
* @desc Repeats firing the passed function while the mouse is clicked down
*
* @name mousehold
* @type jQuery
* @param Number timeout The frequency to repeat the event in milliseconds
* @param Function fn A function to execute
* @cat Plugin
*/
/***
fn.call(thisPerso,params1...)
permet d'appliquer un this passé en parametre et des parametres
***/
/**
ca fait chier, le mousehold lance la même fonction à chaque fois ! mais l'event n'est jamais le même !
**/
jQuery.fn.mousehold = function(timeout, f) {
if (timeout && typeof timeout === 'function') {
f = timeout;
timeout = 100;
}
if (f && typeof f === 'function') {
var timer = 0;
var fireStep = 0;
return this.each(function() {
//jQuery(this).on('mousedown', function() {
fireStep = 1;
var ctr = 0;
var t = this;
timer = setInterval(function() {
ctr++;
f.call(t, ctr);
fireStep = 2;
}, timeout);
//});
clearMousehold = function() {
clearInterval(timer);
if (fireStep == 1)
f.call(this, 1);
fireStep = 0;
// if( $(this).attr('id') === 'mouseholdId') $(this).removeAttr('id');
};
//pour exclure le joystick du mouseout
//si this n'a pas d'id => does'nt work !
// if( typeof $(this).attr('id') === 'undefined' ) $(this).attr('id','mouseholdID');
var $this = $('#'+$(this).attr('id')+':not(#joystick)');
$this.mouseout(clearMousehold);
$(this).mouseup(clearMousehold);
});
}
}
/* joystick
* Fournit un ensemble de listener sur le mousedown+mousemove
* Detecte la slide et calcul le déplacement de la souris par rapport au centre
*
*/
/* effectue le projeté orthogonal du deplacement du joystick
*
* output : sens sur X
* sens sur Y
* cran sur X
* cran sur Y
*/
function orthogonalProjection(current, init, coef) {
var dx = current.pageX - init.pageX;
var dy = current.pageY - init.pageY;
var tolerance = 20;
var d = {
/* explication du bout de code ci dessous :
* condition ternaire, si la distance est inférieur à la tolérance, aucune rotation
* sinon on soustrait à la distance la tolérance doté du signe de la distance (abs(x)/x = signe de x)
*/
y: ((Math.abs(dy) - tolerance) >= 0 ? (dy - tolerance * Math.abs(dy) / dy) : 0),
x: ((Math.abs(dx) - tolerance) >= 0 ? (dx - tolerance * Math.abs(dx) / dx) : 0)
};
//var dist = Math.sqrt(Math.pow(dx, 2) + Math.pow(dy, 2));
var event = {
cranX: 0,
directionX: '',
cranY: 0,
directionY: ''
};
event.cranX = Math.abs(d.x) * coef;
if (dx < 0) {
event.directionX = 'x-';
} else {
event.directionX = 'x+';
}
event.cranY = Math.abs(d.y) / 10;
if (dy < 0) {
event.directionY = 'y-';
} else {
event.directionY = 'y+';
}
return event;
}
function setJoystick($this) {
var heightJ = parseInt($('#joystick').css('height'));
var widthJ = parseInt($('#joystick').css('width'));
$('#joystick').css({
'top': $this.data('posInitMouse').pageY - heightJ / 2,
'left': $this.data('posInitMouse').pageX - widthJ / 2,
'display': 'inline'
});
}
/* Pour faire les choses bien :
* https://github.com/sebastien-p/jquery.hasEventListener
* Permet de savoir si un element du DOM a déjà un listener
*
* Bricollage :
* variable globale pour savoir si le mousemove a été capté
* mouseMove = false -> mousemove libre
*/
simpleHasMouseMove = false;
longHasMouseMove = false;
/* listeners pour le joystick
* Un mousedown n'importe
*
*/
$(document).on('mousedown', function(event) {
if (
$('.sidebar:hover').length === 0 &&
$('#topbar:hover').length === 0 &&
$('.buttonclicked').length === 0) {
joystickHandler(event);
}
});
function joystickHandler(event) {
var $this = $(event.target);
if ($('body')[0] === $this[0]) {
var matricule = 'document';
} else {
var matricule = $this.attr('matricule');
//stockage des données initiales
var composant = findObjectOfComposant(matricule);
$this.data('pos', {
x: composant.pos.x,
y: composant.pos.y,
z: composant.pos.z
});
$this.data('rotate', {
x: composant.rotate.x,
y: composant.rotate.y,
z: composant.rotate.z
});
$this.data('scale', composant.scale);
}
$this.data('posInitMouse', {
pageX: event.pageX,
pageY: event.pageY
});
var objEvt = new ObjectEvent({
matricule: matricule,
action: 'joystick',
event: {
cran: 0,
direction: ''
}
});
setJoystick($this);
/*
* gestion des long press click
*/ /*
if (event.which === 1) { //longpress left
$this.data('checkdown', setTimeout(function() {
//console.log('long left press sur', matricule);
if (simpleHasMouseMove) {
//console.log('long left : mouse move déja pris ');
} else {
$(document).on('mousemove.longLeft', function(event) { //si on move penant un click long
longHasMouseMove = true;
//console.log('mousemove après long left sur', matricule);
});
}
}, 500)).on('mousemove', function() {
clearTimeout($(this).data('checkdown'));
});
}
if (event.which === 3) { //longpress right
$this.data('checkdown', setTimeout(function() {
//console.log('long right press sur', matricule);
if (simpleHasMouseMove) {
//console.log('long right : mouse move déja pris ');
} else {
$(document).on('mousemove.longRight', function(event) { //si on move penant un click long
longHasMouseMove = true;
//console.log('mousemove après long right sur', matricule);
});
}
}, 500)).on('mousemove', function() {
clearTimeout($(this).data('checkdown'));
});
}
*/
/*
* gestion des simple click
*/
if (event.which === 1) {
// $(document).on('mousemove.simpleLeft',function() { //si on move penant un click court
$(document).mousehold(function() { //si on move penant un click court
if (longHasMouseMove) {
console.log('long click a pris le mousemove');
$(this).off('.simpleLeft');
} else {
simpleHasMouseMove = true;
//console.log('mousemove left sur', matricule);
var event = {
pageX: parseInt($('body').data('pageX')),
pageY: parseInt($('body').data('pageY'))
};
var eventXY = orthogonalProjection(event, $this.data('posInitMouse'), 0.1);
//console.log(eventXY.pageX, eventXY.pageY);
objEvt.action = 'move';
//maj position X
objEvt.event.direction = eventXY.directionX;
objEvt.event.cran = eventXY.cranX;
//console.log('avant appel call model', objEvt);
callModel(objEvt);
//maj position Y
objEvt.event.direction = eventXY.directionY;
objEvt.event.cran = eventXY.cranY;
//console.log('avant appel call model', objEvt);
callModel(objEvt);
}
});
}
if (event.which === 3) {
// $(document).on('mousemove.simpleLeft',function(event) { //si on move penant un click court
$(document).mousehold(function(event) { //si on move penant un click court
if (longHasMouseMove) {
//console.log('long click a pris le mousemove');
$(this).off('.simpleRight');
} else {
simpleHasMouseMove = true;
console.log('mousemove right sur', matricule);
var event = {
pageX: parseInt($('body').data('pageX')),
pageY: parseInt($('body').data('pageY'))
};
var eventXY = orthogonalProjection(event, $this.data('posInitMouse'), 0.1);
objEvt.action = 'rotate';
//maj position X
objEvt.event.direction = eventXY.directionX;
objEvt.event.cran = eventXY.cranX;
//console.log('avant appel call model', objEvt);
callModel(objEvt);
//maj position Y
objEvt.event.direction = eventXY.directionY;
objEvt.event.cran = eventXY.cranY;
//console.log('avant appel call model', objEvt);
callModel(objEvt);
}
});
}
//annulation des listeners sur mouseup
$(document).one('mouseup', function(event) {
// $('.buttonclicked"').removeClass("buttonclicked");
simpleHasMouseMove = false;
longHasMouseMove = false;
$(this).off('.longClick');
$(this).off('.longLeft');
$(this).off('.longRight');
//$(this).off('.simpleClick');
//$(this).off('.simpleLeft');
//$(this).off('.simpleRight');
console.log('annulation des events mouse');
$('#joystick').css('display', 'none');
});
}
/*
* pour le mousehold
*/
$(document).on('mousemove', function(event) {
$('body').data('pageX', event.pageX);
$('body').data('pageY', event.pageY);
//console.log($('body').css('cursor'));
});
/* permet de savoir qui de document ou de slide doit capter les touches de clavier en fonction du hover
*
*/
var composantCatchEvent = false;
//var documentCatchEvent = false;
/*
* Gère l'édition de la présentation (des composants, après création) :
* handlerComposant : listener de hover et keyboard pour le deplatement/rotation des composants
*
*
* a couper en deux : mouse et keyboard séparés
*/
/*
* call après l'insertion dans le DOM
*/
function handlerComposant($composant) {
$composant.on('mouseenter', function(event) {
// ('#sidebar:hover').length
composantCatchEvent = true;
event.stopPropagation();
var $target = $(this);
//console.log('hover', $target.attr('matricule'));
//fire contenteditable
if ($target.hasClass('texte')) { //si c'est du texte on place un trigger pour rendre le contenu editable via un click
// $target.on('click', function(){
// //console.log('click texte');
// $target.attr('contenteditable','true');
//
// });
$target.children().one('click', lauchCK);
}
//fire keyboard event
$(document).on('keypress.keySlide', function(event) {
//console.log('key ', $(this).attr('id'));
var matricule = $target.attr('matricule');
var obj = findObjectOfComposant(matricule);
var objEvt = new ObjectEvent({
matricule: matricule,
event: {
}
});
switch (event.which) {
//deplacement composant
case 97:
objEvt.action = 'move';
objEvt.event.direction = 'z+';
break;
case 122:
objEvt.action = 'move';
objEvt.event.direction = 'y-';
break;
case 113:
objEvt.action = 'move';
objEvt.event.direction = 'x-';
break;
case 115:
objEvt.action = 'move';
objEvt.event.direction = 'x+';
break;
case 119:
objEvt.action = 'move';
objEvt.event.direction = 'y+';
break;
case 120:
objEvt.action = 'move';
objEvt.event.direction = 'z-';
break;
//rotation composant
objEvt.event.cran = 10;
case 114:
objEvt.action = 'rotate';
objEvt.event.direction = 'z+';
break;
case 102:
objEvt.action = 'rotate';
objEvt.event.direction = 'y-';
break;
case 118:
objEvt.action = 'rotate';
objEvt.event.direction = 'x-';
break;
case 116:
objEvt.action = 'rotate';
objEvt.event.direction = 'x+';
break;
case 103:
objEvt.action = 'rotate';
objEvt.event.direction = 'y+';
break;
case 98:
objEvt.action = 'rotate';
objEvt.event.direction = 'z-';
break;
}
switch (objEvt.action) {
case 'move':
if (obj.type === 'slide') {
objEvt.event.cran = 100;
} else {
objEvt.event.cran = 10;
}
break;
case 'rotate' :
objEvt.event.cran = 10;
break;
}
callModel(objEvt);
//console.log('key slide ', objEvt);
});
});
$composant.mouseleave(function() {
var $target = $(this);
var matricule = $target.attr('matricule');
$(document).off('.keySlide');
//console.log('fin hover', $target.attr('matricule'));
composantCatchEvent = false;
if ($target.hasClass('texte') && $target.children().children('textarea').length !== 0) { //si c'est du texte on place un trigger pour rendre le contenu editable via un click
// $target.attr('contenteditable', 'false');
var $textarea = $(this).children().children('textarea');
var txt = CKEDITOR.instances[$textarea.attr('id')].getData();
//console.log('leave cke', txt);
$(this).children().one('click', lauchCK);
$(this).children().html(txt);
var slideMother = getSlideMother(matricule);
console.log('maj texte dans container', slideMother, matricule,txt);
//mise à jour de l'objet dans le conteneur
container.getSlide(slideMother).element[matricule].properties.content = txt;
}
});
}<file_sep>/nbproject/project.properties
config.folder=
file.reference.GitHub-ImpressEditV2=.
files.encoding=UTF-8
site.root.folder=${file.reference.GitHub-ImpressEditV2}
test.folder=
<file_sep>/js/test_presentation.js
/*
* n'a pas suivi les changements du container.slide objet à conteneur.slide array
*
*
*/
//function test(){
function testcos() {
var i = 0;
var X = 1000;
var Y = 1000;
var Z = 0;
var RX = 0;
var RY = 0;
var RZ = 0;
var r = 5000; //rayon
var alpha = 0; //cran de position des slides sur le cercle //en degré
var beta; //orientation de la slide (en z) pour qu'elle soit tangente au cercle // en degré
var prevX = 0;
var prevY = 0;
truc = [];
while (i < 100) {
//calcul de la position sur la fonction
//verification de la distance minimum
while (Math.sqrt(Math.pow((X - prevX), 2) + Math.pow((Y - prevY), 2)) < 1000) {
X = X + 100;
Y = 3000 * Math.cos(X / 1000);
//console.log(X, Y, Math.sqrt(Math.pow((X - prevX), 2) + Math.pow((Y - prevY), 2)));
//alert( X+' '+ Y+' '+Math.sqrt( Math.pow((X-prevX),2) + Math.pow((Y-prevY),2) ) );
}
truc.push([X, Y]);
var slide = new Slide({
pos: {
x: X,
y: Y,
z: Z
},
rotate: {
x: RX,
y: RY,
z: RZ
}
});
Z = Z - 1000;
prevX = X;
prevY = Y;
i++;
}
}
function testCircle() {
//function test(){
var i = 0;
var X = 0;
var Y = 0;
var Z = 0;
var RX = 0;
var RY = 0;
var RZ = 0;
var r = 5000; //rayon
var alpha = 0; //cran de position des slides sur le cercle //en degré
var beta; //orientation de la slide (en z) pour qu'elle soit tangente au cercle // en degré
var smaller = true;
truc = [];
while (i < 100) {
//calcul de la position autour du cercle
var alphaRad = alpha * Math.PI / 180;
X = Math.sin(alphaRad) * r;
Y = Math.cos(alphaRad) * r;
//calcul de la rotation en Z
// var signeX = X / Math.abs(X);
// var a = 2*X / Math.sqrt( Math.pow(r,2) - Math.pow(X,2) ); //coef directeur de la tangent
//
// beta = Math.acos( signeX / Math.sqrt( Math.pow(a,2) + 1 ) );
//// beta = Math.atan( coef * 2*X / Math.sqrt( Math.pow(r,2) - Math.pow(X,2) ) ) * 180/ Math.PI ;
// RZ = beta * 180/Math.PI;
// truc.push(beta);
var slide = new Slide({
pos: {
x: X,
y: Y,
z: Z
},
rotate: {
x: RX,
y: RY,
z: RZ
}
});
Z = Z - 1000;
if (smaller) {
r = r - 300;
} else {
r = r + 300;
}
var limite = 1000;
if (r < limite) {
smaller = false;
// r = limite + 10;
}
//console.log('RAYON ', r);
alpha += 20; //en degré
i++;
}
}
function test3() {
var i = 0;
var X = 0;
var Y = 0;
var Z = 0;
var RX = 0;
var RY = 0;
var RZ = 0;
while (i < 10) {
var slide = new Slide({
pos: {
x: X,
y: Y,
z: Z
},
rotate: {
x: RX,
y: RY,
z: RZ
}
});
Y += 1000;
RY += 45;
i++;
}
}
//function automaticDynamic(){
function test() {
var cranY = 1000;
var cranZ = -1000;
var cranX = 1200;
new Slide({pos: {x: 0, y: 0, z: 0}, type: 'overview', scale: 2});
//1
var slide1 = new Slide({pos: {x: -1500, y: 0, z: 0}});
new Text(slide1.matricule, {properties: {hierachy: 'H1Text', content: '1'}});
var nbSoeur = 3;
new Slide({pos: {x: slide1.pos.x + (nbSoeur - 1) * cranX / 2, y: slide1.pos.y + cranY, z: slide1.pos.z + cranZ}, type: 'overview', scale: 2});
var slide11 = new Slide({pos: {x: slide1.pos.x, y: slide1.pos.y + cranY, z: slide1.pos.z + cranZ}});
new Text(slide11.matricule, {properties: {hierachy: 'H1Text', content: '1.1'}});
var slide11content = new Slide({pos: {x: slide11.pos.x, y: slide11.pos.y, z: slide11.pos.z + cranZ * 2}});
new Text(slide11content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
new Slide({pos: {x: slide11.pos.x + cranX + (nbSoeur - 2) * cranX / 2, y: slide1.pos.y + cranY, z: slide1.pos.z + cranZ}, type: 'overview', scale: 2});
var slide12 = new Slide({pos: {x: slide11.pos.x + cranX, y: slide1.pos.y + cranY, z: slide1.pos.z + cranZ}});
new Text(slide12.matricule, {properties: {hierachy: 'H1Text', content: '1.2'}});
var slide12content = new Slide({pos: {x: slide12.pos.x, y: slide12.pos.y, z: slide12.pos.z + cranZ * 2}});
new Text(slide12content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide13 = new Slide({pos: {x: slide12.pos.x + cranX, y: slide1.pos.y + cranY, z: slide1.pos.z + cranZ}});
new Text(slide13.matricule, {properties: {hierachy: 'H1Text', content: '1.3'}});
var slide13content = new Slide({pos: {x: slide13.pos.x, y: slide13.pos.y, z: slide13.pos.z + cranZ * 2}});
new Text(slide13content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
new Slide({pos: {x: 750, y: 0, z: 0}, type: 'overview', scale: 2});
//2
var slide2 = new Slide({pos: {x: 0, y: 0, z: 0}});
new Text(slide2.matricule, {properties: {hierachy: 'H1Text', content: '2'}});
var nbSoeur = 2;
new Slide({pos: {x: slide2.pos.x + (nbSoeur - 1) * cranX / 2, y: slide2.pos.y + cranY, z: slide2.pos.z + cranZ}, type: 'overview', scale: 2});
var slide21 = new Slide({pos: {x: slide2.pos.x, y: slide2.pos.y + cranY, z: slide2.pos.z + cranZ}});
new Text(slide21.matricule, {properties: {hierachy: 'H1Text', content: '2.1'}});
var slide21content = new Slide({pos: {x: slide21.pos.x, y: slide21.pos.y, z: slide21.pos.z + cranZ * 2}});
new Text(slide21content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide22 = new Slide({pos: {x: slide21.pos.x + cranX, y: slide2.pos.y + cranY, z: slide2.pos.z + cranZ}});
new Text(slide22.matricule, {properties: {hierachy: 'H1Text', content: '2.2'}});
var nbSoeur = 2;
new Slide({pos: {x: slide22.pos.x + (nbSoeur - 1) * cranX / 2, y: slide22.pos.y + cranY, z: slide22.pos.z + cranZ}, type: 'overview', scale: 2});
var slide221 = new Slide({pos: {x: slide22.pos.x, y: slide22.pos.y + cranY, z: slide22.pos.z + cranZ}});
new Text(slide221.matricule, {properties: {hierachy: 'H1Text', content: '2.2.1'}});
var slide221content = new Slide({pos: {x: slide221.pos.x, y: slide221.pos.y, z: slide221.pos.z + cranZ * 2}});
new Text(slide221content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide222 = new Slide({pos: {x: slide221.pos.x + cranX, y: slide22.pos.y + cranY, z: slide22.pos.z + cranZ}});
new Text(slide222.matricule, {properties: {hierachy: 'H1Text', content: '2.2.2'}});
var slide222content = new Slide({pos: {x: slide222.pos.x, y: slide222.pos.y, z: slide222.pos.z + cranZ * 2}});
new Text(slide222content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
//3
var slide3 = new Slide({pos: {x: 1500, y: 0, z: 0}});
new Text(slide3.matricule, {properties: {hierachy: 'H1Text', content: '3'}});
var slide3content = new Slide({pos: {x: slide3.pos.x, y: slide3.pos.y, z: slide3.pos.z + cranZ * 2}});
new Text(slide3content.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
}
function automaticEditor() {
//function test() {
var slide = new Slide({pos: {
x: 1500,
y: 0,
z: 0
}, scale: 2, type: 'overview'});
var slide = new Slide({pos: {
x: 0,
y: 0,
z: 0
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '1'}});
var slide = new Slide({pos: {
x: 0,
y: 1000,
z: 0
}, scale: 2, type: 'overview'});
var slide = new Slide({pos: {
x: -600,
y: 1000,
z: 0
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '1.1'}});
var slide = new Slide({pos: {
x: -600,
y: 2000,
z: 0
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide = new Slide({pos: {
x: 600,
y: 1000,
z: -2000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '1.2'}});
var slide = new Slide({pos: {
x: 600,
y: 2000,
z: -2000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide = new Slide({pos: {
x: 2200,
y: 0,
z: -4000
}, scale: 2, type: 'overview'});
var slide = new Slide({pos: {
x: 1500,
y: 0,
z: -4000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '2'}});
var slide = new Slide({pos: {
x: 1100,
y: 1000,
z: -4000
}, scale: 2, type: 'overview'});
var slide = new Slide({pos: {
x: 600,
y: 1000,
z: -4000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '2.1'}});
var slide = new Slide({pos: {
x: 600,
y: 2000,
z: -4000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide = new Slide({pos: {
x: 1800,
y: 1000,
z: -6000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '2.2'}});
var slide = new Slide({pos: {
x: 1800,
y: 2000,
z: -6000
}, scale: 2, type: 'overview'});
var slide = new Slide({pos: {
x: 1200,
y: 2000,
z: -6000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '2.2.1'}});
var slide = new Slide({pos: {
x: 1200,
y: 3000,
z: -6000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide = new Slide({pos: {
x: 2400,
y: 2000,
z: -8000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '2.2.2'}});
var slide = new Slide({pos: {
x: 2400,
y: 3000,
z: -8000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
var slide = new Slide({pos: {
x: 3000,
y: 0,
z: -10000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: '3'}});
var slide = new Slide({pos: {
x: 3000,
y: 1000,
z: -10000
}});
new Text(slide.matricule, {properties: {hierachy: 'H1Text', content: 'content'}});
}
//function test() {
function test2() {
//differents moyen de créer une slide
//juste avec les coord
//$('#slideArea>').html('');
var X = 0;
var Y = 0;
var I = 0;
var slide = new Slide({pos: {x: X, y: Y, z: 0}});
new Text(slide.matricule, {properties: {content: 'slide : ' + I}});
var slide = new Slide({pos: {x: X, y: Y + 1000, z: 0}});
new Text(slide.matricule, {properties: {content: 'slide : ' + I + 2}});
var slide = new Slide({pos: {x: X + 1000, y: Y, z: 0}});
new Text(slide.matricule, {properties: {content: 'slide : ' + I + 4}});
var slide = new Slide({pos: {x: X + 1000, y: Y + 1000, z: 0}});
new Text(slide.matricule, {properties: {content: 'slide : ' + I + 6}});
var slide = new Slide({pos: {x: X + 1000, y: Y + 2000, z: 0}});
new Text(slide.matricule, {properties: {content: 'slide : ' + I + 8}});
var slide = new Slide({pos: {x: X + 1000, y: Y + 3000, z: 0}});
new Text(slide.matricule, {properties: {content: 'slide : ' + I + 10}});
// new Image({pos: {y: 500}}, 'slide0');
//s1.show();
//s1.pos.x = 100;
// container.slide['slide0'].show();
//// //watch des modifications
//// watch(container.slide['slide0'].pos, 'x', function(attr, action, newVal, oldVal) {
//// //console.log('màj', this.matricule, ' propriété ', attr, ' avec ', newVal);
//// });
//
//
// container.slide['slide1'].show();
}
;<file_sep>/js/gui_keyboard.js
/*
* gestion de la partie interface utilisateur de l'editeur (navigable, creation) :
* listerner keyboard :
* creation
* navigable
* class Transform3D
*
*/
$(document).on('keypress',function(event){
//si la souris est sur le body, et que le body, call keyboarGUI
if( $('.slide:hover').length === 0 &&
$('.sidebar:hover').length === 0 &&
$('#topbar:hover').length === 0 &&
$('.buttonclicked').length === 0 ){
keyboardGUI(event);
}
});
function keyboardGUI(event) {
var objEvt = new ObjectEvent({
matricule: '',
action: '',
event: {
cran: 300 //puour le navigable
}
});
switch (event.which) {
//creation composant
case 106 :
objEvt.action = 'createSlide';
break;
case 107 :
objEvt.action = 'createH1Text';
break;
case 108 :
objEvt.action = 'createH2Text';
break;
case 109 :
objEvt.action = 'createH3Text';
break;
case 249 :
objEvt.action = 'createBodyText';
break;
case 42 :
objEvt.action = 'createImage';
break;
}
switch (event.which) {
//gestion navigation
//deplacement
case 97:
objEvt.action = 'navigable';
objEvt.event.direction = 'z+';
break;
case 122:
objEvt.action = 'navigable';
objEvt.event.direction = 'y-';
break;
case 113:
objEvt.action = 'navigable';
objEvt.event.direction = 'x-';
break;
case 115:
objEvt.action = 'navigable';
objEvt.event.direction = 'x+';
break;
case 119:
objEvt.action = 'navigable';
objEvt.event.direction = 'y+';
break;
case 120:
objEvt.action = 'navigable';
objEvt.event.direction = 'z-';
break;
//rotation
objEvt.event.cran = 10;
case 114:
objEvt.action = 'rotate';
objEvt.event.direction = 'z+';
break;
case 102:
objEvt.action = 'rotate';
objEvt.event.direction = 'y-';
break;
case 118:
objEvt.action = 'rotate';
objEvt.event.direction = 'x-';
break;
case 116:
objEvt.action = 'rotate';
objEvt.event.direction = 'x+';
break;
case 103:
objEvt.action = 'rotate';
objEvt.event.direction = 'y+';
break;
case 98:
objEvt.action = 'rotate';
objEvt.event.direction = 'z-';
break;
}
switch (objEvt.action) {
case 'navigable':
objEvt.event.cran = 100;
break;
case 'rotate' :
objEvt.event.cran = 10;
break;
}
callModelGUI(objEvt);
}
<file_sep>/js/auto_dynamic.js
/*
* Manage la présentation en live, lié à 'automaticEditor'
*
* Affiche/masque les slides
* Deplace les slides de titres,
* Gere le début et la fin
*
*
*
*/
/*
* A partir de la présentation init par Jmpress, réarrange la position des slides afin d'obtenir un état initial
*
*/
function initDynamic() {
//on masque les li content du tree
// $('#tree li').each(function(){
// if( $(this).attr('type') === 'content') $(this).css('display','none');
// });
//on affiche le tree
$('#tree').fadeIn(1000);
//au départ, on cache toutes les slides sauf les premiers titres
//les premiers titres
var firstTitle = [];
$('#tree').children('ol').children().each(function() {
firstTitle.push($(this).attr('matricule'));
});
$('.slide').each(function() {
if (firstTitle.indexOf($(this).attr('matricule')) !== -1)
$(this).fadeIn(1000);
else
$(this).fadeOut(2000);
});
// //on deplace les slides à leur véritable positions (utile lorsqu'on loop)
$('.slide').each(function() {
if ($(this).attr('matricule') === 'questions')
return; //deplacement étrange de la slide de questions
$(this).translate3d({
x: parseInt($(this).attr('data-x')),
y: parseInt($(this).attr('data-y')),
z: parseInt($(this).attr('data-z'))
}, 1000);
});
//au depart tout le plan est en 'futur'
$('#tree li').each(function() {
$(this).addClass('li-slide');
$(this).addClass('future-slide');
});
//au depart tous les plus grands titres sont au niveau de leur grande soeur
$('#tree li').each(function() {
if ($(this).attr('type') === 'title') {
if ($('#' + $(this).attr('matricule')).attr('matricule') === 'questions' || $('#' + $(this).attr('matricule')).attr('matricule') === 'end')
return; //on va chercher l'info dans les slides car la li n'a pas trace de ces slides là (qui ne font pas parties du plan)
var $slideRef = $('#' + $(this).attr('matricule'));
if( typeof $slideRef === 'undefined') return;
var dicoRef = getTrans3D($slideRef);
$(this).siblings().each(function() {
if (typeof $(this).attr('matricule') === 'undefined')
return;
var $slide = $('#' + $(this).attr('matricule'));
var dico = getTrans3D($slide);
dico.translate3d[2] = dicoRef.translate3d[2];
setTrans3D(dico, $slide); //pas besoin de deplacement smooth ici alors on change directement le transform3D des slides
});
}
});
}
/*
* A partir de la présentation init par Jmpress, réarrange la position des slides afin d'obtenir un état final
*/
function endDynamic() {
//on cache le tree
$('#tree').fadeOut(1000);
//on montre la dernière slide (qui ne fait pas parti de la présentation donc est exclu du circuit normal
$('#questions').fadeIn(6000);
//container.getSlide($('#questions').attr('matricule')).pos.x = container.getSlide($('#questions').attr('matricule')).pos.x + 1;
/* parcours de la liste pour deplacer les slides jusqu'à la position de fin */
$('#tree li').each(function() {
if (typeof $(this).attr('data-end-x') !== 'undefined') { //si la slide doit être déplacée
var $slide = $('#' + $(this).attr('matricule'));
$slide.fadeIn(10);
$slide.translate3d({
x: parseInt($(this).attr('data-end-x')), //-700,
y: parseInt($(this).attr('data-end-y')), //-600 ,
z: parseInt($(this).attr('data-end-z'))
}, 5000);
} else { //si la slide ne doit pas etre déplacée, on la cache
$('#' + $(this).attr('matricule')).fadeOut(1000);
}
});
}
/*
* Ajoute un listener sur la touche 'space' pour gérer la présentation en dynamique
* Affichage/cache, deplace, smooth !, loop
*
*/
function dynamic() {
initDynamic();
console.log('retour init dynamic');
$(document).on('keypress', function(event) {
if (event.which == 32) { //je n'ai pu récupérer que l'espace
/*
* each time space bar is pressed on cherche la slide qui a la classe 'active'
*/
var currentMatricule = $('#slideArea').attr('class').split(' ')[0].replace('step-', '');
var currentSlide = $('#' + currentMatricule);
var liCurrent = $('#tree #li_' + currentMatricule);
/*
* Réinit de la présentation lorsqu'on arrive à la premiere slide
*/
if (currentMatricule === 'home') {
initDynamic();
}
/*
* Mise en place finale lorsqu'on arrive à la fin
*/
if (currentMatricule === 'end') {
endDynamic();
}
/*
* remerciement final
*/
if (currentMatricule === 'questions') {
var slide = container.getSlide(currentMatricule);
var texte = slide.element['questionstexte'];
texte.properties.content = "Thanks for watching";
}
//petit effet sur le tree
$('.present-slide').removeClass('present-slide').addClass('past-slide');
liCurrent.removeClass('future-slide').addClass('present-slide');
//on cache les filles et leurs filles (et ainsi de suite) de ses soeurs
//pour effectuer le fadeout plus tôt, sur l'arrivée à un overview s'il y en a un
if (currentSlide.hasClass('overview'))
var aliCurrent = $('#tree #li_' + currentSlide.next().attr('matricule')); //on recupère le matricule de la slide suivant le matricule et on recupere la li correspondante
else
var aliCurrent = liCurrent;
aliCurrent.siblings().each(function() { //toutes les soeurs
var allChildren = getChildren($(this), []); // on recupere un tableau des matricules de toutes les filles
for (var child in allChildren) {
var matriculeChild = allChildren[child];
$('#' + matriculeChild).fadeOut(1600);
}
});
/* affiche certaines slides */
//ses soeurs
liCurrent.siblings().each(function() {
var matricule = $(this).attr('matricule'); //on recuperer le matricule stocké dans la li
$('#' + matricule).fadeIn(1600); //on agit sur la slide qui a ce matricule
});
//ses filles directes
$(liCurrent.children('ol')[0]).children().each(function() {
var matricule = $(this).attr('matricule');
$('#' + matricule).fadeIn(1600);
});
//elle meme
$('#' + currentMatricule).fadeIn(1600);
/* mise à niveau des petites soeurs sur la petite soeur de la current*/
var littleHilly = $('#' + liCurrent.next().attr('matricule'));
//console.log('dynamicSlideShow : call getTrans3D for dicoRef');
var dicoRef = getTrans3D(littleHilly);
$('#tree #li_' + currentMatricule + ' ~').each(function() { //pour obtenir les next siblings de la current li
var $slide = $('#' + $(this).attr('matricule'));
//console.log('dynamicSlideShow : call getTrans3D for dico');
var dico = getTrans3D($slide);
// //console.log(littleHilly,dicoRef.translate3d,$slide,dico.translate3d);
// dico.translate3d[2] = littleHilly.attr('data-z');
$slide.translate3d({
x: parseInt($slide.attr('data-x')), //-700,
y: parseInt($slide.attr('data-y')), //-600 ,
z: littleHilly.attr('data-z')
}, 1000);
// setTrans3D(dico,$slide);
});
}
});
}
<file_sep>/js/gui_controler.js
/*
* Recupère objEvt et, en fonction de ses paramètres, effectue le traitement adéquat directement sur les instances des
* classes composants (Slide, Texte et Image héritées de Element
*
* Contient :
* findObjectOfComposant(matricule) return object
* callModel(objectEvent)
* createComposant($target, objectEvent) appelé une fois la Slide cible sélectionnée
* callModelGUI(objectEvent)
* test
*
*/
/* Controler de slideshowEditor
* permet de diriger les interactions avec les composants
* deplacement, rotation, édition (texte, image)
*
*/
function callModel(objectEvent) {
// //console.log(objectEvent);
if (objectEvent.matricule === '' || objectEvent.matricule === 'document') {
//creation de composant
//console.log('warning : bad function call (callModel instead of callModelGui), redirectinnf proceded');
callModelGUI(objectEvent);
} else {
//modification de composant
var composant = findObjectOfComposant(objectEvent.matricule);
// //console.log('avant', composant.show());
if (objectEvent.action === 'move') {
var attr;
var val = objectEvent.event.cran;
switch (objectEvent.event.direction) {
case 'z+':
attr = 'z';
val = val;
break;
case 'z-':
attr = 'z';
val = -val;
break;
case 'x+':
attr = 'x';
val = val;
break;
case 'x-':
attr = 'x';
val = -val;
break;
case 'y+':
attr = 'y';
val = val;
break;
case 'y-':
attr = 'y';
val = -val;
break;
}
composant.pos[attr] += val;
// //console.log('après', composant.show());
} else if (objectEvent.action === 'rotate') {
var attr;
var val = objectEvent.event.cran;
switch (objectEvent.event.direction) {
case 'z+':
attr = 'z';
val = val;
break;
case 'z-':
attr = 'z';
val = -val;
break;
case 'x+':
attr = 'x';
val = val;
break;
case 'x-':
attr = 'x';
val = -val;
break;
case 'y+':
attr = 'y';
val = val;
break;
case 'y-':
attr = 'y';
val = -val;
break;
}
composant.rotate[attr] += val;
// //console.log('après', composant.show());
}
}
}
/* Class gerant la navigation en agissant sur l'attribut CSS transform
*
*/
//transform3D = new Transform3D();
/*
* probleme d'asynchronisme lors de la selection de la slide cible avec le clavier
* du coup, petite magouille, createComposant est appellée une fois la slide selectionnée
*
*/
function createComposant($target, objectEvent) {
if (objectEvent.action === 'createH1Text') {
new Text($target, {properties: {hierarchy: 'H1Text'}});
// //console.log('new text H1');
} else if (objectEvent.action === 'createH2Text') {
new Text($target, {properties: {hierarchy: 'H2Text'}});
// //console.log('new text');
} else if (objectEvent.action === 'createH3Text') {
new Text($target, {properties: {hierarchy: 'H2Text'}});
// //console.log('new text');
} else if (objectEvent.action === 'createBodyText') {
new Text($target, {});
// //console.log('new text');
} else if (objectEvent.action === 'createImage') {
new Image($target, {source: objectEvent.source});
// //console.log('new ');
}
}
/* Controler de gestion de l'interface
* navigable, bouton creation
*
*/
function callModelGUI(objectEvent) {
//en attente de trouver une meilleure méthode pour récupérer la slide destination d'un element
if (objectEvent.action === 'createImage' || objectEvent.action === 'createH3Text' || objectEvent.action === 'createH2Text' || objectEvent.action === 'createH1Text' || objectEvent.action === 'createBodyText') {
if (objectEvent.action === 'createImage') { //infos suplémentaires propre aux images
var source = prompt('Sélectionner l\'adresse de votre image (adresse fichier, ou adresse url', 'images/bleu_twitter.png');
objectEvent.source = source;
}
//s'il y a un matricule
if (objectEvent.matricule !== '') {
//si ce matricule existe
var slide = findObjectOfComposant(objectEvent.matricule);
if (typeof slide !== 'undefined') { // pas de !== ivi
//si le composant est une slide
if (slide.type === 'slide') {
//alors on peut ajouter directement l'element
createComposant(objectEvent.matricule, objectEvent);
return;
}
}
}
var $target = selectSlide(createComposant, objectEvent);
//console.log('after selectc', $target);
return;
}
// //console.log(objectEvent);
if (objectEvent.action === 'createSlide') {
new Slide();
//console.log('new slide');
}
// else if (objectEvent.action === 'createH1Text') { //voir fonction createComposant
// new Text({properties: {hierarchy: 'H1Text'}}, $target);
// //console.log('new text H1');
// } else if (objectEvent.action === 'createH2Text') {
// new Text({properties: {hierarchy: 'H2Text'}}, $target);
// //console.log('new text');
// } else if (objectEvent.action === 'createH3Text') {
// new Text({properties: {hierarchy: 'H2Text'}}, $target);
// //console.log('new text');
// } else if (objectEvent.action === 'createBodyText') {
// new Text({}, $target);
// //console.log('new text');
// }
else if (objectEvent.action === 'move' || objectEvent.action === 'navigable') {
var attr;
var val = objectEvent.event.cran * 10;
switch (objectEvent.event.direction) {
case 'z+':
attr = 'z';
val = val;
break;
case 'z-':
attr = 'z';
val = -val;
break;
case 'x+':
attr = 'x';
val = val;
break;
case 'x-':
attr = 'x';
val = -val;
break;
case 'y+':
attr = 'y';
val = val;
break;
case 'y-':
attr = 'y';
val = -val;
break;
}
transform3D.pos[attr] = val;
//console.log('navigable');
} else if (objectEvent.action === 'rotate') {
var attr;
var val = objectEvent.event.cran;
switch (objectEvent.event.direction) {
case 'z+':
attr = 'z';
val = val;
break;
case 'z-':
attr = 'z';
val = -val;
break;
case 'x+':
attr = 'x';
val = val;
break;
case 'x-':
attr = 'x';
val = -val;
break;
case 'y+':
attr = 'y';
val = val;
break;
case 'y-':
attr = 'y';
val = -val;
break;
}
transform3D.rotate[attr] = val;
//console.log('rotate');
}
}
/* selection d'une slide par click,
*
* @returns {string}
*/
function selectSlide(callback, param1, composant) {
$('#sidebar').parent().addClass("buttonclicked"); //pour empecher le joystick d'apparaitre
alert('Il faut selectionner une slide');
$('.slide').one('click', function(event) {
$('.buttonclicked').removeClass("buttonclicked"); //pour redonner le droit au joystick d'apparaitre
var slide = $(this).attr('matricule');
alert('slide selectionnée' + slide);
if (typeof callback !== 'undefined') {
callback(slide, param1, composant);
////console.log('in select', slide);
return slide;
} else {
////console.log('in select', slide);
return slide;
}
});
}
/* classe objetEvent
* matricule
* action
* event
* des infos
*
* Est ce que je stocke les objetEvent ?
*
*
*/
ObjectEvent = Class.extend({
init: function(params) {
//default value
//matricule du composant
//sinon c'est de la création
this.matricule = '';
/* ce qu'il faut faire sur le composant
* +move
*
*/
this.action = '';
/* caracterique de l'event
* type : keyboard, souris
* si move -> direction : z+, z-, y+...
* si rotate -> sens : x, -x, y, -y
* si create -> pos : {}, rotate{}
*/
this.event = {
};
if (typeof params !== 'undefined') {
for (var param in params) {
if (typeof params[param] === 'object') {
for (var paramNested in param) {
this[param][paramNested] = param[paramNested];
}
}
this[param] = params[param];
}
}
},
show: function(i) {
if (typeof i === 'undefined') {
//console.log('objectEvent');
} else {
//console.log('objectEvent');
}
},
destroy: function() {
//console.log('nothing to delete');
}
}); | 4b1699fd7f9ce6db5c3f70e74c0e0f045439c8c1 | [
"JavaScript",
"Markdown",
"INI"
] | 10 | JavaScript | remipichon/jmpresseditor2.js | 73d9374d8e849d1f9d5488fe36a526c48da1830f | 0b3c48e0978ea0d702df7a04c476d999c29780b2 |
refs/heads/master | <file_sep>import os
import pickle
import numpy as np
def unpickle(file):
with open(file, 'rb') as f:
dict = pickle.load(f, encoding='bytes')
return dict
def create_master_dataset(directory):
trainFormat = os.path.join(directory, 'data_batch_{}')
testFile = os.path.join(directory, 'test_batch')
trainDicts = []
for i in range(1, 6):
data = unpickle(trainFormat.format(i))
trainDicts.append(data)
trainSize = sum(len(l[b'labels']) for l in trainDicts)
trainLabels = np.zeros((trainSize,))
trainData = np.zeros((trainSize,3072))
nextEmpty = 0
for d in trainDicts:
n = len(trainDicts[0][b'labels'])
trainLabels[nextEmpty:(nextEmpty+n)] = d[b'labels']
trainData[nextEmpty:(nextEmpty+n),] = d[b'data']
nextEmpty += n
testDict = unpickle(testFile)
testLabels = np.array(testDict[b'labels'])
testData = np.array(testDict[b'data'])
return trainData, trainLabels, testData, testLabels
outputs = create_master_dataset('../../data/raw/cifar-10-batches-py')
<file_sep># CIFAR-10 Image Classifier
This project is trained on the CIFAR-10 dataset. The dataset tries to classifiy 10 different types of objects.
The url for the dataset is found here: https://www.cs.toronto.edu/~kriz/cifar.html
<file_sep>from loaddata import create_master_dataset
import numpy as np
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras import layers, optimizers
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.preprocessing.image import ImageDataGenerator
dataDir = '../../data/raw/cifar-10-batches-py'
trainXTemp, trainYTemp, testXTemp, testYTemp = create_master_dataset(dataDir)
trainSize = int(trainXTemp.shape[0] * 0.8)
valSize = trainXTemp.shape[0] - trainSize
testSize = testXTemp.shape[0]
allTrainSet = np.moveaxis(trainXTemp.reshape([trainSize+valSize, 3, 32, 32]), 1, -1)
trainX = allTrainSet[:trainSize] / 255
trainY = to_categorical(trainYTemp[:trainSize])
valX = allTrainSet[trainSize:] / 255
valY = to_categorical(trainYTemp[trainSize:])
testX = np.moveaxis(testXTemp.reshape([testSize, 3, 32, 32]) / 255, 1, -1)
testY = to_categorical(testYTemp)
# data augmentation
imageGenerator = ImageDataGenerator(
horizontal_flip=True)
trainAug = imageGenerator.flow(trainX, trainY)
modelNum = 9
if modelNum == 1:
model = Sequential()
model.add(layers.Input(shape=(32, 32, 3)))
model.add(layers.Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(layers.Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu', padding='same'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(32, activation='relu'))
model.add(layers.Dense(10, activation='softmax'))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=40, validation_data=(valX, valY), verbose=1)
elif modelNum == 2:
# BatchNormalization after activation
model = Sequential()
model.add(layers.Input(shape=(32, 32, 3)))
model.add(layers.Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(32, activation='relu'))
model.add(layers.BatchNormalization())
model.add(layers.Dense(10, activation='softmax'))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=30, validation_data=(valX, valY), verbose=1)
elif modelNum == 3:
# BatchNormalization before activation
model = Sequential()
model.add(layers.Input(shape=(32, 32, 3)))
model.add(layers.Conv2D(32, (3, 3), padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.Conv2D(32, (3, 3), padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.Conv2D(64, (3, 3), padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(32))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.Dense(10))
model.add(layers.BatchNormalization())
model.add(layers.Activation('softmax'))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=30, validation_data=(valX, valY), verbose=1)
elif modelNum == 4:
# After the activation performed better for batch normalization
# Add dropout after all layers expect input
model = Sequential()
model.add(layers.Input(shape=(32, 32, 3)))
model.add(layers.Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (3, 3), activation='relu', padding='same'))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dropout(0.3))
model.add(layers.Dense(32, activation='relu'))
model.add(layers.BatchNormalization())
model.add(layers.Dropout(0.3))
model.add(layers.Dense(10, activation='softmax'))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=30, validation_data=(valX, valY), verbose=1)
elif modelNum == 5:
# Tries different architecture using inception modules
# Let's not use dropout for now
input_ = layers.Input(shape=(32, 32, 3))
conv0 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(input_)
batch0 = layers.BatchNormalization()(conv0)
dropout1 = layers.Dropout(0.3)(batch0)
conv1 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout1)
batch1 = layers.BatchNormalization()(conv1)
pool1 = layers.MaxPooling2D((2, 2))(batch1)
dropout2 = layers.Dropout(0.3)(pool1)
conv2a = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout2)
conv2b = layers.Conv2D(16, (3, 3), activation='relu', padding='same')(dropout2)
conv2c = layers.Conv2D(16, (5, 5), activation='relu', padding='same')(dropout2)
conv2 = layers.Concatenate()([conv2a, conv2b, conv2c])
batch2 = layers.BatchNormalization()(conv2)
pool2 = layers.MaxPooling2D((2, 2))(batch2)
dropout3 = layers.Dropout(0.3)(pool2)
conv3a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout3)
conv3b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout3)
conv3c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(dropout3)
conv3 = layers.Concatenate()([conv3a, conv3b, conv3c])
batch3 = layers.BatchNormalization()(conv3)
dropout4 = layers.Dropout(0.3)(batch3)
conv4a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout4)
conv4b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout4)
conv4c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(dropout4)
conv4 = layers.Concatenate()([conv4a, conv4b, conv4c])
batch4 = layers.BatchNormalization()(conv4)
pool4 = layers.MaxPooling2D((2, 2))(batch4)
flat4 = layers.Flatten()(pool4)
dropout5 = layers.Dropout(0.3)(flat4)
dense5 = layers.Dense(32, activation='relu')(dropout5)
batch5 = layers.BatchNormalization()(dense5)
output = layers.Dense(10, activation='softmax')(batch5)
model = Model(inputs=[input_], outputs=[output])
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=30, validation_data=(valX, valY), verbose=1)
elif modelNum == 6:
# try reducing parameters by adding 1x1 convolutions in layer
# 4 inception module
input_ = layers.Input(shape=(32, 32, 3))
conv0 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(input_)
batch0 = layers.BatchNormalization()(conv0)
dropout1 = layers.Dropout(0.3)(batch0)
conv1 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout1)
batch1 = layers.BatchNormalization()(conv1)
pool1 = layers.MaxPooling2D((2, 2))(batch1)
dropout2 = layers.Dropout(0.3)(pool1)
conv2a = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout2)
conv2b = layers.Conv2D(16, (3, 3), activation='relu', padding='same')(dropout2)
conv2c = layers.Conv2D(16, (5, 5), activation='relu', padding='same')(dropout2)
conv2 = layers.Concatenate()([conv2a, conv2b, conv2c])
batch2 = layers.BatchNormalization()(conv2)
pool2 = layers.MaxPooling2D((2, 2))(batch2)
dropout3 = layers.Dropout(0.3)(pool2)
conv3a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout3)
conv3b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout3)
conv3c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(dropout3)
conv3 = layers.Concatenate()([conv3a, conv3b, conv3c])
batch3 = layers.BatchNormalization()(conv3)
dropout4 = layers.Dropout(0.3)(batch3)
conv4a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout4)
conv4b1 = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout4)
conv4b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(conv4b1)
conv4c1 = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout4)
conv4c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(conv4c1)
conv4 = layers.Concatenate()([conv4a, conv4b, conv4c])
batch4 = layers.BatchNormalization()(conv4)
pool4 = layers.MaxPooling2D((2, 2))(batch4)
flat4 = layers.Flatten()(pool4)
dropout5 = layers.Dropout(0.3)(flat4)
dense5 = layers.Dense(32, activation='relu')(dropout5)
batch5 = layers.BatchNormalization()(dense5)
output = layers.Dense(10, activation='softmax')(batch5)
model = Model(inputs=[input_], outputs=[output])
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=45, validation_data=(valX, valY), verbose=1)
model.evaluate(testX, testY) # 0.8317
elif modelNum == 7:
# removing layer 4 of previous
input_ = layers.Input(shape=(32, 32, 3))
conv0 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(input_)
batch0 = layers.BatchNormalization()(conv0)
dropout1 = layers.Dropout(0.3)(batch0)
conv1 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout1)
batch1 = layers.BatchNormalization()(conv1)
pool1 = layers.MaxPooling2D((2, 2))(batch1)
dropout2 = layers.Dropout(0.3)(pool1)
conv2a = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout2)
conv2b = layers.Conv2D(16, (3, 3), activation='relu', padding='same')(dropout2)
conv2c = layers.Conv2D(16, (5, 5), activation='relu', padding='same')(dropout2)
conv2 = layers.Concatenate()([conv2a, conv2b, conv2c])
batch2 = layers.BatchNormalization()(conv2)
pool2 = layers.MaxPooling2D((2, 2))(batch2)
dropout3 = layers.Dropout(0.3)(pool2)
conv3a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout3)
conv3b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout3)
conv3c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(dropout3)
conv3 = layers.Concatenate()([conv3a, conv3b, conv3c])
batch3 = layers.BatchNormalization()(conv3)
pool4 = layers.MaxPooling2D((2, 2))(batch3)
flat4 = layers.Flatten()(pool4)
dropout5 = layers.Dropout(0.3)(flat4)
dense5 = layers.Dense(32, activation='relu')(dropout5)
batch5 = layers.BatchNormalization()(dense5)
output = layers.Dense(10, activation='softmax')(batch5)
model = Model(inputs=[input_], outputs=[output])
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=45, validation_data=(valX, valY), verbose=1)
elif modelNum == 8:
# Reduce parameters further by changing padding in first two layers
input_ = layers.Input(shape=(32, 32, 3))
conv0 = layers.Conv2D(32, (3, 3), activation='relu', padding='valid')(input_)
batch0 = layers.BatchNormalization()(conv0)
dropout1 = layers.Dropout(0.3)(batch0)
conv1 = layers.Conv2D(32, (3, 3), activation='relu', padding='valid')(dropout1)
batch1 = layers.BatchNormalization()(conv1)
pool1 = layers.MaxPooling2D((2, 2))(batch1)
dropout2 = layers.Dropout(0.3)(pool1)
conv2a = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout2)
conv2b = layers.Conv2D(16, (3, 3), activation='relu', padding='same')(dropout2)
conv2c = layers.Conv2D(16, (5, 5), activation='relu', padding='same')(dropout2)
conv2 = layers.Concatenate()([conv2a, conv2b, conv2c])
batch2 = layers.BatchNormalization()(conv2)
pool2 = layers.MaxPooling2D((2, 2))(batch2)
dropout3 = layers.Dropout(0.3)(pool2)
conv3a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout3)
conv3b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout3)
conv3c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(dropout3)
conv3 = layers.Concatenate()([conv3a, conv3b, conv3c])
batch3 = layers.BatchNormalization()(conv3)
dropout4 = layers.Dropout(0.3)(batch3)
conv4a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout4)
conv4b1 = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout4)
conv4b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(conv4b1)
conv4c1 = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout4)
conv4c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(conv4c1)
conv4 = layers.Concatenate()([conv4a, conv4b, conv4c])
batch4 = layers.BatchNormalization()(conv4)
pool4 = layers.MaxPooling2D((2, 2))(batch4)
flat4 = layers.Flatten()(pool4)
dropout5 = layers.Dropout(0.3)(flat4)
dense5 = layers.Dense(32, activation='relu')(dropout5)
batch5 = layers.BatchNormalization()(dense5)
output = layers.Dense(10, activation='softmax')(batch5)
model = Model(inputs=[input_], outputs=[output])
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainX, trainY, epochs=10, validation_data=(valX, valY), verbose=1)
elif modelNum == 9:
# Add data augmentation
input_ = layers.Input(shape=(32, 32, 3))
conv0 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(input_)
batch0 = layers.BatchNormalization()(conv0)
dropout1 = layers.Dropout(0.3)(batch0)
conv1 = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout1)
batch1 = layers.BatchNormalization()(conv1)
pool1 = layers.MaxPooling2D((2, 2))(batch1)
dropout2 = layers.Dropout(0.3)(pool1)
conv2a = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout2)
conv2b = layers.Conv2D(16, (3, 3), activation='relu', padding='same')(dropout2)
conv2c = layers.Conv2D(16, (5, 5), activation='relu', padding='same')(dropout2)
conv2 = layers.Concatenate()([conv2a, conv2b, conv2c])
batch2 = layers.BatchNormalization()(conv2)
pool2 = layers.MaxPooling2D((2, 2))(batch2)
dropout3 = layers.Dropout(0.3)(pool2)
conv3a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout3)
conv3b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(dropout3)
conv3c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(dropout3)
conv3 = layers.Concatenate()([conv3a, conv3b, conv3c])
batch3 = layers.BatchNormalization()(conv3)
dropout4 = layers.Dropout(0.3)(batch3)
conv4a = layers.Conv2D(64, (1, 1), activation='relu', padding='same')(dropout4)
conv4b1 = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout4)
conv4b = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(conv4b1)
conv4c1 = layers.Conv2D(32, (1, 1), activation='relu', padding='same')(dropout4)
conv4c = layers.Conv2D(32, (5, 5), activation='relu', padding='same')(conv4c1)
conv4 = layers.Concatenate()([conv4a, conv4b, conv4c])
batch4 = layers.BatchNormalization()(conv4)
pool4 = layers.MaxPooling2D((2, 2))(batch4)
flat4 = layers.Flatten()(pool4)
dropout5 = layers.Dropout(0.3)(flat4)
dense5 = layers.Dense(32, activation='relu')(dropout5)
batch5 = layers.BatchNormalization()(dense5)
output = layers.Dense(10, activation='softmax')(batch5)
model = Model(inputs=[input_], outputs=[output])
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['categorical_accuracy'])
model.fit(trainAug, epochs=45, validation_data=(valX, valY), verbose=1)<file_sep>wget -c https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz -O - | tar -xz
mv cifar-10-batches-py ../../data/raw/
<file_sep>mkdir -p data
mkdir -p data/raw
(cd src/data/ ; sh download-data.sh)
| 8c6779f2ecceeb1704e5389ece6497b379d6412e | [
"Markdown",
"Python",
"Shell"
] | 5 | Python | Brian-Paulsen/CIFAR-10-Classification | f9431d5a97975cea45d589fa105658d4078bea22 | ef8df0393171b4ac688e5a822db67eb5fec9980a |
refs/heads/master | <repo_name>theloneplant/GameRev<file_sep>/routes/index.js
var path = require('path');
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
module.exports = function(req, res) {
var createReviewHref = 'href="/login"';
console.log(Parse.User.current());
if (Parse.User.current()) {
createReviewHref = '';
};
var Review = Parse.Object.extend('Reviews');
var query = new Parse.Query(Review);
query.addDescending('views');
query.limit(12);
query.find({
success: function(reviews) {
var gameQuery = [];
var data = {
featuredSlider: [
{
active: "active",
id: "featured-1",
href: "/games/dragon-age-inquisition",
src: "/images/dai-cover.jpg"
},
{
id: "featured-2",
href: "/games/infamous",
src: "/images/infamous_banner.jpg"
},
{
id: "featured-3",
href: "/games/skyrim",
src: "/images/skyrim_banner.jpg"
},
{
id: "featured-4",
href: "/games/final-fantasy-XV",
src: "/images/finalfantasyxv_banner.jpg"
},
{
id: "featured-5",
href: "/games/revelations",
src: "/images/revelations_banner.png"
}
],
createReviewHref: createReviewHref,
reviews: []
};
reviews.forEach(function(review, i) {
gameQuery.push(review.relation('game').query().find());
data.reviews.push({
id: review.id,
title: review.get('title'),
rating: review.get('rating'),
summary: createSummary(review.get('reviewBody'), 170),
tldr: review.get('tldr'),
bannerImage: review.get('bannerImage'),
game: {
title: '',
ref: ''
}
});
});
Parse.Promise.when(gameQuery).then(function(poop) {
for (var i = 0; i < arguments.length; i++) {
// Format a summary for each review
data.reviews[i].game.title = arguments[i][0].get('title');
data.reviews[i].game.ref = arguments[i][0].get('ref');
}
require(path.join(__dirname, 'header')).renderPage(req, res, 'index', data);
});
},
error: function(review, error) {
console.log(error);
}
});
};
var createSummary = function(review, length) {
var summary = review.replace(/<(?:.|\n)*?>/gm, '');
if (summary.length > length)
{
summary = summary.substring(0, length);
summary = summary.split(' ');
summary.pop();
summary = summary.join(' ');
summary += '...';
}
return summary;
};<file_sep>/routes/api/database.js
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
this.addReview = function(req, res) {
var user = Parse.User.current();
var Review = Parse.Object.extend('Reviews');
var review = new Review();
review.set('title', req.body.newReview['title']);
review.set('rating', req.body.newReview['rating']);
review.set('reviewBody', req.body.newReview['reviewBody']);
review.set('theGood', req.body.newReview['theGood']);
review.set('theOkay', req.body.newReview['theOkay']);
review.set('theBad', req.body.newReview['theBad']);
review.set('likes', parseInt(req.body.newReview['likes']));
review.set('bannerImage', req.body.newReview['bannerImage']);
var Games = Parse.Object.extend('Games');
var gameQuery = new Parse.Query(Games);
console.log(req.body.game);
// Find the game that this review will have a relation with
gameQuery.equalTo('title', req.body.game);
gameQuery.first({
success: function(game) {
console.log(game);
var userReviewRelation = user.relation('reviews');
var gameReviewRelation = game.relation('reviews');
// Add the relation from the review to the user
var reviewUserRelation = review.relation('user');
reviewUserRelation.add(user);
// Add the relation from the review to the game
var reviewGameRelation = review.relation('game');
reviewGameRelation.add(game);
if (req.body.newReview.rating === 'Good') {
game.increment('totalRating', 1);
}
else if (req.body.newReview.rating === 'Okay') {
game.increment('totalRating', 0.5);
}
// Otherwise it's a bad review, don't increment
game.relation('reviews').query().find({
success: function(reviews) {
var totalRating = (100 * game.get('totalRating') / reviews.length).toFixedDown(1);
if (totalRating < 34) {
game.set('verdict', 'Bad');
}
else if (totalRating < 67) {
game.set('verdict', 'Okay');
}
else {
game.set('verdict', 'Good');
}
console.log(game.get('totalRating'), ' - ', reviews.length);
// Save the review and add relations going back from user and game to review
review.save(null, {
success: function(result) {
console.log(result.id);
// Add relation from user to review
userReviewRelation.add(result);
user.save();
// Add relation from game to review
gameReviewRelation.add(result);
game.save();
},
error: function(result, error) {
console.log("review error: ", error.message);
}
});
}
});
},
error: function(error) {
console.log("game error: ", error.message);
}
});
basicResponse(res, true);
};
var basicResponse = function(res, $success, $data) {
res.send({
success : $success,
data : $data
});
};
Number.prototype.toFixedDown = function(digits) {
var re = new RegExp("(\\d+\\.\\d{" + digits + "})(\\d)"),
m = this.toString().match(re);
return m ? parseFloat(m[1]) : this.valueOf();
};<file_sep>/routes/reviews.js
var path = require('path');
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
module.exports = function(req, res) {
var Review = Parse.Object.extend('Reviews');
var query = new Parse.Query(Review);
query.get(req.params.review, {
success: function(review) {
review.relation('user').query().find({
success: function(user) {
review.relation('game').query().find({
success: function(game) {
var goodArr = review.get('theGood').split('\t');
var okayArr = review.get('theOkay').split('\t');
var badArr = review.get('theBad').split('\t');
var data = {
id: review.id,
title: review.get('title'),
rating: review.get('rating'),
reviewBody: review.get('reviewBody'),
theGood: [],
theOkay: [],
theBad: [],
bannerImage: review.get('bannerImage'),
user: {
username: user.username
// Add more to user later on
},
game: {
title: game.title,
ref: game.ref
// Add more to game later on
},
timestamp: review.createdAt
}
goodArr.forEach(function(e) {
data.theGood.push({
value: e
});
});
data.theGood.pop(); // Remove trailing tab
okayArr.forEach(function(e) {
data.theOkay.push({
value: e
});
});
data.theOkay.pop();
badArr.forEach(function(e) {
data.theBad.push({
value: e
});
});
data.theBad.pop();
console.log(data.theGood);
require(path.join(__dirname, 'header')).renderPage(req, res, 'reviews', data);
}
});
}
});
},
error: function(review, error) {
console.log(error);
}
});
};<file_sep>/routes/api/users.js
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
this.processSignup = function(req, res) {
var user = new Parse.User();
user.set("username", req.body.username);
user.set("password", <PASSWORD>);
user.set("email", req.body.email);
user.signUp(null, {
success: function(user) {
// Hooray! Let them use the app now.
basicResponse(res, true);
},
error: function(user, error) {
basicResponse(res, false, { error: error.message });
}
});
};
this.processLogin = function(req, res) {
Parse.User.logIn(req.body.username, req.body.password, {
success: function(user) {
// Hooray! Let them use the app now.
//var banana = user.relation('reviews');
//banana.add();
/*var derp = Parse.Object.extend('_User');
var herp = new Parse.Query(derp);
herp.equalTo('objectId', user.id);
herp.find({
success: function(result) {
console.log("result: ", result);
var shlerp = result[0];
shlerp.set('reviews', []);
},
error: function(error) {
console.log(error);
}
});
*/
basicResponse(res, true, user);
},
error: function(user, error) {
basicResponse(res, false, { error: error.code });
// Show the error message somewhere and let the user try again.
console.log("Error: " + error.code + " " + error.message);
}
});
};
this.processLogout = function(req, res) {
Parse.User.logOut();
basicResponse(res, true);
};
var basicResponse = function(res, $success, $data) {
res.send({
success : $success,
info : $data
});
};<file_sep>/public/js/new-reviews-controller.js
$(document).ready(function() {
console.log("IM A CONTAINER");
// Initialize editor with custom theme and modules
var fullEditor = new Quill('#editor', {
modules: {
'toolbar': { container: '#toolbar' },
'link-tooltip': true
},
theme: 'snow'
});
/*
$('#summary ul').on('keydown', 'li input', function (e) {
var hasEmpty = false;
$(this).parent().parent().children().each(function (i, e) {
if ($(e).children().val() === '') {
hasEmpty = true;
}
});
console.log(e.which);
if (!hasEmpty && e.which === 13 && $(this).val() !== '') {
console.log($(this).val());
$(this).parent().after('<li><input type="text"></li>');
$(this).parent().parent().children().last().children().focus();
}
else if (e.which === 8 && $(this).val() === '') {
$(this).parent().remove();
e.preventDefault();
$(this).parent().parent().children().last().children().focus();
}
});
*/
$('#publish-button').click(function(e) {
var good = '', okay = '', bad = '';
$('#the-good ul li').each(function(i, e) {
good += $(this).children().val() + '\t';
});
$('#the-okay ul li').each(function(i, e) {
okay += $(this).children().val() + '\t';
});
$('#the-bad ul li').each(function(i, e) {
bad += $(this).children().val() + '\t';
});
var reviewJSON = {
game: $('#hidden-game-title').text(),
newReview: {
title: $('input[name=title]').val(),
reviewBody: fullEditor.getHTML(),
theGood: good,
theOkay: okay,
theBad: bad,
rating: $('input[name=rating]:checked').val(),
likes: 0,
bannerImage: $('#hidden-game-banner').text()
}
}
console.log(reviewJSON);
$.ajax({
type: 'POST',
url: '/api/addReview',
data: reviewJSON,
success: function(data) {
console.log(data);
window.location.href = '/';
}
});
});
});<file_sep>/routes/signup.js
var path = require('path');
module.exports = function(req, res) {
require(path.join(__dirname, 'header')).renderPage(req, res, 'signup');
};<file_sep>/routes/header.js
var fs = require('fs');
var path = require('path');
var handlebars = require('express3-handlebars');
var helpers = handlebars.create({
helpers: {
toLowerCase: function(str) {
return str.toLowerCase();
}
}
});
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
this.renderPage = function(req, res, route, moarDatum) {
var data = {};
data.username = '';
data.loggedIn = false;
data.header = path.join(__dirname, '../views/templates/header.handlebars');
data.footer = path.join(__dirname, '../views/templates/footer.handlebars');
data.createReviewModal = path.join(__dirname, '../views/templates/create-review-modal.handlebars');
data.page = route;
var currentUser = Parse.User.current();
try {
fs.readFileSync(path.join(__dirname, '../public/js/' + data.page + '-controller.js')).toString();
data.loadScript = true;
}
catch (e) {
data.loadScript = false;
}
if (moarDatum) {
mergeJSON(data, moarDatum);
}
if (currentUser) {
data.loggedIn = true;
data.username = currentUser.getUsername();
}
else {
data.loggedIn = false;
}
console.log(data.title);
var Game = Parse.Object.extend("Games");
var query = new Parse.Query(Game);
query.select("ref", "title");
query.find().then(function(gt) {
console.log(gt);
data.gameTitles = [];
gt.forEach(function(e) {
data.gameTitles.push({
title: e.get('title'),
ref: e.get('ref')
});
});
console.log(data.gameTitles);
helpers.render(data.createReviewModal, data, function(err, result) {
data.createReviewModal = result;
helpers.render(data.header, data, function(err, result) {
data.header = result;
helpers.render(data.footer, data, function(err, result) {
data.footer = result;
res.render(route, data);
});
});
});
});
};
var mergeJSON = function(obj1, obj2) {
Object.keys(obj2).forEach(function(p){
try {
if (obj2[p].constructor === Object) {
obj1[p] = MergeRecursive(obj1[p], obj2[p]);
}
else {
obj1[p] = obj2[p];
}
}
catch( e) {
obj1[p] = obj2[p];
}
});
return obj1;
}<file_sep>/routes/user-settings.js
var path = require('path');
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
module.exports = function(req, res) {
var query = new Parse.Query(Parse.User);
query.equalTo('username', Parse.User.current().getUsername());
query.first({
success: function(user) {
console.log(user);
var data = {
id: user.id,
myUsername: user.getUsername(),
myUserPic: user.get('userPic'),
email: user.get('email')
};
console.log(data);
require(path.join(__dirname, 'header')).renderPage(req, res, 'user-settings', data);
}
});
};<file_sep>/routes/api/main.js
var path = require('path');
var users = require(path.join(__dirname, 'users'));
var database = require(path.join(__dirname, 'database'));
module.exports = function(req, res) {
if( req.params.call === 'processSignup' )
users.processSignup (req, res);
else if( req.params.call === 'processLogin' )
users.processLogin (req, res);
else if( req.params.call === 'processLogout' )
users.processLogout (req, res);
else if( req.params.call === 'addReview')
database.addReview(req, res);
else
error (req, res);
};
var error = function(req, res) {
res.send('Invalid API call');
};<file_sep>/public/js/index-controller.js
$(document).ready(function() {
$('#staff-picks li').hover(function() {
$(this).children().each(function(i, e) {
$(e).addClass('show');
});
}, function() {
$(this).children().removeClass('show');
});
$('#staff-picks li').click(function() {
window.location.href = $(this).attr('data');
});
});<file_sep>/routes/users.js
var path = require('path');
var Parse = require('parse').Parse;
Parse.initialize('8ND8FWpNrWD1j2zkGymXBFAGWebC7xiuA2GT7zAk', 'tYcMRGV7XEpjFv782VzQ2ezItHVuU40vsCMZ71DU');
module.exports = function(req, res) {
var query = new Parse.Query(Parse.User);
query.equalTo('username', req.params.user);
query.first({
success: function(user) {
console.log(user);
user.relation('reviews').query().find({
success: function(reviews) {
var data = {
id: user.id,
myUsername: user.getUsername(),
myUserPic: user.get('userPic'),
email: user.get('email'),
reviews: []
};
reviews.forEach(function(review) {
// Format a summary for each review
var summary = review.get('reviewBody').replace(/<(?:.|\n)*?>/gm, '');
if (summary.length > 200)
{
summary = summary.substring(0, 200);
summary = summary.split(' ');
summary.pop();
summary = summary.join(' ');
summary += '...';
}
data.reviews.push({
id: review.id,
title: review.get('title'),
rating: review.get('rating'),
summary: summary,
review: review.get('review'),
tldr: review.get('tldr'),
timestamp: review.createdAt
});
});
console.log(data);
require(path.join(__dirname, 'header')).renderPage(req, res, 'users', data);
}
});
},
error: function(review, error) {
console.log(error);
}
});
}; | 9167842a73bedba6d910a3e5e83cad51385e26fc | [
"JavaScript"
] | 11 | JavaScript | theloneplant/GameRev | de9026862daa3d2e603fbafa76d318455e12c8d6 | 381b60b99c6b779ceb8066b868e378fb90a022ad |
refs/heads/master | <file_sep>#!/usr/bin/env python
# coding: utf-8
# In[94]:
import csv
import os
# In[96]:
input_file = open("budget_data.csv")
csv_read = csv.reader(input_file)
next(csv_read)
# In[97]:
count = 0
total = 0
# In[98]:
newL = []
# In[99]:
for row in csv_read:
if row[0] != 0:
count = 1 + count
n = int(row[1])
total = n + total
newL.append(row)
# In[100]:
greatest_inc = 0
inc_date = ""
greatest_dec = 0
dec_date = ""
# In[101]:
i = 0
change = 0
total_change = 0
while i < len(newL) - 1:
change = int(newL[i+1][1]) - int(newL[i][1])
total_change = change + total_change
if change < greatest_dec:
greatest_dec = change
dec_date = newL[i+1][0]
if change > greatest_inc:
greatest_inc = change
inc_date = newL[i+1][0]
i = i + 1
# In[102]:
avg_change = total_change/count
print("Financial Analysis")
print("----------------------------")
print("Total Months: {}".format(count))
print("Total: ${}".format(total))
print("Average Change: ${:.2f}".format(avg_change))
print("Greatest Increase in Profits: {} (${})".format(inc_date, greatest_inc))
print("Greatest Decrease in Profits: {} (${})".format(dec_date, greatest_dec))
# In[103]:
with open("budget_data.txt", "w") as new_file:
new_file.write("Financial Analysis \n")
new_file.write("---------------------------- \n")
new_file.write("Total Months: {} \n".format(count))
new_file.write("Total: ${} \n".format(total))
new_file.write("Average Change: ${:.2f} \n".format(avg_change))
new_file.write("Greatest Increase in Profits: {} (${})\n".format(inc_date, greatest_inc))
new_file.write("Greatest Decrease in Profits: {} (${}) \n".format(dec_date, greatest_dec))
<file_sep>#!/usr/bin/env python
# coding: utf-8
# In[1]:
import csv
# In[2]:
input_file = open("election_data.csv")
csv_read = csv.reader(input_file)
next(csv_read)
# In[3]:
total = 0
all_rows = []
cand = []
# In[4]:
for row in csv_read:
total = 1 + total
if row[2] not in cand:
cand.append(row[2])
all_rows.append(row)
# In[5]:
newDict = {i : 0 for i in cand}
# In[6]:
for key in newDict.keys():
for row in all_rows:
if row[2] == key:
newDict[key] += 1
# In[7]:
per_votes = []
for key in newDict.keys():
p = newDict[key]
per_votes.append(p/total)
# In[8]:
max_votes = 0
winner = ""
for key in newDict.keys():
if newDict[key] > max_votes:
max_votes = newDict[key]
winner = key
# In[9]:
print("Election Results")
print("-------------------------")
print("Total Votes: {}".format(total))
print("-------------------------")
print("Khan: {:.3f}% ({})".format(per_votes[0]*100, newDict["Khan"]))
print("Correy: {:.3f}% ({})".format(per_votes[1]*100, newDict["Correy"]))
print("Li: {:.3f}% ({})".format(per_votes[2]*100, newDict["Li"]))
print("O'Tooley: {:.3f}% ({})".format(per_votes[3]*100, newDict["O'Tooley"]))
print("-------------------------")
print("Winner: {}".format(winner))
print("-------------------------")
# In[10]:
with open("election_results2.txt", "w") as out_file:
out_file.write("Election Results \n")
out_file.write("------------------------- \n")
out_file.write("Total Votes: {} \n".format(total))
out_file.write("------------------------- \n")
out_file.write("Khan: {:.3f}% ({}) \n".format(per_votes[0]*100, newDict["Khan"]))
out_file.write("Correy: {:.3f}% ({}) \n".format(per_votes[1]*100, newDict["Correy"]))
out_file.write("Li: {:.3f}% ({}) \n".format(per_votes[2]*100, newDict["Li"]))
out_file.write("O'Tooley: {:.3f}% ({}) \n".format(per_votes[3]*100, newDict["O'Tooley"]))
out_file.write("------------------------- \n")
out_file.write("Winner: {} \n".format(winner))
out_file.write("------------------------- \n")
| ede395d4be56a4d84d5d915c322d5a8e0e79e949 | [
"Python"
] | 2 | Python | gemmavo/python-challenge | 19f6cacd04831e5b5f1776d10c80c44679986b45 | c7b757baac1b28b174cad9a34afda9dfef3093b7 |
refs/heads/main | <file_sep>WARNING!
THE AUTHOR DISCLAIMS LIABILITY FOR LOSSES CAUSED BY THIS PROGRAM.
YOU RUN IT AT YOUR OWN RISK!
<file_sep>import os,getpass
path = 'C:/Users/'+getpass.getuser()+'/AppData/Roaming/Microsoft/Windows/Start Menu/Programs/Startup/CREEPER.bat'
open(path, 'w').write('shutdown /r /t 0')
os.system('shutdown /r /t 30 /c "IM CREEPER CATH ME IF YOU CAN!"')
| f5f1d96f1427d5bba422539c85f1678d9af8caf9 | [
"Python",
"Text"
] | 2 | Text | ItMasterAntoni/CREEPER-virus | 5f8214af6e1c0cb36f33ea35359ce5ace2b5ea4f | 857fae6e4b89e4629f6ca3b1ee4a014a6769e940 |
refs/heads/master | <file_sep>// Copyright (c) FIRST and other WPILib contributors.
// Open Source Software; you can modify and/or share it under the terms of
// the WPILib BSD license file in the root directory of this project.
package frc.robot;
import edu.wpi.first.networktables.NetworkTable;
import edu.wpi.first.networktables.NetworkTableInstance;
import edu.wpi.first.wpilibj.GenericHID;
import edu.wpi.first.wpilibj.Joystick;
import edu.wpi.first.wpilibj.SlewRateLimiter;
import edu.wpi.first.wpilibj.TimedRobot;
import edu.wpi.first.wpilibj.Joystick;
import edu.wpi.first.wpilibj.shuffleboard.Shuffleboard;
import edu.wpi.first.wpilibj.smartdashboard.SmartDashboard;
public class Robot extends TimedRobot {
private final Joystick m_controller = new Joystick(0);
private final Drivetrain m_swerve = new Drivetrain();
// Slew rate limiters to make joystick inputs more gentle; 1/3 sec from 0 to 1.
private final SlewRateLimiter m_xspeedLimiter = new SlewRateLimiter(3);
private final SlewRateLimiter m_yspeedLimiter = new SlewRateLimiter(3);
private final SlewRateLimiter m_rotLimiter = new SlewRateLimiter(3);
@Override
public void robotInit() {
// NetworkTable table = NetworkTableInstance.getDefault().getTable("MyTable");
// table.getEntry("")
SmartDashboard.putBoolean("Override", false);
SmartDashboard.putBoolean("ZERO", false);
}
@Override
public void teleopInit() {
m_swerve.m_frontLeft.m_yawEncoder.reset(); // for debugging testing as the encoder is not absolute
}
// @Override
// public void autonomousPeriodic() {
// driveWithJoystick(false);
// m_swerve.updateOdometry();
// }
@Override
public void teleopPeriodic() {
driveWithJoystick(true);
}
private void driveWithJoystick(boolean fieldRelative) {
// Get the x speed. We are inverting this because Xbox controllers return
// negative values when we push forward.
double xSpeed = -m_controller.getY();
// -m_xspeedLimiter.calculate(m_controller.getY(GenericHID.Hand.kLeft))
// * frc.robot.Drivetrain.kMaxSpeed;
// Get the y speed or sideways/strafe speed. We are inverting this because
// we want a positive value when we pull to the left. Xbox controllers
// return positive values when you pull to the right by default.
double ySpeed = -m_controller.getX();
// -m_yspeedLimiter.calculate(m_controller.getX(GenericHID.Hand.kLeft))
// * frc.robot.Drivetrain.kMaxSpeed;
// Get the rate of angular rotation. We are inverting this because we want a
// positive value when we pull to the left (remember, CCW is positive in
// mathematics). Xbox controllers return positive values when you pull to
// the right by default.
double rot = m_controller.getZ();
// -m_rotLimiter.calculate()
// * frc.robot.Drivetrain.kMaxAngularSpeed;
if (Math.abs(rot) < .1) {
rot = 0;
}
if (Math.abs(xSpeed) < .1) {
xSpeed = 0;
}
if (Math.abs(ySpeed) < .1) {
ySpeed = 0;
}
if (SmartDashboard.getBoolean("Override", false)) {
xSpeed = SmartDashboard.getNumber("xSpeed", 0);
ySpeed = SmartDashboard.getNumber("ySpeed", 0);
rot = SmartDashboard.getNumber("rot", 0);
}
if (SmartDashboard.getBoolean("ZERO", false)) {
SmartDashboard.putBoolean("ZERO", false);
xSpeed = 0;
ySpeed = 0;
rot = 0;
}
SmartDashboard.putNumber("xSpeed", xSpeed);
SmartDashboard.putNumber("ySpeed", ySpeed);
SmartDashboard.putNumber("rot", rot);
SmartDashboard.putNumber("Encoder Angle", m_swerve.m_frontLeft.m_yawEncoder.getDistance());
m_swerve.drive(xSpeed, ySpeed, rot);
}
}
| 1c4e162fe3a1c19241ba6cd56f1fbbaf16e6f34d | [
"Java"
] | 1 | Java | team4909/WPI-Swerve | ec983ad326b21faaa97ddf4620a4c5a0c5e2b1c6 | a74e6a730c590ece4b5292bcb0356302a4f8448f |
refs/heads/master | <repo_name>carriercomm/chatServer<file_sep>/chat_client
#!/bin/bash
#usage: chat_client ipaddress
#example: chat_client localhost
#example: chat_client 192.168.3.11
telnet $1 $CHAT_SERVER_PORT
<file_sep>/README.md
# chatServer By <NAME>
* Basic Chat Server implemented in haskell
* chat room like functionality
* github link: github link: https://github.com/allonsy/chatServer
* email: <EMAIL> or <EMAIL>
##Build instructions:
* Clone the repository or untar the cabal sdist tar.gz file
* You will need the Network and Hspec packages so make sure that you build
from a cabal sandbox with these packages installed or have these
packages globally installed.
run
* `cabal configure`
* `cabal build`
to run the tests, run:
* `cabal configure --enable-tests`
* `cabal test`
##Executing instructions
* set the CHAT_SERVER_PORT environment variable
* to run it execute `dist/build/chat/chat`
* or you may execute `cabal run`
* You may use telnet as a client, go ahead and connect to the ip of
the server at the port pointed to by the environment variable
* I have provided in the github repo a quick bash script that
calls telnet with the desired parameters. Its syntax is:
`./chat_client 172.16.31.10` or `./chat_client localhost`
## Using telnet as a client
* just connect via telnet as mentioned above
* type messages as normal and hit enter to send them to the server
* quit telnet or type ":q" to quit
| bbbbaa5fc3b880437d08cb04014f895a652f0f26 | [
"Markdown",
"Shell"
] | 2 | Shell | carriercomm/chatServer | ef48a781ee410742fece1de3f0cd9b0fee748f87 | bddfef9c514d8bc5f280791610d6fd0dc0aacac9 |
refs/heads/main | <repo_name>hfdpx/boltdb<file_sep>/freelist.go
package bolt
import (
"fmt"
"sort"
"unsafe"
)
//freelist 空闲page列表
type freelist struct {
// 空闲页列表,db文件中一组连续的页,用于保存中db使用过程中由于修改操作而释放的页的id列表
ids []pgid // 可以分配的空闲页列表id
pending map[txid][]pgid // 按照事务id 分别记录了中 对应事务期间新增的空闲页列表
// pending需要单独记录的原因:
// 1.某个写事务(txid=7)已经提交,但可能仍然有一些读事务(如txid<7)仍然中使用其刚释放的页,因此不能立即用于分配
cache map[pgid]bool // (pending释放标记,用于验证页面是否空闲)(快速查找所有空闲和待处理页面ID)
}
//newFreelist 返回一个空的初始化好的 freelist
func newFreelist() *freelist {
return &freelist{
pending: make(map[txid][]pgid),
cache: make(map[pgid]bool),
}
}
//size 返回freelist size
func (f *freelist) size() int {
n := f.count()
// 如果溢出了,那么按照规定,真正的存储计数存储中数据部分的第一个元素,所以n需要+1
if n >= 0xFFFF {
n++
}
return pageHeaderSize + (int(unsafe.Sizeof(pgid(0))) * n) // 头部+元素大小*元素数量
}
//count 返回 freelist 的page数量
func (f *freelist) count() int {
return f.freeCount() + f.pendingCount()
}
//freeCount 返回 freelist 中free page 数量
func (f *freelist) freeCount() int {
return len(f.ids)
}
//pendingCount 返回 freelist 中 pending page 数量
func (f *freelist) pendingCount() int {
var count int
for _, list := range f.pending {
count += len(list)
}
return count
}
//copyall 合并 ids 和 pending 并排序
func (f *freelist) copyall(dst []pgid) {
m := make(pgids, 0, f.pendingCount())
for _, list := range f.pending {
m = append(m, list...)
}
sort.Sort(m)
mergepgids(dst, f.ids, m)
}
//allocate 分配n个空闲的page
func (f *freelist) allocate(n int) pgid {
// 如果可以找到连续的n个空闲页,则返回起始页id,否则返回0
// 分配规则:分配单位是页,分配策略是首次适应:即从排序好的空闲列表ids中,找到第一段等于指定长度的连续空闲页,然后返回起始页id
if len(f.ids) == 0 {
return 0
}
// 遍历寻找连续空闲页,并且判断是否等于n
var initial, previd pgid
for i, id := range f.ids {
if id <= 1 {
panic(fmt.Sprintf("invalid page allocation: %d", id))
}
// 非连续,重置initial
if previd == 0 || id-previd != 1 {
initial = id
}
// 找到了n个连续的page
if (id-initial)+1 == pgid(n) {
// 当正好分配到ids中的前n个page时,仅仅简单的往前调整 f.ids 切片即可
// 虽然这样会造成空间浪费,但是中对f.ids 进行appned/free时,会重新进行空间分配,而这些被浪费的空间会被GC
if (i + 1) == n {
f.ids = f.ids[i+1:]
} else {
// 把n个连续page后面的page都往前挪n个位置
copy(f.ids[i-n+1:], f.ids[i+1:])
f.ids = f.ids[:len(f.ids)-n]
}
// 从f.cache中删除这n个page的id
for i := pgid(0); i < pgid(n); i++ {
delete(f.cache, initial+i)
}
//返回起始 page id
return initial
}
previd = id
}
return 0
}
// free 释放page空间到pending缓存上面
func (f *freelist) free(txid txid, p *page) {
if p.id <= 1 {
panic(fmt.Sprintf("cannot free page 0 or 1: %d", p.id))
}
// 释放从[p.id, p.id+p.overflow]的的page(标记释放而已)
var ids = f.pending[txid]
for id := p.id; id <= p.id+pgid(p.overflow); id++ {
// 验证页面是否空闲
if f.cache[id] {
panic(fmt.Sprintf("page %d already freed", id))
}
ids = append(ids, id)
f.cache[id] = true // 打上释放标记
}
f.pending[txid] = ids
}
//release 将事务ID小于txid的事务的page 从f.pending 移动到 f.ids
func (f *freelist) release(txid txid) {
m := make(pgids, 0)
for tid, ids := range f.pending {
if tid <= txid {
m = append(m, ids...)
delete(f.pending, tid)
}
}
sort.Sort(m)
f.ids = pgids(f.ids).merge(m)
}
// rollback 回滚,将特定事务期间的 pages ID 删除
func (f *freelist) rollback(txid txid) {
// 从f.pending中移除 事务的一些page
for _, id := range f.pending[txid] {
delete(f.cache, id)
}
// 在f.pending中删除对应事务的page id 列表
delete(f.pending, txid)
}
// freed 判断给定 page 是否在 空闲列表 中。
func (f *freelist) freed(pgid pgid) bool {
return f.cache[pgid]
}
//read 从page中加载freeList
func (f *freelist) read(p *page) {
// 在数据库重启时,会从仅有的两个元信息页中恢复一个合法的元信息,然后根据元信息中的freelist字段,找到存储freelist页的起始位置,然后恢复到内存中
// 如果p.count溢出了,那么其真正的值存储在数据部分的第一个元素中
idx, count := 0, int(p.count)
if count == 0xFFFF {
idx = 1
count = int(((*[maxAllocSize]pgid)(unsafe.Pointer(&p.ptr)))[0])
}
// 构建 f.ids
if count == 0 {
f.ids = nil
} else {
ids := ((*[maxAllocSize]pgid)(unsafe.Pointer(&p.ptr)))[idx:count]
f.ids = make([]pgid, len(ids))
copy(f.ids, ids)
// 保证f.ids有序
sort.Sort(pgids(f.ids))
}
// 构建 f.cache
f.reindex()
}
// reindex 根据f.ids和f.pending 来重建 f.cache
func (f *freelist) reindex() {
f.cache = make(map[pgid]bool, len(f.ids))
for _, id := range f.ids {
f.cache[id] = true
}
for _, pendingIDs := range f.pending {
for _, pendingID := range pendingIDs {
f.cache[pendingID] = true
}
}
}
//write freeList 写入page
func (f *freelist) write(p *page) error {
// freelist通过wirte函数,在事务提交时,将自己写入给定的页,进行持久化,在写入时,将pending和ids合并后写入
// todo 待理解!
// 合并写入的原因如下:
// 1.write用于写事务提交时调用,写事务是串行的,因此pending中对应的写事务都已提交,所以可以合并
// 2.写入文件是为了应对崩溃后重启,而重启时没有任何读操作,自然不用担心还有读事务还在使用刚释放的页
// 注意:本步骤只是将freelist转化为内存中的页结构,需要额外的操作才能将对应的页持久化到文件(比如tx.write)
// 设置 页 类型
p.flags |= freelistPageFlag
//p.count 是uint16类型,范围是[0,64k-1],0xFFFF=64k
lenids := f.count()
if lenids == 0 {
p.count = uint16(lenids)
} else if lenids < 0xFFFF {
p.count = uint16(lenids)
// 通过capyall将 pending和ids 合并并且排序
f.copyall(((*[maxAllocSize]pgid)(unsafe.Pointer(&p.ptr)))[:])
} else {
p.count = 0xFFFF
((*[maxAllocSize]pgid)(unsafe.Pointer(&p.ptr)))[0] = pgid(lenids)
f.copyall(((*[maxAllocSize]pgid)(unsafe.Pointer(&p.ptr)))[1:]) //如果空闲页id超出了64k,则将其放在数据部分的第一个位置上
}
return nil
}
// reload 从page读取freeList并且过滤掉f.pending
func (f *freelist) reload(p *page) {
f.read(p)
// Build a cache of only pending pages.
pcache := make(map[pgid]bool)
for _, pendingIDs := range f.pending {
for _, pendingID := range pendingIDs {
pcache[pendingID] = true
}
}
// Check each page in the freelist and build a new available freelist
// with any pages not in the pending lists.
var a []pgid
for _, id := range f.ids {
if !pcache[id] {
a = append(a, id)
}
}
f.ids = a
// Once the available list is rebuilt then rebuild the free cache so that
// it includes the available and pending free pages.
f.reindex()
}
<file_sep>/README.md
# boltdb
### 基础理论
* 在文件系统上,boltdb采用页的组织形式,将一切数据都对齐到页
* 在内存中,boltdb采用B+树组织数据,其基本单元树节点
* 一个内存中到树节点对应文件系统上一个或者多个连续的页
### 数据组织形式(自上而下)
* 每个db对应一个文件
* 在逻辑上,一个db包含多个bucket,相当于多个namespace,每个bucket对应一颗B+树,每个bucket的root节点在顶层也组织成一颗树,但不一定是B+树
* 在物理上,一个db文件按页为单位进行顺序存储,一个页的大小和操作系统页的大小保持一致(4KB)
### page和节点(node)的对应关系:
* page是文件存储的基本单位,node是 B+ tree 的基本构成节点
* 一个数据node对应一到多个连续的数据page
* 连续的数据page序列化加载到内存中就成为一个数据node
##### 总结:在文件系统上线性组织的数据page,通过页内指针,在逻辑上组成来一棵二维的B+tree,该树的树根保证在元信息page中,而文件中所有其他没有用到的页的ID列表,保存在空闲列表page中
<file_sep>/page.go
package bolt
import (
"fmt"
"os"
"sort"
"unsafe"
)
//------------------------------------------------------ 常量 -----------------------------------------------------------//
// page头部大小
const pageHeaderSize = int(unsafe.Offsetof(((*page)(nil)).ptr))
// 每个page中最少的key数量
const minKeysPerPage = 2
// 分支页元素头部 大小
const branchPageElementSize = int(unsafe.Sizeof(branchPageElement{}))
// 叶子页元素头部 大小
const leafPageElementSize = int(unsafe.Sizeof(leafPageElement{}))
const (
bucketLeafFlag = 0x01
)
// page类型
const (
branchPageFlag = 0x01 // 分支节点page: 数据page,存放B+树中的中间节点
leafPageFlag = 0x02 // 叶子节点page: 数据page,存放B+树中的叶子节点
metaPageFlag = 0x04 // 元信息page: 全局仅有两个元信息page,保存在文件,是实现事务的关键
freelistPageFlag = 0x10 // 空闲列表page: 存放空闲页ID列表,在文件中表现为一段一段的连续的page
)
//------------------------------------------------------ page -----------------------------------------------------------//
type pgid uint64
type page struct {
id pgid // page id
flags uint16 // page 类型
count uint16 // 对应节点包含的元素个数,比如说包含的KV对
overflow uint32 // 对应节点元素溢出到其他page的page数量,即使用 overflow+1 个page来保存对应节点的信息
//以上四个字段构成了 page 定长header
ptr uintptr //数据指针,指向数据对应的byte数组,当 overflow>0 时会跨越多个连续的物理page,不过多个物理page在内存中也只会用一个page结构体来表示
}
// PageInfo 人类可读的page结构
type PageInfo struct {
ID int // ID
Type string // 类型: branch/leaf/meta/freelist/unkonw
Count int // 元素数量
OverflowCount int // 溢出page数量
}
//为了避免载入内存和写入文件系统时的序列化和反序列化操作,使用了大量的unsafe包中的指针操作
// 返回一个表示page类型的字符串
func (p *page) typ() string {
if (p.flags & branchPageFlag) != 0 {
return "branch"
} else if (p.flags & leafPageFlag) != 0 {
return "leaf"
} else if (p.flags & metaPageFlag) != 0 {
return "meta"
} else if (p.flags & freelistPageFlag) != 0 {
return "freelist"
}
return fmt.Sprintf("unknown<%02x>", p.flags)
}
// 返回一个指向page数据部分的指针
func (p *page) meta() *meta {
return (*meta)(unsafe.Pointer(&p.ptr))
}
// 通过index获取叶子节点
func (p *page) leafPageElement(index uint16) *leafPageElement {
n := &((*[0x7FFFFFF]leafPageElement)(unsafe.Pointer(&p.ptr)))[index]
return n
}
// 获取叶子节点列表
func (p *page) leafPageElements() []leafPageElement {
if p.count == 0 {
return nil
}
return ((*[0x7FFFFFF]leafPageElement)(unsafe.Pointer(&p.ptr)))[:]
}
// 通过index获取中间节点
func (p *page) branchPageElement(index uint16) *branchPageElement {
return &((*[0x7FFFFFF]branchPageElement)(unsafe.Pointer(&p.ptr)))[index]
}
// 获取中间节点列表
func (p *page) branchPageElements() []branchPageElement {
if p.count == 0 {
return nil
}
return ((*[0x7FFFFFF]branchPageElement)(unsafe.Pointer(&p.ptr)))[:]
}
// 将页面的 n 个字节作为 16进制 输出写入到stderr
func (p *page) hexdump(n int) {
buf := (*[maxAllocSize]byte)(unsafe.Pointer(p))[:n]
_, _ = fmt.Fprintf(os.Stderr, "%x\n", buf)
}
type pages []*page
func (s pages) Len() int { return len(s) }
func (s pages) Swap(i, j int) { s[i], s[j] = s[j], s[i] }
func (s pages) Less(i, j int) bool { return s[i].id < s[j].id }
type pgids []pgid
func (s pgids) Len() int { return len(s) }
func (s pgids) Swap(i, j int) { s[i], s[j] = s[j], s[i] }
func (s pgids) Less(i, j int) bool { return s[i] < s[j] }
// merge 合并a和b,并排序
func (a pgids) merge(b pgids) pgids {
// Return the opposite slice if one is nil.
if len(a) == 0 {
return b
}
if len(b) == 0 {
return a
}
merged := make(pgids, len(a)+len(b))
mergepgids(merged, a, b)
return merged
}
func mergepgids(dst, a, b pgids) {
if len(dst) < len(a)+len(b) {
panic(fmt.Errorf("mergepgids bad len %d < %d + %d", len(dst), len(a), len(b)))
}
// Copy in the opposite slice if one is nil.
if len(a) == 0 {
copy(dst, b)
return
}
if len(b) == 0 {
copy(dst, a)
return
}
// Merged will hold all elements from both lists.
merged := dst[:0]
// Assign lead to the slice with a lower starting value, follow to the higher value.
lead, follow := a, b
if b[0] < a[0] {
lead, follow = b, a
}
// Continue while there are elements in the lead.
for len(lead) > 0 {
// Merge largest prefix of lead that is ahead of follow[0].
n := sort.Search(len(lead), func(i int) bool { return lead[i] > follow[0] })
merged = append(merged, lead[:n]...)
if n >= len(lead) {
break
}
// Swap lead and follow.
lead, follow = follow, lead[n:]
}
// Append what's left in follow.
_ = append(merged, follow...)
}
//------------------------------------------------------ page 元素 -----------------------------------------------------------//
//branchPageElement 分支页元素 头部
type branchPageElement struct {
// 只存储key,不存储value本身
pos uint32 // key偏移量
ksize uint32 // key字节数
pgid pgid // 子节点的pageID(相当于value,只是没有存储value本身)
}
// key 返回key
func (n *branchPageElement) key() []byte {
buf := (*[maxAllocSize]byte)(unsafe.Pointer(n))
return (*[maxAllocSize]byte)(unsafe.Pointer(&buf[n.pos]))[:n.ksize]
}
// 叶子页元素 header
type leafPageElement struct {
// 对应:某个bucket的B+树中的叶子节点,存储是某个bucket中的一条用户数据
// 对应:顶层bucket树中的叶子节点,存储的是该db中的某个subbucket
// &leafPageElement + pos == &key
// &leafPageElement + pos + ksize == &value
flags uint32 // 标志位:指明是普通kv还是subbucket
pos uint32 // 偏移位:kv header与对应的 kv 的距离
ksize uint32 // 统计位:key字节数
vsize uint32 // 统计位:value字节数
}
// key 返回key
func (n *leafPageElement) key() []byte {
buf := (*[maxAllocSize]byte)(unsafe.Pointer(n))
return (*[maxAllocSize]byte)(unsafe.Pointer(&buf[n.pos]))[:n.ksize:n.ksize]
}
// value 返回value
func (n *leafPageElement) value() []byte {
buf := (*[maxAllocSize]byte)(unsafe.Pointer(n))
return (*[maxAllocSize]byte)(unsafe.Pointer(&buf[n.pos+n.ksize]))[:n.vsize:n.vsize]
}
<file_sep>/tx.go
package bolt
import (
"fmt"
"io"
"os"
"sort"
"strings"
"time"
"unsafe"
)
//------------------------------------------------------ Tx -----------------------------------------------------------//
type txid uint64
//Tx 事务
type Tx struct {
//Tx 表示数据库上的只读或读写事务。只读事务可用于检索键的值和创建游标。读写事务可以创建和删除存储桶以及创建和删除键。重要提示:处理完事务后,必须提交或回滚事务。
// 全部对数据的操作必须发生在一个事务之内,bolt的并发读取都在此实现
writable bool // 标志位:读写事务 or 仅读事务
managed bool // 标志位:当前事务是否被db托管,即通过db.Update()或db.View来写或读数据库
db *DB // 指向当前db
meta *meta // 事务初始化时 从db复制过来的meta 信息
root Bucket // 事务的 根bucket
pages map[pgid]*page // 事务的读或写page
stats TxStats // 事务操作状态统计
commitHandlers []func() // commit时的回调函数
WriteFlag int // 复制或者打开数据库文件时,指定文件打开模式
}
//init 事务 初始化
func (tx *Tx) init(db *DB) {
tx.db = db
tx.pages = nil
// 创建一个空的mate对象,并初始化为tx.meta,再讲db中的mate复制到刚刚创建的mate对象中(对象拷贝而非指针拷贝)
tx.meta = &meta{}
db.meta().copy(tx.meta)
// 为当前事务创建一个bucket,并将其设为根bucket,同时用mate中保存的根bucket头部来初始化事务的根bucket头部
tx.root = newBucket(tx)
tx.root.bucket = &bucket{}
*tx.root.bucket = tx.meta.root
// 如果是读写事务,就将mate中的txid+1,这样当该读写事务commit之后,mate会更新到数据库文件中,数据库的修改版本号就增加了
if tx.writable {
tx.pages = make(map[pgid]*page) // 读写事务需要一个page cache
tx.meta.txid += txid(1)
}
}
//ID 返回事务ID
func (tx *Tx) ID() int {
return int(tx.meta.txid)
}
//DB 返回一个对db的引用
func (tx *Tx) DB() *DB {
return tx.db
}
//Size 返回当前db size
func (tx *Tx) Size() int64 {
// 当前事务使用到的最大page ID * page 大小
return int64(tx.meta.pgid) * int64(tx.db.pageSize)
}
//Writable 判断是否是 读写事务
func (tx *Tx) Writable() bool {
return tx.writable
}
//Cursor 返回一个新建的游标
func (tx *Tx) Cursor() *Cursor {
return tx.root.Cursor()
}
//Stats 返回 事务的操作状态统计 副本
func (tx *Tx) Stats() TxStats {
return tx.stats
}
//Bucket 返回一个特定名称的bucket,该bucket仅仅在事务生存期间有效
func (tx *Tx) Bucket(name []byte) *Bucket {
return tx.root.Bucket(name)
}
// CreateBucket 创建一个特定名称的bucket,该bucket仅仅在事务生存期间有效
func (tx *Tx) CreateBucket(name []byte) (*Bucket, error) {
return tx.root.CreateBucket(name)
}
// CreateBucketIfNotExists 创建一个特定名称的bucket,条件是如果该bucket不存在,该bucket仅仅在事务生存期间有效
func (tx *Tx) CreateBucketIfNotExists(name []byte) (*Bucket, error) {
return tx.root.CreateBucketIfNotExists(name)
}
// DeleteBucket 删除特定名称 bucket
func (tx *Tx) DeleteBucket(name []byte) error {
return tx.root.DeleteBucket(name)
}
// ForEach 对每个bucket执行指定函数,一个出错,则停止执行剩下的
func (tx *Tx) ForEach(fn func(name []byte, b *Bucket) error) error {
return tx.root.ForEach(func(k, v []byte) error {
if err := fn(k, tx.root.Bucket(k)); err != nil {
return err
}
return nil
})
}
// OnCommit 添加一个事务commit后的回调函数
func (tx *Tx) OnCommit(fn func()) {
tx.commitHandlers = append(tx.commitHandlers, fn)
}
// Commit 提交事务:将所有更改写入磁盘并更新元页。如果发生磁盘写入错误,或者在只读事务上调用commit,则返回错误
func (tx *Tx) Commit() error {
// 具体:
// 1. 从根bucket开始,对访问过的bucket进行合并和分裂,让进行过插入和删除操作的B+树重新达到平衡状态
// 2. 更新freeList页
// 3. 将由当前事务分配的page 写入 disk,需要分配page的地方有:a.节点分裂产生新节点,b.freeList页重新分配
// 4. 将meta 写入 disk
// 处理异常情况
_assert(!tx.managed, "managed tx commit not allowed") // 保证没有被db托管
if tx.db == nil {
// db 空
return ErrTxClosed
} else if !tx.writable {
// 事务不可写
return ErrTxNotWritable
}
// TODO(benbjohnson): Use vectorized I/O to write out dirty pages.
// 1.1 节点合并:对整个db的根bucket进行重新平衡,涉及节点:当前事务读写访问过的bucket中的有删除操作的节点
var startTime = time.Now()
tx.root.rebalance()
if tx.stats.Rebalance > 0 {
tx.stats.RebalanceTime += time.Since(startTime)
}
// 1.2 节点分裂:对整个db的根bucket进行溢出操作,涉及节点:key数量超过限制的节点,进行溢出操作(分裂节点)
startTime = time.Now()
if err := tx.root.spill(); err != nil {
tx.rollback()
return err
}
tx.stats.SpillTime += time.Since(startTime)
tx.meta.root.root = tx.root.root // 更新根bucket的根节点的page id,因为旋转和分裂后根bucket的根节点可能发生了变化
opgid := tx.meta.pgid
// 2.1 对于freelist占用的page:释放,重新分配,写入(freelist写入新分配的page)
tx.db.freelist.free(tx.meta.txid, tx.db.page(tx.meta.freelist))
p, err := tx.allocate((tx.db.freelist.size() / tx.db.pageSize) + 1)
if err != nil {
tx.rollback()
return err
}
if err := tx.db.freelist.write(p); err != nil {
tx.rollback()
return err
}
tx.meta.freelist = p.id
// 当page数量增加时,同步增长db大小
if tx.meta.pgid > opgid {
// 因为向文件写数据时,文件系统上该文件节点的元数据可能不会立刻刷新,导致文件的size不会立刻更新,当进程crash时,可能会出现写文件结束但是文件大小没有更新的情况,为了防止这种情况,主动增长文件大小
if err := tx.db.grow(int(tx.meta.pgid+1) * tx.db.pageSize); err != nil {
tx.rollback()
return err
}
}
// 3.1 将事务中所有的脏页写入磁盘
startTime = time.Now()
if err := tx.write(); err != nil {
tx.rollback()
return err
}
// 启用了严格模式
if tx.db.StrictMode {
// 开始一致性检查
ch := tx.Check()
var errs []string
for {
err, ok := <-ch
if !ok {
break
}
errs = append(errs, err.Error())
}
if len(errs) > 0 {
panic("check fail: " + strings.Join(errs, "\n"))
}
}
// 4.元数据 写入 disk,因为进行读写操作后,mate中的txid已经改变,root,freeList和paid也有可能已经更新了
if err := tx.writeMeta(); err != nil {
tx.rollback()
return err
}
tx.stats.WriteTime += time.Since(startTime) // 记录整个事务耗时
// 关闭事务,清空相关字段
tx.close()
// 执行事务提交后的回调函数
for _, fn := range tx.commitHandlers {
fn()
}
return nil
}
//Rollback 回滚,只读事务不必提交,回滚即可
func (tx *Tx) Rollback() error {
_assert(!tx.managed, "managed tx rollback not allowed")
if tx.db == nil {
return ErrTxClosed
}
tx.rollback()
return nil
}
//rollback
func (tx *Tx) rollback() {
if tx.db == nil {
return
}
// 如果是读写事务,则从freelist中删除这个txid对应的所有paid,并且重新刷新freelist
if tx.writable {
tx.db.freelist.rollback(tx.meta.txid)
tx.db.freelist.reload(tx.db.page(tx.db.meta().freelist))
}
// 最后,将事务关闭
tx.close()
}
//close 关闭事务
func (tx *Tx) close() {
if tx.db == nil {
return
}
// 如果当前事务是读写事务
if tx.writable {
// freelist 状态统计
var freelistFreeN = tx.db.freelist.freeCount()
var freelistPendingN = tx.db.freelist.pendingCount()
var freelistAlloc = tx.db.freelist.size()
// 移除db当前读写事务引用 & 读写锁解锁
tx.db.rwtx = nil
tx.db.rwlock.Unlock()
// 记录状态
tx.db.statlock.Lock()
tx.db.stats.FreePageN = freelistFreeN
tx.db.stats.PendingPageN = freelistPendingN
tx.db.stats.FreeAlloc = (freelistFreeN + freelistPendingN) * tx.db.pageSize
tx.db.stats.FreelistInuse = freelistAlloc
tx.db.stats.TxStats.add(&tx.stats)
tx.db.statlock.Unlock()
} else {
// 如果当前事务是仅读事务,
tx.db.removeTx(tx) // 在db上移除此事务
}
// 清除事务上的所有引用
tx.db = nil
tx.meta = nil
tx.root = Bucket{tx: tx}
tx.pages = nil
}
// Copy 整个db写入writer,用于向后兼容,不推荐使用
func (tx *Tx) Copy(w io.Writer) error {
_, err := tx.WriteTo(w)
return err
}
// WriteTo 整个db写入writer
func (tx *Tx) WriteTo(w io.Writer) (n int64, err error) {
// 打开 db 文件
f, err := os.OpenFile(tx.db.path, os.O_RDONLY|tx.WriteFlag, 0)
if err != nil {
return 0, err
}
defer func() { _ = f.Close() }()
// 生成一个meta page
buf := make([]byte, tx.db.pageSize)
page := (*page)(unsafe.Pointer(&buf[0]))
page.flags = metaPageFlag
*page.meta() = *tx.meta
// Write meta 0.
page.id = 0
page.meta().checksum = page.meta().sum64()
nn, err := w.Write(buf)
n += int64(nn)
if err != nil {
return n, fmt.Errorf("meta 0 copy: %s", err)
}
// Write meta 1
page.id = 1
page.meta().txid -= 1
page.meta().checksum = page.meta().sum64()
nn, err = w.Write(buf)
n += int64(nn)
if err != nil {
return n, fmt.Errorf("meta 1 copy: %s", err)
}
// 偏移 两个page 位置
if _, err := f.Seek(int64(tx.db.pageSize*2), os.SEEK_SET); err != nil {
return n, fmt.Errorf("seek: %s", err)
}
// 复制数据页
wn, err := io.CopyN(w, f, tx.Size()-int64(tx.db.pageSize*2))
n += wn
if err != nil {
return n, err
}
return n, f.Close()
}
// CopyFile 整个 db 拷贝到file
func (tx *Tx) CopyFile(path string, mode os.FileMode) error {
f, err := os.OpenFile(path, os.O_RDWR|os.O_CREATE|os.O_TRUNC, mode)
if err != nil {
return err
}
err = tx.Copy(f)
if err != nil {
_ = f.Close()
return err
}
return f.Close()
}
// Check 一致性检查
func (tx *Tx) Check() <-chan error {
ch := make(chan error)
go tx.check(ch)
return ch
}
//check
func (tx *Tx) check(ch chan error) {
// 检查是否有任何page被 双重释放 (即freelist中是否有相同的pageID)
freed := make(map[pgid]bool)
all := make([]pgid, tx.db.freelist.count())
tx.db.freelist.copyall(all)
for _, id := range all {
if freed[id] {
ch <- fmt.Errorf("page %d: already freed", id)
}
freed[id] = true
}
// 追踪每个可达page
reachable := make(map[pgid]*page)
reachable[0] = tx.page(0) // meta0
reachable[1] = tx.page(1) // meta1
for i := uint32(0); i <= tx.page(tx.meta.freelist).overflow; i++ {
reachable[tx.meta.freelist+pgid(i)] = tx.page(tx.meta.freelist)
}
// 递归检查bucket
tx.checkBucket(&tx.root, reachable, freed, ch)
// 看看有没有 page不可达并且没有被释放
for i := pgid(0); i < tx.meta.pgid; i++ {
_, isReachable := reachable[i]
if !isReachable && !freed[i] {
ch <- fmt.Errorf("page %d: unreachable unfreed", int(i))
}
}
//关闭通道,发出完成信号
close(ch)
}
//checkBucket 检查bucket
func (tx *Tx) checkBucket(b *Bucket, reachable map[pgid]*page, freed map[pgid]bool, ch chan error) {
// 忽略 inline bucket.
if b.root == 0 {
return
}
// 检查 bucket 使用的每一个page(非freelist中的page)
b.tx.forEachPage(b.root, 0, func(p *page, _ int) {
// 超出了当前最大pageID
if p.id > tx.meta.pgid {
ch <- fmt.Errorf("page %d: out of bounds: %d", int(p.id), int(b.tx.meta.pgid))
}
// 确保每个page 只被引用一次
for i := pgid(0); i <= pgid(p.overflow); i++ {
var id = p.id + i
if _, ok := reachable[id]; ok {
ch <- fmt.Errorf("page %d: multiple references", int(id))
}
reachable[id] = p
}
// bucket 使用的page里 存在已释放的page 或者 存在非叶子,非分支的page
if freed[p.id] {
ch <- fmt.Errorf("page %d: reachable freed", int(p.id))
} else if (p.flags&branchPageFlag) == 0 && (p.flags&leafPageFlag) == 0 {
ch <- fmt.Errorf("page %d: invalid type: %s", int(p.id), p.typ())
}
})
// 检查 该bucket下的 每个bucket
_ = b.ForEach(func(k, v []byte) error {
if child := b.Bucket(k); child != nil {
tx.checkBucket(child, reachable, freed, ch)
}
return nil
})
}
//allocate 分配 指定数量的 page
func (tx *Tx) allocate(count int) (*page, error) {
p, err := tx.db.allocate(count)
if err != nil {
return nil, err
}
// 将该page 归属于该
tx.pages[p.id] = p
// 更新 统计记录
tx.stats.PageCount++
tx.stats.PageAlloc += count * tx.db.pageSize
return p, nil
}
//write 将脏页写入disk
func (tx *Tx) write() error {
// 将tx中的脏页引用复制一份,并且释放原来的引用,tx对象并不是线程安全的,而接下来的写文件操作会比较耗时,此时应该避免tx.pages被修改
// 复制 事务的 page cache表
pages := make(pages, 0, len(tx.pages))
for _, p := range tx.pages {
pages = append(pages, p)
}
// 清空原有cache表,对新cache表按照di排序
tx.pages = make(map[pgid]*page)
sort.Sort(pages)
// 按照顺序将page写入disk
for _, p := range pages {
// 先得到page 实际大小
size := (int(p.overflow) + 1) * tx.db.pageSize
offset := int64(p.id) * int64(tx.db.pageSize)
// 将p转化成 最大限制大小的byte数组,ptr指向这个[]byte
ptr := (*[maxAllocSize]byte)(unsafe.Pointer(p))
for {
// size 不允许超出上限
sz := size
if sz > maxAllocSize-1 {
sz = maxAllocSize - 1
}
// 截取合适大小的块 写入
buf := ptr[:sz]
if _, err := tx.db.ops.writeAt(buf, offset); err != nil {
return err
}
// 更新计数
tx.stats.Write++
// 这个page全部写完了 则退出,写下一个page
size -= sz
if size == 0 {
break
}
// 指向page中的下一块 & prt指向p的剩余部分
offset += int64(sz)
ptr = (*[maxAllocSize]byte)(unsafe.Pointer(&ptr[sz]))
}
}
// 判断是否进行文件同步
if !tx.db.NoSync || IgnoreNoSync {
if err := fdatasync(tx.db); err != nil {
return err
}
}
// 将 小page放入 page池
for _, p := range pages {
// Ignore page sizes over 1 page.
// These are allocated using make() instead of the page pool.
if int(p.overflow) != 0 {
continue
}
buf := (*[maxAllocSize]byte)(unsafe.Pointer(p))[:tx.db.pageSize]
// See https://go.googlesource.com/go/+/f03c9202c43e0abb130669852082117ca50aa9b1
for i := range buf {
buf[i] = 0
}
tx.db.pagePool.Put(buf)
}
return nil
}
//writeMeta 将元数据 写入disk
func (tx *Tx) writeMeta() error {
// 为元数据创建临时缓冲区:申请一块buf,buf转化成page,p指向这块page
buf := make([]byte, tx.db.pageSize)
p := tx.db.pageInBuffer(buf, 0)
// 将元数据写入p page
tx.meta.write(p)
// 将 p page 写入file
if _, err := tx.db.ops.writeAt(buf, int64(p.id)*int64(tx.db.pageSize)); err != nil {
return err
}
if !tx.db.NoSync || IgnoreNoSync {
if err := fdatasync(tx.db); err != nil {
return err
}
}
// Update statistics.
tx.stats.Write++
return nil
}
//page 返回指定page的引用,不存在则创建一个再返回
func (tx *Tx) page(id pgid) *page {
// 先在事务使用的 pages中查找,实在找不到再从db映射的page中查找
if tx.pages != nil {
if p, ok := tx.pages[id]; ok {
return p
}
}
return tx.db.page(id)
}
//forEachPage 迭代给定page中的每个page并执行函数。
func (tx *Tx) forEachPage(pgid pgid, depth int, fn func(*page, int)) {
// 查找 page
p := tx.page(pgid)
// 执行函数
fn(p, depth)
// 子对象递归
if (p.flags & branchPageFlag) != 0 {
for i := 0; i < int(p.count); i++ {
elem := p.branchPageElement(uint16(i))
tx.forEachPage(elem.pgid, depth+1, fn)
}
}
}
//Page 返回特定的pageInfo(人为可读的page结构)
func (tx *Tx) Page(id int) (*PageInfo, error) {
// This is only safe for concurrent use when used by a writable transaction.
// 处理异常
if tx.db == nil {
return nil, ErrTxClosed
} else if pgid(id) >= tx.meta.pgid {
return nil, nil
}
// 构建 pageInfo
p := tx.db.page(pgid(id))
info := &PageInfo{
ID: id,
Count: int(p.count),
OverflowCount: int(p.overflow),
}
// 确定 pageInfo type
if tx.db.freelist.freed(pgid(id)) {
info.Type = "free"
} else {
info.Type = p.typ()
}
return info, nil
}
//------------------------------------------------------ TxStats -----------------------------------------------------------//
// TxStats 事务 操作状态统计
type TxStats struct {
// page 统计
PageCount int // page 分配数
PageAlloc int // 分配的总字节数
// cursor 统计
CursorCount int // 创建的游标数量
// Node 统计
NodeCount int // node 分配数
NodeDeref int // node 取消引用数
// Rebalance 统计
Rebalance int // node 重新平衡数
RebalanceTime time.Duration // 重新平衡 耗费的总时间
// Split/Spill 统计
Split int // number of nodes split
Spill int // number of nodes spilled
SpillTime time.Duration // total time spent spilling
// Write 统计
Write int // number of writes performed
WriteTime time.Duration // total time spent writing to disk
}
func (s *TxStats) add(other *TxStats) {
s.PageCount += other.PageCount
s.PageAlloc += other.PageAlloc
s.CursorCount += other.CursorCount
s.NodeCount += other.NodeCount
s.NodeDeref += other.NodeDeref
s.Rebalance += other.Rebalance
s.RebalanceTime += other.RebalanceTime
s.Split += other.Split
s.Spill += other.Spill
s.SpillTime += other.SpillTime
s.Write += other.Write
s.WriteTime += other.WriteTime
}
func (s *TxStats) Sub(other *TxStats) TxStats {
var diff TxStats
diff.PageCount = s.PageCount - other.PageCount
diff.PageAlloc = s.PageAlloc - other.PageAlloc
diff.CursorCount = s.CursorCount - other.CursorCount
diff.NodeCount = s.NodeCount - other.NodeCount
diff.NodeDeref = s.NodeDeref - other.NodeDeref
diff.Rebalance = s.Rebalance - other.Rebalance
diff.RebalanceTime = s.RebalanceTime - other.RebalanceTime
diff.Split = s.Split - other.Split
diff.Spill = s.Spill - other.Spill
diff.SpillTime = s.SpillTime - other.SpillTime
diff.Write = s.Write - other.Write
diff.WriteTime = s.WriteTime - other.WriteTime
return diff
}
<file_sep>/node.go
package bolt
import (
"bytes"
"fmt"
"sort"
"unsafe"
)
// 文件概述:
// 对node所存元素和node间关系的相关操作,节点内所存元素的增删,加载和落盘,访问孩子兄弟元素,拆分与合并的详细逻辑
// node和page的对应关系:文件系统中一组连续的物理page,加载到内存中成为一个逻辑page,进而转化为一个node
//------------------------------------------------------ node -----------------------------------------------------------//
// node 表示内存中一个反序列化后的 page
type node struct {
bucket *Bucket //指针:指向所在的bucket
isLeaf bool //标志位:是否为叶子节点
// 用于调整,维持B+树
unbalanced bool // 标志位:是否需要进行合并
spilled bool // 标志位:是否需要进行拆分和落盘
key []byte // 所含第一个元素的key
pgid pgid // 对应的page id
parent *node // 父节点指针
children nodes // 子节点们(只包含加载到内存中到部分孩子)
inodes inodes // 所存元素的元信息,对于分支节点是key+pgid的数组,对于叶子节点是kv数组
}
//inode 表示node所含的内部元素
type inode struct {
// 指向的元素不一定加载到了内存
flags uint32 // 用于leaf node,区分是正常value还是subbucket
pgid pgid
key []byte // 分支节点使用:指向的分支/叶子节点的page id
value []byte // 叶子节点使用:叶子节点所存储的数据
}
type inodes []inode
//root 返回node的root节点
func (n *node) root() *node {
if n.parent == nil {
return n
}
return n.parent.root()
}
//minKeys 返回 node inodes 的最小数量
func (n *node) minKeys() int {
if n.isLeaf {
return 1 // 叶子节点,只有一个节点
}
return 2 // 非叶节点,最少有它本身和它的一个叶子节点
}
//size 返回序列化后node的大小
func (n *node) size() int {
sz, elsz := pageHeaderSize, n.pageElementSize()
for i := 0; i < len(n.inodes); i++ {
item := &n.inodes[i]
sz += elsz + len(item.key) + len(item.value)
}
return sz
}
//sizeLessThan 判断序列化后node的大小,如果超过v,返回false,否则返回true
func (n *node) sizeLessThan(v int) bool {
sz, elsz := pageHeaderSize, n.pageElementSize()
for i := 0; i < len(n.inodes); i++ {
item := &n.inodes[i]
sz += elsz + len(item.key) + len(item.value)
if sz >= v {
return false
}
}
return true
}
//pageElementSize 根据node类型返回page大小
func (n *node) pageElementSize() int {
// node只有两种类型:leaf和brach
if n.isLeaf {
return leafPageElementSize
}
return branchPageElementSize
}
//childAt 返回给定索引处的子节点
func (n *node) childAt(index int) *node {
// 叶子节点,不可能挂子节点
if n.isLeaf {
panic(fmt.Sprintf("invalid childAt(%d) on a leaf node", index))
}
// 创建一个子节点
return n.bucket.node(n.inodes[index].pgid, n)
}
//childIndex 返回给定节点的索引
func (n *node) childIndex(child *node) int {
// 在inodes里面匹配child的key相同的节点的下标
index := sort.Search(len(n.inodes), func(i int) bool { return bytes.Compare(n.inodes[i].key, child.key) != -1 })
return index
}
//numChildren 返回节点的子节点数量
func (n *node) numChildren() int {
return len(n.inodes)
}
//nextSibling 返回具有相同父亲节点的下一个节点
func (n *node) nextSibling() *node {
if n.parent == nil {
return nil
}
index := n.parent.childIndex(n)
if index >= n.parent.numChildren()-1 {
return nil
}
return n.parent.childAt(index + 1)
}
//prevSibling 返回具有相同父节点的前一个节点
func (n *node) prevSibling() *node {
if n.parent == nil {
return nil
}
index := n.parent.childIndex(n)
if index == 0 {
return nil
}
return n.parent.childAt(index - 1)
}
//put 新增元素
func (n *node) put(oldKey, newKey, value []byte, pgid pgid, flags uint32) {
if pgid >= n.bucket.tx.meta.pgid {
panic(fmt.Sprintf("pgid (%d) above high water mark (%d)", pgid, n.bucket.tx.meta.pgid))
} else if len(oldKey) <= 0 {
panic("put: zero-length old key")
} else if len(newKey) <= 0 {
panic("put: zero-length new key")
}
// 找到插入点下标
index := sort.Search(len(n.inodes), func(i int) bool { return bytes.Compare(n.inodes[i].key, oldKey) != -1 }) // 字典序:a >= b
//fmt.Printf("插入点下标:%v\n\n",index)
// 如果key是新增而非替换,则需要为待插入节点腾出空间(index后面元素整体右移,腾出一个位置)
exact := len(n.inodes) > 0 && index < len(n.inodes) && bytes.Equal(n.inodes[index].key, oldKey)
if !exact {
n.inodes = append(n.inodes, inode{})
copy(n.inodes[index+1:], n.inodes[index:])
}
// 给插入/替换的元素赋值
inode := &n.inodes[index]
inode.flags = flags
inode.key = newKey
inode.value = value
inode.pgid = pgid
_assert(len(inode.key) > 0, "put: zero-length inode key")
}
//del 在node中删除一个key
func (n *node) del(key []byte) {
// 寻找key的下表
index := sort.Search(len(n.inodes), func(i int) bool { return bytes.Compare(n.inodes[i].key, key) != -1 })
// 如果节点找不到,直接返回
if index >= len(n.inodes) || !bytes.Equal(n.inodes[index].key, key) {
return
}
// 删除node的inodes中index位置的元素
n.inodes = append(n.inodes[:index], n.inodes[index+1:]...)
// 将节点标记为 需要重新平衡
n.unbalanced = true
}
//read 读取逻辑page转化为逻辑node
func (n *node) read(p *page) {
// 初始化元信息
n.pgid = p.id
n.isLeaf = (p.flags & leafPageFlag) != 0
n.inodes = make(inodes, int(p.count))
// 加载所包含元素 inodes
for i := 0; i < int(p.count); i++ {
inode := &n.inodes[i]
if n.isLeaf {
elem := p.leafPageElement(uint16(i))
inode.flags = elem.flags
inode.key = elem.key()
inode.value = elem.value()
} else {
elem := p.branchPageElement(uint16(i))
inode.pgid = elem.pgid
inode.key = elem.key()
}
_assert(len(inode.key) > 0, "read: zero-length inode key")
}
// 用第一个元素的key作为该node的key,以便父节点以此作为索引,进行查找和路由
if len(n.inodes) > 0 {
n.key = n.inodes[0].key
_assert(len(n.key) > 0, "read: zero-length node key")
} else {
n.key = nil
}
}
//write 将 node 写入 1个或多个 page
func (n *node) write(p *page) {
// 初始化page
if n.isLeaf {
p.flags |= leafPageFlag
} else {
p.flags |= branchPageFlag
}
// 防止溢出
if len(n.inodes) >= 0xFFFF {
panic(fmt.Sprintf("inode overflow: %d (pgid=%d)", len(n.inodes), p.id))
}
p.count = uint16(len(n.inodes))
if p.count == 0 {
return
}
// 遍历,写入
b := (*[maxAllocSize]byte)(unsafe.Pointer(&p.ptr))[n.pageElementSize()*len(n.inodes):]
for i, item := range n.inodes {
_assert(len(item.key) > 0, "write: zero-length inode key")
// Write the page element.
if n.isLeaf {
elem := p.leafPageElement(uint16(i))
elem.pos = uint32(uintptr(unsafe.Pointer(&b[0])) - uintptr(unsafe.Pointer(elem)))
elem.flags = item.flags
elem.ksize = uint32(len(item.key))
elem.vsize = uint32(len(item.value))
} else {
elem := p.branchPageElement(uint16(i))
elem.pos = uint32(uintptr(unsafe.Pointer(&b[0])) - uintptr(unsafe.Pointer(elem)))
elem.ksize = uint32(len(item.key))
elem.pgid = item.pgid
_assert(elem.pgid != p.id, "write: circular dependency occurred")
}
// key+value 长度太长,重新分配指针
klen, vlen := len(item.key), len(item.value)
if len(b) < klen+vlen {
b = (*[maxAllocSize]byte)(unsafe.Pointer(&b[0]))[:] // 扩容
}
// 写入b
copy(b[0:], item.key)
b = b[klen:]
copy(b[0:], item.value)
b = b[vlen:]
}
// DEBUG ONLY: n.dump()
}
//split node 切分成数个指定大小的node(大小不同)
func (n *node) split(pageSize int) []*node {
var nodes []*node
node := n
for {
// Split node into two.
a, b := node.splitTwo(pageSize)
// a添加到列表中,b留下来继续切
nodes = append(nodes, a)
// If we can't split then exit the loop.
if b == nil {
break
}
// Set node to b so it gets split on the next iteration.
node = b
}
return nodes
}
//splitTwo node 切分成两个
func (n *node) splitTwo(pageSize int) (*node, *node) {
// node足够小,不必再切
if len(n.inodes) <= (minKeysPerPage*2) || n.sizeLessThan(pageSize) {
return n, nil
}
// 确定阈值
var fillPercent = n.bucket.FillPercent
if fillPercent < minFillPercent {
fillPercent = minFillPercent
} else if fillPercent > maxFillPercent {
fillPercent = maxFillPercent
}
threshold := int(float64(pageSize) * fillPercent)
// 根据阈值确定切分的index
splitIndex, _ := n.splitIndex(threshold)
// 如果被拆分的node没有父节点,则需要新建一个
if n.parent == nil {
n.parent = &node{bucket: n.bucket, children: []*node{n}}
}
// 创建一个空节点next,next和node是兄弟节点
next := &node{bucket: n.bucket, isLeaf: n.isLeaf, parent: n.parent}
n.parent.children = append(n.parent.children, next)
// 将node节点的部分元素拆分到next
next.inodes = n.inodes[splitIndex:]
n.inodes = n.inodes[:splitIndex]
// 更新分裂计数
n.bucket.tx.stats.Split++
return n, next
}
//splitIndex 根据阈值给定切分的index
func (n *node) splitIndex(threshold int) (index, sz int) {
sz = pageHeaderSize
// Loop until we only have the minimum number of keys required for the second page.
for i := 0; i < len(n.inodes)-minKeysPerPage; i++ {
index = i
inode := n.inodes[i]
elsize := n.pageElementSize() + len(inode.key) + len(inode.value)
// minimum number of keys or 增加其他key将超过阈值
if i >= minKeysPerPage && sz+elsize > threshold {
break
}
// Add the element size to the total size.
sz += elsize
}
return
}
// spill 溢出操作,拆分节点
func (n *node) spill() error {
var tx = n.bucket.tx
// 溢出?
if n.spilled {
return nil
}
// 不能使用range,因为n.children的切片会一直变化
sort.Sort(n.children) // key sort
for i := 0; i < len(n.children); i++ {
// 递归给各个child进行spill
if err := n.children[i].spill(); err != nil {
return err
}
}
// 不再需要children,避免向上递归调用spill形成回路死循环
n.children = nil
// 按照 pagesize 将node 分裂出若干个新node,新node们和当前node恭喜同一个父node,返回的nodes中包含当前node
var nodes = n.split(tx.db.pageSize)
// 释放这些node占用的page,因为随后要分配新的page
// transaction commit 是只会向磁盘写入当前transacation分配的脏页
for _, node := range nodes {
if node.pgid > 0 {
tx.db.freelist.free(tx.meta.txid, tx.page(node.pgid))
node.pgid = 0
}
//分配page
p, err := tx.allocate((node.size() / tx.db.pageSize) + 1)
if err != nil {
return err
}
// node 写入 page
if p.id >= tx.meta.pgid { // 分配的page id 大于当前使用的最大page id
panic(fmt.Sprintf("pgid (%d) above high water mark (%d)", p.id, tx.meta.pgid))
}
node.pgid = p.id
node.write(p)
node.spilled = true //标志该node已经spill过了
// Insert into parent inodes.
if node.parent != nil {
var key = node.key
if key == nil {
key = node.inodes[0].key
}
// ???
// key属性下沉,向父节点更新或者添加key和pointer,以指向分裂产生的新node,将父node的key设为第一个子node的第一个key
node.parent.put(key, node.inodes[0].key, nil, node.pgid, 0)
node.key = node.inodes[0].key
_assert(len(node.key) > 0, "spill: zero-length node key")
}
// 更新spill 计数
tx.stats.Spill++
}
// 从根节点处递归处理完所有子节点的spill的过程后,若是根节点需要分裂,则它分裂后将产生新的根
// 对于新产生的根,我们也需要将其spill,并且同样对其孩子spill
if n.parent != nil && n.parent.pgid == 0 {
n.children = nil
return n.parent.spill() //防止spill过程又有新的parent节点产生
}
return nil
}
//rebalance 重新平衡操作,合并节点
func (n *node) rebalance() {
//已重新平衡过
if !n.unbalanced {
return
}
n.unbalanced = false
// 总平衡次数+1
n.bucket.tx.stats.Rebalance++
// 该节点的容量超过一个页的25% || inode的个数少于最少key数 则需要重新平衡
var threshold = n.bucket.tx.db.pageSize / 4
if n.size() > threshold && len(n.inodes) > n.minKeys() {
return
}
// n是根节点 特殊处理
if n.parent == nil {
// n 非叶子节点 && n只有一个子节点
if !n.isLeaf && len(n.inodes) == 1 {
// 将 n的孩子节点child 向上移 顶替掉n的位置
// 具体操作如下:
// 将n的子节点child的inodes 拷贝到 根节点上
child := n.bucket.node(n.inodes[0].pgid, n)
n.isLeaf = child.isLeaf
n.inodes = child.inodes[:]
n.children = child.children
// 将子节点的child的孩子节点的父亲 更新为 根节点
for _, inode := range n.inodes {
if child, ok := n.bucket.nodes[inode.pgid]; ok {
child.parent = n
}
}
// 移除 child
child.parent = nil
delete(n.bucket.nodes, child.pgid)
child.free()
}
return
}
// 如果n没有任何子节点,则删除n
// 1.先将父节点上 n的key和指向n的pointer删除
// 2.然后在删除n
// 3.由于父节点出现了删除,对父节点重新进行平衡操作
if n.numChildren() == 0 {
n.parent.del(n.key)
n.parent.removeChild(n)
delete(n.bucket.nodes, n.pgid)
n.free()
n.parent.rebalance()
return
}
// 确保 n 至少有一个兄弟节点
_assert(n.parent.numChildren() > 1, "parent must have at least 2 children")
// 找到兄弟节点
var target *node
var useNextSibling = n.parent.childIndex(n) == 0
if useNextSibling {
target = n.nextSibling()
} else {
target = n.prevSibling()
}
// 根据n和target的位置关系,决定究竟上谁合并到谁:位置靠右的节点合并到位置靠左的节点
if useNextSibling {
// 对target 的子节点进行 re parent 操作,新parent:n
for _, inode := range target.inodes {
if child, ok := n.bucket.nodes[inode.pgid]; ok {
child.parent.removeChild(child)
child.parent = n
child.parent.children = append(child.parent.children, child)
}
}
// target的inodes 移动到 n的inodes
// 删除释放 target
n.inodes = append(n.inodes, target.inodes...)
n.parent.del(target.key)
n.parent.removeChild(target)
delete(n.bucket.nodes, target.pgid)
target.free()
} else {
// 对n的inodes 进行re parent 操作,新parent:target
for _, inode := range n.inodes {
if child, ok := n.bucket.nodes[inode.pgid]; ok {
child.parent.removeChild(child)
child.parent = target
child.parent.children = append(child.parent.children, child)
}
}
// n的inodes 移动到 target的inodes
// 删除释放n
target.inodes = append(target.inodes, n.inodes...)
n.parent.del(n.key)
n.parent.removeChild(n)
delete(n.bucket.nodes, n.pgid)
n.free()
}
// 进行了删除释放操作,所以要对父节点进行重新平衡操作
n.parent.rebalance()
}
//removeChild 移除特定的child,不影响inodes
func (n *node) removeChild(target *node) {
for i, child := range n.children {
if child == target {
n.children = append(n.children[:i], n.children[i+1:]...)
return
}
}
}
//dereference 利用mmap进行解引用,可以避免指向旧数据
func (n *node) dereference() {
if n.key != nil {
key := make([]byte, len(n.key))
copy(key, n.key)
n.key = key
_assert(n.pgid == 0 || len(n.key) > 0, "dereference: zero-length node key on existing node")
}
for i := range n.inodes {
inode := &n.inodes[i]
key := make([]byte, len(inode.key))
copy(key, inode.key)
inode.key = key
_assert(len(inode.key) > 0, "dereference: zero-length inode key")
value := make([]byte, len(inode.value))
copy(value, inode.value)
inode.value = value
}
// Recursively dereference children.
for _, child := range n.children {
child.dereference()
}
// Update statistics.
n.bucket.tx.stats.NodeDeref++
}
// free 将节点的page 加入到 freeList中
func (n *node) free() {
if n.pgid != 0 {
n.bucket.tx.db.freelist.free(n.bucket.tx.meta.txid, n.bucket.tx.page(n.pgid))
n.pgid = 0
}
}
//------------------------------------------------------ nodes -----------------------------------------------------------//
type nodes []*node
func (s nodes) Len() int { return len(s) }
func (s nodes) Swap(i, j int) { s[i], s[j] = s[j], s[i] }
func (s nodes) Less(i, j int) bool {
return bytes.Compare(s[i].inodes[0].key, s[j].inodes[0].key) == -1
}
<file_sep>/test/main.go
package main
import (
"bolt"
"fmt"
"log"
"strconv"
)
func main() {
// 创建数据库,数据库文件为test.db,所有人可读写执行,采用默认的数据库配置
db, err := bolt.Open("test.db", 0777, nil)
defer db.Close()
if err != nil {
log.Fatal(err)
}
key := "testKey"
value := "testValue"
bucketName := "testBucket"
// 执行读写事务,采用Update
err = db.Update(func(tx *bolt.Tx) error {
// 在此读写事务中,如果该bucket不存在,则先创建bucket
b, err := tx.CreateBucketIfNotExists([]byte(bucketName))
if err != nil {
return fmt.Errorf("create bucket: %s", err)
}
for i := 0; i < 50; i++ {
newKey := key + "-" + strconv.Itoa(i)
newValue := value + "-" + strconv.Itoa(i)
//fmt.Println(newKey,newValue)
err = b.Put([]byte(newKey), []byte(newValue))
if err != nil {
return fmt.Errorf("insert kv: %s", err)
}
}
fmt.Printf("bucket 头部:%+v\n", b)
return nil
})
if err != nil {
log.Fatal(err)
}
//err = db.View(func(tx *bolt.Tx) error {
// b := tx.Bucket([]byte(bucketName))
// v := b.Get([]byte(key))
// fmt.Printf("The answer is: %s\n", v)
// return nil
//})
//if err != nil {
// log.Fatal(err)
// return
//}
}
<file_sep>/go.mod
module bolt
go 1.14
require (
github.com/boltdb/bolt v1.3.1
github.com/dustin/go-humanize v1.0.0 // indirect
github.com/kataras/tablewriter v0.0.0-20180708051242-e063d29b7c23 // indirect
github.com/lensesio/tableprinter v0.0.0-20201125135848-89e81fc956e7
github.com/mattn/go-runewidth v0.0.13 // indirect
golang.org/x/sys v0.0.0-20210809222454-d867a43fc93e
)
<file_sep>/bucket.go
package bolt
import (
"bytes"
"fmt"
"unsafe"
)
//------------------------------------------------------ 常量 -----------------------------------------------------------//
const (
// MaxKeySize key的最大长度,单位:字节
MaxKeySize = 32768
// MaxValueSize value的最大长度,单位:字节
MaxValueSize = (1 << 31) - 2
)
const (
maxUint = ^uint(0)
minUint = 0
maxInt = int(^uint(0) >> 1)
minInt = -maxInt - 1
)
// bucket头部大小
const bucketHeaderSize = int(unsafe.Sizeof(bucket{}))
const (
// 最小填充百分比
minFillPercent = 0.1
// 最大填充百分比
maxFillPercent = 1.0
)
// DefaultFillPercent 默认的bucket中节点的填充百分比阈值,当node中key的个数或者size超过整个node容量的某个百分比阈值之后,节点必须分裂为两个节点,这是为了防止B+树中插入kv对时引发频繁的再平衡操作
const DefaultFillPercent = 0.5
//------------------------------------------------------ bucket -----------------------------------------------------------//
// bucket 头部
type bucket struct {
root pgid // bucket 根节点 pageID
sequence uint64 // 序列号,自增
}
// Bucket db中一组kv对的集合
type Bucket struct {
*bucket // 头部
tx *Tx // 当前bucket所属的事务
buckets map[string]*Bucket // 子bucket cache
page *page // inline page reference:当bucket很小的时候,可以存储在此内联page中
rootNode *node // bucket 根节点
nodes map[pgid]*node // node cache
FillPercent float64 // node分裂阈值:当确定大多数写入操作是中尾部添加时,增大此阈值是有帮助的
}
//newBucket 返回一个与事务关联的新bucket
func newBucket(tx *Tx) Bucket {
var b = Bucket{tx: tx, FillPercent: DefaultFillPercent}
if tx.writable {
b.buckets = make(map[string]*Bucket)
b.nodes = make(map[pgid]*node)
}
return b
}
// Tx 返回bucket的tx
func (b *Bucket) Tx() *Tx {
return b.tx
}
// Root 返回bucket的根节点
func (b *Bucket) Root() pgid {
//之所以是返回pageID,因为执行的node不一定加载到了内存,当你需要访问此node的时候,会按需将page转化为node
return b.root
}
// Writable 判断该bucket事务是否是读写事务
func (b *Bucket) Writable() bool {
return b.tx.writable
}
// Cursor 创建一个该bucket上的游标【此游标仅事务存活期间有效】
func (b *Bucket) Cursor() *Cursor {
// 事务游标计数+1
b.tx.stats.CursorCount++
// 返回一个游标
return &Cursor{
bucket: b,
stack: make([]elemRef, 0),
}
}
// Bucket 根据name查找bucket,bucket不存在则返回nil【该bucket也仅仅在事务生存期间有效】
func (b *Bucket) Bucket(name []byte) *Bucket {
// 先看buckets缓存记录有没有,有则直接返回
if b.buckets != nil {
if child := b.buckets[string(name)]; child != nil {
return child
}
}
// 寻找bucket
c := b.Cursor()
k, v, flags := c.seek(name)
// 没找到 或者 找到的不是bucket,返回nil
if !bytes.Equal(name, k) || (flags&bucketLeafFlag) == 0 {
return nil
}
// 找到数据后将其转化成bucket,并且记录到buckets缓存中
var child = b.openBucket(v)
if b.buckets != nil {
b.buckets[string(name)] = child
}
return child
}
//openBucket 通过原始数据打开一个bucket
func (b *Bucket) openBucket(value []byte) *Bucket {
var child = newBucket(b.tx)
// 判断是否对齐
unaligned := brokenUnaligned && uintptr(unsafe.Pointer(&value[0]))&3 != 0
// value没有对齐则clone一个一摸一样的
if unaligned {
value = cloneBytes(value)
}
// 在读写tx中,将value深度拷贝到新bucket头部
// 在只读tx中,将新bucket头指向value即可
if b.tx.writable && !unaligned {
child.bucket = &bucket{}
*child.bucket = *(*bucket)(unsafe.Pointer(&value[0]))
} else {
child.bucket = (*bucket)(unsafe.Pointer(&value[0]))
}
// 如果bucket是内联的,则保存对内联页面的引用:即将新bucket的page字段指向value中内置page的起始位置
if child.root == 0 {
child.page = (*page)(unsafe.Pointer(&value[bucketHeaderSize]))
}
return &child
}
// CreateBucket 创建一个指定名称的bucket【仅在事务生存期间有效】
func (b *Bucket) CreateBucket(key []byte) (*Bucket, error) {
if b.tx.db == nil { // tx 关闭
return nil, ErrTxClosed
} else if !b.tx.writable { // 非可写tx
return nil, ErrTxNotWritable
} else if len(key) == 0 { // bucket name长度不符合要求
return nil, ErrBucketNameRequired
}
// 通过游标寻找bucket
c := b.Cursor()
k, _, flags := c.seek(key)
// 已存在该bucket则返回error
if bytes.Equal(key, k) {
if (flags & bucketLeafFlag) != 0 {
return nil, ErrBucketExists
}
return nil, ErrIncompatibleValue
}
// 创建一个空的bucket
var bucket = Bucket{
bucket: &bucket{},
rootNode: &node{isLeaf: true},
FillPercent: DefaultFillPercent,
}
var value = bucket.write()
// 将新bucket写入node
key = cloneBytes(key)
c.node().put(key, key, value, 0, bucketLeafFlag)
// 因为当前bucket已经包含了刚创建的新bucket,所以是非内联bucket,得将内联引用b.page置为nil
b.page = nil
// 这样查找一遍再返回,可以缓存该bucket,下次查起来更快
return b.Bucket(key), nil
}
// CreateBucketIfNotExists 创建一个指定名称的bucket,如果它还不存在的话【仅在事务生存期间有效】
func (b *Bucket) CreateBucketIfNotExists(key []byte) (*Bucket, error) {
child, err := b.CreateBucket(key)
if err == ErrBucketExists {
return b.Bucket(key), nil
} else if err != nil {
return nil, err
}
return child, nil
}
// DeleteBucket 删除指定bucket
func (b *Bucket) DeleteBucket(key []byte) error {
// tx 已关闭或非可写
if b.tx.db == nil {
return ErrTxClosed
} else if !b.Writable() {
return ErrTxNotWritable
}
// 查找该bucket
c := b.Cursor()
k, _, flags := c.seek(key)
// 不存在或者不是bucket,返回错误
if !bytes.Equal(key, k) {
return ErrBucketNotFound
} else if (flags & bucketLeafFlag) == 0 {
return ErrIncompatibleValue
}
// 递归删除其所有的子bucket
child := b.Bucket(key)
err := child.ForEach(func(k, v []byte) error {
if v == nil {
if err := child.DeleteBucket(k); err != nil {
return fmt.Errorf("delete bucket: %s", err)
}
}
return nil
})
if err != nil {
return err
}
// 删除该bucket在缓存中的信息
delete(b.buckets, string(key))
// 释放该bucket上的所有page 到 freelist
child.nodes = nil
child.rootNode = nil
child.free()
// 在node中删除该key
c.node().del(key)
return nil
}
// Get 查找此bucket中的kv对,同样:返回的值仅中事务生存期间有效
func (b *Bucket) Get(key []byte) []byte {
// 查找
k, v, flags := b.Cursor().seek(key)
// 找到的是bucket,返回nil
if (flags & bucketLeafFlag) != 0 {
return nil
}
// 没找到,同样返回nil
if !bytes.Equal(key, k) {
return nil
}
return v
}
// Put 添加一个kv对
func (b *Bucket) Put(key []byte, value []byte) error {
if b.tx.db == nil { // tx 关闭
return ErrTxClosed
} else if !b.Writable() { // tx 非可写
return ErrTxNotWritable
} else if len(key) == 0 { // key非法
return ErrKeyRequired
} else if len(key) > MaxKeySize { // key 非法
return ErrKeyTooLarge
} else if int64(len(value)) > MaxValueSize { // value 非法
return ErrValueTooLarge
}
// 查找key
c := b.Cursor()
k, _, flags := c.seek(key)
// 当前欲插入的key与当前bucket中已有一个子bucket的key相同时,拒绝写入,从而保护嵌套的子bucket的引用不会被擦除,防止子bucket变成孤儿
if bytes.Equal(key, k) && (flags&bucketLeafFlag) != 0 {
return ErrIncompatibleValue
}
//fmt.Printf("key %v 是否已存在: %v \n", string(key), bytes.Equal(key, k))
// key-value 插入node
key = cloneBytes(key) // 深度拷贝key
c.node().put(key, key, value, 0, 0)
return nil
}
// Delete 删除一个kv对
func (b *Bucket) Delete(key []byte) error {
if b.tx.db == nil { // tx关闭
return ErrTxClosed
} else if !b.Writable() { // tx 非可写
return ErrTxNotWritable
}
// 查找
c := b.Cursor()
_, _, flags := c.seek(key)
// 已存在,但是一个bucket
if (flags & bucketLeafFlag) != 0 {
return ErrIncompatibleValue
}
// 删除key
c.node().del(key)
return nil
}
// Sequence 返回bucket当前序列号
func (b *Bucket) Sequence() uint64 { return b.bucket.sequence }
// SetSequence 更新bucket序列号
func (b *Bucket) SetSequence(v uint64) error {
if b.tx.db == nil {
return ErrTxClosed
} else if !b.Writable() {
return ErrTxNotWritable
}
// 具体化根节点(如果还没有),以便在提交期间保存该bucket。
if b.rootNode == nil {
_ = b.node(b.root, nil)
}
// 更新序列号
b.bucket.sequence = v
return nil
}
// NextSequence 递增bucket序列号并返回
func (b *Bucket) NextSequence() (uint64, error) {
if b.tx.db == nil {
return 0, ErrTxClosed
} else if !b.Writable() {
return 0, ErrTxNotWritable
}
// 具体化根节点(如果还没有),以便在提交期间保存该bucket。
if b.rootNode == nil {
_ = b.node(b.root, nil)
}
b.bucket.sequence++
return b.bucket.sequence, nil
}
// ForEach 为bucket中的每一个kv对执行一个函数,函数返回错误则停止迭代
func (b *Bucket) ForEach(fn func(k, v []byte) error) error {
if b.tx.db == nil {
return ErrTxClosed
}
c := b.Cursor()
for k, v := c.First(); k != nil; k, v = c.Next() {
if err := fn(k, v); err != nil {
return err
}
}
return nil
}
// Stats 返回bucket的状态信息
func (b *Bucket) Stats() BucketStats {
var s, subStats BucketStats
pageSize := b.tx.db.pageSize
s.BucketN += 1 // bucket数量+1
if b.root == 0 {
s.InlineBucketN += 1 // 内联bucket数量+1
}
b.forEachPage(func(p *page, depth int) {
if (p.flags & leafPageFlag) != 0 { // 该page为 叶子节点类型的page
s.KeyN += int(p.count)
// 此page 使用的 总字节数
used := pageHeaderSize
if p.count != 0 {
// kv结构体头部大小 * kv数量
used += leafPageElementSize * int(p.count-1)
lastElement := p.leafPageElement(p.count - 1) // 最后一个kv对
used += int(lastElement.pos + lastElement.ksize + lastElement.vsize) // 最后一个kv对位置=之前所有kv对大小+最后一个kv对头部
}
if b.root == 0 { // 内联bucket
s.InlineBucketInuse += used
} else {
// 非内联bucket 叶子节点类型page 状态 更新(因为该page是叶子节点类型的page)
s.LeafPageN++
s.LeafInuse += used
s.LeafOverflowN += int(p.overflow)
// 从子bucket收集状态信息
for i := uint16(0); i < p.count; i++ {
e := p.leafPageElement(i)
if (e.flags & bucketLeafFlag) != 0 { // 该元素是子bucket
subStats.Add(b.openBucket(e.value()).Stats()) // 递归迭代
}
}
}
} else if (p.flags & branchPageFlag) != 0 { // 该page是 分支节点类型page
s.BranchPageN++
lastElement := p.branchPageElement(p.count - 1)
used := pageHeaderSize + (branchPageElementSize * int(p.count-1)) // page 头部大小+ page中所有元素的头部大小
used += int(lastElement.pos + lastElement.ksize) // 最后一个元素位置+ 最后一个元素key的大小 (分支节点类型的page 不存储value值,只提供索引功能,所以没有加上vsize)
s.BranchInuse += used
s.BranchOverflowN += int(p.overflow)
}
if depth+1 > s.Depth { // 记录最大深度
s.Depth = depth + 1
}
})
s.BranchAlloc = (s.BranchPageN + s.BranchOverflowN) * pageSize // (bucket的分支节点类型page数量 + bucket的分支节点类型page溢出数量)* page 大小
s.LeafAlloc = (s.LeafPageN + s.LeafOverflowN) * pageSize
// 加上子bucket的深度
s.Depth += subStats.Depth
// 加上子bucket状态
s.Add(subStats)
return s
}
//forEachPage 对 bucket 内的每一个page 执行fn
func (b *Bucket) forEachPage(fn func(*page, int)) {
// 内联page 直接执行fn
if b.page != nil {
fn(b.page, 0)
return
}
// 对每一个page 执行fn
b.tx.forEachPage(b.root, 0, fn)
}
//forEachPageNode 对 bucket内的每一个page或node 执行 fn
func (b *Bucket) forEachPageNode(fn func(*page, *node, int)) {
// 内联page or node,直接执行fn
if b.page != nil {
fn(b.page, nil, 0)
return
}
b._forEachPageNode(b.root, 0, fn)
}
func (b *Bucket) _forEachPageNode(pgid pgid, depth int, fn func(*page, *node, int)) {
var p, n = b.pageNode(pgid)
fn(p, n, depth)
// 子对象上递归执行
if p != nil {
if (p.flags & branchPageFlag) != 0 {
for i := 0; i < int(p.count); i++ {
elem := p.branchPageElement(uint16(i))
b._forEachPageNode(elem.pgid, depth+1, fn)
}
}
} else {
if !n.isLeaf {
for _, inode := range n.inodes {
b._forEachPageNode(inode.pgid, depth+1, fn)
}
}
}
}
//spill 将大小超过阈值的node 分解为多个node,避免引发频繁的再平衡操作
func (b *Bucket) spill() error {
// 遍历 子bucket
for name, child := range b.buckets {
//如果子bucket足够小并且没有子bucket,那么将其内联写入父bucket的页面。否则,将其像普通桶一样溢出,并使父值成为指向页面的指针。
var value []byte
if child.inlineable() { // 该子bucket是内联bucket:那么释放子bucket,将其写入到父bucket
child.free()
value = child.write()
} else {
// 该子bucket是普通bucket,同样进行spill操作
if err := child.spill(); err != nil {
return err
}
// 更新 子bucket的header
value = make([]byte, unsafe.Sizeof(bucket{}))
var bucket = (*bucket)(unsafe.Pointer(&value[0]))
*bucket = *child.bucket
}
// 没有具化成node 跳过
if child.rootNode == nil {
continue
}
//更新父节点
var c = b.Cursor()
k, _, flags := c.seek([]byte(name)) // 创建游标,寻找子bucket
if !bytes.Equal([]byte(name), k) { // 没有找到该子bucket
panic(fmt.Sprintf("misplaced bucket header: %x -> %x", []byte(name), k))
}
if flags&bucketLeafFlag == 0 {
panic(fmt.Sprintf("unexpected bucket header flag: %x", flags))
}
// 将子bucket的新value值更新到父bucket中
c.node().put([]byte(name), []byte(name), value, 0, bucketLeafFlag)
}
//如果没有具体化的根节点,则忽略。
if b.rootNode == nil {
return nil
}
// Spill nodes.
// spill所有子bucket后,开始spill自己:从rootNode开始,完成后更新一下rootNode
if err := b.rootNode.spill(); err != nil {
return err
}
b.rootNode = b.rootNode.root()
// 处理异常
if b.rootNode.pgid >= b.tx.meta.pgid {
panic(fmt.Sprintf("pgid (%d) above high water mark (%d)", b.rootNode.pgid, b.tx.meta.pgid))
}
// 更新bucket头中的根节点页号
b.root = b.rootNode.pgid
return nil
}
//inlineable 判断该bucket是否是内联bucket
func (b *Bucket) inlineable() bool {
// 通常情况下,父bucket中只保存了subbucket的bucket header,每个subbucket至少占据一个page,
// 若subbucket中的数据很少,这样会造成磁盘空间的浪费,所以可以将该subbucket做成inline bucket,
// 具体的做法是:将小的sunbucket的值完整的放在父bucket的leaf node上,从而减少占用的page数量)
var n = b.rootNode
// Bucket只能包含一个叶节点。
// n == nil || n是分支节点
if n == nil || !n.isLeaf {
return false
}
//如果Bucket包含子Bucket,或者超过了内联Bucket大小的阈值,则Bucket是不可内联的。
var size = pageHeaderSize
for _, inode := range n.inodes {
size += leafPageElementSize + len(inode.key) + len(inode.value)
if inode.flags&bucketLeafFlag != 0 { // 存在subbucket,非inline
return false
} else if size > b.maxInlineBucketSize() { // size 超过了inline bucket阈值,非inline
return false
}
}
return true
}
//maxInlineBucketSize 返回 inline bucket 阈值
func (b *Bucket) maxInlineBucketSize() int {
return b.tx.db.pageSize / 4
}
//write bucket 转 []byte:其实是将bucket写入page中
func (b *Bucket) write() []byte {
var n = b.rootNode // 只要将bucket 根节点写入page中就好了,其他节点可以通过索引找到
var value = make([]byte, bucketHeaderSize+n.size()) // 分配适当大小(就是bucketheader+根节点大小)
// 向value中写入一个bucket header
var bucket = (*bucket)(unsafe.Pointer(&value[0]))
*bucket = *b.bucket
// p是page指针,指向写入n的page
var p = (*page)(unsafe.Pointer(&value[bucketHeaderSize]))
n.write(p)
return value
}
//rebalance 再平衡操作:合并节点
func (b *Bucket) rebalance() {
// 先对bucket中的node 进行再平衡操作
for _, n := range b.nodes {
n.rebalance()
}
// 然后对子bucket递归调用再平衡
for _, child := range b.buckets {
child.rebalance()
}
}
//node 创建一个node,从指定的page和指定的父节点
func (b *Bucket) node(pgid pgid, parent *node) *node {
// 处理异常
_assert(b.nodes != nil, "nodes map expected")
// 判断要创建的节点是否已存在
if n := b.nodes[pgid]; n != nil {
return n
}
// 创建一个node
n := &node{bucket: b, parent: parent}
if parent == nil { // 如果其parent is nil,则将新建节点n置为bucket的根节点
b.rootNode = n
} else { // 在parent中缓存新建的n节点
parent.children = append(parent.children, n)
}
// 定位到对应page
var p = b.page
if p == nil {
p = b.tx.page(pgid)
}
// node n 读取 page 内容,并且记录在bucket的nodes信息中
n.read(p)
b.nodes[pgid] = n
// Update statistics.
b.tx.stats.NodeCount++
return n
}
//free 释放bucket中的page
func (b *Bucket) free() {
if b.root == 0 {
return
}
var tx = b.tx
b.forEachPageNode(func(p *page, n *node, _ int) {
if p != nil {
tx.db.freelist.free(tx.meta.txid, p) // 释放page
} else {
n.free() // 释放node:其实底层是将node 对应的page释放
}
})
b.root = 0
}
//dereference 解引用
func (b *Bucket) dereference() {
if b.rootNode != nil {
b.rootNode.root().dereference() // 根节点 解引用
}
for _, child := range b.buckets { // 子bucket,递归解引用
child.dereference()
}
}
//pageNode 根据pageID 查找对应page和node,node存在则先返回node,不存在则返回page
func (b *Bucket) pageNode(id pgid) (*page, *node) {
// inline buckeet,rootNode存在则返回rootNode,不存在则返回page
if b.root == 0 {
if id != 0 {
panic(fmt.Sprintf("inline bucket non-zero page access(2): %d != 0", id))
}
if b.rootNode != nil {
return nil, b.rootNode
}
return b.page, nil
}
// 从node cache中查找node,存在则返回
if b.nodes != nil {
if n := b.nodes[id]; n != nil {
return nil, n
}
}
// 没找到node,返回page
return b.tx.page(id), nil
}
//------------------------------------------------------ BucketStats -----------------------------------------------------------//
// BucketStats bucket状态统计
type BucketStats struct {
// page statistics
BranchPageN int // 分支节点类型的page 数
BranchOverflowN int // 分支节点类型page的 溢出数
LeafPageN int // 叶子节点类型page的 数
LeafOverflowN int // 叶子节点类型page的 数
// Tree statistics.
KeyN int // kv对的数量
Depth int // B+树深度
// Page size 利用率.
BranchAlloc int // 为 分支节点类型page 分配的字节数
BranchInuse int // 分支节点类型page 实际使用的字节数
LeafAlloc int // 为叶子节点类型page 分配的字节数
LeafInuse int // 叶子节点类型page 实际使用的字节数
// Bucket statistics
BucketN int // 总bucket数量:包括根bucket在内
InlineBucketN int // 内联bucket数量
InlineBucketInuse int // 内联bucket 使用的字节数
}
func (s *BucketStats) Add(other BucketStats) {
s.BranchPageN += other.BranchPageN
s.BranchOverflowN += other.BranchOverflowN
s.LeafPageN += other.LeafPageN
s.LeafOverflowN += other.LeafOverflowN
s.KeyN += other.KeyN
if s.Depth < other.Depth {
s.Depth = other.Depth
}
s.BranchAlloc += other.BranchAlloc
s.BranchInuse += other.BranchInuse
s.LeafAlloc += other.LeafAlloc
s.LeafInuse += other.LeafInuse
s.BucketN += other.BucketN
s.InlineBucketN += other.InlineBucketN
s.InlineBucketInuse += other.InlineBucketInuse
}
//cloneBytes 返回给定切片的副本
func cloneBytes(v []byte) []byte {
var clone = make([]byte, len(v))
copy(clone, v)
return clone
}
<file_sep>/errors.go
package bolt
import "errors"
// 在数据库上Open()或调用方法时,可能发生的错误
var (
// ErrDatabaseNotOpen db没有打开
ErrDatabaseNotOpen = errors.New("database not open")
// ErrDatabaseOpen db已打开
ErrDatabaseOpen = errors.New("database already open")
// ErrInvalid 无效db:db文件上的两个mate信息都无效
ErrInvalid = errors.New("invalid database")
// ErrVersionMismatch db和dbfile 版本不匹配
ErrVersionMismatch = errors.New("version mismatch")
// ErrChecksum 校验和不匹配
ErrChecksum = errors.New("checksum error")
// ErrTimeout 获取db独占锁超时
ErrTimeout = errors.New("timeout")
)
// 开始或提交事务时,可能发生的错误
var (
// ErrTxNotWritable 该事务非读写事务
ErrTxNotWritable = errors.New("tx not writable")
// ErrTxClosed 事务已关闭
ErrTxClosed = errors.New("tx closed")
// ErrDatabaseReadOnly 数据库是只读模式
ErrDatabaseReadOnly = errors.New("database is in read-only mode")
)
// These errors can occur when putting or deleting a value or a bucket.
var (
// ErrBucketNotFound 没有找到对应的bucket
ErrBucketNotFound = errors.New("bucket not found")
// ErrBucketExists bucket已存在
ErrBucketExists = errors.New("bucket already exists")
// ErrBucketNameRequired bucket name 不符合要求
ErrBucketNameRequired = errors.New("bucket name required")
// ErrKeyRequired bucket key name 不符合要求
ErrKeyRequired = errors.New("key required")
// ErrKeyTooLarge bucket key 太大
ErrKeyTooLarge = errors.New("key too large")
// ErrValueTooLarge bucket value 太大
ErrValueTooLarge = errors.New("value too large")
// ErrIncompatibleValue 非法操作
ErrIncompatibleValue = errors.New("incompatible value")
)
<file_sep>/db.go
package bolt
import (
"errors"
"fmt"
"hash/fnv"
"log"
"os"
"runtime"
"runtime/debug"
"strings"
"sync"
"time"
"unsafe"
)
//------------------------------------------------------ 常量值 -----------------------------------------------------------//
// 最大 mmp 步长
const maxMmapStep = 1 << 30 // 1GB
// 数据文件的格式版本
const version = 2
// 一个文件标记值,表示该文件是db file
const magic uint32 = 0xED0CDAED
// IgnoreNoSync 指定将更改同步到文件时是否忽略数据库的NoSync字段。这是必需的,因为某些操作系统(如OpenBSD)没有统一缓冲区缓存(UBC),必须使用msync(2)系统调用同步写入。
const IgnoreNoSync = runtime.GOOS == "openbsd"
// DB 默认值
const (
DefaultMaxBatchSize int = 1000 // 默认批处理最大size
DefaultMaxBatchDelay = 10 * time.Millisecond // 默认批处理最大延时
DefaultAllocSize = 16 * 1024 * 1024
)
// DB 默认page大小(操作系统page size)
var defaultPageSize = os.Getpagesize()
// Options 表示打开数据库时可以设置的选项
type Options struct {
//Timeout 等待获取文件锁的时间量,等于0则无限期等待
Timeout time.Duration
//NoGrowSync 非增长同步模式,是否启用
NoGrowSync bool
//ReadOnly 只读模式,是否启用
ReadOnly bool
// 在内存映射文件之前设置DB.MmapFlags标志。
MmapFlags int
//InitialMmapSize 数据库初始mmap大小
InitialMmapSize int
}
//DefaultOptions db默认配置
var DefaultOptions = &Options{
Timeout: 0,
NoGrowSync: false,
}
type Info struct {
Data uintptr
PageSize int
}
//------------------------------------------------------ DB -----------------------------------------------------------//
//DB 保存到磁盘上文件的存储桶的集合
type DB struct {
// 严格模式,启用后会在每次commit之后check,严重影响性能,应仅用于调试模式
StrictMode bool
// 非同步模式,启用后会在每次commit时跳过fsync函数的调用,这在批量加载数据时非常有用
NoSync bool
// 非增长同步模式,启用后会在数据库增长时跳过对 truncate 的调用
NoGrowSync bool
// If you want to read the entire database fast, you can set MmapFlag to
// syscall.MAP_POPULATE on Linux 2.6.23+ for sequential read-ahead.
MmapFlags int
// 批处理的最大大小,<=0则禁用批处理
MaxBatchSize int
// 批处理开始前的最大延时,<=0则禁用批处理
MaxBatchDelay time.Duration
//AllocSize 数据库需要创建新页面时分配的空间量。这样做是为了在增长数据文件时分摊truncate()和fsync()的成本。
AllocSize int
path string // db file 存储路径
file *os.File // 指向db file,文件描述符fd
lockfile *os.File // windows only
dataref []byte // mmap'ed readonly, write throws SEGV
data *[maxMapSize]byte // db data
datasz int // db data size
filesz int // current on disk file size
meta0 *meta // 元数据 0
meta1 *meta // 元数据 1
pageSize int // page size
opened bool // db 是否是 打开状态
rwtx *Tx // 当前读写事务,唯一!
txs []*Tx // 所有已打开的只读事务
freelist *freelist // 空闲page 列表
stats Stats // db 统计信息
pagePool sync.Pool // page 缓冲池
batchMu sync.Mutex // 批处理 锁
batch *batch // 批处理
rwlock sync.Mutex // 读写锁
metalock sync.Mutex // meta page 锁
mmaplock sync.RWMutex // mmap 锁
statlock sync.RWMutex // stats 锁
ops struct {
writeAt func(b []byte, off int64) (n int, err error)
}
// 只读模式
readOnly bool
}
// Path 返回当前打开的数据库文件路径
func (db *DB) Path() string {
return db.path
}
// GoString 返回数据库的字符串表示
func (db *DB) GoString() string {
return fmt.Sprintf("bolt.DB{path:%q}", db.path)
}
// String 返回数据库的字符串表示
func (db *DB) String() string {
return fmt.Sprintf("DB<%q>", db.path)
}
// Open 通过给定的路径,创建和打开一个数据库【如果文件不存在则自动创建,传入nil参数将使用默认值】
func Open(path string, mode os.FileMode, options *Options) (*DB, error) {
// 数据库标记为已打开
var db = &DB{opened: true}
// 初始化一些配置
if options == nil {
options = DefaultOptions
}
db.NoGrowSync = options.NoGrowSync
db.MmapFlags = options.MmapFlags
db.MaxBatchSize = DefaultMaxBatchSize
db.MaxBatchDelay = DefaultMaxBatchDelay
db.AllocSize = DefaultAllocSize
flag := os.O_RDWR
if options.ReadOnly {
flag = os.O_RDONLY
db.readOnly = true
}
// 打开数据文件,文件不存在则直接创建
db.path = path
var err error
if db.file, err = os.OpenFile(db.path, flag|os.O_CREATE, mode); err != nil {
_ = db.close()
return nil, err
}
// 锁定数据库文件,如果数据库是仅读模式,则是共享锁,否则是独占锁,保证其他boltdb进程无法获取
if err := flock(db, mode, !db.readOnly, options.Timeout); err != nil {
_ = db.close()
return nil, err
}
// 用于测试钩子函数
db.ops.writeAt = db.file.WriteAt
// 如果 db 不存在 ,初始化db
if info, err := db.file.Stat(); err != nil {
return nil, err
} else if info.Size() == 0 {
// Initialize new files with meta pages.
if err := db.init(); err != nil {
return nil, err
}
} else {
// 读取第一个mata page,用于确定page size
var buf [0x1000]byte
if _, err := db.file.ReadAt(buf[:], 0); err == nil {
m := db.pageInBuffer(buf[:], 0).meta()
// 如果无法读取第一个mata page,则用当前os 系统 page size
if err := m.validate(); err != nil {
db.pageSize = os.Getpagesize()
} else {
db.pageSize = int(m.pageSize)
}
}
}
// 初始化 page pool
db.pagePool = sync.Pool{
New: func() interface{} {
return make([]byte, db.pageSize)
},
}
// mmap db file
if err := db.mmap(options.InitialMmapSize); err != nil {
_ = db.close()
return nil, err
}
// 读入 freeList
db.freelist = newFreelist()
db.freelist.read(db.page(db.meta().freelist))
return db, nil
}
//mmap 打开 底层内存映射文件 并 初始化元page
func (db *DB) mmap(minsz int) error {
// 锁住,确保独占
db.mmaplock.Lock()
defer db.mmaplock.Unlock()
// 获取db file 基础信息(FileInfo 结构)
info, err := db.file.Stat()
if err != nil {
return fmt.Errorf("mmap stat error: %s", err)
} else if int(info.Size()) < db.pageSize*2 {
return fmt.Errorf("file size too small")
}
// 保证 size 下限
var size = int(info.Size())
if size < minsz {
size = minsz
}
// 得到真正的mmap size
size, err = db.mmapSize(size)
if err != nil {
return err
}
// 在 unmaping之前 把所有的mmap内容 解引用,拷贝到内存里
if db.rwtx != nil {
db.rwtx.root.dereference()
}
// unmap , 取消原有的db file 映射,如果有的话
if err := db.munmap(); err != nil {
return err
}
// 将磁盘上的 db file 映射到内存中,这样在内存中修改了db file 都会同步到磁盘,减少了内存的拷贝次数(文件的read和write会产生频繁的内存拷贝)
if err := mmap(db, size); err != nil {
return err
}
// 保存对 meta page 的引用:读取db file 的第0页和第1页来初始化db.mata0和db.mate1
db.meta0 = db.page(0).meta()
db.meta1 = db.page(1).meta()
// 验证mate page 信息是否正确
//(如果两个元页面都未通过验证,我们只会返回一个错误,因为 meta0 验证失败意味着它没有正确保存——但我们可以使用 meta1 进行恢复。反之亦然。)
err0 := db.meta0.validate()
err1 := db.meta1.validate()
if err0 != nil && err1 != nil {
return err0
}
return nil
}
// munmap 取消映射 db file
func (db *DB) munmap() error {
if err := munmap(db); err != nil {
return fmt.Errorf("unmap error: " + err.Error())
}
return nil
}
//mmapSize 根据传入的mmap size大小返回一个合适的mmap size 大小。最小大小为 32KB,并加倍直至达到 1GB。如果新的 mmap 大小大于允许的最大值,则返回错误。
func (db *DB) mmapSize(size int) (int, error) {
// 当 32KB <= size <= 1GB 时,直接翻倍返回
for i := uint(15); i <= 30; i++ {
if size <= 1<<i {
return 1 << i, nil
}
}
// 不允许超过最大值
if size > maxMapSize {
return 0, fmt.Errorf("mmap too large")
}
// size >= 1GB,每次增加到1GB的倍数,比如1.2GB 就增长到 2GB
sz := int64(size)
if remainder := sz % int64(maxMmapStep); remainder > 0 {
sz += int64(maxMmapStep) - remainder
}
// 确保 mmap 大小是页面大小的倍数。这应该总是正确的,因为我们以 MB 为单位递增。
pageSize := int64(db.pageSize)
if (sz % pageSize) != 0 {
sz = ((sz / pageSize) + 1) * pageSize
}
// 保证不超过最大size 上线
if sz > maxMapSize {
sz = maxMapSize
}
return int(sz), nil
}
//init 数据库初始化
func (db *DB) init() error {
// 前面四个page:| meta | meta | freelist | leafpage |
// 此时只包含一个空的叶子节点,该叶子节点即为root bucket的root node
// 将页面大小设置为操作系统的页面大小
db.pageSize = os.Getpagesize()
// 申请四个 page 空间
buf := make([]byte, db.pageSize*4)
// 在pageID 0和1 上创建两个元信息页
for i := 0; i < 2; i++ {
p := db.pageInBuffer(buf[:], pgid(i))
p.id = pgid(i)
p.flags = metaPageFlag
// 初始化元信息页
m := p.meta()
m.magic = magic // 魔数
m.version = version //版本号
m.pageSize = uint32(db.pageSize)
m.freelist = 2 // 指向freelist的pageID 2
m.root = bucket{root: 3} //指向bucket的pageID 3
m.pgid = 4 //指向正式数据的pageID 4
m.txid = txid(i) // 事务序列号
m.checksum = m.sum64() // check sum
}
// 在pageID 2 写入一个空的 freelist页
p := db.pageInBuffer(buf[:], pgid(2))
p.id = pgid(2)
p.flags = freelistPageFlag
p.count = 0
// 在pageID 4 写入一个空的 叶子节点页
p = db.pageInBuffer(buf[:], pgid(3))
p.id = pgid(3)
p.flags = leafPageFlag
p.count = 0
// 将buf中的数据写入文件,同时将磁盘缓冲页立即写入磁盘
if _, err := db.ops.writeAt(buf, 0); err != nil {
return err
}
if err := fdatasync(db); err != nil {
return err
}
return nil
}
//Close 关闭db,释放所有db资源:要求所有tx都已被关闭
func (db *DB) Close() error {
// 锁住所有需要竞争的资源
db.rwlock.Lock()
defer db.rwlock.Unlock()
db.metalock.Lock()
defer db.metalock.Unlock()
db.mmaplock.RLock()
defer db.mmaplock.RUnlock()
return db.close()
}
func (db *DB) close() error {
// db 标记为 关闭
if !db.opened {
return nil
}
db.opened = false
db.freelist = nil
// Clear ops.
db.ops.writeAt = nil
// Close the mmap.
if err := db.munmap(); err != nil {
return err
}
// Close file handles.
if db.file != nil {
// 只读文件无需 unlock
if !db.readOnly {
// Unlock the file.
if err := funlock(db); err != nil {
log.Printf("bolt.Close(): funlock error: %s", err)
}
}
// 关闭 fd(文件描述符)
if err := db.file.Close(); err != nil {
return fmt.Errorf("db file close: %s", err)
}
db.file = nil
}
db.path = ""
return nil
}
//Begin 开始一个新的事务。
func (db *DB) Begin(writable bool) (*Tx, error) {
// 可以同时使用多个只读事务,但一次只能使用一个写事务。
// 启动多个写事务将导致调用阻塞并序列化,直到当前写事务完成。
// 事务不应相互依赖。在同一个goroutine中打开读事务和写事务可能会导致写入程序死锁,因为数据库在增长时需要周期性地重新映射自身,而在读事务打开时不能这样做。
// 如果需要长时间运行的读取事务(例如,快照事务),则可能需要将DB.InitialMmapSize设置为足够大的值,以避免潜在的写入事务阻塞。
// 重要提示:完成后必须关闭只读事务,否则数据库将无法回收旧页。
if writable {
return db.beginRWTx()
}
return db.beginTx()
}
//beginTx 开始一个 只读事务
func (db *DB) beginTx() (*Tx, error) {
// 锁住 mata page
db.metalock.Lock()
// 在mmap上获取只读锁。
// 当mmap被重新映射时,它将获得一个写锁.
db.mmaplock.RLock()
// mmap被重新映射的两种情况:
// 1.db file文件在第一次创建或打开时
// 2.在写入db后且db file需要增大时,需要重新mmap将新的文件映射入进程的地址空间
// 具体情况分析:当db在不同线程中进行读写时,可能存在其中一个线程的读写事务写入了大量的数据,在commit时,由于已映射区的空闲page不够,
// 会重新进行mmap,此时若有未关闭的只读事务,由于此事务占用着mmap的读锁,那么mmap操作会阻塞在争用mmap写锁的地方
// 也就是说,如果存在着耗时的只读事务,同时写事务需要remmap时,写操作会被读操作阻塞
// 所以:在使用boltdb时,应尽量避免耗时的读操作,同时在写操作时,应避免频繁的remmap
// db 不是打开状态,return并释放锁
if !db.opened {
db.mmaplock.RUnlock()
db.metalock.Unlock()
return nil, ErrDatabaseNotOpen
}
// 创建一个与db关联的只读事务
t := &Tx{}
t.init(db)
// 将创建好的事务 加入到 db的仅读事务列表 中
db.txs = append(db.txs, t)
n := len(db.txs)
// 仅释放meta page 锁!
db.metalock.Unlock()
// 更新db 统计数据
db.statlock.Lock()
db.stats.TxN++
db.stats.OpenTxN = n
db.statlock.Unlock()
return t, nil
}
//beginRWTx 开始一个 读写事务
func (db *DB) beginRWTx() (*Tx, error) {
if db.readOnly {
return nil, ErrDatabaseReadOnly
}
// 读写锁,锁住
db.rwlock.Lock()
// 在获得写锁之后,锁住meta page,来修改meta
db.metalock.Lock()
defer db.metalock.Unlock()
// Exit if the database is not open yet.
if !db.opened {
db.rwlock.Unlock()
return nil, ErrDatabaseNotOpen
}
// 创建一个与db关联的读写事务
t := &Tx{writable: true}
t.init(db)
db.rwtx = t
// boltdb同时只能有一个读写事务,但是可以同时有多个只读事务!
// 在所有打开的只读事务中,找到最小的txid,释放该事务用到的page
// todo ???????????? 为什么要释放这个只读事务?
var minid txid = 0xFFFFFFFFFFFFFFFF
for _, t := range db.txs {
if t.meta.txid < minid {
minid = t.meta.txid
}
}
if minid > 0 {
db.freelist.release(minid - 1)
}
return t, nil
}
// removeTx 移除事务
func (db *DB) removeTx(tx *Tx) {
// 释放 mmap 上 的读锁
db.mmaplock.RUnlock()
// 锁住 meta page
db.metalock.Lock()
// 移除此事务
for i, t := range db.txs {
if t == tx {
// 将最后一个事务放到此事务的位置上,然后txs缩短一个长度,就算是删除了此事务
last := len(db.txs) - 1
db.txs[i] = db.txs[last]
db.txs[last] = nil
db.txs = db.txs[:last]
break
}
}
n := len(db.txs)
// Unlock the meta pages.
db.metalock.Unlock()
// Merge statistics.
db.statlock.Lock()
db.stats.OpenTxN = n
db.stats.TxStats.add(&tx.stats)
db.statlock.Unlock()
}
//Update 在读写管理事务的上下文中执行函数。如果函数未返回错误,则提交事务。如果返回错误,则回滚整个事务。函数返回或提交返回的任何错误都将从Update()方法返回。尝试在函数内手动提交或回滚将导致死机。
func (db *DB) Update(fn func(*Tx) error) error {
// 开启一个读写事务
t, err := db.Begin(true)
if err != nil {
return err
}
// 确保事务在紧急状态下回滚
defer func() {
if t.db != nil {
t.rollback()
}
}()
// 标记为托管发送,以便内部函数无法手动提交。
t.managed = true
// 如果函数返回错误,则回滚并返回错误。
err = fn(t)
t.managed = false
if err != nil {
_ = t.Rollback()
return err
}
// 提交此事务
return t.Commit()
}
//View 在只读事务的上下文中执行函数,函数返回的任何错误都将从View()方法返回。尝试在函数内手动回滚将导致死机
func (db *DB) View(fn func(*Tx) error) error {
// 开启一个只读事务
t, err := db.Begin(false)
if err != nil {
return err
}
// 确保事务在紧急情况下回滚
defer func() {
if t.db != nil {
t.rollback()
}
}()
// 标记为托管发送,以便内部功能无法手动回滚。
t.managed = true
// 执行 fn,错误则回滚
err = fn(t)
t.managed = false
if err != nil {
_ = t.Rollback()
return err
}
// 不需要调用Commit(),而是通过调用Rollback()来结束Transaction
if err := t.Rollback(); err != nil {
return err
}
return nil
}
//Batch 批处理
func (db *DB) Batch(fn func(*Tx) error) error {
// 批处理作为批处理的一部分调用fn。它的行为类似于更新,
// 只是:1.并发批处理调用可以组合到单个Bolt事务中。
// 2.传递给Batch的函数可能会被多次调用,无论它是否返回错误。这意味着批处理函数的副作用必须是幂等的,
// 并且只有在调用者中看到成功的返回后才会永久生效。最大批量大小和延迟可分别使用DB.MaxBatchSize和DB.MaxBatchDelay进行调整。批处理只有在有多个goroutine调用它时才有用。
errCh := make(chan error, 1)
// 获取 批处理锁
db.batchMu.Lock()
// 没有批处理任务 或者 批处理中任务数量已满,开启一个新的批处理
if (db.batch == nil) || (db.batch != nil && len(db.batch.calls) >= db.MaxBatchSize) {
db.batch = &batch{
db: db,
}
db.batch.timer = time.AfterFunc(db.MaxBatchDelay, db.batch.trigger)
}
db.batch.calls = append(db.batch.calls, call{fn: fn, err: errCh})
// 只有当批处理中任务数量达到一定要求,才执行批处理
if len(db.batch.calls) >= db.MaxBatchSize {
// wake up batch, it's ready to run
go db.batch.trigger()
}
// 释放批处理锁
db.batchMu.Unlock()
// 批处理产生错误,应该单独重新运行
err := <-errCh
if err == trySolo {
err = db.Update(fn)
}
return err
}
//Sync 针对数据库文件句柄执行fdatasync
//(在正常操作下,这是不必要的,但是,如果您使用NoSync,那么它允许您强制数据库文件与磁盘同步)
func (db *DB) Sync() error { return fdatasync(db) }
//Stats 检索数据库的持续性能统计信息。仅在事务关闭时更新。
func (db *DB) Stats() Stats {
db.statlock.RLock()
defer db.statlock.RUnlock()
return db.stats
}
//Info 返回db的Info信息
func (db *DB) Info() *Info {
return &Info{uintptr(unsafe.Pointer(&db.data[0])), db.pageSize}
}
//page 返回db中的特定page
func (db *DB) page(id pgid) *page {
pos := id * pgid(db.pageSize)
return (*page)(unsafe.Pointer(&db.data[pos]))
}
//pageInBuffer 从给定的[]byte 查找 指定page
func (db *DB) pageInBuffer(b []byte, id pgid) *page {
return (*page)(unsafe.Pointer(&b[id*pgid(db.pageSize)]))
}
// meta 返回db mate page
func (db *DB) meta() *meta {
// 返回具有最高txid的meta,这不会导致验证失败。否则,当数据库实际上处于一致状态时,我们可能会导致错误。
// metaA是txid较高的一种。
metaA := db.meta0
metaB := db.meta1
if db.meta1.txid > db.meta0.txid {
metaA = db.meta1
metaB = db.meta0
}
// Use higher meta page if valid. Otherwise fallback to previous, if valid.
if err := metaA.validate(); err == nil {
return metaA
} else if err := metaB.validate(); err == nil {
return metaB
}
// 这永远不应该达到,因为meta1和meta0都在mmap()上进行了验证,并且我们在每次写入时都执行fsync()。
panic("bolt.DB.meta(): invalid meta pages")
}
// allocate 分配给定数量的page
func (db *DB) allocate(count int) (*page, error) {
// 临时缓冲区
var buf []byte
// 如果需要的page数量只有一个,则从pagePool缓冲池中分配,以减少分配内存带来的时间开销
if count == 1 {
buf = db.pagePool.Get().([]byte)
} else {
buf = make([]byte, count*db.pageSize)
}
// buf转化为page,提供指针
p := (*page)(unsafe.Pointer(&buf[0]))
p.overflow = uint32(count - 1)
// 从freelist查看有没有可用的页号,如果有则分配给刚刚申请到的页缓存,并返回
if p.id = db.freelist.allocate(count); p.id != 0 {
return p, nil
}
// 将新申请的第一个page的ID 设置为文件内容结尾处的page ID
p.id = db.rwtx.meta.pgid
// 如果需要的page数量大于已经映射到内存的文件总page数,则触发remmap
var minsz = int((p.id+pgid(count))+1) * db.pageSize
if minsz >= db.datasz {
if err := db.mmap(minsz); err != nil {
return nil, fmt.Errorf("mmap allocate error: %s", err)
}
}
// 分析:如果打开db文件时,设定的初始文件映射长度足够长,
// 可以减少读写事务需要remmap的概率,从而降低读写事务被阻塞的概率(因为读写事务在remmap时,需要等待所有已经打开的只读事务的结束),提高读写并发
// 更新db已使用的最大page ID
db.rwtx.meta.pgid += pgid(count)
return p, nil
}
// grow db大小增长到给定sz
func (db *DB) grow(sz int) error {
// 给定sz比目前size小,忽略
if sz <= db.filesz {
return nil
}
// 如果数据小于alloc大小,则只分配所需的数据。一旦超过分配大小,则分块分配。
// todo ????????????
if db.datasz < db.AllocSize {
sz = db.datasz
} else {
sz += db.AllocSize
}
// Truncate and fsync to ensure file size metadata is flushed.
// https://github.com/boltdb/bolt/issues/284
// 同步增长模式 && db可写模式
if !db.NoGrowSync && !db.readOnly {
if runtime.GOOS != "windows" {
// 更改文件大小
if err := db.file.Truncate(int64(sz)); err != nil {
return fmt.Errorf("file resize error: %s", err)
}
}
// 安全起见,需要在更改文件大小后进行fsync
if err := db.file.Sync(); err != nil {
return fmt.Errorf("file sync error: %s", err)
}
}
db.filesz = sz
return nil
}
//IsReadOnly db是否只读模式
func (db *DB) IsReadOnly() bool {
return db.readOnly
}
//------------------------------------------------------ 批处理 -----------------------------------------------------------//
//call 函数封装
type call struct {
fn func(*Tx) error
err chan<- error
}
//batch 批处理封装
type batch struct {
db *DB
timer *time.Timer // 定时器
start sync.Once // 只执行一次操作的对象
calls []call
}
// trigger 触发批处理
func (b *batch) trigger() {
b.start.Do(b.run)
}
//run 执行批处理中的事务,并将结果传回DB.batch
func (b *batch) run() {
b.db.batchMu.Lock()
b.timer.Stop()
// 确保没有新工作添加到此批次,但不要中断其他批次。
if b.db.batch == b {
b.db.batch = nil
}
b.db.batchMu.Unlock()
retry:
for len(b.calls) > 0 {
var failIdx = -1
err := b.db.Update(func(tx *Tx) error {
for i, c := range b.calls {
if err := safelyCall(c.fn, tx); err != nil {
failIdx = i
return err
}
}
return nil
})
if failIdx >= 0 {
// 从批处理中取出失败的事务。在这里缩短b.calls是安全的,因为db.batch不再指向我们,而且我们仍然保持互斥。
c := b.calls[failIdx]
b.calls[failIdx], b.calls = b.calls[len(b.calls)-1], b.calls[:len(b.calls)-1]
// 告诉提交者重新单独运行,继续批处理的其余部分
c.err <- trySolo
continue retry
}
// 将成功或内部错误传递给所有调用者
for _, c := range b.calls {
c.err <- err
}
break retry
}
}
//trySolo 是一个特殊的sentinel错误值,用于发出重新运行事务功能的信号
var trySolo = errors.New("batch function returned an error and should be re-run solo")
type panicked struct {
reason interface{}
}
func (p panicked) Error() string {
if err, ok := p.reason.(error); ok {
return err.Error()
}
return fmt.Sprintf("panic: %v", p.reason)
}
func safelyCall(fn func(*Tx) error, tx *Tx) (err error) {
defer func() {
if p := recover(); p != nil {
err = panicked{p}
}
}()
return fn(tx)
}
//------------------------------------------------------ Stats -----------------------------------------------------------//
// Stats db 统计信息
type Stats struct {
// Freelist 统计信息
FreePageN int // freelist上 free page 数量
PendingPageN int // freelist上 pending page 数量
FreeAlloc int // free pages 中 分配的总字节数
FreelistInuse int // freelsit 使用的总字节数
// Transaction 统计信息
TxN int // 仅读事务 总数
OpenTxN int // 当前打开的仅读事务 数量
TxStats TxStats // 事务操作状态 统计
}
// Sub 计算并返回两组数据库统计数据之间的差异
func (s *Stats) Sub(other *Stats) Stats {
if other == nil {
return *s
}
var diff Stats
diff.FreePageN = s.FreePageN
diff.PendingPageN = s.PendingPageN
diff.FreeAlloc = s.FreeAlloc
diff.FreelistInuse = s.FreelistInuse
diff.TxN = s.TxN - other.TxN
diff.TxStats = s.TxStats.Sub(&other.TxStats)
return diff
}
func (s *Stats) add(other *Stats) {
s.TxStats.add(&other.TxStats)
}
//------------------------------------------------------ meta -----------------------------------------------------------//
// 元信息page加载到内存后的数据结构
type meta struct {
magic uint32 // 魔数
version uint32 // 数据文件版本号
pageSize uint32 // 该db的page大小,通过syscall.Getpagesize()获取,通常是4k
flags uint32
root bucket // 各个bucket的根组成的树
freelist pgid // 空闲列表存储的起始页ID
pgid pgid // 当前用到的最大page ID,也即用到的page数量
txid txid // 事务序列号
checksum uint64 // 校验和,用于校验元信息页是否完整
}
// validate 检查 mate page 信息是否正确
func (m *meta) validate() error {
if m.magic != magic {
return ErrInvalid
} else if m.version != version {
return ErrVersionMismatch
} else if m.checksum != 0 && m.checksum != m.sum64() {
return ErrChecksum
}
return nil
}
// copy copies one meta object to another.
func (m *meta) copy(dest *meta) {
*dest = *m
}
// write 元数据 写入 page(此page就成为了元page)
func (m *meta) write(p *page) {
// 保证没有超过 高水位线(当前使用的最大pageID)
if m.root.root >= m.pgid {
panic(fmt.Sprintf("root bucket pgid (%d) above high water mark (%d)", m.root.root, m.pgid))
} else if m.freelist >= m.pgid {
panic(fmt.Sprintf("freelist pgid (%d) above high water mark (%d)", m.freelist, m.pgid))
}
// 通过事务ID来确定Page ID,0或者1
p.id = pgid(m.txid % 2)
// 将page 设置为 mate page
p.flags |= metaPageFlag
// 计算 校验和
m.checksum = m.sum64()
// p的mate 指向 m
m.copy(p.meta())
}
//sum64 生成校验和
func (m *meta) sum64() uint64 {
var h = fnv.New64a()
_, _ = h.Write((*[unsafe.Offsetof(meta{}.checksum)]byte)(unsafe.Pointer(m))[:])
return h.Sum64()
}
//_assert 如果给定的条件为false,则断言将在给定的格式化消息中死机。
func _assert(condition bool, msg string, v ...interface{}) {
if !condition {
panic(fmt.Sprintf("assertion failed: "+msg, v...))
}
}
func warn(v ...interface{}) { fmt.Fprintln(os.Stderr, v...) }
func warnf(msg string, v ...interface{}) { fmt.Fprintf(os.Stderr, msg+"\n", v...) }
func printstack() {
stack := strings.Join(strings.Split(string(debug.Stack()), "\n")[2:], "\n")
fmt.Fprintln(os.Stderr, stack)
}
<file_sep>/bolt_arm64.go
// +build arm64
package bolt
// maxMapSize 表示 Bol 支持的最大 mmap 大小
const maxMapSize = 0xFFFFFFFFFFFF // 256TB
// maxAllocSize 创建数组指针时允许使用的最大大小
const maxAllocSize = 0x7FFFFFFF
// Are unaligned load/stores broken on this arch?
var brokenUnaligned = false
| ff1cc39e104e68d61cad47e339e250911d2b44b4 | [
"Markdown",
"Go Module",
"Go"
] | 11 | Go | hfdpx/boltdb | 7e20f4eb8d5ba25ac101b2c32c1a83d9df96cfa1 | 25f6c7805018ff962bee39d0fcf6681743186db4 |
refs/heads/master | <file_sep>import socks
import socket
socks.setdefaultproxy(socks.PROXY_TYPE_SOCKS5, "127.0.0.1", 9050)
socket.socket = socks.socksocket
import urllib2
#print(urllib2.urlopen("http://icanhazip.com").read())
<file_sep>import socks
import socket
socks.setdefaultproxy(socks.PROXY_TYPE_SOCKS5, "127.0.0.1", 9050)
socket.socket = socks.socksocket
import urllib2
url = "http://www.streetartutopia.com/wp-content/uploads/2012/11/Street-Art-by-David-Walker-in-London-England.jpg"
file_name = url.split('/')[-1]
u = urllib2.urlopen(url)
f = open(file_name, 'wb')
meta = u.info()
file_size = int(meta.getheaders("Content-Length")[0])
print "Downloading: %s Bytes: %s" % (file_name, file_size)
file_size_dl = 0
block_sz = 8192
while True:
buffer = u.read(block_sz)
if not buffer:
break
file_size_dl += len(buffer)
f.write(buffer)
status = r"%10d [%3.2f%%]" % (file_size_dl, file_size_dl * 100. / file_size)
status = status + chr(8)*(len(status)+1)
print status,
f.close()
print(urllib2.urlopen("http://www.ifconfig.me/ip").read())
<file_sep>import socks
import socket
socks.setdefaultproxy(socks.PROXY_TYPE_SOCKS5, "127.0.0.1", 9050)
socket.socket = socks.socksocket
import urllib2
print(urllib2.urlopen("http://www.wtfismyip.com").read())
<file_sep>import socksTor
print(urllib2.urlopen("http://icanhazip.com").read())
<file_sep>import urllib2
response = urllib2.urlopen('http://python.org/')
print "Response:", response
#print "URL:", response.geturl()
#print "This gets the code:", response.code
#print "the headers are:", response.info()
#print "Date:", response.info()['date']
#print "Server:", response.info()['server']
| 1757b39e78f545c75b82e33b3e266c8a6e572b8b | [
"Python"
] | 5 | Python | dot-Sean/PythonHacking | 2711e52110229e1fe061299bd27e30d0f9c157e6 | 8e80032612d02797242b6fa98ef3aa816281260b |
refs/heads/master | <repo_name>michele-deluca/ita.deluca.tools.ldap.framework<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/functions/dao/ModifyRequestDAO.java
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.functions.dao;
import java.util.ArrayList;
import java.util.List;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.ldap.framework.commons.bean.LDAPModification;
import ita.deluca.tools.ldap.framework.commons.bean.ModifyRequest;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.exception.LdapEntityDaoException;
/**
* This class contains all the function that allows interaction with the database via JPA.
*
* @author <NAME>
*/
public class ModifyRequestDAO
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( ModifyRequestDAO.class );
/**
* This method persist the object into the database.
*
* @param entity
* the object to persist.
* @return The persisted entity.
* @throws LdapEntityDaoException
* this exception was raised whenever an error occurred during the persist of the object.
*/
protected static ModifyRequest persist( final ModifyRequest entity ) throws LdapEntityDaoException
{
if ( ModifyRequestDAO.LOG.isDebugEnabled() )
{
ModifyRequestDAO.LOG.debug( "Persist the entity:\n" + entity );
}
final ModifyRequest persistedEntity = LDAPEntityDAO.getAlreadyPersistEntity( entity );
if ( persistedEntity != null )
{
return persistedEntity;
}
final List<LDAPModification> persistedList = new ArrayList<>();
for ( final LDAPModification obj : entity.getModifications() )
{
persistedList.add( LDAPModificationDAO.persist( obj ) );
}
entity.setModifications( persistedList );
return LDAPEntityDAO.persistEntity( entity );
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/EntryChangeNotificationControl.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1Element;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1OctetString;
/**
* This class defines the entry change notification control, which may be used in conjunction with the persistent search control to provide details on the type of change that occurred with an entry.
*
* @author <NAME>
*/
@Entity @Cacheable
public class EntryChangeNotificationControl extends LDAPControl
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The OID of the entry change notification control.
*/
public static final String ENTRY_CHANGE_NOTIFICATION_CONTROL_OID = "2.16.840.1.113730.3.4.7";
/**
* The change number associated with this change, if applicable.
*/
private int changeNumber;
/**
* The type of change that occurred to the entry.
*/
private int changeType;
/**
* The previous DN for this entry if the change was a modify DN.
*/
private String previousDN;
/**
* Creates a new entry change notification control.
*/
public EntryChangeNotificationControl()
{
super( EntryChangeNotificationControl.ENTRY_CHANGE_NOTIFICATION_CONTROL_OID );
}
/**
* Creates a new entry change notification control by decoding the provided value.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param controlValue
* The encoded value for this control.
* @throws ProtocolException
* If a problem occurs while decoding the control value.
*/
public EntryChangeNotificationControl( final boolean isCritical, final ASN1OctetString controlValue ) throws ProtocolException
{
super( EntryChangeNotificationControl.ENTRY_CHANGE_NOTIFICATION_CONTROL_OID, isCritical );
final ASN1Element[] sequenceElements;
try
{
sequenceElements = ASN1Element.decodeAsSequence( controlValue.getValue() ).getElements();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode entry change notification control value sequence", e );
}
if ( ( sequenceElements.length < 1 ) || ( sequenceElements.length > 3 ) )
{
throw new ProtocolException( "There must be between 1 and 3 elements in an entry change notification value sequence" );
}
try
{
this.changeType = sequenceElements[ 0 ].decodeAsEnumerated().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode change type from entry change notification control", e );
}
this.previousDN = null;
this.changeNumber = -1;
if ( this.changeType == PersistentSearchControl.CHANGE_TYPE_MODIFY_DN )
{
try
{
this.previousDN = sequenceElements[ 1 ].decodeAsOctetString().getStringValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode previous DN from entry change notification control", e );
}
if ( sequenceElements.length == 3 )
{
try
{
this.changeNumber = sequenceElements[ 2 ].decodeAsInteger().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode change number from entry change notification control", e );
}
}
} else
{
if ( sequenceElements.length == 2 )
{
try
{
this.changeNumber = sequenceElements[ 1 ].decodeAsInteger().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode change number from entry change notification control", e );
}
}
}
}
/**
* Creates a new entry change notification control.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param changeType
* The type of change that occurred to the entry.
* @param previousDN
* The previous DN of the entry if the change was a modify DN operation.
* @param changeNumber
* The change number associated with this change, if applicable. A negative value should be used to indicate that no change number is available.
*/
public EntryChangeNotificationControl( final boolean isCritical, final int changeType, final String previousDN, final int changeNumber )
{
super( EntryChangeNotificationControl.ENTRY_CHANGE_NOTIFICATION_CONTROL_OID, isCritical );
this.changeType = changeType;
this.previousDN = previousDN;
this.changeNumber = changeNumber;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final EntryChangeNotificationControl other = ( EntryChangeNotificationControl ) obj;
if ( this.changeNumber != other.changeNumber )
{
return false;
}
if ( this.changeType != other.changeType )
{
return false;
}
if ( this.previousDN == null )
{
if ( other.previousDN != null )
{
return false;
}
} else if ( !this.previousDN.equals( other.previousDN ) )
{
return false;
}
return true;
}
/**
* Retrieving the change number associated with this change, if applicable.
*
* @return The change number associated with this change, if applicable.
*/
public int getChangeNumber()
{
return this.changeNumber;
}
/**
* Retrieving the type of change that occurred to the entry.
*
* @return The type of change that occurred to the entry.
*/
public int getChangeType()
{
return this.changeType;
}
/**
* Retrieving the previous DN for this entry if the change was a modify DN.
*
* @return The previous DN for this entry if the change was a modify DN.
*/
public String getPreviousDN()
{
return this.previousDN;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + this.changeNumber;
result = ( prime * result ) + this.changeType;
result = ( prime * result ) + ( this.previousDN == null ? 0 : this.previousDN.hashCode() );
return result;
}
/**
* Setting the change number associated with this change, if applicable.
*
* @param changeNumber
* The change number associated with this change, if applicable.
*/
public void setChangeNumber( final int changeNumber )
{
this.changeNumber = changeNumber;
}
/**
* Setting the type of change that occurred to the entry.
*
* @param changeType
* The type of change that occurred to the entry.
*/
public void setChangeType( final int changeType )
{
this.changeType = changeType;
}
/**
* Setting the previous DN for this entry if the change was a modify DN.
*
* @param previousDN
* The previous DN for this entry if the change was a modify DN.
*/
public void setPreviousDN( final String previousDN )
{
this.previousDN = previousDN;
}
/**
* Retrieves a string representation of this control with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this control with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( "LDAP Entry Change Notification Control" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " OID: " ).append( this.getControlOID() ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Criticality: " ).append( this.isCritical() ).append( LDAPMessage.EOL );
final String changeTypeStr;
switch ( this.changeType )
{
case PersistentSearchControl.CHANGE_TYPE_ADD:
changeTypeStr = "add";
break;
case PersistentSearchControl.CHANGE_TYPE_DELETE:
changeTypeStr = "delete";
break;
case PersistentSearchControl.CHANGE_TYPE_MODIFY:
changeTypeStr = "modify";
break;
case PersistentSearchControl.CHANGE_TYPE_MODIFY_DN:
changeTypeStr = "modify DN";
break;
default:
changeTypeStr = "invalid change type (" + this.changeType + ')';
break;
}
buffer.append( indentBuf ).append( " Change Type: " ).append( this.changeType ).append( " (" ).append( changeTypeStr ).append( ')' ).append( LDAPMessage.EOL );
if ( this.changeType == PersistentSearchControl.CHANGE_TYPE_MODIFY_DN )
{
buffer.append( indentBuf ).append( " Previous DN: " ).append( this.previousDN ).append( LDAPMessage.EOL );
}
if ( this.changeNumber >= 0 )
{
buffer.append( indentBuf ).append( " Change Number: " ).append( this.changeNumber ).append( LDAPMessage.EOL );
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + this.changeNumber;
this.checksum = ( prime * this.checksum ) + this.changeType;
this.checksum = ( prime * this.checksum ) + ( this.previousDN == null ? 0 : this.previousDN.hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/LDAPMessage.java
package ita.deluca.tools.ldap.framework.commons.bean;
import java.util.ArrayList;
import java.util.Date;
import java.util.List;
import javax.persistence.Cacheable;
import javax.persistence.CascadeType;
import javax.persistence.Entity;
import javax.persistence.FetchType;
import javax.persistence.JoinTable;
import javax.persistence.ManyToMany;
import javax.persistence.ManyToOne;
import javax.persistence.Table;
import org.hibernate.annotations.Fetch;
import org.hibernate.annotations.FetchMode;
import ita.deluca.tools.application.framework.data.datetime.model.DateTimeFacilitators;
/**
* This class defines an LDAP message, which is the envelope that encompasses all communication using the LDAP protocol.
*
* @author <NAME>
*/
@Entity
@Cacheable
@Table( schema = "ldap_tools", name = "ldap_message" )
public class LDAPMessage extends LDAPEntity
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The end-of-line character for this platform.
*/
public static final String EOL = System.getProperty( "line.separator" );
/**
* The host where the capture was execute.
*/
private String captureHost;
/**
* The connection id.
*/
private String connectionId;
/**
* The set of controls associated with this LDAP message.
*/
@ManyToMany( fetch = FetchType.EAGER )
@JoinTable( schema = "ldap_tools", name = "ldap_message_controls_join" )
@Fetch( FetchMode.SELECT )
private List<LDAPControl> controls = new ArrayList<>();
/**
* The message destination host.
*/
private String destHost;
/**
* The message destination port.
*/
private int destPort;
/**
* The message ID for this LDAP message.
*/
private int messageID;
/**
* The message length.
*/
private int messageLength;
/**
* The protocol op for this LDAP message.
*/
@ManyToOne( fetch = FetchType.EAGER, cascade = CascadeType.ALL )
private ProtocolOp protocolOp;
/**
* The message source host.
*/
private String sourceHost;
/**
* The message source port.
*/
private int sourcePort;
/**
* The message timestamp.
*/
private Date timestamp;
/**
* Standard constructor.
*/
public LDAPMessage()
{
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( obj == null )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final LDAPMessage other = ( LDAPMessage ) obj;
if ( this.captureHost == null )
{
if ( other.captureHost != null )
{
return false;
}
} else if ( !this.captureHost.equals( other.captureHost ) )
{
return false;
}
if ( this.connectionId == null )
{
if ( other.connectionId != null )
{
return false;
}
} else if ( !this.connectionId.equals( other.connectionId ) )
{
return false;
}
if ( this.controls == null )
{
if ( other.controls != null )
{
return false;
}
} else if ( this.controls.size() != other.controls.size() )
{
return false;
} else
{
for ( final LDAPControl uniqueStr : this.controls )
{
if ( !other.controls.contains( uniqueStr ) )
{
return false;
}
}
}
if ( this.destHost == null )
{
if ( other.destHost != null )
{
return false;
}
} else if ( !this.destHost.equals( other.destHost ) )
{
return false;
}
if ( this.destPort != other.destPort )
{
return false;
}
if ( this.messageID != other.messageID )
{
return false;
}
if ( this.messageLength != other.messageLength )
{
return false;
}
if ( this.protocolOp == null )
{
if ( other.protocolOp != null )
{
return false;
}
} else if ( !this.protocolOp.equals( other.protocolOp ) )
{
return false;
}
if ( this.sourceHost == null )
{
if ( other.sourceHost != null )
{
return false;
}
} else if ( !this.sourceHost.equals( other.sourceHost ) )
{
return false;
}
if ( this.sourcePort != other.sourcePort )
{
return false;
}
if ( this.timestamp == null )
{
if ( other.timestamp != null )
{
return false;
}
} else if ( !this.timestamp.equals( other.timestamp ) )
{
return false;
}
return true;
}
/**
* Retrieves the host where the capture was execute.
*
* @return The host where the capture was execute.
*/
public String getCaptureHost()
{
return this.captureHost;
}
/**
* Retrieves the connection id.
*
* @return The connection id.
*/
public String getConnectionId()
{
return this.connectionId;
}
/**
* Retrieves the set of controls for this LDAP message.
*
* @return The set of controls for this LDAP message, or <CODE>null</CODE> if there are no controls.
*/
public List<LDAPControl> getControls()
{
return this.controls;
}
/**
* This method return the message destination host.
*
* @return the message destination host.
*/
public String getDestHost()
{
return this.destHost;
}
/**
* This method return the message destination port.
*
* @return the message destination port.
*/
public int getDestPort()
{
return this.destPort;
}
/**
* Retrieves the message ID for this LDAP message.
*
* @return The message ID for this LDAP message.
*/
public int getMessageID()
{
return this.messageID;
}
/**
* Retrieves the message length.
*
* @return The message length.
*/
public int getMessageLength()
{
return this.messageLength;
}
/**
* Retrieves the protocol op for this LDAP message.
*
* @return The protocol op for this LDAP message.
*/
public ProtocolOp getProtocolOp()
{
return this.protocolOp;
}
/**
* This method return the message source host.
*
* @return the message source host.
*/
public String getSourceHost()
{
return this.sourceHost;
}
/**
* This method return the message source port.
*
* @return the message source port.
*/
public int getSourcePort()
{
return this.sourcePort;
}
/**
* This method return the message datetime.
*
* @return the message datetime.
*/
public Date getTimestamp()
{
return this.timestamp;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = 1;
result = ( prime * result ) + ( this.captureHost == null ? 0 : this.captureHost.hashCode() );
result = ( prime * result ) + ( this.connectionId == null ? 0 : this.connectionId.hashCode() );
for ( final LDAPControl control : this.controls )
{
result += control.hashCode();
}
result = ( prime * result ) + ( this.destHost == null ? 0 : this.destHost.hashCode() );
result = ( prime * result ) + this.destPort;
result = ( prime * result ) + this.messageID;
result = ( prime * result ) + this.messageLength;
result = ( prime * result ) + ( this.protocolOp == null ? 0 : this.protocolOp.hashCode() );
result = ( prime * result ) + ( this.sourceHost == null ? 0 : this.sourceHost.hashCode() );
result = ( prime * result ) + this.sourcePort;
result = ( prime * result ) + ( this.timestamp == null ? 0 : DateTimeFacilitators.parseToString( this.timestamp, "yyyy-MM-dd HH:mm:ss.SSS" ).hashCode() );
return result;
}
/**
* Setting the host where the capture was execute.
*
* @param captureHost
* The host where the capture was execute.
*/
public void setCaptureHost( final String captureHost )
{
this.captureHost = captureHost;
}
/**
* Setting the connection id.
*
* @param connectionId
* The connection id.
*/
public void setConnectionId( final String connectionId )
{
this.connectionId = connectionId;
}
/**
* Settings the set of controls for this LDAP message.
*
* @param controls
* The set of controls for this LDAP message, or <CODE>null</CODE> if there are no controls.
*/
public void setControls( final List<LDAPControl> controls )
{
this.controls = controls;
}
/**
* This method set the message destination host.
*
* @param destHost
* he message destination host.
*/
public void setDestHost( final String destHost )
{
this.destHost = destHost;
}
/**
* This method set the message destination port.
*
* @param destPort
* the message destination port.
*/
public void setDestPort( final int destPort )
{
this.destPort = destPort;
}
/**
* Settings the message ID for this LDAP message.
*
* @param messageID
* The message ID for this LDAP message.
*/
public void setMessageID( final int messageID )
{
this.messageID = messageID;
}
/**
* Setting the message length.
*
* @param messageLength
* The message length.
*/
public void setMessageLength( final int messageLength )
{
this.messageLength = messageLength;
}
/**
* Setting the protocol op for this LDAP message.
*
* @param protocolOp
* The protocol op for this LDAP message.
*/
public void setProtocolOp( final ProtocolOp protocolOp )
{
this.protocolOp = protocolOp;
}
/**
* This method set the message source host.
*
* @param sourceHost
* the message source host.
*/
public void setSourceHost( final String sourceHost )
{
this.sourceHost = sourceHost;
}
/**
* This method set the message source port.
*
* @param sourcePort
* the message source port.
*/
public void setSourcePort( final int sourcePort )
{
this.sourcePort = sourcePort;
}
/**
* This method set the message timestamp.
*
* @param timestamp
* the message timestamp.
*/
public void setTimestamp( final Date timestamp )
{
this.timestamp = timestamp;
}
/**
* Retrieves a string representation of this LDAP message.
*
* @return A string representation of this LDAP message.
*/
@Override
public String toString()
{
return this.toString( 0 );
}
/**
* Retrieves a string representation of this LDAP message using the specified indent.
*
* @param indent
* The number of spaces to indent the message output.
* @return A string representation of this LDAP message.
*/
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( this.protocolOp.retrieveProtocolOpType() ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Message Capture Host: " ).append( this.captureHost ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Connection ID: " ).append( this.connectionId ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Message ID: " ).append( this.messageID ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Message Length: " ).append( this.messageLength ).append( " byte" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Message Timestamp: " ).append( DateTimeFacilitators.parseToString( this.timestamp, "yyyy-MM-dd HH:mm:ss.SSS" ) ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Message Source: " ).append( this.sourceHost ).append( ":" ).append( this.sourcePort ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Message Destination: " ).append( this.destHost ).append( ":" ).append( this.destPort ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " " ).append( this.protocolOp.retrieveProtocolOpType() ).append( " Protocol Op" ).append( LDAPMessage.EOL );
buffer.append( this.protocolOp.toString( indent + 8 ) );
if ( ( this.controls != null ) && !this.controls.isEmpty() )
{
for ( final LDAPControl control : this.controls )
{
buffer.append( control.toString( indent + 4 ) );
}
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + ( this.captureHost == null ? 0 : this.captureHost.hashCode() );
this.checksum = ( prime * this.checksum ) + ( this.connectionId == null ? 0 : this.connectionId.hashCode() );
for ( final LDAPControl control : this.controls )
{
control.updateChecksum();
this.checksum = ( prime * this.checksum ) + control.checksum;
}
this.checksum = ( prime * this.checksum ) + ( this.destHost == null ? 0 : this.destHost.hashCode() );
this.checksum = ( prime * this.checksum ) + this.destPort;
this.checksum = ( prime * this.checksum ) + this.messageID;
this.checksum = ( prime * this.checksum ) + this.messageLength;
if ( this.protocolOp != null )
{
this.protocolOp.updateChecksum();
this.checksum = ( prime * this.checksum ) + this.protocolOp.checksum;
}
this.checksum = ( prime * this.checksum ) + ( this.sourceHost == null ? 0 : this.sourceHost.hashCode() );
this.checksum = ( prime * this.checksum ) + this.sourcePort;
this.checksum = ( prime * this.checksum ) + ( this.timestamp == null ? 0 : DateTimeFacilitators.parseToString( this.timestamp, "yyyy-MM-dd HH:mm:ss.SSS" ).hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/edirectory/logininformation/model/DumpUserInformationFunctions.java
package ita.deluca.tools.ldap.framework.edirectory.logininformation.model;
import java.io.UnsupportedEncodingException;
import java.security.KeyManagementException;
import java.security.NoSuchAlgorithmException;
import java.security.NoSuchProviderException;
import java.security.Security;
import java.util.ArrayList;
import java.util.Arrays;
import java.util.Enumeration;
import java.util.Iterator;
import java.util.List;
import javax.net.ssl.SSLContext;
import javax.net.ssl.TrustManager;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import com.novell.ldap.LDAPAttribute;
import com.novell.ldap.LDAPAttributeSet;
import com.novell.ldap.LDAPConnection;
import com.novell.ldap.LDAPConstraints;
import com.novell.ldap.LDAPEntry;
import com.novell.ldap.LDAPException;
import com.novell.ldap.LDAPJSSEStartTLSFactory;
import com.novell.ldap.util.Base64;
import com.novell.security.nmas.mgmt.NMASPwdException;
import com.novell.security.nmas.mgmt.NMASPwdMgr;
import com.novell.security.nmas.mgmt.NMASPwdStatus;
import com.novell.security.nmas.mgmt.PwdJLdapTransport;
import com.sun.net.ssl.internal.ssl.Provider;
import ita.deluca.tools.application.framework.properties.exception.PropertiesException;
import ita.deluca.tools.application.framework.properties.model.PropertiesManager;
import ita.deluca.tools.ldap.framework.properties.Constants;
import ita.deluca.tools.ldap.framework.edirectory.bean.FakeX509TrustManager;
import ita.deluca.tools.ldap.framework.edirectory.bean.UserInformation;
import ita.deluca.tools.ldap.framework.edirectory.logininformation.exception.DumpUserInformationException;
/**
* This class dump the user information.
*
* @author <NAME>
*/
public class DumpUserInformationFunctions
{
/**
* The class logging.
*/
private static final Logger LOG = LogManager.getLogger( DumpUserInformationFunctions.class );
/**
* Extract the user information.
*
* @param environment
* The environment where user is inserted.
* @param userDn
* The user fullDN
* @return The user information.
* @throws DumpUserInformationException
* This exception was raised whenever an error occurred during the extraction of user information.
*/
public static UserInformation dumpUser( final String environment, final String userDn ) throws DumpUserInformationException
{
DumpUserInformationFunctions.LOG.info( "Extract information of user '" + userDn + "'" );
Security.addProvider( new Provider() );
final TrustManager[] tms = { new FakeX509TrustManager() };
SSLContext context = null;
try
{
context = SSLContext.getInstance( "TLS", "SunJSSE" );
context.init( null, tms, null );
} catch ( NoSuchProviderException | NoSuchAlgorithmException | KeyManagementException ex )
{
final String msg = "!!! It was impossible to create a TLS context !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
LDAPConnection.setSocketFactory( new LDAPJSSEStartTLSFactory( context.getSocketFactory() ) );
PropertiesManager pMgr;
try
{
pMgr = PropertiesManager.getPropertiesInstance( Constants.LDAP_TOOL_PROPERTIES_PATH );
} catch ( final PropertiesException ex )
{
final String msg = "!!! Error in retrieves of the properties manager !!!";
DumpUserInformationFunctions.LOG.error( msg );
throw new DumpUserInformationException( msg, ex );
}
final String server = pMgr.getPropertiesStringValue( "commons", environment, "server", "master" );
final int port = pMgr.getPropertiesIntegerValue( "commons", environment, "server", "port" );
final String username = pMgr.getPropertiesStringValue( "commons", environment, "adminuser", "username" );
final String password = pMgr.getPropertiesStringValue( "commons", environment, "adminuser", "password" );
final UserInformation userInfo = new UserInformation();
final LDAPConnection ldap = new LDAPConnection();
final LDAPConstraints cons = new LDAPConstraints();
cons.setTimeLimit( 30000 );
try
{
DumpUserInformationFunctions.LOG.debug( "Initialize ldap connection." );
ldap.connect( server, port );
ldap.startTLS();
ldap.bind( LDAPConnection.LDAP_V3, username, password.getBytes( "Utf8" ), cons );
} catch ( final LDAPException | UnsupportedEncodingException ex )
{
final String msg = "!!! The bind operation [server: " + server + " -- username:" + username + "] was ended in error !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
LDAPAttributeSet attrSet = null;
final String[] returnAttrs = { LDAPConnection.ALL_USER_ATTRS, "pwdChangedTime" };
try
{
DumpUserInformationFunctions.LOG.debug( "Extract user information : " + Arrays.toString( returnAttrs ) + "." );
final LDAPEntry entry = ldap.read( userDn, returnAttrs );
attrSet = entry.getAttributeSet();
} catch ( final LDAPException ex )
{
final String msg = "!!! It was impossible to read the login attribute of the user '" + userDn + "' !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
userInfo.setDn( userDn );
final Iterator<?> allAttributes = attrSet.iterator();
while ( allAttributes.hasNext() )
{
final LDAPAttribute attribute = ( LDAPAttribute ) allAttributes.next();
final List<String> attrValueList = new ArrayList<>();
final Enumeration<?> allValues = attribute.getStringValues();
if ( allValues != null )
{
while ( allValues.hasMoreElements() )
{
final String attrValue = ( String ) allValues.nextElement();
attrValueList.add( Base64.isLDIFSafe( attrValue ) ? attrValue : Base64.encode( attrValue.getBytes() ) );
}
}
userInfo.getAttributeMap().put( attribute.getName(), attrValueList );
}
final NMASPwdMgr pwdMgr = new NMASPwdMgr( new PwdJLdapTransport( ldap ) );
try
{
DumpUserInformationFunctions.LOG.debug( "Extract the password." );
userInfo.setPassword( pwdMgr.getPwd( "", userDn ) );
} catch ( final NMASPwdException ex )
{
String msg = "";
if ( ex.getNmasRetCode() == -16049 )
{
msg = "!!! The user '" + userDn + "' has no Universal Password value !!!";
} else if ( ex.getNmasRetCode() == -1659 )
{
msg = "!!! Requester does not have sufficient rights to perform operation !!!";
} else if ( ex.getNmasRetCode() == -1697 )
{
msg = "!!! NMAS is enabled on the server but Universal Password is not enabled for the user '" + userDn + "' !!!";
} else
{
msg = "!!! It was impossible to retrieves the password of the user '" + userDn + "' !!!";
}
throw new DumpUserInformationException( msg, ex );
}
try
{
DumpUserInformationFunctions.LOG.debug( "Extract the password policy." );
userInfo.setPasswordPolicy( pwdMgr.getPwdPolicyDN( "", userDn ) );
} catch ( final NMASPwdException ex )
{
final String msg = "!!! It was impossible to read the password policy of the user '" + userDn + "' !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
DumpUserInformationFunctions.LOG.debug( "Extract the password information." );
try
{
userInfo.setPwdRespectPolicy( pwdMgr.pwdPolicyCheck( "", userDn, null ) == 0 ? true : false );
} catch ( final NMASPwdException ex )
{
final String msg = "!!! It was impossible to test if the password of user '" + userDn + "' respect the password policy !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
NMASPwdStatus nmasPwdStatus = null;
try
{
nmasPwdStatus = pwdMgr.getPwdStatus( "", userDn );
userInfo.setUniversalPasswordEnabled( nmasPwdStatus.isSpmUpwdEnabled() );
userInfo.setUniversalPasswordMatchNdsPassword( nmasPwdStatus.isSpmUpwdMatchesNDS() );
userInfo.setUniversalPasswordMatchSimplePassword( nmasPwdStatus.isSpmUpwdMatchesSPWD() );
userInfo.setUniversalPasswordOlderThanNdsPassword( nmasPwdStatus.isSpmUpwdOlderThanNDS() );
} catch ( final NMASPwdException ex )
{
final String msg = "!!! It was impossible to retrieves the password status of user '" + userDn + "' !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
try
{
ldap.disconnect();
} catch ( final LDAPException ex )
{
final String msg = "!!! The disconnection to the ldap server throw an error !!!";
DumpUserInformationFunctions.LOG.error( msg, ex );
throw new DumpUserInformationException( msg, ex );
}
DumpUserInformationFunctions.LOG.debug( "User information:\n" + userInfo );
return userInfo;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/LDAPControl.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import javax.persistence.Inheritance;
import javax.persistence.InheritanceType;
import javax.persistence.Table;
/**
* This class defines an LDAP control, which provides additional information that may be used when processing an LDAP operation.
*
* @author <NAME>
*/
@Entity
@Cacheable
@Table( schema = "ldap_tools", name = "ldap_control" )
@Inheritance( strategy = InheritanceType.JOINED )
public abstract class LDAPControl extends LDAPEntity
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The ASN.1 type that should be used for a sequence of controls.
*/
public static final byte CONTROL_SEQUENCE_TYPE = ( byte ) 0xA0;
/**
* The OID for this LDAP control.
*/
private String controlOID;
/**
* Indicates whether this control should be considered critical.
*/
private boolean critical;
/**
* Creates a new LDAP control with the specified OID. It will not be critical and will not have a value.
*
* @param controlOID
* The OID for this control.
*/
LDAPControl( final String controlOID )
{
this.controlOID = controlOID;
this.critical = false;
}
/**
* Creates a new LDAP control with the specified OID, criticality, and value.
*
* @param controlOID
* The OID for this control.
* @param critical
* Indicates whether this control should be marked critical.
*/
LDAPControl( final String controlOID, final boolean critical )
{
this.controlOID = controlOID;
this.critical = critical;
}
/**
* Standard constructor.
*/
public LDAPControl()
{
}
/**
* Retrieves the OID for this control.
*
* @return The OID for this control.
*/
String getControlOID()
{
return this.controlOID;
}
/**
* Indicates whether this control is marked critical.
*
* @return <CODE>true</CODE> if this control is marked critical, or <CODE>false</CODE> if not.
*/
boolean isCritical()
{
return this.critical;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( obj == null )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final LDAPControl other = ( LDAPControl ) obj;
if ( this.controlOID == null )
{
if ( other.controlOID != null )
{
return false;
}
} else if ( !this.controlOID.equals( other.controlOID ) )
{
return false;
}
return this.critical == other.critical;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = 1;
result = ( prime * result ) + ( this.controlOID == null ? 0 : this.controlOID.hashCode() );
result = ( prime * result ) + ( this.critical ? 1231 : 1237 );
return result;
}
/**
* Setting the OID for this control.
*
* @param controlOID
* The OID for this control.
*/
public void setControlOID( final String controlOID )
{
this.controlOID = controlOID;
}
/**
* Settings whether this control is marked critical.
*
* @param critical
* <CODE>true</CODE> if this control is marked critical, or <CODE>false</CODE> if not.
*/
public void setCritical( final boolean critical )
{
this.critical = critical;
}
/**
* Retrieves a string representation of this control.
*
* @return A string representation of this control.
*/
@Override
public String toString()
{
return this.toString( 0 );
}
/**
* Retrieves a string representation of this control with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this control with the specified indent.
*/
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
return String.valueOf( indentBuf ) + "LDAP Control" + LDAPMessage.EOL + indentBuf + " OID: " + this.controlOID + LDAPMessage.EOL + indentBuf + " Criticality: " + this.critical + LDAPMessage.EOL;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/BindRequest.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
/**
* This class defines an LDAP bind request, which is used to authenticate to a directory server.
*
* @author <NAME>
*/
@Entity @Cacheable
public class BindRequest extends ProtocolOp
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The authentication type that indicates that SASL authentication is to be performed.
*/
public static final byte AUTH_TYPE_SASL = ( byte ) 0xA3;
/**
* The authentication type that indicates that simple authentication is to be performed.
*/
public static final byte AUTH_TYPE_SIMPLE = ( byte ) 0x80;
/**
* The authentication type contained in this bind request.
*/
private byte authType;
/**
* The base DN of the user performing the bind.
*/
private String baseDN;
/**
* /** The DN of the user performing the bind.
*/
private String bindDN;
/**
* The user password.
*/
private String bindPassword;
/**
* The LDAP protocol version contained in this bind request.
*/
private int protocolVersion;
/**
* The SASL credentials for this bind request.
*/
private String saslCredentials;
/**
* The SASL mechanism used for this bind request.
*/
private String saslMechanism;
/**
* Creates a new bind request using simple authentication with the provided information.
*/
public BindRequest()
{
this.authType = BindRequest.AUTH_TYPE_SIMPLE;
}
/**
* Creates a new bind request using simple authentication with the provided information.
*
* @param protocolVersion
* The LDAP protocol version to use in the bind request.
* @param baseDN
* The base DN of the user performing the bind.
* @param bindDN
* The DN of the user performing the bind.
* @param bindPassword
* The user <PASSWORD>.
*/
public BindRequest( final int protocolVersion, final String baseDN, final String bindDN, final String bindPassword )
{
this.protocolVersion = protocolVersion;
this.baseDN = baseDN;
this.bindDN = bindDN;
this.bindPassword = <PASSWORD>;
this.authType = BindRequest.AUTH_TYPE_SIMPLE;
}
/**
* Creates a new bind request using SASL authentication with the provided information.
*
* @param protocolVersion
* The LDAP protocol version to use in the bind request.
* @param baseDN
* The base DN of the user performing the bind.
* @param bindDN
* The DN of the user performing the bind.
* @param saslMechanism
* The SASL mechanism used to perform the bind.
* @param saslCredentials
* The SASL credentials to use in the bind.
*/
public BindRequest( final int protocolVersion, final String baseDN, final String bindDN, final String saslMechanism, final String saslCredentials )
{
this.protocolVersion = protocolVersion;
this.baseDN = baseDN;
this.bindDN = bindDN;
this.saslMechanism = saslMechanism;
this.saslCredentials = saslCredentials;
this.authType = BindRequest.AUTH_TYPE_SASL;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final BindRequest other = ( BindRequest ) obj;
if ( this.authType != other.authType )
{
return false;
}
if ( this.baseDN == null )
{
if ( other.baseDN != null )
{
return false;
}
} else if ( !this.baseDN.equals( other.baseDN ) )
{
return false;
}
if ( this.bindDN == null )
{
if ( other.bindDN != null )
{
return false;
}
} else if ( !this.bindDN.equals( other.bindDN ) )
{
return false;
}
if ( this.bindPassword == null )
{
if ( other.bindPassword != null )
{
return false;
}
} else if ( !this.bindPassword.equals( other.bindPassword ) )
{
return false;
}
if ( this.protocolVersion != other.protocolVersion )
{
return false;
}
if ( this.saslCredentials == null )
{
if ( other.saslCredentials != null )
{
return false;
}
} else if ( !this.saslCredentials.equals( other.saslCredentials ) )
{
return false;
}
if ( this.saslMechanism == null )
{
if ( other.saslMechanism != null )
{
return false;
}
} else if ( !this.saslMechanism.equals( other.saslMechanism ) )
{
return false;
}
return true;
}
/**
* Retrieves the type of authentication contained in this bind request.
*
* @return <CODE>AUTH_TYPE_SIMPLE</CODE> if simple authentication is to be performed, or <CODE>AUTH_TYPE_SASL</CODE> if SASL authentication should be used.
*/
public byte getAuthType()
{
return this.authType;
}
/**
* Retrieves the base DN of the user performing the bind.
*
* @return The base DN of the user performing the bind.
*/
public String getBaseDN()
{
return this.baseDN;
}
/**
* Retrieves the DN of the user performing the bind.
*
* @return The DN of the user performing the bind.
*/
public String getBindDN()
{
return this.bindDN;
}
/**
* Retrieves the user password.
*
* @return The user password.
*/
public String getBindPassword()
{
return this.bindPassword;
}
/**
* Retrieves the LDAP protocol version used in this bind request.
*
* @return The LDAP protocol version used in this bind request.
*/
public int getProtocolVersion()
{
return this.protocolVersion;
}
/**
* Retrieves the credentials used for SASL authentication.
*
* @return The credentials used for SASL authentication.
*/
public String getSaslCredentials()
{
return this.saslCredentials;
}
/**
* Retrieves the mechanism used for SASL authentication.
*
* @return The mechanism used for SASL authentication.
*/
public String getSaslMechanism()
{
return this.saslMechanism;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + this.authType;
result = ( prime * result ) + ( ( this.baseDN == null ) ? 0 : this.baseDN.hashCode() );
result = ( prime * result ) + ( ( this.bindDN == null ) ? 0 : this.bindDN.hashCode() );
result = ( prime * result ) + ( ( this.bindPassword == null ) ? 0 : this.bindPassword.hashCode() );
result = ( prime * result ) + this.protocolVersion;
result = ( prime * result ) + ( ( this.saslCredentials == null ) ? 0 : this.saslCredentials.hashCode() );
result = ( prime * result ) + ( ( this.saslMechanism == null ) ? 0 : this.saslMechanism.hashCode() );
return result;
}
/**
* Retrieves a user-friendly name for this protocol op.
*
* @return A user-friendly name for this protocol op.
*/
@Override
public String retrieveProtocolOpType()
{
return "LDAP Bind Request";
}
/**
* Setting the type of authentication contained in this bind request.
*
* @param authType
* <CODE>AUTH_TYPE_SIMPLE</CODE> if simple authentication is to be performed, or <CODE>AUTH_TYPE_SASL</CODE> if SASL authentication should be used.
*/
public void setAuthType( final byte authType )
{
this.authType = authType;
}
/**
* Setting the base DN of the user performing the bind.
*
* @param baseDN
* The base DN of the user performing the bind.
*/
public void setBaseDN( final String baseDN )
{
this.baseDN = baseDN;
}
/**
* Setting the DN of the user performing the bind.
*
* @param bindDN
* The DN of the user performing the bind.
*/
public void setBindDN( final String bindDN )
{
this.bindDN = bindDN;
}
/**
* Setting the user password.
*
* @param bindPassword
* The user password.
*/
public void setBindPassword( final String bindPassword )
{
this.bindPassword = bindPassword;
}
/**
* Setting the LDAP protocol version used in this bind request.
*
* @param protocolVersion
* The LDAP protocol version used in this bind request.
*/
public void setProtocolVersion( final int protocolVersion )
{
this.protocolVersion = protocolVersion;
}
/**
* Setting the credentials used for SASL authentication.
*
* @param saslCredentials
* The credentials used for SASL authentication.
*/
public void setSaslCredentials( final String saslCredentials )
{
this.saslCredentials = saslCredentials;
}
/**
* Setting the mechanism used for SASL authentication.
*
* @param saslMechanism
* The mechanism used for SASL authentication.
*/
public void setSaslMechanism( final String saslMechanism )
{
this.saslMechanism = saslMechanism;
}
/**
* Retrieves a string representation of this protocol op with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this protocol op with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( "LDAP Version: " ).append( this.protocolVersion ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( "Base DN: " ).append( this.baseDN ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( "Bind DN: " ).append( this.bindDN ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( "Password : " ).append( this.bindPassword ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( "Authentication Data:" ).append( LDAPMessage.EOL );
if ( this.authType == BindRequest.AUTH_TYPE_SIMPLE )
{
buffer.append( indentBuf ).append( " Authentication Type: Simple" ).append( LDAPMessage.EOL );
} else if ( this.authType == BindRequest.AUTH_TYPE_SASL )
{
buffer.append( indentBuf ).append( " Authentication Type: SASL" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " SASL Mechanism: " ).append( this.saslMechanism ).append( LDAPMessage.EOL );
if ( this.saslCredentials != null )
{
buffer.append( indentBuf ).append( " SASL Credentials:" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " " ).append( this.saslCredentials );
}
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + this.authType;
this.checksum = ( prime * this.checksum ) + ( ( this.baseDN == null ) ? 0 : this.baseDN.hashCode() );
this.checksum = ( prime * this.checksum ) + ( ( this.bindDN == null ) ? 0 : this.bindDN.hashCode() );
this.checksum = ( prime * this.checksum ) + ( ( this.bindPassword == null ) ? 0 : this.bindPassword.hashCode() );
this.checksum = ( prime * this.checksum ) + this.protocolVersion;
this.checksum = ( prime * this.checksum ) + ( ( this.saslCredentials == null ) ? 0 : this.saslCredentials.hashCode() );
this.checksum = ( prime * this.checksum ) + ( ( this.saslMechanism == null ) ? 0 : this.saslMechanism.hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/IntermediateResponse.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
/**
* This class defines an LDAP extended response, which provides information about the result of processing an intermediate response.
*
* @author <NAME>
*/
@Entity @Cacheable
public class IntermediateResponse extends ProtocolOp
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The ASN.1 type that should be used to encode a response OID.
*/
public static final byte RESPONSE_OID_TYPE = ( byte ) 0x80;
/**
* The ASN.1 type that should be used to encode a response value.
*/
public static final byte RESPONSE_VALUE_TYPE = ( byte ) 0x81;
/**
* The OID for the extended response.
*/
private String responseOID;
/**
* The value for the extended response.
*/
private String responseValue;
/**
* Creates a new intermediate response with the provided information.
*/
public IntermediateResponse()
{
}
/**
* Creates a new intermediate response with the provided information.
*
* @param responseOID
* The OID for this intermediate response.
* @param responseValue
* The value for this intermediate response.
*/
public IntermediateResponse( final String responseOID, final String responseValue )
{
this.responseOID = responseOID;
this.responseValue = responseValue;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final IntermediateResponse other = ( IntermediateResponse ) obj;
if ( this.responseOID == null )
{
if ( other.responseOID != null )
{
return false;
}
} else if ( !this.responseOID.equals( other.responseOID ) )
{
return false;
}
if ( this.responseValue == null )
{
if ( other.responseValue != null )
{
return false;
}
} else if ( !this.responseValue.equals( other.responseValue ) )
{
return false;
}
return true;
}
/**
* Retrieves the OID for this extended response.
*
* @return The OID for this extended response.
*/
public String getResponseOID()
{
return this.responseOID;
}
/**
* Retrieves the value for this extended response.
*
* @return The value for this extended response.
*/
public String getResponseValue()
{
return this.responseValue;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + ( this.responseOID == null ? 0 : this.responseOID.hashCode() );
result = ( prime * result ) + ( this.responseValue == null ? 0 : this.responseValue.hashCode() );
return result;
}
/**
* Retrieves a user-friendly name for this protocol op.
*
* @return A user-friendly name for this protocol op.
*/
@Override
public String retrieveProtocolOpType()
{
return "LDAP Intermediate Response";
}
/**
* Setting the OID for this extended response.
*
* @param responseOID
* The OID for this extended response.
*/
public void setResponseOID( final String responseOID )
{
this.responseOID = responseOID;
}
/**
* Setting the value for this extended response.
*
* @param responseValue
* The value for this extended response.
*/
public void setResponseValue( final String responseValue )
{
this.responseValue = responseValue;
}
/**
* Retrieves a string representation of this protocol op with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this protocol op with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
if ( this.responseOID != null )
{
buffer.append( indentBuf ).append( "Response OID: " ).append( this.responseOID ).append( LDAPMessage.EOL );
}
if ( this.responseValue != null )
{
buffer.append( indentBuf ).append( "Response Value:" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " " ).append( this.responseValue );
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + ( this.responseOID == null ? 0 : this.responseOID.hashCode() );
this.checksum = ( prime * this.checksum ) + ( this.responseValue == null ? 0 : this.responseValue.hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/META-INF/Create_LdapAttribute.sql
DROP TABLE IF EXISTS ldap_tools.ldap_attribute_values_join;
DROP TABLE IF EXISTS ldap_tools.ldap_attribute_value;
DROP TABLE IF EXISTS ldap_tools.ldap_attribute;
CREATE TABLE ldap_tools.ldap_attribute_values_join (
ldapattribute_id int8 NOT NULL,
valuelist_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_attribute_value (
id bigserial NOT NULL,
checksum int4 NOT NULL,
value varchar(6144) NULL,
CONSTRAINT ldap_attribute_value_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_attribute (
id bigserial NOT NULL,
checksum int4 NOT NULL,
totalvalues int4 NOT NULL,
"type" varchar(255) NULL,
CONSTRAINT ldap_attribute_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
ALTER TABLE ldap_tools.ldap_attribute_values_join ADD CONSTRAINT fkphkp0no9iistkvjeisqd4jav6 FOREIGN KEY (valuelist_id) REFERENCES ldap_tools.ldap_attribute_value(id);
ALTER TABLE ldap_tools.ldap_attribute_values_join ADD CONSTRAINT fkrf67fjr710xt4hfkplo6s3cuf FOREIGN KEY (ldapattribute_id) REFERENCES ldap_tools.ldap_attribute(id);
CREATE INDEX ldap_attribute_values_join__ldapattribute_id_index ON ldap_tools.ldap_attribute_values_join (ldapattribute_id);
CREATE INDEX ldap_attribute_values_join__valuelist_id_index ON ldap_tools.ldap_attribute_values_join (valuelist_id);
CREATE INDEX ldap_attribute_value_checksum_index ON ldap_tools.ldap_attribute_value (checksum);
CREATE INDEX ldap_attribute_checksum_index ON ldap_tools.ldap_attribute (checksum);<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/functions/dao/BindRequestDAO.java
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.functions.dao;
import java.util.Date;
import java.util.HashMap;
import java.util.List;
import java.util.Map;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.database.jpa.exception.JpaFacilitatorException;
import ita.deluca.tools.application.framework.database.jpa.model.JpaFacilitator;
import ita.deluca.tools.ldap.framework.commons.bean.BindRequest;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.exception.LdapEntityDaoException;
import ita.deluca.tools.ldap.framework.properties.Constants;
/**
* This class contains all the function that allows interaction with the database via JPA.
*
* @author <NAME>
*/
public class BindRequestDAO
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( BindRequestDAO.class );
/**
* This method persist the object into the database.
*
* @param entity
* the object to persist.
* @return The persisted entity.
* @throws LdapEntityDaoException
* this exception was raised whenever an error occurred during the persist of the object.
*/
protected static BindRequest persist( final BindRequest entity ) throws LdapEntityDaoException
{
if ( BindRequestDAO.LOG.isDebugEnabled() )
{
BindRequestDAO.LOG.debug( "Persist the entity:\n" + entity );
}
final BindRequest persistedEntity = LDAPEntityDAO.getAlreadyPersistEntity( entity );
if ( persistedEntity != null )
{
return persistedEntity;
}
return LDAPEntityDAO.persistEntity( entity );
}
/**
* Extract the number of milliseconds that was necessary to make a bind.
*
* @param connectionId
* The connection id that execute the bind.
* @param messageId
* The message id that execute the bind.
* @return The number of milliseconds that was necessary to make a bind.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the calculation.
*/
public static int getTimeToMake( final String connectionId, final int messageId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
parameter.put( "messageId", Integer.valueOf( messageId ) );
String query = "select message.timestamp from LDAPMessage message, BindRequest request where (message.protocolOp = request and message.connectionId = :connectionId and message.messageID = :messageId)";
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the bind request start date thrown an error !!!";
BindRequestDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
final Date startDate = ( Date ) res.get( 0 );
query = "select message.timestamp from LDAPMessage message, BindResponse result where (message.protocolOp = result and message.connectionId = :connectionId and message.messageID = :messageId)";
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the bind end date thrown an error !!!";
BindRequestDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
final Date endDate = ( Date ) res.get( 0 );
return Long.valueOf( endDate.getTime() - startDate.getTime() ).intValue();
}
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/responsetime/model/ResponseTimeFunctions.java
package ita.deluca.tools.ldap.framework.commons.responsetime.model;
import java.net.InetAddress;
import java.net.UnknownHostException;
import java.util.ArrayList;
import java.util.Arrays;
import java.util.Date;
import java.util.List;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.data.datetime.model.DateTimeFacilitators;
import ita.deluca.tools.application.framework.database.jpa.exception.JpaFacilitatorException;
import ita.deluca.tools.application.framework.database.jpa.model.JpaFacilitator;
import ita.deluca.tools.application.framework.ldap.bean.LdapSearchResultBean;
import ita.deluca.tools.application.framework.ldap.exception.LdapException;
import ita.deluca.tools.application.framework.ldap.models.LdapOperations;
import ita.deluca.tools.application.framework.properties.exception.PropertiesException;
import ita.deluca.tools.application.framework.properties.model.PropertiesManager;
import ita.deluca.tools.application.framework.tabulate.TabulateFacilitator;
import ita.deluca.tools.ldap.framework.commons.bean.ResponseTimeResult;
import ita.deluca.tools.ldap.framework.commons.responsetime.exception.ResponseTimeCheckException;
import ita.deluca.tools.ldap.framework.properties.Constants;
/**
* This class implements all the functions related to the testing of ldap server response time.
*
* @author <NAME>
*/
public class ResponseTimeFunctions
{
/**
* The class logging.
*/
private static final Logger LOG = LogManager.getLogger( ResponseTimeFunctions.class );
/**
* Standard constructor.
*/
private ResponseTimeFunctions()
{
}
/**
* This method verify the response time of the ldap server making the connect-search-unbind operations.
*
* @param environment
* The environment where verify the response time.
* @param server
* The ldap server where make the test of the response time.
* @return The response time check result.
* @throws ResponseTimeCheckException
* this exception was raised whenever occurred an error during the check of the response time.
*/
public static ResponseTimeResult checkServer( final String environment, final String server ) throws ResponseTimeCheckException
{
ResponseTimeFunctions.LOG.info( "Starting the test of response time on ldap server '" + server + "'" );
PropertiesManager pMgr;
try
{
pMgr = PropertiesManager.getPropertiesInstance( Constants.LDAP_TOOL_PROPERTIES_PATH );
} catch ( final PropertiesException ex )
{
final String msg = "!!! Error in retrieves of the properties manager !!!";
ResponseTimeFunctions.LOG.error( msg );
throw new RuntimeException( msg, ex );
}
final int port = pMgr.getPropertiesIntegerValue( "commons", environment, "server", "port" );
final String username = pMgr.getPropertiesStringValue( "commons", environment, "adminuser", "username" );
final String password = pMgr.getPropertiesStringValue( "commons", environment, "adminuser", "password" );
final String basepath = pMgr.getPropertiesStringValue( "responsetime", environment, "usersearch", "basepath" );
final String filter = pMgr.getPropertiesStringValue( "responsetime", environment, "usersearch", "filter" );
final String scope = pMgr.getPropertiesStringValue( "responsetime", environment, "usersearch", "scope" );
final String[] attributes = pMgr.getPropertiesArrayValue( "responsetime", environment, "usersearch", "attribute" );
final ResponseTimeResult result = new ResponseTimeResult();
result.setServer( server );
result.setData( new Date() );
final LdapOperations ldap = new LdapOperations();
try
{
try
{
ldap.connect( server, port, username, password );
} finally
{
result.setConnectElaps( ldap.getElaps() );
}
final List<LdapSearchResultBean> ldapResult;
try
{
ldapResult = ldap.search( basepath, scope, filter, attributes );
} finally
{
result.setSearchElaps( ldap.getElaps() );
}
result.setExtractedEntry( ldapResult.size() );
result.setReturnCode( 0 );
} catch ( final LdapException ex )
{
final String msg = ex.getMessage();
ResponseTimeFunctions.LOG.error( msg );
result.setReturnCode( ex.getErrorCode() );
} finally
{
try
{
ldap.close();
} catch ( final LdapException ex )
{
final String msg = ex.getMessage();
ResponseTimeFunctions.LOG.error( msg );
result.setReturnCode( ex.getErrorCode() );
} finally
{
result.setUnbindElaps( ldap.getElaps() );
}
}
try
{
result.setExecutorHost( InetAddress.getLocalHost().getHostName() );
} catch ( final UnknownHostException ex )
{
final String msg = "!!! Error in extraction of the localhost hostname !!!";
ResponseTimeFunctions.LOG.error( msg, ex );
throw new ResponseTimeCheckException( msg, ex );
}
ResponseTimeFunctions.LOG.info( "The request was processed in " + ( result.getConnectElaps() + result.getSearchElaps() + result.getUnbindElaps() ) + " milliseconds" );
return result;
}
/**
* This method verify the response time of the list of the ldap server making the connect-search-unbind operations.
*
* @param environment
* The environment where verify the response time.
* @return The response time check result list.
* @throws ResponseTimeCheckException
* this exception was raised whenever an error occurred during the test of the response time.
*/
public static List<ResponseTimeResult> checkServerList( final String environment ) throws ResponseTimeCheckException
{
ResponseTimeFunctions.LOG.info( "Starting the test of the response time." );
PropertiesManager pMgr;
try
{
pMgr = PropertiesManager.getPropertiesInstance( Constants.LDAP_TOOL_PROPERTIES_PATH );
} catch ( final PropertiesException ex )
{
final String msg = "!!! Error in retrieves of the properties manager !!!";
ResponseTimeFunctions.LOG.error( msg );
throw new RuntimeException( msg, ex );
}
final String[] serverList;
serverList = pMgr.getPropertiesArrayValue( "commons", environment, "server", "list" );
if ( serverList == null )
{
final String msg = "!!! The properties 'server -list' represents the ldap server list was not found !!!";
final ResponseTimeCheckException ex = new ResponseTimeCheckException( msg );
ResponseTimeFunctions.LOG.error( msg, ex );
throw ex;
}
return ResponseTimeFunctions.checkServerList( environment, Arrays.asList( serverList ) );
}
/**
* This method verify the response time of the list of the ldap server making the connect-search-unbind operations.
*
* @param environment
* The environment where verify the response time.
* @param serverList
* The server list.
* @return The response time check result list.
* @throws ResponseTimeCheckException
* this exception was raised whenever an error occurred during the test of the response time.
*/
public static List<ResponseTimeResult> checkServerList( final String environment, final List<String> serverList ) throws ResponseTimeCheckException
{
ResponseTimeFunctions.LOG.info( "Starting the test of the response time." );
final List<ResponseTimeResult> resultList = new ArrayList<>();
for ( final String server : serverList )
{
final ResponseTimeResult res = ResponseTimeFunctions.checkServer( environment, server );
resultList.add( res );
}
return resultList;
}
/**
* This function create the ASCII table relative to the test results.
*
* @param resultList
* The response time check result list.
* @return The ASCII table.
*/
public static String createResultAsciiTable( final List<ResponseTimeResult> resultList )
{
ResponseTimeFunctions.LOG.info( "Create the table contains the test results." );
final List<String> headerList = new ArrayList<>();
headerList.add( "SERVER" );
headerList.add( "DATA" );
headerList.add( "CONNECT ELAPS (Ms)" );
headerList.add( "SEARCH ELAPS (Ms)" );
headerList.add( "UNBIND ELAPS (Ms)" );
headerList.add( "TOTAL ELAPS (Ms)" );
headerList.add( "EXTRACTED ENTRY" );
ResponseTimeFunctions.LOG.debug( "The header contents : " + Arrays.toString( headerList.toArray() ) );
final List<List<Object>> contentsList = new ArrayList<>();
for ( final ResponseTimeResult result : resultList )
{
ResponseTimeFunctions.LOG.debug( "Insert into table contents the object : " + result + "" );
final List<Object> rowColumnList = new ArrayList<>();
rowColumnList.add( result.getServer() );
rowColumnList.add( result.getData() );
rowColumnList.add( Long.valueOf( result.getConnectElaps() ) );
rowColumnList.add( Long.valueOf( result.getSearchElaps() ) );
rowColumnList.add( Long.valueOf( result.getUnbindElaps() ) );
rowColumnList.add( Long.valueOf( result.getConnectElaps() + result.getSearchElaps() + result.getUnbindElaps() ) );
rowColumnList.add( Long.valueOf( result.getExtractedEntry() ) );
contentsList.add( rowColumnList );
ResponseTimeFunctions.LOG.debug( "The row contents : " + Arrays.toString( headerList.toArray() ) + "" );
}
return TabulateFacilitator.tabulate( headerList, contentsList );
}
/**
* This function create the stdout output.
*
* @param resultList
* The response time check result list.
* @return The stdout output.
*/
public static String createResultStdOutTable( final List<ResponseTimeResult> resultList )
{
ResponseTimeFunctions.LOG.info( "Create the table contains the test results." );
String outstr = "";
for ( final ResponseTimeResult result : resultList )
{
outstr += result.getServer() + ";" + DateTimeFacilitators.parseToString( result.getData(), "yyyy.MM.dd HH:mm:ss" ) + ";" + ( result.getConnectElaps() + result.getSearchElaps() + result.getUnbindElaps() ) + "\n";
}
return outstr.endsWith( "\n" ) ? outstr.substring( 0, outstr.length() - 1 ) : outstr;
}
/**
* This method makes the save on database of the list of the result of the response time test.
*
* @param resultList
* The response time check result list.
* @throws ResponseTimeCheckException
* this exception is raised whenever an errors occurs during the save of the results on database.
*/
public static void saveResponseTimeCheckResultList( final List<ResponseTimeResult> resultList ) throws ResponseTimeCheckException
{
ResponseTimeFunctions.LOG.info( "Saving on database the results of the response time test." );
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.RESPONSE_TIME_JPA_ENTITY_MANAGER ); )
{
jpa.openTransaction();
try
{
for ( final ResponseTimeResult result : resultList )
{
ResponseTimeFunctions.LOG.info( "Saving on database the results of the response time test make on the server '" + result.getServer() + "' on '" + result.getData() + "'." );
jpa.persist( result );
}
jpa.commit();
} catch ( final JpaFacilitatorException ex )
{
jpa.rollback();
final String msg = "!!! Error during the persist of the response time result : " + resultList + " !!!";
ResponseTimeFunctions.LOG.error( msg, ex );
throw new ResponseTimeCheckException( msg, ex );
}
}
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/GetEffectiveRightsControl.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import java.util.ArrayList;
import java.util.List;
import javax.persistence.Cacheable;
import javax.persistence.ElementCollection;
import javax.persistence.Entity;
import javax.persistence.FetchType;
import org.hibernate.annotations.Fetch;
import org.hibernate.annotations.FetchMode;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1Element;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1OctetString;
/**
* This class defines the LDAP get effective rights control, which may be used to make determinations about the access permissions for a given user.
*
* @author <NAME>
*/
@Entity
@Cacheable
public class GetEffectiveRightsControl extends LDAPControl
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The OID of the get effective rights control.
*/
public static final String GET_EFFECTIVE_RIGHTS_CONTROL_OID = "1.3.6.1.4.1.42.2.27.9.5.2";
/**
* The set of attribute types for which to make the determination.
*/
@ElementCollection( fetch = FetchType.EAGER )
@Fetch( FetchMode.SELECT )
private List<String> attributeTypes;
/**
* The authorization ID that specifies the user for which to make the determination.
*/
private String authzID;
/**
* Creates a new get effective rights control.
*/
public GetEffectiveRightsControl()
{
super( GetEffectiveRightsControl.GET_EFFECTIVE_RIGHTS_CONTROL_OID );
}
/**
* Creates a new get effective rights control.
*
* @param isCritical
* Indicates whether this control should be marked critical.
*/
public GetEffectiveRightsControl( final boolean isCritical )
{
super( GetEffectiveRightsControl.GET_EFFECTIVE_RIGHTS_CONTROL_OID, isCritical );
}
/**
* Creates a new get effective rights control by decoding the provided control value.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param controlValue
* The control value that may be decoded to obtain additional information about the way the request should be processed.
* @throws ProtocolException
* If a problem occurs while attempting to decode the control value.
*/
public GetEffectiveRightsControl( final boolean isCritical, final ASN1OctetString controlValue ) throws ProtocolException
{
super( GetEffectiveRightsControl.GET_EFFECTIVE_RIGHTS_CONTROL_OID, isCritical );
if ( controlValue == null )
{
this.authzID = null;
this.attributeTypes = null;
} else
{
final ASN1Element[] elements;
try
{
final byte[] valueBytes = controlValue.getValue();
elements = ASN1Element.decodeAsSequence( valueBytes ).getElements();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode get effective rights control sequence", e );
}
if ( elements.length != 2 )
{
throw new ProtocolException( "There must be exactly 2 elements in a get effective rights value sequence" );
}
try
{
this.authzID = elements[ 0 ].decodeAsOctetString().getStringValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode the authzID from the get effective rights control value", e );
}
try
{
final ASN1Element[] attrElements = elements[ 1 ].decodeAsSequence().getElements();
this.attributeTypes = new ArrayList<>();
for ( final ASN1Element attrElement : attrElements )
{
this.attributeTypes.add( attrElement.decodeAsOctetString().getStringValue() );
}
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode attribute types from the get effective rights control value", e );
}
}
}
/**
* Creates a new get effective rights control using the provided information.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param authzID
* The authorization ID that specifies the user for which to determine the effective rights.
* @param attributeTypes
* The set of attributes for which to retrieve the effective rights.
*/
public GetEffectiveRightsControl( final boolean isCritical, final String authzID, final List<String> attributeTypes )
{
super( GetEffectiveRightsControl.GET_EFFECTIVE_RIGHTS_CONTROL_OID, isCritical );
this.authzID = authzID;
this.attributeTypes = attributeTypes;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final GetEffectiveRightsControl other = ( GetEffectiveRightsControl ) obj;
if ( this.attributeTypes == null )
{
if ( other.attributeTypes != null )
{
return false;
}
} else if ( this.attributeTypes.size() != other.attributeTypes.size() )
{
return false;
} else
{
for ( final String uniqueStr : this.attributeTypes )
{
if ( !other.attributeTypes.contains( uniqueStr ) )
{
return false;
}
}
}
if ( this.authzID == null )
{
if ( other.authzID != null )
{
return false;
}
} else if ( !this.authzID.equals( other.authzID ) )
{
return false;
}
return true;
}
/**
* Retrieves the attribute types for which to make the effective rights determination.
*
* @return The attribute types for which to make the effective rights determination.
*/
public List<String> getAttributeTypes()
{
return this.attributeTypes;
}
/**
* Retrieves the authzID that specifies the user for which to make the effective rights determination.
*
* @return The authzID that specifies the user for which to make the effective rights determination.
*/
public String getAuthzID()
{
return this.authzID;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
if ( this.attributeTypes != null )
{
for ( final String str : this.attributeTypes )
{
result += str.hashCode();
}
}
result = ( prime * result ) + ( this.authzID == null ? 0 : this.authzID.hashCode() );
return result;
}
/**
* Setting the attribute types for which to make the effective rights determination.
*
* @param attributeTypes
* The attribute types for which to make the effective rights determination.
*/
public void setAttributeTypes( final List<String> attributeTypes )
{
this.attributeTypes = attributeTypes;
}
/**
* Setting the authzID that specifies the user for which to make the effective rights determination.
*
* @param authzID
* The authzID that specifies the user for which to make the effective rights determination.
*/
public void setAuthzID( final String authzID )
{
this.authzID = authzID;
}
/**
* Retrieves a string representation of this control with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this control with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( "LDAP Get Effective Rights Control" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " OID: " ).append( this.getControlOID() ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Criticality: " ).append( this.isCritical() ).append( LDAPMessage.EOL );
if ( this.authzID != null )
{
buffer.append( indentBuf ).append( " Authorization ID: " ).append( this.authzID ).append( LDAPMessage.EOL );
}
if ( ( this.attributeTypes != null ) && !this.attributeTypes.isEmpty() )
{
buffer.append( indentBuf ).append( " Attribute Types:" ).append( LDAPMessage.EOL );
for ( final String attributeType : this.attributeTypes )
{
buffer.append( indentBuf ).append( " " ).append( attributeType ).append( LDAPMessage.EOL );
}
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
if ( this.attributeTypes != null )
{
for ( final String str : this.attributeTypes )
{
this.checksum = ( prime * this.checksum ) + str.hashCode();
}
}
this.checksum = ( prime * this.checksum ) + ( this.authzID == null ? 0 : this.authzID.hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/LDAPAttribute.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import java.util.ArrayList;
import java.util.List;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import javax.persistence.FetchType;
import javax.persistence.JoinTable;
import javax.persistence.ManyToMany;
import javax.persistence.Table;
import org.hibernate.annotations.Fetch;
import org.hibernate.annotations.FetchMode;
/**
* This class defines an LDAP attribute, which has a type and zero or more values.
*
* @author <NAME>
*/
@Entity
@Cacheable
@Table( schema = "ldap_tools", name = "ldap_attribute" )
public class LDAPAttribute extends LDAPEntity
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The total number of value in this attribute.
*/
private int totalValues;
/**
* The type for this attribute.
*/
private String type;
/**
* The attribute value list.
*/
@ManyToMany( fetch = FetchType.EAGER )
@JoinTable( schema = "ldap_tools", name = "ldap_attribute_values_join" )
@Fetch( FetchMode.SELECT )
private List<LDAPAttributeValue> valueList = new ArrayList<>();
/**
* Creates a new LDAP attribute with the provided type and values.
*/
public LDAPAttribute()
{
}
/**
* Creates a new LDAP attribute with the provided type and values.
*
* @param type
* The attribute type for this attribute.
* @param totalValues
* The total number of value in this attribute.
* @param valueList
* The attribute value list.
*/
public LDAPAttribute( final String type, final int totalValues, final List<LDAPAttributeValue> valueList )
{
this.type = type;
this.totalValues = totalValues;
this.valueList = valueList;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( ( obj == null ) || ( this.getClass() != obj.getClass() ) )
{
return false;
}
final LDAPAttribute that = ( LDAPAttribute ) obj;
if ( this.valueList == null )
{
if ( that.valueList != null )
{
return false;
}
} else if ( this.valueList.size() != that.valueList.size() )
{
return false;
} else
{
for ( final LDAPAttributeValue value : this.valueList )
{
if ( !that.valueList.contains( value ) )
{
return false;
}
}
}
return ( this.totalValues == that.totalValues ) && this.type.equals( that.type );
}
/**
* Retrieves the total number of value in this attribute.
*
* @return The total number of value in this attribute.
*/
public int getTotalValues()
{
return this.totalValues;
}
/**
* Retrieves the attribute type for this attribute.
*
* @return The attribute type for this attribute.
*/
public String getType()
{
return this.type;
}
/**
* Retrieves the attribute value list.
*
* @return The attribute value list.
*/
public List<LDAPAttributeValue> getValueList()
{
return this.valueList;
}
@Override
public int hashCode()
{
int result = this.totalValues;
for ( final LDAPAttributeValue value : this.valueList )
{
result += value.hashCode();
}
result = ( 31 * result ) + this.type.hashCode();
return result;
}
/**
* Setting the total number of value in this attribute.
*
* @param totalValues
* The total number of value in this attribute.
*/
public void setTotalValues( final int totalValues )
{
this.totalValues = totalValues;
}
/**
* Setting the attribute type for this attribute.
*
* @param type
* The attribute type for this attribute.
*/
public void setType( final String type )
{
this.type = type;
}
/**
* Setting the attribute value list.
*
* @param valueList
* The attribute value list.
*/
public void setValueList( final List<LDAPAttributeValue> valueList )
{
this.valueList = valueList;
}
/**
* Retrieves a string representation of this LDAP attribute.
*
* @return A string representation of this LDAP attribute.
*/
@Override
public String toString()
{
return this.toString( 0 );
}
/**
* Retrieves a string representation of this LDAP attribute with the specified indent.
*
* @param indent
* The number of spaces to the left of the value.
* @return A string representation of this LDAP attribute.
*/
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( this.type ).append( ": " ).append( this.totalValues ).append( " values" ).append( LDAPMessage.EOL );
for ( int i = 0; ( this.valueList != null ) && ( i < this.valueList.size() ); i++ )
{
buffer.append( indentBuf ).append( this.type ).append( ": " ).append( this.valueList.get( i ) );
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
for ( final LDAPAttributeValue value : this.valueList )
{
value.updateChecksum();
this.checksum = ( prime * this.checksum ) + value.checksum;
}
this.checksum = ( prime * this.checksum ) + ( this.type == null ? 0 : this.type.hashCode() );
this.checksum = ( prime * this.checksum ) + this.totalValues;
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/SQL/CreateDatabase.sql
DROP TABLE IF EXISTS ldapcheck.responsetime;
DROP SEQUENCE IF EXISTS ldapcheck.responsetime_id_seq;
CREATE SEQUENCE ldapcheck.responsetime_id_seq;
CREATE TABLE ldapcheck.responsetime (
id INT4 NOT NULL DEFAULT nextval('responsetime_id_seq' :: REGCLASS),
connect_elaps INT8 NULL,
"date" TIMESTAMP NULL,
executor_host VARCHAR(255) NULL,
extracted_entry_count INT4 NULL,
search_elaps INT8 NULL,
server VARCHAR(255) NULL,
unbind_elaps INT8 NULL,
CONSTRAINT responsetime_pkey PRIMARY KEY (id)
)
WITH (
OIDS = FALSE
);
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/LDAPAttributeValue.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import javax.persistence.Table;
/**
* This class defines an LDAP attribute value, which has a type and zero or more values.
*
* @author <NAME>
*/
@Entity
@Cacheable
@Table( schema = "ldap_tools", name = "ldap_attribute_value" )
public class LDAPAttributeValue extends LDAPEntity
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The type for this attribute.
*/
private String value;
/**
* Standard constructor.
*/
public LDAPAttributeValue()
{
}
/**
* Standard constructor.
*
* @param value
* The value.
*/
public LDAPAttributeValue( final String value )
{
this.value = value;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( obj == null )
{
return false;
}
if ( !( obj instanceof LDAPAttributeValue ) )
{
return false;
}
final LDAPAttributeValue other = ( LDAPAttributeValue ) obj;
if ( this.value == null )
{
if ( other.value != null )
{
return false;
}
} else if ( !this.value.equals( other.value ) )
{
return false;
}
return true;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = 1;
result = ( prime * result ) + ( ( this.value == null ) ? 0 : this.value.hashCode() );
return result;
}
/**
* Retrieves a string representation of this LDAP attribute.
*
* @return A string representation of this LDAP attribute.
*/
@Override
public String toString()
{
return this.toString( 0 );
}
/**
* Retrieves a string representation of this LDAP attribute with the specified indent.
*
* @param indent
* The number of spaces to the left of the value.
* @return A string representation of this LDAP attribute.
*/
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
return String.valueOf( indentBuf ) + this.value + LDAPMessage.EOL;
}
@Override
public void updateChecksum()
{
this.checksum = this.hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/userconnection/model/UserConnectionCheckFunctions.java
package ita.deluca.tools.ldap.framework.commons.userconnection.model;
import java.util.ArrayList;
import java.util.Arrays;
import java.util.List;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.ldap.exception.LdapException;
import ita.deluca.tools.application.framework.ldap.models.LdapOperations;
import ita.deluca.tools.application.framework.properties.exception.PropertiesException;
import ita.deluca.tools.application.framework.properties.model.PropertiesManager;
import ita.deluca.tools.application.framework.tabulate.TabulateFacilitator;
import ita.deluca.tools.ldap.framework.commons.bean.UserConnectionCheckResult;
import ita.deluca.tools.ldap.framework.commons.userconnection.exception.UserConnectionCheckException;
import ita.deluca.tools.ldap.framework.properties.Constants;
/**
* This class contains all the utility function relative to the test of user connections.
*
* @author <NAME>
*/
public class UserConnectionCheckFunctions
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( UserConnectionCheckFunctions.class );
/**
* Standard constructor.
*/
private UserConnectionCheckFunctions()
{
}
/**
* This function create the ASCII table relative to the test of the user connection.
*
* @param userConnResultList
* The list of the user connection result.
* @return The ASCII table.
*/
public static String createUserConnectionCheckResultAsciiTable( final List<UserConnectionCheckResult> userConnResultList )
{
UserConnectionCheckFunctions.LOG.info( "Create the table contains the result of the user connection test." );
final List<String> headerList = new ArrayList<>();
headerList.add( "SERVER" );
headerList.add( "DATA" );
headerList.add( "USER FULL DN" );
headerList.add( "CONNECTION OK" );
UserConnectionCheckFunctions.LOG.debug( "The header contents : " + Arrays.toString( headerList.toArray() ) );
final List<List<Object>> contentsList = new ArrayList<>();
for ( final UserConnectionCheckResult userConnResult : userConnResultList )
{
UserConnectionCheckFunctions.LOG.debug( "Insert into table contents the object : " + userConnResult + "" );
final List<Object> rowColumnList = new ArrayList<>();
rowColumnList.add( userConnResult.getServer() );
rowColumnList.add( userConnResult.getData() );
rowColumnList.add( userConnResult.getUserFullDn() );
rowColumnList.add( Boolean.valueOf( userConnResult.isConnectionOk() ) );
contentsList.add( rowColumnList );
UserConnectionCheckFunctions.LOG.debug( "The row contents : " + Arrays.toString( headerList.toArray() ) + "" );
}
return TabulateFacilitator.tabulate( headerList, contentsList );
}
/**
* This function test the user connection on a server list.
*
* @param environment
* The environment where make the test.
* @param user
* The user to test connection.
* @param password
* The user <PASSWORD>.
* @return The list of the user connection result.
* @throws UserConnectionCheckException
* this exception was raised whenever an error occurred during the test of user connection.
*/
public static List<UserConnectionCheckResult> testUserConnection( final String environment, final String user, final String password ) throws UserConnectionCheckException
{
UserConnectionCheckFunctions.LOG.info( "Test the user connection of user '" + user + "' on server list." );
final PropertiesManager pMgr;
try
{
pMgr = PropertiesManager.getPropertiesInstance( Constants.LDAP_TOOL_PROPERTIES_PATH );
} catch ( final PropertiesException ex )
{
final String msg = "!!! Error in retrieves of the properties manager !!!";
UserConnectionCheckFunctions.LOG.error( msg );
throw new UserConnectionCheckException( msg, ex );
}
final String[] serverList = pMgr.getPropertiesArrayValue( "commons", environment, "server", "list" );
if ( serverList == null )
{
final String msg = "!!! The properties 'server -list' represents the ldap server list was not found !!!";
final UserConnectionCheckException ex = new UserConnectionCheckException( msg );
UserConnectionCheckFunctions.LOG.error( msg, ex );
throw ex;
}
final List<UserConnectionCheckResult> userConnResultList = new ArrayList<>();
for ( final String server : serverList )
{
UserConnectionCheckFunctions.LOG.info( "Test the user connection of user '" + user + "' on server '" + server + "'." );
try
{
userConnResultList.add( UserConnectionCheckFunctions.testUserConnection( environment, server, user, password ) );
} catch ( final UserConnectionCheckException ex )
{
final String msg = "!!! An error was occurred during the the test of the user '" + user + "' connection on server '" + server + "' !!!";
UserConnectionCheckFunctions.LOG.error( msg );
throw new UserConnectionCheckException( msg, ex );
}
}
return userConnResultList;
}
/**
* This function test the user connection on a specific server.
*
* @param environment
* The environment where make the test.
* @param server
* the server where test the user connection.
* @param user
* The user to test connection.
* @param password
* <PASSWORD>.
* @return The user connection result.
* @throws UserConnectionCheckException
* this exception was raised whenever an error occurred during the test of user connection.
*/
public static UserConnectionCheckResult testUserConnection( final String environment, final String server, final String user, final String password ) throws UserConnectionCheckException
{
UserConnectionCheckFunctions.LOG.info( "Test the user connection of user '" + user + "' on server '" + server + "'." );
final PropertiesManager pMgr;
try
{
pMgr = PropertiesManager.getPropertiesInstance( Constants.LDAP_TOOL_PROPERTIES_PATH );
} catch ( final PropertiesException ex )
{
final String msg = "!!! Error in retrieves of the properties manager !!!";
UserConnectionCheckFunctions.LOG.error( msg );
throw new UserConnectionCheckException( msg, ex );
}
final int port = pMgr.getPropertiesIntegerValue( "commons", environment, "server", "port" );
final UserConnectionCheckResult userConnResult = new UserConnectionCheckResult();
final LdapOperations ldap = new LdapOperations();
try
{
ldap.connect( server, port, user, password );
userConnResult.setData( ldap.getOperationDate() );
ldap.close();
userConnResult.setConnectionOk( true );
} catch ( final LdapException ex )
{
if ( ex.getErrorCode() == LdapException.AUTHENTICATION_ERROR )
{
UserConnectionCheckFunctions.LOG.error( "!!! The username/password was incorrect !!!" );
userConnResult.setConnectionOk( false );
if ( userConnResult.getData() == null )
{
userConnResult.setData( ldap.getOperationDate() );
}
} else
{
final String msg = "!!! An error was occurred during the connect !!!";
UserConnectionCheckFunctions.LOG.error( msg );
throw new UserConnectionCheckException( msg, ex );
}
}
userConnResult.setServer( server );
userConnResult.setUserFullDn( user );
return userConnResult;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/edirectory/bean/FakeX509TrustManager.java
package ita.deluca.tools.ldap.framework.edirectory.bean;
import java.security.cert.CertificateException;
import java.security.cert.X509Certificate;
import javax.net.ssl.X509TrustManager;
/**
* This class represents a fake 509 Trust manager.
*
* @author <NAME>
*/
public class FakeX509TrustManager implements X509TrustManager
{
@Override
public void checkClientTrusted( final X509Certificate[] arg0, final String arg1 ) throws CertificateException
{
}
@Override
public void checkServerTrusted( final X509Certificate[] arg0, final String arg1 ) throws CertificateException
{
}
@Override
public X509Certificate[] getAcceptedIssuers()
{
return null;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/ProtocolException.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import ita.deluca.tools.application.framework.main.exception.MainException;
/**
* This class defines an exception that may be thrown if a problem is encountered while decoding an LDAP operation.
*
* @author <NAME>
*/
public class ProtocolException extends MainException
{
/**
* The serial version UID for this serializable class.
*/
private static final long serialVersionUID = 1885306391093027694L;
/**
* Creates a new protocol exception with the provided message.
*
* @param message
* The message that explains the reason for this exception.
*/
public ProtocolException( final String message )
{
super( message );
}
/**
* Creates a new protocol exception with the provided message.
*
* @param message
* The message that explains the reason for this exception.
* @param cause
* The underlying cause for this exception.
*/
public ProtocolException( final String message, final Throwable cause )
{
super( message, cause );
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/asn1/ASN1Sequence.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1;
import java.util.ArrayList;
/**
* This class defines an ASN.1 element that serves as a sequence, whose value is an ordered set of other ASN.1 elements.
*
* @author <NAME>
*/
public class ASN1Sequence extends ASN1Element
{
/**
* The set of elements that will be used if there are no elements.
*/
public static final ASN1Element[] NO_ELEMENTS = new ASN1Element[ 0 ];
/**
* The set of ASN.1 elements associated with this sequence.
*/
ASN1Element[] elements;
/**
* Creates a new ASN.1 sequence with the given type to hold the specified set of ASN.1 elements.
*
* @param type
* The type to use for this ASN.1 sequence.
* @param elements
* The set of ASN.1 elements to encode in the value of this sequence.
*/
private ASN1Sequence( final byte type, final ArrayList<ASN1Element> elements )
{
super( type );
if ( ( elements == null ) || elements.isEmpty() )
{
this.elements = ASN1Sequence.NO_ELEMENTS;
} else
{
final ASN1Element[] elementArray = new ASN1Element[ elements.size() ];
elements.toArray( elementArray );
this.replaceElements( elementArray );
}
}
/**
* Creates a new ASN.1 sequence with the given type to hold the specified set of ASN.1 elements.
*
* @param type
* The type to use for this ASN.1 sequence.
* @param elements
* The set of ASN.1 elements to encode in the value of this sequence.
*/
private ASN1Sequence( final byte type, final ASN1Element[] elements )
{
super( type );
this.elements = elements;
if ( ( elements == null ) || ( elements.length == 0 ) )
{
this.elements = ASN1Sequence.NO_ELEMENTS;
}
this.replaceElements( elements );
}
/**
* Creates a new ASN.1 sequence with no elements encoded in the value.
*/
public ASN1Sequence()
{
this( ASN1Element.ASN1_SEQUENCE_TYPE, ASN1Sequence.NO_ELEMENTS );
}
/**
* Creates a new ASN.1 sequence to hold the specified set of ASN.1 elements.
*
* @param elements
* The set of ASN.1 elements to encode in the value of this sequence.
*/
public ASN1Sequence( final ArrayList<ASN1Element> elements )
{
this( ASN1Element.ASN1_SEQUENCE_TYPE, elements );
}
/**
* Creates a new ASN.1 sequence to hold the specified set of ASN.1 elements.
*
* @param elements
* The set of ASN.1 elements to encode in the value of this sequence.
*/
public ASN1Sequence( final ASN1Element[] elements )
{
this( ASN1Element.ASN1_SEQUENCE_TYPE, elements );
}
/**
* Creates a new ASN.1 sequence with the specified type and no elements encoded in the value.
*
* @param type
* The type to use for this ASN.1 sequence.
*/
public ASN1Sequence( final byte type )
{
this( type, ASN1Sequence.NO_ELEMENTS );
}
/**
* Decodes the provided byte array as if it were a set of ASN.1 elements.
*
* @param sequenceValue
* The set of encoded ASN.1 elements.
* @return The ASN.1 elements that were decoded from the provided byte array.
* @throws ASN1Exception
* If the provided byte array cannot be decoded into a set of ASN.1 elements.
*/
public static ASN1Element[] decodeSequenceElements( final byte[] sequenceValue ) throws ASN1Exception
{
final ArrayList<ASN1Element> elementList = new ArrayList<>();
int startPos = 0;
while ( startPos < sequenceValue.length )
{
/*
* The first byte is going to be the type. Make sure it's not a multibyte type.
*/
final byte type = sequenceValue[ startPos ];
if ( ( type & 0x1F ) == 0x1F )
{
throw new ASN1Exception( "Multibyte type detected (not supported in this package)" );
}
// The length starts at the second position, but may extend beyond that
final byte firstLengthByte = sequenceValue[ startPos + 1 ];
int length;
int valueStartPos = startPos + 2;
if ( ( firstLengthByte & 0x7F ) == firstLengthByte )
{
length = firstLengthByte;
} else if ( ( firstLengthByte & 0x7F ) == 0x00 )
{
length = 128;
} else
{
// There are multiple bytes in the length. Figure out how many.
final int numLengthBytes = firstLengthByte & 0x7F;
length = 0x00000000;
switch ( numLengthBytes )
{
case 1:
length |= 0x000000FF & sequenceValue[ startPos + 2 ];
valueStartPos++;
break;
case 2:
length |= ( ( 0x000000FF & sequenceValue[ startPos + 2 ] ) << 8 ) | ( 0x000000FF & sequenceValue[ startPos + 3 ] );
valueStartPos += 2;
break;
case 3:
length |= ( ( 0x000000FF & sequenceValue[ startPos + 2 ] ) << 16 ) | ( ( 0x000000FF & sequenceValue[ startPos + 3 ] ) << 8 ) | ( 0x000000FF & sequenceValue[ startPos + 4 ] );
valueStartPos += 3;
break;
case 4:
length |= ( ( 0x000000FF & sequenceValue[ startPos + 2 ] ) << 24 ) | ( ( 0x000000FF & sequenceValue[ startPos + 3 ] ) << 16 ) | ( ( 0x000000FF & sequenceValue[ startPos + 4 ] ) << 8 ) | ( 0x000000FF & sequenceValue[ startPos + 5 ] );
valueStartPos += 4;
break;
default:
throw new ASN1Exception( "Specified length cannot be represented as a Java int" );
}
}
// Make sure that the specified number of bytes actually exist.
if ( ( valueStartPos + length ) > sequenceValue.length )
{
throw new ASN1Exception( "There are not enough bytes in the value to hold the indicated length of " + length );
}
// Copy the value in place
final byte[] value = new byte[ length ];
System.arraycopy( sequenceValue, valueStartPos, value, 0, length );
// Create the new ASN.1 element and add it to the list
final ASN1Element element = new ASN1Element( type, value );
elementList.add( element );
// Reset the start position for the next element in the list
startPos = valueStartPos + length;
}
// Convert the list elements to an array and return it
final ASN1Element[] elements = new ASN1Element[ elementList.size() ];
elementList.toArray( elements );
return elements;
}
/**
* Replaces the current set of elements with the provided set.
*
* @param elementsTmp
* The set of ASN.1 elements to use to replace the existing set of elements encoded in this sequence.
*/
private void replaceElements( final ASN1Element[] elementsTmp )
{
this.elements = elementsTmp == null ? ASN1Sequence.NO_ELEMENTS : elementsTmp;
// Figure out the total length of the encoded value
int totalLength = 0;
for ( final ASN1Element element : this.elements )
{
totalLength += element.encodedElement.length;
}
final byte[] encodedValue = new byte[ totalLength ];
int startPos = 0;
for ( final ASN1Element element : this.elements )
{
startPos += element.encode( encodedValue, startPos );
}
this.setValue( encodedValue );
}
/**
* Adds the specified ASN.1 element to the set of elements encoded in this sequence.
*
* @param element
* The ASN.1 element to include in the set of elements encoded in this sequence.
*/
public void addElement( final ASN1Element element )
{
this.elements = ASN1Element.addElementToSet( this.elements, element );
this.value = this.appendNewElementsByteToValue( element );
}
/**
* Retrieves the set of elements that are encoded in this ASN.1 sequence.
*
* @return The set of elements that are encoded in this ASN.1 sequence.
*/
public ASN1Element[] getElements()
{
return this.elements;
}
/**
* Removes all elements encoded in the value of this ASN.1 element.
*/
public void removeAllElements()
{
this.setValue( ASN1Element.EMPTY_BYTES );
this.elements = ASN1Sequence.NO_ELEMENTS;
}
/**
* Replaces the current set of elements with the provided ASN.1 element.
*
* @param element
* The ASN.1 element to use to replace the existing set of elements encoded in this sequence.
*/
public void replaceElements( final ASN1Element element )
{
this.replaceElements( new ASN1Element[] { element } );
}
/**
* Retrieves a string representation of this ASN.1 sequence. It will recursively display string representations for each of the elements.
*
* @param indent
* The number of spaces to indent the information in the returned string.
* @return A string representation of this ASN.1 sequence.
*/
@Override( )
public String toString( final int indent )
{
String indentStr = "";
for ( int i = 0; i < indent; i++ )
{
indentStr += " ";
}
String elementsStr = "";
for ( int i = 0; i < this.elements.length; i++ )
{
final ASN1Element element = this.elements[ i ];
elementsStr += indentStr + " Element " + i + ASN1Element.EOL + element.toString( indent + 2 );
}
return indentStr + "Type: " + this.type + ASN1Element.EOL + ASN1Element.byteArrayToString( new byte[] { this.type }, 2 + indent ) + ASN1Element.EOL + indentStr + "Length: " + this.value.length + ASN1Element.EOL + ASN1Element.byteArrayToString( ASN1Element.encodeLength( this.value.length ), 2 + indent ) + ASN1Element.EOL + indentStr + "Value: " + new String( this.value ) + ASN1Element.EOL + ASN1Element.byteArrayToString( this.value, 2 + indent ) + ASN1Element.EOL + "Elements: " + ASN1Element.EOL + elementsStr;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/functions/dao/SearchResultEntryDAO.java
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.functions.dao;
import java.util.ArrayList;
import java.util.HashMap;
import java.util.List;
import java.util.Map;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.database.jpa.exception.JpaFacilitatorException;
import ita.deluca.tools.application.framework.database.jpa.model.JpaFacilitator;
import ita.deluca.tools.ldap.framework.commons.bean.LDAPAttribute;
import ita.deluca.tools.ldap.framework.commons.bean.LDAPMessage;
import ita.deluca.tools.ldap.framework.commons.bean.SearchResultEntry;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.exception.LdapEntityDaoException;
import ita.deluca.tools.ldap.framework.properties.Constants;
/**
* This class contains all the function that allows interaction with the database via JPA.
*
* @author <NAME>
*/
public class SearchResultEntryDAO
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( SearchResultEntryDAO.class );
/**
* This method persist the object into the database.
*
* @param entity
* the object to persist.
* @return The persisted entity.
* @throws LdapEntityDaoException
* this exception was raised whenever an error occurred during the persist of the object.
*/
protected static SearchResultEntry persist( final SearchResultEntry entity ) throws LdapEntityDaoException
{
if ( SearchResultEntryDAO.LOG.isDebugEnabled() )
{
SearchResultEntryDAO.LOG.debug( "Persist the entity:\n" + entity );
}
final SearchResultEntry persistedEntity = LDAPEntityDAO.getAlreadyPersistEntity( entity );
if ( persistedEntity != null )
{
return persistedEntity;
}
final List<LDAPAttribute> persistedList = new ArrayList<>();
for ( final LDAPAttribute obj : entity.getAttributes() )
{
final int index = persistedList.indexOf( obj );
if ( index == -1 )
{
persistedList.add( LDAPAttributeDAO.persist( obj ) );
} else
{
persistedList.add( persistedList.get( index ) );
}
}
entity.setAttributes( persistedList );
return LDAPEntityDAO.persistEntity( entity );
}
/**
* Extract the results that has a number of attribute major of the selected values.
*
* @param maxattribute
* The maximum number of attribute.
* @return The list of the search results that has the number of attributes major than the selected values.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the search result.
*/
public static List<LDAPMessage> getEntryWithAttributeNumberMajorOf( final int maxattribute ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select LDAPMessage from LDAPMessage LDAPMessage, SearchResultEntry result join result.attributes attribute where (LDAPMessage.protocolOp = result AND attribute.totalValues > :maxattribute )";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "maxattribute", Integer.valueOf( maxattribute ) );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the list search result having a number of attribute greater than '" + maxattribute + "' thrown an error !!!";
SearchResultEntryDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
final List<LDAPMessage> retRes = ( List<LDAPMessage> ) res;
return retRes;
}
}
/**
* Extract the results of the search request.
*
* @param connectionId
* The connection id of the request.
* @param messageId
* The search message id.
* @return The list of the search results of the search request.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the search result.
*/
public static List<LDAPMessage> getFromConnectionIdAndMessageId( final String connectionId, final int messageId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select LDAPMessage from LDAPMessage LDAPMessage, SearchResultEntry result where (LDAPMessage.protocolOp = result AND LDAPMessage.connectionId = :connectionId and LDAPMessage.messageID = :messageId )";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
parameter.put( "messageId", Integer.valueOf( messageId ) );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the list search result of the search with connection id '" + connectionId + "' and message id '" + messageId + "' thrown an error !!!";
SearchResultEntryDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
final List<LDAPMessage> retRes = ( List<LDAPMessage> ) res;
return retRes;
}
}
/**
* Extract the total bytes of the search results of the search request.
*
* @param connectionId
* The connection id of the request.
* @param messageId
* The search message id.
* @return The total bytes of the search results of the search request.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the search result.
*/
public static int getTotalBytesFromConnectionIdAndMessageId( final String connectionId, final int messageId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select SUM(LDAPMessage.messageLength) from LDAPMessage LDAPMessage, SearchResultEntry result where (LDAPMessage.protocolOp = result AND LDAPMessage.connectionId = :connectionId and LDAPMessage.messageID = :messageId )";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
parameter.put( "messageId", Integer.valueOf( messageId ) );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the total byte of the search result with connection id '" + connectionId + "' and message id '" + messageId + "' thrown an error !!!";
SearchResultEntryDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
return ( ( Long ) res.get( 0 ) ).intValue();
}
}
/**
* Extract the total number of results of the search request.
*
* @param connectionId
* The connection id of the request.
* @param messageId
* The search message id.
* @return The total number of the search results of the search request.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the search result.
*/
public static int getTotalFromConnectionIdAndMessageId( final String connectionId, final int messageId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select COUNT(LDAPMessage) from LDAPMessage LDAPMessage, SearchResultEntry result where (LDAPMessage.protocolOp = result AND LDAPMessage.connectionId = :connectionId and LDAPMessage.messageID = :messageId )";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
parameter.put( "messageId", Integer.valueOf( messageId ) );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the total search result of the search with connection id '" + connectionId + "' and message id '" + messageId + "' thrown an error !!!";
SearchResultEntryDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
return ( ( Long ) res.get( 0 ) ).intValue();
}
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/SearchResultReference.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import java.util.ArrayList;
import java.util.List;
import javax.persistence.Cacheable;
import javax.persistence.ElementCollection;
import javax.persistence.Entity;
import javax.persistence.FetchType;
import org.hibernate.annotations.Fetch;
import org.hibernate.annotations.FetchMode;
/**
* This class defines an LDAP search result reference, which provides a link to one or more other locations in which the search should be performed.
*
* @author <NAME>
*/
@Entity
@Cacheable
public class SearchResultReference extends ProtocolOp
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The set of referral URLs for this search result reference.
*/
@ElementCollection( fetch = FetchType.EAGER )
@Fetch( FetchMode.SELECT )
private List<String> referralURLs = new ArrayList<>();
/**
* Creates a new search result reference with the provided referral URLs.
*/
public SearchResultReference()
{
}
/**
* Creates a new search result reference with the provided referral URLs.
*
* @param referralURLs
* The referral URLs for use in this search result reference.
*/
public SearchResultReference( final ArrayList<String> referralURLs )
{
this.referralURLs = referralURLs;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final SearchResultReference other = ( SearchResultReference ) obj;
if ( this.referralURLs == null )
{
if ( other.referralURLs != null )
{
return false;
}
} else if ( this.referralURLs.size() != other.referralURLs.size() )
{
return false;
} else
{
for ( final String uniqueStr : this.referralURLs )
{
if ( !other.referralURLs.contains( uniqueStr ) )
{
return false;
}
}
}
return true;
}
/**
* Retrieves the set of referral URLs associated with this search result reference.
*
* @return The set of referral URLs associated with this search result reference.
*/
public List<String> getReferralURLs()
{
return this.referralURLs;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
for ( final String str : this.referralURLs )
{
result += str.hashCode();
}
result = prime * result;
return result;
}
/**
* Retrieves a user-friendly name for this protocol op.
*
* @return A user-friendly name for this protocol op.
*/
@Override
public String retrieveProtocolOpType()
{
return "LDAP SearchResultReference";
}
/**
* Setting the set of referral URLs associated with this search result reference.
*
* @param referralURLs
* The set of referral URLs associated with this search result reference.
*/
public void setReferralURLs( final List<String> referralURLs )
{
this.referralURLs = referralURLs;
}
/**
* Retrieves a string representation of this protocol op with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this protocol op with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( "Referral URLs: " ).append( LDAPMessage.EOL );
for ( int i = 0; ( this.referralURLs != null ) && ( i < this.referralURLs.size() ); i++ )
{
buffer.append( indentBuf ).append( " " ).append( this.referralURLs.get( i ) ).append( LDAPMessage.EOL );
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
for ( final String str : this.referralURLs )
{
this.checksum = ( prime * this.checksum ) + str.hashCode();
}
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/PagedResultsControl.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1Element;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1OctetString;
/**
* This class defines the simple paged results control, as defined in RFC 2696. This control is used to retrieve the results of a search operation a "page" at a time.
*
* @author <NAME>
*/
@Entity @Cacheable
public class PagedResultsControl extends LDAPControl
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The OID of the paged results control.
*/
public static final String PAGED_RESULTS_CONTROL_OID = "1.2.840.113556.1.4.319";
/**
* The opaque cookie value for this control.
*/
private String cookie;
/**
* The page size or estimated result set size for this control.
*/
private int size;
/**
* Creates a new simple paged results control.
*/
public PagedResultsControl()
{
super( PagedResultsControl.PAGED_RESULTS_CONTROL_OID );
}
/**
* Creates a new simple paged results control.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param controlValue
* The encoded value for the paged results control.
* @throws ProtocolException
* If the provided control value cannot be decoded appropriately for a paged results control.
*/
public PagedResultsControl( final boolean isCritical, final ASN1OctetString controlValue ) throws ProtocolException
{
super( PagedResultsControl.PAGED_RESULTS_CONTROL_OID, isCritical );
final ASN1Element[] sequenceElements;
try
{
final byte[] valueBytes = controlValue.getValue();
sequenceElements = ASN1Element.decodeAsSequence( valueBytes ).getElements();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode paged results control sequence", e );
}
if ( sequenceElements.length != 2 )
{
throw new ProtocolException( "There must be exactly 2 elements in a paged results control sequence" );
}
try
{
this.size = sequenceElements[ 0 ].decodeAsInteger().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode size from paged results control sequence", e );
}
try
{
this.cookie = ASN1Element.byteArrayToStringWithASCII( sequenceElements[ 1 ].decodeAsOctetString().getValue() );
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode cookie from paged results control sequence", e );
}
}
/**
* Creates a new simple paged results control.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param size
* The size for this control.
* @param cookie
* The opaque cookie for this control.
*/
public PagedResultsControl( final boolean isCritical, final int size, final String cookie )
{
super( PagedResultsControl.PAGED_RESULTS_CONTROL_OID, isCritical );
this.size = size;
this.cookie = cookie;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final PagedResultsControl other = ( PagedResultsControl ) obj;
if ( this.cookie == null )
{
if ( other.cookie != null )
{
return false;
}
} else if ( !this.cookie.equals( other.cookie ) )
{
return false;
}
return this.size == other.size;
}
/**
* Retrieves the opaque cookie for this paged results control.
*
* @return The opaque cookie for this paged results control.
*/
public String getCookie()
{
return this.cookie;
}
/**
* Retrieves the size for this paged results control, which is either the requested page size or the estimated result set size.
*
* @return The size for this paged results control.
*/
public int getSize()
{
return this.size;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + ( this.cookie == null ? 0 : this.cookie.hashCode() );
result = ( prime * result ) + this.size;
return result;
}
/**
* Setting the opaque cookie for this paged results control.
*
* @param cookie
* The opaque cookie for this paged results control.
*/
public void setCookie( final String cookie )
{
this.cookie = cookie;
}
/**
* Setting the size for this paged results control, which is either the requested page size or the estimated result set size.
*
* @param size
* The size for this paged results control.
*/
public void setSize( final int size )
{
this.size = size;
}
/**
* Retrieves a string representation of this control with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this control with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( "LDAP Simple Paged Results Control" ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " OID: " ).append( this.getControlOID() ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Criticality: " ).append( this.isCritical() ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Size: " ).append( this.size ).append( LDAPMessage.EOL );
buffer.append( indentBuf ).append( " Cookie: " ).append( LDAPMessage.EOL );
if ( this.cookie != null )
{
buffer.append( indentBuf ).append( " " ).append( this.cookie );
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + ( this.cookie == null ? 0 : this.cookie.hashCode() );
this.checksum = ( prime * this.checksum ) + this.size;
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/DeleteResponse.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import java.util.List;
import javax.persistence.Cacheable;
import javax.persistence.ElementCollection;
import javax.persistence.Entity;
import javax.persistence.FetchType;
import org.hibernate.annotations.Fetch;
import org.hibernate.annotations.FetchMode;
/**
* This class defines an LDAP delete response, which provides information about the result of processing a delete request.
*
* @author <NAME>
*/
@Entity
@Cacheable
public class DeleteResponse extends ProtocolOp
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The error message associated with this result.
*/
private String errorMessage;
/**
* The matched DN for this result.
*/
private String matchedDN;
/**
* The set of referrals for this result.
*/
@ElementCollection( fetch = FetchType.EAGER )
@Fetch( FetchMode.SELECT )
private List<String> referrals;
/**
* The result code for the operation.
*/
private int resultCode;
/**
* Creates a new delete response protocol op with the provided information.
*/
public DeleteResponse()
{
}
/**
* Creates a new delete response protocol op with the provided information.
*
* @param resultCode
* The result code for this response.
* @param matchedDN
* The matched DN for this response.
* @param errorMessage
* The error message for this response.
*/
public DeleteResponse( final int resultCode, final String matchedDN, final String errorMessage )
{
this.resultCode = resultCode;
this.matchedDN = matchedDN;
this.errorMessage = errorMessage;
this.referrals = null;
}
/**
* Creates a new delete response protocol op with the provided information.
*
* @param resultCode
* The result code for this response.
* @param matchedDN
* The matched DN for this response.
* @param errorMessage
* The error message for this response.
* @param referrals
* The set of referrals for this response.
*/
public DeleteResponse( final int resultCode, final String matchedDN, final String errorMessage, final List<String> referrals )
{
this.resultCode = resultCode;
this.matchedDN = matchedDN;
this.errorMessage = errorMessage;
this.referrals = referrals;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final DeleteResponse other = ( DeleteResponse ) obj;
if ( this.errorMessage == null )
{
if ( other.errorMessage != null )
{
return false;
}
} else if ( !this.errorMessage.equals( other.errorMessage ) )
{
return false;
}
if ( this.matchedDN == null )
{
if ( other.matchedDN != null )
{
return false;
}
} else if ( !this.matchedDN.equals( other.matchedDN ) )
{
return false;
}
if ( this.referrals == null )
{
if ( other.referrals != null )
{
return false;
}
} else if ( this.referrals.size() != other.referrals.size() )
{
return false;
} else
{
for ( final String uniqueStr : this.referrals )
{
if ( !other.referrals.contains( uniqueStr ) )
{
return false;
}
}
}
return this.resultCode == other.resultCode;
}
/**
* Retrieves the error message for this result.
*
* @return The error message for this result.
*/
public String getErrorMessage()
{
return this.errorMessage;
}
/**
* Retrieves the matched DN for this result.
*
* @return The matched DN for this result.
*/
public String getMatchedDN()
{
return this.matchedDN;
}
/**
* Retrieves the set of referrals for this result.
*
* @return The set of referrals for this result, or <CODE>null</CODE> if there were no referrals contained in the result.
*/
public List<String> getReferrals()
{
return this.referrals;
}
/**
* Retrieves the result code for the operation.
*
* @return The result code for the operation.
*/
public int getResultCode()
{
return this.resultCode;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + ( this.errorMessage == null ? 0 : this.errorMessage.hashCode() );
result = ( prime * result ) + ( this.matchedDN == null ? 0 : this.matchedDN.hashCode() );
for ( final String str : this.referrals )
{
result += str.hashCode();
}
result = ( prime * result ) + this.resultCode;
return result;
}
/**
* Retrieves a user-friendly name for this protocol op.
*
* @return A user-friendly name for this protocol op.
*/
@Override
public String retrieveProtocolOpType()
{
return "LDAP Delete Response";
}
/**
* Setting the error message for this result.
*
* @param errorMessage
* The error message for this result.
*/
public void setErrorMessage( final String errorMessage )
{
this.errorMessage = errorMessage;
}
/**
* Setting the matched DN for this result.
*
* @param matchedDN
* The matched DN for this result.
*/
public void setMatchedDN( final String matchedDN )
{
this.matchedDN = matchedDN;
}
/**
* Setting the set of referrals for this result.
*
* @param referrals
* The set of referrals for this result, or <CODE>null</CODE> if there were no referrals contained in the result.
*/
public void setReferrals( final List<String> referrals )
{
this.referrals = referrals;
}
/**
* Setting the result code for the operation.
*
* @param resultCode
* The result code for the operation.
*/
public void setResultCode( final int resultCode )
{
this.resultCode = resultCode;
}
/**
* Retrieves a string representation of this protocol op with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this protocol op with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
buffer.append( indentBuf ).append( "Result Code: " ).append( this.resultCode ).append( " (" ).append( LDAPResultCode.resultCodeToString( this.resultCode ) ).append( ')' ).append( LDAPMessage.EOL );
if ( ( this.matchedDN != null ) && !this.matchedDN.isEmpty() )
{
buffer.append( indentBuf ).append( "Matched DN: " ).append( this.matchedDN ).append( LDAPMessage.EOL );
}
if ( ( this.errorMessage != null ) && !this.errorMessage.isEmpty() )
{
buffer.append( indentBuf ).append( "Error Message: " ).append( this.errorMessage ).append( LDAPMessage.EOL );
}
if ( ( this.referrals != null ) && !this.referrals.isEmpty() )
{
buffer.append( indentBuf ).append( "Referrals:" ).append( LDAPMessage.EOL );
for ( final String referral : this.referrals )
{
buffer.append( indentBuf ).append( " " ).append( referral ).append( LDAPMessage.EOL );
}
}
return buffer.toString();
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + ( this.errorMessage == null ? 0 : this.errorMessage.hashCode() );
this.checksum = ( prime * this.checksum ) + ( this.matchedDN == null ? 0 : this.matchedDN.hashCode() );
for ( final String str : this.referrals )
{
this.checksum = ( prime * this.checksum ) + str.hashCode();
}
this.checksum = ( prime * this.checksum ) + this.resultCode;
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/asn1/ASN1Set.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1;
import java.util.ArrayList;
/**
* This class defines an ASN.1 element that serves as a set, whose value is an unordered set of other ASN.1 elements.
*
* @author <NAME>
*/
public class ASN1Set extends ASN1Element
{
/**
* The set of elements that will be used if there are no elements.
*/
private static final ASN1Element[] NO_ELEMENTS = new ASN1Element[ 0 ];
/**
* The set of ASN.1 elements associated with this set.
*/
ASN1Element[] elements;
/**
* Creates a new ASN.1 sequence with the given type to hold the specified set of ASN.1 elements.
*
* @param type
* The type to use for this ASN.1 sequence.
* @param elements
* The set of ASN.1 elements to encode in the value of this sequence.
*/
private ASN1Set( final byte type, final ArrayList<ASN1Element> elements )
{
super( type );
if ( ( elements == null ) || elements.isEmpty() )
{
this.elements = ASN1Set.NO_ELEMENTS;
} else
{
final ASN1Element[] elementArray = new ASN1Element[ elements.size() ];
elements.toArray( elementArray );
this.replaceElements( elementArray );
}
}
/**
* Creates a new ASN.1 set with the given type to hold the specified set of ASN.1 elements.
*
* @param type
* The type to use for this ASN.1 set.
* @param elements
* The set of ASN.1 elements to encode in the value of this set.
*/
private ASN1Set( final byte type, final ASN1Element[] elements )
{
super( type );
this.elements = elements == null ? ASN1Set.NO_ELEMENTS : elements;
int totalLength = 0;
final byte[][] encodedValues = new byte[ this.elements.length ][];
for ( int i = 0; i < this.elements.length; i++ )
{
encodedValues[ i ] = this.elements[ i ].encode();
totalLength += encodedValues[ i ].length;
}
final byte[] encodedValue = new byte[ totalLength ];
int startPos = 0;
for ( int i = 0; i < this.elements.length; i++ )
{
System.arraycopy( encodedValues[ i ], 0, encodedValue, startPos, encodedValues[ i ].length );
startPos += encodedValues[ i ].length;
}
this.setValue( encodedValue );
}
/**
* Creates a new ASN.1 set with no elements encoded in the value.
*/
public ASN1Set()
{
this( ASN1Element.ASN1_SET_TYPE, ASN1Set.NO_ELEMENTS );
}
/**
* Creates a new ASN.1 sequence to hold the specified set of ASN.1 elements.
*
* @param elements
* The set of ASN.1 elements to encode in the value of this sequence.
*/
public ASN1Set( final ArrayList<ASN1Element> elements )
{
this( ASN1Element.ASN1_SEQUENCE_TYPE, elements );
}
/**
* Creates a new ASN.1 set to hold the specified set of ASN.1 elements.
*
* @param elements
* The set of ASN.1 elements to encode in the value of this set.
*/
public ASN1Set( final ASN1Element[] elements )
{
this( ASN1Element.ASN1_SET_TYPE, elements );
}
/**
* Creates a new ASN.1 set with the specified type and no elements encoded in the value.
*
* @param type
* The type to use for this ASN.1 set.
*/
public ASN1Set( final byte type )
{
this( type, ASN1Set.NO_ELEMENTS );
}
/**
* Replaces the current set of elements with the provided set.
*
* @param elementsTmp
* The set of ASN.1 elements to use to replace the existing set of elements encoded in this set.
*/
private void replaceElements( final ASN1Element[] elementsTmp )
{
this.elements = elementsTmp == null ? ASN1Set.NO_ELEMENTS : elementsTmp;
// Re-encode the data in this set.
int totalLength = 0;
final byte[][] encodedValues = new byte[ this.elements.length ][];
for ( int i = 0; i < this.elements.length; i++ )
{
encodedValues[ i ] = this.elements[ i ].encode();
totalLength += encodedValues[ i ].length;
}
final byte[] encodedValue = new byte[ totalLength ];
int startPos = 0;
for ( int i = 0; i < this.elements.length; i++ )
{
System.arraycopy( encodedValues[ i ], 0, encodedValue, startPos, encodedValues[ i ].length );
startPos += encodedValues[ i ].length;
}
this.setValue( encodedValue );
}
/**
* Adds the specified ASN.1 element to the set of elements encoded in this set.
*
* @param element
* The ASN.1 element to include in the set of elements encoded in this set.
*/
public void addElement( final ASN1Element element )
{
this.elements = ASN1Element.addElementToSet( this.elements, element );
this.value = this.appendNewElementsByteToValue( element );
}
/**
* Retrieves the set of elements that are encoded in this ASN.1 set.
*
* @return The set of elements that are encoded in this ASN.1 set.
*/
public ASN1Element[] getElements()
{
return this.elements;
}
/**
* Removes all elements encoded in the value of this ASN.1 element.
*/
public void removeAllElements()
{
this.setValue( ASN1Element.EMPTY_BYTES );
this.elements = ASN1Set.NO_ELEMENTS;
}
/**
* Replaces the current set of elements with the provided ASN.1 element.
*
* @param element
* The ASN.1 element to use to replace the existing set of elements encoded in this set.
*/
public void replaceElements( final ASN1Element element )
{
this.replaceElements( new ASN1Element[] { element } );
}
/**
* Retrieves a string representation of this ASN.1 set. It will recursively display string representations for each of the elements.
*
* @param indent
* The number of spaces to indent the information in the returned string.
* @return A string representation of this ASN.1 set
*/
@Override( )
public String toString( final int indent )
{
String indentStr = "";
for ( int i = 0; i < indent; i++ )
{
indentStr += " ";
}
String elementsStr = "";
for ( int i = 0; i < this.elements.length; i++ )
{
final ASN1Element element = this.elements[ i ];
elementsStr += indentStr + " Element " + i + ASN1Element.EOL + element.toString( indent + 2 );
}
return indentStr + "Type: " + this.type + ASN1Element.EOL + ASN1Element.byteArrayToString( new byte[] { this.type }, 2 + indent ) + ASN1Element.EOL + indentStr + "Length: " + this.value.length + ASN1Element.EOL + ASN1Element.byteArrayToString( ASN1Element.encodeLength( this.value.length ), 2 + indent ) + ASN1Element.EOL + indentStr + "Value: " + new String( this.value ) + ASN1Element.EOL + ASN1Element.byteArrayToString( this.value, 2 + indent ) + ASN1Element.EOL + "Elements: " + ASN1Element.EOL + elementsStr;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/ProxiedAuthV1Control.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1Element;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1OctetString;
/**
* This class defines the first version of the proxied authorization control, which is used to perform an operation under the authority of one user while authenticated as another. Although it was replaced by a second version, the original version is still in use in some cases.
*
* @author <NAME>
*/
@Entity @Cacheable
public class ProxiedAuthV1Control extends LDAPControl
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The OID of the proxied auth v1 control.
*/
public static final String PROXIED_AUTH_V1_CONTROL_OID = "2.16.840.1.113730.3.4.12";
/**
* The DN of the user whose authority the requested operation should be performed.
*/
private String proxyDN;
/**
* Creates a new proxied auth v1 control with the provided information.
*/
public ProxiedAuthV1Control()
{
super( ProxiedAuthV1Control.PROXIED_AUTH_V1_CONTROL_OID );
}
/**
* Creates a new proxied auth v1 control by decoding the provided value.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param controlValue
* The encoded value for this control.
* @throws ProtocolException
* If a problem occurs while decoding the value for the control.
*/
public ProxiedAuthV1Control( final boolean isCritical, final ASN1OctetString controlValue ) throws ProtocolException
{
super( ProxiedAuthV1Control.PROXIED_AUTH_V1_CONTROL_OID, isCritical );
final ASN1Element[] elements;
try
{
final byte[] valueBytes = controlValue.getValue();
elements = ASN1Element.decodeAsSequence( valueBytes ).getElements();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode proxied auth v1 value sequence", e );
}
if ( elements.length != 1 )
{
throw new ProtocolException( "There must be exactly one element in a proxied auth v1 value sequence" );
}
try
{
this.proxyDN = elements[ 0 ].decodeAsOctetString().getStringValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode the proxy DN from the proxied authorization control", e );
}
}
/**
* Creates a new proxied auth v1 control with the provided information.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param proxyDN
* The DN of the user under whose authority the requested operation should be performed.
*/
public ProxiedAuthV1Control( final boolean isCritical, final String proxyDN )
{
super( ProxiedAuthV1Control.PROXIED_AUTH_V1_CONTROL_OID, isCritical );
this.proxyDN = proxyDN;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final ProxiedAuthV1Control other = ( ProxiedAuthV1Control ) obj;
if ( this.proxyDN == null )
{
if ( other.proxyDN != null )
{
return false;
}
} else if ( !this.proxyDN.equals( other.proxyDN ) )
{
return false;
}
return true;
}
/**
* Retrieves the proxy DN for this proxied authorization control.
*
* @return The proxy DN for this proxied authorization control.
*/
public String getProxyDN()
{
return this.proxyDN;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + ( this.proxyDN == null ? 0 : this.proxyDN.hashCode() );
return result;
}
/**
* Setting the proxy DN for this proxied authorization control.
*
* @param proxyDN
* The proxy DN for this proxied authorization control.
*/
public void setProxyDN( final String proxyDN )
{
this.proxyDN = proxyDN;
}
/**
* Retrieves a string representation of this control with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this control with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
return String.valueOf( indentBuf ) + "LDAP Proxied Authorization (v1) Control" + LDAPMessage.EOL + indentBuf + " OID: " + this.getControlOID() + LDAPMessage.EOL + indentBuf + " Criticality: " + this.isCritical() + LDAPMessage.EOL + indentBuf + " Proxy DN: " + this.proxyDN + LDAPMessage.EOL;
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + ( this.proxyDN == null ? 0 : this.proxyDN.hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/functions/dao/OpenConnectionRequestDAO.java
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.functions.dao;
import java.util.Date;
import java.util.HashMap;
import java.util.List;
import java.util.Map;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.database.jpa.exception.JpaFacilitatorException;
import ita.deluca.tools.application.framework.database.jpa.model.JpaFacilitator;
import ita.deluca.tools.ldap.framework.commons.bean.OpenConnectionRequest;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.exception.LdapEntityDaoException;
import ita.deluca.tools.ldap.framework.properties.Constants;
/**
* This class contains all the function that allows interaction with the database via JPA.
*
* @author <NAME>
*/
public class OpenConnectionRequestDAO
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( OpenConnectionRequestDAO.class );
/**
* This method persist the object into the database.
*
* @param entity
* the object to persist.
* @return The persisted entity.
* @throws LdapEntityDaoException
* this exception was raised whenever an error occurred during the persist of the object.
*/
protected static OpenConnectionRequest persist( final OpenConnectionRequest entity ) throws LdapEntityDaoException
{
if ( OpenConnectionRequestDAO.LOG.isDebugEnabled() )
{
OpenConnectionRequestDAO.LOG.debug( "Persist the entity:\n" + entity );
}
final OpenConnectionRequest persistedEntity = LDAPEntityDAO.getAlreadyPersistEntity( entity );
if ( persistedEntity != null )
{
return persistedEntity;
}
return LDAPEntityDAO.persistEntity( entity );
}
/**
* Extract the id of the connection that was opened by a server in a certain period of time.
*
* @param clientHost
* The host that make the open connection request.
* @param ldapHost
* The ldap host that receive the open connection command.
* @param fromDate
* The search start date.
* @param toDate
* The search end date.
* @return The list of the message having the same connection id.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the message.
*/
public static List<String> getConnectionID( final String clientHost, final String ldapHost, final Date fromDate, final Date toDate ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select message.connectionId from LDAPMessage message, OpenConnectionRequest request where (message.protocolOp = request and message.sourceHost = :clientHost and message.destHost = :ldapHost and message.timestamp between :fromDate and :toDate)";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "clientHost", clientHost );
parameter.put( "ldapHost", ldapHost );
parameter.put( "fromDate", fromDate );
parameter.put( "toDate", toDate );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the connection id of the connection that having source host '" + clientHost + "' and ldap host '" + ldapHost + "' thrown an error !!!";
OpenConnectionRequestDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
final List<String> retRes = ( List<String> ) res;
return retRes;
}
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/asn1/ASN1DecodeResult.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1;
/**
* This class provides a data structure that can hold information about the result of attempting to decode a byte array as an ASN.1 element. If the array contained at least a complete ASN.1 element, then that element will be available, along with any remaining data (if there was more data beyond the end of the element). It is also possible to use this result to determine if there was not enough data in the original array to hold a complete element.
*/
public final class ASN1DecodeResult
{
/**
* The ASN.1 element that was decoded.
*/
private final ASN1Element decodedElement;
/**
* The size of the data message.
*/
private final int length;
/**
* The data left over after decoding the element.
*/
private final byte[] remainingData;
/**
* Creates a new ASN.1 decode result with the provided information.
*
* @param decodedElement
* The ASN.1 element that was decoded. This should be {@code null} if the associated byte array did not contain a complete element.
* @param remainingData
* A byte array containing any data from the associated byte array that were left over after decoding the element. This should be {@code null} if the associated byte array did not contain a complete element, or if there were no bytes left over.
* @param length
* The size of the data message.
*/
ASN1DecodeResult( final ASN1Element decodedElement, final byte[] remainingData, final int length )
{
this.decodedElement = decodedElement;
this.remainingData = remainingData;
this.length = length;
}
/**
* Retrieves the ASN.1 element decoded from the original byte array, if available.
*
* @return The ASN.1 element decoded from the original byte array, or {@code null} if the array did not contain a complete ASN.1 element.
*/
public ASN1Element getDecodedElement()
{
return this.decodedElement;
}
/**
* Retrieves the size of the data message.
*
* @return The size of the data message.
*/
public int getLength()
{
return this.length;
}
/**
* Retrieves a byte array containing data that was left over after decoding the ASN.1 element, if any.
*
* @return A byte array containing data that was left over after decoding the ASN.1 element, or {@code null} if the array did not contain a complete ASN.1 element, or if there were no bytes left over after decoding the element.
*/
public byte[] getRemainingData()
{
return this.remainingData;
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/asn1/ASN1Integer.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1;
/**
* This class defines an ASN.1 element that can hold an integer value.
*
* @author <NAME>
*/
public class ASN1Integer extends ASN1Element
{
/**
* The integer value 0 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_0 = new byte[] { 0x00 };
/**
* The integer value 1 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_1 = new byte[] { 0x01 };
/**
* The integer value 10 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_10 = new byte[] { 0x0A };
/**
* The integer value 100 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_100 = new byte[] { 0x64 };
/**
* The integer value 101 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_101 = new byte[] { 0x65 };
/**
* The integer value 102 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_102 = new byte[] { 0x66 };
/**
* The integer value 103 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_103 = new byte[] { 0x67 };
/**
* The integer value 104 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_104 = new byte[] { 0x68 };
/**
* The integer value 105 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_105 = new byte[] { 0x69 };
/**
* The integer value 106 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_106 = new byte[] { 0x6A };
/**
* The integer value 107 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_107 = new byte[] { 0x6B };
/**
* The integer value 108 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_108 = new byte[] { 0x6C };
/**
* The integer value 109 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_109 = new byte[] { 0x6D };
/**
* The integer value 11 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_11 = new byte[] { 0x0B };
/**
* The integer value 110 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_110 = new byte[] { 0x6E };
/**
* The integer value 111 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_111 = new byte[] { 0x6F };
/**
* The integer value 112 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_112 = new byte[] { 0x70 };
/**
* The integer value 113 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_113 = new byte[] { 0x71 };
/**
* The integer value 114 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_114 = new byte[] { 0x72 };
/**
* The integer value 115 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_115 = new byte[] { 0x73 };
/**
* The integer value 116 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_116 = new byte[] { 0x74 };
/**
* The integer value 117 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_117 = new byte[] { 0x75 };
/**
* The integer value 118 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_118 = new byte[] { 0x76 };
/**
* The integer value 119 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_119 = new byte[] { 0x77 };
/**
* The integer value 12 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_12 = new byte[] { 0x0C };
/**
* The integer value 120 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_120 = new byte[] { 0x78 };
/**
* The integer value 121 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_121 = new byte[] { 0x79 };
/**
* The integer value 122 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_122 = new byte[] { 0x7A };
/**
* The integer value 123 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_123 = new byte[] { 0x7B };
/**
* The integer value 124 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_124 = new byte[] { 0x7C };
/**
* The integer value 125 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_125 = new byte[] { 0x7D };
/**
* The integer value 126 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_126 = new byte[] { 0x7E };
/**
* The integer value 127 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_127 = new byte[] { 0x7F };
/**
* The integer value 13 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_13 = new byte[] { 0x0D };
/**
* The integer value 14 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_14 = new byte[] { 0x0E };
/**
* The integer value 15 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_15 = new byte[] { 0x0F };
/**
* The integer value 16 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_16 = new byte[] { 0x10 };
/**
* The integer value 17 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_17 = new byte[] { 0x11 };
/**
* The integer value 18 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_18 = new byte[] { 0x12 };
/**
* The integer value 19 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_19 = new byte[] { 0x13 };
/**
* The integer value 2 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_2 = new byte[] { 0x02 };
/**
* The integer value 20 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_20 = new byte[] { 0x14 };
/**
* The integer value 21 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_21 = new byte[] { 0x15 };
/**
* The integer value 22 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_22 = new byte[] { 0x16 };
/**
* The integer value 23 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_23 = new byte[] { 0x17 };
/**
* The integer value 24 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_24 = new byte[] { 0x18 };
/**
* The integer value 25 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_25 = new byte[] { 0x19 };
/**
* The integer value 26 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_26 = new byte[] { 0x1A };
/**
* The integer value 27 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_27 = new byte[] { 0x1B };
/**
* The integer value 28 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_28 = new byte[] { 0x1C };
/**
* The integer value 29 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_29 = new byte[] { 0x1D };
/**
* The integer value 3 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_3 = new byte[] { 0x03 };
/**
* The integer value 30 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_30 = new byte[] { 0x1E };
/**
* The integer value 31 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_31 = new byte[] { 0x1F };
/**
* The integer value 32 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_32 = new byte[] { 0x20 };
/**
* The integer value 33 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_33 = new byte[] { 0x21 };
/**
* The integer value 34 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_34 = new byte[] { 0x22 };
/**
* The integer value 35 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_35 = new byte[] { 0x23 };
/**
* The integer value 36 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_36 = new byte[] { 0x24 };
/**
* The integer value 37 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_37 = new byte[] { 0x25 };
/**
* The integer value 38 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_38 = new byte[] { 0x26 };
/**
* The integer value 39 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_39 = new byte[] { 0x27 };
/**
* The integer value 4 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_4 = new byte[] { 0x04 };
/**
* The integer value 40 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_40 = new byte[] { 0x28 };
/**
* The integer value 41 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_41 = new byte[] { 0x29 };
/**
* The integer value 42 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_42 = new byte[] { 0x2A };
/**
* The integer value 43 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_43 = new byte[] { 0x2B };
/**
* The integer value 44 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_44 = new byte[] { 0x2C };
/**
* The integer value 45 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_45 = new byte[] { 0x2D };
/**
* The integer value 46 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_46 = new byte[] { 0x2E };
/**
* The integer value 47 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_47 = new byte[] { 0x2F };
/**
* The integer value 48 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_48 = new byte[] { 0x30 };
/**
* The integer value 49 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_49 = new byte[] { 0x31 };
/**
* The integer value 5 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_5 = new byte[] { 0x05 };
/**
* The integer value 50 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_50 = new byte[] { 0x32 };
/**
* The integer value 51 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_51 = new byte[] { 0x33 };
/**
* The integer value 52 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_52 = new byte[] { 0x34 };
/**
* The integer value 53 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_53 = new byte[] { 0x35 };
/**
* The integer value 54 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_54 = new byte[] { 0x36 };
/**
* The integer value 55 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_55 = new byte[] { 0x37 };
/**
* The integer value 56 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_56 = new byte[] { 0x38 };
/**
* The integer value 57 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_57 = new byte[] { 0x39 };
/**
* The integer value 58 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_58 = new byte[] { 0x3A };
/**
* The integer value 59 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_59 = new byte[] { 0x3B };
/**
* The integer value 6 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_6 = new byte[] { 0x06 };
/**
* The integer value 60 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_60 = new byte[] { 0x3C };
/**
* The integer value 61 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_61 = new byte[] { 0x3D };
/**
* The integer value 62 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_62 = new byte[] { 0x3E };
/**
* The integer value 63 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_63 = new byte[] { 0x3F };
/**
* The integer value 64 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_64 = new byte[] { 0x40 };
/**
* The integer value 65 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_65 = new byte[] { 0x41 };
/**
* The integer value 66 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_66 = new byte[] { 0x42 };
/**
* The integer value 67 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_67 = new byte[] { 0x43 };
/**
* The integer value 68 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_68 = new byte[] { 0x44 };
/**
* The integer value 69 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_69 = new byte[] { 0x45 };
/**
* The integer value 7 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_7 = new byte[] { 0x07 };
/**
* The integer value 70 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_70 = new byte[] { 0x46 };
/**
* The integer value 71 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_71 = new byte[] { 0x47 };
/**
* The integer value 72 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_72 = new byte[] { 0x48 };
/**
* The integer value 73 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_73 = new byte[] { 0x49 };
/**
* The integer value 74 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_74 = new byte[] { 0x4A };
/**
* The integer value 75 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_75 = new byte[] { 0x4B };
/**
* The integer value 76 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_76 = new byte[] { 0x4C };
/**
* The integer value 77 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_77 = new byte[] { 0x4D };
/**
* The integer value 78 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_78 = new byte[] { 0x4E };
/**
* The integer value 79 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_79 = new byte[] { 0x4F };
/**
* The integer value 8 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_8 = new byte[] { 0x08 };
/**
* The integer value 80 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_80 = new byte[] { 0x50 };
/**
* The integer value 81 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_81 = new byte[] { 0x51 };
/**
* The integer value 82 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_82 = new byte[] { 0x52 };
/**
* The integer value 83 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_83 = new byte[] { 0x53 };
/**
* The integer value 84 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_84 = new byte[] { 0x54 };
/**
* The integer value 85 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_85 = new byte[] { 0x55 };
/**
* The integer value 86 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_86 = new byte[] { 0x56 };
/**
* The integer value 87 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_87 = new byte[] { 0x57 };
/**
* The integer value 88 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_88 = new byte[] { 0x58 };
/**
* The integer value 89 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_89 = new byte[] { 0x59 };
/**
* The integer value 9 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_9 = new byte[] { 0x09 };
/**
* The integer value 90 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_90 = new byte[] { 0x5A };
/**
* The integer value 91 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_91 = new byte[] { 0x5B };
/**
* The integer value 92 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_92 = new byte[] { 0x5C };
/**
* The integer value 93 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_93 = new byte[] { 0x5D };
/**
* The integer value 94 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_94 = new byte[] { 0x5E };
/**
* The integer value 95 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_95 = new byte[] { 0x5F };
/**
* The integer value 96 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_96 = new byte[] { 0x60 };
/**
* The integer value 97 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_97 = new byte[] { 0x61 };
/**
* The integer value 98 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_98 = new byte[] { 0x62 };
/**
* The integer value 99 encoded as a byte array in the appropriate ASN.1 format.
*/
public static final byte[] INT_VALUE_99 = new byte[] { 0x63 };
/**
* The Java int value that corresponds to the value of this ASN.1 integer element
*/
private int intValue;
/**
* Creates a new ASN.1 integer element with the specified value.
*
* @param intValue
* The Java int value to use in creating this ASN.1 integer element.
*/
private ASN1Integer( final int intValue )
{
this( ASN1Element.ASN1_INTEGER_TYPE, intValue );
}
/**
* Creates a new ASN.1 integer element with the specified value.
*/
public ASN1Integer()
{
this( ASN1Element.ASN1_INTEGER_TYPE );
}
/**
* Creates a new ASN.1 integer element with the specified type and value.
*
* @param type
* The type to use for this ASN.1 integer value.
* @param intValue
* The Java int value to use in creating this ASN.1 integer element.
*/
public ASN1Integer( final byte type, final int intValue )
{
super( type );
this.setValue( ASN1Integer.encodeIntValue( intValue ) );
this.intValue = intValue;
}
/**
* Encodes the provided int value in the appropriate manner for an ASN.1 integer value.
*
* @param intValue
* The Java int value to encode as an ASN.1 integer value.
* @return A byte array that contains the encoded integer value.
*/
public static byte[] encodeIntValue( final int intValue )
{
// First, see if the int value is within the first 127 values. If so, then
// just return the pre-encoded version.
byte[] returnValue = ASN1Element.returnPreEncodedLengthVersion( intValue );
if ( returnValue != null )
{
return returnValue;
}
// It's greater than 127, so do it the "long" way.
if ( ( intValue & 0xFF800000 ) != 0 )
{
returnValue = new byte[ 4 ];
returnValue[ 0 ] = ( byte ) ( ( intValue & 0xFF000000 ) >>> 24 );
returnValue[ 1 ] = ( byte ) ( ( intValue & 0x00FF0000 ) >>> 16 );
returnValue[ 2 ] = ( byte ) ( ( intValue & 0x0000FF00 ) >>> 8 );
returnValue[ 3 ] = ( byte ) ( intValue & 0x000000FF );
return returnValue;
} else if ( ( intValue & 0x00FF8000 ) != 0 )
{
returnValue = new byte[ 3 ];
returnValue[ 0 ] = ( byte ) ( ( intValue & 0x00FF0000 ) >>> 16 );
returnValue[ 1 ] = ( byte ) ( ( intValue & 0x0000FF00 ) >>> 8 );
returnValue[ 2 ] = ( byte ) ( intValue & 0x000000FF );
return returnValue;
} else
{
returnValue = new byte[ 2 ];
returnValue[ 0 ] = ( byte ) ( ( intValue & 0x0000FF00 ) >>> 8 );
returnValue[ 1 ] = ( byte ) ( intValue & 0x000000FF );
return returnValue;
}
}
/**
* Retrieves the Java int that corresponds to the value of this ASN.1 integer value.
*
* @return The Java int that corresponds to the value of this ASN.1 integer value.
*/
public int getIntValue()
{
return this.intValue;
}
/**
* Setting the Java int that corresponds to the value of this ASN.1 integer value.
*
* @param intValue
* The Java int that corresponds to the value of this ASN.1 integer value.
*/
public void setIntValue( final int intValue )
{
this.setValue( ASN1Integer.encodeIntValue( intValue ) );
this.intValue = intValue;
}
}
<file_sep>/src/META-INF/Create_TcpDecoder.sql
DROP TABLE IF EXISTS ldap_tools.ldap_add_request_attributes_join;
DROP TABLE IF EXISTS ldap_tools.ldap_message_controls_join;
DROP TABLE IF EXISTS ldap_tools.ldap_modify_request_modifications_join;
DROP TABLE IF EXISTS ldap_tools.ldap_search_request_attributes_join;
DROP TABLE IF EXISTS ldap_tools.ldap_search_result_entry_attributes_join;
DROP TABLE IF EXISTS ldap_tools.ldap_server_sort_request_control_sort_keys_join;
DROP TABLE IF EXISTS ldap_tools.managedsaitcontrol;
DROP TABLE IF EXISTS ldap_tools.addresponse_referrals;
DROP TABLE IF EXISTS ldap_tools.bindresponse_referrals;
DROP TABLE IF EXISTS ldap_tools.compareresponse_referrals;
DROP TABLE IF EXISTS ldap_tools.deleteresponse_referrals;
DROP TABLE IF EXISTS ldap_tools.extendedresponse_referralurls;
DROP TABLE IF EXISTS ldap_tools.modifydnresponse_referrals;
DROP TABLE IF EXISTS ldap_tools.modifyresponse_referrals;
DROP TABLE IF EXISTS ldap_tools.searchresultdone_referrals;
DROP TABLE IF EXISTS ldap_tools.searchresultreference_referralurls;
DROP TABLE IF EXISTS ldap_tools.geteffectiverightscontrol_attributetypes;
DROP TABLE IF EXISTS ldap_tools.authorizationidresponsecontrol;
DROP TABLE IF EXISTS ldap_tools.entrychangenotificationcontrol;
DROP TABLE IF EXISTS ldap_tools.geteffectiverightscontrol;
DROP TABLE IF EXISTS ldap_tools.pagedresultscontrol;
DROP TABLE IF EXISTS ldap_tools.passwordexpiredcontrol;
DROP TABLE IF EXISTS ldap_tools.passwordexpiringcontrol;
DROP TABLE IF EXISTS ldap_tools.passwordpolicycontrol;
DROP TABLE IF EXISTS ldap_tools.persistentsearchcontrol;
DROP TABLE IF EXISTS ldap_tools.proxiedauthv1control;
DROP TABLE IF EXISTS ldap_tools.proxiedauthv2control;
DROP TABLE IF EXISTS ldap_tools.realattributesonlycontrol;
DROP TABLE IF EXISTS ldap_tools.serversortrequestcontrol;
DROP TABLE IF EXISTS ldap_tools.serversortresponsecontrol;
DROP TABLE IF EXISTS ldap_tools.vlvrequestcontrol;
DROP TABLE IF EXISTS ldap_tools.vlvresponsecontrol;
DROP TABLE IF EXISTS ldap_tools.abandonrequest;
DROP TABLE IF EXISTS ldap_tools.addrequest;
DROP TABLE IF EXISTS ldap_tools.authorizationidrequestcontrol;
DROP TABLE IF EXISTS ldap_tools.bindrequest;
DROP TABLE IF EXISTS ldap_tools.comparerequest;
DROP TABLE IF EXISTS ldap_tools.deleterequest;
DROP TABLE IF EXISTS ldap_tools.modifydnrequest;
DROP TABLE IF EXISTS ldap_tools.modifyrequest;
DROP TABLE IF EXISTS ldap_tools.openconnectionrequest;
DROP TABLE IF EXISTS ldap_tools.searchrequest;
DROP TABLE IF EXISTS ldap_tools.unbindrequest;
DROP TABLE IF EXISTS ldap_tools.addresponse;
DROP TABLE IF EXISTS ldap_tools.bindresponse;
DROP TABLE IF EXISTS ldap_tools.compareresponse;
DROP TABLE IF EXISTS ldap_tools.deleteresponse;
DROP TABLE IF EXISTS ldap_tools.extendedrequest;
DROP TABLE IF EXISTS ldap_tools.extendedresponse;
DROP TABLE IF EXISTS ldap_tools.intermediateresponse;
DROP TABLE IF EXISTS ldap_tools.modifydnresponse;
DROP TABLE IF EXISTS ldap_tools.modifyresponse;
DROP TABLE IF EXISTS ldap_tools.openconnectionresponse;
DROP TABLE IF EXISTS ldap_tools.ldap_message;
DROP TABLE IF EXISTS ldap_tools.ldap_modification;
DROP TABLE IF EXISTS ldap_tools.ldap_server_sort_key;
DROP TABLE IF EXISTS ldap_tools.searchresultdone;
DROP TABLE IF EXISTS ldap_tools.searchresultentry;
DROP TABLE IF EXISTS ldap_tools.searchresultreference;
DROP TABLE IF EXISTS ldap_tools.ldap_protocol_operation;
DROP TABLE IF EXISTS ldap_tools.ldap_control;
CREATE TABLE ldap_tools.abandonrequest (
idtoabandon int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT abandonrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.addrequest (
dn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT addrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.addresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT addresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.addresponse_referrals (
addresponse_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.authorizationidrequestcontrol (
id bigserial NOT NULL,
CONSTRAINT authorizationidrequestcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.authorizationidresponsecontrol (
authzid varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT authorizationidresponsecontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.bindrequest (
authtype int2 NOT NULL,
basedn varchar(255) NULL,
binddn varchar(255) NULL,
bindpassword varchar(255) NULL,
protocolversion int4 NOT NULL,
saslcredentials varchar(255) NULL,
saslmechanism varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT bindrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.bindresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
serversaslcredentials varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT bindresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.bindresponse_referrals (
bindresponse_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.comparerequest (
assertionvalue varchar(255) NULL,
attributetype varchar(255) NULL,
dn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT comparerequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.compareresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT compareresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.compareresponse_referrals (
compareresponse_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.deleterequest (
dn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT deleterequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.deleteresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT deleteresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.deleteresponse_referrals (
deleteresponse_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.entrychangenotificationcontrol (
changenumber int4 NOT NULL,
changetype int4 NOT NULL,
previousdn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT entrychangenotificationcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.extendedrequest (
requestoid varchar(255) NULL,
requestvalue varchar(5120) NULL,
id bigserial NOT NULL,
CONSTRAINT extendedrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.extendedresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
responseoid varchar(255) NULL,
responsevalue varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT extendedresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.extendedresponse_referralurls (
extendedresponse_id int8 NOT NULL,
referralurls varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.geteffectiverightscontrol (
authzid varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT geteffectiverightscontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.geteffectiverightscontrol_attributetypes (
geteffectiverightscontrol_id int8 NOT NULL,
attributetypes varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.intermediateresponse (
responseoid varchar(255) NULL,
responsevalue varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT intermediateresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_add_request_attributes_join (
addrequest_id int8 NOT NULL,
attributes_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_control (
id bigserial NOT NULL,
checksum int4 NOT NULL,
controloid varchar(255) NULL,
critical bool NOT NULL,
CONSTRAINT ldap_control_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_message (
id bigserial NOT NULL,
checksum int4 NOT NULL,
capturehost varchar(255) NULL,
connectionid varchar(255) NULL,
desthost varchar(255) NULL,
destport int4 NOT NULL,
messageid int4 NOT NULL,
messagelength int4 NOT NULL,
sourcehost varchar(255) NULL,
sourceport int4 NOT NULL,
"timestamp" timestamp NULL,
protocolop_id int8 NULL,
CONSTRAINT ldap_message_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_message_controls_join (
ldapmessage_id int8 NOT NULL,
controls_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_modification (
id bigserial NOT NULL,
checksum int4 NOT NULL,
modtype int4 NOT NULL,
attribute_id int8 NULL,
CONSTRAINT ldap_modification_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_modify_request_modifications_join (
modifyrequest_id int8 NOT NULL,
modifications_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_protocol_operation (
id bigserial NOT NULL,
checksum int4 NOT NULL,
CONSTRAINT ldap_protocol_operation_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_search_request_attributes_join (
searchrequest_id int8 NOT NULL,
attributes_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_search_result_entry_attributes_join (
searchresultentry_id int8 NOT NULL,
attributes_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_server_sort_key (
id bigserial NOT NULL,
checksum int4 NOT NULL,
attributetype varchar(255) NULL,
matchingruleid varchar(255) NULL,
reverseorder bool NOT NULL,
CONSTRAINT ldap_server_sort_key_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.ldap_server_sort_request_control_sort_keys_join (
serversortrequestcontrol_id int8 NOT NULL,
sortkeys_id int8 NOT NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.managedsaitcontrol (
id bigserial NOT NULL,
CONSTRAINT managedsaitcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.modifydnrequest (
deleteoldrdn bool NOT NULL,
dn varchar(255) NULL,
newrdn varchar(255) NULL,
newsuperior varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT modifydnrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.modifydnresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT modifydnresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.modifydnresponse_referrals (
modifydnresponse_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.modifyrequest (
dn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT modifyrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.modifyresponse (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT modifyresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.modifyresponse_referrals (
modifyresponse_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.openconnectionrequest (
id bigserial NOT NULL,
CONSTRAINT openconnectionrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.openconnectionresponse (
id bigserial NOT NULL,
CONSTRAINT openconnectionresponse_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.pagedresultscontrol (
cookie varchar(255) NULL,
"size" int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT pagedresultscontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.passwordexpiredcontrol (
id bigserial NOT NULL,
CONSTRAINT passwordexpiredcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.passwordexpiringcontrol (
secondsuntilexpiration int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT passwordexpiringcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.passwordpolicycontrol (
controltype int4 NOT NULL,
errortype int4 NOT NULL,
warningtype int2 NOT NULL,
warningvalue int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT passwordpolicycontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.persistentsearchcontrol (
changecontrols bool NOT NULL,
changetypes int4 NOT NULL,
changesonly bool NOT NULL,
id bigserial NOT NULL,
CONSTRAINT persistentsearchcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.proxiedauthv1control (
proxydn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT proxiedauthv1control_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.proxiedauthv2control (
authzid varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT proxiedauthv2control_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.realattributesonlycontrol (
id bigserial NOT NULL,
CONSTRAINT realattributesonlycontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.searchrequest (
basedn varchar(255) NULL,
derefpolicy int4 NOT NULL,
"filter" varchar(5120) NULL,
"scope" int4 NOT NULL,
sizelimit int4 NOT NULL,
timelimit int4 NOT NULL,
typesonly bool NOT NULL,
id bigserial NOT NULL,
CONSTRAINT searchrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.searchresultdone (
errormessage varchar(255) NULL,
matcheddn varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT searchresultdone_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.searchresultdone_referrals (
searchresultdone_id int8 NOT NULL,
referrals varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.searchresultentry (
dn varchar(255) NULL,
id bigserial NOT NULL,
CONSTRAINT searchresultentry_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.searchresultreference (
id bigserial NOT NULL,
CONSTRAINT searchresultreference_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.searchresultreference_referralurls (
searchresultreference_id int8 NOT NULL,
referralurls varchar(255) NULL
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.serversortrequestcontrol (
id bigserial NOT NULL,
CONSTRAINT serversortrequestcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.serversortresponsecontrol (
attributetype varchar(255) NULL,
resultcode int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT serversortresponsecontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.unbindrequest (
id bigserial NOT NULL,
CONSTRAINT unbindrequest_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.vlvrequestcontrol (
aftercount int4 NOT NULL,
assertionvalue varchar(255) NULL,
beforecount int4 NOT NULL,
contentcount int4 NOT NULL,
entryoffset int4 NOT NULL,
selecttype int2 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT vlvrequestcontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
CREATE TABLE ldap_tools.vlvresponsecontrol (
contentcount int4 NOT NULL,
resultcode int4 NOT NULL,
targetposition int4 NOT NULL,
id bigserial NOT NULL,
CONSTRAINT vlvresponsecontrol_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
);
ALTER TABLE ldap_tools.bindresponse_referrals ADD CONSTRAINT fkr04v53kw5ryojs6cwndnd5wvm FOREIGN KEY (bindresponse_id) REFERENCES ldap_tools.bindresponse(id);
ALTER TABLE ldap_tools.geteffectiverightscontrol ADD CONSTRAINT fk8fq5rh7203xn0hiipenxk5500 FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.geteffectiverightscontrol_attributetypes ADD CONSTRAINT fkf13o5y5n8rdp1dimu0xbods78 FOREIGN KEY (geteffectiverightscontrol_id) REFERENCES ldap_tools.geteffectiverightscontrol(id);
ALTER TABLE ldap_tools.intermediateresponse ADD CONSTRAINT fk1nbv0kk5416437ixn2aw1793j FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.ldap_add_request_attributes_join ADD CONSTRAINT fkbwds2nm6x53vg5nwt3802g2fy FOREIGN KEY (attributes_id) REFERENCES ldap_tools.ldap_attribute(id);
ALTER TABLE ldap_tools.ldap_add_request_attributes_join ADD CONSTRAINT fknjevel8td0wf7g5sbv5igaher FOREIGN KEY (addrequest_id) REFERENCES ldap_tools.addrequest(id);
ALTER TABLE ldap_tools.ldap_message ADD CONSTRAINT fka6ojhjo398xftv2m854ogq1sr FOREIGN KEY (protocolop_id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.ldap_message_controls_join ADD CONSTRAINT fkclw5f78g7abafwrbcs09lbwoy FOREIGN KEY (ldapmessage_id) REFERENCES ldap_tools.ldap_message(id);
ALTER TABLE ldap_tools.ldap_message_controls_join ADD CONSTRAINT fkoaaramyckfxelbk1m7e07y2n2 FOREIGN KEY (controls_id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.ldap_modification ADD CONSTRAINT fke5238yrtxeicsd0a78owfu0yt FOREIGN KEY (attribute_id) REFERENCES ldap_tools.ldap_attribute(id);
ALTER TABLE ldap_tools.ldap_modify_request_modifications_join ADD CONSTRAINT fkelgqgmmp15q6umg0e6uxaf0ir FOREIGN KEY (modifyrequest_id) REFERENCES ldap_tools.modifyrequest(id);
ALTER TABLE ldap_tools.ldap_modify_request_modifications_join ADD CONSTRAINT fksc1i6dmxlt7if50hlx709cbxe FOREIGN KEY (modifications_id) REFERENCES ldap_tools.ldap_modification(id);
ALTER TABLE ldap_tools.ldap_search_request_attributes_join ADD CONSTRAINT fkdk90oncj80n3um7tcv3bgg30e FOREIGN KEY (searchrequest_id) REFERENCES ldap_tools.searchrequest(id);
ALTER TABLE ldap_tools.ldap_search_request_attributes_join ADD CONSTRAINT fkh7771527e9x2m7avcxt9ge623 FOREIGN KEY (attributes_id) REFERENCES ldap_tools.ldap_attribute(id);
ALTER TABLE ldap_tools.ldap_search_result_entry_attributes_join ADD CONSTRAINT fk208uhvt0s5syj4s95esegdks9 FOREIGN KEY (attributes_id) REFERENCES ldap_tools.ldap_attribute(id);
ALTER TABLE ldap_tools.ldap_search_result_entry_attributes_join ADD CONSTRAINT fkbuqdvmpiagawd4e3ws4hq7bfy FOREIGN KEY (searchresultentry_id) REFERENCES ldap_tools.searchresultentry(id);
ALTER TABLE ldap_tools.ldap_server_sort_request_control_sort_keys_join ADD CONSTRAINT fka0dvn4apuvareh41drwiwmb6j FOREIGN KEY (serversortrequestcontrol_id) REFERENCES ldap_tools.serversortrequestcontrol(id);
ALTER TABLE ldap_tools.ldap_server_sort_request_control_sort_keys_join ADD CONSTRAINT fklobfye0r0fs2awdcy3ygik82y FOREIGN KEY (sortkeys_id) REFERENCES ldap_tools.ldap_server_sort_key(id);
ALTER TABLE ldap_tools.managedsaitcontrol ADD CONSTRAINT fko4qaeeojxlrbwcat2dqwtyw2d FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.modifydnrequest ADD CONSTRAINT fks9qplajmj8oj2lwct8lk4uqdu FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.modifydnresponse ADD CONSTRAINT fkahdun65579lr03d1pgrwpuc6c FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.modifydnresponse_referrals ADD CONSTRAINT fkayqhi74vxifbvmitbwipg1vy3 FOREIGN KEY (modifydnresponse_id) REFERENCES ldap_tools.modifydnresponse(id);
ALTER TABLE ldap_tools.modifyrequest ADD CONSTRAINT fktf0wc3xf7wm9oid2gfbbtdxjn FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.modifyresponse ADD CONSTRAINT fkm5emygjqd329mdn5yo72qc2qf FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.modifyresponse_referrals ADD CONSTRAINT fk8ay6lhs4ggqbdmbpx7cge7awf FOREIGN KEY (modifyresponse_id) REFERENCES ldap_tools.modifyresponse(id);
ALTER TABLE ldap_tools.openconnectionrequest ADD CONSTRAINT fkknpk163mn99gdfb2gxux9wwy0 FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.openconnectionresponse ADD CONSTRAINT fkom3crn10pdvngqo7eptrumkos FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.pagedresultscontrol ADD CONSTRAINT fksopvnexvw89jfeov4qv9ngqyu FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.passwordexpiredcontrol ADD CONSTRAINT fkeeatk48ep783ec7mul06gu4b9 FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.passwordexpiringcontrol ADD CONSTRAINT fk1f5guhfwt1ql5xf98c7rxb21d FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.passwordpolicycontrol ADD CONSTRAINT fkkqxq0yjdtcr8djnt6spjx9dj9 FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.persistentsearchcontrol ADD CONSTRAINT fk3qcd6w82x8nh4o17eqo5gw512 FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.proxiedauthv1control ADD CONSTRAINT fkirifsu5b1dn3gtq5r8de4hcmx FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.proxiedauthv2control ADD CONSTRAINT fkh5uqyvqxfdqi8q5yj1qhn8c9w FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.realattributesonlycontrol ADD CONSTRAINT fkbym2voq6vj4xp55juha35pxpy FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.searchrequest ADD CONSTRAINT fknnhokocu08u0cn33lgqdsulhn FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.searchresultdone ADD CONSTRAINT fk5nvgexk36emq74sm3tltnxyyt FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.searchresultdone_referrals ADD CONSTRAINT fkg49hs05ll8jvmxyf9uv9eh6xv FOREIGN KEY (searchresultdone_id) REFERENCES ldap_tools.searchresultdone(id);
ALTER TABLE ldap_tools.searchresultentry ADD CONSTRAINT fkprdimc5k030blyos729qh80nw FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.searchresultreference ADD CONSTRAINT fknu2eth34oo4fk76bvad5vne3j FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.searchresultreference_referralurls ADD CONSTRAINT fkhpof73a1f2dkp81pqsoijkiig FOREIGN KEY (searchresultreference_id) REFERENCES ldap_tools.searchresultreference(id);
ALTER TABLE ldap_tools.serversortrequestcontrol ADD CONSTRAINT fkescf3e3nwvrn07dtxnhj8yqiw FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.serversortresponsecontrol ADD CONSTRAINT fkn1opjamrg40lg4y2h04w1ccwi FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.unbindrequest ADD CONSTRAINT fkfkfr3tj3ohpskid0tfq23xg8x FOREIGN KEY (id) REFERENCES ldap_tools.ldap_protocol_operation(id);
ALTER TABLE ldap_tools.vlvrequestcontrol ADD CONSTRAINT fkt20svqro7effea1132di0qfb2 FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
ALTER TABLE ldap_tools.vlvresponsecontrol ADD CONSTRAINT fkilgoabn0toj6iq1kxtgok8url FOREIGN KEY (id) REFERENCES ldap_tools.ldap_control(id);
CREATE INDEX ldap_control_checksum_index ON ldap_tools.ldap_control (checksum);
CREATE INDEX ldap_message_checksum_index ON ldap_tools.ldap_message (checksum);
CREATE INDEX ldap_modification_checksum_index ON ldap_tools.ldap_modification (checksum);
CREATE INDEX ldap_protocol_operation_checksum_index ON ldap_tools.ldap_protocol_operation (checksum);
CREATE INDEX ldap_server_sort_key_checksum_index ON ldap_tools.ldap_server_sort_key (checksum);
CREATE INDEX bindresponse_referrals__bindresponse_id_index ON ldap_tools.bindresponse_referrals(bindresponse_id);
CREATE INDEX geteffectiverightscontrol__id_index ON ldap_tools.geteffectiverightscontrol (id);
CREATE INDEX geteffectiverightscontrol_attributetypes__geteffectiverightscontrol_id_index ON ldap_tools.geteffectiverightscontrol_attributetypes (geteffectiverightscontrol_id);
CREATE INDEX intermediateresponse__id_index ON ldap_tools.intermediateresponse (id);
CREATE INDEX ldap_add_request_attributes_join__attributes_id_index ON ldap_tools.ldap_add_request_attributes_join (attributes_id);
CREATE INDEX ldap_add_request_attributes_join__addrequest_id_index ON ldap_tools.ldap_add_request_attributes_join (addrequest_id);
CREATE INDEX ldap_message__protocolop_id_index ON ldap_tools.ldap_message (protocolop_id);
CREATE INDEX ldap_message_controls_join__ldapmessage_id_index ON ldap_tools.ldap_message_controls_join (ldapmessage_id);
CREATE INDEX ldap_message_controls_join__controls_id_index ON ldap_tools.ldap_message_controls_join (controls_id);
CREATE INDEX ldap_modification__attribute_id_index ON ldap_tools.ldap_modification (attribute_id);
CREATE INDEX ldap_modify_request_modifications_join__modifyrequest_id_index ON ldap_tools.ldap_modify_request_modifications_join (modifyrequest_id);
CREATE INDEX ldap_modify_request_modifications_join__modifications_id_index ON ldap_tools.ldap_modify_request_modifications_join (modifications_id);
CREATE INDEX ldap_search_request_attributes_join__searchrequest_id_index ON ldap_tools.ldap_search_request_attributes_join (searchrequest_id);
CREATE INDEX ldap_search_request_attributes_join__attributes_id_index ON ldap_tools.ldap_search_request_attributes_join (attributes_id);
CREATE INDEX ldap_search_result_entry_attributes_join__attributes_id_index ON ldap_tools.ldap_search_result_entry_attributes_join (attributes_id);
CREATE INDEX ldap_search_result_entry_attributes_join__searchresultentry_id_index ON ldap_tools.ldap_search_result_entry_attributes_join (searchresultentry_id);
CREATE INDEX ldap_server_sort_request_control_sort_keys_join__serversortrequestcontrol_id_index ON ldap_tools.ldap_server_sort_request_control_sort_keys_join (serversortrequestcontrol_id);
CREATE INDEX ldap_server_sort_request_control_sort_keys_join__sortkeys_id_index ON ldap_tools.ldap_server_sort_request_control_sort_keys_join (sortkeys_id);
CREATE INDEX managedsaitcontrol__id_index ON ldap_tools.managedsaitcontrol (id);
CREATE INDEX modifydnrequest__id_index ON ldap_tools.modifydnrequest (id);
CREATE INDEX modifydnresponse__id_index ON ldap_tools.modifydnresponse (id);
CREATE INDEX modifydnresponse_referrals__modifydnresponse_id_index ON ldap_tools.modifydnresponse_referrals (modifydnresponse_id);
CREATE INDEX modifyrequest__id_index ON ldap_tools.modifyrequest (id);
CREATE INDEX modifyresponse__id_index ON ldap_tools.modifyresponse (id);
CREATE INDEX modifyresponse_referrals__modifyresponse_id_index ON ldap_tools.modifyresponse_referrals (modifyresponse_id);
CREATE INDEX openconnectionrequest__id_index ON ldap_tools.openconnectionrequest (id);
CREATE INDEX openconnectionresponse__id_index ON ldap_tools.openconnectionresponse (id);
CREATE INDEX pagedresultscontrol__id_index ON ldap_tools.pagedresultscontrol (id);
CREATE INDEX passwordexpiredcontrol__id_index ON ldap_tools.passwordexpiredcontrol (id);
CREATE INDEX passwordexpiringcontrol__id_index ON ldap_tools.passwordexpiringcontrol (id);
CREATE INDEX passwordpolicycontrol__id_index ON ldap_tools.passwordpolicycontrol (id);
CREATE INDEX persistentsearchcontrol__id_index ON ldap_tools.persistentsearchcontrol (id);
CREATE INDEX proxiedauthv1control__id_index ON ldap_tools.proxiedauthv1control (id);
CREATE INDEX proxiedauthv2control__id_index ON ldap_tools.proxiedauthv2control (id);
CREATE INDEX realattributesonlycontrol__id_index ON ldap_tools.realattributesonlycontrol (id);
CREATE INDEX searchrequest__id_index ON ldap_tools.searchrequest (id);
CREATE INDEX searchresultdone__id_index ON ldap_tools.searchresultdone (id);
CREATE INDEX searchresultdone_referrals__searchresultdone_id_index ON ldap_tools.searchresultdone_referrals (searchresultdone_id);
CREATE INDEX searchresultentry__id_index ON ldap_tools.searchresultentry (id);
CREATE INDEX searchresultreference__id_index ON ldap_tools.searchresultreference (id);
CREATE INDEX searchresultreference_referralurls__searchresultreference_id_index ON ldap_tools.searchresultreference_referralurls (searchresultreference_id);
CREATE INDEX serversortrequestcontrol__id_index ON ldap_tools.serversortrequestcontrol (id);
CREATE INDEX serversortresponsecontrol__id_index ON ldap_tools.serversortresponsecontrol (id);
CREATE INDEX unbindrequest__id_index ON ldap_tools.unbindrequest (id);
CREATE INDEX vlvrequestcontrol__id_index ON ldap_tools.vlvrequestcontrol (id);
CREATE INDEX vlvresponsecontrol__id_index ON ldap_tools.vlvresponsecontrol (id);<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/functions/callback/MessageParsedCallbackLoggerInfoPrint.java
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.functions.callback;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.ldap.framework.commons.bean.LDAPMessage;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.exception.MessageParsedCallbackException;
/**
* This class represents the callback utility with the JMS send implementation.
*
* @author <NAME>
*/
public class MessageParsedCallbackLoggerInfoPrint implements MessageParsedCallback
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( MessageParsedCallbackLoggerInfoPrint.class );
/**
* Standard constructor.
*/
public MessageParsedCallbackLoggerInfoPrint()
{
}
@Override
public void messageParsed( final LDAPMessage message )
{
MessageParsedCallbackLoggerInfoPrint.LOG.info( message.toString() );
}
@Override
public void startParsing() throws MessageParsedCallbackException
{
}
@Override
public void stopParsing() throws MessageParsedCallbackException
{
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/VLVResponseControl.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1Element;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1.ASN1OctetString;
/**
* This class defines the virtual list view (VLV) response control, which is returned by the server after processing a search with the VLV request control to provide information on what was actually returned.
*
* @author <NAME>
*/
@Entity @Cacheable
public class VLVResponseControl extends LDAPControl
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The OID of the VLV response control.
*/
public static final String VLV_RESPONSE_CONTROL_OID = "2.16.840.1.113730.3.4.10";
/**
* The total number of entries in the result set.
*/
private int contentCount;
/**
* The result code for the VLV operation.
*/
private int resultCode;
/**
* The position of the target entry in the overall result set.
*/
private int targetPosition;
/**
* Creates a VLV response control.
*/
public VLVResponseControl()
{
super( VLVResponseControl.VLV_RESPONSE_CONTROL_OID );
}
/**
* Creates a new VLV response control by decoding the provided value.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param controlValue
* The encoded value for this control.
* @throws ProtocolException
* If a problem occurs while decoding the value for the control.
*/
public VLVResponseControl( final boolean isCritical, final ASN1OctetString controlValue ) throws ProtocolException
{
super( VLVResponseControl.VLV_RESPONSE_CONTROL_OID, isCritical );
final ASN1Element[] sequenceElements;
try
{
final byte[] valueBytes = controlValue.getValue();
sequenceElements = ASN1Element.decodeAsSequence( valueBytes ).getElements();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode VLV response control sequence", e );
}
if ( sequenceElements.length != 3 )
{
throw new ProtocolException( "There must be exactly 3 elements in a VLV response control sequence" );
}
try
{
this.targetPosition = sequenceElements[ 0 ].decodeAsInteger().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode target position from VLV response control", e );
}
try
{
this.contentCount = sequenceElements[ 1 ].decodeAsInteger().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode content count from VLV response control", e );
}
try
{
this.resultCode = sequenceElements[ 2 ].decodeAsEnumerated().getIntValue();
} catch ( final Exception e )
{
throw new ProtocolException( "Unable to decode result code from VLV response control", e );
}
}
/**
* Creates a VLV response control.
*
* @param isCritical
* Indicates whether this control should be marked critical.
* @param targetPosition
* The position of the target entry in the result set.
* @param contentCount
* The total number of entries in the result set.
* @param resultCode
* The result code for the VLV operation.
*/
public VLVResponseControl( final boolean isCritical, final int targetPosition, final int contentCount, final int resultCode )
{
super( VLVResponseControl.VLV_RESPONSE_CONTROL_OID, isCritical );
this.targetPosition = targetPosition;
this.contentCount = contentCount;
this.resultCode = resultCode;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final VLVResponseControl other = ( VLVResponseControl ) obj;
return ( this.contentCount == other.contentCount ) && ( this.resultCode == other.resultCode ) && ( this.targetPosition == other.targetPosition );
}
/**
* Retrieves the total number of entries in the result set.
*
* @return The total number of entries in the result set.
*/
public int getContentCount()
{
return this.contentCount;
}
/**
* Retrieves the result code for the VLV operation.
*
* @return The result code for the VLV operation.
*/
public int getResultCode()
{
return this.resultCode;
}
/**
* Retrieves the position of the target entry in the result set.
*
* @return The position of the target entry in the result set.
*/
public int getTargetPosition()
{
return this.targetPosition;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + this.contentCount;
result = ( prime * result ) + this.resultCode;
result = ( prime * result ) + this.targetPosition;
return result;
}
/**
* This method set the total number of entries in the result set.
*
* @param contentCount
* The total number of entries in the result set.
*/
public void setContentCount( final int contentCount )
{
this.contentCount = contentCount;
}
/**
* This method set the result code for the VLV operation.
*
* @param resultCode
* The result code for the VLV operation.
*/
public void setResultCode( final int resultCode )
{
this.resultCode = resultCode;
}
/**
* This method set the position of the target entry in the overall result set.
*
* @param targetPosition
* The position of the target entry in the overall result set.
*/
public void setTargetPosition( final int targetPosition )
{
this.targetPosition = targetPosition;
}
/**
* Retrieves a string representation of this control with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this control with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
return String.valueOf( indentBuf ) + "LDAP VLV Response Control" + LDAPMessage.EOL + indentBuf + " OID: " + this.getControlOID() + LDAPMessage.EOL + indentBuf + " Criticality: " + this.isCritical() + LDAPMessage.EOL + indentBuf + " Target Position: " + this.targetPosition + LDAPMessage.EOL + indentBuf + " Content Count: " + this.contentCount + LDAPMessage.EOL + indentBuf + " VLV Result Code: " + this.resultCode + " (" + LDAPResultCode.resultCodeToString( this.resultCode ) + ')' + LDAPMessage.EOL;
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + this.contentCount;
this.checksum = ( prime * this.checksum ) + this.resultCode;
this.checksum = ( prime * this.checksum ) + this.targetPosition;
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/functions/dao/LDAPMessageDAO.java
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.functions.dao;
import java.util.ArrayList;
import java.util.HashMap;
import java.util.List;
import java.util.Map;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.database.jpa.exception.JpaFacilitatorException;
import ita.deluca.tools.application.framework.database.jpa.model.JpaFacilitator;
import ita.deluca.tools.ldap.framework.commons.bean.AbandonRequest;
import ita.deluca.tools.ldap.framework.commons.bean.AddRequest;
import ita.deluca.tools.ldap.framework.commons.bean.AddResponse;
import ita.deluca.tools.ldap.framework.commons.bean.AuthorizationIDRequestControl;
import ita.deluca.tools.ldap.framework.commons.bean.AuthorizationIDResponseControl;
import ita.deluca.tools.ldap.framework.commons.bean.BindRequest;
import ita.deluca.tools.ldap.framework.commons.bean.BindResponse;
import ita.deluca.tools.ldap.framework.commons.bean.CompareRequest;
import ita.deluca.tools.ldap.framework.commons.bean.CompareResponse;
import ita.deluca.tools.ldap.framework.commons.bean.DeleteRequest;
import ita.deluca.tools.ldap.framework.commons.bean.DeleteResponse;
import ita.deluca.tools.ldap.framework.commons.bean.EntryChangeNotificationControl;
import ita.deluca.tools.ldap.framework.commons.bean.ExtendedRequest;
import ita.deluca.tools.ldap.framework.commons.bean.ExtendedResponse;
import ita.deluca.tools.ldap.framework.commons.bean.GetEffectiveRightsControl;
import ita.deluca.tools.ldap.framework.commons.bean.IntermediateResponse;
import ita.deluca.tools.ldap.framework.commons.bean.LDAPControl;
import ita.deluca.tools.ldap.framework.commons.bean.LDAPMessage;
import ita.deluca.tools.ldap.framework.commons.bean.ManageDSAITControl;
import ita.deluca.tools.ldap.framework.commons.bean.ModifyDNRequest;
import ita.deluca.tools.ldap.framework.commons.bean.ModifyDNResponse;
import ita.deluca.tools.ldap.framework.commons.bean.ModifyRequest;
import ita.deluca.tools.ldap.framework.commons.bean.ModifyResponse;
import ita.deluca.tools.ldap.framework.commons.bean.OpenConnectionRequest;
import ita.deluca.tools.ldap.framework.commons.bean.OpenConnectionResponse;
import ita.deluca.tools.ldap.framework.commons.bean.PagedResultsControl;
import ita.deluca.tools.ldap.framework.commons.bean.PasswordExpiredControl;
import ita.deluca.tools.ldap.framework.commons.bean.PasswordExpiringControl;
import ita.deluca.tools.ldap.framework.commons.bean.PasswordPolicyControl;
import ita.deluca.tools.ldap.framework.commons.bean.PersistentSearchControl;
import ita.deluca.tools.ldap.framework.commons.bean.ProxiedAuthV1Control;
import ita.deluca.tools.ldap.framework.commons.bean.ProxiedAuthV2Control;
import ita.deluca.tools.ldap.framework.commons.bean.RealAttributesOnlyControl;
import ita.deluca.tools.ldap.framework.commons.bean.SearchRequest;
import ita.deluca.tools.ldap.framework.commons.bean.SearchResultDone;
import ita.deluca.tools.ldap.framework.commons.bean.SearchResultEntry;
import ita.deluca.tools.ldap.framework.commons.bean.SearchResultReference;
import ita.deluca.tools.ldap.framework.commons.bean.ServerSortRequestControl;
import ita.deluca.tools.ldap.framework.commons.bean.ServerSortResponseControl;
import ita.deluca.tools.ldap.framework.commons.bean.UnbindRequest;
import ita.deluca.tools.ldap.framework.commons.bean.VLVRequestControl;
import ita.deluca.tools.ldap.framework.commons.bean.VLVResponseControl;
import ita.deluca.tools.ldap.framework.commons.tcpdecoder.exception.LdapEntityDaoException;
import ita.deluca.tools.ldap.framework.properties.Constants;
/**
* This class contains all the function that allows interaction with the database via JPA.
*
* @author <NAME>
*/
public class LDAPMessageDAO
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( LDAPMessageDAO.class );
/**
* Retrieves the connection client host.
*
* @param connectionId
* The connection id.
* @return The connection client host.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the client host.
*/
public static String getConnectionClientHost( final String connectionId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select LDAPMessage.destHost from LDAPMessage LDAPMessage, BindResponse bindresponse where (LDAPMessage.protocolOp = bindresponse and LDAPMessage.connectionId = :connectionId)";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the client host referred by the connection id '" + connectionId + "' thrown an error !!!";
LDAPMessageDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
return ( String ) res.get( 0 );
}
}
/**
* Retrieves the connection ldap host.
*
* @param connectionId
* The connection id.
* @return The connection ldap host.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the ldap host.
*/
public static String getConnectionLdapHost( final String connectionId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select LDAPMessage.destHost from LDAPMessage LDAPMessage, BindRequest bindrequest where (LDAPMessage.protocolOp = bindrequest and LDAPMessage.connectionId = :connectionId)";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the ldap host referred by the connection id '" + connectionId + "' thrown an error !!!";
LDAPMessageDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
return ( String ) res.get( 0 );
}
}
/**
* Extract the message that have the same message id.
*
* @param connectionId
* The connection id of the message.
* @return The list of the message having the same connection id.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the message.
*/
public static List<LDAPMessage> getFromConnectionID( final String connectionId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select LDAPMessage from LDAPMessage LDAPMessage where LDAPMessage.connectionId = :connectionId order by LDAPMessage.timestamp";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the list of the ldap message referred by the connection id '" + connectionId + "' thrown an error !!!";
LDAPMessageDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
final List<LDAPMessage> retRes = ( List<LDAPMessage> ) res;
return retRes;
}
}
/**
* Extract the message from this id.
*
* @param id
* The id of the message.
* @return The message having the passed id.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the search of the message.
*/
public static LDAPMessage getFromID( final long id ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
try
{
return jpa.restore( id, LDAPMessage.class );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the message with id '" + id + "' thrown an error !!!";
LDAPMessageDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
}
}
/**
* Retrieves the total bytes transfered in one connection.
*
* @param connectionId
* The connection id.
* @return The total bytes transfered.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the calculation of the transfered bytes.
*/
public static int getTotalByteTransfered( final String connectionId ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select LDAPMessage.messageLength from LDAPMessage LDAPMessage where LDAPMessage.connectionId = :connectionId";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the total byte transfered by the connection '" + connectionId + "' thrown an error !!!";
LDAPMessageDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
int totalLength = 0;
for ( final Object length : res )
{
totalLength += ( ( Integer ) length ).intValue();
}
return totalLength;
}
}
/**
* Retrieves the total bytes transfered from the client to the ldap in one connection.
*
* @param connectionId
* The connection id.
* @return The total bytes transfered from the client to the ldap.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the calculation of the transfered bytes.
*/
public static int getTotalByteTransferedByClientHost( final String connectionId ) throws LdapEntityDaoException
{
final String clientHost = LDAPMessageDAO.getConnectionClientHost( connectionId );
return LDAPMessageDAO.getTotalByteTransferedByHost( connectionId, clientHost );
}
/**
* Retrieves the total bytes transfered from a host in one connection.
*
* @param connectionId
* The connection id.
* @param sourceHost
* The source host.
* @return The total bytes transfered from a host.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the calculation of the transfered bytes.
*/
public static int getTotalByteTransferedByHost( final String connectionId, final String sourceHost ) throws LdapEntityDaoException
{
try ( final JpaFacilitator jpa = JpaFacilitator.getThreadLocalInstance( Constants.TCP_DECODER_JPA_ENTITY_MANAGER ) )
{
final String query = "select SUM(LDAPMessage.messageLength) from LDAPMessage LDAPMessage where LDAPMessage.connectionId = :connectionId and LDAPMessage.sourceHost = :sourcehost";
final Map<String, Object> parameter = new HashMap<>();
parameter.put( "connectionId", connectionId );
parameter.put( "sourcehost", sourceHost );
List<?> res;
try
{
res = jpa.execute( query, parameter );
} catch ( final JpaFacilitatorException ex )
{
final String msg = "!!! The extraction of the total byte transfered by the connection '" + connectionId + "' from the host '" + sourceHost + "' thrown an error !!!";
LDAPMessageDAO.LOG.error( msg, ex );
throw new LdapEntityDaoException( msg, ex );
}
return ( ( Long ) res.get( 0 ) ).intValue();
}
}
/**
* Retrieves the total bytes transfered from the ldap to the client in one connection.
*
* @param connectionId
* The connection id.
* @return The total bytes transfered from the ldap to the client.
* @throws LdapEntityDaoException
* This exception was raised whenever an error occurred during the calculation of the transfered bytes.
*/
public static int getTotalByteTransferedByLdapHost( final String connectionId ) throws LdapEntityDaoException
{
final String ldapHost = LDAPMessageDAO.getConnectionLdapHost( connectionId );
return LDAPMessageDAO.getTotalByteTransferedByHost( connectionId, ldapHost );
}
/**
* This method persist the object into the database.
*
* @param entity
* the object to persist.
* @throws LdapEntityDaoException
* this exception was raised whenever an error occurred during the persist of the object.
*/
public static void persist( final LDAPMessage entity ) throws LdapEntityDaoException
{
if ( LDAPMessageDAO.LOG.isDebugEnabled() )
{
LDAPMessageDAO.LOG.debug( "Persist the entity:\n" + entity );
}
entity.updateChecksum();
final List<LDAPControl> persistedControl = new ArrayList<>();
for ( final LDAPControl control : entity.getControls() )
{
if ( control instanceof ManageDSAITControl )
{
persistedControl.add( ManageDSAITControlDAO.persist( ( ManageDSAITControl ) control ) );
} else if ( control instanceof PersistentSearchControl )
{
persistedControl.add( PersistentSearchControlDAO.persist( ( PersistentSearchControl ) control ) );
} else if ( control instanceof EntryChangeNotificationControl )
{
persistedControl.add( EntryChangeNotificationControlDAO.persist( ( EntryChangeNotificationControl ) control ) );
} else if ( control instanceof PasswordExpiredControl )
{
persistedControl.add( PasswordExpiredControlDAO.persist( ( PasswordExpiredControl ) control ) );
} else if ( control instanceof PasswordExpiringControl )
{
persistedControl.add( PasswordExpiringControlDAO.persist( ( PasswordExpiringControl ) control ) );
} else if ( control instanceof PasswordPolicyControl )
{
persistedControl.add( PasswordPolicyControlDAO.persist( ( PasswordPolicyControl ) control ) );
} else if ( control instanceof ServerSortRequestControl )
{
persistedControl.add( ServerSortRequestControlDAO.persist( ( ServerSortRequestControl ) control ) );
} else if ( control instanceof ServerSortResponseControl )
{
persistedControl.add( ServerSortResponseControlDAO.persist( ( ServerSortResponseControl ) control ) );
} else if ( control instanceof VLVRequestControl )
{
persistedControl.add( VLVRequestControlDAO.persist( ( VLVRequestControl ) control ) );
} else if ( control instanceof VLVResponseControl )
{
persistedControl.add( VLVResponseControlDAO.persist( ( VLVResponseControl ) control ) );
} else if ( control instanceof ProxiedAuthV1Control )
{
persistedControl.add( ProxiedAuthV1ControlDAO.persist( ( ProxiedAuthV1Control ) control ) );
} else if ( control instanceof ProxiedAuthV2Control )
{
persistedControl.add( ProxiedAuthV2ControlDAO.persist( ( ProxiedAuthV2Control ) control ) );
} else if ( control instanceof RealAttributesOnlyControl )
{
persistedControl.add( RealAttributesOnlyControlDAO.persist( ( RealAttributesOnlyControl ) control ) );
} else if ( control instanceof GetEffectiveRightsControl )
{
persistedControl.add( GetEffectiveRightsControlDAO.persist( ( GetEffectiveRightsControl ) control ) );
} else if ( control instanceof AuthorizationIDRequestControl )
{
persistedControl.add( AuthorizationIDRequestControlDAO.persist( ( AuthorizationIDRequestControl ) control ) );
} else if ( control instanceof AuthorizationIDResponseControl )
{
persistedControl.add( AuthorizationIDResponseControlDAO.persist( ( AuthorizationIDResponseControl ) control ) );
} else if ( control instanceof PagedResultsControl )
{
persistedControl.add( PagedResultsControlDAO.persist( ( PagedResultsControl ) control ) );
}
}
entity.setControls( persistedControl );
if ( entity.getProtocolOp() instanceof AbandonRequest )
{
entity.setProtocolOp( AbandonRequestDAO.persist( ( AbandonRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof AddRequest )
{
entity.setProtocolOp( AddRequestDAO.persist( ( AddRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof AddResponse )
{
entity.setProtocolOp( AddResponseDAO.persist( ( AddResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof BindRequest )
{
entity.setProtocolOp( BindRequestDAO.persist( ( BindRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof BindResponse )
{
entity.setProtocolOp( BindResponseDAO.persist( ( BindResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof CompareRequest )
{
entity.setProtocolOp( CompareRequestDAO.persist( ( CompareRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof CompareResponse )
{
entity.setProtocolOp( CompareResponseDAO.persist( ( CompareResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof DeleteRequest )
{
entity.setProtocolOp( DeleteRequestDAO.persist( ( DeleteRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof DeleteResponse )
{
entity.setProtocolOp( DeleteResponseDAO.persist( ( DeleteResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof ExtendedRequest )
{
entity.setProtocolOp( ExtendedRequestDAO.persist( ( ExtendedRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof ExtendedResponse )
{
entity.setProtocolOp( ExtendedResponseDAO.persist( ( ExtendedResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof IntermediateResponse )
{
entity.setProtocolOp( IntermediateResponseDAO.persist( ( IntermediateResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof ModifyDNRequest )
{
entity.setProtocolOp( ModifyDNRequestDAO.persist( ( ModifyDNRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof ModifyDNResponse )
{
entity.setProtocolOp( ModifyDNResponseDAO.persist( ( ModifyDNResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof ModifyRequest )
{
entity.setProtocolOp( ModifyRequestDAO.persist( ( ModifyRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof ModifyResponse )
{
entity.setProtocolOp( ModifyResponseDAO.persist( ( ModifyResponse ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof SearchRequest )
{
entity.setProtocolOp( SearchRequestDAO.persist( ( SearchRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof SearchResultDone )
{
entity.setProtocolOp( SearchResultDoneDAO.persist( ( SearchResultDone ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof SearchResultEntry )
{
entity.setProtocolOp( SearchResultEntryDAO.persist( ( SearchResultEntry ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof SearchResultReference )
{
entity.setProtocolOp( SearchResultReferenceDAO.persist( ( SearchResultReference ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof UnbindRequest )
{
entity.setProtocolOp( UnbindRequestDAO.persist( ( UnbindRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof OpenConnectionRequest )
{
entity.setProtocolOp( OpenConnectionRequestDAO.persist( ( OpenConnectionRequest ) entity.getProtocolOp() ) );
} else if ( entity.getProtocolOp() instanceof OpenConnectionResponse )
{
entity.setProtocolOp( OpenConnectionResponseDAO.persist( ( OpenConnectionResponse ) entity.getProtocolOp() ) );
}
LDAPEntityDAO.persistEntity( entity );
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/bean/DeleteRequest.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.bean;
import javax.persistence.Cacheable;
import javax.persistence.Entity;
/**
* This class defines an LDAP delete request, which is used to remove an entry from a directory server.
*
* @author <NAME>
*/
@Entity @Cacheable
public class DeleteRequest extends ProtocolOp
{
/**
* The class serial version uid.
*/
private static final long serialVersionUID = 1L;
/**
* The DN of the entry to delete.
*/
private String dn;
/**
* Creates a new delete request to delete the entry with the given DN.
*/
public DeleteRequest()
{
}
/**
* Creates a new delete request to delete the entry with the given DN.
*
* @param dn
* The DN of the entry to delete.
*/
public DeleteRequest( final String dn )
{
this.dn = dn;
}
@Override
public boolean equals( final Object obj )
{
if ( this == obj )
{
return true;
}
if ( !super.equals( obj ) )
{
return false;
}
if ( this.getClass() != obj.getClass() )
{
return false;
}
final DeleteRequest other = ( DeleteRequest ) obj;
if ( this.dn == null )
{
if ( other.dn != null )
{
return false;
}
} else if ( !this.dn.equals( other.dn ) )
{
return false;
}
return true;
}
/**
* Retrieves the DN of the entry to delete.
*
* @return The DN of the entry to delete.
*/
public String getDn()
{
return this.dn;
}
@Override
public int hashCode()
{
final int prime = 31;
int result = super.hashCode();
result = ( prime * result ) + ( this.dn == null ? 0 : this.dn.hashCode() );
return result;
}
/**
* Retrieves a user-friendly name for this protocol op.
*
* @return A user-friendly name for this protocol op.
*/
@Override
public String retrieveProtocolOpType()
{
return "LDAP Delete Request";
}
/**
* Setting the DN of the entry to delete.
*
* @param dn
* The DN of the entry to delete.
*/
public void setDn( final String dn )
{
this.dn = dn;
}
/**
* Retrieves a string representation of this protocol op with the specified indent.
*
* @param indent
* The number of spaces to indent the output.
* @return A string representation of this protocol op with the specified indent.
*/
@Override
public String toString( final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
return String.valueOf( indentBuf ) + "dn: " + this.dn + LDAPMessage.EOL;
}
@Override
public void updateChecksum()
{
final int prime = 31;
this.checksum = 1;
this.checksum = ( prime * this.checksum ) + ( this.dn == null ? 0 : this.dn.hashCode() );
this.checksum = ( prime * this.checksum ) + this.getClass().getCanonicalName().hashCode();
}
}
<file_sep>/src/ita/deluca/tools/ldap/framework/commons/tcpdecoder/asn1/ASN1Element.java
/*
* Sun Public License The contents of this file are subject to the Sun Public License Version 1.0 (the "License"). You may not use this file except in compliance with the License. A copy of the License is available at http://www.sun.com/ The Original Code is the SLAMD Distributed Load Generation Engine. The Initial Developer of the Original Code is <NAME>. Portions created by <NAME> are Copyright (C) 2004-2010. Some preexisting portions Copyright (C) 2002-2006 Sun Microsystems, Inc. All Rights Reserved. Contributor(s): <NAME>
*/
package ita.deluca.tools.ldap.framework.commons.tcpdecoder.asn1;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
/**
* This class defines a generic ASN.1 element and a set of core methods for dealing with them. Subclasses may deal with more specific kinds of ASN.1 elements.
*
* @author <NAME>
*/
public class ASN1Element
{
/**
* Class logger.
*/
private static final Logger LOG = LogManager.getLogger( ASN1Element.class );
/**
* The standard ASN.1 type for boolean values.
*/
static final byte ASN1_BOOLEAN_TYPE = 0x01;
/**
* The standard ASN.1 type for enumerated values.
*/
static final byte ASN1_ENUMERATED_TYPE = 0x0A;
/**
* The standard ASN.1 type for integer values.
*/
static final byte ASN1_INTEGER_TYPE = 0x02;
/**
* The standard ASN.1 type for null values.
*/
static final byte ASN1_NULL_TYPE = 0x05;
/**
* The standard ASN.1 type for sequence values.
*/
static final byte ASN1_SEQUENCE_TYPE = 0x30;
/**
* The standard ASN.1 type for set values.
*/
static final byte ASN1_SET_TYPE = 0x31;
/**
* The end of line character(s) for this platform.
*/
static final String EOL = System.getProperty( "line.separator" );
/**
* The standard ASN.1 type for octet string values.
*/
public static final byte ASN1_OCTET_STRING_TYPE = 0x04;
/**
* An empty byte array, used to prevent multiple allocations for empty arrays.
*/
public static final byte[] EMPTY_BYTES = new byte[ 0 ];
/**
* The id of the element.
*/
private long id;
/**
* The encoded form of this ASN.1 element.
*/
byte[] encodedElement;
/**
* The type of this ASN.1 element. This implementation only supports single-byte type values (up through "APPLICATION 31").
*/
byte type;
/**
* The encoded value for this ASN.1 element.
*/
byte[] value;
/**
* Creates a new ASN.1 element with the specified information.
*
* @param type
* The type of this ASN.1 element.
* @param value
* The value for this ASN.1 element.
* @param encodedElement
* The ASN.1 element encoded as a byte array.
*/
private ASN1Element( final byte type, final byte[] value, final byte[] encodedElement )
{
this.type = type;
this.value = value;
this.encodedElement = encodedElement;
}
/**
* Standard constructor.
*/
ASN1Element()
{
}
/**
* Creates a new ASN.1 element with the specified type and no value.
*
* @param type
* The type of this ASN.1 element.
*/
ASN1Element( final byte type )
{
this( type, ASN1Element.EMPTY_BYTES );
}
/**
* Creates a new ASN.1 element with the specified type and value.
*
* @param type
* The type of this ASN.1 element.
* @param value
* The encoded value for this ASN.1 element.
*/
public ASN1Element( final byte type, final byte[] value )
{
this.type = type;
this.value = value == null ? ASN1Element.EMPTY_BYTES : value;
final byte[] encodedLength = ASN1Element.encodeLength( this.value.length );
this.encodedElement = new byte[ 1 + encodedLength.length + this.value.length ];
this.encodedElement[ 0 ] = type;
System.arraycopy( encodedLength, 0, this.encodedElement, 1, encodedLength.length );
System.arraycopy( this.value, 0, this.encodedElement, 1 + encodedLength.length, this.value.length );
}
/**
* Converts the provided byte array into a Java int value. Note that this method assumes that the provided array contains no more than four bytes.
*
* @param byteArray
* The byte array containing the encoded integer value.
* @return The Java int value decoded from the byte array.
*/
private static int byteArrayToInt( final byte[] byteArray )
{
int value = 0x00000000;
switch ( byteArray.length )
{
case 1:
value |= 0x000000FF & byteArray[ 0 ];
break;
case 2:
value |= ( ( 0x000000FF & byteArray[ 0 ] ) << 8 ) | ( 0x000000FF & byteArray[ 1 ] );
break;
case 3:
value |= ( ( 0x000000FF & byteArray[ 0 ] ) << 16 ) | ( ( 0x000000FF & byteArray[ 1 ] ) << 8 ) | ( 0x000000FF & byteArray[ 2 ] );
break;
case 4:
value |= ( ( 0x000000FF & byteArray[ 0 ] ) << 24 ) | ( ( 0x000000FF & byteArray[ 1 ] ) << 16 ) | ( ( 0x000000FF & byteArray[ 2 ] ) << 8 ) | ( 0x000000FF & byteArray[ 3 ] );
break;
default:
break;
}
return value;
}
/**
* Retrieves a string representation of the provided byte array (including the ASCII equivalent) using the specified indent.
*
* @param byteArray
* The byte array to be displayed as a string.
* @param indent
* The number of spaces to indent the output.
* @return A string representation of the provided byte array with the ASCII equivalent.
*/
private static String byteArrayToStringWithASCII( final byte[] byteArray, final int indent )
{
final StringBuilder indentBuf = new StringBuilder( indent );
for ( int i = 0; i < indent; i++ )
{
indentBuf.append( ' ' );
}
final StringBuilder buffer = new StringBuilder();
StringBuilder hexBuf = new StringBuilder();
StringBuilder asciiBuf = new StringBuilder();
for ( int i = 0; i < byteArray.length; i++ )
{
switch ( byteArray[ i ] )
{
case 0x00:
hexBuf.append( "00 " );
asciiBuf.append( '.' );
break;
case 0x01:
hexBuf.append( "01 " );
asciiBuf.append( '.' );
break;
case 0x02:
hexBuf.append( "02 " );
asciiBuf.append( '.' );
break;
case 0x03:
hexBuf.append( "03 " );
asciiBuf.append( '.' );
break;
case 0x04:
hexBuf.append( "04 " );
asciiBuf.append( '.' );
break;
case 0x05:
hexBuf.append( "05 " );
asciiBuf.append( '.' );
break;
case 0x06:
hexBuf.append( "06 " );
asciiBuf.append( '.' );
break;
case 0x07:
hexBuf.append( "07 " );
asciiBuf.append( '.' );
break;
case 0x08:
hexBuf.append( "08 " );
asciiBuf.append( '.' );
break;
case 0x09:
hexBuf.append( "09 " );
asciiBuf.append( '.' );
break;
case 0x0A:
hexBuf.append( "0A " );
asciiBuf.append( '.' );
break;
case 0x0B:
hexBuf.append( "0B " );
asciiBuf.append( '.' );
break;
case 0x0C:
hexBuf.append( "0C " );
asciiBuf.append( '.' );
break;
case 0x0D:
hexBuf.append( "0D " );
asciiBuf.append( '.' );
break;
case 0x0E:
hexBuf.append( "0E " );
asciiBuf.append( '.' );
break;
case 0x0F:
hexBuf.append( "0F " );
asciiBuf.append( '.' );
break;
case 0x10:
hexBuf.append( "10 " );
asciiBuf.append( '.' );
break;
case 0x11:
hexBuf.append( "11 " );
asciiBuf.append( '.' );
break;
case 0x12:
hexBuf.append( "12 " );
asciiBuf.append( '.' );
break;
case 0x13:
hexBuf.append( "13 " );
asciiBuf.append( '.' );
break;
case 0x14:
hexBuf.append( "14 " );
asciiBuf.append( '.' );
break;
case 0x15:
hexBuf.append( "15 " );
asciiBuf.append( '.' );
break;
case 0x16:
hexBuf.append( "16 " );
asciiBuf.append( '.' );
break;
case 0x17:
hexBuf.append( "17 " );
asciiBuf.append( '.' );
break;
case 0x18:
hexBuf.append( "18 " );
asciiBuf.append( '.' );
break;
case 0x19:
hexBuf.append( "19 " );
asciiBuf.append( '.' );
break;
case 0x1A:
hexBuf.append( "1A " );
asciiBuf.append( '.' );
break;
case 0x1B:
hexBuf.append( "1B " );
asciiBuf.append( '.' );
break;
case 0x1C:
hexBuf.append( "1C " );
asciiBuf.append( '.' );
break;
case 0x1D:
hexBuf.append( "1D " );
asciiBuf.append( '.' );
break;
case 0x1E:
hexBuf.append( "1E " );
asciiBuf.append( '.' );
break;
case 0x1F:
hexBuf.append( "1F " );
asciiBuf.append( '.' );
break;
case 0x20:
hexBuf.append( "20 " );
asciiBuf.append( ' ' );
break;
case 0x21:
hexBuf.append( "21 " );
asciiBuf.append( '!' );
break;
case 0x22:
hexBuf.append( "22 " );
asciiBuf.append( '"' );
break;
case 0x23:
hexBuf.append( "23 " );
asciiBuf.append( '#' );
break;
case 0x24:
hexBuf.append( "24 " );
asciiBuf.append( '$' );
break;
case 0x25:
hexBuf.append( "25 " );
asciiBuf.append( '%' );
break;
case 0x26:
hexBuf.append( "26 " );
asciiBuf.append( '&' );
break;
case 0x27:
hexBuf.append( "27 " );
asciiBuf.append( '\'' );
break;
case 0x28:
hexBuf.append( "28 " );
asciiBuf.append( '(' );
break;
case 0x29:
hexBuf.append( "29 " );
asciiBuf.append( ')' );
break;
case 0x2A:
hexBuf.append( "2A " );
asciiBuf.append( '*' );
break;
case 0x2B:
hexBuf.append( "2B " );
asciiBuf.append( '+' );
break;
case 0x2C:
hexBuf.append( "2C " );
asciiBuf.append( ',' );
break;
case 0x2D:
hexBuf.append( "2D " );
asciiBuf.append( '-' );
break;
case 0x2E:
hexBuf.append( "2E " );
asciiBuf.append( '.' );
break;
case 0x2F:
hexBuf.append( "2F " );
asciiBuf.append( '/' );
break;
case 0x30:
hexBuf.append( "30 " );
asciiBuf.append( '0' );
break;
case 0x31:
hexBuf.append( "31 " );
asciiBuf.append( '1' );
break;
case 0x32:
hexBuf.append( "32 " );
asciiBuf.append( '2' );
break;
case 0x33:
hexBuf.append( "33 " );
asciiBuf.append( '3' );
break;
case 0x34:
hexBuf.append( "34 " );
asciiBuf.append( '4' );
break;
case 0x35:
hexBuf.append( "35 " );
asciiBuf.append( '5' );
break;
case 0x36:
hexBuf.append( "36 " );
asciiBuf.append( '6' );
break;
case 0x37:
hexBuf.append( "37 " );
asciiBuf.append( '7' );
break;
case 0x38:
hexBuf.append( "38 " );
asciiBuf.append( '8' );
break;
case 0x39:
hexBuf.append( "39 " );
asciiBuf.append( '9' );
break;
case 0x3A:
hexBuf.append( "3A " );
asciiBuf.append( ':' );
break;
case 0x3B:
hexBuf.append( "3B " );
asciiBuf.append( ';' );
break;
case 0x3C:
hexBuf.append( "3C " );
asciiBuf.append( '<' );
break;
case 0x3D:
hexBuf.append( "3D " );
asciiBuf.append( '=' );
break;
case 0x3E:
hexBuf.append( "3E " );
asciiBuf.append( '>' );
break;
case 0x3F:
hexBuf.append( "3F " );
asciiBuf.append( '?' );
break;
case 0x40:
hexBuf.append( "40 " );
asciiBuf.append( '@' );
break;
case 0x41:
hexBuf.append( "41 " );
asciiBuf.append( 'A' );
break;
case 0x42:
hexBuf.append( "42 " );
asciiBuf.append( 'B' );
break;
case 0x43:
hexBuf.append( "43 " );
asciiBuf.append( 'C' );
break;
case 0x44:
hexBuf.append( "44 " );
asciiBuf.append( 'D' );
break;
case 0x45:
hexBuf.append( "45 " );
asciiBuf.append( 'E' );
break;
case 0x46:
hexBuf.append( "46 " );
asciiBuf.append( 'F' );
break;
case 0x47:
hexBuf.append( "47 " );
asciiBuf.append( 'G' );
break;
case 0x48:
hexBuf.append( "48 " );
asciiBuf.append( 'H' );
break;
case 0x49:
hexBuf.append( "49 " );
asciiBuf.append( 'I' );
break;
case 0x4A:
hexBuf.append( "4A " );
asciiBuf.append( 'J' );
break;
case 0x4B:
hexBuf.append( "4B " );
asciiBuf.append( 'K' );
break;
case 0x4C:
hexBuf.append( "4C " );
asciiBuf.append( 'L' );
break;
case 0x4D:
hexBuf.append( "4D " );
asciiBuf.append( 'M' );
break;
case 0x4E:
hexBuf.append( "4E " );
asciiBuf.append( 'N' );
break;
case 0x4F:
hexBuf.append( "4F " );
asciiBuf.append( 'O' );
break;
case 0x50:
hexBuf.append( "50 " );
asciiBuf.append( 'P' );
break;
case 0x51:
hexBuf.append( "51 " );
asciiBuf.append( 'Q' );
break;
case 0x52:
hexBuf.append( "52 " );
asciiBuf.append( 'R' );
break;
case 0x53:
hexBuf.append( "53 " );
asciiBuf.append( 'S' );
break;
case 0x54:
hexBuf.append( "54 " );
asciiBuf.append( 'T' );
break;
case 0x55:
hexBuf.append( "55 " );
asciiBuf.append( 'U' );
break;
case 0x56:
hexBuf.append( "56 " );
asciiBuf.append( 'V' );
break;
case 0x57:
hexBuf.append( "57 " );
asciiBuf.append( 'W' );
break;
case 0x58:
hexBuf.append( "58 " );
asciiBuf.append( 'X' );
break;
case 0x59:
hexBuf.append( "59 " );
asciiBuf.append( 'Y' );
break;
case 0x5A:
hexBuf.append( "5A " );
asciiBuf.append( 'Z' );
break;
case 0x5B:
hexBuf.append( "5B " );
asciiBuf.append( '[' );
break;
case 0x5C:
hexBuf.append( "5C " );
asciiBuf.append( '\\' );
break;
case 0x5D:
hexBuf.append( "5D " );
asciiBuf.append( ']' );
break;
case 0x5E:
hexBuf.append( "5E " );
asciiBuf.append( '^' );
break;
case 0x5F:
hexBuf.append( "5F " );
asciiBuf.append( '_' );
break;
case 0x60:
hexBuf.append( "60 " );
asciiBuf.append( '`' );
break;
case 0x61:
hexBuf.append( "61 " );
asciiBuf.append( 'a' );
break;
case 0x62:
hexBuf.append( "62 " );
asciiBuf.append( 'b' );
break;
case 0x63:
hexBuf.append( "63 " );
asciiBuf.append( 'c' );
break;
case 0x64:
hexBuf.append( "64 " );
asciiBuf.append( 'd' );
break;
case 0x65:
hexBuf.append( "65 " );
asciiBuf.append( 'e' );
break;
case 0x66:
hexBuf.append( "66 " );
asciiBuf.append( 'f' );
break;
case 0x67:
hexBuf.append( "67 " );
asciiBuf.append( 'g' );
break;
case 0x68:
hexBuf.append( "68 " );
asciiBuf.append( 'h' );
break;
case 0x69:
hexBuf.append( "69 " );
asciiBuf.append( 'i' );
break;
case 0x6A:
hexBuf.append( "6A " );
asciiBuf.append( 'j' );
break;
case 0x6B:
hexBuf.append( "6B " );
asciiBuf.append( 'k' );
break;
case 0x6C:
hexBuf.append( "6C " );
asciiBuf.append( 'l' );
break;
case 0x6D:
hexBuf.append( "6D " );
asciiBuf.append( 'm' );
break;
case 0x6E:
hexBuf.append( "6E " );
asciiBuf.append( 'n' );
break;
case 0x6F:
hexBuf.append( "6F " );
asciiBuf.append( 'o' );
break;
case 0x70:
hexBuf.append( "70 " );
asciiBuf.append( 'p' );
break;
case 0x71:
hexBuf.append( "71 " );
asciiBuf.append( 'q' );
break;
case 0x72:
hexBuf.append( "72 " );
asciiBuf.append( 'r' );
break;
case 0x73:
hexBuf.append( "73 " );
asciiBuf.append( 's' );
break;
case 0x74:
hexBuf.append( "74 " );
asciiBuf.append( 't' );
break;
case 0x75:
hexBuf.append( "75 " );
asciiBuf.append( 'u' );
break;
case 0x76:
hexBuf.append( "76 " );
asciiBuf.append( 'v' );
break;
case 0x77:
hexBuf.append( "77 " );
asciiBuf.append( 'w' );
break;
case 0x78:
hexBuf.append( "78 " );
asciiBuf.append( 'x' );
break;
case 0x79:
hexBuf.append( "79 " );
asciiBuf.append( 'y' );
break;
case 0x7A:
hexBuf.append( "7A " );
asciiBuf.append( 'z' );
break;
case 0x7B:
hexBuf.append( "7B " );
asciiBuf.append( '{' );
break;
case 0x7C:
hexBuf.append( "7C " );
asciiBuf.append( '|' );
break;
case 0x7D:
hexBuf.append( "7D " );
asciiBuf.append( '}' );
break;
case 0x7E:
hexBuf.append( "7E " );
asciiBuf.append( '~' );
break;
case 0x7F:
hexBuf.append( "7F " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x80:
hexBuf.append( "80 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x81:
hexBuf.append( "81 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x82:
hexBuf.append( "82 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x83:
hexBuf.append( "83 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x84:
hexBuf.append( "84 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x85:
hexBuf.append( "85 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x86:
hexBuf.append( "86 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x87:
hexBuf.append( "87 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x88:
hexBuf.append( "88 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x89:
hexBuf.append( "89 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x8A:
hexBuf.append( "8A " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x8B:
hexBuf.append( "8B " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x8C:
hexBuf.append( "8C " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x8D:
hexBuf.append( "8D " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x8E:
hexBuf.append( "8E " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x8F:
hexBuf.append( "8F " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x90:
hexBuf.append( "90 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x91:
hexBuf.append( "91 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x92:
hexBuf.append( "92 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x93:
hexBuf.append( "93 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x94:
hexBuf.append( "94 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x95:
hexBuf.append( "95 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x96:
hexBuf.append( "96 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x97:
hexBuf.append( "97 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x98:
hexBuf.append( "98 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x99:
hexBuf.append( "99 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x9A:
hexBuf.append( "9A " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x9B:
hexBuf.append( "9B " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x9C:
hexBuf.append( "9C " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x9D:
hexBuf.append( "9D " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x9E:
hexBuf.append( "9E " );
asciiBuf.append( '.' );
break;
case ( byte ) 0x9F:
hexBuf.append( "9F " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA0:
hexBuf.append( "A0 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA1:
hexBuf.append( "A1 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA2:
hexBuf.append( "A2 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA3:
hexBuf.append( "A3 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA4:
hexBuf.append( "A4 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA5:
hexBuf.append( "A5 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA6:
hexBuf.append( "A6 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA7:
hexBuf.append( "A7 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA8:
hexBuf.append( "A8 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xA9:
hexBuf.append( "A9 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xAA:
hexBuf.append( "AA " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xAB:
hexBuf.append( "AB " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xAC:
hexBuf.append( "AC " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xAD:
hexBuf.append( "AD " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xAE:
hexBuf.append( "AE " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xAF:
hexBuf.append( "AF " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB0:
hexBuf.append( "B0 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB1:
hexBuf.append( "B1 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB2:
hexBuf.append( "B2 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB3:
hexBuf.append( "B3 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB4:
hexBuf.append( "B4 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB5:
hexBuf.append( "B5 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB6:
hexBuf.append( "B6 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB7:
hexBuf.append( "B7 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB8:
hexBuf.append( "B8 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xB9:
hexBuf.append( "B9 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xBA:
hexBuf.append( "BA " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xBB:
hexBuf.append( "BB " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xBC:
hexBuf.append( "BC " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xBD:
hexBuf.append( "BD " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xBE:
hexBuf.append( "BE " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xBF:
hexBuf.append( "BF " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC0:
hexBuf.append( "C0 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC1:
hexBuf.append( "C1 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC2:
hexBuf.append( "C2 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC3:
hexBuf.append( "C3 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC4:
hexBuf.append( "C4 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC5:
hexBuf.append( "C5 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC6:
hexBuf.append( "C6 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC7:
hexBuf.append( "C7 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC8:
hexBuf.append( "C8 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xC9:
hexBuf.append( "C9 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xCA:
hexBuf.append( "CA " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xCB:
hexBuf.append( "CB " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xCC:
hexBuf.append( "CC " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xCD:
hexBuf.append( "CD " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xCE:
hexBuf.append( "CE " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xCF:
hexBuf.append( "CF " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD0:
hexBuf.append( "D0 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD1:
hexBuf.append( "D1 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD2:
hexBuf.append( "D2 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD3:
hexBuf.append( "D3 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD4:
hexBuf.append( "D4 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD5:
hexBuf.append( "D5 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD6:
hexBuf.append( "D6 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD7:
hexBuf.append( "D7 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD8:
hexBuf.append( "D8 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xD9:
hexBuf.append( "D9 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xDA:
hexBuf.append( "DA " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xDB:
hexBuf.append( "DB " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xDC:
hexBuf.append( "DC " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xDD:
hexBuf.append( "DD " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xDE:
hexBuf.append( "DE " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xDF:
hexBuf.append( "DF " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE0:
hexBuf.append( "E0 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE1:
hexBuf.append( "E1 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE2:
hexBuf.append( "E2 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE3:
hexBuf.append( "E3 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE4:
hexBuf.append( "E4 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE5:
hexBuf.append( "E5 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE6:
hexBuf.append( "E6 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE7:
hexBuf.append( "E7 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE8:
hexBuf.append( "E8 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xE9:
hexBuf.append( "E9 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xEA:
hexBuf.append( "EA " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xEB:
hexBuf.append( "EB " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xEC:
hexBuf.append( "EC " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xED:
hexBuf.append( "ED " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xEE:
hexBuf.append( "EE " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xEF:
hexBuf.append( "EF " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF0:
hexBuf.append( "F0 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF1:
hexBuf.append( "F1 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF2:
hexBuf.append( "F2 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF3:
hexBuf.append( "F3 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF4:
hexBuf.append( "F4 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF5:
hexBuf.append( "F5 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF6:
hexBuf.append( "F6 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF7:
hexBuf.append( "F7 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF8:
hexBuf.append( "F8 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xF9:
hexBuf.append( "F9 " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xFA:
hexBuf.append( "FA " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xFB:
hexBuf.append( "FB " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xFC:
hexBuf.append( "FC " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xFD:
hexBuf.append( "FD " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xFE:
hexBuf.append( "FE " );
asciiBuf.append( '.' );
break;
case ( byte ) 0xFF:
hexBuf.append( "FF " );
asciiBuf.append( '.' );
break;
default:
break;
}
if ( ( i % 16 ) == 15 )
{
buffer.append( indentBuf ).append( hexBuf ).append( ' ' ).append( asciiBuf ).append( ASN1Element.EOL );
hexBuf = new StringBuilder();
asciiBuf = new StringBuilder();
} else if ( ( i % 8 ) == 7 )
{
hexBuf.append( ' ' );
asciiBuf.append( ' ' );
}
}
final int charsLeft = 16 - ( byteArray.length % 16 );
if ( charsLeft < 16 )
{
for ( int i = 0; i < charsLeft; i++ )
{
hexBuf.append( " " );
}
if ( charsLeft > 8 )
{
hexBuf.append( ' ' );
}
}
buffer.append( indentBuf ).append( hexBuf ).append( ' ' ).append( asciiBuf ).append( ASN1Element.EOL );
return buffer.toString();
}
/**
* Decodes the provided byte array as an ASN.1 Boolean element.
*
* @param encodedValue
* The encoded byte array to decode as an ASN.1 Boolean element.
* @return The decoded ASN.1 Boolean element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 Boolean element.
*/
private static ASN1Boolean decodeAsBoolean( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
if ( encodedValue.length != 3 )
{
final String msg = "!!! Expected 3 bytes in encoded value, but " + encodedValue.length + " bytes exist !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
if ( encodedValue[ 1 ] != 0x01 )
{
final String msg = "!!! Length of a Boolean element is not 1 !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
final boolean booleanValue = encodedValue[ 2 ] != 0x00;
return new ASN1Boolean( type, booleanValue );
}
/**
* Decodes the provided byte array as an ASN.1 enumerated element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 enumerated element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 enumerated element.
*/
private static ASN1Enumerated decodeAsEnumerated( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 3 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final byte[] value = ASN1Element.decodeValue( encodedValue );
final int intValue = ASN1Element.decodeIntValue( value );
final ASN1Enumerated enumeratedElement = new ASN1Enumerated( type );
enumeratedElement.intValue = intValue;
enumeratedElement.value = value;
return enumeratedElement;
}
/**
* Decodes the provided byte array as an ASN.1 null element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 null element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 null element.
*/
private static ASN1Null decodeAsNull( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 2 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
if ( encodedValue[ 1 ] != 0x00 )
{
final String msg = "!!! Length of null should be zero, not " + encodedValue[ 1 ] + " !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
return new ASN1Null( encodedValue[ 0 ] );
}
/**
* Decodes the provided byte array as an ASN.1 octet string element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 octet string element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 octet string element.
*/
private static ASN1OctetString decodeAsOctetString( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 2 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final byte[] value = ASN1Element.decodeValue( encodedValue );
return new ASN1OctetString( type, value );
}
/**
* Decodes the provided byte array as an ASN.1 set element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 set element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 set element.
*/
private static ASN1Set decodeAsSet( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 2 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final byte[] value = ASN1Element.decodeValue( encodedValue );
ASN1Element[] elements = new ASN1Element[ 0 ];
if ( value.length > 0 )
{
elements = ASN1Sequence.decodeSequenceElements( value );
}
final ASN1Set set = new ASN1Set( type );
set.setValue( value );
set.elements = elements;
return set;
}
/**
* This method converts the provided byte array into a Java int.
*
* @param encodedValue
* The byte array containing the value to decode as an integer.
* @return The Java int decoded from the provided byte array.
* @throws ASN1Exception
* If the provided byte array cannot be converted into a Java int.
*/
private static int decodeIntValue( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
int value = 0x00000000;
switch ( encodedValue.length )
{
case 1:
value |= 0x000000FF & encodedValue[ 0 ];
break;
case 2:
value |= ( ( 0x000000FF & encodedValue[ 0 ] ) << 8 ) | ( 0x000000FF & encodedValue[ 1 ] );
break;
case 3:
value |= ( ( 0x000000FF & encodedValue[ 0 ] ) << 16 ) | ( ( 0x000000FF & encodedValue[ 1 ] ) << 8 ) | ( 0x000000FF & encodedValue[ 2 ] );
break;
case 4:
value |= ( ( 0x000000FF & encodedValue[ 0 ] ) << 24 ) | ( ( 0x000000FF & encodedValue[ 1 ] ) << 16 ) | ( ( 0x000000FF & encodedValue[ 2 ] ) << 8 ) | ( 0x000000FF & encodedValue[ 3 ] );
break;
default:
final String msg = "!!! The provided value cannot be represented as a Java int !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
return value;
}
/**
* Decodes the provided byte array as a length.
*
* @param encodedLength
* The encoded value to decode as a length.
* @return The length decoded from the provided byte array.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 length.
*/
private static int decodeLength( final byte[] encodedLength ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedLength );
if ( encodedLength.length == 1 )
{
if ( ( encodedLength[ 0 ] & 0x7F ) == encodedLength[ 0 ] )
{
return encodedLength[ 0 ];
} else if ( ( encodedLength[ 0 ] & 0x7F ) == 0 )
{
return 128;
} else
{
final String msg = "!!! Only one byte in length, but it is an invalid value !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
}
if ( ( encodedLength[ 0 ] & 0x7F ) != encodedLength[ 0 ] )
{
final int numLengthBytes = encodedLength[ 0 ] & 0x7F;
if ( numLengthBytes == ( encodedLength.length - 1 ) )
{
if ( numLengthBytes <= 4 )
{
final byte[] byteArray = new byte[ numLengthBytes ];
System.arraycopy( encodedLength, 1, byteArray, 0, byteArray.length );
return ASN1Element.byteArrayToInt( byteArray );
}
final String msg = "!!! Unable to represent length as a Java int !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
final String msg = "!!! Encoded length indicates " + numLengthBytes + " bytes in length, but " + ( encodedLength.length - 1 ) + " bytes were provided !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
final String msg = "!!! Multiple bytes in length, but first byte does not have 0x80 bit set !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
/**
* Decode the byte array.
*
* @param encodedValue
* The byte array to decode.
* @return The decoded byte array.
* @throws ASN1Exception
* This exception was raised whenever an error occurred during the decode process.
*/
private static byte[] decodeValue( final byte[] encodedValue ) throws ASN1Exception
{
final int length;
int valueStartPos = 2;
if ( ( encodedValue[ 1 ] & 0x7F ) != encodedValue[ 1 ] )
{
if ( ( encodedValue[ 1 ] & 0x7F ) == 0x00 )
{
length = 128;
} else
{
final int numLengthBytes = encodedValue[ 1 ] & 0x7F;
if ( encodedValue.length < ( numLengthBytes + 2 ) )
{
final String msg = "!!! Determined the length is encoded in " + numLengthBytes + " bytes, but not enough bytes exist in the encoded value !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
final byte[] lengthArray = new byte[ numLengthBytes + 1 ];
lengthArray[ 0 ] = encodedValue[ 1 ];
System.arraycopy( encodedValue, 2, lengthArray, 1, numLengthBytes );
length = ASN1Element.decodeLength( lengthArray );
valueStartPos += numLengthBytes;
}
} else
{
length = encodedValue[ 1 ];
}
final byte[] value = new byte[ length ];
System.arraycopy( encodedValue, valueStartPos, value, 0, length );
return value;
}
/**
* Throw an error if the byte array to encode is multibyte.
*
* @param encodedValue
* The byte array to encode.
* @throws ASN1Exception
* This exception was raised if the byte array to encode is multibyte.
*/
private static void throwErrorIfIsMultibyte( final byte[] encodedValue ) throws ASN1Exception
{
if ( ( encodedValue[ 0 ] & 0x1F ) == 0x1F )
{
final String msg = "!!! Multibyte type detected (not supported in this package) !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
}
/**
* Throw an exception if there is no data to encode.
*
* @param encodedValue
* The byte array data.
* @throws ASN1Exception
* This exception was raised if there is no data to encode.
*/
private static void throwErrorIfNoData( final byte[] encodedValue ) throws ASN1Exception
{
if ( ( encodedValue == null ) || ( encodedValue.length == 0 ) )
{
final String msg = "!!! No data to decode !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
}
/**
* Throw an exception if there was not enough data to encode.
*
* @param encodedValue
* The byte array where verify the length.
* @param totaldata
* The total data length
* @throws ASN1Exception
* This exception was raised if there was not enough data.
*/
private static void throwIfNoEnoughData( final byte[] encodedValue, final int totaldata ) throws ASN1Exception
{
if ( encodedValue.length < totaldata )
{
final String msg = "!!! Not enough data to make a valid ASN.1 element !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
}
/**
* Throw an exception if there was a wrong length.
*
* @param encodedValue
* The byte array where verify length.
* @param valueStartPos
* The value start position in the array.
* @param length
* The length of the data.
* @throws ASN1Exception
* This exception was raised if length was wrong.
*/
private static void throwIfWrongLength( final byte[] encodedValue, final int valueStartPos, final int length ) throws ASN1Exception
{
if ( ( encodedValue.length - valueStartPos ) != length )
{
final String msg = "!!! Expected a value of " + length + " bytes, but " + ( encodedValue.length - valueStartPos ) + " bytes exist !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
}
/**
* Add a new element to into the set.
*
* @param elements
* The array where insert the new element.
* @param element
* The new element to insert.
* @return The array where was insert the new element.
*/
protected static ASN1Element[] addElementToSet( final ASN1Element[] elements, final ASN1Element element )
{
final ASN1Element[] newElements = new ASN1Element[ elements.length + 1 ];
System.arraycopy( elements, 0, newElements, 0, elements.length );
newElements[ elements.length ] = element;
return newElements;
}
/**
* Retrieves a string containing the hexadecimal digits contained in the provided byte array. Each line will be indented the specified number of spaces.
*
* @param byteArray
* The byte array containing the information to be output.
* @param indent
* The number of spaces to indent each line of the output.
* @return A string containing the hexadecimal digits contained in the provided byte array.
*/
static String byteArrayToString( final byte[] byteArray, final int indent )
{
String indentStr = "";
for ( int i = 0; i < indent; i++ )
{
indentStr += " ";
}
String returnStr = indentStr;
for ( int i = 0; i < byteArray.length; i++ )
{
final String hexStr = Integer.toHexString( 0x000000FF & byteArray[ i ] );
if ( hexStr.length() == 1 )
{
returnStr += "0";
}
returnStr += hexStr;
returnStr += ( ( i + 1 ) % 16 ) == 0 ? ASN1Element.EOL + indentStr : " ";
}
if ( ( byteArray.length % 16 ) != 0 )
{
returnStr += ASN1Element.EOL;
}
return returnStr;
}
/**
* Encodes the specified length as a byte array as it should appear in an ASN.1 element.
*
* @param length
* The length value to be encoded.
* @return The encoded length as a byte array.
*/
static byte[] encodeLength( final int length )
{
// First, see if the int value is within the first 128 values. If so, then
// just return the pre-encoded version.
byte[] returnArray = ASN1Element.returnPreEncodedLengthVersion( length );
if ( returnArray != null )
{
return returnArray;
}
if ( ( length & 0xFF000000 ) != 0 )
{
returnArray = new byte[ 5 ];
returnArray[ 0 ] = ( byte ) 0x84;
returnArray[ 1 ] = ( byte ) ( ( length & 0xFF000000 ) >>> 24 );
returnArray[ 2 ] = ( byte ) ( ( length & 0x00FF0000 ) >>> 16 );
returnArray[ 3 ] = ( byte ) ( ( length & 0x0000FF00 ) >>> 8 );
returnArray[ 4 ] = ( byte ) ( length & 0x000000FF );
return returnArray;
} else if ( ( length & 0x00FF0000 ) != 0 )
{
returnArray = new byte[ 4 ];
returnArray[ 0 ] = ( byte ) 0x83;
returnArray[ 1 ] = ( byte ) ( ( length & 0x00FF0000 ) >>> 16 );
returnArray[ 2 ] = ( byte ) ( ( length & 0x0000FF00 ) >>> 8 );
returnArray[ 3 ] = ( byte ) ( length & 0x000000FF );
return returnArray;
} else if ( ( length & 0x0000FF00 ) != 0 )
{
returnArray = new byte[ 3 ];
returnArray[ 0 ] = ( byte ) 0x82;
returnArray[ 1 ] = ( byte ) ( ( length & 0x0000FF00 ) >>> 8 );
returnArray[ 2 ] = ( byte ) ( length & 0x000000FF );
return returnArray;
} else
{
returnArray = new byte[ 2 ];
returnArray[ 0 ] = ( byte ) 0x81;
returnArray[ 1 ] = ( byte ) ( length & 0x000000FF );
return returnArray;
}
}
/**
* Retrieves a byte array containing the binary representation of the provided string. If the provided string is 7-bit clean, then this method is about 5 times faster than the standard Java <CODE>String.getBytes()</CODE> method.
*
* @param stringValue
* The string for which to retrieve the binary representation.
* @return A byte array containing the binary representation of the provided string.
*/
static byte[] getBytes( final String stringValue )
{
if ( stringValue == null )
{
return new byte[ 0 ];
}
byte b;
char c;
final byte[] returnArray = new byte[ stringValue.length() ];
for ( int i = 0; i < returnArray.length; i++ )
{
c = stringValue.charAt( i );
b = ( byte ) ( c & 0x0000007F );
if ( b == c )
{
returnArray[ i ] = b;
} else
{
return stringValue.getBytes();
}
}
return returnArray;
}
/**
* Return the pre-encoded length.
*
* @param length
* The length to check.
* @return The pre-encoded length.
*/
static byte[] returnPreEncodedLengthVersion( final int length )
{
switch ( length )
{
case 0:
return ASN1Integer.INT_VALUE_0;
case 1:
return ASN1Integer.INT_VALUE_1;
case 2:
return ASN1Integer.INT_VALUE_2;
case 3:
return ASN1Integer.INT_VALUE_3;
case 4:
return ASN1Integer.INT_VALUE_4;
case 5:
return ASN1Integer.INT_VALUE_5;
case 6:
return ASN1Integer.INT_VALUE_6;
case 7:
return ASN1Integer.INT_VALUE_7;
case 8:
return ASN1Integer.INT_VALUE_8;
case 9:
return ASN1Integer.INT_VALUE_9;
case 10:
return ASN1Integer.INT_VALUE_10;
case 11:
return ASN1Integer.INT_VALUE_11;
case 12:
return ASN1Integer.INT_VALUE_12;
case 13:
return ASN1Integer.INT_VALUE_13;
case 14:
return ASN1Integer.INT_VALUE_14;
case 15:
return ASN1Integer.INT_VALUE_15;
case 16:
return ASN1Integer.INT_VALUE_16;
case 17:
return ASN1Integer.INT_VALUE_17;
case 18:
return ASN1Integer.INT_VALUE_18;
case 19:
return ASN1Integer.INT_VALUE_19;
case 20:
return ASN1Integer.INT_VALUE_20;
case 21:
return ASN1Integer.INT_VALUE_21;
case 22:
return ASN1Integer.INT_VALUE_22;
case 23:
return ASN1Integer.INT_VALUE_23;
case 24:
return ASN1Integer.INT_VALUE_24;
case 25:
return ASN1Integer.INT_VALUE_25;
case 26:
return ASN1Integer.INT_VALUE_26;
case 27:
return ASN1Integer.INT_VALUE_27;
case 28:
return ASN1Integer.INT_VALUE_28;
case 29:
return ASN1Integer.INT_VALUE_29;
case 30:
return ASN1Integer.INT_VALUE_30;
case 31:
return ASN1Integer.INT_VALUE_31;
case 32:
return ASN1Integer.INT_VALUE_32;
case 33:
return ASN1Integer.INT_VALUE_33;
case 34:
return ASN1Integer.INT_VALUE_34;
case 35:
return ASN1Integer.INT_VALUE_35;
case 36:
return ASN1Integer.INT_VALUE_36;
case 37:
return ASN1Integer.INT_VALUE_37;
case 38:
return ASN1Integer.INT_VALUE_38;
case 39:
return ASN1Integer.INT_VALUE_39;
case 40:
return ASN1Integer.INT_VALUE_40;
case 41:
return ASN1Integer.INT_VALUE_41;
case 42:
return ASN1Integer.INT_VALUE_42;
case 43:
return ASN1Integer.INT_VALUE_43;
case 44:
return ASN1Integer.INT_VALUE_44;
case 45:
return ASN1Integer.INT_VALUE_45;
case 46:
return ASN1Integer.INT_VALUE_46;
case 47:
return ASN1Integer.INT_VALUE_47;
case 48:
return ASN1Integer.INT_VALUE_48;
case 49:
return ASN1Integer.INT_VALUE_49;
case 50:
return ASN1Integer.INT_VALUE_50;
case 51:
return ASN1Integer.INT_VALUE_51;
case 52:
return ASN1Integer.INT_VALUE_52;
case 53:
return ASN1Integer.INT_VALUE_53;
case 54:
return ASN1Integer.INT_VALUE_54;
case 55:
return ASN1Integer.INT_VALUE_55;
case 56:
return ASN1Integer.INT_VALUE_56;
case 57:
return ASN1Integer.INT_VALUE_57;
case 58:
return ASN1Integer.INT_VALUE_58;
case 59:
return ASN1Integer.INT_VALUE_59;
case 60:
return ASN1Integer.INT_VALUE_60;
case 61:
return ASN1Integer.INT_VALUE_61;
case 62:
return ASN1Integer.INT_VALUE_62;
case 63:
return ASN1Integer.INT_VALUE_63;
case 64:
return ASN1Integer.INT_VALUE_64;
case 65:
return ASN1Integer.INT_VALUE_65;
case 66:
return ASN1Integer.INT_VALUE_66;
case 67:
return ASN1Integer.INT_VALUE_67;
case 68:
return ASN1Integer.INT_VALUE_68;
case 69:
return ASN1Integer.INT_VALUE_69;
case 70:
return ASN1Integer.INT_VALUE_70;
case 71:
return ASN1Integer.INT_VALUE_71;
case 72:
return ASN1Integer.INT_VALUE_72;
case 73:
return ASN1Integer.INT_VALUE_73;
case 74:
return ASN1Integer.INT_VALUE_74;
case 75:
return ASN1Integer.INT_VALUE_75;
case 76:
return ASN1Integer.INT_VALUE_76;
case 77:
return ASN1Integer.INT_VALUE_77;
case 78:
return ASN1Integer.INT_VALUE_78;
case 79:
return ASN1Integer.INT_VALUE_79;
case 80:
return ASN1Integer.INT_VALUE_80;
case 81:
return ASN1Integer.INT_VALUE_81;
case 82:
return ASN1Integer.INT_VALUE_82;
case 83:
return ASN1Integer.INT_VALUE_83;
case 84:
return ASN1Integer.INT_VALUE_84;
case 85:
return ASN1Integer.INT_VALUE_85;
case 86:
return ASN1Integer.INT_VALUE_86;
case 87:
return ASN1Integer.INT_VALUE_87;
case 88:
return ASN1Integer.INT_VALUE_88;
case 89:
return ASN1Integer.INT_VALUE_89;
case 90:
return ASN1Integer.INT_VALUE_90;
case 91:
return ASN1Integer.INT_VALUE_91;
case 92:
return ASN1Integer.INT_VALUE_92;
case 93:
return ASN1Integer.INT_VALUE_93;
case 94:
return ASN1Integer.INT_VALUE_94;
case 95:
return ASN1Integer.INT_VALUE_95;
case 96:
return ASN1Integer.INT_VALUE_96;
case 97:
return ASN1Integer.INT_VALUE_97;
case 98:
return ASN1Integer.INT_VALUE_98;
case 99:
return ASN1Integer.INT_VALUE_99;
case 100:
return ASN1Integer.INT_VALUE_100;
case 101:
return ASN1Integer.INT_VALUE_101;
case 102:
return ASN1Integer.INT_VALUE_102;
case 103:
return ASN1Integer.INT_VALUE_103;
case 104:
return ASN1Integer.INT_VALUE_104;
case 105:
return ASN1Integer.INT_VALUE_105;
case 106:
return ASN1Integer.INT_VALUE_106;
case 107:
return ASN1Integer.INT_VALUE_107;
case 108:
return ASN1Integer.INT_VALUE_108;
case 109:
return ASN1Integer.INT_VALUE_109;
case 110:
return ASN1Integer.INT_VALUE_110;
case 111:
return ASN1Integer.INT_VALUE_111;
case 112:
return ASN1Integer.INT_VALUE_112;
case 113:
return ASN1Integer.INT_VALUE_113;
case 114:
return ASN1Integer.INT_VALUE_114;
case 115:
return ASN1Integer.INT_VALUE_115;
case 116:
return ASN1Integer.INT_VALUE_116;
case 117:
return ASN1Integer.INT_VALUE_117;
case 118:
return ASN1Integer.INT_VALUE_118;
case 119:
return ASN1Integer.INT_VALUE_119;
case 120:
return ASN1Integer.INT_VALUE_120;
case 121:
return ASN1Integer.INT_VALUE_121;
case 122:
return ASN1Integer.INT_VALUE_122;
case 123:
return ASN1Integer.INT_VALUE_123;
case 124:
return ASN1Integer.INT_VALUE_124;
case 125:
return ASN1Integer.INT_VALUE_125;
case 126:
return ASN1Integer.INT_VALUE_126;
case 127:
return ASN1Integer.INT_VALUE_127;
default:
return null;
}
}
/**
* Retrieves a string containing the hexadecimal digits contained in the provided byte array.
*
* @param byteArray
* The byte array containing the information to be output.
* @return A string containing the hexadecimal digits contained in the provided byte array.
*/
public static String byteArrayToString( final byte[] byteArray )
{
return ASN1Element.byteArrayToString( byteArray, 0 );
}
/**
* Retrieves a string representation of the provided byte array (including the ASCII equivalent).
*
* @param byteArray
* The byte array to be displayed as a string.
* @return A string representation of the provided byte array with the ASCII equivalent.
*/
public static String byteArrayToStringWithASCII( final byte[] byteArray )
{
return ASN1Element.byteArrayToStringWithASCII( byteArray, 0 );
}
/**
* Decodes the provided byte array as a generic ASN.1 element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 element.
*/
public static ASN1Element decode( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 2 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final int length;
int valueStartPos = 2;
if ( ( encodedValue[ 1 ] & 0x7F ) != encodedValue[ 1 ] )
{
if ( ( encodedValue[ 1 ] & 0x7F ) == 0x00 )
{
length = 128;
} else
{
final int numLengthBytes = encodedValue[ 1 ] & 0x7F;
if ( encodedValue.length < ( numLengthBytes + 2 ) )
{
final String msg = "!!! Determined the length is encoded in " + numLengthBytes + " bytes, but only " + encodedValue.length + "bytes exist in the encoded value !!!";
final ASN1Exception ex = new ASN1Exception( msg );
ASN1Element.LOG.error( msg, ex );
throw ex;
}
final byte[] lengthArray = new byte[ numLengthBytes + 1 ];
lengthArray[ 0 ] = encodedValue[ 1 ];
System.arraycopy( encodedValue, 2, lengthArray, 1, numLengthBytes );
length = ASN1Element.decodeLength( lengthArray );
valueStartPos += numLengthBytes;
}
} else
{
length = encodedValue[ 1 ];
}
ASN1Element.throwIfWrongLength( encodedValue, valueStartPos, length );
final byte[] value = new byte[ length ];
System.arraycopy( encodedValue, valueStartPos, value, 0, length );
return new ASN1Element( type, value, encodedValue );
}
/**
* Decodes the provided byte array as an ASN.1 Integer element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 Integer element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 integer element.
*/
public static ASN1Integer decodeAsInteger( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 3 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final byte[] value = ASN1Element.decodeValue( encodedValue );
final int intValue = ASN1Element.decodeIntValue( value );
return new ASN1Integer( type, intValue );
}
/**
* Decodes the provided byte array as an ASN.1 sequence element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The decoded ASN.1 sequence element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 sequence element.
*/
public static ASN1Sequence decodeAsSequence( final byte[] encodedValue ) throws ASN1Exception
{
ASN1Element.throwErrorIfNoData( encodedValue );
ASN1Element.throwIfNoEnoughData( encodedValue, 2 );
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final byte[] value = ASN1Element.decodeValue( encodedValue );
ASN1Element[] elements = ASN1Sequence.NO_ELEMENTS;
if ( value.length > 0 )
{
elements = ASN1Sequence.decodeSequenceElements( value );
}
final ASN1Sequence sequence = new ASN1Sequence( type );
sequence.setValue( value );
sequence.elements = elements;
return sequence;
}
/**
* Decodes the provided byte array as a generic ASN.1 element. The provided array may contain a partial element, exactly a complete element, or a complete element plus some additional data. The beginning of the data in the array must be the beginning of an ASN.1 element.
*
* @param encodedValue
* The encoded ASN.1 element.
* @return The result of attempting to decode the element.
* @throws ASN1Exception
* If the provided byte array cannot be decoded as an ASN.1 element.
*/
public static ASN1DecodeResult decodePartial( final byte[] encodedValue ) throws ASN1Exception
{
if ( ( encodedValue == null ) || ( encodedValue.length < 2 ) )
{
return new ASN1DecodeResult( null, null, 0 );
}
ASN1Element.throwErrorIfIsMultibyte( encodedValue );
final byte type = encodedValue[ 0 ];
final int length;
int valueStartPos = 2;
if ( ( encodedValue[ 1 ] & 0x7F ) != encodedValue[ 1 ] )
{
if ( ( encodedValue[ 1 ] & 0x7F ) == 0x00 )
{
length = 128;
} else
{
final int numLengthBytes = encodedValue[ 1 ] & 0x7F;
if ( encodedValue.length < ( numLengthBytes + 2 ) )
{
return new ASN1DecodeResult( null, null, 0 );
}
final byte[] lengthArray = new byte[ numLengthBytes + 1 ];
lengthArray[ 0 ] = encodedValue[ 1 ];
System.arraycopy( encodedValue, 2, lengthArray, 1, numLengthBytes );
length = ASN1Element.decodeLength( lengthArray );
valueStartPos += numLengthBytes;
}
} else
{
length = encodedValue[ 1 ];
}
final int numExtraBytes = encodedValue.length - valueStartPos - length;
if ( numExtraBytes < 0 )
{
return new ASN1DecodeResult( null, null, 0 );
}
final byte[] value = new byte[ length ];
System.arraycopy( encodedValue, valueStartPos, value, 0, length );
final ASN1Element element = new ASN1Element( type, value, encodedValue );
final byte[] remainingData;
if ( numExtraBytes > 0 )
{
remainingData = new byte[ numExtraBytes ];
System.arraycopy( encodedValue, valueStartPos + length, remainingData, 0, numExtraBytes );
} else
{
remainingData = null;
}
return new ASN1DecodeResult( element, remainingData, length );
}
/**
* Append the new element into the value.
*
* @param element
* The new element to insert into the value.
* @return The new value after element insertion.
*/
protected byte[] appendNewElementsByteToValue( final ASN1Element element )
{
final byte[] encodedElementTmp = element.encode();
final byte[] newSequenceValue = new byte[ this.value.length + encodedElementTmp.length ];
System.arraycopy( this.value, 0, newSequenceValue, 0, this.value.length );
System.arraycopy( encodedElementTmp, 0, newSequenceValue, this.value.length, encodedElementTmp.length );
return newSequenceValue;
}
/**
* Encodes this ASN.1 element into the provided byte array at the indicated position.
*
* @param byteArray
* The byte array into which the value is to be encoded.
* @param startPos
* The position in the byte array at which to start writing the encoded value.
* @return The number of bytes written into the array, or -1 if the array was not big enough to hold the encoded value.
*/
int encode( final byte[] byteArray, final int startPos )
{
// First, make sure there is enough space in the provided byte array.
final int length = this.encodedElement.length;
if ( ( startPos + length ) > byteArray.length )
{
return -1;
}
// Encode the value into the array.
System.arraycopy( this.encodedElement, 0, byteArray, startPos, this.encodedElement.length );
return length;
}
/**
* Specifies the value for this ASN.1 element.
*
* @param value
* The value for this ASN.1 element.
*/
void setValue( final byte[] value )
{
this.value = value == null ? ASN1Element.EMPTY_BYTES : value;
final byte[] encodedLength = ASN1Element.encodeLength( this.value.length );
this.encodedElement = new byte[ 1 + encodedLength.length + this.value.length ];
this.encodedElement[ 0 ] = this.type;
System.arraycopy( encodedLength, 0, this.encodedElement, 1, encodedLength.length );
System.arraycopy( this.value, 0, this.encodedElement, 1 + encodedLength.length, this.value.length );
}
/**
* Retrieves a string representation of this ASN.1 element. The information will be indented the specified number of spaces.
*
* @param indent
* The number of spaces to indent each line of the output.
* @return A string representation of this ASN.1 element.
*/
String toString( final int indent )
{
String indentStr = "";
for ( int i = 0; i < indent; i++ )
{
indentStr += " ";
}
return indentStr + "Type: " + this.type + ASN1Element.EOL + ASN1Element.byteArrayToString( new byte[] { this.type }, 2 + indent ) + ASN1Element.EOL + indentStr + "Length: " + this.value.length + ASN1Element.EOL + ASN1Element.byteArrayToString( ASN1Element.encodeLength( this.value.length ), 2 + indent ) + ASN1Element.EOL + indentStr + "Value: " + new String( this.value ) + ASN1Element.EOL + ASN1Element.byteArrayToString( this.value, 2 + indent ) + ASN1Element.EOL;
}
/**
* Decodes this element as an ASN.1 Boolean element.
*
* @return The decoded ASN.1 Boolean element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 Boolean element.
*/
public ASN1Boolean decodeAsBoolean() throws ASN1Exception
{
return ASN1Element.decodeAsBoolean( this.encode() );
}
/**
* Decodes this element as an ASN.1 enumerated element.
*
* @return The decoded ASN.1 enumerated element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 enumerated element.
*/
public ASN1Enumerated decodeAsEnumerated() throws ASN1Exception
{
return ASN1Element.decodeAsEnumerated( this.encodedElement );
}
/**
* Decodes this element as an ASN.1 integer element.
*
* @return The decoded ASN.1 integer element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 integer element.
*/
public ASN1Integer decodeAsInteger() throws ASN1Exception
{
return ASN1Element.decodeAsInteger( this.encodedElement );
}
/**
* Decodes this element as an ASN.1 null element.
*
* @return The decoded ASN.1 null element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 null element.
*/
public ASN1Null decodeAsNull() throws ASN1Exception
{
return ASN1Element.decodeAsNull( this.encodedElement );
}
/**
* Decodes this element as an ASN.1 octet string element.
*
* @return The decoded ASN.1 octet string element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 octet string element.
*/
public ASN1OctetString decodeAsOctetString() throws ASN1Exception
{
return ASN1Element.decodeAsOctetString( this.encodedElement );
}
/**
* Decodes this element as an ASN.1 sequence element.
*
* @return The decoded ASN.1 sequence element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 sequence element.
*/
public ASN1Sequence decodeAsSequence() throws ASN1Exception
{
return ASN1Element.decodeAsSequence( this.encodedElement );
}
/**
* Decodes this element as an ASN.1 set element.
*
* @return The decoded ASN.1 set element.
* @throws ASN1Exception
* If this element cannot be decoded as an ASN.1 set element.
*/
public ASN1Set decodeAsSet() throws ASN1Exception
{
return ASN1Element.decodeAsSet( this.encodedElement );
}
/**
* Encodes this ASN.1 element into a byte array.
*
* @return This ASN.1 element encoded as a byte array.
*/
public byte[] encode()
{
return this.encodedElement;
}
/**
* Retrieves the id of the element.
*
* @return The id of the element.
*/
public long getId()
{
return this.id;
}
/**
* Gets the type of this ASN.1 element.
*
* @return The type of this ASN.1 element.
*/
public byte getType()
{
return this.type;
}
/**
* Gets the type of this ASN.1 element without either of the class bits or the primitive/constructed bit set (i.e., the least significant five bits).
*
* @return The type of this ASN.1 element without any of the flag bits set.
*/
public byte getTypeWithoutFlags()
{
return ( byte ) ( this.type & 0x1F );
}
/**
* Gets the encoded value for this ASN.1 element.
*
* @return The encoded value for this ASN.1 element.
*/
public byte[] getValue()
{
return this.value;
}
/**
* Setting the id of the element.
*
* @param id
* The id of the element.
*/
public void setId( final long id )
{
this.id = id;
}
/**
* Specifies the type for this ASN.1 element.
*
* @param type
* The type for this ASN.1 element.
*/
public void setType( final byte type )
{
this.type = type;
}
/**
* Retrieves a string representation of this ASN.1 element.
*
* @return A string representation of this ASN.1 element.
*/
@Override( )
public String toString()
{
return this.toString( 0 );
}
}
<file_sep>/src/META-INF/Create_RootdseStatistics.sql
CREATE TABLE ldap_tools.rootdsestatisticsbean (
id int8 NOT NULL DEFAULT nextval('ldap_tools.rootdsestatisticsbean_id_seq'::regclass),
caller varchar(255) NULL,
"data" timestamp NULL,
server varchar(255) NULL,
"type" varchar(255) NULL,
value int8 NOT NULL,
CONSTRAINT rootdsestatisticsbean_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
) ;
CREATE INDEX rootdsestatisticsbean_data_idx ON ldap_tools.rootdsestatisticsbean (data DESC) ;
CREATE INDEX rootdsestatisticsbean_server_idx ON ldap_tools.rootdsestatisticsbean (server DESC) ;
CREATE INDEX rootdsestatisticsbean_type_idx ON ldap_tools.rootdsestatisticsbean (type DESC) ;
<file_sep>/src/META-INF/Create_Responsetime.sql
DROP TABLE IF EXISTS ldap_tools.responsetime;
CREATE TABLE ldap_tools.responsetime (
id int8 NOT NULL DEFAULT nextval('ldap_tools.responsetime_id_seq'::regclass),
connectelaps int8 NOT NULL,
"data" timestamp NULL,
executorhost varchar(255) NULL,
extractedentry int4 NOT NULL,
returncode int4 NOT NULL,
searchelaps int8 NOT NULL,
server varchar(255) NULL,
unbindelaps int8 NOT NULL,
CONSTRAINT responsetime_pkey PRIMARY KEY (id)
)
WITH (
OIDS=FALSE
) ;
CREATE INDEX responsetime_data_idx ON ldap_tools.responsetime (data DESC) ;
CREATE INDEX responsetime_server_idx ON ldap_tools.responsetime (server DESC) ;
<file_sep>/src/ita/deluca/tools/ldap/framework/edirectory/passwordpolicy/model/PasswordPolicyFacilitator.java
package ita.deluca.tools.ldap.framework.edirectory.passwordpolicy.model;
import java.util.HashMap;
import java.util.List;
import java.util.Map;
import org.apache.logging.log4j.LogManager;
import org.apache.logging.log4j.Logger;
import ita.deluca.tools.application.framework.ldap.bean.LdapSearchResultBean;
import ita.deluca.tools.application.framework.ldap.exception.LdapException;
import ita.deluca.tools.application.framework.ldap.models.LdapOperations;
import ita.deluca.tools.application.framework.properties.exception.PropertiesException;
import ita.deluca.tools.application.framework.properties.model.PropertiesManager;
import ita.deluca.tools.ldap.framework.edirectory.passwordpolicy.exception.PasswordPolicyFacilitatorException;
/**
* This class contains all the function that enables the user to use the password policy.
*
* @author <NAME>
*/
public class PasswordPolicyFacilitator
{
/**
* The class logger.
*/
private static final Logger LOG = LogManager.getLogger( PasswordPolicyFacilitator.class );
/**
* The selected environment.
*/
private final String environment;
/**
* Ldap operation framework.
*/
private LdapOperations ldap;
/**
* The properties manager.
*/
PropertiesManager properties;
/**
* Standard constructor.
*
* @param environment
* The selected environment.
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever an error occurred during the initialization.
*/
public PasswordPolicyFacilitator( final String environment ) throws PasswordPolicyFacilitatorException
{
this.environment = environment;
try
{
this.properties = PropertiesManager.getPropertiesInstance( "LdapToolConfiguration.xml" );
} catch ( final PropertiesException ex )
{
final String msg = "!!! Error in initialization of the properties manager !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
}
/**
* Execute the preliminary check into the entry where activate the policy.
*
* @param entry
* The entry where activate the policy.
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever the preliminary check fails.
*/
private void entryPreliminaryCheck( final String entry ) throws PasswordPolicyFacilitatorException
{
PasswordPolicyFacilitator.LOG.info( "Verify if the entry '" + entry + "' exist." );
final boolean entryExist;
try
{
entryExist = this.ldap.entryExist( entry );
} catch ( final LdapException ex )
{
final String msg = "!!! Error in search of existence of the entry !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
if ( !entryExist )
{
final String msg = "!!! The entry '" + entry + "' was not found !!!";
final PasswordPolicyFacilitatorException ex = new PasswordPolicyFacilitatorException( msg );
PasswordPolicyFacilitator.LOG.error( msg, ex );
throw ex;
}
PasswordPolicyFacilitator.LOG.info( "Verify if the entry '" + entry + "' has already active policy." );
final List<LdapSearchResultBean> searchResultList;
try
{
searchResultList = this.ldap.search( entry, "OBJECT_SCOPE", "nspmPasswordPolicyDN=*", new String[] { "nspmPasswordPolicyDN" } );
} catch ( final LdapException ex )
{
final String msg = "!!! Error the verification of the entry '" + entry + "' already have a policy !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
if ( !searchResultList.isEmpty() )
{
final String msg = "!!! The entry '" + entry + "' have already active a password policy !!!";
final PasswordPolicyFacilitatorException ex = new PasswordPolicyFacilitatorException( msg );
PasswordPolicyFacilitator.LOG.error( msg, ex );
throw ex;
}
}
/**
* Initialize the ldap connection.
*
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever an error occurred during the ldap initialization.
*/
private void initLdap() throws PasswordPolicyFacilitatorException
{
final String server = this.properties.getPropertiesStringValue( "commons", this.environment, "server", "master" );
final int port = this.properties.getPropertiesIntegerValue( "commons", this.environment, "server", "port" );
final String username = this.properties.getPropertiesStringValue( "commons", this.environment, "adminuser", "username" );
final String password = this.properties.getPropertiesStringValue( "commons", this.environment, "adminuser", "password" );
try
{
this.ldap.connect( server, port, username, password );
} catch ( final LdapException ex )
{
final String msg = "!!! The connection on the ldap server '" + server + "' raised an error !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
}
/**
* Make the effective policy assignments.
*
* @param policyDn
* The policy dn to add.
* @param entry
* The entry where add the policy.
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever an error occurred during the adds of the policy.
*/
private void makePolicyAssignment( final String policyDn, final String entry ) throws PasswordPolicyFacilitatorException
{
PasswordPolicyFacilitator.LOG.info( "Adding the password policy '" + policyDn + "' to the entry '" + entry + "'." );
final Map<String, Object> attributesOnEntry = new HashMap<>();
final Map<String, Object> attributesOnPolicy = new HashMap<>();
attributesOnEntry.put( "nspmPasswordPolicyDN", policyDn );
attributesOnPolicy.put( "nsimAssignments", entry );
try
{
this.ldap.attributeModify( entry, attributesOnEntry, "ADD_ATTRIBUTE" );
this.ldap.attributeModify( policyDn, attributesOnPolicy, "ADD_ATTRIBUTE" );
} catch ( final LdapException ex )
{
final String msg = "!!! The assignment of the policy raised an error !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
}
/**
* Extract the full dn of the policy.
*
* @param policy
* The policy that want the full dn.
* @return The policy full dn.
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever an error occurred during the search of the full dn of the policy.
*/
private String passwordPolicyFullDn( final String policy ) throws PasswordPolicyFacilitatorException
{
PasswordPolicyFacilitator.LOG.info( "Retrieves the full dn of the policy" );
final String basepath = this.properties.getPropertiesStringValue( this.environment, "passwordpolicy", "policy_existence", "basepath" );
final String filter = this.properties.getPropertiesStringValue( this.environment, "passwordpolicy", "policy_existence", "filter" ).replace( "?policy_name?", policy );
final List<String> policyDnList;
try
{
policyDnList = this.ldap.entryFullDn( basepath, "SUBTREE_SCOPE", filter );
} catch ( final LdapException ex )
{
final String msg = "!!! The search of the full dn of the policy '" + policy + "' was raised an error !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
if ( policyDnList.isEmpty() )
{
final String msg = "!!! No policy with name '" + policy + "' was found in the container '" + basepath + "' !!!";
final PasswordPolicyFacilitatorException ex = new PasswordPolicyFacilitatorException( msg );
PasswordPolicyFacilitator.LOG.error( msg, ex );
throw ex;
}
return policyDnList.get( 0 );
}
/**
* Adds the policy to the entry passed as parameter.
*
* @param policy
* The policy to add on the entry.
* @param entry
* The entry where add the policy.
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever an error occurred during the add of the policy.
*/
public void addPasswordPolicy( final String policy, final String entry ) throws PasswordPolicyFacilitatorException
{
this.addPasswordPolicy( policy, new String[] { entry } );
}
/**
* Adds the policy to the entry passed as parameter.
*
* @param policy
* The policy to add on the entry.
* @param entries
* The list of the entry where add the policy.
* @throws PasswordPolicyFacilitatorException
* This exception was raised whenever an error occurred during the add of the policy.
*/
public void addPasswordPolicy( final String policy, final String[] entries ) throws PasswordPolicyFacilitatorException
{
this.initLdap();
try
{
final String policyDn = this.passwordPolicyFullDn( policy );
for ( final String entry : entries )
{
this.entryPreliminaryCheck( entry );
this.makePolicyAssignment( policyDn, entry );
}
} finally
{
try
{
this.ldap.close();
} catch ( final LdapException ex )
{
final String msg = "!!! The close connection raised an error !!!";
PasswordPolicyFacilitator.LOG.error( msg );
throw new PasswordPolicyFacilitatorException( msg, ex );
}
}
}
}
| 45f7a978709acd582622f0b293c945c571859ae6 | [
"Java",
"SQL"
] | 36 | Java | michele-deluca/ita.deluca.tools.ldap.framework | 1be36b5bdbf5e2a2096191c706a9388537e73dcc | ca4ff12a7e07714c7809a0179d6e6537e2eb24e4 |
refs/heads/master | <repo_name>pefarrell/firedrake<file_sep>/tests/regression/test_function_spaces.py
import pytest
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module')
def mesh2():
return UnitSquareMesh(1, 1)
@pytest.fixture(scope='module', params=['cg1cg1', 'cg1vcg1', 'cg1dg0', 'cg2dg1'])
def fs(request, cg1cg1, cg1vcg1, cg1dg0, cg2dg1):
return {'cg1cg1': cg1cg1,
'cg1vcg1': cg1vcg1,
'cg1dg0': cg1dg0,
'cg2dg1': cg2dg1}[request.param]
def test_function_space_cached(mesh):
"FunctionSpaces defined on the same mesh and element are cached."
assert FunctionSpace(mesh, "CG", 1) is FunctionSpace(mesh, "CG", 1)
def test_function_space_different_mesh_differ(mesh, mesh2):
"FunctionSpaces defined on different meshes differ."
assert FunctionSpace(mesh, "CG", 1) is not FunctionSpace(mesh2, "CG", 1)
def test_function_space_different_degree_differ(mesh):
"FunctionSpaces defined with different degrees differ."
assert FunctionSpace(mesh, "CG", 1) is not FunctionSpace(mesh, "CG", 2)
def test_function_space_different_family_differ(mesh):
"FunctionSpaces defined with different element families differ."
assert FunctionSpace(mesh, "CG", 1) is not FunctionSpace(mesh, "DG", 1)
def test_function_space_vector_function_space_differ(mesh):
"""A FunctionSpace and a VectorFunctionSpace defined with the same
family and degree differ."""
assert FunctionSpace(mesh, "CG", 1) is not VectorFunctionSpace(mesh, "CG", 1)
def test_indexed_function_space_index(fs):
assert [s.index for s in fs] == range(2)
# Create another mixed space in reverse order
fs0, fs1 = fs.split()
assert [s.index for s in (fs1 * fs0)] == range(2)
# Verify the indices of the original IndexedFunctionSpaces haven't changed
assert fs0.index == 0 and fs1.index == 1
def test_mixed_function_space_split(fs):
assert fs.split() == list(fs)
<file_sep>/tests/extrusion/test_extruded_1d_cohomology.py
"""
This demo verifies that the various FEEC operators can reproduce the
Betti numbers of the extruded interval, both periodic and non-periodic.
It also verifies that the various FEEC operators with strong Dirichlet
boundary conditions can reproduce the Betti numbers of the extruded
interval, obtained from Poincare duality, which says that the
dimension of the kth cohomology group with Dirichlet boundary
conditions is equal to the dimension of the (n-k)th cohomology group
without boundary conditions.
"""
import numpy.linalg as linalg
import numpy
from firedrake import *
import pytest
from tests.common import *
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti0(horiz_complex, vert_complex):
"""
Verify that the 0-form Hodge Laplacian has kernel of dimension
equal to the 0th Betti number of the extruded mesh, i.e. 1. Also
verify that the 0-form Hodge Laplacian with Dirichlet boundary
conditions has kernel of dimension equal to the 2nd Betti number
of the extruded mesh, i.e. 0.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = UnitIntervalMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
W0_elt = OuterProductElement(U0, V0)
W0 = FunctionSpace(mesh, W0_elt)
u = TrialFunction(W0)
v = TestFunction(W0)
L = assemble(inner(grad(u), grad(v))*dx)
uvecs, s, vvecs = linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
bcs = [DirichletBC(W0, 0., x) for x in ["top", "bottom", 1, 2]]
L = assemble(inner(grad(u), grad(v))*dx, bcs=bcs)
uvecs, s, vvecs = linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti0_periodic(horiz_complex, vert_complex):
"""
Verify that the 0-form Hodge Laplacian has kernel of dimension
equal to the 0th Betti number of the periodic extruded interval,
i.e. 1. Also verify that the 0-form Hodge Laplacian with
Dirichlet boundary conditions has kernel of dimension equal to the
2nd Betti number of the extruded mesh, i.e. 0.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = PeriodicUnitIntervalMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
W0_elt = OuterProductElement(U0, V0)
W0 = FunctionSpace(mesh, W0_elt)
u = TrialFunction(W0)
v = TestFunction(W0)
L = assemble(inner(grad(u), grad(v))*dx)
uvecs, s, vvecs = linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
bcs = [DirichletBC(W0, 0., x) for x in ["top", "bottom"]]
L = assemble(inner(grad(u), grad(v))*dx, bcs=bcs)
uvecs, s, vvecs = linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti1(horiz_complex, vert_complex):
"""
Verify that the 1-form Hodge Laplacian has kernel of dimension
equal to the 1st Betti number of the extruded mesh, i.e. 0. Also
verify that the 1-form Hodge Laplacian with Dirichlet boundary
conditions has kernel of dimension equal to the 2nd Betti number
of the extruded mesh, i.e. 0.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = UnitIntervalMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
U1 = FiniteElement(U1[0], "interval", U1[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W0_elt = OuterProductElement(U0, V0)
W1_a = HDiv(OuterProductElement(U1, V0))
W1_b = HDiv(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W0 = FunctionSpace(mesh, W0_elt)
W1 = FunctionSpace(mesh, W1_elt)
W = W0*W1
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((sigma*tau - inner(rot(tau), u) + inner(rot(sigma), v) +
div(u)*div(v))*dx)
dW0 = W0.dof_count
dW1 = W1.dof_count
A = numpy.zeros((dW0+dW1, dW0+dW1))
A[:dW0, :dW0] = L.M[0, 0].values
A[:dW0, dW0:dW0+dW1] = L.M[0, 1].values
A[dW0:dW0+dW1, :dW0] = L.M[1, 0].values
A[dW0:dW0+dW1, dW0:dW0+dW1] = L.M[1, 1].values
uvecs, s, vvecs = linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
bc0 = [DirichletBC(W.sub(0), 0., x) for x in [1, 2, "top", "bottom"]]
bc1 = [DirichletBC(W.sub(1), Expression(("0.", "0.")), x)
for x in [1, 2, "top", "bottom"]]
L0 = assemble((sigma*tau - inner(rot(tau), u) + inner(rot(sigma), v) +
div(u)*div(v))*dx, bcs=(bc0 + bc1))
A0 = numpy.zeros((dW0+dW1, dW0+dW1))
A0[:dW0, :dW0] = L0.M[0, 0].values
A0[:dW0, dW0:dW0+dW1] = L0.M[0, 1].values
A0[dW0:dW0+dW1, :dW0] = L0.M[1, 0].values
A0[dW0:dW0+dW1, dW0:dW0+dW1] = L0.M[1, 1].values
u, s, v = linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti1_periodic(horiz_complex, vert_complex):
"""
Verify that the 1-form Hodge Laplacian has kernel of dimension
equal to the 1st Betti number of the periodic extruded interval,
i.e. 1. Also verify that the 1-form Hodge Laplacian with
Dirichlet boundary conditions has kernel of dimension equal to the
2nd Betti number of the periodic extruded interval mesh, i.e. 1.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = PeriodicUnitIntervalMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
U1 = FiniteElement(U1[0], "interval", U1[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W0_elt = OuterProductElement(U0, V0)
W1_a = HDiv(OuterProductElement(U1, V0))
W1_b = HDiv(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W0 = FunctionSpace(mesh, W0_elt)
W1 = FunctionSpace(mesh, W1_elt)
W = W0*W1
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((sigma*tau - inner(rot(tau), u) + inner(rot(sigma), v) +
div(u)*div(v))*dx)
dW0 = W0.dof_count
dW1 = W1.dof_count
A = numpy.zeros((dW0+dW1, dW0+dW1))
A[:dW0, :dW0] = L.M[0, 0].values
A[:dW0, dW0:dW0+dW1] = L.M[0, 1].values
A[dW0:dW0+dW1, :dW0] = L.M[1, 0].values
A[dW0:dW0+dW1, dW0:dW0+dW1] = L.M[1, 1].values
uvecs, s, vvecs = linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
bc0 = [DirichletBC(W.sub(0), 0., x) for x in ["top", "bottom"]]
bc1 = [DirichletBC(W.sub(1), Expression(("0.", "0.")), x)
for x in ["top", "bottom"]]
L0 = assemble((sigma*tau - inner(rot(tau), u) + inner(rot(sigma), v) +
div(u)*div(v))*dx, bcs=(bc0 + bc1))
A0 = numpy.zeros((dW0+dW1, dW0+dW1))
A0[:dW0, :dW0] = L0.M[0, 0].values
A0[:dW0, dW0:dW0+dW1] = L0.M[0, 1].values
A0[dW0:dW0+dW1, :dW0] = L0.M[1, 0].values
A0[dW0:dW0+dW1, dW0:dW0+dW1] = L0.M[1, 1].values
u, s, v = linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti2(horiz_complex, vert_complex):
"""
Verify that the 2-form Hodge Laplacian has kernel of dimension
equal to the 2nd Betti number of the extruded mesh, i.e. 0. Also
verify that the 2-form Hodge Laplacian with Dirichlet boundary
conditions has kernel of dimension equal to the 0th Betti number
of the extruded mesh, i.e. 1.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = UnitIntervalMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
U1 = FiniteElement(U1[0], "interval", U1[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W1_a = HDiv(OuterProductElement(U1, V0))
W1_b = HDiv(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W2_elt = OuterProductElement(U1, V1)
W1 = FunctionSpace(mesh, W1_elt)
W2 = FunctionSpace(mesh, W2_elt)
W = W1*W2
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx)
bc1 = [DirichletBC(W.sub(0), Expression(("0.", "0.")), x)
for x in [1, 2, "top", "bottom"]]
L0 = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx, bcs=bc1)
dW1 = W1.dof_count
dW2 = W2.dof_count
A = numpy.zeros((dW1+dW2, dW1+dW2))
A[:dW1, :dW1] = L.M[0, 0].values
A[:dW1, dW1:dW1+dW2] = L.M[0, 1].values
A[dW1:dW1+dW2, :dW1] = L.M[1, 0].values
A[dW1:dW1+dW2, dW1:dW1+dW2] = L.M[1, 1].values
u, s, v = linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
A0 = numpy.zeros((dW1+dW2, dW1+dW2))
A0[:dW1, :dW1] = L0.M[0, 0].values
A0[:dW1, dW1:dW1+dW2] = L0.M[0, 1].values
A0[dW1:dW1+dW2, :dW1] = L0.M[1, 0].values
A0[dW1:dW1+dW2, dW1:dW1+dW2] = L0.M[1, 1].values
u, s, v = linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti2_periodic(horiz_complex, vert_complex):
"""
Verify that the 2-form Hodge Laplacian has kernel of dimension
equal to the 2nd Betti number of the periodic extruded interval
mesh, i.e. 0. Also verify that the 2-form Hodge Laplacian with
Dirichlet boundary conditions has kernel of dimension equal to the
0th Betti number of the periodic extruded interval mesh, i.e. 1.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = PeriodicUnitIntervalMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
U1 = FiniteElement(U1[0], "interval", U1[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W1_a = HDiv(OuterProductElement(U1, V0))
W1_b = HDiv(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W2_elt = OuterProductElement(U1, V1)
W1 = FunctionSpace(mesh, W1_elt)
W2 = FunctionSpace(mesh, W2_elt)
W = W1*W2
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx)
bc1 = [DirichletBC(W.sub(0), Expression(("0.", "0.")), x)
for x in ["top", "bottom"]]
L0 = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx, bcs=bc1)
dW1 = W1.dof_count
dW2 = W2.dof_count
A = numpy.zeros((dW1+dW2, dW1+dW2))
A[:dW1, :dW1] = L.M[0, 0].values
A[:dW1, dW1:dW1+dW2] = L.M[0, 1].values
A[dW1:dW1+dW2, :dW1] = L.M[1, 0].values
A[dW1:dW1+dW2, dW1:dW1+dW2] = L.M[1, 1].values
u, s, v = linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
A0 = numpy.zeros((dW1+dW2, dW1+dW2))
A0[:dW1, :dW1] = L0.M[0, 0].values
A0[:dW1, dW1:dW1+dW2] = L0.M[0, 1].values
A0[dW1:dW1+dW2, :dW1] = L0.M[1, 0].values
A0[dW1:dW1+dW2, dW1:dW1+dW2] = L0.M[1, 1].values
u, s, v = linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_poisson_sphere.py
import pytest
from tests.common import longtest
from firedrake import *
import numpy as np
def run_hdiv_l2(refinement, hdiv_space, degree):
mesh = UnitIcosahedralSphereMesh(refinement_level=refinement)
mesh.init_cell_orientations(Expression(('x[0]', 'x[1]', 'x[2]')))
Ve = FunctionSpace(mesh, "DG", max(3, degree + 1))
V = FunctionSpace(mesh, hdiv_space, degree)
Q = FunctionSpace(mesh, "DG", degree - 1)
W = V*Q
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
g = Function(Q).interpolate(Expression("x[0]*x[1]*x[2]"))
u_exact = Function(Ve, name="exact").interpolate(Expression("-x[0]*x[1]*x[2]/12.0"))
a = (inner(sigma, tau) + div(sigma)*v + div(tau)*u)*dx
L = g*v*dx
w = Function(W)
nullspace = MixedVectorSpaceBasis(W, [W[0], VectorSpaceBasis(constant=True)])
solve(a == L, w, nullspace=nullspace, solver_parameters={'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'fieldsplit_0_pc_type': 'lu',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_1_pc_factor_shift_type': 'INBLOCKS',
'fieldsplit_0_ksp_max_it': 100})
sigma, u = w.split()
L2_error_u = errornorm(u_exact, u, degree_rise=1)
h = 1.0/(2**refinement * sin(2*pi/5))
return L2_error_u, h, assemble(u*dx)
@longtest
@pytest.mark.parametrize(('hdiv_space', 'degree', 'conv_order'),
[('RT', 1, 0.75),
('BDM', 1, 0.8)])
def test_hdiv_l2(hdiv_space, degree, conv_order):
errors = [run_hdiv_l2(r, hdiv_space, degree) for r in range(1, 4)]
errors = np.asarray(errors)
l2err = errors[:, 0]
l2conv = np.log2(l2err[:-1] / l2err[1:])
assert (l2conv > conv_order).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_steady_advection_2D.py
"""This demo program solves the steady-state advection equation
div(u0*D) = 0, for a prescribed velocity field u0. An upwind
method is used, which stress-tests both interior and exterior
facet integrals.
"""
import pytest
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module')
def DG0(mesh):
return FunctionSpace(mesh, "DG", 0)
@pytest.fixture(scope='module')
def DG1(mesh):
return FunctionSpace(mesh, "DG", 1)
@pytest.fixture(scope='module')
def W(mesh):
return FunctionSpace(mesh, "BDM", 1)
def run_left_to_right(mesh, DG0, W):
velocity = Expression(("1.0", "0.0"))
u0 = project(velocity, W)
inflowexpr = Expression("x[1] > 0.25 && x[1] < 0.75 ? 1.0 : 0.5")
inflow = Function(DG0)
inflow.interpolate(inflowexpr)
n = FacetNormal(mesh)
un = 0.5*(dot(u0, n) + abs(dot(u0, n)))
D = TrialFunction(DG0)
phi = TestFunction(DG0)
a1 = -D*dot(u0, grad(phi))*dx
a2 = jump(phi)*(un('+')*D('+') - un('-')*D('-'))*dS
a3 = phi*un*D*ds(2) # outflow at right-hand wall
a = a1 + a2 + a3
L = -inflow*phi*dot(u0, n)*ds(1) # inflow at left-hand wall
out = Function(DG0)
solve(a == L, out)
# we only use inflow at the left wall, but since the velocity field
# is parallel to the coordinate axis, the exact solution matches
# the inflow function
assert max(abs(out.dat.data - inflow.dat.data)) < 1.2e-7
def test_left_to_right(mesh, DG0, W):
run_left_to_right(mesh, DG0, W)
@pytest.mark.parallel
def test_left_to_right_parallel():
m = mesh()
dg0 = DG0(m)
w = W(m)
run_left_to_right(m, dg0, w)
def run_up_to_down(mesh, DG1, W):
velocity = Expression(("0.0", "-1.0"))
u0 = project(velocity, W)
inflowexpr = Expression("1.0 + x[0]")
inflow = Function(DG1)
inflow.interpolate(inflowexpr)
n = FacetNormal(mesh)
un = 0.5*(dot(u0, n) + abs(dot(u0, n)))
D = TrialFunction(DG1)
phi = TestFunction(DG1)
a1 = -D*dot(u0, grad(phi))*dx
a2 = jump(phi)*(un('+')*D('+') - un('-')*D('-'))*dS
a3 = phi*un*D*ds(3) # outflow at lower wall
a = a1 + a2 + a3
L = -inflow*phi*dot(u0, n)*ds(4) # inflow at upper wall
out = Function(DG1)
solve(a == L, out)
assert max(abs(out.dat.data - inflow.dat.data)) < 1.1e-6
def test_up_to_down(mesh, DG1, W):
run_up_to_down(mesh, DG1, W)
@pytest.mark.parallel
def test_up_to_down_parallel():
m = mesh()
dg1 = DG1(m)
w = W(m)
run_up_to_down(m, dg1, w)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/evtk/README.txt
INTRODUCTION:
=============
EVTK (Export VTK) package allows exporting data to binary VTK files for
visualization and data analysis with any of the visualization packages that
support VTK files, e.g. Paraview, VisIt and Mayavi. EVTK does not depend on any
external library (e.g. VTK), so it is easy to install in different systems.
The package is composed of a set of Python files and a small C/Cython library
that provides performance critical routines. EVTK provides low and high level
interfaces. While the low level interface can be used to export data that is
stored in any type of container, the high level functions make easy to export
data stored in Numpy arrays.
INSTALLATION:
=============
Go to the source directory and type:
python setup.py install
DOCUMENTATION:
==============
This file together with the included examples in the examples directory in the
source tree provide enough information to start using the package.
DESIGN GUIDELINES:
==================
The design of the package considered the following objectives:
1. Self-contained. The package does not require any external library with
the exception of Numpy, which is becoming a standard package in many Python
installations.
2. Flexibility. It is possible to use EVTK to export data stored in any
container and in any of the grid formats supported by VTK by using the low level
interface.
3. Easy of use. The high level interface makes very easy to export data stored
in Numpy arrays. The high level interface provides functions to export most of
the grids supported by VTK: image data, rectilinear and structured grids. It
also includes a function to export point sets and associated data that can be
used to export results from particle and meshless numerical simulations.
4. Performance. The aim of the package is to be used as a part of
post-processing tools. Thus, good performance is important to handle the results
of large simulations. To achieve this goal, performance critical routines are
implemented as part of a small C extension.
REQUIREMENTS:
=============
- Numpy. Tested with Numpy 1.5.0.
- Cython 0.12. Cython is only required to update the included C file but
not to compile the package.
The package has been tested on:
- MacOSX 10.6 x86-64.
- Ubuntu 10.04 x86-64 guest running on VMWare Fusion.
DEVELOPER NOTES:
================
It is useful to build and install the package to a temporary location without
touching the global python site-packages directory while developing. To do
this, while in the root directory, one can type:
1. python setup.py build --debug install --prefix=./tmp
2. export PYTHONPATH=./tmp/lib/python2.6/site-packages/:$PYTHONPATH
NOTE: you may have to change the Python version depending of the installed
version on your system.
To test the package one can run some of the examples, e.g.:
./tmp/lib/python2.6/site-packages/examples/points.py
That should create a points.vtu file in the current directory.
<file_sep>/tests/conftest.py
"""Global test configuration."""
# Insert the parent directory into the module path so we can find the common
# module whichever directory we are calling py.test from.
#
# Note that this will ONLY work when tests are run by calling py.test, not when
# calling them as a module. In that case it is required to have the Firedrake
# root directory on your PYTYHONPATH to be able to call tests from anywhere.
import os
import sys
sys.path.insert(0, os.path.abspath(os.path.join(os.path.dirname(__file__), os.pardir)))
from subprocess import check_call
from sys import executable
from functools import wraps
from inspect import getsourcefile
from mpi4py import MPI
def parallel(nprocs=3):
"""Run a test in parallel
:arg nprocs: The number of processes to run.
.. note ::
Parallel tests need to either be in the same folder as the utils
module or the test folder needs to be on the PYTHONPATH."""
def _parallel_test(fn):
@wraps(fn)
def wrapper(*args, **kwargs):
if MPI.COMM_WORLD.size > 1:
fn(*args, **kwargs)
else:
check_call(' '.join(['mpiexec', '-n', '%d' % nprocs, executable,
'-c', '"import %s; %s.%s()"' %
(fn.__module__, fn.__module__, fn.__name__)]),
cwd=os.path.dirname(getsourcefile(fn)), shell=True)
return wrapper
return _parallel_test
def pytest_addoption(parser):
parser.addoption("--short", action="store_true", default=False,
help="Skip long tests")
def pytest_configure(config):
"""Register an additional marker."""
config.addinivalue_line(
"markers",
"parallel(nprocs): mark test to run in parallel on nprocs processors")
def pytest_runtest_setup(item):
run_parallel = item.keywords.get("parallel", None)
if run_parallel:
item._obj = parallel(run_parallel.kwargs.get('nprocs', 3))(item._obj)
def pytest_cmdline_preparse(config, args):
if 'PYTEST_VERBOSE' in os.environ and '-v' not in args:
args.insert(0, '-v')
if 'PYTEST_EXITFIRST' in os.environ and '-x' not in args:
args.insert(0, '-x')
if 'PYTEST_NOCAPTURE' in os.environ and '-s' not in args:
args.insert(0, '-s')
if 'PYTEST_TBNATIVE' in os.environ:
args.insert(0, '--tb=native')
if 'PYTEST_WATCH' in os.environ and '-f' not in args:
args.insert(0, '-f')
<file_sep>/tests/regression/test_zero_forms.py
import pytest
import numpy as np
import itertools
from firedrake import *
@pytest.fixture(scope='module')
def mesh():
return UnitSquareMesh(10, 10)
@pytest.fixture(scope='module')
def one(mesh):
return Constant(1, domain=mesh.ufl_domain())
domains = [(1, 2),
(2, 3),
(3, 4),
(4, 1),
(1, 2, 3, 4)]
def test_ds_dx(one):
assert np.allclose(assemble(one*dx + one*ds), 5.0)
@pytest.mark.parametrize('domains', domains)
def test_dsn(one, domains):
assert np.allclose(assemble(one*ds(domains)), len(domains))
form = one*ds(domains[0])
for d in domains[1:]:
form += one*ds(d)
assert np.allclose(assemble(form), len(domains))
@pytest.mark.parallel
def test_dsn_parallel():
c = one(mesh())
for d in domains:
assert np.allclose(assemble(c*ds(d)), len(d))
for domain in domains:
form = c*ds(domain[0])
for d in domain[1:]:
form += c*ds(d)
assert np.allclose(assemble(form), len(domain))
@pytest.mark.parametrize(['expr', 'value'],
itertools.product(['f',
'2*f',
'tanh(f)',
'2 * tanh(f)',
'f + tanh(f)',
'cos(f) + sin(f)',
'cos(f)*cos(f) + sin(f)*sin(f)',
'tanh(f) + cos(f) + sin(f)'],
[1, 10, 20, -1, -10, -20]))
def test_math_functions(mesh, expr, value):
V = FunctionSpace(mesh, 'CG', 1)
f = Function(V)
f.assign(value)
actual = assemble(eval(expr)*dx)
from math import *
f = value
expect = eval(expr)
assert np.allclose(actual, expect)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_ufl.py
import pytest
from firedrake import *
@pytest.mark.parametrize('n', [1, 3, 16])
def test_cellsize_1d(n):
assert abs(assemble(CellSize(UnitIntervalMesh(n))*dx) - 1.0/n) < 1e-14
@pytest.mark.parametrize('n', [1, 3, 16])
def test_cellsize_2d(n):
assert abs(assemble(CellSize(UnitSquareMesh(n, n))*dx) - sqrt(2)/n) < 1e-14
@pytest.mark.parametrize('n', [1, 3, 16])
def test_cellsize_3d(n):
assert abs(assemble(CellSize(UnitCubeMesh(n, n, n))*dx) - sqrt(3)/n) < 5e-14
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/expression.py
import numpy as np
import ufl
from pyop2 import op2
import utils
__all__ = ['Expression']
class Expression(ufl.Coefficient):
"""A code snippet or Python function that may be evaluated on a
:class:`.FunctionSpace`. This provides a mechanism for setting
:class:`.Function` values to user-determined values.
To use an Expression, we can either :meth:`~Function.interpolate`
it onto a :class:`.Function`, or :func:`.project` it into a
:class:`.FunctionSpace`. Note that not all
:class:`.FunctionSpace`\s support interpolation, but all do
support projection.
:class:`Expression`\s may be provided as snippets of C code, which
results in fast execution but offers limited functionality to the
user, or as a Python function, which is more flexible but slower,
since a Python function is called for every cell in the mesh.
**The C interface**
The code in an :class:`Expression` has access to the coordinates
in the variable ``x``, with ``x[0]`` corresponding to the x
component, ``x[1]`` to the y component and so forth. You can use
mathematical functions from the C library, along with the variable
``pi`` for :math:`\\pi`.
For example, to build an expression corresponding to
.. math::
\\sin(\\pi x)\\sin(\\pi y)\\sin(\\pi z)
we use:
.. code-block:: python
expr = Expression('sin(pi*x[0])*sin(pi*x[1])*sin(pi*x[2])')
If the :class:`.FunctionSpace` the expression will be applied to is
vector valued, a list of code snippets of length matching the
number of components in the function space must be provided.
**The Python interface**
The Python interface is accessed by creating a subclass of
:class:`Expression` with a user-specified :meth:`eval` method. For
example, the following expression sets the output
:class:`.Function` to the square of the magnitude of the
coordinate:
.. code-block:: python
class MyExpression(Expression):
def eval(self, value, X):
value[:] = numpy.dot(X, X)
Observe that the (single) entry of the ``value`` parameter is written to,
not the parameter itself.
This :class:`Expression` could be interpolated onto the
:class:`.Function` ``f`` by executing:
.. code-block:: python
f.interpolate(MyExpression())
Note the brackets required to instantiate the :class:`MyExpression` object.
If a Python :class:`Expression` is to set the value of a
vector-valued :class:`Function` then it is necessary to explicitly
override the :meth:`value_shape` method of that
:class:`Expression`. For example:
.. code-block:: python
class MyExpression(Expression):
def eval(self, value, X):
value[:] = X
def value_shape(self):
return (2,)
"""
def __init__(self, code=None, element=None, cell=None, degree=None, **kwargs):
"""
:param code: a string C statement, or list of statements.
:param element: a :class:`~ufl.finiteelement.finiteelement.FiniteElement`, optional
(currently ignored)
:param cell: a :class:`~ufl.geometry.Cell`, optional (currently ignored)
:param degree: the degree of quadrature to use for evaluation (currently ignored)
:param kwargs: user-defined values that are accessible in the
Expression code. These values maybe updated by
accessing the property of the same name. This can be
used, for example, to pass in the current timestep to
an Expression without necessitating recompilation. For
example:
.. code-block:: python
f = Function(V)
e = Expression('sin(x[0]*t)', t=t)
while t < T:
f.interpolate(e)
...
t += dt
e.t = t
The currently ignored parameters are retained for API compatibility with Dolfin.
"""
# Init also called in mesh constructor, but expression can be built without mesh
utils._init()
self.code = code
self._shape = ()
if code is not None:
shape = np.array(code).shape
self._shape = shape
if self.rank() == 0:
# Make code slot iterable even for scalar expressions
self.code = [code]
self.cell = cell
self.degree = degree
# These attributes are required by ufl.Coefficient to render the repr
# of an Expression. Since we don't call the ufl.Coefficient constructor
# (since we don't yet know the element) we need to set them ourselves
self._element = element
self._repr = None
self._count = 0
self._user_args = []
# Changing counter used to record when user changes values
self._state = 0
# Save the kwargs so that when we rebuild an expression we can
# reconstruct the user arguments.
self._kwargs = {}
if len(kwargs) == 0:
# No need for magic, since there are no user arguments.
return
# We have to build a new class to add these properties to
# since properties work on classes not instances and we don't
# want every Expression to have all the properties of all
# Expressions.
cls = type(self.__class__.__name__, (self.__class__, ), {})
for slot, val in kwargs.iteritems():
# Save the argument for later reconstruction
self._kwargs[slot] = val
# Scalar arguments have to be treated specially
val = np.array(val, dtype=np.float64)
shape = val.shape
rank = len(shape)
if rank == 0:
shape = 1
val = op2.Global(shape, val, dtype=np.float64, name=slot)
# Record the Globals in a known order (for later passing
# to a par_loop). Remember their "name" too, so we can
# construct a kwarg dict when applying python expressions.
self._user_args.append((slot, val))
# And save them as an attribute
setattr(self, '_%s' % slot, val)
# We have to do this because of the worthlessness of
# Python's support for closing over variables.
def make_getx(slot):
def getx(self):
glob = getattr(self, '_%s' % slot)
return glob.data_ro
return getx
def make_setx(slot):
def setx(self, value):
glob = getattr(self, '_%s' % slot)
glob.data = value
self._kwargs[slot] = value
# Bump state
self._state += 1
return setx
# Add public properties for the user-defined variables
prop = property(make_getx(slot), make_setx(slot))
setattr(cls, slot, prop)
# Set the class on this instance to the newly built class with
# properties attached.
self.__class__ = cls
def rank(self):
"""Return the rank of this :class:`Expression`"""
return len(self.value_shape())
def value_shape(self):
"""Return the shape of this :class:`Expression`.
This is the number of values the code snippet in the
expression contains.
"""
return self._shape
def to_expression(val, **kwargs):
"""Convert val to an :class:`Expression`.
:arg val: an iterable of values suitable for a code snippet in an
:class:`Expression`.
:arg \*\*kwargs: keyword arguments passed to the
:class:`Expression` constructor (which see).
"""
try:
expr = ["%s" % v for v in val]
except TypeError:
# Not iterable
expr = "%s" % val
return Expression(code=expr, **kwargs)
<file_sep>/tests/regression/test_assembly_cache.py
import pytest
import numpy as np
from firedrake import *
@pytest.fixture(scope='module')
def mesh():
return UnitSquareMesh(5, 5)
@pytest.fixture(scope='module')
def cg1(mesh):
return FunctionSpace(mesh, "CG", 1)
def test_eviction(cg1):
cache = assembly_cache.AssemblyCache()
cache.clear()
old_limit = parameters["assembly_cache"]["max_bytes"]
try:
parameters["assembly_cache"]["max_bytes"] = 5000
u = TrialFunction(cg1)
v = TestFunction(cg1)
# The mass matrix should be 1648 bytes, so 3 of them fit in
# cache, and inserting a 4th will cause two to be evicted.
for i in range(1, 5):
# Scaling the mass matrix by i causes cache misses.
assemble(i*u*v*dx).M
finally:
parameters["assembly_cache"]["max_bytes"] = old_limit
assert 3000 < cache.nbytes < 5000
assert cache.num_objects == 2
@pytest.mark.parallel(nprocs=2)
def test_eviction_parallel():
cache = assembly_cache.AssemblyCache()
cache.clear()
mesh = UnitSquareMesh(5, 5)
cg1 = FunctionSpace(mesh, "Lagrange", 1)
old_limit = parameters["assembly_cache"]["max_bytes"]
try:
parameters["assembly_cache"]["max_bytes"] = 5000
u = TrialFunction(cg1)
v = TestFunction(cg1)
# In the parallel case it's harder to ascertain exactly how
# much cache we will use, so we do this enough times that we
# can prove that we must have triggered eviction.
for i in range(1, 15):
# Scaling the mass matrix by i causes cache misses.
assemble(i*u*v*dx).M
finally:
parameters["assembly_cache"]["max_bytes"] = old_limit
assert 3000 < cache.nbytes < 5000
def test_hit(cg1):
cache = assembly_cache.AssemblyCache()
cache.clear()
u = TrialFunction(cg1)
v = TestFunction(cg1)
assemble(u*v*dx).M
assemble(u*v*dx).M
assert cache.num_objects == 1
assert cache._hits == 1
def test_assemble_rhs_with_without_constant(cg1):
cache = assembly_cache.AssemblyCache()
cache.clear()
v = TestFunction(cg1)
f = Function(cg1)
f = assemble(v*dx, f)
f = assemble(Constant(2)*v*dx, f)
assert cache.num_objects == 2
assert np.allclose(f.dat.data_ro, 2 * assemble(v*dx).dat.data_ro)
assert cache.num_objects == 2
def test_repeated_assign(cg1):
cache = assembly_cache.AssemblyCache()
cache.clear()
u = Function(cg1)
g = Function(cg1)
f = Function(cg1)
assert np.allclose(assemble(g*g*dx), 0)
assert cache.num_objects == 1
f.assign(1)
u.assign(g)
u.assign(f)
g.assign(u)
assert cache.num_objects == 1
assert np.allclose(assemble(g*g*dx), 1.0)
assert cache.num_objects == 1
@pytest.mark.parallel
def test_repeated_project():
cache = assembly_cache.AssemblyCache()
cache.clear()
mesh = UnitCubeMesh(2, 2, 2)
V2 = FunctionSpace(mesh, "DG", 0)
D0 = project(Expression('x[0]'), V2)
assert cache.num_objects == 2
D1 = project(Expression('x[0]'), V2)
assert cache.num_objects == 2
assert np.allclose(assemble((D0 - D1)*(D0 - D1)*dx), 0)
assert cache.num_objects == 3
@pytest.mark.parallel
def test_repeated_mixed_solve():
cache = assembly_cache.AssemblyCache()
cache.clear()
n = 4
mesh = UnitSquareMesh(n, n)
V1 = FunctionSpace(mesh, 'RT', 1)
V2 = FunctionSpace(mesh, 'DG', 0)
W = V1 * V2
lmbda = 1
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
f = Function(V2)
f.interpolate(Expression('1e-7'))
a = (p*q - q*div(u) + lmbda*inner(v, u) + div(v)*p)*dx
L = f*q*dx
solver_parameters = {'ksp_type': 'cg',
'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_0_ksp_type': 'cg',
'fieldsplit_1_ksp_type': 'cg'}
solution1 = Function(W)
solve(a == L, solution1, solver_parameters=solver_parameters)
assert cache.num_objects == 2
solution2 = Function(W)
solve(a == L, solution2, solver_parameters=solver_parameters)
assert cache.num_objects == 2
assert np.allclose(errornorm(solution1, solution2, degree_rise=0), 0)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_function.py
import pytest
import numpy as np
from firedrake import *
@pytest.fixture
def V():
mesh = UnitIntervalMesh(2)
V = FunctionSpace(mesh, "CG", 1)
return V
def test_firedrake_function(V):
f = Function(V)
f.interpolate(Expression("1"))
assert (f.dat.data_ro == 1.0).all()
g = Function(f)
assert (g.dat.data_ro == 1.0).all()
# Check that g is indeed a deep copy
f.interpolate(Expression("2"))
assert (f.dat.data_ro == 2.0).all()
assert (g.dat.data_ro == 1.0).all()
def test_mismatching_rank_interpolation(V):
f = Function(V)
with pytest.raises(RuntimeError):
f.interpolate(Expression(('1', '2')))
VV = VectorFunctionSpace(V.mesh(), 'CG', 1)
f = Function(VV)
with pytest.raises(RuntimeError):
f.interpolate(Expression(('1', '2')))
def test_mismatching_shape_interpolation(V):
VV = VectorFunctionSpace(V.mesh(), 'CG', 1)
f = Function(VV)
with pytest.raises(RuntimeError):
f.interpolate(Expression(['1'] * (VV.ufl_element().value_shape()[0] + 1)))
def test_function_val(V):
"""Initialise a Function with a NumPy array."""
f = Function(V, np.ones((V.node_count, V.dim)))
assert (f.dat.data_ro == 1.0).all()
def test_function_dat(V):
"""Initialise a Function with an op2.Dat."""
f = Function(V, op2.Dat(V.node_set**V.dim))
f.interpolate(Expression("1"))
assert (f.dat.data_ro == 1.0).all()
def test_function_name(V):
f = Function(V, name="foo")
assert f.name() == "foo"
f.rename(label="bar")
assert f.label() == "bar" and f.name() == "foo"
f.rename(name="baz")
assert f.name() == "baz" and f.label() == "bar"
f.rename("foo", "quux")
assert f.name() == "foo" and f.label() == "quux"
f.rename(name="bar", label="baz")
assert f.name() == "bar" and f.label() == "baz"
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_assemble.py
import pytest
import numpy as np
from firedrake import *
from tests.common import *
# FIXME: cg1vcg1 is not supported yet
@pytest.fixture(scope='module', params=['cg1', 'vcg1',
'cg1cg1', 'cg1cg1[0]', 'cg1cg1[1]',
'cg1vcg1[0]', 'cg1vcg1[1]',
'cg1dg0', 'cg1dg0[0]', 'cg1dg0[1]',
'cg2dg1', 'cg2dg1[0]', 'cg2dg1[1]'])
def fs(request, cg1, vcg1, cg1cg1, cg1vcg1, cg1dg0, cg2dg1):
return {'cg1': cg1,
'vcg1': vcg1,
'cg1cg1': cg1cg1,
'cg1cg1[0]': cg1cg1[0],
'cg1cg1[1]': cg1cg1[1],
'cg1vcg1': cg1vcg1,
'cg1vcg1[0]': cg1vcg1[0],
'cg1vcg1[1]': cg1vcg1[1],
'cg1dg0': cg1dg0,
'cg1dg0[0]': cg1dg0[0],
'cg1dg0[1]': cg1dg0[1],
'cg2dg1': cg2dg1,
'cg2dg1[0]': cg2dg1[0],
'cg2dg1[1]': cg2dg1[1]}[request.param]
@pytest.fixture
def f(fs):
f = Function(fs, name="f")
if isinstance(fs, (MixedFunctionSpace, VectorFunctionSpace)):
f.interpolate(Expression(("x[0]",) * fs.cdim))
else:
f.interpolate(Expression("x[0]"))
return f
@pytest.fixture
def one(fs):
one = Function(fs, name="one")
if isinstance(fs, (MixedFunctionSpace, VectorFunctionSpace)):
one.interpolate(Expression(("1",) * fs.cdim))
else:
one.interpolate(Expression("1"))
return one
@pytest.fixture
def M(fs):
uhat = TrialFunction(fs)
v = TestFunction(fs)
return inner(uhat, v) * dx
def test_one_form(M, f):
one_form = assemble(action(M, f))
assert isinstance(one_form, Function)
for f in one_form.split():
assert abs(f.dat.data.sum() - 0.5*f.function_space().dim) < 1.0e-12
def test_zero_form(M, f, one):
zero_form = assemble(action(action(M, f), one))
assert isinstance(zero_form, float)
assert abs(zero_form - 0.5 * np.prod(f.shape())) < 1.0e-12
def test_assemble_with_tensor(cg1):
v = TestFunction(cg1)
L = v*dx
f = Function(cg1)
# Assemble a form into f
f = assemble(L, f)
# Assemble a different form into f
f = assemble(Constant(2)*L, f)
# Make sure we get the result of the last assembly
assert np.allclose(f.dat.data, 2*assemble(L).dat.data, rtol=1e-14)
def test_assemble_mat_with_tensor(dg0):
u = TestFunction(dg0)
v = TrialFunction(dg0)
a = u*v*dx
M = assemble(a)
# Assemble a different form into M
M = assemble(Constant(2)*a, M)
# Make sure we get the result of the last assembly
assert np.allclose(M.M.values, 2*assemble(a).M.values, rtol=1e-14)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_embedded_sphere.py
# Test that integrals over the surface of a sphere do the right thing
import pytest
import numpy as np
from tests.common import longtest
from firedrake import *
def integrate_sphere_area(radius=1, refinement=2):
mesh = IcosahedralSphereMesh(radius=radius, refinement_level=refinement)
fs = FunctionSpace(mesh, "CG", 1, name="fs")
f = Function(fs)
f.assign(1)
exact = 4*pi*radius**2
return np.abs(assemble(f * dx) - exact) / exact
@longtest
@pytest.mark.parametrize(('radius', 'refinement', 'error'),
[(1, 2, 0.02),
(10, 2, 0.02),
(1, 5, 0.0004),
(10, 5, 0.0004)])
def test_surface_area_sphere(radius, refinement, error):
assert integrate_sphere_area(radius=radius, refinement=refinement) < error
<file_sep>/tests/regression/test_2dcohomology.py
"""
This demo verifies that the various FEEC operators can reproduce the
Betti numbers of the 2D annulus.
It also verifies that the various FEEC operators with strong Dirichlet
boundary conditions can reproduce the Betti numbers of the 2D annulus,
obtained from Poincare duality, which says that the dimension of the
kth cohomology group with Dirichlet boundary conditions is equal to
the dimension of the (n-k)th cohomology group without boundary
conditions.
"""
from os.path import abspath, dirname, join
import numpy.linalg as linalg
import numpy
from firedrake import *
import pytest
from tests.common import *
cwd = abspath(dirname(__file__))
@pytest.mark.parametrize(('space'),
[(("CG", 1), ("RT", 1), ("DG", 0)),
(("CG", 2), ("RT", 2), ("DG", 1)),
(("CG", 3), ("RT", 3), ("DG", 2)),
(("CG", 2), ("BDM", 1), ("DG", 0)),
(("CG", 3), ("BDM", 2), ("DG", 1)),
(("CG", 4), ("BDM", 3), ("DG", 2)),
(("CG", 2, "B", 3), ("BDFM", 2), ("DG", 1))])
def test_betti0(space):
"""
Verify that the 0-form Hodge Laplacian with strong Dirichlet
boundary conditions has kernel of dimension equal to the 2nd Betti
number of the annulus mesh, i.e. 0.
"""
mesh = Mesh(join(cwd, "annulus.msh"))
V0tag, V1tag, V2tag = space
if(len(V0tag) == 2):
V0 = FunctionSpace(mesh, V0tag[0], V0tag[1])
else:
V0a = FiniteElement(V0tag[0], "triangle", V0tag[1])
V0b = FiniteElement(V0tag[2], "triangle", V0tag[3])
V0 = FunctionSpace(mesh, V0a + V0b)
#V0 Hodge Laplacian
u = TrialFunction(V0)
v = TestFunction(V0)
L = assemble(inner(nabla_grad(u), nabla_grad(v))*dx)
bc0 = DirichletBC(V0, 0., 9)
L0 = assemble(inner(nabla_grad(u), nabla_grad(v))*dx, bcs=[bc0])
u, s, v = linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
u, s, v = linalg.svd(L0.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
@pytest.mark.parametrize(('space'),
[(("CG", 1), ("RT", 1), ("DG", 0)),
(("CG", 2), ("RT", 2), ("DG", 1)),
(("CG", 3), ("RT", 3), ("DG", 2)),
(("CG", 2), ("BDM", 1), ("DG", 0)),
(("CG", 3), ("BDM", 2), ("DG", 1)),
(("CG", 4), ("BDM", 3), ("DG", 2)),
(("CG", 2, "B", 3), ("BDFM", 2), ("DG", 1))])
def test_betti1(space):
"""
Verify that the 1-form Hodge Laplacian with strong Dirichlet
boundary conditions has kernel of dimension equal to the 1st Betti
number of the annulus mesh, i.e. 1.
"""
mesh = Mesh(join(cwd, "annulus.msh"))
V0tag, V1tag, V2tag = space
if(len(V0tag) == 2):
V0 = FunctionSpace(mesh, V0tag[0], V0tag[1])
else:
V0a = FiniteElement(V0tag[0], "triangle", V0tag[1])
V0b = FiniteElement(V0tag[2], "triangle", V0tag[3])
V0 = FunctionSpace(mesh, V0a + V0b)
V1 = FunctionSpace(mesh, V1tag[0], V1tag[1])
W = V0*V1
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((sigma*tau - inner(rot(tau), u) + inner(rot(sigma), v) +
div(u)*div(v))*dx)
bc0 = DirichletBC(W.sub(0), 0., 9)
bc1 = DirichletBC(W.sub(1), Expression(("0.0", "0.0")), 9)
L0 = assemble((sigma*tau - inner(rot(tau), u) + inner(rot(sigma), v) +
div(u)*div(v))*dx, bcs=[bc0, bc1])
dV0 = V0.dof_count
dV1 = V1.dof_count
A = numpy.zeros((dV0+dV1, dV0+dV1))
A[:dV0, :dV0] = L.M[0, 0].values
A[:dV0, dV0:dV0+dV1] = L.M[0, 1].values
A[dV0:dV0+dV1, :dV0] = L.M[1, 0].values
A[dV0:dV0+dV1, dV0:dV0+dV1] = L.M[1, 1].values
u, s, v = linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
dV0 = V0.dof_count
dV1 = V1.dof_count
A0 = numpy.zeros((dV0+dV1, dV0+dV1))
A0[:dV0, :dV0] = L0.M[0, 0].values
A0[:dV0, dV0:dV0+dV1] = L0.M[0, 1].values
A0[dV0:dV0+dV1, :dV0] = L0.M[1, 0].values
A0[dV0:dV0+dV1, dV0:dV0+dV1] = L0.M[1, 1].values
u, s, v = linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
@pytest.mark.parametrize(('space'),
[(("CG", 1), ("RT", 1), ("DG", 0)),
(("CG", 2), ("RT", 2), ("DG", 1)),
(("CG", 3), ("RT", 3), ("DG", 2)),
(("CG", 2), ("BDM", 1), ("DG", 0)),
(("CG", 3), ("BDM", 2), ("DG", 1)),
(("CG", 4), ("BDM", 3), ("DG", 2)),
(("CG", 2, "B", 3), ("BDFM", 2), ("DG", 1))])
def test_betti2(space):
"""
Verify that the 2-form Hodge Laplacian with strong Dirichlet
boundary conditions has kernel of dimension equal to the 2nd Betti
number of the annulus mesh, i.e. 1.
"""
mesh = Mesh(join(cwd, "annulus.msh"))
V0tag, V1tag, V2tag = space
V1 = FunctionSpace(mesh, V1tag[0], V1tag[1])
V2 = FunctionSpace(mesh, V2tag[0], V2tag[1])
W = V1*V2
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx)
bc1 = DirichletBC(W.sub(0), Expression(("0.0", "0.0")), 9)
L0 = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx, bcs=[bc1])
dV1 = V1.dof_count
dV2 = V2.dof_count
A = numpy.zeros((dV1+dV2, dV1+dV2))
A[:dV1, :dV1] = L.M[0, 0].values
A[:dV1, dV1:dV1+dV2] = L.M[0, 1].values
A[dV1:dV1+dV2, :dV1] = L.M[1, 0].values
A[dV1:dV1+dV2, dV1:dV1+dV2] = L.M[1, 1].values
u, s, v = linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
print nharmonic, V1tag[0]
assert(nharmonic == 0)
A0 = numpy.zeros((dV1+dV2, dV1+dV2))
A0[:dV1, :dV1] = L0.M[0, 0].values
A0[:dV1, dV1:dV1+dV2] = L0.M[0, 1].values
A0[dV1:dV1+dV2, :dV1] = L0.M[1, 0].values
A0[dV1:dV1+dV2, dV1:dV1+dV2] = L0.M[1, 1].values
u, s, v = linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/wave_explicit.py
# NOTE This is a demo, not a regression test
from firedrake import *
output = True
mesh = UnitSquareMesh(100, 100)
# Plumb the space filling curve into UnitSquareMesh after the call to
# gmsh. Doru knows how to do this.
T = 10
# Note to Kaho: Ensure dt<dx for stability.
dt = 0.001
t = 0
fs = FunctionSpace(mesh, 'Lagrange', 1)
p = Function(fs)
phi = Function(fs)
u = TrialFunction(fs)
v = TestFunction(fs)
p.interpolate(Expression("exp(-40*((x[0]-.5)*(x[0]-.5)+(x[1]-.5)*(x[1]-.5)))"))
if output:
outfile = File("out.pvd")
phifile = File("phi.pvd")
outfile << p
phifile << phi
# Mass matrix
m = u * v * dx
lump_mass = False
step = 0
while t <= T:
step += 1
phi -= dt / 2 * p
if lump_mass:
p += (assemble(dt * inner(nabla_grad(v), nabla_grad(phi)) * dx)
/ assemble(v * dx))
else:
solve(u * v * dx == v * p * dx + dt * inner(
nabla_grad(v), nabla_grad(phi)) * dx, p)
phi -= dt / 2 * p
t += dt
if output:
print t
outfile << p
phifile << phi
<file_sep>/firedrake/solving.py
# Copyright (C) 2011 <NAME>
# Copyright (C) 2012 <NAME>, <NAME>
# Copyright (C) 2013 Imperial College London and others.
#
# This file is part of Firedrake, modified from the corresponding file in DOLFIN
#
# Firedrake is free software: you can redistribute it and/or modify
# it under the terms of the GNU Lesser General Public License as published by
# the Free Software Foundation, either version 3 of the License, or
# (at your option) any later version.
#
# Firedrake is distributed in the hope that it will be useful,
# but WITHOUT ANY WARRANTY; without even the implied warranty of
# MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# GNU Lesser General Public License for more details.
#
# You should have received a copy of the GNU Lesser General Public License
# along with DOLFIN. If not, see <http://www.gnu.org/licenses/>.
__all__ = ["solve"]
import ufl
from pyop2.logger import progress, INFO
from pyop2.profiling import profile
import linear_solver as ls
import variational_solver as vs
@profile
def solve(*args, **kwargs):
"""Solve linear system Ax = b or variational problem a == L or F == 0.
The Firedrake solve() function can be used to solve either linear
systems or variational problems. The following list explains the
various ways in which the solve() function can be used.
*1. Solving linear systems*
A linear system Ax = b may be solved by calling
.. code-block:: python
solve(A, x, b, bcs=bcs, solver_parameters={...})
where `A` is a :class:`.Matrix` and `x` and `b` are :class:`.Function`\s.
If present, `bcs` should be a list of :class:`.DirichletBC`\s
specifying the strong boundary conditions to apply. For the
format of `solver_parameters` see below.
*2. Solving linear variational problems*
A linear variational problem a(u, v) = L(v) for all v may be
solved by calling solve(a == L, u, ...), where a is a bilinear
form, L is a linear form, u is a :class:`.Function` (the
solution). Optional arguments may be supplied to specify boundary
conditions or solver parameters. Some examples are given below:
.. code-block:: python
solve(a == L, u)
solve(a == L, u, bcs=bc)
solve(a == L, u, bcs=[bc1, bc2])
solve(a == L, u, bcs=bcs,
solver_parameters={"ksp_type": "gmres"})
The linear solver uses PETSc under the hood and accepts all PETSc
options as solver parameters. For example, to solve the system
using direct factorisation use:
.. code-block:: python
solve(a == L, u, bcs=bcs,
solver_parameters={"ksp_type": "preonly", "pc_type": "lu"})
*3. Solving nonlinear variational problems*
A nonlinear variational problem F(u; v) = 0 for all v may be
solved by calling solve(F == 0, u, ...), where the residual F is a
linear form (linear in the test function v but possibly nonlinear
in the unknown u) and u is a :class:`.Function` (the
solution). Optional arguments may be supplied to specify boundary
conditions, the Jacobian form or solver parameters. If the
Jacobian is not supplied, it will be computed by automatic
differentiation of the residual form. Some examples are given
below:
The nonlinear solver uses a PETSc SNES object under the hood. To
pass options to it, use the same options names as you would for
pure PETSc code. See :class:`NonlinearVariationalSolver` for more
details.
.. code-block:: python
solve(F == 0, u)
solve(F == 0, u, bcs=bc)
solve(F == 0, u, bcs=[bc1, bc2])
solve(F == 0, u, bcs, J=J,
# Use Newton-Krylov iterations to solve the nonlinear
# system, using direct factorisation to solve the linear system.
solver_parameters={"snes_type": "newtonls",
"ksp_type" : "preonly",
"pc_type" : "lu"})
In all three cases, if the operator is singular you can pass a
:class:`.VectorSpaceBasis` (or :class:`.MixedVectorSpaceBasis`)
spanning the null space of the operator to the solve call using
the ``nullspace`` keyword argument.
"""
assert(len(args) > 0)
# Call variational problem solver if we get an equation
if isinstance(args[0], ufl.classes.Equation):
_solve_varproblem(*args, **kwargs)
else:
# Solve pre-assembled system
return _la_solve(*args, **kwargs)
def _solve_varproblem(*args, **kwargs):
"Solve variational problem a == L or F == 0"
# Extract arguments
eq, u, bcs, J, Jp, M, form_compiler_parameters, solver_parameters, nullspace \
= _extract_args(*args, **kwargs)
# Solve linear variational problem
if isinstance(eq.lhs, ufl.Form) and isinstance(eq.rhs, ufl.Form):
# Create problem
problem = vs.LinearVariationalProblem(eq.lhs, eq.rhs, u, bcs, Jp,
form_compiler_parameters=form_compiler_parameters)
# Create solver and call solve
solver = vs.LinearVariationalSolver(problem, solver_parameters=solver_parameters,
nullspace=nullspace)
with progress(INFO, 'Solving linear variational problem'):
solver.solve()
# Solve nonlinear variational problem
else:
# Create problem
problem = vs.NonlinearVariationalProblem(eq.lhs, u, bcs, J, Jp,
form_compiler_parameters=form_compiler_parameters)
# Create solver and call solve
solver = vs.NonlinearVariationalSolver(problem, solver_parameters=solver_parameters,
nullspace=nullspace)
with progress(INFO, 'Solving nonlinear variational problem'):
solver.solve()
# destroy snes part of solver so everything can be gc'd
solver.destroy()
def _la_solve(A, x, b, **kwargs):
"""Solve a linear algebra problem.
:arg A: the assembled bilinear form, a :class:`.Matrix`.
:arg x: the :class:`.Function` to write the solution into.
:arg b: the :class:`.Function` defining the right hand side values.
:kwarg bcs: an optional list of :class:`.DirichletBC`\s to apply.
:kwarg solver_parameters: optional solver parameters.
:kwarg nullspace: an optional :class:`.VectorSpaceBasis` (or
:class:`.MixedVectorSpaceBasis`) spanning the null space of
the operator.
.. note::
Any boundary conditions passed in as an argument here override the
boundary conditions set when the bilinear form was assembled.
That is, in the following example:
.. code-block:: python
A = assemble(a, bcs=[bc1])
solve(A, x, b, bcs=[bc2])
the boundary conditions in `bc2` will be applied to the problem
while `bc1` will be ignored.
Example usage:
.. code-block:: python
_la_solve(A, x, b, solver_parameters=parameters_dict)."""
bcs, solver_parameters, nullspace = _extract_linear_solver_args(A, x, b, **kwargs)
if bcs is not None:
A.bcs = bcs
solver = ls.LinearSolver(A, solver_parameters=solver_parameters,
nullspace=nullspace)
solver.solve(x, b)
def _extract_linear_solver_args(*args, **kwargs):
valid_kwargs = ["bcs", "solver_parameters", "nullspace"]
if len(args) != 3:
raise RuntimeError("Missing required arguments, expecting solve(A, x, b, **kwargs)")
for kwarg in kwargs.iterkeys():
if kwarg not in valid_kwargs:
raise RuntimeError("Illegal keyword argument '%s'; valid keywords are %s" %
(kwarg, ", ".join("'%s'" % kw for kw in valid_kwargs)))
bcs = kwargs.get("bcs", None)
solver_parameters = kwargs.get("solver_parameters", None)
nullspace = kwargs.get("nullspace", None)
return bcs, solver_parameters, nullspace
def _extract_args(*args, **kwargs):
"Extraction of arguments for _solve_varproblem"
# Check for use of valid kwargs
valid_kwargs = ["bcs", "J", "Jp", "M",
"form_compiler_parameters", "solver_parameters",
"nullspace"]
for kwarg in kwargs.iterkeys():
if not kwarg in valid_kwargs:
raise RuntimeError("Illegal keyword argument '%s'; valid keywords \
are %s" % (kwarg, ", ".join("'%s'" % kwarg
for kwarg in valid_kwargs)))
# Extract equation
if not len(args) >= 2:
raise RuntimeError("Missing arguments, expecting solve(lhs == rhs, u, \
bcs=bcs), where bcs is optional")
if len(args) > 3:
raise RuntimeError("Too many arguments, expecting solve(lhs == rhs, \
u, bcs=bcs), where bcs is optional")
# Extract equation
eq = _extract_eq(args[0])
# Extract solution function
u = _extract_u(args[1])
# Extract boundary conditions
bcs = _extract_bcs(args[2] if len(args) > 2 else kwargs.get("bcs"))
# Extract Jacobian
J = kwargs.get("J", None)
if J is not None and not isinstance(J, ufl.Form):
raise RuntimeError("Expecting Jacobian J to be a UFL Form")
Jp = kwargs.get("Jp", None)
if Jp is not None and not isinstance(Jp, ufl.Form):
raise RuntimeError("Expecting PC Jacobian Jp to be a UFL Form")
# Extract functional
M = kwargs.get("M", None)
if M is not None and not isinstance(M, ufl.Form):
raise RuntimeError("Expecting goal functional M to be a UFL Form")
nullspace = kwargs.get("nullspace", None)
# Extract parameters
form_compiler_parameters = kwargs.get("form_compiler_parameters", {})
solver_parameters = kwargs.get("solver_parameters", {})
return eq, u, bcs, J, Jp, M, form_compiler_parameters, solver_parameters, nullspace
def _extract_eq(eq):
"Extract and check argument eq"
if not isinstance(eq, ufl.classes.Equation):
raise RuntimeError("Expecting first argument to be an Equation")
return eq
def _extract_u(u):
"Extract and check argument u"
if not isinstance(u, ufl.Coefficient):
raise RuntimeError("Expecting second argument to be a Coefficient")
return u
def _extract_bcs(bcs):
"Extract and check argument bcs"
if bcs is None:
return []
try:
return tuple(bcs)
except TypeError:
return (bcs,)
<file_sep>/firedrake/solving_utils.py
from pyop2 import op2
from petsc import PETSc
def update_parameters(obj, petsc_obj):
"""Update parameters on a petsc object
:arg obj: An object with a parameters dict (mapping to petsc options).
:arg petsc_obj: The PETSc object to set parameters on."""
# Skip if parameters haven't changed
if hasattr(obj, '_set_parameters') and obj.parameters == obj._set_parameters:
return
opts = PETSc.Options(obj._opt_prefix)
for k, v in obj.parameters.iteritems():
if type(v) is bool:
if v:
opts[k] = None
else:
opts[k] = v
petsc_obj.setFromOptions()
obj._set_parameters = obj.parameters.copy()
def set_fieldsplits(mat, pc, names=None):
"""Set up fieldsplit splits
:arg mat: a :class:`~.Matrix` (the operator)
:arg pc: a PETSc PC to set splits on
:kwarg names: (optional) list of names for each split.
If not provided, splits are numbered from 0.
Returns a list of (name, IS) pairs (for later use with nullspace),
or None if no fieldsplit was set up."""
# No splits if not mixed
if mat.sparsity.shape == (1, 1):
return None
rows, cols = mat.sparsity.shape
ises = []
nlocal_rows = 0
for i in range(rows):
if i < cols:
nlocal_rows += mat[i, i].sparsity.nrows * mat[i, i].dims[0]
offset = 0
if op2.MPI.comm.rank == 0:
op2.MPI.comm.exscan(nlocal_rows)
else:
offset = op2.MPI.comm.exscan(nlocal_rows)
for i in range(rows):
if i < cols:
nrows = mat[i, i].sparsity.nrows * mat[i, i].dims[0]
name = names[i] if names is not None else str(i)
ises.append((name, PETSc.IS().createStride(nrows, first=offset, step=1)))
offset += nrows
pc.setFieldSplitIS(*ises)
return ises
<file_sep>/tests/extrusion/test_offset_computation.py
import pytest
from firedrake import *
def test_no_offset_zero():
m = UnitSquareMesh(1, 1)
m = ExtrudedMesh(m, layers=2)
V = FunctionSpace(m, 'CG', 2)
assert (V.exterior_facet_boundary_node_map("topological").offset != 0).all()
def test_offset_p2():
m = UnitSquareMesh(1, 1)
m = ExtrudedMesh(m, layers=1)
V = FunctionSpace(m, 'CG', 2)
assert (V.exterior_facet_boundary_node_map("topological").offset == 2).all()
def test_offset_enriched():
m = UnitSquareMesh(1, 1)
m = ExtrudedMesh(m, layers=1)
ele = OuterProductElement(FiniteElement("CG", "triangle", 2),
FiniteElement("CG", "interval", 1)) + \
OuterProductElement(FiniteElement("CG", "triangle", 1),
FiniteElement("DG", "interval", 0))
V = FunctionSpace(m, ele)
# On each facet we have:
#
# o--x--o
# | |
# o o
# | |
# o--x--o
#
# Where the numbering is such that the two "x" dofs are numbered last.
assert (V.exterior_facet_boundary_node_map("topological").offset ==
[2, 2, 2, 2, 2, 2, 1, 1]).all()
def run_offset_parallel():
m = UnitSquareMesh(20, 20)
m = ExtrudedMesh(m, layers=1)
V = FunctionSpace(m, 'CG', 2)
offset = V.exterior_facet_boundary_node_map("topological").offset
offsets = op2.MPI.comm.allgather(offset)
assert all((o == offset).all() for o in offsets)
@pytest.mark.parallel(nprocs=6)
def test_offset_parallel_indexerror():
run_offset_parallel()
@pytest.mark.parallel(nprocs=2)
def test_offset_parallel_allsame():
run_offset_parallel()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/linear_solver.py
import function
import solving_utils
from petsc import PETSc
__all__ = ["LinearSolver"]
class LinearSolver(object):
_id = 0
def __init__(self, A, P=None, solver_parameters=None,
nullspace=None):
"""A linear solver for assembled systems (Ax = b).
:arg A: a :class:`~.Matrix` (the operator).
:arg P: an optional :class:`~.Matrix` to construct any
preconditioner from; if none is supplied :data:`A` is
used to construct the preconditioner.
:kwarg parameters: (optional) dict of solver parameters.
:kwarg nullspace: an optional :class:`~.VectorSpaceBasis` (or
:class:`~.MixedVectorSpaceBasis` spanning the null space
of the operator.
.. note::
Any boundary conditions for this solve *must* have been
applied when assembling the operator.
"""
self.A = A
self.P = P if P is not None else A
parameters = solver_parameters.copy() if solver_parameters is not None else {}
parameters.setdefault("ksp_rtol", "1e-7")
if self.P._M.sparsity.shape != (1, 1):
parameters.setdefault('pc_type', 'jacobi')
self._opt_prefix = "firedrake_ksp_%d_" % LinearSolver._id
LinearSolver._id += 1
self.ksp = PETSc.KSP().create()
self.ksp.setOptionsPrefix(self._opt_prefix)
self.parameters = parameters
pc = self.ksp.getPC()
pmat = self.P._M
ises = solving_utils.set_fieldsplits(pmat, pc)
if nullspace is not None:
nullspace._apply(self.A.M, ises=ises)
if P is not None:
nullspace._apply(self.P.M, ises=ises)
# Operator setting must come after null space has been
# applied
# Force evaluation here
self.ksp.setOperators(A=self.A.M.handle, P=self.P.M.handle)
@property
def parameters(self):
return self._parameters
@parameters.setter
def parameters(self, val):
assert isinstance(val, dict), "Must pass a dict to set parameters"
self._parameters = val
solving_utils.update_parameters(self, self.ksp)
def __del__(self):
if hasattr(self, '_opt_prefix'):
opts = PETSc.Options()
for k in self.parameters.iterkeys():
del opts[self._opt_prefix + k]
delattr(self, '_opt_prefix')
def solve(self, x, b):
# User may have updated parameters
solving_utils.update_parameters(self, self.ksp)
if self.A.has_bcs:
b_bc = function.Function(b.function_space())
for bc in self.A.bcs:
bc.apply(b_bc)
# rhs = b - action(A, b_bc)
b_bc.assign(b - self.A._form_action(b_bc))
# Now we need to apply the boundary conditions to the "RHS"
for bc in self.A.bcs:
bc.apply(b_bc)
# don't want to write into b itself, because that would confuse user
b = b_bc
with b.dat.vec_ro as rhs:
with x.dat.vec as solution:
self.ksp.solve(rhs, solution)
r = self.ksp.getConvergedReason()
if r < 0:
reasons = self.ksp.ConvergedReason()
reasons = dict([(getattr(reasons, reason), reason)
for reason in dir(reasons) if not reason.startswith('_')])
raise RuntimeError("LinearSolver failed to converge after %d iterations with reason: %s", self.ksp.getIterationNumber(), reasons[r])
<file_sep>/tests/extrusion/test_extrusion_0_dg_coords.py
from firedrake import *
import pytest
import ufl
def test_extruded_interval_area():
m = UnitIntervalMesh(10)
DG = VectorFunctionSpace(m, 'DG', 1)
new_coords = project(m.coordinates, DG)
m._coordinate_fs = new_coords.function_space()
m.coordinates = new_coords
ufl.dx._subdomain_data = m.coordinates
V = FunctionSpace(m, 'CG', 1)
u = Function(V)
u.assign(1)
assert abs(assemble(u*dx) - 1.0) < 1e-12
e = ExtrudedMesh(m, layers=4, layer_height=0.25)
V = FunctionSpace(e, 'CG', 1)
u = Function(V)
u.assign(1)
assert abs(assemble(u*dx) - 1.0) < 1e-12
def test_extruded_periodic_interval_area():
m = PeriodicUnitIntervalMesh(10)
V = FunctionSpace(m, 'CG', 1)
u = Function(V)
u.assign(1)
assert abs(assemble(u*dx) - 1.0) < 1e-12
e = ExtrudedMesh(m, layers=4, layer_height=0.25)
V = FunctionSpace(e, 'CG', 1)
u = Function(V)
u.assign(1)
assert abs(assemble(u*dx) - 1.0) < 1e-12
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_circle_manifold.py
from firedrake import *
import numpy as np
import pytest
@pytest.mark.parametrize('i', range(3, 11))
def test_circumference(i):
eps = 1e-12
mesh = CircleManifoldMesh(i, radius=i*i) # noqa: need for dx
f = Constant(1.0)
# 2 * radius * sin(pi/i) * number of sides
circumference = 2*i*i*np.sin(np.pi/i)*i
assert np.abs(assemble(f*dx) - circumference) < eps
def test_pi():
len = 10
errors = np.zeros(len)
for i in range(2, 2+len):
mesh = CircleManifoldMesh(2**i) # noqa: need for dx
f = Constant(1.0)
errors[i-2] = np.abs(assemble(f*dx) - 2*np.pi)
# circumference converges quadratically to 2*pi
for i in range(len-1):
assert ln(errors[i]/errors[i+1])/ln(2) > 1.95
if __name__ == '__main__':
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/utils.py
# Some generic python utilities not really specific to our work.
from decorator import decorator
# after https://micheles.googlecode.com/hg/decorator/documentation.html and
# http://code.activestate.com/recipes/577452-a-memoize-decorator-for-instance-methods/
def _memoize(func, obj, *args, **kw):
try:
cache = obj.__cache
except AttributeError:
cache = obj.__cache = {}
if kw:
key = func, args, tuple(kw.iteritems())
else:
key = func, args
if key in cache:
return cache[key]
else:
cache[key] = result = func(obj, *args, **kw)
return result
def memoize(f):
return decorator(_memoize, f)
# from http://www.toofishes.net/blog/python-cached-property-decorator/
class cached_property(object):
'''A read-only @property that is only evaluated once. The value is cached
on the object itself rather than the function or class; this should prevent
memory leakage.'''
def __init__(self, fget, doc=None):
self.fget = fget
self.__doc__ = doc or fget.__doc__
self.__name__ = fget.__name__
self.__module__ = fget.__module__
def __get__(self, obj, cls):
if obj is None:
return self
obj.__dict__[self.__name__] = result = self.fget(obj)
return result
_current_uid = 0
def _new_uid():
global _current_uid
_current_uid += 1
return _current_uid
def _init():
"""Cause :func:`pyop2.init` to be called in case the user has not done it
for themselves. The result of this is that the user need only call
:func:`pyop2.init` if she wants to set a non-default option, for example
to switch the backend or the debug or log level."""
from pyop2 import op2
from parameters import parameters
if not op2.initialised():
op2.init(log_level='INFO',
compiler=parameters["coffee"]["compiler"],
simd_isa=parameters["coffee"]["simd_isa"])
def unique_name(name, nameset):
"""Return name if name is not in nameset, or a deterministic
uniquified name if name is in nameset. The new name is inserted into
nameset to prevent further name clashes."""
if name not in nameset:
nameset.add(name)
return name
idx = 0
while True:
newname = "%s_%d" % (name, idx)
if newname in nameset:
idx += 1
else:
nameset.add(name)
return newname
<file_sep>/tests/regression/test_poisson_strong_bcs.py
"""This demo program solves Poisson's equation
- div grad u(x, y) = 0
on the unit square with boundary conditions given by:
u(0, y) = 0
v(1, y) = 42
Homogeneous Neumann boundary conditions are applied naturally on the
other two sides of the domain.
This has the analytical solution
u(x, y) = 42*x[1]
"""
import pytest
import numpy as np
from firedrake import *
def run_test(x, degree, parameters={}, quadrilateral=False):
# Create mesh and define function space
mesh = UnitSquareMesh(2 ** x, 2 ** x, quadrilateral=quadrilateral)
V = FunctionSpace(mesh, "CG", degree)
# Define variational problem
u = Function(V)
v = TestFunction(V)
a = dot(grad(v), grad(u)) * dx
bcs = [DirichletBC(V, 0, 3),
DirichletBC(V, 42, 4)]
# Compute solution
solve(a == 0, u, solver_parameters=parameters, bcs=bcs)
f = Function(V)
f.interpolate(Expression("42*x[1]"))
return sqrt(assemble(dot(u - f, u - f) * dx))
def run_test_linear(x, degree, parameters={}, quadrilateral=False):
# Create mesh and define function space
mesh = UnitSquareMesh(2 ** x, 2 ** x, quadrilateral=quadrilateral)
V = FunctionSpace(mesh, "CG", degree)
# Define variational problem
u = TrialFunction(V)
v = TestFunction(V)
a = dot(grad(v), grad(u)) * dx
L = v*0*dx
bcs = [DirichletBC(V, 0, 3),
DirichletBC(V, 42, 4)]
# Compute solution
u = Function(V)
solve(a == L, u, solver_parameters=parameters, bcs=bcs)
f = Function(V)
f.interpolate(Expression("42*x[1]"))
return sqrt(assemble(dot(u - f, u - f) * dx))
def run_test_preassembled(x, degree, parameters={}, quadrilateral=False):
mesh = UnitSquareMesh(2 ** x, 2 ** x, quadrilateral=quadrilateral)
V = FunctionSpace(mesh, "CG", degree)
u = TrialFunction(V)
v = TestFunction(V)
a = dot(grad(v), grad(u)) * dx
f = Function(V)
f.assign(0)
L = v*f*dx
bcs = [DirichletBC(V, 0, 3),
DirichletBC(V, 42, 4)]
u = Function(V)
A = assemble(a)
b = assemble(L)
for bc in bcs:
bc.apply(A)
bc.apply(b)
solve(A, u, b, solver_parameters=parameters)
expected = Function(V)
expected.interpolate(Expression("42*x[1]"))
method_A = sqrt(assemble(dot(u - expected, u - expected) * dx))
A = assemble(a)
b = assemble(L)
solve(A, u, b, bcs=bcs, solver_parameters=parameters)
method_B = sqrt(assemble(dot(u - expected, u - expected) * dx))
A = assemble(a, bcs=bcs)
b = assemble(L, bcs=bcs)
solve(A, u, b, solver_parameters=parameters)
method_C = sqrt(assemble(dot(u - expected, u - expected) * dx))
A = assemble(a, bcs=bcs)
b = assemble(L)
solve(A, u, b, solver_parameters=parameters)
method_D = sqrt(assemble(dot(u - expected, u - expected) * dx))
A = assemble(a)
b = assemble(L)
# Don't actually need to apply the bcs to b explicitly since it's
# done in the solve if A has any.
for bc in bcs:
bc.apply(A)
solve(A, u, b, solver_parameters=parameters)
method_E = sqrt(assemble(dot(u - expected, u - expected) * dx))
A = assemble(a, bcs=[bcs[0]])
b = assemble(L)
# This will not give the right answer
solve(A, u, b, solver_parameters=parameters)
bcs[1].apply(A)
b = assemble(L)
# This will, because we reassemble using the new set of bcs
solve(A, u, b, solver_parameters=parameters)
method_F = sqrt(assemble(dot(u - expected, u - expected) * dx))
return np.asarray([method_A, method_B, method_C, method_D, method_E, method_F])
@pytest.mark.parametrize(['params', 'degree', 'quadrilateral'],
[(p, d, q)
for p in [{}, {'snes_type': 'ksponly', 'ksp_type': 'preonly', 'pc_type': 'lu'}]
for d in (1, 2)
for q in [False, True]])
def test_poisson_analytic(params, degree, quadrilateral):
assert (run_test(2, degree, parameters=params, quadrilateral=quadrilateral) < 1.e-9)
@pytest.mark.parametrize(['params', 'degree', 'quadrilateral'],
[(p, d, q)
for p in [{}, {'snes_type': 'ksponly', 'ksp_type': 'preonly', 'pc_type': 'lu'}]
for d in (1, 2)
for q in [False, True]])
def test_poisson_analytic_linear(params, degree, quadrilateral):
assert (run_test_linear(2, degree, parameters=params, quadrilateral=quadrilateral) < 5.e-6)
@pytest.mark.parametrize(['params', 'degree', 'quadrilateral'],
[(p, d, q)
for p in [{}, {'snes_type': 'ksponly', 'ksp_type': 'preonly', 'pc_type': 'lu'}]
for d in (1, 2)
for q in [False, True]])
def test_poisson_analytic_preassembled(params, degree, quadrilateral):
assert (run_test_preassembled(2, degree, parameters=params, quadrilateral=quadrilateral) < 5.e-6).all()
@pytest.mark.parallel(nprocs=2)
def test_poisson_analytic_linear_parallel():
from mpi4py import MPI
error = run_test_linear(1, 1)
print '[%d]' % MPI.COMM_WORLD.rank, 'error:', error
assert error < 5e-6
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/test_0init.py
import pytest
from firedrake import *
def test_pyop2_not_initialised():
"""Check that PyOP2 has not been initialised yet."""
assert not op2.initialised()
def test_pyop2_custom_init():
"""PyOP2 init parameters set by the user should be retained."""
op2.init(debug=3, log_level='CRITICAL')
UnitIntervalMesh(2)
from pyop2.logger import logger
assert logger.getEffectiveLevel() == CRITICAL
assert op2.configuration['debug'] == 3
op2.configuration.reset()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/setup.py
from distutils.core import setup
from distutils.extension import Extension
from glob import glob
from os import environ as env, path
import sys
import numpy as np
import petsc4py
def get_petsc_dir():
try:
petsc_arch = env.get('PETSC_ARCH', '')
petsc_dir = env['PETSC_DIR']
if petsc_arch:
return (petsc_dir, path.join(petsc_dir, petsc_arch))
return (petsc_dir,)
except KeyError:
try:
import petsc
return (petsc.get_petsc_dir(), )
except ImportError:
sys.exit("""Error: Could not find PETSc library.
Set the environment variable PETSC_DIR to your local PETSc base
directory or install PETSc from PyPI as described in the manual:
http://firedrakeproject.org/obtaining_pyop2.html#petsc
""")
import versioneer
versioneer.versionfile_source = 'firedrake/_version.py'
versioneer.versionfile_build = 'firedrake/_version.py'
versioneer.tag_prefix = 'v'
versioneer.parentdir_prefix = 'firedrake-'
versioneer.VCS = "git"
cmdclass = versioneer.get_cmdclass()
try:
from Cython.Distutils import build_ext
cmdclass['build_ext'] = build_ext
dmplex_sources = ["firedrake/dmplex.pyx"]
evtk_sources = ['evtk/cevtk.pyx']
except ImportError:
# No cython, dmplex.c must be generated in distributions.
dmplex_sources = ["firedrake/dmplex.c"]
evtk_sources = ['evtk/cevtk.c']
if 'CC' not in env:
env['CC'] = "mpicc"
petsc_dirs = get_petsc_dir()
include_dirs = [np.get_include(), petsc4py.get_include()]
include_dirs += ["%s/include" % d for d in petsc_dirs]
setup(name='firedrake',
version=versioneer.get_version(),
cmdclass=cmdclass,
description="""Firedrake is an automated system for the portable solution
of partial differential equations using the finite element method
(FEM)""",
author="<NAME> and others",
author_email="<EMAIL>",
url="http://firedrakeproject.org",
packages=["firedrake", "evtk"],
package_data={"firedrake": ["firedrake_geometry.h"]},
scripts=glob('scripts/*'),
ext_modules=[Extension('firedrake.dmplex',
sources=dmplex_sources,
include_dirs=include_dirs,
libraries=["petsc"],
extra_link_args=["-L%s/lib" % d for d in petsc_dirs] +
["-Wl,-rpath,%s/lib" % d for d in petsc_dirs] +
["-Wl,-rpath,%s/lib" % sys.prefix]),
Extension('evtk.cevtk', evtk_sources,
include_dirs=[np.get_include()])])
<file_sep>/tests/regression/test_hybridisation.py
"""Solve a mixed Helmholtz problem
sigma + grad(u) = 0
u + div(sigma) = f
using hybridisation. The corresponding weak (variational problem)
<tau, sigma> - <div(tau), u> + <<[tau.n], lambda>> = 0 for all tau
<v, u> + <v, div(sigma)> = <v, f> for all v
<<gamma, [sigma.n]>> = 0 for all gamma
is solved using broken RT (Raviart-Thomas) elements of degree k for
(sigma, tau), DG (discontinuous Galerkin) elements of degree k - 1
for (u, v), and Trace-RT elements for (lambda, gamma).
No strong boundary conditions are enforced. A weak boundary condition on
u is enforced implicitly, setting <<u, tau.n>> = 0 for all tau.
The forcing function is chosen as
(1+8*pi*pi)*sin(x[0]*pi*2)*sin(x[1]*pi*2)
which reproduces the known analytical solution
sin(x[0]*pi*2)*sin(x[1]*pi*2)
"""
import pytest
from firedrake import *
@pytest.mark.parametrize('degree', range(1, 3))
def test_hybridisation(degree):
# Create mesh
mesh = UnitSquareMesh(8, 8)
# Define function spaces and mixed (product) space
RT_elt = FiniteElement("RT", triangle, degree)
BrokenRT = FunctionSpace(mesh, BrokenElement(RT_elt))
DG = FunctionSpace(mesh, "DG", degree-1)
TraceRT = FunctionSpace(mesh, TraceElement(RT_elt))
W = MixedFunctionSpace([BrokenRT, DG, TraceRT])
# Define trial and test functions
sigma, u, lambdar = TrialFunctions(W)
tau, v, gammar = TestFunctions(W)
# Mesh normal
n = FacetNormal(mesh)
# Define source function
f = Function(DG)
f.interpolate(Expression("(1+8*pi*pi)*sin(x[0]*pi*2)*sin(x[1]*pi*2)"))
# Define variational form
a_dx = (dot(tau, sigma) - div(tau)*u + v*u + v*div(sigma))*dx
a_dS = (jump(tau, n=n)*lambdar('+') + gammar('+')*jump(sigma, n=n))*dS
a = a_dx + a_dS
L = f*v*dx
bcs = DirichletBC(W.sub(2), Constant(0), (1, 2, 3, 4))
# Compute solution
w = Function(W)
solve(a == L, w, solver_parameters={'ksp_rtol': 1e-14,
'ksp_max_it': 30000},
bcs=bcs)
Hsigma, Hu, Hlambdar = w.split()
# Compare result to non-hybridised calculation
RT = FunctionSpace(mesh, "RT", degree)
W2 = RT * DG
sigma, u = TrialFunctions(W2)
tau, v = TestFunctions(W2)
w2 = Function(W2)
a = (dot(tau, sigma) - div(tau)*u + v*u + v*div(sigma))*dx
L = f*v*dx
solve(a == L, w2, solver_parameters={'ksp_rtol': 1e-14})
NHsigma, NHu = w2.split()
# Return L2 norm of error
# (should be identical, i.e. comparable with solver tol)
uerr = sqrt(assemble((Hu-NHu)*(Hu-NHu)*dx))
sigerr = sqrt(assemble(dot(Hsigma-NHsigma, Hsigma-NHsigma)*dx))
assert uerr < 1e-11
assert sigerr < 4e-11
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_nullspace.py
from firedrake import *
import pytest
@pytest.fixture(scope='module')
def V():
m = UnitSquareMesh(25, 25)
return FunctionSpace(m, 'CG', 1)
def test_nullspace(V):
u = TrialFunction(V)
v = TestFunction(V)
a = inner(grad(u), grad(v))*dx
L = -v*ds(3) + v*ds(4)
nullspace = VectorSpaceBasis(constant=True)
u = Function(V)
solve(a == L, u, nullspace=nullspace)
exact = Function(V)
exact.interpolate(Expression('x[1] - 0.5'))
assert sqrt(assemble((u - exact)*(u - exact)*dx)) < 5e-8
def test_nullspace_preassembled(V):
u = TrialFunction(V)
v = TestFunction(V)
a = inner(grad(u), grad(v))*dx
L = -v*ds(3) + v*ds(4)
nullspace = VectorSpaceBasis(constant=True)
u = Function(V)
A = assemble(a)
b = assemble(L)
solve(A, u, b, nullspace=nullspace)
exact = Function(V)
exact.interpolate(Expression('x[1] - 0.5'))
assert sqrt(assemble((u - exact)*(u - exact)*dx)) < 5e-8
def test_nullspace_mixed():
m = UnitSquareMesh(5, 5)
BDM = FunctionSpace(m, 'BDM', 1)
DG = FunctionSpace(m, 'DG', 0)
W = BDM * DG
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
a = (dot(sigma, tau) + div(tau)*u + div(sigma)*v)*dx
bcs = [DirichletBC(W.sub(0), (0, 0), (1, 2)),
DirichletBC(W.sub(0), (0, 1), (3, 4))]
w = Function(W)
f = Function(DG)
f.assign(0)
L = f*v*dx
# Null space is constant functions in DG and empty in BDM.
nullspace = MixedVectorSpaceBasis(W, [W.sub(0), VectorSpaceBasis(constant=True)])
solve(a == L, w, bcs=bcs, nullspace=nullspace)
exact = Function(DG)
exact.interpolate(Expression('x[1] - 0.5'))
sigma, u = w.split()
assert sqrt(assemble((u - exact)*(u - exact)*dx)) < 1e-7
# Now using a Schur complement
w.assign(0)
solve(a == L, w, bcs=bcs, nullspace=nullspace,
solver_parameters={'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'ksp_type': 'cg',
'pc_fieldsplit_schur_fact_type': 'full',
'fieldsplit_0_ksp_type': 'preonly',
'fieldsplit_0_pc_type': 'lu',
'fieldsplit_1_ksp_type': 'cg',
'fieldsplit_1_pc_type': 'none'})
sigma, u = w.split()
assert sqrt(assemble((u - exact)*(u - exact)*dx)) < 5e-8
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/demos/extruded_upwind_advection/upwind_advection.py.rst
Steady-state advection equation with upwinding
==============================================
We next consider the equation
.. math::
\nabla\cdot(\vec{u_0}D) = 0
in a domain :math:`\Omega`, where :math:`\vec{u_0}` is a prescribed vector
field, and :math:`D` is an unknown scalar field. The value of :math:`D` is known
on the subset of the boundary :math:`\Gamma` in which :math:`\vec{u_0}` is
directed towards the interior of the domain:
.. math::
D = D_0 \quad \mathrm{on} \ \Gamma_\mathrm{inflow}
where :math:`\Gamma_\mathrm{inflow}` is defined appropriately. :math:`D` can
be interpreted as the steady-state distribution of a passive tracer carried by a
fluid with velocity field :math:`\vec{u_0}`.
A weak form of the continuous equation is
.. math::
\int_\Omega \! \phi \nabla \cdot (\vec{u_0} D) \, \mathrm{d} x &= 0 \quad
\forall \ \phi \in D(\Omega), \\
D &= D_0 \quad \mathrm{on} \ \Gamma_\mathrm{inflow}
where :math:`D(\Omega)` is the space of smooth *test functions* with compact
support in :math:`\Omega`. We will look for a solution :math:`D` in a space of
*discontinuous* functions :math:`V`. This suggests using integration by parts to
avoid taking the derivative of a discontinuous quantity:
.. math::
\sum_e \left( \int_{\partial e} \! \phi_e D \vec{u_0} \cdot \vec{n} \,
\mathrm{d} S - \int_e \! D \vec{u_0} \cdot \nabla \phi \, \mathrm{d} x \right) = 0
\quad \forall \ \phi \in V, \\
D = D_0 \quad \mathrm{on} \ \Gamma_\mathrm{inflow}
where the sum is taken over all elements. Since :math:`D` is discontinuous, we
have to make a choice about how it is defined on facets in order to evaluate
the first integral. We will use upwinding: the *upstream* value of :math:`D` is
used on the facet. In light of this, there are three distinct situations we may
encounter:
1. Boundary facets where :math:`\vec{u_0}` points towards the interior of the
domain. Here, the prescribed boundary value :math:`D_0` is used.
2. Boundary facets where :math:`\vec{u_0}` points away from the interior of the
domain. Here, the (unknown) interior solution value :math:`D` is used.
3. Interior facets. Here, the upstream value of :math:`D`,
:math:`\widetilde{D}`, is used.
Note that each of the interior facets contributes to the integral twice. The two
contributions differ in the choice of test function: a subscript :math:`\phi_e`
was used to make this explicit. The full set of equations are then
.. math::
-\int_\Omega \! D \vec{u_0} \cdot \nabla \phi \, \mathrm{d} x
+ \int_{\Gamma_\rlap{\mathrm{ext, outflow}}} \! \phi D \vec{u_0} \cdot \vec{n}
\, \mathrm{d} s
+ \int_{\Gamma_\mathrm{int}} \! (\phi_+ - \phi_-) \widetilde{D}
\vec{u_0} \cdot \vec{n} \, \mathrm{d} S
\quad = \quad
-\int_{\Gamma_\rlap{\mathrm{ext, inflow}}} \phi D_0 \vec{u_0} \cdot
\vec{n} \, \mathrm{d} s \quad \forall \ \phi \in V,
D = D_0 \quad \mathrm{on} \ \Gamma_\mathrm{inflow}
In this worked example, we will take the domain :math:`\Omega` to be the cuboid
:math:`\Omega = [0,1] \times [0,1] \times [0,0.2]`. We will use the constant
velocity field :math:`\vec{u_0} = (0, 0, 1)`. :math:`\Gamma_\mathrm{inflow}`
is therefore the base of the cuboid, while :math:`\Gamma_\mathrm{outflow}`
is the top. The four vertical sides can be ignored, since
:math:`\vec{u_0} \cdot \vec{n} = 0` on these faces.
Firedrake code for this example is as follows:
We will use an *extruded* mesh, where the base mesh is a 20 by 20 unit square,
with 10 evenly-spaced vertical layers. This gives prism-shaped cells. ::
from firedrake import *
m = UnitSquareMesh(20, 20)
mesh = ExtrudedMesh(m, layers=10, layer_height=0.02)
We will use a simple piecewise-constant function space for the unknown scalar
:math:`D`: ::
V = FunctionSpace(mesh, "DG", 0)
Our velocity will live in a low-order Raviart-Thomas space. The construction of
this is more complicated than element spaces you will have seen previously. The
horizontal and vertical components of the field are specified separately. They
are combined into a single element which is used to build a FunctionSpace. ::
# RT1 element on a prism
W0_h = FiniteElement("RT", "triangle", 1)
W0_v = FiniteElement("DG", "interval", 0)
W0 = HDiv(OuterProductElement(W0_h, W0_v))
W1_h = FiniteElement("DG", "triangle", 0)
W1_v = FiniteElement("CG", "interval", 1)
W1 = HDiv(OuterProductElement(W1_h, W1_v))
W_elt = W0 + W1
W = FunctionSpace(mesh, W_elt)
As an aside, since our prescibed velocity is purely in the vertical direction, a
simpler space would have sufficed: ::
# Vertical part of RT1 element
# W_h = FiniteElement("DG", "triangle", 0)
# W_v = FiniteElement("CG", "interval", 1)
# W_elt = HDiv(OuterProductElement(W_h, W_v))
# W = FunctionSpace(mesh, W_elt)
Or even: ::
# Why can't everything in life be this easy?
# W = VectorFunctionSpace(mesh, "CG", 1)
Next, we set the prescribed velocity field: ::
velocity = Expression(("0.0", "0.0", "1.0"))
u0 = project(velocity, W)
# if we had used W = VectorFunctionSpace(mesh, "CG", 1), we could have done
# u0 = Function(W)
# u0.interpolate(velocity)
Next, we will set the boundary value on our scalar to be a simple indicator
function over part of the bottom of the domain: ::
inflow = Expression("(x[2] < 0.02) && (x[0] > 0.5) ? 1.0 : -1.0")
D0 = Function(V)
D0.interpolate(inflow)
Now we will define our forms. There are several new concepts here. Firstly, we
will define a new variable ``un`` which takes the value
:math:`\vec{u_0} \cdot \vec{n}` when this is positive, otherwise `0`. This
will be useful for our upwind terms. ::
n = FacetNormal(mesh)
un = 0.5*(dot(u0, n) + abs(dot(u0, n)))
We define our trial and test functions in the usual way: ::
D = TrialFunction(V)
phi = TestFunction(V)
Since we are on an extruded mesh, we have several new integral types at our
disposal. An integral over the interior of the domain is still denoted by
``dx``. Boundary integrals now come in several varieties: ``ds_b`` denotes an
integral over the base of the mesh, while ``ds_t`` denotes an integral over the
top of the mesh. ``ds_v`` denotes an integral over the sides of a mesh, though
we will not use that here.
Similiarly, interior facet integrals are split into ``dS_h`` and ``dS_v``, over
*horizontal* interior facets and *vertical* interior facets respectively. Since
our velocity field is purely in the vertical direction, we will omit the
integral over vertical interior facets, since we know
:math:`\vec{u_0} \cdot \vec{n}` is zero for these. ::
a1 = -D*dot(u0, grad(phi))*dx
a2 = dot(jump(phi), un('+')*D('+') - un('-')*D('-'))*dS_h
a3 = dot(phi, un*D)*ds_t # outflow at top wall
a = a1 + a2 + a3
L = -D0*phi*dot(u0, n)*ds_b # inflow at bottom wall
Finally, we will compute the solution: ::
out = Function(V)
solve(a == L, out)
By construction, the exact solution is quite simple: ::
exact = Function(V)
exact.interpolate(Expression("(x[0] > 0.5) ? 1.0 : -1.0"))
We finally compare our solution to the expected solution: ::
assert max(abs(out.dat.data - exact.dat.data)) < 1e-10
This demo can be found as a script in
`upwind_advection.py <upwind_advection.py>`__.
<file_sep>/tests/regression/test_dg_advection.py
from firedrake import *
import numpy as np
import pytest
def run_test():
mesh = UnitIcosahedralSphereMesh(refinement_level=3)
mesh.init_cell_orientations(Expression(("x[0]", "x[1]", "x[2]")))
V = FunctionSpace(mesh, "DG", 0)
M = VectorFunctionSpace(mesh, "CG", 1)
# advecting velocity
u0 = Expression(('-x[1]*(1 - x[2]*x[2])', 'x[0]*(1 - x[2]*x[2])', '0'))
u = Function(M).interpolate(u0)
dt = (pi/3) * 0.006
phi = TestFunction(V)
D = TrialFunction(V)
n = FacetNormal(mesh)
un = 0.5 * (dot(u, n) + abs(dot(u, n)))
a_mass = phi*D*dx
a_int = dot(grad(phi), -u*D)*dx
a_flux = dot(jump(phi), un('+')*D('+') - un('-')*D('-'))*dS
arhs = a_mass - dt * (a_int + a_flux)
dD1 = Function(V)
D1 = Function(V)
D0 = Expression("x[0] < 0 ? 1: 0")
D = Function(V).interpolate(D0)
t = 0.0
T = 10*dt
problem = LinearVariationalProblem(a_mass, action(arhs, D1), dD1)
solver = LinearVariationalSolver(problem, parameters={'ksp_type': 'cg'})
L2_0 = norm(D)
Dbar_0 = assemble(D*dx)
while t < (T - dt/2):
D1.assign(D)
solver.solve()
D1.assign(dD1)
solver.solve()
D1.assign(0.75*D + 0.25*dD1)
solver.solve()
D.assign((1.0/3.0)*D + (2.0/3.0)*dD1)
t += dt
L2_T = norm(D)
Dbar_T = assemble(D*dx)
# L2 norm decreases
assert L2_T < L2_0
# Mass conserved
assert np.allclose(Dbar_T, Dbar_0)
def test_dg_advection():
run_test()
@pytest.mark.parallel(nprocs=3)
def test_dg_advection_parallel():
run_test()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_adv_diff.py
"""Firedrake P1 advection-diffusion with operator splitting demo
This demo solves the advection-diffusion equation by splitting the advection
and diffusion terms. The advection term is advanced in time using an Euler
method and the diffusion term is advanced in time using a theta scheme with
theta = 0.5.
"""
import pytest
from firedrake import *
def adv_diff(x, quadrilateral=False, advection=True, diffusion=True):
dt = 0.0001
T = 0.01
# Create mesh and define function space
mesh = UnitSquareMesh(2 ** x, 2 ** x, quadrilateral=quadrilateral)
V = FunctionSpace(mesh, "CG", 1)
U = VectorFunctionSpace(mesh, "CG", 1)
p = TrialFunction(V)
q = TestFunction(V)
t = Function(V)
u = Function(U)
diffusivity = 0.1
adv = p * q * dx
adv_rhs = (q * t + dt * dot(grad(q), u) * t) * dx
d = -dt * diffusivity * dot(grad(q), grad(p)) * dx
diff = adv - 0.5 * d
diff_rhs = action(adv + 0.5 * d, t)
if advection:
A = assemble(adv)
if diffusion:
D = assemble(diff)
# Set initial condition:
# A*(e^(-r^2/(4*D*T)) / (4*pi*D*T))
# with normalisation A = 0.1, diffusivity D = 0.1
r2 = "(pow(x[0]-(0.45+%(T)f), 2.0) + pow(x[1]-0.5, 2.0))"
fexpr = "0.1 * (exp(-" + r2 + "/(0.4*%(T)f)) / (0.4*pi*%(T)f))"
t.interpolate(Expression(fexpr % {'T': T}))
u.interpolate(Expression([1.0, 0.0]))
while T < 0.012:
# Advection
if advection:
b = assemble(adv_rhs)
solve(A, t, b)
# Diffusion
if diffusion:
b = assemble(diff_rhs)
solve(D, t, b)
T = T + dt
# Analytical solution
a = Function(V).interpolate(Expression(fexpr % {'T': T}))
return sqrt(assemble(dot(t - a, t - a) * dx))
def run_adv_diff():
import numpy as np
diff = np.array([adv_diff(i) for i in range(5, 8)])
convergence = np.log2(diff[:-1] / diff[1:])
assert all(convergence > [1.8, 1.95])
def test_adv_diff_serial():
run_adv_diff()
@pytest.mark.parallel
def test_adv_diff_parallel():
run_adv_diff()
def run_adv_diff_on_quadrilaterals():
import numpy as np
diff = np.array([adv_diff(i, quadrilateral=True) for i in range(5, 8)])
convergence = np.log2(diff[:-1] / diff[1:])
assert all(convergence > [1.8, 1.95])
def test_adv_diff_on_quadrilaterals_serial():
run_adv_diff_on_quadrilaterals()
@pytest.mark.parallel
def test_adv_diff_on_quadrilaterals_parallel():
run_adv_diff_on_quadrilaterals()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_0_mixed_mats.py
import pytest
import numpy as np
from firedrake import *
xfail = pytest.mark.xfail
@pytest.fixture(scope='module')
def m(request):
return ExtrudedMesh(UnitTriangleMesh(), layers=2, layer_height=0.5)
@pytest.fixture(scope='module')
def V(m):
return FunctionSpace(m, 'DG', 0)
@pytest.fixture(scope='module')
def Q(m):
return FunctionSpace(m, 'DG', 0)
@pytest.fixture(scope='module')
def W(V, Q):
return V*Q
# NOTE: these tests make little to no mathematical sense, they are
# here to exercise corner cases in PyOP2's handling of mixed spaces.
def test_massVW0(V, W):
u = TrialFunction(V)
v = TestFunction(W)[0]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 1)
# DGxDG block
assert not np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block (0, since test function was restricted to DG block)
assert np.allclose(A.M[1, 0].values, 0.0)
def test_massVW1(V, W):
u = TrialFunction(V)
v = TestFunction(W)[1]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 1)
# DGxDG block (0, since test function was restricted to RT block)
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert not np.allclose(A.M[1, 0].values, 0.0)
def test_massW0W0(W):
u = TrialFunction(W)[0]
v = TestFunction(W)[0]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert not np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert np.allclose(A.M[1, 1].values, 0.0)
def test_massW1W1(W):
u = TrialFunction(W)[1]
v = TestFunction(W)[1]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert not np.allclose(A.M[1, 1].values, 0.0)
def test_massW0W1(W):
u = TrialFunction(W)[0]
v = TestFunction(W)[1]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert not np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert np.allclose(A.M[1, 1].values, 0.0)
def test_massW1W0(W):
u = TrialFunction(W)[1]
v = TestFunction(W)[0]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert not np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert np.allclose(A.M[1, 1].values, 0.0)
def test_massWW(W):
u = TrialFunction(W)
v = TestFunction(W)
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert not np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert not np.allclose(A.M[1, 1].values, 0.0)
<file_sep>/tests/extrusion/test_extrusion_facet_integrals_3D.py
"""Testing assembly of scalars on facets of extruded meshes in 3D"""
import pytest
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module', params=[1, 2])
def f(request):
mesh = extmesh(4, 4, 4)
fspace = FunctionSpace(mesh, "CG", request.param)
return Function(fspace)
@pytest.fixture(scope='module')
def RT2():
mesh = extmesh(4, 4, 4)
U1 = FiniteElement("RT", "triangle", 2)
U2 = FiniteElement("DG", "triangle", 1)
V0 = FiniteElement("CG", "interval", 2)
V1 = FiniteElement("DG", "interval", 1)
W2 = HDiv(OuterProductElement(U1, V1)) + HDiv(OuterProductElement(U2, V0))
return FunctionSpace(mesh, W2)
def test_scalar_area(f):
f.assign(1)
assert abs(assemble(f*ds_t) - 1.0) < 1e-7
assert abs(assemble(f*ds_b) - 1.0) < 1e-7
assert abs(assemble(f*ds_tb) - 2.0) < 1e-7
assert abs(assemble(f*ds_v) - 4.0) < 1e-7
assert abs(assemble(f('+')*dS_h) - 3.0) < 1e-7
assert abs(assemble(f('-')*dS_h) - 3.0) < 1e-7
assert abs(assemble(f('+')*dS_v) - (6.0 + 4*sqrt(2))) < 1e-7
assert abs(assemble(f('-')*dS_v) - (6.0 + 4*sqrt(2))) < 1e-7
def test_scalar_expression(f):
f.interpolate(Expression("x[2]"))
assert abs(assemble(f*ds_t) - 1.0) < 1e-7
assert abs(assemble(f*ds_b) - 0.0) < 1e-7
assert abs(assemble(f*ds_tb) - 1.0) < 1e-7
assert abs(assemble(f*ds_v) - 2.0) < 1e-7
assert abs(assemble(f('+')*dS_h) - 1.5) < 1e-7
assert abs(assemble(f('-')*dS_h) - 1.5) < 1e-7
assert abs(assemble(f('+')*dS_v) - 0.5*(6.0 + 4*sqrt(2))) < 1e-7
assert abs(assemble(f('-')*dS_v) - 0.5*(6.0 + 4*sqrt(2))) < 1e-7
def test_hdiv_area(RT2):
f = project(Expression(("0.0", "0.8", "0.6")), RT2)
assert abs(assemble(dot(f, f)*ds_t) - 1.0) < 1e-7
assert abs(assemble(dot(f, f)*ds_b) - 1.0) < 1e-7
assert abs(assemble(dot(f, f)*ds_tb) - 2.0) < 1e-7
assert abs(assemble(dot(f, f)*ds_v) - 4.0) < 1e-7
assert abs(assemble(dot(f('+'), f('+'))*dS_h) - 3.0) < 1e-7
assert abs(assemble(dot(f('-'), f('-'))*dS_h) - 3.0) < 1e-7
assert abs(assemble(dot(f('+'), f('-'))*dS_h) - 3.0) < 1e-7
assert abs(assemble(dot(f('+'), f('+'))*dS_v) - (6.0 + 4*sqrt(2))) < 1e-7
assert abs(assemble(dot(f('-'), f('-'))*dS_v) - (6.0 + 4*sqrt(2))) < 1e-7
assert abs(assemble(dot(f('+'), f('-'))*dS_v) - (6.0 + 4*sqrt(2))) < 1e-7
def test_exterior_horizontal_normals(RT2):
n = FacetNormal(RT2.mesh())
f = project(Expression(("1.0", "0.0", "0.0")), RT2)
assert abs(assemble(dot(f, n)*ds_t) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_b) - 0.0) < 1e-7
f = project(Expression(("0.0", "0.0", "1.0")), RT2)
assert abs(assemble(dot(f, n)*ds_t) - 1.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_b) - (-1.0)) < 1e-7
def test_exterior_vertical_normals(RT2):
n = FacetNormal(RT2.mesh())
f = project(Expression(("1.0", "0.0", "0.0")), RT2)
assert abs(assemble(dot(f, n)*ds_v(1)) - (-1.0)) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(2)) - 1.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(3)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(4)) - 0.0) < 1e-7
f = project(Expression(("0.0", "1.0", "0.0")), RT2)
assert abs(assemble(dot(f, n)*ds_v(1)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(2)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(3)) - (-1.0)) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(4)) - 1.0) < 1e-7
f = project(Expression(("0.0", "0.0", "1.0")), RT2)
assert abs(assemble(dot(f, n)*ds_v(1)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(2)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(3)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(4)) - 0.0) < 1e-7
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/parameters.py
"""The parameters dictionary contains global parameter settings."""
import os
from ffc import default_parameters
__all__ = ['Parameters', 'parameters']
class Parameters(dict):
def __init__(self, name=None, **kwargs):
self._name = name
for key, value in kwargs.iteritems():
self.add(key, value)
def add(self, key, value=None):
if isinstance(key, Parameters):
self[key.name()] = key
else:
self[key] = value
def name(self):
return self._name
def rename(self, name):
self._name = name
parameters = Parameters()
parameters.add(Parameters("assembly_cache",
enabled=True,
eviction=True,
max_bytes=float("Inf"),
max_factor=0.6,
max_misses=3))
parameters.add(Parameters("coffee",
compiler=os.environ.get('PYOP2_BACKEND_COMPILER', 'gnu'),
simd_isa=os.environ.get('PYOP2_SIMD_ISA', 'sse'),
licm=False,
slice=None,
vect=None,
ap=False,
split=None))
ffc_parameters = default_parameters()
ffc_parameters['write_file'] = False
ffc_parameters['format'] = 'pyop2'
ffc_parameters['representation'] = 'quadrature'
ffc_parameters['pyop2-ir'] = True
parameters.add(Parameters("form_compiler", **ffc_parameters))
parameters["reorder_meshes"] = True
<file_sep>/docs/source/download.rst
Obtaining Firedrake
===================
Firedrake depends on PyOP2_, FFC_, FIAT_, and UFL_. It is easiest to obtain
all of these components on Ubuntu Linux and related distributions such as Mint
or Debian. Installation on other Unix-like operating systems is likely to be
possible, although harder. Installation on a Mac is straightforward using the
commands below.
PyOP2
-----
Instructions for obtaining PyOP2_ and its dependencies are at
:doc:`obtaining_pyop2`. Note that PyOP2_ is updated frequently and Firedrake
requires an up-to-date version.
FFC, FIAT and UFL
-----------------
Firedrake currently requires a fork of FFC_, UFL_ and FIAT_. Note that FFC_
requires a version of Instant_.
FFC_ currently depends on Swig_, which you can install from
package. On Ubuntu and relatives type::
sudo apt-get install swig
while on Mac OS it's::
brew install swig
Install FFC_ and all dependencies via pip::
sudo pip install \
six \
sympy \
git+https://bitbucket.org/mapdes/ffc.git#egg=ffc \
git+https://bitbucket.org/mapdes/ufl.git#egg=ufl \
git+https://bitbucket.org/mapdes/fiat.git#egg=fiat \
git+https://bitbucket.org/fenics-project/instant.git#egg=instant
These dependencies are regularly updated. If you already have the packages
installed and want to upgrade to the latest versions, do the following::
sudo pip install -U --no-deps ...
To install for your user only, which does not require sudo permissions,
modify the pip invocation for either case above as follows::
pip install --user ...
Potential installation errors on Mac OS
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
The installation of FFC_ requires a C++11 compatible compiler and
standard library, some Mac OS systems (for example OS X "Lion")
supply the former, but not the latter. Should you obtain errors
installing FFC_ of the following form:
.. code-block:: c
ufc/ufc_wrap.cpp:3841:8: error: no member named 'shared_ptr' in namespace 'std'
std::shared_ptr< ufc::function > tempshared1 ;
It's possible that you just need to tell the compiler to pick the
correct standard library. To do so, try running with
``CXXFLAGS='-stdlib=libc++'`` when installing::
sudo CXXFLAGS='-stdlib=libc++' pip install -U --no-deps ...
Visualisation software
----------------------
Firedrake can output data in VTK format, suitable for viewing in
Paraview_. On Ubuntu and similar systems, you can obtain Paraview by
installing the ``paraview`` package. On Mac OS, the easiest approach
is to download a binary from the `paraview website <Paraview_>`_.
Firedrake
---------
In addition to PyOP2, you will need to install Firedrake. There are two
routes, depending on whether you intend to contribute to Firedrake
development.
In order to have the form assembly cache operate in the most automatic
fashion possible, you are also advised to install psutil::
sudo pip install psutil
or (to install for your user only)::
pip install --user psutil
Pip instructions for users
~~~~~~~~~~~~~~~~~~~~~~~~~~
If you only wish to use Firedrake, and will not be contributing to
development at all, you can install Firedrake using pip::
sudo pip install git+https://github.com/firedrakeproject/firedrake.git
or (to install for your user only)::
pip install --user git+https://github.com/firedrakeproject/firedrake.git
You're now ready to go. You might like to start with the tutorial
examples on the :doc:`documentation page <documentation>`.
Git instructions for developers
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Next, obtain the Firedrake source from GitHub_ ::
git clone https://github.com/firedrakeproject/firedrake.git
You will also need to point Python at the right directories. You might
want to consider setting this permanently in your
``.bashrc`` or similar::
cd firedrake
export PYTHONPATH=$PWD:$PYTHONPATH
From the Firedrake directory build the relevant modules::
make
Cleaning disk caches after upgrade
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
After upgrading, you may need to clear any disk caches that Firedrake
maintains to ensure that your problem does not pick up any out of date
compiled modules. This can be carried out by executing the
``firedrake-clean`` script. If you carried out a sudo install of
Firedrake using pip, ``firedrake-clean`` should be in your ``PATH``
and so you should just be able to execute it. If you carried out a
user install using pip, you will need to add ``$HOME/.local/bin`` to
your ``PATH`` ::
export PATH=$HOME/.local/bin:$PATH
If you are using a checkout of Firedrake, ``firedrake-clean`` lives in
the ``scripts`` subdirectory.
Additional dependencies for developers
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
If you plan to develop Firedrake then you will require a few more
packages. Building the documention requires Sphinx_
(including the youtube plugin) and wget_. For example on Ubuntu-like
Linux systems::
sudo apt-get install python-sphinx wget
sudo pip install sphinxcontrib.youtube
and on Mac OS::
brew install wget
sudo pip install sphinx sphinxcontrib.youtube
note that the Sphinx in Homebrew is not the python documentation tool!
.. _PyOP2: http://op2.github.io/PyOP2
.. _FFC: https://bitbucket.org/mapdes/ffc
.. _FIAT: https://bitbucket.org/mapdes/fiat
.. _UFL: https://bitbucket.org/mapdes/ufl
.. _Instant: https://bitbucket.org/fenics-project/instant
.. _GitHub: https://github.com/firedrakeproject/firedrake
.. _Paraview: http://www.paraview.org
.. _Sphinx: http://www.sphinx-doc.org
.. _wget: http://www.gnu.org/software/wget/
.. _Swig: http://www.swig.org/
<file_sep>/firedrake/function.py
import numpy as np
import FIAT
import ufl
import pyop2.coffee.ast_base as ast
from pyop2 import op2
import assemble_expressions
import expression as expression_t
import functionspace
import projection
import utils
import vector
__all__ = ['Function']
valuetype = np.float64
class Function(ufl.Coefficient):
"""A :class:`Function` represents a discretised field over the
domain defined by the underlying :class:`.Mesh`. Functions are
represented as sums of basis functions:
.. math::
f = \\sum_i f_i \phi_i(x)
The :class:`Function` class provides storage for the coefficients
:math:`f_i` and associates them with a :class:`FunctionSpace` object
which provides the basis functions :math:`\\phi_i(x)`.
Note that the coefficients are always scalars: if the
:class:`Function` is vector-valued then this is specified in
the :class:`FunctionSpace`.
"""
def __init__(self, function_space, val=None, name=None, dtype=valuetype):
"""
:param function_space: the :class:`.FunctionSpace`, :class:`.VectorFunctionSpace`
or :class:`.MixedFunctionSpace` on which to build this :class:`Function`.
Alternatively, another :class:`Function` may be passed here and its function space
will be used to build this :class:`Function`.
:param val: NumPy array-like (or :class:`op2.Dat`) providing initial values (optional).
:param name: user-defined name for this :class:`Function` (optional).
:param dtype: optional data type for this :class:`Function`
(defaults to :data:`valuetype`).
"""
if isinstance(function_space, Function):
self._function_space = function_space._function_space
elif isinstance(function_space, functionspace.FunctionSpaceBase):
self._function_space = function_space
else:
raise NotImplementedError("Can't make a Function defined on a "
+ str(type(function_space)))
ufl.Coefficient.__init__(self, self._function_space.ufl_element())
self._label = "a function"
self.uid = utils._new_uid()
self._name = name or 'function_%d' % self.uid
if isinstance(val, op2.Dat):
self.dat = val
else:
self.dat = self._function_space.make_dat(val, dtype,
self._name, uid=self.uid)
self._repr = None
self._split = None
if isinstance(function_space, Function):
self.assign(function_space)
def split(self):
"""Extract any sub :class:`Function`\s defined on the component spaces
of this this :class:`Function`'s :class:`FunctionSpace`."""
if self._split is None:
self._split = tuple(Function(fs, dat) for fs, dat in zip(self._function_space, self.dat))
return self._split
def sub(self, i):
"""Extract the ith sub :class:`Function` of this :class:`Function`.
:arg i: the index to extract
See also :meth:`split`"""
return self.split()[i]
@property
def cell_set(self):
"""The :class:`pyop2.Set` of cells for the mesh on which this
:class:`Function` is defined."""
return self._function_space._mesh.cell_set
@property
def node_set(self):
"""A :class:`pyop2.Set` containing the nodes of this
:class:`Function`. One or (for
:class:`.VectorFunctionSpace`\s) more degrees of freedom are
stored at each node.
"""
return self._function_space.node_set
@property
def dof_dset(self):
"""A :class:`pyop2.DataSet` containing the degrees of freedom of
this :class:`Function`."""
return self._function_space.dof_dset
def cell_node_map(self, bcs=None):
return self._function_space.cell_node_map(bcs)
cell_node_map.__doc__ = functionspace.FunctionSpace.cell_node_map.__doc__
def interior_facet_node_map(self, bcs=None):
return self._function_space.interior_facet_node_map(bcs)
interior_facet_node_map.__doc__ = functionspace.FunctionSpace.interior_facet_node_map.__doc__
def exterior_facet_node_map(self, bcs=None):
return self._function_space.exterior_facet_node_map(bcs)
exterior_facet_node_map.__doc__ = functionspace.FunctionSpace.exterior_facet_node_map.__doc__
def project(self, b, *args, **kwargs):
"""Project ``b`` onto ``self``. ``b`` must be a :class:`Function` or an
:class:`Expression`.
This is equivalent to ``project(b, self)``.
Any of the additional arguments to :func:`~firedrake.projection.project`
may also be passed, and they will have their usual effect.
"""
return projection.project(b, self, *args, **kwargs)
def vector(self):
"""Return a :class:`.Vector` wrapping the data in this :class:`Function`"""
return vector.Vector(self.dat)
def function_space(self):
"""Return the :class:`.FunctionSpace`, :class:`.VectorFunctionSpace`
or :class:`.MixedFunctionSpace` on which this :class:`Function` is defined."""
return self._function_space
def name(self):
"""Return the name of this :class:`Function`"""
return self._name
def label(self):
"""Return the label (a description) of this :class:`Function`"""
return self._label
def rename(self, name=None, label=None):
"""Set the name and or label of this :class:`Function`
:arg name: The new name of the `Function` (if not `None`)
:arg label: The new label for the `Function` (if not `None`)
"""
if name is not None:
self._name = name
if label is not None:
self._label = label
def __str__(self):
if self._name is not None:
return self._name
else:
return super(Function, self).__str__()
def interpolate(self, expression, subset=None):
"""Interpolate an expression onto this :class:`Function`.
:param expression: :class:`.Expression` to interpolate
:returns: this :class:`Function` object"""
# Make sure we have an expression of the right length i.e. a value for
# each component in the value shape of each function space
dims = [np.prod(fs.ufl_element().value_shape(), dtype=int)
for fs in self.function_space()]
if np.prod(expression.value_shape(), dtype=int) != sum(dims):
raise RuntimeError('Expression of length %d required, got length %d'
% (sum(dims), np.prod(expression.value_shape(), dtype=int)))
if expression.code:
# Slice the expression and pass in the right number of values for
# each component function space of this function
d = 0
for fs, dat, dim in zip(self.function_space(), self.dat, dims):
idx = d if fs.rank == 0 else slice(d, d+dim)
self._interpolate(fs, dat,
expression_t.Expression(expression.code[idx],
**expression._kwargs),
subset)
d += dim
else:
if isinstance(fs, functionspace.MixedFunctionSpace):
raise NotImplementedError(
"Python expressions for mixed functions are not yet supported.")
self._interpolate(fs, self.dat, expression, subset)
return self
def _interpolate(self, fs, dat, expression, subset):
"""Interpolate expression onto a :class:`FunctionSpace`.
:param fs: :class:`FunctionSpace`
:param dat: :class:`pyop2.Dat`
:param expression: :class:`.Expression`
"""
to_element = fs.fiat_element
to_pts = []
# TODO very soon: look at the mapping associated with the UFL element;
# this needs to be "identity" (updated from "affine")
if to_element.mapping()[0] != "affine":
raise NotImplementedError("Can only interpolate onto elements \
with affine mapping. Try projecting instead")
for dual in to_element.dual_basis():
if not isinstance(dual, FIAT.functional.PointEvaluation):
raise NotImplementedError("Can only interpolate onto point \
evaluation operators. Try projecting instead")
to_pts.append(dual.pt_dict.keys()[0])
if expression.rank() != len(fs.ufl_element().value_shape()):
raise RuntimeError('Rank mismatch: Expression rank %d, FunctionSpace rank %d'
% (expression.rank(), len(fs.ufl_element().value_shape())))
if expression.value_shape() != fs.ufl_element().value_shape():
raise RuntimeError('Shape mismatch: Expression shape %r, FunctionSpace shape %r'
% (expression.value_shape(), fs.ufl_element().value_shape()))
coords = fs.mesh().coordinates
if expression.code:
kernel = self._interpolate_c_kernel(expression,
to_pts, to_element, fs, coords)
args = [kernel, subset or self.cell_set,
dat(op2.WRITE, fs.cell_node_map()[op2.i[0]]),
coords.dat(op2.READ, coords.cell_node_map())]
elif hasattr(expression, "eval"):
kernel = self._interpolate_python_kernel(expression,
to_pts, to_element, fs, coords)
args = [kernel, subset or self.cell_set,
dat(op2.WRITE, fs.cell_node_map()),
coords.dat(op2.READ, coords.cell_node_map())]
else:
raise RuntimeError(
"Attempting to evaluate an Expression which has no value.")
for _, arg in expression._user_args:
args.append(arg(op2.READ))
op2.par_loop(*args)
def _interpolate_python_kernel(self, expression, to_pts, to_element, fs, coords):
"""Produce a :class:`PyOP2.Kernel` wrapping the eval method on the
function provided."""
coords_space = coords.function_space()
coords_element = coords_space.fiat_element
X_remap = coords_element.tabulate(0, to_pts).values()[0]
# The par_loop will just pass us arguments, since it doesn't
# know about keyword args at all so unpack into a dict that we
# can pass to the user's eval method.
def kernel(output, x, *args):
kwargs = {}
for (slot, _), arg in zip(expression._user_args, args):
kwargs[slot] = arg
X = np.dot(X_remap.T, x)
for i in range(len(output)):
# Pass a slice for the scalar case but just the
# current vector in the VFS case. This ensures the
# eval method has a Dolfin compatible API.
expression.eval(output[i:i+1, ...] if np.rank(output) == 1 else output[i, ...],
X[i:i+1, ...] if np.rank(X) == 1 else X[i, ...], **kwargs)
return kernel
def _interpolate_c_kernel(self, expression, to_pts, to_element, fs, coords):
"""Produce a :class:`PyOP2.Kernel` from the c expression provided."""
coords_space = coords.function_space()
coords_element = coords_space.fiat_element
names = {v[0] for v in expression._user_args}
X = coords_element.tabulate(0, to_pts).values()[0]
# Produce C array notation of X.
X_str = "{{"+"},\n{".join([",".join(map(str, x)) for x in X.T])+"}}"
A = utils.unique_name("A", names)
X = utils.unique_name("X", names)
x_ = utils.unique_name("x_", names)
k = utils.unique_name("k", names)
d = utils.unique_name("d", names)
i_ = utils.unique_name("i", names)
# x is a reserved name.
x = "x"
if "x" in names:
raise ValueError("cannot use 'x' as a user-defined Expression variable")
ass_exp = [ast.Assign(ast.Symbol(A, (k,), ((len(expression.code), i),)),
ast.FlatBlock("%s" % code))
for i, code in enumerate(expression.code)]
vals = {
"X": X,
"x": x,
"x_": x_,
"k": k,
"d": d,
"i": i_,
"x_array": X_str,
"dim": coords_space.dim,
"xndof": coords_element.space_dimension(),
# FS will always either be a functionspace or
# vectorfunctionspace, so just accessing dim here is safe
# (we don't need to go through ufl_element.value_shape())
"nfdof": to_element.space_dimension() * fs.dim,
"ndof": to_element.space_dimension(),
"assign_dim": np.prod(expression.value_shape(), dtype=int)
}
init = ast.FlatBlock("""
const double %(X)s[%(ndof)d][%(xndof)d] = %(x_array)s;
double %(x)s[%(dim)d];
const double pi = 3.141592653589793;
""" % vals)
block = ast.FlatBlock("""
for (unsigned int %(d)s=0; %(d)s < %(dim)d; %(d)s++) {
%(x)s[%(d)s] = 0;
for (unsigned int %(i)s=0; %(i)s < %(xndof)d; %(i)s++) {
%(x)s[%(d)s] += %(X)s[%(k)s][%(i)s] * %(x_)s[%(i)s][%(d)s];
};
};
""" % vals)
loop = ast.c_for(k, "%(ndof)d" % vals, ast.Block([block] + ass_exp,
open_scope=True))
user_args = []
user_init = []
for _, arg in expression._user_args:
if arg.shape == (1, ):
user_args.append(ast.Decl("double *", "%s_" % arg.name))
user_init.append(ast.FlatBlock("const double %s = *%s_;" %
(arg.name, arg.name)))
else:
user_args.append(ast.Decl("double *", arg.name))
kernel_code = ast.FunDecl("void", "expression_kernel",
[ast.Decl("double", ast.Symbol(A, (int("%(nfdof)d" % vals),))),
ast.Decl("double**", x_)] + user_args,
ast.Block(user_init + [init, loop],
open_scope=False))
return op2.Kernel(kernel_code, "expression_kernel")
def assign(self, expr, subset=None):
"""Set the :class:`Function` value to the pointwise value of
expr. expr may only contain :class:`Function`\s on the same
:class:`.FunctionSpace` as the :class:`Function` being assigned to.
Similar functionality is available for the augmented assignment
operators `+=`, `-=`, `*=` and `/=`. For example, if `f` and `g` are
both Functions on the same :class:`FunctionSpace` then::
f += 2 * g
will add twice `g` to `f`.
If present, subset must be an :class:`pyop2.Subset` of
:attr:`node_set`. The expression will then only be assigned
to the nodes on that subset.
"""
if isinstance(expr, Function) and \
expr._function_space == self._function_space:
expr.dat.copy(self.dat, subset=subset)
return self
assemble_expressions.evaluate_expression(
assemble_expressions.Assign(self, expr), subset)
return self
def __iadd__(self, expr):
if np.isscalar(expr):
self.dat += expr
return self
if isinstance(expr, Function) and \
expr._function_space == self._function_space:
self.dat += expr.dat
return self
assemble_expressions.evaluate_expression(
assemble_expressions.IAdd(self, expr))
return self
def __isub__(self, expr):
if np.isscalar(expr):
self.dat -= expr
return self
if isinstance(expr, Function) and \
expr._function_space == self._function_space:
self.dat -= expr.dat
return self
assemble_expressions.evaluate_expression(
assemble_expressions.ISub(self, expr))
return self
def __imul__(self, expr):
if np.isscalar(expr):
self.dat *= expr
return self
if isinstance(expr, Function) and \
expr._function_space == self._function_space:
self.dat *= expr.dat
return self
assemble_expressions.evaluate_expression(
assemble_expressions.IMul(self, expr))
return self
def __idiv__(self, expr):
if np.isscalar(expr):
self.dat /= expr
return self
if isinstance(expr, Function) and \
expr._function_space == self._function_space:
self.dat /= expr.dat
return self
assemble_expressions.evaluate_expression(
assemble_expressions.IDiv(self, expr))
return self
<file_sep>/tests/regression/test_helmholtz_sphere.py
# Solve -laplace u + u = f on the surface of the sphere
# with forcing function xyz, this has exact solution xyz/13
import pytest
import numpy as np
from firedrake import *
def run_helmholtz_sphere(r):
m = UnitIcosahedralSphereMesh(refinement_level=r)
m.init_cell_orientations(Expression(('x[0]', 'x[1]', 'x[2]')))
V = FunctionSpace(m, 'RT', 1)
Q = FunctionSpace(m, 'DG', 0)
W = V*Q
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
f = Function(Q)
f.interpolate(Expression("x[0]*x[1]*x[2]"))
a = (p*q - q*div(u) + inner(v, u) + div(v)*p) * dx
L = f*q*dx
soln = Function(W)
solve(a == L, soln)
_, u = soln.split()
f.interpolate(Expression("x[0]*x[1]*x[2]/13.0"))
return errornorm(f, u, degree_rise=0)
def test_helmholtz_sphere():
errors = [run_helmholtz_sphere(r) for r in range(1, 5)]
errors = np.asarray(errors)
l2conv = np.log2(errors[:-1] / errors[1:])
# Note, due to "magic hybridisation stuff" we expect the numerical
# solution to converge to the projection of the exact solution to
# DG0 at second order (ccotter, pers comm).
assert (l2conv > 1.6).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/firedrake_geometry.h
/* --- Computation of Jacobian matrices --- */
/* Compute Jacobian J for interval embedded in R^1 */
#define compute_jacobian_interval_1d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0];
/* Compute Jacobian J for interval embedded in R^2 */
#define compute_jacobian_interval_2d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[3][0] - vertex_coordinates[2][0];
/* Compute Jacobian J for quad embedded in R^2 */
#define compute_jacobian_quad_2d(J, vertex_coordinates) \
J[0] = 0.5*(vertex_coordinates[2][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[1] = 0.5*(vertex_coordinates[1][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[2][0]); \
J[2] = 0.5*(vertex_coordinates[6][0] + vertex_coordinates[7][0] - vertex_coordinates[4][0] - vertex_coordinates[5][0]); \
J[3] = 0.5*(vertex_coordinates[5][0] + vertex_coordinates[7][0] - vertex_coordinates[4][0] - vertex_coordinates[6][0]);
/* Compute Jacobian J for quad embedded in R^3 */
#define compute_jacobian_quad_3d(J, vertex_coordinates) \
J[0] = 0.5*(vertex_coordinates[2][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[1] = 0.5*(vertex_coordinates[1][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[2][0]); \
J[2] = 0.5*(vertex_coordinates[6][0] + vertex_coordinates[7][0] - vertex_coordinates[4][0] - vertex_coordinates[5][0]); \
J[3] = 0.5*(vertex_coordinates[5][0] + vertex_coordinates[7][0] - vertex_coordinates[4][0] - vertex_coordinates[6][0]); \
J[4] = 0.5*(vertex_coordinates[10][0] + vertex_coordinates[11][0] - vertex_coordinates[8][0] - vertex_coordinates[9][0]); \
J[5] = 0.5*(vertex_coordinates[9][0] + vertex_coordinates[11][0] - vertex_coordinates[8][0] - vertex_coordinates[10][0]);
/* Compute Jacobian J for interval embedded in R^3 */
#define compute_jacobian_interval_3d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[3][0] - vertex_coordinates[2][0]; \
J[2] = vertex_coordinates[5][0] - vertex_coordinates[4][0];
/* Compute Jacobian J for triangle embedded in R^2 */
#define compute_jacobian_triangle_2d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[2][0] - vertex_coordinates[0][0]; \
J[2] = vertex_coordinates[4][0] - vertex_coordinates[3][0]; \
J[3] = vertex_coordinates[5][0] - vertex_coordinates[3][0];
/* Compute Jacobian J for triangle embedded in R^3 */
#define compute_jacobian_triangle_3d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[2][0] - vertex_coordinates[0][0]; \
J[2] = vertex_coordinates[4][0] - vertex_coordinates[3][0]; \
J[3] = vertex_coordinates[5][0] - vertex_coordinates[3][0]; \
J[4] = vertex_coordinates[7][0] - vertex_coordinates[6][0]; \
J[5] = vertex_coordinates[8][0] - vertex_coordinates[6][0];
/* Compute Jacobian J for tetrahedron embedded in R^3 */
#define compute_jacobian_tetrahedron_3d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1] [0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[2] [0] - vertex_coordinates[0][0]; \
J[2] = vertex_coordinates[3] [0] - vertex_coordinates[0][0]; \
J[3] = vertex_coordinates[5] [0] - vertex_coordinates[4][0]; \
J[4] = vertex_coordinates[6] [0] - vertex_coordinates[4][0]; \
J[5] = vertex_coordinates[7] [0] - vertex_coordinates[4][0]; \
J[6] = vertex_coordinates[9] [0] - vertex_coordinates[8][0]; \
J[7] = vertex_coordinates[10][0] - vertex_coordinates[8][0]; \
J[8] = vertex_coordinates[11][0] - vertex_coordinates[8][0];
/* Compute Jacobian J for tensor product prism embedded in R^3 */
/* Explanation: the CG1 x CG1 basis functions are, in order,
(1-X-Y)(1-Z), (1-X-Y)Z, X(1-Z), XZ, Y(1-Z), YZ. Each row of the
Jacobian is the derivatives of these w.r.t. X, Y and Z in turn,
evaluated at the midpoint (1/3, 1/3, 1/2). This gives the
coefficients below.*/
#define compute_jacobian_prism_3d(J, vertex_coordinates) \
J[0] = 0.5*(vertex_coordinates[2][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[1] = 0.5*(vertex_coordinates[4][0] + vertex_coordinates[5][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[2] = (vertex_coordinates[1][0] + vertex_coordinates[3][0] + vertex_coordinates[5][0] - vertex_coordinates[0][0] - vertex_coordinates[2][0] - vertex_coordinates[4][0])/3.0; \
J[3] = 0.5*(vertex_coordinates[8][0] + vertex_coordinates[9][0] - vertex_coordinates[6][0] - vertex_coordinates[7][0]); \
J[4] = 0.5*(vertex_coordinates[10][0] + vertex_coordinates[11][0] - vertex_coordinates[6][0] - vertex_coordinates[7][0]); \
J[5] = (vertex_coordinates[7][0] + vertex_coordinates[9][0] + vertex_coordinates[11][0] - vertex_coordinates[6][0] - vertex_coordinates[8][0] - vertex_coordinates[10][0])/3.0; \
J[6] = 0.5*(vertex_coordinates[14][0] + vertex_coordinates[15][0] - vertex_coordinates[12][0] - vertex_coordinates[13][0]); \
J[7] = 0.5*(vertex_coordinates[16][0] + vertex_coordinates[17][0] - vertex_coordinates[12][0] - vertex_coordinates[13][0]); \
J[8] = (vertex_coordinates[13][0] + vertex_coordinates[15][0] + vertex_coordinates[17][0] - vertex_coordinates[12][0] - vertex_coordinates[14][0] - vertex_coordinates[16][0])/3.0;
/* Jacobians for interior facets of different sorts */
/* Compute Jacobian J for interval embedded in R^1 */
#define compute_jacobian_interval_int_1d compute_jacobian_interval_1d
/* Compute Jacobian J for interval embedded in R^2 */
#define compute_jacobian_interval_int_2d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[5][0] - vertex_coordinates[4][0];
/* Compute Jacobian J for quad embedded in R^2 */
#define compute_jacobian_quad_int_2d(J, vertex_coordinates) \
J[0] = 0.5*(vertex_coordinates[2][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[1] = 0.5*(vertex_coordinates[1][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[2][0]); \
J[2] = 0.5*(vertex_coordinates[10][0] + vertex_coordinates[11][0] - vertex_coordinates[8][0] - vertex_coordinates[9][0]); \
J[3] = 0.5*(vertex_coordinates[9][0] + vertex_coordinates[11][0] - vertex_coordinates[8][0] - vertex_coordinates[10][0]);
/* Compute Jacobian J for quad embedded in R^3 */
#define compute_jacobian_quad_int_3d(J, vertex_coordinates) \
J[0] = 0.5*(vertex_coordinates[2][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[1] = 0.5*(vertex_coordinates[1][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[2][0]); \
J[2] = 0.5*(vertex_coordinates[10][0] + vertex_coordinates[11][0] - vertex_coordinates[8][0] - vertex_coordinates[9][0]); \
J[3] = 0.5*(vertex_coordinates[9][0] + vertex_coordinates[11][0] - vertex_coordinates[8][0] - vertex_coordinates[10][0]); \
J[4] = 0.5*(vertex_coordinates[18][0] + vertex_coordinates[19][0] - vertex_coordinates[16][0] - vertex_coordinates[17][0]); \
J[5] = 0.5*(vertex_coordinates[17][0] + vertex_coordinates[19][0] - vertex_coordinates[16][0] - vertex_coordinates[18][0]);
/* Compute Jacobian J for interval embedded in R^3 */
#define compute_jacobian_interval_int_3d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[5][0] - vertex_coordinates[4][0]; \
J[2] = vertex_coordinates[9][0] - vertex_coordinates[8][0];
/* Compute Jacobian J for triangle embedded in R^2 */
#define compute_jacobian_triangle_int_2d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1][0] - vertex_coordinates[0][0]; \
J[1] = vertex_coordinates[2][0] - vertex_coordinates[0][0]; \
J[2] = vertex_coordinates[7][0] - vertex_coordinates[6][0]; \
J[3] = vertex_coordinates[8][0] - vertex_coordinates[6][0];
/* Compute Jacobian J for triangle embedded in R^3 */
#define compute_jacobian_triangle_int_3d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1] [0] - vertex_coordinates[0] [0]; \
J[1] = vertex_coordinates[2] [0] - vertex_coordinates[0] [0]; \
J[2] = vertex_coordinates[7] [0] - vertex_coordinates[6] [0]; \
J[3] = vertex_coordinates[8] [0] - vertex_coordinates[6] [0]; \
J[4] = vertex_coordinates[13][0] - vertex_coordinates[12][0]; \
J[5] = vertex_coordinates[14][0] - vertex_coordinates[12][0];
/* Compute Jacobian J for tetrahedron embedded in R^3 */
#define compute_jacobian_tetrahedron_int_3d(J, vertex_coordinates) \
J[0] = vertex_coordinates[1] [0] - vertex_coordinates[0] [0]; \
J[1] = vertex_coordinates[2] [0] - vertex_coordinates[0] [0]; \
J[2] = vertex_coordinates[3] [0] - vertex_coordinates[0] [0]; \
J[3] = vertex_coordinates[9] [0] - vertex_coordinates[8] [0]; \
J[4] = vertex_coordinates[10][0] - vertex_coordinates[8] [0]; \
J[5] = vertex_coordinates[11][0] - vertex_coordinates[8] [0]; \
J[6] = vertex_coordinates[17][0] - vertex_coordinates[16][0]; \
J[7] = vertex_coordinates[18][0] - vertex_coordinates[16][0]; \
J[8] = vertex_coordinates[19][0] - vertex_coordinates[16][0];
/* Compute Jacobian J for tensor product prism embedded in R^3 */
#define compute_jacobian_prism_int_3d(J, vertex_coordinates) \
J[0] = 0.5*(vertex_coordinates[2][0] + vertex_coordinates[3][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[1] = 0.5*(vertex_coordinates[4][0] + vertex_coordinates[5][0] - vertex_coordinates[0][0] - vertex_coordinates[1][0]); \
J[2] = (vertex_coordinates[1][0] + vertex_coordinates[3][0] + vertex_coordinates[5][0] - vertex_coordinates[0][0] - vertex_coordinates[2][0] - vertex_coordinates[4][0])/3.0; \
J[3] = 0.5*(vertex_coordinates[14][0] + vertex_coordinates[15][0] - vertex_coordinates[12][0] - vertex_coordinates[13][0]); \
J[4] = 0.5*(vertex_coordinates[16][0] + vertex_coordinates[17][0] - vertex_coordinates[12][0] - vertex_coordinates[13][0]); \
J[5] = (vertex_coordinates[13][0] + vertex_coordinates[15][0] + vertex_coordinates[17][0] - vertex_coordinates[12][0] - vertex_coordinates[14][0] - vertex_coordinates[16][0])/3.0; \
J[6] = 0.5*(vertex_coordinates[26][0] + vertex_coordinates[27][0] - vertex_coordinates[24][0] - vertex_coordinates[25][0]); \
J[7] = 0.5*(vertex_coordinates[28][0] + vertex_coordinates[29][0] - vertex_coordinates[24][0] - vertex_coordinates[25][0]); \
J[8] = (vertex_coordinates[25][0] + vertex_coordinates[27][0] + vertex_coordinates[29][0] - vertex_coordinates[24][0] - vertex_coordinates[26][0] - vertex_coordinates[28][0])/3.0;
/* --- Computation of Jacobian inverses --- */
/* Compute Jacobian inverse K for interval embedded in R^1 */
#define compute_jacobian_inverse_interval_1d(K, det, J) \
det = J[0]; \
K[0] = 1.0 / det;
/* Compute Jacobian (pseudo)inverse K for interval embedded in R^2 */
#define compute_jacobian_inverse_interval_2d(K, det, J) \
do { const double det2 = J[0]*J[0] + J[1]*J[1]; \
det = sqrt(det2); \
K[0] = J[0] / det2; \
K[1] = J[1] / det2; } while (0)
/* Compute Jacobian (pseudo)inverse K for interval embedded in R^3 */
#define compute_jacobian_inverse_interval_3d(K, det, J) \
do { const double det2 = J[0]*J[0] + J[1]*J[1] + J[2]*J[2]; \
det = sqrt(det2); \
K[0] = J[0] / det2; \
K[1] = J[1] / det2; \
K[2] = J[2] / det2; } while (0)
/* Compute Jacobian inverse K for triangle embedded in R^2 */
#define compute_jacobian_inverse_triangle_2d(K, det, J) \
det = J[0]*J[3] - J[1]*J[2]; \
K[0] = J[3] / det; \
K[1] = -J[1] / det; \
K[2] = -J[2] / det; \
K[3] = J[0] / det;
/* Compute Jacobian (pseudo)inverse K for triangle embedded in R^3 */
#define compute_jacobian_inverse_triangle_3d(K, det, J) \
do { const double d_0 = J[2]*J[5] - J[4]*J[3]; \
const double d_1 = J[4]*J[1] - J[0]*J[5]; \
const double d_2 = J[0]*J[3] - J[2]*J[1]; \
const double c_0 = J[0]*J[0] + J[2]*J[2] + J[4]*J[4]; \
const double c_1 = J[1]*J[1] + J[3]*J[3] + J[5]*J[5]; \
const double c_2 = J[0]*J[1] + J[2]*J[3] + J[4]*J[5]; \
const double den = c_0*c_1 - c_2*c_2; \
const double det2 = d_0*d_0 + d_1*d_1 + d_2*d_2; \
det = sqrt(det2); \
K[0] = (J[0]*c_1 - J[1]*c_2) / den; \
K[1] = (J[2]*c_1 - J[3]*c_2) / den; \
K[2] = (J[4]*c_1 - J[5]*c_2) / den; \
K[3] = (J[1]*c_0 - J[0]*c_2) / den; \
K[4] = (J[3]*c_0 - J[2]*c_2) / den; \
K[5] = (J[5]*c_0 - J[4]*c_2) / den; } while (0)
/* Compute Jacobian (pseudo)inverse K for quad embedded in R^2 */
#define compute_jacobian_inverse_quad_2d compute_jacobian_inverse_triangle_2d
/* Compute Jacobian (pseudo)inverse K for quad embedded in R^3 */
#define compute_jacobian_inverse_quad_3d compute_jacobian_inverse_triangle_3d
/* Compute Jacobian inverse K for tetrahedron embedded in R^3 */
#define compute_jacobian_inverse_tetrahedron_3d(K, det, J) \
do { const double d_00 = J[4]*J[8] - J[5]*J[7]; \
const double d_01 = J[5]*J[6] - J[3]*J[8]; \
const double d_02 = J[3]*J[7] - J[4]*J[6]; \
const double d_10 = J[2]*J[7] - J[1]*J[8]; \
const double d_11 = J[0]*J[8] - J[2]*J[6]; \
const double d_12 = J[1]*J[6] - J[0]*J[7]; \
const double d_20 = J[1]*J[5] - J[2]*J[4]; \
const double d_21 = J[2]*J[3] - J[0]*J[5]; \
const double d_22 = J[0]*J[4] - J[1]*J[3]; \
det = J[0]*d_00 + J[3]*d_10 + J[6]*d_20; \
K[0] = d_00 / det; \
K[1] = d_10 / det; \
K[2] = d_20 / det; \
K[3] = d_01 / det; \
K[4] = d_11 / det; \
K[5] = d_21 / det; \
K[6] = d_02 / det; \
K[7] = d_12 / det; \
K[8] = d_22 / det; } while(0)
/* Compute Jacobian inverse K for tensor product prism embedded in R^3 - identical to t et */
#define compute_jacobian_inverse_prism_3d(K, det, J) \
do { const double d_00 = J[4]*J[8] - J[5]*J[7]; \
const double d_01 = J[5]*J[6] - J[3]*J[8]; \
const double d_02 = J[3]*J[7] - J[4]*J[6]; \
const double d_10 = J[2]*J[7] - J[1]*J[8]; \
const double d_11 = J[0]*J[8] - J[2]*J[6]; \
const double d_12 = J[1]*J[6] - J[0]*J[7]; \
const double d_20 = J[1]*J[5] - J[2]*J[4]; \
const double d_21 = J[2]*J[3] - J[0]*J[5]; \
const double d_22 = J[0]*J[4] - J[1]*J[3]; \
det = J[0]*d_00 + J[3]*d_10 + J[6]*d_20; \
K[0] = d_00 / det; \
K[1] = d_10 / det; \
K[2] = d_20 / det; \
K[3] = d_01 / det; \
K[4] = d_11 / det; \
K[5] = d_21 / det; \
K[6] = d_02 / det; \
K[7] = d_12 / det; \
K[8] = d_22 / det; } while (0)
/* --- Compute facet edge lengths --- */
#define compute_facet_edge_length_tetrahedron_3d(facet, vertex_coordinates) \
const unsigned int tetrahedron_facet_edge_vertices[4][3][2] = { \
{{2, 3}, {1, 3}, {1, 2}}, \
{{2, 3}, {0, 3}, {0, 2}}, \
{{1, 3}, {0, 3}, {0, 1}}, \
{{1, 2}, {0, 2}, {0, 1}}, \
}; \
double edge_lengths_sqr[3]; \
for (unsigned int edge = 0; edge < 3; ++edge) \
{ \
const unsigned int vertex0 = tetrahedron_facet_edge_vertices[facet][edge][0]; \
const unsigned int vertex1 = tetrahedron_facet_edge_vertices[facet][edge][1]; \
edge_lengths_sqr[edge] = (vertex_coordinates[vertex1 + 0][0] - vertex_coordinates[vertex0 + 0][0])*(vertex_coordinates[vertex1 + 0][0] - vertex_coordinates[vertex0 + 0][0]) \
+ (vertex_coordinates[vertex1 + 4][0] - vertex_coordinates[vertex0 + 4][0])*(vertex_coordinates[vertex1 + 4][0] - vertex_coordinates[vertex0 + 4][0]) \
+ (vertex_coordinates[vertex1 + 8][0] - vertex_coordinates[vertex0 + 8][0])*(vertex_coordinates[vertex1 + 8][0] - vertex_coordinates[vertex0 + 8][0]); \
}
/* Compute min edge length in facet of tetrahedron embedded in R^3 */
#define compute_min_facet_edge_length_tetrahedron_3d(min_edge_length, facet, vertex_coordinates) \
compute_facet_edge_length_tetrahedron_3d(facet, vertex_coordinates); \
min_edge_length = sqrt(fmin(fmin(edge_lengths_sqr[1], edge_lengths_sqr[1]), edge_lengths_sqr[2]));
/* Compute max edge length in facet of tetrahedron embedded in R^3 */
/*
* FIXME: we can't call compute_facet_edge_length_tetrahedron_3d again, so we
* rely on the fact that max is always computed after min
*/
#define compute_max_facet_edge_length_tetrahedron_3d(max_edge_length, facet, vertex_coordinates) \
max_edge_length = sqrt(fmax(fmax(edge_lengths_sqr[1], edge_lengths_sqr[1]), edge_lengths_sqr[2]));
<file_sep>/tests/common.py
import pytest
from firedrake import *
longtest = pytest.mark.skipif("config.option.short")
@pytest.fixture(scope='module')
def mesh():
return UnitSquareMesh(5, 5)
def extmesh(nx, ny, nz):
return ExtrudedMesh(UnitSquareMesh(nx, ny), nz, layer_height=1.0/nz)
def extmesh_2D(nx, ny):
return ExtrudedMesh(UnitIntervalMesh(nx), ny, layer_height=1.0/ny)
@pytest.fixture(scope='module')
def cg1(mesh):
return FunctionSpace(mesh, "CG", 1)
@pytest.fixture(scope='module')
def cg2(mesh):
return FunctionSpace(mesh, "CG", 2)
@pytest.fixture(scope='module')
def dg0(mesh):
return FunctionSpace(mesh, "DG", 0)
@pytest.fixture(scope='module')
def dg1(mesh):
return FunctionSpace(mesh, "DG", 1)
@pytest.fixture(scope='module')
def vcg1(mesh):
return VectorFunctionSpace(mesh, "CG", 1)
@pytest.fixture(scope='module')
def cg1cg1(cg1):
return cg1 * cg1
@pytest.fixture(scope='module')
def cg1dg0(cg1, dg0):
return cg1 * dg0
@pytest.fixture(scope='module')
def cg2dg1(cg2, dg1):
return cg2 * dg1
@pytest.fixture(scope='module')
def cg1vcg1(mesh, cg1, vcg1):
return cg1 * vcg1
<file_sep>/tests/regression/test_solving_interface.py
import pytest
from firedrake import *
from numpy.linalg import norm as np_norm
import gc
def howmany(cls):
return len([x for x in gc.get_objects() if isinstance(x, cls)])
@pytest.fixture
def a_L_out():
mesh = UnitCubeMesh(1, 1, 1)
fs = FunctionSpace(mesh, 'CG', 1)
f = Function(fs)
out = Function(fs)
u = TrialFunction(fs)
v = TestFunction(fs)
return u*v*dx, f*v*dx, out
def test_linear_solver_api(a_L_out):
a, L, out = a_L_out
p = LinearVariationalProblem(a, L, out)
solver = LinearVariationalSolver(p, solver_parameters={'ksp_type': 'cg'})
assert solver.parameters['snes_type'] == 'ksponly'
assert solver.parameters['ksp_rtol'] == 1e-7
assert solver.snes.getType() == solver.snes.Type.KSPONLY
assert solver.snes.getKSP().getType() == solver.snes.getKSP().Type.CG
rtol, _, _, _ = solver.snes.getKSP().getTolerances()
assert rtol == solver.parameters['ksp_rtol']
solver.parameters['ksp_type'] = 'gmres'
solver.parameters['ksp_rtol'] = 1e-8
solver.solve()
assert solver.snes.getKSP().getType() == solver.snes.getKSP().Type.GMRES
assert solver.parameters['ksp_rtol'] == 1e-8
rtol, _, _, _ = solver.snes.getKSP().getTolerances()
assert rtol == solver.parameters['ksp_rtol']
def test_petsc_options_cleared(a_L_out):
a, L, out = a_L_out
from petsc4py import PETSc
opts = PETSc.Options()
original = {}
original.update(opts.getAll())
solve(a == L, out, solver_parameters={'foo': 'bar'})
assert original == opts.getAll()
def test_linear_solver_gced(a_L_out):
a, L, out = a_L_out
gc.collect()
before = howmany(LinearVariationalSolver)
solve(a == L, out)
out.dat.data_ro # force evaluation
gc.collect()
after = howmany(LinearVariationalSolver)
assert before == after
def test_assembled_solver_gced(a_L_out):
a, L, out = a_L_out
A = assemble(a)
b = assemble(L)
gc.collect()
before = howmany(LinearSolver)
solve(A, out, b)
out.dat.data_ro
gc.collect()
after = howmany(LinearSolver)
assert before == after
def test_nonlinear_solver_gced(a_L_out):
a, L, out = a_L_out
gc.collect()
before = howmany(NonlinearVariationalSolver)
F = action(a, out) - L
solve(F == 0, out)
out.dat.data_ro # force evaluation
gc.collect()
after = howmany(NonlinearVariationalSolver)
assert before == after
def test_nonlinear_solver_api(a_L_out):
a, L, out = a_L_out
J = a
F = action(a, out) - L
p = NonlinearVariationalProblem(F, out, J=J)
solver = NonlinearVariationalSolver(p, solver_parameters={'snes_type': 'ksponly'})
assert solver.snes.getType() == solver.snes.Type.KSPONLY
rtol, _, _, _ = solver.snes.getTolerances()
assert rtol == 1e-8
solver.parameters['snes_rtol'] = 1e-3
solver.parameters['snes_type'] = 'newtonls'
solver.solve()
assert solver.parameters['snes_rtol'] == 1e-3
assert solver.snes.getType() == solver.snes.Type.NEWTONLS
rtol, _, _, _ = solver.snes.getTolerances()
assert rtol == solver.parameters['snes_rtol']
def test_linear_solves_equivalent():
"""solve(a == L, out) should return the same as solving with the assembled objects.
This relies on two different code paths agreeing on the same set of solver parameters."""
mesh = UnitSquareMesh(50, 50)
V = FunctionSpace(mesh, "CG", 1)
f = Function(V)
f.assign(1)
f.vector()[:] = 1.
t = TestFunction(V)
q = TrialFunction(V)
a = inner(t, q)*dx
L = inner(f, t)*dx
# Solve the system using forms
sol = Function(V)
solve(a == L, sol)
# And again
sol2 = Function(V)
solve(a == L, sol2)
assert np_norm(sol.vector()[:] - sol2.vector()[:]) == 0
# Solve the system using preassembled objects
sol3 = Function(V)
solve(assemble(a), sol3, assemble(L))
assert np_norm(sol.vector()[:] - sol3.vector()[:]) < 5e-14
def test_constant_jacobian_lvs():
mesh = UnitSquareMesh(2, 2)
V = FunctionSpace(mesh, "CG", 1)
u = TrialFunction(V)
v = TestFunction(V)
q = Function(V)
q.assign(1)
a = q*u*v*dx
f = Function(V)
f.assign(1)
L = f*v*dx
out = Function(V)
# Non-constant jacobian set
lvp = LinearVariationalProblem(a, L, out, constant_jacobian=False)
lvs = LinearVariationalSolver(lvp)
lvs.solve()
assert norm(assemble(out - f)) < 1e-7
q.assign(5)
lvs.solve()
assert norm(assemble(out*5 - f)) < 2e-7
q.assign(1)
# This one should fail (because Jac is wrong)
lvp = LinearVariationalProblem(a, L, out, constant_jacobian=True)
lvs = LinearVariationalSolver(lvp)
lvs.solve()
assert norm(assemble(out - f)) < 1e-7
q.assign(5)
lvs.solve()
assert not (norm(assemble(out*5 - f)) < 2e-7)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/ufl_expr.py
import ufl
import ufl.argument
from ufl.assertions import ufl_assert
from ufl.finiteelement import FiniteElementBase
from ufl.split_functions import split
from ufl.algorithms.analysis import extract_arguments
import function
__all__ = ['Argument', 'TestFunction', 'TrialFunction',
'TestFunctions', 'TrialFunctions',
'derivative', 'adjoint',
'CellSize', 'FacetNormal']
class Argument(ufl.argument.Argument):
"""Representation of the argument to a form,"""
def __init__(self, element, function_space, count):
"""
:arg element: the :class:`ufl.element.FiniteElementBase` this
argument corresponds to.
:arg function_space: the :class:`.FunctionSpace` the argument
corresponds to.
:arg count: the number of the argument being constructed.
.. note::
an :class:`Argument` with a count of ``0`` is used as a
:class:`TestFunction`, with a count of ``1`` it is used as
a :class:`TrialFunction`.
"""
super(Argument, self).__init__(element, count)
self._function_space = function_space
@property
def cell_node_map(self):
return self._function_space.cell_node_map
@property
def interior_facet_node_map(self):
return self._function_space.interior_facet_node_map
@property
def exterior_facet_node_map(self):
return self._function_space.exterior_facet_node_map
def function_space(self):
return self._function_space
def make_dat(self):
return self._function_space.make_dat()
def reconstruct(self, element=None, function_space=None, count=None):
if function_space is None or function_space == self._function_space:
function_space = self._function_space
if element is None or element == self._element:
element = self._element
if count is None or count == self._count:
count = self._count
if count is self._count and element is self._element:
return self
ufl_assert(isinstance(element, FiniteElementBase),
"Expecting an element, not %s" % element)
ufl_assert(isinstance(count, int),
"Expecting an int, not %s" % count)
ufl_assert(element.value_shape() == self._element.value_shape(),
"Cannot reconstruct an Argument with a different value shape.")
return Argument(element, function_space, count)
def TestFunction(function_space):
"""Build a test function on the specified function space.
:arg function_space: the :class:`.FunctionSpaceBase` to build the test
function on."""
return Argument(function_space.ufl_element(), function_space, 0)
def TrialFunction(function_space):
"""Build a trial function on the specified function space.
:arg function_space: the :class:`.FunctionSpaceBase` to build the trial
function on."""
return Argument(function_space.ufl_element(), function_space, 1)
def TestFunctions(function_space):
"""Return a tuple of test functions on the specified function space.
:arg function_space: the :class:`.FunctionSpaceBase` to build the test
functions on.
This returns ``len(function_space)`` test functions, which, if the
function space is a :class:`.MixedFunctionSpace`, are indexed
appropriately.
"""
return split(TestFunction(function_space))
def TrialFunctions(function_space):
"""Return a tuple of trial functions on the specified function space.
:arg function_space: the :class:`.FunctionSpaceBase` to build the trial
functions on.
This returns ``len(function_space)`` trial functions, which, if the
function space is a :class:`.MixedFunctionSpace`, are indexed
appropriately.
"""
return split(TrialFunction(function_space))
def derivative(form, u, du=None):
"""Compute the derivative of a form.
Given a form, this computes its linearization with respect to the
provided :class:`.Function`. The resulting form has one
additional :class:`Argument` in the same finite element space as
the Function.
:arg form: a :class:`ufl.Form` to compute the derivative of.
:arg u: a :class:`.Function` to compute the derivative with
respect to.
:arg du: an optional :class:`Argument` to use as the replacement
in the new form (constructed automatically if not provided).
See also :func:`ufl.derivative`.
"""
if du is None:
if isinstance(u, function.Function):
V = u.function_space()
args = form.arguments()
number = max(a.number() for a in args) if args else -1
du = Argument(V.ufl_element(), V, number + 1)
else:
raise RuntimeError("Can't compute derivative for form")
return ufl.derivative(form, u, du)
def adjoint(form, reordered_arguments=None):
"""UFL form operator:
Given a combined bilinear form, compute the adjoint form by
changing the ordering (count) of the test and trial functions.
By default, new Argument objects will be created with
opposite ordering. However, if the adjoint form is to
be added to other forms later, their arguments must match.
In that case, the user must provide a tuple reordered_arguments=(u2,v2).
"""
# ufl.adjoint creates new Arguments if no reordered_arguments is
# given. To avoid that, always pass reordered_arguments with
# firedrake.Argument objects.
if reordered_arguments is None:
v, u = extract_arguments(form)
reordered_arguments = (Argument(u.element(), u.function_space(),
count=v.count()),
Argument(v.element(), v.function_space(),
count=u.count()))
return ufl.adjoint(form, reordered_arguments)
def CellSize(mesh):
"""A symbolic representation of the cell size of a mesh.
:arg mesh: the mesh for which to calculate the cell size.
"""
return 2.0 * ufl.Circumradius(mesh.ufl_domain())
def FacetNormal(mesh):
"""A symbolic representation of the facet normal on a cell in a mesh.
:arg mesh: the mesh over which the normal should be represented.
"""
return ufl.FacetNormal(mesh.ufl_domain())
<file_sep>/tests/regression/test_vector.py
from firedrake import *
import pytest
@pytest.fixture
def f():
mesh = UnitIntervalMesh(2)
V = FunctionSpace(mesh, "CG", 1)
f = Function(V)
f.interpolate(Expression("1"))
return f
def test_vector_array(f):
v = f.vector()
assert (v.array() == 1.0).all()
def test_vector_setitem(f):
v = f.vector()
v[:] = 2.0
assert(v.array() == 2.0).all()
def test_vector_getitem(f):
v = f.vector()
assert v[0] == 1.0
def test_vector_len(f):
v = f.vector()
assert len(v) == f.dof_dset.size
def test_vector_returns_copy(f):
v = f.vector()
a = v.array()
a[:] = 5.0
assert v.array() is not a
assert (v.array() == 1.0).all()
assert (a == 5.0).all()
def test_vector_gather_works(f):
f.interpolate(Expression("2"))
v = f.vector()
gathered = v.gather([0])
assert len(gathered) == 1 and gathered[0] == 2.0
def test_axpy(f):
f.interpolate(Expression("2"))
v = f.vector()
y = Vector(v)
y[:] = 4
v.axpy(3, y)
assert (v.array() == 14.0).all()
def test_scale(f):
f.interpolate(Expression("3"))
v = f.vector()
v._scale(7)
assert (v.array() == 21.0).all()
@pytest.mark.parallel(nprocs=2)
def test_parallel_gather():
from mpi4py import MPI
mesh = UnitSquareMesh(2, 2)
V = FunctionSpace(mesh, "CG", 1)
f = Function(V)
v = f.vector()
rank = MPI.COMM_WORLD.rank
v[:] = rank
assert (f.dat.data_ro[:f.dof_dset.size] == rank).all()
lsum = sum(v.array())
lsum = MPI.COMM_WORLD.allreduce(lsum, op=MPI.SUM)
gathered = v.gather()
gsum = sum(gathered)
assert lsum == gsum
assert len(gathered) == v.size()
gathered = v.gather([0])
assert len(gathered) == 1 and gathered[0] == 0
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/nullspace.py
from numpy import prod
import function
import functionspace
from petsc import PETSc
__all__ = ['VectorSpaceBasis', 'MixedVectorSpaceBasis']
class VectorSpaceBasis(object):
"""Build a basis for a vector space.
You can use this basis to express the null space of a singular operator.
:arg vecs: a list of :class:`.Vector`\s or :class:`.Functions`
spanning the space. Note that these must be orthonormal
:arg constant: does the null space include the constant vector?
If you pass ``constant=True`` you should not also include the
constant vector in the list of ``vecs`` you supply.
.. warning::
The vectors you pass in to this object are *not* copied. You
should therefore not modify them after instantiation since the
basis will then be incorrect.
"""
def __init__(self, vecs=None, constant=False):
if vecs is None and not constant:
raise RuntimeError("Must either provide a list of null space vectors, or constant keyword (or both)")
self._vecs = vecs or []
self._petsc_vecs = []
for v in self._vecs:
with v.dat.vec_ro as v_:
self._petsc_vecs.append(v_)
if not self.is_orthonormal():
raise RuntimeError("Provided vectors must be orthonormal")
self._nullspace = PETSc.NullSpace().create(constant=constant,
vectors=self._petsc_vecs)
self._constant = constant
@property
def nullspace(self):
"""The PETSc NullSpace object for this :class:`.VectorSpaceBasis`"""
return self._nullspace
def orthogonalize(self, b):
"""Orthogonalize ``b`` with respect to this :class:`.VectorSpaceBasis`.
:arg b: a :class:`.Function`
.. note::
Modifies ``b`` in place."""
for v in self._vecs:
dot = b.dat.inner(v.dat)
b.dat -= dot * v.dat
if self._constant:
s = -b.dat.sum() / b.function_space().dof_count
b.dat += s
def is_orthonormal(self):
"""Is this vector space basis orthonormal?"""
for i, iv in enumerate(self._vecs):
for j, jv in enumerate(self._vecs):
dij = 1 if i == j else 0
# scaled by size of function space
if abs(iv.dat.inner(jv.dat) - dij) / prod(iv.function_space().dof_count) > 1e-10:
return False
return True
def is_orthogonal(self):
"""Is this vector space basis orthogonal?"""
for i, iv in enumerate(self._vecs):
for j, jv in enumerate(self._vecs):
if i == j:
continue
# scaled by size of function space
if abs(iv.dat.inner(jv.dat)) / prod(iv.function_space().dof_count) > 1e-10:
return False
return True
def _apply(self, matrix, ises=None):
"""Set this VectorSpaceBasis as a nullspace for a matrix
:arg matrix: a :class:`pyop2.op2.Mat` whose nullspace should be set.
:arg ises: optional list of PETSc IS objects to compose the
nullspace with (ignored)."""
matrix.handle.setNullSpace(self.nullspace)
def __iter__(self):
"""Yield self when iterated over"""
yield self
class MixedVectorSpaceBasis(object):
"""A basis for a mixed vector space
:arg function_space: the :class:`~MixedFunctionSpace` this vector
space is a basis for.
:arg bases: an iterable of bases for the null spaces of the
subspaces in the mixed space.
You can use this to express the null space of a singular operator
on a mixed space. The bases you supply will be used to set null
spaces for each of the diagonal blocks in the operator. If you
only care about the null space on one of the blocks, you can pass
an indexed function space as a placeholder in the positions you
don't care about.
For example, consider a mixed poisson discretisation with pure
Neumann boundary conditions::
V = FunctionSpace(mesh, "BDM", 1)
Q = FunctionSpace(mesh, "DG", 0)
W = V*Q
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
a = (inner(sigma, tau) + div(sigma)*v + div(tau)*u)*dx
The null space of this operator is a constant function in ``Q``.
If we solve the problem with a Schur complement, we only care
about projecting the null space out of the ``QxQ`` block. We can
do this like so ::
nullspace = MixedVectorSpaceBasis(W, [W[0], VectorSpaceBasis(constant=True)])
solve(a == ..., nullspace=nullspace)
"""
def __init__(self, function_space, bases):
self._function_space = function_space
if not all(isinstance(basis, (VectorSpaceBasis, functionspace.IndexedFunctionSpace))
for basis in bases):
raise RuntimeError("MixedVectorSpaceBasis can only contain vector space bases or indexed function spaces")
for i, basis in enumerate(bases):
if isinstance(basis, functionspace.IndexedFunctionSpace):
if i != basis.index:
raise RuntimeError("FunctionSpace with index %d cannot appear at position %d" % (basis.index, i))
if basis._parent != self._function_space:
raise RuntimeError("FunctionSpace with index %d does not have %s as a parent" % (basis.index, self._function_space))
self._bases = bases
self._nullspace = None
def _build_monolithic_basis(self):
"""Build a basis for the complete mixed space.
The monolithic basis is formed by the cartesian product of the
bases forming each sub part.
"""
from itertools import product
bvecs = [[None] for _ in self]
# Get the complete list of basis vectors for each component in
# the mixed basis.
for idx, basis in enumerate(self):
if isinstance(basis, VectorSpaceBasis):
v = []
if basis._constant:
v = [function.Function(self._function_space[idx]).assign(1)]
bvecs[idx] = basis._vecs + v
# Basis for mixed space is cartesian product of all the basis
# vectors we just made.
allbvecs = [x for x in product(*bvecs)]
vecs = [function.Function(self._function_space) for _ in allbvecs]
# Build the functions representing the monolithic basis.
for vidx, bvec in enumerate(allbvecs):
for idx, b in enumerate(bvec):
if b:
vecs[vidx].sub(idx).assign(b)
for v in vecs:
v /= v.dat.norm
self._vecs = vecs
self._petsc_vecs = []
for v in self._vecs:
with v.dat.vec_ro as v_:
self._petsc_vecs.append(v_)
self._nullspace = PETSc.NullSpace().create(constant=False,
vectors=self._petsc_vecs)
def _apply_monolithic(self, matrix):
"""Set this class:`MixedVectorSpaceBasis` as a nullspace for a
matrix.
:arg matrix: a :class:`pyop2.op2.Mat` whose nullspace should
be set.
Note, this only hangs the nullspace on the Mat, you should
normally be using :meth:`_apply` which also hangs the
nullspace on the appropriate fieldsplit ISes for Schur
complements."""
if self._nullspace is None:
self._build_monolithic_basis()
matrix.handle.setNullSpace(self._nullspace)
def _apply(self, matrix, ises):
"""Set this :class:`MixedVectorSpaceBasis` as a nullspace for a matrix
:arg matrix: a :class:`pyop2.op2.Mat` whose nullspace should be set.
:arg ises: optional list of PETSc IS objects to compose the
nullspace with. You must pass these if you intend to
solve a mixed problem with a nullspace using a Schur
complement."""
rows, cols = matrix.sparsity.shape
if rows != cols:
raise RuntimeError("Can only apply nullspace to square operator")
if rows != len(self):
raise RuntimeError("Shape of matrix (%d, %d) does not match size of nullspace %d" %
(rows, cols, len(self)))
# Hang the expanded nullspace on the big matrix
self._apply_monolithic(matrix)
for i, basis in enumerate(self):
if not isinstance(basis, VectorSpaceBasis):
continue
# Compose appropriate nullspace with IS for schur complement
if ises is not None:
is_ = ises[i][1]
is_.compose("nullspace", basis.nullspace)
def __iter__(self):
"""Yield the individual bases making up this MixedVectorSpaceBasis"""
for basis in self._bases:
yield basis
def __len__(self):
"""The number of bases in this MixedVectorSpaceBasis"""
return len(self._bases)
<file_sep>/firedrake/assemble_expressions.py
import ufl
from ufl.algorithms import ReuseTransformer
from ufl.constantvalue import ConstantValue, Zero, IntValue
from ufl.indexing import MultiIndex
from ufl.core.operator import Operator
from ufl.mathfunctions import MathFunction
from ufl.core.ufl_type import ufl_type as orig_ufl_type
from ufl import classes
import pyop2.coffee.ast_base as ast
from pyop2 import op2
import constant
import function
import functionspace
_to_sum = lambda o: ast.Sum(_ast(o[0]), _to_sum(o[1:])) if len(o) > 1 else _ast(o[0])
_to_prod = lambda o: ast.Prod(_ast(o[0]), _to_sum(o[1:])) if len(o) > 1 else _ast(o[0])
_ast_map = {
MathFunction: (lambda e: ast.FunCall(e._name, _ast(e._argument)), None),
ufl.algebra.Sum: (lambda e: _to_sum(e.ufl_operands)),
ufl.algebra.Product: (lambda e: _to_prod(e.ufl_operands)),
ufl.algebra.Division: (lambda e: ast.Div(_ast(e._a), _ast(e._b))),
ufl.algebra.Abs: (lambda e: ast.FunCall("abs", _ast(e._a))),
ufl.constantvalue.ScalarValue: (lambda e: ast.Symbol(e._value)),
ufl.constantvalue.Zero: (lambda e: ast.Symbol(0))
}
def ufl_type(*args, **kwargs):
"""Decorator mimicing :func:`ufl.core.ufl_type.ufl_type`.
Additionally adds the class decorated to the appropriate set of ufl classes."""
def decorator(cls):
orig_ufl_type(*args, **kwargs)(cls)
classes.all_ufl_classes.add(cls)
if cls._ufl_is_abstract_:
classes.abstract_classes.add(cls)
else:
classes.ufl_classes.add(cls)
if cls._ufl_is_terminal_:
classes.terminal_classes.add(cls)
else:
classes.nonterminal_classes.add(cls)
return cls
return decorator
def _ast(expr):
"""Convert expr to a PyOP2 ast."""
try:
return expr.ast
except AttributeError:
for t, f in _ast_map.iteritems():
if isinstance(expr, t):
return f(expr)
raise TypeError("No ast handler for %s" % str(type(expr)))
class DummyFunction(ufl.Coefficient):
"""A dummy object to take the place of a :class:`.Function` in the
expression. This has the sole role of producing the right strings
when the expression is unparsed and when the arguments are
formatted.
"""
def __init__(self, function, argnum, intent=op2.READ):
ufl.Coefficient.__init__(self, function._element)
self.argnum = argnum
self.function = function
# All arguments in expressions are read, except those on the
# LHS of augmented assignment operators. In those cases, the
# operator will have to change the intent.
self.intent = intent
def __str__(self):
if isinstance(self.function, constant.Constant):
if len(self.function.ufl_element().value_shape()) == 0:
return "fn_%d[0]" % self.argnum
else:
return "fn_%d[dim]" % self.argnum
if isinstance(self.function.function_space(),
functionspace.VectorFunctionSpace):
return "fn_%d[dim]" % self.argnum
else:
return "fn_%d[0]" % self.argnum
@property
def arg(self):
argtype = self.function.dat.ctype + "*"
name = " fn_%r" % self.argnum
return ast.Decl(argtype, ast.Symbol(name))
@property
def ast(self):
# Constant broadcasts across functions if it's a scalar
if isinstance(self.function, constant.Constant) and \
len(self.function.ufl_element().value_shape()) == 0:
return ast.Symbol("fn_%d" % self.argnum, (0, ))
return ast.Symbol("fn_%d" % self.argnum, ("dim",))
class AssignmentBase(Operator):
"""Base class for UFL augmented assignments."""
__slots__ = ("ufl_shape", "_symbol", "_ast", "_visit")
_identity = Zero()
def __init__(self, lhs, rhs):
operands = map(ufl.as_ufl, (lhs, rhs))
super(AssignmentBase, self).__init__(operands)
self.ufl_shape = lhs.ufl_shape
# Sub function assignment, we've put a Zero in the lhs
# indicating we should do nothing.
if type(lhs) is Zero:
return
if not (isinstance(lhs, function.Function)
or isinstance(lhs, DummyFunction)):
raise TypeError("Can only assign to a Function")
def __str__(self):
return (" %s " % self._symbol).join(map(str, self.ufl_operands))
def __repr__(self):
return "%s(%s)" % (self.__class__.__name__,
", ".join(repr(o) for o in self.ufl_operands))
@property
def ast(self):
return self._ast(_ast(self.ufl_operands[0]), _ast(self.ufl_operands[1]))
@ufl_type(num_ops=2, is_abstract=False, is_index_free=True, is_shaping=False)
class Assign(AssignmentBase):
"""A UFL assignment operator."""
_symbol = "="
_ast = ast.Assign
__slots__ = ("ufl_shape", "_symbol", "_ast", "_visit")
def _visit(self, transformer):
lhs = self.ufl_operands[0]
transformer._result = lhs
try:
# If lhs is int the dictionary, this indicates that it is
# also on the RHS and therefore needs to be RW.
new_lhs = transformer._args[lhs]
new_lhs.intent = op2.RW
except KeyError:
if transformer._function_space is None:
transformer._function_space = lhs._function_space
elif transformer._function_space != lhs._function_space:
raise ValueError("Expression has incompatible function spaces")
transformer._args[lhs] = DummyFunction(lhs, len(transformer._args),
intent=op2.WRITE)
new_lhs = transformer._args[lhs]
return [new_lhs, self.ufl_operands[1]]
class AugmentedAssignment(AssignmentBase):
"""Base for the augmented assignment operators `+=`, `-=,` `*=`, `/=`"""
__slots__ = ()
def _visit(self, transformer):
lhs = self.ufl_operands[0]
transformer._result = lhs
try:
new_lhs = transformer._args[lhs]
except KeyError:
if transformer._function_space is None:
transformer._function_space = lhs._function_space
elif transformer._function_space != lhs._function_space:
raise ValueError("Expression has incompatible function spaces")
transformer._args[lhs] = DummyFunction(lhs, len(transformer._args))
new_lhs = transformer._args[lhs]
new_lhs.intent = op2.RW
return [new_lhs, self.ufl_operands[1]]
@ufl_type(num_ops=2, is_abstract=False, is_index_free=True, is_shaping=False)
class IAdd(AugmentedAssignment):
"""A UFL `+=` operator."""
_symbol = "+="
_ast = ast.Incr
__slots__ = ()
@ufl_type(num_ops=2, is_abstract=False, is_index_free=True, is_shaping=False)
class ISub(AugmentedAssignment):
"""A UFL `-=` operator."""
_symbol = "-="
_ast = ast.Decr
__slots__ = ()
@ufl_type(num_ops=2, is_abstract=False, is_index_free=True, is_shaping=False)
class IMul(AugmentedAssignment):
"""A UFL `*=` operator."""
_symbol = "*="
_ast = ast.IMul
_identity = IntValue(1)
__slots__ = ()
@ufl_type(num_ops=2, is_abstract=False, is_index_free=True, is_shaping=False)
class IDiv(AugmentedAssignment):
"""A UFL `/=` operator."""
_symbol = "/="
_ast = ast.IDiv
_identity = IntValue(1)
__slots__ = ()
class Power(ufl.algebra.Power):
"""Subclass of :class:`ufl.algebra.Power` which prints pow(x,y)
instead of x**y."""
def __str__(self):
return "pow(%s, %s)" % self.ufl_operands
@property
def ast(self):
return ast.FunCall("pow", _ast(self.ufl_operands[0], _ast(self.ufl_operands[1])))
class Ln(ufl.mathfunctions.Ln):
"""Subclass of :class:`ufl.mathfunctions.Ln` which prints log(x)
instead of ln(x)."""
def __str__(self):
return "log(%s)" % str(self.ufl_operands[0])
@property
def ast(self):
return ast.FunCall("log", _ast(self.ufl_operands[0]))
class ComponentTensor(ufl.tensors.ComponentTensor):
"""Subclass of :class:`ufl.tensors.ComponentTensor` which only prints the
first operand."""
def __str__(self):
return str(self.ufl_operands[0])
@property
def ast(self):
return _ast(self.ufl_operands[0])
class Indexed(ufl.indexed.Indexed):
"""Subclass of :class:`ufl.indexed.Indexed` which only prints the first
operand."""
def __str__(self):
return str(self.ufl_operands[0])
@property
def ast(self):
return _ast(self.ufl_operands[0])
class ExpressionSplitter(ReuseTransformer):
"""Split an expression tree into a subtree for each component of the
appropriate :class:`.FunctionSpaceBase`."""
def split(self, expr):
"""Split the given expression."""
self._identity = expr._identity
self._trees = None
lhs, rhs = expr.ufl_operands
# If the expression is not an assignment, the function spaces for both
# operands have to match
if not isinstance(expr, AssignmentBase) and \
lhs.function_space() != rhs.function_space():
raise ValueError("Operands of %r must have the same FunctionSpace" % expr)
self._fs = lhs.function_space()
return [expr.reconstruct(*ops) for ops in zip(*map(self.visit, (lhs, rhs)))]
def indexed(self, o, *operands):
"""Reconstruct the :class:`ufl.indexed.Indexed` only if the coefficient
is defined on a :class:`.VectorFunctionSpace`."""
def reconstruct_if_vec(coeff, idx, i):
# If the MultiIndex contains a FixedIndex we only want to return
# the indexed coefficient if its position matches the FixedIndex
# Since we don't split VectorFunctionSpaces, we have to
# reconstruct the fixed index expression for those (and only those)
if isinstance(idx._indices[0], ufl.indexing.FixedIndex):
if idx._indices[0]._value != i:
return self._identity
elif isinstance(coeff.function_space(), functionspace.VectorFunctionSpace):
return o.reconstruct(coeff, idx)
return coeff
return [reconstruct_if_vec(*ops, i=i)
for i, ops in enumerate(zip(*operands))]
def component_tensor(self, o, *operands):
"""Only return the first operand."""
return operands[0]
def terminal(self, o):
if isinstance(o, function.Function):
# A function must either be defined on the same function space
# we're assigning to, in which case we split it into components
if o.function_space() == self._fs:
return o.split()
# If the function space we're assigning into is /not/
# Mixed, o must be indexed and the functionspace component
# much match us.
if not isinstance(self._fs, functionspace.MixedFunctionSpace) \
and self._fs.index is None:
idx = o.function_space().index
if idx is None:
raise ValueError("Coefficient %r is not indexed" % o)
if o.function_space()._fs != self._fs:
raise ValueError("Mismatching function spaces")
return (o,)
# Otherwise the function space must be indexed and we
# return the Function for the indexed component and the
# identity for this assignment for every other
idx = o.function_space().index
# LHS is indexed
if self._fs.index is not None:
# RHS indexed, indexed RHS function space must match
# indexed LHS function space.
if idx is not None and self._fs._fs != o.function_space()._fs:
raise ValueError("Mismatching indexed function spaces")
# RHS not indexed, RHS function space must match
# indexed LHS function space
elif idx is None and self._fs._fs != o.function_space():
raise ValueError("Mismatching function spaces")
# OK, everything checked out. Return RHS
return (o,)
# LHS not indexed, RHS must be indexed and isn't
if idx is None:
raise ValueError("Coefficient %r is not indexed" % o)
# RHS indexed, parent function space must match LHS function space
if self._fs != o.function_space()._parent:
raise ValueError("Mismatching function spaces")
# Return RHS in index slot in expression and
# identity otherwise.
return tuple(o if i == idx else self._identity
for i, _ in enumerate(self._fs))
# We replicate ConstantValue and MultiIndex for each component
elif isinstance(o, (constant.Constant, ConstantValue, MultiIndex)):
# If LHS is indexed, only return a scalar result
if self._fs.index is not None:
return (o,)
return tuple(o for _ in self._fs)
raise NotImplementedError("Don't know what to do with %r" % o)
def product(self, o, *operands):
"""Reconstruct a product on each of the component spaces."""
return [op0 * op1 for op0, op1 in zip(*operands)]
def operator(self, o, *operands):
"""Reconstruct an operator on each of the component spaces."""
ret = []
for ops in zip(*operands):
# Don't try to reconstruct if we've just got the identity
# Stops domain errors when calling Log on Zero (for example)
if len(ops) == 1 and type(ops[0]) is type(self._identity):
ret.append(ops[0])
else:
ret.append(o.reconstruct(*ops))
return ret
class ExpressionWalker(ReuseTransformer):
def __init__(self):
ReuseTransformer.__init__(self)
self._args = {}
self._function_space = None
self._result = None
def walk(self, expr):
"""Walk the given expression and return a tuple of the transformed
expression, the list of coefficients sorted by their count and the
function space the expression is defined on."""
return (self.visit(expr),
sorted(self._args.values(), key=lambda c: c.count()),
self._function_space)
def coefficient(self, o):
if isinstance(o, function.Function):
if self._function_space is None:
self._function_space = o._function_space
else:
# Peel out (potentially indexed) function space of LHS
# and RHS to check for compatibility.
sfs = self._function_space
ofs = o._function_space
if sfs.index is not None:
sfs = sfs._fs
if ofs.index is not None:
ofs = ofs._fs
if sfs != ofs:
raise ValueError("Expression has incompatible function spaces %s and %s" %
(sfs, ofs))
try:
arg = self._args[o]
if arg.intent == op2.WRITE:
# arg occurs on both the LHS and RHS of an assignment.
arg.intent = op2.RW
return arg
except KeyError:
self._args[o] = DummyFunction(o, len(self._args))
return self._args[o]
elif isinstance(o, constant.Constant):
if self._function_space is None:
raise NotImplementedError("Cannot assign to Constant coefficients")
else:
# Constant shape has to match if the constant is not a scalar
# If it is a scalar, it gets broadcast across all of
# the values of the function.
if len(o.ufl_element().value_shape()) > 0:
for fs in self._function_space:
if fs.ufl_element().value_shape() != o.ufl_element().value_shape():
raise ValueError("Constant has mismatched shape for expression function space")
try:
arg = self._args[o]
if arg.intent == op2.WRITE:
arg.intent = op2.RW
return arg
except KeyError:
self._args[o] = DummyFunction(o, len(self._args))
return self._args[o]
elif isinstance(o, DummyFunction):
# Idempotency.
return o
else:
raise TypeError("Operand %s is of unsupported type" % o)
# Prevent AlgebraOperators falling through to the Operator case.
algebra_operator = ReuseTransformer.reuse_if_possible
conditional = ReuseTransformer.reuse_if_possible
condition = ReuseTransformer.reuse_if_possible
math_function = ReuseTransformer.reuse_if_possible
def power(self, o, *operands):
# Need to convert notation to c for exponents.
return Power(*operands)
def ln(self, o, *operands):
# Need to convert notation to c.
return Ln(*operands)
def component_tensor(self, o, *operands):
"""Override string representation to only print first operand."""
return ComponentTensor(*operands)
def indexed(self, o, *operands):
"""Override string representation to only print first operand."""
return Indexed(*operands)
def operator(self, o):
# Need pre-traversal of operators so as to correctly set the
# intent of the lhs function of Assignments.
if isinstance(o, AssignmentBase):
operands = o._visit(self)
# The left operand is special-cased in the assignment
# visit method. The general visitor is applied to the RHS.
operands = [operands[0], self.visit(operands[1])]
else:
# For all other operators, just visit the children.
operands = map(self.visit, o.ufl_operands)
return o.reconstruct(*operands)
def expression_kernel(expr, args):
"""Produce a :class:`pyop2.Kernel` from the processed UFL expression
expr and the corresponding args."""
fs = args[0].function.function_space()
d = ast.Symbol("dim")
if isinstance(fs, functionspace.VectorFunctionSpace):
ast_expr = _ast(expr)
else:
ast_expr = ast.FlatBlock(str(expr) + ";")
body = ast.Block(
(
ast.Decl("int", d),
ast.For(ast.Assign(d, ast.Symbol(0)),
ast.Less(d, ast.Symbol(fs.dof_dset.cdim)),
ast.Incr(d, ast.Symbol(1)),
ast_expr)
)
)
return op2.Kernel(ast.FunDecl("void", "expression",
[arg.arg for arg in args], body),
"expression")
def evaluate_preprocessed_expression(expr, args, subset=None):
# Empty slot indicating assignment to indexed LHS, so don't do anything
if type(expr) is Zero:
return
kernel = expression_kernel(expr, args)
# We need to splice the args according to the components of the
# MixedFunctionSpace if we have one
for j, dats in enumerate(zip(*tuple(a.function.dat for a in args))):
itset = subset or args[0].function._function_space[j].node_set
parloop_args = [dat(args[i].intent) for i, dat in enumerate(dats)]
op2.par_loop(kernel, itset, *parloop_args)
def evaluate_expression(expr, subset=None):
"""Evaluates UFL expressions on :class:`.Function`\s."""
for tree in ExpressionSplitter().split(expr):
e, args, _ = ExpressionWalker().walk(tree)
evaluate_preprocessed_expression(e, args, subset)
def assemble_expression(expr, subset=None):
"""Evaluates UFL expressions on :class:`.Function`\s pointwise and assigns
into a new :class:`.Function`."""
result = function.Function(ExpressionWalker().walk(expr)[2])
evaluate_expression(Assign(result, expr), subset)
return result
<file_sep>/firedrake/vector.py
import numpy as np
from mpi4py import MPI
from pyop2 import op2
from petsc import PETSc
__all__ = ['Vector']
class Vector(object):
def __init__(self, x):
"""Build a `Vector` that wraps a :class:`pyop2.Dat` for Dolfin compatibilty.
:arg x: an :class:`pyop2.Dat` to wrap or a :class:`Vector` to copy.
This copies the underlying data in the :class:`pyop2.Dat`.
"""
if isinstance(x, Vector):
self.dat = op2.Dat(x.dat)
elif isinstance(x, op2.base.Dat): # ugh
self.dat = x
else:
raise RuntimeError("Don't know how to build a Vector from a %r" % type(x))
def axpy(self, a, x):
"""Add a*x to self.
:arg a: a scalar
:arg x: a :class:`Vector` or :class:`.Function`"""
self.dat += a*x.dat
def _scale(self, a):
"""Scale self by `a`.
:arg a: a scalar
"""
self.dat *= a
def array(self):
"""Return a copy of the process local data as a numpy array"""
return np.copy(self.dat.data_ro)
def get_local(self):
"""Return a copy of the process local data as a numpy array"""
return self.array()
def set_local(self, values):
"""Set process local values
:arg values: a numpy array of values of length :func:`Vector.local_size`"""
self.dat.data[:] = values
def local_size(self):
"""Return the size of the process local data (without ghost points)"""
return self.dat.dataset.size
def size(self):
"""Return the global size of the data"""
lsize = self.local_size()
return op2.MPI.comm.allreduce(lsize, op=MPI.SUM)
def gather(self, global_indices=None):
"""Gather a :class:`Vector` to all processes
:arg global_indices: the globally numbered indices to gather
(should be the same on all processes). If
`None`, gather the entire :class:`Vector`."""
if global_indices is None:
N = self.size()
v = PETSc.Vec().createSeq(N, comm=PETSc.COMM_SELF)
is_ = PETSc.IS().createStride(N, 0, 1, comm=PETSc.COMM_SELF)
else:
global_indices = np.asarray(global_indices, dtype=np.int32)
N = len(global_indices)
v = PETSc.Vec().createSeq(N, comm=PETSc.COMM_SELF)
is_ = PETSc.IS().createGeneral(global_indices, comm=PETSc.COMM_SELF)
with self.dat.vec as vec:
vscat = PETSc.Scatter().create(vec, is_, v, None)
vscat.scatterBegin(vec, v, addv=PETSc.InsertMode.INSERT_VALUES)
vscat.scatterEnd(vec, v, addv=PETSc.InsertMode.INSERT_VALUES)
return v.array
def __setitem__(self, idx, value):
"""Set a value or values in the local data
:arg idx: the local idx, or indices to set.
:arg value: the value, or values to give them."""
self.dat.data[idx] = value
def __getitem__(self, idx):
"""Return a value or values in the local data
:arg idx: the local idx, or indices to set."""
return self.dat.data_ro[idx]
def __len__(self):
"""Return the length of the local data (not including ghost points)"""
return self.local_size()
<file_sep>/tests/regression/test_mixed_mats.py
import pytest
import numpy as np
from firedrake import *
@pytest.fixture(scope='module')
def m(request):
return UnitTriangleMesh()
@pytest.fixture(scope='module')
def V(m):
return FunctionSpace(m, 'DG', 0)
@pytest.fixture(scope='module')
def Q(m):
return FunctionSpace(m, 'RT', 1)
@pytest.fixture(scope='module')
def W(V, Q):
return V*Q
# NOTE: these tests make little to no mathematical sense, they are
# here to exercise corner cases in PyOP2's handling of mixed spaces.
def test_massVW0(V, W):
u = TrialFunction(V)
v = TestFunction(W)[0]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 1)
# DGxDG block
assert not np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block (0, since test function was restricted to DG block)
assert np.allclose(A.M[1, 0].values, 0.0)
def test_massVW1(V, W):
u = TrialFunction(V)
v = TestFunction(W)[1]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 1)
# DGxDG block (0, since test function was restricted to RT block)
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert not np.allclose(A.M[1, 0].values, 0.0)
def test_massW0W0(W):
u = TrialFunction(W)[0]
v = TestFunction(W)[0]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert not np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert np.allclose(A.M[1, 1].values, 0.0)
def test_massW1W1(W):
u = TrialFunction(W)[1]
v = TestFunction(W)[1]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert not np.allclose(A.M[1, 1].values, 0.0)
def test_massW0W1(W):
u = TrialFunction(W)[0]
v = TestFunction(W)[1]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert not np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert np.allclose(A.M[1, 1].values, 0.0)
def test_massW1W0(W):
u = TrialFunction(W)[1]
v = TestFunction(W)[0]
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert not np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert np.allclose(A.M[1, 1].values, 0.0)
def test_massWW(W):
u = TrialFunction(W)
v = TestFunction(W)
A = assemble(inner(u, v)*dx)
assert A.M.sparsity.shape == (2, 2)
# DGxDG block
assert not np.allclose(A.M[0, 0].values, 0.0)
# DGxRT block
assert np.allclose(A.M[1, 0].values, 0.0)
# RTxDG block
assert np.allclose(A.M[0, 1].values, 0.0)
# RTxRT block
assert not np.allclose(A.M[1, 1].values, 0.0)
def test_bcs_ordering():
"""Check that application of boundary conditions zeros the correct
rows and columns of a mixed matrix.
The diagonal blocks should get a 1 in the diagonal entries
corresponding to the boundary condition nodes, the corresponding
rows and columns in the whole system should be zeroed."""
m = UnitIntervalMesh(5)
V = FunctionSpace(m, 'CG', 1)
W = V*V
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
bc1 = DirichletBC(W.sub(0), 0, 1)
bc2 = DirichletBC(W.sub(1), 1, 2)
a = (u*v + u*q + p*v + p*q)*dx
A = assemble(a, bcs=[bc1, bc2])
assert np.allclose(A.M[0, 0].values.diagonal()[bc1.nodes], 1.0)
assert np.allclose(A.M[1, 1].values.diagonal()[bc2.nodes], 1.0)
assert np.allclose(A.M[0, 1].values[bc1.nodes, :], 0.0)
assert np.allclose(A.M[1, 0].values[:, bc1.nodes], 0.0)
assert np.allclose(A.M[1, 0].values[bc2.nodes, :], 0.0)
assert np.allclose(A.M[0, 1].values[:, bc2.nodes], 0.0)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/ffc_interface.py
"""Provides the interface to FFC for compiling a form, and transforms the FFC-
generated code in order to make it suitable for passing to the backends."""
from collections import defaultdict
from hashlib import md5
from operator import add
from os import path, environ, getuid, makedirs
import tempfile
import ufl
from ufl import Form, FiniteElement, VectorElement, as_vector
from ufl.measure import Measure
from ufl.algorithms import compute_form_data, ReuseTransformer
from ufl.constantvalue import Zero
from ufl_expr import Argument
from ffc import compile_form as ffc_compile_form
from ffc import constants
from ffc import log
from ffc.quadrature.quadraturetransformerbase import EmptyIntegrandError
from pyop2.caching import DiskCached
from pyop2.op2 import Kernel
from pyop2.mpi import MPI
from pyop2.coffee.ast_base import PreprocessNode, Root
import fiat_utils
import functionspace
from parameters import parameters as default_parameters
_form_cache = {}
# Only spew ffc message on rank zero
if MPI.comm.rank != 0:
log.set_level(log.ERROR)
del log
def _check_version():
from version import __compatible_ffc_version_info__ as compatible_version, \
__compatible_ffc_version__ as version
try:
if constants.PYOP2_VERSION_INFO[:2] == compatible_version[:2]:
return
except AttributeError:
pass
raise RuntimeError("Incompatible PyOP2 version %s and FFC PyOP2 version %s."
% (version, getattr(constants, 'PYOP2_VERSION', 'unknown')))
def sum_integrands(form):
"""Produce a form with the integrands on the same measure summed."""
integrals = defaultdict(list)
for integral in form.integrals():
integrals[(integral.integral_type(), integral.subdomain_id())].append(integral)
return Form([it[0].reconstruct(reduce(add, [i.integrand() for i in it]))
for it in integrals.values()])
class FormSplitter(ReuseTransformer):
"""Split a form into a subtree for each component of the mixed space it is
built on. This is a no-op on forms over non-mixed spaces."""
def split(self, form):
"""Split the given form."""
# Visit each integrand and obtain the tuple of sub forms
args = tuple((a.number(), len(a.function_space()))
for a in form.arguments())
forms_list = []
for it in sum_integrands(form).integrals():
forms = []
def visit(idx):
integrand = self.visit(it.integrand())
if not isinstance(integrand, Zero):
forms.append([(idx, integrand * Measure(it.integral_type(),
domain=it.domain(),
subdomain_id=it.subdomain_id(),
subdomain_data=it.subdomain_data(),
metadata=it.metadata()))])
# 0 form
if not args:
visit((0, 0))
# 1 form
elif len(args) == 1:
count, l = args[0]
for i in range(l):
self._idx = {count: i}
self._args = {}
visit((i, 0))
# 2 form
elif len(args) == 2:
for i in range(args[0][1]):
for j in range(args[1][1]):
self._idx = {args[0][0]: i, args[1][0]: j}
self._args = {}
visit((i, j))
forms_list += forms
return forms_list
def argument(self, arg):
"""Split an argument into its constituent spaces."""
if isinstance(arg.function_space(), functionspace.MixedFunctionSpace):
if arg in self._args:
return self._args[arg]
args = []
for i, fs in enumerate(arg.function_space().split()):
# Look up the split argument in cache since we want it unique
a = Argument(fs.ufl_element(), fs, arg.number())
if a.shape():
if self._idx[arg.number()] == i:
args += [a[j] for j in range(a.shape()[0])]
else:
args += [Zero() for j in range(a.shape()[0])]
else:
if self._idx[arg.number()] == i:
args.append(a)
else:
args.append(Zero())
self._args[arg] = as_vector(args)
return as_vector(args)
return arg
class FFCKernel(DiskCached):
_cache = {}
if MPI.comm.rank == 0:
_cachedir = environ.get('FIREDRAKE_FFC_KERNEL_CACHE_DIR',
path.join(tempfile.gettempdir(),
'firedrake-ffc-kernel-cache-uid%d' % getuid()))
# Include an md5 hash of firedrake_geometry.h in the cache key
with open(path.join(path.dirname(__file__), 'firedrake_geometry.h')) as f:
_firedrake_geometry_md5 = md5(f.read()).hexdigest()
del f
MPI.comm.bcast(_firedrake_geometry_md5, root=0)
else:
# No cache on slave processes
_cachedir = None
# MD5 obtained by broadcast from root
_firedrake_geometry_md5 = MPI.comm.bcast(None, root=0)
@classmethod
def _cache_key(cls, form, name, parameters):
# FIXME Making the COFFEE parameters part of the cache key causes
# unnecessary repeated calls to FFC when actually only the kernel code
# needs to be regenerated
return md5(form.signature() + name + Kernel._backend.__name__ +
cls._firedrake_geometry_md5 + constants.FFC_VERSION +
constants.PYOP2_VERSION + str(default_parameters["coffee"])
+ str(parameters)).hexdigest()
def _needs_orientations(self, elements):
if len(elements) == 0:
return False
cell = elements[0].cell()
if cell.topological_dimension() == cell.geometric_dimension():
return False
for e in elements:
if isinstance(e, ufl.MixedElement) and e.family() != 'Real':
if any("contravariant piola" in fiat_utils.fiat_from_ufl_element(s).mapping()
for s in e.sub_elements()):
return True
else:
if e.family() != 'Real' and \
"contravariant piola" in fiat_utils.fiat_from_ufl_element(e).mapping():
return True
return False
def __init__(self, form, name, parameters):
"""A wrapper object for one or more FFC kernels compiled from a given :class:`~Form`.
:arg form: the :class:`~Form` from which to compile the kernels.
:arg name: a prefix to be applied to the compiled kernel names. This is primarily useful for debugging.
:arg parameters: a dict of parameters to pass to the form compiler.
"""
if self._initialized:
return
incl = PreprocessNode('#include "firedrake_geometry.h"\n')
inc = [path.dirname(__file__)]
try:
ffc_tree = ffc_compile_form(form, prefix=name, parameters=parameters)
kernels = []
# need compute_form_data here to get preproc form integrals
fd = compute_form_data(form)
elements = fd.elements
needs_orientations = self._needs_orientations(elements)
for it, kernel in zip(fd.preprocessed_form.integrals(), ffc_tree):
# Set optimization options
opts = {} if it.integral_type() not in ['cell'] else default_parameters["coffee"]
kernels.append((Kernel(Root([incl, kernel]), '%s_%s_integral_0_%s' %
(name, it.integral_type(), it.subdomain_id()), opts, inc),
needs_orientations))
self.kernels = tuple(kernels)
self._empty = False
except EmptyIntegrandError:
# FFC noticed that the integrand was zero and simplified
# it, catch this here and set a flag telling us to ignore
# the kernel when returning it in compile_form
self._empty = True
self._initialized = True
def compile_form(form, name, parameters=None):
"""Compile a form using FFC.
:arg form: the :class:`ufl.Form` to compile.
:arg name: a prefix for the generated kernel functions.
:arg parameters: optional dict of parameters to pass to the form
compiler. If not provided, parameters are read from the
:data:`form_compiler` slot of the Firedrake
:data:`~.parameters` dictionary (which see).
Returns a tuple of tuples of
(index, integral type, subdomain id, coordinates, coefficients, needs_orientations, :class:`Kernels <pyop2.op2.Kernel>`).
``needs_orientations`` indicates whether the form requires cell
orientation information (for correctly pulling back to reference
elements on embedded manifolds).
The coordinates are extracted from the UFL
:class:`~ufl.domain.Domain` of the integral.
"""
# Check that we get a Form
if not isinstance(form, Form):
raise RuntimeError("Unable to convert object to a UFL form: %s" % repr(form))
if parameters is None:
parameters = default_parameters["form_compiler"]
else:
# Override defaults with user-specified values
_ = parameters
parameters = default_parameters["form_compiler"].copy()
parameters.update(_)
# We stash the compiled kernels on the form so we don't have to recompile
# if we assemble the same form again with the same optimisations
if hasattr(form, "_kernels"):
# Save both kernels and FFC params so we can tell if this
# cached version is valid (the FFC parameters might have changed)
kernels, params = form._kernels
if kernels[0][-1]._opts == default_parameters["coffee"] and \
kernels[0][-1].name.startswith(name) and \
params == parameters:
return kernels
# need compute_form_data since we use preproc. form integrals later
fd = compute_form_data(form)
# If there is no mixed element involved, return the kernels FFC produces
if all(isinstance(e, (FiniteElement, VectorElement)) for e in fd.unique_sub_elements):
kernels = [((0, 0),
it.integral_type(), it.subdomain_id(),
it.domain().data().coordinates,
fd.preprocessed_form.coefficients(), needs_orientations, kernel)
for it, (kernel, needs_orientations) in zip(fd.preprocessed_form.integrals(),
FFCKernel(form, name,
parameters).kernels)]
form._kernels = (kernels, parameters)
return kernels
# Otherwise pre-split the form into mixed blocks before calling FFC
kernels = []
for forms in FormSplitter().split(form):
for (i, j), f in forms:
ffc_kernel = FFCKernel(f, name + str(i) + str(j), parameters)
# FFC noticed the integrand was zero, so don't bother
# using this kernel (it's invalid anyway)
if ffc_kernel._empty:
continue
((kernel, needs_orientations), ) = ffc_kernel.kernels
# need compute_form_data here to get preproc integrals
fd = compute_form_data(f)
it = fd.preprocessed_form.integrals()[0]
kernels.append(((i, j),
it.integral_type(),
it.subdomain_id(),
it.domain().data().coordinates,
fd.preprocessed_form.coefficients(),
needs_orientations, kernel))
form._kernels = (kernels, parameters)
return kernels
def clear_cache():
"""Clear the Firedrake FFC kernel cache."""
if MPI.comm.rank != 0:
return
if path.exists(FFCKernel._cachedir):
import shutil
shutil.rmtree(FFCKernel._cachedir, ignore_errors=True)
_ensure_cachedir()
def _ensure_cachedir():
"""Ensure that the FFC kernel cache directory exists."""
if MPI.comm.rank != 0:
return
if not path.exists(FFCKernel._cachedir):
makedirs(FFCKernel._cachedir)
_check_version()
_ensure_cachedir()
<file_sep>/tests/regression/test_projection.py
import pytest
import numpy as np
from firedrake import *
def run_vector_valued_test(x, degree=1, family='RT'):
m = UnitSquareMesh(2 ** x, 2 ** x)
V = FunctionSpace(m, family, degree)
expr = ['cos(x[0]*pi*2)*sin(x[1]*pi*2)']*2
e = Expression(expr)
exact = Function(VectorFunctionSpace(m, 'CG', 5))
exact.interpolate(e)
# Solve to machine precision.
ret = project(e, V, solver_parameters={'ksp_type': 'preonly', 'pc_type': 'lu'})
return sqrt(assemble(inner((ret - exact), (ret - exact)) * dx))
def run_vector_test(x, degree=1, family='CG'):
m = UnitSquareMesh(2 ** x, 2 ** x)
V = VectorFunctionSpace(m, family, degree)
expr = ['cos(x[0]*pi*2)*sin(x[1]*pi*2)']*2
e = Expression(expr)
exact = Function(VectorFunctionSpace(m, 'CG', 5))
exact.interpolate(e)
# Solve to machine precision. This version of the test uses the
# alternate syntax in which the target Function is already
# available.
ret = Function(V)
project(e, ret, solver_parameters={'ksp_type': 'preonly', 'pc_type': 'lu'})
return sqrt(assemble(inner((ret - exact), (ret - exact)) * dx))
def run_test(x, degree=1, family='CG'):
m = UnitSquareMesh(2 ** x, 2 ** x)
V = FunctionSpace(m, family, degree)
e = Expression('cos(x[0]*pi*2)*sin(x[1]*pi*2)')
exact = Function(FunctionSpace(m, 'CG', 5))
exact.interpolate(e)
# Solve to machine precision. This version of the test uses the
# method version of project.
ret = Function(V)
ret.project(e, solver_parameters={'ksp_type': 'preonly', 'pc_type': 'lu'})
return sqrt(assemble((ret - exact) * (ret - exact) * dx))
@pytest.mark.parametrize(('degree', 'family', 'expected_convergence'), [
(1, 'CG', 1.8),
(2, 'CG', 2.6),
(3, 'CG', 3.8),
(0, 'DG', 0.8),
(1, 'DG', 1.8),
(2, 'DG', 2.8)])
def test_convergence(degree, family, expected_convergence):
l2_diff = np.array([run_test(x, degree, family) for x in range(2, 5)])
conv = np.log2(l2_diff[:-1] / l2_diff[1:])
assert (conv > expected_convergence).all()
@pytest.mark.parametrize(('degree', 'family', 'expected_convergence'), [
(1, 'CG', 1.8),
(2, 'CG', 2.6),
(3, 'CG', 3.8),
(0, 'DG', 0.8),
(1, 'DG', 1.8),
(2, 'DG', 2.8)])
def test_vector_convergence(degree, family, expected_convergence):
l2_diff = np.array([run_vector_test(x, degree, family) for x in range(2, 5)])
conv = np.log2(l2_diff[:-1] / l2_diff[1:])
assert (conv > expected_convergence).all()
@pytest.mark.parametrize(('degree', 'family', 'expected_convergence'), [
(1, 'RT', 0.75),
(2, 'RT', 1.8),
(3, 'RT', 2.8),
(1, 'BDM', 1.8),
(2, 'BDM', 2.8),
(3, 'BDM', 3.8)])
def test_vector_valued_convergence(degree, family, expected_convergence):
l2_diff = np.array([run_vector_valued_test(x, degree, family)
for x in range(2, 6)])
conv = np.log2(l2_diff[:-1] / l2_diff[1:])
assert (conv > expected_convergence).all()
def test_project_mismatched_rank():
m = UnitSquareMesh(2, 2)
V = FunctionSpace(m, 'CG', 1)
U = FunctionSpace(m, 'RT', 1)
v = Function(V)
u = Function(U)
ev = Expression('')
eu = Expression(('', ''))
with pytest.raises(RuntimeError):
project(v, U)
with pytest.raises(RuntimeError):
project(u, V)
with pytest.raises(RuntimeError):
project(ev, U)
with pytest.raises(RuntimeError):
project(eu, V)
def test_project_mismatched_mesh():
m2 = UnitSquareMesh(2, 2)
m3 = UnitCubeMesh(2, 2, 2)
U = FunctionSpace(m2, 'CG', 1)
V = FunctionSpace(m3, 'CG', 1)
u = Function(U)
v = Function(V)
with pytest.raises(RuntimeError):
project(u, V)
with pytest.raises(RuntimeError):
project(v, U)
def test_project_mismatched_shape():
m = UnitSquareMesh(2, 2)
U = VectorFunctionSpace(m, 'CG', 1, dim=3)
V = VectorFunctionSpace(m, 'CG', 1, dim=2)
u = Function(U)
v = Function(V)
with pytest.raises(RuntimeError):
project(u, V)
with pytest.raises(RuntimeError):
project(v, U)
def test_repeatable():
mesh = UnitSquareMesh(1, 1)
Q = FunctionSpace(mesh, 'DG', 1)
V2 = FunctionSpace(mesh, 'DG', 0)
V3 = FunctionSpace(mesh, 'DG', 0)
W = V2 * V3
expr = Expression('1.0')
old = project(expr, Q)
f = project(Expression(('-1.0', '-1.0')), W) # noqa
new = project(expr, Q)
for fd, ud in zip(new.dat.data, old.dat.data):
assert (fd == ud).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/mesh.py
import numpy as np
import os
import FIAT
import ufl
from pyop2 import op2
from pyop2.coffee import ast_base as ast
from pyop2.profiling import timed_function, timed_region, profile
from pyop2.utils import as_tuple
import dmplex
import extrusion_utils as eutils
import fiat_utils
import function
import functionspace
import utility_meshes
import utils
from parameters import parameters
from petsc import PETSc
__all__ = ['Mesh', 'ExtrudedMesh']
class _Facets(object):
"""Wrapper class for facet interation information on a :class:`Mesh`
.. warning::
The unique_markers argument **must** be the same on all processes."""
def __init__(self, mesh, classes, kind, facet_cell, local_facet_number, markers=None,
unique_markers=None):
self.mesh = mesh
classes = as_tuple(classes, int, 4)
self.classes = classes
self.kind = kind
assert(kind in ["interior", "exterior"])
if kind == "interior":
self._rank = 2
else:
self._rank = 1
self.facet_cell = facet_cell
self.local_facet_number = local_facet_number
self.markers = markers
self.unique_markers = [] if unique_markers is None else unique_markers
self._subsets = {}
@utils.cached_property
def set(self):
size = self.classes
halo = None
if isinstance(self.mesh, ExtrudedMesh):
if self.kind == "interior":
base = self.mesh._old_mesh.interior_facets.set
else:
base = self.mesh._old_mesh.exterior_facets.set
return op2.ExtrudedSet(base, layers=self.mesh.layers)
return op2.Set(size, "%s_%s_facets" % (self.mesh.name, self.kind), halo=halo)
@property
def bottom_set(self):
'''Returns the bottom row of cells.'''
return self.mesh.cell_set
@utils.cached_property
def _null_subset(self):
'''Empty subset for the case in which there are no facets with
a given marker value. This is required because not all
markers need be represented on all processors.'''
return op2.Subset(self.set, [])
def measure_set(self, integral_type, subdomain_id):
'''Return the iteration set appropriate to measure. This will
either be for all the interior or exterior (as appropriate)
facets, or for a particular numbered subdomain.'''
# ufl.Measure doesn't have enums for these any more :(
if subdomain_id in ["everywhere", "otherwise"]:
if integral_type == "exterior_facet_bottom":
return [(op2.ON_BOTTOM, self.bottom_set)]
elif integral_type == "exterior_facet_top":
return [(op2.ON_TOP, self.bottom_set)]
elif integral_type == "interior_facet_horiz":
return self.bottom_set
else:
return self.set
else:
return self.subset(subdomain_id)
def subset(self, markers):
"""Return the subset corresponding to a given marker value.
:param markers: integer marker id or an iterable of marker ids"""
if self.markers is None:
return self._null_subset
markers = as_tuple(markers, int)
try:
return self._subsets[markers]
except KeyError:
indices = np.concatenate([np.nonzero(self.markers == i)[0]
for i in markers])
self._subsets[markers] = op2.Subset(self.set, indices)
return self._subsets[markers]
@utils.cached_property
def local_facet_dat(self):
"""Dat indicating which local facet of each adjacent
cell corresponds to the current facet."""
return op2.Dat(op2.DataSet(self.set, self._rank), self.local_facet_number,
np.uintc, "%s_%s_local_facet_number" % (self.mesh.name, self.kind))
@timed_function("Build mesh")
@profile
def Mesh(meshfile, **kwargs):
"""Construct a mesh object.
Meshes may either be created by reading from a mesh file, or by
providing a PETSc DMPlex object defining the mesh topology.
:param meshfile: Mesh file name (or DMPlex object) defining
mesh topology. See below for details on supported mesh
formats.
:param dim: optional specification of the geometric dimension
of the mesh (ignored if not reading from mesh file).
If not supplied the geometric dimension is deduced from
the topological dimension of entities in the mesh.
:param reorder: optional flag indicating whether to reorder
meshes for better cache locality. If not supplied the
default value in :data:`parameters["reorder_meshes"]`
is used.
:param periodic_coords: optional numpy array of coordinates
used to replace those in the mesh object. These are
only supported in 1D and must have enough entries to be
used as a DG1 field on the mesh. Not supported when
reading from file.
When the mesh is read from a file the following mesh formats
are supported (determined, case insensitively, from the
filename extension):
* GMSH: with extension `.msh`
* Exodus: with extension `.e`, `.exo`
* CGNS: with extension `.cgns`
* Triangle: with extension `.node`
.. note::
When the mesh is created directly from a DMPlex object,
the :data:`dim` parameter is ignored (the DMPlex already
knows its geometric and topological dimensions).
"""
utils._init()
dim = kwargs.get("dim", None)
reorder = kwargs.get("reorder", parameters["reorder_meshes"])
periodic_coords = kwargs.get("periodic_coords", None)
if isinstance(meshfile, PETSc.DMPlex):
name = "plexmesh"
plex = meshfile
else:
name = meshfile
basename, ext = os.path.splitext(meshfile)
if periodic_coords is not None:
raise RuntimeError("Periodic coordinates are unsupported when reading from file")
if ext.lower() in ['.e', '.exo']:
plex = _from_exodus(meshfile)
elif ext.lower() == '.cgns':
plex = _from_cgns(meshfile)
elif ext.lower() == '.msh':
plex = _from_gmsh(meshfile)
elif ext.lower() == '.node':
plex = _from_triangle(meshfile, dim)
else:
raise RuntimeError("Mesh file %s has unknown format '%s'."
% (meshfile, ext[1:]))
# Distribute the dm to all ranks
if op2.MPI.comm.size > 1:
plex.distribute(overlap=1)
topological_dim = plex.getDimension()
cStart, cEnd = plex.getHeightStratum(0) # cells
cell_facets = plex.getConeSize(cStart)
if topological_dim + 1 == cell_facets:
MeshClass = SimplexMesh
elif topological_dim == 2 and cell_facets == 4:
MeshClass = QuadrilateralMesh
else:
raise RuntimeError("Unsupported mesh type.")
return MeshClass(name, plex, dim, reorder,
periodic_coords=periodic_coords)
def _from_gmsh(filename):
"""Read a Gmsh .msh file from `filename`"""
# Create a read-only PETSc.Viewer
gmsh_viewer = PETSc.Viewer().create()
gmsh_viewer.setType("ascii")
gmsh_viewer.setFileMode("r")
gmsh_viewer.setFileName(filename)
gmsh_plex = PETSc.DMPlex().createGmsh(gmsh_viewer)
if gmsh_plex.hasLabel("Face Sets"):
boundary_ids = gmsh_plex.getLabelIdIS("Face Sets").getIndices()
gmsh_plex.createLabel("boundary_ids")
for bid in boundary_ids:
faces = gmsh_plex.getStratumIS("Face Sets", bid).getIndices()
for f in faces:
gmsh_plex.setLabelValue("boundary_ids", f, bid)
return gmsh_plex
def _from_exodus(filename):
"""Read an Exodus .e or .exo file from `filename`"""
plex = PETSc.DMPlex().createExodusFromFile(filename)
boundary_ids = dmplex.getLabelIdIS("Face Sets").getIndices()
plex.createLabel("boundary_ids")
for bid in boundary_ids:
faces = plex.getStratumIS("Face Sets", bid).getIndices()
for f in faces:
plex.setLabelValue("boundary_ids", f, bid)
return plex
def _from_cgns(filename):
"""Read a CGNS .cgns file from `filename`"""
plex = PETSc.DMPlex().createCGNSFromFile(filename)
#TODO: Add boundary IDs
return plex
def _from_triangle(filename, dim):
"""Read a set of triangle mesh files from `filename`"""
basename, ext = os.path.splitext(filename)
if op2.MPI.comm.rank == 0:
try:
facetfile = open(basename+".face")
tdim = 3
except:
try:
facetfile = open(basename+".edge")
tdim = 2
except:
facetfile = None
tdim = 1
if dim is None:
dim = tdim
op2.MPI.comm.bcast(tdim, root=0)
with open(basename+".node") as nodefile:
header = np.fromfile(nodefile, dtype=np.int32, count=2, sep=' ')
nodecount = header[0]
nodedim = header[1]
assert nodedim == dim
coordinates = np.loadtxt(nodefile, usecols=range(1, dim+1), skiprows=1, delimiter=' ')
assert nodecount == coordinates.shape[0]
with open(basename+".ele") as elefile:
header = np.fromfile(elefile, dtype=np.int32, count=2, sep=' ')
elecount = header[0]
eledim = header[1]
eles = np.loadtxt(elefile, usecols=range(1, eledim+1), dtype=np.int32, skiprows=1, delimiter=' ')
assert elecount == eles.shape[0]
cells = map(lambda c: c-1, eles)
else:
tdim = op2.MPI.comm.bcast(None, root=0)
cells = None
coordinates = None
plex = utility_meshes._from_cell_list(tdim, cells, coordinates, comm=op2.MPI.comm)
# Apply boundary IDs
if op2.MPI.comm.rank == 0:
facets = None
try:
header = np.fromfile(facetfile, dtype=np.int32, count=2, sep=' ')
edgecount = header[0]
facets = np.loadtxt(facetfile, usecols=range(1, tdim+2), dtype=np.int32, skiprows=0, delimiter=' ')
assert edgecount == facets.shape[0]
finally:
facetfile.close()
if facets is not None:
vStart, vEnd = plex.getDepthStratum(0) # vertices
for facet in facets:
bid = facet[-1]
vertices = map(lambda v: v + vStart - 1, facet[:-1])
join = plex.getJoin(vertices)
plex.setLabelValue("boundary_ids", join[0], bid)
return plex
class MeshBase(object):
"""A representation of mesh topology and geometry."""
def __init__(self, name, plex, geometric_dim,
reorder, periodic_coords=None):
""" Create mesh from DMPlex object """
# A cache of function spaces that have been built on this mesh
self._cache = {}
self.parent = None
self.name = name
self._plex = plex
self.uid = utils._new_uid()
# Mark exterior and interior facets
# Note. This must come before distribution, because otherwise
# DMPlex will consider facets on the domain boundary to be
# exterior, which is wrong.
with timed_region("Mesh: label facets"):
dmplex.label_facets(self._plex)
topological_dim = plex.getDimension()
if geometric_dim is None:
geometric_dim = topological_dim
if reorder:
with timed_region("Mesh: reorder"):
old_to_new = self._plex.getOrdering(PETSc.Mat.OrderingType.RCM).indices
reordering = np.empty_like(old_to_new)
reordering[old_to_new] = np.arange(old_to_new.size, dtype=old_to_new.dtype)
else:
# No reordering
reordering = None
# Mark OP2 entities and derive the resulting Plex renumbering
with timed_region("Mesh: renumbering"):
dmplex.mark_entity_classes(self._plex)
self._plex_renumbering = dmplex.plex_renumbering(self._plex, reordering)
cStart, cEnd = self._plex.getHeightStratum(0) # cells
cell_facets = self._plex.getConeSize(cStart)
self._ufl_cell = ufl.Cell(fiat_utils._cells[topological_dim][cell_facets],
geometric_dimension=geometric_dim)
self._ufl_domain = ufl.Domain(self.ufl_cell(), data=self)
dim = self._plex.getDimension()
self._cells, self.cell_classes = dmplex.get_cells_by_class(self._plex)
with timed_region("Mesh: cell numbering"):
# Derive a cell numbering from the Plex renumbering
cell_entity_dofs = np.zeros(dim+1, dtype=np.int32)
cell_entity_dofs[-1] = 1
try:
# Old style createSection
self._cell_numbering = self._plex.createSection(1, [1], cell_entity_dofs,
perm=self._plex_renumbering)
except:
# New style
self._cell_numbering = self._plex.createSection([1], cell_entity_dofs,
perm=self._plex_renumbering)
self.interior_facets = None
self.exterior_facets = None
# Note that for bendy elements, this needs to change.
with timed_region("Mesh: coordinate field"):
if periodic_coords is not None:
if self.ufl_cell().geometric_dimension() != 1:
raise NotImplementedError("Periodic coordinates in more than 1D are unsupported")
# We've been passed a periodic coordinate field, so use that.
self._coordinate_fs = functionspace.VectorFunctionSpace(self, "DG", 1)
self.coordinates = function.Function(self._coordinate_fs,
val=periodic_coords,
name="Coordinates")
else:
self._coordinate_fs = functionspace.VectorFunctionSpace(self, "Lagrange", 1)
coordinates = dmplex.reordered_coords(self._plex, self._coordinate_fs._global_numbering,
(self.num_vertices(), geometric_dim))
self.coordinates = function.Function(self._coordinate_fs,
val=coordinates,
name="Coordinates")
self._ufl_domain = ufl.Domain(self.coordinates)
# Build a new ufl element for this function space with the
# correct domain. This is necessary since this function space
# is in the cache and will be picked up by later
# VectorFunctionSpace construction.
self._coordinate_fs._ufl_element = self._coordinate_fs.ufl_element().reconstruct(domain=self.ufl_domain())
# HACK alert!
# Replace coordinate Function by one that has a real domain on it (but don't copy values)
self.coordinates = function.Function(self._coordinate_fs, val=self.coordinates.dat)
# Add domain and subdomain_data to the measure objects we store with the mesh.
self._dx = ufl.Measure('cell', domain=self, subdomain_data=self.coordinates)
self._ds = ufl.Measure('exterior_facet', domain=self, subdomain_data=self.coordinates)
self._dS = ufl.Measure('interior_facet', domain=self, subdomain_data=self.coordinates)
# Set the subdomain_data on all the default measures to this
# coordinate field. Also set the domain on the measure.
for measure in [ufl.dx, ufl.ds, ufl.dS]:
measure._subdomain_data = self.coordinates
measure._domain = self.ufl_domain()
@property
def coordinates(self):
"""The :class:`.Function` containing the coordinates of this mesh."""
return self._coordinate_function
@coordinates.setter
def coordinates(self, value):
self._coordinate_function = value
# If the new coordinate field has a different dimension from
# the geometric dimension of the existing cell, replace the
# cell with one with the correct dimension.
ufl_cell = self.ufl_cell()
if value.element().value_shape()[0] != ufl_cell.geometric_dimension():
if isinstance(ufl_cell, ufl.OuterProductCell):
self._ufl_cell = ufl.OuterProductCell(ufl_cell._A, ufl_cell._B, value.element().value_shape()[0])
else:
self._ufl_cell = ufl.Cell(ufl_cell.cellname(),
geometric_dimension=value.element().value_shape()[0])
self._ufl_domain = ufl.Domain(self.ufl_cell(), data=self)
@property
def layers(self):
"""Return the number of layers of the extruded mesh
represented by the number of occurences of the base mesh."""
return self._layers
def cell_orientations(self):
"""Return the orientation of each cell in the mesh.
Use :func:`init_cell_orientations` to initialise this data."""
if not hasattr(self, '_cell_orientations'):
raise RuntimeError("No cell orientations found, did you forget to call init_cell_orientations?")
return self._cell_orientations
def init_cell_orientations(self, expr):
"""Compute and initialise :attr:`cell_orientations` relative to a specified orientation.
:arg expr: an :class:`.Expression` evaluated to produce a
reference normal direction.
"""
if expr.value_shape()[0] != 3:
raise NotImplementedError('Only implemented for 3-vectors')
if self.ufl_cell() not in (ufl.Cell('triangle', 3), ufl.OuterProductCell(ufl.Cell('interval', 3), ufl.Cell('interval')), ufl.OuterProductCell(ufl.Cell('interval', 2), ufl.Cell('interval'), gdim=3)):
raise NotImplementedError('Only implemented for triangles embedded in 3d')
if hasattr(self, '_cell_orientations'):
raise RuntimeError("init_cell_orientations already called, did you mean to do so again?")
v0 = lambda x: ast.Symbol("v0", (x,))
v1 = lambda x: ast.Symbol("v1", (x,))
n = lambda x: ast.Symbol("n", (x,))
x = lambda x: ast.Symbol("x", (x,))
coords = lambda x, y: ast.Symbol("coords", (x, y))
body = []
body += [ast.Decl("double", v(3)) for v in [v0, v1, n, x]]
body.append(ast.Decl("double", "dot"))
body.append(ast.Assign("dot", 0.0))
body.append(ast.Decl("int", "i"))
body.append(ast.For(ast.Assign("i", 0), ast.Less("i", 3), ast.Incr("i", 1),
[ast.Assign(v0("i"), ast.Sub(coords(1, "i"), coords(0, "i"))),
ast.Assign(v1("i"), ast.Sub(coords(2, "i"), coords(0, "i"))),
ast.Assign(x("i"), 0.0)]))
# n = v0 x v1
body.append(ast.Assign(n(0), ast.Sub(ast.Prod(v0(1), v1(2)), ast.Prod(v0(2), v1(1)))))
body.append(ast.Assign(n(1), ast.Sub(ast.Prod(v0(2), v1(0)), ast.Prod(v0(0), v1(2)))))
body.append(ast.Assign(n(2), ast.Sub(ast.Prod(v0(0), v1(1)), ast.Prod(v0(1), v1(0)))))
body.append(ast.For(ast.Assign("i", 0), ast.Less("i", 3), ast.Incr("i", 1),
[ast.Incr(x(j), coords("i", j)) for j in range(3)]))
body.extend([ast.FlatBlock("dot += (%(x)s) * n[%(i)d];\n" % {"x": x_, "i": i})
for i, x_ in enumerate(expr.code)])
body.append(ast.Assign("orientation[0][0]", ast.Ternary(ast.Less("dot", 0), 1, 0)))
kernel = op2.Kernel(ast.FunDecl("void", "cell_orientations",
[ast.Decl("int**", "orientation"),
ast.Decl("double**", "coords")],
ast.Block(body)),
"cell_orientations")
# Build the cell orientations as a DG0 field (so that we can
# pass it in for facet integrals and the like)
fs = functionspace.FunctionSpace(self, 'DG', 0)
cell_orientations = function.Function(fs, name="cell_orientations", dtype=np.int32)
op2.par_loop(kernel, self.cell_set,
cell_orientations.dat(op2.WRITE, cell_orientations.cell_node_map()),
self.coordinates.dat(op2.READ, self.coordinates.cell_node_map()))
self._cell_orientations = cell_orientations
def cells(self):
return self._cells
def ufl_id(self):
return id(self)
def ufl_domain(self):
return self._ufl_domain
def ufl_cell(self):
"""The UFL :class:`~ufl.cell.Cell` associated with the mesh."""
return self._ufl_cell
def num_cells(self):
cStart, cEnd = self._plex.getHeightStratum(0)
return cEnd - cStart
def num_facets(self):
fStart, fEnd = self._plex.getHeightStratum(1)
return fEnd - fStart
def num_faces(self):
fStart, fEnd = self._plex.getDepthStratum(2)
return fEnd - fStart
def num_edges(self):
eStart, eEnd = self._plex.getDepthStratum(1)
return eEnd - eStart
def num_vertices(self):
vStart, vEnd = self._plex.getDepthStratum(0)
return vEnd - vStart
def num_entities(self, d):
eStart, eEnd = self._plex.getDepthStratum(d)
return eEnd - eStart
def size(self, d):
return self.num_entities(d)
@utils.cached_property
def cell_set(self):
size = self.cell_classes
return self.parent.cell_set if self.parent else \
op2.Set(size, "%s_cells" % self.name)
class SimplexMesh(MeshBase):
"""A mesh class providing functionality specific to simplex meshes.
Not part of the public API.
"""
@utils.cached_property
def cell_closure(self):
"""2D array of ordered cell closures
Each row contains ordered cell entities for a cell, one row per cell.
"""
dm = self._plex
a_cell = dm.getHeightStratum(0)[0]
a_closure = dm.getTransitiveClosure(a_cell)[0]
topological_dimension = dm.getDimension()
entity_per_cell = np.zeros(topological_dimension + 1, dtype=np.int32)
for dim in xrange(topological_dimension + 1):
start, end = dm.getDepthStratum(dim)
entity_per_cell[dim] = sum(map(lambda idx: start <= idx < end, a_closure))
return dmplex.closure_ordering(dm, dm.getDefaultGlobalSection(),
self._cell_numbering, entity_per_cell)
def create_cell_node_list(self, global_numbering, fiat_element, dofs_per_cell):
"""Builds the DoF mapping.
:arg global_numbering: Section describing the global DoF numbering
:arg fiat_element: The FIAT element for the cell
:arg dofs_per_cell: Number of DoFs associated with each mesh cell
"""
return dmplex.get_cell_nodes(global_numbering,
self.cell_closure,
dofs_per_cell)
def facet_dimensions(self):
"""Returns a singleton list containing the facet dimension."""
# Facets have co-dimension 1
return [self.ufl_cell().topological_dimension() - 1]
class QuadrilateralMesh(MeshBase):
"""A mesh class providing functionality specific to quadrilateral meshes.
Not part of the public API.
"""
@utils.cached_property
def _closure_ordering(self):
"""Pair of the cell closure and edge directions."""
return dmplex.quadrilateral_closure_ordering(self._plex, self._cell_numbering)
@property
def cell_closure(self):
"""2D array of ordered cell closures
Each row contains ordered cell entities for a cell, one row per cell.
"""
return self._closure_ordering[0]
def create_cell_node_list(self, global_numbering, fiat_element, dofs_per_cell):
"""Builds the DoF mapping.
:arg global_numbering: Section describing the global DoF numbering
:arg fiat_element: The FIAT element for the cell
:arg dofs_per_cell: Number of DoFs associated with each mesh cell
"""
edge_directions = self._closure_ordering[1]
return dmplex.get_quadrilateral_cell_nodes(global_numbering,
self.cell_closure,
edge_directions,
fiat_element,
dofs_per_cell)
def facet_dimensions(self):
"""Returns a list containing the facet dimensions."""
return [(0, 1), (1, 0)]
class ExtrudedMesh(MeshBase):
"""Build an extruded mesh from an input mesh
:arg mesh: the unstructured base mesh
:arg layers: number of extruded cell layers in the "vertical"
direction.
:arg layer_height: the layer height, assuming all layers are evenly
spaced. If this is omitted, the value defaults to
1/layers (i.e. the extruded mesh has total height 1.0)
unless a custom kernel is used.
:arg extrusion_type: the algorithm to employ to calculate the extruded
coordinates. One of "uniform", "radial",
"radial_hedgehog" or "custom". See below.
:arg kernel: a :class:`pyop2.Kernel` to produce coordinates for
the extruded mesh. See :func:`~.make_extruded_coords`
for more details.
:arg gdim: number of spatial dimensions of the
resulting mesh (this is only used if a
custom kernel is provided)
The various values of ``extrusion_type`` have the following meanings:
``"uniform"``
the extruded mesh has an extra spatial
dimension compared to the base mesh. The layers exist
in this dimension only.
``"radial"``
the extruded mesh has the same number of
spatial dimensions as the base mesh; the cells are
radially extruded outwards from the origin. This
requires the base mesh to have topological dimension
strictly smaller than geometric dimension.
``"radial_hedgehog"``
similar to `radial`, but the cells
are extruded in the direction of the outward-pointing
cell normal (this produces a P1dgxP1 coordinate field).
In this case, a radially extruded coordinate field
(generated with ``extrusion_type="radial"``) is
available in the :attr:`radial_coordinates` attribute.
``"custom"``
use a custom kernel to generate the extruded coordinates
For more details see the :doc:`manual section on extruded meshes <extruded-meshes>`.
"""
@timed_function("Build extruded mesh")
@profile
def __init__(self, mesh, layers, layer_height=None, extrusion_type='uniform', kernel=None, gdim=None):
# A cache of function spaces that have been built on this mesh
self._cache = {}
self._old_mesh = mesh
if layers < 1:
raise RuntimeError("Must have at least one layer of extruded cells (not %d)" % layers)
# All internal logic works with layers of base mesh (not layers of cells)
self._layers = layers + 1
self._cells = mesh._cells
self.parent = mesh.parent
self.uid = mesh.uid
self.name = mesh.name
self._plex = mesh._plex
self._plex_renumbering = mesh._plex_renumbering
self._cell_numbering = mesh._cell_numbering
interior_f = self._old_mesh.interior_facets
self._interior_facets = _Facets(self, interior_f.classes,
"interior",
interior_f.facet_cell,
interior_f.local_facet_number)
exterior_f = self._old_mesh.exterior_facets
self._exterior_facets = _Facets(self, exterior_f.classes,
"exterior",
exterior_f.facet_cell,
exterior_f.local_facet_number,
exterior_f.markers)
self.ufl_cell_element = ufl.FiniteElement("Lagrange",
domain=mesh.ufl_cell(),
degree=1)
self.ufl_interval_element = ufl.FiniteElement("Lagrange",
domain=ufl.Cell("interval", 1),
degree=1)
self.fiat_base_element = fiat_utils.fiat_from_ufl_element(self.ufl_cell_element)
self.fiat_vert_element = fiat_utils.fiat_from_ufl_element(self.ufl_interval_element)
fiat_element = FIAT.tensor_finite_element.TensorFiniteElement(self.fiat_base_element, self.fiat_vert_element)
if extrusion_type == "uniform":
# *must* add a new dimension
self._ufl_cell = ufl.OuterProductCell(mesh.ufl_cell(), ufl.Cell("interval", 1), gdim=mesh.ufl_cell().geometric_dimension() + 1)
elif extrusion_type in ("radial", "radial_hedgehog"):
# do not allow radial extrusion if tdim = gdim
if mesh.ufl_cell().geometric_dimension() == mesh.ufl_cell().topological_dimension():
raise RuntimeError("Cannot radially-extrude a mesh with equal geometric and topological dimension")
# otherwise, all is fine, so make cell
self._ufl_cell = ufl.OuterProductCell(mesh.ufl_cell(), ufl.Cell("interval", 1))
else:
# check for kernel
if kernel is None:
raise RuntimeError("If the custom extrusion_type is used, a kernel must be provided")
# otherwise, use the gdim that was passed in
if gdim is None:
raise RuntimeError("The geometric dimension of the mesh must be specified if a custom extrusion kernel is used")
self._ufl_cell = ufl.OuterProductCell(mesh.ufl_cell(), ufl.Cell("interval", 1), gdim=gdim)
self._ufl_domain = ufl.Domain(self.ufl_cell(), data=self)
flat_temp = fiat_element.flattened_element()
# Calculated dofs_per_column from flattened_element and layers.
# The mirrored elements have to be counted only once.
# Then multiply by layers and layers - 1 accordingly.
self.dofs_per_column = eutils.compute_extruded_dofs(fiat_element, flat_temp.entity_dofs(),
layers)
#Compute Coordinates of the extruded mesh
if layer_height is None:
# Default to unit
layer_height = 1.0 / layers
if extrusion_type == 'radial_hedgehog':
hfamily = "DG"
else:
hfamily = mesh.coordinates.element().family()
hdegree = mesh.coordinates.element().degree()
self._coordinate_fs = functionspace.VectorFunctionSpace(self, hfamily,
hdegree,
vfamily="CG",
vdegree=1)
self.coordinates = function.Function(self._coordinate_fs)
self._ufl_domain = ufl.Domain(self.coordinates)
eutils.make_extruded_coords(self, layer_height, extrusion_type=extrusion_type,
kernel=kernel)
if extrusion_type == "radial_hedgehog":
fs = functionspace.VectorFunctionSpace(self, "CG", hdegree, vfamily="CG", vdegree=1)
self.radial_coordinates = function.Function(fs)
eutils.make_extruded_coords(self, layer_height, extrusion_type="radial",
output_coords=self.radial_coordinates)
# Build a new ufl element for this function space with the
# correct domain. This is necessary since this function space
# is in the cache and will be picked up by later
# VectorFunctionSpace construction.
self._coordinate_fs._ufl_element = self._coordinate_fs.ufl_element().reconstruct(domain=self.ufl_domain())
# HACK alert!
# Replace coordinate Function by one that has a real domain on it (but don't copy values)
self.coordinates = function.Function(self._coordinate_fs, val=self.coordinates.dat)
self._dx = ufl.Measure('cell', domain=self, subdomain_data=self.coordinates)
self._ds = ufl.Measure('exterior_facet', domain=self, subdomain_data=self.coordinates)
self._dS = ufl.Measure('interior_facet', domain=self, subdomain_data=self.coordinates)
self._ds_t = ufl.Measure('exterior_facet_top', domain=self, subdomain_data=self.coordinates)
self._ds_b = ufl.Measure('exterior_facet_bottom', domain=self, subdomain_data=self.coordinates)
self._ds_v = ufl.Measure('exterior_facet_vert', domain=self, subdomain_data=self.coordinates)
self._dS_h = ufl.Measure('interior_facet_horiz', domain=self, subdomain_data=self.coordinates)
self._dS_v = ufl.Measure('interior_facet_vert', domain=self, subdomain_data=self.coordinates)
# Set the subdomain_data on all the default measures to this coordinate field.
for measure in [ufl.ds, ufl.dS, ufl.dx, ufl.ds_t, ufl.ds_b, ufl.ds_v, ufl.dS_h, ufl.dS_v]:
measure._subdomain_data = self.coordinates
measure._domain = self.ufl_domain()
@property
def cell_closure(self):
"""2D array of ordered cell closures
Each row contains ordered cell entities for a cell, one row per cell.
"""
return self._old_mesh.cell_closure
def create_cell_node_list(self, global_numbering, fiat_element, dofs_per_cell):
"""Builds the DoF mapping.
:arg global_numbering: Section describing the global DoF numbering
:arg fiat_element: The FIAT element for the cell
:arg dofs_per_cell: Number of DoFs associated with each mesh cell
"""
return dmplex.get_extruded_cell_nodes(self._plex,
global_numbering,
self.cell_closure,
fiat_element,
dofs_per_cell)
@property
def layers(self):
"""Return the number of layers of the extruded mesh
represented by the number of occurences of the base mesh."""
return self._layers
@utils.cached_property
def cell_set(self):
return self.parent.cell_set if self.parent else \
op2.ExtrudedSet(self._old_mesh.cell_set, layers=self._layers)
@property
def exterior_facets(self):
return self._exterior_facets
@property
def interior_facets(self):
return self._interior_facets
@property
def geometric_dimension(self):
return self.ufl_cell().geometric_dimension()
def facet_dimensions(self):
"""Returns a singleton list containing the facet dimension.
.. note::
This only returns the dimension of the "side" (vertical) facets,
not the "top" or "bottom" (horizontal) facets.
"""
# The facet is indexed by (base-ele-codim 1, 1) for
# extruded meshes.
# e.g. for the two supported options of
# triangle x interval interval x interval it's (1, 1) and
# (0, 1) respectively.
if self.geometric_dimension == 3:
return [(1, 1)]
elif self.geometric_dimension == 2:
return [(0, 1)]
else:
raise RuntimeError("Dimension computation for other than 2D or 3D extruded meshes not supported.")
<file_sep>/tests/extrusion/test_laplace_neumann.py
"""This demo program solves Laplace's equation
- div grad u(x, y, z) = 0
in a unit square or unit cube, with Dirichlet boundary
conditions on 2/4 sides and Neumann boundary conditions
on the other 2, opposite, sides.
"""
import pytest
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module')
def P2():
mesh = extmesh(4, 4, 4)
return FunctionSpace(mesh, "CG", 2)
@pytest.fixture(scope='module')
def P2_2D():
mesh = extmesh_2D(4, 4)
return FunctionSpace(mesh, "CG", 2)
def test_bottom_and_top(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = 10*v*ds_b - 10*v*ds_t
bc_expr = Expression("-10*x[2]")
bcs = [DirichletBC(P2, bc_expr, 1),
DirichletBC(P2, bc_expr, 2),
DirichletBC(P2, bc_expr, 3),
DirichletBC(P2, bc_expr, 4)]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.1e-6
def test_top_and_bottom(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = 10*v*ds_t - 10*v*ds_b
bc_expr = Expression("10*x[2]")
bcs = [DirichletBC(P2, bc_expr, 1),
DirichletBC(P2, bc_expr, 2),
DirichletBC(P2, bc_expr, 3),
DirichletBC(P2, bc_expr, 4)]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.1e-6
def test_left_right(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = 10*v*ds_v(2) - 10*v*ds_v(1)
bc_expr = Expression("10*x[0]")
bcs = [DirichletBC(P2, bc_expr, "top"),
DirichletBC(P2, bc_expr, "bottom"),
DirichletBC(P2, bc_expr, 3),
DirichletBC(P2, bc_expr, 4)]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.1e-6
def test_near_far(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = 10*v*ds_v(4) - 10*v*ds_v(3)
bc_expr = Expression("10*x[1]")
bcs = [DirichletBC(P2, bc_expr, 1),
DirichletBC(P2, bc_expr, 2),
DirichletBC(P2, bc_expr, "top"),
DirichletBC(P2, bc_expr, "bottom")]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.0e-6
def test_2D_bottom_top(P2_2D):
u = TrialFunction(P2_2D)
v = TestFunction(P2_2D)
a = dot(grad(u), grad(v))*dx
L = 10*v*ds_t - 10*v*ds_b
bc_expr = Expression("10*x[1]")
bcs = [DirichletBC(P2_2D, bc_expr, 1),
DirichletBC(P2_2D, bc_expr, 2)]
u = Function(P2_2D)
solve(a == L, u, bcs)
u_exact = Function(P2_2D)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.0e-6
def test_2D_left_right(P2_2D):
u = TrialFunction(P2_2D)
v = TestFunction(P2_2D)
a = dot(grad(u), grad(v))*dx
L = 10*v*ds_v(2) - 10*v*ds_v(1)
bc_expr = Expression("10*x[0]")
bcs = [DirichletBC(P2_2D, bc_expr, "top"),
DirichletBC(P2_2D, bc_expr, "bottom")]
u = Function(P2_2D)
solve(a == L, u, bcs)
u_exact = Function(P2_2D)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.0e-6
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_mixed_bcs.py
from firedrake import *
import pytest
@pytest.mark.parametrize('degree', [1, 2, 3])
def test_multiple_poisson_Pn(degree):
m = UnitSquareMesh(4, 4)
mesh = ExtrudedMesh(m, 4)
V = FunctionSpace(mesh, 'CG', degree)
W = V*V
w = Function(W)
u, p = split(w)
v, q = TestFunctions(W)
# Solve 2 independent Poisson problems with strong boundary
# conditions applied to the top and bottom for the first and on x
# == 0 and x == 1 for the second.
a = dot(grad(u), grad(v))*dx + dot(grad(p), grad(q))*dx
# BCs for first problem
bc0 = [DirichletBC(W[0], 10.0, "top"),
DirichletBC(W[0], 1.0, "bottom")]
# BCs for second problem
bc1 = [DirichletBC(W[1], 8.0, 1),
DirichletBC(W[1], 6.0, 2)]
bcs = bc0 + bc1
solve(a == 0, w, bcs=bcs,
# Operator is block diagonal, so we can just do block jacobi
# with lu on each block
solver_parameters={'ksp_type': 'cg',
'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'additive',
'fieldsplit_ksp_type': 'preonly',
'fieldsplit_0_pc_type': 'lu',
'fieldsplit_1_pc_type': 'lu'})
wexact = Function(W)
u, p = wexact.split()
u.interpolate(Expression("1.0 + 9*x[2]"))
p.interpolate(Expression("8.0 - 2*x[0]"))
assert assemble(inner(w - wexact, w - wexact)*dx) < 1e-8
@pytest.mark.parametrize('degree', [1, 2, 3])
def test_multiple_poisson_strong_weak_Pn(degree):
m = UnitSquareMesh(4, 4)
mesh = ExtrudedMesh(m, 4)
V = FunctionSpace(mesh, 'CG', degree)
W = V*V
w = Function(W)
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
# Solve two independent Poisson problems with a strong boundary
# condition on the top and a weak condition on the bottom, and
# vice versa.
a = dot(grad(u), grad(v))*dx + dot(grad(p), grad(q))*dx
L = Constant(1)*v*ds_b + Constant(4)*q*ds_t
# BCs for first problem
bc0 = [DirichletBC(W[0], 10.0, "top")]
# BCs for second problem
bc1 = [DirichletBC(W[1], 2.0, "bottom")]
bcs = bc0 + bc1
solve(a == L, w, bcs=bcs,
# Operator is block diagonal, so we can just do block jacobi
# with lu on each block
solver_parameters={'ksp_type': 'cg',
'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'additive',
'fieldsplit_ksp_type': 'preonly',
'fieldsplit_0_pc_type': 'lu',
'fieldsplit_1_pc_type': 'lu'})
wexact = Function(W)
u, p = wexact.split()
u.interpolate(Expression("11.0 - x[2]"))
p.interpolate(Expression("2.0 + 4*x[2]"))
assert assemble(inner(w - wexact, w - wexact)*dx) < 1e-8
def test_stokes_taylor_hood():
length = 10
m = IntervalMesh(40, length)
mesh = ExtrudedMesh(m, 20)
V = VectorFunctionSpace(mesh, 'CG', 2)
P = FunctionSpace(mesh, 'CG', 1)
W = V*P
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
a = inner(grad(u), grad(v))*dx - div(v)*p*dx + q*div(u)*dx
f = Constant((0, 0))
L = inner(f, v)*dx
# No-slip velocity boundary condition on top and bottom,
# y == 0 and y == 1
noslip = Constant((0, 0))
bc0 = [DirichletBC(W[0], noslip, "top"),
DirichletBC(W[0], noslip, "bottom")]
# Parabolic inflow y(1-y) at x = 0 in positive x direction
inflow = Expression(("x[1]*(1 - x[1])", "0.0"))
bc1 = DirichletBC(W[0], inflow, 1)
# Zero pressure at outlow at x = 1
bc2 = DirichletBC(W[1], 0.0, 2)
bcs = bc0 + [bc1, bc2]
w = Function(W)
u, p = w.split()
solve(a == L, w, bcs=bcs,
solver_parameters={'pc_type': 'fieldsplit',
'ksp_rtol': 1e-15,
'pc_fieldsplit_type': 'schur',
'fieldsplit_schur_fact_type': 'diag',
'fieldsplit_0_pc_type': 'lu',
'fieldsplit_1_pc_type': 'none'})
# We've set up Poiseuille flow, so we expect a parabolic velocity
# field and a linearly decreasing pressure.
uexact = Function(V).interpolate(Expression(("x[1]*(1 - x[1])", "0.0")))
pexact = Function(P).interpolate(Expression("2*(L - x[0])", L=length))
assert errornorm(u, uexact, degree_rise=0) < 1e-7
assert errornorm(p, pexact, degree_rise=0) < 1e-7
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_poisson_mixed_strong_bcs.py
"""Solve the mixed formulation of the Laplacian on the unit square
sigma - grad(u) = 0
div(sigma) = f
The corresponding weak (variational problem)
<sigma, tau> + <div(tau), u> = 0 for all tau
<div(sigma), v> = <f, v> for all v
is solved using BDM (Brezzi-Douglas-Marini) elements of degree k for
(sigma, tau) and DG (discontinuous Galerkin) elements of degree k - 1
for (u, v).
The boundary conditions on the left and right are enforced strongly as
dot(sigma, n) = 0
which corresponds to a Neumann condition du/dn = 0.
The top is fixed to 42 with a Dirichlet boundary condition, which enters
the weak formulation of the right hand side as
42*dot(tau, n)*ds
"""
import pytest
from firedrake import *
def poisson_mixed(size, parameters={}):
# Create mesh
mesh = UnitSquareMesh(2 ** size, 2 ** size)
# Define function spaces and mixed (product) space
BDM = FunctionSpace(mesh, "BDM", 1)
DG = FunctionSpace(mesh, "DG", 0)
W = BDM * DG
# Define trial and test functions
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
# Define source function
f = Function(DG).assign(0)
# Define variational form
a = (dot(sigma, tau) + div(tau)*u + div(sigma)*v)*dx
n = FacetNormal(mesh)
L = -f*v*dx + 42*dot(tau, n)*ds(4)
# Apply dot(sigma, n) == 0 on left and right boundaries strongly
# (corresponding to Neumann condition du/dn = 0)
bcs = DirichletBC(W.sub(0), Expression(('0', '0')), (1, 2))
# Compute solution
w = Function(W)
solve(a == L, w, bcs=bcs, solver_parameters=parameters)
sigma, u = w.split()
# Analytical solution
f.interpolate(Expression("42*x[1]"))
return sqrt(assemble(dot(u - f, u - f) * dx)), u, f
@pytest.mark.parametrize('parameters',
[{}, {'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'ksp_type': 'gmres',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_0_pc_factor_shift_type': 'INBLOCKS',
'fieldsplit_1_pc_factor_shift_type': 'INBLOCKS',
'fieldsplit_0_ksp_type': 'cg',
'fieldsplit_1_ksp_type': 'cg'}])
def test_poisson_mixed(parameters):
assert poisson_mixed(3, parameters)[0] < 2e-5
@pytest.mark.parallel(nprocs=3)
def test_poisson_mixed_parallel_fieldsplit():
x = poisson_mixed(3, parameters={'pc_type': 'fieldsplit',
'snes_type': 'ksponly',
'snes_monitor': True,
'pc_fieldsplit_type': 'schur',
'fieldsplit_schur_fact_type': 'full',
'fieldsplit_0_ksp_type': 'cg',
'fieldsplit_1_ksp_type': 'cg',
'fieldsplit_0_pc_type': 'bjacobi',
'fieldsplit_0_sub_pc_type': 'ilu',
'fieldsplit_1_pc_type': 'none',
'ksp_type': 'bcgs'})[0]
assert x < 2e-5
@pytest.mark.parallel(nprocs=3)
def test_poisson_mixed_parallel():
x = poisson_mixed(3)[0]
assert x < 2e-5
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_kernel_intvar_p0.py
import pytest
import numpy as np
from firedrake import *
import pyop2 as op2
def integrate_var_p0(family, degree):
power = 5
m = UnitSquareMesh(2 ** power, 2 ** power)
layers = 10
# Populate the coordinates of the extruded mesh by providing the
# coordinates as a field.
# TODO: provide a kernel which will describe how coordinates are extruded.
mesh = ExtrudedMesh(m, layers, layer_height=0.1)
fs = FunctionSpace(mesh, family, degree, name="fs")
f = Function(fs)
populate_p0 = op2.Kernel("""
void populate_tracer(double *x[], double *c[])
{
x[0][0] = (c[1][2] + c[0][2]) / 2;
}""", "populate_tracer")
coords = f.function_space().mesh().coordinates
op2.par_loop(populate_p0, f.cell_set,
f.dat(op2.INC, f.cell_node_map()),
coords.dat(op2.READ, coords.cell_node_map()))
volume = op2.Kernel("""
void comp_vol(double A[1], double *x[], double *y[])
{
double area = x[0][0]*(x[2][1]-x[4][1]) + x[2][0]*(x[4][1]-x[0][1])
+ x[4][0]*(x[0][1]-x[2][1]);
if (area < 0)
area = area * (-1.0);
A[0] += 0.5 * area * (x[1][2] - x[0][2]) * y[0][0];
}""", "comp_vol")
g = op2.Global(1, data=0.0, name='g')
op2.par_loop(volume, f.cell_set,
g(op2.INC),
coords.dat(op2.READ, coords.cell_node_map()),
f.dat(op2.READ, f.cell_node_map())
)
return np.abs(g.data[0] - 0.5)
def test_firedrake_extrusion_var_p0():
family = "DG"
degree = 0
assert integrate_var_p0(family, degree) < 1.0e-14
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/docs/source/publications.rst
Firedrake and PyOP2 publications
================================
Some useful slides from a talk on PyOP2 and Firedrake at Compilers for Parallel Computers are
`here <http://florianrathgeber.me/CPC2013/>`_.
Journal papers and conference proceedings about or using Firedrake
------------------------------------------------------------------
.. raw:: html
<script src="http://www.bibbase.org/show?bib=http%3A%2F%2Fwww.firedrakeproject.org%2F_static%2Fbibliography.bib&jsonp=1"></script>
<file_sep>/tests/extrusion/test_extrusion_rhs_side_bcs.py
"""This demo program sets opposite boundary sides to 10 and 42 and
then checks that the exact result has bee achieved.
"""
import pytest
from firedrake import *
def run_test(x, degree, parameters={}, test_mode=False):
# Create mesh and define function space
m = UnitSquareMesh(3, 3)
layers = 10
mesh = ExtrudedMesh(m, layers, layer_height=1.0 / layers)
# Define variational problem
V = FunctionSpace(mesh, "CG", degree)
# Define variational problem
u = Function(V)
bcs = [DirichletBC(V, 10, 1),
DirichletBC(V, 42, 2)]
for bc in bcs:
bc.apply(u)
v = Function(V)
v.interpolate(Expression("x[0] < 0.05 ? 10.0 : x[0] > 0.95 ? 42.0 : 0.0"))
res = sqrt(assemble(dot(u - v, u - v) * dx))
u1 = Function(V)
bcs1 = [DirichletBC(V, 10, 3),
DirichletBC(V, 42, 4)]
for bc in bcs1:
bc.apply(u1)
v1 = Function(V)
v1.interpolate(Expression("x[1] < 0.05 ? 10.0 : x[1] > 0.95 ? 42.0 : 0.0"))
res1 = sqrt(assemble(dot(u1 - v1, u1 - v1) * dx))
if not test_mode:
print "The error is ", res1
file = File("side-bcs-computed.pvd")
file << u1
file = File("side-bcs-expected.pvd")
file << v1
return (res, res1)
def test_extrusion_rhs_bcs():
res1, res2 = run_test(1, 1, test_mode=True)
assert (res1 < 1.e-13 and res2 < 1.e-13)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_strong_bcs_caching.py
from tests.common import extmesh
import numpy as np
from firedrake import *
def test_extrusion_strong_bcs_caching():
m = extmesh(1, 1, 1)
V = FunctionSpace(m, "CG", 1)
bc1 = DirichletBC(V, 0, "bottom")
bc2 = DirichletBC(V, 1, "top")
v = TestFunction(V)
u = TrialFunction(V)
a = u*v*dx
Aboth = assemble(a, bcs=[bc1, bc2])
Aneither = assemble(a)
Abottom = assemble(a, bcs=[bc1])
Atop = assemble(a, bcs=[bc2])
# None of the matrices should be the same
assert not np.allclose(Aboth.M.values, Aneither.M.values)
assert not np.allclose(Aboth.M.values, Atop.M.values)
assert not np.allclose(Aboth.M.values, Abottom.M.values)
assert not np.allclose(Aneither.M.values, Atop.M.values)
assert not np.allclose(Aneither.M.values, Abottom.M.values)
assert not np.allclose(Atop.M.values, Abottom.M.values)
# There should be no zeros on the diagonal
assert not any(Atop.M.values.diagonal() == 0)
assert not any(Abottom.M.values.diagonal() == 0)
assert not any(Aneither.M.values.diagonal() == 0)
# The top/bottom case should just be the identity (since the only
# dofs live on the top and bottom)
assert np.allclose(Aboth.M.values, np.diag(np.ones_like(Aboth.M.values.diagonal())))
<file_sep>/firedrake/parloops.py
"""This module implements parallel loops reading and writing
:class:`.Function`\s. This provides a mechanism for implementing
non-finite element operations such as slope limiters."""
from ufl.indexed import Indexed
from pyop2 import READ, WRITE, RW, INC # NOQA get flake8 to ignore unused import.
import pyop2
import pyop2.coffee.ast_base as ast
import constant
__all__ = ['par_loop', 'direct', 'READ', 'WRITE', 'RW', 'INC']
class _DirectLoop(object):
"""A singleton object which can be used in a :func:`par_loop` in place
of the measure in order to indicate that the loop is a direct loop
over degrees of freedom."""
def integral_type(self):
return "direct"
def __repr__(self):
return "direct"
direct = _DirectLoop()
"""A singleton object which can be used in a :func:`par_loop` in place
of the measure in order to indicate that the loop is a direct loop
over degrees of freedom."""
"""Map a measure to the correct maps."""
_maps = {
'cell': {
'nodes': lambda x: x.cell_node_map(),
'itspace': lambda mesh, measure: mesh.cell_set
},
'interior_facet': {
'nodes': lambda x: x.interior_facet_node_map(),
'itspace': lambda mesh, measure: mesh.interior_facets.measure_set(measure.integral_type(), measure.subdomain_id())
},
'exterior_facet': {
'nodes': lambda x: x.exterior_facet_node_map(),
'itspace': lambda mesh, measure: mesh.exterior_facets.measure_set(measure.integral_type(), measure.subdomain_id())
},
'direct': {
'nodes': lambda x: None,
'itspace': lambda mesh, measure: mesh
}
}
def _form_kernel(kernel, measure, args, **kwargs):
kargs = []
lkernel = kernel
for var, (func, intent) in args.iteritems():
if isinstance(func, constant.Constant):
if intent is not READ:
raise RuntimeError("Only READ access is allowed to Constant")
# Constants modelled as Globals, so no need for double
# indirection
ndof = func.dat.cdim
kargs.append(ast.Decl("double", ast.Symbol(var, (ndof, )),
qualifiers=["const"]))
else:
# Do we have a component of a mixed function?
if isinstance(func, Indexed):
c, i = func.operands()
idx = i._indices[0]._value
ndof = c.function_space()[idx].fiat_element.space_dimension()
else:
ndof = func.function_space().fiat_element.space_dimension()
if measure.integral_type() == 'interior_facet':
ndof *= 2
if measure is direct:
kargs.append(ast.Decl("double", ast.Symbol(var, (ndof,))))
else:
kargs.append(ast.Decl("double *", ast.Symbol(var, (ndof,))))
lkernel = lkernel.replace(var+".dofs", str(ndof))
body = ast.FlatBlock(lkernel)
return pyop2.Kernel(ast.FunDecl("void", "par_loop_kernel", kargs, body),
"par_loop_kernel", **kwargs)
def par_loop(kernel, measure, args, **kwargs):
"""A :func:`par_loop` is a user-defined operation which reads and
writes :class:`.Function`\s by looping over the mesh cells or facets
and accessing the degrees of freedom on adjacent entities.
:arg kernel: is a string containing the C code to be executed.
:arg measure: is a UFL :class:`~ufl.measure.Measure` which determines the
manner in which the iteration over the mesh is to occur.
Alternatively, you can pass :data:`direct` to designate a direct loop.
:arg args: is a dictionary mapping variable names in the kernel to
:class:`.Function`\s or components of mixed :class:`.Function`\s and
indicates how these :class:`.Function`\s are to be accessed.
:arg kwargs: additional keyword arguments are passed to the
:class:`~pyop2.op2.Kernel` constructor
**Example**
Assume that `A` is a :class:`.Function` in CG1 and `B` is a
:class:`.Function` in DG0. Then the following code sets each DoF in
`A` to the maximum value that `B` attains in the cells adjacent to
that DoF::
A.assign(numpy.finfo(0.).min)
par_loop('for (int i=0; i<A.dofs; i++;) A[i][0] = fmax(A[i][0], B[0][0]);', dx,
{'A' : (A, RW), 'B': (B, READ)})
**Argument definitions**
Each item in the `args` dictionary maps a string to a tuple
containing a :class:`.Function` or :class:`.Constant` and an
argument intent. The string is the c language variable name by
which this function will be accessed in the kernel. The argument
intent indicates how the kernel will access this variable:
`READ`
The variable will be read but not written to.
`WRITE`
The variable will be written to but not read. If multiple kernel
invocations write to the same DoF, then the order of these writes
is undefined.
`RW`
The variable will be both read and written to. If multiple kernel
invocations access the same DoF, then the order of these accesses
is undefined, but it is guaranteed that no race will occur.
`INC`
The variable will be added into using +=. As before, the order in
which the kernel invocations increment the variable is undefined,
but there is a guarantee that no races will occur.
.. note::
Only `READ` intents are valid for :class:`.Constant`
coefficients, and an error will be raised in other cases.
**The measure**
The measure determines the mesh entities over which the iteration
will occur, and the size of the kernel stencil. The iteration will
occur over the same mesh entities as if the measure had been used
to define an integral, and the stencil will likewise be the same
as the integral case. That is to say, if the measure is a volume
measure, the kernel will be called once per cell and the DoFs
accessible to the kernel will be those associated with the cell,
its facets, edges and vertices. If the measure is a facet measure
then the iteration will occur over the corresponding class of
facets and the accessible DoFs will be those on the cell(s)
adjacent to the facet, and on the facets, edges and vertices
adjacent to those facets.
For volume measures the DoFs are guaranteed to be in the FIAT
local DoFs order. For facet measures, the DoFs will be in sorted
first by the cell to which they are adjacent. Within each cell,
they will be in FIAT order. Note that if a continuous
:class:`.Function` is accessed via an internal facet measure, the
DoFs on the interface between the two facets will be accessible
twice: once via each cell. The orientation of the cell(s) relative
to the current facet is currently arbitrary.
A direct loop over nodes without any indirections can be specified
by passing :data:`direct` as the measure. In this case, all of the
arguments must be :class:`.Function`\s in the same
:class:`.FunctionSpace` or in the corresponding
:class:`.VectorFunctionSpace`.
**The kernel code**
The kernel code is plain C in which the variables specified in the
`args` dictionary are available to be read or written in according
to the argument intent specified. Most basic C operations are
permitted. However there are some restrictions:
* Only functions from `math.h
<http://pubs.opengroup.org/onlinepubs/9699919799/basedefs/math.h.html>`_
may be called.
* Pointer operations other than dereferencing arrays are prohibited.
Indirect free variables referencing :class:`.Function`\s are all
of type `double**` in which the first index is the local node
number, while the second index is the vector component. The latter
only applies to :class:`.Function`\s over a
:class:`.VectorFunctionSpace`, for :class:`.Function`\s over a
plain :class:`.FunctionSpace` the second index will always be 0.
In a direct :func:`par_loop`, the variables will all be of type
`double*` with the single index being the vector component.
:class:`.Constant`\s are always of type `double*`, both for
indirect and direct :func:`par_loop` calls.
"""
_map = _maps[measure.integral_type()]
if measure is direct:
mesh = None
for (func, intent) in args.itervalues():
if isinstance(func, Indexed):
c, i = func.operands()
idx = i._indices[0]._value
if mesh and c.node_set[idx] is not mesh:
raise ValueError("Cannot mix sets in direct loop.")
mesh = c.node_set[idx]
else:
try:
if mesh and func.node_set is not mesh:
raise ValueError("Cannot mix sets in direct loop.")
mesh = func.node_set
except AttributeError:
# Argument was a Global.
pass
if not mesh:
raise TypeError("No Functions passed to direct par_loop")
else:
mesh = measure.subdomain_data().function_space().mesh()
op2args = [_form_kernel(kernel, measure, args, **kwargs)]
op2args.append(_map['itspace'](mesh, measure))
def mkarg(f, intent):
if isinstance(func, Indexed):
c, i = func.operands()
idx = i._indices[0]._value
m = _map['nodes'](c)
return c.dat[idx](intent, m.split[idx] if m else None)
return f.dat(intent, _map['nodes'](f))
op2args += [mkarg(func, intent) for (func, intent) in args.itervalues()]
return pyop2.par_loop(*op2args)
<file_sep>/tests/regression/test_parameters.py
from firedrake import *
# These Parameter tests are a cut down version of the unit Parameter tests in DOLFIN
def test_simple():
# Create some parameters
p0 = Parameters("test")
p0.add("filename", "foo.txt")
p0.add("maxiter", 100)
p0.add("tolerance", 0.001)
p0.add("monitor_convergence", True)
# Check values
assert p0.name() == "test"
assert p0["filename"] == "foo.txt"
assert p0["maxiter"] == 100
assert p0["tolerance"] == 0.001
assert p0["monitor_convergence"] is True
def test_nested():
# Create some nested parameters
p0 = Parameters("test")
p00 = Parameters("sub0")
p00.add("filename", "foo.txt")
p00.add("maxiter", 100)
p00.add("tolerance", 0.001)
p00.add("monitor_convergence", True)
p0.add("foo", "bar")
p01 = Parameters(p00)
p01.rename("sub1")
p0.add(p00)
p0.add(p01)
# Check values
assert p0.name() == "test"
assert p0["foo"] == "bar"
assert p0["sub0"]["filename"] == "foo.txt"
assert p0["sub0"]["maxiter"] == 100
assert p0["sub0"]["tolerance"] == 0.001
assert p0["sub0"]["monitor_convergence"] is True
if __name__ == "__main__":
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/README.rst
Firedrake is an automated system for the portable solution of partial
differential equations using the finite element method (FEM). Firedrake
enables users to employ a wide range of discretisations to an infinite
variety of PDEs and employ either conventional CPUs or GPUs to obtain
the solution.
For more information on Firedrake, please see http://www.firedrakeproject.org.
<file_sep>/docs/source/index.rst
.. title:: The Firedrake project
.. only:: html
.. sidebar:: Latest commits to the Firedrake master branch on Github
.. raw:: html
<div class="latest-commit" data-github="firedrakeproject/firedrake" data-commits="5"></div>
<script type="text/javascript" src="_static/jquery.latest-commit.js"></script>
.. only:: latex
Introduction
------------
Firedrake is an automated system for the portable solution of partial
differential equations using the finite element method (FEM). Firedrake
enables users to employ a wide range of discretisations to an infinite
variety of PDEs and employ either conventional CPUs or GPUs to obtain
the solution.
Firedrake employs the Unifed Form Language (UFL) and FEniCS Form
Compiler (FFC) from `the FEniCS Project <http://fenicsproject.org>`_
while the parallel execution of FEM assembly is accomplished by the
`PyOP2 <http://op2.github.io/PyOP2/>`_ system. The global mesh data
structures, as well as linear and non-linear solvers, are provided by
`PETSc <https://www.mcs.anl.gov/petsc/>`_.
.. only:: html
.. container:: youtube
.. youtube:: xhxvM1N8mDQ?modestbranding=1;controls=0;rel=0
:width: 400px
.. only:: latex
.. toctree::
documentation
firedrake
funding
team
<file_sep>/tests/extrusion/test_extrusion_3_galerkinproj.py
"""Tests for Galerkin projection convergence on extruded meshes"""
import numpy as np
import pytest
from firedrake import *
from tests.common import *
@pytest.mark.parametrize(('testcase', 'convrate'),
[(("CG", 1), 1.5), (("CG", 2), 2.6),
(("DG", 0), 0.9), (("DG", 1), 1.7)])
def test_scalar_convergence(testcase, convrate):
family, degree = testcase
l2err = np.zeros(2)
for ii in range(len(l2err)):
mesh = extmesh(2**(ii+1), 2**(ii+1), 2**ii)
fspace = FunctionSpace(mesh, family, degree, vfamily=family, vdegree=degree)
exactfspace = FunctionSpace(mesh, "Lagrange", 3)
u = TrialFunction(fspace)
v = TestFunction(fspace)
expr = Expression("x[0]*x[0]*x[1]*x[2]")
exact = project(expr, exactfspace)
out = Function(fspace)
solve(u*v*dx == exact*v*dx, out)
l2err[ii] = sqrt(assemble((out-exact)*(out-exact)*dx))
assert (np.array([np.log2(l2err[i]/l2err[i+1]) for i in range(len(l2err)-1)]) > convrate).all()
@pytest.mark.parametrize(('testcase', 'convrate'),
[(("RT", 1, "DG", 0, "h"), 0.9),
(("RT", 2, "DG", 1, "h"), 1.94),
(("RT", 3, "DG", 2, "h"), 2.9),
(("BDM", 1, "DG", 1, "h"), 1.8),
(("BDM", 2, "DG", 2, "h"), 2.8),
(("BDFM", 2, "DG", 1, "h"), 1.95),
(("N1curl", 1, "DG", 0, "h"), 0.9),
(("N1curl", 2, "DG", 1, "h"), 1.9),
(("N2curl", 1, "DG", 1, "h"), 1.8),
(("N2curl", 2, "DG", 2, "h"), 2.85),
(("DG", 1, "CG", 1, "v"), 1.84),
(("DG", 2, "CG", 2, "v"), 2.98)])
def test_hdiv_convergence(testcase, convrate):
hfamily, hdegree, vfamily, vdegree, orientation = testcase
l2err = np.zeros(2)
for ii in range(len(l2err)):
mesh = extmesh(2**(ii+1), 2**(ii+1), 2**(ii+1))
exactfspace = VectorFunctionSpace(mesh, "Lagrange", 3)
horiz_elt = FiniteElement(hfamily, "triangle", hdegree)
vert_elt = FiniteElement(vfamily, "interval", vdegree)
product_elt = HDiv(OuterProductElement(horiz_elt, vert_elt))
fspace = FunctionSpace(mesh, product_elt)
u = TrialFunction(fspace)
v = TestFunction(fspace)
if orientation == "h":
expr = Expression(("x[0]*x[1]*x[2]*x[2]", "x[0]*x[0]*x[1]*x[2]", "0.0"))
elif orientation == "v":
expr = Expression(("0.0", "0.0", "x[0]*x[1]*x[1]*x[2]"))
exact = Function(exactfspace)
exact.interpolate(expr)
out = Function(fspace)
solve(dot(u, v)*dx == dot(exact, v)*dx, out)
l2err[ii] = sqrt(assemble(dot((out-exact), (out-exact))*dx))
assert (np.array([np.log2(l2err[i]/l2err[i+1]) for i in range(len(l2err)-1)]) > convrate).all()
@pytest.mark.parametrize(('testcase', 'convrate'),
[(("BDM", 1, "CG", 1, "h"), 1.82),
(("BDM", 2, "CG", 2, "h"), 2.9),
(("RT", 2, "CG", 1, "h"), 1.87),
(("RT", 3, "CG", 2, "h"), 2.95),
(("BDFM", 2, "CG", 1, "h"), 1.77),
(("N1curl", 2, "CG", 1, "h"), 1.87),
(("N2curl", 1, "CG", 1, "h"), 1.82),
(("N2curl", 2, "CG", 2, "h"), 2.9),
(("CG", 1, "DG", 1, "v"), 1.6),
(("CG", 2, "DG", 2, "v"), 2.7)])
def test_hcurl_convergence(testcase, convrate):
hfamily, hdegree, vfamily, vdegree, orientation = testcase
l2err = np.zeros(2)
for ii in range(len(l2err)):
mesh = extmesh(2**(ii+1), 2**(ii+1), 2**(ii+1))
exactfspace = VectorFunctionSpace(mesh, "Lagrange", 3)
horiz_elt = FiniteElement(hfamily, "triangle", hdegree)
vert_elt = FiniteElement(vfamily, "interval", vdegree)
product_elt = HCurl(OuterProductElement(horiz_elt, vert_elt))
fspace = FunctionSpace(mesh, product_elt)
u = TrialFunction(fspace)
v = TestFunction(fspace)
if orientation == "h":
expr = Expression(("x[0]*x[1]*x[2]*x[2]", "x[0]*x[0]*x[1]*x[2]", "0.0"))
elif orientation == "v":
expr = Expression(("0.0", "0.0", "x[0]*x[1]*x[1]*x[2]"))
exact = Function(exactfspace)
exact.interpolate(expr)
out = Function(fspace)
solve(dot(u, v)*dx == dot(exact, v)*dx, out)
l2err[ii] = sqrt(assemble(dot((out-exact), (out-exact))*dx))
assert (np.array([np.log2(l2err[i]/l2err[i+1]) for i in range(len(l2err)-1)]) > convrate).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_kernel_intrhs.py
import pytest
import numpy as np
from firedrake import *
import pyop2 as op2
import ufl
def integrate_rhs(family, degree):
power = 5
m = UnitSquareMesh(2 ** power, 2 ** power)
layers = 10
# Populate the coordinates of the extruded mesh by providing the
# coordinates as a field.
# TODO: provide a kernel which will describe how coordinates are extruded.
mesh = ExtrudedMesh(m, layers, layer_height=0.1)
horiz = ufl.FiniteElement(family, "triangle", degree)
vert = ufl.FiniteElement(family, "interval", degree)
prod = ufl.OuterProductElement(horiz, vert)
fs = FunctionSpace(mesh, prod, name="fs")
f = Function(fs)
populate_p0 = op2.Kernel("""
void populate_tracer(double *x[], double *c[])
{
x[0][0] = ((c[1][2] + c[0][2]) / 2);
}""", "populate_tracer")
coords = f.function_space().mesh().coordinates
op2.par_loop(populate_p0, f.cell_set,
f.dat(op2.INC, f.cell_node_map()),
coords.dat(op2.READ, coords.cell_node_map()))
g = assemble(f * dx)
return np.abs(g - 0.5)
def test_firedrake_extrusion_rhs():
family = "DG"
degree = 0
assert integrate_rhs(family, degree) < 1.0e-14
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/docs/source/funding.rst
.. only:: html
Firedrake is supported by:
--------------------------
.. only:: latex
Funding
=======
.. |NERC| image:: /images/nerc.*
:height: 60px
:target: http://www.nerc.ac.uk
.. |EPSRC| image:: /images/epsrc.*
:height: 60px
:target: http://www.epsrc.ac.uk
.. |Imperial College London| image:: /images/imperial.*
:height: 60px
:target: http://www.imperial.ac.uk
.. |Grantham| image:: /images/grantham.*
:height: 60px
:target: http://www.imperial.ac.uk/climatechange
.. only:: html
+---------------------------+------------+---------+--------+
| |Imperial College London| | |Grantham| | |EPSRC| | |NERC| |
+---------------------------+------------+---------+--------+
and in particular by the following grants:
.. only:: latex
Firedrake is supported by Imperial College London, the Grantham
Institute for Climate Change, the Engineering and Physical Sciences
Research Council and the Natural Environment research council.
NERC fellowship `NE/K008951/1 <http://gtr.rcuk.ac.uk/project/10179C8D-1FE9-48C1-AC82-8D549D6EF8F5>`_:
Abstracting the environment: automating geoscientific simulation
NERC grant `NE/K006789/1 <http://gtr.rcuk.ac.uk/project/68AD0B6D-91D1-45D5-9C8A-991518BF028E>`_:
Gung Ho Phase 2 - developing a new UK weather and climate model.
EPSRC grant `EP/L000407/1 <http://gow.epsrc.ac.uk/NGBOViewGrant.aspx?GrantRef=EP/L000407/1>`_:
Platform: underpinning technologies for finite element simulation.
EPSRC Grant `EP/K008730/1 <http://gow.epsrc.ac.uk/NGBOViewGrant.aspx?GrantRef=EP/K008730/1>`_:
PAMELA: a Panoramic Approach to the Many-CorE LAnd-sape - from end-user to end-device: a holistic game-changing approach.
EPSRC Grant `EP/I00677X/1 <http://gow.epsrc.ac.uk/NGBOViewGrant.aspx?GrantRef=EP/I00677X/1>`_:
Multi-layered abstractions for PDEs.
NERC studentship `NE/G523512/1 <http://gtr.rcuk.ac.uk/project/C997B5F6-99AF-45E1-8ED3-9DE2BD0DD964>`_:
Multi-layered abstractions for PDEs.
<file_sep>/tests/regression/test_identity.py
import numpy as np
import pytest
from firedrake import *
def identity(family, degree):
mesh = UnitCubeMesh(1, 1, 1)
fs = FunctionSpace(mesh, family, degree)
f = Function(fs)
out = Function(fs)
u = TrialFunction(fs)
v = TestFunction(fs)
a = u * v * dx
f.interpolate(Expression("x[0]"))
L = f * v * dx
solve(a == L, out)
return np.max(np.abs(out.dat.data - f.dat.data))
def vector_identity(family, degree):
mesh = UnitSquareMesh(2, 2)
fs = VectorFunctionSpace(mesh, family, degree)
f = Function(fs)
out = Function(fs)
u = TrialFunction(fs)
v = TestFunction(fs)
f.interpolate(Expression(("x[0]", "x[1]")))
solve(inner(u, v)*dx == inner(f, v)*dx, out)
return np.max(np.abs(out.dat.data - f.dat.data))
def run_test():
family = "Lagrange"
degree = range(1, 5)
return np.array([identity(family, d) for d in degree])
def run_vector_test():
family = "Lagrange"
degree = range(1, 5)
return np.array([vector_identity(family, d) for d in degree])
def test_identity():
assert (run_test() < 1e-6).all()
def test_vector_identity():
assert (run_vector_test() < 1e-6).all()
@pytest.mark.parallel
def test_identity_parallel():
from mpi4py import MPI
error = run_test()
MPI.COMM_WORLD.allreduce(MPI.IN_PLACE, error, MPI.MAX)
print '[%d]' % MPI.COMM_WORLD.rank, 'error:', error
assert (error < np.array([1.0e-11, 1.0e-6, 1.0e-6, 1.0e-5])).all()
@pytest.mark.parallel(nprocs=2)
def test_vector_identity_parallel():
from mpi4py import MPI
error = run_vector_test()
MPI.COMM_WORLD.allreduce(MPI.IN_PLACE, error, MPI.MAX)
print '[%d]' % MPI.COMM_WORLD.rank, 'error:', error
assert (error < 1e-6).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_nonlinear_helmholtz.py
"""This demo program solves Helmholtz's equation
- div grad u(x, y) + u(x,y) = f(x, y)
on the unit square with source f given by
f(x, y) = (1.0 + 8.0*pi**2)*cos(x[0]*2*pi)*cos(x[1]*2*pi)
and the analytical solution
u(x, y) = cos(x[0]*2*pi)*cos(x[1]*2*pi)
"""
import pytest
from firedrake import *
def run_test(x, parameters={}):
# Create mesh and define function space
mesh = UnitSquareMesh(2 ** x, 2 ** x)
V = FunctionSpace(mesh, "CG", 2)
# Define variational problem
lmbda = 1
u = Function(V)
v = TestFunction(V)
f = Function(V)
f.interpolate(Expression("(1+8*pi*pi)*cos(x[0]*pi*2)*cos(x[1]*pi*2)"))
a = (dot(grad(v), grad(u)) + lmbda * v * u) * dx
L = f * v * dx
# Compute solution
solve(a - L == 0, u, solver_parameters=parameters)
f.interpolate(Expression("cos(x[0]*2*pi)*cos(x[1]*2*pi)"))
return sqrt(assemble(dot(u - f, u - f) * dx))
def run_convergence_test(parameters={}):
import numpy as np
diff = np.array([run_test(i, parameters) for i in range(3, 6)])
return np.log2(diff[:-1] / diff[1:])
@pytest.mark.parametrize('params', [{}, {'snes_type': 'ksponly', 'ksp_type': 'preonly', 'pc_type': 'lu'}])
def test_l2_conv(params):
assert (run_convergence_test(parameters=params) > 2.8).all()
@pytest.mark.parallel
def test_l2_conv_parallel():
from mpi4py import MPI
l2_conv = run_convergence_test()
print '[%d]' % MPI.COMM_WORLD.rank, 'convergence rate:', l2_conv
assert (l2_conv > 2.8).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/scripts/firedrake-clean
#!/usr/bin/env python
from firedrake.ffc_interface import clear_cache, FFCKernel
from firedrake.utility_meshes import _clear_cachedir as clear_mesh_cache, _cachedir as mesh_cachedir
from pyop2.compilation import clear_cache as pyop2_clear_cache
if __name__ == '__main__':
print 'Removing cached ffc kernels from %s' % FFCKernel._cachedir
clear_cache()
print 'Removing cached generated meshes from %s' % mesh_cachedir
clear_mesh_cache()
pyop2_clear_cache(prompt=True)
<file_sep>/firedrake/matrix.py
import copy
import ufl
from pyop2 import op2
from pyop2.utils import as_tuple, flatten
import assemble
class Matrix(object):
"""A representation of an assembled bilinear form.
:arg a: the bilinear form this :class:`Matrix` represents.
:arg bcs: an iterable of boundary conditions to apply to this
:class:`Matrix`. May be `None` if there are no boundary
conditions to apply.
A :class:`pyop2.Mat` will be built from the remaining
arguments, for valid values, see :class:`pyop2.Mat`.
.. note::
This object acts to the right on an assembled :class:`.Function`
and to the left on an assembled cofunction (currently represented
by a :class:`.Function`).
"""
def __init__(self, a, bcs, *args, **kwargs):
self._a = a
self._M = op2.Mat(*args, **kwargs)
self._thunk = None
self._assembled = False
# Iteration over bcs must be in a parallel consistent order
# (so we can't use a set, since the iteration order may differ
# on different processes)
self._bcs = [bc for bc in bcs] if bcs is not None else []
self._bcs_at_point_of_assembly = []
def assemble(self):
"""Actually assemble this :class:`Matrix`.
This calls the stashed assembly callback or does nothing if
the matrix is already assembled.
.. note::
If the boundary conditions stashed on the :class:`Matrix` have
changed since the last time it was assembled, this will
necessitate reassembly. So for example:
.. code-block:: python
A = assemble(a, bcs=[bc1])
solve(A, x, b)
bc2.apply(A)
solve(A, x, b)
will apply boundary conditions from `bc1` in the first
solve, but both `bc1` and `bc2` in the second solve.
"""
if self._assembly_callback is None:
raise RuntimeError('Trying to assemble a Matrix, but no thunk found')
if self._assembled:
if self._needs_reassembly:
assemble._assemble(self.a, tensor=self, bcs=self.bcs)
return self.assemble()
return
self._bcs_at_point_of_assembly = copy.copy(self.bcs)
self._assembly_callback(self.bcs)
self._assembled = True
@property
def _assembly_callback(self):
"""Return the callback for assembling this :class:`Matrix`."""
return self._thunk
@_assembly_callback.setter
def _assembly_callback(self, thunk):
"""Set the callback for assembling this :class:`Matrix`.
:arg thunk: the callback, this should take one argument, the
boundary conditions to apply (pass None for no boundary
conditions).
Assigning to this property sets the :attr:`assembled` property
to False, necessitating a re-assembly."""
self._thunk = thunk
self._assembled = False
@property
def assembled(self):
"""Return True if this :class:`Matrix` has been assembled."""
return self._assembled
@property
def has_bcs(self):
"""Return True if this :class:`Matrix` has any boundary
conditions attached to it."""
return self._bcs != []
@property
def bcs(self):
"""The set of boundary conditions attached to this
:class:`Matrix` (may be empty)."""
return self._bcs
@bcs.setter
def bcs(self, bcs):
"""Attach some boundary conditions to this :class:`Matrix`.
:arg bcs: a boundary condition (of type
:class:`.DirichletBC`), or an iterable of boundary
conditions. If bcs is None, erase all boundary conditions
on the :class:`Matrix`.
"""
self._bcs = []
if bcs is not None:
try:
for bc in bcs:
self._bcs.append(bc)
except TypeError:
# BC instance, not iterable
self._bcs.append(bcs)
@property
def a(self):
"""The bilinear form this :class:`Matrix` was assembled from"""
return self._a
@property
def M(self):
"""The :class:`pyop2.Mat` representing the assembled form
.. note ::
This property forces an actual assembly of the form, if you
just need a handle on the :class:`pyop2.Mat` object it's
wrapping, use :attr:`_M` instead."""
self.assemble()
# User wants to see it, so force the evaluation.
self._M._force_evaluation()
return self._M
@property
def _needs_reassembly(self):
"""Does this :class:`Matrix` need reassembly.
The :class:`Matrix` needs reassembling if the subdomains over
which boundary conditions were applied the last time it was
assembled are different from the subdomains of the current set
of boundary conditions.
"""
old_subdomains = set(flatten(as_tuple(bc.sub_domain)
for bc in self._bcs_at_point_of_assembly))
new_subdomains = set(flatten(as_tuple(bc.sub_domain)
for bc in self.bcs))
return old_subdomains != new_subdomains
def add_bc(self, bc):
"""Add a boundary condition to this :class:`Matrix`.
:arg bc: the :class:`.DirichletBC` to add.
If the subdomain this boundary condition is applied over is
the same as the subdomain of an existing boundary condition on
the :class:`Matrix`, the existing boundary condition is
replaced with this new one. Otherwise, this boundary
condition is added to the set of boundary conditions on the
:class:`Matrix`.
"""
new_bcs = [bc]
for existing_bc in self.bcs:
# New BC doesn't override existing one, so keep it.
if bc.sub_domain != existing_bc.sub_domain:
new_bcs.append(existing_bc)
self.bcs = new_bcs
def _form_action(self, u):
"""Assemble the form action of this :class:`Matrix`' bilinear form
onto the :class:`Function` ``u``.
.. note::
This is the form **without** any boundary conditions."""
if not hasattr(self, '_a_action'):
self._a_action = ufl.action(self._a, u)
if hasattr(self, '_a_action_coeff'):
self._a_action = ufl.replace(self._a_action, {self._a_action_coeff: u})
self._a_action_coeff = u
# Since we assemble the cached form, the kernels will already have
# been compiled and stashed on the form the second time round
return assemble._assemble(self._a_action)
def __repr__(self):
return '%sassembled firedrake.Matrix(form=%r, bcs=%r)' % \
('' if self._assembled else 'un',
self.a,
self.bcs)
def __str__(self):
return '%sassembled firedrake.Matrix(form=%s, bcs=%s)' % \
('' if self._assembled else 'un',
self.a,
self.bcs)
<file_sep>/firedrake/norms.py
from pyop2.logger import warning
from ufl import inner, div, grad, curl, sqrt
import assemble
import function
import functionspace
import projection
__all__ = ['errornorm', 'norm']
def errornorm(u, uh, norm_type="L2", degree_rise=3, mesh=None):
"""Compute the error :math:`e = u - u_h` in the specified norm.
:arg u: a :class:`.Function` containing an "exact" solution
:arg uh: a :class:`.Function` containing the approximate solution
:arg norm_type: the type of norm to compute, see :func:`.norm` for
details of supported norm types.
:arg degree_rise: increase in polynomial degree to use as the
approximation space for computing the error.
:arg mesh: an optional mesh on which to compute the error norm
(currently ignored).
This function works by :func:`.project`\ing ``u`` and ``uh`` into
a space of degree ``degree_rise`` higher than the degree of ``uh``
and computing the error there.
"""
urank = len(u.shape())
uhrank = len(uh.shape())
rank = urank
if urank != uhrank:
raise RuntimeError("Mismatching rank between u and uh")
degree = uh.function_space().ufl_element().degree()
if isinstance(degree, tuple):
degree = max(degree) + degree_rise
else:
degree += degree_rise
# The exact solution might be an expression, in which case this test is irrelevant.
if isinstance(u, function.Function):
degree_u = u.function_space().ufl_element().degree()
if degree > degree_u:
warning("Degree of exact solution less than approximation degree")
mesh = uh.function_space().mesh()
if rank == 0:
V = functionspace.FunctionSpace(mesh, 'DG', degree)
elif rank == 1:
V = functionspace.VectorFunctionSpace(mesh, 'DG', degree,
dim=u.shape()[0])
else:
raise RuntimeError("Don't know how to compute error norm for tensor valued functions")
u_ = projection.project(u, V)
uh_ = projection.project(uh, V)
uh_ -= u_
return norm(uh_, norm_type=norm_type, mesh=mesh)
def norm(v, norm_type="L2", mesh=None):
"""Compute the norm of ``v``.
:arg v: a :class:`.Function` to compute the norm of
:arg norm_type: the type of norm to compute, see below for
options.
:arg mesh: an optional mesh on which to compute the norm
(currently ignored).
Available norm types are:
* L2
.. math::
||v||_{L^2}^2 = \int (v, v) \mathrm{d}x
* H1
.. math::
||v||_{H^1}^2 = \int (v, v) + (\\nabla v, \\nabla v) \mathrm{d}x
* Hdiv
.. math::
||v||_{H_\mathrm{div}}^2 = \int (v, v) + (\\nabla\cdot v, \\nabla \cdot v) \mathrm{d}x
* Hcurl
.. math::
||v||_{H_\mathrm{curl}}^2 = \int (v, v) + (\\nabla \wedge v, \\nabla \wedge v) \mathrm{d}x
"""
assert isinstance(v, function.Function)
typ = norm_type.lower()
mesh = v.function_space().mesh()
dx = mesh._dx
if typ == 'l2':
form = inner(v, v)*dx
elif typ == 'h1':
form = inner(v, v)*dx + inner(grad(v), grad(v))*dx
elif typ == "hdiv":
form = inner(v, v)*dx + div(v)*div(v)*dx
elif typ == "hcurl":
form = inner(v, v)*dx + inner(curl(v), curl(v))*dx
else:
raise RuntimeError("Unknown norm type '%s'" % norm_type)
return sqrt(assemble.assemble(form))
<file_sep>/demos/helmholtz/helmholtz.py.rst
Simple Helmholtz equation
=========================
Let's start by considering the Helmholtz equation on a unit square,
:math:`\Omega`, with boundary :math:`\Gamma`:
.. math::
-\nabla^2 u + u = f
\nabla u \cdot \vec{n} = 0 \ \textrm{on}\ \Gamma
for some known function :math:`f`. The solution to this equation will
be some function :math:`u\in V` for some suitable function space
:math:`V` that satisfies these equations. We transform the equation
into weak form by multiplying by an arbitrary test function in
:math:`V`, integrating over the domain and then integrating by
parts. The variational problem so derived reads: find :math:`u\in V`
such that:
.. math::
\require{cancel}
\int_\Omega \nabla u\cdot\nabla v + uv\ \mathrm{d}x = \int_\Omega
vf\ \mathrm{d}x + \cancel{\int_\Gamma v \nabla u \cdot \vec{n} \mathrm{d}s}
Note that the boundary condition has been enforced weakly by removing
the surface term resulting from the integration by parts.
We can choose the function :math:`f`, so we take:
.. math::
f = (1.0 + 8.0\pi^2)\cos(2\pi x)\cos(2\pi y)
which conveniently yields the analytic solution:
.. math::
u = \cos(2\pi x)\cos(2\pi y)
However we wish to employ this as an example for the finite element
method, so lets go ahead and produce a numerical solution.
First, we always need a mesh. Let's have a :math:`10\times10` element unit square::
from firedrake import *
mesh = UnitSquareMesh(10, 10)
We need to decide on the function space in which we'd like to solve the
problem. Let's use piecewise linear functions continuous between
elements::
V = FunctionSpace(mesh, "CG", 1)
We'll also need the test and trial functions corresponding to this
function space::
u = TrialFunction(V)
v = TestFunction(V)
We declare a function over our function space and give it the
value of our right hand side function::
f = Function(V)
f.interpolate(Expression("(1+8*pi*pi)*cos(x[0]*pi*2)*cos(x[1]*pi*2)"))
We can now define the bilinear and linear forms for the left and right
hand sides of our equation respectively::
a = (dot(grad(v), grad(u)) + v * u) * dx
L = f * v * dx
Finally we solve the equation. We redefine `u` to be a function
holding the solution::
u = Function(V)
Since we know that the Helmholtz equation is
symmetric, we instruct PETSc to employ the conjugate gradient method::
solve(a == L, u, solver_parameters={'ksp_type': 'cg'})
For more details on how to specify solver parameters, see the section
of the manual on :doc:`solving PDEs <../solving-interface>`.
Next, we might want to look at the result, so we output our solution
to a file::
File("helmholtz.pvd") << u
This file can be visualised using `paraview <http://www.paraview.org/>`__.
Alternatively, since we have an analytic solution, we can check the
:math:`L_2` norm of the error in the solution::
f.interpolate(Expression("cos(x[0]*pi*2)*cos(x[1]*pi*2)"))
print sqrt(assemble(dot(u - f, u - f) * dx))
A python script version of this demo can be found `here <helmholtz.py>`__.
<file_sep>/tests/regression/test_poisson_mixed_no_bcs.py
"""Solve the mixed formulation of the Laplacian described in section 2.3.1 of
<NAME> 2010 "Finite Element Exterior Calculus: From Hodge Theory
to Numerical Stability":
sigma - grad(u) = 0
div(sigma) = f
The corresponding weak (variational problem)
<sigma, tau> + <div(tau), u> = 0 for all tau
<div(sigma), v> = <f, v> for all v
is solved using BDM (Brezzi-Douglas-Marini) elements of degree k for
(sigma, tau) and DG (discontinuous Galerkin) elements of degree k - 1
for (u, v).
No strong boundary conditions are enforced. The forcing function is chosen as
-2*(x[0]-1)*x[0] - 2*(x[1]-1)*x[1]
which reproduces the known analytical solution
x[0]*(1-x[0])*x[1]*(1-x[1])
"""
import pytest
import numpy as np
from firedrake import *
def poisson_mixed(size, parameters={}, quadrilateral=False):
# Create mesh
mesh = UnitSquareMesh(2 ** size, 2 ** size, quadrilateral=quadrilateral)
# Define function spaces and mixed (product) space
if quadrilateral:
S0 = FiniteElement("CG", "interval", 1)
S1 = FiniteElement("DG", "interval", 0)
T0 = FiniteElement("CG", "interval", 1)
T1 = FiniteElement("DG", "interval", 0)
DG_elt = OuterProductElement(S1, T1)
BDM_elt_h = HDiv(OuterProductElement(S1, T0))
BDM_elt_v = HDiv(OuterProductElement(S0, T1))
BDM_elt = BDM_elt_h + BDM_elt_v
# spaces for calculation
DG = FunctionSpace(mesh, DG_elt)
BDM = FunctionSpace(mesh, BDM_elt)
else:
BDM = FunctionSpace(mesh, "BDM", 1)
DG = FunctionSpace(mesh, "DG", 0)
W = BDM * DG
# Define trial and test functions
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
# Define source function
f = Function(DG).interpolate(Expression("-2*(x[0]-1)*x[0] - 2*(x[1]-1)*x[1]"))
# Define variational form
a = (dot(sigma, tau) + div(tau)*u + div(sigma)*v)*dx
L = - f*v*dx
# Compute solution
w = Function(W)
solve(a == L, w, solver_parameters=parameters)
sigma, u = w.split()
# Analytical solution
f.interpolate(Expression("x[0]*(1-x[0])*x[1]*(1-x[1])"))
return sqrt(assemble(dot(u - f, u - f) * dx)), u, f
@pytest.mark.parametrize('parameters',
[{}, {'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'ksp_type': 'gmres',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_0_ksp_type': 'cg',
'fieldsplit_0_pc_factor_shift_type': 'INBLOCKS',
'fieldsplit_1_pc_factor_shift_type': 'INBLOCKS',
'fieldsplit_1_ksp_type': 'cg'}])
def test_poisson_mixed(parameters):
"""Test second-order convergence of the mixed poisson formulation."""
diff = np.array([poisson_mixed(i, parameters)[0] for i in range(3, 6)])
print "l2 error norms:", diff
conv = np.log2(diff[:-1] / diff[1:])
print "convergence order:", conv
assert (np.array(conv) > 1.9).all()
@pytest.mark.parametrize(('testcase', 'convrate'),
[((3, 6), 1.9)])
def test_hdiv_convergence(testcase, convrate):
"""Test second-order convergence of the mixed poisson formulation
on quadrilaterals with HDiv elements."""
start, end = testcase
l2err = np.zeros(end - start)
for ii in [i + start for i in range(len(l2err))]:
l2err[ii - start] = poisson_mixed(ii, quadrilateral=True)[0]
assert (np.log2(l2err[:-1] / l2err[1:]) > convrate).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/halo.py
import numpy as np
from collections import defaultdict
from mpi4py import MPI
from pyop2 import op2
import utils
class Halo(object):
"""Build a Halo associated with the appropriate FunctionSpace.
The Halo is derived from a PetscSF object and builds the global
to universal numbering map from the respective PetscSections."""
def __init__(self, petscsf, global_numbering, universal_numbering):
self._tag = utils._new_uid()
self._comm = op2.MPI.comm
self._nprocs = self.comm.size
self._sends = defaultdict(list)
self._receives = defaultdict(list)
self._gnn2unn = None
remote_sends = defaultdict(list)
if op2.MPI.comm.size <= 1:
return
# Sort the SF by local indices
nroots, nleaves, local, remote = petscsf.getGraph()
local_new, remote_new = (list(x) for x in zip(*sorted(zip(local, remote), key=lambda x: x[0])))
petscsf.setGraph(nroots, nleaves, local_new, remote_new)
# Derive local receives and according remote sends
nroots, nleaves, local, remote = petscsf.getGraph()
for local, (rank, index) in zip(local, remote):
if rank != self.comm.rank:
self._receives[rank].append(local)
remote_sends[rank].append(index)
# Propagate remote send lists to the actual sender
send_reqs = []
for p in range(self._nprocs):
# send sizes
if p != self._comm.rank:
s = np.array(len(remote_sends[p]), dtype=np.int32)
send_reqs.append(self.comm.Isend(s, dest=p, tag=self.tag))
recv_reqs = []
sizes = [np.zeros(1, dtype=np.int32) for _ in range(self._nprocs)]
for p in range(self._nprocs):
# receive sizes
if p != self._comm.rank:
recv_reqs.append(self.comm.Irecv(sizes[p], source=p, tag=self.tag))
MPI.Request.Waitall(recv_reqs)
MPI.Request.Waitall(send_reqs)
for p in range(self._nprocs):
# allocate buffers
if p != self._comm.rank:
if sizes[p][0] > 0:
self._sends[p] = np.empty(sizes[p][0], dtype=np.int32)
send_reqs = []
for p in range(self._nprocs):
if p != self._comm.rank:
if len(remote_sends[p]) > 0:
send_buf = np.array(remote_sends[p], dtype=np.int32)
send_reqs.append(self.comm.Isend(send_buf, dest=p, tag=self.tag))
recv_reqs = []
for p in range(self._nprocs):
if p != self._comm.rank:
if sizes[p][0] > 0:
recv_reqs.append(self.comm.Irecv(self._sends[p], source=p, tag=self.tag))
MPI.Request.Waitall(send_reqs)
MPI.Request.Waitall(recv_reqs)
# Build Global-To-Universal mapping
pStart, pEnd = global_numbering.getChart()
self._gnn2unn = np.zeros(global_numbering.getStorageSize(), dtype=np.int32)
for p in range(pStart, pEnd):
dof = global_numbering.getDof(p)
goff = global_numbering.getOffset(p)
uoff = universal_numbering.getOffset(p)
if uoff < 0:
uoff = (-1*uoff)-1
for c in range(dof):
self._gnn2unn[goff+c] = uoff+c
@utils.cached_property
def op2_halo(self):
if not self.sends and not self.receives:
return None
return op2.Halo(self.sends, self.receives,
comm=self.comm, gnn2unn=self.gnn2unn)
@property
def comm(self):
return self._comm
@property
def tag(self):
return self._tag
@property
def nprocs(self):
return self._nprocs
@property
def sends(self):
return self._sends
@property
def receives(self):
return self._receives
@property
def gnn2unn(self):
return self._gnn2unn
<file_sep>/Makefile
all: modules
modules:
@echo " Building extension modules"
@python setup.py build_ext --inplace > build.log 2>&1 || cat build.log
lint:
@echo " Linting firedrake codebase"
@flake8 firedrake
@echo " Linting firedrake test suite"
@flake8 tests
clean:
@echo " Cleaning extension modules"
@python setup.py clean > /dev/null 2>&1
@echo " RM firedrake/dmplex.so"
-@rm -f firedrake/dmplex.so > /dev/null 2>&1
THREADS=1
ifeq ($(THREADS), 1)
PYTEST_ARGS=
else
PYTEST_ARGS=-n $(THREADS)
endif
test_regression: modules
@echo " Running non-extruded regression tests"
@py.test tests/regression $(PYTEST_ARGS)
test_extrusion: modules
@echo " Running extruded regression tests"
@py.test tests/extrusion $(PYTEST_ARGS)
test: modules
@echo " Running all regression tests"
@py.test tests $(PYTEST_ARGS)
alltest: modules lint test
shorttest: modules lint
@echo " Running short regression tests"
@py.test --short tests $(PYTEST_ARGS)
<file_sep>/firedrake/assemble.py
import numpy
import ufl
from pyop2 import op2
from pyop2.exceptions import MapValueError
from pyop2.profiling import timed_region, profile
import assembly_cache
import assemble_expressions
import ffc_interface
import function
import functionspace
import matrix
import solving
__all__ = ["assemble"]
@profile
def assemble(f, tensor=None, bcs=None, form_compiler_parameters=None):
"""Evaluate f.
:arg f: a :class:`ufl.Form` or :class:`ufl.core.expr.Expr`.
:arg tensor: an existing tensor object to place the result in
(optional).
:arg bcs: a list of boundary conditions to apply (optional).
:arg form_compiler_parameters: (optional) dict of parameters to pass to
the form compiler. Ignored if not assembling a
:class:`ufl.Form`. Any parameters provided here will be
overridden by parameters set on the :class;`ufl.Measure` in the
form. For example, if a :data:`quadrature_degree` of 4 is
specified in this argument, but a degree of 3 is requested in
the measure, the latter will be used.
If f is a :class:`ufl.Form` then this evaluates the corresponding
integral(s) and returns a :class:`float` for 0-forms, a
:class:`.Function` for 1-forms and a :class:`.Matrix` for 2-forms.
If f is an expression other than a form, it will be evaluated
pointwise on the :class:`.Function`\s in the expression. This will
only succeed if all the Functions are on the same
:class:`.FunctionSpace`
If ``tensor`` is supplied, the assembled result will be placed
there, otherwise a new object of the appropriate type will be
returned.
If ``bcs`` is supplied and ``f`` is a 2-form, the rows and columns
of the resulting :class:`.Matrix` corresponding to boundary nodes
will be set to 0 and the diagonal entries to 1. If ``f`` is a
1-form, the vector entries at boundary nodes are set to the
boundary condition values.
"""
if isinstance(f, ufl.form.Form):
return _assemble(f, tensor=tensor, bcs=solving._extract_bcs(bcs),
form_compiler_parameters=form_compiler_parameters)
elif isinstance(f, ufl.core.expr.Expr):
return assemble_expressions.assemble_expression(f)
else:
raise TypeError("Unable to assemble: %r" % f)
def _assemble(f, tensor=None, bcs=None, form_compiler_parameters=None):
"""Assemble the form f and return a Firedrake object representing the
result. This will be a :class:`float` for 0-forms, a
:class:`.Function` for 1-forms and a :class:`.Matrix` for 2-forms.
:arg bcs: A tuple of :class`.DirichletBC`\s to be applied.
:arg tensor: An existing tensor object into which the form should be
assembled. If this is not supplied, a new tensor will be created for
the purpose.
:arg form_compiler_parameters: (optional) dict of parameters to pass to
the form compiler.
"""
kernels = ffc_interface.compile_form(f, "form", parameters=form_compiler_parameters)
rank = len(f.arguments())
is_mat = rank == 2
is_vec = rank == 1
integrals = f.integrals()
def get_rank(arg):
return arg.function_space().rank
if is_mat:
test, trial = f.arguments()
map_pairs = []
cell_domains = []
exterior_facet_domains = []
interior_facet_domains = []
# For horizontal facets of extrded meshes, the corresponding domain
# in the base mesh is the cell domain. Hence all the maps used for top
# bottom and interior horizontal facets will use the cell to dofs map
# coming from the base mesh as a starting point for the actual dynamic map
# computation.
for integral in integrals:
integral_type = integral.integral_type()
if integral_type == "cell":
cell_domains.append(op2.ALL)
elif integral_type == "exterior_facet":
exterior_facet_domains.append(op2.ALL)
elif integral_type == "interior_facet":
interior_facet_domains.append(op2.ALL)
elif integral_type == "exterior_facet_bottom":
cell_domains.append(op2.ON_BOTTOM)
elif integral_type == "exterior_facet_top":
cell_domains.append(op2.ON_TOP)
elif integral_type == "exterior_facet_vert":
exterior_facet_domains.append(op2.ALL)
elif integral_type == "interior_facet_horiz":
cell_domains.append(op2.ON_INTERIOR_FACETS)
elif integral_type == "interior_facet_vert":
interior_facet_domains.append(op2.ALL)
else:
raise RuntimeError('Unknown integral type "%s"' % integral_type)
# To avoid an extra check for extruded domains, the maps that are being passed in
# are DecoratedMaps. For the non-extruded case the DecoratedMaps don't restrict the
# space over which we iterate as the domains are dropped at Sparsity construction
# time. In the extruded case the cell domains are used to identify the regions of the
# mesh which require allocation in the sparsity.
if cell_domains:
map_pairs.append((op2.DecoratedMap(test.cell_node_map(), cell_domains),
op2.DecoratedMap(trial.cell_node_map(), cell_domains)))
if exterior_facet_domains:
map_pairs.append((op2.DecoratedMap(test.exterior_facet_node_map(), exterior_facet_domains),
op2.DecoratedMap(trial.exterior_facet_node_map(), exterior_facet_domains)))
if interior_facet_domains:
map_pairs.append((op2.DecoratedMap(test.interior_facet_node_map(), interior_facet_domains),
op2.DecoratedMap(trial.interior_facet_node_map(), interior_facet_domains)))
map_pairs = tuple(map_pairs)
if tensor is None:
# Construct OP2 Mat to assemble into
fs_names = (
test.function_space().name, trial.function_space().name)
sparsity = op2.Sparsity((test.function_space().dof_dset,
trial.function_space().dof_dset),
map_pairs,
"%s_%s_sparsity" % fs_names)
result_matrix = matrix.Matrix(f, bcs, sparsity, numpy.float64,
"%s_%s_matrix" % fs_names)
tensor = result_matrix._M
else:
result_matrix = tensor
# Replace any bcs on the tensor we passed in
result_matrix.bcs = bcs
tensor = tensor._M
def mat(testmap, trialmap, i, j):
return tensor[i, j](op2.INC,
(testmap(test.function_space()[i])[op2.i[0]],
trialmap(trial.function_space()[j])[op2.i[1]]),
flatten=True)
result = lambda: result_matrix
elif is_vec:
test = f.arguments()[0]
if tensor is None:
result_function = function.Function(test.function_space())
tensor = result_function.dat
else:
result_function = tensor
tensor = result_function.dat
def vec(testmap, i):
return tensor[i](op2.INC,
testmap(test.function_space()[i])[op2.i[0]],
flatten=True)
result = lambda: result_function
else:
# 0-forms are always scalar
if tensor is None:
tensor = op2.Global(1, [0.0])
result = lambda: tensor.data[0]
# Since applying boundary conditions to a matrix changes the
# initial assembly, to support:
# A = assemble(a)
# bc.apply(A)
# solve(A, ...)
# we need to defer actually assembling the matrix until just
# before we need it (when we know if there are any bcs to be
# applied). To do so, we build a closure that carries out the
# assembly and stash that on the Matrix object. When we hit a
# solve, we funcall the closure with any bcs the Matrix now has to
# assemble it.
def thunk(bcs):
try:
tensor.zero()
except AttributeError:
pass
for (i, j), integral_type, subdomain_id, coords, coefficients, needs_orientations, kernel in kernels:
m = coords.function_space().mesh()
if needs_orientations:
cell_orientations = m.cell_orientations()
# Extract block from tensor and test/trial spaces
# FIXME Ugly variable renaming required because functions are not
# lexical closures in Python and we're writing to these variables
if is_mat and tensor.sparsity.shape > (1, 1):
tsbc = [bc for bc in bcs if bc.function_space().index == i]
trbc = [bc for bc in bcs if bc.function_space().index == j]
elif is_mat:
tsbc, trbc = bcs, bcs
if integral_type == 'cell':
with timed_region("Assemble cells"):
if is_mat:
tensor_arg = mat(lambda s: s.cell_node_map(tsbc),
lambda s: s.cell_node_map(trbc),
i, j)
elif is_vec:
tensor_arg = vec(lambda s: s.cell_node_map(), i)
else:
tensor_arg = tensor(op2.INC)
itspace = m.cell_set
args = [kernel, itspace, tensor_arg,
coords.dat(op2.READ, coords.cell_node_map(),
flatten=True)]
if needs_orientations:
args.append(cell_orientations.dat(op2.READ,
cell_orientations.cell_node_map(),
flatten=True))
for c in coefficients:
args.append(c.dat(op2.READ, c.cell_node_map(),
flatten=True))
try:
op2.par_loop(*args)
except MapValueError:
raise RuntimeError("Integral measure does not match measure of all coefficients/arguments")
elif integral_type in ['exterior_facet', 'exterior_facet_vert']:
with timed_region("Assemble exterior facets"):
if is_mat:
tensor_arg = mat(lambda s: s.exterior_facet_node_map(tsbc),
lambda s: s.exterior_facet_node_map(trbc),
i, j)
elif is_vec:
tensor_arg = vec(lambda s: s.exterior_facet_node_map(), i)
else:
tensor_arg = tensor(op2.INC)
args = [kernel, m.exterior_facets.measure_set(integral_type,
subdomain_id),
tensor_arg,
coords.dat(op2.READ, coords.exterior_facet_node_map(),
flatten=True)]
if needs_orientations:
args.append(cell_orientations.dat(op2.READ,
cell_orientations.exterior_facet_node_map(),
flatten=True))
for c in coefficients:
args.append(c.dat(op2.READ, c.exterior_facet_node_map(),
flatten=True))
args.append(m.exterior_facets.local_facet_dat(op2.READ))
try:
op2.par_loop(*args)
except MapValueError:
raise RuntimeError("Integral measure does not match measure of all coefficients/arguments")
elif integral_type in ['exterior_facet_top', 'exterior_facet_bottom']:
with timed_region("Assemble exterior facets"):
# In the case of extruded meshes with horizontal facet integrals, two
# parallel loops will (potentially) get created and called based on the
# domain id: interior horizontal, bottom or top.
# Get the list of sets and globals required for parallel loop construction.
set_global_list = m.exterior_facets.measure_set(integral_type, subdomain_id)
# Iterate over the list and assemble all the args of the parallel loop
for (index, set) in set_global_list:
if is_mat:
tensor_arg = mat(lambda s: op2.DecoratedMap(s.cell_node_map(tsbc), index),
lambda s: op2.DecoratedMap(s.cell_node_map(trbc), index),
i, j)
elif is_vec:
tensor_arg = vec(lambda s: s.cell_node_map(), i)
else:
tensor_arg = tensor(op2.INC)
# Add the kernel, iteration set and coordinate fields to the loop args
args = [kernel, set, tensor_arg,
coords.dat(op2.READ, coords.cell_node_map(),
flatten=True)]
if needs_orientations:
args.append(cell_orientations.dat(op2.READ,
cell_orientations.cell_node_map(),
flatten=True))
for c in coefficients:
args.append(c.dat(op2.READ, c.cell_node_map(),
flatten=True))
try:
op2.par_loop(*args, iterate=index)
except MapValueError:
raise RuntimeError("Integral measure does not match measure of all coefficients/arguments")
elif integral_type in ['interior_facet', 'interior_facet_vert']:
with timed_region("Assemble interior facets"):
if is_mat:
tensor_arg = mat(lambda s: s.interior_facet_node_map(tsbc),
lambda s: s.interior_facet_node_map(trbc),
i, j)
elif is_vec:
tensor_arg = vec(lambda s: s.interior_facet_node_map(), i)
else:
tensor_arg = tensor(op2.INC)
args = [kernel, m.interior_facets.set, tensor_arg,
coords.dat(op2.READ, coords.interior_facet_node_map(),
flatten=True)]
if needs_orientations:
args.append(cell_orientations.dat(op2.READ,
cell_orientations.interior_facet_node_map(),
flatten=True))
for c in coefficients:
args.append(c.dat(op2.READ, c.interior_facet_node_map(),
flatten=True))
args.append(m.interior_facets.local_facet_dat(op2.READ))
try:
op2.par_loop(*args)
except MapValueError:
raise RuntimeError("Integral measure does not match measure of all coefficients/arguments")
elif integral_type == 'interior_facet_horiz':
with timed_region("Assemble interior facets"):
if is_mat:
tensor_arg = mat(lambda s: op2.DecoratedMap(s.cell_node_map(tsbc),
op2.ON_INTERIOR_FACETS),
lambda s: op2.DecoratedMap(s.cell_node_map(trbc),
op2.ON_INTERIOR_FACETS),
i, j)
elif is_vec:
tensor_arg = vec(lambda s: s.cell_node_map(), i)
else:
tensor_arg = tensor(op2.INC)
args = [kernel, m.interior_facets.measure_set(integral_type, subdomain_id),
tensor_arg,
coords.dat(op2.READ, coords.cell_node_map(),
flatten=True)]
if needs_orientations:
args.append(cell_orientations.dat(op2.READ,
cell_orientations.cell_node_map(),
flatten=True))
for c in coefficients:
args.append(c.dat(op2.READ, c.cell_node_map(),
flatten=True))
try:
op2.par_loop(*args, iterate=op2.ON_INTERIOR_FACETS)
except MapValueError:
raise RuntimeError("Integral measure does not match measure of all coefficients/arguments")
else:
raise RuntimeError('Unknown integral type "%s"' % integral_type)
# Must apply bcs outside loop over kernels because we may wish
# to apply bcs to a block which is otherwise zero, and
# therefore does not have an associated kernel.
if bcs is not None and is_mat:
with timed_region('DirichletBC apply'):
for bc in bcs:
fs = bc.function_space()
if isinstance(fs, functionspace.MixedFunctionSpace):
raise RuntimeError("""Cannot apply boundary conditions to full mixed space. Did you forget to index it?""")
shape = tensor.sparsity.shape
for i in range(shape[0]):
for j in range(shape[1]):
# Set diagonal entries on bc nodes to 1 if the current
# block is on the matrix diagonal and its index matches the
# index of the function space the bc is defined on.
if i == j and (fs.index is None or fs.index == i):
tensor[i, j].inc_local_diagonal_entries(bc.nodes)
if bcs is not None and is_vec:
for bc in bcs:
bc.apply(result_function)
if is_mat:
# Queue up matrix assembly (after we've done all the other operations)
tensor.assemble()
return result()
thunk = assembly_cache._cache_thunk(thunk, f, result(), form_compiler_parameters)
if is_mat:
result_matrix._assembly_callback = thunk
return result()
else:
return thunk(bcs)
<file_sep>/tests/extrusion/test_extrusion_poisson_strong_bcs.py
"""This demo solves Poisson's equation
- div grad u(x, y) = 0
on an extruded unit cube with boundary conditions given by:
u(x, y, 0) = 0
v(x, y, 1) = 42
Homogeneous Neumann boundary conditions are applied naturally on the
other sides of the domain.
This has the analytical solution
u(x, y, z) = 42*z
"""
import pytest
from firedrake import *
def run_test(layers):
# Create mesh and define function space
m = UnitSquareMesh(1, 1)
mesh = ExtrudedMesh(m, layers, layer_height=1.0 / layers)
V = FunctionSpace(mesh, "CG", 1)
bcs = [DirichletBC(V, 0, "bottom"),
DirichletBC(V, 42, "top")]
v = TestFunction(V)
u = TrialFunction(V)
a = dot(grad(u), grad(v)) * dx
f = Function(V)
f.assign(0)
L = v * f * dx
u = Function(V)
exact = Function(V)
exact.interpolate(Expression('42*x[2]'))
solve(a == L, u, bcs=bcs)
res = sqrt(assemble(dot(u - exact, u - exact) * dx))
return res
def test_extrusion_poisson_strong_bcs():
for layers in [1, 2, 10]:
assert (run_test(layers) < 1.e-6)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_cylinder.py
from firedrake import *
import numpy as np
import pytest
def test_area():
len = 7
errors = np.zeros(len)
for i in range(2, 2+len):
m = CircleManifoldMesh(2**i)
mesh = ExtrudedMesh(m, layers=2**i, layer_height=1.0/(2**i))
fs = FunctionSpace(mesh, "DG", 0)
f = Function(fs).assign(1)
# surface area is 2*pi*r*h = 2*pi
errors[i-2] = np.abs(assemble(f*dx) - 2*np.pi)
# area converges quadratically to 2*pi
for i in range(len-1):
assert ln(errors[i]/errors[i+1])/ln(2) > 1.95
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti0_cylinder(horiz_complex, vert_complex):
"""
Verify that the 0-form Hodge Laplacian has kernel of dimension
equal to the 0th Betti number of the periodic extruded interval,
i.e. 1. Also verify that the 0-form Hodge Laplacian with
Dirichlet boundary conditions has kernel of dimension equal to the
2nd Betti number of the extruded mesh, i.e. 0.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = CircleManifoldMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
U0 = FiniteElement(U0[0], "interval", U0[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
W0_elt = OuterProductElement(U0, V0)
W0 = FunctionSpace(mesh, W0_elt)
u = TrialFunction(W0)
v = TestFunction(W0)
L = assemble(inner(grad(u), grad(v))*dx)
uvecs, s, vvecs = np.linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
bcs = [DirichletBC(W0, 0., x) for x in ["top", "bottom"]]
L = assemble(inner(grad(u), grad(v))*dx, bcs=bcs)
uvecs, s, vvecs = np.linalg.svd(L.M.values)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti1_cylinder(horiz_complex, vert_complex):
"""
Verify that the 1-form Hodge Laplacian has kernel of dimension
equal to the 1st Betti number of the periodic extruded interval,
i.e. 1. Also verify that the 1-form Hodge Laplacian with
Dirichlet boundary conditions has kernel of dimension equal to the
2nd Betti number of the periodic extruded interval mesh, i.e. 1.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = CircleManifoldMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
mesh.init_cell_orientations(Expression(('x[0]', 'x[1]', '0.0')))
U0 = FiniteElement(U0[0], "interval", U0[1])
U1 = FiniteElement(U1[0], "interval", U1[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W0_elt = OuterProductElement(U0, V0)
W1_a = HDiv(OuterProductElement(U1, V0))
W1_b = HDiv(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W0 = FunctionSpace(mesh, W0_elt)
W1 = FunctionSpace(mesh, W1_elt)
outward_normal = Function(VectorFunctionSpace(mesh, "DG", 0)).interpolate(Expression(('x[0]/sqrt(x[0]*x[0] + x[1]*x[1])', 'x[1]/sqrt(x[0]*x[0] + x[1]*x[1])', '0.0')))
W = W0*W1
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((sigma*tau - inner(cross(outward_normal, grad(tau)), u) + inner(cross(outward_normal, grad(sigma)), v) +
div(u)*div(v))*dx)
dW0 = W0.dof_count
dW1 = W1.dof_count
A = np.zeros((dW0+dW1, dW0+dW1))
A[:dW0, :dW0] = L.M[0, 0].values
A[:dW0, dW0:dW0+dW1] = L.M[0, 1].values
A[dW0:dW0+dW1, :dW0] = L.M[1, 0].values
A[dW0:dW0+dW1, dW0:dW0+dW1] = L.M[1, 1].values
uvecs, s, vvecs = np.linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
bc0 = [DirichletBC(W.sub(0), 0., x) for x in ["top", "bottom"]]
bc1 = [DirichletBC(W.sub(1), Expression(("0.", "0.", "0.")), x)
for x in ["top", "bottom"]]
L0 = assemble((sigma*tau - inner(cross(outward_normal, grad(tau)), u) + inner(cross(outward_normal, grad(sigma)), v) +
div(u)*div(v))*dx, bcs=(bc0 + bc1))
A0 = np.zeros((dW0+dW1, dW0+dW1))
A0[:dW0, :dW0] = L0.M[0, 0].values
A0[:dW0, dW0:dW0+dW1] = L0.M[0, 1].values
A0[dW0:dW0+dW1, :dW0] = L0.M[1, 0].values
A0[dW0:dW0+dW1, dW0:dW0+dW1] = L0.M[1, 1].values
u, s, v = np.linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("DG", 0)),
(("CG", 1), ("DG", 0)))])
def test_betti2_cylinder(horiz_complex, vert_complex):
"""
Verify that the 2-form Hodge Laplacian has kernel of dimension
equal to the 2nd Betti number of the periodic extruded interval
mesh, i.e. 0. Also verify that the 2-form Hodge Laplacian with
Dirichlet boundary conditions has kernel of dimension equal to the
0th Betti number of the periodic extruded interval mesh, i.e. 1.
"""
U0, U1 = horiz_complex
V0, V1 = vert_complex
m = CircleManifoldMesh(5)
mesh = ExtrudedMesh(m, layers=4, layer_height=0.25)
mesh.init_cell_orientations(Expression(('x[0]', 'x[1]', '0.0')))
U0 = FiniteElement(U0[0], "interval", U0[1])
U1 = FiniteElement(U1[0], "interval", U1[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W1_a = HDiv(OuterProductElement(U1, V0))
W1_b = HDiv(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W2_elt = OuterProductElement(U1, V1)
W1 = FunctionSpace(mesh, W1_elt)
W2 = FunctionSpace(mesh, W2_elt)
W = W1*W2
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
L = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx)
bc1 = [DirichletBC(W.sub(0), Expression(("0.", "0.", "0.")), x)
for x in ["top", "bottom"]]
L0 = assemble((inner(sigma, tau) - div(tau)*u + div(sigma)*v)*dx, bcs=bc1)
dW1 = W1.dof_count
dW2 = W2.dof_count
A = np.zeros((dW1+dW2, dW1+dW2))
A[:dW1, :dW1] = L.M[0, 0].values
A[:dW1, dW1:dW1+dW2] = L.M[0, 1].values
A[dW1:dW1+dW2, :dW1] = L.M[1, 0].values
A[dW1:dW1+dW2, dW1:dW1+dW2] = L.M[1, 1].values
u, s, v = np.linalg.svd(A)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 0)
A0 = np.zeros((dW1+dW2, dW1+dW2))
A0[:dW1, :dW1] = L0.M[0, 0].values
A0[:dW1, dW1:dW1+dW2] = L0.M[0, 1].values
A0[dW1:dW1+dW2, :dW1] = L0.M[1, 0].values
A0[dW1:dW1+dW2, dW1:dW1+dW2] = L0.M[1, 1].values
u, s, v = np.linalg.svd(A0)
nharmonic = sum(s < 1.0e-5)
assert(nharmonic == 1)
if __name__ == '__main__':
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_annulus.py
from firedrake import *
import numpy as np
import pytest
import os
def test_pi():
len = 7
errors = np.zeros(len)
for i in range(2, 2+len):
m = CircleManifoldMesh(2**i)
mesh = ExtrudedMesh(m, layers=2**i, layer_height=1.0/(2**i), extrusion_type="radial")
fs = FunctionSpace(mesh, "DG", 0)
f = Function(fs).assign(1)
# area is pi*(2^2) - pi*(1^2) = 3*pi
errors[i-2] = np.abs(assemble(f*dx) - 3*np.pi)
# area converges linearly to 3*pi
for i in range(len-1):
assert ln(errors[i]/errors[i+1])/ln(2) > 0.95
def test_poisson():
# u = x^2 + y^2 is a solution to the Poisson equation
# -div(grad(u)) = -4 on the annulus with inner radius
# 2, outer radius 5
len = 4
errors = np.zeros(len)
for i in range(4, 4+len):
m = CircleManifoldMesh(2**i, radius=2.0)
mesh = ExtrudedMesh(m, layers=2**i, layer_height=3.0/(2**i), extrusion_type="radial")
V = FunctionSpace(mesh, "CG", 1)
u = TrialFunction(V)
v = TestFunction(V)
f = Constant(-4)
bcs = [DirichletBC(V, 4, "bottom"),
DirichletBC(V, 25, "top")]
out = Function(V)
solve(dot(grad(u), grad(v))*dx == f*v*dx, out, bcs=bcs)
exactfs = FunctionSpace(mesh, "CG", 2)
exact = Function(exactfs).interpolate(Expression("x[0]*x[0] + x[1]*x[1]"))
errors[i-4] = sqrt(assemble((out-exact)*(out-exact)*dx))/sqrt(21*np.pi) # normalised
# we seem to get second-order convergence...
for i in range(len-1):
assert ln(errors[i]/errors[i+1])/ln(2) > 1.7
if __name__ == '__main__':
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_geometric_strong_bcs.py
import pytest
from firedrake import *
@pytest.mark.parametrize(('degree'), range(5))
def test_dg_advection(degree):
m = UnitSquareMesh(10, 10)
V = FunctionSpace(m, "DG", degree)
V_u = VectorFunctionSpace(m, "CG", 1)
t = Function(V)
v = TestFunction(V)
u = Function(V_u)
u.assign(Constant((1, 0)))
def upw(t, un):
return t('+')*un('+') - t('-')*un('-')
n = FacetNormal(m)
un = 0.5 * (dot(u, n) + abs(dot(u, n)))
F = -dot(grad(v), u) * t * dx + dot(jump(v), upw(t, un)) * dS + dot(v, un*t) * ds
bc = DirichletBC(V, 1., 1, method="geometric")
solve(F == 0, t, bcs=bc)
assert errornorm(Constant(1.0), t) < 1.e-12
<file_sep>/tests/extrusion/test_extrusion_facet_integrals_2D.py
"""Testing assembly of scalars on facets of extruded meshes in 2D"""
import pytest
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module', params=[1, 2])
def f(request):
mesh = extmesh_2D(4, 4)
fspace = FunctionSpace(mesh, "CG", request.param)
return Function(fspace)
@pytest.fixture(scope='module')
def RT2():
mesh = extmesh_2D(4, 4)
U0 = FiniteElement("CG", "interval", 2)
U1 = FiniteElement("DG", "interval", 1)
W1 = HDiv(OuterProductElement(U1, U0)) + HDiv(OuterProductElement(U0, U1))
return FunctionSpace(mesh, W1)
def test_scalar_area(f):
f.assign(1)
assert abs(assemble(f*ds_t) - 1.0) < 1e-7
assert abs(assemble(f*ds_b) - 1.0) < 1e-7
assert abs(assemble(f*ds_tb) - 2.0) < 1e-7
assert abs(assemble(f*ds_v) - 2.0) < 1e-7
assert abs(assemble(f('+')*dS_h) - 3.0) < 1e-7
assert abs(assemble(f('-')*dS_h) - 3.0) < 1e-7
assert abs(assemble(f('+')*dS_v) - 3.0) < 1e-7
assert abs(assemble(f('-')*dS_v) - 3.0) < 1e-7
def test_scalar_expression(f):
f.interpolate(Expression("x[1]"))
assert abs(assemble(f*ds_t) - 1.0) < 1e-7
assert abs(assemble(f*ds_b) - 0.0) < 1e-7
assert abs(assemble(f*ds_tb) - 1.0) < 1e-7
assert abs(assemble(f*ds_v) - 1.0) < 1e-7
assert abs(assemble(f('+')*dS_h) - 1.5) < 1e-7
assert abs(assemble(f('-')*dS_h) - 1.5) < 1e-7
assert abs(assemble(f('+')*dS_v) - 1.5) < 1e-7
assert abs(assemble(f('-')*dS_v) - 1.5) < 1e-7
def test_hdiv_area(RT2):
f = project(Expression(("0.8", "0.6")), RT2)
assert abs(assemble(dot(f, f)*ds_t) - 1.0) < 1e-7
assert abs(assemble(dot(f, f)*ds_b) - 1.0) < 1e-7
assert abs(assemble(dot(f, f)*ds_tb) - 2.0) < 1e-7
assert abs(assemble(dot(f, f)*ds_v) - 2.0) < 1e-7
assert abs(assemble(dot(f('+'), f('+'))*dS_h) - 3.0) < 1e-7
assert abs(assemble(dot(f('-'), f('-'))*dS_h) - 3.0) < 1e-7
assert abs(assemble(dot(f('+'), f('-'))*dS_h) - 3.0) < 1e-7
assert abs(assemble(dot(f('+'), f('+'))*dS_v) - 3.0) < 1e-7
assert abs(assemble(dot(f('-'), f('-'))*dS_v) - 3.0) < 1e-7
assert abs(assemble(dot(f('+'), f('-'))*dS_v) - 3.0) < 1e-7
def test_exterior_horizontal_normals(RT2):
n = FacetNormal(RT2.mesh())
f = project(Expression(("1.0", "0.0")), RT2)
assert abs(assemble(dot(f, n)*ds_t) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_b) - 0.0) < 1e-7
f = project(Expression(("0.0", "1.0")), RT2)
assert abs(assemble(dot(f, n)*ds_t) - 1.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_b) - (-1.0)) < 1e-7
def test_exterior_vertical_normals(RT2):
n = FacetNormal(RT2.mesh())
f = project(Expression(("1.0", "0.0")), RT2)
assert abs(assemble(dot(f, n)*ds_v(1)) - (-1.0)) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(2)) - 1.0) < 1e-7
f = project(Expression(("0.0", "1.0")), RT2)
assert abs(assemble(dot(f, n)*ds_v(1)) - 0.0) < 1e-7
assert abs(assemble(dot(f, n)*ds_v(2)) - 0.0) < 1e-7
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_info.py
from firedrake import *
def test_info():
info("This is normal text")
info_red("This is red text")
info_green("This is green text")
info_blue("This is blue text")
set_log_level(ERROR)
info("You should not see this")
log(ERROR, "You should see this error message")
log(WARNING, "You should NOT see this warning")
if __name__ == "__main__":
test_info()
<file_sep>/tests/regression/test_python_parloop.py
import numpy as np
import pytest
from firedrake import *
def test_python_parloop():
m = UnitSquareMesh(4, 4)
fs = FunctionSpace(m, "CG", 2)
f = Function(fs)
class MyExpression(Expression):
def eval(self, value, X):
value[:] = np.dot(X, X)
f.interpolate(MyExpression())
X = m.coordinates
assert assemble((f-dot(X, X))**2*dx)**.5 < 1.e-15
def test_python_parloop_vector():
m = UnitSquareMesh(4, 4)
fs = VectorFunctionSpace(m, "CG", 1)
f = Function(fs)
class MyExpression(Expression):
def eval(self, value, X):
value[:] = X
def value_shape(self):
return (2,)
f.interpolate(MyExpression())
X = m.coordinates
assert assemble((f - X)**2*dx)**.5 < 1.e-15
def test_python_parloop_vector_1D():
m = UnitIntervalMesh(4)
fs = VectorFunctionSpace(m, "CG", 1)
f = Function(fs)
class MyExpression(Expression):
def eval(self, value, X):
value[:] = X
def value_shape(self):
return (1,)
f.interpolate(MyExpression())
X = m.coordinates
assert assemble((f - X)**2*dx)**.5 < 1.e-15
def test_python_parloop_user_kwarg():
m = UnitSquareMesh(4, 4)
fs = FunctionSpace(m, "CG", 2)
f = Function(fs)
class MyExpression(Expression):
def eval(self, value, X, t=None):
value[:] = t
f.interpolate(MyExpression(t=10.0))
assert np.allclose(assemble(f*dx), 10.0)
def test_python_parloop_vector_user_kwarg():
m = UnitSquareMesh(4, 4)
fs = VectorFunctionSpace(m, "CG", 1)
f = Function(fs)
class MyExpression(Expression):
def eval(self, value, X, b=None, a=None):
value[0] = a
value[1] = b
def value_shape(self):
return (2,)
f.interpolate(MyExpression(a=1.0, b=2.0))
exact = Function(fs)
exact.interpolate(Expression(("1.0", "2.0")))
assert np.allclose(assemble((f - exact)**2*dx), 0.0)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/extrusion_utils.py
import numpy as np
from pyop2 import op2
import fiat_utils
import functionspace as fs
def compute_extruded_dofs(fiat_element, flat_dofs, layers):
"""Compute the number of dofs in a column."""
size = len(flat_dofs)
dofs_per_column = np.zeros(size, np.int32)
for i in range(size):
for j in range(2): # 2 is due to the process of extrusion
dofs_per_column[i] += (layers - j) * len(fiat_element.entity_dofs()[(i, j)][0])
return dofs_per_column
def compute_vertical_offsets(ent_dofs, flat_dofs):
"""Compute the offset between corresponding dofs in layers.
offsets[i] is the offset from the bottom of the stack to the
corresponding dof in the ith layer.
"""
size = len(flat_dofs)
offsets_per_vertical = np.zeros(size, np.int32)
for i in range(size):
if len(flat_dofs[i][0]) > 0:
offsets_per_vertical[i] = len(flat_dofs[i][0]) - len(ent_dofs[(i, 0)][0])
return offsets_per_vertical
def compute_offset(ent_dofs, flat_dofs, total_dofs):
"""Compute extruded offsets for flattened element.
offsets[i] is the number of dofs in the vertical for the ith
column of flattened mesh entities."""
size = len(flat_dofs)
res = np.zeros(total_dofs, np.int32)
vert_dofs = compute_vertical_offsets(ent_dofs, flat_dofs)
for i in range(size):
elems = len(flat_dofs[i])
dofs_per_elem = len(flat_dofs[i][0])
for j in range(elems):
for k in range(dofs_per_elem):
res[flat_dofs[i][j][k]] = vert_dofs[i]
return res
def total_num_dofs(flat_dofs):
"""Compute the total number of degrees of freedom in the extruded mesh."""
size = len(flat_dofs)
total = 0
for i in range(size):
total += len(flat_dofs[i]) * len(flat_dofs[i][0])
return total
def make_flat_fiat_element(ufl_cell_element, ufl_cell, flattened_entity_dofs):
"""Create a modified FIAT-style element.
Transform object from 3D-Extruded to 2D-flattened FIAT-style object."""
# Create base element
base_element = fiat_utils.fiat_from_ufl_element(ufl_cell_element)
# Alter base element
base_element.dual.entity_ids = flattened_entity_dofs
base_element.poly_set.num_members = total_num_dofs(flattened_entity_dofs)
return base_element
def make_extruded_coords(extruded_mesh, layer_height,
extrusion_type='uniform', kernel=None,
output_coords=None):
"""
Given either a kernel or a (fixed) layer_height, compute an
extruded coordinate field for an extruded mesh.
:arg extruded_mesh: an :class:`ExtrudedMesh` to extrude a
coordinate field for.
:arg layer_height: an equi-spaced height for each layer.
:arg extrusion_type: the type of extrusion to use. Predefined
options are either "uniform" (creating equi-spaced layers by
extruding in the (n+1)dth direction), "radial" (creating
equi-spaced layers by extruding in the outward direction from
the origin) or "radial_hedgehog" (creating equi-spaced layers
by extruding coordinates in the outward cell-normal
direction, needs a P1dgxP1 coordinate field).
:arg kernel: an optional kernel to carry out coordinate extrusion.
:arg output_coords: an optional :class:`~.Function` to write the
extruded coordinates into. If not provided, the coordinate
field in the :data:`extruded_mesh` will be written to.
The kernel signature (if provided) is::
void kernel(double **base_coords, double **ext_coords,
int **layer, double *layer_height)
The kernel iterates over the cells of the mesh and receives as
arguments the coordinates of the base cell (to read), the
coordinates on the extruded cell (to write to), the layer number
of each cell and the fixed layer height.
"""
base_coords = extruded_mesh._old_mesh.coordinates
if output_coords is None:
ext_coords = extruded_mesh.coordinates
else:
ext_coords = output_coords
vert_space = ext_coords.function_space().ufl_element()._B
if kernel is None and not (vert_space.degree() == 1 and
vert_space.family() in ['Lagrange',
'Discontinuous Lagrange']):
raise RuntimeError('Extrusion of coordinates is only possible for a P1 or P1dg interval unless a custom kernel is provided')
if kernel is not None:
pass
elif extrusion_type == 'uniform':
kernel = op2.Kernel("""
void uniform_extrusion_kernel(double **base_coords,
double **ext_coords,
int **layer,
double *layer_height) {
for ( int d = 0; d < %(base_map_arity)d; d++ ) {
for ( int c = 0; c < %(base_coord_dim)d; c++ ) {
ext_coords[2*d][c] = base_coords[d][c];
ext_coords[2*d+1][c] = base_coords[d][c];
}
ext_coords[2*d][%(base_coord_dim)d] = *layer_height * (layer[0][0]);
ext_coords[2*d+1][%(base_coord_dim)d] = *layer_height * (layer[0][0] + 1);
}
}""" % {'base_map_arity': base_coords.cell_node_map().arity,
'base_coord_dim': base_coords.function_space().cdim},
"uniform_extrusion_kernel")
elif extrusion_type == 'radial':
kernel = op2.Kernel("""
void radial_extrusion_kernel(double **base_coords,
double **ext_coords,
int **layer,
double *layer_height) {
for ( int d = 0; d < %(base_map_arity)d; d++ ) {
double norm = 0.0;
for ( int c = 0; c < %(base_coord_dim)d; c++ ) {
norm += base_coords[d][c] * base_coords[d][c];
}
norm = sqrt(norm);
for ( int c = 0; c < %(base_coord_dim)d; c++ ) {
ext_coords[2*d][c] = base_coords[d][c] * (1 + (*layer_height * layer[0][0])/norm);
ext_coords[2*d+1][c] = base_coords[d][c] * (1 + (*layer_height * (layer[0][0]+1))/norm);
}
}
}""" % {'base_map_arity': base_coords.cell_node_map().arity,
'base_coord_dim': base_coords.function_space().cdim},
"radial_extrusion_kernel")
elif extrusion_type == 'radial_hedgehog':
# Only implemented for interval in 2D and triangle in 3D.
# gdim != tdim already checked in ExtrudedMesh constructor.
if base_coords.cell().topological_dimension() not in [1, 2]:
raise NotImplementedError("Hedgehog extrusion not implemented for %s" % base_coords.cell())
kernel = op2.Kernel("""
void radial_hedgehog_extrusion_kernel(double **base_coords,
double **ext_coords,
int **layer,
double *layer_height) {
double v0[%(base_coord_dim)d];
double v1[%(base_coord_dim)d];
double n[%(base_map_arity)d];
double x[%(base_map_arity)d] = {0};
double dot = 0.0;
double norm = 0.0;
int i, c, d;
if (%(base_coord_dim)d == 2) {
/*
* normal is:
* (0 -1) (x2 - x1)
* (1 0) (y2 - y1)
*/
n[0] = -(base_coords[1][1] - base_coords[0][1]);
n[1] = base_coords[1][0] - base_coords[0][0];
} else if (%(base_coord_dim)d == 3) {
/*
* normal is
* v0 x v1
*
* /\
* v0/ \
* / \
* /------\
* v1
*/
for (i = 0; i < 3; ++i) {
v0[i] = base_coords[1][i] - base_coords[0][i];
v1[i] = base_coords[2][i] - base_coords[0][i];
}
n[0] = v0[1] * v1[2] - v0[2] * v1[1];
n[1] = v0[2] * v1[0] - v0[0] * v1[2];
n[2] = v0[0] * v1[1] - v0[1] * v1[0];
}
for (i = 0; i < %(base_map_arity)d; ++i) {
for (c = 0; c < %(base_coord_dim)d; ++c) {
x[i] += base_coords[c][i];
}
}
for (i = 0; i < %(base_map_arity)d; ++i) {
dot += x[i] * n[i];
norm += n[i] * n[i];
}
/*
* Make inward-pointing normals point out
*/
norm = sqrt(norm);
norm *= (dot < 0 ? -1 : 1);
for (d = 0; d < %(base_coord_dim)d; ++d) {
for (c = 0; c < %(base_map_arity)d; ++c ) {
ext_coords[2*d][c] = base_coords[d][c] + n[c] * layer_height[0] * layer[0][0] / norm;
ext_coords[2*d+1][c] = base_coords[d][c] + n[c] * layer_height[0] * (layer[0][0] + 1)/ norm;
}
}
}""" % {'base_map_arity': base_coords.cell_node_map().arity,
'base_coord_dim': base_coords.function_space().cdim},
"radial_hedgehog_extrusion_kernel")
else:
raise NotImplementedError('Unsupported extrusion type "%s"' % extrusion_type)
# Dat to hold layer number
layer_fs = fs.FunctionSpace(extruded_mesh, 'DG', 0)
layers = extruded_mesh.layers
layer = op2.Dat(layer_fs.dof_dset,
np.repeat(np.arange(layers-1, dtype=np.int32),
extruded_mesh.cell_set.total_size).reshape(layers-1, extruded_mesh.cell_set.total_size).T.ravel(), dtype=np.int32)
height = op2.Global(1, layer_height, dtype=float)
op2.par_loop(kernel,
ext_coords.cell_set,
base_coords.dat(op2.READ, base_coords.cell_node_map()),
ext_coords.dat(op2.WRITE, ext_coords.cell_node_map()),
layer(op2.READ, layer_fs.cell_node_map()),
height(op2.READ))
<file_sep>/tests/regression/test_upwind_flux.py
"""
This code does the following.
First, obtains an upwind DG0 approximation to div(u*D).
Then, tries to find a BDM1 flux F such that div F is
equal to this upwind approximation.
we have
\int_e phi D_1 dx = -\int_e grad phi . u D dx
+ \int_{\partial e} phi u.n \tilde{D} ds
where \tilde{D} is the value of D on the upwind face. For
DG0, grad phi = 0.
Then, if we define F such that
\int_f phi F.n ds = \int_f phi u.n \tilde{D} ds
then
\int_e phi div(F) ds = \int_{\partial e} phi u.n \tilde{D} ds
as required.
"""
from firedrake import *
import pytest
def run_test():
mesh = UnitIcosahedralSphereMesh(refinement_level=2)
global_normal = Expression(("x[0]/sqrt(x[0]*x[0]+x[1]*x[1]+x[2]*x[2])",
"x[1]/sqrt(x[0]*x[0]+x[1]*x[1]+x[2]*x[2])",
"x[2]/sqrt(x[0]*x[0]+x[1]*x[1]+x[2]*x[2])"))
mesh.init_cell_orientations(global_normal)
# Define function spaces and basis functions
V_dg = FunctionSpace(mesh, "DG", 0)
M = FunctionSpace(mesh, "RT", 1)
# advecting velocity
u0 = Expression(('-x[1]', 'x[0]', '0'))
u = Function(M).project(u0)
# Mesh-related functions
n = FacetNormal(mesh)
# ( dot(v, n) + |dot(v, n)| )/2.0
un = 0.5*(dot(u, n) + abs(dot(u, n)))
# D advection equation
phi = TestFunction(V_dg)
D = TrialFunction(V_dg)
a_mass = phi*D*dx
a_int = dot(grad(phi), -u*D)*dx
a_flux = (dot(jump(phi), un('+')*D('+') - un('-')*D('-')))*dS
arhs = (a_int + a_flux)
D1 = Function(V_dg)
D0 = Expression("exp(-pow(x[2],2) - pow(x[1],2))")
D = Function(V_dg).interpolate(D0)
D1problem = LinearVariationalProblem(a_mass, action(arhs, D), D1)
D1solver = LinearVariationalSolver(D1problem)
D1solver.solve()
# Surface Flux equation
V1 = FunctionSpace(mesh, "RT", 1)
w = TestFunction(V1)
Ft = TrialFunction(V1)
Fs = Function(V1)
aFs = (inner(w('+'), n('+'))*inner(Ft('+'), n('+')) +
inner(w('-'), n('-'))*inner(Ft('-'), n('-')))*dS
LFs = 2.0*(inner(w('+'), n('+'))*un('+')*D('+')
+ inner(w('-'), n('-'))*un('-')*D('-'))*dS
Fsproblem = LinearVariationalProblem(aFs, LFs, Fs)
Fssolver = LinearVariationalSolver(Fsproblem,
solver_parameters={'ksp_type': 'preonly'})
Fssolver.solve()
divFs = Function(V_dg)
solve(a_mass == phi*div(Fs)*dx, divFs)
assert errornorm(divFs, D1, degree_rise=0) < 1e-12
def test_upwind_flux():
run_test()
@pytest.mark.parallel
def test_upwind_flux_parallel():
run_test()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_par_loops.py
import pytest
import numpy as np
from firedrake import *
@pytest.fixture
def f():
m = UnitIntervalMesh(2)
cg = FunctionSpace(m, "CG", 1)
dg = FunctionSpace(m, "DG", 0)
c = Function(cg)
d = Function(dg)
return c, d
@pytest.fixture
def f_mixed():
m = UnitIntervalMesh(2)
cg = FunctionSpace(m, "CG", 1)
dg = FunctionSpace(m, "DG", 0)
return Function(cg*dg)
@pytest.fixture
def const(f):
return Constant(1.0, domain=f[0].function_space().mesh().ufl_domain())
@pytest.fixture
def f_extruded():
i = UnitIntervalMesh(2)
m = ExtrudedMesh(i, 2, layer_height=0.1)
cg = FunctionSpace(m, "CG", 1)
dg = FunctionSpace(m, "DG", 0)
c = Function(cg)
d = Function(dg)
return c, d
def test_direct_par_loop(f):
c, _ = f
par_loop("""*c = 1;""", direct, {'c': (c, WRITE)})
assert all(c.dat.data == 1)
@pytest.mark.xfail
def test_mixed_direct_par_loop(f_mixed):
par_loop("""*c = 1;""", direct, {'c': (f_mixed, WRITE)})
assert all(f_mixed.dat.data == 1)
@pytest.mark.parametrize('idx', [0, 1])
def test_mixed_direct_par_loop_components(f_mixed, idx):
par_loop("""*c = 1;""", direct, {'c': (f_mixed[idx], WRITE)})
assert all(f_mixed.dat[idx].data == 1)
def test_direct_par_loop_read_const(f, const):
c, _ = f
const.assign(10.0)
par_loop("""*c = *constant;""", direct, {'c': (c, WRITE), 'constant': (const, READ)})
assert np.allclose(c.dat.data, const.dat.data)
def test_indirect_par_loop_read_const(f, const):
_, d = f
const.assign(10.0)
par_loop("""for (int i = 0; i < d.dofs; i++) d[0][0] = *constant;""",
dx, {'d': (d, WRITE), 'constant': (const, READ)})
assert np.allclose(d.dat.data, const.dat.data)
@pytest.mark.xfail
def test_indirect_par_loop_read_const_mixed(f_mixed, const):
const.assign(10.0)
par_loop("""for (int i = 0; i < d.dofs; i++) d[0][0] = *constant;""",
dx, {'d': (f_mixed, WRITE), 'constant': (const, READ)})
assert np.allclose(f_mixed.dat.data, const.dat.data)
# FIXME: this is supposed to work, but for unknown reasons fails with
# MapValueError: Iterset of arg 1 map 0 doesn't match ParLoop iterset.
@pytest.mark.xfail
@pytest.mark.parametrize('idx', [0, 1])
def test_indirect_par_loop_read_const_mixed_component(f_mixed, const, idx):
const.assign(10.0)
par_loop("""for (int i = 0; i < d.dofs; i++) d[0][0] = *constant;""",
dx, {'d': (f_mixed[idx], WRITE), 'constant': (const, READ)})
assert np.allclose(f_mixed.dat[idx].data, const.dat.data)
def test_par_loop_const_write_error(f, const):
_, d = f
with pytest.raises(RuntimeError):
par_loop("""c[0] = d[0];""", direct, {'c': (const, WRITE), 'd': (d, READ)})
def test_cg_max_field(f):
c, d = f
d.interpolate(Expression("x[0]"))
par_loop("""
for (int i=0; i<c.dofs; i++)
c[i][0] = fmax(c[i][0], d[0][0]);""",
dx, {'c': (c, RW), 'd': (d, READ)})
assert (c.dat.data == [1./4, 3./4, 3./4]).all()
def test_cg_max_field_extruded(f_extruded):
c, d = f_extruded
d.interpolate(Expression("x[0]"))
par_loop("""
for (int i=0; i<c.dofs; i++)
c[i][0] = (c[i][0] > d[0][0] ? c[i][0] : d[0][0]);""",
dx, {'c': (c, RW), 'd': (d, READ)})
assert (c.dat.data == [1./4, 1./4, 1./4,
3./4, 3./4, 3./4,
3./4, 3./4, 3./4]).all()
def test_walk_facets_rt():
m = UnitSquareMesh(3, 3)
V = FunctionSpace(m, 'RT', 1)
f1 = Function(V)
f2 = Function(V)
project(Expression(('x[0]', 'x[1]')), f1)
par_loop("""
for (int i = 0; i < f1.dofs; i++) {
f2[i][0] = f1[i][0];
}""", dS, {'f1': (f1, READ), 'f2': (f2, WRITE)})
par_loop("""
for (int i = 0; i < f1.dofs; i++) {
f2[i][0] = f1[i][0];
}""", ds, {'f1': (f1, READ), 'f2': (f2, WRITE)})
assert errornorm(f1, f2, degree_rise=0) < 1e-10
<file_sep>/tests/test_ffc_interface.py
import pytest
from firedrake import *
import os
@pytest.fixture(scope='module')
def fs():
mesh = UnitSquareMesh(1, 1)
return FunctionSpace(mesh, 'CG', 1)
@pytest.fixture
def mass(fs):
u = TrialFunction(fs)
v = TestFunction(fs)
return u * v * dx
@pytest.fixture
def mixed_mass(fs):
u, r = TrialFunctions(fs*fs)
v, s = TestFunctions(fs*fs)
return (u*v + r*s) * dx
@pytest.fixture
def laplace(fs):
u = TrialFunction(fs)
v = TestFunction(fs)
return inner(grad(u), grad(v)) * dx
@pytest.fixture
def rhs(fs):
v = TestFunction(fs)
g = Function(fs)
return g * v * ds
@pytest.fixture
def rhs2(fs):
v = TestFunction(fs)
f = Function(fs)
g = Function(fs)
return f * v * dx + g * v * ds
@pytest.fixture
def cache_key(mass):
return ffc_interface.FFCKernel(mass, 'mass', parameters["form_compiler"]).cache_key
@pytest.mark.xfail("not hasattr(ffc_interface.constants, 'PYOP2_VERSION')")
class TestFFCCache:
"""FFC code generation cache tests."""
def test_ffc_cache_dir_exists(self):
"""Importing ffc_interface should create FFC Kernel cache dir."""
assert os.path.exists(ffc_interface.FFCKernel._cachedir)
def test_ffc_cache_persist_on_disk(self, cache_key):
"""FFCKernel should be persisted on disk."""
assert os.path.exists(
os.path.join(ffc_interface.FFCKernel._cachedir, cache_key))
def test_ffc_cache_read_from_disk(self, cache_key):
"""Loading an FFCKernel from disk should yield the right object."""
assert ffc_interface.FFCKernel._read_from_disk(
cache_key).cache_key == cache_key
def test_ffc_same_form(self, mass):
"""Compiling the same form twice should load kernels from cache."""
k1 = ffc_interface.compile_form(mass, 'mass')
k2 = ffc_interface.compile_form(mass, 'mass')
assert k1 is k2
assert all(k1_[-1] is k2_[-1] for k1_, k2_ in zip(k1, k2))
def test_ffc_same_mixed_form(self, mixed_mass):
"""Compiling a mixed form twice should load kernels from cache."""
k1 = ffc_interface.compile_form(mixed_mass, 'mixed_mass')
k2 = ffc_interface.compile_form(mixed_mass, 'mixed_mass')
assert k1 is k2
assert all(k1_[-1] is k2_[-1] for k1_, k2_ in zip(k1, k2))
def test_ffc_different_forms(self, mass, laplace):
"""Compiling different forms should not load kernels from cache."""
k1, = ffc_interface.compile_form(mass, 'mass')
k2, = ffc_interface.compile_form(laplace, 'mass')
assert k1[-1] is not k2[-1]
def test_ffc_different_names(self, mass):
"""Compiling different forms should not load kernels from cache."""
k1, = ffc_interface.compile_form(mass, 'mass')
k2, = ffc_interface.compile_form(mass, 'laplace')
assert k1[-1] is not k2[-1]
def test_ffc_cell_kernel(self, mass):
k = ffc_interface.compile_form(mass, 'mass')
assert 'cell_integral' in k[0][-1].code and len(k) == 1
def test_ffc_exterior_facet_kernel(self, rhs):
k = ffc_interface.compile_form(rhs, 'rhs')
assert 'exterior_facet_integral' in k[0][-1].code and len(k) == 1
def test_ffc_cell_exterior_facet_kernel(self, rhs2):
k = ffc_interface.compile_form(rhs2, 'rhs2')
assert 'cell_integral' in k[0][-1].code and \
'exterior_facet_integral' in k[1][-1].code and len(k) == 2
if __name__ == '__main__':
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_helmholtz_nonlinear_diffusion.py
"""This demo program solves Helmholtz's equation
- div D(u) grad u(x, y) + kappa u(x,y) = f(x, y)
with
D(u) = 1 + alpha * u**2
alpha = 0.1
kappa = 1
on the unit square with source f given by
f(x, y) = -8*pi^2*alpha*cos(2*pi*x)*cos(2*pi*y)^3*sin(2*pi*x)^2
- 8*pi^2*alpha*cos(2*pi*x)^3*cos(2*pi*y)*sin(2*pi*y)^2
+ 8*pi^2*(alpha*cos(2*pi*x)^2*cos(2*pi*y)^2 + 1)
*cos(2*pi*x)*cos(2*pi*y)
+ kappa*cos(2*pi*x)*cos(2*pi*y)
and the analytical solution
u(x, y) = cos(x*2*pi)*cos(y*2*pi)
"""
import pytest
from firedrake import *
def helmholtz(x, quadrilateral=False, parameters={}):
# Create mesh and define function space
mesh = UnitSquareMesh(2 ** x, 2 ** x, quadrilateral=quadrilateral)
V = FunctionSpace(mesh, "CG", 1)
# Define variational problem
kappa = 1
alpha = 0.1
u = Function(V)
v = TestFunction(V)
f = Function(V)
D = 1 + alpha * u * u
f.interpolate(
Expression("-8*pi*pi*%(alpha)s*cos(2*pi*x[0])*cos(2*pi*x[1])\
*cos(2*pi*x[1])*cos(2*pi*x[1])*sin(2*pi*x[0])*sin(2*pi*x[0])\
- 8*pi*pi*%(alpha)s*cos(2*pi*x[0])*cos(2*pi*x[0])\
*cos(2*pi*x[0])*cos(2*pi*x[1])*sin(2*pi*x[1])*sin(2*pi*x[1])\
+ 8*pi*pi*(%(alpha)s*cos(2*pi*x[0])*cos(2*pi*x[0])\
*cos(2*pi*x[1])*cos(2*pi*x[1]) + 1)*cos(2*pi*x[0])*cos(2*pi*x[1])\
+ %(kappa)s*cos(2*pi*x[0])*cos(2*pi*x[1])"
% {'alpha': alpha, 'kappa': kappa}))
a = (dot(grad(v), D * grad(u)) + kappa * v * u) * dx
L = f * v * dx
solve(a - L == 0, u, solver_parameters=parameters)
f.interpolate(Expression("cos(x[0]*2*pi)*cos(x[1]*2*pi)"))
return sqrt(assemble(dot(u - f, u - f) * dx))
def run_convergence_test(quadrilateral=False, parameters={}):
import numpy as np
l2_diff = np.array([helmholtz(i, quadrilateral, parameters) for i in range(3, 6)])
return np.log2(l2_diff[:-1] / l2_diff[1:])
def run_l2_conv():
assert (run_convergence_test() > 1.8).all()
def test_l2_conv_serial():
run_l2_conv()
@pytest.mark.parallel
def test_l2_conv_parallel():
run_l2_conv()
def run_l2_conv_on_quadrilaterals():
assert (run_convergence_test(quadrilateral=True) > 1.8).all()
def test_l2_conv_on_quadrilaterals_serial():
run_l2_conv_on_quadrilaterals()
@pytest.mark.parallel
def test_l2_conv_on_quadrilaterals_parallel():
run_l2_conv_on_quadrilaterals()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/fiat_utils.py
from weakref import WeakKeyDictionary
import ffc
_fiat_element_cache = WeakKeyDictionary()
_cells = {
1: {2: "interval"},
2: {3: "triangle", 4: "quadrilateral"},
3: {4: "tetrahedron"}
}
def fiat_from_ufl_element(ufl_element):
try:
return _fiat_element_cache[ufl_element]
except KeyError:
fiat_element = ffc.create_actual_fiat_element(ufl_element)
_fiat_element_cache[ufl_element] = fiat_element
return fiat_element
def flat_entity_dofs(fiat_element):
"""Returns entity dofs with flattened dimensions.
For outer product elements, dimensions are pairs instead of integers.
Parts of the pairs are added, and their corresponding values are merged.
For example:
{(0, 0): {0: [0], 1: [1], 2: [2], 3: [3]},
(0, 1): {0: [4], 1: [5]},
(1, 0): {0: [6], 1: [7]},
(1, 1): {0: [8]}}
is flattened into:
{0: {0: [0], 1: [1], 2: [2], 3: [3]},
1: {0: [4], 1: [5], 2: [6], 3: [7]},
2: {0: [8]}}
Maps with integer dimensions are unaffected.
"""
entity_dofs = fiat_element.entity_dofs()
f = lambda x: sum(x) if isinstance(x, tuple) else x
indexless = dict((dim, []) for dim in set(map(f, entity_dofs.keys())))
for (dim, entity) in sorted(entity_dofs.iteritems()):
indexless[f(dim)].extend(entity.itervalues())
return dict((dim, dict((i, indices) for i, indices in enumerate(e)))
for dim, e in indexless.iteritems())
<file_sep>/tests/extrusion/test_extrusion_interval.py
import pytest
import numpy as np
from firedrake import *
def integrate_one(intervals):
m = UnitIntervalMesh(intervals)
layers = intervals
mesh = ExtrudedMesh(m, layers, layer_height=1.0 / layers)
V = FunctionSpace(mesh, 'CG', 1)
u = Function(V)
u.interpolate(Expression("1"))
return assemble(u * dx)
def test_unit_interval():
assert abs(integrate_one(5) - 1) < 1e-12
def test_interval_div_free():
m = UnitIntervalMesh(50)
mesh = ExtrudedMesh(m, 50)
V = VectorFunctionSpace(mesh, 'CG', 3)
u = Function(V)
u.interpolate(Expression(('x[0]*x[0]*x[1]', '-x[0]*x[1]*x[1]')))
# u is pointwise divergence free, so the integral should also be
# div-free.
assert np.allclose(assemble(div(u)*dx), 0)
L2 = FunctionSpace(mesh, 'DG', 2)
v = TestFunction(L2)
f = assemble(div(u)*v*dx)
# Check pointwise div-free
assert np.allclose(f.dat.data, 0)
def test_periodic_interval_div_free():
m = PeriodicUnitIntervalMesh(50)
mesh = ExtrudedMesh(m, 50)
V = VectorFunctionSpace(mesh, 'CG', 3)
u = Function(V)
u.interpolate(Expression(('sin(2*pi*x[0])',
'-2*pi*x[1]*cos(2*pi*x[0])')))
# u is pointwise divergence free, so the integral should also be
# div-free.
assert np.allclose(assemble(div(u)*dx), 0)
L2 = FunctionSpace(mesh, 'DG', 2)
v = TestFunction(L2)
f = assemble(div(u)*v*dx)
# Check pointwise div-free
assert np.allclose(f.dat.data, 0)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_side_strong_bcs.py
"""This demo program sets the top, bottom and side boundaries
of an extruded unit square. We then check against the actual solution
of the equation.
"""
import pytest
from firedrake import *
def run_test_3D(size, parameters={}, test_mode=False):
# Create mesh and define function space
m = UnitSquareMesh(size, size)
layers = size
mesh = ExtrudedMesh(m, layers, layer_height=1.0 / layers)
# Define variational problem
V = FunctionSpace(mesh, "CG", 1)
exp = Expression('x[0]*x[0] - x[1]*x[1] - x[2]*x[2]')
bcs = [DirichletBC(V, exp, "bottom"),
DirichletBC(V, exp, "top"),
DirichletBC(V, exp, 1),
DirichletBC(V, exp, 2),
DirichletBC(V, exp, 3),
DirichletBC(V, exp, 4)]
v = TestFunction(V)
u = TrialFunction(V)
a = dot(grad(u), grad(v)) * dx
f = Function(V)
f.assign(2)
L = v * f * dx
out = Function(V)
exact = Function(V)
exact.interpolate(exp)
solve(a == L, out, bcs=bcs)
res = sqrt(assemble(dot(out - exact, out - exact) * dx))
if not test_mode:
print "The error is ", res
file = File("side-bcs-computed.pvd")
file << out
file = File("side-bcs-expected.pvd")
file << exact
return res
def run_test_2D(intervals, parameters={}, test_mode=False):
# Create mesh and define function space
m = UnitIntervalMesh(intervals)
layers = intervals
mesh = ExtrudedMesh(m, layers, layer_height=1.0 / layers)
# Define variational problem
V = FunctionSpace(mesh, "CG", 1)
exp = Expression('x[0]*x[0] - 2*x[1]*x[1]')
bcs = [DirichletBC(V, exp, "bottom"),
DirichletBC(V, exp, "top"),
DirichletBC(V, exp, 1),
DirichletBC(V, exp, 2)]
v = TestFunction(V)
u = TrialFunction(V)
a = dot(grad(u), grad(v)) * dx
f = Function(V)
f.assign(2)
L = v * f * dx
out = Function(V)
exact = Function(V)
exact.interpolate(exp)
solve(a == L, out, bcs=bcs)
res = sqrt(assemble(dot(out - exact, out - exact) * dx))
if not test_mode:
print "The error is ", res
file = File("side-bcs-computed.pvd")
file << out
file = File("side-bcs-expected.pvd")
file << exact
return res
def test_extrusion_side_strong_bcs():
assert (run_test_3D(3, test_mode=True) < 1.e-13)
def test_extrusion_side_strong_bcs_large():
assert (run_test_3D(6, test_mode=True) < 1.3e-08)
def test_extrusion_side_strong_bcs_2D():
assert (run_test_2D(2, test_mode=True) < 1.e-13)
def test_extrusion_side_strong_bcs_2D_large():
assert (run_test_2D(4, test_mode=True) < 1.e-12)
def test_get_all_bc_nodes():
m = UnitSquareMesh(1, 1)
m = ExtrudedMesh(m, layers=2)
V = FunctionSpace(m, 'CG', 2)
bc = DirichletBC(V, 0, 1)
# Exterior facet nodes on a single column are:
# o--o--o
# | |
# o o o
# | |
# o--o--o
# | |
# o o o
# | |
# o--o--o
#
# And there is 1 base facet with the "1" marker. So we expect to
# see 15 dofs in the bc object.
assert len(bc.nodes) == 15
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/projection.py
import ufl
import expression
import function
import functionspace
import solving
import ufl_expr
__all__ = ['project']
# Store the solve function to use in a variable so external packages
# (dolfin-adjoint) can override it.
_solve = solving.solve
def project(v, V, bcs=None, mesh=None,
solver_parameters=None,
form_compiler_parameters=None,
name=None):
"""Project an :class:`.Expression` or :class:`.Function` into a :class:`.FunctionSpace`
:arg v: the :class:`.Expression`, :class:`ufl.Expr` or
:class:`.Function` to project
:arg V: the :class:`.FunctionSpace` or :class:`.Function` to project into
:arg bcs: boundary conditions to apply in the projection
:arg mesh: the mesh to project into
:arg solver_parameters: parameters to pass to the solver used when
projecting.
:arg form_compiler_parameters: parameters to the form compiler
:arg name: name of the resulting :class:`.Function`
If ``V`` is a :class:`.Function` then ``v`` is projected into
``V`` and ``V`` is returned. If `V` is a :class:`.FunctionSpace`
then ``v`` is projected into a new :class:`.Function` and that
:class:`.Function` is returned.
The ``bcs``, ``mesh`` and ``form_compiler_parameters`` are
currently ignored."""
if isinstance(V, functionspace.FunctionSpaceBase):
ret = function.Function(V, name=name)
elif isinstance(V, function.Function):
ret = V
V = V.function_space()
else:
raise RuntimeError(
'Can only project into functions and function spaces, not %r'
% type(V))
if isinstance(v, expression.Expression):
shape = v.value_shape()
# Build a function space that supports PointEvaluation so that
# we can interpolate into it.
if isinstance(V.ufl_element().degree(), tuple):
deg = max(V.ufl_element().degree())
else:
deg = V.ufl_element().degree()
if v.rank() == 0:
fs = functionspace.FunctionSpace(V.mesh(), 'DG', deg+1)
elif v.rank() == 1:
fs = functionspace.VectorFunctionSpace(V.mesh(), 'DG',
deg+1,
dim=shape[0])
else:
raise NotImplementedError(
"Don't know how to project onto tensor-valued function spaces")
f = function.Function(fs)
f.interpolate(v)
v = f
elif isinstance(v, function.Function):
if v.function_space().mesh() != ret.function_space().mesh():
raise RuntimeError("Can't project between mismatching meshes")
elif not isinstance(v, ufl.core.expr.Expr):
raise RuntimeError("Can't only project from expressions and functions, not %r" % type(v))
if v.shape() != ret.shape():
raise RuntimeError('Shape mismatch between source %s and target function spaces %s in project' % (v.shape(), ret.shape()))
p = ufl_expr.TestFunction(V)
q = ufl_expr.TrialFunction(V)
a = ufl.inner(p, q) * V.mesh()._dx
L = ufl.inner(p, v) * V.mesh()._dx
# Default to 1e-8 relative tolerance
if solver_parameters is None:
solver_parameters = {'ksp_type': 'cg', 'ksp_rtol': 1e-8}
else:
solver_parameters.setdefault('ksp_type', 'cg')
solver_parameters.setdefault('ksp_rtol', 1e-8)
_solve(a == L, ret, bcs=bcs,
solver_parameters=solver_parameters,
form_compiler_parameters=form_compiler_parameters)
return ret
<file_sep>/tests/regression/test_change_coordinates.py
"""This tests that exchanging the coordinate field for one of a different dimension does the right thing."""
import pytest
from firedrake import *
@pytest.mark.parametrize("dim", [2, 3])
def test_immerse_1d(dim):
m = UnitIntervalMesh(5)
cfs = VectorFunctionSpace(m, "Lagrange", 1, dim)
new_coords = Function(cfs)
m.coordinates = new_coords
assert m.ufl_cell().geometric_dimension() == dim
def test_immerse_2d():
m = UnitSquareMesh(2, 2)
cfs = VectorFunctionSpace(m, "Lagrange", 1, 3)
new_coords = Function(cfs)
m.coordinates = new_coords
assert m.ufl_cell().geometric_dimension() == 3
def test_project_2d():
m = CircleManifoldMesh(5)
cfs = VectorFunctionSpace(m, "Lagrange", 1, 1)
new_coords = Function(cfs)
m.coordinates = new_coords
assert m.ufl_cell().geometric_dimension() == 1
def test_immerse_extruded():
m1 = UnitIntervalMesh(5)
m = ExtrudedMesh(m1, 10)
cfs = VectorFunctionSpace(m, "Lagrange", 1, 3)
new_coords = Function(cfs)
m.coordinates = new_coords
assert m.ufl_cell().geometric_dimension() == 3
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/assembly_cache.py
"""Firedrake by default caches assembled forms. This means that it is
generally not necessary for users to manually lift assembly of linear
operators out of timestepping loops; the same performance benefit will
be realised automatically by the assembly cache.
In order to prevent the assembly cache leaking memory, a simple cache
eviction strategy is implemented. This is documented below. In
addition, the following parameters control the operation of the
assembly_cache:
:data:`parameters["assembly_cache"]["enabled"]`
a boolean value used to disable the assembly cache if required.
:data:`parameters["assembly_cache"]["eviction"]`
a boolean value used to disable the cache eviction
strategy. Disabling cache eviction can lead to memory leaks so is
discouraged in almost all circumstances.
:data:`parameters["assembly_cache"]["max_misses"]`
attempting to cache objects whose inputs change every time they are
assembled is a waste of memory. This parameter sets a maximum number
of consecutive misses beyond which a form will be marked as
uncachable.
:data:`parameters["assembly_cache"]["max_bytes"]`
absolute limit on the size of the assembly cache in bytes. This
defaults to :data:`float("inf")`.
:data:`parameters["assembly_cache"]["max_factor"]`
limit on the size of the assembly cache relative to the amount of
memory per core on the current system. This defaults to 0.6.
"""
import numpy as np
import weakref
from collections import defaultdict
from pyop2.logger import debug, warning
from pyop2.mpi import MPI, _MPI
import function
import matrix
from parameters import parameters
from petsc import PETSc
try:
# Estimate the amount of memory per core may use.
import psutil
memory = np.array([psutil.virtual_memory().total/psutil.cpu_count()])
if MPI.comm.size > 1:
MPI.comm.Allreduce(_MPI.IN_PLACE, memory, _MPI.MIN)
except ImportError:
memory = None
class _DependencySnapshot(object):
"""Record the dependencies of a form at a particular point in order to
establish whether a cached form is valid."""
def __init__(self, form):
# For each dependency, we store a weak reference and the
# current version number.
ref = lambda dep: (weakref.ref(dep), dep.dat._version)
deps = []
coords = form.integrals()[0].domain().data().coordinates
deps.append(ref(coords))
for c in form.coefficients():
deps.append(ref(c))
self.dependencies = tuple(deps)
def valid(self, form):
"""Check whether form is valid with respect to this dependency snapshot."""
original_coords = self.dependencies[0][0]()
if original_coords:
coords = form.integrals()[0].domain().data().coordinates
if coords is not original_coords or \
coords.dat._version != self.dependencies[0][1]:
return False
else:
return False
# Since UFL sorts the coefficients by count (creation index),
# further sorting here is not required.
deps = form.coefficients()
for original_d, dep in zip(self.dependencies[1:], deps):
original_dep = original_d[0]()
if original_dep:
if dep is not original_dep or dep.dat._version != original_d[1]:
return False
else:
return False
return True
class _BCSnapshot(object):
"""Record the boundary conditions which were applied to a form."""
def __init__(self, bcs):
self.bcs = map(weakref.ref, bcs) if bcs is not None else None
def valid(self, bcs):
if len(bcs) != len(self.bcs):
return False
for bc, wbc in zip(bcs, self.bcs):
if bc != wbc():
return False
return True
class _CacheEntry(object):
"""This is the basic caching unit. The form signature forms the key for
each CacheEntry, while a reference to the main data object is kept.
Additionally a list of Snapshot objects are kept in self.dependencies that
together form a snapshot of all the data objects used during assembly.
The validity of each CacheEntry object depends on the validity of its
dependencies (i.e., that none of the referred objects have changed)."""
def __init__(self, obj, form, bcs):
self.form = form
self.dependencies = _DependencySnapshot(form)
self.bcs = _BCSnapshot(bcs)
if isinstance(obj, float):
self.obj = np.float64(obj)
else:
self.obj = obj.duplicate()
global _assemble_count
self._assemble_count += 1
self.value = self._assemble_count
if MPI.comm.size > 1:
tmp = np.array([obj.nbytes])
MPI.comm.Allreduce(_MPI.IN_PLACE, tmp, _MPI.MAX)
self.nbytes = tmp[0]
else:
self.nbytes = obj.nbytes
_assemble_count = 0
def is_valid(self, form, bcs):
return self.dependencies.valid(form) and self.bcs.valid(bcs)
def get_object(self):
return self.obj
class AssemblyCache(object):
"""This is the central point of the assembly cache subsystem. This is a
singleton object so all the stored cache entries will reside in the single
instance object returned.
It is not usually necessary for users to access the
:class:`AssemblyCache` object directly, but this may occassionally
be useful when studying performance problems.
"""
_instance = None
def __new__(cls):
if not cls._instance:
cls._instance = super(AssemblyCache, cls).__new__(cls)
cls._instance._hits = 0
cls._instance._hits_size = 0
cls._instance.cache = {}
cls._instance.invalid_count = defaultdict(int)
cls._instance.evictwarned = False
return cls._instance
def _lookup(self, form, bcs, ffc_parameters):
form_sig = form.signature()
parms, cache_entry = self.cache.get(form_sig, (None, None))
retval = None
if cache_entry is not None:
if parms != str(ffc_parameters) or not cache_entry.is_valid(form, bcs):
self.invalid_count[form_sig] += 1
del self.cache[form_sig]
return None
else:
self.invalid_count[form_sig] = 0
retval = cache_entry.get_object()
self._hits += 1
self._hits_size += retval.nbytes
return retval
def _store(self, obj, form, bcs, ffc_parameters):
form_sig = form.signature()
if self.invalid_count[form_sig] > parameters["assembly_cache"]["max_misses"]:
if self.invalid_count[form_sig] == \
parameters["assembly_cache"]["max_misses"] + 1:
debug("form %s missed too many times, excluding from cache." % form)
else:
cache_entry = _CacheEntry(obj, form, bcs)
self.cache[form_sig] = str(ffc_parameters), cache_entry
self.evict()
def evict(self):
"""Run the cache eviction algorithm. This works out the permitted
cache size and deletes objects until it is achieved. Cache values are
assumed to have a :attr:`value` attribute and eviction occurs in
increasing :attr:`value` order. Currently :attr:`value` is an index of
the assembly operation, so older operations are evicted first.
The cache will be evicted down to 90% of permitted size.
The permitted size is either the explicit
:data:`parameters["assembly_cache"]["max_bytes"]` or it is the amount of
memory per core scaled by :data:`parameters["assembly_cache"]["max_factor"]`
(by default the scale factor is 0.6).
In MPI parallel, the nbytes of each cache entry is set to the maximum
over all processes, while the available memory is set to the
minimum. This produces a conservative caching policy which is
guaranteed to result in the same evictions on each processor.
"""
if not parameters["assembly_cache"]["eviction"]:
return
max_cache_size = min(parameters["assembly_cache"]["max_bytes"] or float("inf"),
(memory or float("inf"))
* parameters["assembly_cache"]["max_factor"]
)
if max_cache_size == float("inf"):
if not self.evictwarned:
warning("No maximum assembly cache size. Install psutil or risk leaking memory!")
self.evictwarned = True
return
cache_size = self.nbytes
if cache_size < max_cache_size:
return
debug("Cache eviction triggered. %s bytes in cache, %s bytes allowed" %
(cache_size, max_cache_size))
# Evict down to 90% full.
bytes_to_evict = cache_size - 0.9 * max_cache_size
sorted_cache = sorted(self.cache.items(), key=lambda x: x[1][1].value)
nbytes = lambda x: x[1][1].nbytes
candidates = []
while bytes_to_evict > 0:
next = sorted_cache.pop(0)
candidates.append(next)
bytes_to_evict -= nbytes(next)
for c in reversed(candidates):
if bytes_to_evict + nbytes(c) < 0:
# We may have been overzealous.
bytes_to_evict += nbytes(c)
else:
del self.cache[c[0]]
def clear(self):
"""Clear the cache contents."""
self.cache = {}
self._hits = 0
self._hits_size = 0
self.invalid_count = defaultdict(int)
@property
def num_objects(self):
"""The number of objects currently in the cache."""
return len(self.cache)
@property
def cache_stats(self):
"""Consolidated statistics for the cache contents"""
stats = "OpCache statistics: \n"
stats += "\tnum_stored=%d\tbytes=%d\trealbytes=%d\thits=%d\thit_bytes=%d" % \
(self.num_objects, self.nbytes, self.realbytes, self._hits,
self._hits_size)
return stats
@property
def nbytes(self):
"""An estimate of the total number of bytes in the cached objects."""
return sum([entry.nbytes for _, entry in self.cache.values()])
@property
def realbytes(self):
"""An estimate of the total number of bytes for which the cache holds
the sole reference to an object."""
tot_bytes = 0
for _, entry in self.cache.values():
obj = entry.get_object()
if not (hasattr(obj, "_cow_is_copy_of") and obj._cow_is_copy_of):
tot_bytes += entry.nbytes
return tot_bytes
def _cache_thunk(thunk, form, result, form_compiler_parameters=None):
"""Wrap thunk so that thunk is only executed if its target is not in
the cache."""
if form_compiler_parameters is None:
form_compiler_parameters = parameters["form_compiler"]
def inner(bcs):
cache = AssemblyCache()
if not parameters["assembly_cache"]["enabled"]:
return thunk(bcs)
obj = cache._lookup(form, bcs, form_compiler_parameters)
if obj is not None:
if isinstance(result, float):
# 0-form case
assert isinstance(obj, float)
r = obj
elif isinstance(result, function.Function):
# 1-form
result.dat = obj
r = result
elif isinstance(result, matrix.Matrix):
# 2-form
if obj.handle is not result._M.handle:
obj.handle.copy(result._M.handle,
PETSc.Mat.Structure.DIFFERENT_NONZERO_PATTERN)
# Ensure result matrix is assembled (MatCopy_Nest bug)
if not result._M.handle.assembled:
result._M.handle.assemble()
r = result
else:
raise TypeError("Unknown result type")
return r
r = thunk(bcs)
if isinstance(r, float):
# 0-form case
cache._store(r, form, bcs, form_compiler_parameters)
elif isinstance(r, function.Function):
# 1-form
cache._store(r.dat, form, bcs, form_compiler_parameters)
elif isinstance(r, matrix.Matrix):
# 2-form
cache._store(r._M, form, bcs, form_compiler_parameters)
else:
raise TypeError("Unknown result type")
return r
return inner
<file_sep>/tox.ini
[flake8]
ignore = E501,F403,E226,E265,E713,E112
exclude = .git,__pycache__,build,.tox,dist,./evtk,./pylit
<file_sep>/tests/regression/test_facets.py
import pytest
import numpy as np
from firedrake import *
@pytest.fixture
def f():
m = UnitSquareMesh(1, 1)
fs = FunctionSpace(m, "CG", 1)
f = Function(fs)
f.interpolate(Expression("x[0]"))
return f
@pytest.fixture(scope='module')
def dg_trial_test():
# Interior facet tests hard code order in which cells were
# numbered, so don't reorder this mesh.
m = UnitSquareMesh(1, 1, reorder=False)
V = FunctionSpace(m, "DG", 0)
u = TrialFunction(V)
v = TestFunction(V)
return u, v
def test_external_integral(f):
assert abs(assemble(f * ds) - 2.0) < 1.0e-14
def test_bottom_external_integral(f):
assert abs(assemble(f * ds(3)) - 0.5) < 1.0e-14
def test_top_external_integral(f):
assert abs(assemble(f * ds(4)) - 0.5) < 1.0e-14
def test_left_external_integral(f):
assert abs(assemble(f * ds(1))) < 1.0e-14
def test_right_external_integral(f):
assert abs(assemble(f * ds(2)) - 1.0) < 1.0e-14
def test_internal_integral(f):
assert abs(assemble(f('+') * dS) - 1.0 / (2.0 ** 0.5)) < 1.0e-14
def test_facet_integral_with_argument(f):
v = TestFunction(f.function_space())
assert np.allclose(assemble(f*v*ds).dat.data_ro.sum(), 2.0)
def test_bilinear_cell_integral(dg_trial_test):
u, v = dg_trial_test
cell = assemble(u*v*dx).M.values
# each diagonal entry should be volume of cell
assert np.allclose(np.diag(cell), 0.5)
# all off-diagonals should be zero
cell[range(2), range(2)] = 0.0
assert np.allclose(cell, 0.0)
def test_bilinear_exterior_facet_integral(dg_trial_test):
u, v = dg_trial_test
outer_facet = assemble(u*v*ds).M.values
# each diagonal entry should be length of exterior facet in this
# cell (2)
assert np.allclose(np.diag(outer_facet), 2.0)
# all off-diagonals should be zero
outer_facet[range(2), range(2)] = 0.0
assert np.allclose(outer_facet, 0.0)
@pytest.mark.parametrize('restrictions',
# ((trial space restrictions), (test space restrictions))
[(('+', ), ('+', )),
(('+', ), ('-', )),
(('-', ), ('+', )),
(('-', '+'), ('+', '+')),
(('-', '+'), ('-', '+')),
(('-', '+'), ('+', '-')),
(('-', '+'), ('-', '-')),
(('+', '+'), ('+', '+')),
(('+', '+'), ('-', '+')),
(('+', '+'), ('+', '-')),
(('+', '+'), ('-', '-')),
(('-', '-'), ('+', '+')),
(('-', '-'), ('-', '+')),
(('-', '-'), ('+', '-')),
(('-', '-'), ('-', '-')),
(('+', '-'), ('+', '+')),
(('+', '-'), ('-', '+')),
(('+', '-'), ('+', '-')),
(('+', '-'), ('-', '-')),
(('+', '+', '-', '-'), ('+', '-', '+', '-'))])
def test_bilinear_interior_facet_integral(dg_trial_test, restrictions):
u, v = dg_trial_test
trial_r, test_r = restrictions
idx = {'+': 0, '-': 1}
exact = np.zeros((2, 2), dtype=float)
form = 0
for u_r, v_r in zip(trial_r, test_r):
form = form + u(u_r)*v(v_r)*dS
exact[idx[v_r], idx[u_r]] += sqrt(2)
interior_facet = assemble(form).M.values
assert np.allclose(interior_facet - exact, 0.0)
@pytest.mark.parametrize('space', ["RT", "BDM"])
def test_contravariant_piola_facet_integral(space):
m = UnitSquareMesh(1, 1)
V = FunctionSpace(m, space, 1)
u = project(Expression(("0.0", "1.0")), V)
assert abs(assemble(dot(u('+'), u('+'))*dS) - sqrt(2)) < 1.0e-13
assert abs(assemble(dot(u('-'), u('-'))*dS) - sqrt(2)) < 1.0e-13
assert abs(assemble(dot(u('+'), u('-'))*dS) - sqrt(2)) < 1.0e-13
@pytest.mark.parametrize('space', ["N1curl", "N2curl"])
def test_covariant_piola_facet_integral(space):
m = UnitSquareMesh(1, 1)
V = FunctionSpace(m, space, 1)
u = project(Expression(("0.0", "1.0")), V)
assert abs(assemble(dot(u('+'), u('+'))*dS) - sqrt(2)) < 1.0e-13
assert abs(assemble(dot(u('-'), u('-'))*dS) - sqrt(2)) < 1.0e-13
assert abs(assemble(dot(u('+'), u('-'))*dS) - sqrt(2)) < 1.0e-13
def test_internal_integral_unit_tri():
t = UnitTriangleMesh()
V = FunctionSpace(t, 'CG', 1)
u = Function(V)
u.interpolate(Expression("x[0]"))
assert abs(assemble(u('+') * dS)) < 1.0e-14
def test_internal_integral_unit_tet():
t = UnitTetrahedronMesh()
V = FunctionSpace(t, 'CG', 1)
u = Function(V)
u.interpolate(Expression("x[0]"))
assert abs(assemble(u('+') * dS)) < 1.0e-14
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/requirements.txt
six
sympy
git+https://bitbucket.org/fenics-project/instant.git#egg=instant
git+https://bitbucket.org/mapdes/ufl.git#egg=ufl
git+https://bitbucket.org/mapdes/fiat.git#egg=fiat
git+https://bitbucket.org/mapdes/ffc.git#egg=ffc
git+https://github.com/OP2/PyOP2
<file_sep>/firedrake/__init__.py
# Ensure petsc is initialised by us before anything else gets in there.
import petsc
del petsc
# UFL Exprs come with a custom __del__ method, but we hold references
# to them /everywhere/, some of which are circular (the Mesh object
# holds a ufl.Domain that references the Mesh). The Python2 GC
# explicitly DOES NOT collect such reference cycles (even though it
# can deal with normal cycles). Quoth the documentation:
#
# Objects that have __del__() methods and are part of a reference
# cycle cause the entire reference cycle to be uncollectable,
# including objects not necessarily in the cycle but reachable
# only from it.
#
# To get around this, since the default __del__ on Expr is just
# "pass", we just remove the method from the definition of Expr.
import ufl
try:
del ufl.core.expr.Expr.__del__
except AttributeError:
pass
del ufl
from ufl import *
from pyop2.logger import set_log_level, info_red, info_green, info_blue, log # noqa
from pyop2.logger import debug, info, warning, error, critical # noqa
from pyop2.logger import DEBUG, INFO, WARNING, ERROR, CRITICAL # noqa
from pyop2 import op2 # noqa
from assemble import *
from bcs import *
from constant import *
from expression import *
from function import *
from functionspace import *
from io import *
from linear_solver import *
from mesh import *
from norms import *
from nullspace import *
from parameters import *
from parloops import *
from projection import *
from solving import *
from ufl_expr import *
from utility_meshes import *
from variational_solver import *
from vector import *
from version import __version__ as ver, __version_info__, check # noqa
# Set default log level
set_log_level(INFO)
check()
del check
from ._version import get_versions
__version__ = get_versions(default={"version": ver, "full": ""})['version']
del get_versions
<file_sep>/tests/extrusion/test_extrusion_1_assembly.py
"""Tests for successful assembly of forms on extruded meshes"""
import pytest
from firedrake import *
from tests.common import *
CG = [("CG", 1), ("CG", 2)]
DG = [("DG", 0), ("DG", 1)]
hdiv = [("RT", 1), ("RT", 2), ("RT", 3), ("BDM", 1), ("BDM", 2), ("BDFM", 2)]
hcurl = [("N1curl", 1), ("N1curl", 2), ("N2curl", 1), ("N2curl", 2)]
@pytest.mark.parametrize(('hfamily', 'hdegree', 'vfamily', 'vdegree'),
[(f, d, vf, vd) for (vf, vd) in CG + DG for (f, d) in CG + DG])
def test_scalar_assembly(hfamily, hdegree, vfamily, vdegree):
mesh = extmesh(4, 4, 2)
fspace = FunctionSpace(mesh, hfamily, hdegree, vfamily=vfamily, vdegree=vdegree)
u = TrialFunction(fspace)
v = TestFunction(fspace)
assemble(u*v*dx).M._force_evaluation()
assemble(dot(grad(u), grad(v))*dx).M._force_evaluation()
# three valid combinations for hdiv: 1) hdiv x DG, 2) hcurl x DG, 3) DG x CG
@pytest.mark.parametrize(('hfamily', 'hdegree', 'vfamily', 'vdegree'),
[(f, d, vf, vd) for (vf, vd) in DG for (f, d) in hdiv]
+ [(f, d, vf, vd) for (vf, vd) in DG for (f, d) in hcurl]
+ [(f, d, vf, vd) for (vf, vd) in CG for (f, d) in DG])
def test_hdiv_assembly(hfamily, hdegree, vfamily, vdegree):
mesh = extmesh(4, 4, 2)
horiz_elt = FiniteElement(hfamily, "triangle", hdegree)
vert_elt = FiniteElement(vfamily, "interval", vdegree)
product_elt = HDiv(OuterProductElement(horiz_elt, vert_elt))
fspace = FunctionSpace(mesh, product_elt)
u = TrialFunction(fspace)
v = TestFunction(fspace)
assemble(dot(u, v)*dx).M._force_evaluation()
assemble(inner(grad(u), grad(v))*dx).M._force_evaluation()
# three valid combinations for hcurl: 1) hcurl x CG, 1) hdiv x CG, 3) CG x DG
@pytest.mark.parametrize(('hfamily', 'hdegree', 'vfamily', 'vdegree'),
[(f, d, vf, vd) for (vf, vd) in CG for (f, d) in hcurl]
+ [(f, d, vf, vd) for (vf, vd) in CG for (f, d) in hdiv]
+ [(f, d, vf, vd) for (vf, vd) in DG for (f, d) in CG])
def test_hcurl_assembly(hfamily, hdegree, vfamily, vdegree):
mesh = extmesh(4, 4, 2)
horiz_elt = FiniteElement(hfamily, "triangle", hdegree)
vert_elt = FiniteElement(vfamily, "interval", vdegree)
product_elt = HCurl(OuterProductElement(horiz_elt, vert_elt))
fspace = FunctionSpace(mesh, product_elt)
u = TrialFunction(fspace)
v = TestFunction(fspace)
assemble(dot(u, v)*dx).M._force_evaluation()
assemble(inner(grad(u), grad(v))*dx).M._force_evaluation()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/constant.py
import numpy as np
import ufl
from pyop2 import op2
from pyop2.exceptions import DataTypeError, DataValueError
import utils
__all__ = ['Constant']
class Constant(ufl.Coefficient):
"""A "constant" coefficient
A :class:`Constant` takes one value over the whole
:class:`~.Mesh`. The advantage of using a :class:`Constant` in a
form rather than a literal value is that the constant will be
passed as an argument to the generated kernel which avoids the
need to recompile the kernel if the form is assembled for a
different value of the constant.
:arg value: the value of the constant. May either be a scalar, an
iterable of values (for a vector-valued constant), or an iterable
of iterables (or numpy array with 2-dimensional shape) for a
tensor-valued constant.
:arg domain: an optional UFL :class:`~ufl.domain.Domain` on which the constant is defined.
"""
def __init__(self, value, domain=None):
# Init also called in mesh constructor, but constant can be built without mesh
utils._init()
data = np.array(value, dtype=np.float64)
shape = data.shape
rank = len(shape)
if rank == 0:
e = ufl.FiniteElement("Real", domain, 0)
self.dat = op2.Global(1, data)
elif rank == 1:
e = ufl.VectorElement("Real", domain, 0, shape[0])
self.dat = op2.Global(shape, data)
elif rank == 2:
e = ufl.TensorElement("Real", domain, 0, shape=shape)
self.dat = op2.Global(shape, data)
else:
raise RuntimeError("Don't know how to make Constant from data with rank %d" % rank)
super(Constant, self).__init__(e)
self._ufl_element = self.element()
self._repr = 'Constant(%r)' % self._ufl_element
def ufl_element(self):
"""Return the UFL element on which this Constant is built."""
return self._ufl_element
def function_space(self):
"""Return a null function space."""
return None
def cell_node_map(self, bcs=None):
"""Return a null cell to node map."""
if bcs is not None:
raise RuntimeError("Can't apply boundary conditions to a Constant")
return None
def interior_facet_node_map(self, bcs=None):
"""Return a null interior facet to node map."""
if bcs is not None:
raise RuntimeError("Can't apply boundary conditions to a Constant")
return None
def exterior_facet_node_map(self, bcs=None):
"""Return a null exterior facet to node map."""
if bcs is not None:
raise RuntimeError("Can't apply boundary conditions to a Constant")
return None
def assign(self, value):
"""Set the value of this constant.
:arg value: A value of the appropriate shape"""
try:
self.dat.data = value
return self
except (DataTypeError, DataValueError) as e:
raise ValueError(e)
def __iadd__(self, o):
raise NotImplementedError("Augmented assignment to Constant not implemented")
def __isub__(self, o):
raise NotImplementedError("Augmented assignment to Constant not implemented")
def __imul__(self, o):
raise NotImplementedError("Augmented assignment to Constant not implemented")
def __idiv__(self, o):
raise NotImplementedError("Augmented assignment to Constant not implemented")
<file_sep>/demos/extruded_shallow_water/test_extrusion_lsw.py
# FIXME: document properly
"""Demo of Linear Shallow Water, with Strang timestepping and silly BCs, but
a sin(x)*sin(y) solution that doesn't care about the silly BCs"""
from firedrake import *
power = 5
# Create mesh and define function space
m = UnitSquareMesh(2 ** power, 2 ** power)
layers = 5
# Populate the coordinates of the extruded mesh by providing the
# coordinates as a field.
mesh = ExtrudedMesh(m, layers, layer_height=0.25)
horiz = FiniteElement("BDM", "triangle", 1)
vert = FiniteElement("DG", "interval", 0)
prod = HDiv(OuterProductElement(horiz, vert))
W = FunctionSpace(mesh, prod)
X = FunctionSpace(mesh, "DG", 0, vfamily="DG", vdegree=0)
Xplot = FunctionSpace(mesh, "CG", 1, vfamily="Lagrange", vdegree=1)
# Define starting field
u_0 = Function(W)
u_h = Function(W)
u_1 = Function(W)
p_0 = Function(X)
p_1 = Function(X)
p_plot = Function(Xplot)
p_0.interpolate(Expression("sin(4*pi*x[0])*sin(2*pi*x[1])"))
T = 0.5
t = 0
dt = 0.0025
file = File("lsw3d.pvd")
p_trial = TrialFunction(Xplot)
p_test = TestFunction(Xplot)
solve(p_trial * p_test * dx == p_0 * p_test * dx, p_plot)
file << p_plot, t
E_0 = assemble(0.5 * p_0 * p_0 * dx + 0.5 * dot(u_0, u_0) * dx)
while t < T:
u = TrialFunction(W)
w = TestFunction(W)
a_1 = dot(w, u) * dx
L_1 = dot(w, u_0) * dx + 0.5 * dt * div(w) * p_0 * dx
solve(a_1 == L_1, u_h)
p = TrialFunction(X)
phi = TestFunction(X)
a_2 = phi * p * dx
L_2 = phi * p_0 * dx - dt * phi * div(u_h) * dx
solve(a_2 == L_2, p_1)
u = TrialFunction(W)
w = TestFunction(W)
a_3 = dot(w, u) * dx
L_3 = dot(w, u_h) * dx + 0.5 * dt * div(w) * p_1 * dx
solve(a_3 == L_3, u_1)
u_0.assign(u_1)
p_0.assign(p_1)
t += dt
# project into P1 x P1 for plotting
p_trial = TrialFunction(Xplot)
p_test = TestFunction(Xplot)
solve(p_trial * p_test * dx == p_0 * p_test * dx, p_plot)
file << p_plot, t
print t
E_1 = assemble(0.5 * p_0 * p_0 * dx + 0.5 * dot(u_0, u_0) * dx)
print 'Initial energy', E_0
print 'Final energy', E_1
<file_sep>/tests/regression/test_quadrature.py
import pytest
from firedrake import *
from tests.common import *
def test_hand_specified_quadrature(mesh):
V = FunctionSpace(mesh, 'CG', 2)
v = TestFunction(V)
a = v*dx
norm_q0 = norm(assemble(a, form_compiler_parameters={'quadrature_degree': 0}))
norm_q2 = norm(assemble(a, form_compiler_parameters={'quadrature_degree': 2}))
assert norm_q0 != norm_q2
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_helmholtz.py
"""This demo program solves Helmholtz's equation
- div grad u(x, y) + u(x,y) = f(x, y)
on the unit square with source f given by
f(x, y) = (1.0 + 8.0*pi**2)*cos(x[0]*2*pi)*cos(x[1]*2*pi)
and the analytical solution
u(x, y) = cos(x[0]*2*pi)*cos(x[1]*2*pi)
"""
from os.path import abspath, dirname, join
import numpy as np
import pytest
from firedrake import *
cwd = abspath(dirname(__file__))
def helmholtz(x, quadrilateral=False, degree=2, mesh=None):
# Create mesh and define function space
if mesh is None:
mesh = UnitSquareMesh(2 ** x, 2 ** x, quadrilateral=quadrilateral)
V = FunctionSpace(mesh, "CG", degree)
# Define variational problem
lmbda = 1
u = TrialFunction(V)
v = TestFunction(V)
f = Function(V)
f.interpolate(Expression("(1+8*pi*pi)*cos(x[0]*pi*2)*cos(x[1]*pi*2)"))
a = (dot(grad(v), grad(u)) + lmbda * v * u) * dx
L = f * v * dx
# Compute solution
assemble(a)
assemble(L)
x = Function(V)
solve(a == L, x, solver_parameters={'ksp_type': 'cg'})
# Analytical solution
f.interpolate(Expression("cos(x[0]*pi*2)*cos(x[1]*pi*2)"))
return sqrt(assemble(dot(x - f, x - f) * dx)), x, f
def run_firedrake_helmholtz():
diff = np.array([helmholtz(i)[0] for i in range(3, 6)])
print "l2 error norms:", diff
conv = np.log2(diff[:-1] / diff[1:])
print "convergence order:", conv
assert (np.array(conv) > 2.8).all()
def test_firedrake_helmholtz_serial():
run_firedrake_helmholtz()
@pytest.mark.parallel
def test_firedrake_helmholtz_parallel():
run_firedrake_helmholtz()
@pytest.mark.parametrize(('testcase', 'convrate'),
[((1, (4, 6)), 1.9),
((2, (3, 6)), 2.9),
((3, (2, 4)), 3.9),
((4, (2, 4)), 4.7)])
def test_firedrake_helmholtz_scalar_convergence_on_quadrilaterals(testcase, convrate):
degree, (start, end) = testcase
l2err = np.zeros(end - start)
for ii in [i + start for i in range(len(l2err))]:
l2err[ii - start] = helmholtz(ii, quadrilateral=True, degree=degree)[0]
assert (np.array([np.log2(l2err[i]/l2err[i+1]) for i in range(len(l2err)-1)]) > convrate).all()
def run_firedrake_helmholtz_on_quadrilateral_mesh_from_file():
meshfile = join(cwd, "unitsquare_unstructured_quadrilaterals.msh")
assert helmholtz(None, mesh=Mesh(meshfile))[0] <= 0.01
def test_firedrake_helmholtz_on_quadrilateral_mesh_from_file_serial():
run_firedrake_helmholtz_on_quadrilateral_mesh_from_file()
@pytest.mark.parallel
def test_firedrake_helmholtz_on_quadrilateral_mesh_from_file_parallel():
run_firedrake_helmholtz_on_quadrilateral_mesh_from_file()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_4_helmholtz_scalar.py
"""Tests for scalar Helmholtz convergence on extruded meshes"""
import numpy as np
import pytest
from firedrake import *
from tests.common import *
@pytest.mark.parametrize(('testcase', 'convrate'),
[(("CG", 1, (4, 6)), 1.9),
(("CG", 2, (3, 5)), 2.9),
(("CG", 3, (1, 3)), 3.9)])
def test_scalar_convergence(testcase, convrate):
family, degree, (start, end) = testcase
l2err = np.zeros(end - start)
for ii in [i + start for i in range(len(l2err))]:
mesh = extmesh(2**ii, 2**ii, 2**ii)
fspace = FunctionSpace(mesh, family, degree, vfamily=family, vdegree=degree)
u = TrialFunction(fspace)
v = TestFunction(fspace)
f = Function(fspace)
f.interpolate(Expression("(1+12*pi*pi)*cos(2*pi*x[0])*cos(2*pi*x[1])*cos(2*pi*x[2])"))
out = Function(fspace)
solve(dot(grad(u), grad(v))*dx + u*v*dx == f*v*dx, out)
exact = Function(fspace)
exact.interpolate(Expression("cos(2*pi*x[0])*cos(2*pi*x[1])*cos(2*pi*x[2])"))
l2err[ii - start] = sqrt(assemble((out-exact)*(out-exact)*dx))
assert (np.array([np.log2(l2err[i]/l2err[i+1]) for i in range(len(l2err)-1)]) > convrate).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/docs/source/variational-problems.rst
.. only:: html
.. contents::
Defining variational problems
=============================
Firedrake uses a high-level language, `UFL`_, to describe variational
problems. To do this, we need a number of pieces. We need a
representation of the domain we're solving the :abbr:`PDE (partial
differential equation)` on: Firedrake uses a
:py:class:`~.Mesh` for this. On top of this mesh,
we build :py:class:`~.FunctionSpace`\s which
define the space in which the solutions to our equation live. Finally
we define :py:class:`~.Function`\s in those
function spaces to actually hold the solutions.
Constructing meshes
-------------------
Firedrake can read meshes in `Gmsh`_, `triangle`_, `CGNS`_, and
`Exodus`_ formats. To build a mesh one uses the :py:class:`~.Mesh`
constructor, passing the name of the file as an argument, which see
for more details. The mesh type is determined by the file extension,
for example if the provided filename is ``coastline.msh`` the mesh is
assumed to be in Gmsh format, in which case you can construct a mesh
object like so:
.. code-block:: python
coastline = Mesh("coastline.msh")
This works in both serial and parallel, Firedrake takes care of
decomposing the mesh among processors transparently.
Reordering meshes for better performance
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Most mesh generators produce badly numbered meshes (with bad data
locality) which can reduce the performance of assembling and solving
finite element problems. By default then, Firedrake reorders input
meshes to improve data locality by performing reverse Cuthill-McKee
reordering on the adjacency matrix of the input mesh. If you know
your mesh has a good numbering (perhaps your mesh generator uses space
filling curves to number entities) then you can switch off this
reordering by passing :py:data:`reorder=False` to the appropriate
:py:class:`~.Mesh` constructor. You can control Firedrake's default
behaviour in reordering meshes with the :py:data:`"reorder_meshes"`
parameter. For example, to turn off mesh reordering globally:
.. code-block:: python
from firedrake import *
parameters["reorder_meshes"] = False
The parameter passed in to the mesh constructor overrides this default
value.
.. note::
Firedrake numbers degrees of freedom in a function space by
visiting each cell in order and performing a depth first numbering
of all degrees of freedom on that cell. Hence, if your mesh has a
good numbering, the degrees of freedom will too.
.. _utility_mesh_functions:
Utility mesh functions
~~~~~~~~~~~~~~~~~~~~~~
As well as offering the ability to read mesh information from a file,
Firedrake also provides a number of built in mesh types for a number
of standard shapes. 1-dimensional intervals may be constructed with
:func:`~.IntervalMesh`; 2-dimensional rectangles with
:func:`~.RectangleMesh`; and 3-dimensional boxes with
:func:`~.BoxMesh`. There are also more specific constructors (for
example to build unit square meshes). See
:mod:`~firedrake.utility_meshes` for full details.
Immersed manifolds
~~~~~~~~~~~~~~~~~~
In addition to the simple meshes described above, Firedrake also has
support for solving problems on orientable `immersed manifolds
<submanifold_>`_. That is, meshes in which the entities are
*immersed* in a higher dimensional space. For example, the surface of
a sphere in 3D.
If your mesh is such an immersed manifold, you need to tell Firedrake
that the geometric dimension of the coordinate field (defining where
the points in mesh are) is not the same as the topological dimension
of the mesh entities. This is done by passing an optional second
argument to the mesh constructor which specifies the geometric
dimension. For example, for the surface of a sphere embedded in 3D we
use:
.. code-block:: python
sphere_mesh = Mesh('sphere_mesh.node', 3)
Firedrake provides utility meshes for the surfaces of spheres immersed
in 3D that are approximated using an `icosahedral mesh`_. You can
either build a mesh of the unit sphere with
:py:func:`~.UnitIcosahedralSphereMesh`, or a mesh of a
sphere with specified radius using
:py:func:`~.IcosahedralSphereMesh`. The meshes are
constructed by recursively refining a `regular icosahedron
<icosahedron_>`_, you can specify the refinement level by passing a
non-zero ``refinement_level`` to the constructor. For example, to
build a sphere mesh that approximates the surface of the Earth (with a
radius of 6371 km) that has subdivided the original icosahedron 7
times we would write:
.. code-block:: python
earth = IcosahedralSphereMesh(radius=6371, refinement_level=7)
Ensuring consistent cell orientations
+++++++++++++++++++++++++++++++++++++
Variational forms that include particular function spaces (those
requiring a *contravariant Piola transform*), require information
about the orientation of the cells. For normal meshes, this can be
deduced automatically. However, when using immersed meshes, Firedrake
needs extra information to calculate the orientation of each cell
relative to some global orientation. This
is used by Firedrake to ensure that the cell normal on,
say, the surface of a sphere, uniformly points outwards. To do this,
after constructing an immersed mesh, we must initialise the cell
orientation information. This is carried out with the function
:py:meth:`~.Mesh.init_cell_orientations`, which
takes an :py:class:`~.Expression` used to produce
the reference normal direction. For example, on the sphere mesh of
the earth defined above we can initialise the cell orientations
relative to vector pointing out from the origin:
.. code-block:: python
earth.init_cell_orientations(Expression(('x[0]', 'x[1]', 'x[2]')))
However, a more complicated expression would be needed to initialise
the cell orientations on a toroidal mesh.
Semi-structured extruded meshes
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Firedrake has special support for solving PDEs on high-aspect ratio
domains, such as in the ocean or atmosphere, where the numerics
dictate that the "short" dimension should be structured. These are
termed *extruded meshes* and have a :doc:`separate section
<extruded-meshes>` in the manual.
Building function spaces
------------------------
Now that we have a mesh of our domain, we need to build the function
spaces the solution to our :abbr:`PDE (partial differential equation)`
will live in, along with the spaces for the trial and test functions.
To do so, we normally use the :py:class:`~.FunctionSpace` constructor.
This is the only way to obtain a function space for a scalar variable,
such as pressure, which has a single value at each point in the
domain.
To construct a function space, you must specify its family and
polynomial degree. To build a scalar-valued function space of
continuous piecewise-cubic polynomials, we write:
.. code-block:: python
V = FunctionSpace(mesh, "Lagrange", 3)
There are two main routes to obtaining a function space for a
vector-valued variable such as velocity. Firstly, you can pass the
:py:class:`~.FunctionSpace` constructor a natively *vector-valued*
family such as ``"Raviart-Thomas"``. Secondly, you may use the
:py:class:`~.VectorFunctionSpace` constructor with a *scalar-valued*
family, which gives a vector-valued space where each component is
identical to the appropriate scalar-valued :py:class:`~.FunctionSpace`.
To build a vector-valued function space using the lowest-order
``Raviart-Thomas`` elements, we write
.. code-block:: python
V = FunctionSpace(mesh, "Raviart-Thomas", 1)
To build a vector-valued function space for which each component
is a discontinuous piecewise-quadratic polynomial, we write
.. code-block:: python
V = VectorFunctionSpace(mesh, "Discontinuous Lagrange", 2)
Firedrake supports the use of all function spaces generated by
`FFC`_ and `FIAT`_.
Advanced usage of ``VectorFunctionSpace``
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
By default, the number of components of a
:py:class:`~.VectorFunctionSpace` is the geometric dimension of the
mesh (e.g. 3, if the mesh is 3D). However, sometimes we might want
the number of components in the vector to differ from the geometric
dimension of the mesh. We can do this by passing a value for the
``dim`` argument to the :py:class:`~.VectorFunctionSpace` constructor.
For example, if we wanted a vector-valued function space on the surface
of a unit sphere mesh with only 2 components, we might write:
.. code-block:: python
mesh = UnitIcosahedralSphereMesh(refinement_level=3)
V = VectorFunctionSpace(mesh, "Lagrange", 1, dim=2)
Mixed function spaces
~~~~~~~~~~~~~~~~~~~~~
Many :abbr:`PDE (partial differential equation)`\s are posed in terms
of multiple, coupled, variables. The variational problem for such a
PDE uses a so-called *mixed* function space. In Firedrake, this is
represented by a :py:class:`~.MixedFunctionSpace`. We can either
build such a space by invoking the constructor directly, or, more
readably, by taking existing function spaces and multiplying them
together using the ``*`` operator. For example:
.. code-block:: python
V = FunctionSpace(mesh, 'RT', 1)
Q = FunctionSpace(mesh, 'DG', 0)
W = V*Q
is equivalent to:
.. code-block:: python
V = FunctionSpace(mesh, 'RT', 1)
Q = FunctionSpace(mesh, 'DG', 0)
W = MixedFunctionSpace([V, Q])
Function spaces on extruded meshes
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
On :doc:`extruded meshes <extruded-meshes>`, we build function spaces
by taking a tensor product of the base ("horizontal") space and the
extruded ("vertical") space. Firedrake allows us to separately choose
the horizontal and vertical spaces when building a function space on
an extruded mesh. We refer the reader to the :doc:`manual section on
extrusion <extruded-meshes>` for details.
Expressing a variational problem
--------------------------------
Firedrake uses the UFL language to express variational problems. For
complete documentation, we refer the reader to `the UFL package
documentation <UFL_package_>`_ and the description of the language in
`TOMS <UFL_>`_. We present a brief overview of the syntax here,
for a more didactic introduction, we refer the reader to the
:ref:`Firedrake tutorial examples <firedrake_tutorials>`.
Building test and trial spaces
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Now that we have function spaces that our solution will live in, the
next step is to actually write down the variational form of the
problem we wish to solve. To do this, we will need a test function in
an appropriate space along with a function to hold the solution and
perhaps a trial function. Test functions are obtained via a call to
:py:class:`~firedrake.ufl_expr.TestFunction`, trial functions via
:py:class:`~firedrake.ufl_expr.TrialFunction` and functions with
:py:class:`~.Function`. The former two are purely
symbolic objects, the latter contains storage for the coefficients of
the basis functions in the function space. We use them as follows:
.. code-block:: python
u = TrialFunction(V)
v = TestFunction(V)
f = Function(V)
.. note::
A newly allocated :py:class:`~.Function` has
coefficients which are all zero.
If ``V`` above were a
:py:class:`~.MixedFunctionSpace`, the test and
trial functions we obtain are for the combined mixed space. Often, we
would like to have test and trial functions for the subspaces of the
mixed space. We can do this by asking for
:py:class:`~firedrake.ufl_expr.TrialFunctions` and
:py:class:`~firedrake.ufl_expr.TestFunctions`, which return an ordered
tuple of test and trial functions for the underlying spaces. For
example, if we write:
.. code-block:: python
V = FunctionSpace(mesh, 'RT', 1)
Q = FunctionSpace(mesh, 'DG', 0)
W = V * Q
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
then ``u`` and ``v`` will be, respectively, trial and test
functions for ``V``, while ``p`` and ``q`` will be trial and test
functions for ``Q``.
.. note::
If we intend to build a variational problem on a mixed space, we
cannot build the individual test and trial functions on the
function spaces that were used to construct the mixed space
directly. The functions that we build must "know" that they come
from a mixed space or else Firedrake will not be able to assemble
the correct system of equations.
A first variational form
~~~~~~~~~~~~~~~~~~~~~~~~
With our test and trial functions defined, we can write down our first
variational form. Let us consider solving the identity equation:
.. math::
u = f \quad \mathrm{on} \, \Omega
where :math:`\Omega` is the unit square, using piecewise linear
polynomials for our solution. We start with a mesh and build a
function space on it:
.. code-block:: python
mesh = UnitSquareMesh(10, 10)
V = FunctionSpace(mesh, "CG", 1)
now we need a test function, and since ``u`` is unknown, a trial
function:
.. code-block:: python
u = TrialFunction(V)
v = TestFunction(V)
finally we need a function to hold the right hand side :math:`f` which
we will populate with the x component of the coordinate field.
.. code-block:: python
f = Function(V)
f.interpolate(Expression('x[0]'))
For details on how :py:class:`~.Expression`\s and
:py:meth:`~.Function.interpolate` work, see the
:doc:`appropriate section in the manual <expressions>`. The
variational problem is to find :math:`u \in V` such that
.. math::
\int_\Omega \! u v \, \mathrm{d}x = \int_\Omega \! f v \, \mathrm{d}x \quad
\forall v \in V
we define the variational problem in UFL with:
.. code-block:: python
a = u*v*dx
L = f*v*dx
Where the ``dx`` indicates that the integration should be carried out
over the cells of the mesh. UFL can also express integrals over the
boundary of the domain, using ``ds``, and the interior facets of the
domain, using ``dS``.
How to solve such variational problems is the subject of the
:doc:`next section <solving-interface>`, but for completeness we show
how to do it here. First we define a function to hold the solution
.. code-block:: python
s = Function(V)
and call :py:func:`~.solve` to solve the variational
problem:
.. code-block:: python
solve(a == L, s)
Forms with constant coefficients
--------------------------------
Many PDEs will contain values that are constant over the whole mesh,
but may vary in time. For example, a time-varying diffusivity, or a
time-dependent forcing function. Although you can create a new form
for each new value of this constant, this will not be efficient, since
Firedrake must generate new code each time the value changes. A
better option is to use a :py:class:`~.Constant` coefficient. This
object behaves exactly like a :py:class:`~.Function`, except that it
has a single value over the whole mesh. One may assign a new value to
the :py:class:`~.Constant` using the :py:meth:`~.Constant.assign`
method. As an example, let us consider a form which contains a time
varying constant which we wish to assemble in a time loop. We can use
a :py:class:`~.Constant` to do this:
.. code-block:: python
...
t = 0
dt = 0.1
from math import exp
c = Constant(exp(-t))
# Exponentially decaying RHS
L = f*v*c*dx
while t < tend:
solve(a == L, ...)
t += dt
c.assign(exp(-t))
.. warning::
Although UFL supports computing the derivative of a form with
respect to a :py:class:`~.Constant`, the resulting form will have
an unknown in the reals, which is currently unsupported by
Firedrake.
Incorporating boundary conditions
---------------------------------
Boundary conditions enter the variational problem in one of two ways.
`Natural` (often termed `Neumann` or `weak`) boundary conditions,
which prescribe values of the derivative of the solution, are
incorporated into the variational form. `Essential` (often termed
`Dirichlet` or `strong`) boundary conditions, which prescribe values
of the solution, become prescriptions on the function space. In
Firedrake, the former are naturally expressed as part of the
formulation of the variational problem, the latter are represented as
:py:class:`~.DirichletBC` objects and are applied when
solving the variational problem. Construction of such a strong
boundary condition requires a function space (to impose the boundary
condition in), a value and a subdomain to apply the boundary condition
over:
.. code-block:: python
bc = DirichletBC(V, value, subdomain_id)
The ``subdomain_id`` is an integer indicating which section of the
mesh the boundary condition should be applied to. The subdomain ids
for the various :ref:`utility meshes <utility_mesh_functions>` are
described in their respective constructor documentation. For
externally generated meshes, Firedrake just uses whichever ids the
mesh generator provided. The ``value`` may be either a scalar, or
more generally an :py:class:`~.Expression`, :py:class:`~.Function` or
:py:class:`~.Constant` of the appropriate shape. You may also supply
an iterable of literal constants, which will be converted to an
:py:class:`~.Expression`. Hence the following two are equivalent:
.. code-block:: python
bc1 = DirichletBC(V, Expression(('1.0', '2.0')), 1)
bc2 = DirichletBC(V, (1.0, 2.0), 1)
Strong boundary conditions are applied in the solve by passing a list
of boundary condition objects:
.. code-block:: python
solve(a == L, bcs=[bc])
See the :doc:`next section <solving-interface>` for a more complete
description of the interface Firedrake provides to solve PDEs. The
details of how Firedrake applies strong boundary conditions are
slightly involved and therefore have :doc:`their own section
<boundary_conditions>` in the manual.
Boundary conditions in discontinuous spaces
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
The default method Firedrake uses to determine where to apply strong
boundary conditions is :py:data:`"topological"`, meaning that nodes
topologically associated with a boundary facet will be included. In
discontinuous spaces, however, the nodes to be included do not all
live on boundary facets, in this case, you should use the
:py:data:`"geometric"` method for determining boundary condition
nodes. In this case, nodes associated with basis functions that do
not vanish on the boundary are included. This method can be used to
impose strong boundary conditions on discontinuous galerkin spaces, or
no-slip conditions on HDiv spaces. To select the method used for
determining boundary condition nodes, use the :py:data:`method`
argument to the :py:class:`DirichletBC` constructor. For example, to
select geometric boundary node determination we would write:
.. code-block:: python
V = FunctionSpace(mesh, 'DG', 2)
bc = DirichletBC(V, 1.0, subdomain_id, method="geometric")
...
Time dependent boundary conditions
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Imposition of time-dependent boundary conditions can by carried out by
modifying the value in the appropriate :py:class:`~.DirichletBC`
object. Note that if you use a literal value to initialise the
boundary condition object within the timestepping loop, this will
necessitate a recompilation of code every time the boundary condition
changes. For this reason we either recommend using a
:py:class:`~.Constant` if the boundary condition is spatially uniform,
or a :py:class:`~.Expression` if it has both space and
time-dependence. For example, a purely time-varying boundary
condition might be implemented as:
.. code-block:: python
c = Constant(sin(t))
bc = DirichletBC(V, c, 1)
while t < T:
solve(F == 0, bcs=[bc])
t += dt
c.assign(sin(t))
If the boundary condition instead has both space and time dependence
we can write:
.. code-block:: python
e = Expression('sin(x[0]*t'), t=t)
bc = DirichletBC(V, e, 1)
while t < T:
solve(F == 0, bcs=[bc])
t += dt
e.t = t
More complicated forms
----------------------
UFL is a fully-fledged language for expressing variational problems,
and hence has operators for all appropriate vector calculus operations
along with special support for discontinuous galerkin methods in the
form of symbolic expressions for facet averages and jumps. For an
introduction to these concepts we refer the user to the `UFL manual
<UFL_package_>`_ as well as the :ref:`Firedrake tutorials
<firedrake_tutorials>` which cover a wider variety of different
problems.
.. _icosahedral mesh: http://en.wikipedia.org/wiki/Geodesic_grid
.. _icosahedron: http://en.wikipedia.org/wiki/Icosahedron
.. _triangle: http://www.cs.cmu.edu/~quake/triangle.html
.. _Gmsh: http://geuz.org/gmsh/
.. _CGNS: http://www.cgns.org/
.. _Exodus: http://sourceforge.net/projects/exodusii/
.. _UFL: http://arxiv.org/abs/1211.4047
.. _UFL_package: http://fenicsproject.org/documentation/ufl/1.2.0/ufl.html
.. _FIAT: https://bitbucket.org/mapdes/fiat
.. _FFC: https://bitbucket.org/mapdes/ffc
.. _submanifold: http://en.wikipedia.org/wiki/Submanifold
<file_sep>/firedrake/io.py
from evtk import hl
from evtk.vtk import _get_byte_order
from evtk.hl import _requiresLargeVTKFileSize
from ufl import Cell, OuterProductCell
import numpy as np
import os
from pyop2.logger import warning, RED
from pyop2.mpi import MPI
import functionspace as fs
import projection
__all__ = ['File']
# Dictionary used to translate the cellname of firedrake
# to the celltype of evtk module.
_cells = {}
_cells[Cell("interval")] = hl.VtkLine
_cells[Cell("interval", 2)] = hl.VtkLine
_cells[Cell("interval", 3)] = hl.VtkLine
_cells[Cell("triangle")] = hl.VtkTriangle
_cells[Cell("triangle", 3)] = hl.VtkTriangle
_cells[Cell("tetrahedron")] = hl.VtkTetra
_cells[OuterProductCell(Cell("triangle"), Cell("interval"))] = hl.VtkWedge
_cells[OuterProductCell(Cell("triangle", 3), Cell("interval"))] = hl.VtkWedge
_cells[Cell("quadrilateral")] = hl.VtkQuad
_cells[OuterProductCell(Cell("interval"), Cell("interval"))] = hl.VtkQuad
_cells[OuterProductCell(Cell("interval", 2), Cell("interval"))] = hl.VtkQuad
_cells[OuterProductCell(Cell("interval", 2), Cell("interval"), gdim=3)] = hl.VtkQuad
_cells[OuterProductCell(Cell("interval", 3), Cell("interval"))] = hl.VtkQuad
_points_per_cell = {}
_points_per_cell[Cell("interval")] = 2
_points_per_cell[Cell("interval", 2)] = 2
_points_per_cell[Cell("interval", 3)] = 2
_points_per_cell[Cell("triangle")] = 3
_points_per_cell[Cell("triangle", 3)] = 3
_points_per_cell[Cell("quadrilateral")] = 4
_points_per_cell[Cell("tetrahedron")] = 4
_points_per_cell[OuterProductCell(Cell("triangle"), Cell("interval"))] = 6
_points_per_cell[OuterProductCell(Cell("triangle", 3), Cell("interval"))] = 6
_points_per_cell[OuterProductCell(Cell("interval"), Cell("interval"))] = 4
_points_per_cell[OuterProductCell(Cell("interval", 2), Cell("interval"))] = 4
_points_per_cell[OuterProductCell(Cell("interval", 2), Cell("interval"), gdim=3)] = 4
_points_per_cell[OuterProductCell(Cell("interval", 3), Cell("interval"))] = 4
class File(object):
"""A pvd file object to which :class:`~.Function`\s can be output.
Parallel output is handled automatically.
File output is achieved using the left shift operator:
.. code-block:: python
a = Function(...)
f = File("foo.pvd")
f << a
.. note::
A single :class:`File` object only supports output in a single
function space. The supported function spaces for output are
CG1 or DG1; any functions which do not live in these spaces
will automatically be projected to one or the other as
appropriate. The selection of which space is used for output
in this :class:`File` depends on both the continuity of the
coordinate field and the continuity of the output function.
The logic for selecting the output space is as follows:
* If the both the coordinate field and the output function are
in :math:`H^1`, the output will be in CG1.
* Otherwise, both the coordinate field and the output function
will be in DG1.
"""
def __init__(self, filename):
# Ensure output directory exists
outdir = os.path.dirname(os.path.abspath(filename))
if MPI.comm.rank == 0:
if not os.path.exists(outdir):
os.makedirs(outdir)
MPI.comm.barrier()
# Parallel
if MPI.comm.size > 1:
new_file = os.path.splitext(os.path.abspath(filename))[0]
# If the rank of process is 0, then create PVD file that can create
# PVTU file and VTU file.
if MPI.comm.rank == 0:
self._file = _PVDFile(new_file)
# Else, create VTU file.
elif os.path.splitext(filename)[1] == ".pvd":
self._file = _VTUFile(new_file)
else:
raise ValueError("On parallel writing, the filename written\
must be vtu file.")
else:
new_file = os.path.splitext(os.path.abspath(filename))[0]
if os.path.splitext(filename)[1] == ".vtu":
self._file = _VTUFile(new_file)
elif os.path.splitext(filename)[1] == ".pvd":
self._file = _PVDFile(new_file)
else:
raise ValueError("File name is wrong. It must be either vtu\
file or pvd file.")
def __lshift__(self, data):
self._file << data
class _VTUFile(object):
"""Class that represents a VTU file."""
def __init__(self, filename, warnings=None):
#_filename : full path to the file without extension.
self._filename = filename
if warnings:
self._warnings = warnings
else:
self._warnings = [None, None]
if MPI.parallel:
self._time_step = -1
# If _generate_time, _time_step would be incremented by
# one everytime.
self._generate_time = False
def __lshift__(self, data):
"""It allows file << function syntax for writing data out to disk.
In the case of parallel, it would also accept (function, timestep)
tuple as an argument. If only function is given, then the timestep
will be automatically generated."""
# If parallel, it needs to keep track of its timestep.
if MPI.parallel:
# if statements to keep the consistency of how to update the
# timestep.
if isinstance(data, tuple):
if self._time_step == -1 or not self._generate_time:
function = data[0]
self._time_step = data[1]
else:
raise TypeError("Expected function, got tuple.")
else:
if self._time_step != -1 and not self._generate_time:
raise TypeError("Expected tuple, got function.")
function = data
self._time_step += 1
self._generate_time = True
else:
function = data
def is_family1(e, family):
import ufl.finiteelement.hdivcurl as hc
if isinstance(e, (hc.HDiv, hc.HCurl)):
return False
if e.family() == 'OuterProductElement':
if e.degree() == (1, 1):
if e._A.family() == family \
and e._B.family() == family:
return True
elif e.family() == family and e.degree() == 1:
return True
return False
def is_cgN(e):
import ufl.finiteelement.hdivcurl as hc
if isinstance(e, (hc.HDiv, hc.HCurl)):
return False
if e.family() == 'OuterProductElement':
if e._A.family() == 'Lagrange' \
and e._B.family() == 'Lagrange':
return True
elif e.family() == 'Lagrange':
return True
return False
mesh = function.function_space().mesh()
e = function.function_space().ufl_element()
if len(e.value_shape()) > 1:
raise RuntimeError("Can't output tensor valued functions")
ce = mesh.coordinates.function_space().ufl_element()
coords_p1 = is_family1(ce, 'Lagrange')
coords_p1dg = is_family1(ce, 'Discontinuous Lagrange')
coords_cgN = is_cgN(ce)
function_p1 = is_family1(e, 'Lagrange')
function_p1dg = is_family1(e, 'Discontinuous Lagrange')
function_cgN = is_cgN(e)
project_coords = False
project_function = False
discontinuous = False
# We either output in P1 or P1dg.
if coords_cgN and function_cgN:
family = 'CG'
project_coords = not coords_p1
project_function = not function_p1
else:
family = 'DG'
project_coords = not coords_p1dg
project_function = not function_p1dg
discontinuous = True
if project_function:
if len(e.value_shape()) == 0:
Vo = fs.FunctionSpace(mesh, family, 1)
elif len(e.value_shape()) == 1:
Vo = fs.VectorFunctionSpace(mesh, family, 1, dim=e.value_shape()[0])
else:
# Never reached
Vo = None
if not self._warnings[0]:
warning(RED % "*** Projecting output function to %s1", family)
self._warnings[0] = True
output = projection.project(function, Vo, name=function.name())
else:
output = function
Vo = output.function_space()
if project_coords:
Vc = fs.VectorFunctionSpace(mesh, family, 1, dim=mesh._coordinate_fs.dim)
if not self._warnings[1]:
warning(RED % "*** Projecting coordinates to %s1", family)
self._warnings[1] = True
coordinates = projection.project(mesh.coordinates, Vc, name=mesh.coordinates.name())
else:
coordinates = mesh.coordinates
Vc = coordinates.function_space()
num_points = Vo.node_count
layers = mesh.layers - 1 if isinstance(e.cell(), OuterProductCell) else 1
num_cells = mesh.num_cells() * layers
if not isinstance(e.cell(), OuterProductCell) and e.cell() != Cell("quadrilateral"):
connectivity = Vc.cell_node_map().values_with_halo.flatten()
else:
# Connectivity of bottom cell in extruded mesh
base = Vc.cell_node_map().values_with_halo
if _cells[mesh.ufl_cell()] == hl.VtkQuad:
# Quad is
#
# 1--3
# | |
# 0--2
#
# needs to be
#
# 3--2
# | |
# 0--1
base = base[:, [0, 2, 3, 1]]
points_per_cell = 4
elif _cells[mesh.ufl_cell()] == hl.VtkWedge:
# Wedge is
#
# 5
# /|\
# / | \
# 1----3
# | 4 |
# | /\ |
# |/ \|
# 0----2
#
# needs to be
#
# 5
# /|\
# / | \
# 3----4
# | 2 |
# | /\ |
# |/ \|
# 0----1
#
base = base[:, [0, 2, 4, 1, 3, 5]]
points_per_cell = 6
# Repeat up the column
connectivity_temp = np.repeat(base, layers, axis=0)
if discontinuous:
scale = points_per_cell
else:
scale = 1
offsets = np.arange(layers) * scale
# Add offsets going up the column
connectivity_temp += np.tile(offsets.reshape(-1, 1), (mesh.num_cells(), 1))
connectivity = connectivity_temp.flatten()
if isinstance(output.function_space(), fs.VectorFunctionSpace):
tmp = output.dat.data_ro_with_halos
vdata = [None]*3
if output.dat.dim[0] == 1:
vdata[0] = tmp.flatten()
else:
for i in range(output.dat.dim[0]):
vdata[i] = tmp[:, i].flatten()
for i in range(output.dat.dim[0], 3):
vdata[i] = np.zeros_like(vdata[0])
data = tuple(vdata)
# only for checking large file size
flat_data = {function.name(): tmp.flatten()}
else:
data = output.dat.data_ro_with_halos.flatten()
flat_data = {function.name(): data}
coordinates = self._fd_to_evtk_coord(coordinates.dat.data_ro_with_halos)
cell_types = np.empty(num_cells, dtype="uint8")
# Assume that all cells are of same shape.
cell_types[:] = _cells[mesh.ufl_cell()].tid
p_c = _points_per_cell[mesh.ufl_cell()]
# This tells which are the last nodes of each cell.
offsets = np.arange(start=p_c, stop=p_c * (num_cells + 1), step=p_c,
dtype='int32')
large_file_flag = _requiresLargeVTKFileSize("VtkUnstructuredGrid",
numPoints=num_points,
numCells=num_cells,
pointData=flat_data,
cellData=None)
new_name = self._filename
# When vtu file makes part of a parallel process, aggregated by a
# pvtu file, the output is : filename_timestep_rank.vtu
if MPI.parallel:
new_name += "_" + str(self._time_step) + "_" + str(MPI.comm.rank)
self._writer = hl.VtkFile(
new_name, hl.VtkUnstructuredGrid, large_file_flag)
self._writer.openGrid()
self._writer.openPiece(ncells=num_cells, npoints=num_points)
# openElement allows the stuff in side of the tag <arg></arg>
# to be editted.
self._writer.openElement("Points")
# addData adds the DataArray in the tag <arg1>
self._writer.addData("Points", coordinates)
self._writer.closeElement("Points")
self._writer.openElement("Cells")
self._writer.addData("connectivity", connectivity)
self._writer.addData("offsets", offsets)
self._writer.addData("types", cell_types)
self._writer.closeElement("Cells")
self._writer.openData("Point", scalars=function.name())
self._writer.addData(function.name(), data)
self._writer.closeData("Point")
self._writer.closePiece()
self._writer.closeGrid()
# Create the AppendedData
self._writer.appendData(coordinates)
self._writer.appendData(connectivity)
self._writer.appendData(offsets)
self._writer.appendData(cell_types)
self._writer.appendData(data)
self._writer.save()
def _fd_to_evtk_coord(self, fdcoord):
"""In firedrake function, the coordinates are represented by the
array."""
if len(fdcoord.shape) == 1:
# 1D case.
return (fdcoord,
np.zeros(fdcoord.shape[0]),
np.zeros(fdcoord.shape[0]))
if len(fdcoord[0]) == 3:
return (fdcoord[:, 0].ravel(),
fdcoord[:, 1].ravel(),
fdcoord[:, 2].ravel())
else:
return (fdcoord[:, 0].ravel(),
fdcoord[:, 1].ravel(),
np.zeros(fdcoord.shape[0]))
class _PVTUFile(object):
"""Class that represents PVTU file."""
def __init__(self, filename):
# filename is full path to the file without the extension.
# eg: /home/dir/dir1/filename
self._filename = filename
self._writer = PVTUWriter(self._filename)
def __del__(self):
self._writer.save()
def _update(self, function):
"""Add all the vtu to be added to pvtu file."""
for i in xrange(0, MPI.comm.size):
new_vtk_name = os.path.splitext(
self._filename)[0] + "_" + str(i) + ".vtu"
self._writer.addFile(new_vtk_name, function)
class PVTUWriter(object):
"""Class that is responsible for writing the PVTU file."""
def __init__(self, filename):
self.xml = hl.XmlWriter(filename + ".pvtu")
self.root = os.path.dirname(filename)
self.xml.openElement("VTKFile")
self.xml.addAttributes(type="PUnstructuredGrid", version="0.1",
byte_order=_get_byte_order())
self.xml.openElement("PUnstructuredGrid")
self._initialised = False
def save(self):
"""Close up the File by completing the tag."""
self.xml.closeElement("PUnstructuredGrid")
self.xml.closeElement("VTKFile")
def addFile(self, filepath, function):
"""Add VTU files to the PVTU file given in the filepath. For now, the
attributes in vtu is assumed e.g. connectivity, offsets."""
# I think I can improve this part by creating PVTU file
# from VTU file, passing the dictionary of
#{attribute_name : (data type, number of components)}
# but for now it is quite pointless since writing vtu
# is not dynamic either.
assert filepath[-4:] == ".vtu"
if not self._initialised:
self.xml.openElement("PPointData")
if len(function.shape()) == 1:
self.addData("Float64", function.name(), num_of_components=3)
elif len(function.shape()) == 0:
self.addData("Float64", function.name(), num_of_components=1)
else:
raise RuntimeError("Don't know how to write data with shape %s\n",
function.shape())
self.xml.closeElement("PPointData")
self.xml.openElement("PCellData")
self.addData("Int32", "connectivity")
self.addData("Int32", "offsets")
self.addData("UInt8", "types")
self.xml.closeElement("PCellData")
self.xml.openElement("PPoints")
self.addData("Float64", "Points", 3)
self.xml.closeElement("PPoints")
self._initialised = True
vtu_name = os.path.relpath(filepath, start=self.root)
self.xml.stream.write('<Piece Source="%s"/>\n' % vtu_name)
def addData(self, dtype, name, num_of_components=1):
"""Adds data array description of PDataArray. The header is as follows:
<PDataArray type="dtype" Name="name"
NumberOfComponents=num_of_components/>"""
self.xml.openElement("PDataArray")
self.xml.addAttributes(type=dtype, Name=name,
NumberOfComponents=num_of_components)
self.xml.closeElement("PDataArray")
class _PVDFile(object):
"""Class that represents PVD file."""
def __init__(self, filename):
# Full path to the file without extension.
self._filename = filename
self._writer = hl.VtkGroup(self._filename)
self._warnings = [False, False]
# Keep the index of child file
#(parallel -> pvtu, else vtu)
self._child_index = 0
self._time_step = -1
#_generate_time -> This file does not accept (function, time) tuple
# for __lshift__, and it generates the integer
# time step by itself instead.
self._generate_time = False
def __lshift__(self, data):
if isinstance(data, tuple):
if self._time_step == -1 or not self._generate_time:
self._time_step = data[1]
self._update_PVD(data[0])
else:
raise TypeError(
"You cannot start setting the time by giving a tuple.")
else:
if self._time_step == -1:
self._generate_time = True
if self._generate_time:
self._time_step += 1
self._update_PVD(data)
else:
raise TypeError("You need to provide time stamp")
def __del__(self):
self._writer.save()
def _update_PVD(self, function):
"""Update a pvd file.
* In parallel: create a vtu file and update it with the function given.
Then it will create a pvtu file that includes all the vtu file
produced in the parallel writing.
* In serial: a VTU file is created and is added to PVD file."""
if not MPI.parallel:
new_vtk_name = self._filename + "_" + str(self._child_index)
new_vtk = _VTUFile(new_vtk_name, warnings=self._warnings)
new_vtk << function
self._writer.addFile(new_vtk_name + ".vtu", self._time_step)
self._child_index += 1
else:
new_pvtu_name = self._filename + "_" + str(self._time_step)
new_vtk = _VTUFile(self._filename, warnings=self._warnings)
new_pvtu = _PVTUFile(new_pvtu_name)
# The new_vtk object has its timestep initialised to -1 each time,
# so we need to provide the timestep ourselves here otherwise
# the VTU of timestep 0 (belonging to the process with rank 0)
# will be over-written each time _update_PVD is called.
new_vtk << (function, self._time_step)
new_pvtu._update(function)
self._writer.addFile(new_pvtu_name + ".pvtu", self._time_step)
<file_sep>/firedrake/bcs.py
# A module implementing strong (Dirichlet) boundary conditions.
import numpy as np
from ufl import as_ufl, UFLException
import pyop2 as op2
from pyop2.profiling import timed_function
import expression
import function
import matrix
import projection
import utils
__all__ = ['DirichletBC']
class DirichletBC(object):
'''Implementation of a strong Dirichlet boundary condition.
:arg V: the :class:`.FunctionSpace` on which the boundary condition
should be applied.
:arg g: the boundary condition values. This can be a :class:`.Function` on
``V``, a :class:`.Constant`, an :class:`.Expression`, an
iterable of literal constants (converted to an
:class:`.Expression`), or a literal constant which can be
pointwise evaluated at the nodes of
``V``. :class:`.Expression`\s are projected onto ``V`` if it
does not support pointwise evaluation.
:arg sub_domain: the integer id of the boundary region over which the
boundary condition should be applied. In the case of extrusion
the ``top`` and ``bottom`` strings are used to flag the bcs application on
the top and bottom boundaries of the extruded mesh respectively.
:arg method: the method for determining boundary nodes. The default is
"topological", indicating that nodes topologically associated with a
boundary facet will be included. The alternative value is "geometric",
which indicates that nodes associated with basis functions which do not
vanish on the boundary will be included. This can be used to impose
strong boundary conditions on DG spaces, or no-slip conditions on HDiv spaces.
.. warning::
Geometric boundary conditions are not yet supported on extruded meshes
'''
def __init__(self, V, g, sub_domain, method="topological"):
self._function_space = V
# Save the original value the user passed in. If the user
# passed in an Expression that has user-defined variables in
# it, we need to remember it so that we can re-interpolate it
# onto the function_arg if its state has changed. Note that
# the function_arg assignment is actually a property setter
# which in the case of expressions interpolates it onto a
# function and then throws the expression away.
self._original_val = g
self.function_arg = g
self._original_arg = self.function_arg
self.sub_domain = sub_domain
self._currently_zeroed = False
if method not in ["topological", "geometric"]:
raise ValueError("Unknown boundary condition method %s" % method)
self.method = method
if V.extruded and method == "geometric":
raise ValueError("Geometric boundary conditions are not yet supported on extruded meshes")
@property
def function_arg(self):
'''The value of this boundary condition.'''
if isinstance(self._original_val, expression.Expression):
if not self._currently_zeroed and \
self._original_val._state != self._expression_state:
# Expression values have changed, need to reinterpolate
self.function_arg = self._original_val
return self._function_arg
@function_arg.setter
def function_arg(self, g):
'''Set the value of this boundary condition.'''
if isinstance(g, function.Function) and g.function_space() != self._function_space:
raise RuntimeError("%r is defined on incompatible FunctionSpace!" % g)
if not isinstance(g, expression.Expression):
try:
# Bare constant?
as_ufl(g)
except UFLException:
try:
# List of bare constants? Convert to Expression
g = expression.to_expression(g)
except:
raise ValueError("%r is not a valid DirichletBC expression" % (g,))
if isinstance(g, expression.Expression):
self._expression_state = g._state
try:
g = function.Function(self._function_space).interpolate(g)
# Not a point evaluation space, need to project onto V
except NotImplementedError:
g = projection.project(g, self._function_space)
self._function_arg = g
self._currently_zeroed = False
def function_space(self):
'''The :class:`.FunctionSpace` on which this boundary condition should
be applied.'''
return self._function_space
def homogenize(self):
'''Convert this boundary condition into a homogeneous one.
Set the value to zero.
'''
self.function_arg = 0
self._currently_zeroed = True
def restore(self):
'''Restore the original value of this boundary condition.
This uses the value passed on instantiation of the object.'''
self._function_arg = self._original_arg
self._currently_zeroed = False
def set_value(self, val):
'''Set the value of this boundary condition.
:arg val: The boundary condition values. See
:class:`.DirichletBC` for valid values.
'''
self.function_arg = val
self._original_arg = self.function_arg
self._original_val = val
@utils.cached_property
def nodes(self):
'''The list of nodes at which this boundary condition applies.'''
fs = self._function_space
if self.sub_domain == "bottom":
return fs.bottom_nodes()
elif self.sub_domain == "top":
return fs.top_nodes()
else:
if fs.extruded:
base_maps = fs.exterior_facet_boundary_node_map(
self.method).values_with_halo.take(
fs._mesh._old_mesh.exterior_facets.subset(self.sub_domain).indices,
axis=0)
facet_offset = fs.exterior_facet_boundary_node_map(self.method).offset
return np.unique(np.concatenate([base_maps + i * facet_offset
for i in range(fs._mesh.layers - 1)]))
return np.unique(
fs.exterior_facet_boundary_node_map(
self.method).values_with_halo.take(
fs._mesh.exterior_facets.subset(self.sub_domain).indices,
axis=0))
@utils.cached_property
def node_set(self):
'''The subset corresponding to the nodes at which this
boundary condition applies.'''
return op2.Subset(self._function_space.node_set, self.nodes)
@timed_function('DirichletBC apply')
def apply(self, r, u=None):
"""Apply this boundary condition to ``r``.
:arg r: a :class:`.Function` or :class:`.Matrix` to which the
boundary condition should be applied.
:arg u: an optional current state. If ``u`` is supplied then
``r`` is taken to be a residual and the boundary condition
nodes are set to the value ``u-bc``. Supplying ``u`` has
no effect if ``r`` is a :class:`.Matrix` rather than a
:class:`.Function`. If ``u`` is absent, then the boundary
condition nodes of ``r`` are set to the boundary condition
values.
If ``r`` is a :class:`.Matrix`, it will be assembled with a 1
on diagonals where the boundary condition applies and 0 in the
corresponding rows and columns.
"""
if isinstance(r, matrix.Matrix):
r.add_bc(self)
return
fs = self._function_space
# Check if the FunctionSpace of the Function r to apply is compatible.
# If fs is an IndexedFunctionSpace and r is defined on a
# MixedFunctionSpace, we need to compare the parent of fs
if not (fs == r.function_space() or (hasattr(fs, "_parent") and
fs._parent == r.function_space())):
raise RuntimeError("%r defined on incompatible FunctionSpace!" % r)
# If this BC is defined on a subspace of a mixed function space, make
# sure we only apply to the appropriate subspace of the Function r
if fs.index is not None:
r = function.Function(self._function_space, r.dat[fs.index])
if u:
if fs.index is not None:
u = function.Function(fs, u.dat[fs.index])
r.assign(u - self.function_arg, subset=self.node_set)
else:
r.assign(self.function_arg, subset=self.node_set)
def zero(self, r):
"""Zero the boundary condition nodes on ``r``.
:arg r: a :class:`.Function` to which the
boundary condition should be applied.
"""
if isinstance(r, matrix.Matrix):
raise NotImplementedError("Zeroing bcs on a Matrix is not supported")
# Record whether we are homogenized on entry.
currently_zeroed = self._currently_zeroed
self.homogenize()
self.apply(r)
if not currently_zeroed:
self.restore()
<file_sep>/firedrake/version.py
__version_info__ = (0, 11, 0)
__version__ = '.'.join(map(str, __version_info__))
__compatible_ffc_version_info__ = (0, 6, 0)
__compatible_ffc_version__ = '.'.join(map(str, __compatible_ffc_version_info__))
def check():
from pyop2.version import __version_info__ as pyop2_version_info
from pyop2.version import __version__ as pyop2_version
if pyop2_version_info[:2] != __version_info__[:2]:
raise RuntimeError(
"""Firedrake version %s and PyOP2 version %s are incompatible.""" %
(__version__, pyop2_version))
<file_sep>/docs/source/team.rst
The Firedrake team
==================
Firedrake is brought to you by the departments of `Computing <http://www.imperial.ac.uk/computing/>`_, `Mathematics <http://www3.imperial.ac.uk/mathematics>`_, and `Earth Science and Engineering <http://www.imperial.ac.uk/ese/>`_ at `Imperial College London <http://www3.imperial.ac.uk>`_.
.. include:: teamgrid.rst
Summer students 2013
--------------------
<NAME>
<NAME>
<NAME>
`<NAME>ister <http://www5.in.tum.de/wiki/index.php/Dipl.-Inf._Oliver_Meister>`_
<NAME>
<file_sep>/tests/regression/test_matrix.py
from firedrake import *
from firedrake import matrix
import pytest
@pytest.fixture
def V():
mesh = UnitIntervalMesh(2)
V = FunctionSpace(mesh, "CG", 1)
return V
@pytest.fixture
def a(V):
u = TrialFunction(V)
v = TestFunction(V)
return u*v*dx
def test_assemble_returns_matrix(a):
A = assemble(a)
assert isinstance(A, matrix.Matrix)
def test_assemble_is_lazy(a):
A = assemble(a)
assert not A.assembled
assert A._assembly_callback is not None
assert (A._M.values == 0.0).all()
def test_M_forces_assemble(a):
A = assemble(a)
assert not A.assembled
assert not (A.M.values == 0.0).all()
assert A.assembled
def test_solve_forces_assemble(a, V):
A = assemble(a)
v = TestFunction(V)
f = Function(V)
b = assemble(f*v*dx)
assert not A.assembled
solve(A, f, b)
assert A.assembled
assert not (A._M.values == 0.0).all()
def test_adding_bcs(a, V):
bc1 = DirichletBC(V, 0, 1)
A = assemble(a, bcs=[bc1])
assert set(A.bcs) == set([bc1])
bc2 = DirichletBC(V, 1, 1)
bc2.apply(A)
assert set(A.bcs) == set([bc2])
bc3 = DirichletBC(V, 1, 0)
bc3.apply(A)
assert set(A.bcs) == set([bc2, bc3])
def test_assemble_with_bcs(a, V):
bc1 = DirichletBC(V, 0, 1)
A = assemble(a, bcs=[bc1])
A.assemble()
assert A.assembled
assert not A._needs_reassembly
# Same subdomain, should not need reassembly
bc2 = DirichletBC(V, 1, 1)
bc2.apply(A)
assert A.assembled
assert not A._needs_reassembly
bc3 = DirichletBC(V, 1, 0)
A.bcs = bc3
assert A.assembled
assert A._needs_reassembly
A.assemble()
assert A.assembled
assert not A._needs_reassembly
bc2.apply(A)
assert A.assembled
assert A._needs_reassembly
A.assemble()
assert A.assembled
assert not A._needs_reassembly
def test_assemble_with_bcs_then_not(a, V):
bc1 = DirichletBC(V, 0, 1)
A = assemble(a, bcs=[bc1])
Abcs = A.M.values
A = assemble(a)
assert not A.has_bcs
Anobcs = A.M.values
assert (Anobcs != Abcs).any()
A = assemble(a, bcs=[bc1])
Abcs = A.M.values
assemble(a, tensor=A)
Anobcs = A.M.values
assert not A.has_bcs
assert (Anobcs != Abcs).any()
def test_assemble_with_bcs_multiple_subdomains(a, V):
bc1 = DirichletBC(V, 0, [0, 1])
A = assemble(a, bcs=[bc1])
assert not A.assembled
assert A._needs_reassembly
A.assemble()
assert A.assembled
assert not A._needs_reassembly
def test_form_action(a, V):
A = assemble(a)
u1 = A._form_action(Function(V).assign(1.0))
u2 = A._form_action(Function(V).assign(2.0))
assert (2.0*u1.dat.data == u2.dat.data).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_embedded_sphere.py
import pytest
import numpy as np
from tests.common import longtest
from firedrake import *
def integrate_spherical_annulus_volume(radius=1000, refinement=2):
m = IcosahedralSphereMesh(radius=radius, refinement_level=refinement)
layers = 10
layer_height = 1.0 / (radius * layers)
mesh = ExtrudedMesh(m, layers, layer_height=layer_height, extrusion_type='radial')
fs = FunctionSpace(mesh, 'CG', 1, name="fs")
f = Function(fs)
f.assign(1)
exact = 4 * pi * ((radius + 1.0/radius)**3 - radius**3) / 3
return np.abs(assemble(f * dx) - exact) / exact
@longtest
@pytest.mark.parametrize(('radius', 'refinement', 'error'),
[(1000, 2, 0.04),
(10000, 2, 0.04),
(1000, 5, 0.0006),
(10000, 5, 0.0006)])
def test_volume_spherical_annulus(radius, refinement, error):
assert integrate_spherical_annulus_volume(radius=radius, refinement=refinement) < error
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_constant.py
from firedrake import *
import numpy as np
import pytest
def test_scalar_constant():
for m in [UnitIntervalMesh(5), UnitSquareMesh(2, 2), UnitCubeMesh(2, 2, 2)]:
c = Constant(1, domain=m.ufl_domain())
assert abs(assemble(c*m._dx) - 1.0) < 1e-10
def test_scalar_constant_assign():
for m in [UnitIntervalMesh(5), UnitSquareMesh(2, 2), UnitCubeMesh(2, 2, 2)]:
c = Constant(1, domain=m.ufl_domain())
assert abs(assemble(c*m._dx) - 1.0) < 1e-10
c.assign(4)
assert abs(assemble(c*m._dx) - 4.0) < 1e-10
@pytest.mark.parametrize(('init', 'new_vals'),
((1, ([1, 1], "x", [[1, 1], [1, 1]])),
([1, 1], ([1, "x"], "x", 1, [[1, 1], [1, 1]])),
([[1], [1]], ([1, "x"], "x", 1, [[1, 1], [1, 1]]))))
def test_constant_assign_mismatch(init, new_vals):
c = Constant(init)
for v in new_vals:
with pytest.raises(ValueError):
c.assign(v)
def test_vector_constant_2d():
m = UnitSquareMesh(1, 1)
n = FacetNormal(m)
c = Constant([1, -1])
# Mesh is:
# ,---.
# |\ |
# | \ |
# | \|
# `---'
# Normal is in (1, 1) direction
assert abs(assemble(dot(c('+'), n('+'))*dS)) < 1e-10
assert abs(assemble(dot(c('-'), n('+'))*dS)) < 1e-10
# Normal is in (-1, -1) direction
assert abs(assemble(dot(c('+'), n('-'))*dS)) < 1e-10
assert abs(assemble(dot(c('-'), n('-'))*dS)) < 1e-10
c.assign([1, 1])
assert abs(assemble(dot(c('+'), n('+'))*dS) - 2) < 1e-10
assert abs(assemble(dot(c('-'), n('+'))*dS) - 2) < 1e-10
# Normal is in (-1, -1) direction
assert abs(assemble(dot(c('+'), n('-'))*dS) + 2) < 1e-10
assert abs(assemble(dot(c('-'), n('-'))*dS) + 2) < 1e-10
def test_tensor_constant():
mesh = UnitSquareMesh(4, 4)
V = VectorFunctionSpace(mesh, "CG", 1)
v = Function(V)
v.assign(1.0)
sigma = Constant(((1., 0.), (0., 2.)))
val = assemble(inner(v, dot(sigma, v))*dx)
assert abs(val-3.0) < 1.0e-10
def test_constant_scalar_assign_distributes():
m = UnitSquareMesh(1, 1)
V = VectorFunctionSpace(m, 'CG', 1)
f = Function(V)
c = Constant(11)
f.assign(c)
assert np.allclose(f.dat.data_ro, 11)
def test_constant_vector_assign_works():
m = UnitSquareMesh(1, 1)
V = VectorFunctionSpace(m, 'CG', 1)
f = Function(V)
c = Constant([10, 11])
f.assign(c)
assert np.allclose(f.dat.data_ro[:, 0], 10)
assert np.allclose(f.dat.data_ro[:, 1], 11)
def test_constant_vector_assign_to_scalar_error():
m = UnitSquareMesh(1, 1)
V = FunctionSpace(m, 'CG', 1)
f = Function(V)
c = Constant([10, 11])
with pytest.raises(ValueError):
f.assign(c)
def test_constant_vector_assign_to_vector_mismatch_error():
m = UnitSquareMesh(1, 1)
V = VectorFunctionSpace(m, 'CG', 1)
f = Function(V)
c = Constant([10, 11, 12])
with pytest.raises(ValueError):
f.assign(c)
def test_constant_assign_to_mixed():
m = UnitSquareMesh(1, 1)
V = VectorFunctionSpace(m, 'CG', 1)
W = V*V
f = Function(W)
c = Constant([10, 11])
f.assign(c)
for d in f.dat.data_ro:
assert np.allclose(d[:, 0], 10)
assert np.allclose(d[:, 1], 11)
def test_constant_multiplies_function():
m = UnitSquareMesh(1, 1)
V = FunctionSpace(m, 'CG', 1)
u = Function(V)
u.assign(10)
f = Function(V)
c = Constant(11)
f.assign(u * c)
assert np.allclose(f.dat.data_ro, 110)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_bcs.py
import pytest
import numpy as np
from firedrake import *
@pytest.fixture(scope='module')
def mesh():
return UnitSquareMesh(2, 2)
@pytest.fixture(scope='module', params=[FunctionSpace, VectorFunctionSpace])
def V(request, mesh):
return request.param(mesh, "CG", 1)
@pytest.fixture
def u(V):
return Function(V)
@pytest.fixture
def a(u, V):
v = TestFunction(V)
return inner(grad(v), grad(u)) * dx
@pytest.fixture
def f(V):
return Function(V).assign(10)
@pytest.fixture
def f2(mesh):
return Function(FunctionSpace(mesh, 'CG', 2))
@pytest.mark.parametrize('v', [0, 1.0])
def test_init_bcs(V, v):
"Initialise a DirichletBC."
assert DirichletBC(V, v, 0).function_arg == v
@pytest.mark.parametrize('v', [(0, 0), 'foo'])
def test_init_bcs_illegal(mesh, v):
"Initialise a DirichletBC with illegal values."
with pytest.raises(RuntimeError):
DirichletBC(FunctionSpace(mesh, "CG", 1), v, 0)
@pytest.mark.parametrize('measure', [dx, ds])
def test_assemble_bcs_wrong_fs(V, measure):
"Assemble a Matrix with a DirichletBC on an incompatible FunctionSpace."
u, v = TestFunction(V), TrialFunction(V)
W = FunctionSpace(V.mesh(), "CG", 2)
A = assemble(dot(u, v)*measure, bcs=[DirichletBC(W, 32, 1)])
with pytest.raises(RuntimeError):
A.M.values
def test_assemble_bcs_wrong_fs_interior(V):
"Assemble a Matrix with a DirichletBC on an incompatible FunctionSpace."
u, v = TestFunction(V), TrialFunction(V)
W = FunctionSpace(V.mesh(), "CG", 2)
n = FacetNormal(V.mesh())
A = assemble(inner(jump(u, n), jump(v, n))*dS, bcs=[DirichletBC(W, 32, 1)])
with pytest.raises(RuntimeError):
A.M.values
def test_apply_bcs_wrong_fs(V, f2):
"Applying a DirichletBC to a Function on an incompatible FunctionSpace."
bc = DirichletBC(V, 32, 1)
with pytest.raises(RuntimeError):
bc.apply(f2)
def test_zero_bcs_wrong_fs(V, f2):
"Zeroing a DirichletBC on a Function on an incompatible FunctionSpace."
bc = DirichletBC(V, 32, 1)
with pytest.raises(RuntimeError):
bc.zero(f2)
def test_init_bcs_wrong_fs(V, f2):
"Initialise a DirichletBC with a Function on an incompatible FunctionSpace."
with pytest.raises(RuntimeError):
DirichletBC(V, f2, 1)
def test_set_bcs_wrong_fs(V, f2):
"Set a DirichletBC to a Function on an incompatible FunctionSpace."
bc = DirichletBC(V, 32, 1)
with pytest.raises(RuntimeError):
bc.set_value(f2)
def test_homogeneous_bcs(a, u, V):
bcs = [DirichletBC(V, 32, 1)]
[bc.homogenize() for bc in bcs]
# Compute solution - this should have the solution u = 0
solve(a == 0, u, bcs=bcs)
assert abs(u.vector().array()).max() == 0.0
def test_homogenize_doesnt_overwrite_function(a, u, V, f):
bc = DirichletBC(V, f, 1)
bc.homogenize()
assert (f.vector().array() == 10.0).all()
solve(a == 0, u, bcs=[bc])
assert abs(u.vector().array()).max() == 0.0
def test_restore_bc_value(a, u, V, f):
bc = DirichletBC(V, f, 1)
bc.homogenize()
solve(a == 0, u, bcs=[bc])
assert abs(u.vector().array()).max() == 0.0
bc.restore()
solve(a == 0, u, bcs=[bc])
assert np.allclose(u.vector().array(), 10.0)
def test_set_bc_value(a, u, V, f):
bc = DirichletBC(V, f, 1)
bc.set_value(7)
solve(a == 0, u, bcs=[bc])
assert np.allclose(u.vector().array(), 7.0)
def test_update_bc_expression(a, u, V, f):
if isinstance(V, VectorFunctionSpace):
e = Expression(['t', 't'], t=1.0)
else:
e = Expression('t', t=1.0)
bc = DirichletBC(V, e, 1)
solve(a == 0, u, bcs=[bc])
# We should get the value in the expression
assert np.allclose(u.vector().array(), 1.0)
e.t = 2.0
solve(a == 0, u, bcs=[bc])
# Updating the expression value should give new value.
assert np.allclose(u.vector().array(), 2.0)
e.t = 3.0
bc.homogenize()
solve(a == 0, u, bcs=[bc])
# Homogenized bcs shouldn't be overridden by the expression
# changing.
assert np.allclose(u.vector().array(), 0.0)
bc.restore()
solve(a == 0, u, bcs=[bc])
# Restoring the bcs should give the new expression value.
assert np.allclose(u.vector().array(), 3.0)
bc.set_value(7)
solve(a == 0, u, bcs=[bc])
# Setting a value should replace the expression
assert np.allclose(u.vector().array(), 7.0)
e.t = 4.0
solve(a == 0, u, bcs=[bc])
# And now we should just have the new value (since the expression
# is gone)
assert np.allclose(u.vector().array(), 7.0)
def test_update_bc_constant(a, u, V, f):
if isinstance(V, VectorFunctionSpace):
# Don't bother with the VFS case
return
c = Constant(1)
bc = DirichletBC(V, c, 1)
solve(a == 0, u, bcs=[bc])
# We should get the value in the constant
assert np.allclose(u.vector().array(), 1.0)
c.assign(2.0)
solve(a == 0, u, bcs=[bc])
# Updating the constant value should give new value.
assert np.allclose(u.vector().array(), 2.0)
c.assign(3.0)
bc.homogenize()
solve(a == 0, u, bcs=[bc])
# Homogenized bcs shouldn't be overridden by the constant
# changing.
assert np.allclose(u.vector().array(), 0.0)
bc.restore()
solve(a == 0, u, bcs=[bc])
# Restoring the bcs should give the new constant value.
assert np.allclose(u.vector().array(), 3.0)
bc.set_value(7)
solve(a == 0, u, bcs=[bc])
# Setting a value should replace the constant
assert np.allclose(u.vector().array(), 7.0)
c.assign(4.0)
solve(a == 0, u, bcs=[bc])
# And now we should just have the new value (since the constant
# is gone)
assert np.allclose(u.vector().array(), 7.0)
def test_preassembly_change_bcs(V, f):
v = TestFunction(V)
u = TrialFunction(V)
a = dot(u, v)*dx
bc = DirichletBC(V, f, 1)
A = assemble(a, bcs=[bc])
L = dot(v, f)*dx
b = assemble(L)
y = Function(V)
y.assign(7)
bc1 = DirichletBC(V, y, 1)
u = Function(V)
solve(A, u, b)
assert np.allclose(u.vector().array(), 10.0)
u.assign(0)
b = assemble(dot(v, y)*dx)
solve(A, u, b, bcs=[bc1])
assert np.allclose(u.vector().array(), 7.0)
def test_preassembly_doesnt_modify_assembled_rhs(V, f):
v = TestFunction(V)
u = TrialFunction(V)
a = dot(u, v)*dx
bc = DirichletBC(V, f, 1)
A = assemble(a, bcs=[bc])
L = dot(v, f)*dx
b = assemble(L)
b_vals = b.vector().array()
u = Function(V)
solve(A, u, b)
assert np.allclose(u.vector().array(), 10.0)
assert np.allclose(b_vals, b.vector().array())
def test_preassembly_bcs_caching(V):
bc1 = DirichletBC(V, 0, 1)
bc2 = DirichletBC(V, 1, 2)
v = TestFunction(V)
u = TrialFunction(V)
a = dot(u, v)*dx
Aboth = assemble(a, bcs=[bc1, bc2])
Aneither = assemble(a)
A1 = assemble(a, bcs=[bc1])
A2 = assemble(a, bcs=[bc2])
assert not np.allclose(Aboth.M.values, Aneither.M.values)
assert not np.allclose(Aboth.M.values, A2.M.values)
assert not np.allclose(Aboth.M.values, A1.M.values)
assert not np.allclose(Aneither.M.values, A2.M.values)
assert not np.allclose(Aneither.M.values, A1.M.values)
assert not np.allclose(A2.M.values, A1.M.values)
# There should be no zeros on the diagonal
assert not any(A2.M.values.diagonal() == 0)
assert not any(A1.M.values.diagonal() == 0)
assert not any(Aneither.M.values.diagonal() == 0)
def test_assemble_mass_bcs_2d(V):
u = TrialFunction(V)
v = TestFunction(V)
f = Function(V).interpolate(Expression(['x[0]'] * V.dim))
bcs = [DirichletBC(V, 0.0, 1),
DirichletBC(V, 1.0, 2)]
w = Function(V)
solve(dot(u, v)*dx == dot(f, v)*dx, w, bcs=bcs)
assert assemble(dot((w - f), (w - f))*dx) < 1e-12
def test_mixed_bcs():
m = UnitSquareMesh(2, 2)
V = FunctionSpace(m, 'CG', 1)
W = V*V
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
bc = DirichletBC(W.sub(1), 0.0, (1, 2, 3, 4))
A = assemble(inner(u, v)*dx, bcs=bc)
A11 = A.M[1, 1].values
assert np.allclose(A11.diagonal()[bc.nodes], 1.0)
def test_bcs_rhs_assemble(a, V):
bcs = [DirichletBC(V, 1.0, 1), DirichletBC(V, 2.0, 3)]
b1 = assemble(a)
for bc in bcs:
bc.apply(b1)
b2 = assemble(a, bcs=bcs)
assert np.allclose(b1.dat.data, b2.dat.data)
@pytest.mark.parallel(nprocs=3)
def test_empty_exterior_facet_node_list():
mesh = UnitIntervalMesh(6)
V = FunctionSpace(mesh, 'CG', 1)
bc = DirichletBC(V, 1, 1)
assert V.exterior_facet_node_map([bc])
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_extrusion_two_step.py
"""Testing extruded RT elements."""
import pytest
import numpy as np
from firedrake import *
def two_step():
power = 4
# Create mesh and define function space
m = UnitSquareMesh(2 ** power, 2 ** power)
layers = 10
# Populate the coordinates of the extruded mesh by providing the
# coordinates as a field.
mesh = ExtrudedMesh(m, layers, layer_height=0.1)
V = FunctionSpace(mesh, "Lagrange", 2, vfamily="DG", vdegree=0)
horiz = FiniteElement("BDM", "triangle", 1)
vert = FiniteElement("DG", "interval", 0)
prod = HDiv(OuterProductElement(horiz, vert))
W = FunctionSpace(mesh, prod)
X = FunctionSpace(mesh, "DG", 0, vfamily="DG", vdegree=0)
# Define starting field
f0 = Function(V)
f0.interpolate(Expression("1 + x[0]*x[0] + x[1]*x[1]"))
# DO IN ONE STEP
u = TrialFunction(X)
v = TestFunction(X)
a = u * v * dx
L = div(grad(f0)) * v * dx
assemble(a)
assemble(L)
f_e = Function(X)
solve(a == L, f_e)
# DO IN TWO STEPS
u = TrialFunction(W)
v = TestFunction(W)
a = dot(u, v) * dx
L = dot(grad(f0), v) * dx
# Compute solution
assemble(a)
assemble(L)
f1 = Function(W)
solve(a == L, f1)
# FIXME x should be (2x, 2y) but we have no way of checking
u = TrialFunction(X)
v = TestFunction(X)
a = u * v * dx
L = div(f1) * v * dx
# Compute solution
assemble(a)
assemble(L)
f2 = Function(X)
solve(a == L, f2)
return np.max(np.abs(f2.dat.data - f_e.dat.data))
def test_firedrake_extrusion_two_step():
assert two_step() < 1.0e-4
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/utility_meshes.py
import numpy as np
import os
import tempfile
from shutil import rmtree
from pyop2.mpi import MPI
from pyop2.profiling import profile
import mesh
from petsc import PETSc
__all__ = ['IntervalMesh', 'UnitIntervalMesh',
'PeriodicIntervalMesh', 'PeriodicUnitIntervalMesh',
'UnitTriangleMesh',
'RectangleMesh', 'SquareMesh', 'UnitSquareMesh',
'CircleMesh', 'UnitCircleMesh',
'CircleManifoldMesh',
'UnitTetrahedronMesh',
'BoxMesh', 'CubeMesh', 'UnitCubeMesh',
'IcosahedralSphereMesh', 'UnitIcosahedralSphereMesh']
_cachedir = os.path.join(tempfile.gettempdir(),
'firedrake-mesh-cache-uid%d' % os.getuid())
def _ensure_cachedir():
if MPI.comm.rank == 0 and not os.path.exists(_cachedir):
os.makedirs(_cachedir)
_ensure_cachedir()
def _clear_cachedir():
if MPI.comm.rank == 0 and os.path.exists(_cachedir):
rmtree(_cachedir, ignore_errors=True)
_ensure_cachedir()
def _msh_exists(name):
f = os.path.join(_cachedir, name)
return os.path.exists(f + '.msh')
def _build_msh_file(input, output, dimension):
try:
# Must occur after mpi4py import due to:
# 1) MPI initialisation issues
# 2) LD_PRELOAD issues
import gmshpy
gmshpy.Msg.SetVerbosity(-1)
# We've got the gmsh python interface available, so
# use that, rather than spawning the gmsh binary.
m = gmshpy.GModel()
m.readGEO(input)
m.mesh(dimension)
m.writeMSH(output + ".msh")
return
except ImportError:
raise RuntimeError('Creation of gmsh meshes requires gmshpy')
def _get_msh_file(source, name, dimension, meshed=False):
"""Given a source code, name and dimension of the mesh,
returns the name of the file that contains necessary information to build
a mesh class. The mesh class would call _from_file method on this file
to contruct itself.
"""
if MPI.comm.rank == 0:
input = os.path.join(_cachedir, name + '.geo')
if not meshed:
if not os.path.exists(input):
with open(input, 'w') as f:
f.write(source)
output = os.path.join(_cachedir, name)
if not _msh_exists(name):
if meshed:
with file(output + '.msh', 'w') as f:
f.write(source)
else:
_build_msh_file(input, output, dimension)
MPI.comm.bcast(output, root=0)
else:
output = MPI.comm.bcast(None, root=0)
return output + '.msh'
def _from_cell_list(dim, cells, coords, comm=None):
"""
Create a DMPlex from a list of cells and coords.
:arg dim: The topological dimension of the mesh
:arg cells: The vertices of each cell
:arg coords: The coordinates of each vertex
:arg comm: An optional MPI communicator to build the plex on
(defaults to ``COMM_WORLD``)
"""
if comm is None:
comm = MPI.comm
if comm.rank == 0:
cells = np.asarray(cells, dtype=PETSc.IntType)
coords = np.asarray(coords, dtype=float)
comm.bcast(cells.shape, root=0)
comm.bcast(coords.shape, root=0)
# Provide the actual data on rank 0.
return PETSc.DMPlex().createFromCellList(dim, cells, coords, comm=comm)
cell_shape = list(comm.bcast(None, root=0))
coord_shape = list(comm.bcast(None, root=0))
cell_shape[0] = 0
coord_shape[0] = 0
# Provide empty plex on other ranks
# A subsequent call to plex.distribute() takes care of parallel partitioning
return PETSc.DMPlex().createFromCellList(dim,
np.zeros(cell_shape, dtype=PETSc.IntType),
np.zeros(coord_shape, dtype=float),
comm=comm)
@profile
def IntervalMesh(ncells, length):
"""
Generate a uniform mesh of the interval [0,L].
:arg ncells: The number of the cells over the interval.
:arg length: The length of the interval.
The left hand (:math:`x=0`) boundary point has boundary marker 1,
while the right hand (:math:`x=L`) point has marker 2.
"""
dx = float(length) / ncells
# This ensures the rightmost point is actually present.
coords = np.arange(0, length + 0.01 * dx, dx).reshape(-1, 1)
cells = np.dstack((np.arange(0, len(coords) - 1, dtype=np.int32),
np.arange(1, len(coords), dtype=np.int32))).reshape(-1, 2)
plex = _from_cell_list(1, cells, coords)
# Apply boundary IDs
plex.createLabel("boundary_ids")
coordinates = plex.getCoordinates()
coord_sec = plex.getCoordinateSection()
vStart, vEnd = plex.getDepthStratum(0) # vertices
for v in range(vStart, vEnd):
vcoord = plex.vecGetClosure(coord_sec, coordinates, v)
if vcoord[0] == coords[0]:
plex.setLabelValue("boundary_ids", v, 1)
if vcoord[0] == coords[-1]:
plex.setLabelValue("boundary_ids", v, 2)
return mesh.Mesh(plex, reorder=False)
def UnitIntervalMesh(ncells):
"""
Generate a uniform mesh of the interval [0,1].
:arg ncells: The number of the cells over the interval.
The left hand (:math:`x=0`) boundary point has boundary marker 1,
while the right hand (:math:`x=1`) point has marker 2.
"""
return IntervalMesh(ncells, length=1.0)
@profile
def PeriodicIntervalMesh(ncells, length):
"""Generate a periodic mesh of an interval.
:arg ncells: The number of cells over the interval.
:arg length: The length the interval."""
if MPI.comm.size > 1:
raise NotImplementedError("Periodic intervals not yet implemented in parallel")
nvert = ncells
nedge = ncells
plex = PETSc.DMPlex().create()
plex.setDimension(1)
plex.setChart(0, nvert+nedge)
for e in range(nedge):
plex.setConeSize(e, 2)
plex.setUp()
for e in range(nedge-1):
plex.setCone(e, [nedge+e, nedge+e+1])
plex.setConeOrientation(e, [0, 0])
# Connect v_(n-1) with v_0
plex.setCone(nedge-1, [nedge+nvert-1, nedge])
plex.setConeOrientation(nedge-1, [0, 0])
plex.symmetrize()
plex.stratify()
# Build coordinate section
dx = float(length) / ncells
coords = [x for x in np.arange(0, length + 0.01 * dx, dx)]
coordsec = plex.getCoordinateSection()
coordsec.setChart(nedge, nedge+nvert)
for v in range(nedge, nedge+nvert):
coordsec.setDof(v, 1)
coordsec.setUp()
size = coordsec.getStorageSize()
coordvec = PETSc.Vec().createWithArray(coords, size=size)
plex.setCoordinatesLocal(coordvec)
dx = length / ncells
# HACK ALERT!
# Almost certainly not right when symbolic geometry stuff lands.
# Hopefully DMPlex will eventually give us a DG coordinate
# field. Until then, we build one by hand.
coords = np.dstack((np.arange(dx, length + dx*0.01, dx),
np.arange(0, length - dx*0.01, dx))).flatten()
# Last cell is back to front.
coords[-2:] = coords[-2:][::-1]
return mesh.Mesh(plex, periodic_coords=coords, reorder=False)
def PeriodicUnitIntervalMesh(ncells):
"""Generate a periodic mesh of the unit interval
:arg ncells: The number of cells in the interval.
"""
return PeriodicIntervalMesh(ncells, length=1.0)
def UnitTriangleMesh():
"""Generate a mesh of the reference triangle"""
coords = [[0., 0.], [1., 0.], [0., 1.]]
cells = [[0, 1, 2]]
plex = _from_cell_list(2, cells, coords)
return mesh.Mesh(plex, reorder=False)
@profile
def RectangleMesh(nx, ny, Lx, Ly, quadrilateral=False, reorder=None):
"""Generate a rectangular mesh
:arg nx: The number of cells in the x direction
:arg ny: The number of cells in the y direction
:arg Lx: The extent in the x direction
:arg Ly: The extent in the y direction
:kwarg quadrilateral: (optional), creates quadrilateral mesh, defaults to False
:kwarg reorder: (optional), should the mesh be reordered
The boundary edges in this mesh are numbered as follows:
* 1: plane x == 0
* 2: plane x == Lx
* 3: plane y == 0
* 4: plane y == Ly
"""
if quadrilateral:
dx = float(Lx) / nx
dy = float(Ly) / ny
xcoords = np.arange(0.0, Lx + 0.01 * dx, dx)
ycoords = np.arange(0.0, Ly + 0.01 * dy, dy)
coords = np.asarray(np.meshgrid(xcoords, ycoords)).swapaxes(0, 2).reshape(-1, 2)
# cell vertices
i, j = np.meshgrid(np.arange(nx), np.arange(ny))
cells = [i*(ny+1) + j, i*(ny+1) + j+1, (i+1)*(ny+1) + j+1, (i+1)*(ny+1) + j]
cells = np.asarray(cells).swapaxes(0, 2).reshape(-1, 4)
plex = _from_cell_list(2, cells, coords)
else:
boundary = PETSc.DMPlex().create(MPI.comm)
boundary.setDimension(1)
boundary.createSquareBoundary([0., 0.], [float(Lx), float(Ly)], [nx, ny])
boundary.setTriangleOptions("pqezQYSl")
plex = PETSc.DMPlex().generate(boundary)
# mark boundary facets
plex.createLabel("boundary_ids")
plex.markBoundaryFaces("boundary_faces")
coords = plex.getCoordinates()
coord_sec = plex.getCoordinateSection()
if plex.getStratumSize("boundary_faces", 1) > 0:
boundary_faces = plex.getStratumIS("boundary_faces", 1).getIndices()
xtol = float(Lx)/(2*nx)
ytol = float(Ly)/(2*ny)
for face in boundary_faces:
face_coords = plex.vecGetClosure(coord_sec, coords, face)
if abs(face_coords[0]) < xtol and abs(face_coords[2]) < xtol:
plex.setLabelValue("boundary_ids", face, 1)
if abs(face_coords[0] - Lx) < xtol and abs(face_coords[2] - Lx) < xtol:
plex.setLabelValue("boundary_ids", face, 2)
if abs(face_coords[1]) < ytol and abs(face_coords[3]) < ytol:
plex.setLabelValue("boundary_ids", face, 3)
if abs(face_coords[1] - Ly) < ytol and abs(face_coords[3] - Ly) < ytol:
plex.setLabelValue("boundary_ids", face, 4)
return mesh.Mesh(plex, reorder=reorder)
def SquareMesh(nx, ny, L, reorder=None, quadrilateral=False):
"""Generate a square mesh
:arg nx: The number of cells in the x direction
:arg ny: The number of cells in the y direction
:arg L: The extent in the x and y directions
:kwarg quadrilateral: (optional), creates quadrilateral mesh, defaults to False
:kwarg reorder: (optional), should the mesh be reordered
The boundary edges in this mesh are numbered as follows:
* 1: plane x == 0
* 2: plane x == L
* 3: plane y == 0
* 4: plane y == L
"""
return RectangleMesh(nx, ny, L, L, reorder=reorder, quadrilateral=quadrilateral)
def UnitSquareMesh(nx, ny, reorder=None, quadrilateral=False):
"""Generate a unit square mesh
:arg nx: The number of cells in the x direction
:arg ny: The number of cells in the y direction
:kwarg quadrilateral: (optional), creates quadrilateral mesh, defaults to False
:kwarg reorder: (optional), should the mesh be reordered
The boundary edges in this mesh are numbered as follows:
* 1: plane x == 0
* 2: plane x == 1
* 3: plane y == 0
* 4: plane y == 1
"""
return SquareMesh(nx, ny, 1, reorder=reorder, quadrilateral=quadrilateral)
@profile
def CircleMesh(radius, resolution, reorder=None):
"""Generate a structured triangular mesh of a circle.
:arg radius: The radius of the circle.
:arg resolution: The number of cells lying along the radius and
the arc of the quadrant.
:kwarg reorder: (optional), should the mesh be reordered?
"""
source = """
lc = %g;
Point(1) = {0, -0.5, 0, lc};
Point(2) = {0, 0.5, 0, lc};
Line(1) = {1, 2};
surface[] = Extrude{{0, 0, %g},{0, 0, 0}, 0.9999 * Pi}{
Line{1};Layers{%d};
};
Physical Surface(2) = { surface[1] };
""" % (0.5 / resolution, radius, resolution * 4)
output = _get_msh_file(source, "circle_%g_%d" % (radius, resolution), 2)
return mesh.Mesh(output, reorder=reorder)
def UnitCircleMesh(resolution, reorder=None):
"""Generate a structured triangular mesh of a unit circle.
:arg resolution: The number of cells lying along the radius and
the arc of the quadrant.
:kwarg reorder: (optional), should the mesh be reordered?
"""
return CircleMesh(1.0, resolution, reorder=reorder)
def CircleManifoldMesh(ncells, radius=1):
"""Generated a 1D mesh of the circle, immersed in 2D.
:arg ncells: number of cells the circle should be
divided into (min 3)
:kwarg radius: (optional) radius of the circle to approximate
(defaults to 1).
"""
if ncells < 3:
raise ValueError("CircleManifoldMesh must have at least three cells")
vertices = radius*np.column_stack((np.cos(np.arange(ncells)*(2*np.pi/ncells)),
np.sin(np.arange(ncells)*(2*np.pi/ncells))))
cells = np.column_stack((np.arange(0, ncells, dtype=np.int32),
np.roll(np.arange(0, ncells, dtype=np.int32), -1)))
plex = _from_cell_list(1, cells, vertices)
return mesh.Mesh(plex, dim=2, reorder=False)
def UnitTetrahedronMesh():
"""Generate a mesh of the reference tetrahedron"""
coords = [[0., 0., 0.], [1., 0., 0.], [0., 1., 0.], [0., 0., 1.]]
cells = [[0, 1, 2, 3]]
plex = _from_cell_list(3, cells, coords)
return mesh.Mesh(plex, reorder=False)
@profile
def BoxMesh(nx, ny, nz, Lx, Ly, Lz, reorder=None):
"""Generate a mesh of a 3D box.
:arg nx: The number of cells in the x direction
:arg ny: The number of cells in the y direction
:arg nz: The number of cells in the z direction
:arg Lx: The extent in the x direction
:arg Ly: The extent in the y direction
:arg Lz: The extent in the z direction
:kwarg reorder: (optional), should the mesh be reordered?
The boundary surfaces are numbered as follows:
* 1: plane x == 0
* 2: plane x == Lx
* 3: plane y == 0
* 4: plane y == Ly
* 5: plane z == 0
* 6: plane z == Lz
"""
# Create mesh from DMPlex
boundary = PETSc.DMPlex().create(MPI.comm)
boundary.setDimension(2)
boundary.createCubeBoundary([0., 0., 0.], [Lx, Ly, Lz], [nx, ny, nz])
plex = PETSc.DMPlex().generate(boundary)
# Apply boundary IDs
plex.createLabel("boundary_ids")
plex.markBoundaryFaces("boundary_faces")
coords = plex.getCoordinates()
coord_sec = plex.getCoordinateSection()
if plex.getStratumSize("boundary_faces", 1) > 0:
boundary_faces = plex.getStratumIS("boundary_faces", 1).getIndices()
xtol = float(Lx)/(2*nx)
ytol = float(Ly)/(2*ny)
ztol = float(Lz)/(2*nz)
for face in boundary_faces:
face_coords = plex.vecGetClosure(coord_sec, coords, face)
if abs(face_coords[0]) < xtol and abs(face_coords[3]) < xtol and abs(face_coords[6]) < xtol:
plex.setLabelValue("boundary_ids", face, 1)
if abs(face_coords[0] - Lx) < xtol and abs(face_coords[3] - Lx) < xtol and abs(face_coords[6] - Lx) < xtol:
plex.setLabelValue("boundary_ids", face, 2)
if abs(face_coords[1]) < ytol and abs(face_coords[4]) < ytol and abs(face_coords[7]) < ytol:
plex.setLabelValue("boundary_ids", face, 3)
if abs(face_coords[1] - Ly) < ytol and abs(face_coords[4] - Ly) < ytol and abs(face_coords[7] - Ly) < ytol:
plex.setLabelValue("boundary_ids", face, 4)
if abs(face_coords[2]) < ztol and abs(face_coords[5]) < ztol and abs(face_coords[8]) < ztol:
plex.setLabelValue("boundary_ids", face, 5)
if abs(face_coords[2] - Lz) < ztol and abs(face_coords[5] - Lz) < ztol and abs(face_coords[8] - Lz) < ztol:
plex.setLabelValue("boundary_ids", face, 6)
return mesh.Mesh(plex, reorder=reorder)
def CubeMesh(nx, ny, nz, L, reorder=None):
"""Generate a mesh of a cube
:arg nx: The number of cells in the x direction
:arg ny: The number of cells in the y direction
:arg nz: The number of cells in the z direction
:arg L: The extent in the x, y and z directions
:kwarg reorder: (optional), should the mesh be reordered?
The boundary surfaces are numbered as follows:
* 1: plane x == 0
* 2: plane x == L
* 3: plane y == 0
* 4: plane y == L
* 5: plane z == 0
* 6: plane z == L
"""
return BoxMesh(nx, ny, nz, L, L, L, reorder=reorder)
def UnitCubeMesh(nx, ny, nz, reorder=None):
"""Generate a mesh of a unit cube
:arg nx: The number of cells in the x direction
:arg ny: The number of cells in the y direction
:arg nz: The number of cells in the z direction
:kwarg reorder: (optional), should the mesh be reordered?
The boundary surfaces are numbered as follows:
* 1: plane x == 0
* 2: plane x == 1
* 3: plane y == 0
* 4: plane y == 1
* 5: plane z == 0
* 6: plane z == 1
"""
return CubeMesh(nx, ny, nz, 1, reorder=reorder)
@profile
def IcosahedralSphereMesh(radius, refinement_level=0, reorder=None):
"""Generate an icosahedral approximation to the surface of the
sphere.
:arg radius: The radius of the sphere to approximate.
For a radius R the edge length of the underlying
icosahedron will be.
.. math::
a = \\frac{R}{\\sin(2 \\pi / 5)}
:kwarg refinement_level: optional number of refinements (0 is an
icosahedron).
:kwarg reorder: (optional), should the mesh be reordered?
"""
from math import sqrt
phi = (1 + sqrt(5)) / 2
# vertices of an icosahedron with an edge length of 2
vertices = np.array([[-1, phi, 0],
[1, phi, 0],
[-1, -phi, 0],
[1, -phi, 0],
[0, -1, phi],
[0, 1, phi],
[0, -1, -phi],
[0, 1, -phi],
[phi, 0, -1],
[phi, 0, 1],
[-phi, 0, -1],
[-phi, 0, 1]])
# faces of the base icosahedron
faces = np.array([[0, 11, 5],
[0, 5, 1],
[0, 1, 7],
[0, 7, 10],
[0, 10, 11],
[1, 5, 9],
[5, 11, 4],
[11, 10, 2],
[10, 7, 6],
[7, 1, 8],
[3, 9, 4],
[3, 4, 2],
[3, 2, 6],
[3, 6, 8],
[3, 8, 9],
[4, 9, 5],
[2, 4, 11],
[6, 2, 10],
[8, 6, 7],
[9, 8, 1]], dtype=np.int32)
plex = _from_cell_list(2, faces, vertices)
plex.setRefinementUniform(True)
for i in range(refinement_level):
plex = plex.refine()
vStart, vEnd = plex.getDepthStratum(0)
nvertices = vEnd - vStart
coords = plex.getCoordinatesLocal().array.reshape(nvertices, 3)
scale = (radius / np.linalg.norm(coords, axis=1)).reshape(-1, 1)
coords *= scale
return mesh.Mesh(plex, dim=3, reorder=reorder)
def UnitIcosahedralSphereMesh(refinement_level=0, reorder=None):
"""Generate an icosahedral approximation to the unit sphere.
:kwarg refinement_level: optional number of refinements (0 is an
icosahedron).
:kwarg reorder: (optional), should the mesh be reordered?
"""
return IcosahedralSphereMesh(1.0, refinement_level=refinement_level,
reorder=reorder)
<file_sep>/tests/extrusion/test_extrusion_5_helmholtz_vector.py
"""Tests for mixed Helmholtz convergence on extruded meshes"""
import numpy as np
import pytest
from firedrake import *
from tests.common import *
@pytest.mark.parametrize(('testcase', 'convrate'),
[(("RT", 1, "DG", 0, "h", "DG", 0, (2, 5)), 0.9),
(("RT", 2, "DG", 0, "h", "DG", 1, (2, 4)), 1.55),
(("RT", 3, "DG", 0, "h", "DG", 2, (2, 4)), 2.55),
(("BDM", 1, "DG", 0, "h", "DG", 0, (2, 5)), 0.9),
(("BDM", 2, "DG", 0, "h", "DG", 1, (2, 4)), 1.59),
(("BDFM", 2, "DG", 0, "h", "DG", 1, (2, 4)), 1.55),
(("DG", 0, "CG", 1, "v", "DG", 0, (2, 5)), 0.9),
(("DG", 0, "CG", 2, "v", "DG", 1, (2, 5)), 1.9)])
def test_scalar_convergence(testcase, convrate):
hfamily, hdegree, vfamily, vdegree, ori, altfamily, altdegree, (start, end) = testcase
l2err = np.zeros(end - start)
for ii in [i + start for i in range(len(l2err))]:
mesh = extmesh(2**ii, 2**ii, 2**ii)
horiz_elt = FiniteElement(hfamily, "triangle", hdegree)
vert_elt = FiniteElement(vfamily, "interval", vdegree)
product_elt = HDiv(OuterProductElement(horiz_elt, vert_elt))
V1 = FunctionSpace(mesh, product_elt)
if ori == "h":
# use same vertical variation, but different horizontal
# (think about product of complexes...)
horiz_elt = FiniteElement(altfamily, "triangle", altdegree)
vert_elt = FiniteElement(vfamily, "interval", vdegree)
elif ori == "v":
# opposite
horiz_elt = FiniteElement(hfamily, "triangle", hdegree)
vert_elt = FiniteElement(altfamily, "interval", altdegree)
product_elt = OuterProductElement(horiz_elt, vert_elt)
V2 = FunctionSpace(mesh, product_elt)
f = Function(V2)
exact = Function(V2)
if ori == "h":
f.interpolate(Expression("(1+8*pi*pi)*sin(x[0]*pi*2)*sin(x[1]*pi*2)"))
exact.interpolate(Expression("sin(x[0]*pi*2)*sin(x[1]*pi*2)"))
elif ori == "v":
f.interpolate(Expression("(1+4*pi*pi)*sin(x[2]*pi*2)"))
exact.interpolate(Expression("sin(x[2]*pi*2)"))
W = V1 * V2
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
a = (p*q - q*div(u) + dot(v, u) + div(v)*p)*dx
L = f*q*dx
out = Function(W)
solve(a == L, out, solver_parameters={'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'ksp_type': 'cg',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_0_ksp_type': 'cg',
'fieldsplit_1_ksp_type': 'cg'})
l2err[ii - start] = sqrt(assemble((out[3]-exact)*(out[3]-exact)*dx))
assert (np.array([np.log2(l2err[i]/l2err[i+1]) for i in range(len(l2err)-1)]) > convrate).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_bubble.py
"""Test bubble function space"""
import pytest
from firedrake import *
def test_simple():
mesh = UnitTriangleMesh()
V = FunctionSpace(mesh, "B", 3)
u = project(Expression("27*x[0]*x[1]*(1-x[0]-x[1])"), V)
assert (u.dat.data[0] - 1.0) < 1e-14
def test_enrichment():
mesh = UnitTriangleMesh()
P2 = FiniteElement("CG", "triangle", 2)
Bubble = FiniteElement("Bubble", "triangle", 3)
P2B3 = P2 + Bubble
V = FunctionSpace(mesh, P2B3)
W = FunctionSpace(mesh, "CG", 3)
u = project(Expression("27*x[0]*x[1]*(1-x[0]-x[1])"), V)
exact = Function(W)
exact.interpolate(Expression("27*x[0]*x[1]*(1-x[0]-x[1])"))
# make sure that these are the same
assert sqrt(assemble((u-exact)*(u-exact)*dx)) < 1e-14
def test_BDFM():
mesh = UnitTriangleMesh()
P2 = FiniteElement("CG", "triangle", 2)
Bubble = FiniteElement("Bubble", "triangle", 3)
P2B3 = P2 + Bubble
V0 = FunctionSpace(mesh, P2B3)
V1 = FunctionSpace(mesh, "BDFM", 2)
u = project(Expression("27*x[0]*x[1]*(1-x[0]-x[1])"), V0)
v = TrialFunction(V1)
w = TestFunction(V1)
out = Function(V1)
solve(dot(v, w)*dx == dot(curl(u), w)*dx, out)
# testing against known result where the interior DOFS of BDFM are excited
a = out.dat.data
a.sort()
assert (abs(a[1:7]) < 1e-12).all()
assert abs(a[0] + 6.75) < 1e-12
assert abs(a[7] - 6.75) < 1e-12
assert abs(a[8] - 13.5) < 1e-12
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/petsc.py
# Utility module that imports and initialises petsc4py
import petsc4py
import sys
petsc4py.init(sys.argv)
from petsc4py import PETSc # NOQA get flake8 to ignore unused import.
<file_sep>/tests/regression/test_mesh_generation.py
from math import pi
import pytest
from firedrake import *
# Must come after firedrake import (that loads MPI)
try:
import gmshpy
except ImportError:
gmshpy = None
def integrate_one(m):
V = FunctionSpace(m, 'CG', 1)
u = Function(V)
u.interpolate(Expression("1"))
return assemble(u * dx)
def test_unit_interval():
assert abs(integrate_one(UnitIntervalMesh(3)) - 1) < 1e-3
def test_interval():
assert abs(integrate_one(IntervalMesh(3, 5.0)) - 5.0) < 1e-3
def test_periodic_unit_interval():
assert abs(integrate_one(PeriodicUnitIntervalMesh(3)) - 1) < 1e-3
def test_periodic_interval():
assert abs(integrate_one(PeriodicIntervalMesh(3, 5.0)) - 5.0) < 1e-3
def test_unit_square():
assert abs(integrate_one(UnitSquareMesh(3, 3)) - 1) < 1e-3
def test_rectangle():
assert abs(integrate_one(RectangleMesh(3, 3, 10, 2)) - 20) < 1e-3
def test_unit_cube():
assert abs(integrate_one(UnitCubeMesh(3, 3, 3)) - 1) < 1e-3
def test_box():
assert abs(integrate_one(BoxMesh(3, 3, 3, 1, 2, 3)) - 6) < 1e-3
def test_unit_circle():
pytest.importorskip('gmshpy')
assert abs(integrate_one(UnitCircleMesh(4)) - pi * 0.5 ** 2) < 0.02
def test_unit_triangle():
assert abs(integrate_one(UnitTriangleMesh()) - 0.5) < 1e-3
def test_unit_tetrahedron():
assert abs(integrate_one(UnitTetrahedronMesh()) - 0.5 / 3) < 1e-3
@pytest.mark.parallel
def test_unit_interval_parallel():
assert abs(integrate_one(UnitIntervalMesh(30)) - 1) < 1e-3
@pytest.mark.parallel
def test_interval_parallel():
assert abs(integrate_one(IntervalMesh(30, 5.0)) - 5.0) < 1e-3
@pytest.mark.xfail(reason='Periodic intervals not implemented in parallel')
@pytest.mark.parallel
def test_periodic_unit_interval_parallel():
assert abs(integrate_one(PeriodicUnitIntervalMesh(30)) - 1) < 1e-3
@pytest.mark.xfail(reason='Periodic intervals not implemented in parallel')
@pytest.mark.parallel
def test_periodic_interval_parallel():
assert abs(integrate_one(PeriodicIntervalMesh(30, 5.0)) - 5.0) < 1e-3
@pytest.mark.parallel
def test_unit_square_parallel():
assert abs(integrate_one(UnitSquareMesh(5, 5)) - 1) < 1e-3
@pytest.mark.parallel
def test_unit_cube_parallel():
assert abs(integrate_one(UnitCubeMesh(3, 3, 3)) - 1) < 1e-3
@pytest.mark.skipif("gmshpy is None", reason='gmshpy not available')
@pytest.mark.parallel
def test_unit_circle_parallel():
assert abs(integrate_one(UnitCircleMesh(4)) - pi * 0.5 ** 2) < 0.02
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/demos/burgers/burgers.py.rst
Burgers equation
================
The Burgers equation is a non-linear equation for the advection and
diffusion of momentum. Here we choose to write the Burgers equation in
two dimensions to demonstrate the use of vector function spaces:
.. math::
\frac{\partial u}{\partial t} + (u\cdot\nabla) u - \nu\nabla^2 u = 0
(n\cdot \nabla) u = 0 \ \textrm{on}\ \Gamma
where :math:`\Gamma` is the domain boundary and :math:`\nu` is a
constant scalar viscosity. The solution :math:`u` is sought in some
suitable vector-valued function space :math:`V`. We take the inner
product with an arbitrary test function :math:`v\in V` and integrate
the viscosity term by parts:
.. math::
\int_\Omega\frac{\partial u}{\partial t}\cdot v +
((u\cdot\nabla) u)\cdot v + \nu\nabla u\cdot\nabla v \ \mathrm d x = 0.
The boundary condition has been used to discard the surface
integral. Next, we need to discretise in time. For simplicity and
stability we elect to use a backward Euler discretisation:
.. math::
\int_\Omega\frac{u^{n+1}-u^n}{dt}\cdot v +
((u^{n+1}\cdot\nabla) u^{n+1})\cdot v + \nu\nabla u^{n+1}\cdot\nabla v \ \mathrm d x = 0.
We can now proceed to set up the problem. We choose a resolution and set up a square mesh::
from firedrake import *
n = 30
mesh = UnitSquareMesh(n, n)
We choose degree 2 continuous Lagrange polynomials. We also need a
piecewise linear space for output purposes::
V = VectorFunctionSpace(mesh, "CG", 2)
V_out = VectorFunctionSpace(mesh, "CG", 1)
We also need solution functions for the current and the next
timestep. Note that, since this is a nonlinear problem, we don't
define trial functions::
u_ = Function(V, name="Velocity")
u = Function(V, name="VelocityNext")
v = TestFunction(V)
For this problem we need an initial condition::
ic = project(Expression(["sin(pi*x[0])", 0]), V)
We start with current value of u set to the initial condition, but we
also use the initial condition as our starting guess for the next
value of u::
u_.assign(ic)
u.assign(ic)
:math:`\nu` is set to a (fairly arbitrary) small constant value::
nu = 0.0001
The timestep is set to produce an advective Courant number of
around 1. Since we are employing backward Euler, this is stricter than
is required for stability, but ensures good temporal resolution of the
system's evolution::
timestep = 1.0/n
Here we finally get to define the residual of the equation. In the advection
term we need to contract the test function :math:`v` with
:math:`(u\cdot\nabla)u`, which is the derivative of the velocity in the
direction :math:`u`. This directional derivative can be written as
``dot(u,nabla_grad(u))`` since ``nabla_grad(u)[i,j]``:math:`=\partial_i u_j`.
Note once again that for a nonlinear problem, there are no trial functions in
the formulation. These will be created automatically when the residual
is differentiated by the nonlinear solver::
F = (inner((u - u_)/timestep, v)
+ inner(dot(u,nabla_grad(u)), v) + nu*inner(grad(u), grad(v)))*dx
To output the initial conditions, we project them into P1 and pipe
them into the output file::
outfile = File("burgers.pvd")
outfile << project(u, V_out, name="Velocity")
Finally, we loop over the timesteps solving the equation each time and
outputting each result::
t = 0.0
end = 0.5
while (t <= end):
solve(F == 0, u)
u_.assign(u)
t += timestep
outfile << project(u, V_out, name="Velocity")
A python script version of this demo can be found `here <burgers.py>`__.
<file_sep>/CONTRIBUTING.md
# Contributing to Firedrake
We value third-party contributions. To keep things simple for you and us,
please adhere to the following contributing guidelines. We are happy
to help if you have any [questions][4].
## Getting Started
* You will need a [GitHub account](https://github.com/signup/free).
* Submit a [ticket for your issue][0], assuming one does not already exist.
* Clearly describe the issue including steps to reproduce when it is a bug.
* Make sure you specify the version that you know has the issue.
* Bonus points for submitting a failing test along with the ticket.
* If you don't have push access, fork the repository on GitHub.
## Making Changes
* Create a topic branch for your feature or bug fix.
* Make commits of logical units.
* Make sure your commits adhere to the coding guidelines below.
* Make sure your commit messages are in the [proper format][1]: The first line
of the message should have 50 characters or less, separated by a blank line
from the (optional) body. The body should be wrapped at 70 characters and
paragraphs separated by blank lines. Bulleted lists are also fine.
* Make sure you have added the necessary tests for your changes.
* Run _all_ the tests to assure nothing else was accidentally broken.
## Coding guidelines
[PEP 0008][2] is enforced, with the exception of [E501][3] and [E226][3]:
* Indent by 4 spaces, tabs are *strictly forbidden*.
* Lines should not exceed 79 characters where possible without severely
impacting legibility. If breaking a line would make the code much less
readable it's fine to overrun by a little bit.
* No trailing whitespace at EOL or trailing blank lines at EOF.
## Checking your commit conforms to coding guidelines
Install a Git pre-commit hook automatically checking for tab and whitespace
errors before committing and also calls `flake8` on your changed files. In the
`.git/hooks` directory of your local Git repository, run the following:
```
git config --local core.whitespace "space-before-tab, tab-in-indent, trailing-space, tabwidth=4"
wget https://gist.github.com/kynan/d233073b66e860c41484/raw/pre-commit
chmod +x pre-commit
```
Make sure the `pre-commit.sample` hook is still in place, since it is required.
## Submitting Changes
* We can only accept your contribution if you have signed the Contributor
License Agreement (CLA).
* Push your changes to a topic branch in your fork of the repository.
* Submit a pull request to the repository in the firedrakeproject organization.
[0]: https://github.com/firedrakeproject/firedrake/issues
[1]: http://tbaggery.com/2008/04/19/a-note-about-git-commit-messages.html
[2]: http://www.python.org/dev/peps/pep-0008/
[3]: http://pep8.readthedocs.org/en/latest/intro.html#error-codes
[4]: http://www.firedrakeproject.org/contact.html
<file_sep>/tests/extrusion/test_enrichment_1_feec.py
"""Test curl-grad = 0 and div-curl = 0, using enriched function spaces"""
import pytest
from firedrake import *
from tests.common import *
@pytest.mark.parametrize(('horiz_complex', 'vert_complex'),
[((("CG", 1), ("RT", 1), ("DG", 0)), (("CG", 3), ("DG", 2))),
((("CG", 2), ("RT", 2), ("DG", 1)), (("CG", 2), ("DG", 1))),
((("CG", 3), ("RT", 3), ("DG", 2)), (("CG", 1), ("DG", 0))),
((("CG", 2), ("BDM", 1), ("DG", 0)), (("CG", 1), ("DG", 0))),
((("CG", 3), ("BDM", 2), ("DG", 1)), (("CG", 2), ("DG", 1))),
((("CG", 2, "B", 3), ("BDFM", 2), ("DG", 1)), (("CG", 2), ("DG", 1)))])
def test_feec(horiz_complex, vert_complex):
U0, U1, U2 = horiz_complex
V0, V1 = vert_complex
# U0, U1, U2 is our horizontal complex
# V0, V1 is be our vertical complex
# W0, W1, W2, W3 will be our product complex, where
# W0 = U0 x V0
# W1 = HCurl(U1 x V0) + HCurl(U0 x V1)
# W2 = HDiv(U2 x V0) + HDiv(U1 x V1)
# W3 = U2 x V1
mesh = extmesh(2, 2, 4)
if len(U0) == 2:
U0 = FiniteElement(U0[0], "triangle", U0[1])
else:
# make bubble space for BDFM
U0_a = FiniteElement(U0[0], "triangle", U0[1])
U0_b = FiniteElement(U0[2], "triangle", U0[3])
U0 = U0_a + U0_b
U1 = FiniteElement(U1[0], "triangle", U1[1])
U2 = FiniteElement(U2[0], "triangle", U2[1])
V0 = FiniteElement(V0[0], "interval", V0[1])
V1 = FiniteElement(V1[0], "interval", V1[1])
W0_elt = OuterProductElement(U0, V0)
W1_a = HCurl(OuterProductElement(U1, V0))
W1_b = HCurl(OuterProductElement(U0, V1))
W1_elt = W1_a + W1_b
W2_a = HDiv(OuterProductElement(U2, V0))
W2_b = HDiv(OuterProductElement(U1, V1))
W2_elt = W2_a + W2_b
W3_elt = OuterProductElement(U2, V1)
W0 = FunctionSpace(mesh, W0_elt)
W1 = FunctionSpace(mesh, W1_elt)
W2 = FunctionSpace(mesh, W2_elt)
W3 = FunctionSpace(mesh, W3_elt)
parms = {'snes_type': 'ksponly', 'ksp_type': 'preonly', 'pc_type': 'lu'}
### TEST CURL(GRAD(u)) = 0, for u in W0 ###
u = Function(W0)
u.interpolate(Expression("x[0]*x[1] - x[1]*x[2]"))
v1 = TrialFunction(W1)
v2 = TestFunction(W1)
a = dot(v1, v2)*dx
L = dot(grad(u), v2)*dx
v = Function(W1)
solve(a == L, v, solver_parameters=parms)
w1 = TrialFunction(W2)
w2 = TestFunction(W2)
a = dot(w1, w2)*dx
L = dot(curl(v), w2)*dx
w = Function(W2)
solve(a == L, w, solver_parameters=parms)
maxcoeff = max(abs(w.dat.data))
assert maxcoeff < 1e-11
### TEST DIV(CURL(v)) = 0, for v in W1 ###
v = project(Expression(("x[0]*x[1]", "-x[1]*x[2]", "x[0]*x[2]")), W1)
w1 = TrialFunction(W2)
w2 = TestFunction(W2)
a = dot(w1, w2)*dx
L = dot(curl(v), w2)*dx
w = Function(W2)
solve(a == L, w, solver_parameters=parms)
y1 = TrialFunction(W3)
y2 = TestFunction(W3)
a = y1*y2*dx
L = div(w)*y2*dx
y = Function(W3)
solve(a == L, y, solver_parameters=parms)
maxcoeff = max(abs(y.dat.data))
assert maxcoeff < 1e-11
### TEST WEAKCURL(WEAKGRAD(y)) = 0, for y in W3 ###
y = Function(W3)
y.interpolate(Expression("x[0]*x[1] - x[1]*x[2]"))
w1 = TrialFunction(W2)
w2 = TestFunction(W2)
a = dot(w1, w2)*dx
L = -y*div(w2)*dx
w = Function(W2)
solve(a == L, w, solver_parameters=parms)
v1 = TrialFunction(W1)
v2 = TestFunction(W1)
a = dot(v1, v2)*dx
L = -dot(w, curl(v2))*dx
v = Function(W1)
solve(a == L, v, solver_parameters=parms)
maxcoeff = max(abs(v.dat.data))
assert maxcoeff < 1e-11
### TEST WEAKDIV(WEAKCURL(w)) = 0, for w in W2 ###
w = project(Expression(("x[0]*x[1]", "-x[1]*x[2]", "x[0]*x[2]")), W2)
v1 = TrialFunction(W1)
v2 = TestFunction(W1)
a = dot(v1, v2)*dx
L = -dot(w, curl(v2))*dx
v = Function(W1)
solve(a == L, v, solver_parameters=parms)
u1 = TrialFunction(W0)
u2 = TestFunction(W0)
a = dot(u1, u2)*dx
L = -dot(v, grad(u2))*dx
u = Function(W0)
solve(a == L, u, solver_parameters=parms)
maxcoeff = max(abs(u.dat.data))
assert maxcoeff < 3e-11
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/docs/source/documentation.rst
.. only:: html
.. sidebar:: Current development information.
Firedrake and PyOP2 are continually tested using a `local buildbot
<http://buildbot-ocean.ese.ic.ac.uk:8080/builders/firedrake-trunk>`__
and the `Travis continuous integration system
<https://travis-ci.org>`__ respectively.
Latest Firedrake status can be found `here
<http://buildbot-ocean.ese.ic.ac.uk:8080/builders/firedrake-trunk>`__.
Latest PyOP2 status: |pyop2build|
.. |pyop2build| image:: https://travis-ci.org/OP2/PyOP2.png?branch=master
:target: https://travis-ci.org/OP2/PyOP2
Firedrake and PyOP2 are developed on `GitHub <http://github.com>`__ while we also maintain Firedrake-ready versions of the `FEniCS <http://fenicsproject.org>`__ tools on `Bitbucket <http://bitbucket.org>`__.
* `Firedrake on GitHub <https://github.com/firedrakeproject/firedrake/>`__
* `PyOp2 on GitHub <https://github.com/OP2/PyOP2>`__
* `Firedrake version of FFC on Bitbucket <https://bitbucket.org/mapdes/ffc>`__
* `Firedrake version of UFL on Bitbucket <https://bitbucket.org/mapdes/ufl>`__
* `Firedrake version of FIAT on Bitbucket <https://bitbucket.org/mapdes/fiat>`__
Getting started
===============
The first step is to download and install Firedrake and its
dependencies. For full instructions, see :doc:`obtaining Firedrake
<download>`.
.. _firedrake_tutorials:
Tutorials
=========
Once you've built Firedrake, you'll want to actually solve some
PDEs. Below are a few tutorial examples to get you started.
.. toctree::
A basic Helmholtz equation.<demos/helmholtz.py>
The Burgers equation, a non-linear, unsteady example.<demos/burgers.py>
A mixed formulation of the Poisson equation.<demos/poisson_mixed.py>
A steady-state advection equation using upwinding, on an extruded mesh.<demos/upwind_advection.py>
Manual
======
Once you have worked through the tutorials, the user manual is the
next step. It goes in to more detail on how to set up and solve
finite element problems in Firedrake.
.. toctree::
:maxdepth: 2
variational-problems
solving-interface
boundary_conditions
extruded-meshes
.. only:: html
API documentation
=================
The complete list of all the classes and methods in Firedrake is
available at the :doc:`firedrake` page. The same information is
:ref:`indexed <genindex>` in alphabetical order. Another very
effective mechanism is the site :ref:`search engine <search>`.
<file_sep>/tests/regression/test_piola_mixed_fn.py
import numpy as np
import pytest
from firedrake import *
from tests.common import *
def test_project(mesh):
U = FunctionSpace(mesh, "RT", 1)
V = FunctionSpace(mesh, "N1curl", 1)
W = U*V
f = Function(W)
f.assign(1)
out = Function(W)
u1, u2 = TrialFunctions(W)
v1, v2 = TestFunctions(W)
f1, f2 = split(f)
a = dot(u1, v1)*dx + dot(u2, v2)*dx
L = dot(f1, v1)*dx + dot(f2, v2)*dx
solve(a == L, out)
assert np.allclose(out.dat.data[0], f.dat.data[0], rtol=1e-5)
assert np.allclose(out.dat.data[1], f.dat.data[1], rtol=1e-5)
def test_sphere_project():
mesh = UnitIcosahedralSphereMesh(0)
mesh.init_cell_orientations(Expression(("x[0]", "x[1]", "x[2]")))
U1 = FunctionSpace(mesh, "RT", 1)
U2 = FunctionSpace(mesh, "CG", 2)
U3 = FunctionSpace(mesh, "N1curl", 1)
W = U1*U2*U3
f = Function(W)
f1, f2, f3 = f.split()
f1.assign(1)
f2.assign(2)
f3.assign(3)
out = Function(W)
u1, u2, u3 = TrialFunctions(W)
v1, v2, v3 = TestFunctions(W)
f1, f2, f3 = split(f)
a = dot(u1, v1)*dx + dot(u2, v2)*dx + dot(u3, v3)*dx
L = dot(f1, v1)*dx + dot(f2, v2)*dx + dot(f3, v3)*dx
solve(a == L, out)
assert np.allclose(out.dat.data[0], f.dat.data[0], rtol=1e-5)
assert np.allclose(out.dat.data[1], f.dat.data[1], rtol=1e-5)
assert np.allclose(out.dat.data[2], f.dat.data[2], rtol=1e-5)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_vector_laplace_on_quadrilaterals.py
import numpy as np
import pytest
from firedrake import *
def vector_laplace(n, degree):
mesh = UnitSquareMesh(n, n, quadrilateral=True)
S0 = FiniteElement("CG", interval, degree)
S1 = FiniteElement("DG", interval, degree - 1)
T0 = FiniteElement("CG", interval, degree)
T1 = FiniteElement("DG", interval, degree - 1)
V0_elt = OuterProductElement(S0, T0)
V1_elt_h = HCurl(OuterProductElement(S1, T0))
V1_elt_v = HCurl(OuterProductElement(S0, T1))
V1_elt = V1_elt_h + V1_elt_v
# spaces for calculation
V0 = FunctionSpace(mesh, V0_elt)
V1 = FunctionSpace(mesh, V1_elt)
V = V0*V1
# spaces to store 'analytic' functions
W0 = FunctionSpace(mesh, "CG", degree + 1)
W1 = VectorFunctionSpace(mesh, "CG", degree + 1)
# constants
k = 1.0
l = 2.0
f_expr = Expression(("pi*pi*(kk*kk + ll*ll)*sin(kk*pi*x[0])*cos(ll*pi*x[1])", "pi*pi*(kk*kk + ll*ll)*cos(kk*pi*x[0])*sin(ll*pi*x[1])"), kk=k, ll=l)
exact_s_expr = Expression("-(kk+ll)*pi*cos(kk*pi*x[0])*cos(ll*pi*x[1])", kk=k, ll=l)
exact_u_expr = Expression(("sin(kk*pi*x[0])*cos(ll*pi*x[1])", "cos(kk*pi*x[0])*sin(ll*pi*x[1])"), kk=k, ll=l)
f = Function(W1).interpolate(f_expr)
exact_s = Function(W0).interpolate(exact_s_expr)
exact_u = Function(W1).interpolate(exact_u_expr)
sigma, u = TrialFunctions(V)
tau, v = TestFunctions(V)
a = (sigma*tau - dot(u, grad(tau)) + dot(grad(sigma), v) + dot(curl(u), curl(v)))*dx
L = dot(f, v)*dx
out = Function(V)
# preconditioner for H1 x H(curl)
aP = (dot(grad(sigma), grad(tau)) + sigma*tau + dot(curl(u), curl(v)) + dot(u, v))*dx
solve(a == L, out, Jp=aP,
solver_parameters={'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'additive',
'fieldsplit_0_pc_type': 'lu',
'fieldsplit_1_pc_type': 'lu',
'ksp_monitor': True})
out_s, out_u = out.split()
return (sqrt(assemble(dot(out_u - exact_u, out_u - exact_u)*dx)),
sqrt(assemble((out_s - exact_s)*(out_s - exact_s)*dx)))
@pytest.mark.parametrize(('testcase', 'convrate'),
[((1, (2, 4)), 0.9),
((2, (2, 4)), 1.9),
((3, (2, 4)), 2.9),
((4, (2, 4)), 3.9)])
def test_hcurl_convergence(testcase, convrate):
degree, (start, end) = testcase
l2err = np.zeros((end - start, 2))
for ii in [i + start for i in range(len(l2err))]:
l2err[ii - start, :] = vector_laplace(2 ** ii, degree)
assert (np.log2(l2err[:-1, :] / l2err[1:, :]) > convrate).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_manifolds.py
from firedrake import *
import pytest
import numpy as np
from os.path import abspath, dirname, join
cwd = abspath(dirname(__file__))
# This test solves a mixed formulation of the Poisson equation with
# inhomogeneous Neumann boundary conditions such that the exact
# solution is p(x, y) = x - 0.5. First on a 2D mesh, and then again
# on a 2D mesh embedded in 3D.
def run_no_manifold():
mesh = UnitSquareMesh(1, 1)
V0 = FunctionSpace(mesh, "RT", 2)
V1 = FunctionSpace(mesh, "DG", 1)
V = V0 * V1
bc = DirichletBC(V.sub(0), (-1, 0), (1, 2, 3, 4))
u, p = TrialFunctions(V)
v, q = TestFunctions(V)
a = (dot(u, v) - p*div(v) - div(u)*q)*dx
f = Function(V1)
f.assign(0)
L = -f*q*dx
up = Function(V)
null_vec = Function(V)
null_vec.dat[1].data[:] = 1/sqrt(V1.dof_count)
nullspace = VectorSpaceBasis(vecs=[null_vec])
solve(a == L, up, bcs=bc, nullspace=nullspace)
exact = Function(V1).interpolate(Expression('x[0] - 0.5'))
u, p = up.split()
assert errornorm(exact, p, degree_rise=0) < 1e-8
def run_manifold():
mesh = Mesh(join(cwd, "unitsquare_in_3d.node"), dim=3)
mesh.init_cell_orientations(Expression(('0', '0', '1')))
V0 = FunctionSpace(mesh, "RT", 2)
V1 = FunctionSpace(mesh, "DG", 1)
V = V0 * V1
bc = DirichletBC(V.sub(0), (-1, 0, 0), (1, 2, 3, 4))
u, p = TrialFunctions(V)
v, q = TestFunctions(V)
a = (dot(u, v) - p*div(v) - div(u)*q)*dx
f = Function(V1)
f.assign(0)
L = -f*q*dx
up = Function(V)
null_vec = Function(V)
null_vec.dat[1].data[:] = 1/sqrt(V1.dof_count)
nullspace = VectorSpaceBasis(vecs=[null_vec])
solve(a == L, up, bcs=bc, nullspace=nullspace)
exact = Function(V1).interpolate(Expression('x[0] - 0.5'))
u, p = up.split()
assert errornorm(exact, p, degree_rise=0) < 1e-8
def test_no_manifold_serial():
run_no_manifold()
def test_manifold_serial():
run_manifold()
@pytest.mark.parallel(nprocs=2)
def test_no_manifold_parallel():
run_no_manifold()
@pytest.mark.parallel(nprocs=2)
def test_manifold_parallel():
run_manifold()
@pytest.mark.parametrize('space', ["RT", "BDM"])
def test_contravariant_piola_facet_integral(space):
mesh = UnitIcosahedralSphereMesh(refinement_level=2)
global_normal = Expression(("x[0]", "x[1]", "x[2]"))
mesh.init_cell_orientations(global_normal)
V = FunctionSpace(mesh, space, 1)
# Some non-zero function
u = project(Expression(('x[0]', '-x[1]', '0')), V)
n = FacetNormal(mesh)
pos = inner(u('+'), n('+'))*dS
neg = inner(u('-'), n('-'))*dS
assert np.allclose(assemble(pos) + assemble(neg), 0)
assert np.allclose(assemble(pos + neg), 0)
@pytest.mark.parametrize('space', ["N1curl", "N2curl"])
def test_covariant_piola_facet_integral(space):
mesh = UnitIcosahedralSphereMesh(refinement_level=2)
global_normal = Expression(("x[0]", "x[1]", "x[2]"))
mesh.init_cell_orientations(global_normal)
V = FunctionSpace(mesh, space, 1)
# Some non-zero function
u = project(Expression(('x[0]', '-x[1]', '0')), V)
n = FacetNormal(mesh)
pos = inner(u('+'), n('+'))*dS
neg = inner(u('-'), n('-'))*dS
assert np.allclose(assemble(pos) + assemble(neg), 0, atol=1e-7)
assert np.allclose(assemble(pos + neg), 0, atol=1e-7)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_split.py
import pytest
from firedrake import *
def test_assemble_split_derivative():
"""Assemble the derivative of a form with a zero block."""
mesh = UnitSquareMesh(1, 1)
V1 = FunctionSpace(mesh, "BDM", 1, name="V")
V2 = FunctionSpace(mesh, "DG", 0, name="P")
W = V1 * V2
x = Function(W)
u, p = split(x)
v, q = TestFunctions(W)
F = (inner(u, v) + v[1]*p)*dx
assert assemble(derivative(F, x))
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_poisson_strong_bcs_nitsche.py
# coding=utf-8
"""
Solve
- div grad u(x, y) = 0
with u(0, y) = u_0 = 0 (\Gamma_0)
u(1, y) = u_1 = 42 (\Gamma_1)
and du/dn = 0 on the other two sides
we impose the strong boundary conditions weakly using Nitsche's method:
<NAME>, Über ein Variationsprinzip zur Lösung von
Dirichlet-Problemen bei Verwendung von Teilräumen, die keinen
Randbedingungen unterworfen sind. Abh. Math. Sem. Univ. Hamburg 36
(1971), 9–15. (http://www.ams.org/mathscinet-getitem?mr=0341903)
In particular we follow the method described in:
<NAME> and <NAME>, Nitsche's method for general boundary
conditions. Mathematics of Computation 78(267):1353-1374 (2009)
That is, on \Gamma_0 we impose
du/dn = 1/\epsilon (u - u_0)
and on \Gamma_1
du/dn = 1/\epsilon (u - u_1)
and take \lim_{\epsilon \rightarrow 0}
"""
import pytest
from firedrake import *
def run_test(x, degree):
mesh = UnitSquareMesh(2 ** x, 2 ** x)
V = FunctionSpace(mesh, "CG", degree)
u = TrialFunction(V)
v = TestFunction(V)
a = dot(grad(v), grad(u)) * dx
f = Function(V)
f.assign(0)
L = v*f*dx
# This value of the stabilisation parameter gets us about 4 sf
# accuracy.
h = 0.25
gamma = 0.00001
n = FacetNormal(mesh)
B = a - \
inner(dot(grad(u), n), v)*(ds(3) + ds(4)) - \
inner(u, dot(grad(v), n))*(ds(3) + ds(4)) + \
(1.0/(h*gamma))*u*v*(ds(3) + ds(4))
u_0 = Function(V)
u_0.assign(0)
u_1 = Function(V)
u_1.assign(42)
F = L - \
inner(u_0, dot(grad(v), n))*ds(3) - \
inner(u_1, dot(grad(v), n))*ds(4) + \
(1.0/(h*gamma))*u_0*v*ds(3) + \
(1.0/(h*gamma))*u_1*v*ds(4)
u = Function(V)
solve(B == F, u)
f = Function(V)
f.interpolate(Expression("42*x[1]"))
return sqrt(assemble(dot(u - f, u - f)*dx))
@pytest.mark.parametrize('degree', (1, 2))
def test_poisson_nitsche(degree):
assert run_test(2, degree) < 1e-3
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_mixed_interior_facets.py
import pytest
from firedrake import *
@pytest.fixture(scope='module')
def mesh2D():
# .---.
# |\ |
# | \ |
# | \|
# '---'
return UnitSquareMesh(1, 1)
@pytest.mark.parametrize('degree', [1, 2, 3])
def test_vfs(mesh2D, degree):
V = VectorFunctionSpace(mesh2D, 'CG', degree)
u = Function(V)
u.interpolate(Expression(('1.0', '1.0')))
n = FacetNormal(mesh2D)
# Unit '+' normal is (1, 1)/sqrt2, and diagonal has length sqrt2.
assert abs(assemble(dot(u('-'), n('-'))*dS) + 2.0) < 1e-10
assert abs(assemble(dot(u('+'), n('-'))*dS) + 2.0) < 1e-10
assert abs(assemble(dot(u('+'), n('+'))*dS) - 2.0) < 1e-10
u.interpolate(Expression(('1.0', '-1.0')))
assert abs(assemble(dot(u('+'), n('+'))*dS)) < 1e-10
def test_mfs(mesh2D):
V1 = FunctionSpace(mesh2D, 'BDM', 1)
V2 = FunctionSpace(mesh2D, 'CG', 2)
V3 = FunctionSpace(mesh2D, 'CG', 3)
W = V3 * V1 * V2
u = project(Expression(('1.0', '-1.0', '-1.0', '1.0')), W)
n = FacetNormal(mesh2D)
# Unit '+' normal is (1, 1)/sqrt2, and diagonal has length sqrt2.
# This is (dot((1, 1), n+) + 10*dot((-1, -1), n-)) * dS = 2 + 20 = 22
a = (u[0]('+')*n[0]('+') + u[3]('-')*n[1]('+')
+ 10*u[1]('+')*n[0]('-') + 10*u[2]('-')*n[1]('-'))*dS
assert abs(assemble(a) - 22.0) < 1e-9
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_expressions.py
from operator import iadd, isub, imul, idiv
from functools import partial
from itertools import permutations
import pytest
import numpy as np
import ufl # noqa: used in eval'd expressions
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module', params=['cg1', 'cg1cg1[0]', 'cg1cg1[1]',
'cg1vcg1[0]', 'cg1dg0[0]', 'cg1dg0[1]',
'cg2dg1[0]', 'cg2dg1[1]'])
def sfs(request, cg1, cg1cg1, cg1vcg1, cg1dg0, cg2dg1):
"""A parametrized fixture for scalar function spaces."""
return {'cg1': cg1,
'cg1cg1[0]': cg1cg1[0],
'cg1cg1[1]': cg1cg1[1],
'cg1vcg1[0]': cg1vcg1[0],
'cg1dg0[0]': cg1dg0[0],
'cg1dg0[1]': cg1dg0[1],
'cg2dg1[0]': cg2dg1[0],
'cg2dg1[1]': cg2dg1[1]}[request.param]
@pytest.fixture(scope='module', params=['vcg1', 'cg1vcg1[1]'])
def vfs(request, vcg1, cg1vcg1):
"""A parametrized fixture for vector function spaces."""
return {'vcg1': vcg1,
'cg1vcg1[1]': cg1vcg1[1]}[request.param]
@pytest.fixture(scope='module', params=['cg1cg1', 'cg1vcg1', 'cg1dg0', 'cg2dg1'])
def mfs(request, cg1cg1, cg1vcg1, cg1dg0, cg2dg1):
"""A parametrized fixture for mixed function spaces."""
return {'cg1cg1': cg1cg1,
'cg1vcg1': cg1vcg1,
'cg1dg0': cg1dg0,
'cg2dg1': cg2dg1}[request.param]
def func_factory(fs):
f = Function(fs, name="f")
one = Function(fs, name="one").assign(1)
two = Function(fs, name="two").assign(2)
minusthree = Function(fs, name="minusthree").assign(-3)
return f, one, two, minusthree
@pytest.fixture()
def functions(request, sfs):
return func_factory(sfs)
@pytest.fixture()
def vfunctions(request, vfs):
return func_factory(vfs)
@pytest.fixture()
def mfunctions(request, mfs):
return func_factory(mfs)
@pytest.fixture
def msfunctions(request, mfs):
return Function(mfs), Function(mfs[0]).assign(1), Function(mfs[1]).assign(2)
@pytest.fixture
def sf(cg1):
return Function(cg1, name="sf")
@pytest.fixture
def vf(vcg1):
return Function(vcg1, name="vf")
@pytest.fixture
def mf(cg1, vcg1):
return Function(cg1 * vcg1, name="mf")
@pytest.fixture(params=permutations(['sf', 'vf', 'mf'], 2))
def fs_combinations(sf, vf, mf, request):
funcs = {'sf': sf, 'vf': vf, 'mf': mf}
return [funcs[p] for p in request.param]
def evaluate(v, x):
try:
assert len(v) == len(x)
except TypeError:
x = (x,) * len(v)
try:
return all(np.all(v_ == x_) for v_, x_ in zip(v, x))
except:
return v == x
def ioptest(f, expr, x, op):
return evaluate(op(f, expr).dat.data, x)
def interpolatetest(f, expr, x):
if f.function_space().cdim > 1:
expr = (expr,) * f.function_space().cdim
return evaluate(f.interpolate(Expression(expr)).dat.data, x)
exprtest = lambda expr, x: evaluate(assemble(expr).dat.data, x)
assigntest = lambda f, expr, x: evaluate(f.assign(expr).dat.data, x)
iaddtest = partial(ioptest, op=iadd)
isubtest = partial(ioptest, op=isub)
imultest = partial(ioptest, op=imul)
idivtest = partial(ioptest, op=idiv)
common_tests = [
'assigntest(f, 1, 1)',
'exprtest(one + one, 2)',
'exprtest(3 * one, 3)',
'exprtest(one + two, 3)',
'assigntest(f, one + two, 3)',
'iaddtest(one, one, 2)',
'iaddtest(one, two, 3)',
'iaddtest(f, 2, 2)',
'isubtest(two, 1, 1)',
'imultest(one, 2, 2)',
'imultest(one, two, 2)',
'idivtest(two, 2, 1)',
'idivtest(one, two, 0.5)',
'isubtest(one, one, 0)',
'assigntest(f, 2 * one, 2)',
'assigntest(f, one - one, 0)']
scalar_tests = common_tests + [
'interpolatetest(f, 0.0, 0)',
'interpolatetest(f, "sin(pi/2)", 1)',
'exprtest(ufl.ln(one), 0)',
'exprtest(two ** minusthree, 0.125)',
'exprtest(ufl.sign(minusthree), -1)',
'exprtest(one + two / two ** minusthree, 17)']
mixed_tests = common_tests + [
'interpolatetest(f, "sin(pi/2)", (1, 1))',
'exprtest(one[0] + one[1], (1, 1))',
'exprtest(one[1] + two[0], (2, 1))',
'exprtest(one[0] - one[1], (1, -1))',
'exprtest(one[1] - two[0], (-2, 1))',
'assigntest(f, one[0], (1, 0))',
'assigntest(f, one[1], (0, 1))',
'assigntest(two, one[0], (1, 0))',
'assigntest(two, one[1], (0, 1))',
'assigntest(two, one[0] + two[0], (3, 0))',
'assigntest(two, two[1] - one[1], (0, 1))',
'assigntest(f, one[0] + two[1], (1, 2))',
'iaddtest(one, one[0], (2, 1))',
'iaddtest(one, one[1], (1, 2))',
'assigntest(f, 2 * two[1] + 2 * minusthree[0], (-6, 4))']
indexed_fs_tests = [
'assigntest(f, one, (1, 0))',
'assigntest(f, two, (0, 2))',
'iaddtest(f, one, (1, 0))',
'iaddtest(f, two, (0, 2))',
'isubtest(f, one, (-1, 0))',
'isubtest(f, two, (0, -2))']
@pytest.mark.parametrize('expr', scalar_tests)
def test_scalar_expressions(expr, functions):
f, one, two, minusthree = functions
assert eval(expr)
@pytest.mark.parametrize('expr', common_tests)
def test_vector_expressions(expr, vfunctions):
f, one, two, minusthree = vfunctions
assert eval(expr)
@pytest.mark.parametrize('expr', mixed_tests)
def test_mixed_expressions(expr, mfunctions):
f, one, two, minusthree = mfunctions
assert eval(expr)
@pytest.mark.parametrize('expr', indexed_fs_tests)
def test_mixed_expressions_indexed_fs(expr, msfunctions):
f, one, two = msfunctions
assert eval(expr)
def test_different_fs_asign_fails(fs_combinations):
"""Assigning to a Function on a different function space should raise
ValueError."""
f1, f2 = fs_combinations
with pytest.raises(ValueError):
f1.assign(f2)
def test_asign_to_nonindexed_subspace_fails(mfs):
"""Assigning a Function on a non-indexed sub space of a mixed function
space to a function on the mixed function space should fail."""
for fs in mfs:
with pytest.raises(ValueError):
Function(mfs).assign(Function(fs._fs))
def test_assign_mixed_no_nan(mfs):
w = Function(mfs)
vs = w.split()
vs[0].assign(2)
w /= vs[0]
assert np.allclose(vs[0].dat.data_ro, 1.0)
for v in vs[1:]:
assert not np.isnan(v.dat.data_ro).any()
def test_assign_mixed_no_zero(mfs):
w = Function(mfs)
vs = w.split()
w.assign(2)
w *= vs[0]
assert np.allclose(vs[0].dat.data_ro, 4.0)
for v in vs[1:]:
assert np.allclose(v.dat.data_ro, 2.0)
def test_assign_to_mfs_sub(cg1, vcg1):
W = cg1*vcg1
w = Function(W)
u = Function(cg1)
v = Function(vcg1)
u.assign(4)
v.assign(10)
w.sub(0).assign(u)
assert np.allclose(w.sub(0).dat.data_ro, 4)
assert np.allclose(w.sub(1).dat.data_ro, 0)
w.sub(1).assign(v)
assert np.allclose(w.sub(0).dat.data_ro, 4)
assert np.allclose(w.sub(1).dat.data_ro, 10)
Q = vcg1*cg1
q = Function(Q)
q.assign(11)
w.sub(1).assign(q.sub(0))
assert np.allclose(w.sub(1).dat.data_ro, 11)
assert np.allclose(w.sub(0).dat.data_ro, 4)
with pytest.raises(ValueError):
w.sub(1).assign(q.sub(1))
with pytest.raises(ValueError):
w.sub(1).assign(w.sub(0))
with pytest.raises(ValueError):
w.sub(1).assign(u)
with pytest.raises(ValueError):
w.sub(0).assign(v)
w.sub(0).assign(ufl.ln(q.sub(1)))
assert np.allclose(w.sub(0).dat.data_ro, ufl.ln(11))
with pytest.raises(ValueError):
w.assign(q.sub(1))
def test_assign_from_mfs_sub(cg1, vcg1):
W = cg1*vcg1
w = Function(W)
u = Function(cg1)
v = Function(vcg1)
w1, w2 = w.split()
w1.assign(4)
w2.assign(10)
u.assign(w1)
assert np.allclose(u.dat.data_ro, w1.dat.data_ro)
v.assign(w2)
assert np.allclose(v.dat.data_ro, w2.dat.data_ro)
Q = vcg1*cg1
q = Function(Q)
q1, q2 = q.split()
q1.assign(11)
q2.assign(12)
v.assign(q1)
assert np.allclose(v.dat.data_ro, q1.dat.data_ro)
u.assign(q2)
assert np.allclose(u.dat.data_ro, q2.dat.data_ro)
with pytest.raises(ValueError):
u.assign(q1)
with pytest.raises(ValueError):
v.assign(q2)
with pytest.raises(ValueError):
u.assign(w2)
with pytest.raises(ValueError):
v.assign(w1)
@pytest.mark.parametrize("uservar", ["A", "X", "x_", "k", "d", "i"])
def test_scalar_user_defined_values(uservar):
m = UnitSquareMesh(2, 2)
V = FunctionSpace(m, 'CG', 1)
f = Function(V)
e = Expression(uservar, **{uservar: 1.0})
f.interpolate(e)
assert np.allclose(f.dat.data_ro, 1.0)
setattr(e, uservar, 2.0)
f.interpolate(e)
assert np.allclose(f.dat.data_ro, 2.0)
def test_vector_user_defined_values():
m = UnitSquareMesh(2, 2)
V = FunctionSpace(m, 'CG', 1)
f = Function(V)
e = Expression('n[0] + n[1]', n=[1.0, 2.0])
f.interpolate(e)
assert np.allclose(f.dat.data_ro, 3.0)
e.n = [2.0, 4.0]
f.interpolate(e)
assert np.allclose(f.dat.data_ro, 6.0)
def test_scalar_increment_fails():
e = Expression('n', n=1.0)
# Some versions of numpy raise RuntimeError on access to read-only
# array view, rather than ValueError.
with pytest.raises((ValueError, RuntimeError)):
e.n += 1
with pytest.raises((ValueError, RuntimeError)):
e.n[0] += 2
assert np.allclose(e.n, 1.0)
def test_vector_increment_fails():
e = Expression('n', n=[1.0, 1.0])
with pytest.raises((ValueError, RuntimeError)):
e.n += 1
with pytest.raises((ValueError, RuntimeError)):
e.n[0] += 2
assert np.allclose(e.n, 1.0)
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/extrusion/test_poisson_neumann.py
"""This demo program solves Poisson's equation
- div grad u(x, y, z) = f
in a unit cube, with Dirichlet boundary conditions on 4 sides
and Neumann boundary conditions (one explicit, one implicit)
on the other 2, opposite, sides.
"""
import pytest
from firedrake import *
from tests.common import *
@pytest.fixture(scope='module')
def P2():
mesh = extmesh(4, 4, 4)
return FunctionSpace(mesh, "CG", 2)
def test_bottom(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = -20*v*dx + 20*v*ds_b
bc_expr = Expression("10*(x[2]-1)*(x[2]-1)")
bcs = [DirichletBC(P2, bc_expr, 1),
DirichletBC(P2, bc_expr, 2),
DirichletBC(P2, bc_expr, 3),
DirichletBC(P2, bc_expr, 4)]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.0e-6
def test_top(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = -20*v*dx + 20*v*ds_t
bc_expr = Expression("10*x[2]*x[2]")
bcs = [DirichletBC(P2, bc_expr, 1),
DirichletBC(P2, bc_expr, 2),
DirichletBC(P2, bc_expr, 3),
DirichletBC(P2, bc_expr, 4)]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.0e-6
def test_topbottom(P2):
u = TrialFunction(P2)
v = TestFunction(P2)
a = dot(grad(u), grad(v))*dx
L = -20*v*dx + 10*v*ds_tb
bc_expr = Expression("10*(x[2]-0.5)*(x[2]-0.5)")
bcs = [DirichletBC(P2, bc_expr, 1),
DirichletBC(P2, bc_expr, 2),
DirichletBC(P2, bc_expr, 3),
DirichletBC(P2, bc_expr, 4)]
u = Function(P2)
solve(a == L, u, bcs)
u_exact = Function(P2)
u_exact.interpolate(bc_expr)
assert max(abs(u.dat.data - u_exact.dat.data)) < 1.0e-6
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_fs_caching.py
import pytest
from firedrake import *
from firedrake.mesh import MeshBase
import gc
def howmany(cls):
return len([x for x in gc.get_objects() if isinstance(x, cls)])
def test_same_fs_hits_cache():
m = UnitSquareMesh(1, 1)
V1 = FunctionSpace(m, 'CG', 2)
V2 = FunctionSpace(m, 'CG', 2)
assert V1 is V2
def test_different_fs_misses_cache():
m = UnitSquareMesh(1, 1)
V1 = FunctionSpace(m, 'CG', 2)
V2 = FunctionSpace(m, 'DG', 2)
assert V1 is not V2
def test_extruded_fs_hits_cache():
m = UnitSquareMesh(1, 1)
e = ExtrudedMesh(m, 2, layer_height=1)
V1 = FunctionSpace(e, 'CG', 1)
V2 = FunctionSpace(e, 'CG', 1)
assert V1 is V2
assert V1 not in m._cache.values()
assert V1 in e._cache.values()
def test_extruded_fs_misses_cache():
m = UnitSquareMesh(1, 1)
e = ExtrudedMesh(m, 2, layer_height=1)
V1 = FunctionSpace(e, 'CG', 1)
V2 = FunctionSpace(e, 'DG', 1)
assert V1 is not V2
def test_extruded_ope_hits_cache():
m = UnitSquareMesh(1, 1)
e = ExtrudedMesh(m, 2, layer_height=1)
U0 = FiniteElement('DG', 'triangle', 0)
U1 = FiniteElement('CG', 'interval', 2)
W0 = OuterProductElement(U0, U1)
W1 = FunctionSpace(e, HDiv(W0))
U0 = FiniteElement('DG', 'triangle', 0)
U1 = FiniteElement('CG', 'interval', 2)
W0 = OuterProductElement(U0, U1)
W2 = FunctionSpace(e, HDiv(W0))
assert W1 is W2
def test_extruded_ope_misses_cache():
m = UnitSquareMesh(1, 1)
e = ExtrudedMesh(m, 2, layer_height=1)
U0 = FiniteElement('DG', 'triangle', 0)
U1 = FiniteElement('CG', 'interval', 2)
W0 = OuterProductElement(U0, U1)
W1 = FunctionSpace(e, HDiv(W0))
U0 = FiniteElement('CG', 'triangle', 1)
U1 = FiniteElement('DG', 'interval', 2)
W0 = OuterProductElement(U0, U1)
W2 = FunctionSpace(e, HCurl(W0))
assert W1 is not W2
def test_mixed_fs_hits_cache():
m = UnitSquareMesh(1, 1)
V1 = FunctionSpace(m, 'DG', 1)
Q1 = FunctionSpace(m, 'RT', 2)
W1 = V1*Q1
V2 = FunctionSpace(m, 'DG', 1)
Q2 = FunctionSpace(m, 'RT', 2)
W2 = V2*Q2
assert W1 is W2
def test_mixed_fs_misses_cache():
m = UnitSquareMesh(1, 1)
V1 = FunctionSpace(m, 'DG', 1)
Q1 = FunctionSpace(m, 'RT', 2)
W1 = V1*Q1
V2 = FunctionSpace(m, 'DG', 1)
Q2 = FunctionSpace(m, 'RT', 2)
W2 = Q2*V2
assert W1 is not W2
def test_extruded_mixed_fs_hits_cache():
m = UnitSquareMesh(1, 1)
e = ExtrudedMesh(m, 2, layer_height=1)
U0 = FiniteElement('DG', 'triangle', 0)
U1 = FiniteElement('CG', 'interval', 2)
V0 = OuterProductElement(U0, U1)
V1 = FunctionSpace(e, HDiv(V0))
U0 = FiniteElement('CG', 'triangle', 1)
U1 = FiniteElement('DG', 'interval', 2)
V0 = OuterProductElement(U0, U1)
V2 = FunctionSpace(e, HCurl(V0))
W1 = V1*V2
W2 = V1*V2
assert W1 is W2
def test_extruded_mixed_fs_misses_cache():
m = UnitSquareMesh(1, 1)
e = ExtrudedMesh(m, 2, layer_height=1)
U0 = FiniteElement('DG', 'triangle', 0)
U1 = FiniteElement('CG', 'interval', 2)
V0 = OuterProductElement(U0, U1)
V1 = FunctionSpace(e, HDiv(V0))
U0 = FiniteElement('CG', 'triangle', 1)
U1 = FiniteElement('DG', 'interval', 2)
V0 = OuterProductElement(U0, U1)
V2 = FunctionSpace(e, HCurl(V0))
W1 = V1*V2
W2 = V2*V1
assert W1 is not W2
def test_different_meshes_miss_cache():
m1 = UnitSquareMesh(1, 1)
V1 = FunctionSpace(m1, 'CG', 1)
m2 = UnitSquareMesh(1, 1)
V2 = FunctionSpace(m2, 'CG', 1)
assert V1 is not V2
# A bit of a weak test, but the gc is slightly non-deterministic
def test_mesh_fs_gced():
from firedrake.functionspace import FunctionSpaceBase
gc.collect()
gc.collect()
nmesh = howmany(MeshBase)
nfs = howmany(FunctionSpaceBase)
for i in range(10):
m = UnitIntervalMesh(5)
for fs in ['CG', 'DG']:
V = FunctionSpace(m, fs, 1)
del m, V
gc.collect()
gc.collect()
nmesh1 = howmany(MeshBase)
nfs1 = howmany(FunctionSpaceBase)
assert nmesh1 - nmesh < 5
assert nfs1 - nfs < 10
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/tests/regression/test_helmholtz_mixed.py
import pytest
from firedrake import *
def helmholtz_mixed(x, V1, V2, action=False):
# Create mesh and define function space
mesh = UnitSquareMesh(2**x, 2**x)
V1 = FunctionSpace(mesh, *V1, name="V")
V2 = FunctionSpace(mesh, *V2, name="P")
W = V1 * V2
# Define variational problem
lmbda = 1
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
f = Function(V2)
f.interpolate(Expression("(1+8*pi*pi)*sin(x[0]*pi*2)*sin(x[1]*pi*2)"))
a = (p*q - q*div(u) + lmbda*inner(v, u) + div(v)*p) * dx
L = f*q*dx
# Compute solution
x = Function(W)
if action:
system = action(a, x) - L == 0
else:
system = a == L
# Block system is:
# V Ct
# Ch P
# Eliminate V by forming a schur complement
solve(system, x, solver_parameters={'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'ksp_type': 'cg',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_V_ksp_type': 'cg',
'fieldsplit_P_ksp_type': 'cg'})
# Analytical solution
f.interpolate(Expression("sin(x[0]*pi*2)*sin(x[1]*pi*2)"))
return sqrt(assemble(dot(x[2] - f, x[2] - f) * dx))
@pytest.mark.parametrize(('V1', 'V2', 'threshold', 'action'),
[(('RT', 1), ('DG', 0), 1.9, False),
(('BDM', 1), ('DG', 0), 1.89, False),
(('BDM', 1), ('DG', 0), 1.89, True),
(('BDFM', 2), ('DG', 1), 1.9, False)])
def test_firedrake_helmholtz(V1, V2, threshold, action):
import numpy as np
diff = np.array([helmholtz_mixed(i, V1, V2) for i in range(3, 6)])
print "l2 error norms:", diff
conv = np.log2(diff[:-1] / diff[1:])
print "convergence order:", conv
assert (np.array(conv) > threshold).all()
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/docs/source/extruded-meshes.rst
.. only:: html
.. contents::
Extruded Meshes in Firedrake
============================
Introduction
------------
Firedrake provides several utility functions for the creation of
semi-structured meshes from an unstructured base mesh. Firedrake also
provides a wide range of finite element spaces, both simple and sophisticated,
for use with such meshes.
These meshes may be particularly appropriate when carrying out simulations
on high aspect ratio domains. More mundanely, they allow a two-dimensional
mesh to be built from square or rectangular cells.
The partial structure can be exploited to give performance advantages when
iterating over the mesh, relative to a fully unstructured traversal of the
same mesh. Firedrake exploits these benefits when extruded meshes are used.
Structured, Unstructured and Semi-Structured Meshes
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Structured and unstructured meshes differ in the way the topology of the mesh
is specified.
In a *fully structured* mesh, the array indices of mesh entities can be
computed directly. For example, given the index of the current cell, the
indices of the cell's vertices can be computed using a simple mathematical
expression. This means that data can be directly addressed, using expressions
of the form A[i].
In a *fully unstructured* mesh, there is no simple relation between the
indices of different mesh entities. Instead, the relationships have to be
explicitly stored. For example, given the index of the current cell, the
indices of the cell's vertices can only be found by looking up the information
in a separate array. It follows that data must be indirectly addressed, using
expressions of the form A[B[i]].
Memory access latency makes indirect addressing more expensive than direct
addressing: it is usually more efficient to compute the array index directly
than to look it up from memory.
The characteristics of a *semi-structured* or *extruded* mesh lie somewhere
between the two extremes above. An extruded mesh has an unstructured *base*
mesh. Each cell of the base mesh corresponds to a *column* of cells in the
extruded mesh. Visiting the first cell in each column requires indirect
addressing. However, visiting subsequent cells in the column can be done
using direct addressing. As the number of cells in the column increases,
the performance should approach that of a fully structured mesh.
Generating Extruded Meshes in Firedrake
---------------------------------------
Extruded meshes are built using the :py:class:`~.ExtrudedMesh` class. There
are several built-in extrusion types that generate commonly-used extruded
meshes. To create a more complicated extruded mesh, one can either pass a
hand-written kernel into the :py:class:`~.ExtrudedMesh` constructor, or one
can use a built-in extrusion type and modify the coordinate field afterwards.
The following information may be passed in to the constructor:
- a :py:class:`~.Mesh` object, which will be used as the base mesh.
- the desired number of cell layers in the extruded mesh.
- the ``extrusion_type``, which can be one of the built-in "uniform",
"radial" or "radial_hedgehog" -- these are described below -- or "custom".
If this argument is omitted, the "uniform" extrusion type will be used.
- the ``layer_height``, which is needed for the built-in extrusion types.
- a ``kernel``, only if the custom extrusion type is used
- the appropriate ``gdim``, describing the geometric dimension of the mesh,
only if the custom extrusion type is used.
Uniform Extrusion
~~~~~~~~~~~~~~~~~
Uniform extrusion adds another spatial dimension to the mesh. For example, a
2D base mesh becomes a 3D extruded mesh. The coordinates of the extruded mesh
are computed on the assumption that the layers are evenly spaced (hence the
word 'uniform').
Let ``m`` be a standard :py:class:`~.UnitSquareMesh`. The following code
produces the extruded mesh, whose base mesh is ``m``, with 5 mesh layers and
a layer thickness of 0.2:
.. code-block:: python
m = UnitSquareMesh(4, 4)
mesh = ExtrudedMesh(m, 5, layer_height=0.2, extrusion_type='uniform')
This can be simplified slightly. The extrusion_type defaults to 'uniform', so
this can be omitted. Furthermore, the layer_height, if omitted, defaults to
the reciprocal of the number of layers. The following code therefore has the
same effect:
.. code-block:: python
m = UnitSquareMesh(4, 4)
mesh = ExtrudedMesh(m, 5)
The base mesh and extruded mesh are shown below.
.. figure:: images/UnitSquare44.png
:scale: 63 %
:align: left
.. figure:: images/UnifExt.png
:scale: 57 %
:align: right
Radial Extrusion
~~~~~~~~~~~~~~~~
Radial extrusion extrudes cells radially outwards from the origin, without
increasing the number of spatial dimensions. An example in 2 dimensions, in
which a circle is extruded into an annulus, is:
.. code-block:: python
m = CircleManifoldMesh(20, radius=2)
mesh = ExtrudedMesh(m, 5, extrusion_type='radial')
The base mesh and extruded mesh are shown below.
.. figure:: images/CircleMM20.png
:scale: 67 %
:align: left
.. figure:: images/RadExt2D.png
:scale: 68 %
:align: right
An example in 3 dimensions, in which a sphere is extruded into a spherical
annulus, is:
.. code-block:: python
m = IcosahedralSphereMesh(radius=3, refinement_level=3)
mesh = ExtrudedMesh(m, 5, layer_height=0.1, extrusion_type='radial')
The base mesh and part of the extruded mesh are shown below.
.. figure:: images/Icos3.png
:scale: 68 %
:align: left
.. figure:: images/RadExt3D.png
:scale: 72 %
:align: right
Hedgehog Extrusion
~~~~~~~~~~~~~~~~~~
Hedgehog extrusion is similar to radial extrusion, but the cells are extruded
outwards in a direction normal to the base cell. This produces a discontinuous
coordinate field.
.. code-block:: python
m = CircleManifoldMesh(20, radius=2)
mesh = ExtrudedMesh(m, 5, extrusion_type='radial_hedgehog')
An example in 3 dimensions, in which a sphere is extruded into a spherical
annulus, is:
.. code-block:: python
m = UnitIcosahedralSphereMesh(refinement_level=2)
mesh = ExtrudedMesh(m, 5, layer_height=0.1, extrusion_type='radial_hedgehog')
The 2D and 3D hedgehog-extruded meshes are shown below.
.. figure:: images/HedgeExt2D.png
:scale: 65 %
:align: left
.. figure:: images/HedgeExt3D.png
:scale: 70 %
:align: right
Custom Extrusion
~~~~~~~~~~~~~~~~
For a more complicated extruded mesh, a custom *kernel* can be given by the
user. Since this is a mesh-wide operation, a PyOP2 parallel loop is
constructed by Firedrake.
.. code-block:: python
m = UnitSquareMesh(5, 5)
kernel = op2.Kernel("""
void extrusion_kernel(double **base_coords, double **ext_coords,
int **layer, double *layer_height) {
ext_coords[0][0] = base_coords[0][0]; // X
ext_coords[0][1] = base_coords[0][1]; // Y
ext_coords[0][2] = 0.1 * layer[0][0] + base_coords[0][1]; // Z
}
""", "extrusion_kernel")
mesh = ExtrudedMesh(m, 5, extrusion_type='custom', kernel=kernel, gdim=3)
Function Spaces on Extruded Meshes
----------------------------------
The syntax for building a :py:class:`~.FunctionSpace` on an extruded mesh is
an extension of the existing syntax used with normal meshes. On a
non-extruded mesh, the following syntax is used:
.. code-block:: python
mesh = UnitSquareMesh(4, 4)
V = FunctionSpace(mesh, "RT", 1)
To allow maximal flexibility in constructing function spaces, Firedrake
supports a more general syntax:
.. code-block:: python
V = FunctionSpace(mesh, element)
where ``element`` is a UFL :py:class:`~ufl.finiteelement.finiteelement.FiniteElement` object. This
requires generation and manipulation of FiniteElement objects.
Geometrically, an extruded mesh cell is the *product* of a base, "horizontal",
cell with a "vertical" interval. The construction of function spaces on
extruded meshes makes use of this. Firedrake supports all function spaces
whose local element can be expressed as the product of an element defined on
the base cell with an element defined on an interval.
We will now introduce the new operators which act on FiniteElement objects.
The OuterProductElement operator
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
To create an Element compatible with an extruded mesh, one should use
the :py:class:`~ufl.finiteelement.outerproductelement.OuterProductElement`
operator. For example,
.. code-block:: python
horiz_elt = FiniteElement("CG", triangle, 1)
vert_elt = FiniteElement("CG", interval, 1)
elt = OuterProductElement(horiz_elt, vert_elt)
V = FunctionSpace(mesh, elt)
will give a continuous, scalar-valued function space. The resulting space
contains functions which vary linearly in the horizontal direction and
linearly in the vertical direction.
.. figure:: images/cg1cg1_prism.svg
:align: center
The product of a CG1 triangle element with a CG1 interval element
The degree and continuity may differ; for example
.. code-block:: python
horiz_elt = FiniteElement("DG", triangle, 0)
vert_elt = FiniteElement("CG", interval, 2)
elt = OuterProductElement(horiz_elt, vert_elt)
V = FunctionSpace(mesh, elt)
will give a function space which is continuous between cells in a column,
but discontinuous between horizontally-neighbouring cells. In addition,
the function may vary piecewise-quadratically in the vertical direction,
but is piecewise constant horizontally.
.. figure:: images/dg0cg2_prism.svg
:align: center
The product of a DG0 triangle element with a CG2 interval element
A more complicated element, like a Mini horizontal element with linear
variation in the vertical direction, may be built using the
:py:class:`~ufl.finiteelement.enrichedelement.EnrichedElement` functionality
in either of the following ways:
.. code-block:: python
mini_horiz_1 = FiniteElement("CG", triangle, 1)
mini_horiz_2 = FiniteElement("B", triangle, 3)
mini_horiz = mini_horiz_1 + mini_horiz_2 # Enriched element
mini_vert = FiniteElement("CG", interval, 1)
mini_elt = OuterProductElement(mini_horiz, mini_vert)
V = FunctionSpace(mesh, mini_elt)
or
.. code-block:: python
mini_horiz_1 = FiniteElement("CG", triangle, 1)
mini_horiz_2 = FiniteElement("B", triangle, 3)
mini_vert = FiniteElement("CG", interval, 1)
mini_elt_1 = OuterProductElement(mini_horiz_1, mini_vert)
mini_elt_2 = OuterProductElement(mini_horiz_2, mini_vert)
mini_elt = mini_elt_1 + mini_elt_2 # Enriched element
V = FunctionSpace(mesh, mini_elt)
.. figure:: images/mini_prism.svg
:align: center
The product of a Mini triangle element with a CG1 interval element
The HDiv and HCurl operators
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
For moderately complicated vector-valued elements,
:py:class:`~ufl.finiteelement.outerproductelement.OuterProductElement`
does not give enough information to unambiguously produce the desired
space. As an example, consider the lowest-order *Raviart-Thomas* element on a
quadrilateral. The degrees of freedom live on the facets, and consist of
a single evaluation of the component of the vector field normal to each facet.
The following element is closely related to the desired Raviart-Thomas element:
.. code-block:: python
CG_1 = FiniteElement("CG", interval, 1)
DG_0 = FiniteElement("DG", interval, 0)
P1P0 = OuterProductElement(CG_1, DG_0)
P0P1 = OuterProductElement(DG_0, CG_1)
elt = P1P0 + P0P1
.. figure:: images/rt_quad_pre.svg
:align: center
The element created above
However, this is only scalar-valued. There are two natural vector-valued
elements that can be generated from this: one of them preserves tangential
continuity between elements, and the other preserves normal continuity
between elements. To obtain the Raviart-Thomas element, we must use the
:py:class:`~ufl.finiteelement.hdivcurl.HDiv` operator:
.. code-block:: python
CG_1 = FiniteElement("CG", interval, 1)
DG_0 = FiniteElement("DG", interval, 0)
P1P0 = OuterProductElement(CG_1, DG_0)
RT_horiz = HDiv(P1P0)
P0P1 = OuterProductElement(DG_0, CG_1)
RT_vert = HDiv(P0P1)
elt = RT_horiz + RT_vert
.. figure:: images/rt_quad_post.svg
:align: center
The RT quadrilateral element, requiring the use
of :py:class:`~ufl.finiteelement.hdivcurl.HDiv`
Another reason to use these operators is when expanding a vector into a
higher dimensional space. Consider the lowest-order Nedelec element of the
2nd kind on a triangle:
.. code-block:: python
N2_1 = FiniteElement("N2curl", triangle, 1)
This is naturally vector-valued, and has two components. Suppose we form
the product of this with a continuous element on an interval:
.. code-block:: python
CG_2 = FiniteElement("CG", interval, 2)
N2CG = OuterProductElement(N2_1, CG_2)
This element still only has two components. To expand this into a
three-dimensional curl-conforming element, we must use the
:py:class:`~ufl.finiteelement.hdivcurl.HCurl` operator; the syntax is:
.. code-block:: python
Ned_horiz = HCurl(N2CG)
.. figure:: images/ned_prism.svg
:align: center
This gives the horizontal part of a Nedelec edge element on a triangular
prism. The full element can be built as follows:
.. code-block:: python
N2_1 = FiniteElement("N2curl", triangle, 1)
CG_2 = FiniteElement("CG", interval, 2)
N2CG = OuterProductElement(N2_1, CG_2)
Ned_horiz = HCurl(N2CG)
P2tr = FiniteElement("CG", triangle, 2)
P1dg = FiniteElement("DG", interval, 1)
P2P1 = OuterProductElement(P2tr, P1dg)
Ned_vert = HCurl(P2P1)
Ned_wedge = Ned_horiz + Ned_vert
V = FunctionSpace(mesh, Ned_wedge)
Shortcuts for simple spaces
~~~~~~~~~~~~~~~~~~~~~~~~~~~
Simple scalar-valued spaces can be created using a variation on the existing
syntax, if the ``HDiv``, ``HCurl`` and enrichment operations
are not required. To create a function space of degree 2 in the horizontal
direction, degree 1 in the vertical direction and possibly discontinuous
between layers, the short syntax is
.. code-block:: python
fspace = FunctionSpace(mesh, "CG", 2, vfamily="DG", vdegree=1)
If the horizontal and vertical parts have the same ``family`` and ``degree``,
the ``vfamily`` and ``vdegree`` arguments may be omitted. If ``mesh`` is an
:py:class:`~.ExtrudedMesh` object then the following are equivalent:
.. code-block:: python
fspace = FunctionSpace(mesh, "Lagrange", 1)
and
.. code-block:: python
fspace = FunctionSpace(mesh, "Lagrange", 1, vfamily="Lagrange", vdegree=1)
Solving Equations on Extruded Meshes
------------------------------------
Once the mesh and function spaces have been declared, extruded meshes behave
almost identically to normal meshes. However, there are some small differences,
which are listed below.
1. Surface integrals are no longer denoted by ``ds``. Since extruded meshes have
multiple types of surfaces, the following notation is used:
* ``ds_v`` is used to denote an integral over *side* facets of the mesh.
This can be combined with boundary markers from the base mesh, such as
``ds_v(1)``.
* ``ds_t`` is used to denote an integral over the *top* surface of the mesh.
* ``ds_b`` is used to denote an integral over the *bottom* surface of the mesh.
* ``ds_tb`` is used to denote an integral over both the *top* and *bottom*
surfaces of the mesh.
2. Interior facet integrals are no longer denoted by ``dS``. The *horizontal*
and *vertical* interior facets may require different numerical treatment.
To facilitate this, the following notation is used:
* ``dS_v`` is used to denote an integral over *horizontal* interior facets.
* ``dS_h`` is used to denote an integral over *vertical* interior facets.
3. When setting strong boundary conditions, the boundary markers from the base
mesh can be used to set boundary conditions on the relevant side of the
extruded mesh. To set boundary conditions on the top or bottom, the label
is replaced by:
* ``top``, to set a boundary condition on the top surface.
* ``bottom``, to set a boundary condition on the bottom surface.
<file_sep>/docs/source/r-space.rst
.. default-role:: math
A plausibly efficient implementation of `R`
===========================================
The function space `R` (for "Real" or, possibly, `Rognes
<http://home.simula.no/~meg/>`_) is employed to model concepts such as
global constraints. When employed as an unknown in an equation, it
presents implementation difficulties because it couples with all of
the other degrees of freedom. This results in a dense row in the
resulting matrix. Using the distributed CSR format which Firedrake
employs for other function spaces, both the assembly and action of
this row will require the entire system state to be gathered onto one
MPI process. This is clearly a horribly non-performant option.
Representing matrices involving `R`
-----------------------------------
Instead, we can observe that a dense matrix row (or column) is
isomorphic to a :class:`~firedrake.types.Function` and model these
blocks of the matrix accordingly. A non-trivial system
involving a function over `R` will always be a mixed system. The
resulting matrix will have four blocks, one a conventional sparse
matrix, one a dense column, one a dense row, and one a single
double. The dense row and column blocks can be implemented as matrix
shells. The row block will implement matrix multiplication as a dot
product returning a :class:`~pyop2.base.Global` while the column block
will implement matrix multiplication by pointwise scaling the input
:class:`~pyop2.base.Dat` and will return another
:class:`~pyop2.base.Dat`. This arrangement enables both the row block
and the column block to have the same parallel data distribution as a
:class:`~pyop2.base.Dat`, which removes the key scalability problem.
Assembling matrices involving `R`
---------------------------------
Assembling the column block will be as simple as replacing the trial
function with the constant 1, thereby transforming a 2-form into a
1-form, and assembling.
Similarly, assembling the row block simply requires the replacement of
the test function with the constant 1, and assembling.
The one by one block in the corner can be assembled by replacing both
the test and trial functions of the corresponding form with 1 and
assembling.
Clearly the remaining block does not involve `R` and can be assembled
as usual.
<file_sep>/tests/extrusion/test_extrusion_wedge_analytic.py
from __future__ import division
from firedrake import *
import pytest
import numpy as np
@pytest.fixture(scope='module')
def u_v():
m = ExtrudedMesh(UnitTriangleMesh(), layers=1, layer_height=1)
V = FunctionSpace(m, 'CG', 1)
u = TrialFunction(V)
v = TestFunction(V)
return u, v
def test_analytic_laplacian(u_v):
u, v = u_v
a = dot(grad(u), grad(v))*dx
vals = assemble(a).M.values
# Computed via sage:
# sage: x, y, z = var('x', 'y', 'z')
# sage: phi0 = (1 - x - y)*(1 - z)
# sage: phi1 = (1 - x - y)*z
# sage: phi2 = x*(1-z)
# sage: phi3 = x*z
# sage: phi4 = y*(1-z)
# sage: phi5 = y*z
# sage: phi = [phi0, phi1, phi2, phi3, phi4, phi5]
# sage: def grad(f):
# ... return [f.derivative(x), f.derivative(y), f.derivative(z)]
# sage: def dot(a, b):
# ... return sum(a_ * b_ for a_, b_ in zip(a, b))
# sage: def laplace(phi_i, phi_j):
# ... return integral(integral(integral(dot(grad(phi_i), grad(phi_j)), x, 0, 1 - y), y, 0, 1), z, 0, 1)
# sage: [[w_laplace(phi_i, phi_j) for phi_i in phi] for phi_j in phi]
analytic = np.asarray([[5/12, 1/12, -1/8, -1/8, -1/8, -1/8],
[1/12, 5/12, -1/8, -1/8, -1/8, -1/8],
[-1/8, -1/8, 1/4, 0, 1/24, -1/24],
[-1/8, -1/8, 0, 1/4, -1/24, 1/24],
[-1/8, -1/8, 1/24, -1/24, 1/4, 0],
[-1/8, -1/8, -1/24, 1/24, 0, 1/4]])
assert np.allclose(sorted(np.linalg.eigvals(vals)),
sorted(np.linalg.eigvals(analytic)))
def test_analytic_mass(u_v):
u, v = u_v
a = u*v*dx
vals = assemble(a).M.values
# sage: def w_mass(phi_i, phi_j):
# ... return integral(integral(integral(phi_i*phi_j, x, 0, 1 - y), y, 0, 1), z, 0, 1)
# sage: [[w_mass(phi_i, phi_j) for phi_i in phi] for phi_j in phi]
analytic = np.asarray([[1/36, 1/72, 1/72, 1/144, 1/72, 1/144],
[1/72, 1/36, 1/144, 1/72, 1/144, 1/72],
[1/72, 1/144, 1/36, 1/72, 1/72, 1/144],
[1/144, 1/72, 1/72, 1/36, 1/144, 1/72],
[1/72, 1/144, 1/72, 1/144, 1/36, 1/72],
[1/144, 1/72, 1/144, 1/72, 1/72, 1/36]])
assert np.allclose(sorted(np.linalg.eigvals(vals)),
sorted(np.linalg.eigvals(analytic)))
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/firedrake/functionspace.py
import numpy as np
import ufl
import pyop2.coffee.ast_base as ast
from pyop2 import op2
from pyop2.caching import ObjectCached
from pyop2.utils import flatten, as_tuple
import dmplex
import extrusion_utils as eutils
import fiat_utils
import mesh as mesh_t
import halo
import utils
__all__ = ['FunctionSpace', 'VectorFunctionSpace',
'MixedFunctionSpace', 'IndexedFunctionSpace']
class FunctionSpaceBase(ObjectCached):
"""Base class for :class:`.FunctionSpace`, :class:`.VectorFunctionSpace` and
:class:`.MixedFunctionSpace`.
.. note ::
Users should not directly create objects of this class, but one of its
derived types.
"""
def __init__(self, mesh, element, name=None, dim=1, rank=0):
"""
:param mesh: :class:`Mesh` to build this space on
:param element: :class:`ufl.FiniteElementBase` to build this space from
:param name: user-defined name for this space
:param dim: vector space dimension of a :class:`.VectorFunctionSpace`
:param rank: rank of the space, not the value rank
"""
self._ufl_element = element
# Compute the FIAT version of the UFL element above
self.fiat_element = fiat_utils.fiat_from_ufl_element(element)
if isinstance(mesh, mesh_t.ExtrudedMesh):
# Set up some extrusion-specific things
# The bottom layer maps will come from element_dof_list
# dof_count is the total number of dofs in the extruded mesh
# Get the flattened version of the FIAT element
self.flattened_element = self.fiat_element.flattened_element()
entity_dofs = self.flattened_element.entity_dofs()
self._dofs_per_cell = [len(entity)*len(entity[0]) for d, entity in entity_dofs.iteritems()]
# Compute the number of DoFs per dimension on top/bottom and sides
entity_dofs = self.fiat_element.entity_dofs()
top_dim = mesh._plex.getDimension()
self._xtr_hdofs = [len(entity_dofs[(d, 0)][0]) for d in range(top_dim+1)]
self._xtr_vdofs = [len(entity_dofs[(d, 1)][0]) for d in range(top_dim+1)]
# Compute the dofs per column
self.dofs_per_column = eutils.compute_extruded_dofs(self.fiat_element,
self.flattened_element.entity_dofs(),
mesh._layers)
# Compute the offset for the extrusion process
self.offset = eutils.compute_offset(self.fiat_element.entity_dofs(),
self.flattened_element.entity_dofs(),
self.fiat_element.space_dimension())
# Compute the top and bottom masks to identify boundary dofs
b_mask = self.fiat_element.get_lower_mask()
t_mask = self.fiat_element.get_upper_mask()
self.bt_masks = (b_mask, t_mask)
self.extruded = True
self._dofs_per_entity = self.dofs_per_column
else:
# If not extruded specific, set things to None/False, etc.
self.offset = None
self.bt_masks = None
self.dofs_per_column = np.zeros(1, np.int32)
self.extruded = False
entity_dofs = fiat_utils.flat_entity_dofs(self.fiat_element)
self._dofs_per_entity = [len(entity[0]) for d, entity in entity_dofs.iteritems()]
self._dofs_per_cell = [len(entity)*len(entity[0]) for d, entity in entity_dofs.iteritems()]
self.name = name
self._dim = dim
self._mesh = mesh
self._index = None
# Create the PetscSection mapping topological entities to DoFs
try:
# Old style createSection
self._global_numbering = mesh._plex.createSection(1, [1], self._dofs_per_entity,
perm=mesh._plex_renumbering)
except:
# New style
self._global_numbering = mesh._plex.createSection([1], self._dofs_per_entity,
perm=mesh._plex_renumbering)
mesh._plex.setDefaultSection(self._global_numbering)
self._universal_numbering = mesh._plex.getDefaultGlobalSection()
# Re-initialise the DefaultSF with the numbering for this FS
mesh._plex.createDefaultSF(self._global_numbering,
self._universal_numbering)
# Derive the Halo from the DefaultSF
self._halo = halo.Halo(mesh._plex.getDefaultSF(),
self._global_numbering,
self._universal_numbering)
# Compute entity class offsets
self.dof_classes = [0, 0, 0, 0]
for d in range(mesh._plex.getDimension()+1):
ncore = mesh._plex.getStratumSize("op2_core", d)
nowned = mesh._plex.getStratumSize("op2_non_core", d)
nhalo = mesh._plex.getStratumSize("op2_exec_halo", d)
nnonexec = mesh._plex.getStratumSize("op2_non_exec_halo", d)
ndofs = self._dofs_per_entity[d]
self.dof_classes[0] += ndofs * ncore
self.dof_classes[1] += ndofs * (ncore + nowned)
self.dof_classes[2] += ndofs * (ncore + nowned + nhalo)
self.dof_classes[3] += ndofs * (ncore + nowned + nhalo + nnonexec)
self._node_count = self._global_numbering.getStorageSize()
self.cell_node_list = mesh.create_cell_node_list(self._global_numbering,
self.fiat_element,
sum(self._dofs_per_cell))
if mesh._plex.getStratumSize("interior_facets", 1) > 0:
# Compute the facet_numbering and store with the parent mesh
if mesh.interior_facets is None:
# Order interior facets by OP2 entity class
interior_facets, interior_facet_classes = \
dmplex.get_facets_by_class(mesh._plex, "interior_facets")
interior_local_facet_number, interior_facet_cell = \
dmplex.facet_numbering(mesh._plex, "interior",
interior_facets,
mesh._cell_numbering,
mesh.cell_closure)
mesh.interior_facets = mesh_t._Facets(mesh, interior_facet_classes,
"interior",
interior_facet_cell,
interior_local_facet_number)
interior_facet_cells = mesh.interior_facets.facet_cell
self.interior_facet_node_list = \
dmplex.get_facet_nodes(interior_facet_cells,
self.cell_node_list)
else:
self.interior_facet_node_list = np.array([], dtype=np.int32)
if mesh.interior_facets is None:
mesh.interior_facets = mesh_t._Facets(self, 0, "exterior", None, None)
if mesh._plex.getStratumSize("exterior_facets", 1) > 0:
# Compute the facet_numbering and store with the parent mesh
if mesh.exterior_facets is None:
# Order exterior facets by OP2 entity class
exterior_facets, exterior_facet_classes = \
dmplex.get_facets_by_class(mesh._plex, "exterior_facets")
# Derive attached boundary IDs
if mesh._plex.hasLabel("boundary_ids"):
boundary_ids = np.zeros(exterior_facets.size, dtype=np.int32)
for i, facet in enumerate(exterior_facets):
boundary_ids[i] = mesh._plex.getLabelValue("boundary_ids", facet)
unique_ids = np.sort(mesh._plex.getLabelIdIS("boundary_ids").indices)
else:
boundary_ids = None
unique_ids = None
exterior_local_facet_number, exterior_facet_cell = \
dmplex.facet_numbering(mesh._plex, "exterior",
exterior_facets,
mesh._cell_numbering,
mesh.cell_closure)
mesh.exterior_facets = mesh_t._Facets(mesh, exterior_facet_classes,
"exterior",
exterior_facet_cell,
exterior_local_facet_number,
boundary_ids,
unique_markers=unique_ids)
exterior_facet_cells = mesh.exterior_facets.facet_cell
self.exterior_facet_node_list = \
dmplex.get_facet_nodes(exterior_facet_cells,
self.cell_node_list)
else:
self.exterior_facet_node_list = np.array([], dtype=np.int32)
if mesh.exterior_facets is None:
if mesh._plex.hasLabel("boundary_ids"):
unique_ids = np.sort(mesh._plex.getLabelIdIS("boundary_ids").indices)
else:
unique_ids = None
mesh.exterior_facets = mesh_t._Facets(self, 0, "exterior", None, None,
unique_markers=unique_ids)
# Note: this is the function space rank. The value rank may be different.
self.rank = rank
# Empty map caches. This is a sui generis cache
# implementation because of the need to support boundary
# conditions.
self._cell_node_map_cache = {}
self._exterior_facet_map_cache = {}
self._interior_facet_map_cache = {}
@property
def index(self):
"""Position of this :class:`FunctionSpaceBase` in the
:class:`.MixedFunctionSpace` it was extracted from."""
return self._index
@property
def node_count(self):
"""The number of global nodes in the function space. For a
plain :class:`.FunctionSpace` this is equal to
:attr:`dof_count`, however for a :class:`.VectorFunctionSpace`,
the :attr:`dof_count`, is :attr:`dim` times the
:attr:`node_count`."""
return self._node_count
@property
def dof_count(self):
"""The number of global degrees of freedom in the function
space. Cf. :attr:`node_count`."""
return self._node_count*self._dim
@utils.cached_property
def node_set(self):
"""A :class:`pyop2.Set` containing the nodes of this
:class:`.FunctionSpace`. One or (for
:class:`.VectorFunctionSpace`\s) more degrees of freedom are
stored at each node.
"""
name = "%s_nodes" % self.name
if self._halo:
s = op2.Set(self.dof_classes, name,
halo=self._halo.op2_halo)
if self.extruded:
return op2.ExtrudedSet(s, layers=self._mesh.layers)
return s
else:
s = op2.Set(self.node_count, name)
if self.extruded:
return op2.ExtrudedSet(s, layers=self._mesh.layers)
return s
@utils.cached_property
def dof_dset(self):
"""A :class:`pyop2.DataSet` containing the degrees of freedom of
this :class:`.FunctionSpace`."""
return op2.DataSet(self.node_set, self.dim)
def make_dat(self, val=None, valuetype=None, name=None, uid=None):
"""Return a newly allocated :class:`pyop2.Dat` defined on the
:attr:`dof_dset` of this :class:`.Function`."""
return op2.Dat(self.dof_dset, val, valuetype, name, uid=uid)
def cell_node_map(self, bcs=None):
"""Return the :class:`pyop2.Map` from interior facets to
function space nodes. If present, bcs must be a tuple of
:class:`.DirichletBC`\s. In this case, the facet_node_map will return
negative node indices where boundary conditions should be
applied. Where a PETSc matrix is employed, this will cause the
corresponding values to be discarded during matrix assembly."""
if bcs:
parent = self.cell_node_map()
else:
parent = None
return self._map_cache(self._cell_node_map_cache,
self._mesh.cell_set,
self.cell_node_list,
self.fiat_element.space_dimension(),
bcs,
"cell_node",
self.offset,
parent)
def interior_facet_node_map(self, bcs=None):
"""Return the :class:`pyop2.Map` from interior facets to
function space nodes. If present, bcs must be a tuple of
:class:`.DirichletBC`\s. In this case, the facet_node_map will return
negative node indices where boundary conditions should be
applied. Where a PETSc matrix is employed, this will cause the
corresponding values to be discarded during matrix assembly."""
if bcs:
parent = self.interior_facet_node_map()
else:
parent = None
offset = self.cell_node_map().offset
return self._map_cache(self._interior_facet_map_cache,
self._mesh.interior_facets.set,
self.interior_facet_node_list,
2*self.fiat_element.space_dimension(),
bcs,
"interior_facet_node",
offset=np.append(offset, offset),
parent=parent)
def exterior_facet_node_map(self, bcs=None):
"""Return the :class:`pyop2.Map` from exterior facets to
function space nodes. If present, bcs must be a tuple of
:class:`.DirichletBC`\s. In this case, the facet_node_map will return
negative node indices where boundary conditions should be
applied. Where a PETSc matrix is employed, this will cause the
corresponding values to be discarded during matrix assembly."""
if bcs:
parent = self.exterior_facet_node_map()
else:
parent = None
facet_set = self._mesh.exterior_facets.set
if isinstance(self._mesh, mesh_t.ExtrudedMesh):
name = "extruded_exterior_facet_node"
offset = self.offset
else:
name = "exterior_facet_node"
offset = None
return self._map_cache(self._exterior_facet_map_cache,
facet_set,
self.exterior_facet_node_list,
self.fiat_element.space_dimension(),
bcs,
name,
parent=parent,
offset=offset)
def bottom_nodes(self):
"""Return a list of the bottom boundary nodes of the extruded mesh.
The bottom mask is applied to every bottom layer cell to get the
dof ids."""
return np.unique(self.cell_node_list[:, self.bt_masks[0]])
def top_nodes(self):
"""Return a list of the top boundary nodes of the extruded mesh.
The top mask is applied to every top layer cell to get the dof ids."""
voffs = self.offset.take(self.bt_masks[1])*(self._mesh.layers-2)
return np.unique(self.cell_node_list[:, self.bt_masks[1]] + voffs)
def _map_cache(self, cache, entity_set, entity_node_list, map_arity, bcs, name,
offset=None, parent=None):
if bcs is not None:
# Separate explicit bcs (we just place negative entries in
# the appropriate map values) from implicit ones (extruded
# top and bottom) that require PyOP2 code gen.
explicit_bcs = [bc for bc in bcs if bc.sub_domain not in ['top', 'bottom']]
implicit_bcs = [bc.sub_domain for bc in bcs if bc.sub_domain in ['top', 'bottom']]
if len(explicit_bcs) == 0:
# Implicit bcs are not part of the cache key for the
# map (they only change the generated PyOP2 code),
# hence rewrite bcs here.
bcs = None
if len(implicit_bcs) == 0:
implicit_bcs = None
else:
implicit_bcs = None
if bcs is None:
# Empty tuple if no bcs found. This is so that matrix
# assembly, which uses a set to keep track of the bcs
# applied to matrix hits the cache when that set is
# empty. tuple(set([])) == tuple().
lbcs = tuple()
else:
if not all(bc.function_space() == self for bc in bcs):
raise RuntimeError("DirichletBC defined on a different FunctionSpace!")
# Ensure bcs is a tuple in a canonical order for the hash key.
lbcs = tuple(sorted(bcs, key=lambda bc: bc.__hash__()))
try:
# Cache hit
val = cache[lbcs]
# In the implicit bc case, we decorate the cached map with
# the list of implicit boundary conditions so PyOP2 knows
# what to do.
if implicit_bcs:
val = op2.DecoratedMap(val, implicit_bcs=implicit_bcs)
return val
except KeyError:
# Cache miss.
# Any top and bottom bcs (for the extruded case) are handled elsewhere.
nodes = [bc.nodes for bc in lbcs if bc.sub_domain not in ['top', 'bottom']]
if nodes:
bcids = reduce(np.union1d, nodes)
node_list_bc = np.arange(self.node_count)
node_list_bc[bcids] = -10000000
new_entity_node_list = node_list_bc.take(entity_node_list)
else:
new_entity_node_list = entity_node_list
val = op2.Map(entity_set, self.node_set,
map_arity,
new_entity_node_list,
("%s_"+name) % (self.name),
offset,
parent,
self.bt_masks)
cache[lbcs] = val
if implicit_bcs:
return op2.DecoratedMap(val, implicit_bcs=implicit_bcs)
return val
@utils.memoize
def exterior_facet_boundary_node_map(self, method):
'''The :class:`pyop2.Map` from exterior facets to the nodes on
those facets. Note that this differs from
:meth:`exterior_facet_node_map` in that only surface nodes
are referenced, not all nodes in cells touching the surface.
:arg method: The method for determining boundary nodes. See
:class:`~.bcs.DirichletBC`.
'''
el = self.fiat_element
# Facet dimension becomes a bit more complicated
# for quadrilaterals, as their dimension is (1, 1),
# so facets have dimensions (0, 1) AND (1, 0),
# which forces us to deal with multiple dimension values.
dims = self._mesh.facet_dimensions()
if method == "topological":
boundary_dofs = dict(enumerate(value
for dim in dims
for value in el.entity_closure_dofs()[dim].values()))
elif method == "geometric":
boundary_dofs = el.facet_support_dofs()
nodes_per_facet = \
len(boundary_dofs[0])
# HACK ALERT
# The facet set does not have a halo associated with it, since
# we only construct halos for DoF sets. Fortunately, this
# loop is direct and we already have all the correct
# information available locally. So We fake a set of the
# correct size and carry out a direct loop
facet_set = op2.Set(self._mesh.exterior_facets.set.total_size)
fs_dat = op2.Dat(facet_set**el.space_dimension(),
data=self.exterior_facet_node_map().values_with_halo)
facet_dat = op2.Dat(facet_set**nodes_per_facet,
dtype=np.int32)
local_facet_nodes = np.array(
[dofs for e, dofs in boundary_dofs.iteritems()])
# Helper function to turn the inner index of an array into c
# array literals.
c_array = lambda xs: "{"+", ".join(map(str, xs))+"}"
body = ast.Block([ast.Decl("int",
ast.Symbol("l_nodes",
(sum(len(el.get_reference_element().topology[dim]) for dim in dims),
nodes_per_facet)),
init=ast.ArrayInit(c_array(map(c_array, local_facet_nodes))),
qualifiers=["const"]),
ast.For(ast.Decl("int", "n", 0),
ast.Less("n", nodes_per_facet),
ast.Incr("n", 1),
ast.Assign(ast.Symbol("facet_nodes", ("n",)),
ast.Symbol("cell_nodes", ("l_nodes[facet[0]][n]",))))
])
kernel = op2.Kernel(ast.FunDecl("void", "create_bc_node_map",
[ast.Decl("int*", "cell_nodes"),
ast.Decl("int*", "facet_nodes"),
ast.Decl("unsigned int*", "facet")],
body),
"create_bc_node_map")
local_facet_dat = op2.Dat(facet_set ** self._mesh.exterior_facets._rank,
self._mesh.exterior_facets.local_facet_dat.data_ro_with_halos,
dtype=np.uintc)
op2.par_loop(kernel, facet_set,
fs_dat(op2.READ),
facet_dat(op2.WRITE),
local_facet_dat(op2.READ))
if isinstance(self._mesh, mesh_t.ExtrudedMesh):
offset = self.offset[boundary_dofs[0]]
else:
offset = None
return op2.Map(facet_set, self.node_set,
nodes_per_facet,
facet_dat.data_ro_with_halos,
name="exterior_facet_boundary_node",
offset=offset)
@property
def dim(self):
"""The vector dimension of the :class:`.FunctionSpace`. For a
:class:`.FunctionSpace` this is always one. For a
:class:`.VectorFunctionSpace` it is the value given to the
constructor, and defaults to the geometric dimension of the :class:`Mesh`. """
return self._dim
@property
def cdim(self):
"""The sum of the vector dimensions of the :class:`.FunctionSpace`. For a
:class:`.FunctionSpace` this is always one. For a
:class:`.VectorFunctionSpace` it is the value given to the
constructor, and defaults to the geometric dimension of the :class:`Mesh`. """
return self._dim
def ufl_element(self):
"""The :class:`ufl.FiniteElement` used to construct this
:class:`FunctionSpace`."""
return self._ufl_element
def mesh(self):
"""The :class:`Mesh` used to construct this :class:`.FunctionSpace`."""
return self._mesh
def __len__(self):
return 1
def __iter__(self):
yield self
def __getitem__(self, i):
"""Return ``self`` if ``i`` is 0 or raise an exception."""
if i != 0:
raise IndexError("Only index 0 supported on a FunctionSpace")
return self
def __mul__(self, other):
"""Create a :class:`.MixedFunctionSpace` composed of this
:class:`.FunctionSpace` and other"""
return MixedFunctionSpace((self, other))
class FunctionSpace(FunctionSpaceBase):
"""Create a function space
:arg mesh: :class:`.Mesh` to build the function space on
:arg family: string describing function space family, or an
:class:`~ufl.finiteelement.outerproductelement.OuterProductElement`
:arg degree: degree of the function space
:arg name: (optional) name of the function space
:arg vfamily: family of function space in vertical dimension
(:class:`.ExtrudedMesh`\es only)
:arg vdegree: degree of function space in vertical dimension
(:class:`.ExtrudedMesh`\es only)
If the mesh is an :class:`.ExtrudedMesh`, and the ``family``
argument is a
:class:`~ufl.finiteelement.outerproductelement.OuterProductElement`,
``degree``, ``vfamily`` and ``vdegree`` are ignored, since the
``family`` provides all necessary information, otherwise a
:class:`~ufl.finiteelement.outerproductelement.OuterProductElement`
is built from the (``family``, ``degree``) and (``vfamily``,
``vdegree``) pair. If the ``vfamily`` and ``vdegree`` are not
provided, the vertical element defaults to the same as the
(``family``, ``degree``) pair.
If the mesh is not an :class:`.ExtrudedMesh`, the ``family`` must be
a string describing the finite element family to use, and the
``degree`` must be provided, ``vfamily`` and ``vdegree`` are ignored in
this case.
"""
def __init__(self, mesh, family, degree=None, name=None, vfamily=None, vdegree=None):
if self._initialized:
return
# Two choices:
# 1) pass in mesh, family, degree to generate a simple function space
# 2) set up the function space using FiniteElement, EnrichedElement,
# OuterProductElement and so on
if isinstance(family, ufl.FiniteElementBase):
# Second case...
element = family.reconstruct(domain=mesh.ufl_domain())
else:
# First case...
if isinstance(mesh, mesh_t.ExtrudedMesh):
# if extruded mesh, make the OPE
la = _ufl_finite_element(family,
domain=mesh._old_mesh.ufl_cell(),
degree=degree)
if vfamily is None or vdegree is None:
# if second element was not passed in, assume same as first
# (only makes sense for CG or DG)
lb = ufl.FiniteElement(family,
domain=ufl.Cell("interval", 1),
degree=degree)
else:
# if second element was passed in, use in
lb = ufl.FiniteElement(vfamily,
domain=ufl.Cell("interval", 1),
degree=vdegree)
# now make the OPE
element = ufl.OuterProductElement(la, lb, domain=mesh.ufl_domain())
else:
# if not an extruded mesh, just make the element
element = _ufl_finite_element(family, domain=mesh.ufl_domain(), degree=degree)
super(FunctionSpace, self).__init__(mesh, element, name, dim=1)
self._initialized = True
@classmethod
def _process_args(cls, *args, **kwargs):
return (args[0], ) + args, kwargs
@classmethod
def _cache_key(cls, mesh, family, degree=None, name=None, vfamily=None, vdegree=None):
return family, degree, vfamily, vdegree
def __getitem__(self, i):
"""Return self if ``i`` is 0, otherwise raise an error."""
assert i == 0, "Can only extract subspace 0 from %r" % self
return self
class VectorFunctionSpace(FunctionSpaceBase):
"""A vector finite element :class:`FunctionSpace`."""
def __init__(self, mesh, family, degree, dim=None, name=None, vfamily=None, vdegree=None):
if self._initialized:
return
# VectorFunctionSpace dimension defaults to the geometric dimension of the mesh.
dim = dim or mesh.ufl_cell().geometric_dimension()
if isinstance(mesh, mesh_t.ExtrudedMesh):
if isinstance(family, ufl.OuterProductElement):
raise NotImplementedError("Not yet implemented")
la = _ufl_finite_element(family,
domain=mesh._old_mesh.ufl_cell(),
degree=degree)
if vfamily is None or vdegree is None:
lb = ufl.FiniteElement(family, domain=ufl.Cell("interval", 1),
degree=degree)
else:
lb = ufl.FiniteElement(vfamily, domain=ufl.Cell("interval", 1),
degree=vdegree)
element = ufl.OuterProductVectorElement(la, lb, dim=dim, domain=mesh.ufl_domain())
else:
element = _ufl_vector_element(family, domain=mesh.ufl_domain(), degree=degree, dim=dim)
super(VectorFunctionSpace, self).__init__(mesh, element, name, dim=dim, rank=1)
self._initialized = True
@classmethod
def _process_args(cls, *args, **kwargs):
return (args[0], ) + args, kwargs
@classmethod
def _cache_key(cls, mesh, family, degree=None, dim=None, name=None, vfamily=None, vdegree=None):
return family, degree, dim, vfamily, vdegree
def __getitem__(self, i):
"""Return self if ``i`` is 0, otherwise raise an error."""
assert i == 0, "Can only extract subspace 0 from %r" % self
return self
class MixedFunctionSpace(FunctionSpaceBase):
"""A mixed finite element :class:`FunctionSpace`."""
def __init__(self, spaces, name=None):
"""
:param spaces: a list (or tuple) of :class:`FunctionSpace`\s
The function space may be created as ::
V = MixedFunctionSpace(spaces)
``spaces`` may consist of multiple occurances of the same space: ::
P1 = FunctionSpace(mesh, "CG", 1)
P2v = VectorFunctionSpace(mesh, "Lagrange", 2)
ME = MixedFunctionSpace([P2v, P1, P1, P1])
"""
if self._initialized:
return
self._spaces = [IndexedFunctionSpace(s, i, self)
for i, s in enumerate(flatten(spaces))]
self._mesh = self._spaces[0].mesh()
self._ufl_element = ufl.MixedElement(*[fs.ufl_element() for fs in self._spaces])
self.name = name or '_'.join(str(s.name) for s in self._spaces)
self.rank = 1
self._index = None
self._initialized = True
@classmethod
def _process_args(cls, *args, **kwargs):
"""Convert list of spaces to tuple (to make it hashable)"""
mesh = args[0][0].mesh()
pargs = tuple(as_tuple(arg) for arg in args)
return (mesh, ) + pargs, kwargs
@classmethod
def _cache_key(cls, *args, **kwargs):
return args
def split(self):
"""The list of :class:`FunctionSpace`\s of which this
:class:`MixedFunctionSpace` is composed."""
return self._spaces
def sub(self, i):
"""Return the `i`th :class:`FunctionSpace` in this
:class:`MixedFunctionSpace`."""
return self[i]
def num_sub_spaces(self):
"""Return the number of :class:`FunctionSpace`\s of which this
:class:`MixedFunctionSpace` is composed."""
return len(self)
def __len__(self):
"""Return the number of :class:`FunctionSpace`\s of which this
:class:`MixedFunctionSpace` is composed."""
return len(self._spaces)
def __getitem__(self, i):
"""Return the `i`th :class:`FunctionSpace` in this
:class:`MixedFunctionSpace`."""
return self._spaces[i]
def __iter__(self):
for s in self._spaces:
yield s
@property
def dim(self):
"""Return a tuple of :attr:`FunctionSpace.dim`\s of the
:class:`FunctionSpace`\s of which this :class:`MixedFunctionSpace` is
composed."""
return tuple(fs.dim for fs in self._spaces)
@property
def cdim(self):
"""Return the sum of the :attr:`FunctionSpace.dim`\s of the
:class:`FunctionSpace`\s this :class:`MixedFunctionSpace` is
composed of."""
return sum(fs.dim for fs in self._spaces)
@property
def node_count(self):
"""Return a tuple of :attr:`FunctionSpace.node_count`\s of the
:class:`FunctionSpace`\s of which this :class:`MixedFunctionSpace` is
composed."""
return tuple(fs.node_count for fs in self._spaces)
@property
def dof_count(self):
"""Return a tuple of :attr:`FunctionSpace.dof_count`\s of the
:class:`FunctionSpace`\s of which this :class:`MixedFunctionSpace` is
composed."""
return tuple(fs.dof_count for fs in self._spaces)
@utils.cached_property
def node_set(self):
"""A :class:`pyop2.MixedSet` containing the nodes of this
:class:`MixedFunctionSpace`. This is composed of the
:attr:`FunctionSpace.node_set`\s of the underlying
:class:`FunctionSpace`\s this :class:`MixedFunctionSpace` is
composed of one or (for VectorFunctionSpaces) more degrees of freedom
are stored at each node."""
return op2.MixedSet(s.node_set for s in self._spaces)
@utils.cached_property
def dof_dset(self):
"""A :class:`pyop2.MixedDataSet` containing the degrees of freedom of
this :class:`MixedFunctionSpace`. This is composed of the
:attr:`FunctionSpace.dof_dset`\s of the underlying
:class:`FunctionSpace`\s of which this :class:`MixedFunctionSpace` is
composed."""
return op2.MixedDataSet(s.dof_dset for s in self._spaces)
def cell_node_map(self, bcs=None):
"""A :class:`pyop2.MixedMap` from the :attr:`Mesh.cell_set` of the
underlying mesh to the :attr:`node_set` of this
:class:`MixedFunctionSpace`. This is composed of the
:attr:`FunctionSpace.cell_node_map`\s of the underlying
:class:`FunctionSpace`\s of which this :class:`MixedFunctionSpace` is
composed."""
# FIXME: these want caching of sorts
bc_list = [[] for _ in self]
if bcs:
for bc in bcs:
bc_list[bc.function_space().index].append(bc)
return op2.MixedMap(s.cell_node_map(bc_list[i])
for i, s in enumerate(self._spaces))
def interior_facet_node_map(self, bcs=None):
"""Return the :class:`pyop2.MixedMap` from interior facets to
function space nodes. If present, bcs must be a tuple of
:class:`.DirichletBC`\s. In this case, the facet_node_map will return
negative node indices where boundary conditions should be
applied. Where a PETSc matrix is employed, this will cause the
corresponding values to be discarded during matrix assembly."""
# FIXME: these want caching of sorts
bc_list = [[] for _ in self]
if bcs:
for bc in bcs:
bc_list[bc.function_space().index].append(bc)
return op2.MixedMap(s.interior_facet_node_map(bc_list[i])
for i, s in enumerate(self._spaces))
def exterior_facet_node_map(self, bcs=None):
"""Return the :class:`pyop2.Map` from exterior facets to
function space nodes. If present, bcs must be a tuple of
:class:`.DirichletBC`\s. In this case, the facet_node_map will return
negative node indices where boundary conditions should be
applied. Where a PETSc matrix is employed, this will cause the
corresponding values to be discarded during matrix assembly."""
# FIXME: these want caching of sorts
bc_list = [[] for _ in self]
if bcs:
for bc in bcs:
bc_list[bc.function_space().index].append(bc)
return op2.MixedMap(s.exterior_facet_node_map(bc_list[i])
for i, s in enumerate(self._spaces))
@utils.cached_property
def exterior_facet_boundary_node_map(self):
'''The :class:`pyop2.MixedMap` from exterior facets to the nodes on
those facets. Note that this differs from
:meth:`exterior_facet_node_map` in that only surface nodes
are referenced, not all nodes in cells touching the surface.'''
return op2.MixedMap(s.exterior_facet_boundary_node_map for s in self._spaces)
def make_dat(self, val=None, valuetype=None, name=None, uid=None):
"""Return a newly allocated :class:`pyop2.MixedDat` defined on the
:attr:`dof_dset` of this :class:`MixedFunctionSpace`."""
if val is not None:
assert len(val) == len(self)
else:
val = [None for _ in self]
return op2.MixedDat(s.make_dat(v, valuetype, name, utils._new_uid())
for s, v in zip(self._spaces, val))
class IndexedFunctionSpace(FunctionSpaceBase):
"""A :class:`.FunctionSpaceBase` with an index to indicate which position
it has as part of a :class:`MixedFunctionSpace`."""
def __init__(self, fs, index, parent):
"""
:param fs: the :class:`.FunctionSpaceBase` that was extracted
:param index: the position in the parent :class:`MixedFunctionSpace`
:param parent: the parent :class:`MixedFunctionSpace`
"""
if self._initialized:
return
# If the function space was extracted from a mixed function space,
# extract the underlying component space
if isinstance(fs, IndexedFunctionSpace):
fs = fs._fs
# Override the __class__ to make instance checks on the type of the
# wrapped function space work as expected
self.__class__ = type(fs.__class__.__name__,
(self.__class__, fs.__class__), {})
self._fs = fs
self._index = index
self._parent = parent
self._initialized = True
@classmethod
def _process_args(cls, fs, index, parent, **kwargs):
return (fs.mesh(), fs, index, parent), kwargs
@classmethod
def _cache_key(cls, *args, **kwargs):
return args
def __getattr__(self, name):
return getattr(self._fs, name)
def __repr__(self):
return "<IndexFunctionSpace: %r at %d>" % (FunctionSpaceBase.__repr__(self._fs), self._index)
@property
def node_set(self):
"""A :class:`pyop2.Set` containing the nodes of this
:class:`FunctionSpace`. One or (for VectorFunctionSpaces) more degrees
of freedom are stored at each node."""
return self._fs.node_set
@property
def dof_dset(self):
"""A :class:`pyop2.DataSet` containing the degrees of freedom of
this :class:`FunctionSpace`."""
return self._fs.dof_dset
@property
def exterior_facet_boundary_node_map(self):
'''The :class:`pyop2.Map` from exterior facets to the nodes on
those facets. Note that this differs from
:meth:`exterior_facet_node_map` in that only surface nodes
are referenced, not all nodes in cells touching the surface.'''
return self._fs.exterior_facet_boundary_node_map
def _ufl_finite_element(family, domain, degree):
if isinstance(domain, ufl.Domain):
cell = domain.cell()
elif isinstance(domain, ufl.Cell):
cell = domain
else:
raise ValueError("Illegal domain or cell type")
if cell == ufl.Cell("quadrilateral"):
return ufl.OuterProductElement(
ufl.FiniteElement(family, domain=ufl.Cell("interval", 1), degree=degree),
ufl.FiniteElement(family, domain=ufl.Cell("interval", 1), degree=degree),
domain=domain)
else:
return ufl.FiniteElement(family, domain=domain, degree=degree)
def _ufl_vector_element(family, domain, degree, dim):
if isinstance(domain, ufl.Domain):
cell = domain.cell()
elif isinstance(domain, ufl.Cell):
cell = domain
else:
raise ValueError("Illegal domain or cell type")
if cell == ufl.Cell("quadrilateral"):
return ufl.OuterProductVectorElement(
ufl.FiniteElement(family, domain=ufl.Cell("interval", 1), degree=degree),
ufl.FiniteElement(family, domain=ufl.Cell("interval", 1), degree=degree),
dim=dim, domain=domain)
else:
return ufl.VectorElement(family, domain=domain, degree=degree, dim=dim)
<file_sep>/tests/regression/test_facet_normal.py
import pytest
from firedrake import *
def test_facet_normal_unit_interval():
"""Compute facet normals on the sides of the unit square."""
m = UnitIntervalMesh(2)
V = VectorFunctionSpace(m, 'CG', 1)
x_hat = Function(V).interpolate(Expression(('1.0',)))
n = FacetNormal(m)
assert assemble(dot(x_hat, n)*ds(1)) == -1.0 # x = 0
assert assemble(dot(x_hat, n)*ds(2)) == 1.0 # x = 1
def test_facet_normal_unit_square():
"""Compute facet normals on the sides of the unit square."""
m = UnitSquareMesh(2, 2)
V = VectorFunctionSpace(m, 'CG', 1)
x_hat = Function(V).interpolate(Expression(('1', '0')))
y_hat = Function(V).interpolate(Expression(('0', '1')))
n = FacetNormal(m)
assert assemble(dot(x_hat, n)*ds(1)) == -1.0 # x = 0
assert assemble(dot(x_hat, n)*ds(2)) == 1.0 # x = 1
assert assemble(dot(x_hat, n)*ds(3)) == 0.0 # y = 0
assert assemble(dot(x_hat, n)*ds(4)) == 0.0 # y = 1
assert assemble(dot(y_hat, n)*ds(1)) == 0.0 # x = 0
assert assemble(dot(y_hat, n)*ds(2)) == 0.0 # x = 1
assert assemble(dot(y_hat, n)*ds(3)) == -1.0 # y = 0
assert assemble(dot(y_hat, n)*ds(4)) == 1.0 # y = 1
def test_facet_normal_unit_cube():
"""Compute facet normals on the sides of the unit cube."""
m = UnitCubeMesh(1, 1, 1)
V = VectorFunctionSpace(m, 'CG', 1)
x_hat = Function(V).interpolate(Expression(('1', '0', '0')))
y_hat = Function(V).interpolate(Expression(('0', '1', '0')))
z_hat = Function(V).interpolate(Expression(('0', '0', '1')))
n = FacetNormal(m)
assert abs(assemble(dot(x_hat, n)*ds(1)) + 1.0) < 1e-14 # x = 0
assert abs(assemble(dot(x_hat, n)*ds(2)) - 1.0) < 1e-14 # x = 1
assert abs(assemble(dot(x_hat, n)*ds(3)) - 0.0) < 1e-14 # y = 0
assert abs(assemble(dot(x_hat, n)*ds(4)) - 0.0) < 1e-14 # y = 1
assert abs(assemble(dot(x_hat, n)*ds(5)) - 0.0) < 1e-14 # z = 0
assert abs(assemble(dot(x_hat, n)*ds(6)) - 0.0) < 1e-14 # z = 1
assert abs(assemble(dot(y_hat, n)*ds(1)) - 0.0) < 1e-14 # x = 0
assert abs(assemble(dot(y_hat, n)*ds(2)) - 0.0) < 1e-14 # x = 1
assert abs(assemble(dot(y_hat, n)*ds(3)) + 1.0) < 1e-14 # y = 0
assert abs(assemble(dot(y_hat, n)*ds(4)) - 1.0) < 1e-14 # y = 1
assert abs(assemble(dot(y_hat, n)*ds(5)) - 0.0) < 1e-14 # z = 0
assert abs(assemble(dot(y_hat, n)*ds(6)) - 0.0) < 1e-14 # z = 1
assert abs(assemble(dot(z_hat, n)*ds(1)) - 0.0) < 1e-14 # x = 0
assert abs(assemble(dot(z_hat, n)*ds(2)) - 0.0) < 1e-14 # x = 1
assert abs(assemble(dot(z_hat, n)*ds(3)) - 0.0) < 1e-14 # y = 0
assert abs(assemble(dot(z_hat, n)*ds(4)) - 0.0) < 1e-14 # y = 1
assert abs(assemble(dot(z_hat, n)*ds(5)) + 1.0) < 1e-14 # z = 0
assert abs(assemble(dot(z_hat, n)*ds(6)) - 1.0) < 1e-14 # z = 1
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
<file_sep>/docs/source/solving-interface.rst
.. only:: html
.. contents::
Solving PDEs
============
Introduction
------------
Now that we have learnt how to define weak variational problems, we
will move on to how to actually solve them using Firedrake. Let us
consider a weak variational problem
.. math::
a(u, v) = L(v) \; \forall v \in V \mathrm{on}\: \Omega
u = u_0 \; \mathrm{on}\: \partial\Omega
we will call the bilinear and linear parts of this form ``a`` and
``L`` respectively. The strongly imposed boundary condition, :math:`u
= u_0 \;\mathrm{on}\:\partial\Omega` will be represented by a variable
of type :py:class:`~.DirichletBC`, ``bc``.
Now that we have all the pieces of our variational problem, we can
move forward to solving it.
Solving the variational problem
-------------------------------
The function used to solve PDEs defined as above is
:py:func:`~firedrake.solving.solve`. This is a unified interface for
solving both linear and non-linear variational problems along with
linear systems (where the arguments are already assembled matrices and
vectors, rather than `UFL`_ forms). We will treat the variational
interface first.
Linear variational problems
~~~~~~~~~~~~~~~~~~~~~~~~~~~
If the problem is linear, that is ``a`` is linear in both the test and
trial functions and ``L`` is linear in the test function, we can use
the linear variational problem interface to ``solve``. To start, we
need a :py:class:`~.Function` to hold the value of
the solution:
.. code-block:: python
s = Function(V)
We can then solve the problem, placing the solution in ``s`` with:
.. code-block:: python
solve(a == L, s)
To apply boundary conditions, one passes a list of
:py:class:`~.DirichletBC` objects using the ``bcs``
keyword argument. For example, if there are two boundary conditions,
in ``bc1`` and ``bc2``, we write:
.. code-block:: python
solve(a == L, s, bcs=[bc1, bc2])
Nonlinear variational problems
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
For nonlinear problems, the interface is similar. In this case, we
solve a problem:
.. math::
F(u; v) = 0 \; \forall v \in V \mathrm{on}\: \Omega
u = u_0 \; \mathrm{on}\: \partial\Omega
where the *residual* :math:`F(u; v)` is linear in the test function
:math:`v` but possibly non-linear in the unknown
:py:class:`~.Function` :math:`u`. To solve such a
problem we write, if ``F`` is the residual form:
.. code-block:: python
solve(F == 0, u)
to apply strong boundary conditions, as before, we provide a list of
``DirichletBC`` objects using the ``bcs`` keyword:
.. code-block:: python
solve(F == 0, u, bcs=[bc1, bc2])
Nonlinear problems in Firedrake are solved using Newton-like methods.
That is, we compute successive approximations to the solution using
.. math::
u_{k+1} = u_{k} - J(u_k)^{-1} F(u_k) \; k = 0, 1, \dots
where :math:`u_0` is an initial guess for the solution and
:math:`J(u_k) = \frac{\partial F(u_k)}{\partial u_k}` is the
*Jacobian* of the residual, which should be non-singular at each
iteration. Notice how in the above examples, we did not explicitly
supply a Jacobian. If it is not supplied, it will be computed by
automatic differentiation of the residual form ``F`` with respect to the
solution variable ``u``. However, we may also supply the Jacobian
explicitly, using the keyword argument ``J``:
.. code-block:: python
solve(F == 0, u, J=user_supplied_jacobian_form)
The initial guess for the Newton iterations is provided in ``u``, for
example, to provide a non-zero guess that the solution is the value of
the ``x`` coordinate everywhere:
.. code-block:: python
u.interpolate(Expression('x[0]'))
solve(F == 0, u)
Solving linear systems
----------------------
Often, we might be solving a time-dependent linear system. In this
case, the bilinear form ``a`` does not change between timesteps, whereas
the linear form ``L`` does. Since assembly of the bilinear form is a
potentially costly process, Firedrake offers the ability to
"pre-assemble" forms in such systems and then reuse the assembled
operator in successive linear solves. Again, we use the same ``solve``
interface to do this, but must build slightly different objects to
pass in. In the pre-assembled case, we are solving a linear system:
.. math::
A\vec{x} = \vec{b}
Where :math:`A` is a known matrix, :math:`\vec{b}` is a known right
hand side vector and :math:`\vec{x}` is the unknown solution vector.
In Firedrake, :math:`A` is represented as a
:py:class:`~.Matrix`, while :math:`\vec{b}` and
:math:`\vec{x}` are both :py:class:`~.Function`\s.
We build these values by calling ``assemble`` on the UFL forms that
define our problem, which, as before are denoted ``a`` and ``L``.
Similarly to the linear variational case, we first need a function in
which to place our solution:
.. code-block:: python
x = Function(V)
We then :py:func:`~firedrake.solving.assemble` the left hand side
matrix ``A`` and known right hand side ``b`` from the bilinear and
linear forms respectively:
.. code-block:: python
A = assemble(a)
b = assemble(L)
Finally, we can solve the problem placing the solution in ``x``:
.. code-block:: python
solve(A, x, b)
to apply boundary conditions to the problem, we can assemble the
linear operator ``A`` with boundary conditions using the ``bcs``
keyword argument to :py:func:`~firedrake.solving.assemble` (and then
not supply them in solve call):
.. code-block:: python
A = assemble(a, bcs=[bc1, bc2])
b = assemble(L)
solve(A, x, b)
alternately, we can supply boundary conditions in
:py:func:`~firedrake.solving.solve` as before:
.. code-block:: python
A = assemble(a)
b = assemble(L)
solve(A, x, b, bcs=[bc1, bc2])
If boundary conditions have been supplied both in the assemble and
solve calls, then those provided for the solve take precedence, for
example, in the following, the system is solved only applying ``bc1``:
.. code-block:: python
A = assemble(a, bcs=[bc1, bc2])
b = assemble(L)
solve(A, x, b, bcs=[bc1])
Note that after the call to solve, ``A`` will be an assembled system
with only ``bc1`` applied, hence subsequent calls to ``solve`` that do
not change the boundary conditions again will not require a further
re-assembly.
Specifying solution methods
---------------------------
Not all linear and non-linear systems defined by PDEs are created
equal, and we therefore need ways of specifying which solvers to use
and options to pass to them. Firedrake uses `PETSc`_ to solve both
linear and non-linear systems and presents a uniform interface in
``solve`` to set PETSc solver options. In all cases, we set options
in the solve call by passing a dictionary to the ``solver_parameters``
keyword argument. To set options we use the same names that PETSc
uses in its command-line option setting interface (having removed the
leading ``-``). For more complete details on PETSc option naming we
recommend looking in the `PETSc manual`_. We describe some of the
more common options here.
Linear solver options
~~~~~~~~~~~~~~~~~~~~~
We use a PETSc `KSP`_ object to solve linear systems. This is a
uniform interface for solving linear systems using Krylov subspace
methods. By default, the solve call will use GMRES using an
incomplete LU factorisation to precondition the problem. To change
the Krylov method used in solving the problem, we set the
``'ksp_type'`` option. For example, if we want to solve a Helmholtz
equation, we know the operator is symmetric positive definite, and
therefore can choose the conjugate gradient method, rather than
GMRES.
.. code-block:: python
solve(a == L, solver_parameters={'ksp_type': 'cg'})
To change the preconditioner used, we set the ``'pc_type'`` option.
For example, if PETSc has been installed with the `Hypre`_ package, we
can use its algebraic multigrid preconditioner, BoomerAMG, to
precondition the system with:
.. code-block:: python
solve(a == L,
solver_parameters={'pc_type': 'hypre',
'pc_hypre_type': 'boomeramg'})
Although the `KSP` name suggests that only Krylov methods are
supported, this is not the case. We may, for example, solve the
system directly by computing an LU factorisation of the problem. To
do this, we set the ``pc_type`` to ``'lu'`` and tell PETSc to use a
"preconditioner only" Krylov method:
.. code-block:: python
solve(a == L,
solver_parameters={'ksp_type': 'preonly',
'pc_type': 'lu'})
In a similar manner, we can use Jacobi preconditioned Richardson
iterations with:
.. code-block:: python
solve(a == L,
solver_parameters={'ksp_type': 'richardson',
'pc_type': 'jacobi'}
.. note::
We note in passing that the method Firedrake utilises internally
for applying strong boundary conditions does not destroy the
symmetry of the linear operator. If the system without boundary
conditions is symmetric, it will continue to be so after the
application of any boundary conditions.
.. _linear_solver_tols:
Setting solver tolerances
+++++++++++++++++++++++++
In an iterative solver, such as Krylov method, we iterate until some
specified tolerance is reached. The measure of how much the current
solution :math:`\vec{x}_i` differs from the true solution is called
the residual and is calculated as:
.. math::
r = |\vec{b} - A \vec{x}_i|
PETSc allows us to set three different tolerance options for solving
the system. The *absolute tolerance* tells us we should stop if
:math:`r` drops below some given value. The *relative tolerance*
tells us we should stop if :math:`\frac{r}{|\vec{b}|}` drops below
some given value. Finally, PETSc can detect divergence in a linear
solve, that is, if :math:`r` increases above some specified value.
These values are set with the options ``'ksp_atol'`` for the absolute
tolerance, ``'ksp_rtol'`` for the relative tolerance, and
``'ksp_divtol'`` for the divergence tolerance. The values provided to
these options should be floats. For example, to set the absolute
tolerance to :math:`10^{-30}`, the relative tolerance to
:math:`10^{-9}` and the divergence tolerance to :math:`10^4` we would
use:
.. code-block:: python
solver_parameters={'ksp_atol': 1e-30,
'ksp_rtol': 1e-9,
'ksp_divtol': 1e4}
.. note::
By default, PETSc (and hence Firedrake) check for the convergence
in the preconditioned norm, that is, if the system is
preconditioned with a matrix :math:`P` the residual is calculated
as:
.. math::
r = |P^{-1}(\vec{b} - A \vec{x}_i)|
to check for convergence in the unpreconditioned norm set the
``'ksp_norm_type'`` option to ``'unpreconditioned'``.
Finally, we can set the maximum allowed number of iterations for the
Krylov method by using the ``'ksp_max_it'`` option.
.. _mixed_preconditioning:
Preconditioning mixed finite element systems
++++++++++++++++++++++++++++++++++++++++++++
PETSc provides an interface to composing "physics-based"
preconditioners for mixed systems which Firedrake exploits when it
assembles linear systems. In particular, for systems with two
variables (for example Navier-Stokes where we solve for the velocity
and pressure of the fluid), we can exploit PETSc's ability to build
preconditioners from Schur complements. This is one type of
preconditioner based on PETSc's `fieldsplit`_ technology. To take a
concrete example, let us consider solving the dual form of the
Helmholtz equation:
.. math::
\langle p, q \rangle - \langle q, \mathrm{div} u \rangle + \lambda
\langle v, u \rangle + \langle \mathrm{div}v, p \rangle =
\langle f, q \rangle \; \forall v \in V_1, q \in V_2
This has a stable solution if, for example, :math:`V_1` is the lowest order
Raviart-Thomas space and :math:`V_2` is the lowest order discontinuous
space.
.. code-block:: python
V1 = FunctionSpace(mesh, 'RT', 1)
V2 = FunctionSpace(mesh, 'DG', 0)
W = V1 * V2
lmbda = 1
u, p = TrialFunctions(W)
v, q = TestFunctions(W)
f = Function(V2)
a = (p*q - q*div(u) + lmbda*inner(v, u) + div(v)*p)*dx
L = f*q*dx
u = Function(W)
solve(a == L, u,
solver_parameters={'ksp_type': 'cg',
'pc_type': 'fieldsplit',
'pc_fieldsplit_type': 'schur',
'pc_fieldsplit_schur_fact_type': 'FULL',
'fieldsplit_0_ksp_type': 'cg',
'fieldsplit_1_ksp_type': 'cg'})
We refer to section 4.5 of the `PETSc manual`_ for more complete
details, but briefly describe the options in use here. The monolithic
system is conceptually a :math:`2\times2` block matrix:
.. math::
\left(\begin{matrix}
\lambda \langle v, u \rangle & -\langle q, \mathrm{div} u \rangle \\
\langle \mathrm{div} v, p \rangle & \langle p, q \rangle
\end{matrix}
\right) = \left(\begin{matrix} A & B \\ C & D \end{matrix}\right).
We can factor this block matrix in the following way:
.. math::
\left(\begin{matrix} I & 0 \\ C A^{-1} & I\end{matrix}\right)
\left(\begin{matrix}A & 0 \\ 0 & S\end{matrix}\right)
\left(\begin{matrix} I & A^{-1} B \\ 0 & I\end{matrix}\right).
This is the *Schur complement factorisation* of the block system, its
inverse is:
.. math::
P = \left(\begin{matrix} I & -A^{-1}B \\ 0 & I \end{matrix}\right)
\left(\begin{matrix} A^{-1} & 0 \\ 0 & S^{-1}\end{matrix}\right)
\left(\begin{matrix} I & 0 \\ -CA^{-1} & I\end{matrix}\right).
Where :math:`S` is the *Schur complement*:
.. math::
S = D - C A^{-1} B.
The options in the example above use an approximation to :math:`P` to
precondition the system. To do so, we tell PETSc that the
preconditioner should be of type ``'fieldsplit'``, and the the
fieldsplit's type should be ``'schur'``. We then select a
factorisation type for the Schur complement. The option ``'FULL'`` as
used above preconditions using an approximation to :math:`P`. We can
also use ``'diag'`` which uses an approximation to:
.. math::
\left(\begin{matrix} A^{-1} & 0 \\ 0 & -S^{-1} \end{matrix}\right).
Note the minus sign in front of :math:`S^{-1}` which is there such
that this preconditioner is positive definite. Two other options are
``'lower'``, where the preconditioner is an approximation to:
.. math::
\left(\begin{matrix}A & 0 \\ C & S\end{matrix}\right)^{-1} =
\left(\begin{matrix}A^{-1} & 0 \\ 0 & S^{-1}\end{matrix}\right)
\left(\begin{matrix}I & 0 \\ -C A^{-1} & I\end{matrix}\right)
and ``'upper'`` which uses:
.. math::
\left(\begin{matrix}A & B \\ 0 & S\end{matrix}\right)^{-1} =
\left(\begin{matrix}I & -A^{-1}B \\ 0 & I\end{matrix}\right)
\left(\begin{matrix}A^{-1} & 0 \\ 0 & S^{-1}\end{matrix}\right).
Note that the inverses of :math:`A` and :math:`S` are never formed
explicitly by PETSc, instead their actions are computed approximately
using a Krylov method. The choice of method is selected using the
``'fieldsplit_0_ksp_type'`` option (for the Krylov solver computing
:math:`A^{-1}`) and ``'fieldsplit_1_ksp_type'`` (for the Krylov solver
computing :math:`S^{-1}`).
.. note::
If you have given your
:py:class:`~.FunctionSpace`\s names, then
instead of 0 and 1, you should use the name of the function space
in these options.
By default PETSc uses an approximation to :math:`D^{-1}` to
precondition the Krylov system solving for :math:`S`, you can also use
a `least squares commutator <LSC_>`_, see the relevant section of the
`PETSc manual pages <fieldsplit_>`_ for more details.
More block preconditioners
++++++++++++++++++++++++++
As well as physics-based Schur complement preconditioners for block
systems, PETSc also allows us to use preconditioners formed from block
Jacobi (``'pc_fieldsplit_type': 'additive'``) and block Gauss-Seidel
(``'multiplicative'`` or ``'symmetric_multiplicative'``)
inverses of the block system. These work for any number of blocks,
whereas the Schur complement approach mentioned above only works for
two by two blocks.
.. note::
PETSc offers support for composing fieldsplit preconditioners
recursively. That is, defining a :math:`3\times3` block system as
composed of a :math:`2\times2` piece and a :math:`1\times1` piece.
However, the Firedrake interface to the solver options does not
currently support this. At present, we cannot tell PETSc that the
blocks should be split recursively.
Future versions of Firedrake may offer a symbolic language for
describing the composition of such physics-like preconditioners,
rather than having to specify everything using PETSc solver
options.
Nonlinear solver options
~~~~~~~~~~~~~~~~~~~~~~~~
As for linear systems, we use a PETSc object to solve nonlinear
systems. This time it is a `SNES`_. This offers a uniform interface
to Newton-like and quasi-Newton solution schemes. To select the SNES
type to use, we use the ``'snes_type'`` option. Recall that each
Newton iteration is the solution of a linear system, options for the
inner linear solve may be set in the same way as described above for
linear problems. For example, to solve a nonlinear problem using
Newton-Krylov iterations using a line search and direct factorisation
to solve the linear system we would write:
.. code-block:: python
solve(F == 0, u,
solver_parameters={'snes_type': 'newtonls',
'ksp_type': 'preonly',
'pc_type': 'lu'}
.. note::
Not all of PETSc's SNES types are currently supported by Firedrake,
since some of them require extra information which we do not
currently provide.
Setting convergence criteria
++++++++++++++++++++++++++++
In addition to setting the tolerances for the inner, linear solve in a
nonlinear system, which is done in exactly the same way as for
:ref:`linear problems <linear_solver_tols>`, we can also set
convergence tolerances on the outer SNES object. These are the
*absolute tolerance* (``'snes_atol'``), *relative tolerance*
(``'snes_rtol'``), *step tolerance* (``'snes_stol'``) along with the
maximum number of nonlinear iterations (``'snes_max_it'``) and the
maximum number of allowed function evaluations (``'snes_max_func'``).
The step tolerance checks for convergence due to:
.. math::
|\Delta x_k| < \mathrm{stol} \, |x_k|
The maximum number of allowed function evaluations limits the number
of times the residual may be evaluated before returning a
non-convergence error, and defaults to 1000.
Providing an operator for preconditioning
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
By default, Firedrake uses the Jacobian of the residual (or equally
the bilinear form for linear problems) to construct preconditioners
for the linear systems it solves. That is, it does not directly
solve:
.. math::
A \vec{x} = \vec{b}
but rather
.. math::
\tilde{A}^{-1} A \vec{x} = \tilde{A}^{-1} \vec{b}
where :math:`\tilde{A}^{-1}` is an approximation to :math:`A^{-1}`. If we
know something about the structure of our problem, we may be able to
construct an operator :math:`P` explicitly which is "easy" to invert,
and whose inverse approximates :math:`A^{-1}` well. Firedrake allows
you to provide this operator when solving variational problems by
passing an explicit :py:data:`Jp` keyword argument to the solve call,
the provided form will then be used to construct an approximate
inverse when preconditioning the problem, rather than the form we're
solving with.
.. code-block:: python
a = ...
L = ...
Jp = ...
# Use the approximate inverse of Jp to precondition solves
solve(a == L, ..., Jp=Jp)
Default solver options
~~~~~~~~~~~~~~~~~~~~~~
If no parameters are passed to a solve call, we use, in most cases,
the defaults that PETSc supplies for solving the linear or nonlinear
system. We describe the most commonly modified options (along with
their defaults in Firedrake) here. For linear variational solves we
use:
* ``ksp_type``: GMRES, with a restart (``ksp_gmres_restart``) of 30
* ``ksp_rtol``: 1e-7
* ``ksp_atol``: 1e-50
* ``ksp_divtol`` 1e4
* ``ksp_max_it``: 10000
* ``pc_type``: ILU (Jacobi preconditioning for mixed problems)
For nonlinear variational solves we have:
* ``snes_type``: Newton linesearch
* ``ksp_type``: GMRES, with a restart (``ksp_gmres_restart``) of 30
* ``snes_rtol``: 1e-8
* ``snes_atol``: 1e-50
* ``snes_stol``: 1e-8
* ``snes_max_it``: 50
* ``ksp_rtol``: 1e-5
* ``ksp_atol``: 1e-50
* ``ksp_divtol``: 1e4
* ``ksp_max_it``: 10000
* ``pc_type``: ILU (Jacobi preconditioning for mixed problems)
To see the full view that PETSc has of solver objects, you can pass a
view flag to the solve call. For linear solves pass:
.. code-block:: python
solver_parameters={'ksp_view': True}
For nonlinear solves use:
.. code-block:: python
solver_parameters={'snes_view': True}
PETSc will then print its view of the solver objects that Firedrake
has constructed. This is especially useful for debugging complicated
preconditioner setups for mixed problems.
Solving singular systems
------------------------
Some systems of PDEs, for example the Poisson equation with pure
Neumann boundary conditions, have an operator which is singular. That
is, we have :math:`Ae = 0` with :math:`e \neq 0`. The vector space
spanned by the set of vectors :math:`{e}` for which :math:`Ae = 0` is
termed the *null space* of :math:`A`. If we wish to solve such a
system, we must remove the null space from the solution. To do this
in Firedrake, we first must define the null space, and then inform the
solver of its existance. We use a
:class:`~firedrake.nullspace.VectorSpaceBasis` to hold the vectors
which span the null space. We must provide a list of
:class:`~.Function`\s or
:class:`~.Vector`\s spanning the space. Additionally,
since removing a constant null space is such a common operation, we
can pass ``constant=True`` to the constructor (rather than
constructing the constant vector by hand). Note that the vectors we
pass in must be *orthonormal*. Once the null space is built, we just
need to inform the solve about it (using the ``nullspace`` keyword
argument).
As an example, consider the Poisson equation with pure Neumann
boundary conditions:
.. math::
-\nabla^2 u &= 0 \quad \mathrm{in}\;\Omega\\
\nabla u \cdot n &= g \quad \mathrm{on}\;\Gamma.
We will solve this problem on the unit square applying homogeneous
Neumann boundary conditions on the planes :math:`x = 0` and :math:`x =
1`. On :math:`y = 0` we set :math:`g = -1` while on :math:`y = 1` we
set :math:`g = 1`. The null space of the operator we form is the set
of constant functions, and thus the problem has solution
:math:`u(x, y) = y + c` where :math:`c` is a constant. To solve the
problem, we will inform the solver of this constant null space, fixing
the solution to be :math:`u(x, y) = y - 0.5`.
.. code-block:: python
m = UnitSquareMesh(25, 25)
V = FunctionSpace(m, 'CG', 1)
u = TrialFunction(V)
v = TestFunction(V)
a = inner(grad(u), grad(v))*dx
L = -v*ds(3) + v*ds(4)
nullspace = VectorSpaceBasis(constant=True)
u = Function(V)
solve(a == L, u, nullspace=nullspace)
exact = Function(V)
exact.interpolate(Expression('x[1] - 0.5'))
print sqrt(assemble((u - exact)*(u - exact)*dx))
Singular operators in mixed spaces
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
If you have an operator in a mixed space, you may well precondition
the system using a `Schur complement <mixed_preconditioning>`_. If
the operator is singular, you will therefore have to tell the solver
about the null space of each diagonal block separately. To do this in
Firedrake, we build a
:class:`~firedrake.nullspace.MixedVectorSpaceBasis` instead of a
:class:`~firedrake.nullspace.VectorSpaceBasis` and then inform the
solver about it as before. A
:class:`~firedrake.nullspace.MixedVectorSpaceBasis` takes a list of
:class:`~firedrake.nullspace.VectorSpaceBasis` objects defining the
null spaces of each of the diagonal blocks in the mixed operator. In
addition, as a first argument, you must provide the
:class:`~.MixedFunctionSpace` you're building a basis for. You do not
have to provide a null space for all blocks. For those you don't care
about, you can pass an indexed function space at the appropriate
position. For example, imagine we have a mixed space :math:`W = V
\times Q` and an operator which has a null space of constant functions
in :math:`V` (this occurs, for example, for a discretisation of the
mixed poisson problem on the surface of a sphere). We can specify the
null space (indicating that we only really care about the constant
function) as:
.. code-block:: python
V = ...
Q = ...
W = V*Q
v_basis = VectorSpaceBasis(constant=True)
nullspace = MixedVectorSpaceBasis(W, [v_basis, W.sub(1)])
Debugging convergence failures
------------------------------
Occasionally, we will set up a problem and call solve only to be
confronted with an error that the solve failed to converge. Here, we
discuss some useful techniques to try and understand the reason. Much
of the advice in the `PETSc FAQ`_ is useful here, especially the
sections on `SNES nonconvergence`_ and `KSP nonconvergence`_. We
first consider linear problems.
Linear convergence failures
~~~~~~~~~~~~~~~~~~~~~~~~~~~
If the linear operator is correct, but the solve fails to converge, it
is likely the case that the problem is badly conditioned (leading to
slow convergence) or a symmetric method is being used (such as
conjugate gradient) where the problem is non-symmetric. The first
thing to check is what happened to the residual (error) term. To
monitor this in the solution we pass the "flag" options
``'ksp_converged_reason'`` and ``'ksp_monitor_true_residual'``,
additionally, we pass ``ksp_view`` so that PETSc prints its idea of
what the solver object contains (this is useful to debug the where
options are not being passed in correctly):
.. code-block:: python
solver_parameters={'ksp_converged_reason': True,
'ksp_monitor_true_residual': True,
'ksp_view': True}
If the problem is converging, but only slowly, it may be that it is
badly conditioned. If the problem is small, we can try using a direct
solve to see if the solution obtained is correct:
.. code-block:: python
solver_parameters={'ksp_type': 'preonly', 'pc_type': 'lu'}
If this approach fails with a "zero-pivot" error, it is likely that
the equations are singular, or nearly so, check to see if boundary
conditions have been imposed correctly.
If the problem converges with a direct method to the correct solution
but does not converge with a Krylov method, it's probable that the
conditioning is bad. If it's a mixed problem, try using a
physics-based preconditioner as described above, if not maybe try
using an algebraic multigrid preconditioner. If PETSc was installed
with Hypre use:
.. code-block:: python
solver_parameters={'pc_type': 'hypre', 'pc_hypre_type': 'boomeramg'}
If you're using a symmetric method, such as conjugate gradient, check
that the linear operator is actually symmetric, which you can compute
with the following:
.. code-block:: python
A = assemble(a) # use bcs keyword if there are boundary conditions
print A.M.handle.isSymmetric(tol=1e-13)
If the problem is not symmetric, try using a method such as GMRES
instead. PETSc uses restarted GMRES with a default restart of 30, for
difficult problems this might be too low, in which case, you can
increase the restart length with:
.. code-block:: python
solver_parameters={'ksp_gmres_restart': 100}
Nonlinear convergence failures
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Much of the advice for linear systems applies to nonlinear systems as
well. If you have a convergence failure for a nonlinear problem, the
first thing to do is run with monitors to see what is going on, and
view the SNES object with ``snes_view`` to ensure that PETSc is seeing
the correct options:
.. code-block:: python
solver_parameters={'snes_monitor': True,
'snes_view': True,
'ksp_monitor_true_residual': True,
'snes_converged_reason': True,
'ksp_converged_reason': True}
If the linear solve fails to converge, debug the problem as above for
linear systems. If the linear solve converges but the outer Newton
iterations do not, the problem is likely a bad Jacobian. If you
provided the Jacobian by hand, is it correct? If no Jacobian was
provided in the solve call, it is likely a bug in Firedrake and you
should `report it to us <firedrake_bugs_>`_.
.. _Hypre: http://acts.nersc.gov/hypre/
.. _PETSc: http://www.mcs.anl.gov/petsc/
.. _PETSc manual: http://www.mcs.anl.gov/petsc/petsc-current/docs/manual.pdf
.. _KSP: http://www.mcs.anl.gov/petsc/petsc-current/docs/manualpages/KSP/
.. _SNES: http://www.mcs.anl.gov/petsc/petsc-current/docs/manualpages/SNES/
.. _fieldsplit: http://www.mcs.anl.gov/petsc/petsc-current/docs/manualpages/PC/PCFIELDSPLIT.html
.. _PETSc FAQ: http://www.mcs.anl.gov/petsc/documentation/faq.html
.. _SNES nonconvergence: http://www.mcs.anl.gov/petsc/documentation/faq.html#newton
.. _KSP nonconvergence: http://www.mcs.anl.gov/petsc/documentation/faq.html#kspdiverged
.. _LSC: http://www.mcs.anl.gov/petsc/petsc-current/docs/manualpages/PC/PCLSC.html
.. _UFL: http://fenicsproject.org/documentation/ufl/1.2.0/ufl.html
.. _firedrake_bugs: mailto:<EMAIL>
<file_sep>/demos/poisson/poisson_mixed.py.rst
Mixed formulation for the Poisson equation
==========================================
We're considering the Poisson equation :math:`\nabla^2 u = -f` using a mixed
formulation on two coupled fields. We start by introducing the negative flux
:math:`\sigma = \nabla u` as an auxiliary vector-valued variable. This leaves
us with the PDE on a unit square :math:`\Omega = [0,1] \times [0,1]` with
boundary :math:`\Gamma`
.. math::
\sigma - \nabla u = 0 \ \textrm{on}\ \Omega
\nabla \cdot \sigma = -f \ \textrm{on}\ \Omega
u = u_0 \ \textrm{on}\ \Gamma_D
\sigma \cdot n = g \ \textrm{on}\ \Gamma_N
for some known function :math:`f`. The solution to this equation will be some
functions :math:`u\in V` and :math:`\sigma\in \Sigma` for some suitable
function space :math:`V` and :math:`\Sigma` that satisfy these equations. We
multiply by arbitrary test functions :math:`\tau \in V` and :math:`\nu \in
\Sigma`, integrate over the domain and then integrate by parts to obtain a
weak formulation of the variational problem: find :math:`\sigma\in \Sigma` and
:math:`\nu\in V` such that:
.. math::
\int_{\Omega} (\sigma \cdot \tau + \nabla \cdot \tau \ u) \ {\rm d} x
&= \int_{\Gamma} \tau \cdot n \ u \ {\rm d} s
\quad \forall \ \tau \in \Sigma, \\
\int_{\Omega} \nabla \cdot \sigma v \ {\rm d} x
&= - \int_{\Omega} f \ v \ {\rm d} x
\quad \forall \ v \in V.
The flux boundary condition :math:`\sigma \cdot n = g` becomes an *essential*
boundary condition to be enforced on the function space, while the boundary
condition :math:`u = u_0` turn into a *natural* boundary condition which
enters into the variational form, such that the variational problem can be
written as: find :math:`(\sigma, u)\in \Sigma_g \times V` such that
.. math::
a((\sigma, u), (\tau, v)) = L((\tau, v))
\quad \forall \ (\tau, v) \in \Sigma_0 \times V
with the variational forms :math:`a` and :math:`L` defined as
.. math::
a((\sigma, u), (\tau, v)) &=
\int_{\Omega} \sigma \cdot \tau + \nabla \cdot \tau \ u
+ \nabla \cdot \sigma \ v \ {\rm d} x \\
L((\tau, v)) &= - \int_{\Omega} f v \ {\rm d} x
+ \int_{\Gamma_D} u_0 \tau \cdot n \ {\rm d} s
The essential boundary condition is reflected in function spaces
:math:`\Sigma_g = \{ \tau \in H({\rm div}) \text{ such that } \tau \cdot
n|_{\Gamma_N} = g \}` and :math:`V = L^2(\Omega)`.
We need to choose a stable combination of discrete function spaces
:math:`\Sigma_h \subset \Sigma` and :math:`V_h \subset V` to form a mixed
function space :math:`\Sigma_h \times V_h`. One such choice is
Brezzi-Douglas-Marini elements of polynomial order :math:`k` for
:math:`\Sigma_h` and discontinuous elements of polynomial order :math:`k-1`
for :math:`V_h`.
For the remaining functions and boundaries we choose:
.. math::
\Gamma_{D} = \{(0, y) \cup (1, y) \in \partial \Omega\},
\Gamma_{N} = \{(x, 0) \cup (x, 1) \in \partial \Omega\}
u_0 = 0,
g = \sin(5x)
f = 10~e^{-\frac{(x - 0.5)^2 + (y - 0.5)^2}{0.02}}
To produce a numerical solution to this PDE in Firedrake we procede as
follows:
The mesh is chosen as a :math:`32\times32` element unit square. ::
from firedrake import *
mesh = UnitSquareMesh(32, 32)
As argued above, a stable choice of function spaces for our problem is the
combination of order :math:`k` Brezzi-Douglas-Marini (BDM) elements and order
:math:`k - 1` discontinuous Galerkin elements (DG). We use :math:`k = 1` and
combine the BDM and DG spaces into a mixed function space ``W``. ::
BDM = FunctionSpace(mesh, "BDM", 1)
DG = FunctionSpace(mesh, "DG", 0)
W = BDM * DG
We obtain test and trial functions on the subspaces of the mixed function
spaces as follows: ::
sigma, u = TrialFunctions(W)
tau, v = TestFunctions(W)
Next we declare our source function ``f`` over the DG space and initialise it
with our chosen right hand side function value. ::
f = Function(DG).interpolate(Expression(
"10*exp(-(pow(x[0] - 0.5, 2) + pow(x[1] - 0.5, 2)) / 0.02)"))
After dropping the vanishing boundary term on the right hand side, the
bilinear and linear forms of the variational problem are defined as: ::
a = (dot(sigma, tau) + div(tau)*u + div(sigma)*v)*dx
L = - f*v*dx
The strongly enforced boundary conditions on the BDM space on the top and
bottom of the domain are declared as: ::
bc0 = DirichletBC(W.sub(0), Expression(("0.0", "-sin(5*x[0])")), 1)
bc1 = DirichletBC(W.sub(0), Expression(("0.0", "sin(5*x[0])")), 2)
Note that it is necessary to apply these boundary conditions to the first
subspace of the mixed function space using ``W.sub(0)``. This way the
association with the mixed space is preserved. Declaring it on the BDM space
directly is *not* the same and would in fact cause the application of the
boundary condition during the later solve to fail.
Now we're ready to solve the variational problem. We define `w` to be a function
to hold the solution on the mixed space. ::
w = Function(W)
Then we solve the linear variational problem ``a == L`` for ``w`` under the
given boundary conditions ``bc0`` and ``bc1``. Afterwards we extract the
components ``sigma`` and ``u`` on each of the subspaces with ``split``. ::
solve(a == L, w, bcs=[bc0, bc1])
sigma, u = w.split()
Lastly we write the component of the solution corresponding to the primal
variable on the DG space to a file in VTK format for later inspection with a
visualisation tool such as `ParaView <http://www.paraview.org/>`__ ::
File("poisson_mixed.pvd") << u
This demo is based on the corresponding `DOLFIN mixed Poisson demo
<http://fenicsproject.org/documentation/dolfin/1.3.0/python/demo/documented/mixed-poisson/python/documentation.html>`__
and can be found as a script in `poisson_mixed.py <poisson_mixed.py>`__.
<file_sep>/firedrake/variational_solver.py
import ufl
from pyop2.logger import warning, RED
from pyop2.profiling import timed_function, profile
import assemble
import function
import solving
import solving_utils
import ufl_expr
from petsc import PETSc
__all__ = ["LinearVariationalProblem",
"LinearVariationalSolver",
"NonlinearVariationalProblem",
"NonlinearVariationalSolver"]
class NonlinearVariationalProblem(object):
"""Nonlinear variational problem F(u; v) = 0."""
def __init__(self, F, u, bcs=None, J=None,
Jp=None,
form_compiler_parameters=None):
"""
:param F: the nonlinear form
:param u: the :class:`.Function` to solve for
:param bcs: the boundary conditions (optional)
:param J: the Jacobian J = dF/du (optional)
:param Jp: a form used for preconditioning the linear system,
optional, if not supplied then the Jacobian itself
will be used.
:param dict form_compiler_parameters: parameters to pass to the form
compiler (optional)
"""
# Extract and check arguments
u = solving._extract_u(u)
bcs = solving._extract_bcs(bcs)
# Store input UFL forms and solution Function
self.F_ufl = F
# Use the user-provided Jacobian. If none is provided, derive
# the Jacobian from the residual.
self.J_ufl = J or ufl_expr.derivative(F, u)
self.Jp = Jp
self.u_ufl = u
self.bcs = bcs
# Store form compiler parameters
self.form_compiler_parameters = form_compiler_parameters
self._constant_jacobian = False
class NonlinearVariationalSolver(object):
"""Solves a :class:`NonlinearVariationalProblem`."""
_id = 0
def __init__(self, *args, **kwargs):
"""
:arg problem: A :class:`NonlinearVariationalProblem` to solve.
:kwarg nullspace: an optional :class:`.VectorSpaceBasis` (or
:class:`.MixedVectorSpaceBasis`) spanning the null
space of the operator.
:kwarg solver_parameters: Solver parameters to pass to PETSc.
This should be a dict mapping PETSc options to values. For
example, to set the nonlinear solver type to just use a linear
solver:
.. code-block:: python
{'snes_type': 'ksponly'}
PETSc flag options should be specified with `bool` values. For example:
.. code-block:: python
{'snes_monitor': True}
.. warning ::
Since this object contains a circular reference and a
custom ``__del__`` attribute, you *must* call :meth:`.destroy`
on it when you are done, otherwise it will never be
garbage collected.
"""
assert isinstance(args[0], NonlinearVariationalProblem)
self._problem = args[0]
# Build the jacobian with the correct sparsity pattern. Note
# that since matrix assembly is lazy this doesn't actually
# force an additional assembly of the matrix since in
# form_jacobian we call assemble again which drops this
# computation on the floor.
self._jac_tensor = assemble.assemble(self._problem.J_ufl, bcs=self._problem.bcs,
form_compiler_parameters=self._problem.form_compiler_parameters)
if self._problem.Jp is not None:
self._jac_ptensor = assemble.assemble(self._problem.Jp, bcs=self._problem.bcs,
form_compiler_parameters=self._problem.form_compiler_parameters)
else:
self._jac_ptensor = self._jac_tensor
test = self._problem.F_ufl.arguments()[0]
self._F_tensor = function.Function(test.function_space())
# Function to hold current guess
self._x = function.Function(self._problem.u_ufl)
self._problem.F_ufl = ufl.replace(self._problem.F_ufl, {self._problem.u_ufl:
self._x})
self._problem.J_ufl = ufl.replace(self._problem.J_ufl, {self._problem.u_ufl:
self._x})
if self._problem.Jp is not None:
self._problem.Jp = ufl.replace(self._problem.Jp, {self._problem.u_ufl:
self._x})
self._jacobian_assembled = False
self.snes = PETSc.SNES().create()
self._opt_prefix = 'firedrake_snes_%d_' % NonlinearVariationalSolver._id
NonlinearVariationalSolver._id += 1
self.snes.setOptionsPrefix(self._opt_prefix)
parameters = kwargs.get('solver_parameters', None)
if 'parameters' in kwargs:
warning(RED % "The 'parameters' keyword to %s is deprecated, use 'solver_parameters' instead.",
self.__class__.__name__)
parameters = kwargs['parameters']
if 'solver_parameters' in kwargs:
warning(RED % "'parameters' and 'solver_parameters' passed to %s, using the latter",
self.__class__.__name__)
parameters = kwargs['solver_parameters']
# Make sure we don't stomp on a dict the user has passed in.
parameters = parameters.copy() if parameters is not None else {}
# Mixed problem, use jacobi pc if user has not supplied one.
if self._jac_tensor._M.sparsity.shape != (1, 1):
parameters.setdefault('pc_type', 'jacobi')
self.parameters = parameters
ksp = self.snes.getKSP()
pc = ksp.getPC()
pmat = self._jac_ptensor._M
names = [fs.name if fs.name else str(i)
for i, fs in enumerate(test.function_space())]
ises = solving_utils.set_fieldsplits(pmat, pc, names=names)
with self._F_tensor.dat.vec as v:
self.snes.setFunction(self.form_function, v)
self.snes.setJacobian(self.form_jacobian, J=self._jac_tensor._M.handle,
P=self._jac_ptensor._M.handle)
nullspace = kwargs.get('nullspace', None)
if nullspace is not None:
self.set_nullspace(nullspace, ises=ises)
def set_nullspace(self, nullspace, ises=None):
"""Set the null space for this solver.
:arg nullspace: a :class:`.VectorSpaceBasis` spanning the null
space of the operator.
This overwrites any existing null space."""
nullspace._apply(self._jac_tensor._M, ises=ises)
if self._problem.Jp is not None:
nullspace._apply(self._jac_ptensor._M, ises=ises)
def form_function(self, snes, X_, F_):
# X_ may not be the same vector as the vec behind self._x, so
# copy guess in from X_.
with self._x.dat.vec as v:
if v != X_:
with v as _v, X_ as _x:
_v[:] = _x[:]
assemble.assemble(self._problem.F_ufl, tensor=self._F_tensor,
form_compiler_parameters=self._problem.form_compiler_parameters)
for bc in self._problem.bcs:
bc.zero(self._F_tensor)
# F_ may not be the same vector as self._F_tensor, so copy
# residual out to F_.
with self._F_tensor.dat.vec_ro as v:
if F_ != v:
with v as _v, F_ as _f:
_f[:] = _v[:]
def form_jacobian(self, snes, X_, J_, P_):
if self._problem._constant_jacobian and self._jacobian_assembled:
# Don't need to do any work with a constant jacobian
# that's already assembled
return
self._jacobian_assembled = True
# X_ may not be the same vector as the vec behind self._x, so
# copy guess in from X_.
with self._x.dat.vec as v:
if v != X_:
with v as _v, X_ as _x:
_v[:] = _x[:]
assemble.assemble(self._problem.J_ufl,
tensor=self._jac_tensor,
bcs=self._problem.bcs,
form_compiler_parameters=self._problem.form_compiler_parameters)
self._jac_tensor.M._force_evaluation()
if self._problem.Jp is not None:
assemble.assemble(self._problem.Jp,
tensor=self._jac_ptensor,
bcs=self._problem.bcs,
form_compiler_parameters=self._problem.form_compiler_parameters)
self._jac_ptensor.M._force_evaluation()
return PETSc.Mat.Structure.DIFFERENT_NONZERO_PATTERN
return PETSc.Mat.Structure.SAME_NONZERO_PATTERN
def __del__(self):
# Remove stuff from the options database
# It's fixed size, so if we don't it gets too big.
if hasattr(self, '_opt_prefix'):
opts = PETSc.Options()
for k in self.parameters.iterkeys():
del opts[self._opt_prefix + k]
delattr(self, '_opt_prefix')
def destroy(self):
"""Destroy the SNES object inside the solver.
You must call this explicitly, because the SNES holds a
reference to the solver it lives inside, defeating the garbage
collector."""
if self.snes is not None:
self.snes.destroy()
self.snes = None
@property
def parameters(self):
return self._parameters
@parameters.setter
def parameters(self, val):
assert isinstance(val, dict), 'Must pass a dict to set parameters'
self._parameters = val
solving_utils.update_parameters(self, self.snes)
@timed_function("SNES solver execution")
@profile
def solve(self):
# Apply the boundary conditions to the initial guess.
for bc in self._problem.bcs:
bc.apply(self._problem.u_ufl)
# User might have updated parameters dict before calling
# solve, ensure these are passed through to the snes.
solving_utils.update_parameters(self, self.snes)
with self._problem.u_ufl.dat.vec as v:
self.snes.solve(None, v)
reasons = self.snes.ConvergedReason()
reasons = dict([(getattr(reasons, r), r)
for r in dir(reasons) if not r.startswith('_')])
r = self.snes.getConvergedReason()
try:
reason = reasons[r]
inner = False
except KeyError:
kspreasons = self.snes.getKSP().ConvergedReason()
kspreasons = dict([(getattr(kspreasons, kr), kr)
for kr in dir(kspreasons) if not kr.startswith('_')])
r = self.snes.getKSP().getConvergedReason()
try:
reason = kspreasons[r]
inner = True
except KeyError:
reason = 'unknown reason (petsc4py enum incomplete?)'
if r < 0:
if inner:
msg = "Inner linear solve failed to converge after %d iterations with reason: %s" % \
(self.snes.getKSP().getIterationNumber(), reason)
else:
msg = reason
raise RuntimeError("""Nonlinear solve failed to converge after %d nonlinear iterations.
Reason:
%s""" % (self.snes.getIterationNumber(), msg))
class LinearVariationalProblem(NonlinearVariationalProblem):
"""Linear variational problem a(u, v) = L(v)."""
def __init__(self, a, L, u, bcs=None, aP=None,
form_compiler_parameters=None,
constant_jacobian=True):
"""
:param a: the bilinear form
:param L: the linear form
:param u: the :class:`.Function` to solve for
:param bcs: the boundary conditions (optional)
:param aP: an optional operator to assemble to precondition
the system (if not provided a preconditioner may be
computed from ``a``)
:param dict form_compiler_parameters: parameters to pass to the form
compiler (optional)
:param constant_jacobian: (optional) flag indicating that the
Jacobian is constant (i.e. does not depend on
varying fields). If your Jacobian can change, set
this flag to :data:`False`.
"""
# In the linear case, the Jacobian is the equation LHS.
J = a
F = ufl.action(J, u) - L
super(LinearVariationalProblem, self).__init__(F, u, bcs, J, aP,
form_compiler_parameters=form_compiler_parameters)
self._constant_jacobian = constant_jacobian
class LinearVariationalSolver(NonlinearVariationalSolver):
"""Solves a :class:`LinearVariationalProblem`."""
def __init__(self, *args, **kwargs):
"""
:arg problem: A :class:`LinearVariationalProblem` to solve.
:kwarg solver_parameters: Solver parameters to pass to PETSc.
This should be a dict mapping PETSc options to values.
:kwarg nullspace: an optional :class:`.VectorSpaceBasis` (or
:class:`.MixedVectorSpaceBasis`) spanning the null
space of the operator.
.. warning ::
Since this object contains a circular reference and a
custom ``__del__`` attribute, you *must* call :meth:`.destroy`
on it when you are done, otherwise it will never be
garbage collected.
"""
super(LinearVariationalSolver, self).__init__(*args, **kwargs)
self.parameters.setdefault('snes_type', 'ksponly')
self.parameters.setdefault('ksp_rtol', 1.0e-7)
solving_utils.update_parameters(self, self.snes)
<file_sep>/tests/extrusion/test_extrusion_unit_square.py
import pytest
import numpy as np
from firedrake import *
import pyop2 as op2
from pyop2.profiling import *
def integrate_unit_square(family, degree):
power = 5
m = UnitIntervalMesh(2 ** power)
layers = 10
# Populate the coordinates of the extruded mesh by providing the
# coordinates as a field.
# A kernel which describes how coordinates are extruded.
mesh = ExtrudedMesh(m, layers, layer_height=0.1)
fs = FunctionSpace(mesh, family, degree, name="fs")
f = Function(fs)
area = op2.Kernel("""
void comp_area(double A[1], double *x[], double *y[])
{
double area = (x[1][1]-x[0][1])*(x[2][0]-x[0][0]);
if (area < 0)
area = area * (-1.0);
A[0] += area;
}""", "comp_area")
g = op2.Global(1, data=0.0, name='g')
coords = f.function_space().mesh().coordinates
op2.par_loop(area, f.cell_set,
g(op2.INC),
coords.dat(op2.READ, coords.cell_node_map()),
f.dat(op2.READ, f.cell_node_map())
)
return np.abs(g.data[0] - 1.0)
def test_firedrake_extrusion_unit_square():
family = "Lagrange"
degree = 1
assert integrate_unit_square(family, degree) < 1.0e-12
if __name__ == '__main__':
import os
pytest.main(os.path.abspath(__file__))
| 2dd1deb89047bdc540f5ec568567c89d5f58fa2c | [
"reStructuredText",
"Markdown",
"Makefile",
"INI",
"Python",
"Text",
"C"
] | 138 | Python | pefarrell/firedrake | 673e1120e55c0eab0b5b447d0337385d571f6775 | d7dfd4d0df44ab45544fd8509fe6c32a04b18ef2 |
refs/heads/master | <repo_name>Tigana13/legitimate-tech-test<file_sep>/src/app/modules/app-common/models/column-config/column-config.ts
export interface ColumnConfig {
definition?: string;
heading?: string;
sorting?: boolean;
type?: string;
class?: string;
}
<file_sep>/src/app/_services/chamas.service.ts
import { Injectable } from '@angular/core';
import {ApiService} from "./api/api.service";
import {Router} from "@angular/router";
import {HttpParams} from "@angular/common/http";
@Injectable({
providedIn: 'root'
})
export class ChamasService {
constructor(protected apiService: ApiService, protected router: Router) {}
fetchChamas(params?: HttpParams | any) {
return this.apiService.get('/chamas/');
}
createChama(chama: {} | any) {
return this.apiService.post('/chamas/', chama);
}
createChamaSchedule(schedule: {} | any) {
return this.apiService.post('/chamas/contributions/create_schedule/', schedule);
}
generateContributionRequest(payload: {} | any) {
return this.apiService.post('/chamas/contributions/generate_request/', payload);
}
}
<file_sep>/src/app/_services/excel/excel.service.spec.ts
import { TestBed } from '@angular/core/testing';
import { ExcelService } from './excel.service';
import {of} from 'rxjs';
describe('ExcelService', () => {
beforeEach(() => TestBed.configureTestingModule({}));
it('should be created', () => {
const service: ExcelService = TestBed.get(ExcelService);
expect(service).toBeTruthy();
});
it('should save a file to excel', () => {
const service: ExcelService = TestBed.get(ExcelService);
const serviceSpy: any = jasmine.createSpyObj('ExcelService', ['saveAsExcelFile']);
serviceSpy.saveAsExcelFile.and.returnValue(of({})); expect(service.saveAsExcelFile).toBeTruthy();
});
it('should export a file to CSV', () => {
const service: ExcelService = TestBed.get(ExcelService);
const serviceSpy: any = jasmine.createSpyObj('ExcelService', ['exportToCsv']);
serviceSpy.exportToCsv.and.returnValue(of({})); expect(service.exportToCsv).toBeTruthy();
});
});
<file_sep>/src/app/_helpers/snackbar/snackbar.service.ts
import { Injectable } from '@angular/core';
import {MatSnackBar, MatSnackBarConfig, MatSnackBarHorizontalPosition, MatSnackBarVerticalPosition} from '@angular/material/snack-bar';
@Injectable({
providedIn: 'root'
})
export class SnackbarService {
constructor(private snackBar: MatSnackBar) {
}
/*****************
* @param message
* @param action
* @param duration
* @param verticalPos
* @param horizontalPos
* @param panelClass
*********************************
********* SAMPLE USAGE **********
* *******************************
* import {SnackbarService} from '../_helpers/snackbar/snackbar.service';
*
******* initialize in constructor *********
* private snackBarService: SnackbarService
*
****** open snackBar **********************
*****(pass 'undefined' to function call to opt out of a parameter)
* this.snackBarService.open('Welcome', 'Dismiss', undefined, undefined, undefined, ['snack-bar', 'snack-bar-success']);
*/
openSnackBar(message: string,
action?: string,
duration?: number,
verticalPos?: MatSnackBarVerticalPosition,
horizontalPos?: MatSnackBarHorizontalPosition,
panelClass?: string[]) {
const config: any = new MatSnackBarConfig();
config.duration = duration || 150000;
config.horizontalPosition = horizontalPos || 'center';
config.verticalPosition = verticalPos || 'top';
config.panelClass = panelClass || ['snack-bar', 'snack-bar-success'];
return this.snackBar.open(message, action, config);
}
closeSnackBar() {
return this.snackBar.dismiss();
}
}
<file_sep>/src/app/modules/app-common/side-nav-item/side-nav-item.component.ts
import {AfterViewInit, Component, EventEmitter, HostBinding, Input, OnInit, Output} from '@angular/core';
import {Router} from '@angular/router';
import {animate, state, style, transition, trigger} from '@angular/animations';
import {Observable} from 'rxjs';
import {SidenavToggleService} from '../../../_services/sidenav/sidenav-toggle.service';
import {MatIconRegistry} from '@angular/material/icon';
import {DomSanitizer} from '@angular/platform-browser';
@Component({
selector: 'app-side-nav-item',
templateUrl: './side-nav-item.component.html',
styleUrls: ['./side-nav-item.component.scss'],
animations: [
trigger('indicatorRotate', [
state('collapsed', style({transform: 'rotate(0deg)'})),
state('expanded', style({transform: 'rotate(180deg)'})),
transition('expanded <=> collapsed',
animate('225ms cubic-bezier(0.4,0.0,0.2,1)')
),
])
]
})
export class SideNavItemComponent implements OnInit, AfterViewInit {
expanded: boolean;
@HostBinding('attr.aria-expanded') ariaExpanded = this.expanded;
@Input() item;
@Input() depth: number;
@Input() toggleEvent: EventEmitter<any>;
sideNavToggled: any = true;
constructor(
private matIconRegistry: MatIconRegistry,
private domSanitizer: DomSanitizer,
public router: Router,
private sidenavToggleService: SidenavToggleService,
) {
if (this.depth === undefined) {
this.depth = 0;
}
}
ngOnInit() {
this.registerMatIcon();
this.expanded = this.router.isActive(this.item.route, false);
this.toggleEvent.subscribe(toggleEvent => {
this.sideNavToggled = toggleEvent.open;
});
this.sidenavToggleService.toggleState.subscribe(state => {
this.sideNavToggled = state;
});
}
ngAfterViewInit() {
}
onItemSelected(item) {
if (!item.children || !item.children.length) {
this.router.navigate([item.route]);
}
if (item.children && item.children.length) {
this.expanded = !this.expanded;
}
}
registerMatIcon() {
this.matIconRegistry.addSvgIcon(this.item.title, this.domSanitizer.bypassSecurityTrustResourceUrl(this.item.iconURL));
}
}
<file_sep>/src/app/_helpers/directives/modals/add-product-modal/add-product-modal.directive.spec.ts
import { AddProductModalDirective } from './add-product-modal.directive';
describe('AddStoreModalDirective', () => {
it('should create an instance', () => {
const directive = new AddProductModalDirective();
expect(directive).toBeTruthy();
});
});
<file_sep>/src/app/_services/users.service.ts
import { Injectable } from '@angular/core';
import {ApiService} from "./api/api.service";
import {Router} from "@angular/router";
import {HttpParams} from "@angular/common/http";
@Injectable({
providedIn: 'root'
})
export class UsersService {
constructor(protected apiService: ApiService, protected router: Router) {}
fetchUsers(params?: HttpParams | any) {
return this.apiService.get('/users/', params);
}
}
<file_sep>/src/app/modules/app-common/side-nav/side-nav.component.ts
import {Component, EventEmitter, OnInit} from '@angular/core';
import {Observable, of} from 'rxjs';
import { BreakpointObserver, Breakpoints } from '@angular/cdk/layout';
import { map } from 'rxjs/operators';
import {animate, state, style, transition, trigger} from '@angular/animations';
import { Router } from '@angular/router';
import {SidenavToggleService} from '../../../_services/sidenav/sidenav-toggle.service';
import {MatIconRegistry} from '@angular/material/icon';
import {DomSanitizer} from '@angular/platform-browser';
@Component({
selector: 'app-side-nav',
templateUrl: './side-nav.component.html',
styleUrls: ['./side-nav.component.scss'],
animations: [
trigger('indicatorRotate', [
state('collapsed', style({transform: 'rotate(0deg)'})),
state('expanded', style({transform: 'rotate(180deg)'})),
transition('expanded <=> collapsed',
animate('225ms cubic-bezier(0.4,0.0,0.2,1)')
),
])
]
})
export class SideNavComponent implements OnInit {
sidenavWidth: Observable<any> = of(20);
toggleEvent: EventEmitter<any> = new EventEmitter<any>();
sidenavToggled: Observable<any> | any = this.sidenavToggleService.toggleState.value;
singleProfileSideItem: any = {
title: 'Profile',
icon: 'person',
route: '/jobs/available',
iconURL: '/assets/SVG/user-white.svg'
}
routes: any[] = [
{
title: 'Dashboard',
icon: 'compass',
route: '/jobs/available',
iconURL: '/assets/SVG/dashboard.svg'
},
{
title: 'Make a request',
icon: 'hour-glass',
route: '/jobs/current',
iconURL: '/assets/SVG/comment.svg',
},
{
title: 'Bulk request',
icon: 'referral',
route: '/jobs/revision',
iconURL: '/assets/SVG/conversation.svg'
},
{
title: 'Reports',
icon: 'court',
route: '/jobs/disputed',
iconURL: '/assets/SVG/paper.svg'
},
{
title: 'Payments',
icon: 'delivery-complete',
route: '/jobs/finished',
iconURL: '/assets/SVG/dollar.svg'
},
{
title: 'Downloads',
icon: 'gavel',
route: '/jobs/bids',
iconURL: '/assets/SVG/download.svg',
},
{
title: 'FAQs',
icon: 'gavel',
route: '/trips/list',
iconURL: '/assets/SVG/faq.svg',
},
{
title: 'APIs',
icon: 'gavel',
route: '/trips/list',
iconURL: '/assets/SVG/api.svg',
},
];
isHandset$: Observable<boolean> = this.breakpointObserver.observe(Breakpoints.Handset).pipe(map(result => result.matches));
constructor(
private breakpointObserver: BreakpointObserver,
private sidenavToggleService: SidenavToggleService,
public router: Router
) {}
ngOnInit() {
const membership = JSON.parse(localStorage.getItem('membership'));
this.routes = this.routes.filter(route => {
if (route.title === 'Merchandising') {
if (membership && membership.is_merchandiser) {
return route;
}
return;
}
if (route.title === 'Distributors') {
if (membership && membership.business_category === 'MANUFACTURER') {
return route;
}
return;
}
if (route.title === 'Customers') {
if (membership && membership.customer_categorization === 'INDIVIDUAL') {
return route;
}
if (membership && membership.customer_categorization === 'BOTH') {
return route;
}
return;
}
if (route.title === 'Outlets') {
if (membership && membership.customer_categorization === 'BUSINESS') {
return route;
}
if (membership && membership.customer_categorization === 'BOTH') {
return route;
}
return;
}
return route;
});
}
toggleSidenav() {
this.sidenavToggleService.toggleSidenavStatus();
this.sidenavToggleService.toggleState.subscribe(status => {
this.sidenavToggled = status;
this.toggleEvent.emit({open: status});
});
}
// Return observable form of a value
of(value) {
return of(value);
}
}
<file_sep>/src/app/_helpers/sanitizers/phoneNumber-sanitizer.directive.ts
export function sanitizePhoneNumber(phoneNumber: string) {
// convert phoneNumber to string
phoneNumber = phoneNumber.toString();
return phoneNumber.replace(/ +/g, '').replace(' ', '').replace(/[-]/, '').replace(/[)]/, '');
}
<file_sep>/src/app/_helpers/directives/modals/receive-product/receive-product.directive.spec.ts
import { ReceiveProductDirective } from './receive-product.directive';
describe('ReceiveProductDirective', () => {
it('should create an instance', () => {
const directive = new ReceiveProductDirective();
expect(directive).toBeTruthy();
});
});
<file_sep>/src/app/_helpers/directives/modals/receive-product/receive-product.directive.ts
import { Directive } from '@angular/core';
@Directive({
selector: '[appReceiveProduct]'
})
export class ReceiveProductDirective {
constructor() { }
}
<file_sep>/src/app/modules/app-common/top-nav/top-nav.component.ts
import {Component, OnInit, Output, EventEmitter, Input} from '@angular/core';
import {Router, ActivatedRoute} from '@angular/router';
import {BreadCrumbService} from '../../../_services/bread-crumb/bread-crumb.service';
import { DomSanitizer } from '@angular/platform-browser';
import {SidenavToggleService} from '../../../_services/sidenav/sidenav-toggle.service';
import {MatDialog} from '@angular/material/dialog';
import {MatIconRegistry} from '@angular/material/icon';
import {SnackbarService} from '../../../_helpers/snackbar/snackbar.service';
@Component({
selector: 'app-top-nav',
templateUrl: './top-nav.component.html',
styleUrls: ['./top-nav.component.scss']
})
export class TopNavComponent implements OnInit {
public currentUserDetails: any;
public title: string;
@Input() toggleEvent: EventEmitter<any>;
@Output() toggleSidenav = new EventEmitter<void>();
sideNavToggled: any = true;
constructor(
private snackBar: SnackbarService,
private router: Router,
private dialog: MatDialog,
private route: ActivatedRoute,
public breadCrumbService: BreadCrumbService,
private matIconRegistry: MatIconRegistry,
private domSanitizer: DomSanitizer,
private sidenavToggleService: SidenavToggleService
) {
this.matIconRegistry.addSvgIcon('Help', this.domSanitizer.bypassSecurityTrustResourceUrl('/assets/SVG/Help.svg') );
this.matIconRegistry.addSvgIcon('Chat', this.domSanitizer.bypassSecurityTrustResourceUrl('/assets/SVG/chat.svg') );
this.matIconRegistry.addSvgIcon('Person', this.domSanitizer.bypassSecurityTrustResourceUrl('/assets/SVG/Blue/avatar.svg') );
this.matIconRegistry.addSvgIcon('Books', this.domSanitizer.bypassSecurityTrustResourceUrl('/assets/SVG/books.svg') ); }
ngOnInit() {
this.breadCrumbService.title.subscribe(title => {
this.title = title;
});
this.currentUserDetails = JSON.parse(localStorage.getItem('currentUser'));
this.toggleEvent.subscribe(toggleEvent => {
this.sideNavToggled = toggleEvent.open;
});
this.sidenavToggleService.toggleState.subscribe(state => {
this.sideNavToggled = state;
});
}
openConfirmLogoutDialog() {
}
currentUser() {
return this.currentUserDetails = {};
}
}
<file_sep>/src/app/_helpers/directives/modals/add-order-modal/add-order-modal.directive.spec.ts
import { AddOrderModalDirective } from './add-order-modal.directive';
describe('AddOrderModalDirective', () => {
it('should create an instance', () => {
const directive = new AddOrderModalDirective();
expect(directive).toBeTruthy();
});
});
<file_sep>/src/app/modules/app-common/models/column-config/column-config.spec.ts
import { ColumnConfig } from './column-config';
describe('ColumnConfig', () => {
it('should create an instance', () => {
expect(new ColumnConfig()).toBeTruthy();
});
});
<file_sep>/src/app/modules/app-common/app-common.module.ts
import {NgModule, NO_ERRORS_SCHEMA} from '@angular/core';
import {CommonModule, DatePipe} from '@angular/common';
import {LayoutModule} from '@angular/cdk/layout';
import { FlexLayoutModule } from '@angular/flex-layout';
import { RouterModule} from '@angular/router';
import { FormsModule, ReactiveFormsModule} from '@angular/forms';
import { CdkTreeModule} from '@angular/cdk/tree';
import { SideNavComponent } from './side-nav/side-nav.component';
import { TopNavComponent } from './top-nav/top-nav.component';
import { CardsWidgetComponent } from './cards-widget/cards-widget.component';
import { LoadingScreenComponent } from './loading-screen/loading-screen.component';
import { PermissionDeniedComponent } from './permission-denied/permission-denied.component';
import { SideNavItemComponent } from './side-nav-item/side-nav-item.component';
import {MatListModule} from '@angular/material/list';
import {MatToolbarModule} from '@angular/material/toolbar';
import {MatIconModule} from '@angular/material/icon';
import {MatFormFieldModule} from '@angular/material/form-field';
import {MatTreeModule} from '@angular/material/tree';
import {MatProgressBarModule} from '@angular/material/progress-bar';
import {MatSlideToggleModule} from '@angular/material/slide-toggle';
import {MatPaginatorModule} from '@angular/material/paginator';
import {MatInputModule} from '@angular/material/input';
import {MatDatepickerModule} from '@angular/material/datepicker';
import {MatSnackBarModule} from '@angular/material/snack-bar';
import {MatMenuModule} from '@angular/material/menu';
import {MatSortModule} from '@angular/material/sort';
import {MatCheckboxModule} from '@angular/material/checkbox';
import {MatTableModule} from '@angular/material/table';
import {MatAutocompleteModule} from '@angular/material/autocomplete';
import {MatCardModule} from '@angular/material/card';
import {MatTooltipModule} from '@angular/material/tooltip';
import {MatButtonModule} from '@angular/material/button';
import {MatSelectModule} from '@angular/material/select';
import {MatSidenavModule} from '@angular/material/sidenav';
import {MatStepperModule} from '@angular/material/stepper';
import {MatProgressSpinnerModule} from '@angular/material/progress-spinner';
import {MatRippleModule} from '@angular/material/core';
import {MatBottomSheetModule} from '@angular/material/bottom-sheet';
import {MatDialogModule} from '@angular/material/dialog';
@NgModule({
declarations: [
SideNavComponent,
TopNavComponent,
CardsWidgetComponent,
LoadingScreenComponent,
PermissionDeniedComponent,
SideNavItemComponent,
],
imports: [
CommonModule,
FormsModule,
ReactiveFormsModule,
// Material
LayoutModule,
MatToolbarModule,
MatButtonModule,
MatSidenavModule,
MatIconModule,
MatListModule,
RouterModule,
MatSnackBarModule,
MatDialogModule,
MatFormFieldModule,
MatDatepickerModule,
MatCardModule,
MatSnackBarModule,
MatTreeModule,
CdkTreeModule,
MatIconModule,
MatStepperModule,
MatCheckboxModule,
MatMenuModule,
FlexLayoutModule,
MatInputModule,
MatSelectModule,
MatTableModule,
MatProgressBarModule,
MatBottomSheetModule,
MatRippleModule,
MatSlideToggleModule,
MatAutocompleteModule,
MatSortModule,
MatPaginatorModule,
MatProgressSpinnerModule,
MatTooltipModule
],
exports: [
CardsWidgetComponent,
LoadingScreenComponent,
PermissionDeniedComponent,
SideNavItemComponent,
TopNavComponent,
],
entryComponents: [
],
providers: [
DatePipe
],
schemas: [NO_ERRORS_SCHEMA]
})
export class AppCommonModule { }
<file_sep>/src/app/modules/app-common/cards-widget/cards-widget.component.spec.ts
import { async, ComponentFixture, TestBed } from '@angular/core/testing';
import { CardsWidgetComponent } from './cards-widget.component';
import {CUSTOM_ELEMENTS_SCHEMA, NO_ERRORS_SCHEMA} from '@angular/core';
describe('CardsWidgetComponent', () => {
let component: CardsWidgetComponent;
let fixture: ComponentFixture<CardsWidgetComponent>;
beforeEach(async(() => {
TestBed.configureTestingModule({
declarations: [ CardsWidgetComponent ],
schemas: [ CUSTOM_ELEMENTS_SCHEMA, NO_ERRORS_SCHEMA ]
})
.compileComponents();
}));
beforeEach(() => {
fixture = TestBed.createComponent(CardsWidgetComponent);
component = fixture.componentInstance;
fixture.detectChanges();
});
});
<file_sep>/src/app/modules/app-common/cards-widget/cards-widget.component.ts
import { Component, OnInit, Input, Output, EventEmitter } from '@angular/core';
@Component({
selector: 'app-cards-widget',
templateUrl: './cards-widget.component.html',
styleUrls: ['./cards-widget.component.scss']
})
export class CardsWidgetComponent implements OnInit {
@Input() cards: any[];
@Output() filter = new EventEmitter<any>();
constructor() { }
ngOnInit() {
//
}
clicked(clickedCard: any) {
this.filter.emit(clickedCard);
}
}
<file_sep>/src/app/_helpers/directives/modals/add-product-modal/add-product-modal.directive.ts
import {Directive, ViewContainerRef} from '@angular/core';
@Directive({
selector: '[appAddProductModalDirective]'
})
export class AddProductModalDirective {
constructor(public viewContainerRef?: ViewContainerRef) { }
}
<file_sep>/src/app/_services/contributions.service.ts
import { Injectable } from '@angular/core';
import {ApiService} from "./api/api.service";
import {Router} from "@angular/router";
import {HttpParams} from "@angular/common/http";
@Injectable({
providedIn: 'root'
})
export class ContributionsService {
constructor(protected apiService: ApiService, protected router: Router) {}
fetchContributions(params?: HttpParams | any) {
return this.apiService.get('/group-contributions/');
}
createContribution(donation: {} | any) {
return this.apiService.post('/group-contributions/', donation);
}
}
<file_sep>/src/app/modules/app-common/models/field-config/field-config.spec.ts
import { FieldConfig } from './field-config';
describe('FieldConfig', () => {
it('should create an instance', () => {
expect(new FieldConfig()).toBeTruthy();
});
});
<file_sep>/src/app/_services/donation.service.ts
import { Injectable } from '@angular/core';
import {ApiService} from "./api/api.service";
import {Router} from "@angular/router";
import {HttpParams} from "@angular/common/http";
@Injectable({
providedIn: 'root'
})
export class DonationService {
constructor(protected apiService: ApiService, protected router: Router) {}
fetchDonations(params?: HttpParams | any) {
return this.apiService.get('/donations/');
}
createDonation(donation: {} | any) {
return this.apiService.post('/donations/', donation);
}
}
<file_sep>/src/app/_services/excel/excel.service.ts
import { Injectable } from '@angular/core';
import * as FileSaver from 'file-saver';
import * as XLSX from 'xlsx';
const EXCEL_TYPE = 'application/vnd.openxmlformats-officedocument.spreadsheetml.sheet;charset=UTF-8';
const EXCEL_EXTENSION = '.xlsx';
@Injectable({
providedIn: 'root'
})
export class ExcelService {
constructor() { }
exportAsExcelFile(json: any[], excelFileName: string): void {
const worksheet: XLSX.WorkSheet = XLSX.utils.json_to_sheet(json);
const workbook: XLSX.WorkBook = { Sheets: { 'data': worksheet }, SheetNames: ['data'] };
const excelBuffer: any = XLSX.write(workbook, { bookType: 'xlsx', type: 'array' });
this.saveAsExcelFile(excelBuffer, excelFileName);
}
exportToCsv(filename, rows) {
var processRow = function (row) {
var finalVal = '';
for (var j = 0; j < row.length; j++) {
var innerValue = row[j] === null ? '' : row[j].toString();
if (row[j] instanceof Date) {
innerValue = row[j].toLocaleString();
};
var result = innerValue.replace(/"/g, '""');
if (result.search(/("|,|\n)/g) >= 0)
result = '"' + result + '"';
if (j > 0)
finalVal += ',';
finalVal += result;
}
return finalVal + '\n';
};
var csvFile = '';
for (var i = 0; i < rows.length; i++) {
csvFile += processRow(rows[i]);
}
var blob = new Blob([csvFile], { type: 'text/csv;charset=utf-8;' });
if (navigator.msSaveBlob) { // IE 10+
navigator.msSaveBlob(blob, filename);
} else {
var link = document.createElement("a");
if (link.download !== undefined) { // feature detection
// Browsers that support HTML5 download attribute
var url = URL.createObjectURL(blob);
link.setAttribute("href", url);
link.setAttribute("download", filename);
link.style.visibility = 'hidden';
document.body.appendChild(link);
link.click();
document.body.removeChild(link);
}
}
}
saveAsExcelFile(buffer: any, fileName: string): void {
const data: Blob = new Blob([buffer], {type: EXCEL_TYPE});
FileSaver.saveAs(data, fileName + '_export_' + new Date().getTime() + EXCEL_EXTENSION);
}
}
<file_sep>/src/app/_helpers/directives/validators/custom-validator.directive.ts
import {AbstractControl, ValidatorFn} from '@angular/forms';
export function forbiddenTextPattern( pattern: RegExp): ValidatorFn {
return (control: AbstractControl): {[key: string]: any} | null => {
const forbidden = pattern.test(control.value);
return forbidden ? {forbiddenPattern: {value: control.value}} : null;
};
}
<file_sep>/src/app/modules/app-common/side-nav-item/side-nav-item.component.spec.ts
import { async, ComponentFixture, TestBed } from '@angular/core/testing';
import { SideNavItemComponent } from './side-nav-item.component';
import {CUSTOM_ELEMENTS_SCHEMA, NO_ERRORS_SCHEMA} from '@angular/core';
import {RouterTestingModule} from '@angular/router/testing';
fdescribe('SideNavItemComponent', () => {
let component: SideNavItemComponent;
let fixture: ComponentFixture<SideNavItemComponent>;
beforeEach(async(() => {
TestBed.configureTestingModule({
imports: [
RouterTestingModule.withRoutes([])
],
declarations: [ SideNavItemComponent ],
schemas: [ CUSTOM_ELEMENTS_SCHEMA, NO_ERRORS_SCHEMA ]
})
.compileComponents();
component.item = {route: '', children: {}};
}));
beforeEach(() => {
fixture = TestBed.createComponent(SideNavItemComponent);
component = fixture.componentInstance;
fixture.detectChanges();
});
it('should create', () => {
expect(component).toBeTruthy();
});
});
<file_sep>/src/app/_services/excel-export/excel-export.service.ts
import { Injectable } from '@angular/core';
import { saveAs } from 'file-saver';
import { ApiService } from '../api/api.service';
import { HttpHeaders, HttpParams } from '@angular/common/http';
import { SnackbarService } from 'src/app/_helpers/snackbar/snackbar.service';
@Injectable({
providedIn: 'root'
})
export class ExcelExportService {
constructor(
private apiService: ApiService,
private snackBar: SnackbarService,
) { }
export(url, fileName, fileType = 'application/vnd.openxmlformats-officedocument.spreadsheetml.sheet; charset=utf-8;', responseType = 'blob') {
let params = new HttpParams();
let headers = new HttpHeaders();
headers = headers.set('Content-Type', fileType);
this.snackBar.openSnackBar(
'Export successfully initiated you will be notified to save the file when done',
'',
8000,
undefined,
undefined,
['snack-bar', 'snack-bar-success']
);
this.apiService.get(url, true, params, headers, responseType).subscribe(response => {
const blob = new Blob([response], { type: fileType });
saveAs(blob, fileName);
});
}
}
<file_sep>/src/app/modules/home/home.module.ts
import { NgModule } from '@angular/core';
import { CommonModule } from '@angular/common';
import { HomeRoutingModule } from './home-routing.module';
import { DashboardComponent } from './components/dashboard/dashboard.component';
import {NgbModule} from '@ng-bootstrap/ng-bootstrap';
import {MatFormFieldModule} from '@angular/material/form-field';
import {MatSelectModule} from '@angular/material/select';
@NgModule({
declarations: [DashboardComponent],
imports: [
CommonModule,
HomeRoutingModule,
NgbModule,
MatFormFieldModule,
MatSelectModule
]
})
export class HomeModule { }
<file_sep>/src/app/_helpers/directives/analytics/sample-chart.directive.ts
import {Directive, ViewContainerRef} from '@angular/core';
@Directive({
selector: '[appSampleChart]'
})
export class SampleChartDirective {
constructor(public viewContainerRef?: ViewContainerRef) { }
}
<file_sep>/src/app/_services/registration/registration.service.ts
import { Injectable } from '@angular/core';
import {ApiService} from '../api/api.service';
import {Router} from '@angular/router';
@Injectable({
providedIn: 'root'
})
export class RegistrationService {
constructor(protected apiService: ApiService, protected router: Router) {}
currentUser() {
return this.apiService.get('/auth/user/', true);
}
register(payload) {
return this.apiService.post('/register/', payload);
}
}
<file_sep>/src/app/_services/registration/registration.service.spec.ts
import { TestBed } from '@angular/core/testing';
import { RegistrationService } from './registration.service';
import {HttpClientTestingModule} from '@angular/common/http/testing';
import {RouterTestingModule} from '@angular/router/testing';
import {of} from 'rxjs';
describe('RegistrationService', () => {
beforeEach(() => TestBed.configureTestingModule({
imports: [
HttpClientTestingModule,
RouterTestingModule
]
}));
});
<file_sep>/src/app/modules/app-common/models/field-config/field-config.ts
import { Validator } from '../validator/validator';
export interface FieldConfig {
label?: string;
name?: string;
inputType?: string;
options?: string[];
collections?: any;
type: string;
value?: any;
validations?: Validator[];
hint?: string;
class?: string;
}<file_sep>/src/app/_services/chamas.service.spec.ts
import { TestBed } from '@angular/core/testing';
import { ChamasService } from './chamas.service';
describe('ChamasService', () => {
beforeEach(() => TestBed.configureTestingModule({}));
it('should be created', () => {
const service: ChamasService = TestBed.get(ChamasService);
expect(service).toBeTruthy();
});
});
| 679522259a2c71f6e1039298110e2e3a803c10e8 | [
"TypeScript"
] | 31 | TypeScript | Tigana13/legitimate-tech-test | 3ff06eac7e5d56fbb201b80635185301a5e7e240 | 9c1feee5600cf9ca020bd269113a2ba8417b8af4 |
refs/heads/master | <repo_name>aolteanu00/blog<file_sep>/blog.wsgi
#!/usr/bin/python3
import sys
sys.path.insert(0,"/var/www/blog/")
sys.path.insert(0,"/var/www/blog/blog/")
import logging
logging.basicConfig(stream=sys.stderr)
from blog import app as application
<file_sep>/blog/README.md
# QUACKHEADS- BLOG
## TEAM
##### <NAME>- Project Manager and Python Backend
##### <NAME>- Python Backend
##### <NAME>- HTML Frontend
##### <NAME>- SQL Handling
## Installation
- `pip3 install flask sqlite3`
- `git clone <EMAIL>:aolteanu00/blog.git`
- You can also download the repo to your preference
- `python3 app.py`
- Open your preferred browser and go to `localhost:5000`
- You can also port forward this however you wish
- Register and login
- :pizza::pizza::beers::pizza::pizza:
## Dependencies
- Flask
- SQLite3
<file_sep>/blog/htmlTest.py
from flask import Flask
from flask import render_template
from flask import request
from flask import redirect
from flask import url_for
from flask import session
from flask import flash
import time
import sqlite3 #enable control of an sqlite database
import sqldb
app = Flask(__name__)
app.secret_key = '<KEY>'
@app.route('/')
def test():
print(__name__)
return render_template("viewBlog.html", username = "alex", posts = {"I am the title":"this is where post content will go.", "I am post2":"beep boop"})##For html testing purposes
##@app.route('/editPost')
##def editPost():
print(__name__)
##title = request.args["postTitle"]
##title = title.substring(5)
## return render_template("editPost.html", post = ["I was the title","this is where old post content will go."])
if __name__ == '__main__':
app.debug = True
app.run()
<file_sep>/blog/__init__.py
from flask import Flask
from flask import render_template
from flask import request
from flask import redirect
from flask import url_for
from flask import session
from flask import flash
import time
import sqlite3 # enable control of an sqlite database
import sqldb
app = Flask(__name__)
app.secret_key = '<KEY>'
@app.route('/test')
def test():
print(__name__)
return render_template("results.html") # For html testing purposes
@app.route('/')
def hello():
print(__name__)
if "username" in session:
return redirect("/welcome")
return render_template("login.html")
@app.route('/debug')
def hddd():
print(__name__)
return session["username"]
# REdirect to hmer
@app.route('/register')
def register():
if len(request.args) == 0:
return render_template("register.html")
else:
existencecommand = "SELECT * FROM userinfo WHERE username = '{}'".format(
request.args["usernamein"])
if(len(runsqlcommand(existencecommand)) == 0):
command = "INSERT INTO userinfo VALUES('{}','{}');".format(
request.args["usernamein"], request.args["passwordin"])
runsqlcommand(command)
session["usernamein"] = request.args["passwordin"]
return redirect("/")
else:
flash("username already exists")
return redirect("/register")
@app.route('/auth')
def auth():
if len(request.args) == 0:
session.pop("username")
return redirect("/")
command = "SELECT * FROM userinfo where username = '{}'".format(
request.args["username"])
pair = runsqlcommand(command)
if len(pair) == 0:
return render_template("login.html", error="Your username is unfortunately WRONG")
else:
if (request.args["password"] == pair[0][1]):
session["username"] = request.args["username"]
return redirect("/welcome")
else:
return render_template("login.html", error="Your password is unfortunately WRONG")
@app.route('/welcome')
def welcome():
command = "SELECT * FROM bloginfo"
data = runsqlcommand(command)
allBlogs = []
for row in data:
if row[0] == session["username"] and not(row[3] in allBlogs):
allBlogs.append(row[3])
return render_template("welcome.html", blogNames=allBlogs, username=session["username"])
@app.route('/createPost')
def createPost():
blogName = request.args["blogName"]
return render_template("createPost.html", blogName=blogName)
@app.route('/addpost')
def postadd():
blogName = request.args["blogName"]
title = request.args["postTitle"]
content = request.args["postContent"]
command = "SELECT * FROM bloginfo"
dict = runsqlcommand(command)
for row in dict:
if (row[1] == title):
flash("Title already exists. Change Title")
return redirect(url_for("createPost", blogName = blogName))
command = "INSERT INTO bloginfo VALUES('{}','{}', '{}', '{}')".format(
session["username"], title, content, blogName)
runsqlcommand(command)
flash("added post alright")
return redirect(url_for("viewBlog", blogName=blogName))
@app.route("/delete")
def delete():
entrytodelete = request.args["delete"]
command = "SELECT * FROM bloginfo"
data = runsqlcommand(command)
blogname = ""
for row in data:
if entrytodelete == row[1]:
blogname = row[3]
command = "DELETE FROM bloginfo WHERE title = '{}'".format(entrytodelete)
runsqlcommand(command)
print(blogname)
return redirect(url_for("viewBlog", blogName=blogname))
@app.route('/search')
def search():
return render_template("search.html")
@app.route('/results')
def results():
searchInput = request.args["searchInput"].lower()
length = len(searchInput)
command = "SELECT * FROM bloginfo where lower(title) LIKE '%{}%'".format(
searchInput)
allPosts = runsqlcommand(command)
print(allPosts)
# results = {}
# for post in allPosts:
# title = post[0]
# content = post[1]
# titlecomp = title.lower()
# if (titlecomp.__contains__(searchInput)):
# results[title] = content
return render_template("results.html", results=allPosts)
@app.route('/addBlog')
def addblog():
#username = session["username"]
blogname = request.args["blogName"]
title = ""
content = ""
command = "SELECT * FROM bloginfo"
dict = runsqlcommand(command)
print(dict)
for row in dict:
if (row[3] == blogname):
flash("Blog name already exists. Change it to add it")
return redirect("/createBlog")
command = "INSERT INTO bloginfo VALUES('{}','{}', '{}','{}')".format(
session["username"], title, content, blogname)
runsqlcommand(command)
return redirect("/welcome")
@app.route('/createBlog')
def createBlog():
return render_template("createBlog.html")
@app.route('/edit')
def edit():
postTitle = request.args["edit"]
command = "SELECT * FROM bloginfo"
data = runsqlcommand(command)
post = []
for row in data:
if postTitle == row[1]:
post.append(row[1])
post.append(row[2])
blogName = row[3]
return render_template("editPost.html", post=post, blogName=blogName)
@app.route('/editPost')
def editPost():
newTitle = request.args["postTitle"]
newContent = request.args["postContent"]
oldTitle = request.args["oldTitle"]
blogName = ""
command = "SELECT * FROM bloginfo"
data = runsqlcommand(command)
for row in data:
if oldTitle == row[1]:
command = "UPDATE bloginfo SET title = '{}', content = '{}' WHERE title = '{}'".format(newTitle, newContent, oldTitle)
blogName = row[3]
runsqlcommand(command)
return redirect(url_for("viewBlog", blogName = blogName))
@app.route('/showall')
def showall():
command = "SELECT * FROM bloginfo"
all = runsqlcommand(command)
blogTitles = []
for row in all:
if not(row[3] in blogTitles):
blogTitles.append(row[3])
return render_template("showall.html", blogTitles=blogTitles)
@app.route('/viewBlog')
def viewBlog():
# if not(blogName in locals()):
blogName = request.args["blogName"]
command = "SELECT * FROM bloginfo"
data = runsqlcommand(command)
dict = {}
for row in data:
if row[3] == blogName:
user = row[0]
if row[3] == blogName and not(row[1] == "" and row[2] == ""):
dict.update({row[1]: row[2]})
if user == session["username"]:
return render_template("viewYourBlog.html", posts=dict, blogName=blogName, username=session["username"])
else:
return render_template("viewBlog.html", posts=dict, blogName=blogName, username=user)
def runsqlcommand(command):
DB_FILE = "glit.db"
db = sqlite3.connect(DB_FILE) # open if file exists, otherwise create
c = db.cursor() # facilitate db ops
c.execute(command)
if "select" in command.lower():
return c.fetchall()
db.commit() # save changes
db.close() # close database
if __name__ == '__main__':
app.debug = False
app.run()
<file_sep>/blog/sqldb.py
import sqlite3 #enable control of an sqlite database
import csv #facilitate CSV I/O
DB_FILE= "glit.db"
db = sqlite3.connect("glit.db") #open if file exists, otherwise create
c = db.cursor() #facilitate db ops
#==============================================================================
#create table to store user information
#row[0] = username
#row[1] = password
#command = "CREATE TABLE userinfo(username TEXT, password TEXT);"
#c.execute(command) # run SQL statement
#create table to store blog information
#row[0] = username of blog creator
#row[1] = blogid
#row[2] = blog title
#row[3] = blog content
#command = "CREATE TABLE bloginfo(username TEXT, title TEXT, content TEXT, blogName TEXT);"
#c.execute(command) # run SQL statement
#------------------------------------------------------------------------------
#USERINFO TABLE COMMANDS
#returns username as string given corresponding password
#if password is not present in userinfo database returns None
def fetchUsername(password):
command = "SELECT * FROM userinfo"
c.execute(command)
data = c.fetchall()
for row in data:
if row[1] == password:
return row[0]
else:
return None
#returns password given corresponding username
#if username is not present in userinfo returns None
def fetchPassword(user):
command = "SELECT * FROM userinfo"
c.execute(command)
data = c.fetchall()
for row in data:
if row[0] == user:
return row[1]
else:
return None
#returns all users in userinfo database as a list
def fetchAllUsers():
command = "SELECT * FROM userinfo"
c.execute(command)
data = c.fetchall()
list = []
for row in data:
list.append(row[0])
return list
#returns all passwords in userinfo database as a list
def fetchAllPasswords():
command = "SELECT * FROM userinfo"
c.execute(command)
data = c.fetchall()
list = []
for row in data:
list.append(row[1])
return list
#adds a user to the userinfo database
def addUser(user, password):
command = "INSERT INTO userinfo VALUES('{}', '{}')".format(user, password)
c.execute(command)
db.commit()
#deletes a user from the userinfo databse
def deleteUser(user):
command = "DELETE FROM userinfo WHERE username = '{}' ;".format(user)
c.execute(command)
data = c.fetchall()
#prints userinfo database
def printUserT():
command = "SELECT * FROM userinfo"
c.execute(command)
data = c.fetchall()
print(data)
#------------------------------------------------------------------------------
#BLOGINFO TABLE COMMANDS
#returns all post titles a user has created as a list
def fetchUserTitles(user):
command = "SELECT * FROM bloginfo"
c.execute(command)
data = c.fetchall()
dict = {}
for row in data:
if row[0] == user:
dict.update( {row[1] : row[2]})
return dict.keys()
#returns all post content a user has created as a list
def fetchUserContent(user):
command = "SELECT * FROM bloginfo"
c.execute(command)
data = c.fetchall()
dict = {}
for row in data:
if row[0] == user:
dict.update( {row[1] : row[2]})
return dict.values()
#returns all posts and corresponding content a user has created as a dictionary
def fetchUserBlog(user):
command = "SELECT * FROM bloginfo"
c.execute(command)
data = c.fetchall()
dict = {}
for row in data:
if row[0] == user:
dict.update( {row[1] : row[2]} )
return dict
#returns all blog titles as a list
def fetchAllBlogTitles():
command = "SELECT * FROM bloginfo"
c.execute(command)
data = c.fetchall()
#returns all blog content as a list
def fetchAllBlogContent():
command = "SELECT * FROM bloginfo"
c.execute(command)
data = c.fetchall()
#adds a post to bloginfo database
def addPost(user, title, content):
command = "INSERT INTO bloginfo VALUES('{}', '{}', '{}')".format(user, title, content)
c.execute(command)
db.commit()
#deletes all posts(a blog) of a user from bloginfo database
def deleteBlog(user):
command = "DELETE FROM userinfo WHERE username = '{}' ;".format(user)
c.execute(command)
data = c.fetchall()
#prints bloginfo database
def printBlogT():
command = "SELECT * FROM bloginfo"
c.execute(command)
data = c.fetchall()
print(data)
#------------------------------------------------------------------------------
db.commit()
db.close()
| 8f27c9792c2214172951bea732e0774a9e576385 | [
"Markdown",
"Python"
] | 5 | Python | aolteanu00/blog | a3966ddbac64e6425968dc600c14dea539b2df47 | fc24016df32a4b8368a5a6e6cf2d037b50d3967d |
refs/heads/master | <repo_name>meganecummings/legislators<file_sep>/spreadsheet.py
import gspread
from oauth2client.service_account import ServiceAccountCredentials
import pprint
scope = ['https://spreadsheets.google.com/feeds', 'https://www.googleapis.com/auth/drive']
creds = ServiceAccountCredentials.from_json_keyfile_name('client_secret.json', scope)
client = gspread.authorize(creds)
sheet = client.open("116th_congress_190103").sheet1
pp = pprint.PrettyPrinter()
result = sheet.row_values(43)
result_cols = sheet.col_values(26)
result_cell = sheet.cell(43,13).value
pp.pprint(result_cell)
sheet.update_cell(43,13, '555-555-5555')
result_cell = sheet.cell(43,13).value
pp.pprint(result_cell)
row = ["I'm", "updating", "a", "spreadsheet", "from", "python"]
index = 3
sheet.delete_row(index)
pp.pprint(sheet.row_count)
| b082620d77ba295359fffe8a03965c07fa1eec63 | [
"Python"
] | 1 | Python | meganecummings/legislators | 485d73240866684eac18e80cba31e7078e018685 | 7f842898298c5052370f8302e1bf2f48e760d82c |
refs/heads/master | <file_sep>package com.example.mvphelloworld.view;
import android.os.Bundle;
import android.support.v7.app.AppCompatActivity;
import android.view.View;
import android.widget.Button;
import android.widget.TextView;
import com.example.mvphelloworld.R;
import com.example.mvphelloworld.contract.MainActivityContract;
import com.example.mvphelloworld.presenter.MainActivityPresenter;
public class MainActivity extends AppCompatActivity implements MainActivityContract.View {
private MainActivityContract.Presenter presenter;
private TextView tvSample;
@Override
protected void onCreate(Bundle savedInstanceState) {
super.onCreate(savedInstanceState);
setContentView(R.layout.activity_main);
presenter = new MainActivityPresenter(this);
}
@Override
public void initViews() {
tvSample = findViewById(R.id.tvSample);
Button btnLoadData = findViewById(R.id.btnLoadData);
btnLoadData.setOnClickListener(new View.OnClickListener() {
@Override
public void onClick(View v) {
presenter.onClick();
}
});
}
@Override
public void setViewData(String data) {
tvSample.setText(data);
}
}
<file_sep>package com.example.mvphelloworld.contract;
public interface MainActivityContract {
interface View {
/*Define Rules to perform in your view class*/
void initViews();
void setViewData(String data);
}
interface Presenter {
/*Define rules to perform in your presenter class*/
void onClick();
}
interface Model {
/*Define rules to perform in your model class*/
String getData();
}
}
| a79efdf9eb811dd41be9f33afe20461548204bab | [
"Java"
] | 2 | Java | RajputNaveenLalTiwari/MVPHelloWorld | 977a0b405b2b4ddef71e5a5a8e96110c07b6acda | a00960c32f712dd216469dd646b7e33bc91acda9 |
refs/heads/main | <file_sep>/** Object Containers
* Objects are typically Constants
* We can change the properties of the object inside the container
* We can't remove or replace the object from the container
*/
/** Object Properties
* Describe different aspect of the object
* Syntax: Key: "Value",
* Property names can only consists of letters, numbers, dollar signs, and underscores. Any others cause JS to crash
* use camelCase
*/
/** Accessing Objects
* To access, call the name
*/
| 5e2fe4400e062d0ab62cd7b34f0d0428790d0e7b | [
"JavaScript"
] | 1 | JavaScript | anibidar/JavaScript | 3bd60171625ddb287da63329958b23de5490e6b1 | c82104511603d22f0e9281bbf9466c05f5736709 |
refs/heads/master | <repo_name>rafa761/decision-tree-example<file_sep>/README.md
# Decision-Tree-1<file_sep>/Decision_tree.py
# Decision Tree
# Fist 2 Columns is the Features
# The last Column is the Label
# Each Line is an example
training_data = [["Verde", 3, "Manga"],
["Amarelo", 3, "Manga"],
["Vermelho", 1, "Uva"],
["Vermelho", 1, "Uva"],
["Amarelo", 3, 'Limao'],]
# Print the Tree - used only to print
header = ["cor", "tamanho", "label"]
def unique_vals(rows, col):
"""Find the unique values for the column in the dataset
Example: unique_vals(training_data, 0)
unique_vals(training_data, 1)"""
return set([row[col] for row in rows])
def class_counts(rows):
"""Counts the number of each type of example in a dataset
Example: class_counts(training_data)"""
counts = {} # A Dictionary of label -> count.
for row in rows:
# in our dataset format, the label is always the last column
label = row[-1]
if label not in counts:
counts[label] = 0
counts[label] += 1
return counts
def is_numeric(value):
"""Test if a value is Numeric.
Example: is_numeric(7)
is_numeric("Vermelho")"""
return isinstance(value, int) or isinstance(value, float)
class Question:
"""A Question is used to partition a dataset
This Class just records a 'Colum Number' (e.g., 0 for color) and a
'Colum value' (e.g., Verde). The 'match' method is used to compare
the feature value in an example to the feature value stored in the
question. """
def __init__(self, column, value):
self.column = column
self.value = value
def match(self, example):
# Compare the feature value in an example to The
# feature stored in this question
val = example[self.column]
if is_numeric(val):
return val >= self.value
else:
return val == self.value
def __repr__(self):
# This is just a helper method to print
# the result in an readable format
condition = "=="
if is_numeric(self.value):
condition = ">="
return "{} é {} {}".format(header[self.column], condition, str(self.value) )
def partition(rows, question):
"""Partition a dataset
For each row in the dataset, check if it matches the question. If
so, add it to 'true rows', otherwise, add it to 'false rows' """
true_rows, false_rows = [], []
for row in rows:
if question.match(row):
true_rows.append(row)
else:
false_rows.append(row)
return true_rows, false_rows
def gini(rows):
"""Calculate the Gini Impurity for a list of rows."""
counts = class_counts(rows)
impurity = 1
for lbl in counts:
prob_of_lbl = counts[lbl] / float(len(rows))
impurity -= prob_of_lbl**2
return impurity
def info_gain(left, right, current_uncertainty):
"""Information Gain
The uncertainty of the starting node, minus the wighted impurity of
two child nodes"""
p = float(len(left)) / (len(left) + len(right))
return current_uncertainty - p * gini(left) - (1 - p) * gini(right)
def find_best_split(rows):
"""Find the best question to ask by iterating over every feature / value
and calculating the information gain."""
best_gain = 0 # keep track of the best information Gain
best_question = None # keep train ofthe feature / value that produce it
current_uncertainty = gini(rows)
n_features = len(rows[0]) - 1 # number of Columns
for col in range(n_features): # for each Features
values = set([row[col] for row in rows]) # Unique values in column
for val in values: # for each values
question = Question(col, val)
# Try Splitting the dataset
true_rows, false_rows = partition(rows, question)
# Skip this split if doesn't divide the dataset
if len(true_rows) == 0 or len(false_rows) == 0:
continue
# Calculate the information gain from this split
gain = info_gain(true_rows, false_rows, current_uncertainty)
# You actually can use '>' instead of '>=' here
# but I wanted the tree to look a certain way for our toy dataset
if gain >= best_gain:
best_gain, best_question = gain, question
return best_gain, best_question
class Leaf:
"""A leaf node classifies data.
This holds a dictionary of class (e.g., 'Manga') -> number of times
it appears in the rows from the training data that reach this leaf"""
def __init__(self, rows):
self.predictions = class_counts(rows)
class Decision_Node:
"""A Decision Node ask a question.
This holds a reference to the question, and to the two child nodes."""
def __init__(self, question, true_branch, false_branch):
self.question = question
self.true_branch = true_branch
self.false_branch = false_branch
def build_tree(rows):
"""Build the tree"""
# Try partitioning the dataset on each of the unique attribute,
# calculate the information gain,
# and return the question that produces the highest gain
gain, question = find_best_split(rows)
# Base case: no further information gain
# Since we can ask o futher questions,
# we'll retun a leaf
if gain == 0:
return Leaf(rows)
# If we reach here, we have found a useful feature / value
# to partition on
true_rows , false_rows = partition(rows, question)
# Recursively build the true branch
true_branch = build_tree(true_rows)
# Recursively build the false branch
false_branch = build_tree(false_rows)
# Return a Question node
# This records the best feature / value to ask at this point,
# as well as the branches o follow depending on the answer
return Decision_Node(question, true_branch, false_branch)
def print_tree(node, spacing=""):
# Base case: we've reached a leaf
if isinstance(node, Leaf):
print(spacing + "Predict", node.predictions)
return
# Print the question at this node
print(spacing + str(node.question))
# Call this function recursively on the true branch
print(spacing + '--> True:')
print_tree(node.true_branch, spacing + " ")
# Call this function recursively on the false branch
print(spacing + '--> False:')
print_tree(node.false_branch, spacing + " ")
def classify(row, node):
# Base case: we've reached a leaf
if isinstance(node, Leaf):
return node.predictions
# Decide whether to follow the true-branch or the false-branch.
# Compare the feature / value stored in the node,
# to the example we're considering
if node.question.match(row):
return classify(row, node.true_branch)
else:
return classify(row, node.false_branch)
def print_leaf(counts):
"""Print the predictions at a leaf"""
total = sum(counts.values()) * 1.0
probs = {}
for lbl in counts.keys():
probs[lbl] = str(int(counts[lbl] / total * 100)) + "%"
return probs
if __name__ == "__main__":
my_tree = build_tree(training_data)
print_tree(my_tree)
# Evaluate
testing_data = [
["Verde", 3, "Manga"],
["Amarelo", 4, "Manga"],
["Vermelho", 2, "Uva"],
["Vermelho", 1, "Uva"],
["Amarelo", 3, "Limao"],
]
for row in testing_data:
print("Actual: {}. Predicted: {}".format(row[-1], print_leaf(classify(row, my_tree)) ))
| 7585d5d3eba71c084827210c8f247f494a6f183f | [
"Markdown",
"Python"
] | 2 | Markdown | rafa761/decision-tree-example | 25e153521200cbf331f57d73c5afaf03224f5ca7 | e659d503af690e54307d082180fc512c0ac9eb99 |
refs/heads/master | <repo_name>Jomzi/3rdYearProjectWhatsApp<file_sep>/ChattingApplication/ChattingClient/ClientCallback.cs
using ChattingInterfaces;
using System;
using System.Collections.Generic;
using System.Linq;
using System.ServiceModel;
using System.Text;
using System.Threading.Tasks;
using System.Windows;
namespace ChattingClient
{
[CallbackBehavior(ConcurrencyMode = ConcurrencyMode.Multiple)]
public class ClientCallback : IClient
{
public void GetMessage(string message, string userName)
{
//casting main window
((MainWindow)Application.Current.MainWindow).TakeMessage(message, userName);
}
public void GetUpdate(int value, string userName)
{
switch(value)
{
case 0:
{
((MainWindow)Application.Current.MainWindow).AddUserToList(userName);
break;
}
case 1:
{
((MainWindow)Application.Current.MainWindow).RemoveUserFromList(userName);
break;
}
}
}
}
}
<file_sep>/README.md
# 3rdYearProjectWhatsApp
3rd-Year-Project
This is my 3rd year project for college. It incorporates material and topics from all the modules I studied in college. There was no specification provided as it was up to the student to choose the project.
Introduction
Since my first attempt at the messaging project was unsuccessful, the reason being…I could not get the WhatsApp API working, as everyone online was running into the same problem when I searched for a solution. So I had to change my app around slightly.
I set my goal to make a chatting app on visual studios 2015 using c#. First step I took was creating 3 projects in 1 containing a Client, Server and Interface using WPF (Windows Presentation Foundation) application.
Inside this project the client is going to be communicating with server and the server is going to be communicating with the clients, to create an instant messaging app. Also when a client goes offline other clients will pick up on this and see the client has gone offline at a certain time.
I done all of my projects design in Visual Studio “Blend”.
How to run the project
GitHub
Project link: https://github.com/Jomzi/3rdYearProjectWhatsApp
1. Import the project from github into visual studios.
2. Run the project.
3. When the project is running a client window will pop up and also a the console server.
4. Insert a username into the client.
5. Run another client by right clicking the “ChattingClient” or press f5 (I think also works).
6. Insert a different username into second client as a username that has already being used wont work!
7. Once all this is done the app is very easy to use. (Screenshot below should help you)
ScreenShots provided in word doc!
<file_sep>/ChattingApplication/ChattingClient/MainWindow.xaml.cs
using ChattingInterfaces;
using System;
using System.Collections.Generic;
using System.Linq;
using System.ServiceModel;
using System.Text;
using System.Threading.Tasks;
using System.Windows;
using System.Windows.Controls;
using System.Windows.Data;
using System.Windows.Documents;
using System.Windows.Input;
using System.Windows.Media;
using System.Windows.Media.Imaging;
using System.Windows.Navigation;
using System.Windows.Shapes;
namespace ChattingClient
{
/// <summary>
/// Interaction logic for MainWindow.xaml
/// </summary>
public partial class MainWindow : Window
{
public static IChattingService Server;
private static DuplexChannelFactory<IChattingService> _channelFactory;
public MainWindow()
{
InitializeComponent();
_channelFactory = new DuplexChannelFactory<IChattingService>(new ClientCallback(), "ChattingServiceEndPoint");
Server = _channelFactory.CreateChannel();
}
public void TakeMessage(string message, string userName)
{
ChatDisplayTextBox.Text += userName + ": " + message + "\n";
//Scrolls automatically to text at the bottom of the message box
ChatDisplayTextBox.ScrollToEnd();
}
//Sends the message in the chat textbox when send is clicked
private void SendButton_Click(object sender, RoutedEventArgs e)
{
if(ChatTextBox.Text.Length == 0 )
{
return;
}
Server.SendMessageToAll(ChatTextBox.Text, UserNameTextBox.Text);
TakeMessage(ChatTextBox.Text, "You");
ChatTextBox.Text = "";
}
private void LoginButton_Click(object sender, RoutedEventArgs e)
{
int returnValue = Server.Login(UserNameTextBox.Text);
if(returnValue == 1)
{
MessageBox.Show("You are already logged in. Try again");
}
else if (returnValue == 0)
{
//Displays the name of the user who logged on
MessageBox.Show("You successfully logged in!");
WelcomeUserLbl.Content = "Welcome " + UserNameTextBox.Text + "!";
UserNameTextBox.IsEnabled = false;
LoginButton.IsEnabled = false;
//loads users
LoadUserList(Server.GetCurrentUsers());
}
}
private void Window_Closing(object sender, System.ComponentModel.CancelEventArgs e)
{
Server.Logout();
}
//Adds online users to show there online
public void AddUserToList(string userName)
{
if(OnlineUserListBox.Items.Contains(userName))
{
return;
}
OnlineUserListBox.Items.Add(userName);
}
public void RemoveUserFromList(string userName)
{
if (OnlineUserListBox.Items.Contains(userName))
{
OnlineUserListBox.Items.Remove(userName);
}
}
private void LoadUserList(List<string>users)
{
foreach(var user in users)
{
AddUserToList(user);
}
}
}
}
<file_sep>/ChattingApplication/ChattingServer/ChattingService.cs
using ChattingInterfaces;
using System;
using System.Collections.Concurrent;
using System.Collections.Generic;
using System.Linq;
using System.Runtime.Serialization;
using System.ServiceModel;
using System.Text;
namespace ChattingServer
{
[ServiceBehavior(ConcurrencyMode = ConcurrencyMode.Multiple, InstanceContextMode = InstanceContextMode.Single)]
public class ChattingService : IChattingService
{
public ConcurrentDictionary<string, ConnectedClient> _connectedClients = new ConcurrentDictionary<string,
ConnectedClient>();
public int Login(string userName)
{
//is anyone else logged in with my name?
{ foreach (var client in _connectedClients)
{
if(client.Key.ToLower()==userName.ToLower())
{
//if yes
return 1;
}
}
}
var establishedUserConnection = OperationContext.Current.GetCallbackChannel<IClient>();
//username is unqiue...so cannot log in with the same username.
ConnectedClient newClient = new ConnectedClient();
newClient.connection = establishedUserConnection;
newClient.UserName = userName;
_connectedClients.TryAdd(userName, newClient);
updateHelper(0, userName);
//shows information on the console window of the user login
Console.ForegroundColor = ConsoleColor.Green;
Console.WriteLine("Client Logged In: {0} at {1}", newClient.UserName, System.DateTime.Now);
Console.ResetColor();
return 0;
}
public void Logout()
{
ConnectedClient client = GetMyClient();
if(client != null)
{
ConnectedClient removedClient;
_connectedClients.TryRemove(client.UserName, out removedClient);
updateHelper(1, removedClient.UserName);
//shows information on the console window of the user logged out
Console.ForegroundColor = ConsoleColor.Red;
Console.WriteLine("Client Logged Off: {0} at {1}", removedClient.UserName, System.DateTime.Now);
Console.ResetColor();
}
}
public ConnectedClient GetMyClient()
{
var establishedUserConnection = OperationContext.Current.GetCallbackChannel<IClient>();
foreach(var client in _connectedClients)
{
if(client.Value.connection == establishedUserConnection)
{
return client.Value;
}
}
return null;
}
//Sends message to all users
public void SendMessageToAll(string message, string userName)
{
foreach(var client in _connectedClients)
{
if(client.Key.ToLower() != userName.ToLower())
{
client.Value.connection.GetMessage(message, userName);
}
}
}
private void updateHelper(int Value, string userName)
{
foreach (var client in _connectedClients)
{
if (client.Value.UserName.ToLower() != userName.ToLower())
{
client.Value.connection.GetUpdate(Value, userName);
}
}
}
public List<string> GetCurrentUsers()
{
List<string> listOfClients = new List<string>();
foreach (var client in _connectedClients)
{
listOfClients.Add(client.Value.UserName);
}
{
return listOfClients;
}
}
}
}
| b39e1dbeb3671962d0c0feeca90e51de3912f490 | [
"Markdown",
"C#"
] | 4 | C# | Jomzi/3rdYearProjectWhatsApp | 7ec2d0705358c1dab1f11d4781b4020bd2d76ade | 1b4631fcf6b0ec7a5d537591bb4afbc337c4e8dc |
refs/heads/master | <file_sep>=== Debt Reduction Calculator ===
Contributors: prasunsen, <NAME>
Tags: debt reduction, debt payoff, calculator
Requires at least: 2.0.2
Tested up to: 3.4.1
Stable tag: trunk
This plugin displays debt reduction calculator
== Description ==
This plugin displays functional debt reduction/debt payoff calculator. It shows people how they can pay debt faster or how much they need to pay extra each month in order to be debt free after chosen period of time.
To display the calculator simply place `{{debt-reduction-calculator}}` mask in the content of a page or post. Of course the plugin should be activated first.
== Installation ==
1. Upload debtpayoffcalc.php` to the `/wp-content/plugins/` directory
2. Activate the plugin through the 'Plugins' menu in WordPress
3. Configure the CSS class in the admin screen in Plugins/Debt Reduction Calculator. This is optional, the calculator will work fine without any CSS classes assigned.
4. Place `{{debt-reduction-calculator}}` mask in the content of a page or post. The calculator will appear on the place of this tag
== Frequently Asked Questions ==
= Can I change the CSS? =
Yes, you can change it in Plugins/Debt Reduction Calculator page
= Can I put Ads On the Page With The Calculator? =
Yes, you can put Adsense or any other ads
= Should I link back to the plugin page =
It's not required but a link to http://calendarscripts.info/debt-reduction-calculator.html will be highly appreciated.
= Why doesn't this use Wordpress shortcodes? =
Because they weren't available when I first wrote the plugin, and I have to keep the format for backward compatibility. Technically it makes no difference, really.
<file_sep><?php
/*
Plugin Name: Debt Reduction Calculator
Plugin URI: http://calendarscripts.info/debt-reduction-wp.html
Description: This plugin displays functional debt reduction/debt payoff calculator. It helps the user calculate how a debt can be paid faster and the total amount paid to the bank be reduced.
Author: <NAME>
Version: 1.1
Author URI: http://calendarscripts.info
*/
/* Copyright 2008 <NAME> (email : <EMAIL>)
This program is free software; you can redistribute it and/or modify
it under the terms of the GNU General Public License as published by
the Free Software Foundation; either version 2 of the License, or
(at your option) any later version.
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
GNU General Public License for more details.
You should have received a copy of the GNU General Public License
along with this program; if not, write to the Free Software
Foundation, Inc., 51 Franklin St, Fifth Floor, Boston, MA 02110-1301 USA
*/
function debtpayoff_add_page()
{
add_submenu_page('plugins.php', 'Debt Reduction Calculator', 'Debt Reduction Calculator', 8, __FILE__, 'debtpayoff_options');
}
// ovpredct_options() displays the page content for the Ovpredct Options submenu
function debtpayoff_options()
{
// Read aan existing option value from database
$drc_table = stripslashes( get_option( 'drc_table' ) );
// See if the user has posted us some information
// If they did, this hidden field will be set to 'Y'
if( $_POST[ 'drc_update' ] == 'Y' )
{
// Read their posted value
$drc_table = $_POST[ 'drc_table' ];
// Save the posted value in the database
update_option( 'drc_table', $drc_table );
// Put an options updated message on the screen
?>
<div class="updated"><p><strong><?php _e('Options saved.', 'drc_domain' ); ?></strong></p></div>
<?php
}
// Now display the options editing screen
echo '<div class="wrap">';
// header
echo "<h2>" . __( 'Debt Reduction Calculator', 'drc_domain' ) . "</h2>";
// options form
?>
<form name="form1" method="post" action="<?php echo str_replace( '%7E', '~', $_SERVER['REQUEST_URI']); ?>">
<input type="hidden" name="drc_update" value="Y">
<p><?php _e("CSS class definition for the debt reduction calculator <div>:", 'drc_domain' ); ?>
<textarea name="drc_table" rows='5' cols='70'><?php echo stripslashes ($drc_table); ?></textarea>
</p><hr />
<p class="submit">
<input type="submit" name="Submit" value="<?php _e('Update Options', 'drc_domain' ) ?>" />
</p>
</form>
</div>
<?php
}
// This just echoes the text
function debtpayoffcalc($content)
{
if(!strstr($content,"{{debt-reduction-calculator}}")) return $content;
//construct the calculator page
$css=get_option('drc_table');
$dr_calc="";
if(!empty($css))
{
$dr_calc.="<style type=\"text/css\">
.calculator_table
{
$css
}
</style>\n\n";
}
if(empty($css))
{
$inline_style="style='margin:auto;padding:5px;width:450px;text-align:left;'";
}
if(!empty($_POST['calculator_ok']))
{
// save in session to be used when the link "calculate again" is clicked
foreach($_POST as $key=>$var) $_SESSION["calc_".$key]=$var;
// first let's calculate how much the user is going to pay
// with the current payment plan
$monthly_interest=round($_POST['interest']/12,2)/100;
$num_months=0;
$total_interest=0;
$principal=$_POST['debt'];
// make sure monthly payment is not < of the first interest payment
$minimum_payment=$principal*$monthly_interest;
if($minimum_payment>=$_POST['payment'])
{
die("Your monthly payment is too low, you can never pay this debt!");
}
while($principal>0)
{
$this_month_interest=$principal*$monthly_interest;
$total_interest+=$this_month_interest;
$num_months++;
$current_total=$principal+$this_month_interest;
// reduce principal with the difference
$principal=$current_total-$_POST['payment'];
}
// months and years text
if($num_months>12)
{
$left=$num_months%12;
$num_years=($num_months-$left)/12;
if($left>0) $current_debt_free_text="$num_years years and $left months";
else $current_debt_free_text="$num_years years";
}
else
{
$current_debt_free_text="$num_months months";
}
// start reduction calculations
if($_POST['mode']=='extra')
{
// we need the same loop as above but using the new amount
$num_months_extra=0;
$total_interest_extra=0;
$principal=$_POST['debt'];
while($principal>0)
{
$this_month_interest=$principal*$monthly_interest;
$total_interest_extra+=$this_month_interest;
$num_months_extra++;
$current_total=$principal+$this_month_interest;
// reduce principal with the difference
$principal=$current_total-$_POST['payment']-$_POST['extra_payment'];
}
// months and years text
if($num_months_extra>12)
{
$left_extra=$num_months_extra%12;
$num_years_extra=($num_months_extra-$left_extra)/12;
if($left_extra>0) $debt_free_text_extra="$num_years_extra years and $left_extra months";
else $debt_free_text_extra="$num_years_extra years";
}
else
{
$debt_free_text_extra="$num_months_extra months";
}
}
else // $_POST[mode]==target
{
// number of months
$target_months=$_POST['target_years']*12 + $_POST['target_months'];
// calculate the monthly payment using this formula
// http://www.vertex42.com/ExcelArticles/amortization-calculation.html
$monthly_payment=$_POST['debt'] *
$monthly_interest * pow(1+$monthly_interest, $target_months)
/
(pow(1+$monthly_interest, $target_months) - 1);
}
//the result is here
$dr_calc.='<div class="drc_table" '.$inline_style.'>';
$dr_calc.=' <h2>Calculation Results</h2>
<p>Following your current payment plan you will be debt free after <b>'.$current_debt_free_text.'</b>. For this period you will pay total interest of <b>$'.number_format($total_interest).'</b> to the bank.</p>';
if($_POST['mode']=='extra')
{
$dr_calc.='<p>By paying extra $'.$_POST['extra_payment'].' monthly you will be debt free after <b>'.$debt_free_text_extra.'</b>.
The interest paid to the bank in this case will be <b>$'.number_format($total_interest_extra).'</b>.</p>';
}
else
{
$dr_calc.='<p>If you want to be debt free after <b>'.$_POST['target_years'].' years and '.number_format($_POST['target_months']).' months</b> you need to pay <b>$'.number_format($monthly_payment).' monthly</b>.</p>';
}
$dr_calc.='<p align="center"><a href="http://'.$_SERVER['HTTP_HOST'].$_SERVER['REQUEST_URI'].'">Calculate again</a></p></div>';
}
else
{
$dr_calc.='<div class="calculator_table">
<form method="post" onsubmit="return validateDebtCalculator(this);">
<fieldset>
<legend>Debt Details:</legend>
<p><label>Debt Balance:</label> $<input type="text" name="debt" value="'.$_SESSION['calc_debt'].'"></p>
<p><label>Annual Interest Rate:</label> <input type="text" name="interest" size="5" value="'.$_SESSION['calc_interest'].'">%</p>
<p><label>Current Monthly Payment:</label> $<input type="text" size="5" name="payment" value="'.$_SESSION['calc_payment'].'"></p>
</fieldset>
<fieldset>
<legend>I want to know:</legend>
<p><input type="radio" id="extraMode" name="mode" value="extra" '.((!isset($_SESSION['calc_mode']) or $_SESSION['calc_mode']=="extra")?"checked":"").'>
What will happen if I pay extra $<input type="text" name="extra_payment" size="5" value="'.((@$_SESSION['calc_mode']=='extra') ?@$_SESSION['calc_extra_payment']:'').'"> monthly</p>
<p><input type="radio" id="targetMode" name="mode" value="target" '.((@$_SESSION['calc_mode']=='target')?'checked':'').'> How to be debt-free after <input type="text" name="target_years" size="3" value="'.(($_SESSION['calc_mode']=='target')?$_SESSION['calc_target_years']:'').'"> years & <input type="text" name="target_months" size="2" value="'.((@$_SESSION['calc_mode']=='target')?@$_SESSION['calc_target_months']:'').'"> months</p>
</fieldset>
<p align="center"><input type="submit" name="calculator_ok" value="Calculate"></p>
</form>
</div>
<script language="javascript">
function validateDebtCalculator(frm)
{
if(isNaN(frm.debt.value) || frm.debt.value=="")
{
alert("Please enter your debt balance, numbers only");
frm.debt.focus();
return false;
}
if(isNaN(frm.interest.value) || frm.interest.value=="")
{
alert("Please enter your debt annual interest rate, numberic");
frm.interest.focus();
return false;
}
if(isNaN(frm.payment.value))
{
alert("Please enter only numbers for your current monthly payment");
frm.payment.focus();
return false;
}
if(document.getElementById('."'extraMode'".').checked)
{
if(isNaN(frm.extra_payment.value) || frm.extra_payment.value=="")
{
alert("Please enter your debt extra monthly payment, numbers only");
frm.extra_payment.focus();
return false;
}
}
if(document.getElementById('."'targetMode'".').checked)
{
if(isNaN(frm.target_years.value) || frm.target_years.value=="")
{
alert("Please enter target years, numbers only");
frm.target_years.focus();
return false;
}
if(isNaN(frm.target_months.value) || frm.target_months.value=="")
{
alert("Please enter target months, numbers only");
frm.target_months.focus();
return false;
}
}
}
</script>';
}
$content=str_replace("{{debt-reduction-calculator}}",$dr_calc,$content);
return $content;
}
add_action('admin_menu','debtpayoff_add_page');
add_filter('the_content', 'debtpayoffcalc'); | 573569ab9e5c51bfd04601f92037121a97f7b779 | [
"Text",
"PHP"
] | 2 | Text | kazi-shahin/debt-reduction-calculator | 2884700f30769b5dea5f5eea22e799cfdbfa5203 | b52eeb745d98df25824f42d760cf12a506984aeb |
refs/heads/master | <repo_name>jacknichao/test-reference<file_sep>/src/jmetal/nichao/testreferences/test.java
package jmetal.nichao.testreferences;
import java.io.File;
import java.io.PrintWriter;
import java.nio.file.FileStore;
import java.nio.file.Files;
import java.nio.file.Path;
import java.nio.file.Paths;
import java.util.Scanner;
import java.util.StringTokenizer;
public class test {
public static void main(String[] args) throws Exception {
// Scanner scanner=new Scanner(new File("/tmp/multi-object/results-random-1/RelinkSixMethods/data/NSGAII/ApacheJ48/VAR-NSGAII.0"));
/* while(scanner.hasNextLine()){
String str=scanner.nextLine();
StringTokenizer stringTokenizer=new StringTokenizer(str," ");
while(stringTokenizer.hasMoreTokens()){
System.out.println(stringTokenizer.nextToken());
}
break;
}
scanner.close();*/
/* File f=new File("/home/jacknichao/datasets/Relink");
System.out.println(f.getName());
System.out.println(f.getPath());*/
System.out.println(1-Double.valueOf(0.23123214243434));
}
}
<file_sep>/README.md
# test-reference
this project is used to generate references fronts on testing instances.
this project can only be executed after "**multi-object**" project.
**multi-project** generates reference front of training instances, while **test-reference** generates references front of test instances under the help of **multi-object**;
##Steps to execute this project:
###first :
run FilterParetoFront.java to filter out those fronts which can not effectively select any features.
###second:
run CheckOnTest.java to check on test instances and evaluate the performance with paretofront feature.
###third:
run GenerateTestParetoFront.java to generate references fronts of test instances with the help of the fronts of trainning instances.<file_sep>/src/jmetal/nichao/testreferences/CheckOnTest.java
package jmetal.nichao.testreferences;
import jmetal.nichao.testreferences.core.BaseClassiferEnum;
import jmetal.nichao.testreferences.core.ClassifierValidation;
import weka.classifiers.Evaluation;
import weka.core.Instances;
import weka.core.converters.ConverterUtils;
import weka.filters.Filter;
import weka.filters.unsupervised.attribute.Remove;
import java.io.File;
import java.io.PrintWriter;
import java.util.*;
public class CheckOnTest {
/**
* 将源项目分成70%的训练集和30%的验证集,对两者进行特征过滤,
* 在70%训练数据上进行预测,然后在30%测试数据上进行判断,得到预测结果;
*
* @param trainning 训练数据的实例
* @param testing 测试数据的实例
* @param baseClassiferEnum 基分类器枚举
* @return 返回Evaluation:表示预测得到的结果
*/
public static Evaluation doValidation(Instances trainning, Instances testing, BaseClassiferEnum baseClassiferEnum, int[] toDeletedFeature) {
Evaluation evaluations = null;
try {
trainning.setClassIndex(trainning.numAttributes() - 1);
testing.setClassIndex(testing.numAttributes() - 1);
Remove remove = new Remove();
remove.setAttributeIndicesArray(toDeletedFeature);
remove.setInputFormat(trainning);
//Filter默认会产生新的Instances因此需要将trainning和testing指向新产生的Instances
trainning = Filter.useFilter(trainning, remove);
//删除训练集中的属性
testing = Filter.useFilter(testing, remove);
trainning.setClassIndex(trainning.numAttributes() - 1);
testing.setClassIndex(testing.numAttributes() - 1);
//ClassifierValidation类完成在训练集上构建模型,在测试集上进行预测的计算
evaluations = ClassifierValidation.classify(trainning, testing, baseClassiferEnum, null);
} catch (Exception e) {
e.printStackTrace();
}
return evaluations;
}
/**
* 完成内层十次循环
*
* @param fileName 文件名称
* @param outerInstances 被外层随机化后的项目实例
* @param outerIterate 外层循环的编号
*/
public static void doDefectPrediciton(String fileName, Instances outerInstances, int outerIterate) {
//计算前70%的实例的索引
int index = (int) Math.round(outerInstances.numInstances() * 0.7);
Instances train = new Instances(outerInstances, 0, index);
Instances test = new Instances(outerInstances, index, outerInstances.numInstances() - index);
train.setClassIndex(train.numAttributes() - 1);
test.setClassIndex(test.numAttributes() - 1);
//实验结果的根目录
String experimentBaseDirectory = MyTools.getBaseInfo("experimentBaseDirectory");
String enableDataset = MyTools.getBaseInfo("enableDataset").trim();
String experimentName = null;
if (enableDataset.equals("Relink")) {
experimentName = MyTools.getBaseInfo("experimentNameRelink");
} else if (enableDataset.equals("PROMISE")) {
experimentName = MyTools.getBaseInfo("experimentNamePROMISE");
}
String subDir = experimentName + "/data/NSGAII/";
Evaluation evaluation = null;
//遍历每一个集分类
for (BaseClassiferEnum baseClassiferEnum : BaseClassiferEnum.values()) {
//数据集和对应的集分类器方法构成的目录
String dirName = fileName.split("\\.")[0] + baseClassiferEnum;
//遍历里面的10个文件
for (int i = 0; i < 10; i++) {
String tmpFile = experimentBaseDirectory + "-random-" + outerIterate + "/" + subDir + dirName + "/VAR-NSGAII." + i;
String tmpOut = experimentBaseDirectory + "-random-" + outerIterate + "/" + subDir + dirName + "/FUN-NSGAII-TEST." + i;
Scanner scanner = null;
PrintWriter printWriter = null;
try {
scanner = new Scanner(new File(tmpFile));
printWriter = new PrintWriter(new File(tmpOut));
while (scanner.hasNextLine()) {
String line = scanner.nextLine();
if(line.trim().equals("")) continue;
Instances currentTrain = new Instances(train);
Instances currentTest = new Instances(test);
int[] toDeletedFeature=MyTools.findToDeletedFeature(line);
//断言检验
assert toDeletedFeature!=null &&toDeletedFeature.length!=0;
evaluation = doValidation(currentTrain, currentTest, baseClassiferEnum, toDeletedFeature);
//因为再求解paretofront的时候,我们需要是目标最小化
printWriter.println(new Double(train.numAttributes()-toDeletedFeature.length-1) + " " + (1-evaluation.weightedAreaUnderROC()));
}//while
} catch (Exception e) {
e.printStackTrace();
} finally {
scanner.close();
printWriter.close();
}
}
}
}
public static void main(String[] args) {
ArrayList<File> currenctDataSet = null;
String datasetName = MyTools.getBaseInfo("enableDataset");
try {
//遍历每一个数据集中的所有项目
//String[] strs = datasetName.split(",");
String[] strs = new String[]{datasetName};
for (String dataset : strs) {
System.out.println("开始在测试集上验证:\t" + dataset + "\t" + new Date().toString());
//得到该数据集中的所有项目
currenctDataSet = MyTools.getProjects(dataset);
for (File file : currenctDataSet) {
Instances originalInstances = ConverterUtils.DataSource.read(file.toString());
//外层10次随机循环
for (int outerIterate = 1; outerIterate <= 10; outerIterate++) {
System.out.println("正在测试:\t"+file.getName()+"\t 轮次:"+outerIterate);
//新建一个外层循环的实例集合
Instances outerInstances = new Instances(originalInstances);
//先进行外层的随机化,为了避免内层随机化与外层的一样,内层的随机因子将在外层的随机因子上加1
outerInstances.setClassIndex(outerInstances.numAttributes() - 1);
outerInstances.randomize(new Random(outerIterate));
if (outerInstances.classAttribute().isNominal()) {
outerInstances.stratify(10);
}
doDefectPrediciton(file.getName(), outerInstances, outerIterate);
}
}
System.out.println("结束运行项目:\t" + dataset + "\t" + new Date().toString());
}
} catch (Exception e) {
e.printStackTrace();
}
}
}
<file_sep>/src/jmetal/nichao/testreferences/GenerateTestParetoFront.java
package jmetal.nichao.testreferences;
import jmetal.qualityIndicator.util.MetricsUtil;
import jmetal.util.NonDominatedSolutionList;
import java.io.File;
import java.nio.file.Files;
import java.nio.file.Paths;
public class GenerateTestParetoFront {
private static String experimentBaseDirectory = null;
private static String enableDataset = null;
private static String experimentName = null;
private static String subDir = null;
static {
//实验结果的根目录
experimentBaseDirectory = MyTools.getBaseInfo("experimentBaseDirectory");
enableDataset = MyTools.getBaseInfo("enableDataset").trim();
experimentName = null;
if (enableDataset.equals("Relink")) {
experimentName = MyTools.getBaseInfo("experimentNameRelink");
} else if (enableDataset.equals("PROMISE")) {
experimentName = MyTools.getBaseInfo("experimentNamePROMISE");
}
subDir = experimentName + "/data/NSGAII/";
}
/**
* 生成引用前沿
*
* @param problemFulPath 目标问题的目录
* @param outParentDir 输出的引用前沿跟目录
*/
public static void generateReferenceFronts(String problemFulPath, String outParentDir) {
MetricsUtil metricsUtils = new MetricsUtil();
NonDominatedSolutionList solutionSet = new NonDominatedSolutionList();
for (int numRun = 0; numRun < 10; numRun++) {
String outputParetoFrontFilePath = problemFulPath + "/FUN-NSGAII-TEST." + numRun;
metricsUtils.readNonDominatedSolutionSet(outputParetoFrontFilePath, solutionSet);
} // for
solutionSet.printObjectivesToFile(outParentDir + "/" + new File(problemFulPath).getName() + ".rf");
} // generateReferenceFronts
/**
*生成NSGAII的引用的参照前沿
*/
public static void generateReferenceNSGAIIFronts() {
System.out.println("Starting generate NSGAII referencesFront");
//10个轮次的独立运行结果
for (int i = 1; i <= 10; i++) {
String parent = experimentBaseDirectory + "-random-" + i + "/" + subDir;
String outParentDir = experimentBaseDirectory + "-random-" + i +"/"+experimentName +"/referenceFrontsNSGAII";
System.out.println("Generating:\t"+experimentName+"\tindenpendentIndex:\t"+i);
File rfDirectory = new File(outParentDir);
if (!rfDirectory.exists()) {
boolean result = new File(outParentDir).mkdirs();
System.out.println("Creating " + outParentDir);
}
//遍历每一个问题,即每一个目录
for (File problemDir : new File(parent).listFiles()) {
//进入每一个问题的文件夹
if (problemDir.isDirectory()) {
generateReferenceFronts(problemDir.getPath(), outParentDir);
}
}
}
} // generateReferenceFronts
public static void main(String[] args) {
generateReferenceNSGAIIFronts();
}
}
| 4e694edb876a57280c4be25568f44eb1cdf27eb5 | [
"Markdown",
"Java"
] | 4 | Java | jacknichao/test-reference | 882b2c3b0fbe37c6295c961d06c70c492c0d26b1 | 0a826dfdeb97b8bf0c16ba51f628cabe4eb9220f |
refs/heads/master | <file_sep><?php
namespace Models;
use Illuminate\Database\Eloquent\Model as Eloquent;
class User extends Eloquent
{
const CREATED_AT = 'date_created';
const UPDATED_AT = 'date_modified';
protected $table = 'users';
protected $dateFormat = 'U';
protected $fillable = [
'name',
'date_created'
];
}<file_sep><?php
require_once 'bootstrap.php';
use Models\User;
$users = User::pluck('name');
print_r($users);
| 7c9dba47c4839e4dabb41fea5d13fd6d7fa9fe5a | [
"PHP"
] | 2 | PHP | dmokhtari/laravel-db | 4ef04b5f46c30fd3e66958ecb1122c05349373ba | 66697bc8194a94520587846d2d129869997d06a8 |
refs/heads/master | <file_sep>from __future__ import print_function, division
import nltk
import sys, os
import random
from collections import Counter
from nltk import word_tokenize, WordNetLemmatizer
from nltk.corpus import stopwords
from nltk import NaiveBayesClassifier, classify
import pickle
import smtplib
import time
import imaplib
import email
import base64
stoplist = stopwords.words('english') #stopwords sind Füllwörter wie der die das the and und...
stoplist = stopwords.words('german')
# Aufruf des Ordners und Sammeln der darin beinhalteten Dateien
def init_lists(Ordner):
a_Liste = []
#Ordner welcher übergeben wurde wird ausgelesen
Dateienliste = os.listdir(Ordner)
for a_Datei in Dateienliste:
f = open(Ordner + a_Datei, 'r', encoding="utf-8", errors='ignore')
#wird in a Liste gepackt und gelesen, dann geschlossen
a_Liste.append(f.read())
f.close()
return a_Liste
def vorprozess(satz):
lemmatizer = WordNetLemmatizer()
#gibt die kleingeschriebenen wörter zurück
return [lemmatizer.lemmatize(word.lower()) for word in word_tokenize(satz)]
def get_features(text, setting):
# bag-of-words model (BoW model) can be applied to image classification, by treating image features as words
if setting=='bow':
# wort wird gezählt und vorprozess (lemmatizer) übergeben wenn kein stopwort
return {word: count for word, count in Counter(vorprozess(text)).items() if not word in stoplist}
else:
return {word: True for word in vorprozess(text) if not word in stoplist}
def train(features, samples_proportion):
#trainingsgröße = Alle Eigenschaften * Stichprobenanteil
train_size = int(len(features) * samples_proportion)
# Initialisieren des Training und Testsets
#trainingsset bis zur trainingsgröße und testset ab diesem bis zum ende - Grund: die Set Größe ändert sich immer wieder, kann aber auch beloebig angepasst werden
trainingset, testset = features[:train_size], features[train_size:]
print ('Trainingset Größe = ' + str(len(trainingset)) + ' emails')
print ('Testset Größe = ' + str(len(testset)) + ' emails')
# Trainieren des Classifiers
# Eigenschaften werden übergeben
classifier = nltk.NaiveBayesClassifier.train(trainingset)
return trainingset, testset, classifier
def leistung(trainingset, testset, classifier):
# Richtigkeit des Classifiers auf dem Training und Testsets testen
print ('Accuracy on the training set = ' + str(nltk.classify.accuracy(classifier, trainingset)))
print ('Accuracy of the test set = ' + str(nltk.classify.accuracy(classifier, testset)))
# Die ersten 20 wichtigsten Wörter für den Classifier testen
classifier.show_most_informative_features(20)
def read_email_from_gmail():
try:
ORG_EMAIL = "@gmail.com"
FROM_EMAIL = "valeriefelixprojekt" + ORG_EMAIL
FROM_PWD = decode('<PASSWORD>', '<PASSWORD>=')
SMTP_SERVER = "imap.gmail.com"
SMTP_PORT = 993
mail = imaplib.IMAP4_SSL(SMTP_SERVER)
mail.login(FROM_EMAIL,FROM_PWD)
mail.select('inbox')
type, data = mail.search(None, 'ALL')
mail_ids = data[0]
id_list = mail_ids.split()
for i in reversed(id_list):
typ, data = mail.fetch(i, '(RFC822)' )
for response_part in data:
if isinstance(response_part, tuple):
msg = email.message_from_string(response_part[1].decode('utf-8'))
email_subject = msg['subject']
email_from = msg['from']
print('From : ' + email_from + '\n')
print('Subject : ' + email_subject + '\n')
kleister = get_features (email_subject, '')
print (kleister)
print (str(classifier.classify (kleister)))
print('\n')
print('\n')
pw = 0
FROM_PWD = 0
except Exception as e:
print(str(e))
def decode(key, enc):
dec = []
enc = base64.urlsafe_b64decode(enc).decode()
for i in range(len(enc)):
key_c = key[i % len(key)]
dec_c = chr((256 + ord(enc[i]) - ord(key_c)) % 256)
dec.append(dec_c)
return "".join(dec)
# Initialisieren
spam = init_lists('/code/MeineSpam/')
ham = init_lists('/code/MeineHam/')
allemails = [(email, 'spam') for email in spam]
allemails += [(email, 'ham') for email in ham]
random.shuffle(allemails)
#Anzahl aller Mails wird ausgegeben
print ('Corpus size = ' + str(len(allemails)) + ' emails')
# Ausgabe der wichtigsten Wörter
# holt zuerst WÖrter aus der Email raus, übergibt sie an den Lemmatizer und zählt diese
alleEigenschaften = [(get_features(email, ''), label) for (email, label) in allemails]
#gibt die gesammelten Wörter zuvor aus
print ('Collected ' + str(len(alleEigenschaften)) + ' feature sets')
# Aufruf des Trainings des Classifiers
#alle Eigenschaften werden an train übergeben und einen Stichprobenanteil von 80%
trainingset, testset, classifier = train(alleEigenschaften, 0.8)
# Aufruf der Leistungsbewertung
leistung(trainingset, testset, classifier)
f = open('/code/klasse.txt', 'wb')
pickle.dump(classifier, f)
f.close()
#nutzen des classifiers ohne die einspeisung
f = open('/code/klasse.txt', 'rb')
classifier = pickle.load(f)
f.close()
read_email_from_gmail()
<file_sep>FROM python:3
ADD . /code
RUN apt-get update
RUN apt-get install -y python3 python-dev python-pip
RUN pip3 install -U nltk
RUN apt-get install -y python-scipy
RUN python3 -m nltk.downloader stopwords
RUN python3 -m nltk.downloader punkt
RUN python3 -m nltk.downloader wordnet
WORKDIR /code
ENV DISPLAY :0
COPY . .
CMD [ "python3", "./FertigesProjektKrommMäder.py" ]<file_sep><H1>SPAMHINATOR</H1>
Dieser Spamfilter wurde im Rahmen eines DHBW-Projektes, für Neue Konzepte, ein Modul des Kurses WWI15AMB entwickelt.
Dieser Spamfilter soll maximal im Privaten genutzt werden.
Verbesserungen sind möglich.
Die Hauptdatei ist FertigesProjektKrommMäder.py
Der Spamfilter wurde in Python entwickelt mit dem Hintergrund dessen, dass zum einen die Wahl der Programmiersprache frei war und zum anderen nützliche Packages bereits vorhanden waren, wodurch umständliche Entwicklungen, besonders die benötigten Algorithmen betreffend, nicht notwendig waren. Weitere vorhandene Dateien sind klasse.txt, welche sogenannte "classifier" beinhaltet. Dieses sind einzelne Wörter, welche auf Spam-Emails hinweisen. Weiterhin gibt es im MeineHam und MeineSpam Ordner entsprechende Training- und Testsets zum Einspeisen des classifiers.
Der angesprochene Algorithmus ist der NaiveBayes Algorithmus. Dieser ordnet, zumindest in Bezug auf Emails, die ausgewählten Mails anhand der Trefferzahlen (Statistiken) in Ham (Kein Spam) oder Spam.
Zur Nutzung dieses Codes müssen zuvor einige Anpassungen am Code erfolgen. Besonders die Pfade müssen angepasst werden. Um den Spamfilter individueller zu gestalten, können die genutzten Testsets um eigene Mails erweitert werden. Diese sollten das Dateiformat .txt oder .rtf benutzen. Andere Formate könnten eventuell nicht erkannt werden.
Weiterhin kann durch den Originalcode nur Gmail verwendet werden. Die API wurde nicht eingebunden, da sie äußerst viele Fehlermeldungen, welche nicht auf Dauer behoben werden konnte, ausgeworfen hat. Die hier genutzte Lösung ist für den Privatzweck jedoch mehr als ausreichend.
Um den Spamfilter erfolgreich nutzen zu können muss Python 3 installiert werden. Ebenso ein Editor für Python da die Datei bearbeitet werden muss. Zu empfehlen ist Idle. Falls bereits Python installiert ist sollte ein Upgrade durchgeführt werden.
Für den Bayer Spamfilter mittels Python werden im Normalfall Sklearn oder NLTK genutzt. Hier wurde NLTK genutzt.
Sollte es nicht ausführbar sein muss die Python Version überprüft werden.
Zusätzlich wurde im Rahmen des Projektes ein Dockerfile erstellt.
Zu beachten ist, dass die Sets lediglich beim erstmaligen Gebrauch genutzt werden müssen. Im Anschluss kann der Code zum initialisieren auskommentiert werden. Jedoch sollte der Code, welcher die Datei klasse.txt aufruft nicht auskommentiert werden, da andernfalls keine Spam oder Ham Einordnung stattfinden kann.
<H2>Anleitung</H2>
Die einfachste Nutzung dieses Spamfilters ist die Ausführungs mittels eines Editors, z.B. Idle.
Andernfalls kann es ebenfalls durch Docker ausgeführt werden.
<H3>!BITTE BEACHTEN!</H3>
Sollte es mittels Docker ausgeführt werden, dann muss (im Normalfall) kein Pfad angepasst werden.
Wird es jedoch über einen Editor genutzt lokal auf dem Rechner, dann müssen die entsprechenden Pfade in der Hauptdatei "FertigesProjektKrommMäder.py" angepasst werden.
<H3>!BITTE BEACHTEN!</H3>
Zunächst muss dafür Docker installiert werden. Sämtliche Dateien, inklusive des Dockerfile sollen bitte in einen Ordner lokal heruntergeladen werden.
Anschließend wird mittels des Dockerfiles ein Dockerimage erstellt. Dazu muss zunächst das Terminal geöffnet werden, anschließend der Ordner mittels 'cd' ausgewählt werden und folgender Befehl eingegeben werden:
<p align="center">docker build -t "deine namensgebung":"versionsnummer" .<p align="center">
<p align="center">Bsp: docker build -t meinimage:1.0 .</p align="center">
Sollte es nicht funktionieren bitte folgenden Befehl ausführen und den oberen nochmals ausführen:
<p align="center">docker pull python3 (oder python)</p align="center">
Nach erfolgreichem erstellen des Images kann ein Container erstellt und zum laufen gebracht werden.
Hierzu folgende Befehle ausführen:
<p align="center">docker run --name "mein toller containername" -p "":"" "deine namensgebung":"versionsnummer" (Vgl. oben)</p align="center">
<p align="center">Bsp: docker run --name containername -p 80:80 meinimage:1.0</p align="center">
Um die Ausführung des Containers zu beenden kann folgender Befehl genutzt werden:
<p align="center">docker stop "mein toller containername"</p align="center">
Und für die Löschung des Container:
<p align="center">docker rm "mein toller containername"</p align="center">
<H3>Abschließend</H3>
Python ist eine sehr eigenwillige Sprache. Um ein Python Programm, unabhängig davon wie aufwendig es ist, zum Laufen zu bringen, sollte beachtet werden, dass kein Zeitdruck bestehen soll. Das Installieren entsprechender Packete, um das Programm ausführen zu können, nimmt viel Zeit in Anspruch.
| 314184f82e96a74b553f585a7ae4a434a8e7c12d | [
"Markdown",
"Python",
"Dockerfile"
] | 3 | Python | Valeriekrm/valfelprojekt | ff88081f289d1394a2e6b2e5e1c902fc81b40740 | 213cb59e24a8c9967cd08b3a6ac036fdc848380e |
refs/heads/master | <file_sep>import collections
import operator
import re
def make_histogram(d):
histogram = [(l, len(i)) for l, i in d.iteritems()]
histogram = sorted(histogram, key=operator.itemgetter(1))
return histogram
class Hex(object):
"""
Data we store about a hex.
"""
def __init__(self, location, settlement, author, description, url, themes):
self.location = location
self.settlement = settlement.upper().strip()
self.themes = [t.upper().strip() for t in themes.split(',')]
self.author = author
self.description = description or '-'.encode('utf-8')
self.url = url
class HexMap(object):
def __init__(self, csvfile):
"""
Expects a neatly formated CSV file and generates a data structure
storing descriptions and information about the area described. The
expected fields, in order, are:
0 1 2 3 4 5 6 7
X, Y, UNUSED, Hex Key, Terrain, Settlement(s), UNUSED, Author, ...
8 9 10
... Description, URL, Themes
"""
self.hexes = collections.defaultdict(list)
self.themes = collections.defaultdict(list)
self.settlements = {}
for h in csvfile:
h = Hex(h[3], h[5], h[7], h[8], h[9], h[10])
if not h.location.isdigit() or not h.author:
# skip empty / junky hexes
continue
if h.settlement:
self.settlements[h.settlement] = h.location
for t in h.themes:
self.themes[t].append(h.location)
self.hexes[h.location].append(h)
# Yank out all the authors
self.authors = [d.author
for l, details in self.hexes.iteritems() for d in details
if d.author]
self.author_histogram = collections.Counter(self.authors)
# Yank out all references
self.references = collections.defaultdict(set)
for l, details in self.hexes.iteritems():
for d in details:
for m in re.finditer(r"\[\[(\d\d\d\d)\]\]", d.description):
if l != m.group(1):
self.references[m.group(1)].add(l)
for m in re.finditer(r"\[\[(.*?)\]\]", d.description):
settlement = m.group(1).upper().strip()
if not settlement.isdigit() and settlement in self.settlements:
location = self.settlements[settlement]
if l != location:
self.references[location].add(l)
@property
def reference_histogram(self):
return make_histogram(self.references)
@property
def themes_histogram(self):
return make_histogram(self.themes)
| f5acf36d60ff19348fa9c4a05811ca5c82de177b | [
"Python"
] | 1 | Python | dauten/hexenbracken | 5aa0010310ec9c946d91864df5cba882536bcf53 | 0971bc21549a6eadb423e91f4e58b58b6f455cd5 |
refs/heads/master | <file_sep>/**
* @file 工具函数集合
*/
/**
* @name 生成[0, n]范围内的随机数
* @param {Number} 范围上限
* @return {Number} 随机数
*/
const rand = (n) => {
return parseInt(Math.random() * n)
}
module.exports = {rand}<file_sep>/**
* 小程序样例 - 首页
*/
const app = getApp()
const fakeData = require('../../utils/mock')
const { rand } = require('../../utils/utils')
Page({
data: {
motto: '这个人很懒,什么都没有留下', // 个性签名
username: '', // 用户名
userImg: '/images/avatar.jpg', // 头像图片地址
hasUserInfo: false,
listContent: []
},
onLoad: function () {
// 加载用户数据以及模拟数据
if (app.globalData.userInfo) {
const {nickName, avatarUrl} = app.globalData.userInfo; // 优先加载缓存中数据
this.setData({
hasUserInfo: true,
username: nickName,
userImg: avatarUrl
})
} else {
wx.getUserInfo({
success: res => {
app.globalData.userInfo = res.userInfo // 做一个数据缓存
const { nickName, avatarUrl } = res.userInfo;
this.setData({
hasUserInfo: true,
username: nickName,
userImg: avatarUrl
})
}
})
}
setTimeout(() => {
// 模拟网络环境
this.setData({
listContent: fakeData
})
}, rand(1000))
}
})
<file_sep>/**
* @file 生成模拟数据
*/
const { rand } = require('./utils')
const imgUrl = '/images/user-bg.jpg'
const fakeText = '这里风景很不错'
const today = new Date().getTime();
/**
* @name 生成假图片动态
* @return {
* type: {String} 动态种类
* text: {String} 动态内容
* img: {String} 动态图片url
* }
*/
const fakeImage = () => {
return {
type: 'image',
text: new Array(rand(5) + 1).fill(fakeText),
img: imgUrl
}
}
/**
* @name 生成假分享数据
* @return {
* type: {String} 动态类型
* text: {String} 动态文字
* shareText: {String} 分享文字
* shareImg: {String} 分享缩略图地址
* }
*/
const fakeShare = () => {
return {
type: 'share',
text: new Array(rand(2) + 1).fill(fakeText),
shareImg: imgUrl,
shareText: new Array(rand(3) + 1).fill(fakeText)
}
}
/**
* @name 生成假日期
* @param {Number} n 当前日往前数n天
* @reuturn {String, String} {month, day}
*/
const fakeDate = (n) => {
const date = new Date(today - 24 * 3600 * 1000 * n);
const day = date.getDate(); // 获取日期
const month = (date.getMonth() + 1).toString(); // 获取月份
return {
month,
day: day < 10 ? `0${day}` : `${day}` // 个位数前面补零
}
}
module.exports = new Array(rand(10) + 3).fill(0).map((v, i) => {
return {
date: fakeDate(i),
content: new Array(rand(3)).fill(0).map((v, i) => {
// 每天最多生成3条动态
return Math.random() > 0.6 ? fakeShare() : fakeImage()
})
}
}).filter(v => v.content.length > 0) // 筛选掉无动态的日子
<file_sep>## 小白健康小程序代码阅读心得
> 可能会有不准确或错误的地方,希望大家交流指正
### 序言
经过这两天的阅读,心里基本对整个项目有了一个较为清晰的认识,项目其实并不复杂,但是受限于微信小程序官方api的限制,很多功能不得不自建轮子,比如宋亮说的实现地不太优雅的组件化系统,为了方便对比,本文拿当下比较完善的小程序框架`wepy`来进行对比,对比内容也着重放在组件化的实现上。此外,笔者也会将阅读代码时发现的可改进之处提出来,供大家交流讨论。
### 代码结构与编码规范
其实前端工程化的实践经由`React`, `Angular`和`Vue`几大框架历练至今,已经形成了一个相对成熟的形态,文件路径的设置以及文件夹的结构和命名基本上时大同小异,所以只要稍微对前端工程化熟悉一点,就可以很快上手一个新的项目。
本项目也不例外,各个文件夹职责分明,结构清晰,从文件夹名基本就可以判断出其职责所在,这一点上也无可挑剔。
继而讨论编码规范,这里有两点需要吐槽的,下面一一来讲。
1. 驼峰式与下划线式命名方式共存。这个问题其实说大不大,说小不小,命名方式这种东西带有很强的个人色彩,经过笔者观察,本项目所有的自建函数都是采用下划线式命名,但W3C标准制定的 HTML Dom api 却无一例外采用的驼峰式命名,前端社区也基本都是提倡使用驼峰式命名来写js。但鉴于项目本身代码量已经不少了,为这个问题大动干戈也有些不实际,这里只是稍微提一下。
2. 注释缺失或格式不统一。相比上面命名问题,笔者认为这个问题更重要一点,因为注释的质量直接关系到了整个代码的可读性,良好的注释习惯可以大幅降低沟通成本。经笔者观察,项目中大部分文件是**没有一行注释的**,可能开发者在进行开发时认为这个文件逻辑简单,不需要太多注释,但注释是一个编码习惯问题,人在编码时对复杂度的判断标准是时刻波动的,举个例子,你在精力充沛时可以对一个包含200+行代码和10多个函数的文件了如指掌,但只要过个半天回来看,会发现还要花上一段时间来熟悉代码。所以笔者建议对本项目的注释加强规范,注释标准可参考[JSdoc](http://www.css88.com/doc/jsdoc/index.html)。一些简单的例子可以查看附录1。
### 组件化实践——对比`wepy`
笔者认为,小程序的组件化绝对是小程序开发中最大的痛点,可能官方对小程序的定位是仅承载轻量交互的即开即用的载体,但现在显然越来越多的初创公司选择将交互重心放在了webapp和小程序这类易于分发的载体上了,项目复杂度也随之上升,更完善的组件化也愈发重要。
小程序原生提供的组件化很奇怪,引用官方的描述:
> wxs 模块均为单例,wxs 模块在第一次被引用时,会自动初始化为单例对象。多个页面,多个地方,多次引用,使用的都是同一个 wxs 模块对象。
由于小程序的框架是采用与vue类似的数据与视图绑定逻辑,但小程序把数据、视图和样式剖分成了三个文件,运行时通过文件名将三者绑定在一起,其中`wxml`的模块化逻辑是,模板的引用相当于把被引用模板的代码拷贝到引用处,也就是说,被引用模块与引用模块共用一个数据作用域,被引用的模块没有自己的作用域,这就从根本上决定了组件嵌套在小程序框架下是无法实现的。
对于上述模块化出现的问题,`wepy`采用了预处理的思路,编码时采用类似`vue`的单页面组件,视图、数据和逻辑写在同一`.wpy`文件中,然后通过一个预处理程序将`.wpy`文件解耦为`wxml/wxss/wxs`,有了预处理层,才有了真正意义上的模块化,如下图所示:

`wepy`通过对方法和数据加前缀预处理的方式,实现了作用域隔离,从而实现了组件嵌套,这其实也是`React/Vue`一直在做的事情,这也是为什么`React/Vue`在渲染列表子元素的时候一定要指定`key`的原因。
然后再说一说本项目采用的组件化思想,本项目直接将所有的可复用组件放在一个大模块(`components/component`)里面,需要用的时候只需要分别引用这个`component`里的三大文件。`components/base`用纯js方法对模块进行了封装,利用`component_id`对各个组件的数据进行分割,这样只需要在`wxml`中将`component_id`与对应的template进行捆绑,即可实现数据作用域的组件化,相当于把`wepy`中的预处理操作手动实现了。

### 交互改进
一个小地方需要注意,就是在用户第一次进入小白健康的时候,需要选择个人信息,笔者在填年龄那一节时,发现无论前一步骤选的是“欧巴”还是“大叔”,年龄的年份选择框总是从1975年开始,这里可以稍微优化下,比如用户选了“欧巴”,年份就从1990年开始;用户选了“大叔”,就从1980年开始。
### 附录-1:一份符合JSdoc规范的样例代码
```javascript
/**
* Book类,代表一个书本.
* @constructor
* @param {string} title - 书本的标题.
* @param {string} author - 书本的作者.
*/
function Book(title, author) {
this.title=title;
this.author=author;
}
Book.prototype={
/**
* 获取书本的标题
* @returns {string|*}
*/
getTitle:function(){
return this.title;
},
/**
* 设置书本的页数
* @param pageNum {number} 页数
*/
setPageNum:function(pageNum){
this.pageNum=pageNum;
}
};
```
### 附录-2:一页朋友圈UI预览图
 | 6e18e8ec97f91c1c5da6d2df7225629b60ceb3bc | [
"JavaScript",
"Markdown"
] | 4 | JavaScript | Yidadaa/miniapp-example | ce9370c3fccfefe07631194279df9496a271ffe3 | 0e7b980a26548c8ed3c475a440838504c103bab0 |
refs/heads/master | <repo_name>golsby/async<file_sep>/README.md
async
=====
A simple example of how to use the async/await pattern in C# 4.5.
Goal: actually perform a time-consuming operation that isn't already wrapped up in a task. That is, I've found myriad examples of how to write code that does asynchronous HTTP requests using code somebody else wrote to do the heavy lifting.
But for Rhino, we need to know how to write and call tasks that do things like intersections and boolean operations.
##Running the Example
1. Clone this repository
2. Open async.sln in Visual Studio 2012
3. Compile and Run
4. Select a Computation Method
5. Click Compute...
6. While the computation is happening, click the Change Background button.
##Examples
###Synchronous
This demonstrates and unresponsive UI because the computation is happening on the UI thread.
###Asynchronous
The UI is alive - you can change the background color and move the dialog - but there's no indication of what's going on.
###Asynchronous with Progress
The UI is alive, and periodic progress reports come back from the computation thread.
###Asynchronous with Progress and Cancel
A cancellable operation that reports progress, and of course, keeps the UI alive.
##The Meat
The nugget in this entire example is calling **Task.Run()** to spawn the task I've written in a separate thread. By default, .NET doesn't spawn threads, it uses [cooperative multitasking](http://stackoverflow.com/questions/13993750/the-async-and-await-keywords-dont-cause-additional-threads-to-be-created) on the same thread.
<file_sep>/FactorialDialog.cs
using System;
using System.Drawing;
using System.Threading.Tasks;
using System.Threading;
using System.Windows.Forms;
namespace async
{
public partial class FactorialDialog : Form
{
CancellationTokenSource m_cancellation_token_source;
public FactorialDialog()
{
InitializeComponent();
}
#region Button Handlers
// Note that this method has to be decorated with async, otherwise I can't
// call async methods inside it.
private async void buttonCompute_Click(object sender, EventArgs e)
{
// Initialize the Result field to show we're computing.
labelResult.Text = "Wait...";
Int64 n = Convert.ToInt32(txtInput.Text);
Int64 result = 0;
Progress<Int64> progress = new Progress<Int64>();
switch (comboComputeMethod.Text)
{
case "Synchronous":
// Notice how the UI is unresponsive when using this call.
result = ComputeFactorial(n, null, CancellationToken.None);
labelResult.Text = "Answer: " + result.ToString();
return;
case "Asynchronous":
// Here we have a responsive UI, but no indication of progress, and no way to cancel.
result = await ComputeFactorialAsync(n);
break;
case "Async with Progress":
progress.ProgressChanged += ProgressChanged;
result = await ComputeFactorialAsync(n, progress);
break;
case "Async with Progress and Cancel":
progress.ProgressChanged += ProgressChanged;
m_cancellation_token_source = new CancellationTokenSource();
try
{
buttonCancel.Enabled = true;
result = await ComputeFactorialAsync(n, progress, m_cancellation_token_source.Token);
}
catch (OperationCanceledException)
{
labelResult.Text = "Computation Cancelled";
progress.ProgressChanged -= ProgressChanged;
return;
}
finally
{
buttonCancel.Enabled = false;
}
break;
}
// Display the answer.
labelResult.Text = "Answer: " + result.ToString();
}
/// <summary>
/// Handler for progress reports while computing Factorial
/// </summary>
/// <param name="sender"></param>
/// <param name="e"></param>
void ProgressChanged(object sender, Int64 e)
{
labelResult.Text = "Computing... " + e.ToString();
}
/// <summary>
/// Toggle background color of the form. This is useful to test if the UI is still alive.
/// </summary>
/// <param name="sender"></param>
/// <param name="e"></param>
private void buttonChangeBackground_Click(object sender, EventArgs e)
{
this.BackColor = this.BackColor == Color.White ? Color.Gainsboro : Color.White;
}
// Handler for cancel button.
private void buttonCancel_Click(object sender, EventArgs e)
{
if (m_cancellation_token_source != null)
m_cancellation_token_source.Cancel();
}
#endregion
/// <summary>
/// Compute Factorial of n on a separate thread.
/// </summary>
/// <param name="n">Number to compute factorial from</param>
/// <returns>The factorial of input 'n'</returns>
Task<Int64> ComputeFactorialAsync(Int64 n)
{
return Task.Run(
() => ComputeFactorial(n, null, CancellationToken.None)
);
}
/// <summary>
/// Compute Factorial of n on a separate thread and report progress.
/// </summary>
/// <param name="n">Number to compute factorial from</param>
/// <param name="progress">Optional class to report progress. Pass null if you don't care to receive progress.</param>
/// <returns>The factorial of input 'n'</returns>
Task<Int64> ComputeFactorialAsync(Int64 n, IProgress<Int64> progress)
{
return Task.Run(
() => ComputeFactorial(n, progress, CancellationToken.None)
);
}
/// <summary>
/// Compute Factorial of n on a separate thread, supporting cancellation and reporting of progress.
/// </summary>
/// <param name="n">Number to compute factorial from</param>
/// <param name="progress">Optional class to report progress. Pass null if you don't care to receive progress.</param>
/// <param name="cancellationToken">Optional CancellationToken. Pass CancellationToken.None if you don't care to cancel.</param>
/// <returns>The factorial of input 'n'</returns>
Task<Int64> ComputeFactorialAsync(Int64 n, IProgress<Int64> progress, CancellationToken cancellationToken)
{
return Task.Run(
() => ComputeFactorial(n, progress, cancellationToken)
);
}
/// <summary>
/// ComputeFactorial is a time consuming function (mostly because it sleeps for half a second at each iteration)
/// that reports progress and optionally can be cancelled.
/// </summary>
/// <param name="n">Number to compute factorial from</param>
/// <param name="progress">Optional class to report progress. Pass null if you don't care to receive progress.</param>
/// <param name="cancellationToken">Optional CancellationToken. Pass CancellationToken.None if you don't care to cancel.</param>
/// <returns>The factorial of input 'n'</returns>
Int64 ComputeFactorial(Int64 n, IProgress<Int64> progress, CancellationToken cancellationToken)
{
Int64 result = 1;
for (Int64 i = n; i > 1; i--)
{
Thread.Sleep(500);
result *= i;
if (progress != null)
progress.Report(result);
if (cancellationToken.IsCancellationRequested)
throw new OperationCanceledException();
}
return result;
}
}
}
| 209029999855658662aae5eedd6bd3f844e3689b | [
"Markdown",
"C#"
] | 2 | Markdown | golsby/async | 53b3c3169901bec2ff17c6950c522748c44fa86a | e3435f85ede6693d81fbc1b4b2fc57d93421c885 |
refs/heads/master | <repo_name>echoywp/getWordContent<file_sep>/test.php
<?php
use Word\action;
require 'vendor/autoload.php';
$class = new action();
$class->setLoadPath('./test.docx');
$res = $class->getHtml();
print_r($res);
<file_sep>/README.md
# getWordContent
PHP获取Word内容
<file_sep>/src/Word/action.php
<?php
namespace Word;
use PhpOffice\PhpWord\IOFactory;
class action {
private $load_path;
private $file;
public function __construct() {
$this->setFileName();
}
protected $error = [
10001 => 'File not found',
10002 => 'Wrong file format',
];
protected $mineType = [
'application/zip',
];
public function handle() {
$check = $this->check();
if ($check) {
return $this->error[$check];
}
$word = IOFactory::load($this->load_path);
$html = IOFactory::createWriter($word, "HTML");
$html->save($this->file);
}
/**
* @param $load_path
* 设置读取文件路径
*/
public function setLoadPath($load_path) {
$this->load_path = $load_path;
}
/**
* @param $path
* @param $filename
* 设置临时文件名
* @return string
*/
public function setFileName($path = '', $filename = '') {
$suffix = '.html';
$filename = $filename ? $filename : ('TMP' . time() . rand(1000, 9999) . $suffix);
$save_path = $path ? $path : (__DIR__. './');
$this->file = $save_path . $filename;
}
/**
* @return bool|int
* 基础校验
*/
protected function check() {
if (!file_exists($this->load_path)) {
return 10001;
}
$file_info = mime_content_type($this->load_path);
if (!in_array($file_info, $this->mineType)) {
return 10002;
}
return false;
}
/**
* @return bool|mixed
* 获取html内容
*/
public function getHtml() {
self::handle();
$content = file_get_contents($this->file);
preg_match_all('/<body>(.*)<\/body>/iUs', $content, $res);
if (array_key_exists(1, $res)) {
return $res[1][0];
}
return false;
}
/**
* 删除文件
*/
public function __destruct() {
unlink($this->file);
}
}
| 0b884fb35e778f477a9b9c33dcdde3918464119e | [
"Markdown",
"PHP"
] | 3 | PHP | echoywp/getWordContent | 55b3fdb3bf3338f88ba6201ded8e45eaa89ffff6 | 889a4e7db06476338184e077bac8cefefb5181d4 |
refs/heads/master | <repo_name>marcofuentes05/Figuras<file_sep>/src/Comportamientos/Describible.kt
package Comportamientos
interface Describible {
//Un objeto describible ha de tener un atributo alto y un metodo pedirDatos()
var alto: Int
fun pedirDatos()
}<file_sep>/src/Figuras/Cuadrado.kt
package Figuras
import Comportamientos.Describible
import Comportamientos.Dibujable
open class Cuadrado: Dibujable, Describible {
override var alto =0
//El constructor no hace nada
constructor(){
alto = 0
}
//El metodo pedirDatos solo solicita los datos -_-
override fun pedirDatos() {
println("Ingrese el alto del cuadrado")
try{
alto = readLine()!!.toInt()
}catch(e : Exception){
println("Ese no es un dato valido, por eso, el valor por defecto es 5")
alto = 5
}
}
// El metodo dibujar dibuja cada figura segun sus propiedades, usando ciclos
override fun dibujar(): String{
var a : String = ""
for (i in 1..alto){
for (b in 1..alto){
a = a + "* "
}
a = a + "\n"
}
return a
}
}
<file_sep>/src/main.kt
import Comportamientos.Describible
import Comportamientos.Dibujable
import Figuras.Cuadrado
import Figuras.Rectangulo
import Figuras.Triangulo
fun main (args : Array <String>){
// Se definen los strings que se imprimirán
var menu : String = """
Menu Principal
---------------
1. Dibujar Figura
2. Salir
""".trimIndent()
var menuF : String = """
Menu de Figuras
---------------
1. Dibujar un Cuadrado
2. Dibujar un Rectángulo
3. Dibujar un Triángulo
4. Regresar al menu principal
""".trimIndent()
// Estas variables funcionan de control para los ciclos
var seguir : Boolean = true
var respuesta : String
while (seguir){
println(menu)
respuesta = readLine()!!.toString()
var seguir1 : Boolean = true
//Si el usuario desea dibujar una figura,se imprime el siguiente menu, hasta que decida salir
if (respuesta == "1"){
while (seguir1){
//Se imprime el segundo menu y se solicita una respuesta
println(menuF)
respuesta = readLine()!!.toString()
//Se evalua su respuesta y para cada caso se instancia un objeto, y se le aplica el metodo pedirDatos y dibujarFigura de
// las respectivas interfaces
when (respuesta){
"1" ->{
var cuadrado = Cuadrado()
pedirDatos(cuadrado)
println(dibujarFigura(cuadrado))
}
"2"->{
var rectangulo = Rectangulo()
pedirDatos(rectangulo)
println(dibujarFigura(rectangulo))
}
"3"->{
var triangulo = Triangulo()
pedirDatos(triangulo)
println(dibujarFigura(triangulo))
}
"4"->seguir1 =false
}
}
}else{
seguir = false
}
}
}
//Estas funciones unen los objetos a sus interfaces en la clase principal
fun dibujarFigura(dibujable: Dibujable): String{
return dibujable.dibujar()
}
fun pedirDatos(describible: Describible){
describible.pedirDatos()
}<file_sep>/src/Comportamientos/Dibujable.kt
package Comportamientos
interface Dibujable {
//Un objeto dibujable ha de tener un metodo dibujar()
fun dibujar(): String
}<file_sep>/src/Figuras/Rectangulo.kt
package Figuras
import Comportamientos.Describible
import Comportamientos.Dibujable
class Rectangulo: Dibujable, Describible {
override var alto =0
var ancho: Int = 0
override fun pedirDatos() {
println("Ingrese el alto del rectángulo: ")
try{
alto = readLine()!!.toInt()}
catch(e: Exception){
println("Ese no es un dato valido, por eso, el valor por defecto es 5")
alto = 5
}
println("Ingrese el ancho del rectángulo: ")
try{
ancho = readLine()!!.toInt()}
catch(e: Exception){
println("Ese no es un dato valido, por eso, el valor por defecto es 5")
ancho = 5
}
}
override fun dibujar(): String{
var a : String = ""
for (i in 1..alto){
for (j in 1..ancho){
a = a + "* "
}
a = a + "\n"
}
return a
}
}<file_sep>/src/Figuras/Triangulo.kt
package Figuras
import Comportamientos.Describible
import Comportamientos.Dibujable
class Triangulo: Dibujable, Describible {
override var alto = 0
override fun pedirDatos() {
println("Ingrese el alto del triángulo: ")
try{
alto = readLine()!!.toInt()}
catch(e:Exception){
println("Ese no es un dato valido, por eso, el valor por defecto es 5")
alto = 5
}
}
override fun dibujar(): String {
var a : String = ""
var c : Int = 1
for (i in 1..alto){
for(k in 1..(alto-c)){
a = a + " "
}
for (j in 1..c){
a = a + "* "
}
c = c + 1
a = a + "\n"
}
return a
}
} | 263a91cd3085676d5218e542ad1f4628a96db283 | [
"Kotlin"
] | 6 | Kotlin | marcofuentes05/Figuras | 8c695a8319e1256a5bbacdc5d1b8dfd02c5dbf1a | ea45425115770547bb38d6fcc4964047d5edb95b |
refs/heads/master | <file_sep>MQTT Practice
========
This if for [OpenIRC](https://github.com/openirc) project
```bash
# Install dependencies
bundle
# Set username & password
cp secret.toml.example secret.toml
vim secret.toml
# Try it!
./run pub
./run sub
```
###### Prerequisite
- Ruby *≥2.0.0*
<file_sep>#!/bin/bash
# Change to script directory
cd "$( dirname "${BASH_SOURCE[0]}" )"
# Execute
bundle exec ruby src/main.rb "$@"
<file_sep># frozen_string_literal: true
require 'mqtt'
require 'tomlrb'
# Parse ARGV
if ARGV.length == 1 and ARGV[0] == 'pub'
MODE = :publish
elsif ARGV.length == 1 and ARGV[0] == 'sub'
MODE = :subscribe
else
puts <<END
usage: run [pub | sub]
Commands:
pub Run publishing sample
sub Run subscribing sample
END
exit 1
end
# Load configs
config = Tomlrb.load_file('secret.toml', symbolize_keys: true)
host = config.dig(:config, :hostname)
port = config.dig(:config, :port)
username = config.dig(:credentials, :username)
password = config.dig(:credentials, :password)
# Connect to MQTT broker
print '연결중 ... '
MQTT::Client.connect(host: host, port: port, username: username, password: <PASSWORD>) do |c|
puts "\e[32m완료!\e[0m"
case MODE
when :publish
loop.with_index do |_, i|
c.publish('mqtt-practice', "야호 #{i}")
sleep 1
end
when :subscribe
c.get('mqtt-practice') do |topic, message|
puts "#{topic}: #{message}"
end
end
end
<file_sep># frozen_string_literal: true
source 'https://rubygems.org'
gem 'mqtt'
gem 'tomlrb'
<file_sep># vim: ft=toml
# Configs about MQTT broker
[config]
hostname = 'test.mosquitto.org'
port = 1883
# If required, uncomment the lines below and write your username and password to
# connect with MQTT broker
# [credentials]
# username = "account_name_here"
# password = "<PASSWORD>"
| 0d3422e07fb0062ede95c55245e71536da8c73ef | [
"Markdown",
"TOML",
"Ruby",
"Shell"
] | 5 | Markdown | simnalamburt/mqtt-practice | 9a5fec8132311a364d451dff4e1d0ad4e71ad070 | 976871b42bee302f52bb5a4d8c8777333ce3eb7a |
refs/heads/main | <repo_name>franklaguy/react-app<file_sep>/README.md
# react-app
React App Concept
| ebaf9f38f9ba221e817d04fd6b0770af2b057ce8 | [
"Markdown"
] | 1 | Markdown | franklaguy/react-app | 5e7284df5d5c5b83ed3328f83505e27e9fe6f3cd | af0ae2978420a0bff08a63e6e8f284c17d3ed8fc |
refs/heads/master | <file_sep># SCpersonal
Aplicación de desarrollo web con PHP, Mysql y Bootstrap.
<h3><font color='blue'>Sistema de Gestión de Personal.</font></h3>
Es una aplicación realizada con tecnología Bootstrap. la cual se integra con PHP, Mysql, Jquery y mas...
Para iniciar debes de tener el la guia de desarrollo del sistemas el cual esta publicado en Slideshare.
1.- Instalar el servidor web con la aplicación XAAMP<br>
2.- Crear la BD bd_planilla, Descargar script copleto aqui.<br>
3.- Crear las estrutura de carpetas y organizar las librerias de css y jquery que estan en "assets".<br>
4.- Inciar a crear los archivos segun Manual.<br>
Creado por <NAME>.<br>
<EMAIL><br>
www.jeefperu.com<br>
La siguiente aplicación se puede utilizar y realizar las mejoras necesarias<br>
Derecho reservado.
2018
<file_sep>/*
SQLyog Enterprise Trial - MySQL GUI v5.01
Host - 5.5.5-10.1.36-MariaDB : Database - planilla
*********************************************************************
Server version : 5.5.5-10.1.36-MariaDB
*/
/*Table structure for table `areas` */
CREATE TABLE `areas` (
`id_area` tinyint(4) NOT NULL,
`nom_area` varchar(50) NOT NULL,
`est_area` char(1) NOT NULL,
PRIMARY KEY (`id_area`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `areas` */
insert into `areas` values (1,'Recursos Humanos','1');
insert into `areas` values (2,'Producción','1');
insert into `areas` values (3,'Contabilidad','1');
/*Table structure for table `bonificacion_personal` */
CREATE TABLE `bonificacion_personal` (
`idbonfi` int(11) NOT NULL,
`id_tipo_bonifi` int(11) DEFAULT NULL,
`idpersonal` int(11) DEFAULT NULL,
`idplanilla` int(11) DEFAULT NULL,
`idperiodo` int(11) DEFAULT NULL,
`nummes` varchar(2) DEFAULT NULL,
`valor_bonfi` decimal(10,2) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idbonfi`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Table structure for table `cargos` */
CREATE TABLE `cargos` (
`id_cargo` tinyint(4) NOT NULL,
`nom_cargo` varchar(50) NOT NULL,
`est_cargo` char(1) NOT NULL,
PRIMARY KEY (`id_cargo`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `cargos` */
insert into `cargos` values (1,'Conserje','1');
insert into `cargos` values (2,'Secretaria','1');
insert into `cargos` values (3,'Contador','1');
insert into `cargos` values (4,'Administrador','1');
insert into `cargos` values (5,'Programador','0');
/*Table structure for table `configurar_planilla` */
CREATE TABLE `configurar_planilla` (
`idconfig` int(11) NOT NULL,
`idperiodo` int(11) DEFAULT NULL,
`idtipopla` int(11) DEFAULT NULL,
`numes` varchar(2) DEFAULT NULL,
`obs` text,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idconfig`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `configurar_planilla` */
insert into `configurar_planilla` values (1,1,1,'01','Pago de Trabajadores','1');
insert into `configurar_planilla` values (2,1,1,'02','','0');
insert into `configurar_planilla` values (3,1,1,'03','','0');
insert into `configurar_planilla` values (4,1,2,'03','Utilidades','0');
insert into `configurar_planilla` values (5,2,1,'01','','0');
/*Table structure for table `descuento_personal` */
CREATE TABLE `descuento_personal` (
`idtdsto` int(11) NOT NULL,
`id_tipo_dscto` int(11) NOT NULL,
`id_personal` int(11) DEFAULT NULL,
`id_planilla` int(11) DEFAULT NULL,
`id_periodo` int(11) DEFAULT NULL,
`nummes` varchar(2) DEFAULT NULL,
`valor_dscto` decimal(10,2) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idtdsto`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
CREATE TABLE `periodo` (
`idperiodo` int(11) NOT NULL,
`nameperiodo` varchar(50) DEFAULT NULL,
`namecorto` varchar(20) DEFAULT NULL,
`fechareg` date DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idperiodo`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `periodo` */
insert into `periodo` values (1,'2019','19','2019-01-01','1');
/*Table structure for table `personal` */
CREATE TABLE `personal` (
`id` int(11) NOT NULL AUTO_INCREMENT,
`codigoper` varchar(100) NOT NULL,
`dno` char(8) DEFAULT NULL,
`nombre` varchar(150) NOT NULL,
`apellido` varchar(150) NOT NULL,
`email` varchar(200) NOT NULL,
`Passwordp` varchar(180) NOT NULL,
`sexo` varchar(100) NOT NULL,
`fechanac` varchar(100) NOT NULL,
`direccion` varchar(255) NOT NULL,
`ciudad` varchar(255) NOT NULL,
`pais` varchar(200) NOT NULL,
`telefono` varchar(11) NOT NULL,
`area` int(11) NOT NULL,
`cargo` int(11) DEFAULT NULL,
`sueldo` decimal(10,2) DEFAULT NULL,
`estado` int(1) NOT NULL,
`fecIngreso` date DEFAULT NULL,
`sistema` char(3) NOT NULL,
`numero` varchar(25) DEFAULT NULL,
`tipouser` tinyint(4) DEFAULT '0',
PRIMARY KEY (`id`)
) ENGINE=InnoDB AUTO_INCREMENT=7 DEFAULT CHARSET=latin1;
/*Data for the table `personal` */
insert into `personal` values (1,'EMP10806121','34343433','Jose','Fernandez','<EMAIL>','45e340b2402d2f6271a6c49e8e7ba06d','M','2019-02-15','Av las palmeras N° 546','Chiclayo','sochi','6587976',1,4,'434.00',1,'2017-11-10','ONP','',1);
insert into `personal` values (3,'09907887','14567890','Romina','<NAME>','<EMAIL>','','F','2019-01-30','Jr. Cusco N° 344','Trujillo','cusco','343434',1,2,'122.00',1,'2018-06-26','ONP','',0);
insert into `personal` values (5,'','12362514','Carlos','<NAME>','<EMAIL>','','M','2019-02-13','Jr la Unión N° 3434','Monsefu','','979343451',2,3,'1250.00',1,'2019-02-14','ONP','0001',0);
insert into `personal` values (6,'0006','34543543','Jacinto','<NAME>','<EMAIL>','202cb962ac59075b964b07152d234b70','M','2019-02-14','Los treboles','Tuman','','77878',1,3,'3434.00',1,'2019-02-14','AFP','34',1);
/*Table structure for table `planilla` */
CREATE TABLE `planilla` (
`idplanilla` int(11) NOT NULL,
`fecha` date DEFAULT NULL,
`hora` time DEFAULT NULL,
`idperiodo` int(11) DEFAULT NULL,
`idtipoplanilla` int(11) DEFAULT NULL,
`idtrabajador` int(11) DEFAULT NULL,
`sueldo` decimal(10,2) DEFAULT NULL,
`total_dscto` decimal(10,2) DEFAULT NULL,
`total_bonf` decimal(10,2) DEFAULT NULL,
`netopagar` decimal(10,2) DEFAULT NULL,
`estadoplanilla` char(1) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idplanilla`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `planilla` */
/*Table structure for table `sistema_pension` */
CREATE TABLE `sistema_pension` (
`idtipo` int(11) NOT NULL,
`name_sp` varchar(150) DEFAULT NULL,
`valor_sp` decimal(10,2) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idtipo`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `sistema_pension` */
/*Table structure for table `tipo_bonificacion` */
CREATE TABLE `tipo_bonificacion` (
`idbonificacion` int(11) NOT NULL,
`namebonificacion` varchar(125) DEFAULT NULL,
`namecorto` varchar(5) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idbonificacion`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `tipo_bonificacion` */
insert into `tipo_bonificacion` values (1,'Horas extras',NULL,'1');
insert into `tipo_bonificacion` values (2,'Famillar',NULL,'1');
insert into `tipo_bonificacion` values (3,'Comisiones',NULL,'1');
/*Table structure for table `tipo_descuento` */
CREATE TABLE `tipo_descuento` (
`iddescuento` int(11) NOT NULL,
`namedescuento` varchar(125) DEFAULT NULL,
`nombrecorto` varchar(5) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`iddescuento`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `tipo_descuento` */
insert into `tipo_descuento` values (1,'AFP',NULL,'1');
insert into `tipo_descuento` values (2,'Seguro SIS',NULL,'1');
insert into `tipo_descuento` values (3,'Asignacion Familiar',NULL,'0');
insert into `tipo_descuento` values (4,'Seguro EsSalud',NULL,'1');
/*Table structure for table `tipoplanilla` */
CREATE TABLE `tipoplanilla` (
`idtipoplanilla` int(11) NOT NULL,
`nameplanilla` varchar(50) DEFAULT NULL,
`estado` char(1) DEFAULT NULL,
PRIMARY KEY (`idtipoplanilla`)
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
/*Data for the table `tipoplanilla` */
insert into `tipoplanilla` values (1,'Empleados','1');
insert into `tipoplanilla` values (2,'Obreros','1');
insert into `tipoplanilla` values (3,'Services','1');
insert into `tipoplanilla` values (4,'Externa','1');
| db3fcc9882776be9bac4c33bef768f4e162a79e0 | [
"Markdown",
"SQL"
] | 2 | Markdown | jeefernandez/SCpersonal | 1a751564d89476de9051ba29419e04cfa1d283af | a2b6476c5e5fcc7955f906aa467b8b6c5792b086 |
refs/heads/master | <repo_name>candrec/curso_programacao<file_sep>/src/TesteString.java
public class TesteString {
public static void main(String[] args) {
String original = "abcde FGHIJ ABC abc DEFG ";
String s="Potato apple lemon";
String[] vect=s.split(" ");
String s01=original.toLowerCase();
String s02=original.toUpperCase();
String s03=original.trim();
String s04=original.substring(2);
String s05=original.substring(2, 9);
String s06=original.replace('a', 'x');
String s07=original.replace("abc", "xy");
int i=original.indexOf("bc");
int j=original.lastIndexOf("bc");
System.out.println("original: -"+original+"-");
System.out.println("toLowerCase: -"+s01+"-");
System.out.println("toUpperCase: -"+s02+"-");
System.out.println("trim: -"+s03+"-");
System.out.println("subString(2): -"+s04+"-");
System.out.println("subString(2,9): -"+s05+"-");
System.out.println("replace('a','x'): -"+s06+"-");
System.out.println("replace(\"abc\",\"xy\"): -"+s07+"-");
System.out.println("Index of 'bc': "+i);
System.out.println("Last index of 'bc': "+j);
System.out.println(vect[0]);
System.out.println(vect[1]);
System.out.println(vect[2]);
}
}
<file_sep>/src/TesteFor.java
import java.util.Scanner;
public class TesteFor {
public static void main(String[] args) {
Scanner sc = new Scanner(System.in);
int N = sc.nextInt();
int aux,soma=0;
for(int i=0;i<N;i++) {
aux = sc.nextInt();
soma += aux;
}
System.out.println(soma);
sc.close();
}
}
<file_sep>/src/TesteDoWhile.java
import java.util.Locale;
import java.util.Scanner;
public class TesteDoWhile {
public static void main(String[] args) {
Locale.setDefault(Locale.US);
Scanner sc = new Scanner(System.in);
double celsius;
double fahren;
char condicao;
do {
System.out.print("Digite a temperatura em Celsius: ");
celsius=sc.nextDouble();
fahren=((9*celsius)/5)+32;
System.out.printf("Equivalente em Fahrenheit: %.1f%n",fahren);
System.out.print("Deseja repetir (s/n): ");
condicao=sc.next().charAt(0);
}while(condicao=='s');
sc.close();
}
}
<file_sep>/src/Main2.java
import java.util.Locale;
import java.util.Scanner;
public class Main2 {
public static void main(String[] args) {
Locale.setDefault(Locale.US);
Scanner sc = new Scanner(System.in); // inicia uma variável da classe Scanner para ajudar na entrada de dadosA
String x;
int y;
double z;
char sexo;
System.out.println("Digite uma string");
x = sc.next(); // captura a string do teclado
System.out.println("Digite um inteiro: ");
y = sc.nextInt(); // captura o inteiro do teclado
System.out.println("Digite um ponto flutuante: ");
z = sc.nextDouble(); // captura o ponto flutuante do teclado
System.out.println("Qual o sexo: (F/M)");
sexo = sc.next().charAt(0); // captura o primeiro caractere da string digitada
System.out.println("A string digitada foi " + x);
System.out.println("O inteiro digitado foi " + y);
System.out.println("O ponto flutuante é " + z);
System.out.println("O sexo é " + sexo);
sc.close();
}
}
| df006e14864f37d693ed61d48275b3156b400b99 | [
"Java"
] | 4 | Java | candrec/curso_programacao | d84d7b5652c9e08622573059d6bc5fa365066e26 | bf10072d79419da6922fa21320a1968d94298572 |
refs/heads/master | <file_sep><?php
class Csv {
private $fh;
protected $eol;
protected $delimiter;
protected $enclosure;
function __construct($fileName = '', $savePath = false, $eol = "\n", $delimiter = ",", $enclosure = '"') {
if ($fileName) {
$this->fileName = $fileName;
}
// NOTE: I don't think custom new lines work with direct downloads
// In these cases, you may need to save the file first then manually serve it for download
$this->eol = $eol;
$this->delimiter = $delimiter;
if ($savePath) {
$this->fh = fopen($savePath.$fileName.".csv", 'w');
} else {
$this->fh = fopen('php://output', 'w');
header("Content-type: text/csv");
header("Content-Disposition: attachment; filename={$fileName}.csv");
header("Pragma: no-cache");
header("Expires: 0");
}
}
function write($fields = array()) {
fputcsv($this->fh, $fields, $this->delimiter, $this->enclosure);
// Have we specified a custom EOL? If so, apply to the row
if("\n" != $this->eol && 0 === fseek($this->fh, -1, SEEK_CUR)) {
fwrite($this->fh, $this->eol);
}
}
function close() {
fclose($this->fh);
}
}
<file_sep>PHP EOL CSV Class
=============
This exists for one specific purpose - to provide a class for creating CSV files in PHP, while also allowing you to set a custom end-of-line character.
fputcsv allows you to specify a field delimiter, but strangely doesn't let you set the EOL character - which, when it insists on \n, isn't much use for files destined for Windows based systems.
I may extend this to do more in future, but for now it does exactly what I needed it for.
| 68242827d2192ec62a204750c2823218ac43311b | [
"Markdown",
"PHP"
] | 2 | PHP | matthewfedak/php-eol-csv | 9e125ff95e1edc974e32f0aed664bfe5974aba9b | 58629f7ebc4c609d689a08cf47e2c222b2104337 |
refs/heads/master | <file_sep>using Emgu.CV;
using Emgu.CV.OCR;
using Emgu.CV.Structure;
using System;
using System.Collections.Generic;
using System.ComponentModel;
using System.Data;
using System.Drawing;
using System.IO;
using System.Linq;
using System.Text;
using System.Threading.Tasks;
using System.Windows.Forms;
using System.Xml;
using Tesseract;
using AForge;
using AForge.Imaging;
using AForge.Imaging.Filters;
using System.Drawing.Imaging;
using MySql.Data.MySqlClient;
namespace WindowsFormsApplication29
{
public partial class Form1 : Form
{
public Form1()
{
InitializeComponent();
}
private void button1_Click(object sender, EventArgs e)
{
var img = new Bitmap(pictureBox2.Image);
var ocr = new TesseractEngine(@"C:\Users\User\Desktop\anil_bitir\WindowsFormsApplication29\WindowsFormsApplication29\bin\Debug\tessdata", "eng", EngineMode.TesseractAndCube);
var page = ocr.Process(img);
textBox1.Text = page.GetText();
}
Bitmap KesilenResim;
private void Form1_Load(object sender, EventArgs e)
{
pictureBox4_Click(sender,e);
}
private void button3_Click(object sender, EventArgs e)
{
Bitmap kaynakResmi, filtreliResim;
OtsuThreshold otsuFiltre = new OtsuThreshold();
kaynakResmi = KesilenResim;
//orjinal resim gösteriliyor
pictureBox2.Image = kaynakResmi;
//resmi eğer renkliyse önce griye çeviriyor sonra filtre uyguluyor
//resim zaten griyse direk filtre uyguluyor
filtreliResim = otsuFiltre.Apply(kaynakResmi.PixelFormat != PixelFormat.Format8bppIndexed ? Grayscale.CommonAlgorithms.BT709.Apply(kaynakResmi) : kaynakResmi);
//filtre uygulanan resim gösteriliyor
pictureBox3.Image = filtreliResim;
//Uygulanan Threshold Değeri form başlığında görünüyor
this.Text = "Threshold Değeri : " + otsuFiltre.ThresholdValue.ToString();
button1.Enabled = true;
}
int j = 0;
string[] dizi = new string[600];
public void plakaKes(int a)
{
CascadeClassifier Classifier = new CascadeClassifier(@"C:\Users\User\Desktop\anil_bitir\car4\cascade\cascade.xml");
String image_name = dizi[a];
label1.Text = dizi[a];
Mat img = CvInvoke.Imread(image_name, Emgu.CV.CvEnum.ImreadModes.AnyColor);
Image<Bgr, Byte> imgInput = img.ToImage<Bgr, Byte>();
var imgGray = imgInput.Convert<Gray, byte>();
Rectangle[] rectangles = Classifier.DetectMultiScale(imgGray, 1.2, 1, new Size(10, 10), new Size(1000, 1000));
foreach (var rectangle in rectangles)
{
CvInvoke.Rectangle(img, rectangle, new MCvScalar(0, 0, 255), 2);
Image<Bgr, Byte> img_cut = imgInput;
img_cut.ROI = rectangle;
}
System.Drawing.Image pMyImage = imgInput.ToBitmap(); /// Kesilmiş olan plakayı pictureboxa yükledik.
pictureBox2.Image = pMyImage;
Image<Bgr, Byte> de = img.ToImage<Bgr, Byte>(); /// Araç resmini pictuereBoxa verdik.
System.Drawing.Image pMyImage1 = de.ToBitmap();
pictureBox1.Image = pMyImage1;
KesilenResim = imgInput.ToBitmap();
button3.Enabled = true;
}
string[] ayir=new string[50];
string[] ArananPlakalar;
public void Veritabanı()
{
try
{
string aranan = "";
int i = 1;
MySqlConnection mysqlbaglan = new MySqlConnection("Server=localhost;Database=plakatanima;Uid=root;Pwd='';SslMode=none");
MySqlDataReader dr;
MySqlCommand cmd;
char[] ayrac = { ' ', '\n' };
mysqlbaglan.Open();
cmd = new MySqlCommand("select * from arananaraclar", mysqlbaglan);
dr = cmd.ExecuteReader();
while (dr.Read())
{
aranan = dr["Plaka"].ToString() + " " + aranan;
ayir = aranan.Split(ayrac);
}
string[] bulunan = textBox1.Text.Split(ayrac);
for (int q = 1; q < bulunan.Length; q += 3)
{
if (ayir[q] == bulunan[i])
{
if (ayir[q + 1] == bulunan[i + 1])
{
label2.Visible = true;
timer1.Enabled = true;
break;
}
}
}
mysqlbaglan.Close();
}
catch
{
}
}
private void timer1_Tick(object sender, EventArgs e)
{
timer1.Enabled = false;
label2.Visible = false;
}
private void pictureBox4_Click(object sender, EventArgs e) // ileri butonu
{
int i = 0;
string dosya_yolu = @"C:\Users\User\Desktop\anil_bitir\WindowsFormsApplication29\WindowsFormsApplication29\arabaa.txt";
FileStream fs = new FileStream(dosya_yolu, FileMode.Open, FileAccess.Read);
StreamReader sw = new StreamReader(fs);
//Okuma işlemi için bir StreamReader nesnesi oluşturduk.
string yazi = sw.ReadLine();
while (yazi != null)
{
if (i == 600)
{
break;
}
dizi[i] = yazi.ToString();
i++;
yazi = sw.ReadLine();
}
sw.Close();
fs.Close();
plakaKes(j);
j++;
button3_Click(sender, e);
button1_Click(sender, e);
Veritabanı();
}
private void pictureBox5_Click(object sender, EventArgs e)
{
j--;
plakaKes(j);
}
}
}
<file_sep>-- phpMyAdmin SQL Dump
-- version 4.7.9
-- https://www.phpmyadmin.net/
--
-- Anamakine: 127.0.0.1
-- Üretim Zamanı: 05 Haz 2018, 11:48:04
-- Sunucu sürümü: 10.1.31-MariaDB
-- PHP Sürümü: 7.2.3
SET SQL_MODE = "NO_AUTO_VALUE_ON_ZERO";
SET AUTOCOMMIT = 0;
START TRANSACTION;
SET time_zone = "+00:00";
/*!40101 SET @OLD_CHARACTER_SET_CLIENT=@@CHARACTER_SET_CLIENT */;
/*!40101 SET @OLD_CHARACTER_SET_RESULTS=@@CHARACTER_SET_RESULTS */;
/*!40101 SET @OLD_COLLATION_CONNECTION=@@COLLATION_CONNECTION */;
/*!40101 SET NAMES utf8mb4 */;
--
-- Veritabanı: `plakatanima`
--
-- --------------------------------------------------------
--
-- Tablo için tablo yapısı `arananaraclar`
--
CREATE TABLE `arananaraclar` (
`id` int(11) NOT NULL,
`Plaka` varchar(50) NOT NULL
) ENGINE=InnoDB DEFAULT CHARSET=latin1;
--
-- Tablo döküm verisi `arananaraclar`
--
INSERT INTO `arananaraclar` (`id`, `Plaka`) VALUES
(1, '06 EA 7114'),
(2, '58 TV 036');
--
-- Dökümü yapılmış tablolar için indeksler
--
--
-- Tablo için indeksler `arananaraclar`
--
ALTER TABLE `arananaraclar`
ADD PRIMARY KEY (`id`);
--
-- Dökümü yapılmış tablolar için AUTO_INCREMENT değeri
--
--
-- Tablo için AUTO_INCREMENT değeri `arananaraclar`
--
ALTER TABLE `arananaraclar`
MODIFY `id` int(11) NOT NULL AUTO_INCREMENT, AUTO_INCREMENT=3;
COMMIT;
/*!40101 SET CHARACTER_SET_CLIENT=@OLD_CHARACTER_SET_CLIENT */;
/*!40101 SET CHARACTER_SET_RESULTS=@OLD_CHARACTER_SET_RESULTS */;
/*!40101 SET COLLATION_CONNECTION=@OLD_COLLATION_CONNECTION */;
<file_sep># PlakaOkuma-EmguCv- | 41f20f700a7fc693df3e0614ef3e8d66a5682d7f | [
"Markdown",
"C#",
"SQL"
] | 3 | C# | yesim142536/PlakaOkuma-EmguCv- | ebca5a10f5ccd76ab4dec36823d0023895b447fb | 6be08b802f60bff573348f8d14cb0541bd9a8220 |
refs/heads/master | <file_sep>from mcpi.minecraft import Minecraft
mc = Minecraft.create()
x,y,z=mc.player.getPos()
a=0
while a<20:
mc.setBlocks(x,y-1,z+30,x,y-10,z-30,19)
x=x-5
a=a+1<file_sep>from mcpi.minecraft import Minecraft
mc = Minecraft.create()
x,y,z=mc.player.getPos()
mc.setSign(x,y,z,63,0,"我愛","Minecraft")
<file_sep>import time
import random
from mcpi.minecraft import Minecraft
mc = Minecraft.create()
x,y,z= mc.player.getTilePos()
color=random.randrange(0,16)
mc.setBlocks(x+25,y-1,z+25,x-25,y-1,z-25,95,color)
<file_sep>from mcpi.minecraft import Minecraft
mc = Minecraft.create()
x,y,z=mc.player.getPos()
def plantTree(x,y,z):
mc.setBlock(x+1,y+5,z+1,x-1,y+3,z-1,18)
mc.setBlock(x,y,z,x,y+4,z,17)
for h in range(0,10):
for i in range(0,5):
plantTree(x+i,y,z+h)<file_sep>
from mcpi.minecraft import Minecraft
mc = Minecraft.create()
length=20
width=10
height=5
x,y,z=mc.player.getTilePos()
x2=x+length
y2=y+height
z2=z+width
mc.setBlocks(x,y,z,x2,y2,z2,51)
mc.setBlocks(x+1,y+1,z+1,x2-1,y2-1,z2-1,0) | 56352eac8431205e58b301d07194d09999dd7ebc | [
"Python"
] | 5 | Python | felix20080608/mPython2020-8 | 0c6b90cf3ec3d14c13af52f26ba0ae6f3f995783 | 9e5d434966c4c6a3870e79aa1375a8ad11e3a3a9 |
refs/heads/master | <file_sep>from testpy import myclass
def self(args):
pass
myclass.run(self)
<file_sep>import sfml as sf
mwindow = sf.RenderWindow(sf.VideoMode(640, 480), "SFML Application")
mplayer = sf.CircleShape()
class myclass:
mplayer.radius = 40
mplayer.position = (100, 100)
mplayer.fill_color = sf.Color.CYAN
def run(self):
while mwindow.is_open:
myclass.processevents(self)
myclass.update(self)
myclass.render(self)
def processevents(self):
event = sf.Event
for event in mwindow.events:
if type(event) is sf.CloseEvent:
mwindow.close()
# player movement
if sf.Keyboard.is_key_pressed(sf.Keyboard.W):
mplayer.position += (0,-5)
elif sf.Keyboard.is_key_pressed(sf.Keyboard.A):
mplayer.position -= (5,0)
elif sf.Keyboard.is_key_pressed(sf.Keyboard.S):
mplayer.position -= (0,-5)
elif sf.Keyboard.is_key_pressed(sf.Keyboard.D):
mplayer.position += (5,0)
def update(self):
pass
def render(self):
mwindow.clear()
mwindow.draw(mplayer)
mwindow.display() | 6244f35542a7b404c69cc5bd2cb3e420bdcd9033 | [
"Python"
] | 2 | Python | Thizze/AWPP | adc3adcf191c50992a1c9fdd414fd5d13f273fe1 | 02265dc2114dc4a29c8f4ea450dc3ab7a0db3988 |
refs/heads/master | <repo_name>mguterl/twallaby<file_sep>/dist/js/twallaby.js
this["JST"] = this["JST"] || {};
this["JST"]["app/templates/overview.us"] = function(obj) {
obj || (obj = {});
var __t, __p = '', __e = _.escape;
with (obj) {
__p += '<div id="overview" class="step" data-scale="' +
((__t = ( this.collection.size() / 3 )) == null ? '' : __t) +
'"></div>';
}
return __p
};
this["JST"]["app/templates/tweet.us"] = function(obj) {
obj || (obj = {});
var __t, __p = '', __e = _.escape;
with (obj) {
__p += '<div class="tweet__avatar">\n <img src="' +
((__t = ( get('bigger_profile_image_url') )) == null ? '' : __t) +
'" />\n</div>\n<div class="tweet__content">\n <p class="tweet__author">@' +
((__t = ( get('from_user') )) == null ? '' : __t) +
'</p>\n <p class="tweet__text">' +
((__t = ( get('text') )) == null ? '' : __t) +
'</p>\n <p class="tweet__timestamp" title="' +
((__t = ( get('created_at') )) == null ? '' : __t) +
'"></p>\n</div>\n';
}
return __p
};
(function() {
window.Twallaby = (function() {
function Twallaby(options) {
if (options == null) {
options = {};
}
this.tweetsCollection = new Twallaby.TweetsCollection(options.tweets, {
twitterParams: options.twitterParams,
urlRoot: options.urlRoot,
pollInterval: options.pollInterval
});
this.stepsView = new Twallaby.StepsView({
el: options.element,
collection: this.tweetsCollection
});
this.stepsView.render();
}
Twallaby.prototype.startPolling = function() {
return this.tweetsCollection.startPolling();
};
Twallaby.prototype.fetch = function() {
return this.tweetsCollection.fetch();
};
return Twallaby;
})();
Twallaby.Tweet = Backbone.Model.extend({
initialize: function() {
if (this.get('profile_image_url') != null) {
return this.set('bigger_profile_image_url', this.get('profile_image_url').replace('normal', 'bigger'));
} else {
return this.set('bigger_profile_image_url', void 0);
}
}
});
Twallaby.TweetsCollection = Backbone.Collection.extend({
model: Twallaby.Tweet,
twitterParams: {
page: 1,
rpp: 20,
result_type: 'recent'
},
urlRoot: 'http://search.twitter.com/search.json',
pollInterval: 60000,
initialize: function(models, options) {
if (options == null) {
options = {};
}
this.twitterParams = _.extend({}, this.twitterParams, options.twitterParams);
if (options.urlRoot) {
this.urlRoot = options.urlRoot;
}
if (options.pollInterval) {
return this.pollInterval = options.pollInterval;
}
},
startPolling: function(interval) {
var _this = this;
this.fetch();
return this.interval = setInterval((function() {
return _this.fetch();
}), interval || this.pollInterval);
},
stopPolling: function() {
return clearInterval(this.interval);
},
url: function() {
var key, urlParts, value, _ref;
urlParts = [];
_ref = this.twitterParams;
for (key in _ref) {
value = _ref[key];
urlParts.push("" + key + "=" + value);
}
urlParts.push("callback=?");
return encodeURI("" + this.urlRoot + "?" + (urlParts.join("&")));
},
parse: function(resp, xhr) {
return resp.results;
}
});
Twallaby.cssHelper = {
translate: function(coords) {
return "translate3d(" + (coords.x || 0) + "px," + (coords.y || 0) + "px," + (coords.z || 0) + "px)";
},
rotate: function(coords) {
return "rotateX(" + (coords.x || 0) + "deg) rotateY(" + (coords.y || 0) + "deg) rotateZ(" + (coords.z || 0) + "deg)";
},
scale: function(s) {
return "scale(" + (s || 1) + ")";
},
perspective: function(p) {
return "perspective(" + p + "px)";
}
};
$.fn.applyStyles = function(styles) {
return this.each(function() {
var key, value, _results;
_results = [];
for (key in styles) {
value = styles[key];
_results.push(this.style[Modernizr.prefixed(key)] = value);
}
return _results;
});
};
Twallaby.randomHelper = {
integerBetween: function(from, to) {
if (from == null) {
from = 0;
}
if (to == null) {
to = 0;
}
return Math.floor(Math.random() * (to - from + 1) + from);
}
};
Twallaby.StepView = Backbone.View.extend({
className: "step",
defaultStyles: {
position: "absolute",
transformStyle: "preserve-3d",
transition: "all 0.5s ease-in-out"
},
currentPosition: {
x: 0,
y: 0,
z: 0
},
currentRotation: {
x: 0,
y: 0,
z: 0
},
currentScale: 1,
position: function(positions) {
this.currentPosition = positions;
this.stylize();
return this;
},
scale: function(scale) {
this.currentScale = scale;
this.stylize();
return this;
},
rotate: function(rotations) {
this.currentRotation = rotations;
this.stylize();
return this;
},
stylize: function() {
var tranforms;
tranforms = ["translate(-50%, -50%)", Twallaby.cssHelper.translate(this.currentPosition), Twallaby.cssHelper.rotate(this.currentRotation), Twallaby.cssHelper.scale(this.currentScale)].join(" ");
return this.$el.applyStyles({
transform: tranforms
});
}
});
Twallaby.AnnouncementView = Twallaby.StepView.extend({
className: "step announcement",
render: function() {
this.$el.applyStyles(this.defaultStyles);
this.position({
x: this.$el.data('x') || 0,
y: this.$el.data('y') || 0,
z: this.$el.data('z') || 0
});
this.rotate({
x: this.$el.data('rotate-x') || 0,
y: this.$el.data('rotate-y') || 0,
z: this.$el.data('rotate-z') || 0
});
this.scale(this.$el.data('scale') || 0);
return this;
}
});
Twallaby.StepsView = Backbone.View.extend({
id: "impress",
template: JST['app/templates/overview.us'],
delay: 4000,
announcementInterval: 5,
perspective: 1000,
transitionDuration: 1000,
events: {
'click': 'go'
},
paused: false,
stepCount: 0,
defaultStyles: {
position: "absolute",
transformOrigin: "top left",
transition: "all 1s ease-in-out",
transformStyle: "preserve-3d"
},
initialize: function() {
this.$canvas = this.$el.children(':first');
this.tweetViews = [];
this.announcementViews = [];
return this.listenTo(this.collection, 'add', function(model) {
this.repositionTweetViews();
this.appendTweet(model);
return this.applyPerspective();
});
},
render: function() {
this.$canvas.applyStyles(this.defaultStyles);
this.$el.applyStyles(this.defaultStyles);
this.$el.applyStyles({
top: "50%",
left: "50%"
});
this.applyPerspective();
this.renderAnnouncements();
this.renderTweets();
return this;
},
applyPerspective: function() {
return this.$el.applyStyles({
transform: "" + (Twallaby.cssHelper.scale(1))
});
},
renderTweets: function() {
var model, _i, _len, _ref, _results;
_ref = this.collection.models;
_results = [];
for (_i = 0, _len = _ref.length; _i < _len; _i++) {
model = _ref[_i];
_results.push(this.appendTweet(model));
}
return _results;
},
renderAnnouncements: function() {
var announcementView, element, _i, _len, _ref, _results;
_ref = this.$canvas.children();
_results = [];
for (_i = 0, _len = _ref.length; _i < _len; _i++) {
element = _ref[_i];
announcementView = new Twallaby.AnnouncementView({
el: element
});
announcementView.render();
_results.push(this.announcementViews.push(announcementView));
}
return _results;
},
appendTweet: function(model) {
var tweetView;
tweetView = new Twallaby.TweetView({
model: model
});
this.position(tweetView);
this.$canvas.append(tweetView.render().el);
return this.tweetViews.push(tweetView);
},
position: function(view) {
var index, radius, theta;
radius = 100 * this.collection.size();
theta = Math.PI * 2 / this.collection.size();
index = this.collection.indexOf(view.model);
return view.position({
x: Math.floor(radius * Math.cos(index * theta)),
y: Math.floor(radius * Math.sin(index * theta)),
z: 0
});
},
repositionTweetViews: function() {
var view, _i, _len, _ref, _results;
_ref = this.tweetViews;
_results = [];
for (_i = 0, _len = _ref.length; _i < _len; _i++) {
view = _ref[_i];
_results.push(this.position(view));
}
return _results;
},
go: function() {
if (this.stepCount % this.announcementInterval === 0) {
return this.goToAnnouncement();
} else {
return this.goToTweet();
}
},
goTo: function(view) {
var targetPosition;
targetPosition = {
x: -view.currentPosition.x,
y: -view.currentPosition.y,
z: -view.currentPosition.z
};
this.$canvas.applyStyles({
transform: Twallaby.cssHelper.translate(targetPosition)
});
return this.stepCount += 1;
},
goToTweet: function(index) {
return this.goTo(this.tweetViews[index] || this.tweetViews[this.randomTweetIndex()]);
},
goToAnnouncement: function(index) {
return this.goTo(this.announcementViews[index] || this.announcementViews[this.randomAnnouncementIndex()]);
},
randomAnnouncementIndex: function() {
return Twallaby.randomHelper.integerBetween(0, this.announcementViews.length - 1);
},
randomTweetIndex: function() {
return Twallaby.randomHelper.integerBetween(0, this.tweetViews.length - 1);
}
});
Twallaby.TweetView = Twallaby.StepView.extend({
className: "step tweet",
template: JST['app/templates/tweet.us'],
render: function() {
this.$el.html(this.template(this.model));
this.$('.tweet__timestamp').timeago();
this.$el.applyStyles(this.defaultStyles);
return this;
}
});
}).call(this);
<file_sep>/README.md
# Twallaby
### Install
* `npm install`
* `npm install lineman -g`
### Run
* `lineman run`
### Test
* `lineman spec`
### Build
* `lineman build`
### API
var tweetWall = new Twallaby({
element: $('#slides')
pollInterval: 10000
twitterParams:
q: 'CWTDrinkup'
rpp: 25
})
tweetWall.startPolling() | 8f7c4a56ce65d8ea7ce0521624b478275375b3e4 | [
"JavaScript",
"Markdown"
] | 2 | JavaScript | mguterl/twallaby | bcf9ab1bf728c6e6fc02ba95149e4ffeca9b9a51 | 764b5b2f4f5729f6d844e43f9f7db9762b508750 |
refs/heads/master | <file_sep>// Copyright 2020 The Moov Authors
// Use of this source code is governed by an Apache License
// license that can be found in the LICENSE file.
package imagecashletter
import (
"strings"
"testing"
)
// mockCredit creates a Credit
func mockCredit() *Credit {
cr := NewCredit()
cr.AuxiliaryOnUs = "010910999940910"
cr.ExternalProcessingCode = ""
cr.PayorBankRoutingNumber = "999920060"
cr.CreditAccountNumberOnUs = "50920060509383521210"
cr.ItemAmount = 102088
cr.ECEInstitutionItemSequenceNumber = " "
cr.DocumentationTypeIndicator = "G"
cr.AccountTypeCode = "1"
cr.SourceWorkCode = "3"
cr.WorkType = " "
cr.DebitCreditIndicator = " "
return cr
}
// TestMockCredit creates a CreditItem
func TestMockCredit(t *testing.T) {
ci := mockCredit()
if err := ci.Validate(); err != nil {
t.Error("mockCredit does not validate and will break other tests: ", err)
}
if ci.recordType != "61" {
t.Error("recordType does not validate")
}
if ci.AuxiliaryOnUs != "010910999940910" {
t.Error("AuxiliaryOnUs does not validate")
}
if ci.ExternalProcessingCode != "" {
t.Error("ExternalProcessingCode does not validate")
}
if ci.PayorBankRoutingNumber != "999920060" {
t.Error("PayorBankRoutingNumber does not validate")
}
if ci.CreditAccountNumberOnUs != "50920060509383521210" {
t.Error("CreditAccountNumberOnUs does not validate")
}
if ci.ItemAmount != 102088 {
t.Error("ItemAmount does not validate")
}
if ci.ECEInstitutionItemSequenceNumber != " " {
t.Error("ECEInstitutionItemSequenceNumber does not validate")
}
if ci.DocumentationTypeIndicator != "G" {
t.Error("DocumentationTypeIndicator does not validate")
}
if ci.AccountTypeCode != "1" {
t.Error("AccountTypeCode does not validate")
}
if ci.SourceWorkCode != "3" {
t.Error("SourceWorkCode does not validate")
}
if ci.WorkType != " " {
t.Error("WorkType does not validate")
}
if ci.DebitCreditIndicator != " " {
t.Error("DebitCreditIndicator does not validate")
}
}
func TestCreditCrash(t *testing.T) {
cr := &Credit{}
cr.Parse(`61010910999940910 999920060509200605093835212100000102088 G13 `)
if cr.DocumentationTypeIndicator != "G" {
t.Errorf("expected ci.DocumentationTypeIndicator=G")
}
}
func TestParseCredit(t *testing.T) {
var line = "61010910999940910 999920060509200605093835212100000102088 G13 "
r := NewReader(strings.NewReader(line))
r.line = line
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
cr := mockCredit()
r.currentCashLetter.AddCredit(cr)
if err := r.parseCredit(); err != nil {
t.Errorf("%T: %s", err, err)
}
record := r.currentCashLetter.GetCredits()[0]
if record.recordType != "61" {
t.Errorf("RecordType Expected '61' got: %v", record.recordType)
}
if record.AuxiliaryOnUs != "010910999940910" {
t.Errorf("AuxiliaryOnUs Expected '010910999940910' got: %v", record.AuxiliaryOnUs)
}
if record.ExternalProcessingCode != "" {
t.Errorf("ExternalProcessingCode Expected '' got: %v", record.ExternalProcessingCode)
}
if record.PayorBankRoutingNumber != "999920060" {
t.Errorf("PostingBankRoutingNumber Expected '999920060' got: %v", record.PayorBankRoutingNumber)
}
if record.CreditAccountNumberOnUs != "50920060509383521210" {
t.Errorf("OnUs Expected '50920060509383521210' got: %v", record.CreditAccountNumberOnUs)
}
if record.ItemAmount != 102088 {
t.Errorf("ItemAmount Expected '102088' got: %v", record.ItemAmount)
}
if record.ECEInstitutionItemSequenceNumber != " " {
t.Errorf("ECEInstitutionItemSequenceNumber Expected ' ' got: %v", record.ECEInstitutionItemSequenceNumber)
}
if record.DocumentationTypeIndicator != "G" {
t.Errorf("DocumentationTypeIndicator Expected 'G' got: %v", record.DocumentationTypeIndicator)
}
if record.AccountTypeCode != "1" {
t.Errorf("AccountTypeCode Expected '1' got: %v", record.AccountTypeCode)
}
if record.SourceWorkCode != "3" {
t.Errorf("SourceWorkCode Expected '3' got: %v", record.SourceWorkCode)
}
if record.WorkType != " " {
t.Errorf("WorkType Expected ' ' got: %v", record.WorkType)
}
if record.DebitCreditIndicator != " " {
t.Errorf("DebitCreditIndicator Expected ' ' got: %v", record.DebitCreditIndicator)
}
}
// testCIString validates parsing a CreditItem
func testCRString(t testing.TB) {
var line = "61010910999940910 999920060509200605093835212100000102088 G13 "
r := NewReader(strings.NewReader(line))
r.line = line
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
cr := mockCredit()
r.currentCashLetter.AddCredit(cr)
if err := r.parseCredit(); err != nil {
t.Errorf("%T: %s", err, err)
}
record := r.currentCashLetter.GetCredits()[0]
if record.String() != line {
t.Errorf("Strings do not match")
}
}
// TestCRString tests validating that a known parsed CheckDetail can return to a string of the same value
func TestCRString(t *testing.T) {
testCRString(t)
}
// BenchmarkCRString benchmarks validating that a known parsed Credit
// can return to a string of the same value
func BenchmarkCRString(b *testing.B) {
b.ReportAllocs()
for i := 0; i < b.N; i++ {
testCRString(b)
}
}
// TestCRRecordType validation
func TestCRRecordType(t *testing.T) {
ci := mockCredit()
ci.recordType = "00"
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "recordType" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRDocumentationTypeIndicator validation
func TestCRDocumentationTypeIndicator(t *testing.T) {
ci := mockCredit()
ci.DocumentationTypeIndicator = "P"
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "DocumentationTypeIndicator" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRDocumentationTypeIndicatorZ validation
func TestCRDocumentationTypeIndicatorZ(t *testing.T) {
ci := mockCredit()
ci.DocumentationTypeIndicator = "Z"
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "DocumentationTypeIndicator" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRDocumentationTypeIndicatorM validation
func TestCRDocumentationTypeIndicatorM(t *testing.T) {
ci := mockCredit()
ci.DocumentationTypeIndicator = "M"
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "DocumentationTypeIndicator" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRSourceWorkCode validation
func TestCRSourceWorkCode(t *testing.T) {
ci := mockCredit()
ci.SourceWorkCode = "99"
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "SourceWorkCode" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// Field Inclusion
// TestCRFIRecordType validation
func TestCRFIRecordType(t *testing.T) {
ci := mockCredit()
ci.recordType = ""
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "recordType" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRPayorBankRoutingNumber validation
func TestCRPayorBankRoutingNumber(t *testing.T) {
ci := mockCredit()
ci.PayorBankRoutingNumber = "000000000"
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "PayorBankRoutingNumber" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRCreditAccountNumberOnUs validation
func TestCRCreditAccountNumberOnUs(t *testing.T) {
ci := mockCredit()
ci.CreditAccountNumberOnUs = ""
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "CreditAccountNumberOnUs" {
t.Errorf("%T: %s", err, err)
}
}
}
}
// TestCRItemAmount validation
func TestCRItemAmount(t *testing.T) {
ci := mockCredit()
ci.ItemAmount = 0
if err := ci.Validate(); err != nil {
if e, ok := err.(*FieldError); ok {
if e.FieldName != "ItemAmount" {
t.Errorf("%T: %s", err, err)
}
}
}
}
<file_sep>// Copyright 2020 The Moov Authors
// Use of this source code is governed by an Apache License
// license that can be found in the LICENSE file.
package imagecashletter
import (
"bufio"
"bytes"
"crypto/rand"
"encoding/json"
"os"
"path/filepath"
"strconv"
"strings"
"testing"
)
// TestICLFileRead validates reading an ICL file
func TestICLFileRead(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", "BNK20180905121042882-A.icl"))
if err != nil {
t.Errorf("%T: %s", err, err)
}
defer fd.Close()
r := NewReader(fd, ReadVariableLineLengthOption())
_, err = r.Read()
if err != nil {
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*BundleError); ok {
if e.FieldName != "entries" {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
err2 := r.File.Validate()
if err2 != nil {
if e, ok := err2.(*FileError); ok {
if e.FieldName != "BundleCount" {
t.Errorf("%T: %s", e, e)
}
} else {
t.Errorf("%T: %s", err, err)
}
}
}
// TestICLFile validates reading ICL files
func TestICLFiles(t *testing.T) {
files := []string{"BNK20180905121042882-A.icl", "without-micrValidIndicator.icl"}
for _, f := range files {
t.Run(f, func(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", f))
if err != nil {
t.Fatalf("Can not open local file: %s: \n", err)
}
defer fd.Close()
r := NewReader(fd, ReadVariableLineLengthOption())
ICLFile, err := r.Read()
if err != nil {
t.Errorf("Issue reading file: %+v \n", err)
}
t.Logf("r.File.Header=%#v", r.File.Header)
t.Logf("r.File.Control=%#v", r.File.Control)
// ensure we have a validated file structure
if ICLFile.Validate(); err != nil {
t.Errorf("Could not validate entire read file: %v", err)
}
})
}
}
func TestICL_ReadVariableLineLengthOption(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", "valid-ascii.x937"))
if err != nil {
t.Fatalf("Can not open local file: %s: \n", err)
}
defer fd.Close()
r := NewReader(fd, ReadVariableLineLengthOption())
ICLFile, err := r.Read()
if err != nil {
t.Errorf("Issue reading file: %+v \n", err)
}
t.Logf("r.File.Header=%#v", r.File.Header)
t.Logf("r.File.Control=%#v", r.File.Control)
// ensure we have a validated file structure
if ICLFile.Validate(); err != nil {
t.Errorf("Could not validate entire read file: %v", err)
}
actual, err := json.MarshalIndent(ICLFile, "", " ")
if err != nil {
t.Errorf("Issue marshaling file: %+v \n", err)
}
expected, err := os.ReadFile(filepath.Join("test", "testdata", "valid-x937.json"))
if err != nil {
t.Errorf("Issue loading validation criteria: %+v \n", err)
}
if !bytes.Equal(actual, expected) {
t.Errorf("Read file does not match expected JSON")
}
}
func TestICL_EBCDICEncodingOption(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", "valid-ebcdic.x937"))
if err != nil {
t.Fatalf("Can not open local file: %s: \n", err)
}
defer fd.Close()
r := NewReader(fd, ReadVariableLineLengthOption(), ReadEbcdicEncodingOption())
ICLFile, err := r.Read()
if err != nil {
t.Errorf("Issue reading file: %+v \n", err)
}
t.Logf("r.File.Header=%#v", r.File.Header)
t.Logf("r.File.Control=%#v", r.File.Control)
// ensure we have a validated file structure
if ICLFile.Validate(); err != nil {
t.Errorf("Could not validate entire read file: %v", err)
}
actual, err := json.MarshalIndent(ICLFile, "", " ")
if err != nil {
t.Errorf("Issue marshaling file: %+v \n", err)
}
expected, err := os.ReadFile(filepath.Join("test", "testdata", "valid-x937.json"))
if err != nil {
t.Errorf("Issue loading validation criteria: %+v \n", err)
}
if !bytes.Equal(actual, expected) {
t.Errorf("Read file does not match expected JSON")
}
}
// TestRecordTypeUnknown validates record type unknown
func TestRecordTypeUnknown(t *testing.T) {
var line = "1735T231380104121042882201809051523NCitadel Wells Fargo US "
r := NewReader(strings.NewReader(line))
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.FieldName != "recordType" {
t.Errorf("%T: %s", e, e)
}
} else {
t.Errorf("%T: %s", err, err)
}
}
}
// TestFileLineShort validates file line is short
func TestFileLineShort(t *testing.T) {
line := "1 line is only 70 characters ........................................!"
r := NewReader(strings.NewReader(line))
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.FieldName != "RecordLength" {
t.Errorf("%T: %s", e, e)
}
} else {
t.Errorf("%T: %s", e, e)
}
}
}
func TestReaderCrash__parseBundleControl(t *testing.T) {
r := &Reader{}
if err := r.parseBundleControl(); err == nil {
t.Error("expected error")
}
}
// TestFileFileHeaderErr validates error flows back from the parser
func TestFileFileHeaderErr(t *testing.T) {
fh := mockFileHeader()
fh.ImmediateOrigin = ""
r := NewReader(strings.NewReader(fh.String()))
// necessary to have a file control not nil
r.File.Control = mockFileControl()
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestTwoFileHeaders validates one file header
func TestTwoFileHeaders(t *testing.T) {
var line = "0135T231380104121042882201809051523NCitadel Wells Fargo US "
var twoHeaders = line + "\n" + line
r := NewReader(strings.NewReader(twoHeaders))
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileControl {
t.Errorf("%T: %s", e, e)
}
} else {
t.Errorf("%T: %s", err, err)
}
}
}
// TestCashLetterHeaderErr validates error flows back from the parser
func TestCashLetterHeaderErr(t *testing.T) {
clh := mockCashLetterHeader()
clh.DestinationRoutingNumber = ""
r := NewReader(strings.NewReader(clh.String()))
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCashLetterHeaderDuplicate validates when two CashLetterHeader exists in a current CashLetter
func TestCashLetterHeaderDuplicate(t *testing.T) {
// create a new CashLetter header string
clh := mockCashLetterHeader()
r := NewReader(strings.NewReader(clh.String()))
// instantiate a CashLetter in the reader
r.addCurrentCashLetter(NewCashLetter(clh))
// read should fail because it is parsing a second CashLetter Header and there can only be one.
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileCashLetterInside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestBundleHeaderErr validates error flows back from the parser
func TestBundleHeaderErr(t *testing.T) {
bh := mockBundleHeader()
bh.DestinationRoutingNumber = ""
r := NewReader(strings.NewReader(bh.String()))
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestBundleHeaderDuplicate validates when two BundleHeader exists in a current Bundle
func TestBundleHeaderDuplicate(t *testing.T) {
// create a new CashLetter header string
bh := mockBundleHeader()
r := NewReader(strings.NewReader(bh.String()))
// instantiate a CashLetter in the reader
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bhTwo := mockBundleHeader()
r.addCurrentBundle(NewBundle(bhTwo))
// read should fail because it is parsing a second CashLetter Header and there can only be one.
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleInside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailError validates error flows back from the parser
func TestCheckDetailError(t *testing.T) {
cd := mockCheckDetail()
cd.PayorBankRoutingNumber = ""
r := NewReader(strings.NewReader(cd.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailAddendumABundleError validates error flows back from the parser
func TestCheckDetailAddendumABundleError(t *testing.T) {
cd := mockCheckDetail()
cdaddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdaddendumA)
r := NewReader(strings.NewReader(cdaddendumA.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailAddendumBBundleError validates error flows back from the parser
func TestCheckDetailAddendumBBundleError(t *testing.T) {
cd := mockCheckDetail()
cdaddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdaddendumA)
cdaddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdaddendumB)
r := NewReader(strings.NewReader(cdaddendumB.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailAddendumCBundleError validates error flows back from the parser
func TestCheckDetailAddendumCBundleError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
cdAddendumC := mockCheckDetailAddendumC()
cd.AddCheckDetailAddendumC(cdAddendumC)
r := NewReader(strings.NewReader(cdAddendumC.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailAddendumAError validates error flows back from the parser
func TestCheckDetailAddendumAError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cdAddendumA.ReturnLocationRoutingNumber = ""
cd.AddCheckDetailAddendumA(cdAddendumA)
r := NewReader(strings.NewReader(cdAddendumA.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailAddendumBError validates error flows back from the parser
func TestCheckDetailAddendumBError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cdAddendumB.MicrofilmArchiveSequenceNumber = " "
cd.AddCheckDetailAddendumB(cdAddendumB)
r := NewReader(strings.NewReader(cdAddendumB.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailAddendumCError validates error flows back from the parser
func TestCheckDetailAddendumCError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
cdAddendumC := mockCheckDetailAddendumC()
cdAddendumC.EndorsingBankRoutingNumber = ""
cd.AddCheckDetailAddendumC(cdAddendumC)
r := NewReader(strings.NewReader(cdAddendumC.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailError validates error flows back from the parser
func TestReturnDetailError(t *testing.T) {
rd := mockReturnDetail()
rd.PayorBankRoutingNumber = ""
r := NewReader(strings.NewReader(rd.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumABundleError validates error flows back from the parser
func TestReturnDetailAddendumABundleError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
r := NewReader(strings.NewReader(rdAddendumA.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumBBundleError validates error flows back from the parser
func TestReturnDetailAddendumBBundleError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumB := mockReturnDetailAddendumB()
rd.AddReturnDetailAddendumB(rdAddendumB)
r := NewReader(strings.NewReader(rdAddendumB.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumCBundleError validates error flows back from the parser
func TestReturnDetailAddendumCBundleError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumB := mockReturnDetailAddendumB()
rd.AddReturnDetailAddendumB(rdAddendumB)
rdAddendumC := mockReturnDetailAddendumC()
rd.AddReturnDetailAddendumC(rdAddendumC)
r := NewReader(strings.NewReader(rdAddendumC.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumDBundleError validates error flows back from the parser
func TestReturnDetailAddendumDBundleError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumB := mockReturnDetailAddendumB()
rd.AddReturnDetailAddendumB(rdAddendumB)
rdAddendumC := mockReturnDetailAddendumC()
rd.AddReturnDetailAddendumC(rdAddendumC)
rdAddendumD := mockReturnDetailAddendumD()
rd.AddReturnDetailAddendumD(rdAddendumD)
r := NewReader(strings.NewReader(rdAddendumD.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumAError validates error flows back from the parser
func TestReturnDetailAddendumAError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rdAddendumA.ReturnLocationRoutingNumber = ""
rd.AddReturnDetailAddendumA(rdAddendumA)
r := NewReader(strings.NewReader(rdAddendumA.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumBError validates error flows back from the parser
func TestReturnDetailAddendumBError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumB := mockReturnDetailAddendumB()
rdAddendumB.PayorBankSequenceNumber = " "
rd.AddReturnDetailAddendumB(rdAddendumB)
r := NewReader(strings.NewReader(rdAddendumB.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumCError validates error flows back from the parser
func TestReturnDetailAddendumCError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumB := mockReturnDetailAddendumB()
rd.AddReturnDetailAddendumB(rdAddendumB)
rdAddendumC := mockReturnDetailAddendumC()
rdAddendumC.MicrofilmArchiveSequenceNumber = " "
rd.AddReturnDetailAddendumC(rdAddendumC)
r := NewReader(strings.NewReader(rdAddendumC.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailAddendumDError validates error flows back from the parser
func TestReturnDetailAddendumDError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumB := mockReturnDetailAddendumB()
rd.AddReturnDetailAddendumB(rdAddendumB)
rdAddendumC := mockReturnDetailAddendumC()
rd.AddReturnDetailAddendumC(rdAddendumC)
rdAddendumD := mockReturnDetailAddendumD()
rdAddendumD.EndorsingBankRoutingNumber = "000000000"
rd.AddReturnDetailAddendumD(rdAddendumD)
r := NewReader(strings.NewReader(rdAddendumD.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailBundleError validates error flows back from the parser
func TestCheckDetailBundleError(t *testing.T) {
cd := mockCheckDetail()
r := NewReader(strings.NewReader(cd.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailBundleError validates error flows back from the parser
func TestReturnDetailBundleError(t *testing.T) {
rd := mockReturnDetail()
r := NewReader(strings.NewReader(rd.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailIVDetailError validates error flows back from the parser
func TestCheckDetailIVDetailError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
ivDetail := mockImageViewDetail()
ivDetail.ViewDescriptor = ""
cd.AddImageViewDetail(ivDetail)
r := NewReader(strings.NewReader(ivDetail.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailIVDataError validates error flows back from the parser
func TestCheckDetailIVDataError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
ivd := mockImageViewDetail()
cd.AddImageViewDetail(ivd)
ivData := mockImageViewData()
ivData.EceInstitutionRoutingNumber = "000000000"
cd.AddImageViewData(ivData)
r := NewReader(strings.NewReader(ivData.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestCheckDetailIVAnalysisError validates error flows back from the parser
func TestCheckDetailIVAnalysisError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
ivd := mockImageViewDetail()
cd.AddImageViewDetail(ivd)
ivData := mockImageViewData()
cd.AddImageViewData(ivData)
ivAnalysis := mockImageViewAnalysis()
ivAnalysis.GlobalImageQuality = 9
cd.AddImageViewAnalysis(ivAnalysis)
r := NewReader(strings.NewReader(ivAnalysis.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if e.FieldName != "GlobalImageQuality" {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailIVDetailError validates error flows back from the parser
func TestReturnDetailIVDetailError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumC := mockReturnDetailAddendumC()
rd.AddReturnDetailAddendumC(rdAddendumC)
ivDetail := mockImageViewDetail()
ivDetail.ViewDescriptor = ""
rd.AddImageViewDetail(ivDetail)
r := NewReader(strings.NewReader(ivDetail.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailIVDataError validates error flows back from the parser
func TestReturnDetailIVDataError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumC := mockReturnDetailAddendumC()
rd.AddReturnDetailAddendumC(rdAddendumC)
ivDetail := mockImageViewDetail()
rd.AddImageViewDetail(ivDetail)
ivData := mockImageViewData()
ivData.EceInstitutionRoutingNumber = "000000000"
r := NewReader(strings.NewReader(ivData.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if !strings.Contains(e.Msg, msgFieldInclusion) {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestReturnDetailIVAnalysisError validates error flows back from the parser
func TestReturnDetailIVAnalysisError(t *testing.T) {
rd := mockReturnDetail()
rdAddendumA := mockReturnDetailAddendumA()
rd.AddReturnDetailAddendumA(rdAddendumA)
rdAddendumC := mockReturnDetailAddendumC()
rd.AddReturnDetailAddendumC(rdAddendumC)
ivDetail := mockImageViewDetail()
rd.AddImageViewDetail(ivDetail)
ivData := mockImageViewData()
rd.AddImageViewData(ivData)
ivAnalysis := mockImageViewAnalysis()
ivAnalysis.GlobalImageQuality = 9
rd.AddImageViewAnalysis(ivAnalysis)
r := NewReader(strings.NewReader(ivAnalysis.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
b.AddReturnDetail(rd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FieldError); ok {
if e.FieldName != "GlobalImageQuality" {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestIVDetailBundleError validates error flows back from the parser
func TestIVDetailBundleError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
ivDetail := mockImageViewDetail()
cd.AddImageViewDetail(ivDetail)
r := NewReader(strings.NewReader(ivDetail.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
// b.AddCheckDetail(cd)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestIVDataBundleError validates error flows back from the parser
func TestIVDataBundleError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
ivDetail := mockImageViewDetail()
cd.AddImageViewDetail(ivDetail)
ivData := mockImageViewData()
cd.AddImageViewData(ivData)
r := NewReader(strings.NewReader(ivData.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestIVAnalysisBundleError validates error flows back from the parser
func TestIVAnalysisBundleError(t *testing.T) {
cd := mockCheckDetail()
cdAddendumA := mockCheckDetailAddendumA()
cd.AddCheckDetailAddendumA(cdAddendumA)
cdAddendumB := mockCheckDetailAddendumB()
cd.AddCheckDetailAddendumB(cdAddendumB)
ivDetail := mockImageViewDetail()
cd.AddImageViewDetail(ivDetail)
ivData := mockImageViewData()
cd.AddImageViewData(ivData)
ivAnalysis := mockImageViewAnalysis()
cd.AddImageViewAnalysis(ivAnalysis)
r := NewReader(strings.NewReader(ivAnalysis.String()))
clh := mockCashLetterHeader()
r.addCurrentCashLetter(NewCashLetter(clh))
bh := mockBundleHeader()
b := NewBundle(bh)
r.currentCashLetter.AddBundle(b)
r.addCurrentBundle(b)
_, err := r.Read()
if p, ok := err.(*ParseError); ok {
if e, ok := p.Err.(*FileError); ok {
if e.Msg != msgFileBundleOutside {
t.Errorf("%T: %s", e, e)
}
}
} else {
t.Errorf("%T: %s", err, err)
}
}
// TestICLCreditItemFile validates reading an ICL file with a CreditItem
func TestICLCreditItemFile(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", "BNK20181010121042882-A.icl"))
if err != nil {
t.Fatalf("Can not open local file: %s: \n", err)
}
defer fd.Close()
ICLFile, err := NewReader(fd, ReadVariableLineLengthOption()).Read()
if err != nil {
t.Errorf("Issue reading file: %+v \n", err)
}
// ensure we have a validated file structure
if err = ICLFile.Validate(); err != nil {
t.Errorf("Could not validate entire read file: %v", err)
}
}
// TestICLCreditRecord61File validates reading an ICL file with a Credit record (type 61)
func TestICLCreditRecord61File(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", "creditRecord61.icl"))
if err != nil {
t.Fatalf("Can not open local file: %s: \n", err)
}
defer fd.Close()
ICLFile, err := NewReader(fd, ReadVariableLineLengthOption()).Read()
if err != nil {
t.Errorf("Issue reading file: %+v \n", err)
}
// ensure we have a validated file structure
if err = ICLFile.Validate(); err != nil {
t.Errorf("Could not validate entire read file: %v", err)
}
if len(ICLFile.CashLetters) != 2 {
t.Errorf("File was missing CashLetters")
}
if len(ICLFile.CashLetters[0].Credits) != 1 {
t.Errorf("File was missing Credit record 61")
}
}
func TestICLBase64ImageData(t *testing.T) {
bs, err := os.ReadFile(filepath.Join("test", "testdata", "base64-encoded-images.json"))
if err != nil {
t.Fatal(err)
}
file, err := FileFromJSON(bs)
if err != nil {
t.Fatal(err)
}
var buf bytes.Buffer
if err := NewWriter(&buf).Write(file); err != nil {
t.Fatal(err)
}
if !bytes.Contains(buf.Bytes(), []byte("hello, world")) {
t.Fatalf("unexpected ICL file:\n%s", buf.String())
}
}
// TestICLFile_LargeCheckImage validates that reading a file with a large
// check detail record fails by default with bufio.ErrTooLong, and succeeds
// if a sufficiently-large buffer is created via BufferSizeOption.
//
// It creates this file on the fly to avoid bloating the repository.
func TestICLFile_LargeCheckImage(t *testing.T) {
fd, err := os.Open(filepath.Join("test", "testdata", "BNK20180905121042882-A.icl"))
if err != nil {
t.Fatalf("Can not open local file: %s: \n", err)
}
defer fd.Close()
r := NewReader(fd, ReadVariableLineLengthOption())
ICLFile, err := r.Read()
if err != nil {
t.Errorf("Issue reading file: %+v \n", err)
}
t.Logf("r.File.Header=%#v", r.File.Header)
t.Logf("r.File.Control=%#v", r.File.Control)
// ensure we have a validated file structure
if ICLFile.Validate(); err != nil {
t.Errorf("Could not validate entire read file: %v", err)
}
data := make([]byte, 128*1024)
if _, err = rand.Read(data); err != nil {
t.Errorf("Failed to read random data: %v", err)
}
ICLFile.CashLetters[0].Bundles[0].Checks[0].ImageViewData[0].LengthImageData = strconv.Itoa(len(data))
ICLFile.CashLetters[0].Bundles[0].Checks[0].ImageViewData[0].ImageData = data
var buf bytes.Buffer
w := NewWriter(&buf, WriteVariableLineLengthOption())
if err := w.Write(&ICLFile); err != nil {
t.Errorf("Failed to write file: %v", err)
}
fileReader := bytes.NewReader(buf.Bytes())
r = NewReader(fileReader, ReadVariableLineLengthOption())
_, err = r.Read()
if err == nil {
t.Error("Expected read of file with large check image to fail")
}
var ok bool
var p *ParseError
var e *FileError
if p, ok = err.(*ParseError); ok {
if e, ok = p.Err.(*FileError); ok {
if e.Msg != bufio.ErrTooLong.Error() {
t.Fatalf("Received unexpected error %s, expected %s",
e.Msg, bufio.ErrTooLong.Error())
}
}
}
if !ok {
t.Errorf("Received unexpected error type %T: %v", err, err)
}
fileReader.Reset(buf.Bytes())
r = NewReader(fileReader, ReadVariableLineLengthOption(), BufferSizeOption(256*1024))
_, err = r.Read()
if err != nil {
t.Errorf("Unexpected error while reading file: %v", err)
}
}
| 7c227a3e2f4b653b5d277e530335319df3aea7f5 | [
"Go"
] | 2 | Go | adamdecaf/imagecashletter | 85b706f7edf5beb4218ce6390f88911ee63aa06b | 38f719fff4edc08b75c1e61ad15e72cecf4ccd42 |
refs/heads/master | <repo_name>pmoulos/metaseqr<file_sep>/man/get.bs.organism.Rd
\name{get.bs.organism}
\alias{get.bs.organism}
\title{Return a proper formatted BSgenome organism name}
\usage{
get.bs.organism(org)
}
\arguments{
\item{org}{one of metaseqR supported organisms.}
}
\value{
A proper BSgenome package name.
}
\description{
Returns a properly formatted BSgenome package name
according to metaseqR's supported organism. Internal
use.
}
\examples{
\donttest{
bs.name <- get.bs.organism("hg18")
}
}
\author{
<NAME>
}
<file_sep>/man/filter.high.Rd
\name{filter.high}
\alias{filter.high}
\title{Filtering helper}
\usage{
filter.high(x, f)
}
\arguments{
\item{x}{a data numeric matrix.}
\item{f}{a threshold.}
}
\description{
High score filtering function. Internal use.
}
\examples{
data("mm9.gene.data",package="metaseqR")
counts <- as.matrix(mm9.gene.counts[,9:12])
f <- filter.low(counts,median(counts))
}
\author{
<NAME>
}
<file_sep>/man/diagplot.de.heatmap.Rd
\name{diagplot.de.heatmap}
\alias{diagplot.de.heatmap}
\title{Diagnostic heatmap of differentially expressed genes}
\usage{
diagplot.de.heatmap(x, con = NULL, output = "x11",
path = NULL, ...)
}
\arguments{
\item{x}{the data matrix to create a heatmap for.}
\item{con}{an optional string depicting a name (e.g. the
contrast name) to appear in the title of the volcano
plot.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"}, \code{"ps"}.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filenames of the plots produced in a named list with
names the \code{which.plot} argument. If
\code{output="x11"}, no output filenames are produced.
}
\description{
This function plots a heatmap of the differentially
expressed genes produced by the metaseqr workflow, useful
for quality control, e.g. whether samples belonging to
the same group cluster together.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
M <- normalize.edger(data.matrix,sample.list)
p <- stat.edger(M,sample.list,contrast)
diagplot.de.heatmap(data.matrix[p[[1]]<0.05,])
}
\author{
<NAME>
}
<file_sep>/man/combine.bonferroni.Rd
\name{combine.bonferroni}
\alias{combine.bonferroni}
\title{Combine p-values with Bonferroni's method}
\usage{
combine.bonferroni(p)
}
\arguments{
\item{p}{a p-value matrix (rows are genes,
columns are statistical tests).}
}
\value{
A vector of combined p-values.
}
\description{
This function combines p-values from the
various statistical tests supported by
metaseqR using the Bonferroni's method (see
reference in the main \code{\link{metaseqr}}
help page or in the vignette).
}
\examples{
p <- matrix(runif(300),100,3)
pc <- combine.bonferroni(p)
}
\author{
<NAME>
}
<file_sep>/man/as.class.vector.Rd
\name{as.class.vector}
\alias{as.class.vector}
\title{Create a class vector}
\usage{
as.class.vector(sample.list)
}
\arguments{
\item{sample.list}{the list containing condition names
and the samples under each condition.}
}
\value{
A vector of condition names.
}
\description{
Creates a class vector from a sample list. Internal to
the \code{stat.*} functions. Mostly internal use.
}
\examples{
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
clv <- as.class.vector(sample.list)
}
\author{
<NAME>
}
<file_sep>/man/wapply.Rd
\name{wapply}
\alias{wapply}
\title{List apply helper}
\usage{
wapply(m, ...)
}
\arguments{
\item{m}{a logical indicating whether to execute in
parallel or not.}
\item{...}{the rest arguments to \code{\link{lapply}} (or
\code{mclapply})}
}
\description{
A wrapper around normal and parallel apply
(\code{\link{mclapply}} or multicore package) to avoid
excessive coding for control of single or parallel code
execution. Internal use.
}
\examples{
\donttest{
multic <- check.parallel(0.8)
# Test meaningful only in machines where parallel computation
# supported
if (multic) {
system.time(r<-wapply(TRUE,1:10,function(x) runif(1e+6)))
system.time(r<-wapply(FALSE,1:10,function(x) runif(1e+6)))
}
}
}
\author{
<NAME>
}
<file_sep>/man/diagplot.venn.Rd
\name{diagplot.venn}
\alias{diagplot.venn}
\title{Venn diagrams when performing meta-analysis}
\usage{
diagplot.venn(pmat, fcmat = NULL, pcut = 0.05,
fcut = 0.5, direction = c("dereg", "up", "down"),
nam = as.character(round(1000 * runif(1))),
output = "x11", path = NULL, alt.names = NULL, ...)
}
\arguments{
\item{pmat}{a matrix with p-values corresponding to the
application of each statistical algorithm. The p-value
matrix must have the colnames attribute and the colnames
should correspond to the name of the algorithm used to
fill the specific column (e.g. if
\code{"statistics"=c("deseq","edger","nbpseq")} then
\code{colnames(pmat) <-}
\code{c("deseq","edger","nbpseq")}.}
\item{fcmat}{an optional matrix with fold changes
corresponding to the application of each statistical
algorithm. The fold change matrix must have the colnames
attribute and the colnames should correspond to the name
of the algorithm used to fill the specific column (see
the parameter \code{pmat}).}
\item{pcut}{if \code{fcmat} is supplied, an absolute
fold change cutoff to be applied to \code{fcmat} to
determine the differentially expressed genes for each
algorithm.}
\item{fcut}{a p-value cutoff for statistical
significance. Defaults to \code{0.05}.}
\item{direction}{if \code{fcmat} is supplied, a keyword
to denote which genes to draw in the Venn diagrams with
respect to their direction of regulation. It can be one
of \code{"dereg"} for the total of regulated genes, where
\code{abs(fcmat[,n])>=fcut} (default), \code{"up"} for
the up-regulated genes where \code{fcmat[,n]>=fcut} or
\code{"down"} for the up-regulated genes where
\code{fcmat[,n]<=-fcut}.}
\item{nam}{a name to be appended to the output graphics
file (if \code{"output"} is not \code{"x11"}).}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files. If
\code{"path"} is not \code{NULL}, a file with the
intersections in the Venn diagrams will be produced
and written in \code{"path"}.}
\item{alt.names}{an optional named vector of names, e.g.
HUGO gene symbols, alternative or complementary to the
unique gene names which are the rownames of \code{pmat}.
The names of the vector must be the rownames of
\code{pmat}.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filenames of the plots produced in a named list with
names the \code{which.plot} argument. If
output=\code{"x11"}, no output filenames are produced.
}
\description{
This function uses the R package VennDiagram and plots an
up to 5-way Venn diagram depicting the common and
specific to each statistical algorithm genes, for each
contrast. Mostly for internal use because of its main
argument which is difficult to construct, but can be used
independently if the user grasps the logic.
}
\examples{
require(VennDiagram)
p1 <- 0.01*matrix(runif(300),100,3)
p2 <- matrix(runif(300),100,3)
p <- rbind(p1,p2)
rownames(p) <- paste("gene",1:200,sep="_")
colnames(p) <- paste("method",1:3,sep="_")
venn.contents <- diagplot.venn(p)
}
\author{
<NAME>
}
<file_sep>/man/filter.exons.Rd
\name{filter.exons}
\alias{filter.exons}
\title{Filter gene expression based on exon counts}
\usage{
filter.exons(the.counts, gene.data, sample.list,
exon.filters, restrict.cores = 0.8)
}
\arguments{
\item{the.counts}{a named list created with the
\code{\link{construct.gene.model}} function. See its help
page for details.}
\item{gene.data}{an annotation data frame usually
obtained with \code{\link{get.annotation}} containing the
unique gene accession identifiers.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{exon.filters}{a named list with exon filters and
their parameters. See the main help page of
\code{\link{metaseqr}} for details.}
\item{restrict.cores}{in case of parallel execution of
several subfunctions, the fraction of the available cores
to use. In some cases if all available cores are used
(\code{restrict.cores=1} and the system does not have
sufficient RAM, the running machine might significantly
slow down.}
}
\value{
a named list with two members. The first member
(\code{result} is a named list whose names are the
exon filter names and its members are the filtered
rownames of \code{gene.data}. The second member is a
matrix of binary flags (0 for non-filtered, 1 for
filtered) for each gene. The rownames of the flag
matrix correspond to gene ids.
}
\description{
This function performs the gene expression filtering
based on exon read counts and a set of exon filter rules.
For more details see the main help pages of
\code{\link{metaseqr}}.
}
\examples{
\donttest{
data("hg19.exon.data",package="metaseqR")
exon.counts <- hg19.exon.counts
gene.data <- get.annotation("hg19","gene")
sample.list <- sample.list.hg19
exon.filters <- get.defaults("exon.filter")
the.counts <- construct.gene.model(exon.counts,sample.list,
gene.data)
filter.results <- filter.exons(the.counts,gene.data,
sample.list,exon.filters)
}
}
\author{
<NAME>
}
<file_sep>/man/get.exon.attributes.Rd
\name{get.exon.attributes}
\alias{get.exon.attributes}
\title{Annotation downloader helper}
\usage{
get.exon.attributes(org)
}
\arguments{
\item{org}{one of the supported organisms.}
}
\value{
A character vector of Ensembl exon attributes.
}
\description{
Returns a vector of genomic annotation attributes which
are used by the biomaRt package in order to fetch the
exon annotation for each organism. It has no parameters.
Internal use.
}
\examples{
exon.attr <- get.exon.attributes("mm9")
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.export.R
#' Results export builder
#'
#' This function help build the output files of the metaseqr pipeline based on
#' several elements produced during the pipeline execution. It is intended for
#' internal use and not available to the users.
#'
#' @param gene.data an annotation data frame (such the ones produced by
#' \code{\link{get.annotation}}).
#' @param raw.gene.counts a matrix of un-normalized gene counts.
#' @param norm.gene.counts a matrix of normalized gene counts.
#' @param flags a matrix of filtering flags (0,1), created by the filtering
#' functions.
#' @param sample.list see the documentation of \code{\link{metaseqr}}.
#' @param cnt the statistical contrast for which the export builder is currently
#' running.
#' @param statistics the statistical tests used (see the documentation of
#' \code{\link{metaseqr}}).
#' @param raw.list a list of transformed un-normalized counts, see the documentation
#' of \code{\link{make.transformation}}.
#' @param norm.list a list of transformed normalized counts, see the documentation
#' of \code{\link{make.transformation}}.
#' @param p.mat a matrix of p-values, see the documentation of \code{\link{metaseqr}}.
#' @param adj.p.mat a matrix of adjusted p-values, see the documentation of
#' \code{\link{metaseqr}}.
#' @param sum.p a vector of combined p-values, see the documentation of
#' \code{\link{metaseqr}}.
#' @param adj.sum.p a vector of adjusted combined p-values, see the documentation
#' of \code{\link{metaseqr}}.
#' @param export.what see the documentation of \code{\link{metaseqr}}.
#' @param export.scale see the documentation of \code{\link{metaseqr}}.
#' @param export.values see the documentation of \code{\link{metaseqr}}.
#' @param export.stats see the documentation of \code{\link{metaseqr}}.
#' @param log.offset see the documentation of \code{\link{metaseqr}}.
#' @param report see the documentation of \code{\link{metaseqr}}.
#' @return A list with three members: a data frame to be exported in a text file,
#' a long string with the result in a html formatted table (if \code{report=TRUE})
#' and the column names of the output data frame.
#' @author <NAME>
#' @examples
#' \dontrun{
#' # Not yet available
#'}
build.export <- function(gene.data,raw.gene.counts,norm.gene.counts,flags,
sample.list,cnt,statistics,
raw.list,norm.list,
p.mat=matrix(NA,nrow(gene.data),length(statistics)),
adj.p.mat=matrix(NA,nrow(gene.data),length(statistics)),
sum.p=rep(NA,nrow(gene.data)),
adj.sum.p=rep(NA,nrow(gene.data)),
export.what=c("annotation","p.value","adj.p.value","meta.p.value",
"adj.meta.p.value","fold.change","stats","counts","flags"),
export.scale=c("natural","log2","log10","rpgm","vst"),
export.values=c("raw","normalized"),
export.stats=c("mean","median","sd","mad","cv","rcv"),
log.offset=1,report=TRUE
) {
if (is.null(colnames(p.mat)))
colnames(p.mat) <- statistics
if (is.null(adj.p.mat))
adj.p.mat=matrix(NA,nrow(gene.data),length(statistics))
if (is.null(colnames(adj.p.mat)))
colnames(adj.p.mat) <- statistics
export <- data.frame(row.names=rownames(gene.data))
if (report) export.html <- as.matrix(export)
the.names <- character(0)
if ("annotation" %in% export.what) {
disp(" binding annotation...")
export <- cbind(export,gene.data)
if (report)
export.html <- cbind(export.html,make.html.cells(gene.data,
type="text"))
the.names <- c(the.names,colnames(gene.data))
}
if ("p.value" %in% export.what) {
disp(" binding p-values...")
export <- cbind(export,p.mat)
if (report)
export.html <- cbind(export.html,make.html.cells(p.mat))
the.names <- c(the.names,paste("p-value_",colnames(p.mat),sep=""))
}
if ("adj.p.value" %in% export.what) {
disp(" binding FDRs...")
export <- cbind(export,adj.p.mat)
if (report)
export.html <- cbind(export.html,make.html.cells(adj.p.mat))
the.names <- c(the.names,paste("FDR_",colnames(adj.p.mat),sep=""))
}
if ("meta.p.value" %in% export.what && length(statistics)>1) {
# Otherwise, it does not exist
disp(" binding meta p-values...")
export <- cbind(export,sum.p)
if (report)
export.html <- cbind(export.html,make.html.cells(sum.p))
the.names <- c(the.names,paste("meta_p-value_",cnt,sep=""))
}
if ("adj.meta.p.value" %in% export.what && length(statistics)>1) {
disp(" binding adjusted meta p-values...")
export <- cbind(export,adj.sum.p)
if (report)
export.html <- cbind(export.html,make.html.cells(adj.sum.p))
the.names <- c(the.names,paste("meta_FDR_",cnt,sep=""))
}
if ("fold.change" %in% export.what) {
if ("normalized" %in% export.values) {
tmp <- make.fold.change(cnt,sample.list,norm.gene.counts,log.offset)
if ("natural" %in% export.scale) {
disp(" binding natural normalized fold changes...")
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,
paste("natural_normalized_fold_change_",colnames(tmp),
sep=""))
}
if ("log2" %in% export.scale) {
disp(" binding log2 normalized fold changes...")
export <- cbind(export,log2(tmp))
if (report)
export.html <- cbind(export.html,
make.html.cells(log2(tmp)))
the.names <- c(the.names,paste("log2_normalized_fold_change_",
colnames(tmp),sep=""))
}
}
if ("raw" %in% export.values) {
tmp <- make.fold.change(cnt,sample.list,raw.gene.counts,log.offset)
if ("natural" %in% export.scale) {
disp(" binding natural raw fold changes...")
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste("natural_raw_fold_change_",
colnames(tmp),sep=""))
}
if ("log2" %in% export.scale) {
disp(" binding log2 raw fold changes...")
export <- cbind(export,log2(tmp))
if (report)
export.html <- cbind(export.html,make.html.cells(log2(tmp)))
the.names <- c(the.names,paste("log2_raw_fold_change_",
colnames(tmp),sep=""))
}
}
}
if ("stats" %in% export.what) {
conds <- strsplit(cnt,"_vs_")[[1]]
for (cond in conds) {
if ("normalized" %in% export.values) {
if ("mean" %in% export.stats) {
disp(" binding normalized mean counts...")
tmp <- make.stat(sample.list[[cond]],norm.list,"mean",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_normalized_mean_counts_",cond,sep=""))
}
if ("median" %in% export.stats) {
disp(" binding normalized median counts...")
tmp <- make.stat(sample.list[[cond]],norm.list,"median",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_normalized_median_counts_",cond,sep=""))
}
if ("sd" %in% export.stats) {
disp(" binding normalized count sds...")
tmp <- make.stat(sample.list[[cond]],norm.list,"sd",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_normalized_sd_counts_",cond,sep=""))
}
if ("mad" %in% export.stats) {
disp(" binding normalized count MADs...")
tmp <- make.stat(sample.list[[cond]],norm.list,"mad",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_normalized_mad_counts_",cond,sep=""))
}
if ("cv" %in% export.stats) {
disp(" binding normalized count CVs...")
tmp <- make.stat(sample.list[[cond]],norm.list,"cv",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_normalized_cv_counts_",cond,sep=""))
}
if ("rcv" %in% export.stats) {
disp(" binding normalized counts RCVs...")
tmp <- make.stat(sample.list[[cond]],norm.list,"rcv",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_normalized_rcv_counts_",cond,sep=""))
}
}
if ("raw" %in% export.values) {
if ("mean" %in% export.stats) {
disp(" binding raw mean counts...")
tmp <- make.stat(sample.list[[cond]],raw.list,"mean",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_raw_mean_counts_",cond,sep=""))
}
if ("median" %in% export.stats) {
disp(" binding raw median counts...")
tmp <- make.stat(sample.list[[cond]],raw.list,"median",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_raw_median_counts_",cond,sep=""))
}
if ("sd" %in% export.stats) {
disp(" binding raw counts sds...")
tmp <- make.stat(sample.list[[cond]],raw.list,"sd",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_raw_sd_counts_",cond,sep=""))
}
if ("mad" %in% export.stats) {
disp(" binding raw counts MADs...")
tmp <- make.stat(sample.list[[cond]],raw.list,"mad",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_raw_mad_counts_",cond,sep=""))
}
if ("cv" %in% export.stats) {
disp(" binding raw counts CVs...")
tmp <- make.stat(sample.list[[cond]],raw.list,"cv",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_raw_cv_counts_",cond,sep=""))
}
if ("rcv" %in% export.stats) {
disp(" binding raw counts RCVs...")
tmp <- make.stat(sample.list[[cond]],raw.list,"rcv",
export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
the.names <- c(the.names,paste(colnames(tmp),
"_raw_rcv_counts_",cond,sep=""))
}
}
}
}
if ("counts" %in% export.what) {
conds <- strsplit(cnt,"_vs_")[[1]]
for (cond in conds) {
if ("normalized" %in% export.values) {
disp(" binding all normalized counts for ",cond,"...")
tmp <- make.matrix(sample.list[[cond]],norm.list,export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
part.1 <- rep(paste(export.scale,"_normalized_counts_",sep=""),
each=length(sample.list[[cond]]))
part.2 <- paste(part.1,colnames(tmp),sep="")
the.names <- c(the.names,part.2)
}
if ("raw" %in% export.values) {
disp(" binding all raw counts for ",cond,"...")
tmp <- make.matrix(sample.list[[cond]],raw.list,export.scale)
export <- cbind(export,tmp)
if (report)
export.html <- cbind(export.html,make.html.cells(tmp))
part.1 <- rep(paste(export.scale,"_raw_counts_",sep=""),
each=length(sample.list[[cond]]))
part.2 <- paste(part.1,colnames(tmp),sep="")
the.names <- c(the.names,part.2)
}
}
}
if ("flags" %in% export.what && !is.null(flags)) {
disp(" binding filtering flags...")
export <- cbind(export,as.data.frame(flags))
if (report)
export.html <- cbind(export.html,make.html.cells(flags))
the.names <- c(the.names,colnames(flags))
}
names(export) <- the.names
if (!report)
export.html <- NULL
return (list(text.table=export,html.table=export.html,headers=the.names))
}
<file_sep>/man/diagplot.avg.ftd.Rd
\name{diagplot.avg.ftd}
\alias{diagplot.avg.ftd}
\title{Create average False (or True) Discovery
curves}
\usage{
diagplot.avg.ftd(ftdr.obj, output = "x11",
path = NULL, draw = TRUE, ...)
}
\arguments{
\item{ftdr.obj}{a list with outputs from
\code{\link{diagplot.ftd}}.}
\item{output}{one or more R plotting
device to direct the plot result to.
Supported mechanisms: \code{"x11"} (default),
\code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{draw}{boolean to determine whether
to plot the curves or just return the
calculated values (in cases where the user
wants the output for later averaging
for example). Defaults to \code{TRUE} (make
plots).}
\item{...}{further arguments to be passed to
plot devices, such as parameter from
\code{\link{par}}.}
}
\value{
A named list with two members: the first member
(\code{avg.ftdr}) contains a list with the
means and the standard deviations of the averaged
\code{ftdr.obj} and are used to create the plot.
The second member (\code{path}) contains the
path to the created figure graphic.
}
\description{
This function creates false (or true) discovery
curves using a list containing several outputs
from \code{\link{diagplot.ftd}}.
}
\examples{
p11 <- 0.001*matrix(runif(300),100,3)
p12 <- matrix(runif(300),100,3)
p21 <- 0.001*matrix(runif(300),100,3)
p22 <- matrix(runif(300),100,3)
p31 <- 0.001*matrix(runif(300),100,3)
p32 <- matrix(runif(300),100,3)
p1 <- rbind(p11,p21)
p2 <- rbind(p12,p22)
p3 <- rbind(p31,p32)
rownames(p1) <- rownames(p2) <- rownames(p3) <-
paste("gene",1:200,sep="_")
colnames(p1) <- colnames(p2) <- colnames(p3) <-
paste("method",1:3,sep="_")
truth <- c(rep(1,40),rep(-1,40),rep(0,20),
rep(1,10),rep(2,10),rep(0,80))
names(truth) <- rownames(p1)
ftd.obj.1 <- diagplot.ftd(truth,p1,N=100,draw=FALSE)
ftd.obj.2 <- diagplot.ftd(truth,p2,N=100,draw=FALSE)
ftd.obj.3 <- diagplot.ftd(truth,p3,N=100,draw=FALSE)
ftd.obj <- list(ftd.obj.1,ftd.obj.2,ftd.obj.3)
avg.ftd.obj <- diagplot.avg.ftd(ftd.obj)
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.argcheck.R
#' Main argument validator
#'
#' Checks if the arguments passed to \code{\link{metaseqr}} are valid and throws
#' a warning about the invalid ones (which are ignored anyway because of the
#' \code{...} in \code{\link{metaseqr}}. However, for this reason this function
#' is useful as some important parameter faults might go unnoticed in the beginning
#' and cause a failure afterwards. Internal use.
#'
#' @param main.args a list of parameters with which metaseqr is called (essentially,
#' the output of \code{\link{match.call}}.
#' @author <NAME>
check.main.args <- function(main.args) {
in.args <- names(main.args)[-1] # 1st member name of calling function
valid.args <- c(
"counts","sample.list","exclude.list","file.type","path","contrast",
"libsize.list","id.col","gc.col","name.col","bt.col","annotation",
"gene.file","org","trans.level","count.type","utr.flank","exon.filters",
"gene.filters","when.apply.filter","normalization","norm.args",
"statistics","stat.args","adjust.method","meta.p","weight","nperm",
"reprod","pcut","log.offset","preset","qc.plots","fig.format",
"out.list","export.where","export.what","export.scale","export.values",
"export.stats","export.counts.table","restrict.cores","report","refdb",
"report.top","report.template","verbose","run.log","save.gene.model"
)
invalid <- setdiff(in.args,valid.args)
if (length(invalid) > 0) {
for (i in 1:length(invalid))
warnwrap("Unknown input argument to metaseqr pipeline: ",invalid[i],
" ...Ignoring...",now=TRUE)
}
}
#' Text argument validator
#'
#' Checks if one or more given textual argument(s) is/are member(s) of a list of
#' correct arguments. It's a more package-specific function similar to
#' \code{\link{match.arg}}. Mostly for internal use.
#'
#' @param arg.name the name of the argument that is checked (for display purposes).
#' @param arg.value the value(s) of the argument to be checked.
#' @param arg.list a vector of valid argument values for \code{arg.value} to be
#' matched against.
#' @param multiarg a logical scalar indicating whether \code{arg.name} accepts
#' multiple arguments or not. In that case, all of the values in \code{arg.value}
#' are checked against \code{arg.list}.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' check.text.args("count.type",count.type,c("gene","exon"),multiarg=FALSE)
#' check.text.args("statistics",statistics,c("deseq","edger","noiseq","bayseq",
#' "limma"), multiarg=TRUE)
#'}
check.text.args <- function(arg.name,arg.value,arg.list,multiarg=FALSE) {
if (multiarg) {
arg.value <- tolower(arg.value)
if (!all(arg.value %in% arg.list))
stopwrap("\"",arg.name,"\""," parameter must be one or more of ",
paste(paste("\"",arg.list,sep=""),collapse="\", "),"\"!")
}
else {
arg.save <- arg.value[1]
arg.value <- tolower(arg.value[1])
# An exception must be added for annotation because it can be an external
# file too
if (arg.name=="annotation") {
if (!(arg.value %in% arg.list) && !file.exists(arg.save))
stopwrap("\"",arg.name,"\""," parameter must be one of ",
paste(paste("\"",arg.list,sep=""),collapse="\", "),
"\" or an existing file!")
}
else {
if (!(arg.value %in% arg.list))
stopwrap("\"",arg.name,"\""," parameter must be one of ",
paste(paste("\"",arg.list,sep=""),collapse="\", "),"\"!")
}
}
}
#' Numeric argument validator
#'
#' Checks if one or more given numeric argument(s) satisfy several rules concerning
#' numeric arguments, e.g. proper bounds or proper format (e.g. it must be a number
#' and not a character). Mostly for internal use.
#'
#' @param arg.name the name of the argument that is checked (for display purposes).
#' @param arg.value the value(s) of the argument to be checked.
#' @param arg.type either the string \code{"numeric"} to denote generic double-like
#' R numerics or \code{"integer"} for integer values.
#' @param arg.bounds a numeric or a vector with 2 elements, restraining
#' \code{arg.value} to be within the bounds defined by the input vector or e.g.
#' larger (smaller) than the numeric value. See examples.
#' @param direction a string denoting to which direction the \code{arg.value}
#' should be compared with \code{arg.bounds}. For example, \code{"both"} should
#' be given with a two element vector against which, \code{arg.value} will be
#' checked to see whether it is smaller than the low boundary or larger than the
#' higher boundary. In that case, the function will throw an error. The direction
#' parameter can be one of: \code{"both"} (described above), \code{"botheq"} (as
#' above, but the \code{arg.val} is also checked for equality -closed intervals),
#' \code{"gt"} or \code{"gte"} (check whether \code{arg.val} is smaller or smaller
#' than or equal to the first value of \code{arg.bounds}), \code{"lt"} or \code{"lte"}
#' (check whether \code{arg.val} is larger or larger than or equal to the first
#' value of \code{arg.bounds}).
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' pcut <- 1.2 # A probability cannot be larger than 1! It will throw an error!
#' check.num.args("pcut",pcut,"numeric",c(0,1),"botheq")
#' pcut <- 0.05 # Pass
#' check.num.args("pcut",pcut,"numeric",c(0,1),"botheq")
#' gc.col <- 3.4 # A column in a file cannot be real! It will throw an error!
#' check.num.args("gc.col",gc.col,"integer",0,"gt")
#' gc.col <- 5 # Pass
#' check.num.args("gc.col",gc.col,"integer",0,"gt")
#'}
check.num.args <- function(arg.name,arg.value,arg.type,arg.bounds,direction) {
switch(arg.type,
numeric = {
if (!is.numeric(arg.value))
stopwrap("\"",arg.name,"\"",
" parameter must be a numeric value!")
if (!missing(arg.bounds)) {
switch(direction,
both = {
if (arg.value<=arg.bounds[1] ||
arg.value>=arg.bounds[2])
stopwrap("\"",arg.name,"\""," parameter must be a ",
"numeric ","value larger than or equal to ",
arg.bounds[1]," and smaller than or equal to ",
arg.bounds[2],"!")
},
botheq = {
if (arg.value<arg.bounds[1] || arg.value>arg.bounds[2])
stopwrap("\"",arg.name,"\""," parameter must be a ",
"numeric value larger than ",arg.bounds[1],
" and smaller than ",arg.bounds[2],"!")
},
gt = {
if (arg.value<=arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be a ",
"numeric value greater than ",arg.bounds[1],"!")
},
lt = {
if (arg.value>=arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be a ",
"numeric value lower than ",arg.bounds[1],"!")
},
gte = {
if (arg.value<arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be a ",
"numeric value greater than or equal to ",
arg.bounds[1],"!")
},
lte = {
if (arg.value>arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be a ",
"numeric value lower than or equal to ",
arg.bounds[1],"!")
}
)
}
},
integer = {
if (!is.integer(arg.value))
stopwrap("\"",arg.name,"\""," parameter must be an integer!")
if (!missing(arg.bounds)) {
switch(direction,
both = {
if (arg.value<=arg.bounds[1] ||
arg.value>=arg.bounds[2])
stopwrap("\"",arg.name,"\""," parameter must be ",
"an integer larger than or equal to ",
arg.bounds[1]," and smaller than or equal to ",
arg.bounds[2],"!")
},
botheq = {
if (arg.value<arg.bounds[1] || arg.value>arg.bounds[2])
stopwrap("\"",arg.name,"\""," parameter must be ",
"an integer larger than or equal to ",
arg.bounds[1]," and smaller than or equal to ",
arg.bounds[2],"!")
},
gt = {
if (arg.value<=arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be ",
"an integer greater than ",arg.bounds[1],"!")
},
lt = {
if (arg.value>=arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be ",
"an integer lower than ",arg.bounds[1],"!")
},
gte = {
if (arg.value<arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be ",
"an integer greater than or equal to ",
arg.bounds[1],"!")
},
lte = {
if (arg.value>arg.bounds[1])
stopwrap("\"",arg.name,"\""," parameter must be ",
"an integer lower than or equal to ",
arg.bounds[1],"!")
}
)
}
}
)
}
#' File argument validator
#'
#' Checks if a file exists for specific arguments requiring a file input. Internal
#' use only.
#'
#' @param arg.name argument name to display in a possible error.
#' @param arg.value the filename to check.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' # OK
#' check.file.args("file",system.file("metaseqr_report.html",package="metaseqR"))
#' # Error!
#' check.file.args("file",system.file("metaseqr_report.htm",package="metaseqR"))
#'}
check.file.args <- function(arg.name,arg.value) {
if (!file.exists(arg.value))
stopwrap("\"",arg.name,"\""," parameter must be an existing file!")
}
#' Parallel run validator
#'
#' Checks existence of multiple cores and loads parallel package.
#'
#' @param rc fraction of available cores to use.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
# multic <- check.parallel(0.8)
#'}
check.parallel <- function(rc) {
if (suppressWarnings(!require(parallel)) || .Platform$OS.type!="unix")
multi <- FALSE
else {
multi <- TRUE
ncores <- parallel::detectCores()
if (!missing(rc) || !is.na(rc) || !is.null(rc))
ncores <- ceiling(rc*ncores)
options(cores=ncores)
}
return(multi)
}
#' Contrast validator
#'
#' Checks if the contrast vector follows the specified format. Internal use only.
#'
#' @param cnt contrasts vector.
#' @param sample.list the input sample list.
#' @author <NAME>
#' @export
#' @examples
#' dontrun{
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' cnt <- c("A_vs_B") # Will work
#' #cnt <- c("A_vs_C") ## Will throw error!
#' check.contrast.format(cnt,sample.list)
#}
check.contrast.format <- function(cnt,sample.list) {
# This function will break cnt and check that all contrast counter parts are
# members of the names of the sample.list and contain the string "_vs_" as
# many times as the names of the sample.list minus 1. If satisfied return
# TRUE else error.
cnts <- strsplit(cnt,"_vs_")
#if (length(unique(unlist(cnts))) != length(names(sample.list)))
if (!all(unique(unlist(cnts)) %in% names(sample.list)))
stopwrap("Condition names in sample list and contrast list do not ",
"match! Check if the contrasts follow the appropriate format (e.g.",
" \"_vs_\" separating contrasting conditions...")
if (length(unique(cnt))!=length(cnt))
warnwrap("Duplicates found in the contrasts list! Duplicates will be ",
"ignored...")
}
#' Library size validator
#'
#' Checks the names of the supplied library sizes. Internal use only.
#'
#' @param libsize.list the samples-names library size list.
#' @param sample.list the input sample list.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' libsize.list.1 <- list(A1=1e+6,A2=1.1e+6,B1=1.2e+6,B2=1.3e+6,B3=1.5e+6)
#' libsize.list.2 <- list(A1=1e+6,A2=1.1e+6,B1=1.2e+6,B2=1.3e+6)
#' check.libsize(libsize.list.1,sample.list) # Will work
#' #check.libsize(libsize.list.2,sample.list) # Will throw error!
#'}
check.libsize <- function(libsize.list,sample.list) {
if (!is.null(libsize.list)) {
if (length(intersect(names(libsize.list),unlist(sample.list,
use.names=FALSE)))!=length(unlist(sample.list,
use.names=FALSE))) {
warnwrap("Sample names in \"libsize.list\" and \"sample.list\" do ",
"not match! Library sizes will be estimated from count data...")
return(NULL)
}
else return(libsize.list)
}
else
return(NULL)
}
#' Required packages validator
#'
#' Checks if all the required packages are present according to metaseqr input
#' options. Internal use only.
#'
#' @param m meta-analysis method
#' @param p qc plot types
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' check.packages(c("simes","whitlock"),c("gcbias","correl"))
#}
check.packages <- function(m,p) {
# Check meta-analysis packages
if ("whitlock" %in% m && !require(survcomp))
stopwrap("Bioconductor package survcomp is required for \"whitlock\" ",
"p-value meta analysis!")
if ("venn" %in% p && !require(VennDiagram))
stopwrap("R package VennDiagram is required for some of the selected ",
"QC plots!")
}
<file_sep>/R/metaseqr.norm.R
#' Normalization based on the EDASeq package
#'
#' This function is a wrapper over EDASeq normalization. It accepts a matrix of
#' gene counts (e.g. produced by importing an externally generated table of counts
#' to the main metaseqr pipeline).
#'
#' @param gene.counts a table where each row represents a gene and each column a
#' sample. Each cell contains the read counts for each gene and sample. Such a
#' table can be produced outside metaseqr and is imported during the basic metaseqr
#' workflow.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param norm.args a list of EDASeq normalization parameters. See the result of
#' \code{get.defaults("normalization",} \code{"edaseq")} for an example and how
#' you can modify it.
#' @param gene.data an optional annotation data frame (such the ones produced by
#' \code{get.annotation}) which contains the GC content for each gene and from
#' which the gene lengths can be inferred by chromosome coordinates.
#' @param output the class of the output object. It can be \code{"matrix"} (default)
#' for versatility with other tools or \code{"native"} for the EDASeq native S4
#' object (SeqExpressionSet). In the latter case it should be handled with suitable
#' EDASeq methods.
#' @return A matrix or a SeqExpressionSet with normalized counts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.boxplot(data.matrix,sample.list)
#'
#' lengths <- round(1000*runif(nrow(data.matrix)))
#' starts <- round(1000*runif(nrow(data.matrix)))
#' ends <- starts + lengths
#' gc <- runif(nrow(data.matrix))
#' gene.data <- data.frame(
#' chromosome=c(rep("chr1",nrow(data.matrix)/2),rep("chr2",nrow(data.matrix)/2)),
#' start=starts,end=ends,gene_id=rownames(data.matrix),gc_content=gc
#' )
#' norm.data.matrix <- normalize.edaseq(data.matrix,sample.list,gene.data=gene.data)
#' diagplot.boxplot(norm.data.matrix,sample.list)
#'}
normalize.edaseq <- function(gene.counts,sample.list,norm.args=NULL,
gene.data=NULL,output=c("matrix","native")) {
if (is.null(norm.args))
norm.args <- get.defaults("normalization","edaseq")
if (!is.matrix(gene.counts))
gene.counts <- as.matrix(gene.counts)
if (!is.null(gene.data) && is.null(attr(gene.data,"gene.length")))
attr(gene.data,"gene.length") <- rep(1,nrow(gene.counts))
output <- tolower(output[1])
check.text.args("output",output,c("matrix","native"))
classes <- as.class.vector(sample.list)
if (is.null(gene.data)) {
seq.genes <- newSeqExpressionSet(
gene.counts,
phenoData=AnnotatedDataFrame(
data.frame(
conditions=classes,
row.names=colnames(gene.counts)
)
),
featureData=AnnotatedDataFrame(
data.frame(
length=rep(1,nrow(gene.counts)),
row.names=rownames(gene.counts)
)
)
)
seq.genes <- betweenLaneNormalization(withinLaneNormalization(seq.genes,
"length",which=norm.args$within.which),
which=norm.args$between.which)
}
else {
seq.genes <- newSeqExpressionSet(
gene.counts,
phenoData=AnnotatedDataFrame(
data.frame(
conditions=classes,
row.names=colnames(gene.counts)
)
),
featureData=AnnotatedDataFrame(
data.frame(
gc=gene.data$gc_content,
length=attr(gene.data,"gene.length"),
row.names=rownames(gene.data)
)
)
)
seq.genes <- betweenLaneNormalization(withinLaneNormalization(seq.genes,
"gc",which=norm.args$within.which),which=norm.args$between.which)
}
if (output=="matrix")
return(exprs(seq.genes)) # Class: matrix
else if (output=="native")
return(seq.genes) # Class: SeqExpressionSet
}
#' Normalization based on the DESeq package
#'
#' This function is a wrapper over DESeq normalization. It accepts a matrix of
#' gene counts (e.g. produced by importing an externally generated table of counts
#' to the main metaseqr pipeline).
#'
#' @param gene.counts a table where each row represents a gene and each column a
#' sample. Each cell contains the read counts for each gene and sample. Such a
#' table can be produced outside metaseqr and is imported during the basic metaseqr
#' workflow.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param norm.args a list of DESeq normalization parameters. See the result of
#' \code{get.defaults("normalization",} \code{"deseq")} for an example and how you
#' can modify it.
#' @param output the class of the output object. It can be \code{"matrix"} (default)
#' for versatility with other tools or \code{"native"} for the DESeq native S4
#' object (CountDataSet). In the latter case it should be handled with suitable
#' DESeq methods.
#' @return A matrix or a CountDataSet with normalized counts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.boxplot(data.matrix,sample.list)
#'
#' norm.data.matrix <- normalize.deseq(data.matrix,sample.list)
#' diagplot.boxplot(norm.data.matrix,sample.list)
#'}
normalize.deseq <- function(gene.counts,sample.list,norm.args=NULL,
output=c("matrix","native")) {
if (is.null(norm.args))
norm.args <- get.defaults("normalization","deseq")
output <- tolower(output[1])
check.text.args("output",output,c("matrix","native"))
classes <- as.class.vector(sample.list)
cds <- newCountDataSet(gene.counts,classes)
cds <- estimateSizeFactors(cds,locfunc=norm.args$locfunc)
if (output=="native")
return(cds) # Class: CountDataSet
else if (output=="matrix")
return(round(counts(cds,normalized=TRUE))) # Class: matrix
}
#' Normalization based on the edgeR package
#'
#' This function is a wrapper over edgeR normalization. It accepts a matrix of
#' gene counts (e.g. produced by importing an externally generated table of counts
#' to the main metaseqr pipeline).
#'
#' @param gene.counts a table where each row represents a gene and each column a
#' sample. Each cell contains the read counts for each gene and sample. Such a
#' table can be produced outside metaseqr and is imported during the basic metaseqr
#' workflow.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param norm.args a list of edgeR normalization parameters. See the result of
#' \code{get.defaults("normalization",} \code{"edger")} for an example and how
#' you can modify it.
#' @param output the class of the output object. It can be \code{"matrix"} (default)
#' for versatility with other tools or \code{"native"} for the edgeR native S4
#' object (DGEList). In the latter case it should be handled with suitable edgeR
#' methods.
#' @return A matrix or a DGEList with normalized counts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.boxplot(data.matrix,sample.list)
#'
#' norm.data.matrix <- normalize.edger(data.matrix,sample.list)
#' diagplot.boxplot(norm.data.matrix,sample.list)
#'}
normalize.edger <- function(gene.counts,sample.list,norm.args=NULL,
output=c("matrix","native")) {
if (is.null(norm.args))
norm.args <- get.defaults("normalization","edger")
output <- tolower(output[1])
check.text.args("output",output,c("matrix","native"))
classes <- as.class.vector(sample.list)
dge <- DGEList(counts=gene.counts,group=classes)
dge <- calcNormFactors(dge,method=norm.args$method,
refColumn=norm.args$refColumn,logratioTrim=norm.args$logratioTrim,
sumTrim=norm.args$sumTrim,doWeighting=norm.args$doWeighting,
Acutoff=norm.args$Acutoff,p=norm.args$p)
if (output=="native")
return(dge) # Class: DGEList
else if (output=="matrix") {
scl <- dge$samples$lib.size * dge$samples$norm.factors
return(round(t(t(dge$counts)/scl)*mean(scl)))
#return(round(dge$pseudo.counts)) # Class: matrix
}
}
#' Normalization based on the NOISeq package
#'
#' This function is a wrapper over NOISeq normalization. It accepts a matrix of
#' gene counts (e.g. produced by importing an externally generated table of counts
#' to the main metaseqr pipeline).
#'
#' @param gene.counts a table where each row represents a gene and each column a
#' sample. Each cell contains the read counts for each gene and sample. Such a
#' table can be produced outside metaseqr and is imported during the basic metaseqr
#' workflow.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param norm.args a list of NOISeq normalization parameters. See the result of
#' \code{get.defaults("normalization",} \code{"noiseq")} for an example and how
#' you can modify it.
#' @param gene.data an optional annotation data frame (such the ones produced by
#' \code{get.annotation} which contains the GC content for each gene and from which
#' the gene lengths can be inferred by chromosome coordinates.
#' @param log.offset an offset to use to avoid infinity in logarithmic data
#' transformations.
#' @param output the class of the output object. It can be \code{"matrix"} (default)
#' for versatility with other tools or \code{"native"} for the NOISeq native S4
#' object (SeqExpressionSet). In the latter case it should be handled with suitable
#' NOISeq methods.
#' @return A matrix with normalized counts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.boxplot(data.matrix,sample.list)
#'
#' lengths <- round(1000*runif(nrow(data.matrix)))
#' starts <- round(1000*runif(nrow(data.matrix)))
#' ends <- starts + lengths
#' gc=runif(nrow(data.matrix)),
#' gene.data <- data.frame(
#' chromosome=c(rep("chr1",nrow(data.matrix)/2),rep("chr2",nrow(data.matrix)/2)),
#' start=starts,end=ends,gene_id=rownames(data.matrix),gc_content=gc
#' )
#' norm.data.matrix <- normalize.noiseq(data.matrix,sample.list,gene.data)
#' diagplot.boxplot(norm.data.matrix,sample.list)
#'}
normalize.noiseq <- function(gene.counts,sample.list,norm.args=NULL,
gene.data=NULL,log.offset=1,output=c("matrix","native")) {
if (is.null(norm.args))
norm.args <- get.defaults("normalization","noiseq")
output <- tolower(output[1])
check.text.args("output",output,c("matrix","native"))
classes <- as.class.vector(sample.list)
if (is.null(gene.data)) {
ns.obj <- NOISeq::readData(
data=gene.counts,
factors=data.frame(class=classes)
)
}
else {
gc.content <- gene.data$gc_content
gene.length <- attr(gene.data,"gene.length")
biotype <- as.character(gene.data$biotype)
names(gc.content) <- names(biotype) <- names(gene.length) <-
rownames(gene.data)
ns.obj <- NOISeq::readData(
data=gene.counts,
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)
#norm.args$long=gene.length # Set the gene length feature
}
norm.args$k=log.offset # Set the zero fixing constant
switch(norm.args$method,
rpkm = {
#M <- NOISeq::rpkm(assayData(ns.obj)$exprs,long=norm.args$long,
# k=norm.args$k,lc=norm.args$lc)
M <- rpkm(assayData(ns.obj)$exprs,gene.length=norm.args$long)
},
uqua = {
M <- NOISeq::uqua(assayData(ns.obj)$exprs,long=norm.args$long,
k=norm.args$k,lc=norm.args$lc)
},
tmm = {
M <- NOISeq::tmm(assayData(ns.obj)$exprs,long=norm.args$long,
k=norm.args$k,lc=norm.args$lc,refColumn=norm.args$refColumn,
logratioTrim=norm.args$logratioTrim,sumTrim=norm.args$sumTrim,
doWeighting=norm.args$doWeighting,Acutoff=norm.args$Acutoff)
}
)
if (output=="native") {
if (is.null(gene.data))
return(NOISeq::readData(
data=M,
factors=data.frame(class=classes)
)) # Class: CD
else
return(NOISeq::readData(
data=M,
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)) # Class: CD
}
else if (output=="matrix")
return(as.matrix(round(M))) # Class: matrix
}
#' Normalization based on the NBPSeq package
#'
#' This function is a wrapper over DESeq normalization. It accepts a matrix of gene
#' counts (e.g. produced by importing an externally generated table of counts to
#' the main metaseqr pipeline).
#'
#' @param gene.counts a table where each row represents a gene and each column a
#' sample. Each cell contains the read counts for each gene and sample. Such a
#' table can be produced outside metaseqr and is imported during the basic metaseqr
#' workflow.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param norm.args a list of NBPSeq normalization parameters. See the result of
#' \code{get.defaults("normalization",} \code{"nbpseq")} for an example and how
#' you can modify it.
#' @param libsize.list an optional named list where names represent samples (MUST
#' be the same as the samples in \code{sample.list}) and members are the library
#' sizes (the sequencing depth) for each sample. If not provided, the default is
#' the column sums of the \code{gene.counts} matrix.
#' @param output the class of the output object. It can be \code{"matrix"} (default)
#' for versatility with other tools or \code{"native"} for the NBPSeq native S4
#' object (a specific list). In the latter case it should be handled with suitable
#' NBPSeq methods.
#' @return A matrix with normalized counts or a list with the normalized counts
#' and other NBPSeq specific parameters.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.boxplot(data.matrix,sample.list)
#'
#' norm.data.matrix <- normalize.nbpseq(data.matrix,sample.list)
#' diagplot.boxplot(norm.data.matrix,sample.list)
#'}
normalize.nbpseq <- function(gene.counts,sample.list,norm.args=NULL,
libsize.list=NULL,output=c("matrix","native")) {
if (is.null(norm.args))
norm.args <- get.defaults("normalization","nbpseq")
output <- tolower(output[1])
check.text.args("output",output,c("matrix","native"))
classes <- as.class.vector(sample.list)
if (is.null(libsize.list)) {
libsize.list <- vector("list",length(classes))
names(libsize.list) <- unlist(sample.list,use.names=FALSE)
for (n in names(libsize.list))
libsize.list[[n]] <- sum(gene.counts[,n])
}
lib.sizes <- unlist(libsize.list)
norm.factors <- estimate.norm.factors(gene.counts,lib.sizes=lib.sizes,
method=norm.args$method)
#if (norm.args$main.method=="nbpseq")
# nb.data <- prepare.nb.data(gene.counts,lib.sizes=lib.sizes,
# norm.factors=norm.factors)
#else if (norm.args$main.method=="nbsmyth")
nb.data <- prepare.nbp(gene.counts,classes,lib.sizes=lib.sizes,
norm.factors=norm.factors,thinning=norm.args$thinning)
if (output=="native")
return(nb.data) # Class: list or nbp
else if (output=="matrix") {
#if (norm.args$main.method=="nbpseq") {
# norm.counts <- matrix(0,nrow(gene.counts),ncol(gene.counts))
# for (i in 1:ncol(gene.counts))
# norm.counts[,i] <- norm.factors[i]*gene.counts[,i]
#}
#else if (norm.args$main.method=="nbsmyth")
norm.counts <- nb.data$pseudo.counts
return(as.matrix(round(norm.counts))) # Class: matrix
}
}
<file_sep>/man/make.matrix.Rd
\name{make.matrix}
\alias{make.matrix}
\title{Results output build helper}
\usage{
make.matrix(samples, data.list, export.scale = "natural")
}
\arguments{
\item{samples}{a set of samples from the dataset under
processing. They should match sample names from
\code{sample.list}. See also the main help page of
\code{\link{metaseqr}}.}
\item{data.list}{a list containing natural or transformed
data, typically an output
from \code{\link{make.transformation}}.}
\item{export.scale}{the output transformations used as
input also to \code{\link{make.transformation}}.}
}
\value{
A named list whose names are the elements in
\code{export.scale}. Each list member is the respective
sample subest data matrix.
}
\description{
Returns a list of matrices based on the export scales
that have been chosen from the main function and a subset
of samples based on the sample names provided in the
\code{sample.list} argument of the main
\code{\link{metaseqr}} function. Internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
tr <- make.transformation(data.matrix,c("log2","vst"))
mm <- make.matrix(c("C1","T1"),tr,"log2")
head(tr$vst)
}
}
\author{
<NAME>
}
<file_sep>/man/make.sim.data.tcc.Rd
\name{make.sim.data.tcc}
\alias{make.sim.data.tcc}
\title{Create simulated counts using TCC package}
\usage{
make.sim.data.tcc(...)
}
\arguments{
\item{...}{parameters to the \code{simulateReadCounts}
function.}
}
\value{
A list with the following members: \code{simdata} holding
the simulated dataset complying with metaseqr
requirements, and \code{simparam} holding the simulation
parameters (see TCC documentation). Note that the produced
data are based in an Arabidopsis dataset.
}
\description{
This function creates simulated RNA-Seq gene expression
datasets using the \code{simulateReadCounts} function
from the Bioconductor package TCC and it adds simulated
annoation elements. For further information please
consult the TCC package documentation.
}
\examples{
\donttest{
dd <- make.sim.data.tcc(Ngene=10000,PDEG=0.2,
DEG.assign=c(0.9,0.1),
DEG.foldchange=c(5,5),replicates=c(3,3))
head(dd$simdata)
}
}
\author{
<NAME>
}
<file_sep>/man/check.main.args.Rd
\name{check.main.args}
\alias{check.main.args}
\title{Main argument validator}
\usage{
check.main.args(main.args)
}
\arguments{
\item{main.args}{ a list of parameters
with which metaseqr is called (essentially,
the output of \code{\link{match.call}}.}
}
\description{
Checks if the arguments passed to
\code{\link{metaseqr}} are valid and throws a
warning about the invalid ones (which are
ignored anyway because of the \code{...} in
\code{\link{metaseqr}}. However, for this reason
this function is useful as some important
parameter faults might go unnoticed in the
beginning and cause a failure afterwards.
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.filter.R
#' Filter gene expression based on exon counts
#'
#' This function performs the gene expression filtering based on exon read counts
# and a set of exon filter rules. For more details see the main help pages of
#' \code{\link{metaseqr}}.
#'
#' @param the.counts a named list created with the \code{\link{construct.gene.model}}
#' function. See its help page for details.
#' @param gene.data an annotation data frame usually obtained with
#' \code{\link{get.annotation}} containing the unique gene accession identifiers.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param exon.filters a named list with exon filters and their parameters. See
#' the main help page of \code{\link{metaseqr}} for details.
#' @param restrict.cores in case of parallel execution of several subfunctions,
#' the fraction of the available cores to use. In some cases if all available
#' cores are used (\code{restrict.cores=1} and the system does not have sufficient
#' RAM, the running machine might significantly slow down.
#' @return a named list with two members. The first member (\code{result} is a
#' named list whose names are the exon filter names and its members are the filtered
#' rownames of \code{gene.data}. The second member is a matrix of binary flags
#' (0 for non-filtered, 1 for filtered) for each gene. The rownames of the flag
#' matrix correspond to gene ids.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' data("hg19.exon.data",package="metaseqR")
#' exon.counts <- hg19.exon.counts
#' gene.data <- get.annotation("hg19","gene")
#' sample.list <- sample.list.hg19
#' exon.filters <- get.defaults("exon.filter")
#' the.counts <- construct.gene.model(exon.counts,sample.list,gene.data)
#' filter.results <- filter.exons(the.counts,gene.data,sample.list,exon.filters)
#'}
filter.exons <- function(the.counts,gene.data,sample.list,exon.filters,
restrict.cores=0.8) {
multic <- check.parallel(restrict.cores)
exon.filter.result <- vector("list",length(exon.filters))
names(exon.filter.result) <- names(exon.filters)
flags <- matrix(0,nrow(gene.data),1)
rownames(flags) <- rownames(gene.data)
colnames(flags) <- c("MAE")
the.genes <- as.character(gene.data$gene_id)
if (!is.null(exon.filters)) {
for (xf in names(exon.filters)) {
disp("Applying exon filter ",xf,"...")
switch(xf,
min.active.exons = {
pass <- vector("list",length(unlist(sample.list)))
names(pass) <- names(the.counts)
for (n in names(pass)) {
disp(" Checking read presence in exons for ",n,"...")
pass[[n]] <- the.genes
names(pass[[n]]) <- the.genes
pass[[n]] <- wapply(multic,the.counts[[n]],
function(x,f) {
if (length(x$count) == 1)
if (x$count[1]!=0)
return(FALSE)
else
return(TRUE)
else if (length(x$count) > 1 &&
length(x$count) <= f$exons.per.gene)
if (length(which(x$count!=0)) >= f$min.exons)
return(FALSE)
else
return(TRUE)
else
if (length(which(x$count!=0)) >=
ceiling(length(x$count)*f$frac))
return(FALSE)
else
return(TRUE)
},exon.filters$min.active.exons)
pass[[n]] <- do.call("c",pass[[n]])
}
pass.matrix <- do.call("cbind",pass)
exon.filter.result[[xf]] <- the.genes[which(apply(
pass.matrix,1,function(x) return(all(x))))]
flags[exon.filter.result[[xf]],"MAE"] <- 1
}
# More to come...
# TODO: Write more rules based in exons
)
}
}
return(list(result=exon.filter.result,flags=flags))
}
#' Filter gene expression based on gene counts
#'
#' This function performs the gene expression filtering based on gene read counts
#' and a set of gene filter rules. For more details see the main help pages of
#' \code{\link{metaseqr}}.
#'
#' @param gene.counts a matrix of gene counts, preferably after the normalization
#' procedure.
#' @param gene.data an annotation data frame usually obtained with
#' \code{\link{get.annotation}} containing the unique gene accession identifiers.
#' @param gene.filters a named list with gene filters and their parameters. See
#' the main help page of \code{\link{metaseqr}} for details.
#' @return a named list with three members. The first member (\code{result} is a
#' named list whose names are the gene filter names and its members are the
#' filtered rownames of \code{gene.data}. The second member (\code{cutoff} is a
#' named list whose names are the gene filter names and its members are the cutoff
#' values corresponding to each filter. The third member is a matrix of binary
#' flags (0 for non-filtered, 1 for filtered) for each gene. The rownames of the
#' flag matrix correspond to gene ids.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' data("mm9.gene.data",package="metaseqR")
#' gene.counts <- mm9.gene.counts
#' sample.list <- mm9.sample.list
#' gene.counts <- normalize.edger(as.matrix(gene.counts[,9:12]),sample.list)
#' gene.data <- get.annotation("mm9","gene")
#' gene.filters <- get.defaults("gene.filter","mm9")
#' filter.results <- filter.genes(gene.counts,gene.data,gene.filters)
#'}
filter.genes <- function(gene.counts,gene.data,gene.filters,sample.list) {
gene.filter.result <- gene.filter.cutoff <- vector("list",
length(gene.filters))
names(gene.filter.result) <- names(gene.filter.cutoff) <-
names(gene.filters)
flags <- matrix(0,nrow(gene.counts),9)
rownames(flags) <- rownames(gene.counts)
colnames(flags) <- c("LN","AR","MD","MN","QN","KN","CM","BT","PR")
for (gf in names(gene.filters)) {
disp("Applying gene filter ",gf,"...")
switch(gf,
length = { # This is real gene length independently of exons
if (!is.null(gene.filters$length)) {
gene.filter.result$length <- rownames(gene.data)[which(
gene.data$end - gene.data$start <
gene.filters$length$length
)]
gene.filter.cutoff$length <- gene.filters$length$length
flags[intersect(gene.filter.result$length,
rownames(gene.counts)),"LN"] <- 1
disp(" Threshold below which ignored: ",
gene.filters$length$length)
}
else
gene.filter.cutoff$length <- NULL
},
avg.reads = {
if (!is.null(gene.filters$avg.reads)) {
len <- attr(gene.data,"gene.length")
if (!is.null(len))
len <- len[rownames(gene.counts)]
else {
gg <- gene.data[rownames(gene.counts),]
len <- gg$end - gg$start
}
avg.mat <- sweep(gene.counts,1,
len/gene.filters$avg.reads$average.per.bp,"/")
q.t <- max(apply(avg.mat,2,quantile,
gene.filters$avg.reads$quantile))
gene.filter.result$avg.reads <- rownames(gene.data)[which(
apply(avg.mat,1,filter.low,q.t))]
gene.filter.cutoff$avg.reads <- q.t
flags[intersect(gene.filter.result$avg.reads,
rownames(gene.counts)),"AR"] <- 1
disp(" Threshold below which ignored: ",q.t)
}
else
gene.filter.cutoff$avg.reads <- NULL
},
expression = {
if (!is.null(gene.filters$expression)) {
if (!is.null(gene.filters$expression$median) &&
gene.filters$expression$median) {
md <- median(gene.counts)
the.dead.median <- rownames(gene.counts)[which(
apply(gene.counts,1,filter.low,md))]
disp(" Threshold below which ignored: ",md)
}
else
the.dead.median <- md <- NULL
if (!is.null(gene.filters$expression$mean) &&
gene.filters$expression$mean) {
mn <- mean(gene.counts)
the.dead.mean <- rownames(gene.counts)[which(apply(
gene.counts,1,filter.low,mn))]
disp(" Threshold below which ignored: ",mn)
}
else
the.dead.mean <- mn <- NULL
if (!is.null(gene.filters$expression$quantile) &&
!is.na(gene.filters$expression$quantile)) {
qu <- quantile(gene.counts,
gene.filters$expression$quantile)
the.dead.quantile <- rownames(gene.counts)[which(
apply(gene.counts,1,filter.low,qu))]
disp(" Threshold below which ignored: ",qu)
}
else
the.dead.quantile <- qu <- NULL
if (!is.null(gene.filters$expression$known) &&
!is.na(gene.filters$expression$known)) {
# Think about the case of embedded
bio.cut <- match(gene.filters$expression$known,
gene.data$gene_name)
bio.cut <- bio.cut[-which(is.na(bio.cut))]
bio.cut.counts <- as.vector(gene.counts[bio.cut,])
the.bio.cut <- quantile(bio.cut.counts,0.9)
the.dead.known <- rownames(gene.counts)[which(apply(
gene.counts,1,filter.low,the.bio.cut))]
disp(" Threshold below which ignored: ",the.bio.cut)
}
else
the.dead.known <- the.bio.cut <- NULL
if (!is.null(gene.filters$expression$custom) &&
!is.na(gene.filters$expression$custom)) {
# For future use
the.dead.custom <- NULL
}
else
the.dead.custom <- NULL
# Derive one common expression filter
gene.filter.result$expression$median <- the.dead.median
gene.filter.result$expression$mean <- the.dead.mean
gene.filter.result$expression$quantile <-
the.dead.quantile
gene.filter.result$expression$known <- the.dead.known
gene.filter.result$expression$custom <- the.dead.custom
gene.filter.cutoff$expression$median <- md
gene.filter.cutoff$expression$mean <- mn
gene.filter.cutoff$expression$quantile <- qu
gene.filter.cutoff$expression$known <- the.bio.cut
gene.filter.cutoff$expression$custom <- NULL
if (!is.null(the.dead.median)) flags[the.dead.median,
"MD"] <- 1
if (!is.null(the.dead.mean)) flags[the.dead.mean,"MN"] <- 1
if (!is.null(the.dead.quantile)) flags[the.dead.quantile,
"QN"] <- 1
if (!is.null(the.dead.known)) flags[the.dead.known,
"KN"] <- 1
if (!is.null(the.dead.custom)) flags[the.dead.custom,
"CM"] <- 1
#the.dead <- list(the.dead.median,the.dead.mean,
# the.dead.quantile,the.dead.known,the.dead.custom)
#gene.filter.result$expression <- Reduce("union",the.dead)
}
},
biotype = {
if (!is.null(gene.filters$biotype)) {
filter.out <- names(which(unlist(gene.filters$biotype)))
# Necessary hack because of R naming system
if (length(grep("three_prime_overlapping_ncrna",
filter.out))>0)
filter.out <- sub("three_prime_overlapping_ncrna",
"3prime_overlapping_ncrna",filter.out)
filter.ind <- vector("list",length(filter.out))
names(filter.ind) <- filter.out
for (bt in filter.out)
filter.ind[[bt]] <- rownames(gene.data)[which(
gene.data$biotype==bt)]
gene.filter.result$biotype <- Reduce("union",filter.ind)
gene.filter.cutoff$biotype <- paste(filter.out,
collapse=", ")
disp(" Biotypes ignored: ",paste(filter.out,
collapse=", "))
}
else
gene.filter.result$biotype <- NULL
if (!is.null(gene.filter.result$biotype) &&
length(gene.filter.result$biotype)>0)
flags[gene.filter.result$biotype,"BT"] <- 1
},
presence = {
if (!is.null(gene.filters$presence)) {
frac <- gene.filters$presence$frac
minc <- gene.filters$presence$min.count
pc <- gene.filters$presence$per.condition
if (pc) {
np.tmp <- nam.tmp <- vector("list",length(sample.list))
names(np.tmp) <- names(nam.tmp) <- names(sample.list)
for (n in names(sample.list)) {
tmp <- gene.counts[,sample.list[[n]]]
nreq <- ceiling(frac*ncol(tmp))
np.tmp[[n]] <- apply(tmp,1,function(x,m,n) {
w <- which(x>=m)
if (length(w)>0 && length(w)>n)
return(FALSE)
return(TRUE)
},minc,nreq)
nam.tmp[[n]] <- rownames(tmp[which(np.tmp[[n]]),,
drop=FALSE])
}
the.dead.presence <- Reduce("union",nam.tmp)
}
else {
nreq <- ceiling(frac*ncol(gene.counts))
not.present <- apply(gene.counts,1,function(x,m,n) {
w <- which(x>=m)
if (length(w)>0 && length(w)>n)
return(FALSE)
return(TRUE)
},minc,nreq)
the.dead.presence <-
rownames(gene.counts[which(not.present),,
drop=FALSE])
}
if (length(the.dead.presence)>0) {
gene.filter.result$presence <- the.dead.presence
flags[the.dead.presence,"PR"] <- 1
gene.filter.cutoff$presence <- nreq
disp(" Threshold below which ignored: ",nreq)
}
else
gene.filter.result$presence <-
gene.filter.cutoff$presence <- NULL
}
else
gene.filter.result$presence <- NULL
}
)
}
return(list(result=gene.filter.result,cutoff=gene.filter.cutoff,
flags=flags))
}
<file_sep>/man/check.file.args.Rd
\name{check.file.args}
\alias{check.file.args}
\title{File argument validator}
\usage{
check.file.args(arg.name, arg.value)
}
\arguments{
\item{arg.name}{argument name to display in a possible
error.}
\item{arg.value}{the filename to check.}
}
\description{
Checks if a file exists for specific arguments requiring
a file input. Internal use only.
}
\examples{
# OK
check.file.args("file",system.file("metaseqr_report.html",
package="metaseqR"))
## Error!
#check.file.args("file",system.file("metaseqr_report.htm",
# package="metaseqR"))
}
\author{
<NAME>
}
<file_sep>/man/libsize.list.mm9.Rd
\docType{data}
\name{libsize.list.mm9}
\alias{libsize.list.mm9}
\title{Mouse RNA-Seq data with two conditions, four samples}
\format{a named \code{list} with library sizes.}
\source{
ENCODE (http://genome.ucsc.edu/encode/)
}
\description{
The library size list for \code{mm9.gene.counts}. See the
data set description.
}
\author{
<NAME>
}
\keyword{datasets}
<file_sep>/man/get.preset.opts.Rd
\name{get.preset.opts}
\alias{get.preset.opts}
\title{Return several analysis options given an analysis preset}
\usage{
get.preset.opts(preset, org)
}
\arguments{
\item{preset}{preset can be one of \code{"all.basic"},
\code{"all.normal"}, \code{"all.full"},
\code{"medium.basic"}, \code{"medium.normal"},}
\item{org}{one of the supported organisms. See
\code{\link{metaseqr}} main help page.
\code{"medium.full"}, \code{"strict.basic"},
\code{"strict.normal"} or \code{"strict.full"}, each of
which control the strictness of the analysis and the
amount of data to be exported. For an explanation of the
presets, see the main \code{\link{metaseqr}} help page.}
}
\value{
A named list with names \code{exon.filters},
\code{gene.filters}, \code{pcut}, \code{export.what},
\code{export.scale}, \code{export.values} and
\code{export.stats}, each of which correspond to an
element of the metaseqr pipeline.
}
\description{
This is a helper function which returns a set of metaseqr
pipeline options, grouped together according to a preset
keyword. It is intended mostly for internal use.
}
\examples{
strict.preset <- get.preset.opts("strict.basic","mm9")
}
\author{
<NAME>
}
<file_sep>/man/graphics.open.Rd
\name{graphics.open}
\alias{graphics.open}
\title{Open plotting device}
\usage{
graphics.open(o, f, ...)
}
\arguments{
\item{o}{the plotting device, see main metaseqr function}
\item{f}{a filename, if the plotting device requires it
(e.g. \code{"pdf"})}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\description{
Wrapper function to open a plotting device. Internal use
only.
}
\examples{
\donttest{
graphics.open("pdf","test.pdf",width=12,height=12)
}
}
\author{
<NAME>
}
<file_sep>/man/disp.Rd
\name{disp}
\alias{disp}
\title{Message displayer}
\usage{
disp(...)
}
\arguments{
\item{...}{a vector of elements that compose the display
message.}
}
\description{
Displays a message during execution of the several
functions. Internal use.
}
\examples{
i <- 1
disp("Now running iteration ",i,"...")
}
\author{
<NAME>
}
<file_sep>/man/stat.nbpseq.Rd
\name{stat.nbpseq}
\alias{stat.nbpseq}
\title{Statistical testing with NBPSeq}
\usage{
stat.nbpseq(object, sample.list, contrast.list = NULL,
stat.args = NULL, libsize.list = NULL)
}
\arguments{
\item{object}{a matrix or an object specific to each
normalization algorithm supported by metaseqR, containing
normalized counts. Apart from matrix (also for NOISeq),
the object can be a SeqExpressionSet (EDASeq),
CountDataSet (DESeq), DGEList (edgeR) or list (NBPSeq).}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{contrast.list}{a named structured list of contrasts
as returned by \code{\link{make.contrast.list}} or just
the vector of contrasts as defined in the main help page
of \code{\link{metaseqr}}.}
\item{stat.args}{a list of NBPSeq statistical algorithm
parameters. See the result of
\code{get.defaults("statistics",} \code{"nbpseq")} for an
example and how you can modify it. It is not required
when the input object is already a list from NBPSeq
normalization as the dispersions are already estimated.}
\item{libsize.list}{an optional named list where names
represent samples (MUST be the same as the samples
\code{in sample.list}) and members are the library sizes
(the sequencing depth) for each sample. If not provided,
the default is the column sums of the \code{gene.counts}
matrix.}
}
\value{
A named list of p-values, whose names are the names of
the contrasts.
}
\description{
This function is a wrapper over NBPSeq statistical
testing. It accepts a matrix of normalized gene counts or
an S4 object specific to each normalization algorithm
supported by metaseqR.
}
\note{
There is currently a problem with the NBPSeq package and
the workflow that is specific to the NBPSeq package. The
problem has to do with function exporting as there are
certain functions which are not recognized from the
package internally. For this reason and until it is
fixed, only the Smyth workflow will be available with the
NBPSeq package.
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
norm.data.matrix <- normalize.nbpseq(data.matrix,sample.list)
p <- stat.nbpseq(norm.data.matrix,sample.list,contrast)
}
}
\author{
<NAME>
}
<file_sep>/man/make.highcharts.points.Rd
\name{make.highcharts.points}
\alias{make.highcharts.points}
\title{Interactive volcano plot helper}
\usage{
make.highcharts.points(x, y, a)
}
\arguments{
\item{x}{The x coordinates (should be a named vector!).}
\item{y}{The y coordinates.}
\item{a}{Alternative names for each point.}
}
\value{
A list that is later serialized to JSON.
}
\description{
Creates a list which contains the data series of a
scatterplot, to be used for serialization with highcharts
JavaScript plotting. framework. Internal use only.
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.json.R
#' MDS plot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diagplot.mds}}.
#'
#' @param obj A list holding MDS plot data. See \code{\link{diaplot.mds}}.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
mdsToJSON <- function(obj,jl=c("highcharts")) {
jl <- tolower(jl[1])
x <- obj$x
y <- obj$y
xlim <- obj$xlim
ylim <- obj$ylim
samples <- obj$samples
cols <- getColorScheme()
# Construct series
counter <- 0
series <- vector("list",length(samples))
names(series) <- names(samples)
for (n in names(series)) {
counter <- counter + 1
series[[n]] <- list()
series[[n]]$name=n
series[[n]]$type="scatter"
series[[n]]$color=cols$fill[counter]
series[[n]]$marker <- list(
lineWidth=1,
states=list(
hover=list(
enabled=TRUE,
lineColor=cols$border[counter]
),
select=list(
fillColor=cols$selected[counter],
lineColor=cols$border[counter],
lineWidth=2
)
)
)
m <- match(samples[[n]],names(x))
if (length(m)>0) {
series[[n]]$data <- make.highcharts.points(x[m],y[m])
}
}
switch(jl,
highcharts = {
point.format=paste("<strong>Sample name: </strong>{point.name}<br>",
"<strong>Principal coordinate 1: </strong>{point.x}<br>",
"<strong>Principal coordinate 2: </strong>{point.y}",sep="")
json <- toJSON(
list(
chart=list(
type="scatter",
zoomType="xy"
),
title=list(
text=paste("Multidimensional Scaling")
),
xAxis=list(
title=list(
useHTML=TRUE,
text="1<sup>st</sup> Principal Coordinate",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE,
gridLineWidth=1,
min=round(xlim[1],3),
max=round(xlim[2],3)
),
yAxis=list(
title=list(
useHTML=TRUE,
text="2<sup>nd</sup> Principal Coordinate",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE,
gridLineWidth=1,
min=round(ylim[1],3),
max=round(ylim[2],3)
),
plotOptions=list(
scatter=list(
allowPointSelect=TRUE,
tooltip=list(
headerFormat=paste("<span style=",
"\"font-size:1.1em;color:{series.color};",
"font-weight:bold\">{series.name}<br>",
sep=""),
pointFormat=point.format
)
)
),
series=unname(series)
)
)
}
)
return(json)
}
#' Biodetection counts plot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diaplot.noiseq}}.
#'
#' @param obj A list holding boxplot data. See \code{\link{diaplot.noiseq}}.
#' @param by Can be \code{"sample"} to create biotypes boxplots per sample or
#' \code{"biotype"} to create samples boxplots per biotype.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
countsBioToJSON <- function(obj,by=c("sample","biotype"),jl=c("highcharts")) {
by <- tolower(by[1])
jl <- tolower(jl[1])
samples <- obj$samples
status <- obj$status
altnames <- obj$altnames
counts <- round(2^obj$user$counts - 1)
counts[counts==0] <- 0.001
#counts <- obj$user$counts
covars <- obj$user$covars
biotypes <- unique(as.character(covars$biotype))
if (!is.null(altnames))
names(altnames) <- rownames(counts)
grouped <- FALSE
if (is.null(samples)) {
if (is.null(colnames(counts)))
samplenames <- paste("Sample",1:ncol(counts),sep=" ")
else
samplenames <- colnames(counts)
samples <- list(Samples=nams)
}
else if (is.list(samples)) {
samplenames <- unlist(samples,use.names=FALSE)
grouped <- TRUE
}
# y label formatter for logarithmic axis
y.label.formatter <- paste('function() {if(this.value === 0.001)',
'{return 0;} else {return Highcharts.Axis.prototype.',
'defaultLabelFormatter.call(this);}}',sep="")
tooltip.point.formatter <- paste("function() {",
" var min = this.low === 0.001 ? 0 : this.low;" ,
" var q1 = this.q1 === 0.001 ? 0 : this.q1;" ,
" var med = this.median === 0.001 ? 0 : this.median;",
" var q3 = this.q3 === 0.001 ? 0 : this.q3;",
" var max = this.high === 0.001 ? 0 : this.high;",
" var str = 'Maximum: ' + max + '<br/>' +",
" 'Upper quartile: ' + q3 + '<br/>' +",
" 'Median: ' + med + '<br/>' +",
" 'Lower quartile: ' + q1 + '<br/>' +",
" 'Minimum: ' + min + '<br/>';",
" return str;",
"}",sep="")
# Legend clicker
boxplot.onclick <- paste("function() { ",
" var chart = this.chart;",
" var outlier_id = chart.get(this.name);",
" if (!outlier_id.visible) {",
" outlier_id.show();",
" } else {",
" outlier_id.hide();",
" }",
"}",sep="")
# Outliers tooltip
if (is.null(obj$altnames)) {
outlier.pointformat <- paste(
'<strong>Sample {point.category}</strong><br/>',
'Gene ID: {point.name}<br/>',
'Value: {point.y}<br/>',sep=""
)
}
else {
outlier.pointformat <- paste(
'<strong>Sample {point.category}</strong><br/>',
'Gene ID: {point.name}<br/>',
'Gene name: {point.alt_name}<br/>',
'Value: {point.y}<br/>',
sep=""
)
}
if (by=="sample") {
cols <- getColorScheme(length(biotypes))
box.list <- json <- vector("list",length(samplenames))
names(box.list) <- names(json) <- samplenames
for (n in samplenames) {
box.list[[n]] <- vector("list",length(biotypes))
names(box.list[[n]]) <- biotypes
for (b in biotypes)
box.list[[n]][[b]] <- counts[covars$biotype==b,n]
B <- boxplot(box.list[[n]],plot=FALSE)$stats
colnames(B) <- biotypes
o.list <- lapply(names(box.list[[n]]),function(x,M,b) {
v <- b[,x]
o <- which(M[[x]]<v[1] | M[[x]]>v[5])
if (length(o)>0)
return(M[[x]][o])
else
return(NULL)
},box.list[[n]],B)
names(o.list) <- biotypes
# Data series
BB <- matrix(0,nrow(B),ncol(B)) # Workaround of strange problem...
colnames(BB) <- colnames(B)
for (jj in 1:ncol(B))
BB[,jj] <- round(B[,jj],3)
d <- as.data.frame(BB)
ids <- 0:(ncol(d)-1)
d <- rbind(ids,d)
names(ids) <- colnames(d)
counter <- 0
series <- vector("list",length(biotypes))
names(series) <- biotypes
for (s in names(series)) {
counter <- counter + 1
series[[s]] <- list()
series[[s]]$name <- s
series[[s]]$color <- cols$fill[counter]
#series[[s]]$turboThreshold <- 10000
series[[s]]$data <- list(unname(as.list(d[,s])))
r <- round(d[,s])
series[[s]]$tooltip=list(
pointFormat=paste('<strong>Population: ',
length(box.list[[n]][[s]]),'</strong><br/>',
'Maximum: ',r[6],'<br/>',
'Upper quartile: ',r[5],'<br/>',
'Median: ',r[4],'<br/>',
'Lower quartile: ',r[3],'<br/>',
'Minimum: ',r[2],'<br/>',sep="")
)
}
# Outlier series (if any)
counter <- 0
outliers <- vector("list",length(biotypes))
names(outliers) <- biotypes
for (o in names(outliers)) {
counter <- counter + 1
outliers[[o]] <- list()
outliers[[o]]$id <- o
outliers[[o]]$name <- o
outliers[[o]]$type <- "scatter"
outliers[[o]]$showInLegend <- FALSE
#outliers[[o]]$turboThreshold <- 10000
outliers[[o]]$color <- cols$fill[counter]
outliers[[o]]$marker <- list(
fillColor=cols$fill[counter],
symbol="circle",
lineWidth=1,
lineColor=cols$border[counter]
)
outliers[[o]]$data <- list()
x <- rep(d[1,o],length(o.list[[o]]))
names(x) <- names(o.list[[o]])
if (is.null(obj$altnames)) {
outliers[[o]]$data <-
make.highcharts.points(x,o.list[[o]])
}
else {
outliers[[o]]$data <-
make.highcharts.points(x,o.list[[o]],
unname(altnames[names(x)]))
}
}
json[[n]] <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="boxplot"
),
title=list(
text=paste("Biotype detection for sample ",n,
sep="")
),
legend=list(
enabled=TRUE,
itemHoverStyle=list(
color="#B40000"
)
),
xAxis=list(
categories=biotypes,
title=list(
text="Biotype",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
)
),
yAxis=list(
type="logarithmic",
showFirstLabel=FALSE,
min=1e-4,
tickInterval=1,
title=list(
useHTML=TRUE,
#text="Read count (log<sub>2</sub>)",
text="Expression (read count)",
margin=25,
style=list(
color="#000000",
fontSize="1.1em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
),
formatter=y.label.formatter
)
),
plotOptions=list(
boxplot=list(
fillColor="#F0F0E0",
lineWidth=2,
medianColor="#000000",
medianWidth=3,
stemColor="#000000",
stemDashStyle="dash",
stemWidth=1,
whiskerColor="#000000",
whiskerLength="75%",
whiskerWidth=1,
grouping=FALSE,
tooltip=list(
headerFormat=paste(
'<span style="font-size:1.1em;',
'color:{series.color};',
'font-weight:bold">',
'\u25CF </span>',
'<span style="font-size:1.1em;',
'font-weight:bold">',
'Biotype {series.name}</span><br/>',
sep=""
)
),
events=list(
legendItemClick=boxplot.onclick
)
),
scatter=list(
allowPointSelect=TRUE,
tooltip=list(
headerFormat=paste(
'<span style="font-weight:bold;',
'color:{series.color};">',
'\u25CF </span>',
'<span style="font-weight:bold">',
'Biotype {series.name}</span><br/>',
sep=""
),
pointFormat=outlier.pointformat
),
states=list(
hover=list(
marker=list(
enabled=FALSE
)
)
)
)
),
series=c(unname(series),unname(outliers))
)
)
}
)
}
return(unquote_js_fun(json))
}
else if (by=="biotype") {
cols <- getColorScheme(length(samples))
box.list <- json <- vector("list",length(biotypes))
names(box.list) <- names(json) <- biotypes
for (b in biotypes) {
box.list[[b]] <- vector("list",length(samplenames))
names(box.list[[b]]) <- samplenames
for (n in samplenames)
box.list[[b]][[n]] <- counts[covars$biotype==b,n]
B <- boxplot(box.list[[b]],plot=FALSE)$stats
colnames(B) <- samplenames
o.list <- lapply(names(box.list[[b]]),function(x,M,b) {
v <- b[,x]
o <- which(M[[x]]<v[1] | M[[x]]>v[5])
if (length(o)>0)
return(M[[x]][o])
else
return(NULL)
},box.list[[b]],B)
names(o.list) <- samplenames
# Data series
BB <- matrix(0,nrow(B),ncol(B)) # Workaround of strange problem...
colnames(BB) <- colnames(B)
for (jj in 1:ncol(B))
BB[,jj] <- round(B[,jj],3)
d <- as.data.frame(BB)
ids <- 0:(ncol(d)-1)
d <- rbind(ids,d)
names(ids) <- colnames(d)
counter <- 0
series <- vector("list",length(samples))
names(series) <- names(samples)
for (s in names(series)) {
counter <- counter + 1
series[[s]] <- list()
series[[s]]$name=s
if (grouped)
series[[s]]$color=cols$fill[counter]
else
series[[s]]$color=cols$fill[1]
m <- match(samples[[s]],colnames(d))
series[[s]]$data <- unname(as.list(d[,m]))
}
# Outlier series (if any)
counter <- 0
outliers <- vector("list",length(samples))
names(outliers) <- names(samples)
for (o in names(outliers)) {
counter <- counter + 1
outliers[[o]] <- list()
outliers[[o]]$id <- o
outliers[[o]]$name <- o
outliers[[o]]$type <- "scatter"
outliers[[o]]$showInLegend <- FALSE
if (grouped) {
outliers[[o]]$color <- cols$fill[counter]
outliers[[o]]$marker <- list(
fillColor=cols$fill[counter],
symbol="circle",
lineWidth=1,
lineColor=cols$border[counter]
)
}
else {
outliers[[o]]$color <- cols$fill[1]
outliers[[o]]$marker <- list(
fillColor=cols$fill[1],
symbol="circle",
lineWidth=1,
lineColor=cols$border[1]
)
}
outliers[[o]]$data <- list()
m <- match(samples[[o]],colnames(d))
if (length(m)>0) {
for (i in m) {
x <- rep(d[1,i],length(o.list[[i]]))
names(x) <- names(o.list[[i]])
if (is.null(obj$altnames)) {
outliers[[o]]$data <-
make.highcharts.points(x,o.list[[i]])
}
else {
outliers[[o]]$data <- c(outliers[[o]]$data,
make.highcharts.points(x,o.list[[i]],
unname(altnames)))
}
}
}
}
json[[b]] <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="boxplot"
),
title=list(
text=paste("Detection for biotype ",b,
" (population: ",lengths(box.list[[b]])[1],
")",sep="")
),
legend=list(
enabled=TRUE
),
xAxis=list(
categories=samplenames,
title=list(
text="Sample name",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
)
),
yAxis=list(
type="logarithmic",
showFirstLabel=FALSE,
min=1e-4,
tickInterval=1,
title=list(
text="Expression (read count)",
margin=25,
style=list(
color="#000000",
fontSize="1.1em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
),
formatter=y.label.formatter
)
),
plotOptions=list(
boxplot=list(
fillColor="#F0F0E0",
lineWidth=2,
medianColor="#000000",
medianWidth=3,
stemColor="#000000",
stemDashStyle="dash",
stemWidth=1,
whiskerColor="#000000",
whiskerLength="75%",
whiskerWidth=1,
grouping=FALSE,
tooltip=list(
headerFormat=paste(
'<span style="font-size:1.1em;',
'color:{series.color};',
'font-weight:bold">',
'\u25CF </span>',
'<span style="font-size:1.1em;',
'font-weight:bold">',
'Condition {series.name}</span>',
'<br/>',
'<span style="font-weight:bold;">',
'Sample {point.key}',
'</span><br/>',sep=""
),
pointFormatter=tooltip.point.formatter
),
events=list(
legendItemClick=boxplot.onclick
)
),
scatter=list(
allowPointSelect=TRUE,
tooltip=list(
headerFormat=paste(
'<span style="font-weight:bold;',
'color:{series.color};">',
'\u25CF </span>',
'<span style="font-weight:bold">',
'Condition {series.name}</span>',
'<br/>',sep=""
),
pointFormat=outlier.pointformat
),
states=list(
hover=list(
marker=list(
enabled=FALSE
)
)
)
)
),
series=c(unname(series),unname(outliers))
)
)
}
)
}
return(unquote_js_fun(json))
}
}
#' Biodetection barplot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diaplot.noiseq}}.
#'
#' @param obj A list holding boxplot data. See \code{\link{diaplot.noiseq}}.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
bioDetectionToJSON <- function(obj,jl=c("highcharts")) {
jl <- tolower(jl[1])
samples <- obj$samples
status <- obj$status
plotdata <- obj$user$plotdata
covars <- obj$user$covars
if (!is.null(samples)&& is.list(samples)) {
samplenames <- unlist(samples,use.names=FALSE)
names(plotdata$biotables) <- samplenames
}
# Otherwise we are using the names present in the input object
abu <- which(plotdata$genome>7)
nabu <- which(plotdata$genome<=7)
cols <- getColorScheme()
json <- vector("list",length(samplenames))
names(json) <- samplenames
for (n in samplenames) {
# Data series
series.abu <- vector("list",3)
names(series.abu) <- c("genome","detectionVSgenome","detectionVSsample")
series.abu$genome <- list()
series.abu$genome$id <- "abu_genome"
series.abu$genome$name <- "% in genome"
series.abu$genome$color <- cols$trans[1]
series.abu$genome$pointPlacement <- -0.2
series.abu$genome$data <- round(as.numeric(plotdata$genome[abu]),3)
series.abu$detectionVSgenome <- list()
series.abu$detectionVSgenome$id <- "abu_detected"
series.abu$detectionVSgenome$name <- "% detected"
series.abu$detectionVSgenome$color <- cols$trans[2]
series.abu$detectionVSgenome$pointPlacement <- 0
series.abu$detectionVSgenome$data <- round(as.numeric(
plotdata$biotables[[n]][1,abu]),3)
series.abu$detectionVSsample <- list()
series.abu$detectionVSsample$id <- "abu_sample"
series.abu$detectionVSsample$name <- "% in sample"
series.abu$detectionVSsample$color <- cols$trans[3]
series.abu$detectionVSsample$pointPlacement <- 0.2
series.abu$detectionVSsample$data <- round(as.numeric(
plotdata$biotables[[n]][2,abu]),3)
series.nabu <- vector("list",3)
names(series.nabu) <- c("genome","detectionVSgenome",
"detectionVSsample")
series.nabu$genome <- list()
series.nabu$genome$name <- "% in genome"
series.nabu$genome$yAxis <- 1
series.nabu$genome$pointStart <- length(abu)
series.nabu$genome$linkedTo <- "abu_genome"
series.nabu$genome$color <- cols$trans[1]
series.nabu$genome$pointPlacement <- -0.2
series.nabu$genome$data <- round(as.numeric(plotdata$genome[nabu]),3)
series.nabu$detectionVSgenome <- list()
series.nabu$detectionVSgenome$name <- "% detected"
series.nabu$detectionVSgenome$yAxis <- 1
series.nabu$detectionVSgenome$pointStart <- length(abu)
series.nabu$detectionVSgenome$linkedTo <- "abu_detected"
series.nabu$detectionVSgenome$color <- cols$trans[2]
series.nabu$detectionVSgenome$pointPlacement <- 0
series.nabu$detectionVSgenome$data <- round(as.numeric(
plotdata$biotables[[n]][1,nabu]),3)
series.nabu$detectionVSsample <- list()
series.nabu$detectionVSsample$name <- "% in sample"
series.nabu$detectionVSsample$yAxis <- 1
series.nabu$detectionVSsample$pointStart <- length(abu)
series.nabu$detectionVSsample$linkedTo <- "abu_sample"
series.nabu$detectionVSsample$color <- cols$trans[3]
series.nabu$detectionVSsample$pointPlacement <- 0.2
series.nabu$detectionVSsample$data <- round(as.numeric(
plotdata$biotables[[n]][2,nabu]),3)
json[[n]] <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="column",
alignTicks=FALSE
),
title=list(
text=paste("Comparative biotype detection for ",
"sample ",n,sep="")
),
legend=list(
enabled=TRUE,
itemHoverStyle=list(
color="#B40000"
)
),
tooltip=list(
shared=TRUE
),
xAxis=list(
categories=names(plotdata$genome)[c(abu,nabu)],
title=list(
text="Biotype",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
),
plotLines=list(
list(
color="#8A8A8A",
width=1.5,
dashStyle="Dash",
value=length(abu)-0.5
)
),
plotBands=list(
list(
color="#FFFFE0",
from=-0.5,
to=length(abu)-0.5
),
list(
color="#FFECEB",
from=length(abu)-0.5,
to=length(plotdata$genome)
)
)
),
yAxis=list(
list(
min=0,
max=70,
title=list(
text="% of abundant features",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
)
),
list(
min=0,
max=7,
title=list(
text="% of non-abundant features",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
opposite=TRUE
)
),
plotOptions=list(
column=list(
grouping=FALSE,
shadow=FALSE,
groupPadding=0.3,
pointPadding=0.25,
tooltip=list(
headerFormat=paste(
'<span style="font-size:1.1em;',
'font-weight:bold">',
'{point.key}</span><br/>',sep=""
)
)
)
),
series=c(unname(series.abu),unname(series.nabu))
)
)
}
)
}
return(json)
}
#' Biotype saturation plot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diaplot.noiseq}}.
#'
#' @param obj A list holding boxplot data. See \code{\link{diaplot.noiseq}}.
#' @param by Can be \code{"sample"} to create biotypes boxplots per sample or
#' \code{"biotype"} to create samples boxplots per biotype.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
bioSaturationToJSON <- function(obj,by=c("sample","biotype"),
jl=c("highcharts")) {
by <- tolower(by[1])
jl <- tolower(jl[1])
samples <- obj$samples
plotdata <- obj$user$plotdata
if (!is.null(samples)&& is.list(samples)) {
samplenames <- unlist(samples,use.names=FALSE)
names(plotdata) <- samplenames
}
if (by=="sample") {
json <- vector("list",length(samplenames))
names(json) <- samplenames
for (n in samplenames) {
depth <- round(plotdata[[n]][,1]/1e+6)
global <- round(plotdata[[n]][,2])
M <- plotdata[[n]][,3:ncol(plotdata[[n]])]
# To determine the separation
ord <- sort(M[nrow(M),],decreasing=TRUE,index.return=TRUE)
abu <- ord$ix[1:2]
names(abu) <- names(ord$x[1:2])
nabu <- ord$ix[3:length(ord$ix)]
names(nabu) <- names(ord$x[3:length(ord$x)])
cols <- getColorScheme(ncol(plotdata[[n]])-1)
counter <- 1
global.series <- list(
global=list(
id="global",
name="global",
color=cols$fill[counter],
data=make.highcharts.points(depth,global)
)
)
abu.series <- vector("list",2)
names(abu.series) <- names(abu)
for (s in names(abu.series)) {
counter <- counter + 1
abu.series[[s]] <- list()
abu.series[[s]]$id <- s
abu.series[[s]]$name <- s
abu.series[[s]]$color <- cols$fill[counter]
abu.series[[s]]$data <- make.highcharts.points(depth,
round(M[,s]))
}
nabu.series <- vector("list",length(3:ncol(M)))
names(nabu.series) <- names(nabu)
for (s in names(nabu.series)) {
counter <- counter + 1
nabu.series[[s]] <- list()
nabu.series[[s]]$id <- s
nabu.series[[s]]$name <- s
nabu.series[[s]]$color <- cols$fill[counter]
nabu.series[[s]]$data <- make.highcharts.points(depth,
round(M[,s]))
}
json[[n]] <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="scatter",
zoomType="xy"
),
title=list(
text=paste("Biotype saturations for sample ",n,
sep="")
),
legend=list(
enabled=TRUE,
itemHoverStyle=list(
color="#B40000"
)
),
xAxis=list(
title=list(
text="Read depth (millions of reads)",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
)
),
yAxis=list(
min=0,
max=max(global),
title=list(
text="Detected features",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
)
),
plotOptions=list(
series=list(
lineWidth=2
),
scatter=list(
tooltip=list(
headerFormat=paste(
'<span style="font-weight:bold;',
'color:{series.color};">',
'\u25CF </span>',
'<span style="font-weight:bold">',
'Biotype {series.name}</span><br/>',
sep=""
),
pointFormat=paste(
"Depth: {point.x}M<br/>",
"Detected features: {point.y}",
sep="")
)
)
),
series=c(unname(global.series),
unname(abu.series),unname(nabu.series))
)
)
}
)
}
return(json)
}
else if (by=="biotype") {
biotypes <- colnames(plotdata[[1]])[2:ncol(plotdata[[1]])]
depths <- vector("list",length(plotdata))
names(depths) <- samplenames
for (n in samplenames)
depths[[n]] <- round(plotdata[[n]][,1]/1e+6)
json <- vector("list",length(biotypes))
names(json) <- biotypes
for (b in biotypes) {
series <- vector("list",length(plotdata))
names(series) <- samplenames
cols <- getColorScheme(length(samplenames))
counter <- 0
for (s in names(series)) {
counter <- counter + 1
series[[s]] <- list()
series[[s]]$id <- s
series[[s]]$name <- s
series[[s]]$color <- cols$fill[counter]
series[[s]]$data <- make.highcharts.points(depths[[s]],
round(plotdata[[s]][,b]))
}
json[[b]] <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="scatter",
zoomType="xy"
),
title=list(
text=paste("Sample saturations for biotype ",b,
sep="")
),
legend=list(
enabled=TRUE,
itemHoverStyle=list(
color="#B40000"
)
),
xAxis=list(
title=list(
text="Read depth (millions of reads)",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
)
),
yAxis=list(
title=list(
text="Detected features",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
)
),
plotOptions=list(
series=list(
lineWidth=2
),
scatter=list(
tooltip=list(
headerFormat=paste(
'<span style="font-weight:bold;',
'color:{series.color};">',
'\u25CF </span>',
'<span style="font-weight:bold">',
'Sample {series.name}</span><br/>',
sep=""
),
pointFormat=paste(
"Depth: {point.x}M<br/>",
"Detected features: {point.y}",
sep="")
)
)
),
series=c(unname(series))
)
)
}
)
}
return(json)
}
}
#' Read noise plot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diagplot.noiseq}}.
#'
#' @param obj A list holding plot data. See \code{\link{diaplot.noiseq}}.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
readNoiseToJSON <- function(obj,jl=c("highcharts"),seed=42) {
jl <- tolower(jl[1])
d <- obj$user
samples <- obj$samples
# Too many points for a lot of curves of interactive data
if (nrow(d)>1000) {
set.seed(seed)
ii <- sort(sample(1:nrow(d),998))
ii <- c(1,ii,nrow(d))
d <- cbind(d[ii,1],d[ii,2:ncol(d)])
}
if (is.null(samples))
samples <- 1:(ncol(d)-1)
if (is.numeric(samples))
samplenames = colnames(dat)[samples+1]
if (is.list(samples))
samplenames <- unlist(samples)
cols <- getColorScheme(length(samplenames))
# Construct series
counter <- 0
series <- vector("list",length(samplenames))
names(series) <- samplenames
for (n in names(series)) {
counter <- counter + 1
series[[n]] <- list()
series[[n]]$name=n
series[[n]]$color=cols$fill[counter]
series[[n]]$data <- make.highcharts.points(d[,1],d[,n])
series[[n]]$tooltip=list(
headerFormat=paste("<span style=",
"\"font-size:1.1em;color:{series.color};",
"font-weight:bold\">{series.name}<br>",
sep=""),
pointFormat=NULL
)
}
switch(jl,
highcharts = {
json <- toJSON(list(
chart=list(
type="line",
zoomType="xy"
),
title=list(
text=paste("RNA-Seq mapped reads noise")
),
xAxis=list(
title=list(
text="% detected features",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE,
gridLineWidth=1,
min=0,
max=100
),
yAxis=list(
title=list(
text="% of total reads",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE,
gridLineWidth=1,
tickPositions=seq(0,110,10)
),
plotOptions=list(
line=list(
allowPointSelect=TRUE,
lineWidth=1,
marker=list(
enabled=FALSE
),
tooltip=list(
headerFormat=paste("<span style=",
"\"font-size:1.1em;color:{series.color};",
"font-weight:bold\">{series.name}<br>",
sep=""),
pointFormat=NULL
),
turboThreshold=50000
)
),
series=unname(series)
)
)
}
)
return(json)
}
#' Boxplots JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diaplot.boxplot}}.
#'
#' @param obj A list holding boxplot data. See \code{\link{diaplot.boxplot}}.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
boxplotToJSON <- function(obj,jl=c("highcharts")) {
jl <- tolower(jl[1])
b <- obj$plot
name <- obj$samples
status <- obj$status
altnames <- obj$altnames
o.list <- obj$user
grouped <- FALSE
if (is.null(name)) {
if (is.null(colnames(b$stats)))
nams <- paste("Sample",1:ncol(b$stats),sep=" ")
else
nams <- colnames(b$stats)
name <- list(Samples=nams)
}
else if (length(name)==1 && name=="none") {
nams <- rep("",ncol(b$stats))
name <- list(Samples=nams)
}
else if (is.list(name)) { # Is sample.list
nams <- unlist(name,use.names=FALSE)
grouped <- TRUE
}
cols <- getColorScheme()
# Data series
d <- as.data.frame(round(b$stat,3))
ids <- 0:(ncol(d)-1)
d <- rbind(ids,d)
colnames(d) <- nams
names(ids) <- colnames(d)
counter <- 0
series <- vector("list",length(name))
names(series) <- names(name)
for (n in names(series)) {
counter <- counter + 1
series[[n]] <- list()
series[[n]]$name=n
if (grouped)
series[[n]]$color=cols$fill[counter]
else
series[[n]]$color=cols$fill[1]
m <- match(name[[n]],colnames(d))
series[[n]]$data <- unname(as.list(d[,m]))
}
# Outlier series (if any)
counter <- 0
outliers <- vector("list",length(name))
names(outliers) <- names(name)
for (n in names(outliers)) {
counter <- counter + 1
outliers[[n]] <- list()
outliers[[n]]$id <- n
outliers[[n]]$name <- n
outliers[[n]]$type <- "scatter"
outliers[[n]]$showInLegend <- FALSE
if (grouped) {
outliers[[n]]$color <- cols$fill[counter]
outliers[[n]]$marker <- list(
fillColor=cols$fill[counter],
symbol="circle",
lineWidth=1,
lineColor=cols$border[counter]
)
}
else {
outliers[[n]]$color <- cols$fill[1]
outliers[[n]]$marker <- list(
fillColor=cols$fill[1],
symbol="circle",
lineWidth=1,
lineColor=cols$border[1]
)
}
outliers[[n]]$data <- list()
m <- match(name[[n]],colnames(d))
if (length(m)>0) {
for (i in m)
x <- rep(d[1,i],length(o.list[[i]]))
names(x) <- names(o.list[[i]])
outliers[[n]]$data <- c(outliers[[n]]$data,
make.highcharts.points(x,o.list[[i]],unname(altnames)))
}
}
# Boxplot tooltip point formatter for the case of zeros
tooltip.point.formatter <- paste("function() {",
" var min = this.low === 0.001 ? 0 : this.low;" ,
" var q1 = this.q1 === 0.001 ? 0 : this.q1;" ,
" var med = this.median === 0.001 ? 0 : this.median;",
" var q3 = this.q3 === 0.001 ? 0 : this.q3;",
" var max = this.high === 0.001 ? 0 : this.high;",
" var str = 'Maximum: ' + max + '<br/>' +",
" 'Upper quartile: ' + q3 + '<br/>' +",
" 'Median: ' + med + '<br/>' +",
" 'Lower quartile: ' + q1 + '<br/>' +",
" 'Minimum: ' + min + '<br/>';",
" return str;",
"}",sep="")
# Legend clicker
boxplot.onclick <- paste("function() {",
" var chart = this.chart;",
" var outlier_id = chart.get(this.name);",
" if (!outlier_id.visible) {",
" outlier_id.show();",
" } else {",
" outlier_id.hide();",
" }",
"}",sep="")
if (is.null(obj$altnames)) {
outlier.pointformat=paste(
'<strong>Sample {point.category}</strong><br/>',
'Gene ID: {point.name}<br/>',
'Value: {point.y}<br/>',sep=""
)
}
else {
outlier.pointformat=paste(
'<strong>Sample {point.category}</strong><br/>',
'Gene ID: {point.name}<br/>',
'Gene name: {point.alt_name}<br/>',
'Value: {point.y}<br/>',sep=""
)
}
json <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="boxplot"
),
title=list(
text=paste("Boxplot ",status,sep="")
),
legend=list(
enabled=TRUE
),
xAxis=list(
categories=nams,
title=list(
text="Sample name",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
)
),
yAxis=list(
title=list(
useHTML=TRUE,
text="Read count (log<sub>2</sub>)",
margin=30,
style=list(
color="#000000",
fontSize="1.1em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
)
),
plotOptions=list(
boxplot=list(
fillColor="#F0F0E0",
lineWidth=2,
medianColor="#000000",
medianWidth=3,
stemColor="#000000",
stemDashStyle="dash",
stemWidth=1,
whiskerColor="#000000",
whiskerLength="75%",
whiskerWidth=1,
grouping=FALSE,
tooltip=list(
headerFormat=paste(
'<span style="font-size:1.1em;',
'color:{series.color};',
'font-weight:bold">',
'\u25CF </span>',
'<span style="font-size:1.1em;',
'font-weight:bold">',
'Condition {series.name}</span><br/>',
'<span style="font-weight:bold">',
'Sample {point.key}</span><br/>',sep=""
),
pointFormatter=tooltip.point.formatter
),
events=list(
legendItemClick=boxplot.onclick
)
),
scatter=list(
allowPointSelect=TRUE,
tooltip=list(
headerFormat=paste(
'<span style="font-weight:bold;',
'color:{series.color};">',
'\u25CF </span>',
'<span style="font-weight:bold">',
'Condition {series.name}</span><br/>',
sep=""
),
pointFormat=outlier.pointformat
),
states=list(
hover=list(
marker=list(
enabled=FALSE
)
)
)
)
),
series=c(unname(series),unname(outliers))
)
)
}
)
return(unquote_js_fun(json))
}
#' GC/length bias plot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diagplot.edaseq}}.
#'
#' @param obj A list holding plot data. See \code{\link{diaplot.noiseq}}.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
biasPlotToJSON <- function(obj,jl=c("highcharts"),seed=1) {
jl <- tolower(jl[1])
counts <- round(nat2log(obj$user$counts),3)
status <- obj$status
covar <- obj$user$covar
covarname <- obj$user$covarname
samples <- obj$samples
# Too many points for a lot of curves of interactive data
if (nrow(counts)>2000) {
set.seed(seed)
ii <- sample(1:nrow(counts),2000)
counts <- counts[ii,]
covar <- covar[ii]
}
# If length bias, not nice to have x-axis at -200k
min.x <- ifelse(max(covar>100),0,"undefined")
if (is.null(samples)) {
if (is.null(colnames(x)))
samplenames <- paste("Sample",1:ncol(counts),sep=" ")
else
samplenames <- colnames(counts)
samples <- list(Samples=nams)
cols <- getColorScheme(length(samples))
}
else if (is.list(samples)) { # Is sample.list
samplenames <- unlist(samples,use.names=FALSE)
grouped <- TRUE
}
colnames(counts) <- samplenames
# Construct series
counter <- 0
series <- vector("list",length(samplenames))
names(series) <- samplenames
for (n in names(series)) {
counter <- counter + 1
x <- counts[,n]
fit <- lowess(covar,x)
series[[n]] <- list()
series[[n]]$name <- n
series[[n]]$color <- cols$fill[counter]
series[[n]]$data <- lapply(1:length(x),function(i,x,y) {
return(c(x[i],y[i])) },round(fit$x,3),round(fit$y,3))
}
switch(jl,
highcharts = {
json <- toJSON(list(
chart=list(
type="line",
zoomType="xy"
),
title=list(
text=paste(covarname," bias detection - ",status)
),
xAxis=list(
min=min.x,
title=list(
text=covarname,
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE
),
yAxis=list(
title=list(
useHTML=TRUE,
text="Read count (log<sub>2</sub>)",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE
),
plotOptions=list(
line=list(
marker=list(
enabled=FALSE,
states=list(
hover=list(
enabled=FALSE
)
)
),
tooltip=list(
headerFormat=paste("<span style=",
"\"font-size:1.1em;color:{series.color};",
"font-weight:bold\">{series.name}<br>",
sep=""),
pointFormat=NULL
),
turboThreshold=50000
)
),
series=unname(series)
)
)
}
)
return(json)
}
#' Filtered genes barplot JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diaplot.filtered}}.
#'
#' @param obj A list holding plot data. See \code{\link{diaplot.filered}}.
#' @param by Either \code{"chromosome"} or \code{"biotype"}
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
filteredToJSON <- function(obj,by=c("chromosome","biotype"),
jl=c("highcharts")) {
jl <- tolower(jl[1])
by <- tolower(by[1])
filtered <- obj$user$filtered
total <- obj$user$total
cols <- getColorScheme(2)
if (by=="chromosome") {
chr <- table(as.character(filtered$chromosome))
chr.all <- table(as.character(total$chromosome))
barlab.chr <- as.character(chr)
per.chr <- round(chr/chr.all[names(chr)],3)
per.chr[per.chr>1] <- 1
series <- vector("list",2)
names(series) <- c("number","fraction")
series$number <- list()
series$number$id <- "chr_number"
series$number$name <- "Number of genes"
series$number$color <- cols$fill[1]
series$number$pointPlacement <- -0.2
series$number$data <- unname(chr)
series$fraction <- list()
series$fraction$id <- "chr_fraction"
series$fraction$name <- "Fraction of total genes"
series$fraction$color <- cols$fill[2]
series$fraction$pointPlacement <- 0.2
series$fraction$yAxis <- 1
series$fraction$data <- unname(per.chr)
what <- chr
}
else if (by=="biotype") {
bt <- table(as.character(filtered$biotype))
bt.all <- table(as.character(total$biotype))
barlab.bt <- as.character(bt)
per.bt <- round(bt/bt.all[names(bt)],3)
per.bt[per.bt>1] <- 1
series <- vector("list",2)
names(series) <- c("number","fraction")
series$number <- list()
series$number$id <- "bt_number"
series$number$name <- "Number of genes"
series$number$color <- cols$fill[1]
series$number$pointPlacement <- -0.2
series$number$data <- unname(bt)
series$fraction <- list()
series$fraction$id <- "bt_fraction"
series$fraction$name <- "Fraction of total genes"
series$fraction$color <- cols$fill[2]
series$fraction$pointPlacement <- 0.2
series$fraction$yAxis <- 1
series$fraction$data <- unname(per.bt)
what <- bt
}
json <- switch(jl,
highcharts = {
toJSON(
list(
chart=list(
type="column",
alignTicks=FALSE
),
title=list(
text=paste("Filtered genes per ",by,sep="")
),
legend=list(
enabled=TRUE,
itemHoverStyle=list(
color="#B40000"
)
),
tooltip=list(
shared=TRUE
),
xAxis=list(
categories=names(what),
title=list(
text=paste(toupper(substr(by,1,1)),substr(by,2,
nchar(by)),sep=""),
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontWeight="bold"
)
)
),
yAxis=list(
list(
lineColor=cols$fill[1],
lineWidth=2,
min=0,
tickAmount=11,
title=list(
text="Number of genes",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
offset=10
),
list(
lineColor=cols$fill[2],
lineWidth=2,
min=0,
max=1,
tickAmount=11,
#tickInterval=0.1,
title=list(
text="Fraction of total genes",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
opposite=TRUE,
offset=10
)
),
plotOptions=list(
column=list(
grouping=FALSE,
shadow=FALSE,
groupPadding=0.2,
pointPadding=0.2,
tooltip=list(
headerFormat=paste(
'<span style="font-size:1.1em;',
'font-weight:bold">',
'{point.key}</span><br/>',sep=""
)
)
)
),
series=c(unname(series))
)
)
}
)
return(json)
}
#' Volcano JSON exporter for the metaseqR package
#'
#' Non-exportable JSON exporter for \code{\link{diagplot.volcano}}.
#'
#' @param obj A list holding volcano plot data. See \code{\link{diaplot.volcano}}.
#' @param jl JavaScript charting library to export. Currently only \code{"highcharts"}
#' supported.
#' @return A JSON string.
#' @author <NAME>
volcanoToJSON <- function(obj,jl=c("highcharts")) {
jl <- tolower(jl[1])
f <- obj$x
p <- obj$y
xlim <- obj$xlim
ylim <- obj$ylim
alt.names <- obj$altnames
pcut <- obj$pcut
fcut <- obj$fcut
up <- obj$user$up
down <- obj$user$down
ff <- obj$user$unf
pp <- obj$user$unp
alt.names.neutral <- obj$user$ualt
con <- obj$user$con
switch(jl,
highcharts = {
if (is.null(alt.names))
point.format=paste("<strong>Gene ID: </strong>{point.name}<br>",
"<strong>Fold change: </strong>{point.x}<br>",
"<strong>Significance: </strong>{point.y}",sep="")
else
point.format=paste("<strong>Gene name: </strong>",
"{point.alt_name}<br>",
"<strong>Gene ID: </strong>{point.name}<br>",
"<strong>Fold change: </strong>{point.x}<br>",
"<strong>Significance: </strong>{point.y}",sep="")
json <- toJSON(
list(
chart=list(
type="scatter",
zoomType="xy"
),
title=list(
text=paste("Volcano plot for",con)
),
xAxis=list(
title=list(
text="Fold change",
margin=20,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE,
gridLineWidth=1,
min=round(xlim[1],3),
max=round(xlim[2],3)
),
yAxis=list(
title=list(
useHTML=TRUE,
text="Significance (-log<sub>10</sub>(p-value))",
margin=25,
style=list(
color="#000000",
fontSize="1.2em"
)
),
labels=list(
style=list(
color="#000000",
fontSize="1.1em",
fontWeight="bold"
)
),
startOnTick=TRUE,
endOnTick=TRUE,
showLastLabel=TRUE,
gridLineWidth=1,
min=round(ylim[1]-2,3),
max=round(ylim[2],3)
),
#legend=list(
# layout="vertical",
# align="left",
# verticalAlign="top",
# floating=TRUE,
# backgroundColor="#FFFFFF",
# borderWidth=1
#),
plotOptions=list(
scatter=list(
allowPointSelect=TRUE,
marker=list(
radius=2,
states=list(
hover=list(
enabled=TRUE,
lineColor="#333333"
)
)
),
states=list(
hover=list(
marker=list(
enabled=FALSE
)
)
),
tooltip=list(
headerFormat=paste("<span style=",
"\"font-size:1.1em;color:{series.color};",
"font-weight:bold\">{series.name}<br>",
sep=""),
pointFormat=point.format
),
turboThreshold=50000
)
),
series=list(
list(
name="Up-regulated",
color="#EE0000",
marker=list(
symbol="circle"
),
data=make.highcharts.points(f[up],-log10(p[up]),
unname(alt.names[up]))
),
list(
name="Down-regulated",
marker=list(
symbol="circle"
),
color="#00CD00",
data=make.highcharts.points(f[down],-log10(p[down]),
unname(alt.names[down]))
),
list(
name="Unregulated",
marker=list(
symbol="circle"
),
color="#0000EE",
data=make.highcharts.points(ff,-log10(pp),
unname(alt.names.neutral))
),
list(
name="Downfold threshold",
color="#000000",
type="line",
dashStyle="dash",
marker=list(
enabled=FALSE
),
tooltip=list(
headerFormat=paste('<strong>{series.name}',
'</strong><br/>',sep=""),
pointFormat=paste('<strong>Threshold: ',
'</strong>{point.x}<br/>',sep="")
),
data=list(round(c(-fcut,ylim[1]-5),3),
round(c(-fcut,ylim[2]),3))
),
list(
name="Upfold threshold",
color="#000000",
type="line",
dashStyle="Dash",
marker=list(
enabled=FALSE
),
tooltip=list(
headerFormat=paste('<strong>{series.name}',
'</strong><br/>',sep=""),
pointFormat=paste('<strong>Threshold: ',
'</strong>{point.x}<br/>',sep="")
),
data=list(round(c(fcut,ylim[1]-5),3),
round(c(fcut,ylim[2]),3))
),
list(
name="Significance threshold",
color="#000000",
type="line",
dashStyle="DashDot",
marker=list(
enabled=FALSE
),
tooltip=list(
headerFormat=paste('<strong>{series.name}',
'</strong><br/>',sep=""),
pointFormat=paste('<strong>Threshold: ',
'</strong>{point.y}<br/>',sep="")
),
data=list(round(c(xlim[1],-log10(pcut)),3),
round(c(xlim[2],-log10(pcut)),3))
)
)
)
)
}
)
return(json)
}
unquote_js_fun <- function(js) {
if (is.list(js))
js <- lapply(js,unquote_js_fun)
else {
op <- gregexpr(pattern="function",js)
cl <- gregexpr(pattern="}\\\"",js)
if (length(op)>0) {
starts <- as.numeric(op[[1]])
for (i in 1:length(starts))
substr(js,starts[i]-1,starts[i]-1) <- " "
ends <- as.numeric(cl[[1]])
for (i in 1:length(starts))
substr(js,ends[i]+1,ends[i]+1) <- " "
}
}
return(js)
}
getGroupColorScheme <- function(group) {
cols <- getColorScheme(length(group))
classes <- as.factor(as.class.vector(group))
design <- as.numeric(classes)
return(lapply(cols,function(x,classes,design) {
return(x[1:length(levels(classes))][design])
},classes,design))
}
getColorScheme <- function(n=NULL) {
if (missing(n) || is.null(n))
return(getColors())
else {
cols <- getColors()
if (n > length(cols$fill)) {
cols$fill <- rep(cols$fill,length.out=n)
cols$border <- rep(cols$border,length.out=n)
cols$select <- rep(cols$select,length.out=n)
cols$trans <- rep(cols$trans,length.out=n)
}
return(cols)
}
}
getColors <- function() {
return(list(
fill=c("#CD0000","#00CD00","#0000EE","#FFD700","#87CEEB","#CD8500",
"#DEB887","#FF0000","#0000FF","#00FF00","#FFA500","#A9A9A9",
"#008B00","#313131","#FFC0CB","#A52A2A","#FF00FF","#9ACD32",
"#8B636C","#2E8B57","#008B8B"),
border=c("#850000","#006B00","#000085","#927C00","#156280","#5A3A00",
"#8B7457","#935E18","#000080","#008500","#603E00","#454545",
"#073E07","#000000","#896067","#691111","#7C007C","#3A4D14",
"#5B1726","#0C2517","#062A2A"),
selected=c("#FF0000","#00FF00","#0066FF","#FFD700","#FFEB77","#FFB428",
"#FFD9A5","#FF326D","#0089FF","#B3FF00","#FFC352","#D9D9D9",
"#00EC00","#8E8E8E","#FFDAE0","#F94444","#FF87FF","#C2FF45",
"#EA889D","#4EE590","#00DADA"),
trans=c("rgba(205,0,0,0.6)","rgba(0,205,0,0.6)","rgba(0,0,238,0.6)",
"rgba(255,215,0,0.6)","rgba(135,206,235,0.6)","rgba(205,133,0,0.6)",
"rgba(222,184,135,0.6)","rbga(255,0,0,0.5)","rgba(0,0,255,0.5)",
"rgba(0,255,0,0.5)","rgba(255,165,0,0.6)","rgba(169,169,169,0.5)",
"rgba(0,139,0,0.6)","rgba(49,49,49,0.6)","rgba(255,192,203,0.5)",
"rgba(165,42,42,0.6)","rgba(255,0,255,0.6)","rgba(154,205,50,0.6)",
"rgba(139,99,108,0.6)","rgba(46,139,87,0.6)","rgba(0,139,139,0.6)")
))
}
<file_sep>/man/get.ucsc.query.Rd
\name{get.ucsc.query}
\alias{get.ucsc.query}
\title{Return queries for the UCSC Genome Browser database,
according to organism and source}
\usage{
get.ucsc.query(org, type, refdb="ucsc")
}
\arguments{
\item{org}{one of metaseqR supported organisms.}
\item{type}{either \code{"gene"} or \code{"exon"}.}
\item{refdb}{one of \code{"ucsc"} or \code{"refseq"}
to use the UCSC or RefSeq annotation sources
respectively.}
}
\value{
A valid SQL query.
}
\description{
Returns an SQL query to be used with a connection to
the UCSC Genome Browser database and fetch metaseqR
supported organism annotations. This query is constructed
based on the data source and data type to be returned.
}
\examples{
\donttest{
db.query <- get.ucsc.query("hg18","gene","ucsc")
}
}
\author{
<NAME>
}
<file_sep>/man/make.contrast.list.Rd
\name{make.contrast.list}
\alias{make.contrast.list}
\title{Create contrast lists from contrast vectors}
\usage{
make.contrast.list(contrast, sample.list)
}
\arguments{
\item{contrast}{a vector of contrasts in the form
"ConditionA_vs_ConditionB" or "ConditionA_
vs_ConditionB_vs_ConditionC_vs_...". In case of Control
vs Treatment designs, the Control condition should ALWAYS
be the first.}
\item{sample.list}{the list of samples in the experiment.
See also the main help page of \code{\link{metaseqr}}.}
}
\value{
A named list whose names are the contrasts and its
members are named vectors, where the names are the sample
names and the actual vector members are the condition
names. See the example.
}
\description{
Returns a list, properly structured to be used within the
\code{stat.*} functions of the metaseqr package. See the
main documentation for the structure of this list and the
example below. This function is mostly for internal use,
as the \code{stat.*} functions can be supplied directly
with the contrasts vector which is one of the main
\code{\link{metaseqr}} arguments.
}
\examples{
sample.list <- list(Control=c("C1","C2"),TreatmentA=c("TA1","TA2"),TreatmentB=c("TB1","TB2"))
contrast <- c("Control_vs_TreatmentA","Control_vs_TreatmentA_vs_TreatmentB")
cl <- make.contrast.list(contrast,sample.list)
}
\author{
<NAME>
}
<file_sep>/man/cddat.Rd
\name{cddat}
\alias{cddat}
\title{Old functions from NOISeq}
\usage{
cddat(input)
}
\arguments{
\item{input}{input to cddat.}
}
\value{
a list with data to plot.
}
\description{
Old functions from NOISeq to create the
\code{"readnoise"} plots. Internal use only.
}
\note{
Adopted from an older version of NOISeq package (author:
<NAME>).
}
\author{
<NAME>
}
<file_sep>/R/zzz.R
# Package environment to store a couple of variables that must be global
meta.env <- new.env(parent=emptyenv())
assign("VERBOSE",NULL,envir=meta.env)
assign("LOGGER",NULL,envir=meta.env)
#.onLoad <- function(...) {
# packageStartupMessage("\n\nWelcome to metaseqR 1.5.3. Please do not ",
# "include \"bayseq\" in the statistics\nargument in metaseqr calls. ",
# "There is an incompatibility with the latest\nversion of baySeq which ",
# "we are fixing right now...\n\n")
#}
<file_sep>/man/make.html.cells.Rd
\name{make.html.cells}
\alias{make.html.cells}
\title{HTML report helper}
\usage{
make.html.cells(mat, type = "numeric", digits = 3)
}
\arguments{
\item{mat}{the data matrix (numeric or character)}
\item{type}{the type of data in the matrix
(\code{"numeric"} or \code{"character"})}
\item{digits}{the number of digits on the right of the
decimal points to pass to \code{\link{formatC}}. It has
meaning when \code{type="numeric"}.}
}
\value{
A character matrix with html formatted cells.
}
\description{
Returns a character matrix with html formatted table
cells. Essentially, it converts the input data to text
and places them in a <td></td> tag set. Internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
the.cells <- make.html.cells(data.matrix)
}
}
\author{
<NAME>
}
<file_sep>/man/make.venn.colorscheme.Rd
\name{make.venn.colorscheme}
\alias{make.venn.colorscheme}
\title{Helper for Venn diagrams}
\usage{
make.venn.colorscheme(n)
}
\arguments{
\item{n}{the number of the sets used for the Venn
diagram.}
}
\value{
A list with colors for fill and font.
}
\description{
This function returns a list of colorschemes accroding to
the number of sets. Internal use.
}
\examples{
\donttest{
sets <- c("apple","pear","banana")
cs <- make.venn.colorscheme(length(sets))
}
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.main.R
#' The main metaseqr pipeline
#'
#' This function is the main metaseqr workhorse and implements the main metaseqr
#' workflow which performs data read, filtering, normalization and statistical
#' selection, creates diagnostic plots and exports the results and a report if
#' requested. The metaseqr function is responsible for assembling all the steps
#' of the metaseqr pipeline which i) reads the input gene or exon read count table
#' ii) performs prelimininary filtering of data by removing chrM and other
#' non-essential information for a typical differential gene expression analysis
#' as well as a preliminary expression filtering based on the exon counts, if an
#' exon read count file is provided. iii) performs data normalization with one of
#' currently widely used algorithms, including EDASeq (Risso et al., 2011), DESeq re
#' (<NAME>, 2010), edgeR (Robinson et al., 2010), NOISeq (Tarazona et
#' al., 2012) or no normalization iv) performs a second stage of filtering based
#' on the normalized gene expression according to several gene filters v) performs
#' statistical testing with one or more of currently widely used algorithms,
#' including DESeq (<NAME>, 2010), edgeR (Robinson et al., 2010), NOISeq
#' (Tarazona et al., 2012), limma (Smyth et al., 2005) for RNA-Seq data, baySeq
#' (Hardcastle et al., 2012) vi) in the case of multiple statistical testing
#' algorithms, performs meta-analysis using one of five available methods (see the
#' meta.p argument) vii) exports the resulting differentially expressed gene list
#' in text tab-delimited format viii) creates a set of diagnostic plots either
#' available in the aforementioned packages or metaseqr specific ones and ix)
#' creates a comprehensive HTML report which summarizes the run information, the
#' results and the diagnostic plots. Certain diagnostic plots (e.g. the volcano
#' plot) can be interactive with the use of the external Highcharts
#' (http://www.highcharts.com) JavaScript library for interactive graphs. Although
#' the inputs to the metaseqr workflow are many, in practice, setting only very
#' few of them and accepting the defaults as the rest can result in quite
#' comprehensible results for mainstream organisms like mouse, human, fruitfly and
#' rat.
#'
#' @aliases metaseqr.main
#' @param counts a text tab-delimited file containing gene or exon counts in one
#' of the following formats: i) the first column contains unique gene or exon
#' identifiers and the rest of the columns contain the read counts for each sample.
#' Thus the first cell of each row is a gene or exon accession and the rest are
#' integers representing the counts for that accession. In that case, the
#' \code{annotation} parameter should strictly be \code{"download"} or an external
#' file in proper format. ii) The first n columns should contain gene or exon
#' annotation elements like chromosomal locations, gene accessions, exon accessions,
#' GC content etc. In that case, the \code{annotation} parameter can also be
#' \code{"embedded"}. The ideal embedded annotation contains 8 columns, chromosome,
#' gene or exon start, gene or exon end, gene or exon accession, GC-content
#' (fraction or percentage), strand, HUGO gene symbol and gene biotype (e.g.
#' "protein_coding" or "ncRNA"). When the \code{annotation} parameter is "embedded",
#' certain of these features are mandatory (co-ordinates and accessions). If they
#' are not present, the pipeline will not run. If additional elements are not
#' present (e.g. GC content or biotypes), certain features of metaseqr will not
#' be available. For example, EDASeq normalization will not be performed based on
#' a GC content covariate but based on gene length which is not what the authors
#' of EDASeq suggest. If biotypes are not present, a lot of diagnostic plots will
#' not be available. If the HUGO gene symbols are missing, the final annotation
#' will contain only gene accessions and thus be less comprehensible. Generally,
#' it's best to set the \code{annotation} parameter to \code{"download"} to ensure
#' the most comprehensible results. Counts can be a data frame satisfying the
#' above conditions. It is a data frame by default when \code{read2count} is used.
#' counts can also be an .RData file (output of \code{\link{save}} function
#' which contains static input elements (list containing the gene model (exon
#' counts for each gene constructed by the \code{\link{construct.gene.model}}
#' function, gene and exon annotation to avoid re-downloading and/or gene counts
#' depending on \code{count.type}). This kind of input facilitates the
#' re-analysis of the same experiment, using different filtering, normalization
#' and statistical algorithms. Finally, counts can be a list representing the
#' gene model (exon counts for each gene) constructed by the
#' \code{\link{construct.gene.model}} function (provided for backwards
#' compatibility). This .RData file can be generated by setting
#' \code{save.gene.model=TRUE} when performing data analysis for the first time.
#' @param sample.list a list containing condition names and the samples under each
#' condition. It should have the format \code{sample.list <-}
#' \code{list(ConditionA=c("Sample_A1",} \code{"Sample_A2", "Sample_A3"),}
#' \code{ConditionB=c("Sample_B1", "Sample_B2"),}
#' \code{ConditionC=c("Sample_C1", "Sample_C2"))}. The names of the samples in list
#' members MUST match the column names containing the read counts in the counts
#' file. If they do not match, the pipeline will either crash or at best, ignore
#' several of your samples. Alternative, \code{sample.list} can be a small
#' tab-delimited file structured as follows: the first line of the external tab
#' delimited file should contain column names (names are not important). The first
#' column MUST contain UNIQUE sample names and the second column MUST contain the
#' biological condition where each of the samples in the first column should belong
#' to. In this case, the function \code{\link{make.sample.list}} is used. If the
#' \code{counts} argument is missing, the \code{sample.list} argument MUST be a
#' targets text tab-delimited file which contains the sample names, the BAM/BED
#' file names and the biological conditions/groups for each sample/file. The file
#' should be text tab-delimited and structured as follows: the first line of the
#' external tab delimited file should contain column names (names are not important).
#' The first column MUST contain UNIQUE sample names. The second column MUST contain
#' the raw BAM/BED files WITH their full path. Alternatively, the \code{path}
#' argument should be provided (see below). The third column MUST contain the
#' biological condition where each of the samples in the first column should belong
#' to.
#' @param exclude.list a list of samples to exclude, in the same (list) format
#' as \code{sample.list} above.
#' @param path an optional path where all the BED/BAM files are placed, to be
#' prepended to the BAM/BED file names in the targets file. If not given and if
#' the files in the second column of the targets file do not contain a path to a
#' directory, the current directory is assumed to be the BAM/BED file container.
#' @param file.type the type of raw input files. It can be \code{"auto"} for
#' auto-guessing, \code{"bed"} for BED files, \code{"sam"} for SAM files or
#' \code{"bam"} for BAM files.
#' @param contrast a character vector of contrasts to be tested in the statistical
#' testing step(s) of the metaseqr pipeline. Each element of the should STRICTLY
#' have the format "ConditionA_vs_ConditionB_vs_...". A valid example based on the
#' \code{sample.list} above is \code{contrast <- c("ConditionA_vs_ConditionB",}
#' \code{"ConditionA_vs_ConditionC",} \code{"ConditionA_vs_ConditionB_vs_ConditionC")}.
#' The first element of pairwise contrasts (e.g. "ConditionA" above) MUST be the
#' control condition or any reference that ConditionB is checked against. metaseqr
#' uses this convention to properly calculate fold changes. If it's NULL, a contrast
#' between the first two members of the \code{sample.list} will be auto-generated.
#' @param libsize.list an optional named list where names represent samples (MUST
#' be the same as the samples in \code{sample.list}) and members are the library
#' sizes (the sequencing depth) for each sample. For example
#' \code{libsize.list <- list(Sample_A1=32456913,} \code{Sample_A2=4346818)}.
#' @param id.col an integer denoting the column number in the file (or data frame)
#' provided with the counts argument, where the unique gene or exon accessions are.
#' Default to \code{4} which is the standard feature name column in a BED file.
#' @param gc.col an integer denoting the column number in the file (or data frame)
#' provided with the \code{counts} argument, where each gene's GC content is given.
#' If not provided, GC content normalization provided by EDASeq will not be available.
#' @param name.col an integer denoting the column number in the file (or data frame)
#' provided with the counts argument, where the HUGO gene symbols are given. If
#' not provided, it will not be available when reporting results. In addition, the
#' \code{"known"} gene filter will not be available.
#' @param bt.col an integer denoting the column number in the file (or data frame)
#' provided with the counts argument, where the gene biotypes are given. If not
#' provided, the \code{"biodetection"}, \code{"countsbio"}, \code{"saturation"},
#' \code{"filtered"} and \code{"biodist"} plots will not be available.
#' @param annotation instructs metaseqr where to find the annotation for the given
#' counts file. It can be one of i) \code{"download"} (default) for automatic
#' downloading of the annotation for the organism specified by the org parameter
#' (using biomaRt), ii) \code{"embedded"} if the annotation elements are embedded
#' in the read counts file or iv) a file specified by the user which should be as
#' similar as possible to the \code{"download"} case, in terms of column structure.
#' @param org the supported organisms by metaseqr. These can be, for human genomes
#' \code{"hg18"}, \code{"hg19"} or \code{"hg38"} for mouse genomes \code{"mm9"},
#' \code{"mm10"}, for rat genomes \code{"rn5"}, for drosophila genome \code{"dm6"},
#' for zebrafish genome \code{"danrer7"}, for chimpanzee genome \code{"pantro4"},
#' for pig genome \code{"susScr3"} and for Arabidopsis thaliana genome \code{"tair10"}.
#' Finally, \code{"custom"} will instruct metaseqR to completely ignore the
#' \code{org} argument and depend solely on annotation file provided by the user.
#' @param refdb the reference annotation repository from which to retrieve annotation
#' elements to use with metaseqr. It can be one of \code{"ensembl"} (default),
#' \code{"ucsc"} or \code{"refseq"}.
#' @param count.type the type of reads inside the counts file. It can be one of
#' \code{"gene"} or \code{"exon"}. This is a very important and mandatory parameter
#' as it defines the course of the workflow.
#' @param exon.filters a named list whose names are the names of the supported
#' exon filters and its members the filter parameters. See section "Exon filters"
#' below for details.
#' @param gene.filters a named list whose names are the names of the supported
#' gene filters and its members the filter parameters. See section "Gene filters"
#' below for details.
#' @param when.apply.filter a character string determining when to apply the exon
#' and/or gene filters, relative to normalization. It can be \code{"prenorm"} to
#' apply apply the filters and exclude genes from further processing before
#' normalization, or \code{"postnorm"} to apply the filters after normalization
#' (default). In the case of \code{when.apply.filter="prenorm"}, a first
#' normalization round is applied to a copy of the gene counts matrix in order to
#' derive the proper normalized values that will constitute the several
#' expression-based filtering cutoffs.
#' @param normalization the normalization algorithm to be applied on the count
#' data. It can be one of \code{"edaseq"} (default) for EDASeq normalization,
#' \code{"deseq"} for the normalization algorithm (individual options specified
#' by the \code{norm.args} argument) in the DESeq package, \code{"edger"} for the
#' normalization algorithms present in the edgeR package (specified by the
#' \code{norm.args} argument), \code{"noiseq"} for the normalization algorithms
#' present in the NOISeq package (specified by the \code{norm.args} argument),
#' \code{"nbpseq"} for the normalization algorithms present in the NBPSeq package
#' (specified by the \code{norm.args} argument) or \code{"none"} to not normalize
#' the data (highly unrecommended). It can also be \code{"each"} where in this
#' case, the normalization applied will be specific to each statistical test used
#' (i.e. the normalization method bundled with each package and used in its
#' examples and documentation).
#' @param norm.args a named list whose names are the names of the normalization
#' algorithm parameters and its members parameter values. See section "Normalization
#' parameters" below for details. Leave \code{NULL} for the defaults of
#' \code{normalization}. If \code{normalization="each"}, it must be a named list
#' of lists, where each sub-list contains normalization parameters specific to
#' each statistical test to be used.
#' @param statistics one or more statistical analyses to be performed by the
#' metaseqr pipeline.It can be one or more of \code{"deseq"} (default) to conduct
#' statistical test(s) implemented in the DESeq package, \code{"edger"} to conduct
#' statistical test(s) implemented in the edgeR package, \code{"limma"} to conduct
#' the RNA-Seq version of statistical test(s) implemented in the limma package,
#' \code{"noiseq"} to conduct statistical test(s) implemented in the NOISeq package,
#' \code{"bayseq"} to conduct statistical test(s) implemented in the baySeq package
#' and \code{"nbpseq"} to conduct statistical test(s) implemented in the NBPSeq
#' package. In any case individual algorithm parameters are controlled by the
#' contents of the \code{stat.args} list.
#' @param stat.args a named list whose names are the names of the statistical
#' algorithms used in the pipeline. Each member is another named list whose names
#' are the algorithm parameters and its members are the parameter values. See
#' section "Statistics parameters" below for details. Leave \code{NULL} for the
#' defaults of \code{statistics}.
#' @param adjust.method the multiple testing p-value adjustment method. It can be
#' one of \code{\link{p.adjust.methods}} or \code{"qvalue"} from the qvalue
#' Bioconductor package. Defaults to \code{"BH"} for Benjamini-Hochberg correction.
#' @param meta.p the meta-analysis method to combine p-values from multiple
#' statistical tests \strong{(experimental! see also the second note below,
#' regarding meta-analysis)}. It can be one of \code{"simes"} (default),
#' \code{"bonferroni"}, \code{"minp"}, \code{"maxp"}, \code{"weight"}, \code{"pandora"},
#' \code{"dperm.min"}, \code{"dperm.max"}, \code{"dperm.weight"}, \code{"fisher"},
#' \code{"fperm"}, \code{"whitlock"} or\code{"none"}. For the \code{"fisher"} and
#' \code{"fperm"} methods, see the documentation of the R package MADAM. For the
#' \code{"whitlock"} method, see the documentation of the survcomp Bioconductor
#' package. With the \code{"maxp"} option, the final p-value is the maximum p-value
#' out of those returned by each statistical test. This is equivalent to an
#' "intersection" of the results derived from each algorithm so as to have a final
#' list with the common genes returned by all statistical tests. Similarly, when
#' \code{meta.p="minp"}, is equivalent to a "union" of the results derived from
#' each algorithm so as to have a final list with all the genes returned by all
#' statistical tests. The latter can be used as a very lose statistical threshold
#' to aggregate results from all methods regardless of their False Positive Rate.
#' With the \code{"simes"} option, the method proposed by Simes (<NAME>., 1986)
#' is used. With the \code{"dperm.min"}, \code{"dperm.max"}, \code{"dperm.weight"}
#' options, a permutation procedure is initialed, where \code{nperm} permutations
#' are performed across the samples of the normalized counts matrix, producing
#' \code{nperm} permuted instances of the initital dataset. Then, all the chosen
#' statistical tests are re-executed for each permutation. The final p-value is
#' the number of times that the p-value of the permuted datasets is smaller than
#' the original dataset. The p-value of the original dataset is created based on
#' the choice of one of \code{dperm.min}, \code{dperm.max} or \code{dperm.weight}
#' options. In case of \code{dperm.min}, the intial p-value vector is consists of
#' the minimum p-value resulted from the applied statistical tests for each gene.
#' The maximum p-value is used with the \code{dperm.max} option. With the
#' \code{dperm.weight} option, the \code{weight} weighting vector for each
#' statistical test is used to weight each p-value according to the power of
#' statistical tests (some might work better for a specific dataset). Be careful
#' as the permutation procedure usually requires a lot of time. However, it should
#' be the most accurate. This method will NOT work when there are no replicated
#' samples across biological conditions. In that case, use \code{meta.p="simes"}
#' instead. Finally, there are the \code{"minp"}, \code{"maxp"} and \code{"weight"}
#' options which correspond to the latter three methods but without permutations.
#' Generally, permutations would be accurate to use when the experiment includes
#' >5 samples per condition (or even better 7-10) which is rather rare in RNA-Seq
#' experiments. Finally, \code{"pandora"} is the same as \code{"weight"} and is
#' added to be in accordance with the metaseqR paper.
#' @param weight a vector of weights with the same length as the \code{statistics}
#' vector containing a weight for each statistical test. It should sum to 1.
#' \strong{Use with caution with the} \code{dperm.weight} \strong{parameter!
#' Theoretical background is not yet} \strong{solid and only experience shows
#' improved results!}
#' @param nperm the number of permutations performed to derive the meta p-value
#' when \code{meta.p="fperm"} or \code{meta.p="dperm"}. It defaults to 10000.
#' @param reprod create reproducible permutations when \code{meta.p="dperm.min"},
#' \code{meta.p="dperm.max"} or \code{meta.p="dperm.weight"}. Ideally one would
#' want to create the same set of indices for a given dataset so as to create
#' reproducible p-values. If \code{reprod=TRUE}, a fixed seed is used by
#' \code{meta.perm} for all the datasets analyzed with \code{metaseqr}. If
#' \code{reprod=FALSE}, then the p-values will not be reproducible, although
#' statistical significance is not expected to change for a large number of
#' resamplings. Finally, \code{reprod} can be a numeric vector of seeds with the
#' same length as \code{nperm} so that the user can supply his/her own seeds.
#' @param pcut a p-value cutoff for exporting differentially genes, default is
#' to export all the non-filtered genes.
#' @param log.offset an offset to be added to values during logarithmic
#' transformations in order to avoid Infinity (default is \code{1}).
#' @param preset an analysis strictness preset. \code{preset} can be one of
#' \code{"all.basic"}, \code{"all.normal"}, \code{"all.full"}, \code{"medium.basic"},
#' \code{"medium.normal"}, \code{"medium.full"}, \code{"strict.basic"},
#' \code{"strict.normal"} or \code{"strict.full"}, each of which control the
#' strictness of the analysis and the amount of data to be exported. For an
#' explanation of the presets, see the section "Presets" below.
#' @param qc.plots a set of diagnostic plots to show/create. It can be one or more
#' of \code{"mds"}, \code{"biodetection"}, \code{"rnacomp"}, \code{"countsbio"},
#' \code{"saturation"}, \code{"readnoise"}, \code{"filtered"}, \code{"boxplot"},
#' \code{"gcbias"}, \code{"lengthbias"}, \code{"meandiff"}, \code{"meanvar"},
#' \code{"deheatmap"}, \code{"volcano"}, \code{"biodist"}, \code{"venn"}. The
#' \code{"mds"} stands for Mutlti-Dimensional Scaling and it creates a PCA-like
#' plot but using the MDS dimensionality reduction instead. It has been succesfully
#' used for NGS data (e.g. see the package htSeqTools) and it shows how well
#' samples from the same condition cluster together. For \code{"biodetection"},
#' \code{"countsbio"}, \code{"saturation"}, \code{"rnacomp"}, \code{"readnoise"},
#' \code{"biodist"} see the vignette of NOISeq package. The \code{"saturation"}
#' case has been rewritten in order to display more samples in a more simple
#' way. See the help page of \code{\link{diagplot.noiseq.saturation}}. In addition,
#' the \code{"readnoise"} plots represent an older version or the RNA composition
#' plot included in older versions of NOISeq. For \code{"gcbias"},
#' \code{"lengthbias"}, \code{"meandiff"}, \code{"meanvar"} see the vignette of
#' EDASeq package. \code{"lenghtbias"} is similar to \code{"gcbias"} but using the
#' gene length instead of the GC content as covariate. The \code{"boxplot"} option
#' draws boxplots of log2 transformed gene counts. The \code{"filtered"} option
#' draws a 4-panel figure with the filtered genes per chromosome and per biotype,
#' as absolute numbers and as fractions of the genome. See also the help page of
#' \code{\link{diagplot.filtered}}. The \code{"deheatmap"} option performs
#' hierarchical clustering and draws a heatmap of differentially expressed genes.
#' In the context of diagnostic plots, it's useful to see if samples from the
#' same groups cluster together after statistical testing. The \code{"volcano"}
#' option draws a volcano plot for each contrast and if a report is requested, an
#' interactive volcano plot is presented in the HTML report. The \code{"venn"}
#' option will draw an up to 5-way Venn diagram depicting the common and specific
#' to each statistical algorithm genes and for each contrast, when meta-analysis
#' is performed. The \code{"correl"} option creates two correlation graphs: the
#' first one is a correlation heatmap (a correlation matrix which depicts all the
#' pairwise correlations between each pair of samples in the counts matrix is
#' drawn as a clustered heatmap) and the second one is a correlogram plot, which
#' summarizes the correlation matrix in the form of ellipses (for an explanation
#' please see the vignette/documentation of the R package corrplot. Set
#' \code{qc.plots=NULL} if you don't want any diagnostic plots created.
#' @param fig.format the format of the output diagnostic plots. It can be one or
#' more of \code{"png"}, \code{"jpg"}, \code{"tiff"}, \code{"bmp"}, \code{"pdf"},
#' \code{"ps"}. The native format \code{"x11"} (for direct display) is not provided
#' as an option as it may not render the proper display of some diagnostic plots
#' in some devices.
#' @param out.list a logical controlling whether to export a list with the results
#' in the current running environment.
#' @param export.where an output directory for the project results (report, lists,
#' diagnostic plots etc.).
#' @param export.what the content of the final lists. It can be one or more of
#' \code{"annotation"}, to bind the annoation elements for each gene, \code{"p.value"},
#' to bind the p-values of each method, \code{"adj.p.value"}, to bind the multiple
#' testing adjusted p-values, \code{"meta.p.value"}, to bind the combined p-value
#' from the meta-analysis, \code{"adj.meta.p.value"}, to bind the corrected combined
#' p-value from the meta-analysis, \code{"fold.change"}, to bind the fold changes
#' of each requested contrast, \code{"stats"}, to bind several statistics calclulated
#' on raw and normalized counts (see the \code{export.stats} argument), \code{"counts"},
#' to bind the raw and normalized counts for each sample.
#' @param export.scale export values from one or more transformations applied to
#' the data. It can be one or more of \code{"natural"}, \code{"log2"}, \code{"log10"},
#' \code{"vst"} (Variance Stabilizing Transormation, see the documentation of DESeq
#' package) and \code{"rpgm"} which is ratio of mapped reads per gene model
#' (either the gene length or the sum of exon lengths, depending on \code{count.type}
#' argument). Note that this is not RPKM as reads are already normalized for
#' library size using one of the supported normalization methods. Also, \code{"rpgm"}
#' might be misleading when \code{normalization} is other than \code{"deseq"}.
#' @param export.values It can be one or more of \code{"raw"} to export raw values
#' (counts etc.) and \code{"normalized"} to export normalized counts.
#' @param export.stats calculate and export several statistics on raw and normalized
#' counts, condition-wise. It can be one or more of \code{"mean"}, \code{"median"},
#' \code{"sd"}, \code{"mad"}, \code{"cv"} for the Coefficient of Variation,
#' \code{"rcv"} for a robust version of CV where the median and the MAD are used
#' instead of the mean and the standard deviation.
#' @param export.counts.table exports also the calculated read counts table when
#' input is read from bam files and exports also the normalized count table in
#' all cases. Defaults to \code{FALSE}.
#' @param restrict.cores in case of parallel execution of several subfunctions,
#' the fraction of the available cores to use. In some cases if all available cores
#' are used (\code{restrict.cores=1} and the system does not have sufficient RAM,
#' the pipeline running machine might significantly slow down.
#' @param report a logical value controlling whether to produce a summary report
#' or not. Defaults to \code{TRUE}.
#' @param report.top a fraction of top statistically significant genes to append
#' to the HTML report. This helps in keeping the size of the report as small as
#' possible, as appending the total gene list might create a huge HTML file. Users
#' can always retrieve the whole gene lists from the report links. Defaults to
#' \code{0.1} (top 10% of statistically significant genes). Set to \code{NULL}
#' to report all the statistically significant genes.
#' @param report.template an HTML template to use for the report. Do not change
#' this unless you know what you are doing.
#' @param save.gene.model in case of exon analysis, a list with exon counts for
#' each gene will be saved to the file \code{export.where/data/gene_model.RData}.
#' This file can be used as input to metaseqR for exon count based analysis, in
#' order to avoid the time consuming step of assembling the counts for each gene
#' from its exons.
#' @param verbose print informative messages during execution? Defaults to
#' \code{TRUE}.
#' @param run.log write a log file of the \code{metaseqr} run using package log4r.
#' Defaults to \code{TRUE}. The filename will be auto-generated.
#' @param ... further arguments that may be passed to plotting functions, related
#' to \code{\link{par}}.
#' @return If \code{out.list} is \code{TRUE}, a named list whose length is the same
#' as the number of requested contrasts. Each list member is named according to
#' the corresponding contrast and contains a data frame of differentially expressed
#' genes for that contrast. The contents of the data frame are defined by the
#' \code{export.what, export.scale, export.stats, export.values} parameters. If
#' \code{report} is \code{TRUE}, the output list contains two main elements. The
#' first is described above (the analysis results) and the second contains the same
#' results but in HTML formatted tables.
#' @section Exon filters: The exon filters are a set of filters which are applied
#' after the gene models are assembled from the read counts of individual exons
#' and before the gene expression is summarized from the exons belonging to each
#' gene. These filters can be applied when the input read counts file contains exon
#' reads. It is not applicable when the input file already contains gene counts.
#' Such filters can be for example "accept genes where all the exons contain more
#' than x reads" or "accept genes where there is read presence in at least m/n
#' exons, n being the total exons of the gene". Such filters are NOT meant for
#' detecting differential splicing as also the whole metaseqr pipeline, thus they
#' should not be used in that context. The \code{exon.filters} argument is a named
#' list of filters, where the names are the filter names and the members are the
#' filter parameters (named lists with parameter name, parameter value). See the
#' usage of the \code{metaseqr} function for an example of how these lists are
#' structured. The supported exon filters in the current version are: i)
#' \code{min.active.exons} which implements a filter for demanding m out of n exons
#' of a gene to have a certain read presence with parameters \code{exons.per.gene},
#' \code{min.exons} and \code{frac}. The filter is described as follows: if a gene
#' has up to \code{exons.per.gene} exons, then read presence is required in at
#' least \code{min.exons} of them, else read presence is required in a \code{frac}
#' fraction of the total exons. With the default values, the filter instructs that
#' if a gene has up to 5 exons, read presence is required in at least 2, else in
#' at least 20% of the exons, in order to be accepted. More filters will be
#' implemented in future versions and users are encouraged to propose exon filter
#' ideas to the author by mail. See \code{metaseqr} usage for the defaults. Set
#' \code{exon.filters=NULL} to not apply any exon filtering.
#' @section Gene filters: The gene filters are a set of filters applied to gene
#' expression as this is manifested through the read presence on each gene and
#' are preferably applied after normalization. These filters can be applied both
#' when the input file or data frame contains exon read counts and gene read
#' counts. Such filter can be for example "accept all genes above a certain count
#' threshold" or "accept all genes with expression above the median of the
#' normalized counts distribution" or "accept all with length above a certain
#' threshold in kb" or "exclude the 'pseudogene' biotype from further analysis".
#' The supported gene filters in the current version, which have the same structure
#' as the exon filters (named list of lists with filter names, parameter names and
#' parameter arguments) are: i) \code{length} which implements a length filter
#' where genes are accepted for further analysis if they are above \code{length}
#' (its parameter) kb. ii) \code{avg.reads} which implements a filter where a gene
#' is accepted for further analysis if it has more average reads than the
#' \code{quantile} of the average count distribution per \code{average.per.bp} base
#' pairs. In summary, the reads of each gene are averaged per \code{average.per.bp}
#' based on each gene's length (in case of exons, input the "gene's length" is the
#' sum of the lengths of exons) and the \code{quantile} quantile of the average
#' counts distribution is calculated for each sample. Genes passing the filter
#' should have an average read count larger than the maximum of the vector of the
#' quantiles calculated above. iii) \code{expression} which implements a filter
#' based on the overall expression of a gene. The parameters of this filter are:
#' \code{median}, where genes below the median of the overall count distribution
#' are not accepted for further analysis (this filter has been used to distinguish
#' between "expressed" and "not expressed" genes in several cases, e.g. (Mokry et
#' al., NAR, 2011) with a logical as value, \code{mean} which is the same as
#' \code{median} but using the mean, \code{quantile} which is the same as the
#' previous two but using a specific quantile of the total counts distribution,
#' \code{known}, where in this case, a set of known not-expressed genes in the
#' system under investigation are used to estimate an expression cutoff. This can
#' be quite useful, as the genes are filtered based on a "true biological" cutoff
#' instead of a statistical cutoff. The value of this filter is a character vector
#' of HUGO gene symbols (MUST be contained in the annotation, thus it's better to
#' use \code{annotation="download"}) whose counts are used to build a "null"
#' expression distribution. The 90th quantile of this distribution is then the
#' expression cutoff. This filter can be combined with any other filter. Be careful
#' with gene names as they are case sensitive and must match exactly ("Pten" is
#' different from "PTEN"!). iv) \code{biotype} where in this case, genes with a
#' certain biotype (MUST be contained in the annotation, thus it's better to use
#' \code{annotation="download"}) are excluded from the analysis. This filter is
#' a named list of logical, where names are the biotypes in each genome and values
#' are \code{TRUE} or \code{FALSE}. If the biotype should be excluded, the value
#' should be \code{TRUE} else \code{FALSE}. See the result of
#' \code{get.defaults("biotype.filter","hg19")} for an example. Finally, in future
#' versions there will be support for user-defined filters in the form of a function.
#' @section Normalization parameters: The normalization parameters are passed again
#' as a named list where the names of the members are the normalization parameter
#' names and the values are the normalization parameter values. You should check
#' the documentation of the packages EDASeq, DESeq, edgeR, NOISeq and NBPSeq for
#' the parameter names and parameter values. There are a few exceptions in
#' parameter names: in case of \code{normalization="edaseq"} the only parameter
#' names are \code{within.which} and \code{between.which}, controlling the within
#' lane/sample and between lanes/samples normalization algorithm. In the case of
#' \code{normalization="nbpseq"}, there is one additional parameter called
#' \code{main.method} which can take the calues \code{"nbpseq"} or \code{"nbsmyth"}.
#' These values correspond to the two different workflows available in the NBPSeq
#' package. Please, consult the NBPSeq package documentation for further details.
#' For the rest of the algorithms, the parameter names are the same as the names
#' used in the respective packages. For examples, please use the
#' \code{\link{get.defaults}} function.
#' @section Statistics parameters: The statistics parameters as passed to statistical
#' algorithms in metaseqr, exactly with the same way as the normalization parameters
#' above. In this case, there is one more layer in list nesting. Thus, \code{stat.args}
#' is a named list whose names are the names the algorithms used (see the
#' \code{statistics} parameter). Each member is another named list,with parameters
#' to be used for each statistical algorithm. Again, the names of the member lists
#' are parameter names and the values of the member lists are parameter values.
#' You should check the documentations of DESeq, edgeR, NOISeq, baySeq, limma and
#' NBPSeq for these parameters. There are a few exceptions in parameter names:
#' In case of \code{statistics="edger"}, apart from the rest of the edgeR statistical
#' testing arguments, there is the argument \code{main.method} which can be either
#' \code{"classic"} or \code{"glm"}, defining whether the binomial test or GLMs
#' will be used for statistical testing. For examples, please use the'
#' \code{\link{get.defaults}} function. When \code{statistics="nbpseq"}, apart
#' from the rest arguments of the NBPSeq functions \code{estimate.disp} and
#' \code{estimate.dispersion}, there is the argument \code{main.method} which can
#' be \code{"nbpseq"} or \code{"nbsmyth"}. This argument determines the parameters
#' to be used by the \code{estimate.dispersion} function or by the
#' \code{estimate.disp} function to estimate RNA-Seq count dispersions. The
#' difference between the two is that they constitute different starting points
#' for the two workflows in the package NBPSeq. The first worklfow (with
#' \code{main.method="nbpseq"} and the \code{estimate.dispersion} function is
#' NBPSeq package specific, while the second (with \code{main.method="nbsmyth"}
#' and the \code{estimate.disp} function is similar to the workflow of the edgeR
#' package. For additional information regarding the statistical testing in
#' NBPSeq, please consult the documentation of the NBPSeq package.
#' \strong{Additinally, please note that there is currently a problem with the
#' NBPSeq package and the workflow that is specific to the NBPSeq package. The
#' problem has to do with function exporting as there are certain functions which
#' are not recognized from the package internally. For this reason and until it is
#' fixed, only the Smyth workflow will be available with the NBPSeq package (thus}
#' \code{stat.args$main.method="nbpseq"} \strong{will not be available)!}
#' @section Presets: The analysis presets are a set of keywords (only one can be
#' used) that predefine some of the parameters of the metaseqr pipeline. For the
#' time being they are quite simple and they control i) the strictness of
#' filtering and statistical thresholding with three basic levels ("all", "medium",
#' "strict") and ii) the data columns that are exported, again in three basic ways
#' ("basic", "normal", "full") controlling the amount of data to be exported. These
#' keywords can be combined with a dot in the middle (e.g. \code{"all.basic"} to
#' define an analysis preset. When using analysis presets, the following arguments
#' of metaseqr are overriden: \code{exon.filters}, \code{gene.filters}, \code{pcut},
#' \code{export.what}, \code{export.scale}, \code{export.values}, \code{exon.stats}.
#' If you want to explicitly control the above arguments, the \code{preset} argument
#' should be set to \code{NULL} (default). Following is a synopsis of the different
#' presets and the values of the arguments they moderate:
#' \itemize{
#' \item \code{"all.basic"}: use all genes (do not filter) and export all genes
#' and basic annotation and statistics elements. In this case, the above described
#' arguments become:
#' \itemize{
#' \item \code{exon.filters=NULL}
#' \item \code{gene.filters=NULL}
#' \item \code{pcut=1}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change")}
#' \item \code{export.scale=c("natural","log2")}
#' \item \code{export.values=c("normalized")}
#' \item \code{export.stats=c("mean")}
#' }
#' \item \code{"all.normal"}: use all genes (do not filter) and export all genes
#' and normal annotation and statistics elements. In
#' this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=NULL}
#' \item \code{gene.filters=NULL}
#' \item \code{pcut=1}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change","stats","counts")}
#' \item \code{export.scale=c("natural","log2")}
#' \item \code{export.values=c("normalized")}
#' \item \code{export.stats=c("mean","sd","cv")}
#' }
#' \item \code{"all.full"}: use all genes (do not filter) and export all genes and
#' all available annotation and statistics elements. In this case, the above
#' described arguments become:
#' \itemize{
#' \item \code{exon.filters=NULL}
#' \item \code{gene.filters=NULL}
#' \item \code{pcut=1}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change","stats","counts")}
#' \item \code{export.scale=c("natural","log2","log10","vst")}
#' \item \code{export.values=c("raw","normalized")}
#' \item \code{export.stats=c("mean","median","sd","mad","cv","rcv")}
#' }
#' \item \code{"medium.basic"}: apply a medium set of filters and and export
#' statistically significant genes and basic annotation and statistics elements.
#' In this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=list(min.active.exons=list(exons.per.gene=5,
#' min.exons=2,frac=1/5))}
#' \item \code{gene.filters=list(length=list(length=500),}
#' \code{avg.reads=list(average.per.bp=100,quantile=0.25),}
#' \code{expression=list(median=TRUE,mean=FALSE,quantile=NA,
#' known=NA,custom=NA),}
#' \code{biotype=get.defaults("biotype.filter",org[1]))}
#' \item \code{pcut=0.05}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change")}
#' \item \code{export.scale=c("natural","log2")}
#' \item \code{export.values=c("normalized")}
#' \item \code{export.stats=c("mean")}
#' }
#' \item \code{"medium.normal"}: apply a medium set of filters and and export
#' statistically significant genes and normal annotation and statistics elements.
#' In this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=list(min.active.exons=list(exons.per.gene=5,
#' min.exons=2,frac=1/5))}
#' \item \code{gene.filters=list(length=list(length=500),}
#' \code{avg.reads=list(average.per.bp=100,quantile=0.25),}
#' \code{expression=list(median=TRUE,mean=FALSE,quantile=NA,
#' known=NA,custom=NA),}
#' \code{biotype=get.defaults("biotype.filter",org[1]))}
#' \item \code{pcut=0.05}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change","stats","counts")}
#' \item \code{export.scale=c("natural","log2")}
#' \item \code{export.values=c("normalized")}
#' \item \code{export.stats=c("mean","sd","cv")}
#' }
# \item \code{"medium.full"}: apply a medium set of filters and and export
#' statistically significant genes and all available annotation and statistics
#' elements. In this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=list(min.active.exons=list(exons.per.gene=5,
#' min.exons=2,frac=1/5))}
#' \item \code{gene.filters=list(length=list(length=500),}
#' \code{avg.reads=list(average.per.bp=100,quantile=0.25),}
#' \code{expression=list(median=TRUE,mean=FALSE,quantile=NA,
#' known=NA,custom=NA),}
#' \code{biotype=get.defaults("biotype.filter",org[1]))}
#' \item \code{pcut=0.05}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change","stats","counts")}
#' \item \code{export.scale=c("natural","log2","log10","vst")}
#' \item \code{export.values=c("raw","normalized")}
#' \item \code{export.stats=c("mean","median","sd","mad","cv","rcv")}
#' }
#' \item \code{"strict.basic"}: apply a strict set of filters and and export
#' statistically significant genes and basic annotation and statistics elements.
#' In this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=list(min.active.exons=list(exons.per.gene=4,
#' min.exons=2,frac=1/4))}
#' \item \code{gene.filters=list(length=list(length=750),}
#' \code{avg.reads=list(average.per.bp=100,quantile=0.5),}
#' \code{expression=list(median=TRUE,mean=FALSE,quantile=NA,
#' known=NA,custom=NA),}
#' \code{biotype=get.defaults("biotype.filter",org[1]))}
#' \item \code{pcut=0.01}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change")}
#' \item \code{export.scale=c("natural","log2")}
#' \item \code{export.values=c("normalized")}
#' \item \code{export.stats=c("mean")}
#' }
#' \item \code{"strict.normal"}: apply a strict set of filters and and export
#' statistically significant genes and normal annotation and statistics elements.
#' In this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=list(min.active.exons=list(exons.per.gene=4,
#' min.exons=2,frac=1/4))}
#' \item \code{gene.filters=list(length=list(length=750),}
#' \code{avg.reads=list(average.per.bp=100,quantile=0.5),}
#' \code{expression=list(median=TRUE,mean=FALSE,quantile=NA,
#' known=NA,custom=NA),}
#' \code{biotype=get.defaults("biotype.filter",org[1]))}
#' \item \code{pcut=0.01}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change","stats","counts")}
#' \item \code{export.scale=c("natural","log2")}
#' \item \code{export.values=c("normalized")}
#' \item \code{export.stats=c("mean","sd","cv")}
#' }
# \item \code{"strict.full"}: apply a strict set of filters and and export
#' statistically significant genes and all available annotation and statistics
#' elements. In this case, the above described arguments become:
#' \itemize{
#' \item \code{exon.filters=list(min.active.exons=list(exons.per.gene=4,
#' min.exons=2,frac=1/4))}
#' \item \code{gene.filters=list(length=list(length=750),}
#' \code{avg.reads=list(average.per.bp=100,quantile=0.5),}
#' \code{expression=list(median=TRUE,mean=FALSE,quantile=NA,
#' known=NA,custom=NA),}
#' \code{biotype=get.defaults("biotype.filter",org[1]))}
#' \item \code{pcut=0.01}
#' \item \code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
#' \code{"adj.meta.p.value","fold.change","stats","counts")}
#' \item \code{export.scale=c("natural","log2","log10","vst")}
#' \item \code{export.values=c("raw","normalized")}
#' \item \code{export.stats=c("mean","median","sd","mad","cv","rcv")}
#' }
#' }
#' @note Please note that currently only gene and exon annotation from Ensembl
#' (http://www.ensembl.org) are supported. Thus, the unique gene or exon ids in
#' the counts files should correspond to valid Ensembl gene or exon accessions
#' for the organism of interest. If you are not sure about the source of your
#' counts file or do not know how to produce it, it's better to start from the
#' original BAM/BED files (metaseqr will use the \code{\link{read2count}} function
#' to create a counts file). Keep in mind that in the case of BED files, the
#' performance will be significantly lower and the overall running time significantly
#' higher as the R functions which are used to read BED files to proper structures
#' (GenomicRanges) and calculate the counts are quite slow. An alternative way is
#' maybe the easyRNASeq package (Delhomme et al, 2012). The \code{\link{read2count}}
#' function does not use this package but rather makes use of standard Bioconductor
#' functions to handle NGS data. If you wish to work outside R, you can work with
#' other popular read counters such as the HTSeq read counter
#' (http://www-huber.embl.de/users/anders/HTSeq/doc/overview.html). Please also
#' note that in the current version, the members of the \code{gene.filters} and
#' \code{exon.filters} lists are not checked for validity so be careful to supply
#' with correct names otherwise the pipeline will crash or at the best case scenario,
#' will ignore the filters. Also note that when you are supplying metaseqr with
#' an exon counts table, gene annotation is always downloaded so please be sure
#' to have a working internet connection. In addition to the above, if you have
#' a multiple core system, be very careful on how you are using the
#' \code{restrict.cores} argument and generally how many cores you are using with
#' scripts purely written in R. The analysis with exon read data can very easily
#' cause memory problems, so unless you have more than 64Gb of RAM available,
#' consider setting restrict.cores to something like 0.2 when working with exon
#' data. Finally, if you do not wish to download the same annotation again and
#' again when performing multiple analyses, it is best to use the
#' \code{\link{get.annotation}} function to download and store the resulting
#' data frames in local files and then use these files with the \code{annotation}
#' option.
#' @note Please note that the \strong{meta-analysis} feature provided by metaseqR
#' is currently experimental and does not satisfy the strict definition of
#' "meta-analysis", which is the combination of multiple similar datasets under
#' the same statistical methodology. Instead it is the use of mulitple statistical
#' tests applied to the same data so the results at this point are not guaranteed
#' and should be interpreted appropriately. We are working on a more solid
#' methodology for combining multiple statistical tests based on multiple testing
#' correction and Monte Carlo methods. For the Simes method, please consult also
#' "<NAME>. (1986). "An improved Bonferroni procedure for multiple tests of
#' significance". Biometrika 73 (3): 751–754."
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' # An example pipeline with exon counts
#' data("hg19.exon.data",package="metaseqR")
#' metaseqr(
#' counts=hg19.exon.counts,
#' sample.list=list(normal="normal",paracancerous="paracancerous",
#' cancerous="cancerous"),
#' contrast=c("normal_vs_paracancerous","normal_vs_cancerous",
#' "normal_vs_paracancerous_vs_cancerous"),
#' libsize.list=libsize.list.hg19,
#' id.col=4,
#' annotation="download",
#' org="hg19",
#' count.type="exon",
#' normalization="edaseq",
#' statistics="deseq",
#' pcut=0.05,
#' qc.plots=c("mds", "biodetection", "countsbio", "saturation", "rnacomp",
#' "boxplot", "gcbias", "lengthbias", "meandiff", "readnoise","meanvar",
#' "readnoise", "deheatmap", "volcano", "biodist", "filtered"),
#' fig.format=c("png","pdf"),
#' export.what=c("annotation","p.value","adj.p.value","fold.change","stats",
#' "counts"),
#' export.scale=c("natural","log2","log10","vst"),
#' export.values=c("raw","normalized"),
#' export.stats=c("mean","median","sd","mad","cv","rcv"),
#' restrict.cores=0.8,
#' gene.filters=list(
#' length=list(
#' length=500
#' ),
#' avg.reads=list(
#' average.per.bp=100,
#' quantile=0.25
#' ),
#' expression=list(
#' median=TRUE,
#' mean=FALSE
#' ),
#' biotype=get.defaults("biotype.filter","hg18")
#' )
#' )
#'
#' # An example pipeline with gene counts
#' data("mm9.gene.data",package="metaseqR")
#' result <- metaseqr(
#' counts=mm9.gene.counts,
#' sample.list=list(e14.5=c("e14.5_1","e14.5_2"),
#' adult_8_weeks=c("a8w_1","a8w_2")),
#' contrast=c("e14.5_vs_adult_8_weeks"),
#' libsize.list=libsize.list.mm9,
#' annotation="download",
#' org="mm9",
#' count.type="gene",
#' normalization="edger",
#' statistics=c("deseq","edger","noiseq"),
#' meta.p="fisher",
#' pcut=0.05,
#' fig.format=c("png","pdf"),
#' export.what=c("annotation","p.value","meta.p.value","adj.meta.p.value",
#' "fold.change"),
#' export.scale=c("natural","log2"),
#' export.values="normalized",
#' export.stats=c("mean","sd","cv"),
#' export.where=getwd(),
#' restrict.cores=0.8,
#' gene.filters=list(
#' length=list(
#' length=500
#' ),
#' avg.reads=list(
#' average.per.bp=100,
#' quantile=0.25
#' ),
#' expression=list(
#' median=TRUE,
#' mean=FALSE,
#' quantile=NA,
#' known=NA,
#' custom=NA
#' ),
#' biotype=get.defaults("biotype.filter","mm9")
#' ),
#' out.list=TRUE
#' )
#' head(result$data[["e14.5_vs_adult_8_weeks"]])
#' }
metaseqr <- function(
counts,
sample.list,
exclude.list=NULL,
file.type=c("auto","sam","bam","bed"),
path=NULL,
contrast=NULL,
libsize.list=NULL,
id.col=4,
gc.col=NA,
name.col=NA,
bt.col=NA,
annotation=c("download","embedded"),
gene.file=NULL,
org=c("hg18","hg19","hg38","mm9","mm10","rn5","rn6","dm3","dm6",
"danrer7","pantro4","susscr3","tair10","equcab2","custom"),
refdb=c("ensembl","ucsc","refseq"),
trans.level=c("gene","transcript","exon"),
count.type=c("gene","exon","utr"),
utr.flank=500,
exon.filters=list(
min.active.exons=list(
exons.per.gene=5,
min.exons=2,
frac=1/5
)
),
gene.filters=list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.25
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.defaults("biotype.filter",org[1]),
presence=list(
frac=0.25,
min.count=10,
per.condition=FALSE
)
),
when.apply.filter=c("postnorm","prenorm"),
normalization=c("deseq","edaseq","edger","noiseq","nbpseq","each","none"),
norm.args=NULL,
statistics=c("deseq","edger","noiseq","bayseq","limma","nbpseq"),
stat.args=NULL,
adjust.method=sort(c(p.adjust.methods,"qvalue")), # Brings BH first which is the default
meta.p=if (length(statistics)>1) c("simes","bonferroni","fisher",
"dperm.min","dperm.max","dperm.weight","fperm","whitlock","minp","maxp",
"weight","pandora","none") else "none",
weight=rep(1/length(statistics),length(statistics)),
nperm=10000,
reprod=TRUE,
pcut=NA, # A p-value cutoff for exporting DE genes, default is to export all
log.offset=1, # Logarithmic transformation offset to avoid +/-Inf (log2(a+offset/b+offset))
preset=NULL, # An analysis strictness preset
qc.plots=c(
"mds","biodetection","countsbio","saturation","readnoise","filtered",
"correl","pairwise", # Raw count data
"boxplot","gcbias","lengthbias","meandiff","meanvar","rnacomp", # Pre and post normalization
"deheatmap","volcano","biodist" # Post statistical testing
),
fig.format=c("png","jpg","tiff","bmp","pdf","ps"),
out.list=FALSE,
export.where=NA, # An output directory for the project
export.what=c("annotation","p.value","adj.p.value","meta.p.value",
"adj.meta.p.value","fold.change","stats","counts","flags"),
export.scale=c("natural","log2","log10","vst","rpgm"),
export.values=c("raw","normalized"),
export.stats=c("mean","median","sd","mad","cv","rcv"),
export.counts.table=FALSE,
restrict.cores=0.6,
report=TRUE,
report.top=0.1,
report.template="default",
save.gene.model=TRUE,
verbose=TRUE,
run.log=TRUE,
progress.fun=NULL,
...
)
{
# Check essential arguments
from.raw <- from.previous <- FALSE
if (missing(counts) && (missing(sample.list) || is.list(sample.list)))
stop("You must provide a file with genomic region (gene, exon, etc.) ",
"counts or an input targets file to create input from! If the ",
"counts file is missing, sample.list cannot be missing or it must ",
"be a targets file with at least three columns! See the ",
"read.targets function. counts may also be a gene model list ",
"(see the documentation)")
if (!missing(counts) && !missing(sample.list) && is.character(counts)
&& file.exists(counts) && length(grep(".RData$",counts))>0)
{
warning("When restoring a previous analysis, sample.list argument is ",
"not necessary! Ignoring...")
from.previous <- TRUE
tmp.env <- new.env()
disp("Restoring previous analysis from ",basename(counts))
load(counts,tmp.env)
sample.list <- tmp.env$sample.list
count.type <- tmp.env$count.type
}
if (!missing(counts) && missing(sample.list) && is.character(counts)
&& file.exists(counts) && length(grep(".RData$",counts))>0)
{ # Time to load previous analysis if existing
from.previous <- TRUE
tmp.env <- new.env()
message("Restoring previous analysis from ",basename(counts))
load(counts,tmp.env)
sample.list <- tmp.env$sample.list
count.type <- tmp.env$count.type
}
if (missing(sample.list) && !from.previous || (!is.list(sample.list) &&
!file.exists(sample.list)))
stop("You must provide a list with condition names and sample names ",
"(same as in the counts file) or an input file to create the ",
"sample list from!")
if (!missing(sample.list) && !is.list(sample.list)
&& file.exists(sample.list) && !missing(counts) && !from.previous)
sample.list <- make.sample.list(sample.list)
if (!missing(sample.list) && !is.list(sample.list)
&& file.exists(sample.list) && missing(counts))
{
counts <- NULL
the.list <- read.targets(sample.list,path=path)
sample.list <- the.list$samples
file.list <- the.list$files
if (tolower(file.type[1])=="auto")
file.type <- the.list$type
if (is.null(file.type))
stop(paste("The type of the input files could not be recognized!",
"Please specify (BAM or BED)..."))
from.raw <- TRUE
}
# Initialize environmental variables
HOME <- system.file(package="metaseqR")
TEMPLATE <- HOME
#if (!exists("HOME"))
# init.envar()
# Globalize the project's verbosity and logger
if (from.raw)
PROJECT.PATH <- make.project.path(export.where)
else
PROJECT.PATH <- make.project.path(export.where,counts)
assign("VERBOSE",verbose,envir=meta.env)
if (run.log)
logger <- create.logger(logfile=file.path(PROJECT.PATH$logs,
"metaseqr_run.log"),level=2,logformat="%d %c %m")
else
logger <- NULL
assign("LOGGER",logger,envir=meta.env)
# Check if there are any mispelled or invalid parameters and throw a warning
check.main.args(as.list(match.call()))
# Check if sample names match in file/df and list, otherwise meaningless to proceed
if (!from.raw && !from.previous)
{
if (!is.data.frame(counts) && !is.list(counts))
{
if (file.exists(counts))
{
aline <- read.delim(counts,nrows=5) # Read the 1st lines
aline <- colnames(aline)
}
else
stopwrap("The counts file you provided does not exist!")
}
else if (is.data.frame(counts))
aline <- colnames(counts)
else if (is.list(counts))
aline <- names(counts)
samples <- unlist(sample.list,use.names=FALSE)
if (length(which(!is.na(match(samples,aline)))) != length(samples))
stopwrap("The sample names provided in the counts file/list do ",
"not match with those of the sample.list!")
}
# If exclude list given, check that it's a subset of sample.list, otherwise
# just ignore exclude.list
if (!is.null(exclude.list) && !is.na(exclude.list))
{
sl <- unlist(sample.list)
el <- unlist(exclude.list)
if (length(intersect(sl,el)) != length(el))
{
warnwrap("Some samples in exclude.list do not match those in the ",
"initial sample.list! Ignoring...",now=TRUE)
exclude.list <- NULL
}
}
file.type <- tolower(file.type[1])
annotation <- tolower(annotation[1])
org <- tolower(org[1])
refdb <- tolower(refdb[1])
trans.level <- tolower(trans.level[1])
count.type <- tolower(count.type[1])
when.apply.filter <- tolower(when.apply.filter[1])
normalization <- tolower(normalization[1])
adjust.method <- adjust.method[1]
meta.p <- tolower(meta.p[1])
statistics <- tolower(statistics)
fig.format <- tolower(fig.format)
if (!is.null(qc.plots)) qc.plots <- tolower(qc.plots)
export.what <- tolower(export.what)
export.scale <- tolower(export.scale)
export.values <- tolower(export.values)
export.stats <- tolower(export.stats)
if (!is.null(preset)) preset <- tolower(preset[1])
if (from.raw)
counts.name <- "imported sam/bam/bed files"
else
{
if (!is.data.frame(counts) && !is.null(counts) && !is.list(counts))
{
check.file.args("counts",counts)
if (from.previous)
counts.name <- "previously stored project"
else
counts.name <- basename(counts)
}
else if (is.list(counts) && !is.data.frame(counts))
{
counts.name <- "previously stored gene model"
}
else
{
counts.name <- "imported custom data frame"
}
}
if (is.list(counts) && !is.data.frame(counts) && count.type=="exon"
&& annotation=="embedded")
{
warnwrap("annotation cannot be \"embedded\" when importing a stored ",
"gene model! Switching to \"download\"...")
annotation <- "download"
}
if (meta.p %in% c("weight","pandora","dperm.weight") &&
abs(1-sum(weight))>1e-5)
stopwrap("The weights given for p-value combination should sum to 1!")
check.text.args("file.type",file.type,c("auto","sam","bam","bed"),
multiarg=FALSE)
check.text.args("annotation",annotation,c("embedded","download"),
multiarg=FALSE)
check.text.args("org",org,c("hg18","hg19","hg38","mm9","mm10","rn5","rn6",
"dm3","dm6","danrer7","pantro4","susscr3","tair10","equcab2","custom"),
multiarg=FALSE)
check.text.args("refdb",refdb,c("ensembl","ucsc","refseq"),multiarg=FALSE)
check.text.args("trans.level",trans.level,c("gene","transcript","exon"),
multiarg=FALSE)
check.text.args("count.type",count.type,c("gene","exon","utr"),
multiarg=FALSE)
check.text.args("when.apply.filter",when.apply.filter,c("postnorm",
"prenorm"),multiarg=FALSE)
check.text.args("normalization",normalization,c("edaseq","deseq","edger",
"noiseq","nbpseq","each","none"),multiarg=FALSE)
check.text.args("statistics",statistics,c("deseq","edger","noiseq","bayseq",
"limma","nbpseq"),multiarg=TRUE)
check.text.args("meta.p",meta.p,c("simes","bonferroni","fisher","dperm.min",
"dperm.max","dperm.weight","fperm","whitlock","minp","maxp","weight",
"pandora","none"),multiarg=FALSE)
check.text.args("fig.format",fig.format,c("png","jpg","tiff","bmp","pdf",
"ps"),multiarg=TRUE)
check.text.args("export.what",export.what,c("annotation","p.value",
"adj.p.value","meta.p.value","adj.meta.p.value","fold.change","stats",
"counts","flags"),multiarg=TRUE)
check.text.args("export.scale",export.scale,c("natural","log2","log10",
"rpgm","vst"),multiarg=TRUE)
check.text.args("export.values",export.values,c("raw","normalized"),
multiarg=TRUE)
check.text.args("export.stats",export.stats,c("mean","median","sd","mad",
"cv","rcv"),multiarg=TRUE)
if (!is.null(preset))
check.text.args("preset",preset,c("all.basic","all.normal","all.full",
"medium.basic","medium.normal","medium.full","strict.basic",
"strict.normal","strict.full"),multiarg=FALSE)
if (!is.null(qc.plots))
check.text.args("qc.plots",qc.plots,c("mds","biodetection","countsbio",
"saturation","readnoise","correl","pairwise","boxplot","gcbias",
"lengthbias","meandiff","meanvar","rnacomp","deheatmap","volcano",
"biodist","filtered","venn"),multiarg=TRUE)
if (!is.na(restrict.cores)) check.num.args("restrict.cores",restrict.cores,
"numeric",c(0,1),"botheq")
if (!is.na(pcut)) check.num.args("pcut",pcut,"numeric",c(0,1),"botheq")
if (!is.na(gc.col)) check.num.args("gc.col",gc.col,"numeric",0,"gt")
if (!is.na(name.col)) check.num.args("name.col",name.col,"numeric",0,"gt")
if (!is.na(bt.col)) check.num.args("bt.col",bt.col,"numeric",0,"gt")
if (!is.na(log.offset)) check.num.args("log.offset",log.offset,"numeric",0,
"gt")
check.num.args("nperm",nperm,"numeric",10,"gt")
check.num.args("utr.flank",utr.flank,"numeric")
if (!is.null(report.top))
check.num.args("report.top",report.top,"numeric",c(0,1),"both")
if (!is.null(contrast))
{
check.contrast.format(contrast,sample.list)
contrast <- unique(contrast)
}
if ("bayseq" %in% statistics) libsize.list <- check.libsize(libsize.list,
sample.list)
# Check main functionality packages
check.packages(meta.p,qc.plots)
# Check if parallel processing is available
multic <- check.parallel(restrict.cores)
# Check the case of embedded annotation but not given gc and gene name columns
if (annotation=="embedded")
{
if (is.na(gc.col) && count.type=="gene")
stopwrap("The column that contains the gene GC content ",
"(\"gc.col\") argument is required when \"annotation\" is ",
"\"embedded\"!")
if (is.na(name.col) && !is.na(gene.filters$expression$known))
{
warnwrap("The column that contains the HUGO gene symbols ",
"(\"bt.col\") is missing with embedded annotation! Gene name ",
"expression filter will not be available...")
gene.filters$expression$known=NA
if ("volcano" %in% qc.plots)
warnwrap("The column that contains the HUGO gene symbols ",
"(\"bt.col\") is missing with embedded annotation! ",
"Interactive volcano plots will not contain gene names...")
}
if (is.na(bt.col) && count.type=="gene")
{
warnwrap("The column that contains the gene biotypes (\"bt.col\") ",
"is missing with embedded annotation! Biotype filters and ",
"certain plots will not be available...")
gene.filters$biotype=NULL
to.remove <- match(c("biodetection","countsbio","saturation",
"biodist","filtered"),qc.plots)
no.match <- which(is.na(to.remove))
if (length(no.match)>0)
to.remove <- to.remove[-no.match]
if (length(to.remove)>0)
qc.plots <- qc.plots[-to.remove]
}
}
if (org=="hg18" && (refdb %in% c("ucsc","refseq")))
{
warnwrap("Gene/exon biotypes cannot be retrieved when organism is ",
"\"hg18\" and annotation database is \"ucsc\" or \"refseq\"! ",
"Biotype filters and certain plots will not be available...")
gene.filters$biotype=NULL
to.remove <- match(c("biodetection","countsbio","saturation",
"biodist","filtered"),qc.plots)
no.match <- which(is.na(to.remove))
if (length(no.match)>0)
to.remove <- to.remove[-no.match]
if (length(to.remove)>0)
qc.plots <- qc.plots[-to.remove]
}
# Check if drawing a Venn diagram is possible
if ("venn" %in% qc.plots && length(statistics)==1)
{
warnwrap("The creation of a Venn diagram is possible only when more ",
"than one statistical algorithms are used (meta-analysis)! ",
"Removing from figures list...")
to.remove <- match("venn",qc.plots)
no.match <- which(is.na(to.remove))
if (length(no.match)>0)
to.remove <- to.remove[-no.match]
if (length(to.remove)>0)
qc.plots <- qc.plots[-to.remove]
}
# Check what happens with custom organism and exons/utrs
if (org=="custom" && count.type %in% c("exon","utr")
&& (is.null(gene.file) || !file.exists(gene.file)))
stopwrap("When org=\"custom\" and count.type is not \"gene\", ",
"an additional gene file must be provided with the gene.file ",
"argument!")
# Check additional input arguments for normalization and statistics
alg.args <- validate.alg.args(normalization,statistics,norm.args,stat.args)
norm.args <- alg.args$norm.args
stat.args <- alg.args$stat.args
# Override settigs if a preset is given
if (!is.null(preset))
{
preset.opts <- get.preset.opts(preset,org)
exon.filters <- preset.opts$exon.filters
gene.filters <- preset.opts$gene.filters
pcut <- preset.opts$pcut
export.what <- preset.opts$export.what
export.scale <- preset.opts$export.scale
export.values <- preset.opts$export.values
export.stats <- preset.opts$export.stats
}
if (report)
{
report.messages <- make.report.messages("en")
if (!is.null(qc.plots) && !("png" %in% fig.format))
{
warnwrap("png format is required in order to build a report! ",
"Adding to figure output formats...")
fig.format <- c(fig.format,"png")
}
}
# Display initialization report
TB <- Sys.time()
disp(strftime(Sys.time()),": Data processing started...\n")
############################################################################
disp("Read counts file: ",counts.name)
disp("Conditions: ",paste(names(sample.list),collapse=", "))
disp("Samples to include: ",paste(unlist(sample.list),collapse=", "))
if (!is.null(exclude.list) && !is.na(exclude.list))
disp("Samples to exclude: ",paste(unlist(exclude.list),collapse=", "))
else
disp("Samples to exclude: none")
disp("Requested contrasts: ",paste(contrast,collapse=", "))
if (!is.null(libsize.list))
{
disp("Library sizes: ")
for (n in names(libsize.list))
disp(" ",paste(n,libsize.list[[n]],sep=": "))
}
disp("Annotation: ",annotation)
disp("Organism: ",org)
disp("Reference source: ",refdb)
disp("Count type: ",count.type)
if (count.type == "utr")
disp("3' UTR flanking: ",utr.flank)
if (!is.null(preset))
disp("Analysis preset: ",preset)
disp("Transcriptional level: ",trans.level)
if (!is.null(exon.filters))
{
disp("Exon filters: ",paste(names(exon.filters),collapse=", "))
for (ef in names(exon.filters))
{
disp(" ",ef,": ")
for (efp in names(exon.filters[[ef]]))
{
if (length(exon.filters[[ef]][[efp]])==1 &&
is.function(exon.filters[[ef]][[efp]]))
print(exon.filters[[ef]][[efp]])
else if (length(exon.filters[[ef]][[efp]])==1)
disp(" ",paste(efp,exon.filters[[ef]][[efp]],sep=": "))
else if (length(exon.filters[[ef]][[efp]])>1)
disp(" ",paste(efp,paste(exon.filters[[ef]][[efp]],
collapse=", "),sep=": "))
}
}
}
else
disp("Exon filters: none applied")
if (!is.null(gene.filters))
{
disp("Gene filters: ",paste(names(gene.filters),collapse=", "))
for (gf in names(gene.filters))
{
disp(" ",gf,": ")
for (gfp in names(gene.filters[[gf]]))
{
if (length(gene.filters[[gf]][[gfp]])==1 &&
is.function(gene.filters[[gf]][[gfp]]))
print(gene.filters[[gf]][[gfp]])
else if (length(gene.filters[[gf]][[gfp]])==1)
disp(" ",paste(gfp,gene.filters[[gf]][[gfp]],sep=": "))
else if (length(gene.filters[[gf]][[gfp]])>1)
disp(" ",paste(gfp,paste(gene.filters[[gf]][[gfp]],
collapse=", "),sep=": "))
}
}
}
else
disp("Gene filters: none applied")
disp("Filter application: ",when.apply.filter)
disp("Normalization algorithm: ",normalization)
if (!is.null(norm.args))
{
disp("Normalization arguments: ")
for (na in names(norm.args))
{
if (length(norm.args[[na]])==1 && is.function(norm.args[[na]]))
{
disp(" ",na,": ")
disp(as.character(substitute(norm.args[[na]])))
}
else if (length(norm.args[[na]])==1)
disp(" ",paste(na,norm.args[[na]],sep=": "))
else if (length(norm.args[[na]])>1)
disp(" ",paste(na,paste(norm.args[[na]],collapse=", "),
sep=": "))
}
}
disp("Statistical algorithm: ",paste(statistics,collapse=", "))
if (!is.null(stat.args))
{
disp("Statistical arguments: ")
for (sa in names(stat.args))
{
if (length(stat.args[[sa]])==1 && is.function(stat.args[[sa]]))
{
disp(" ",sa,": ")
disp(as.character(substitute(stat.args[[na]])))
}
else if (length(stat.args[[sa]])==1)
disp(" ",paste(sa,stat.args[[sa]],sep=": "))
else if (length(stat.args[[sa]])>1)
disp(" ",paste(sa,paste(stat.args[[sa]],collapse=", "),
sep=": "))
}
}
disp("Meta-analysis method: ",meta.p)
disp("Multiple testing correction: ",adjust.method)
if (!is.na(pcut)) disp("p-value threshold: ",pcut)
disp("Logarithmic transformation offset: ",log.offset)
if (!is.null(preset)) disp("Analysis preset: ",preset)
disp("Quality control plots: ",paste(qc.plots,collapse=", "))
disp("Figure format: ",paste(fig.format,collapse=", "))
if (!is.na(export.where)) disp("Output directory: ",export.where)
disp("Output data: ",paste(export.what,collapse=", "))
disp("Output scale(s): ",paste(export.scale,collapse=", "))
disp("Output values: ",paste(export.values,collapse=", "))
if ("stats" %in% export.what)
disp("Output statistics: ",paste(export.stats,collapse=", "),"\n")
if (is.function(progress.fun)) {
text <- paste("Starting the analysis...")
progress.fun(detail=text)
}
############################################################################
if (count.type=="exon")
{
# Download gene annotation anyway if not previous analysis restored
if (!from.previous)
{
if (org=="custom")
{
if (!is.null(gene.file) && file.exists(gene.file))
{
disp("Reading custom external gene annotation for from ",
gene.file,"...")
gene.data <- read.delim(gene.file)
rownames(gene.data) <- as.character(gene.data$gene_id)
if (!is.null(gene.data$gc_content) # Already divided
&& max(as.numeric(gene.data$gc_content))<=1)
gene.data$gc_content =
100*as.numeric(gene.data$gc_content)
}
}
else
{
if (trans.level=="gene") {
disp("Downloading gene annotation for ",org,"...")
gene.data <- get.annotation(org,"gene",refdb)
}
else if (trans.level=="transcript") {
disp("Downloading transcript annotation for ",org,"...")
gene.data <- get.annotation(org,"transcript",refdb)
}
}
}
if (!from.previous)
{
if (annotation=="download")
{
disp("Downloading exon annotation for ",org,"...")
exon.data <- get.annotation(org,count.type,refdb,multic)
}
else if (annotation=="embedded")
{
# The following should work if annotation elements are arranged in
# MeV-like data style
# Embedded annotation can NEVER occur when receiving data from
# read2count, so there is no danger here
if (!is.data.frame(counts))
{
disp("Reading counts file ",counts.name,"...")
exon.counts <- read.delim(counts)
}
else
exon.counts <- counts
rownames(exon.counts) <- as.character(exon.counts[,id.col])
all.cols <- 1:ncol(exon.counts)
sam.cols <- match(unlist(sample.list),colnames(exon.counts))
sam.cols <- sam.cols[which(!is.na(sam.cols))]
ann.cols <- all.cols[-sam.cols]
exon.data <- exon.counts[,ann.cols]
exon.counts <- exon.counts[,sam.cols]
colnames(exon.data)[id.col] <- "exon_id"
if (!is.na(name.col)) colnames(exon.data)[name.col] <-
"gene_name"
if (!is.na(bt.col)) colnames(exon.data)[bt.col] <- "biotype"
exon.counts <- cbind(exon.data[rownames(exon.counts),c("start",
"end","exon_id","gene_id")],exon.counts)
}
else # Reading from external file, similar to embedded
{
disp("Reading external exon annotation for ",org," from ",
annotation,"...")
exon.data <- read.delim(annotation)
colnames(exon.data)[id.col] <- "exon_id"
}
}
else
{
counts <- tmp.env$the.counts
exon.data <- tmp.env$exon.data
gene.data <- tmp.env$gene.data
}
# Else everything is provided and done
#if (is.data.frame(counts))
if (annotation!="embedded" & !from.previous)
{
if (!is.null(counts)) # Otherwise it's coming ready from read2count
{
if (!is.data.frame(counts) && !is.list(counts))
{
disp("Reading counts file ",counts.name,"...")
exon.counts <- read.delim(counts)
}
else # Already a data frame as input
exon.counts <- counts
rownames(exon.counts) <- as.character(exon.counts[,id.col])
exon.counts <- exon.counts[,unlist(sample.list,
use.names=FALSE)]
}
else # Coming from read2count
{
if (from.raw) # Double check
{
r2c <- read2count(the.list,exon.data,file.type,
utr.flank,multic=multic)
exon.counts <- r2c$counts
# Merged exon data!
exon.data <- r2c$mergedann
if (is.null(libsize.list))
libsize.list <- r2c$libsize
if (export.counts.table) {
disp("Exporting raw read counts table to ",
file.path(PROJECT.PATH[["lists"]],
"raw_counts_table.txt.gz"))
res.file <- file.path(PROJECT.PATH[["lists"]],
"raw_counts_table.txt.gz")
gzfh <- gzfile(res.file,"w")
write.table(cbind(
exon.data[rownames(exon.counts),],
exon.counts),gzfh,sep="\t",row.names=FALSE,
quote=FALSE)
close(gzfh)
}
}
}
exon.counts <- cbind(exon.data[rownames(exon.counts),c("start",
"end","exon_id","gene_id")],exon.counts[,unlist(sample.list,
use.names=FALSE)])
# Get the exon counts per gene model
disp("Checking chromosomes in exon counts and gene annotation...")
gene.data <- reduce.gene.data(exon.data[rownames(exon.counts),],
gene.data)
disp("Processing exons...")
the.counts <- construct.gene.model(exon.counts,sample.list,
gene.data,multic=multic)
if (save.gene.model)
{
disp("Saving gene model to ",file.path(PROJECT.PATH[["data"]],
"gene_model.RData"))
save(the.counts,exon.data,gene.data,sample.list,count.type,
file=file.path(PROJECT.PATH$data,"gene_model.RData"),
compress=TRUE)
}
}
# Retrieved gene model and/or previous analysis
else if (annotation !="embedded" && from.previous)
the.counts <- counts
else if (annotation=="embedded") {
# First time read, construct gene model
disp("Checking chromosomes in exon counts and gene annotation...")
gene.data <- reduce.gene.data(exon.data[rownames(exon.counts),],
gene.data)
disp("Processing exons...")
the.counts <- construct.gene.model(exon.counts,sample.list,
gene.data,multic=multic)
if (save.gene.model)
{
disp("Saving gene model to ",file.path(PROJECT.PATH[["data"]],
"gene_model.RData"))
save(the.counts,exon.data,gene.data,sample.list,count.type,
file=file.path(PROJECT.PATH$data,"gene_model.RData"),
compress=TRUE)
}
}
# Exclude any samples not wanted (when e.g. restoring a previous project
# and having determined that some samples are of bad quality
if (!is.null(exclude.list) && !is.na(exclude.list))
{
for (n in names(exclude.list)) {
sample.list[[n]] <- setdiff(sample.list[[n]],
exclude.list[[n]])
if (length(sample.list[[n]])==0) # Removed whole condition
sample.list[n] <- NULL
}
the.counts <- the.counts[unlist(sample.list)]
}
# Apply exon filters
if (!is.null(exon.filters))
{
exon.filter.out <- filter.exons(the.counts,gene.data,sample.list,
exon.filters)
exon.filter.result <- exon.filter.out$result
exon.filter.flags <- exon.filter.out$flags
}
else
exon.filter.result <- exon.filter.flags <- NULL
disp("Summarizing count data...")
the.gene.counts <- the.exon.lengths <- vector("list",
length(unlist(sample.list)))
names(the.gene.counts) <- names(the.exon.lengths) <- names(the.counts)
for (n in names(the.gene.counts))
{
the.gene.counts[[n]] <- wapply(multic,the.counts[[n]],
function(x) return(sum(x$count)))
the.exon.lengths[[n]] <- wapply(multic,the.counts[[n]],
function(x) return(sum(x$length)))
the.gene.counts[[n]] <- do.call("c",the.gene.counts[[n]])
the.exon.lengths[[n]] <- do.call("c",the.exon.lengths[[n]])
}
gene.counts <- do.call("cbind",the.gene.counts)
gene.length <- the.exon.lengths[[1]] # Based on the sum of their exon lengths
names(gene.length) <- rownames(gene.data)
# In case there are small differences between annotation data and external
# file, due to e.g. slightly different Ensembl versions
gene.data <- gene.data[rownames(gene.counts),]
total.gene.data <- gene.data # We need this for some total stats
}
if (count.type=="utr")
{
# Download gene annotation anyway if not previous analysis restored
if (!from.previous)
{
if (org=="custom")
{
if (!is.null(gene.file) && file.exists(gene.file))
{
disp("Reading custom external gene annotation for from ",
gene.file,"...")
gene.data <- read.delim(gene.file)
rownames(gene.data) <- as.character(gene.data$gene_id)
if (!is.null(gene.data$gc_content) # Already divided
&& max(as.numeric(gene.data$gc_content))<=1)
gene.data$gc_content =
100*as.numeric(gene.data$gc_content)
}
}
else
{
if (trans.level=="gene") {
disp("Downloading gene annotation for ",org,"...")
gene.data <- get.annotation(org,"gene",refdb)
}
else if (trans.level=="transcript") {
disp("Downloading transcript annotation for ",org,"...")
gene.data <- get.annotation(org,"transcript",refdb)
}
}
}
if (!from.previous)
{
if (annotation=="download")
{
if (trans.level %in% c("gene","transcript")) {
disp("Downloading transcript annotation for ",org,"...")
transcript.data <- get.annotation(org,count.type,refdb,
multic)
}
}
else if (annotation=="embedded")
{
# The following should work if annotation elements are arranged in
# MeV-like data style
# Embedded annotation can NEVER occur when receiving data from
# read2count, so there is no danger here
if (!is.data.frame(counts))
{
disp("Reading counts file ",counts.name,"...")
transcript.counts <- read.delim(counts)
}
else
transcript.counts <- counts
rownames(transcript.counts) <-
as.character(transcript.counts[,id.col])
all.cols <- 1:ncol(transcript.counts)
sam.cols <- match(unlist(sample.list),
colnames(transcript.counts))
sam.cols <- sam.cols[which(!is.na(sam.cols))]
ann.cols <- all.cols[-sam.cols]
transcript.data <- transcript.counts[,ann.cols]
transcript.counts <- transcript.counts[,sam.cols]
colnames(transcript.data)[id.col] <- "transcript_id"
if (!is.na(name.col)) colnames(transcript.data)[name.col] <-
"gene_name"
if (!is.na(bt.col))
colnames(transcript.data)[bt.col] <- "biotype"
transcript.counts <-
cbind(transcript.data[rownames(transcript.counts),c("start",
"end","transcript_id","gene_id")],transcript.counts)
}
else # Reading from external file, similar to embedded
{
disp("Reading external transcript annotation for ",org," from ",
annotation,"...")
transcript.data <- read.delim(annotation)
colnames(transcript.data)[id.col] <- "transcript_id"
}
}
else
{
counts <- tmp.env$the.counts
transcript.data <- tmp.env$transcript.data
gene.data <- tmp.env$gene.data
}
# Else everything is provided and done
#if (is.data.frame(counts))
if (annotation!="embedded" & !from.previous)
{
if (!is.null(counts)) # Otherwise it's coming ready from read2count
{
if (!is.data.frame(counts) && !is.list(counts))
{
disp("Reading counts file ",counts.name,"...")
transcript.counts <- read.delim(counts)
}
else # Already a data frame as input
transcript.counts <- counts
rownames(transcript.counts) <-
as.character(transcript.counts[,id.col])
transcript.counts <- transcript.counts[,unlist(sample.list,
use.names=FALSE)]
}
else # Coming from read2count
{
if (from.raw) # Double check
{
r2c <- read2count(the.list,transcript.data,file.type,
trans.level,utr.flank,multic=multic)
transcript.counts <- r2c$counts
# Merged transcript data!
transcript.data <- r2c$mergedann
if (is.null(libsize.list))
libsize.list <- r2c$libsize
if (export.counts.table) {
disp("Exporting raw read counts table to ",
file.path(PROJECT.PATH[["lists"]],
"raw_counts_table.txt.gz"))
res.file <- file.path(PROJECT.PATH[["lists"]],
"raw_counts_table.txt.gz")
gzfh <- gzfile(res.file,"w")
write.table(cbind(
transcript.data[rownames(transcript.counts),],
transcript.counts),gzfh,sep="\t",row.names=FALSE,
quote=FALSE)
close(gzfh)
}
}
}
transcript.counts <-
cbind(transcript.data[rownames(transcript.counts),c("start",
"end","transcript_id","gene_id")],
transcript.counts[,unlist(sample.list,use.names=FALSE)])
# Get the transcript counts per gene model
disp("Checking chromosomes in transcript counts and gene ",
"annotation...")
gene.data <-
reduce.gene.data(transcript.data[rownames(transcript.counts),],
gene.data)
disp("Processing transcripts...")
if (trans.level=="gene")
the.counts <- construct.utr.model(transcript.counts,sample.list,
gene.data,multic=multic)
else if (trans.level=="transcript")
the.counts <- construct.utr.model(transcript.counts,sample.list,
transcript.data,multic=multic)
if (save.gene.model)
{
disp("Saving gene model to ",file.path(PROJECT.PATH[["data"]],
"gene_model.RData"))
save(the.counts,transcript.data,gene.data,sample.list,
count.type,
file=file.path(PROJECT.PATH$data,"gene_model.RData"),
compress=TRUE)
}
}
# Retrieved gene model and/or previous analysis
else if (annotation !="embedded" && from.previous)
the.counts <- counts
else if (annotation=="embedded") {
# First time read, construct gene model
disp("Checking chromosomes in transcript counts and gene ",
"annotation...")
gene.data <-
reduce.gene.data(transcript.data[rownames(transcript.counts),],
gene.data)
disp("Processing transcripts...")
the.counts <- construct.utr.model(transcript.counts,sample.list,
gene.data,multic=multic)
if (save.gene.model)
{
disp("Saving gene model to ",file.path(PROJECT.PATH[["data"]],
"gene_model.RData"))
save(the.counts,transcript.data,gene.data,sample.list,
count.type,
file=file.path(PROJECT.PATH$data,"gene_model.RData"),
compress=TRUE)
}
}
# Exclude any samples not wanted (when e.g. restoring a previous project
# and having determined that some samples are of bad quality
if (!is.null(exclude.list) && !is.na(exclude.list))
{
for (n in names(exclude.list)) {
sample.list[[n]] <- setdiff(sample.list[[n]],
exclude.list[[n]])
if (length(sample.list[[n]])==0) # Removed whole condition
sample.list[n] <- NULL
}
the.counts <- the.counts[unlist(sample.list)]
}
disp("Summarizing count data...")
the.gene.counts <- the.transcript.lengths <- vector("list",
length(unlist(sample.list)))
names(the.gene.counts) <- names(the.transcript.lengths) <-
names(the.counts)
for (n in names(the.gene.counts))
{
the.gene.counts[[n]] <- wapply(multic,the.counts[[n]],
function(x) return(sum(x$count)))
the.transcript.lengths[[n]] <- wapply(multic,the.counts[[n]],
function(x) return(sum(x$length)))
the.gene.counts[[n]] <- do.call("c",the.gene.counts[[n]])
the.transcript.lengths[[n]] <-
do.call("c",the.transcript.lengths[[n]])
}
gene.counts <- do.call("cbind",the.gene.counts)
gene.length <- the.transcript.lengths[[1]]
# Based on the sum of their transcript lengths
names(gene.length) <- rownames(gene.data)
# In case there are small differences between annotation data and
# external file, due to e.g. slightly different Ensembl versions
gene.data <- gene.data[rownames(gene.counts),]
total.gene.data <- gene.data # We need this for some total stats
exon.filter.result <- NULL
}
else if (count.type=="gene")
{
if (!from.previous)
{
if (annotation=="download")
{
if (trans.level=="gene") {
disp("Downloading gene annotation for ",org,"...")
gene.data <- get.annotation(org,count.type,refdb)
}
else if (trans.level=="transcript") {
disp("Downloading transcript annotation for ",org,"...")
gene.data <- get.annotation(org,"transcript",refdb)
gene.data$gc_content = rep(0.5,nrow(gene.data))
}
else if (trans.level=="exon") {
disp("Downloading exon annotation for ",org,"...")
gene.data <- get.annotation(org,"exon",refdb)
gene.data$gc_content = rep(0.5,nrow(gene.data))
}
}
else if (annotation=="embedded")
{
# The following should work if annotation elements are arranged
# in MeV-like data style
if (!is.data.frame(counts))
{
disp("Reading counts file ",counts.name,"...")
gene.counts <- read.delim(counts)
}
else
gene.counts <- counts
rownames(gene.counts) <- as.character(gene.counts[,id.col])
all.cols <- 1:ncol(gene.counts)
sam.cols <- match(unlist(sample.list),colnames(gene.counts))
sam.cols <- sam.cols[which(!is.na(sam.cols))]
ann.cols <- all.cols[-sam.cols]
gene.data <- gene.counts[,ann.cols]
gene.counts <- gene.counts[,sam.cols]
if (trans.level=="gene")
colnames(gene.data)[id.col] <- "gene_id"
else if (trans.level=="transcript")
colnames(gene.data)[id.col] <- "transcript_id"
else if (trans.level=="exon")
colnames(gene.data)[id.col] <- "exon_id"
if (!is.na(gc.col))
{
colnames(gene.data)[gc.col] <- "gc_content"
if (max(gene.data$gc_content<=1)) # Is already divided
gene.data$gc_content = 100*gene.data$gc_content
}
if (!is.na(name.col)) colnames(gene.data)[name.col] <-
"gene_name"
if (!is.na(bt.col)) colnames(gene.data)[bt.col] <- "biotype"
}
else # Reading from external file, similar to embedded
{
if (!is.data.frame(counts))
{
disp("Reading counts file ",counts.name,"...")
gene.counts <- read.delim(counts)
}
else
gene.counts <- counts
rownames(gene.counts) <- as.character(gene.counts[,id.col])
disp("Reading external ",trans.level," annotation for ",org,
" from ",annotation,"...")
gene.data <- read.delim(annotation)
if (trans.level=="gene")
rownames(gene.data) <- as.character(gene.data$gene_id)
else if (trans.level=="transcript")
rownames(gene.data) <- as.character(gene.data$transcript_id)
else if (trans.level=="exon")
rownames(gene.data) <- as.character(gene.data$exon_id)
gene.data <- gene.data[rownames(gene.counts),]
if (max(gene.data$gc_content)<=1) # Is already divided
gene.data$gc_content = 100*gene.data$gc_content
}
}
else
{
gene.counts <- tmp.env$gene.counts
gene.data <- tmp.env$gene.data
}
total.gene.data <- gene.data # We need this for some total stats
exon.filter.result <- NULL
# Else everything is provided and done
if (annotation!="embedded" & !from.previous)
{
if (!is.null(counts)) # Otherwise it's coming ready from read2count
{
if (!is.data.frame(counts)) # Else it's already here
{
disp("Reading counts file ",counts.name,"...")
gene.counts <- read.delim(counts)
}
else # Already a data frame as input
gene.counts <- counts
rownames(gene.counts) <- as.character(gene.counts[,id.col])
gene.counts <- gene.counts[,unlist(sample.list,
use.names=FALSE)]
}
else # Coming from read2count
{
if (from.raw) # Double check
{
r2c <- read2count(the.list,gene.data,file.type,
utr.flank,multic=multic)
gene.counts <- r2c$counts
if (is.null(libsize.list))
libsize.list <- r2c$libsize
if (export.counts.table) {
disp("Exporting raw read counts table to ",
file.path(PROJECT.PATH[["lists"]],
"raw_counts_table.txt.gz"))
res.file <- file.path(PROJECT.PATH[["lists"]],
"raw_counts_table.txt.gz")
gzfh <- gzfile(res.file,"w")
write.table(cbind(gene.data[rownames(gene.counts),],
gene.counts),gzfh,sep="\t",row.names=FALSE,
quote=FALSE)
close(gzfh)
}
}
}
}
gene.data <- gene.data[rownames(gene.counts),]
gene.length <- gene.data$end - gene.data$start # Based on total gene lengths
names(gene.length) <- rownames(gene.data)
# Exclude any samples not wanted (when e.g. restoring a previous project
# and having determined that some samples are of bad quality
if (!is.null(exclude.list) && !is.na(exclude.list))
{
for (n in names(exclude.list)) {
sample.list[[n]] <- setdiff(sample.list[[n]],
exclude.list[[n]])
if (length(sample.list[[n]])==0) # Removed whole condition
sample.list[n] <- NULL
}
gene.counts <- gene.counts[,unlist(sample.list,use.names=FALSE)]
}
if (save.gene.model)
{
disp("Saving gene model to ",file.path(PROJECT.PATH[["data"]],
"gene_model.RData"))
save(gene.counts,gene.data,sample.list,count.type,
file=file.path(PROJECT.PATH$data,"gene_model.RData"),
compress=TRUE)
}
}
# Transform GC-content and biotype
if (is.null(gene.data$gc_content))
gene.data$gc_content <- rep(0.5,nrow(gene.data))
if (is.null(gene.data$biotype))
gene.data$biotype <- rep(NA,nrow(gene.data))
names(gene.length) <- rownames(gene.counts)
attr(gene.data,"gene.length") <- gene.length
############################################################################
# BEGIN FILTERING SECTION
############################################################################
if (is.function(progress.fun)) {
text <- paste("Filtering...")
progress.fun(detail=text)
}
# GC bias is NOT alleviated if we do not remove the zeros!!!
disp("Removing genes with zero counts in all samples...")
the.zeros <- which(apply(gene.counts,1,filter.low,0))
if (length(the.zeros)>0)
{
# Store the filtered, maybe we do some stats
gene.counts.zero <- gene.counts[the.zeros,]
gene.data.zero <- gene.data[the.zeros,]
attr(gene.data.zero,"gene.length") <- gene.length[the.zeros]
the.zero.names <- rownames(gene.data)[the.zeros]
# Then remove
gene.counts <- gene.counts[-the.zeros,]
gene.data <- gene.data[-the.zeros,]
attr(gene.data,"gene.length") <- gene.length[-the.zeros]
}
else
gene.counts.zero <- gene.data.zero <- the.zero.names <- NULL
# Store un-normalized gene counts for export purposes
gene.counts.unnorm <- gene.counts
# Apply filtering prior to normalization if desired
if (when.apply.filter=="prenorm")
{
# However, a first round of normalization has to be performed in order to
# get proper expression filters
disp("Prefiltering normalization with: ",normalization)
switch(normalization,
edaseq = {
temp.genes <- normalize.edaseq(gene.counts,sample.list,
norm.args,gene.data,output="matrix")
},
deseq = {
temp.genes <- normalize.deseq(gene.counts,sample.list,norm.args,
output="matrix")
},
edger = {
temp.genes <- normalize.edger(gene.counts,sample.list,norm.args,
output="matrix")
},
noiseq = {
temp.genes <- normalize.noiseq(gene.counts,sample.list,
norm.args,gene.data,log.offset,output="matrix")
},
nbpseq = {
temp.genes <- normalize.nbpseq(gene.counts,sample.list,
norm.args,libsize.list,output="matrix")
},
none = {
# In case some external normalization is applied (e.g. equal read
# counts from all samples)
temp.genes <- gene.counts
}
)
# Now filter
if (!is.null(gene.filters))
{
gene.filter.out <- filter.genes(temp.genes,gene.data,gene.filters,
sample.list)
gene.filter.result <- gene.filter.out$result
gene.filter.cutoff <- gene.filter.out$cutoff
gene.filter.flags <- gene.filter.out$flags
}
else
gene.filter.result <- gene.filter.cutoff <-
gene.filter.flags <- NULL
# Unify the filters and filter
the.dead.genes <- list(
gene.filter.result$expression$median,
gene.filter.result$expression$mean,
gene.filter.result$expression$quantile,
gene.filter.result$expression$known,
gene.filter.result$expression$custom
)
the.dead <- unique(unlist(c(gene.filter.result,exon.filter.result)))
# Some genes filtered by zero, were present in exon filters, not yet applied
if (count.type=="exon")
the.dead <- setdiff(the.dead,the.zero.names)
# All method specific objects are row-index subsettable
if (length(the.dead)>0)
{
# Store the filtered for later export or some stats
gene.counts.dead <- gene.counts[the.dead,]
gene.counts.unnorm <- gene.counts.unnorm[the.dead,]
gene.data.dead <- gene.data[the.dead,]
attr(gene.data.dead,"gene.length") <- attr(gene.data,
"gene.length")[the.dead]
# Now filter
the.dead.ind <- match(the.dead,rownames(gene.counts))
gene.counts.expr <- gene.counts[-the.dead.ind,]
gene.data.expr <- gene.data[-the.dead.ind,]
attr(gene.data.expr,"gene.length") <- attr(gene.data,
"gene.length")[-the.dead.ind]
}
else
{
gene.counts.expr <- gene.counts
gene.data.expr <- gene.data
gene.counts.dead <- gene.data.dead <- gene.counts.unnorm <- NULL
}
if (is.function(progress.fun)) {
text <- paste("Normalizing...")
progress.fun(detail=text)
}
disp("Normalizing with: ",normalization)
switch(normalization,
edaseq = {
norm.genes <- normalize.edaseq(gene.counts.expr,sample.list,
norm.args,gene.data.expr,output="matrix")
},
deseq = {
norm.genes <- normalize.deseq(gene.counts.expr,sample.list,
norm.args,output="native")
},
edger = {
norm.genes <- normalize.edger(gene.counts.expr,sample.list,
norm.args,output="native")
},
noiseq = {
norm.genes <- normalize.noiseq(gene.counts.expr,sample.list,
norm.args,gene.data.expr,log.offset,output="matrix")
},
nbpseq = {
norm.genes <- normalize.nbpseq(gene.counts.expr,sample.list,
norm.args,libsize.list,output="native")
},
none = {
norm.genes <- gene.counts.expr
}
)
norm.genes.expr <- norm.genes
}
else if (when.apply.filter=="postnorm")
{
if (is.function(progress.fun)) {
text <- paste("Normalizing...")
progress.fun(detail=text)
}
# Apply filtering after normalization if desired (default)
disp("Normalizing with: ",normalization)
switch(normalization,
edaseq = {
norm.genes <- normalize.edaseq(gene.counts,sample.list,
norm.args,gene.data,output="matrix")
},
deseq = {
norm.genes <- normalize.deseq(gene.counts,sample.list,norm.args,
output="native")
},
edger = {
norm.genes <- normalize.edger(gene.counts,sample.list,norm.args,
output="native")
},
noiseq = {
norm.genes <- normalize.noiseq(gene.counts,sample.list,
norm.args,gene.data,log.offset,output="matrix")
},
nbpseq = {
norm.genes <- normalize.nbpseq(gene.counts,sample.list,
norm.args,libsize.list,output="native")
},
none = {
norm.genes <- gene.counts
}
)
switch(class(norm.genes)[1],
CountDataSet = { # Has been normalized with DESeq
temp.matrix <- round(counts(norm.genes,normalized=TRUE))
},
DGEList = { # Has been normalized with edgeR
# Trick found at http://cgrlucb.wikispaces.com/edgeR+spring2013
scl <- norm.genes$samples$lib.size *
norm.genes$samples$norm.factors
temp.matrix <- round(t(t(norm.genes$counts)/scl)*mean(scl))
},
matrix = { # Has been normalized with EDASeq or NOISeq or nothing
temp.matrix <- norm.genes
},
data.frame = { # Has been normalized with or nothing
temp.matrix <- as.matrix(norm.genes)
},
list = { # Has been normalized with NBPSeq and main method was "nbpseq"
temp.matrix <- as.matrix(round(sweep(norm.genes$counts,2,
norm.genes$norm.factors,"*")))
},
nbp = { # Has been normalized with NBPSeq and main method was "nbsmyth"
temp.matrix <- as.matrix(round(norm.genes$pseudo.counts))
}
)
# Implement gene filters after normalization
if (!is.null(gene.filters)) {
gene.filter.out <- filter.genes(temp.matrix,gene.data,gene.filters,
sample.list)
gene.filter.result <- gene.filter.out$result
gene.filter.cutoff <- gene.filter.out$cutoff
gene.filter.flags <- gene.filter.out$flags
}
else
gene.filter.result <- gene.filter.cutoff <-
gene.filter.flags <- NULL
# Unify the filters and filter
the.dead.genes <- list(
gene.filter.result$expression$median,
gene.filter.result$expression$mean,
gene.filter.result$expression$quantile,
gene.filter.result$expression$known,
gene.filter.result$expression$custom
)
#gene.filter.result$expression <- Reduce("union",the.dead.genes)
the.dead <- unique(unlist(c(gene.filter.result,exon.filter.result)))
# Some genes filtered by zero, were present in exon filters, not yet applied
if (count.type=="exon")
the.dead <- setdiff(the.dead,the.zero.names)
# All method specific object are row-index subsettable
if (length(the.dead)>0)
{
# Store the filtered for later export or some stats
gene.counts.dead <- temp.matrix[the.dead,]
gene.counts.unnorm <- gene.counts.unnorm[the.dead,]
gene.data.dead <- gene.data[the.dead,]
attr(gene.data.dead,"gene.length") <- attr(gene.data,
"gene.length")[the.dead]
# Now filter
the.dead.ind <- match(the.dead,rownames(temp.matrix))
switch(class(norm.genes)[1],
CountDataSet = {
norm.genes.expr <- norm.genes[-the.dead.ind,]
},
DGEList = { # edgeR bug???
norm.genes.expr <- norm.genes[-the.dead.ind,]
norm.genes.expr$AveLogCPM <-
norm.genes.expr$AveLogCPM[-the.dead.ind]
},
matrix = { # Has been normalized with EDASeq or NOISeq
norm.genes.expr <- norm.genes[-the.dead.ind,]
},
data.frame = { # Has been normalized with EDASeq or NOISeq
norm.genes.expr <- as.matrix(norm.genes[-the.dead.ind,])
},
list = { # Has been normalized with NBPSeq, main.method="nbpseq"
norm.genes.expr <- norm.genes
norm.genes.expr$counts <-
as.matrix(norm.genes.expr$counts[-the.dead.ind,])
norm.genes.expr$rel.frequencies <-
norm.genes.expr$rel.frequencies[-the.dead.ind,]
norm.genes.expr$tags <-
as.matrix(norm.genes.expr$tags[-the.dead.ind,])
},
nbp = {
norm.genes.expr <- norm.genes
norm.genes.expr$counts <-
as.matrix(norm.genes.expr$counts[-the.dead.ind,])
norm.genes.expr$pseudo.counts <-
as.matrix(norm.genes.expr$pseudo.counts[-the.dead.ind,])
norm.genes.expr$pseudo.lib.sizes <-
colSums(as.matrix(norm.genes.expr$pseudo.counts))*
rep(1,dim(norm.genes.expr$counts)[2])
}
)
gene.counts.expr <- gene.counts[rownames(norm.genes.expr),]
gene.data.expr <- gene.data[-the.dead.ind,]
attr(gene.data.expr,"gene.length") <-
attr(gene.data,"gene.length")[-the.dead.ind]
}
else
{
norm.genes.expr <- norm.genes
gene.counts.expr <- gene.counts
gene.data.expr <- gene.data
gene.counts.dead <- gene.data.dead <- gene.counts.unnorm <- NULL
}
}
# Store the final filtered, maybe we do some stats
gene.data.filtered <- rbind(gene.data.zero,gene.data.dead)
if (!is.null(gene.data.filtered) && nrow(gene.data.filtered)>0)
{
disp(nrow(gene.data.filtered)," genes filtered out")
if (!is.null(gene.data.zero) && nrow(gene.data.zero)>0)
attr(gene.data.filtered,"gene.length") <- c(attr(gene.data.zero,
"gene.length"),attr(gene.data.dead,"gene.length"))
else
attr(gene.data.filtered,"gene.length") <-
attr(gene.data.dead,"gene.length")
}
if (!is.null(gene.filters) || !is.null(exon.filters))
disp(nrow(gene.data.expr)," genes remain after filtering")
############################################################################
# END FILTERING SECTION
############################################################################
# There is a small case that no genes are left after filtering...
if(any(dim(norm.genes.expr)==0))
stopwrap("No genes left after gene and/or exon filtering! Try again ",
"with no filtering or less strict filter rules...")
if (is.function(progress.fun)) {
text <- paste("Statistical testing...")
progress.fun(detail=text)
}
# Run the statistical test, norm.genes is always a method-specific object,
# handled in the metaseqr.stat.R stat.* functions
cp.list <- vector("list",length(contrast))
names(cp.list) <- contrast
contrast.list <- make.contrast.list(contrast,sample.list)
for (n in names(cp.list))
{
cp.list[[n]] <- vector("list",length(statistics))
names(cp.list[[n]]) <- statistics
}
for (alg in statistics)
{
disp("Running statistical tests with: ",alg)
switch(alg,
deseq = {
p.list <- stat.deseq(norm.genes.expr,sample.list,contrast.list,
stat.args[[alg]])
if (!is.na(pcut)) {
for (con in names(contrast.list))
disp(" Contrast ",con,": found ",
length(which(p.list[[con]]<=pcut))," genes")
}
},
edger = {
p.list <- stat.edger(norm.genes.expr,sample.list,contrast.list,
stat.args[[alg]])
if (!is.na(pcut)) {
for (con in names(contrast.list))
disp(" Contrast ",con,": found ",
length(which(p.list[[con]]<=pcut))," genes")
}
},
noiseq = {
p.list <- stat.noiseq(norm.genes.expr,sample.list,contrast.list,
stat.args[[alg]],gene.data.expr,log.offset)
if (!is.na(pcut)) {
for (con in names(contrast.list))
disp(" Contrast ",con,": found ",
length(which(p.list[[con]]<=pcut))," genes")
}
},
bayseq = {
p.list <- stat.bayseq(norm.genes.expr,sample.list,contrast.list,
stat.args[[alg]],libsize.list)
if (!is.na(pcut)) {
for (con in names(contrast.list))
disp(" Contrast ",con,": found ",
length(which(p.list[[con]]<=pcut))," genes")
}
},
limma = {
p.list <- stat.limma(norm.genes.expr,sample.list,contrast.list,
stat.args[[alg]])
if (!is.na(pcut)) {
for (con in names(contrast.list))
disp(" Contrast ",con,": found ",
length(which(p.list[[con]]<=pcut))," genes")
}
},
nbpseq = {
p.list <- stat.nbpseq(norm.genes.expr,sample.list,contrast.list,
stat.args[[alg]],libsize.list)
if (!is.na(pcut)) {
for (con in names(contrast.list))
disp(" Contrast ",con,": found ",
length(which(p.list[[con]]<=pcut))," genes")
}
}
)
for (n in names(p.list))
cp.list[[n]][[alg]] <- p.list[[n]]
}
for (n in names(cp.list))
cp.list[[n]] <- do.call("cbind",cp.list[[n]])
# Create the adjusted p-value matrices (if needed)
if ("adj.p.value" %in% export.what)
{
adj.cp.list <- wapply(multic,cp.list,
function(x,a) return(apply(x,2,p.adjust,a)),adjust.method)
for (n in names(cp.list))
{
noi <- grep("noiseq",colnames(cp.list[[n]]))
if (length(noi)>0)
{
# DESeq has not run in this case, FDR cannot be calculated
if (length(strsplit(n,"_vs_")[[1]])==2)
adj.cp.list[[n]][,noi] <- rep(NA,nrow(cp.list[[n]]))
}
}
}
else
adj.cp.list <- NULL
# At this point, all method-specific objects must become matrices for exporting
# and plotting
switch(class(norm.genes.expr)[1],
CountDataSet = { # Has been processed with DESeq
norm.genes <- round(counts(norm.genes,normalized=TRUE))
norm.genes.expr <- round(counts(norm.genes.expr,normalized=TRUE))
},
DGEList = { # Has been processed with edgeR
# Trick found at http://cgrlucb.wikispaces.com/edgeR+spring2013
scl.r <- norm.genes$samples$lib.size*norm.genes$samples$norm.factors
norm.genes <- round(t(t(norm.genes$counts)/scl.r)*mean(scl.r))
scl.n <- norm.genes.expr$samples$lib.size *
norm.genes.expr$samples$norm.factors
norm.genes.expr <- round(t(t(norm.genes.expr$counts)/scl.n) *
mean(scl.n))
},
list = {
norm.genes <- as.matrix(round(sweep(norm.genes$counts,2,
norm.genes$norm.factors,"*")))
norm.genes.expr <- as.matrix(round(sweep(norm.genes.expr$counts,2,
norm.genes$norm.factors,"*")))
},
nbp = {
norm.genes <- as.matrix(round(norm.genes$pseudo.counts))
norm.genes.expr <- as.matrix(round(norm.genes.expr$pseudo.counts))
}
# We don't need the matrix case
)
# Now that everything is a matrix, export the normalized counts if asked
if (export.counts.table) {
disp("Exporting and compressing normalized read counts table to ",
file.path(PROJECT.PATH[["lists"]],"normalized_counts_table.txt"))
expo <- cbind(
rbind(gene.data.expr,gene.data.filtered),
rbind(norm.genes.expr,gene.counts.zero,gene.counts.dead)
)
res.file <- file.path(PROJECT.PATH[["lists"]],
"normalized_counts_table.txt.gz")
gzfh <- gzfile(res.file,"w")
write.table(expo,gzfh,sep="\t",row.names=FALSE,quote=FALSE)
close(gzfh)
}
# Calculate meta-statistics, if more than one statistical algorithm has been used
if (length(statistics)>1)
{
sum.p.list <- meta.test(
cp.list=cp.list,
meta.p=meta.p,
counts=norm.genes.expr,
sample.list=sample.list,
statistics=statistics,
stat.args=stat.args,
libsize.list=libsize.list,
nperm=nperm,
weight=weight,
reprod=reprod,
multic=multic
)
}
# We assign the p-values from the only statistic used to sum.p.list in order
# to use it for stat plots
else
sum.p.list <- cp.list
# Useless for one statistics but just for safety
if ("adj.meta.p.value" %in% export.what)
adj.sum.p.list <- wapply(multic,sum.p.list,
function(x,a) return(p.adjust(x,a)),adjust.method)
else
adj.sum.p.list <- NULL
############################################################################
# BEGIN EXPORT SECTION
############################################################################
if (is.function(progress.fun)) {
text <- paste("Exporting...")
progress.fun(detail=text)
}
# Bind all the flags
if (count.type=="gene")
flags <- gene.filter.flags
else if (count.type=="utr")
flags <- gene.filter.flags
else if (count.type=="exon")
{
if (!is.null(exon.filter.flags))
{
flags <- cbind(gene.filter.flags,
as.matrix(exon.filter.flags[rownames(gene.filter.flags),]))
nams <- c(colnames(gene.filter.flags),colnames(exon.filter.flags))
rownames(flags) <- rownames(gene.filter.flags)
colnames(flags) <- nams
}
else
flags <- gene.filter.flags
}
disp("Building output files...")
if (out.list) out <- make.export.list(contrast) else out <- NULL
if (report) html <- make.export.list(contrast) else html <- NULL
if ("rpgm" %in% export.scale)
fa <- attr(gene.data,"gene.length")
else
fa <- NULL
if ("normalized" %in% export.values) {
fac <- fa[rownames(norm.genes.expr)]
norm.list <- make.transformation(norm.genes.expr,export.scale,fac,
log.offset)
}
else
norm.list <- NULL
if ("raw" %in% export.values) {
fac <- fa[rownames(gene.counts.expr)]
raw.list <- make.transformation(gene.counts.expr,export.scale,fac,
log.offset)
}
else
raw.list <- NULL
if ("flags" %in% export.what)
good.flags <- flags[rownames(norm.genes.expr),]
else
good.flags <- NULL
if (!is.null(gene.counts.zero) || !is.null(gene.counts.dead))
{
gene.counts.filtered <- rbind(gene.counts.zero,gene.counts.dead)
gene.counts.unnorm.filtered <- rbind(gene.counts.zero,
gene.counts.unnorm)
if ("normalized" %in% export.values) {
fac <- fa[rownames(gene.counts.filtered)]
norm.list.filtered <- make.transformation(gene.counts.filtered,
export.scale,fac,log.offset)
}
else
norm.list.filtered <- NULL
if ("raw" %in% export.values) {
fac <- fa[rownames(gene.counts.unnorm.filtered)]
raw.list.filtered <- make.transformation(
gene.counts.unnorm.filtered,export.scale,fac,log.offset)
}
else
raw.list.filtered <- NULL
if ("flags" %in% export.what && !is.null(flags))
all.flags <- rbind(
matrix(1,nrow(gene.counts.zero),ncol(flags)),
flags[rownames(gene.counts.dead),]
)
else
all.flags <- NULL
}
else {
gene.counts.filtered <- NULL
gene.counts.unnorm.filtered <- NULL
all.flags <- NULL
}
counter <- 1
for (cnt in contrast)
{
disp(" Contrast: ",cnt)
disp(" Adding non-filtered data...")
the.export <- build.export(
gene.data=gene.data.expr,
raw.gene.counts=gene.counts.expr,
norm.gene.counts=norm.genes.expr,
flags=good.flags,
sample.list=sample.list,
cnt=cnt,
statistics=statistics,
raw.list=raw.list,
norm.list=norm.list,
p.mat=cp.list[[cnt]],
adj.p.mat=adj.cp.list[[cnt]],
sum.p=sum.p.list[[cnt]],
adj.sum.p=adj.sum.p.list[[cnt]],
export.what=export.what,
export.scale=export.scale,
export.values=export.values,
export.stats=export.stats,
log.offset=log.offset,
report=report
)
# Adjust the export based on what statistics have been done and a possible
# p-value cutoff
export <- the.export$text.table
if (report)
export.html <- the.export$html.table
if (!is.na(pcut))
{
if (length(statistics)>1)
{
switch(meta.p,
fisher = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
fperm = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
whitlock = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
dperm.min = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
dperm.max = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
dperm.weight = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
minp = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
maxp = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
weight = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
pandora = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
simes = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
},
none = {
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
}
)
pp <- sum.p.list[[cnt]][cut.ind]
export <- export[cut.ind,]
export <- export[order(pp),]
if (report)
{
export.html <- export.html[cut.ind,]
export.html <- export.html[order(pp),]
}
}
else
{
cut.ind <- which(sum.p.list[[cnt]]<=pcut)
pp <- sum.p.list[[cnt]][cut.ind,]
export <- export[cut.ind,]
export <- export[order(pp),]
if (report)
{
export.html <- export.html[cut.ind,]
export.html <- export.html[order(pp),]
}
}
}
else
{
pp <- sum.p.list[[cnt]]
export <- export[order(pp),]
if (report)
export.html <- export.html[order(pp),]
}
# Final safety trigger
na.ind <- grep("NA",rownames(export))
if (length(na.ind)>0)
{
export <- export[-na.ind,]
if (report) export.html <- export.html[-na.ind,]
}
res.file <- file.path(PROJECT.PATH[["lists"]],
paste("metaseqr_sig_out_",cnt,".txt.gz",sep=""))
disp(" Writing output...")
gzfh <- gzfile(res.file,"w")
write.table(export,gzfh,quote=FALSE,row.names=FALSE,sep="\t")
close(gzfh)
if (out.list)
out[[cnt]] <- export
if (report)
{
the.html.header <- make.html.header(the.export$headers)
if (!is.null(report.top)) {
topi <- ceiling(report.top*nrow(export.html))
the.html.rows <- make.html.rows(export.html[1:topi,])
}
else
the.html.rows <- make.html.rows(export.html)
the.html.body <- make.html.body(the.html.rows)
the.html.table <- make.html.table(the.html.body,the.html.header,
id=paste("table_",counter,sep=""))
html[[cnt]] <- the.html.table
counter <- counter+1
}
if (!is.null(gene.counts.zero) || !is.null(gene.counts.dead))
{
disp(" Adding filtered data...")
the.export.filtered <- build.export(
gene.data=gene.data.filtered,
raw.gene.counts=gene.counts.unnorm.filtered,
norm.gene.counts=gene.counts.filtered,
flags=all.flags,
sample.list=sample.list,
cnt=cnt,
statistics=statistics,
raw.list=raw.list.filtered,
norm.list=norm.list.filtered,
export.what=export.what,
export.scale=export.scale,
export.values=export.values,
export.stats=export.stats,
log.offset=log.offset,
report=FALSE
)
# Now we should be having the.export and the.export.filtered. We do not
# generate html output for filtered or total results just a compressed
# text file. We thus have to append the.export$text.table and
# the.export.filtered$html.table before writing the final output...
export.all <- rbind(the.export$text.table,
the.export.filtered$text.table)
# ...and order them somehow... alphabetically according to row names, as
# the annotation might not have been bundled...
export.all <- export.all[order(rownames(export.all)),]
res.file <- file.path(PROJECT.PATH[["lists"]],paste(
"metaseqr_all_out_",cnt,".txt.gz",sep=""))
disp(" Writing output...")
gzfh <- gzfile(res.file,"w")
write.table(export.all,gzfh,quote=FALSE,row.names=FALSE,sep="\t")
close(gzfh)
}
}
############################################################################
# END EXPORT SECTION
############################################################################
############################################################################
# BEGIN PLOTTING SECTION
############################################################################
if (is.function(progress.fun)) {
text <- paste("Plotting...")
progress.fun(detail=text)
}
if (!is.null(qc.plots))
{
disp("Creating quality control graphs...")
plots <- list(
raw=c("mds","biodetection","countsbio","saturation","readnoise",
"correl","pairwise"),
norm=c("boxplot","gcbias","lengthbias","meandiff","meanvar",
"rnacomp"),
stat=c("deheatmap","volcano","biodist"),
other=c("filtered"),
venn=c("venn")
)
fig.raw <- fig.unorm <- fig.norm <- fig.stat <- fig.other <- fig.venn <-
vector("list",length(fig.format))
names(fig.raw) <- names(fig.unorm) <- names(fig.norm) <-
names(fig.stat) <- names(fig.other) <- names(fig.venn) <-
fig.format
for (fig in fig.format)
{
disp("Plotting in ",fig," format...")
fig.raw[[fig]] <- diagplot.metaseqr(gene.counts,sample.list,
annotation=gene.data,diagplot.type=intersect(qc.plots,
plots$raw),is.norm=FALSE,output=fig,path=PROJECT.PATH$qc)
fig.unorm[[fig]] <- diagplot.metaseqr(gene.counts,sample.list,
annotation=gene.data,diagplot.type=intersect(qc.plots,
plots$norm),is.norm=FALSE,output=fig,
path=PROJECT.PATH$normalization)
if (when.apply.filter=="prenorm") # The annotation dimensions change...
fig.norm[[fig]] <- diagplot.metaseqr(norm.genes,sample.list,
annotation=gene.data.expr,diagplot.type=intersect(qc.plots,
plots$norm),is.norm=TRUE,output=fig,
path=PROJECT.PATH$normalization)
else if (when.apply.filter=="postnorm")
fig.norm[[fig]] <- diagplot.metaseqr(norm.genes,sample.list,
annotation=gene.data,diagplot.type=intersect(qc.plots,
plots$norm),is.norm=TRUE,output=fig,
path=PROJECT.PATH$normalization)
fig.stat[[fig]] <- diagplot.metaseqr(norm.genes.expr,sample.list,
annotation=gene.data.expr,contrast.list=contrast.list,
p.list=sum.p.list,thresholds=list(p=pcut,f=1),
diagplot.type=intersect(qc.plots,plots$stat),is.norm=TRUE,
output=fig,path=PROJECT.PATH$statistics)
if (!is.null(gene.data.filtered))
fig.other[[fig]] <- diagplot.metaseqr(gene.data.filtered,
sample.list,annotation=total.gene.data,
diagplot.type=intersect(qc.plots,plots$other),
is.norm=FALSE,output=fig,path=PROJECT.PATH$qc)
else fig.other[[fig]] <- NULL
if ("venn" %in% qc.plots)
fig.venn[[fig]] <- diagplot.metaseqr(norm.genes.expr,
sample.list,annotation=gene.data.expr,
contrast.list=contrast.list,
p.list=cp.list,thresholds=list(p=pcut,f=1),
diagplot.type=intersect(qc.plots,plots$venn),
output=fig,path=PROJECT.PATH$statistics)
}
}
############################################################################
# END PLOTTING SECTION
############################################################################
############################################################################
# BEGIN REPORTING SECTION
############################################################################
if (report)
{
disp("Creating HTML report...")
if (!is.null(qc.plots))
{
# First create zip archives of the figures
disp("Compressing figures...")
zipfiles <- file.path(PROJECT.PATH$plots,paste("metaseqr_figures_",
fig.format,".zip",sep=""))
names(zipfiles) <- fig.format
for (f in fig.format)
{
files <- c(
dir(PROJECT.PATH$qc,pattern=paste(".",f,sep=""),
full.names=TRUE),
dir(PROJECT.PATH$normalization,pattern=paste(".",f,sep=""),
full.names=TRUE),
dir(PROJECT.PATH$statistics,pattern=paste(".",f,sep=""),
full.names=TRUE)
)
zip(zipfiles[f],files)
}
# Then create the final figure variables which brew will find...
fig.raw <- fig.raw[["png"]]
fig.unorm <- fig.unorm[["png"]]
fig.norm <- fig.norm[["png"]]
fig.stat <- fig.stat[["png"]]
fig.other <- fig.other[["png"]]
fig.venn <- fig.venn[["png"]]
}
else
fig.raw <- fig.unorm <- fig.norm <- fig.stat <- fig.other <-
fig.venn <- NULL
if (tolower(report.template)=="default")
{
if (exists("TEMPLATE"))
{
report.template=list(
html=file.path(TEMPLATE,"metaseqr_report.html"),
css=file.path(TEMPLATE,"styles.css"),
logo=file.path(TEMPLATE,"logo.png"),
loader=file.path(TEMPLATE,"loader.gif")
)
}
else
report.template=list(html=NULL,css=NULL,logo=NULL,
loader=NULL)
}
if (!is.null(report.template$html))
{
if (file.exists(report.template$html))
{
template <- report.template$html
has.template <- TRUE
}
else
{
warnwrap(paste("The template file",report.template$html,
"was not ","found! The HTML report will NOT be generated."))
has.template <- FALSE
}
}
else
{
warnwrap(paste("The report option was enabled but no template ",
"file is provided! The HTML report will NOT be generated."))
has.template <- FALSE
}
if (!is.null(report.template$css))
{
if (file.exists(report.template$css))
file.copy(from=report.template$css,to=PROJECT.PATH$media)
else
warnwrap(paste("The stylesheet file",report.template$css,
"was not ","found! The HTML report will NOT be styled."))
}
else
warnwrap(paste("The report stylesheet file was not provided! The ",
"HTML report will NOT be styled."))
if (!is.null(report.template$logo))
{
if (file.exists(report.template$logo))
file.copy(from=report.template$logo,to=PROJECT.PATH$media)
else
warnwrap(paste("The report logo image",report.template$logo,
"was not found!"))
}
else
warnwrap(paste("The report logo image was not provided!"))
if (!is.null(report.template$loader))
{
if (file.exists(report.template$loader))
file.copy(from=report.template$loader,to=PROJECT.PATH$media)
else
warnwrap(paste("The report logo image",report.template$loader,
"was not found!"))
}
else
warnwrap(paste("The report loader image was not provided!"))
if (has.template)
{
exec.time <- elap2human(TB)
TEMP <- environment()
brew(
file=report.template$html,
#output=file.path(PROJECT.PATH$main,paste(
# basename(PROJECT.PATH$main),"html",sep=".")),
output=file.path(PROJECT.PATH$main,"index.html"),
envir=TEMP
)
}
}
############################################################################
# END REPORTING SECTION
############################################################################
disp("\n",strftime(Sys.time()),": Data processing finished!\n")
exec.time <- elap2human(TB)
disp("\n","Total processing time: ",exec.time,"\n\n")
if (out.list) {
tmp <- rbind(gene.data.expr,gene.data.filtered)
a <- c(attr(gene.data.expr,"gene.length"),
attr(gene.data.filtered,"gene.length"))
names(a) <- rownames(tmp)
attr(tmp,"gene.length") <- a
for (n in names(cp.list)) {
if (!is.null(gene.data.filtered)) {
filler <- matrix(NA,nrow(gene.data.filtered),ncol(cp.list[[n]]))
rownames(filler) <- rownames(gene.data.filtered)
colnames(filler) <- colnames(cp.list[[n]])
}
else
filler <- NULL
cp.list[[n]] <- rbind(cp.list[[n]],filler)
cp.list[[n]] <- cp.list[[n]][rownames(tmp),,drop=FALSE]
}
if (!is.null(adj.cp.list)) {
for (n in names(adj.cp.list)) {
if (!is.null(gene.data.filtered)) {
filler <- matrix(NA,nrow(gene.data.filtered),
ncol(adj.cp.list[[n]]))
rownames(filler) <- rownames(gene.data.filtered)
colnames(filler) <- colnames(cp.list[[n]])
}
else
filler <- NULL
adj.cp.list[[n]] <- rbind(adj.cp.list[[n]],filler)
adj.cp.list[[n]] <- adj.cp.list[[n]][rownames(tmp),,drop=FALSE]
}
}
if (!is.null(sum.p.list)) {
for (n in names(sum.p.list)) {
if (!is.null(gene.data.filtered)) {
filler <- rep(NA,nrow(gene.data.filtered))
names(filler) <- rownames(gene.data.filtered)
}
else
filler <- NULL
if (is.matrix(sum.p.list[[n]])) {
sum.p.list[[n]] <- rbind(sum.p.list[[n]],as.matrix(filler))
sum.p.list[[n]] <- sum.p.list[[n]][rownames(tmp),,
drop=FALSE]
}
else {
sum.p.list[[n]] <- c(sum.p.list[[n]],filler)
sum.p.list[[n]] <- sum.p.list[[n]][rownames(tmp)]
}
}
}
if (!is.null(adj.sum.p.list)) {
for (n in names(adj.sum.p.list)) {
if (!is.null(gene.data.filtered)) {
filler <- rep(NA,nrow(gene.data.filtered))
names(filler) <- rownames(gene.data.filtered)
}
else
filler <- NULL
adj.sum.p.list[[n]] <- c(adj.sum.p.list[[n]],filler)
adj.sum.p.list[[n]] <- adj.sum.p.list[[n]][rownames(tmp)]
}
}
complete <- list(
#call=as.list(match.call()),
params=list(
sample.list=sample.list,
exclude.list=exclude.list,
file.type=file.type,
path=path,
contrast=contrast,
libsize.list=libsize.list,
id.col=id.col,
gc.col=gc.col,
name.col=name.col,
bt.col=bt.col,
annotation=annotation,
org=org,
refdb=refdb,
count.type=count.type,
exon.filters=exon.filters,
gene.filters=gene.filters,
when.apply.filter=when.apply.filter,
normalization=normalization,
norm.args=norm.args,
statistics=statistics,
stat.args=stat.args,
adjust.method=adjust.method,
meta.p=meta.p,
weight=weight,
nperm=nperm,
reprod=reprod,
pcut=pcut,
log.offset=log.offset,
preset=preset,
qc.plots=qc.plots,
fig.format=fig.format,
out.list=out.list,
export.where=export.where,
export.what=export.what,
export.scale=export.scale,
export.values=export.values,
export.stats=export.stats,
export.counts.table=export.counts.table,
restrict.cores=restrict.cores,
report=report,
report.top=report.top,
report.template=report.template,
save.gene.model=save.gene.model,
verbose=verbose,
run.log=run.log
),
filter.cutoffs=list(
exon.filter=list(
min.active.exons=NULL
),
gene.filter=list(
length=gene.filters$length$length,
avg.reads=if (is.null(gene.filter.cutoff)) NULL else
round(gene.filter.cutoff$avg.reads,digits=5),
expression=list(
median=gene.filter.cutoff$expression$median,
mean=gene.filter.cutoff$expression$mean,
quantile=gene.filter.cutoff$expression$quantile,
known=gene.filter.cutoff$expression$known,
custom=gene.filter.cutoff$expression$custom
),
biotype=if (is.null(gene.filters$biotype)) NULL else
paste(names(gene.filters$biotype)[which(
unlist(gene.filters$biotype))],collapse=", ")
),
zero.filtered=length(the.zeros),
exon.filtered=length(unique(unlist(exon.filter.result))),
gene.filtered=length(unique(unlist(gene.filter.result))),
total.filtered=length(the.zeros)+length(the.dead)
),
gene.data=tmp,
raw.counts=rbind(gene.counts.expr,gene.counts.unnorm.filtered),
norm.counts=rbind(norm.genes.expr,gene.counts.filtered),
flags=rbind(good.flags,all.flags),
sample.list=sample.list,
contrast.list=contrast,
p.value=cp.list,
fdr=adj.cp.list,
meta.p.value=sum.p.list,
meta.fdr=adj.sum.p.list
)
return(list(data=out,html=html,complete=complete))
}
} # End metaseqr
#' Assemble a gene model based on exon counts
#'
#' This function assembles gene models (single genes, not isoforms) based on the
#' input exon read counts file and a gene annotation data frame, either from an
#' external file provided by the user, or with the \code{\link{get.annotation}}
#' function. The \code{gene.data} argument should have a specific format and for
#' this reason it's better to use one of the two aforementioned ways to supply it.
#' This function is intended mostly for internal use but can be used if the
#' requirements are met.
#'
#' @param exon.counts the exon counts data frame produced by reading the exon read
#' counts file.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param gene.data an annotation data frame from the same organism as
#' \code{exon.counts} (such the ones produced by \code{get.annotation}).
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not.
#' @return A named list where names represent samples. Each list member is a also
#' a named list where names correspond to gene ids and members are named vectors.
#' Each vector is named according to the exons corresponding to each gene and
#' contains the read counts for each exon. This structure is used for exon filtering
#' and assembling final gene counts in the metaseqr pipeline.
#' @author <NAME>
#' @export
#' @examples
#' @examples
#' \dontrun{
#' data("hg19.exon.data",package="metaseqR")
#' gene.data <- get.annotation("hg19","gene","ensembl")
#' reduced.gene.data <- reduce.gene.data(hg19.exon.counts,gene.data)
#' multic <- check.parallel(0.4)
#' gene.model <- construct.gene.model(hg19.exon.counts,sample.list.hg19,gene.data,
#' multic)
#'}
construct.gene.model <- function(exon.counts,sample.list,
gene.data,multic=FALSE) {
the.counts <- vector("list",length(unlist(sample.list)))
names(the.counts) <- unlist(sample.list,use.names=FALSE)
the.genes <- as.character(unique(gene.data$gene_id))
#the.exons <- as.character(unique(exon.counts$exon_id))
#the.exons <- as.character(unique(exon.data[,id.col]))
for (n in names(the.counts))
{
disp(" Separating exons per gene for ",n,"...")
#the.counts[[n]] <- vector("list",length(the.genes))
the.counts[[n]] <- the.genes
names(the.counts[[n]]) <- the.genes
the.counts[[n]] <- wapply(multic,the.counts[[n]],function(x,d,n) {
tmp <- d[which(d$gene_id==x),c("start","end","exon_id",n)]
xx <- tmp[,n]
yy <- tmp$end - tmp$start
names(xx) <- names(yy) <- tmp$exon_id
return(list(count=xx,length=yy))
},exon.counts,n)
}
return(the.counts)
}
#' Assemble a gene model based on 3' UTR counts for quant-seq data
#'
#' This function assembles gene models (single genes, not isoforms) based on the
#' input read counts file (3' UTRs) and a gene annotation data frame, either from an
#' external file provided by the user, or with the \code{\link{get.annotation}}
#' function. The \code{gene.data} argument should have a specific format and for
#' this reason it's better to use one of the two aforementioned ways to supply it.
#' This function is intended mostly for internal use but can be used if the
#' requirements are met.
#'
#' @param utr.counts the utr counts data frame produced by reading the exon read
#' counts file.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param gene.data an annotation data frame from the same organism as
#' \code{utr.counts} (such the ones produced by \code{get.annotation}).
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not.
#' @return A named list where names represent samples. Each list member is a also
#' a named list where names correspond to gene ids and members are named vectors.
#' Each vector is named according to the transcripts corresponding to each gene and
#' contains the read counts for each UTR regions. This structure is used for
#' assembling final gene counts in the metaseqr pipeline.
#' @author <NAME>
#' @export
#' @examples
#' @examples
#' \dontrun{
#' data("hg19.exon.data",package="metaseqR")
#' gene.data <- get.annotation("hg19","gene","ensembl")
#' reduced.gene.data <- reduce.gene.data(hg19.exon.counts,gene.data)
#' multic <- check.parallel(0.4)
#' gene.model <- construct.gene.model(hg19.exon.counts,sample.list.hg19,gene.data,
#' multic)
#'}
construct.utr.model <- function(utr.counts,sample.list,gene.data,
multic=FALSE) {
the.counts <- vector("list",length(unlist(sample.list)))
names(the.counts) <- unlist(sample.list,use.names=FALSE)
#if (trans.level=="gene")
the.genes <- as.character(unique(gene.data$gene_id))
#else if (trans.level=="transcript")
# the.genes <- as.character(unique(gene.data$transcript_id))
for (n in names(the.counts))
{
disp(" Separating transcripts (UTR regions) per for ",n,"...")
the.counts[[n]] <- the.genes
names(the.counts[[n]]) <- the.genes
#if (trans.level=="gene") {
the.counts[[n]] <- wapply(multic,the.counts[[n]],function(x,d,n) {
tmp <- d[which(d$gene_id==x),c("start","end","transcript_id",n)]
xx <- tmp[,n]
yy <- tmp$end - tmp$start
names(xx) <- names(yy) <- tmp$transcript_id
return(list(count=xx,length=yy))
},utr.counts,n)
#}
#else if (trans.level=="transcript") {
# the.counts[[n]] <- wapply(multic,the.counts[[n]],function(x,d,n) {
# tmp <- d[which(d$transcript_id==x),
# c("start","end","transcript_id",n)]
# xx <- tmp[,n]
# yy <- tmp$end - tmp$start
# names(xx) <- names(yy) <- tmp$transcript_id
# return(list(count=xx,length=yy))
# },utr.counts,n)
#}
}
return(the.counts)
}
#' Reduce the gene annotation in case of not all chromosomes present in counts
#'
#' This function reduces the gene annotation in case of exon reads and when the
#' data to be analyzed do not contain all the standard chromosomes of the genome
#' under investigation. This can greatly reduce processing time in these cases.
#'
#' @param exon.data the exon annotation already reduced to the size of the input
#' exon counts table.
#' @param gene.data an annotation data frame from the same organism as
#' \code{exon.counts} (such the ones produced by \code{get.annotation}).
#' @return The \code{gene.data} annotation, reduced to have the same chromosomes
#' as in \code{exon.data}, or the original \code{gene.data} if \code{exon.data}
#' does contain the standard chromosomes.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' data("hg19.exon.data",package="metaseqR")
#' gene.data <- get.annotation("hg19","gene","ensembl")
#' reduced.gene.data <- reduce.gene.data(hg19.exon.counts,gene.data)
#'}
reduce.gene.data <- function(exon.data,gene.data) {
exon.chrs <- unique(as.character(exon.data$chromosome))
gene.chrs <- unique(as.character(gene.data$chromosome))
if (length(exon.chrs)!=length(gene.chrs)) {
m <- match(gene.data$chromosome,exon.chrs)
gene.data <- gene.data[which(!is.na(m)),]
}
return(gene.data)
}
# Initialize environment
#
# Initializes metaseqr environmental variables. Internal use only.
#
# @author <NAME>
#init.envar <- function() {
# HOME <<- system.file(package="metaseqR")
# SCRIPT <<- file.path(HOME,"R")
# TEMPLATE <<- HOME
# ANNOTATION <<- file.path(HOME,"data")
#}
<file_sep>/man/estimate.aufc.weights.Rd
\name{estimate.aufc.weights}
\alias{estimate.aufc.weights}
\title{Estimate AUFC weights}
\usage{
estimate.aufc.weights(counts, normalization,
statistics, nsim = 10, N = 10000,
samples = c(3, 3), ndeg = c(500, 500),
top = 500, model.org = "mm9", fc.basis=1.5,
seed = NULL, draw.fpc = FALSE, multic = FALSE,
...)
}
\arguments{
\item{counts}{the real raw counts table from
which the simulation parameters will be
estimated. It must not be normalized and must
contain only integer counts, without any other
annotation elements and unique gene identifiers
as the rownames attribute.}
\item{normalization}{same as \code{normalization}
in \code{\link{metaseqr}}.}
\item{statistics}{same as \code{statistics} in
\code{\link{metaseqr}}.}
\item{nsim}{the number of simulations to perform
to estimate the weights. It default to 10.}
\item{N}{the number of genes to produce.
See \code{\link{make.sim.data.sd}}.}
\item{samples}{a vector with 2 integers, which
are the number of samples for each condition
(two conditions currently supported).}
\item{ndeg}{a vector with 2 integers, which
are the number of differentially expressed
genes to be produced. The first element is
the number of up-regulated genes while the
second is the number of down-regulated genes.}
\item{fc.basis}{the minimum fold-change for
deregulation.}
\item{top}{the top \code{top} best ranked
(according to p-value) to use, to calculate
area under the false discovery curve.}
\item{model.org}{the organism from which the
data are derived. It must be one of
\code{\link{metaseqr}} supported organisms.}
\item{seed}{a list of seed for reproducible
simulations. Defaults to \code{NULL}.}
\item{draw.fpc}{draw the averaged false
discovery curves? Default to \code{FALSE}.}
\item{multic}{whether to run in parallel
(if package \code{parallel} is present or
not.}
\item{...}{Further arguments to be passed to
\code{\link{estimate.sim.params}}.}
}
\value{
A vector of weights to be used in
\code{\link{metaseqr}} with the
\code{weights} option.
}
\description{
This function automatically estimates weights
for the \code{"weight"} and \code{"dperm.weight"}
options of metaseqR for combining p-values from
multiple statistical tests. It creates simulated
dataset based on real data and then performs
statistical analysis with metaseqR several times
in order to derive False Discovery Curves. Then,
the average areas under the false discovery curves
are used to construct weights for each algorithm,
according to its performance when using simulated
data.
}
\examples{
\donttest{
data("mm9.gene.data",package="metaseqR")
multic <- check.parallel(0.8)
weights <- estimate.aufc.weights(
counts=as.matrix(mm9.gene.counts[,9:12]),
normalization="edaseq",
statistics=c("deseq","edger"),
nsim=3,N=100,ndeg=c(10,10),top=10,model.org="mm9",
seed=10,multic=multic,libsize.gt=1e+5
)
}
}
\author{
<NAME>
}
<file_sep>/man/get.transcript.utr.attributes.Rd
\name{get.transcript.utr.attributes}
\alias{get.transcript.utr.attributes}
\title{Annotation downloader helper}
\usage{
get.transcript.utr.attributes(org)
}
\arguments{
\item{org}{one of the supported organisms.}
}
\value{
A character vector of Ensembl transcript attributes.
}
\description{
Returns a vector of genomic annotation attributes which
are used by the biomaRt package in order to fetch the
transcript annotation for each organism. It has no
parameters. Internal use.
}
\examples{
tran.attr <- get.transcript.utr.attributes("mm9")
}
\author{
<NAME>
}
<file_sep>/man/read2count.Rd
\name{read2count}
\alias{read2count}
\title{SAM/BAM/BED file reader helper for the metaseqr pipeline}
\usage{
read2count(targets, annotation, file.type = targets$type,
trans.level = "gene", utr.flank = 500,
has.all.fields = FALSE, multic = FALSE)
}
\arguments{
\item{targets}{a named list, the output of
\code{\link{read.targets}}.}
\item{annotation}{see the \code{annotation} argument in
the main \code{\link{metaseqr}} function. The
\code{"annotation"} parameter here is the result of the
same parameter in the main function. See also
\code{\link{get.annotation}}.}
\item{file.type}{the type of raw input files. It can be
\code{"bed"} for BED files or \code{"sam"}, \code{"bam"}
for SAM/BAM files. See the same argument in the main
\code{\link{metaseqr}} function for the case of
auto-guessing.}
\item{utr.flank}{the number of base pairs to flank the
3' UTR of transcripts when analyzing Quant-Seq data.}
\item{trans.level}{see the \code{trans.level} argument
in the main \code{\link{metaseqr}} function.}
\item{has.all.fields}{a logical variable indicating if
all annotation fields used by \code{metaseqr} are
available (that is apart from the main chromosome, start,
end, unique id and strand columns, if also present are
the gene name and biotype columns). The default is
\code{FALSE}.}
\item{multic}{a logical value indicating the presence
of multiple cores. Defaults to \code{FALSE}. Do not
change it if you are not sure whether package parallel
has been loaded or not.}
}
\value{
A data frame with counts for each sample, ready to be
passed to the main \code{\link{metaseqr}} pipeline.
}
\description{
This function is a helper for the \code{metaseqr}
pipeline, for reading SAM/BAM or BED files when a read
counts file is not available.
}
\examples{
\dontrun{
my.targets <- read.targets("my_mm9_study_bam_files.txt")
gene.data <- get.annotation("mm9","gene")
r2c <- read2count(targets=my.targets,
file.type=my.targets$type,annotation=gene.data)
gene.counts <- r2c$counts
libsize.list <- r2s$libsize
}
}
\author{
<NAME>
}
<file_sep>/man/make.venn.counts.Rd
\name{make.venn.counts}
\alias{make.venn.counts}
\title{Helper for Venn diagrams}
\usage{
make.venn.counts(n)
}
\arguments{
\item{n}{the number of the sets used for the Venn
diagram.}
}
\value{
A named list, see descritpion.
}
\description{
This function creates a list with names the arguments of
the Venn diagram construction functions of the R package
VennDiagram and list members are initially \code{NULL}.
They are filled by the \code{\link{diagplot.venn}}
function. Internal use mostly.
}
\examples{
\donttest{
sets <- c("apple","pear","banana")
counts <- make.venn.counts(length(sets))
}
}
\author{
<NAME>
}
<file_sep>/man/combine.maxp.Rd
\name{combine.maxp}
\alias{combine.maxp}
\title{Combine p-values using the maximum p-value}
\usage{
combine.maxp(p)
}
\arguments{
\item{p}{a p-value matrix (rows are genes,
columns are statistical tests).}
}
\value{
A vector of combined p-values.
}
\description{
This function combines p-values from the
various statistical tests supported by
metaseqR by taking the maximum p-value.
}
\examples{
p <- matrix(runif(300),100,3)
pc <- combine.maxp(p)
}
\author{
<NAME>
}
<file_sep>/man/get.arg.Rd
\name{get.arg}
\alias{get.arg}
\title{Argument getter}
\usage{
get.arg(arg.list, arg.name)
}
\arguments{
\item{arg.list}{the initial list of a method's (e.g.
normalization) arguments. Can be created with the
\code{\link{get.defaults}} function.}
\item{arg.name}{the argument name inside the argument
list to fetch its value.}
}
\value{
The argument sub-list.
}
\description{
Get argument(s) from a list of arguments, e.g.
normalization arguments.
}
\examples{
\donttest{
norm.list <- get.defaults("normalization","egder")
a <- get.arg(norm.list,c("main.method","logratioTrim"))
}
}
\author{
<NAME>
}
<file_sep>/man/get.ucsc.annotation.Rd
\name{get.ucsc.annotation}
\alias{get.ucsc.annotation}
\title{UCSC/RefSeq annotation downloader}
\usage{
get.ucsc.annotation(org, type, refdb="ucsc",
multic=FALSE)
}
\arguments{
\item{org}{the organism for which to download
annotation.}
\item{type}{either \code{"gene"} or \code{"exon"}.}
\item{refdb}{either \code{"ucsc"} or
\code{"refseq"}.}
\item{multic}{a logical value indicating the
presence of multiple cores. Defaults to
\code{FALSE}. Do not change it if you are not
sure whether package parallel has been loaded
or not. It is used in the case of
\code{type="exon"} to process the return value of
the query to the UCSC Genome Browser database.}
}
\value{
A data frame with the canonical (not isoforms!) genes or
exons of the requested organism. When
\code{type="genes"}, the data frame has the following
columns: chromosome, start, end, gene_id, gc_content,
strand, gene_name, biotype. When \code{type="exon"} the
data frame has the following columns: chromosome, start,
end, exon_id, gene_id, strand, gene_name, biotype. The
gene_id and exon_id correspond to UCSC or RefSeq gene
and exon accessions respectively. The gene_name corresponds
to HUGO nomenclature gene names.
}
\description{
This function connects to the UCSC Genome Browser public
database and downloads annotation elements (gene
co-ordinates, exon co-ordinates, gene identifications
etc.) for each of the supported organisms, but using UCSC
instead of Ensembl. See the help page of
\code{\link{metaseqr}} for a list of supported organisms.
The function downloads annotation for an organism genes or
exons.
}
\note{
The data frame that is returned contains only "canonical"
chromosomes for each organism. It does not contain
haplotypes or random locations and does not contain
chromosome M. Note also that as the UCSC databases do
not contain biotype classifications like Ensembl. These
will be returned as \code{NA} and as a result, some
quality control plots will not be available.
}
\examples{
\donttest{
hg19.genes <- get.ucsc.annotation("hg19","gene","ucsc")
mm9.exons <- get.ucsc.annotation("mm9","exon","refseq")
}
}
\author{
<NAME>
}
<file_sep>/man/diagplot.roc.Rd
\name{diagplot.roc}
\alias{diagplot.roc}
\title{Create basic ROC curves}
\usage{
diagplot.roc(truth, p, sig = 0.05, x = "fpr",
y = "tpr", output = "x11", path = NULL,
draw = TRUE, ...)
}
\arguments{
\item{truth}{the ground truth differential
expression vector. It should contain only
zero and non-zero elements, with zero denoting
non-differentially expressed genes and non-zero,
differentially expressed genes. Such a vector
can be obtained for example by using the
\code{\link{make.sim.data.sd}} function, which
creates simulated RNA-Seq read counts based on
real data.}
\item{p}{a p-value matrix whose rows correspond
to each element in the \code{truth} vector. If
the matrix has a \code{colnames} attribute, a
legend will be added to the plot using these
names, else a set of column names will be
auto-generated. \code{p} can also be a list or
a data frame.}
\item{sig}{a significance level (0 < \code{sig}
<=1).}
\item{x}{what to plot on x-axis, can be one of
\code{"fpr"}, \code{"fnr"}, \code{"tpr"},
\code{"tnr"} for False Positive Rate, False
Negative Rate, True Positive Rate and True
Negative Rate respectively.}
\item{y}{what to plot on y-axis, same as
\code{x} above.}
\item{output}{one or more R plotting device to
direct the plot result to. Supported mechanisms:
\code{"x11"} (default), \code{"png"}, \code{"jpg"},
\code{"bmp"}, \code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{draw}{boolean to determine whether to
plot the curves or just return the calculated
values (in cases where the user wants the
output for later averaging for example).
Defaults to \code{TRUE} (make plots).}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
A named list with two members. The first member is
a list containing the ROC statistics: \code{TP}
(True Postives), \code{FP} (False Positives),
\code{FN} (False Negatives), \code{TN}
(True Negatives), \code{FPR} (False Positive Rate),
\code{FNR} (False Negative Rate), \code{TPR} (True
Positive Rate), \code{TNR} (True Negative Rate),
\code{AUC} (Area Under the Curve). The second is
the path to the created figure graphic.
}
\description{
This function creates basic ROC curves using a
matrix of p-values (such a matrix can be
derived for example from the result table of
\code{\link{metaseqr}} by subsetting the table
to get the p-values from several algorithms)
given a ground truth vector for differential
expression and a significance level.
}
\examples{
p1 <- 0.001*matrix(runif(300),100,3)
p2 <- matrix(runif(300),100,3)
p <- rbind(p1,p2)
rownames(p) <- paste("gene",1:200,sep="_")
colnames(p) <- paste("method",1:3,sep="_")
truth <- c(rep(1,40),rep(-1,40),rep(0,20),rep(1,10),
rep(2,10),rep(0,80))
names(truth) <- rownames(p)
roc.obj <- diagplot.roc(truth,p)
}
\author{
<NAME>
}
<file_sep>/man/make.report.messages.Rd
\name{make.report.messages}
\alias{make.report.messages}
\title{Initializer of report messages}
\usage{
make.report.messages(lang)
}
\arguments{
\item{lang}{The language of the report. For now, only
english (\code{"en"}) is supported.}
}
\value{
An named list with messages for each input option.
}
\description{
Initializes metaseqR report tmeplate messages output.
Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/combine.simes.Rd
\name{combine.simes}
\alias{combine.simes}
\title{Combine p-values with Simes' method}
\usage{
combine.simes(p)
}
\arguments{
\item{p}{a p-value matrix (rows are genes,
columns are statistical tests).}
}
\value{
A vector of combined p-values.
}
\description{
This function combines p-values from the
various statistical tests supported by
metaseqR using the Simes' method (see
reference in the main \code{\link{metaseqr}}
help page or in the vignette).
}
\examples{
p <- matrix(runif(300),100,3)
pc <- combine.simes(p)
}
\author{
<NAME>
}
<file_sep>/man/check.contrast.format.Rd
\name{check.contrast.format}
\alias{check.contrast.format}
\title{Contrast validator}
\usage{
check.contrast.format(cnt, sample.list)
}
\arguments{
\item{cnt}{contrasts vector.}
\item{sample.list}{the input sample list.}
}
\description{
Checks if the contrast vector follows the specified
format. Internal use only.
}
\examples{
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
cnt <- c("A_vs_B") # Will work
#cnt <- c("A_vs_C") ## Will throw error!
check.contrast.format(cnt,sample.list)
}
\author{
<NAME>
}
<file_sep>/man/fisher.method.Rd
\name{fisher.method}
\alias{fisher.method}
\title{
Perform Fisher's Method for combining p-values
}
\description{
Function for combining p-values by performing Fisher's method.
The approach as described by Fisher's combines p-values to a statistic
\deqn{S = -2 \sum^k \log p}, which follows a \eqn{\chi^2} distribution
with \eqn{2k} degrees of freedom.
}
\usage{
fisher.method(pvals, method = c("fisher"), p.corr = c("bonferroni",
"BH", "none"), zero.sub = 1e-05, na.rm = FALSE, mc.cores=NULL)
}
\arguments{
\item{pvals}{
A matrix or data.frame containing the p-values from the single tests
}
\item{method}{
A string indicating how to combine the p-values for deriving a
sumary p-value. Currently only the classical approach described by
Fisher is implemented.
}
\item{p.corr}{
Method for correcting the summary p-values. BH: Benjamini-Hochberg
(default); Bonferroni's method or no ('none') correction are currently
supported.
}
\item{zero.sub}{
Replacement for p-values of 0
}
\item{na.rm}{
A flag indicating whether NA values should be removed from the analysis.
}
\item{mc.cores}{
Currently ignored
}
}
\details{
As \code{log(0)} results in \code{Inf} we replace p-values of 0 by default
with a small float. If you want to keep them as 0 you have to provide 0
as a parameter in \code{zero.sub}.
Note that only p-values between 0 and 1 are allowed to be passed to this
method.
}
\value{
This method returns a data.frame containing the following columns
\item{S }{The statistic}
\item{num.p }{The number of p-values used to calculate S}
\item{p.value }{The overall p-value}
\item{p.adj}{The adjusted p-value}
}
\note{
This function was copied from the CRAN package MADAM which is no longer
maintained. Recognition goes to the original author(s) below.
}
\references{
<NAME>. (1925). Statistical Methods for Research Workers. Oliver and
Boyd (Edinburgh).
Moreau, Y.et al. (2003). Comparison and meta-analysis of microarray data:
from the bench to the computer desk. Trends in Genetics, 19(10), 570-577.
}
\author{
<NAME> <<EMAIL>>
}
\seealso{
\code{\link{fisher.method.perm}}
}
\examples{
set.seed(123)
pp <- matrix(c(runif(20),c(0.001,0.02,0.03,0.001)), ncol=4)
pp[2,3] <- NA
fisher.method(pp) #returns one NA row
fisher.method(pp, na.rm=TRUE) #ignore NA entry in that row
}
<file_sep>/man/get.host.Rd
\name{get.host}
\alias{get.host}
\title{Annotation downloader helper}
\usage{
get.host(org)
}
\arguments{
\item{org}{the organism for which to return the host
address.}
}
\value{
A string with the host address.
}
\description{
Returns the appropriate Ensembl host address to get
different versions of annotation from. Internal use.
}
\examples{
mm9.hist <- get.host("mm9")
}
\author{
<NAME>
}
<file_sep>/man/estimate.sim.params.Rd
\name{estimate.sim.params}
\alias{estimate.sim.params}
\title{Estimate negative binomial parameters from
real data}
\usage{
estimate.sim.params(real.counts, libsize.gt = 3e+6,
rowmeans.gt = 5,eps = 1e-11,
restrict.cores = 0.1, seed = 42, draw = FALSE)
}
\arguments{
\item{real.counts}{a text tab-delimited file
with real RNA-Seq data. The file should
strictly contain a unique gene name (e.g. Ensembl
accession) in the first column and all other
columns should contain read counts for each
gene. Each column must be named with a unique
sample identifier. See examples in the ReCount
database
\url{http://bowtie-bio.sourceforge.net/recount/}.}
\item{libsize.gt}{a library size below which
samples are excluded from parameter estimation
(default: 3000000).}
\item{rowmeans.gt}{a row means (mean counts
over samples for each gene) below which
genes are excluded from parameter estimation
(default: 5).}
\item{eps}{the tolerance for the convergence
of \code{\link{optimize}} function. Defaults
to 1e-11.}
\item{restrict.cores}{in case of parallel
optimization, the fraction of the available
cores to use.}
\item{seed}{a seed to use with random number
generation for reproducibility.}
\item{draw}{boolean to determine whether to
plot the estimated simulation parameters
(mean and dispersion) or not. Defaults to
\code{FALSE} (do not draw a mean-dispersion
scatterplot).}
}
\value{
A named list with two members: \code{mu.hat}
which contains negative binomial mean
estimates and \code{phi.hat} which contains
dispersion estimates.
}
\description{
This function reads a read counts table
containing real RNA-Seq data (preferebly
with more than 20 samples so as to get as
much accurate as possible estimations) and
calculates a population of count means and
dispersion parameters which can be used to
simulate an RNA-Seq dataset with synthetic
genes by drawing from a negative binomial
distribution. This function works in the
same way as described in (Soneson and
Delorenzi, BMC Bioinformatics, 2013) and
(Robles et al., BMC Genomics, 2012).
}
\examples{
\donttest{
# Dowload locally the file "bottomly_read_counts.txt" from
# the ReCount database
download.file(paste("http://bowtie-bio.sourceforge.net/",
"recount/countTables/bottomly_count_table.txt",sep=""),
destfile="~/bottomly_count_table.txt")
# Estimate simulation parameters
par.list <- estimate.sim.params("~/bottomly_count_table.txt")
}
}
\author{
<NAME>
}
<file_sep>/man/reduce.gene.data.Rd
\name{reduce.gene.data}
\alias{reduce.gene.data}
\title{Reduce the gene annotation in case of not all chromosomes present in counts}
\usage{
reduce.gene.data(exon.data, gene.data)
}
\arguments{
\item{exon.data}{the exon annotation already reduced to
the size of the input exon counts table.}
\item{gene.data}{an annotation data frame from the same
organism as \code{exon.counts} (such the ones produced by
\code{get.annotation}).}
}
\value{
The \code{gene.data} annotation, reduced to have the same
chromosomes as in \code{exon.data}, or the original
\code{gene.data} if \code{exon.data} do contain the
standard chromosomes.
}
\description{
This function reduces the gene annotation in case of exon
reads and when the data to be analyzed do not contain all
the standard chromosomes of the genome under
investigation. This can greatly reduce processing time in
these cases.
}
\examples{
\donttest{
data("hg19.exon.data",package="metaseqR")
gene.data <- get.annotation("hg19","gene","ensembl")
reduced.gene.data <- reduce.gene.data(hg19.exon.counts,
gene.data)
}
}
\author{
<NAME>
}
<file_sep>/man/get.weights.Rd
\name{get.weights}
\alias{get.weights}
\title{Get precalculated statistical test
weights}
\usage{
get.weights(org = c("human", "chimpanzee", "mouse",
"fruitfly", "arabidopsis"))
}
\arguments{
\item{org}{\code{"human"}, \code{"chimpanzee"},
\code{"mouse"}, \code{"fruitfly"} or
\code{"arabidopsis"}.}
}
\value{
A named vector of convex weights.
}
\description{
This function returns pre-calculated weights
for human, chimpanzee, mouse, fruitfly and
arabidopsis based on the performance of
simulated datasets estimated from real data
from the ReCount database
(\url{http://bowtie-bio.sourceforge.net/recount/}).
Currently pre-calculated weights are available only
when all six statistical tests are used and for
normalization with EDASeq. For other combinations,
use the \code{\link{estimate.aufc.weights}} function.
}
\examples{
wh <- get.weights("human")
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.sim.R
#' Estimate AUFC weights
#'
#' This function automatically estimates weights for the \code{"weight"} and
#' \code{"dperm.weight"} options of metaseqR for combining p-values from multiple
#' statistical tests. It creates simulated dataset based on real data and then
#' performs statistical analysis with metaseqR several times in order to derive
#' False Discovery Curves. Then, the average areas under the false discovery curves
#' are used to construct weights for each algorithm, according to its performance
#' when using simulated data.
#'
#' @param counts the real raw counts table from which the simulation parameters
#' will be estimated. It must not be normalized and must contain only integer
#' counts, without any other annotation elements and unique gene identifiers as
#' the rownames attribute.
#' @param normalization same as \code{normalization} in \code{link{metaseqr}}.
#' @param statistics same as \code{statistics} in \code{link{metaseqr}}.
#' @param nsim the number of simulations to perform to estimate the weights. It
#' default to 10.
#' @param N the number of genes to produce. See \code{link{make.sim.data.sd}}.
#' @param samples a vector with 2 integers, which are the number of samples for
#' each condition (two conditions currently supported).
#' @param ndeg a vector with 2 integers, which are the number of differentially
#' expressed genes to be produced. The first element is the number of up-regulated
#' genes while the second is the number of down-regulated genes.
#' @param fc.basis the minimum fold-change for deregulation.
#' @param top the top \code{top} best ranked (according to p-value) to use, to
#' calculate area under the false discovery curve.
#' @param model.org the organism from which the data are derived. It must be one
#' of \code{\link{metaseqr}} supported organisms.
#' @param seed a number to be used as seed for reproducible simulations. Defaults
#' to \code{NULL} (NOT reproducible results!).
#' @param draw.fpc draw the averaged false discovery curves? Default to \code{FALSE}.
#' @param multic whether to run in parallel (if package \code{parallel} is present
#' or not.
#' @param ... Further arguments to be passed to \code{\link{estimate.sim.params}}.
#' @value A vector of weights to be used in \link{\code{metaseqr}} with the
#' \code{weights} option.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data("mm9.gene.data",package="metaseqR")
#' multic <- check.parallel(0.8)
#' weights <- estimate.aufc.weights(
#' counts=as.matrix(mm9.gene.counts[,9:12]),
#' normalization="edaseq",
#' statistics=c("deseq","edger"),
#' nsim=3,N=100,ndeg=c(10,10),top=10,model.org="mm9",
#' seed=100,multic=multic,libsize.gt=1e+5
#' )
#'}
estimate.aufc.weights <- function(counts,normalization,statistics,nsim=10,
N=10000,samples=c(3,3),ndeg=c(500,500),top=500,model.org="mm9",fc.basis=1.5,
seed=NULL,draw.fpc=FALSE,multic=FALSE,...) {
if (!require(zoo))
stopwrap("R pacakage zoo is required in order to estimate AUFC ",
"weights!")
if (is.null(seed)) {
seed.start <- round(100*runif(1))
seed.end <- seed.start + nsim - 1
seed <- as.list(seed.start:seed.end)
}
else {
set.seed(seed)
seed.start <- round(100*runif(1))
seed.end <- seed.start + nsim - 1
seed <- as.list(seed.start:seed.end)
}
if (ncol(counts)<4)
stopwrap("Cannot estimate AUFC weights with an initial dataset with ",
"less than 4 samples!")
else if (ncol(counts)>=4 && ncol(counts)<10) {
set.seed(seed.start)
reind <- sample(1:ncol(counts),20,replace=TRUE)
counts <- counts[,reind]
}
par.list <- estimate.sim.params(counts,...)
disp("Running simulations... This procedure requires time... Please ",
"wait...")
if (nsim==1) {
sim.results <- lapply(seed[1],function(x,normalization,statistics,N,
par.list,samples,ndeg,fc.basis,model.org) {
D <- make.sim.data.sd(N=N,param=par.list,samples=samples,ndeg=ndeg,
fc.basis=fc.basis,model.org=model.org,seed=x)
dd <- D$simdata
if (!is.null(model.org)) {
tmp <- metaseqr(
counts=dd,
sample.list=list(G1=paste("G1_rep",1:samples[1],sep=""),
G2=paste("G2_rep",1:samples[2],sep="")),
contrast=c("G1_vs_G2"),
annotation="embedded",
id.col=4,
gc.col=5,
name.col=7,
bt.col=8,
org=model.org,
count.type="gene",
normalization=normalization,
statistics=statistics,
meta.p="simes",
fig.format="png",
preset="all.basic",
export.where=tempdir(),
qc.plots=NULL,
report=FALSE,
run.log=FALSE,
out.list=TRUE,
restrict.cores=0.1
)
}
else {
tmp <- metaseqr(
counts=dd,
sample.list=list(G1=paste("G1_rep",1:samples[1],sep=""),
G2=paste("G2_rep",1:samples[2],sep="")),
contrast=c("G1_vs_G2"),
annotation="embedded",
id.col=4,
gc.col=5,
name.col=7,
bt.col=8,
count.type="gene",
normalization=normalization,
statistics=statistics,
meta.p="simes",
fig.format="png",
preset="all.basic",
export.where=tempdir(),
qc.plots=NULL,
report=FALSE,
run.log=FALSE,
out.list=TRUE,
restrict.cores=0.1
)
}
# Retrieve several p-values
p.list <- vector("list",length(statistics))
for (s in statistics) {
field <- paste("p-value",s,sep="_")
p.list[[s]] <- tmp$data[[1]][,field]
names(p.list[[s]]) <- rownames(tmp$data[[1]])
}
p.matrix <- do.call("cbind",p.list)
return(list(simdata=D,pvalues=p.matrix))
},normalization,statistics,N,par.list,samples,ndeg,fc.basis,model.org)
}
else {
sim.results <- wapply(multic,seed,function(x,normalization,statistics,N,
par.list,samples,ndeg,fc.basis,model.org) {
D <- make.sim.data.sd(N=N,param=par.list,samples=samples,ndeg=ndeg,
fc.basis=fc.basis,model.org=model.org,seed=x)
dd <- D$simdata
if (!is.null(model.org)) {
tmp <- metaseqr(
counts=dd,
sample.list=list(G1=paste("G1_rep",1:samples[1],sep=""),
G2=paste("G2_rep",1:samples[2],sep="")),
contrast=c("G1_vs_G2"),
annotation="embedded",
id.col=4,
gc.col=5,
name.col=7,
bt.col=8,
org=model.org,
count.type="gene",
normalization=normalization,
statistics=statistics,
meta.p="simes",
fig.format="png",
preset="all.basic",
export.where=tempdir(),
qc.plots=NULL,
report=FALSE,
run.log=FALSE,
out.list=TRUE,
restrict.cores=0.1
)
}
else {
tmp <- metaseqr(
counts=dd,
sample.list=list(G1=paste("G1_rep",1:samples[1],sep=""),
G2=paste("G2_rep",1:samples[2],sep="")),
contrast=c("G1_vs_G2"),
annotation="embedded",
id.col=4,
gc.col=5,
name.col=7,
bt.col=8,
count.type="gene",
normalization=normalization,
statistics=statistics,
meta.p="simes",
fig.format="png",
preset="all.basic",
export.where=tempdir(),
qc.plots=NULL,
report=FALSE,
run.log=FALSE,
out.list=TRUE,
restrict.cores=0.1
)
}
# Retrieve several p-values
p.list <- vector("list",length(statistics))
for (s in statistics) {
field <- paste("p-value",s,sep="_")
p.list[[s]] <- tmp$data[[1]][,field]
names(p.list[[s]]) <- rownames(tmp$data[[1]])
}
p.matrix <- do.call("cbind",p.list)
return(list(simdata=D,pvalues=p.matrix))
},normalization,statistics,N,par.list,samples,ndeg,fc.basis,model.org)
}
disp("Estimating AUFC weights... Please wait...")
#fpc.obj <- wapply(multic,sim.results,function(x) {
fpc.obj <- lapply(sim.results,function(x) {
true.de <- x$simdata$truedeg
names(true.de) <- rownames(x$simdata$simdata)
p.matrix <- x$pvalues
true.de <- true.de[rownames(p.matrix)]
fdc <- diagplot.ftd(true.de,p.matrix,type="fpc",draw=FALSE)
})
avg.fpc <- diagplot.avg.ftd(fpc.obj,draw=draw.fpc)
x <- 1:top
aufc <- apply(avg.fpc$avg.ftdr$means[1:top,],2,function(x,i) {
return(sum(diff(i)*rollmean(x,2)))
},x)
weight.aufc <- (sum(aufc)/aufc)/sum(sum(aufc)/aufc)
return(weight.aufc)
}
#' Create simulated counts using TCC package
#'
#' This function creates simulated RNA-Seq gene expression datasets using the
#' \code{simulateReadCounts} function from the Bioconductor
#' package TCC and it adds simulated annoation elements. For further information
#' please consult the TCC package documentation. Note that the produced data are
#' based in an Arabidopsis dataset.
#'
#' @param ... parameters to the \code{simulateReadCounts} function.
#' @return A list with the following members: \code{simdata} holding the simulated
#' dataset complying with metaseqr requirements, and \code{simparam} holding the
#' simulation parameters (see TCC documentation).
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' dd <- make.sim.data(Ngene=10000,PDEG=0.2,DEG.assign=c(0.9,0.1),
#' DEG.foldchange=c(5,5),replicates=c(3,3))
#' head(dd$simdata)
#'}
make.sim.data.tcc <- function(...) {
if (suppressWarnings(!require(TCC)))
stopwrap("Bioconductor package TCC is required to create simulated data!")
#tcc <- simulateReadCounts(Ngene=Ngene,PDEG=PDEG,DEG.assign=DEG.assign,
# DEG.foldchange=DEG.foldchange,replicates=replicates)
tcc <- TCC::simulateReadCounts(...)
n <- nrow(tcc$count)
# Now we have to simulate annotation
chromosome <- paste("chr",1+round(20*runif(n)),sep="")
start <- 1 + round(1e+6*runif(n))
end <- start + 250 + round(1e+6*runif(n))
gene_id <- gene_name <- rownames(tcc$count)
gc_content <- runif(n)
strand <- sample(c("+","-"),n,replace=TRUE)
biotype <- sample(paste("biotype",1:10),n,replace=TRUE)
sim.data <- data.frame(
chromosome=chromosome,
start=start,
end=end,
gene_id=gene_id,
gc_content=gc_content,
strand=strand,
gene_name=gene_name,
biotype=biotype
)
sim.data <- cbind(sim.data,tcc$count)
return(list(simdata=sim.data,simparam=tcc$simulation))
}
#' Create simulated counts using the Soneson-Delorenzi method
#'
#' This function creates simulated RNA-Seq gene expression datasets using the
#' method presented in (Soneson and Delorenzi, BMC Bioinformatics, 2013). For the
#' time being, it creates only simulated datasets with two conditions.
#'
#' @param N the number of genes to produce.
#' @param param a named list with negative binomial parameter sets to sample from.
#' The first member is the mean parameter to sample from (\code{mu.hat}} and the
#' second the dispersion (\code{phi.hat}). This list can be created with the
#' \code{\link{estimate.sim.params}} function.
#' @param samples a vector with 2 integers, which are the number of samples for
#' each condition (two conditions currently supported).
#' @param ndeg a vector with 2 integers, which are the number of differentially
#' expressed genes to be produced. The first element is the number of up-regulated
#' genes while the second is the number of down-regulated genes.
#' @param fc.basis the minimum fold-change for deregulation.
#' @param libsize.range a vector with 2 numbers (generally small, see the default),
#' as they are multiplied with \code{libsize.mag}.
#' These numbers control the library sized of the synthetic data to be produced.
#' @param libsize.mag a (big) number to multiply the \code{libsize.range} to
#' produce library sizes.
#' @param model.org the organism from which the real data are derived from. It
#' must be one of the supported organisms (see the main \code{\link{metaseqr}}
#' help page). It is used to sample real values for GC content.
#' @param sim.length.bias a boolean to instruct the simulator to create genes
#' whose read counts is proportional to their length. This is achieved by sorting
#' in increasing order the mean parameter of the negative binomial distribution
#' (and the dispersion according to the mean) which will cause an increasing gene
#' count length with the sampling. The sampled lengths are also sorted so that in
#' the final gene list, shorter genes have less counts as compared to the longer
#' ones. The default is FALSE.
#' @param seed a seed to use with random number generation for reproducibility.
#' @return A named list with two members. The first member (\code{simdata})
#' contains the synthetic dataset
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' # File "bottomly_read_counts.txt" from the ReCount database
#' download.file("http://bowtie-bio.sourceforge.net/recount/countTables/bottomly_count_table.txt",
#' destfile="~/bottomly_count_table.txt")
#' N <- 10000
#' par.list <- estimate.sim.params("~/bottomly_read_counts.txt")
#' sim <- make.sim.data.sd(N,par.list)
#' synth.data <- sim$simdata
#' true.deg <- which(sim$truedeg!=0)
#'}
make.sim.data.sd <- function(N,param,samples=c(5,5),ndeg=rep(round(0.1*N),2),
fc.basis=1.5,libsize.range=c(0.7,1.4),libsize.mag=1e+7,model.org=NULL,
sim.length.bias=FALSE,seed=NULL) {
if (!is.null(model.org)) {
model.org <- tolower(model.org)
check.text.args("model.org",model.org,c("hg18","hg19","mm9","mm10",
"rno5","dm3","danrer7","pantro4","tair10"),multiarg=FALSE)
ann <- get.annotation(model.org,"gene")
real.gc <- as.numeric(ann$gc_content)
real.start <- as.numeric(ann$start)
real.end <- as.numeric(ann$end)
real.strand <- as.character(ann$strand)
}
mu.hat <- param$mu.hat
phi.hat <- param$phi.hat
if (!is.null(seed)) set.seed(seed)
if (sim.length.bias) {
sind <- sort(mu.hat,index.return=TRUE)$ix
mu.hat <- mu.hat[sind]
phi.hat <- phi.hat[sind]
if (length(mu.hat)>=N)
ii <- sort(sample(1:length(mu.hat),N))
else
ii <- sort(sample(1:length(mu.hat),N,replace=TRUE))
}
else {
if (length(mu.hat)>=N)
ii <- sample(1:length(mu.hat),N)
else
ii <- sample(1:length(mu.hat),N,replace=TRUE)
}
s1 <- samples[1]
s2 <- samples[2]
if (!is.null(seed)) set.seed(seed)
L1 <- round(libsize.mag*runif(s1,min=libsize.range[1],
max=libsize.range[2]))
if (!is.null(seed)) set.seed(2*seed)
L2 <- round(libsize.mag*runif(s2,min=libsize.range[1],
max=libsize.range[2]))
lambda.1 <- do.call("cbind",rep(list(mu.hat[ii]),s1))
mu.1 <- sweep(lambda.1,2,L1/sum(lambda.1[,1]),"*")
sim.1 <- matrix(0,N,s1)
for (j in 1:s1) {
if (!is.null(seed)) set.seed(seed+j)
sim.1[,j] <- rnbinom(N,size=1/phi.hat[ii],mu=mu.1[,j])
}
v <- numeric(N)
if (sum(ndeg)>0) {
if (!is.null(seed)) set.seed(seed)
i.updown <- sample(1:length(v),sum(ndeg))
reg.dir <- rep(c(1,-1),c(ndeg[1],ndeg[2]))
v[i.updown] <- reg.dir
if (!is.null(seed)) set.seed(seed+19051980)
lambda.2 <- ((fc.basis + rexp(N))^v)*lambda.1
mu.2 <- sweep(lambda.2,2,L2/sum(lambda.2[,1]),"*")
sim.2 <- matrix(0,N,s2)
for (j in 1:s2)
sim.2[,j] <- rnbinom(N,size=1/phi.hat[ii],mu=mu.2[,j])
}
else {
if (!is.null(seed)) set.seed(seed+19051980)
lambda.2 <- lambda.1
mu.2 <- sweep(lambda.2,2,L2/sum(lambda.2[,1]),"*")
sim.2 <- matrix(0,N,s2)
for (j in 1:s2)
sim.2[,j] <- rnbinom(N,size=1/phi.hat[ii],mu=mu.2[,j])
}
# Now we have to simulate annotation
if (!is.null(seed)) set.seed(seed)
chromosome <- paste("chr",1+round(20*runif(N)),sep="")
gene_id <- gene_name <- paste("gene",1:N,sep="_")
if (!is.null(model.org)) {
if (!is.null(seed)) set.seed(seed)
if (length(real.gc)>=N)
sample.ind <- sample(1:length(real.gc),N)
else
sample.ind <- sample(1:length(real.gc),N,replace=TRUE)
gc_content <- real.gc[sample.ind]
start <- real.start[sample.ind]
end <- real.end[sample.ind]
strand <- real.strand[sample.ind]
if (sim.length.bias) {
lenix <- sort(end-start,index.return=TRUE)$ix
start <- start[lenix]
end <- end[lenix]
gc_content <- gc_content[lenix]
strand <- strand[lenix]
}
}
else {
if (!is.null(seed)) set.seed(seed)
gc_content <- runif(N)
if (!is.null(seed)) set.seed(seed)
start <- 1 + round(1e+6*runif(N))
if (!is.null(seed)) set.seed(seed)
end <- start + 250 + round(1e+6*runif(N))
if (!is.null(seed)) set.seed(seed)
strand <- sample(c("+","-"),N,replace=TRUE)
if (sim.length.bias) {
lenix <- sort(end-start,index.return=TRUE)$ix
start <- start[lenix]
end <- end[lenix]
gc_content <- gc_content[lenix]
strand <- strand[lenix]
}
}
if (!is.null(seed)) set.seed(seed)
biotype <- sample(paste("biotype",1:10),N,replace=TRUE)
sim.data <- data.frame(
chromosome=chromosome,
start=start,
end=end,
gene_id=gene_id,
gc_content=gc_content,
strand=strand,
gene_name=gene_name,
biotype=biotype
)
colnames(sim.1) <- paste("G1_rep",1:s1,sep="")
colnames(sim.2) <- paste("G2_rep",1:s2,sep="")
rownames(sim.1) <- rownames(sim.2) <- names(v) <- gene_id
return(list(simdata=cbind(sim.data,sim.1,sim.2),truedeg=v))
}
#' Estimate negative binomial parameters from real data
#'
#' This function reads a read counts table containing real RNA-Seq data (preferebly
#' with more than 20 samples so as to get as much accurate as possible estimations)
#' and calculates a population of count means and dispersion parameters which can
#' be used to simulate an RNA-Seq dataset with synthetic genes by drawing from a
#' negative binomial distribution. This function works in the same way as described
#' in (<NAME>, BMC Bioinformatics, 2013) and (<NAME> al., BMC
#' Genomics, 2012).
#'
#' @param real.counts a text tab-delimited file with real RNA-Seq data. The file
#' should strictly contain a unique gene name (e.g. Ensembl accession) in the
#' first column and all other columns should contain read counts for each gene.
#' Each column must be named with a unique sample identifier. See examples in the
#' ReCount database \link{http://bowtie-bio.sourceforge.net/recount/}.
#' @param libsize.gt a library size below which samples are excluded from parameter
#' estimation (default: 3000000).
#' @param rowmeans.gt a row means (mean counts over samples for each gene) below
#' which genes are excluded from parameter estimation (default: 5).
#' @param eps the tolerance for the convergence of \code{\link{optimize}} function.
#' Defaults to 1e-11.
#' @param restrict.cores in case of parallel optimization, the fraction of the
#' available cores to use.
#' @param seed a seed to use with random number generation for reproducibility.
#' @param draw boolean to determine whether to plot the estimated simulation
#' parameters (mean and dispersion) or not. Defaults to \code{FALSE} (do not draw
#' a mean-dispersion scatterplot).
#' @return A named list with two members: \code{mu.hat} which contains negative
#' binomial mean estimates and \code{phi.hat} which contains dispersion.
#' estimates
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' # Dowload locally the file "bottomly_count_table.txt" from the ReCount datbase
#' download.file("http://bowtie-bio.sourceforge.net/recount/countTables/bottomly_count_table.txt",
#' destfile="~/bottomly_count_table.txt")
#' # Estimate simulation parameters
#' par.list <- estimate.sim.params("~/bottomly_count_table.txt")
#'}
estimate.sim.params <- function(real.counts,libsize.gt=3e+6,rowmeans.gt=5,
eps=1e-11,restrict.cores=0.1,seed=42,draw=FALSE) {
multic <- check.parallel(restrict.cores)
if (is.data.frame(real.counts))
mat <- as.matrix(real.counts)
else if (is.matrix(real.counts))
mat <- real.counts
else if (file.exists(real.counts)) {
real.data <- read.delim(real.counts,row.names=1)
mat <- as.matrix(real.data)
}
else
stopwrap("The input count data must be either a file, a matrix or a ",
"data frame!")
low.lib <- which(apply(mat,2,sum)<libsize.gt)
if (length(low.lib)==ncol(mat))
stopwrap("Cannot estimate simulation parameters as the library sizes ",
"are too small! Try lowering the value of the libsize.gt ",
"parameter...")
if (length(low.lib)>0)
mat <- mat[,-low.lib]
disp("Downsampling counts...")
dmat <- downsample.counts(mat,seed)
low.co <- which(apply(dmat,1,
function(x) if (mean(x)<5) TRUE else FALSE))
if (length(low.co)>0)
dmat <- dmat[-low.co,]
mu.hat <- apply(dmat,1,mean)
disp("Estimating initial dispersion population...")
phi.est <- apply(dmat,1,function(x) {
m <- mean(x)
v <- var(x)
phi <- (v-m)/m^2
return(phi)
})
phi.ind <- which(phi.est>0)
phi.est <- phi.est[phi.ind]
dmat <- dmat[phi.ind,]
disp("Estimating dispersions using log-likelihood...\n")
init <- wapply(multic,seq_along(1:nrow(dmat)),function(i,d,p) {
list(y=d[i,],h=p[i])
},dmat,phi.est)
phi.hat <- unlist(wapply(multic,init,function(x,eps) {
optimize(mlfo,c(x$h-1e-2,x$h+1e-2),y=x$y,tol=eps)$minimum
},eps))
if (draw) {
dev.new()
plot(log10(mu.hat[phi.ind]),log10(phi.hat),col="blue",pch=20,cex=0.5,
xlab="",ylab="")
title(xlab="mean",ylab="dispesion",font=2,cex=0.9)
grid()
}
return(list(mu.hat=mu.hat[phi.ind],phi.hat=phi.hat))
}
#' Downsample read counts
#'
#' This function downsamples the library sizes of a read counts table to the lowest
#' library size, according to the methdology used in (Soneson and Delorenzi,
#' BMC Bioinformatics, 2013).
#'
#' @param counts the read counts table which is subjected to downsampling.
#' @param seed random seed for reproducible downsampling.
#' @return The downsampled counts matrix.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' # Dowload locally the file "bottomly_read_counts.txt" from
#' # the ReCount database
#' download.file(paste("http://bowtie-bio.sourceforge.net/",
#' "recount/countTables/bottomly_count_table.txt",sep=""),
#' destfile="~/bottomly_count_table.txt")
#' M <- as.matrix(read.delim("~/bottomly_count_table.txt",row.names=1))
#' D <- downsample.counts(M)
#'}
downsample.counts <- function(counts,seed=42) {
lib.sizes <- apply(counts,2,sum)
target.size <- min(lib.sizes)
to.remove <- lib.sizes-target.size
ii <- which(to.remove>0)
dcounts <- counts
for (i in ii) {
tmp <- round(to.remove[i]*(counts[,i]/sum(counts[,i])))
victim.size <- sum(tmp)
if (victim.size>to.remove[i]) {
dif <- victim.size - to.remove[i]
#victims <- sample(1:length(tmp),dif)
victims <- sort(tmp,decreasing=TRUE,index.return=TRUE)$ix[1:dif]
tmp[victims] <- tmp[victims] - 1
}
else if (victim.size<to.remove[i]) {
dif <- to.remove[i] - victim.size
#victims <- sample(1:length(tmp),dif)
victims <- sort(tmp,decreasing=TRUE,index.return=TRUE)$ix[1:dif]
tmp[victims] <- tmp[victims] + 1
}
dcounts[,i] <- dcounts[,i] - tmp
}
return(dcounts)
}
#' MLE dispersion estimate
#'
#' MLE function used to estimate negative binomial dispersions from real RNA-Seq
#' data, as in (<NAME> Delorenzi, BMC Bioinformatics, 2013) and (Robles et al.,
#' BMC Genomics, 2012). Internal use.
#'
#' @param phi the parameter to be optimized.
#' @param y count samples used to perform the optimization.
#' @return objective function value.
#' @author <NAME>
#' @examples
#' \dontrun{
#' # Not yet available
#'}
mlfo <- function(phi,y) {
N <- length(y)
mu <- mean(y)
-(sum(lgamma(y+1/phi)) - N*lgamma(1/phi) - sum(lgamma(y+1)) +
sum(y*log(mu*phi/(1+mu*phi))) - (N/phi)*log(1+mu*phi))
}
#' Create counts matrix permutations
#'
#' This function creates a permuted read counts matrix based on the \code{contrast}
#' argument (to define new virtual contrasts of the same number) and on the
#' \code{sample.list} to derive the number of samples for each virtual condition.
#' It is a helper for the \code{\link{meta.perm}} function.
#'
#' @param counts the gene read counts matrix.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast the contrasts vector. See the main \code{\link{metaseqr}} help
#' page.
#' @param repl the same as the replace argument in \code{\link{sample}} function.
#' @return A list with three members: the matrix of permuted per sample read counts,
#' the virtual sample list and the virtual contrast to be used with the \code{stat.*}
#' functions.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data("mm9.gene.data",package="metaseqR")
#' per <- make.permutation(mm9.gene.counts,sample.list.mm9,"e14.5_vs_adult_8_weeks")
#'}
make.permutation <- function(counts,sample.list,contrast,repl=FALSE) {
cnts <- strsplit(contrast,"_vs_")[[1]]
virtual.contrast <- paste(paste("VirtCond",1:length(cnts),sep=""),
collapse="_vs_")
virtual.sample.list <- vector("list",length(sample.list))
names(virtual.sample.list) <- paste("VirtCond",1:length(sample.list),
sep="")
# Avoid the extreme case of returning a vector with all samples the same
if (repl) {
resample <- rep(1,ncol(counts))
while(length(unique(resample))==1)
resample <- sample(1:ncol(counts),ncol(counts),replace=repl)
}
else
resample <- sample(1:ncol(counts),ncol(counts),replace=repl)
virtual.counts <- counts[,resample]
samples <- paste("VirtSamp",1:ncol(counts),sep="")
colnames(virtual.counts) <- samples
nsample <- sapply(sample.list,length)
virtual.samples <- split(samples,rep(1:length(nsample),nsample))
names(virtual.samples) <- names(virtual.sample.list)
for (n in names(virtual.sample.list))
virtual.sample.list[[n]] <- virtual.samples[[n]]
return(list(counts=virtual.counts,sample.list=virtual.sample.list,
contrast=virtual.contrast))
}
#' Calculate the ratio TP/(FP+FN)
#'
#' This function calculates the ratio of True Positives to the sum of False
#' Positives and False Negatives given a matrix of p-values (one for each
#' statistical test used) and a vector of ground truth (DE or non-DE). This
#' function serves as a method evaluation helper.
#'
#' @param truth the ground truth differential expression vector. It should contain
#' only zero and non-zero elements, with zero denoting non-differentially expressed
#' genes and non-zero, differentially expressed genes. Such a vector can be obtained
#' for example by using the \code{\link{make.sim.data.sd}} function, which creates
#' simulated RNA-Seq read counts based on real data. It MUST be named with gene
#' names, the same as in \code{p}.
#' @param p a p-value matrix whose rows correspond to each element in the
#' \code{truth} vector. If the matrix has a \code{colnames} attribute, a legend
#' will be added to the plot using these names, else a set of column names will
#' be auto-generated. \code{p} can also be a list or a data frame. In any case,
#' each row (or element) MUST be named with gene names (the same as in \code{truth}).
#' @param sig a significance level (0 < \code{sig} <=1).
#' @return A named list with two members. The first member is a data frame with
#' the numbers used to calculate the TP/(FP+FN) ratio and the second member is
#' the ratio TP/(FP+FN) for each statistical test.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p1 <- 0.001*matrix(runif(300),100,3)
#' p2 <- matrix(runif(300),100,3)
#' p <- rbind(p1,p2)
#' rownames(p) <- paste("gene",1:200,sep="_")
#' colnames(p) <- paste("method",1:3,sep="_")
#' truth <- c(rep(1,40),rep(-1,40),rep(0,10),rep(1,10),rep(2,10),rep(0,80))
#' names(truth) <- rownames(p)
#' otr <- calc.otr(truth,p)
#'}
calc.otr <- function(truth,p,sig=0.05) {
if (is.list(p))
pmat <- do.call("cbind",p)
else if (is.data.frame(p))
pmat <- as.matrix(p)
else if (is.matrix(p))
pmat <- p
if (is.null(colnames(pmat)))
colnames(pmat) <- paste("p",1:ncol(pmat),sep="_")
sig.genes <- true.isects <- missed <- vector("list",ncol(pmat))
names(sig.genes) <- names(true.isects) <- names(missed) <- colnames(pmat)
for (n in colnames(pmat)) {
sig.genes[[n]] <- names(which(pmat[,n]<sig))
true.isects[[n]] <- intersect(sig.genes[[n]],names(which(truth!=0)))
missed[[n]] <- setdiff(names(which(truth!=0)),true.isects[[n]])
}
result <- data.frame(
P=sapply(sig.genes,length),
TP=sapply(true.isects,length),
FN=sapply(missed,length)
)
result$FP <- result$P - result$TP
otr <- result$TP/(result$FP+result$FN)
names(otr) <- rownames(result)
return(list(result=result,otr=otr))
}
#' Calculate the F1-score
#'
#' This function calculates the F1 score (2*(precision*recall/precision+racall)
#' or 2*TP/(2*TP+FP+FN) given a matrix of p-values (one for each statistical test
#' used) and a vector of ground truth (DE or non-DE). This function serves as a
#' method evaluation helper.
#'
#' @param truth the ground truth differential expression vector. It should contain
#' only zero and non-zero elements, with zero denoting non-differentially expressed
#' genes and non-zero, differentially expressed genes. Such a vector can be obtained
#' for example by using the \code{\link{make.sim.data.sd}} function, which creates
#' simulated RNA-Seq read counts based on real data. It MUST be named with gene
#' names, the same as in \code{p}.
#' @param p a p-value matrix whose rows correspond to each element in the
#' \code{truth} vector. If the matrix has a \code{colnames} attribute, a legend
#' will be added to the plot using these names, else a set of column names will
#' be auto-generated. \code{p} can also be a list or a data frame. In any case,
#' each row (or element) MUST be named with gene names (the same as in \code{truth}).
#' @param sig a significance level (0 < \code{sig} <=1).
#' @return A named list with two members. The first member is a data frame with
#' the numbers used to calculate the F1-score and the second member is the
#' F1-score for each statistical test.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p1 <- 0.001*matrix(runif(300),100,3)
#' p2 <- matrix(runif(300),100,3)
#' p <- rbind(p1,p2)
#' rownames(p) <- paste("gene",1:200,sep="_")
#' colnames(p) <- paste("method",1:3,sep="_")
#' truth <- c(rep(1,40),rep(-1,40),rep(0,10),rep(1,10),rep(2,10),rep(0,80))
#' names(truth) <- rownames(p)
#' f1 <- calc.f1score(truth,p)
#'}
calc.f1score <- function(truth,p,sig=0.05) {
if (is.list(p))
pmat <- do.call("cbind",p)
else if (is.data.frame(p))
pmat <- as.matrix(p)
else if (is.matrix(p))
pmat <- p
if (is.null(colnames(pmat)))
colnames(pmat) <- paste("p",1:ncol(pmat),sep="_")
sig.genes <- true.isects <- missed <- vector("list",ncol(pmat))
names(sig.genes) <- names(true.isects) <- names(missed) <- colnames(pmat)
for (n in colnames(pmat)) {
sig.genes[[n]] <- names(which(pmat[,n]<sig))
true.isects[[n]] <- intersect(sig.genes[[n]],names(which(truth!=0)))
missed[[n]] <- setdiff(names(which(truth!=0)),true.isects[[n]])
}
result <- data.frame(
P=sapply(sig.genes,length),
TP=sapply(true.isects,length),
FN=sapply(missed,length)
)
result$FP <- result$P - result$TP
f1 <- 2*result$TP/(2*result$TP+result$FP+result$FN)
names(f1) <- rownames(result)
return(list(result=result,f1=f1))
}
<file_sep>/man/make.grid.Rd
\name{make.grid}
\alias{make.grid}
\title{Optimize rectangular grid plots}
\usage{
make.grid(n)
}
\arguments{
\item{n}{An integer, denoting the total number of plots
to be created.}
}
\value{
A 2-element vector with the dimensions of the grid.
}
\description{
Returns a vector for an optimized m x m plot grid to be
used with e.g. \code{par(mfrow)}. m x m is as close as
possible to the input n. Of course, there will be empty
grid positions if n < m x m.
}
\examples{
\donttest{
g1 <- make.grid(16) # Returns c(4,4)
g2 <- make.grid(11) # Returns c(4,3)
}
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.annotation.R
#' Annotation downloader
#'
#' This function connects to the EBI's Biomart service using the package biomaRt
#' and downloads annotation elements (gene co-ordinates, exon co-ordinates, gene
#' identifications, biotypes etc.) for each of the supported organisms. See the
#' help page of \code{\link{metaseqr}} for a list of supported organisms. The
#' function downloads annotation for an organism genes or exons.
#'
#' @param org the organism for which to download annotation.
#' @param type either \code{"gene"} or \code{"exon"}.
#' @param refdb the online source to use to fetch annotation. It can be
#' \code{"ensembl"} (default), \code{"ucsc"} or \code{"refseq"}. In the later two
#' cases, an SQL connection is opened with the UCSC public databases.
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not. It is used in the case of \code{type="exon"} to process
#' the return value of the query to the UCSC Genome Browser database.
#' @return A data frame with the canonical (not isoforms!) genes or exons of the
#' requested organism. When \code{type="genes"}, the data frame has the following
#' columns: chromosome, start, end, gene_id, gc_content, strand, gene_name, biotype.
#' When \code{type="exon"} the data frame has the following columns: chromosome,
#' start, end, exon_id, gene_id, strand, gene_name, biotype. The gene_id and exon_id
#' correspond to Ensembl gene and exon accessions respectively. The gene_name
#' corresponds to HUGO nomenclature gene names.
#' @note The data frame that is returned contains only "canonical" chromosomes
#' for each organism. It does not contain haplotypes or random locations and does
#' not contain chromosome M.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' hg19.genes <- get.annotation("hg19","gene","ensembl")
#' mm9.exons <- get.annotation("mm9","exon","ucsc")
#'}
get.annotation <- function(org,type,refdb="ensembl",multic=FALSE) {
org <- tolower(org)
if (type %in% c("utr","transcript") && refdb %in% c("ucsc","refseq")) {
disp("Quant-Seq (utr) and transcript analysis is not yet supported ",
"with UCSC or RefSeq annotation. Switching to Ensembl...")
refdb <- "ensembl"
}
switch(refdb,
ensembl = { return(get.ensembl.annotation(org,type)) },
ucsc = { return(get.ucsc.annotation(org,type,refdb,multic)) },
refseq = { return(get.ucsc.annotation(org,type,refdb,multic)) }
)
}
#' Ensembl annotation downloader
#'
#' This function connects to the EBI's Biomart service using the package biomaRt
#' and downloads annotation elements (gene co-ordinates, exon co-ordinates, gene
#' identifications, biotypes etc.) for each of the supported organisms. See the
#' help page of \code{\link{metaseqr}} for a list of supported organisms. The
#' function downloads annotation for an organism genes or exons.
#'
#' @param org the organism for which to download annotation.
#' @param type either \code{"gene"} or \code{"exon"}.
#' @return A data frame with the canonical (not isoforms!) genes or exons of the
#' requested organism. When \code{type="genes"}, the data frame has the following
#' columns: chromosome, start, end, gene_id, gc_content, strand, gene_name, biotype.
#' When \code{type="exon"} the data frame has the following columns: chromosome,
#' start, end, exon_id, gene_id, strand, gene_name, biotype. The gene_id and exon_id
#' correspond to Ensembl gene and exon accessions respectively. The gene_name
#' corresponds to HUGO nomenclature gene names.
#' @note The data frame that is returned contains only "canonical" chromosomes
#' for each organism. It does not contain haplotypes or random locations and does
#' not contain chromosome M.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' hg19.genes <- get.ensembl.annotation("hg19","gene")
#' mm9.exons <- get.ensembl.annotation("mm9","exon")
#'}
get.ensembl.annotation <- function(org,type) {
if (org=="tair10")
dat <- "plants_mart"
else if (org=="bmori2")
dat <- "metazoa_mart"
else
dat <- "ENSEMBL_MART_ENSEMBL"
mart <- tryCatch({
useMart(biomart=dat,host=get.host(org),dataset=get.dataset(org))
},
error=function(e) {
useMart(biomart=dat,host=get.alt.host(org),
dataset=get.dataset(org))
},
finally={})
chrs.exp <- paste(get.valid.chrs(org),collapse="|")
if (type=="gene") {
bm <- getBM(attributes=get.gene.attributes(org),mart=mart)
ann <- data.frame(
chromosome=paste("chr",bm$chromosome_name,sep=""),
start=bm$start_position,
end=bm$end_position,
gene_id=bm$ensembl_gene_id,
gc_content=if (org %in% c("hg18","mm9","tair10"))
bm$percentage_gc_content else bm$percentage_gene_gc_content,
strand=ifelse(bm$strand==1,"+","-"),
gene_name=if (org %in% c("hg18","mm9","tair10")) bm$external_gene_id
else bm$external_gene_name,
biotype=bm$gene_biotype
)
rownames(ann) <- ann$gene_id
}
else if (type=="exon") {
bm <- getBM(attributes=get.exon.attributes(org),mart=mart)
if (org == "hg19") {
disp(" Bypassing problem with hg19 Ensembl combined gene-exon ",
"annotation... Will take slightly longer...")
bmg <- getBM(attributes=get.gene.attributes(org),mart=mart)
gene_name <- bmg$external_gene_name
names(gene_name) <- bmg$ensembl_gene_id
ann <- data.frame(
chromosome=paste("chr",bm$chromosome_name,sep=""),
start=bm$exon_chrom_start,
end=bm$exon_chrom_end,
exon_id=bm$ensembl_exon_id,
gene_id=bm$ensembl_gene_id,
strand=ifelse(bm$strand==1,"+","-"),
gene_name=gene_name[bm$ensembl_gene_id],
biotype=bm$gene_biotype
)
rownames(ann) <- ann$exon_id
}
else
ann <- data.frame(
chromosome=paste("chr",bm$chromosome_name,sep=""),
start=bm$exon_chrom_start,
end=bm$exon_chrom_end,
exon_id=bm$ensembl_exon_id,
gene_id=bm$ensembl_gene_id,
strand=ifelse(bm$strand==1,"+","-"),
gene_name=if (org %in% c("hg18","mm9","tair10"))
bm$external_gene_id else bm$external_gene_name,
biotype=bm$gene_biotype
)
rownames(ann) <- ann$exon_id
}
else if (type=="utr") {
bm <- getBM(attributes=get.transcript.utr.attributes(org),mart=mart)
ann <- data.frame(
chromosome=paste("chr",bm$chromosome_name,sep=""),
start=bm$`3_utr_start`,
end=bm$`3_utr_end`,
tstart=bm$transcript_start,
tend=bm$transcript_end,
transcript_id=bm$ensembl_transcript_id,
gene_id=bm$ensembl_gene_id,
strand=ifelse(bm$strand==1,"+","-"),
gene_name=if (org %in% c("hg18","mm9","tair10"))
bm$external_gene_id else bm$external_gene_name,
biotype=bm$gene_biotype
)
ann <- correct.transcripts(ann)
ann <- ann[,c("chromosome","start","end","transcript_id","gene_id",
"strand","gene_name","biotype")]
}
else if (type=="transcript") {
bm <- getBM(attributes=get.transcript.attributes(org),mart=mart)
ann <- data.frame(
chromosome=paste("chr",bm$chromosome_name,sep=""),
start=bm$transcript_start,
end=bm$transcript_end,
transcript_id=bm$ensembl_transcript_id,
gene_id=bm$ensembl_gene_id,
strand=ifelse(bm$strand==1,"+","-"),
gene_name=if (org %in% c("hg18","mm9","tair10"))
bm$external_gene_id else bm$external_gene_name,
biotype=bm$gene_biotype
)
rownames(ann) <- as.character(ann$transcript_id)
}
ann <- ann[order(ann$chromosome,ann$start),]
ann <- ann[grep(chrs.exp,ann$chromosome),]
ann$chromosome <- as.character(ann$chromosome)
return(ann)
}
#' UCSC/RefSeq annotation downloader
#'
#' This function connects to the UCSC Genome Browser public database and downloads
#' annotation elements (gene co-ordinates, exon co-ordinates, gene identifications
#' etc.) for each of the supported organisms, but using UCSC instead of Ensembl.
#' See the help page of \code{\link{metaseqr}} for a list of supported organisms.
#' The function downloads annotation for an organism genes or exons.
#'
#' @param org the organism for which to download annotation.
#' @param type either \code{"gene"} or \code{"exon"}.
#' @param refdb either \code{"ucsc"} or \code{"refseq"}.
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not. It is used in the case of \code{type="exon"} to process
#' the return value of the query to the UCSC Genome Browser database.
#' @return A data frame with the canonical (not isoforms!) genes or exons of the
#' requested organism. When \code{type="genes"}, the data frame has the following
#' columns: chromosome, start, end, gene_id, gc_content, strand, gene_name, biotype.
#' When \code{type="exon"} the data frame has the following columns: chromosome,
#' start, end, exon_id, gene_id, strand, gene_name, biotype. The gene_id and exon_id
#' correspond to UCSC or RefSeq gene and exon accessions respectively. The gene_name
#' corresponds to HUGO nomenclature gene names.
#' @note The data frame that is returned contains only "canonical" chromosomes
#' for each organism. It does not contain haplotypes or random locations and does
#' not contain chromosome M. Note also that as the UCSC databases do not contain
#' biotype classification like Ensembl, this will be returned as \code{NA} and
#' as a result, some quality control plots will not be available.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' hg19.genes <- get.ucsc.annotation("hg19","gene","ucsc")
#' mm9.exons <- get.ucsc.annotation("mm9","exon")
#'}
get.ucsc.annotation <- function(org,type,refdb="ucsc",multic=FALSE) {
if (org=="bmori2") {
warnwrap("Bombyx mori (silkworm) annotation is not supported by UCSC ",
"or RefSeq! Will use Ensembl...")
return(get.ensembl.annotation("bmori2",type))
}
if (!require(RMySQL)) {
rmysql.present <- FALSE
warnwrap("R package RMySQL is not present! Annotation will be ",
"retrieved by downloading temporary files from UCSC and the usage
of a temporary SQLite database...")
}
else
rmysql.present <- TRUE
if (!require(RSQLite))
stopwrap("R package RSQLite is required to use annotation from UCSC!")
if (org=="tair10") {
warnwrap("Arabidopsis thaliana genome is not supported by UCSC Genome ",
"Browser database! Switching to Ensembl...")
return(get.ensembl.annotation("tair10",type))
}
if (org=="equcab2") {
warnwrap("Equus cabalus genome is not supported by UCSC Genome ",
"Browser database! Switching to Ensembl...")
return(get.ensembl.annotation("equcab2",type))
}
valid.chrs <- get.valid.chrs(org)
chrs.exp <- paste("^",paste(valid.chrs,collapse="$|^"),"$",sep="")
db.org <- get.ucsc.organism(org)
if (rmysql.present) {
db.creds <- get.ucsc.credentials()
drv <- dbDriver("MySQL")
con <- dbConnect(drv,user=db.creds[2],password=<PASSWORD>,dbname=db.org,
host=db.creds[1])
query <- get.ucsc.query(org,type,refdb)
raw.ann <- dbGetQuery(con,query)
dbDisconnect(con)
}
else {
# This should return the same data frame as the db query
tmp.sqlite <- get.ucsc.dbl(org,type,refdb)
drv <- dbDriver("SQLite")
con <- dbConnect(drv,dbname=tmp.sqlite)
query <- get.ucsc.query(org,type,refdb)
raw.ann <- dbGetQuery(con,query)
dbDisconnect(con)
}
if (type=="gene") {
ann <- raw.ann
ann <- ann[grep(chrs.exp,ann$chromosome,perl=TRUE),]
ann$chromosome <- as.character(ann$chromosome)
rownames(ann) <- ann$gene_id
}
else if (type=="exon") {
raw.ann <- raw.ann[grep(chrs.exp,raw.ann$chromosome,perl=TRUE),]
ex.list <- wapply(multic,as.list(1:nrow(raw.ann)),function(x,d,s) {
r <- d[x,]
starts <- as.numeric(strsplit(r[,"start"],",")[[1]])
ends <- as.numeric(strsplit(r[,"end"],",")[[1]])
nexons <- length(starts)
ret <- data.frame(
rep(r[,"chromosome"],nexons),
starts,ends,
paste(r[,"exon_id"],"_e",1:nexons,sep=""),
rep(r[,"strand"],nexons),
rep(r[,"gene_id"],nexons),
rep(r[,"gene_name"],nexons),
rep(r[,"biotype"],nexons)
)
names(ret) <- names(r)
rownames(ret) <- ret$exon_id
ret <- makeGRangesFromDataFrame(
df=ret,
keep.extra.columns=TRUE,
seqnames.field="chromosome",
seqinfo=s
)
return(ret)
},raw.ann,valid.chrs)
tmp.ann <- do.call("c",ex.list)
ann <- data.frame(
chromosome=as.character(seqnames(tmp.ann)),
start=start(tmp.ann),
end=end(tmp.ann),
exon_id=as.character(tmp.ann$exon_id),
gene_id=as.character(tmp.ann$gene_id),
strand=as.character(strand(tmp.ann)),
gene_name=as.character(tmp.ann$gene_name),
biotype=tmp.ann$biotype
)
rownames(ann) <- ann$exon_id
}
gc.content <- get.gc.content(ann,org)
ann$gc_content <- gc.content
ann <- ann[order(ann$chromosome,ann$start),]
return(ann)
}
#' Return a named vector of GC-content for each genomic region
#'
#' Returns a named numeric vector (names are the genomic region names, e.g. genes)
#' given a data frame which can be converted to a GRanges object (e.g. it has at
#' least chromosome, start, end fields). This function works best when the input
#' annotation data frame has been retrieved using one of the SQL queries generated
#' from \code{\link{get.ucsc.query}}, used in \code{\link{get.ucsc.annotation}}.
#'
#' @param ann a data frame which can be converted to a GRanges object, that means
#' it has at least the chromosome, start, end fields. Preferably, the output of
#' \code{link{get.ucsc.annotation}}.
#' @param org one of metaseqR supported organisms.
#' @return A named numeric vector.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' ann <- get.ucsc.annotation("mm9","gene","ucsc")
#' gc <- get.gc.content(ann,"mm9")
#'}
get.gc.content <- function(ann,org) {
if (missing(ann))
stopwrap("A valid annotation data frame must be provided in order to ",
"retrieve GC-content.")
org <- tolower(org[1])
check.text.args("org",org,c("hg18","hg19","hg38","mm9","mm10","rn5","dm3",
"dm6","danrer7","pantro4","susscr3","tair10"),multiarg=FALSE)
# Convert annotation to GRanges
disp("Converting annotation to GenomicRanges object...")
if (packageVersion("GenomicRanges")<1.14)
ann.gr <- GRanges(
seqnames=Rle(ann[,1]),
ranges=IRanges(start=ann[,2],end=ann[,3]),
strand=Rle(ann[,6]),
name=as.character(ann[,4])
)
else
ann.gr <- makeGRangesFromDataFrame(
df=ann,
keep.extra.columns=TRUE,
seqnames.field="chromosome"
)
bsg <- load.bs.genome(org)
disp("Getting DNA sequences...")
seqs <- getSeq(bsg,names=ann.gr)
disp("Getting GC content...")
freq.matrix <- alphabetFrequency(seqs,as.prob=TRUE,baseOnly=TRUE)
gc.content <- apply(freq.matrix,1,function(x) round(100*sum(x[2:3]),
digits=2))
names(gc.content) <- as.character(ann[,4])
return(gc.content)
}
#' Return a proper formatted organism alias
#'
#' Returns the proper UCSC Genome Browser database organism alias based on what is
#' given to metaseqR. Internal use.
#'
#' @return A proper organism alias.
#' @author <NAME>
#' @examples
#' \dontrun{
#' org <- get.ucsc.organism("danrer7")
#'}
get.ucsc.organism <- function(org) {
switch(org,
hg18 = { return("hg18") },
hg19 = { return("hg19") },
hg38 = { return("hg38") },
mm9 = { return("mm9") },
mm10 = { return("mm10") },
rn5 = { return("rn5") },
rn6 = { return("rn6") },
dm3 = { return("dm3") },
dm6 = { return("dm6") },
danrer7 = { return("danRer7") },
pantro4 = { return("panTro4") },
susscr3 = { return("susScr3") },
tair10 = { return("TAIR10") },
equcab2 = { return("equCab2") }
)
}
#' Return a proper formatted BSgenome organism name
#'
#' Returns a properly formatted BSgenome package name according to metaseqR's
#' supported organism. Internal use.
#'
#' @return A proper BSgenome package name.
#' @author <NAME>
#' @examples
#' \dontrun{
#' bs.name <- get.bs.organism("hg18")
#'}
get.bs.organism <- function(org) {
switch(org,
hg18 = {
return("BSgenome.Hsapiens.UCSC.hg18")
},
hg19 = {
return("BSgenome.Hsapiens.UCSC.hg19")
},
hg38 = {
return("BSgenome.Hsapiens.UCSC.hg38")
},
mm9 = {
return("BSgenome.Mmusculus.UCSC.mm9")
},
mm10 = {
return("BSgenome.Mmusculus.UCSC.mm10")
},
rn5 = {
return("BSgenome.Rnorvegicus.UCSC.rn5")
},
rn6 = {
return("BSgenome.Rnorvegicus.UCSC.rn6")
},
dm3 = {
return("BSgenome.Dmelanogaster.UCSC.dm3")
},
dm6 = {
return("BSgenome.Dmelanogaster.UCSC.dm6")
},
danrer7 = {
return("BSgenome.Drerio.UCSC.danRer7")
},
pantro4 = {
stopwrap("panTro4 is not yet supported by BSgenome! Please use ",
"Ensembl as annoation source.")
},
susscr3 = {
return("BSgenome.Sscrofa.UCSC.susScr3")
},
tair10 = {
stopwrap("TAIR10 is not yet supported by BSgenome! Please use ",
"Ensembl as annoation source.")
}
)
}
#' Loads (or downloads) the required BSGenome package
#'
#' Retrieves the required BSgenome package when the annotation source is \code{"ucsc"}
#' or \code{"refseq"}. These packages are required in order to estimate the
#' GC-content of the retrieved genes from UCSC or RefSeq.
#'
#' @param org one of \code{\link{metaseqr}} supported organisms.
#' @return The BSgenome object for the requested organism.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' bs.obj <- load.bs.genome("mm9")
#'}
load.bs.genome <- function(org) {
if (!require(BiocManager))
stopwrap("The Bioconductor package BiocManager is required to ",
"proceed!")
if (!require(BSgenome))
stopwrap("The Bioconductor package BSgenome is required to ",
"proceed!")
bs.org <- get.bs.organism(org)
if (bs.org %in% installed.genomes())
bs.obj <- getBSgenome(get.ucsc.organism(org))
else {
BiocManager::install(bs.org)
bs.obj <- getBSgenome(get.ucsc.organism(org))
}
return(bs.obj)
}
#' Biotype converter
#'
#' Returns biotypes as character vector. Internal use.
#'
#' @param a the annotation data frame (output of \code{\link{get.annotation}}).
#' @return A character vector of biotypes.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' hg18.genes <- get.annotation("hg18","gene")
#' hg18.bt <- get.biotypes(hg18.genes)
#'}
get.biotypes <- function(a) {
return(as.character(unique(a$biotype)))
}
#' Annotation downloader helper
#'
#' Returns the appropriate Ensembl host address to get different versions of
#' annotation from. Internal use.
#'
#' @param org the organism for which to return the host address.
#' @return A string with the host address.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' mm9.host <- get.host("mm9")
#'}
get.host <- function(org) {
switch(org,
hg18 = { return("may2009.archive.ensembl.org") },
hg19 = { return("grch37.ensembl.org") },
hg38 = { return("www.ensembl.org") },
mm9 = { return("may2012.archive.ensembl.org") },
mm10 = { return("www.ensembl.org") },
rn5 = { return("may2012.archive.ensembl.org") },
rn6 = { return("www.ensembl.org") },
dm3 = { return("grch37.ensembl.org") },
dm6 = { return("www.ensembl.org") },
danrer7 = { return("www.ensembl.org") },
pantro4 = { return("www.ensembl.org") },
susscr3 = { return("www.ensembl.org") },
tair10 = { return("plants.ensembl.org") },
equcab2 = { return("www.ensembl.org") },
bmori2 = { return("metazoa.ensembl.org") }
)
}
#' Annotation downloader helper
#'
#' Returns the appropriate Ensembl host address to get different versions of
#' annotation from (alternative hosts). Internal use.
#'
#' @param org the organism for which to return the host address.
#' @return A string with the host address.
#' @author <NAME>
#' @examples
#' \dontrun{
#' mm9.host <- get.alt.host("mm9")
#'}
get.alt.host <- function(org) {
switch(org,
hg18 = { return("may2009.archive.ensembl.org") },
hg19 = { return("grch37.ensembl.org") },
hg38 = { return("uswest.ensembl.org") },
mm9 = { return("may2012.archive.ensembl.org") },
mm10 = { return("uswest.ensembl.org") },
rn5 = { return("uswest.ensembl.org") },
dm3 = { return("grch37.ensembl.org") },
dm6 = { return("uswest.ensembl.org") },
danrer7 = { return("uswest.ensembl.org") },
pantro4 = { return("uswest.ensembl.org") },
susscr3 = { return("uswest.ensembl.org") },
tair10 = { return("plants.ensembl.org") },
equcab2 = { return("uswest.ensembl.org") },
bmori2 = { return("metazoa.ensembl.org") }
)
}
#' Annotation downloader helper
#'
#' Returns a dataset (gene or exon) identifier for each organism recognized by
#' the Biomart service for Ensembl. Internal use.
#'
#' @param org the organism for which to return the identifier.
#' @return A string with the dataset identifier.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' dm6.id <- get.dataset("dm6")
#'}
get.dataset <- function(org) {
switch(org,
hg18 = { return("hsapiens_gene_ensembl") },
hg19 = { return("hsapiens_gene_ensembl") },
hg38 = { return("hsapiens_gene_ensembl") },
mm9 = { return("mmusculus_gene_ensembl") },
mm10 = { return("mmusculus_gene_ensembl") },
rn5 = { return("rnorvegicus_gene_ensembl") },
rn6 = { return("rnorvegicus_gene_ensembl") },
dm3 = { return("dmelanogaster_gene_ensembl") },
dm6 = { return("dmelanogaster_gene_ensembl") },
danrer7 = { return("drerio_gene_ensembl") },
pantro4 = { return("ptroglodytes_gene_ensembl") },
susscr3 = { return("sscrofa_gene_ensembl") },
tair10 = { return("athaliana_eg_gene") },
equcab2 = { return("ecaballus_gene_ensembl") },
bmori2 = { return("bmori_eg_gene") },
)
}
#' Annotation downloader helper
#'
#' Returns a vector of chromosomes to maintain after annotation download. Internal
#' use.
#'
#' @param org the organism for which to return the chromosomes.
#' @return A character vector of chromosomes.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' hg18.chr <- get.valid.chrs("hg18")
#'}
get.valid.chrs <- function(org)
{
switch(org,
hg18 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr20","chr21","chr22","chr3",
"chr4","chr5","chr6","chr7","chr8","chr9","chrX","chrY"
))
},
hg19 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr20","chr21","chr22","chr3",
"chr4","chr5","chr6","chr7","chr8","chr9","chrX","chrY"
))
},
hg38 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr20","chr21","chr22","chr3",
"chr4","chr5","chr6","chr7","chr8","chr9","chrX","chrY"
))
},
mm9 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr3","chr4","chr5","chr6",
"chr7","chr8","chr9","chrX","chrY"
))
},
mm10 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr3","chr4","chr5","chr6",
"chr7","chr8","chr9","chrX","chrY"
))
},
rn5 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr3","chr4","chr5","chr6",
"chr7","chr8","chr9","chrX"
))
},
rn6 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr3","chr4","chr5","chr6",
"chr7","chr8","chr9","chrX"
))
},
dm3 = {
return(c(
"chr2L","chr2LHet","chr2R","chr2RHet","chr3L","chr3LHet",
"chr3R","chr3RHet","chr4","chrU","chrUextra","chrX","chrXHet",
"chrYHet"
))
},
dm6 = {
return(c(
"chr2L","chr2LHet","chr2R","chr2RHet","chr3L","chr3LHet",
"chr3R","chr3RHet","chr4","chrU","chrUextra","chrX","chrXHet",
"chrYHet"
))
},
danrer7 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr20","chr21","chr22","chr23",
"chr24","chr25","chr3","chr4","chr5","chr6","chr7","chr8","chr9"
))
},
pantro4 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr20","chr21","chr22","chr2A","chr2B",
"chr3","chr4","chr5","chr6","chr7","chr8","chr9","chrX","chrY"
))
},
susscr3 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr2","chr3","chr4","chr5","chr6","chr7",
"chr8","chr9","chrX","chrY"
))
},
tair10 = {
return(c(
"chr1","chr2","chr3","chr4","chr5"
))
},
equcab2 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr20","chr21","chr22","chr23",
"chr24","chr25","chr26","chr27","chr28","chr29","chr3","chr30",
"chr31","chr4","chr5","chr6","chr7","chr8","chr9","chrX","chrY"
))
},
bmori2 = {
return(c(
"chr1","chr10","chr11","chr12","chr13","chr14","chr15","chr16",
"chr17","chr18","chr19","chr2","chr20","chr21","chr22","chr23",
"chr24","chr25","chr26","chr27","chr28","chr3","chr4","chr5",
"chr6","chr7","chr8","chr9"
))
}
)
}
#' Annotation downloader helper
#'
#' Returns a vector of genomic annotation attributes which are used by the biomaRt
#' package in order to fetch the gene annotation for each organism. It has no
#' parameters. Internal use.
#'
#' @param org one of the supported organisms.
#' @return A character vector of Ensembl gene attributes.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' gene.attr <- get.gene.attributes()
#'}
get.gene.attributes <- function(org) {
if (org %in% c("hg18","mm9","tair10","bmori2"))
return(c(
"chromosome_name",
"start_position",
"end_position",
"ensembl_gene_id",
"percentage_gc_content",
"strand",
"external_gene_id",
"gene_biotype"
))
else
return(c(
"chromosome_name",
"start_position",
"end_position",
"ensembl_gene_id",
"percentage_gene_gc_content",
"strand",
"external_gene_name",
"gene_biotype"
))
}
#' Annotation downloader helper
#'
#' Returns a vector of genomic annotation attributes which are used by the biomaRt
#' package in order to fetch the exon annotation for each organism. It has no
#' parameters. Internal use.
#'
#' @param org one of the supported organisms.
#' @return A character vector of Ensembl exon attributes.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' exon.attr <- get.exon.attributes()
#'}
get.exon.attributes <- function(org) {
if (org %in% c("hg18","mm9","tair10","bmori2"))
return(c(
"chromosome_name",
"exon_chrom_start",
"exon_chrom_end",
"ensembl_exon_id",
"strand",
"ensembl_gene_id",
"external_gene_id",
"gene_biotype"
))
else if (org == "hg19")
return(c(
"chromosome_name",
"exon_chrom_start",
"exon_chrom_end",
"ensembl_exon_id",
"strand",
"ensembl_gene_id",
"gene_biotype"
))
else
return(c(
"chromosome_name",
"exon_chrom_start",
"exon_chrom_end",
"ensembl_exon_id",
"strand",
"ensembl_gene_id",
"external_gene_name",
"gene_biotype"
))
}
#' Annotation downloader helper
#'
#' Returns a vector of genomic annotation attributes which are used by the biomaRt
#' package in order to fetch the exon annotation for each organism. It has no
#' parameters. Internal use.
#'
#' @param org one of the supported organisms.
#' @return A character vector of Ensembl transcript attributes.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' trans.attr <- get.transcript.attributes()
#'}
get.transcript.utr.attributes <- function(org) {
if (org %in% c("hg18","mm9","tair10","bmori2"))
return(c(
"chromosome_name",
"transcript_start",
"transcript_end",
"3_utr_start",
"3_utr_end",
"ensembl_transcript_id",
"strand",
"ensembl_gene_id",
"external_gene_id",
"gene_biotype"
))
else
return(c(
"chromosome_name",
"transcript_start",
"transcript_end",
"3_utr_start",
"3_utr_end",
"ensembl_transcript_id",
"strand",
"ensembl_gene_id",
"external_gene_name",
"gene_biotype"
))
}
get.transcript.attributes <- function(org) {
if (org %in% c("hg18","mm9","tair10","bmori2"))
return(c(
"chromosome_name",
"transcript_start",
"transcript_end",
"ensembl_transcript_id",
"strand",
"ensembl_gene_id",
"external_gene_id",
"gene_biotype"
))
else
return(c(
"chromosome_name",
"transcript_start",
"transcript_end",
"ensembl_transcript_id",
"strand",
"ensembl_gene_id",
"external_gene_name",
"gene_biotype"
))
}
correct.transcripts <- function(ann) {
rownames(ann) <- paste("T",1:nrow(ann),sep="_")
len <- ann[,3] - ann[,2]
len <- len[-which(is.na(len))]
len[len==0] <- 1
def.utr.len <- round(2^mean(log2(len)))
nas <- which(is.na(ann$start))
ann.na <- ann[nas,]
ann.na$start <- ann.na$tstart
ann.na$end <- ann.na$tend
tmp <- makeGRangesFromDataFrame(df=ann.na)
tmp <- flank(resize(tmp,width=1,fix="end"),start=FALSE,width=def.utr.len)
ann[names(tmp),"start"] <- start(tmp)
ann[names(tmp),"end"] <- end(tmp)
return(ann)
}
<file_sep>/man/hg19.exon.counts.Rd
\docType{data}
\name{hg19.exon.counts}
\alias{hg19.exon.counts}
\title{Human RNA-Seq data with three conditions, three samples}
\format{a \code{data.frame} with exon read counts and some embedded annotation, one row per exon.}
\source{
GEO (http://www.ncbi.nlm.nih.gov/geo/)
}
\description{
This data set contains RNA-Seq exon read counts for 3
chromosomes. The data are from an experiment studying the
effect of a long non-coding RNA related to the ASCL2 gene
in WNT signaling and intestinal cancer. It has two
conditions (CON, DOX) and four samples (CON_BR1, CON_BR2,
DOX_BR1, DOX_BR2). It also contains a predefined
\code{sample.list} and \code{libsize.list} named
\code{sample.list.hg18} and \code{libsize.list.hg18}.
}
\author{
<NAME>
}
\keyword{datasets}
<file_sep>/inst/unitTests/test_metaseqr.R
test_metaseqr <- function() {
data("mm9.gene.data",package="metaseqR")
ex.dir <- tempdir()
result.1 <- metaseqr(
counts=mm9.gene.counts,
sample.list=sample.list.mm9,
contrast=c("e14.5_vs_adult_8_weeks"),
libsize.list=libsize.list.mm9,
annotation="download",
org="mm9",
count.type="gene",
normalization="edger",
statistics=c("edger","limma"),
meta.p="simes",
preset="medium.basic",
qc.plots="mds",
fig.format="png",
export.where=ex.dir,
out.list=TRUE,
restrict.cores=0.1
)
checkTrue(file.exists(file.path(ex.dir,"index.html")))
checkTrue(file.exists(file.path(ex.dir,"plots","qc","mds.png")))
checkTrue(file.exists(file.path(ex.dir,"lists")))
checkTrue(nrow(result.1[[1]][[1]])>0)
checkEqualsNumeric(ncol(result.1[[1]][[1]]),16)
}
<file_sep>/man/metaseqR-package.Rd
\docType{package}
\name{metaseqR-package}
\alias{metaseqR}
\alias{metaseqR-package}
\title{The metaseqR Package}
\description{
An R package for the analysis and result reporting
of RNA-Seq gene expression data, using multiple
statistical algorithms.
}
\details{
\tabular{ll}{ Package: \tab metaseqR\cr Type: \tab
Package\cr Version: \tab 1.9.1\cr Date: \tab
2015-07-27\cr Depends: \tab R (>= 2.13.0), EDASeq,
DESeq, limma, NOISeq, baySeq\cr Encoding: \tab UTF-8\cr
License: \tab GPL (>= 3)\cr LazyLoad: \tab yes\cr
URL: \tab http://www.fleming.gr\cr }
Provides an interface to several normalization and
statistical testing packages for RNA-Seq gene expression
data. Additionally, it creates several diagnostic plots,
performs meta-analysis by combinining the results of
several statistical tests and reports the results in an
interactive way.
}
\author{
<NAME> \email{<EMAIL>}
}
\keyword{package}
<file_sep>/man/stat.bayseq.Rd
\name{stat.bayseq}
\alias{stat.bayseq}
\title{Statistical testing with baySeq}
\usage{
stat.bayseq(object, sample.list, contrast.list = NULL,
stat.args = NULL, libsize.list = NULL)
}
\arguments{
\item{object}{a matrix or an object specific to each
normalization algorithm supported by metaseqR, containing
normalized counts. Apart from matrix (also for NOISeq),
the object can be a SeqExpressionSet (EDASeq),
CountDataSet (DESeq) or DGEList (edgeR).}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{contrast.list}{a named structured list of contrasts
as returned by \code{\link{make.contrast.list}} or just
the vector of contrasts as defined in the main help page
of \code{\link{metaseqr}}.}
\item{stat.args}{a list of edgeR statistical algorithm
parameters. See the result of
\code{get.defaults("statistics",} \code{"bayseq")} for an
example and how you can modify it.}
\item{libsize.list}{an optional named list where names
represent samples (MUST be the same as the samples in
\code{sample.list}) and members are the library sizes
(the sequencing depth) for each sample. If not provided,
they will be estimated from baySeq.}
}
\value{
A named list of the value 1-likelihood that a gene is
differentially expressed, whose names are the names of
the contrasts.
}
\description{
This function is a wrapper over baySeq statistical
testing. It accepts a matrix of normalized gene counts or
an S4 object specific to each normalization algorithm
supported by metaseqR.
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
norm.data.matrix <- normalize.edaseq(data.matrix,sample.list,gene.data)
p <- stat.bayseq(norm.data.matrix,sample.list,contrast)
}
}
\author{
<NAME>
}
<file_sep>/man/get.dataset.Rd
\name{get.dataset}
\alias{get.dataset}
\title{Annotation downloader helper}
\usage{
get.dataset(org)
}
\arguments{
\item{org}{the organism for which to return the
identifier.}
}
\value{
A string with the dataset identifier.
}
\description{
Returns a dataset (gene or exon) identifier for each
organism recognized by the Biomart service for Ensembl.
Internal use.
}
\examples{
\donttest{
dm3.id <- get.dataset("dm3")
}
}
\author{
<NAME>
}
<file_sep>/man/get.ucsc.organism.Rd
\name{get.ucsc.organism}
\alias{get.ucsc.organism}
\title{Return a proper formatted organism alias}
\usage{
get.ucsc.organism(org)
}
\arguments{
\item{org}{one of the \code{\link{metaseqr}}
supported organism.}
}
\value{
A proper organism alias.
}
\description{
Returns the proper UCSC Genome Browser database
organism alias based on what is given to metaseqR.
Internal use.
}
\examples{
\donttest{
org <- get.ucsc.organism("danrer7")
}
}
\author{
<NAME>
}
<file_sep>/man/get.annotation.Rd
\name{get.annotation}
\alias{get.annotation}
\title{Annotation downloader}
\usage{
get.annotation(org, type, refdb="ensembl",
multic=FALSE)
}
\arguments{
\item{org}{the organism for which to download
annotation.}
\item{type}{either \code{"gene"} or \code{"exon"}.}
\item{refdb}{the online source to use to fetch
annotation. It can be \code{"ensembl"} (default),
\code{"ucsc"} or \code{"refseq"}. In the later two
cases, an SQL connection is opened with the UCSC
public databases.}
\item{multic}{a logical value indicating the
presence of multiple cores. Defaults to
\code{FALSE}. Do not change it if you are not
sure whether package parallel has been loaded
or not. It is used in the case of
\code{type="exon"} to process the return value of
the query to the UCSC Genome Browser database.}
}
\value{
A data frame with the canonical (not isoforms!) genes or
exons of the requested organism. When
\code{type="genes"}, the data frame has the following
columns: chromosome, start, end, gene_id, gc_content,
strand, gene_name, biotype. When \code{type="exon"} the
data frame has the following columns: chromosome, start,
end, exon_id, gene_id, strand, gene_name, biotype. The
gene_id and exon_id correspond to Ensembl gene and exon
accessions respectively. The gene_name corresponds to
HUGO nomenclature gene names.
}
\description{
This function connects to the EBI's Biomart service using
the package biomaRt and downloads annotation elements
(gene co-ordinates, exon co-ordinates, gene
identifications, biotypes etc.) for each of the supported
organisms. See the help page of \code{\link{metaseqr}}
for a list of supported organisms. The function downloads
annotation for an organism genes or exons.
}
\note{
The data frame that is returned contains only "canonical"
chromosomes for each organism. It does not contain
haplotypes or random locations and does not contain
chromosome M.
}
\examples{
\donttest{
hg19.genes <- get.annotation("hg19","gene","ensembl")
mm9.exons <- get.annotation("mm9","exon","ucsc")
}
}
\author{
<NAME>
}
<file_sep>/inst/unitTests/test_estimate_aufc_weights.R
test_estimate_aufc_weights <- function() {
data("mm9.gene.data",package="metaseqR")
#multic <- check.parallel(0.1)
#multic <- FALSE
weights <- estimate.aufc.weights(
counts=as.matrix(mm9.gene.counts[,9:12]),
normalization="edaseq",
statistics=c("edger","limma"),
nsim=1,N=10,ndeg=c(2,2),top=4,model.org="mm9",
seed=42,multic=FALSE,libsize.gt=1e+5
)
#checkEqualsNumeric(weights,c(0.5384615,0.4615385),tolerance=1e-5)
checkEqualsNumeric(weights,c(0.5,0.5),tolerance=1e-5)
checkEqualsNumeric(sum(weights),1,tolerance=1e-9)
}
<file_sep>/man/mm9.gene.counts.Rd
\docType{data}
\name{mm9.gene.counts}
\alias{mm9.gene.counts}
\title{mouse RNA-Seq data with two conditions, four samples}
\format{a \code{data.frame} with gene read counts and some embedded annotation, one row per gene.}
\source{
ENCODE (http://genome.ucsc.edu/encode/)
}
\description{
This data set contains RNA-Seq gene read counts for 3
chromosomes. The data were downloaded from the ENCODE
public repository and are derived from the study of
Mortazavi et al., 2008 (<NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Mapping and quantifying mammalian
transcriptomes by RNA-Seq. Nat Methods. 2008
Jul;5(7):621-8). In their experiment, the authors studied
among others genes expression at two developmental stages
of mouse liver cells. It has two conditions-developmental
stages (e14.5, adult_8_weeks) and four samples (e14.5_1,
e14.5_2, a8w_1, a8w_2). It also contains a predefined
\code{sample.list} and \code{libsize.list} named
\code{sample.list.mm9} and \code{libsize.list.mm9}.
}
\author{
<NAME>
}
\keyword{datasets}
<file_sep>/man/get.ucsc.dbl.Rd
\name{get.ucsc.dbl}
\alias{get.ucsc.dbl}
\title{Download annotation from UCSC servers, according to organism and
source}
\usage{
get.ucsc.dbl(org, type, refdb="ucsc")
}
\arguments{
\item{org}{one of metaseqR supported organisms.}
\item{type}{either \code{"gene"} or \code{"exon"}.}
\item{refdb}{one of \code{"ucsc"} or \code{"refseq"}
to use the UCSC or RefSeq annotation sources
respectively.}
}
\value{
An SQLite database.
}
\description{
Directly downloads UCSC and RefSeq annotation files from UCSC servers
to be used with metaseqR. This functionality is used when the package
RMySQL is not available for some reason, e.g. Windows machines. It
created an SQLite database where the same queries can be used.
}
\examples{
\donttest{
db.file <- get.ucsc.dbl("hg18","gene","ucsc")
}
}
\author{
<NAME>
}
<file_sep>/man/fisher.sum.Rd
\name{fisher.sum}
\alias{fisher.sum}
\title{
A function to calculate Fisher's sum for a set of p-values
}
\description{
This method combines a set of p-values using Fisher's method:
\deqn{S = -2 \sum \log p}
}
\usage{
fisher.sum(p, zero.sub=0.00001, na.rm=FALSE)
}
%- maybe also 'usage' for other objects documented here.
\arguments{
\item{p}{
A vector of p-values
}
\item{zero.sub}{
Replacement for 0 values.
}
\item{na.rm}{
Should NA values be removed before calculating the sum
}
}
\details{
As \code{log(0)} results in \code{Inf} we replace p-values of 0 by
default with a small float. If you want to keep them as 0 you have
to provide 0 as a parameter in \code{zero.sub}.
Note that only p-values between 0 and 1 are allowed to be passed to
this method.
}
\value{
Fisher's sum as described above.
}
\note{
This function was copied from the CRAN package MADAM which is no longer
maintained. Recognition goes to the original author(s) below.
}
\references{
Fisher, R.A. (1925). Statistical Methods for Research Workers. Oliver
and Boyd (Edinburgh).
}
\author{
<NAME> <<EMAIL>>
}
\seealso{
\code{\link{fisher.method}}
}
\examples{
fisher.sum(c(0.2,0.05,0.05))
fisher.sum(c(0.2,0.05,0.05, NA), na.rm=TRUE)
}
<file_sep>/man/make.html.body.Rd
\name{make.html.body}
\alias{make.html.body}
\title{HTML report helper}
\usage{
make.html.body(mat)
}
\arguments{
\item{mat}{the character vector produced by
\code{\link{make.html.rows}}.}
}
\value{
A character vector with the body of mat formatted in
html.
}
\description{
Returns a character vector with an html formatted table.
Essentially, it collapses the input rows to a single
character and puts a <tbody></tbody> tag set around. It
is meant to be applied to the output of
\code{\link{make.html.rows}}. Internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
the.cells <- make.html.cells(data.matrix)
the.header <- make.html.header(the.cells[1,])
the.rows <- make.html.rows(the.cells)
the.body <- make.html.body(the.rows)
}
}
\author{
<NAME>
}
<file_sep>/man/reduce.exons.Rd
\name{reduce.exons}
\alias{reduce.exons}
\title{Merges exons to create a unique set of exons for each gene}
\usage{
reduce.exons(gr, multic = FALSE)
}
\arguments{
\item{gr}{a GRanges object created from the supplied
annotation (see also the \code{\link{read2count}} and
\code{\link{get.annotation}} functions.}
\item{multic}{a logical value indicating the presence
of multiple cores. Defaults to \code{FALSE}. Do not
change it if you are not sure whether package parallel
has been loaded or not.}
}
\value{
A GRanges object with virtual merged exons for each
gene/transcript.
}
\description{
This function uses the \code{"reduce"} function of
IRanges to construct virtual unique exons for each
gene, so as to avoid inflating the read counts for
each gene because of multiple possible transcripts.
If the user wants transcripts instead of genes, they
should be supplied to the original annotation table.
}
\examples{
\donttest{
require(GenomicRanges)
multic <- check.parallel(0.8)
ann <- get.annotation("mm9","exon")
gr <- makeGRangesFromDataFrame(
df=ann,
keep.extra.columns=TRUE,
seqnames.field="chromosome"
)
re <- reduce.exons(gr,multic=multic)
}
}
\author{
<NAME>
}
<file_sep>/man/stat.edger.Rd
\name{stat.edger}
\alias{stat.edger}
\title{Statistical testing with edgeR}
\usage{
stat.edger(object, sample.list, contrast.list = NULL,
stat.args = NULL)
}
\arguments{
\item{object}{a matrix or an object specific to each
normalization algorithm supported by metaseqR, containing
normalized counts. Apart from matrix (also for NOISeq),
the object can be a SeqExpressionSet (EDASeq),
CountDataSet (DESeq) or DGEList (edgeR).}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{contrast.list}{a named structured list of contrasts
as returned by \code{\link{make.contrast.list}} or just
the vector of contrasts as defined in the main help page
of \code{\link{metaseqr}}.}
\item{stat.args}{a list of edgeR statistical algorithm
parameters. See the result of
\code{get.defaults("statistics",} \code{"edger")} for an
example and how you can modify it.}
}
\value{
A named list of p-values, whose names are the names of
the contrasts.
}
\description{
This function is a wrapper over edgeR statistical
testing. It accepts a matrix of normalized gene counts or
an S4 object specific to each normalization algorithm
supported by metaseqR.
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
norm.data.matrix <- normalize.edger(data.matrix,sample.list)
p <- stat.edger(norm.data.matrix,sample.list,contrast)
}
}
\author{
<NAME>
}
<file_sep>/man/nat2log.Rd
\name{nat2log}
\alias{nat2log}
\title{General value transformation}
\usage{
nat2log(x, base = 2, off = 1)
}
\arguments{
\item{x}{input data matrix}
\item{base}{logarithmic base, 2 or 10}
\item{off}{offset to avoid Infinity}
}
\description{
Logarithmic transformation. Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/stat.noiseq.Rd
\name{stat.noiseq}
\alias{stat.noiseq}
\title{Statistical testing with NOISeq}
\usage{
stat.noiseq(object, sample.list, contrast.list = NULL,
stat.args = NULL, gene.data = NULL, log.offset = 1)
}
\arguments{
\item{object}{a matrix or an object specific to each
normalization algorithm supported by metaseqR, containing
normalized counts. Apart from matrix (also for NOISeq),
the object can be a SeqExpressionSet (EDASeq),
CountDataSet (DESeq) or DGEList (edgeR).}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{contrast.list}{a named structured list of contrasts
as returned by \code{\link{make.contrast.list}} or just
the vector of contrasts as defined in the main help page
of \code{\link{metaseqr}}.}
\item{stat.args}{a list of edgeR statistical algorithm
parameters. See the result of
\code{get.defaults("statistics",} \code{"noiseq")} for an
example and how you can modify it.}
\item{gene.data}{an optional annotation data frame (such
the ones produced by \code{get.annotation} which contains
the GC content for each gene and from which the gene
lengths can be inferred by chromosome coordinates.}
\item{log.offset}{a number to be added to each element of
data matrix in order to avoid Infinity on log type data
transformations.}
}
\value{
A named list of NOISeq q-values, whose names are the
names of the contrasts.
}
\description{
This function is a wrapper over NOISeq statistical
testing. It accepts a matrix of normalized gene counts or
an S4 object specific to each normalization algorithm
supported by metaseqR.
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
lengths <- round(1000*runif(nrow(data.matrix)))
starts <- round(1000*runif(nrow(data.matrix)))
ends <- starts + lengths
gc=runif(nrow(data.matrix))
gene.data <- data.frame(
chromosome=c(rep("chr1",nrow(data.matrix)/2),
rep("chr2",nrow(data.matrix)/2)),
start=starts,end=ends,gene_id=rownames(data.matrix),gc_content=gc
)
norm.data.matrix <- normalize.noiseq(data.matrix,sample.list,gene.data)
p <- stat.noiseq(norm.data.matrix,sample.list,contrast,
gene.data=gene.data)
}
}
\author{
<NAME>
}
<file_sep>/man/check.text.args.Rd
\name{check.text.args}
\alias{check.text.args}
\title{Text argument validator}
\usage{
check.text.args(arg.name, arg.value, arg.list,
multiarg=FALSE)
}
\arguments{
\item{arg.name}{the name of the argument that is
checked (for display purposes).}
\item{arg.value}{the value(s) of the argument to be
checked.}
\item{arg.list}{a vector of valid argument values
for \code{arg.value} to be matched against.}
\item{multiarg}{a logical scalar indicating whether
\code{arg.name} accepts multiple arguments or not.
In that case, all of the values in \code{arg.value}
are checked against \code{arg.list}.}
}
\description{
Checks if one or more given textual argument(s)
is/are member(s) of a list of correct arguments.
It's a more package-specific function similar to
\code{\link{match.arg}}. Mostly for internal use.
}
\examples{
# OK
check.text.args("count.type","gene",c("gene","exon"),
multiarg=FALSE)
## Error!
#check.text.args("statistics","ebseq",c("deseq","edger",
# "noiseq","bayseq","limma"), multiarg=TRUE)
}
\author{
<NAME>
}
<file_sep>/man/diagplot.cor.Rd
\name{diagplot.cor}
\alias{diagplot.cor}
\title{Summarized correlation plots}
\usage{
diagplot.cor(mat, type = c("heatmap", "correlogram"),
output = "x11", path = NULL, ...)
}
\arguments{
\item{mat}{the read counts matrix or data frame.}
\item{type}{create heatmap of correlogram plots.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filename of the pairwise comparisons plot produced if
it's a file.
}
\description{
This function uses the read counts matrix to create
heatmap or correlogram correlation plots.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
diagplot.cor(data.matrix,type="heatmap")
diagplot.cor(data.matrix,type="correlogram")
}
\author{
<NAME>
}
<file_sep>/man/diagplot.volcano.Rd
\name{diagplot.volcano}
\alias{diagplot.volcano}
\title{(Interactive) volcano plots of differentially expressed genes}
\usage{
diagplot.volcano(f, p, con = NULL, fcut = 1, pcut = 0.05,
alt.names = NULL, output = "x11", path = NULL, ...)
}
\arguments{
\item{f}{the fold changes which are to be plotted on the
x-axis.}
\item{p}{the p-values whose -log10 transformation is
going to be plotted on the y-axis.}
\item{con}{an optional string depicting a name (e.g. the
contrast name) to appear in the title of the volcano
diagplot.}
\item{fcut}{a fold change cutoff so as to draw two
vertical lines indicating the cutoff threshold for
biological significance.}
\item{pcut}{a p-value cutoff so as to draw a horizontal
line indicating the cutoff threshold for statistical
significance.}
\item{alt.names}{an optional vector of names, e.g. HUGO
gene symbols, alternative or complementary to the unique
names of \code{f} or \code{p} (one of them must be
named!). It is used only in JSON output.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"}, \code{"ps"} or \code{"json"}. The latter is
currently available for the creation of interactive
volcano plots only when reporting the output, through the
highcharts javascript library.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filenames of the plots produced in a named list with
names the \code{which.plot} argument. If
\code{output="x11"}, no output filenames are produced.
}
\description{
This function plots a volcano plot or returns a JSON
string which is used to render aninteractive in case of
HTML reporting.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
M <- normalize.edger(data.matrix,sample.list)
p <- stat.edger(M,sample.list,contrast)
ma <- apply(M[,sample.list$A],1,mean)
mb <- apply(M[,sample.list$B],1,mean)
f <- log2(ifelse(mb==0,1,mb)/ifelse(ma==0,1,ma))
diagplot.volcano(f,p[[1]],con=contrast)
#j <- diagplot.volcano(f,p[[1]],con=contrast,output="json")
}
\author{
<NAME>
}
<file_sep>/man/get.gene.attributes.Rd
\name{get.gene.attributes}
\alias{get.gene.attributes}
\title{Annotation downloader helper}
\usage{
get.gene.attributes(org)
}
\arguments{
\item{org}{one of the supported organisms.}
}
\value{
A character vector of Ensembl gene attributes.
}
\description{
Returns a vector of genomic annotation attributes which
are used by the biomaRt package in order to fetch the
gene annotation for each organism. It has no parameters.
Internal use.
}
\examples{
gene.attr <- get.gene.attributes("mm9")
}
\author{
<NAME>
}
<file_sep>/man/make.transformation.Rd
\name{make.transformation}
\alias{make.transformation}
\title{Calculates several transformation of counts}
\usage{
make.transformation(data.matrix, export.scale,
scf = NULL, log.offset = 1)
}
\arguments{
\item{data.matrix}{the raw or normalized counts matrix.
Each column represents one input sample.}
\item{export.scale}{a character vector containing one of
the supported data transformations (\code{"natural"},
\code{"log2"}, \code{"log10"},\code{"vst"}). See also the
main help page of metaseqr.}
\item{scf}{a scaling factor for the reads of each gene,
for example the sum of exon lengths or the gene length.
Divided by each read count when \code{export.scale="rpgm"}.
It provides an RPKM-like measure but not the actual RPKM
as this normalization is not supported.}
\item{log.offset}{a number to be added to each element of
data.matrix in order to avoid Infinity on log type data
transformations.}
}
\value{
A named list whose names are the elements in
export.scale. Each list member is the respective
transformed data matrix.
}
\description{
Returns a list of transformed (normalized) counts, based
on the input count matrix data.matrix. The data
transformations are passed from the \code{export.scale}
parameter and the output list is named accordingly. This
function is intended mostly for internal use but can also
be used independently.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
tr <- make.transformation(data.matrix,c("log2","vst"))
head(tr$vst)
}
}
\author{
<NAME>
}
<file_sep>/man/make.sim.data.sd.Rd
\name{make.sim.data.sd}
\alias{make.sim.data.sd}
\title{Create simulated counts using the
Soneson-Delorenzi method}
\usage{
make.sim.data.sd(N, param, samples = c(5, 5),
ndeg = rep(round(0.1*N), 2), fc.basis = 1.5,
libsize.range = c(0.7, 1.4), libsize.mag = 1e+7,
model.org = NULL, sim.length.bias = FALSE,
seed = NULL)
}
\arguments{
\item{N}{the number of genes to produce.}
\item{param}{a named list with negative binomial
parameter sets to sample from. The first member is
the mean parameter to sample from (\code{mu.hat})
and the second the dispersion (\code{phi.hat}).
This list can be created with the
\code{\link{estimate.sim.params}} function.}
\item{samples}{a vector with 2 integers,
which are the number of samples for each
condition (two conditions currently supported).}
\item{ndeg}{a vector with 2 integers, which are
the number of differentially expressed genes to
be produced. The first element is the number of
up-regulated genes while the second is the
number of down-regulated genes.}
\item{fc.basis}{the minimum fold-change for
deregulation.}
\item{libsize.range}{a vector with 2 numbers
(generally small, see the default), as they
are multiplied with \code{libsize.mag}. These
numbers control the library sized of the
synthetic data to be produced.}
\item{libsize.mag}{a (big) number to multiply
the \code{libsize.range} to produce library
sizes.}
\item{model.org}{the organism from which the
real data are derived from. It must be one
of the supported organisms (see the main
\code{\link{metaseqr}} help page). It is used
to sample real values for GC content.}
\item{sim.length.bias}{a boolean to instruct
the simulator to create genes whose read counts is
proportional to their length. This is achieved by
sorting in increasing order the mean parameter of
the negative binomial distribution (and the
dispersion according to the mean) which will cause
an increasing gene count length with the sampling.
The sampled lengths are also sorted so that in the
final gene list, shorter genes have less counts as
compared to the longer ones. The default is FALSE.}
\item{seed}{a seed to use with random number
generation for reproducibility.}
}
\value{
A named list with two members. The first
member (\code{simdata}) contains the
synthetic dataset
}
\description{
This function creates simulated RNA-Seq gene
expression datasets using the method presented
in (Soneson and Delorenzi, BMC Bioinformatics,
2013). For the time being, it creates only
simulated datasets with two conditions.
}
\examples{
\donttest{
# File "bottomly_read_counts.txt" from the ReCount database
download.file(paste("http://bowtie-bio.sourceforge.net/recount/",
"countTables/bottomly_count_table.txt",sep=""),
destfile="~/bottomly_count_table.txt")
N <- 10000
par.list <- estimate.sim.params("~/bottomly_read_counts.txt")
sim <- make.sim.data.sd(N,par.list)
synth.data <- sim$simdata
true.deg <- which(sim$truedeg!=0)
}
}
\author{
<NAME>
}
<file_sep>/man/sample.list.mm9.Rd
\docType{data}
\name{sample.list.mm9}
\alias{sample.list.mm9}
\title{Mouse RNA-Seq data with two conditions, four samples}
\format{a named \code{list} with condition and sample names.}
\source{
ENCODE (http://genome.ucsc.edu/encode/)
}
\description{
The sample list for \code{mm9.gene.counts}. See the data
set description.
}
\author{
<NAME>
}
\keyword{datasets}
<file_sep>/man/meta.worker.Rd
\name{meta.worker}
\alias{meta.worker}
\title{Permutation tests helper}
\usage{
meta.worker(x,co,sl,cnt,s,r,sa,ll,
el,w)
}
\arguments{
\item{x}{a virtual list with the random seed and
the permutation index.}
\item{co}{the counts matrix.}
\item{sl}{the sample list.}
\item{cnt}{the contrast name.}
\item{s}{the statistical algorithms.}
\item{sa}{the parameters for each statistical
algorithm.}
\item{ll}{a list with library sizes.}
\item{r}{same as the \code{replace} argument in
the \code{\link{sample}} function.}
\item{el}{min, max or weight.}
\item{w}{a numeric vector of weights for each
statistical algorithm}
}
\value{
A matrix of p-values.
}
\description{
This function performs the statistical test for
each permutation. Internal use only.
}
\examples{
\donttest{
# Not yet available
}
}
\author{
<NAME>
}
<file_sep>/man/calc.otr.Rd
\name{calc.otr}
\alias{calc.otr}
\title{Calculate the ratio TP/(FP+FN)}
\usage{
calc.otr(truth, p, sig = 0.05)
}
\arguments{
\item{truth}{the ground truth differential
expression vector. It should contain only
zero and non-zero elements, with zero denoting
non-differentially expressed genes and non-zero,
differentially expressed genes. Such a vector
can be obtained for example by using the
\code{\link{make.sim.data.sd}} function, which
creates simulated RNA-Seq read counts based on
real data. It MUST be named with gene names,
the same as in \code{p}.}
\item{p}{a p-value matrix whose rows correspond
to each element in the \code{truth} vector. If
the matrix has a \code{colnames} attribute, a
legend will be added to the plot using these
names, else a set of column names will be
auto-generated. \code{p} can also be a list or
a data frame. In any case, each row (or element)
MUST be named with gene names (the same as in
\code{truth}).}
\item{sig}{a significance level (0 < \code{sig}
<=1).}
}
\value{
A named list with two members. The first member
is a data frame with the numbers used to
calculate the TP/(FP+FN) ratio and the second
member is the ratio TP/(FP+FN) for each
statistical test.
}
\description{
This function calculates the ratio of True
Positives to the sum of False Positives and
False Negatives given a matrix of p-values
(one for each statistical test used) and a
vector of ground truth (DE or non-DE). This
function serves as a method evaluation helper.
}
\examples{
p1 <- 0.001*matrix(runif(300),100,3)
p2 <- matrix(runif(300),100,3)
p <- rbind(p1,p2)
rownames(p) <- paste("gene",1:200,sep="_")
colnames(p) <- paste("method",1:3,sep="_")
truth <- c(rep(1,40),rep(-1,40),rep(0,20),rep(1,10),
rep(2,10),rep(0,80))
names(truth) <- rownames(p)
otr <- calc.otr(truth,p)
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.util.R
#' Get precalculated statistical test weights
#'
#' This function returns pre-calculated weights for human, chimpanzee, mouse,
#' fruitfly and arabidopsis based on the performance of simulated datasets estimated
#' from real data from the ReCount database (\url{http://bowtie-bio.sourceforge.net/recount/}).
#' Currently pre-calculated weights are available only when all six statistical
#' tests are used and for normalization with EDASeq. For other combinations, use
#' the \code{\link{estimate.aufc.weights}} function.
#'
#' @param org \code{"human"}, \code{"mouse"}, \code{"chimpanzee"}, \code{"fruitfly"}
#' or \code{"arabidopsis"}.
#' @return A named vector of convex weights.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' wh <- get.weights("human",c("deseq","edger","noiseq"))
#}
get.weights <- function(org=c("human","chimpanzee","mouse","fruitfly",
"arabidopsis")) {
org <- tolower(org)
check.text.args("org",org,c("human","chimpanzee","mouse","fruitfly",
"arabidopsis"))
switch(org,
human = {
return(c(
deseq=0.05772458,
edger=0.14321672,
limma=0.34516089,
nbpseq=0.06108182,
noiseq=0.11595169,
bayseq=0.27686431
))
},
chimpanzee = {
return(c(
deseq=0.06026782,
edger=0.14964358,
limma=0.33500306,
nbpseq=0.05814585,
noiseq=0.11337043,
bayseq=0.28356925
))
},
mouse = {
return(c(
deseq=0.05257695,
edger=0.24161354,
limma=0.29957277,
nbpseq=0.04914485,
noiseq=0.06847809,
bayseq=0.28861381
))
},
fruitfly = {
return(c(
deseq=0.01430269,
edger=0.12923339,
limma=0.38315685,
nbpseq=0.01265952,
noiseq=0.06778537,
bayseq=0.39286218
))
},
arabidopsis = {
return(c(
deseq=0.04926122,
edger=0.10130858,
limma=0.40842011,
nbpseq=0.04596652,
noiseq=0.09336509,
bayseq=0.30167848
))
},
chimp = {
return(c(
deseq=NULL,
edger=NULL,
limma=NULL,
nbpseq=NULL,
noiseq=NULL,
bayseq=NULL
))
}
)
}
#' Default parameters for several metaseqr functions
#'
#' This function returns a list with the default settings for each filtering,
#' statistical and normalization algorithm included in the metaseqR package.
#' See the documentation of the main function and the documentation of each
#' statistical and normalization method for details.
#'
#' @param what a keyword determining the procedure for which to fetch the default
#' settings according to method parameter. It can be one of \code{"normalization"},
#' \code{"statistics"}, \code{"gene.filter"}, \code{"exon.filter"} or
#' \code{"biotype.filter"}.
#' @param method the supported algorithm included in metaseqR for which to fetch
#' the default settings. When \code{what} is \code{"normalization"}, method is one
#' of \code{"edaseq"}, \code{"deseq"}, \code{"edger"}, \code{"noiseq"} or
#' \code{"nbpseq"}. When \code{what} is \code{"statistics"}, method is one of
#' \code{"deseq"}, \code{"edger"}, \code{"noiseq"}, \code{"bayseq"}, \code{"limma"}
#' or \code{"nbpseq"}. When \code{method} is \code{"biotype.filter"}, \code{what}
#' is the input organism (see the main \code{\link{metaseqr}} help page for a list
#' of supported organisms).
#' @return A list with default setting that can be used directly in the call of
#' metaseqr.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' norm.args.edaseq <- get.defaults("normalization","edaseq")
#' stat.args.edger <- get.defaults("statistics","edger")
#'}
get.defaults <- function(what,method=NULL) {
if (what %in% c("normalization","statistics") && is.null(method))
stopwrap("The method argument must be provided when what is ",
"\"normalization\" or \"statistics\"!")
switch(what,
normalization = {
switch(method,
edaseq = {
return(list(within.which="loess",between.which="full"))
},
deseq = {
return(list(locfunc=median))
},
edger = {
return(list(
method="TMM",refColumn=NULL,logratioTrim=0.3,
sumTrim=0.05,doWeighting=TRUE,Acutoff=-1e10,p=0.75
))
},
noiseq = {
return(list(
method="tmm", # which normalization
long=1000,lc=1,k=1, # common arguments
refColumn=1,logratioTrim=0.3,sumTrim=0.05,
doWeighting=TRUE,Acutoff=-1e+10 # TMM normalization arguments
))
},
nbpseq = {
return(list(main.method="nbsmyth",method="AH2010",
thinning=TRUE))
}
)
},
statistics = {
switch(method,
deseq = {
return(list(method="blind",sharingMode="fit-only",
fitType="local"))
},
edger = {
return(list(
main.method="classic", # classic or glm fit
rowsum.filter=5,prior.df=10,
trend="movingave",span=NULL, # classic estimateCommonDisp arguments
tag.method="grid",grid.length=11,grid.range=c(-6,6), # classic estimateTagwiseDisp arguments
offset=NULL,glm.method="CoxReid",subset=10000, # glm estimateGLMCommonDisp and estimateGLMTrendedDisp arguments
AveLogCPM=NULL,trend.method="auto", # glm estimateGLMTagwiseDisp arguments
dispersion=NULL,offset=NULL,weights=NULL, # glmFit arguments
lib.size=NULL,prior.count=0.125,start=NULL,
method="auto",test="chisq", # glmLRT arguments
abundance.trend=TRUE,robust=FALSE,
winsor.tail.p=c(0.05,0.1) # glmLFTest arguments
))
},
noiseq = {
return(list(
k=0.5,norm="n",replicates="biological",
factor="class",conditions=NULL,pnr=0.2,
nss=5,v=0.02,lc=1, # noiseq general and specific arguments
nclust=15,r=100,adj=1.5,
a0per=0.9,filter=0,depth=NULL,
cv.cutoff=500,cpm=1 # noiseqbio specific arguments
))
},
bayseq = {
return(list(samplesize=10000,samplingSubset=NULL,
equalDispersions=TRUE,estimation="QL",zeroML=FALSE,
consensus=FALSE,moderate=TRUE,pET="BIC",
marginalise=FALSE,subset=NULL,priorSubset=NULL,
bootStraps=1,conv=1e-4,nullData=FALSE,returnAll=FALSE,
returnPD=FALSE,discardSampling=FALSE,cl=NULL))
},
limma = {
return(list(normalize.method="none"))
},
nbpseq = {
return(list(
main.method="nbsmyth",
model=list(nbpseq="log-linear-rel-mean",nbsmyth="NBP"),
tests="HOA",
alternative="two.sided"
))
}
)
},
gene.filter = {
return(list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.75
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.defaults("biotype.filter",method[1]),
presence=list(
frac=0.25,
min.count=10,
per.condition=FALSE
)
))
},
exon.filter = {
return(list(
mnrpx=list(
exons.per.gene=5,
min.exons=2,
frac=1/5
)
))
},
biotype.filter = {
switch(method,
hg18 = {
return(list(
unprocessed_pseudogene=TRUE,
pseudogene=FALSE,
miRNA=FALSE,
retrotransposed=FALSE,
protein_coding=FALSE,
processed_pseudogene=FALSE,
snRNA=FALSE,
snRNA_pseudogene=TRUE,
Mt_tRNA_pseudogene=TRUE,
miRNA_pseudogene=TRUE,
misc_RNA=FALSE,
tRNA_pseudogene=TRUE,
snoRNA=FALSE,
scRNA_pseudogene=TRUE,
rRNA_pseudogene=TRUE,
snoRNA_pseudogene=TRUE,
rRNA=TRUE,
misc_RNA_pseudogene=TRUE,
IG_V_gene=FALSE,
IG_D_gene=FALSE,
IG_J_gene=FALSE,
IG_C_gene=FALSE,
IG_pseudogene=TRUE,
scRNA=FALSE
))
},
hg19 = {
return(list(
pseudogene=FALSE,
lincRNA=FALSE,
protein_coding=FALSE,
antisense=FALSE,
processed_transcript=FALSE,
snRNA=FALSE,
sense_intronic=FALSE,
miRNA=FALSE,
misc_RNA=FALSE,
snoRNA=FALSE,
rRNA=TRUE,
polymorphic_pseudogene=FALSE,
sense_overlapping=FALSE,
three_prime_overlapping_ncrna=FALSE,
TR_V_gene=FALSE,
TR_V_pseudogene=TRUE,
TR_D_gene=FALSE,
TR_J_gene=FALSE,
TR_C_gene=FALSE,
TR_J_pseudogene=TRUE,
IG_C_gene=FALSE,
IG_C_pseudogene=TRUE,
IG_J_gene=FALSE,
IG_J_pseudogene=TRUE,
IG_D_gene=FALSE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE
))
},
hg38 = {
return(list(
protein_coding=FALSE,
polymorphic_pseudogene=FALSE,
lincRNA=FALSE,
unprocessed_pseudogene=TRUE,
processed_pseudogene=FALSE,
antisense=FALSE,
processed_transcript=FALSE,
transcribed_unprocessed_pseudogene=FALSE,
sense_intronic=FALSE,
unitary_pseudogene=TRUE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE,
TR_V_gene=FALSE,
sense_overlapping=FALSE,
transcribed_processed_pseudogene=FALSE,
miRNA=FALSE,
snRNA=FALSE,
misc_RNA=FALSE,
rRNA=TRUE,
snoRNA=FALSE,
IG_J_pseudogene=TRUE,
IG_J_gene=FALSE,
IG_D_gene=FALSE,
three_prime_overlapping_ncrna=FALSE,
IG_C_gene=FALSE,
IG_C_pseudogene=TRUE,
pseudogene=TRUE,
TR_V_pseudogene=TRUE,
Mt_tRNA=TRUE,
Mt_rRNA=TRUE,
translated_processed_pseudogene=FALSE,
TR_J_gene=FALSE,
TR_C_gene=FALSE,
TR_D_gene=FALSE,
TR_J_pseudogene=TRUE,
LRG_gene=FALSE
))
},
mm9 = {
return(list(
pseudogene=FALSE,
snRNA=FALSE,
protein_coding=FALSE,
antisense=FALSE,
miRNA=FALSE,
lincRNA=FALSE,
snoRNA=FALSE,
processed_transcript=FALSE,
misc_RNA=FALSE,
rRNA=TRUE,
sense_overlapping=FALSE,
sense_intronic=FALSE,
polymorphic_pseudogene=FALSE,
non_coding=FALSE,
three_prime_overlapping_ncrna=FALSE,
IG_C_gene=FALSE,
IG_J_gene=FALSE,
IG_D_gene=FALSE,
IG_V_gene=FALSE,
ncrna_host=FALSE
))
},
mm10 = {
return(list(
pseudogene=FALSE,
snRNA=FALSE,
protein_coding=FALSE,
antisense=FALSE,
miRNA=FALSE,
snoRNA=FALSE,
lincRNA=FALSE,
processed_transcript=FALSE,
misc_RNA=FALSE,
rRNA=TRUE,
sense_intronic=FALSE,
sense_overlapping=FALSE,
polymorphic_pseudogene=FALSE,
IG_C_gene=FALSE,
IG_J_gene=FALSE,
IG_D_gene=FALSE,
IG_LV_gene=FALSE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE,
TR_V_gene=FALSE,
TR_V_pseudogene=TRUE,
three_prime_overlapping_ncrna=FALSE
))
},
dm3 = {
return(list(
protein_coding=FALSE,
ncRNA=FALSE,
snoRNA=FALSE,
pre_miRNA=FALSE,
pseudogene=FALSE,
snRNA=FALSE,
tRNA=FALSE,
rRNA=TRUE
))
},
rn5 = {
return(list(
protein_coding=FALSE,
pseudogene=FALSE,
processed_pseudogene=FALSE,
miRNA=FALSE,
rRNA=TRUE,
misc_RNA=FALSE
))
},
rn6 = {
return(list(
antisense=FALSE,
lincRNA=FALSE,
miRNA=FALSE,
misc_RNA=FALSE,
processed_pseudogene=FALSE,
processed_transcript=FALSE,
protein_coding=FALSE,
pseudogene=FALSE,
ribozyme=FALSE,
rRNA=TRUE,
scaRNA=FALSE,
sense_intronic=FALSE,
snoRNA=FALSE,
snRNA=FALSE,
sRNA=FALSE,
TEC=FALSE,
transcribed_processed_pseudogene=FALSE,
transcribed_unprocessed_pseudogene=FALSE,
unprocessed_pseudogene=FALSE
))
},
danrer7 = {
return(list(
antisense=FALSE,
protein_coding=FALSE,
miRNA=FALSE,
snoRNA=FALSE,
rRNA=TRUE,
lincRNA=FALSE,
processed_transcript=FALSE,
snRNA=FALSE,
pseudogene=FALSE,
sense_intronic=FALSE,
misc_RNA=FALSE,
polymorphic_pseudogene=FALSE,
IG_V_pseudogene=TRUE,
IG_C_pseudogene=TRUE,
IG_J_pseudogene=TRUE,
non_coding=FALSE,
sense_overlapping=FALSE
))
},
pantro4 = {
return(list(
protein_coding=FALSE,
pseudogene=FALSE,
processed_pseudogene=FALSE,
miRNA=FALSE,
rRNA=TRUE,
snRNA=FALSE,
snoRNA=FALSE,
misc_RNA=FALSE
))
},
susscr3 = {
return(list(
antisense=FALSE,
protein_coding=FALSE,
lincRNA=FALSE,
pseudogene=FALSE,
processed_transcript=FALSE,
miRNA=FALSE,
rRNA=TRUE,
snRNA=FALSE,
snoRNA=FALSE,
misc_RNA=FALSE,
non_coding=FALSE,
IG_C_gene=FALSE,
IG_J_gene=FALSE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE
))
},
tair10 = {
return(list(
miRNA=FALSE,
ncRNA=FALSE,
protein_coding=FALSE,
pseudogene=FALSE,
rRNA=TRUE,
snoRNA=FALSE,
snRNA=FALSE,
transposable_element=FALSE,
tRNA=FALSE
))
},
equcab2 = {
return(list(
miRNA=FALSE,
misc_RNA=FALSE,
protein_coding=FALSE,
pseudogene=FALSE,
processed_pseudogene=FALSE,
rRNA=TRUE,
snoRNA=FALSE,
snRNA=FALSE
))
}
)
}
)
}
#' Validate normalization and statistical algorithm arguments
#'
#' This function checks and validates the arguments passed by the user to the
#' normalization and statistics algorithms supported by metaseqR. As these are
#' given into lists and passed to the algorithms, the list members must be checked
#' for \code{NULL}, valid names etc. This function performs these checks and
#' ignores any invalid arguments.
#'
#' @param normalization a keyword determining the normalization strategy to be
#' performed by metaseqR. See \code{\link{metaseqr}} main help page for details.
#' @param statistics the statistical tests to be performed by metaseqR. See
#' \code{\link{metaseqr}} main help page for details.
#' @param norm.args the user input list of normalization arguments. See
#' \code{\link{metaseqr}} main help page for details.
#' @param stat.args the user input list of statistical test arguments. See
#' \code{\link{metaseqr}} main help page for details.
#' @return A list with two members (\code{norm.args}, \code{stat.args}) with valid
#' arguments to be used as user input for the algorithms supported by metaseqR.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' normalization <- "edaseq"
#' statistics <- "edger"
#' norm.args <- get.defaults("normalization","edaseq")
#' stat.args <- get.defaults("statistics","deseq")
#' # Will return as is
#' val <- validate.alg.args(normalization,statistics,norm.args,stat.args)
#' val$norm.args
#' val$stat.args
#' # but...
#' stat.args <- c(stat.args,my.irrelevant.arg=999)
#' val <- validate.alg.args(normalization,statistics,norm.args,stat.args)
#' # irrelevant argument will be removed
#' val$norm.args
#' val$stat.args
#'}
validate.alg.args <- function(normalization,statistics,norm.args,stat.args) {
if (normalization=="each") {
if (!is.null(norm.args)) {
for (s in statistics) {
if (!is.null(norm.args[[s]])) {
switch(s,
deseq = {
tmp <- norm.args[[s]]
tmp <- validate.list.args("normalization",s,tmp)
norm.args[[s]] <- get.defaults("normalization",s)
if (length(tmp)>0)
norm.args[[s]] <- set.arg(norm.args[[s]],tmp)
},
edger = {
tmp <- norm.args[[s]]
tmp <- validate.list.args("statistics",s,tmp)
norm.args[[s]] <- get.defaults("normalization",s)
if (length(tmp)>0)
norm.args[[s]] <- set.arg(norm.args[[s]],tmp)
},
limma = {
tmp <- norm.args[[s]]
tmp <- validate.list.args("statistics","edger",tmp)
norm.args[[s]] <- get.defaults("normalization",
"edger")
if (length(tmp)>0)
norm.args[[s]] <- set.arg(norm.args[[s]],tmp)
},
nbpseq = {
tmp <- norm.args[[s]]
tmp <- validate.list.args("statistics",s,tmp)
norm.args[[s]] <- get.defaults("normalization",s)
if (length(tmp)>0)
norm.args[[s]] <- set.arg(norm.args[[s]],tmp)
},
noiseq = {
tmp <- norm.args[[s]]
tmp <- validate.list.args("statistics",s,tmp)
norm.args[[s]] <- get.defaults("normalization",s)
if (length(tmp)>0)
norm.args[[s]] <- set.arg(norm.args[[s]],tmp)
},
bayseq = {
tmp <- norm.args[[s]]
tmp <- validate.list.args("statistics","edger",tmp)
norm.args[[s]] <- get.defaults("normalization",
"edger")
if (length(tmp)>0)
norm.args[[s]] <- set.arg(norm.args[[s]],tmp)
}
)
}
else {
switch(s,
deseq = {
norm.args[[s]] <- get.defaults(normalization,
"deseq")
},
edger = {
norm.args[[s]] <- get.defaults(normalization,
"edger")
},
limma = {
norm.args[[s]] <- get.defaults(normalization,
"edger")
},
nbpseq = {
norm.args[[s]] <- get.defaults(normalization,
"nbpseq")
},
noiseq = {
norm.args[[s]] <- get.defaults(normalization,
"noiseq")
},
bayseq = {
norm.args[[s]] <- get.defaults(normalization,
"edger")
}
)
}
}
}
else {
norm.args <- vector("list",length(statistics))
names(norm.args) <- statistics
for (s in statistics) {
switch(s,
deseq = {
norm.args[[s]] <- get.defaults(normalization,"deseq")
},
edger = {
norm.args[[s]] <- get.defaults(normalization,"edger")
},
limma = {
norm.args[[s]] <- get.defaults(normalization,"edger")
},
nbpseq = {
norm.args[[s]] <- get.defaults(normalization,"nbpseq")
},
noiseq = {
norm.args[[s]] <- get.defaults(normalization,"noiseq")
},
bayseq = {
norm.args[[s]] <- get.defaults(normalization,"edger")
}
)
}
}
}
else {
if (!is.null(norm.args))
{
tmp <- norm.args
tmp <- validate.list.args("normalization",normalization,tmp)
norm.args <- get.defaults("normalization",normalization)
if (length(tmp)>0)
norm.args <- set.arg(norm.args,tmp)
}
else
norm.args <- get.defaults("normalization",normalization)
}
for (s in statistics) {
if (!is.null(stat.args[[s]])) {
tmp <- stat.args[[s]]
tmp <- validate.list.args("statistics",s,tmp)
stat.args[[s]] <- get.defaults("statistics",s)
if (length(tmp)>0)
stat.args[[s]] <- set.arg(stat.args[[s]],tmp)
}
else
stat.args[[s]] <- get.defaults("statistics",s)
}
return(list(norm.args=norm.args,stat.args=stat.args))
}
#' Validate list parameters for several metaseqR functions
#'
#' This function validates the arguments passed by the user to the normalization,
#' statistics and filtering algorithms supported by metaseqR. As these are given
#' into lists and passed to the algorithms, the list member names must be valid
#' algorithm arguments for the pipeline not to crash. This function performs these
#' checks and ignores any invalid arguments.
#'
#' @param what a keyword determining the procedure for which to validate arguments.
#' It can be one of \code{"normalization"}, \code{"statistics"}, \code{"gene.filter"},
#' \code{"exon.filter"} or \code{"biotype.filter"}.
#' @param method the normalization/statistics/filtering algorithm included in
#' metaseqR for which to validate user input. When \code{what} is
#' \code{"normalization"}, method is one of \code{"edaseq"}, \code{"deseq"},
#' \code{"edger"}, \code{"noiseq"} or \code{"nbpseq"}. When \code{what} is
#' \code{"statistics"}, method is one of \code{"deseq"}, \code{"edger"},
#' \code{"noiseq"}, \code{"bayseq"}, \code{"limma"} or \code{"nbpseq"}. When
#' \code{method} is \code{"biotype.filter"}, \code{what} is the input organism
#' (see the main \code{\link{metaseqr}} help page for a list of supported organisms).
#' @param arg.list the user input list of arguments.
#' @return A list with valid arguments to be used as user input in the algorithms
#' supported by metaseqR.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' norm.args.edger <- list(method="TMM",refColumn=NULL,
#' logratioTrim=0.3,sumTrim=0.05,doWeighting=TRUE,
#' Bcutoff=-1e10,p=0.75)
#' # Bcutoff does not exist, will throw a warning and ignore it.
#' norm.args.edger <- validate.list.args("normalization","edger",norm.args.edger)
#'}
validate.list.args <- function(what,method=NULL,arg.list) {
what <- tolower(what)
check.text.args("what",what,c("normalization","statistics","gene.filter",
"exon.filter","biotype.filter"))
if (what %in% c("normalization","statistics") && is.null(method))
stopwrap("The method argument must be provided when what is ",
"\"normalization\" or \"statistics\"!")
switch(what,
normalization = {
switch(method,
edaseq = {
valid <- names(arg.list) %in% c("within.which",
"between.which")
not.valid <- which(!valid)
},
deseq = {
valid <- names(arg.list) %in% c("locfunc")
not.valid <- which(!valid)
},
edger = {
valid <- names(arg.list) %in% c("method","refColumn",
"logratioTrim","sumTrim","doWeighting","Acutoff","p")
not.valid <- which(!valid)
},
noiseq = {
valid <- names(arg.list) %in% c("method","long","lc","k",
"refColumn","logratioTrim","sumTrim","doWeighting",
"Acutoff")
not.valid <- which(!valid)
},
nbpseq = {
valid <- names(arg.list) %in% c("main.method","method",
"thinning")
not.valid <- which(!valid)
}
)
if (length(not.valid)>0) {
warnwrap(paste("The following",method,what,"argument names",
"are invalid and will be ignored:",
paste(names(arg.list)[not.valid],collapse=", ")))
arg.list[not.valid] <- NULL
}
return(arg.list)
},
statistics = {
switch(method,
deseq = {
valid <- names(arg.list) %in% c("method","sharingMode",
"fitType")
not.valid <- which(!valid)
},
edger = {
valid <- names(arg.list) %in% c("main.method",
"rowsum.filter","prior.df","trend","span","tag.method",
"grid.length","grid.range","offset","glm.method",
"subset","AveLogCPM","trend.method","dispersion",
"offset","weights","lib.size","prior.count","start",
"method","abundance.trend","robust",
"winsor.tail.p")
not.valid <- which(!valid)
},
noiseq = {
valid <- names(arg.list) %in% c("k","norm","replicates",
"factor","conditions","pnr","nss","v","lc","nclust","r",
"adj","a0per","filter","depth","cv.cutoff","cpm")
not.valid <- which(!valid)
},
bayseq = {
valid <- names(arg.list) %in% c("samplesize",
"samplingSubset","equalDispersions","estimation",
"zeroML","consensus","moderate","pET","marginalise",
"subset","priorSubset","bootStraps","conv","nullData",
"returnAll","returnPD","discardSampling","cl")
not.valid <- which(!valid)
},
limma = {
valid <- names(arg.list) %in% c("normalize.method")
not.valid <- which(!valid)
},
nbpseq = {
valid <- names(arg.list) %in% c("main.method","method",
"tests","alternative")
not.valid <- which(!valid)
}
)
if (length(not.valid)>0) {
warnwrap(paste("The following",method,what,"argument names",
"are invalid and will be ignored:",
paste(names(arg.list)[not.valid],collapse=", ")))
arg.list[not.valid] <- NULL
}
return(arg.list)
},
gene.filter = {
valid.1 <- names(arg.list) %in% c("length","avg.reads","expression",
"biotype","presence")
not.valid.1 <- which(!valid.1)
if (length(not.valid.1)>0) {
warnwrap(paste("The following",method,what,"argument names",
"are invalid and will be ignored:",
paste(names(arg.list)[not.valid.1],collapse=", ")))
arg.list[not.valid.1] <- NULL
}
if (length(arg.list)>0) {
for (n in names(arg.list)) {
switch(n,
length = {
valid.2 <- names(arg.list[[n]]) %in% c("length")
not.valid.2 <- which(!valid.2)
},
avg.reads = {
valid.2 <- names(arg.list[[n]]) %in%
c("average.per.bp","quantile")
not.valid.2 <- which(!valid.2)
},
expression = {
valid.2 <- names(arg.list[[n]]) %in% c("median",
"mean","quantile","known","custom")
not.valid.2 <- which(!valid.2)
},
presence = {
valid.2 <- names(arg.list[[n]]) %in% c("frac",
"min.count","per.condition")
not.valid.2 <- which(!valid.2)
}
)
if (length(not.valid.2)>0) {
warnwrap(paste("The following",method,what,
"sub-argument names are invalid and will be",
"ignored:",paste(names(arg.list[[n]])[not.valid.2],
collapse=", ")))
arg.list[[n]][not.valid.2] <- NULL
}
}
}
return(arg.list)
},
exon.filter = {
valid.1 <- names(arg.list) %in% c("mnrpx")
not.valid.1 <- which(!valid.1)
if (length(not.valid.1)>0) {
warnwrap(paste("The following",method,what,"argument names",
"are invalid and will be ignored:",
paste(names(arg.list)[not.valid.1],collapse=", ")))
arg.list[not.valid.1] <- NULL
}
if (length(arg.list)>0) {
for (n in names(arg.list)) {
switch(n,
mnrpx = {
valid.2 <- names(arg.list[[n]]) %in%
c("exons.per.gene","min.exons","frac")
not.valid.2 <- which(!valid.2)
}
)
if (length(not.valid.2)>0) {
warnwrap(paste("The following",method,what,
"sub-argument names are invalid and will be",
"ignored:",paste(names(arg.list[[n]])[not.valid.2],
collapse=", ")))
arg.list[[n]][not.valid.2] <- NULL
}
}
}
return(arg.list)
},
biotype.filter = {
switch(method,
hg18 = {
valid <- names(arg.list) %in% c("unprocessed_pseudogene",
"pseudogene","miRNA","retrotransposed","protein_coding",
"processed_pseudogene","snRNA","snRNA_pseudogene",
"Mt_tRNA_pseudogene","miRNA_pseudogene","misc_RNA",
"tRNA_pseudogene","snoRNA","scRNA_pseudogene",
"rRNA_pseudogene","snoRNA_pseudogene","rRNA",
"misc_RNA_pseudogene","IG_V_gene","IG_D_gene",
"IG_J_gene","IG_C_gene","IG_pseudogene","scRNA")
not.valid <- which(!valid)
},
hg19 = {
valid <- names(arg.list) %in% c("pseudogene","lincRNA",
"protein_coding","antisense","processed_transcript",
"snRNA","sense_intronic","miRNA","misc_RNA","snoRNA",
"rRNA","polymorphic_pseudogene","sense_overlapping",
"three_prime_overlapping_ncrna","TR_V_gene",
"TR_V_pseudogene","TR_D_gene","TR_J_gene","TR_C_gene",
"TR_J_pseudogene","IG_C_gene","IG_C_pseudogene",
"IG_J_gene","IG_J_pseudogene","IG_D_gene","IG_V_gene",
"IG_V_pseudogene")
not.valid <- which(!valid)
},
mm9 = {
valid <- names(arg.list) %in% c("pseudogene","snRNA",
"protein_coding","antisense","miRNA","lincRNA","snoRNA",
"processed_transcript","misc_RNA","rRNA",
"sense_overlapping","sense_intronic",
"polymorphic_pseudogene","non_coding",
"three_prime_overlapping_ncrna","IG_C_gene","IG_J_gene",
"IG_D_gene","IG_V_gene","ncrna_host")
not.valid <- which(!valid)
},
mm10 = {
valid <- names(arg.list) %in% c("pseudogene","snRNA",
"protein_coding","antisense","miRNA","snoRNA","lincRNA",
"processed_transcript","misc_RNA","rRNA",
"sense_intronic","sense_overlapping",
"polymorphic_pseudogene","IG_C_gene","IG_J_gene",
"IG_D_gene","IG_LV_gene","IG_V_gene","IG_V_pseudogene",
"TR_V_gene","TR_V_pseudogene",
"three_prime_overlapping_ncrna")
not.valid <- which(!valid)
},
dm3 = {
valid <- names(arg.list) %in% c("protein_coding","ncRNA",
"snoRNA","pre_miRNA","pseudogene","snRNA","tRNA","rRNA")
not.valid <- which(!valid)
},
rn5 = {
valid <- names(arg.list) %in% c("protein_coding",
"pseudogene","processed_pseudogene","miRNA","rRNA",
"misc_RNA")
not.valid <- which(!valid)
},
danrer7 = {
valid <- names(arg.list) %in% c("antisense",
"protein_coding","miRNA","snoRNA","rRNA","lincRNA",
"processed_transcript","snRNA","pseudogene",
"sense_intronic","misc_RNA","polymorphic_pseudogene",
"IG_V_pseudogene","IG_C_pseudogene","IG_J_pseudogene",
"non_coding","sense_overlapping")
not.valid <- which(!valid)
},
pantro4 = {
valid <- names(arg.list) %in% c("protein_coding",
"pseudogene","processed_pseudogene","miRNA","rRNA",
"snRNA","snoRNA","misc_RNA")
not.valid <- which(!valid)
},
susscr3 = {
valid <- names(arg.list) %in% c("antisense",
"protein_coding","lincRNA","pseudogene",
"processed_transcript","miRNA","rRNA","snRNA","snoRNA",
"misc_RNA","non_coding","IG_C_gene","IG_J_gene",
"IG_V_gene","IG_V_pseudogene")
not.valid <- which(!valid)
},
tair10 = {
valid <- names(arg.list) %in% c("miRNA","ncRNA",
"protein_coding","pseudogene","rRNA","snoRNA",
"snRNA","transposable_element","tRNA")
not.valid <- which(!valid)
},
equcab2 = {
valid <- names(arg.list) %in% c("miRNA","misc_RNA",
"protein_coding","pseudogene","processed_pseudogene",
"rRNA","snoRNA","snRNA")
}
)
if (length(not.valid)>0) {
warnwrap(paste("The following",method,what,"argument names",
"are invalid and will be ignored:",
paste(names(arg.list)[not.valid],collapse=", ")))
arg.list[not.valid] <- NULL
}
return(arg.list)
}
)
}
#' Group together a more strict biotype filter
#'
#' Returns a list with TRUE/FALSE according to the biotypes that are going to be
#' filtered in a more strict way than the defaults. This is a helper function for
#' the analysis presets of metaseqR. Internal use only.
#'
#' @param org one of the supported organisms.
#' @return A list of booleans, one for each biotype.
#' @author <NAME>
#' @examples
#' \dontrun{
#' sf <- get.strict.biofilter("hg18")
#'}
get.strict.biofilter <- function(org) {
switch(org,
hg18 = {
return(list(
unprocessed_pseudogene=TRUE,
pseudogene=TRUE,
miRNA=FALSE,
retrotransposed=FALSE,
protein_coding=FALSE,
processed_pseudogene=TRUE,
snRNA=FALSE,
snRNA_pseudogene=TRUE,
Mt_tRNA_pseudogene=TRUE,
miRNA_pseudogene=TRUE,
misc_RNA=TRUE,
tRNA_pseudogene=TRUE,
snoRNA=TRUE,
scRNA_pseudogene=TRUE,
rRNA_pseudogene=TRUE,
snoRNA_pseudogene=TRUE,
rRNA=TRUE,
misc_RNA_pseudogene=TRUE,
IG_V_gene=FALSE,
IG_D_gene=FALSE,
IG_J_gene=FALSE,
IG_C_gene=FALSE,
IG_pseudogene=TRUE,
scRNA=FALSE
))
},
hg19 = {
return(list(
pseudogene=TRUE,
lincRNA=FALSE,
protein_coding=FALSE,
antisense=FALSE,
processed_transcript=FALSE,
snRNA=FALSE,
sense_intronic=FALSE,
miRNA=FALSE,
misc_RNA=FALSE,
snoRNA=TRUE,
rRNA=TRUE,
polymorphic_pseudogene=TRUE,
sense_overlapping=FALSE,
three_prime_overlapping_ncrna=FALSE,
TR_V_gene=FALSE,
TR_V_pseudogene=TRUE,
TR_D_gene=FALSE,
TR_J_gene=FALSE,
TR_C_gene=FALSE,
TR_J_pseudogene=TRUE,
IG_C_gene=FALSE,
IG_C_pseudogene=TRUE,
IG_J_gene=FALSE,
IG_J_pseudogene=TRUE,
IG_D_gene=FALSE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE
))
},
hg38 = {
return(list(
protein_coding=FALSE,
polymorphic_pseudogene=TRUE,
lincRNA=FALSE,
unprocessed_pseudogene=TRUE,
processed_pseudogene=TRUE,
antisense=FALSE,
processed_transcript=FALSE,
transcribed_unprocessed_pseudogene=TRUE,
sense_intronic=FALSE,
unitary_pseudogene=TRUE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE,
TR_V_gene=FALSE,
sense_overlapping=FALSE,
transcribed_processed_pseudogene=TRUE,
miRNA=FALSE,
snRNA=FALSE,
misc_RNA=FALSE,
rRNA=TRUE,
snoRNA=TRUE,
IG_J_pseudogene=TRUE,
IG_J_gene=FALSE,
IG_D_gene=FALSE,
three_prime_overlapping_ncrna=FALSE,
IG_C_gene=FALSE,
IG_C_pseudogene=TRUE,
pseudogene=TRUE,
TR_V_pseudogene=TRUE,
Mt_tRNA=TRUE,
Mt_rRNA=TRUE,
translated_processed_pseudogene=TRUE,
TR_J_gene=FALSE,
TR_C_gene=FALSE,
TR_D_gene=FALSE,
TR_J_pseudogene=TRUE,
LRG_gene=FALSE
))
},
mm9 = {
return(list(
pseudogene=TRUE,
snRNA=FALSE,
protein_coding=FALSE,
antisense=FALSE,
miRNA=FALSE,
lincRNA=FALSE,
snoRNA=TRUE,
processed_transcript=FALSE,
misc_RNA=TRUE,
rRNA=TRUE,
sense_overlapping=FALSE,
sense_intronic=FALSE,
polymorphic_pseudogene=TRUE,
non_coding=FALSE,
three_prime_overlapping_ncrna=FALSE,
IG_C_gene=FALSE,
IG_J_gene=FALSE,
IG_D_gene=FALSE,
IG_V_gene=FALSE,
ncrna_host=FALSE
))
},
mm10 = {
return(list(
pseudogene=TRUE,
snRNA=FALSE,
protein_coding=FALSE,
antisense=FALSE,
miRNA=FALSE,
snoRNA=TRUE,
lincRNA=FALSE,
processed_transcript=FALSE,
misc_RNA=TRUE,
rRNA=TRUE,
sense_intronic=FALSE,
sense_overlapping=FALSE,
polymorphic_pseudogene=TRUE,
IG_C_gene=FALSE,
IG_J_gene=FALSE,
IG_D_gene=FALSE,
IG_LV_gene=FALSE,
IG_V_gene=FALSE,
IG_V_pseudogene=TRUE,
TR_V_gene=FALSE,
TR_V_pseudogene=TRUE,
three_prime_overlapping_ncrna=FALSE
))
},
dm3 = {
return(list(
protein_coding=FALSE,
ncRNA=FALSE,
snoRNA=TRUE,
pre_miRNA=FALSE,
pseudogene=TRUE,
snRNA=FALSE,
tRNA=FALSE,
rRNA=TRUE
))
},
rn5 = {
return(list(
protein_coding=FALSE,
pseudogene=TRUE,
processed_pseudogene=FALSE,
miRNA=FALSE,
rRNA=TRUE,
misc_RNA=TRUE
))
},
danrer7 = {
return(list(
antisense=FALSE,
protein_coding=FALSE,
miRNA=FALSE,
snoRNA=TRUE,
rRNA=TRUE,
lincRNA=FALSE,
processed_transcript=FALSE,
snRNA=FALSE,
pseudogene=TRUE,
sense_intronic=FALSE,
misc_RNA=TRUE,
polymorphic_pseudogene=TRUE,
IG_V_pseudogene=TRUE,
IG_C_pseudogene=TRUE,
IG_J_pseudogene=TRUE,
non_coding=FALSE,
sense_overlapping=FALSE
))
},
pantro4 = {
return(list(
protein_coding=FALSE,
pseudogene=TRUE,
processed_pseudogene=TRUE,
miRNA=FALSE,
rRNA=TRUE,
snRNA=TRUE,
snoRNA=TRUE,
misc_RNA=TRUE
))
},
susscr3 = {
return(list(
antisense=FALSE,
protein_coding=FALSE,
lincRNA=FALSE,
pseudogene=TRUE,
processed_transcript=FALSE,
miRNA=FALSE,
rRNA=TRUE,
snRNA=TRUE,
snoRNA=TRUE,
misc_RNA=TRUE,
non_coding=FALSE,
IG_C_gene=TRUE,
IG_J_gene=TRUE,
IG_V_gene=TRUE,
IG_V_pseudogene=FALSE
))
},
tair10 = {
return(list(
miRNA=FALSE,
ncRNA=FALSE,
protein_coding=FALSE,
pseudogene=TRUE,
rRNA=TRUE,
snoRNA=TRUE,
snRNA=TRUE,
transposable_element=FALSE,
tRNA=TRUE
))
},
equcab2 = {
return(list(
miRNA=FALSE,
misc_RNA=TRUE,
protein_coding=FALSE,
pseudogene=FALSE,
processed_pseudogene=FALSE,
rRNA=TRUE,
snoRNA=TRUE,
snRNA=TRUE
))
}
)
}
#' Return several analysis options given an analysis preset
#'
#' This is a helper function which returns a set of metaseqr pipeline options,
#' grouped together according to a preset keyword. It is intended mostly for
#' internal use.
#'
#' @param preset preset can be one of \code{"all.basic"}, \code{"all.normal"},
#' \code{"all.full"}, \code{"medium.basic"}, \code{"medium.normal"},
#' @param org one of the supported organisms. See \code{\link{metaseqr}} main
#' help page.
#' \code{"medium.full"}, \code{"strict.basic"}, \code{"strict.normal"} or
#' \code{"strict.full"}, each of which control the strictness of the analysis and
#' the amount of data to be exported. For an explanation of the presets, see the
#' main \code{\link{metaseqr}} help page.
#' @return A named list with names \code{exon.filters}, \code{gene.filters},
#' \code{pcut}, \code{export.what}, \code{export.scale}, \code{export.values} and
#' \code{export.stats}, each of which correspond to an element of the metaseqr
#' pipeline.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' strict.preset <- get.preset.opts("strict.basic","mm9")
#'}
get.preset.opts <- function(preset,org) {
# Override filter rules and maybe norm.args and stat.args
switch(preset,
all.basic = {
exon.filters <- NULL
gene.filters <- NULL
pcut <- NA
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change")
export.scale <- c("natural","log2")
export.values <- c("normalized")
export.stats <- c("mean")
},
all.normal = {
exon.filters <- NULL
gene.filters <- NULL
pcut <- NA
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change","stats",
"counts")
export.scale <- c("natural","log2")
export.values <- c("normalized")
export.stats <- c("mean","sd","cv")
},
all.full = {
exon.filters <- NULL
gene.filters <- NULL
pcut <- NA
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change","stats",
"counts","flags")
export.scale <- c("natural","log2","log10","vst")
export.values <- c("raw","normalized")
export.stats <- c("mean","median","sd","mad","cv","rcv")
},
medium.basic = {
exon.filters <- list(
min.active.exons=list(
exons.per.gene=5,
min.exons=2,
frac=1/5
)
)
gene.filters <- list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.25
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.defaults("biotype.filter",org[1])
)
pcut <- 0.05
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change")
export.scale <- c("natural","log2")
export.values <- c("normalized")
export.stats <- c("mean")
},
medium.normal = {
exon.filters <- list(
min.active.exons=list(
exons.per.gene=5,
min.exons=2,
frac=1/5
)
)
gene.filters <- list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.25
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.defaults("biotype.filter",org[1])
)
pcut <- 0.05
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change","stats",
"counts")
export.scale <- c("natural","log2")
export.values <- c("normalized")
export.stats <- c("mean","sd","cv")
},
medium.full = {
exon.filters <- list(
min.active.exons=list(
exons.per.gene=5,
min.exons=2,
frac=1/5
)
)
gene.filters <- list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.25
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.defaults("biotype.filter",org[1])
)
pcut <- 0.05
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change","stats",
"counts","flags")
export.scale <- c("natural","log2","log10","vst")
export.values <- c("raw","normalized")
export.stats <- c("mean","median","sd","mad","cv","rcv")
},
strict.basic = {
exon.filters=list(
min.active.exons=list(
exons.per.gene=4,
min.exons=2,
frac=1/4
)
)
gene.filters=list(
length=list(
length=750
),
avg.reads=list(
average.per.bp=100,
quantile=0.5
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.strict.biofilter(org[1])
)
pcut <- 0.01
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change")
export.scale <- c("natural","log2")
export.values <- c("normalized")
export.stats <- c("mean")
},
strict.normal = {
exon.filters=list(
min.active.exons=list(
exons.per.gene=4,
min.exons=2,
frac=1/4
)
)
gene.filters=list(
length=list(
length=750
),
avg.reads=list(
average.per.bp=100,
quantile=0.5
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.strict.biofilter(org[1])
)
pcut <- 0.01
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change","stats",
"counts")
export.scale <- c("natural","log2")
export.values <- c("normalized")
export.stats <- c("mean","sd","cv")
},
strict.full = {
exon.filters=list(
min.active.exons=list(
exons.per.gene=4,
min.exons=2,
frac=1/4
)
)
gene.filters=list(
length=list(
length=750
),
avg.reads=list(
average.per.bp=100,
quantile=0.5
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.strict.biofilter(org[1])
)
pcut <- 0.01
export.what <- c("annotation","p.value","adj.p.value",
"meta.p.value","adj.meta.p.value","fold.change","stats",
"counts","flags")
export.scale <- c("natural","log2","log10","vst")
export.values <- c("raw","normalized")
export.stats <- c("mean","median","sd","mad","cv","rcv")
}
)
preset.opts <- list(
exon.filters=exon.filters,
gene.filters=gene.filters,
pcut=pcut,
export.what=export.what,
export.scale=export.scale,
export.values=export.values,
export.stats=export.stats
)
return(preset.opts)
}
#' Calculates fold changes
#'
#' Returns a matrix of fold changes based on the requested contrast, the list of
#' all samples and the data matrix which is produced by the metaseqr workflow. For
#' details on the \code{contrast}, \code{sample.list} and \code{log.offset}
#' parameters, see the main usage page of metaseqr. This function is intended
#' mostly for internal use but can also be used independently.
#'
#' @param contrast the vector of requested statistical comparison contrasts.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param data.matrix a matrix of gene expression data whose column names are the
#' same as the sample names included in the sample list.
#' @param log.offset a number to be added to each element of data matrix in order
#' to avoid Infinity on log type data transformations.
#' @return A matrix of fold change ratios, treatment to control, as these are
#' parsed from contrast.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' fc <- make.fold.change("Control_vs_Treatment",list(Control=c("C1","C2"),
#' Treatment=c("T1","T2")),data.matrix)
#'}
make.fold.change <- function(contrast,sample.list,data.matrix,log.offset=1) {
conds <- strsplit(contrast,"_vs_")[[1]]
#f (!is.matrix(data.matrix) || !is.data.frame(data.matrix)) { # Vector, nrow=1
# nn <- names(data.matrix)
# data.matrix <- t(as.matrix(data.matrix))
# colnames(data.matrix) <- nn
#}
fold.mat <- matrix(0,nrow(data.matrix),length(conds)-1)
for (i in 2:length(conds)) { # First condition is ALWAYS reference
samples.nom <- sample.list[[conds[i]]]
samples.denom <- sample.list[[conds[1]]]
nom <- data.matrix[,match(samples.nom,colnames(data.matrix)),drop=FALSE]
denom <- data.matrix[,match(samples.denom,colnames(data.matrix)),
drop=FALSE]
if (!is.matrix(nom)) nom <- as.matrix(nom) # Cover the case with no replicates...
if (!is.matrix(denom)) denom <- as.matrix(denom)
mean.nom <- apply(nom,1,mean)
mean.denom <- apply(denom,1,mean)
#mean.nom <- ifelse(mean.nom==0,log.offset,mean.nom)
if (any(mean.nom==0)) mean.nom <- mean.nom + log.offset
#mean.denom <- ifelse(mean.denom==0,log.offset,mean.denom)
if (any(mean.denom==0)) mean.denom <- mean.denom + log.offset
fold.mat[,i-1] <- mean.nom/mean.denom
}
rownames(fold.mat) <- rownames(data.matrix)
colnames(fold.mat) <- paste(conds[1],"_vs_",conds[2:length(conds)],sep="")
return(fold.mat)
}
#' Calculates average expression for an MA plot
#'
#' Returns a matrix of average expressions (A in MA plot) based on the requested
#' contrast, the list of all samples and the data matrix which is produced by
#' the metaseqr workflow. For details on the \code{contrast}, \code{sample.list}
#' and \code{log.offset} parameters, see the main usage page of metaseqr.
#' This function is intended mostly for internal use but can also be used
#' independently.
#'
#' @param contrast the vector of requested statistical comparison contrasts.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param data.matrix a matrix of gene expression data whose column names are the
#' same as the sample names included in the sample list.
#' @param log.offset a number to be added to each element of data matrix in order
#' to avoid Infinity on log type data transformations.
#' @return A matrix of fold change ratios, treatment to control, as these are
#' parsed from contrast.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' a <- make.avg.expression("Control_vs_Treatment",list(Control=c("C1","C2"),
#' Treatment=c("T1","T2")),data.matrix)
#'}
make.avg.expression <- function(contrast,sample.list,data.matrix,log.offset=1) {
conds <- strsplit(contrast,"_vs_")[[1]]
a.mat <- matrix(0,nrow(data.matrix),length(conds)-1)
for (i in 2:length(conds)) { # First condition is ALWAYS reference
samples.nom <- sample.list[[conds[i]]]
samples.denom <- sample.list[[conds[1]]]
nom <- data.matrix[,match(samples.nom,colnames(data.matrix))]
denom <- data.matrix[,match(samples.denom,colnames(data.matrix))]
if (!is.matrix(nom)) nom <- as.matrix(nom) # Cover the case with no replicates...
if (!is.matrix(denom)) denom <- as.matrix(denom)
mean.nom <- apply(nom,1,mean)
mean.denom <- apply(denom,1,mean)
if (any(mean.nom==0)) mean.nom <- mean.nom + log.offset
if (any(mean.denom==0)) mean.denom <- mean.denom + log.offset
a.mat[,i-1] <- 0.5*(log2(mean.nom)+log2(mean.denom))
}
rownames(a.mat) <- rownames(data.matrix)
colnames(a.mat) <- paste(conds[1],"_vs_",conds[2:length(conds)],sep="")
return(a.mat)
}
#' HTML report helper
#'
#' Returns a character matrix with html formatted table cells. Essentially, it
#' converts the input data to text and places them in a <td></td> tag set.
#' Internal use.
#'
#' @param mat the data matrix (numeric or character)
#' @param type the type of data in the matrix (\code{"numeric"} or
#' \code{"character"}).
#' @param digits the number of digits on the right of the decimal points to pass
#' to \code{\link{formatC}}. It has meaning when \code{type="numeric"}.
#' @return A character matrix with html formatted cells.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' the.cells <- make.html.cells(data.matrix)
#'}
make.html.cells <- function(mat,type="numeric",digits=3) {
if (type=="numeric")
tmp <- format(mat,digits=digits)
#tmp <- formatC(mat,digits=digits,format="f")
else
tmp <- mat
if (!is.matrix(tmp)) tmp <- as.matrix(tmp)
tmp <- apply(tmp,c(1,2),function(x) paste("<td>",x,"</td>",sep=""))
return(tmp)
}
#' HTML report helper
#'
#' Returns a character vector with html formatted rows. Essentially, it collapses
#' every row of a matrix to a single character and puts a <tr></tr> tag set around.
#' It is meant to be applied to the output of \code{\link{make.html.cells}}.
#' Internal use.
#'
#' @param mat the data matrix, usually the output of \code{\link{make.html.cells}}
#' function.
#' @return A character vector with html formatted rows of a matrix.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' the.cells <- make.html.cells(data.matrix)
#' the.rows <- make.html.rows(the.cells)
#'}
make.html.rows <- function(mat) {
tmp <- apply(mat,1,paste,collapse="")
tmp <- paste("<tr>",tmp,"</tr>",sep="")
return(tmp)
}
#' HTML report helper
#'
#' Returns a character vector with an html formatted table head row. Essentially,
#' it collapses the input row to a single character and puts a <th></th> tag set
#' around. It is meant to be applied to the output of \code{\link{make.html.cells}}.
#' Internal use.
#'
#' @param h the colnames of a matrix or data frame, usually as output of
#' \code{\link{make.html.cells}} function.
#' @return A character vector with html formatted header of a matrix.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' the.cells <- make.html.cells(data.matrix)
#' the.header <- make.html.header(the.cells[1,])
#'}
make.html.header <- function(h) {
tmp <- paste("<th>",h,"</th>",sep="")
tmp <- paste(tmp,collapse="")
tmp <- paste("<tr>",tmp,"</tr>",sep="")
return(tmp)
}
#' HTML report helper
#'
#' Returns a character vector with an html formatted table. Essentially, it
#' collapses the input rows to a single character and puts a <tbody></tbody>
#' tag set around. It is meant to be applied to the output of
#' \code{\link{make.html.rows}}. Internal use.
#'
#' @param mat the character vector produced by \code{\link{make.html.rows}}.
#' @return A character vector with the body of mat formatted in html.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' the.cells <- make.html.cells(data.matrix)
#' the.header <- make.html.header(the.cells[1,])
#' the.rows <- make.html.rows(the.cells)
#' the.body <- make.html.body(the.rows)
#'}
make.html.body <- function(mat) {
tmp <- paste(mat,collapse="")
return(tmp)
}
#' HTML report helper
#'
#' Returns a character vector with a fully html formatted table. Essentially, it
#' binds the outputs of \code{\link{make.html.cells}}, \code{\link{make.html.rows}},
#' \code{\link{make.html.header}} and \code{\link{make.html.body}} to the final
#' table and optionally assigns an id attribute. The above functions are meant to
#' format a data table so as it can be rendered by external tools such as
#' DataTables.js during a report creation. It is meant for internal use.
#'
#' @param b the table body as produced by \code{\link{make.html.body}}.
#' @param h the table header as produced by \code{\link{make.html.header}}.
#' @param id the table id attribute.
#' @return A fully formatted html table.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' the.cells <- make.html.cells(data.matrix)
#' the.header <- make.html.header(the.cells[1,])
#' the.rows <- make.html.rows(the.cells)
#' the.body <- make.html.body(the.rows)
#' the.table <- make.html.table(the.body,the.header,id="my_table")
#'}
make.html.table <- function(b,h=NULL,id=NULL) {
if (!is.null(id))
html <- paste("<table id=\"",id,"\" class=\"datatable\">",sep="")
else
html <- "<table class=\"datatable\">"
if (!is.null(h))
html <- paste(html,"<thead>",h,"</thead>",sep="")
html <- paste(html,"<tbody>",b,"</tbody></table>",sep="")
return(html)
}
#' Calculates several transformation of counts
#'
#' Returns a list of transformed (normalized) counts, based on the input count
#' matrix data.matrix. The data transformations are passed from the
#' \code{export.scale} parameter and the output list is named accordingly. This
#' function is intended mostly for internal use but can also be used independently.
#'
#' @param data.matrix the raw or normalized counts matrix. Each column represents
#' one input sample.
#' @param export.scale a character vector containing one of the supported data
#' transformations (\code{"natural"}, \code{"log2"}, \code{"log10"},\code{"vst"}).
#' See also the main help page of metaseqr.
#' @param scf a scaling factor for the reads of each gene, for example the sum of
#' exon lengths or the gene length. Devided by each read count when
#' \code{export.scale="rpgm"}. It provides an RPKM-like measure but not the
#' actual RPKM as this normalization is not supported.
#' @param log.offset a number to be added to each element of data.matrix in order
#' to avoid Infinity on log type data transformations.
#' @return A named list whose names are the elements in export.scale. Each list
#' member is the respective transformed data matrix.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' tr <- make.transformation(data.matrix,c("log2","vst"))
#' head(tr$vst)
#'}
make.transformation <- function(data.matrix,export.scale,
scf=NULL,log.offset=1) {
mat <- vector("list",length(export.scale))
names(mat) <- export.scale
if (!is.matrix(data.matrix)) data.matrix <- as.matrix(data.matrix)
if (is.null(scf)) scf <- rep(1,nrow(data.matrix))
for (scl in export.scale) {
switch(scl,
natural = {
mat[[scl]] <- data.matrix
},
log2 = {
mat[[scl]] <- nat2log(data.matrix,base=2,off=log.offset)
},
log10 = {
mat[[scl]] <- nat2log(data.matrix,base=10,off=log.offset)
},
vst = {
fit <- vsn2(data.matrix,verbose=FALSE)
mat[[scl]] <- predict(fit,newdata=data.matrix)
},
rpgm = {
mat[[scl]] <- data.matrix
for (i in 1:ncol(data.matrix))
mat[[scl]][,i] <- data.matrix[,i]/scf
}
)
}
return(mat)
}
#' Calculates several statistices on read counts
#'
#' Returns a matrix of statistics calculated for a set of given samples. Internal
#' use.
#'
#' @param samples a set of samples from the dataset under processing. They should
#' match sample names from \code{sample.list}. See also the main help page of
#' \code{\link{metaseqr}}.
#' @param data.list a list containing natural or transformed data, typically an
#' output from \code{\link{make.transformation}}.
#' @param stat the statistics to calculate. Can be one or more of \code{"mean"},
#' \code{"median"}, \code{"sd"}, \code{"mad"}, \code{"cv"}, \code{"rcv"}. See also
#' the main help page of \code{\link{metaseqr}}.
#' @param export.scale the output transformations used as input also to
#' \code{\link{make.transformation}}.
#' @return A matrix of statistics calculated based on the input sample names. The
#' different data transformnations are appended columnwise.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' tr <- make.transformation(data.matrix,c("log2","vst"))
#' st <- make.stat(c("C1","C2"),tr,c("mean","sd"),c("log2","vst"))
#'}
make.stat <- function(samples,data.list,stat,export.scale) {
stat.result <- vector("list",length(export.scale))
names(stat.result) <- export.scale
for (scl in export.scale) {
stat.data <- data.list[[scl]][,match(samples,
colnames(data.list[[scl]]))]
if (!is.matrix(stat.data)) stat.data <- as.matrix(stat.data)
switch(stat,
mean = {
stat.result[[scl]] <- apply(stat.data,1,function(x,s) {
if (s=="natural")
return(round(mean(x)))
else
return(mean(x))
},scl)
},
median = {
stat.result[[scl]] <- apply(stat.data,1,function(x,s) {
if (s=="natural")
return(round(median(x)))
else
return(median(x))
},scl)
},
sd = {
stat.result[[scl]] <- apply(stat.data,1,function(x,s) {
if (s=="natural")
return(ceiling(sd(x)))
else
return(sd(x))
},scl)
},
mad = {
stat.result[[scl]] <- apply(stat.data,1,function(x,s) {
if (s=="natural")
return(ceiling(mad(x)))
else
return(mad(x))
},scl)
},
cv = {
stat.result[[scl]] <- apply(stat.data,1,function(x,s) {
if (s=="natural")
return(ceiling(sd(x))/round(mean(x)))
else
return(sd(x)/mean(x))
},scl)
},
rcv = {
stat.result[[scl]] <- apply(stat.data,1,function(x,s) {
if (s=="natural")
return(ceiling(mad(x))/round(median(x)))
else
return(mad(x)/median(x))
},scl)
}
)
}
return(do.call("cbind",stat.result))
}
#' Results output build helper
#'
#' Returns a list of matrices based on the export scales that have been chosen
#' from the main function and a subset of samples based on the sample names
#' provided in the \code{sample.list} argument of the main \code{\link{metaseqr}}
#' function. Internal use.
#'
#' @param samples a set of samples from the dataset under processing. They should
#' match sample names from \code{sample.list}. See also the main help page of
#' \code{\link{metaseqr}}.
#' @param data.list a list containing natural or transformed data, typically an
#' output from \code{\link{make.transformation}}.
#' @param export.scale the output transformations used as input also to
#' \code{\link{make.transformation}}.
#' @return A named list whose names are the elements in \code{export.scale}.
#' Each list member is the respective sample subest data matrix.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data.matrix <- round(1000*matrix(runif(400),100,4))
#' rownames(data.matrix) <- paste("gene_",1:100,sep="")
#' colnames(data.matrix) <- c("C1","C2","T1","T2")
#' tr <- make.transformation(data.matrix,c("log2","vst"))
#' mm <- make.matrix(c("C1","T1"),tr,"log2")
#' head(tr$vst)
#'}
make.matrix <- function(samples,data.list,export.scale="natural") {
mat <- vector("list",length(export.scale))
names(mat) <- export.scale
for (scl in export.scale) {
mat.data <- data.list[[scl]][,match(samples,
colnames(data.list[[scl]]))]
if (!is.matrix(mat.data)) {
mat.data <- as.matrix(mat.data)
colnames(mat.data) <- samples
}
mat[[scl]] <- mat.data
}
return(do.call("cbind",mat))
}
#' Create contrast lists from contrast vectors
#'
#' Returns a list, properly structured to be used within the \code{stat.*}
#' functions of the metaseqr package. See the main documentation for the structure
#' of this list and the example below. This function is mostly for internal use,
#' as the \code{stat.*} functions can be supplied directly with the contrasts
#' vector which is one of the main \code{\link{metaseqr}} arguments.
#'
#' @param contrast a vector of contrasts in the form "ConditionA_vs_ConditionB"
#' or "ConditionA_
#' vs_ConditionB_vs_ConditionC_vs_...".
#' In case of Control vs Treatment designs, the Control condition should ALWAYS
#' be the first.
#' @param sample.list the list of samples in the experiment. See also the main
#' help page of \code{\link{metaseqr}}.
#' @return A named list whose names are the contrasts and its members are named
#' vectors, where the names are the sample names and the
#' actual vector members are the condition names. See the example.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' sample.list <- list(Control=c("C1","C2"),TreatmentA=c("TA1","TA2"),
#' TreatmentB=c("TB1","TB2"))
#' contrast <- c("Control_vs_TreatmentA","Control_vs_TreatmentA_vs_TreatmentB"),
#' cl <- make.contrast.list(contrast,sample.list)
#' cl
#'}
make.contrast.list <- function(contrast,sample.list) {
# Construction
contrast.list <- vector("list",length(contrast))
names(contrast.list) <- contrast
# First break the contrast vector
cnts <- strsplit(contrast,"_vs_")
names(cnts) <- names(contrast.list)
# Create list members
for (n in names(contrast.list)) {
contrast.list[[n]] <- vector("list",length(cnts[[n]]))
for (i in 1:length(cnts[[n]])) {
contrast.list[[n]][[i]] <- rep(cnts[[n]][i],
length(sample.list[[cnts[[n]][i]]]))
names(contrast.list[[n]][[i]]) <- sample.list[[cnts[[n]][[i]]]]
}
}
return(contrast.list)
}
#' Creates sample list from file
#'
#' Create the main sample list from an external file.
#'
#' @param input a tab-delimited file structured as follows: the first line of the
#' external tab delimited file should contain column names (names are not important).
#' The first column MUST contain UNIQUE sample names and the second column MUST
#' contain the biological condition where each of the samples in the first column
#' should belong to.
#' @param type one of \code{"simple"} or \code{"targets"} to indicate if the input
#' is a simple two column text file or the targets file used to launch the main
#' analysis pipeline.
#' @return A named list whose names are the conditions of the experiments and its
#' members are the samples belonging to each condition.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' targets <- data.frame(sample=c("C1","C2","T1","T2"),
#' condition=c("Control","Control","Treatment","Treatment"))
#' write.table(targets,file="targets.txt",sep="\t",row.names=FALSE,quote="")
#' sample.list <- make.sample.list("targets.txt")
#'}
make.sample.list <- function(input,type=c("simple","targets")) {
if (missing(input) || !file.exists(input))
stopwrap("File to make sample list from should be a valid existing ",
"text file!")
type <- tolower(type[1])
check.text.args("type",type,c("simple","targets"),multiarg=FALSE)
tab <- read.delim(input)
samples <- as.character(tab[,1])
if (type=="simple") {
ii <- 2
conditions <- unique(as.character(tab[,ii]))
}
else if (type=="targets") {
ii <- 3
conditions <- unique(as.character(tab[,ii]))
}
if (length(samples) != length(unique(samples)))
stopwrap("Sample names must be unique for each sample!")
sample.list <- vector("list",length(conditions))
names(sample.list) <- conditions
for (n in conditions)
sample.list[[n]] <- samples[which(as.character(tab[,ii])==n)]
return(sample.list)
}
#' Project path constructor
#'
#' Create the main metaseqr project path. Internal use only.
#'
#' @param path The desired project path. Can be NULL for auto-generation.
#' @param f The input counts table file.
#' @return A list with project path elements.
#' @author <NAME>
make.project.path <- function(path,f=NULL) {
if (is.na(path) || is.null(path)) {
if (!is.data.frame(f) && !is.null(f) && !is.list(f) && file.exists(f)) {
if (length(grep(".RData$",f))>0)
main.path <- file.path(getwd(),paste("metaseqr_result_",
format(Sys.time(),format="%Y%m%d%H%M%S"),sep=""))
else
main.path <- file.path(dirname(f),paste("metaseqr_result_",
format(Sys.time(),format="%Y%m%d%H%M%S"),sep=""))
}
else
main.path <- file.path(getwd(),paste("metaseqr_result_",
format(Sys.time(),format="%Y%m%d%H%M%S"),sep=""))
project.path <- make.path.struct(main.path)
}
else {
success <- tryCatch(
if (!file.exists(path)) dir.create(path,recursive=TRUE) else TRUE,
error=function(e) {
disp("Cannot create ",path,"! Is it a valid system path? Is ",
"there a write permissions problem? Reverting to ",
"automatic creation...")
return(FALSE)
},
finally=""
)
if (success)
project.path <- make.path.struct(path)
else
project.path <- make.project.path(NA,f)
}
return(project.path)
}
#' Project path constructor helper
#'
#' Helper for \code{make.project.path}. Internal use only.
#'
#' @param main.path The desired project path.
#' @return A named list whose names are the conditions of the experiments and its
#' members are the samples belonging to each condition.
#' @author <NAME>
make.path.struct <- function(main.path) {
project.path <- list(
main=main.path,
media=file.path(main.path,"media"),
data=file.path(main.path,"data"),
logs=file.path(main.path,"logs"),
lists=file.path(main.path,"lists"),
plots=file.path(main.path,"plots"),
qc=file.path(main.path,"plots","qc"),
normalization=file.path(main.path,"plots","normalization"),
statistics=file.path(main.path,"plots","statistics")
)
for (p in names(project.path))
if (!file.exists(project.path[[p]]))
dir.create(project.path[[p]],recursive=TRUE)
return(project.path)
}
#' Intitialize output list
#'
#' Initializes metaseqr R output. Internal use only.
#'
#' @param con The contrasts.
#' @return An empty named list.
#' @author <NAME>
make.export.list <- function(con) {
f <- vector("list",length(con))
names(f) <- con
return(f)
}
#' Optimize rectangular grid plots
#'
#' Returns a vector for an optimized m x m plot grid to be used with e.g.
#' \code{par(mfrow)}. m x m is as close as possible to the input n. Of course,
#' there will be empty grid positions if n < m x m.
#'
#' @param n An integer, denoting the total number of plots to be created.
#' @return A 2-element vector with the dimensions of the grid.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' g1 <- make.grid(16) # Returns c(4,4)
#' g2 <- make.grid(11) # Returns c(4,3)
#'}
make.grid <- function(n) {
m <- 0
while (n > m*m)
m <- m+1
if (n < m*m) {
k <- m-1
if (n > m*k)
k <- k+1
else {
while (n > m*k)
k=k-1
}
}
else
k <- m
return(c(m,k))
}
#' Initializer of report messages
#'
#' Initializes metaseqr report tmeplate messages output. Internal use only.
#'
#' @param lang The language of the report. For now, only english (\code{"en"})
#' is supported.
#' @return An named list with messages for each input option.
#' @author <NAME>
make.report.messages <- function(lang) {
switch(lang,
en = {
messages <- list(
org=list(
hg18=paste("human (<em>Homo sapiens</em>),",
"genome version alias hg18"),
hg19=paste("human (<em>Homo sapiens</em>),",
"genome version alias hg19"),
mm9=paste("mouse (<em>Mus musculus</em>),",
"genome version alias mm9"),
mm10=paste("mouse (<em>Mus musculus</em>),",
"genome version alias mm10"),
rn5=paste("rat (<em>Rattus norvegicus</em>),",
"genome version alias rn5"),
dm3=paste("fruitfly (<em>Drosophila melanogaster</em>),",
"genome version alias dm3"),
danrer7=paste("zebrafish (<em>Danio rerio</em>),",
"genome version alias danRer7"),
pantro5=paste("chimpanzee (<em>Pan troglodytes</em>),",
"genome version alias panTro5"),
susscr3=paste("pig (<em>Sus scrofa</em>),",
"genome version alias susScr3"),
tair10=paste("arabidopsis (<em>Arabidobsis thaliana</em>)",
",","genome version alias TAIR10"),
bmori2=paste("silkworm (<em>Bombyx mori</em>)",
",","genome version alias bmori2")
),
refdb=list(
ensembl="Ensembl genomes",
ucsc="UCSC genomes database",
refseq="RefSeq database"
),
whenfilter=list(
prenorm="before normalization",
postnorm="after normalization"
),
norm=list(
edaseq="EDASeq",
deseq="DESeq",
edger="edgeR",
noiseq="NOISeq",
nbpseq="NBPSeq",
each="the same as the corresponding statistical test"
),
stat=list(
deseq="DESeq",
edger="edgeR",
noiseq="NOISeq",
bayseq="baySeq",
limma="limma",
nbpseq="NBPSeq"
),
meta=list(
intersection="intersection of individual results",
union="union of individual results",
fisher="Fisher's method",
fperm="Fisher's method with permutations",
dperm.min=paste("samples permutation based method with",
"minimum p-values"),
dperm.max=paste("samples permutation based method with",
"maximum p-values"),
dperm.weight=paste("samples permutation based method with",
"weighted p-values"),
minp="minimum p-value across results",
maxp="maximum p-value across results",
weight="PANDORA weighted p-value across results",
pandora="PANDORA weighted p-value across results",
simes="Simes correction and combination method",
whitlock=paste("Whitlock's Z-transformation method",
"(Bioconductor package survcomp)"),
none=paste("no meta-analysis, reported p-values from the",
"first supplied statistical algorithm")
),
adjust=list(
holm="Holm FWER",
hochberg="Hochberg DFR",
hommel="Hommel FWER",
bonferroni="Bonferroni FWER",
bh="Benjamini-Hochberg FDR",
by="Benjamini-Yekutiely FDR",
fdr="Benjamini-Hochberg FDR",
none="no multiple test correction",
qvalue="Storey-Tibshirani FDR"
),
plots=list(
mds="multidimensional scaling",
biodetection="biotype detection",
countsbio="biotype counts",
saturation="sample and biotype saturation",
rnacomp="RNA composition",
boxplot="boxplots",
gcbias="GC-content bias",
lengthbias="transcript length bias",
meandiff="mean-difference plot",
meanvar="mean-variance plot",
deheatmap="DEG heatmap",
volcano="volcano plot",
biodist="DEG biotype detection",
filtered="filtered biotypes",
correl="correlation heatmap and correlogram",
pairwise="pairwise scatterplots between samples",
venn="Venn diagrams"
),
export=list(
annotation="Annotation",
p.value="p-value",
adj.p.value="Adjusted p-value (FDR)",
fold.change="Fold change",
stats="Statistics",
counts="Read counts",
natural="Natural scale",
log2="log2 scale",
log10="log10 scale",
vst="Variance stabilization transformation",
rpgm="Reads per Gene Model",
raw="Raw values",
normalized="Normalized values",
mean="Mean",
median="Median",
sd="Standard deviation",
mad="Median Absolute Deviation (MAD)",
cv="Coefficient of Variation",
rcv="Robust Coefficient of Variation"
),
preset=list(
all.basic=paste("use all genes and export all genes and",
"basic annotation and statistics elements"),
all.normal=paste("use all genes and export all genes and",
"normal annotation and statistics elements"),
all.full=paste("use all genes and export all genes and all",
"available annotation and statistics elements"),
medium.basic=paste("apply a medium set of filters and",
"export statistically significant genes and basic",
"annotation and statistics elements"),
medium.normal=paste("apply a medium set of filters and",
"export statistically significant genes and normal",
"annotation and statistics elements"),
medium.full=paste("apply a medium set of filters and",
"export statistically significant genes and all",
"available annotation and statistics elements"),
strict.basic=paste("apply a strict set of filters and",
"export statistically significant genes and basic",
"annotation and statistics elements"),
strict.normal=paste("apply a medium set of filters and",
"export statistically significant genes and normal",
"annotation and statistics elements"),
strict.full=paste("apply a medium set of filters and",
"export statistically significant genes and all",
"available annotation and statistics elements")
),
explain=list(
mds=paste(
"The Multi-Dimensional Scaling (MDS) plots comprise a means",
"of visualizing the level of similarity of individual cases",
"of a dataset. It is similar to Principal Component Analysis",
"(PCA), but instead of using the covariance matrix to find",
"similarities among cases, MDS uses absolute distance metrics",
"such as the classical Euclidean distance. Because of the",
"relative linear relations among sequencing samples, it",
"provides a more realistic clustering among samples. MDS",
"serves quality control and it can be interpreted as follows:",
"when the distance among samples of the same biological",
"condition in the MDS space is small, this is an indication",
"of high correlation and reproducibility among them. When",
"this distance is larger or heterogeneous (e.g. the 3rd",
"sample of a triplicate set is further from the other 2),",
"this constitutes an indication of low correlation and",
"reproducibility among samples. It can help exclude poor",
"samples from further analysis.",collapse=" "
),
biodetection=paste(
"The biotype detection bar diagrams are a set of quality",
"control charts that show the percentage of each biotype",
"in the genome (i.e. in the whole set of features provided,",
"for example, protein coding genes, non coding RNAs or",
"pseudogenes) in grey bars, which proportion has been",
"detected in a sample before normalization and after a",
"basic filtering by removing features with zero counts in",
"red lined bars, and the percentage of each biotype within",
"the sample in solid red bars. The difference between grey",
"bars and solid red bars is that the grey bars show the",
"percentage of a feature in the genome while the solid red",
"bars show the percentage in the sample. Thus, the solid",
"red bars may be sometimes higher than the grey bars because",
"certain features (e.g. protein coding genes) may be",
"detected within a sample with a higher proportion",
"relatively to their presence in the genome, as compared",
"with other features. For example, while the percentage",
"of protein coding genes in the whole genome is already",
"higher than other biotypes, this percentage is expected",
"to be even higher in an RNA-Seq experiment where one",
"expects protein-coding genes to exhibit greater abundance.",
"The vertical green line separates the most abundant",
"biotypes (on the left-hand side, corresponding to the",
"left axis scale) from the rest (on the right-hand side,",
"corresponding to the right axis scale). Otherwise, the",
"lower abundance biotypes would be indistinguishable.",
"Unexpected outcomes in this quality control chart (e.g.",
"very low detection of protein coding genes) would signify",
"possible low quality of a sample.",collapse=" "
),
countsbio=paste(
"The biotype detection counts boxplots are a set of quality",
"control charts that depict both the biological classification",
"for the detected features and the actual distribution of",
"the read counts for each biological type. The boxplot",
"comprises a means of summarizing the read counts distribution",
"of a sample in the form of a bar with extending lines,",
"as commonly used way of graphically presenting groups of",
"numerical data. A boxplot also indicates which observations,",
"if any, might be considered outliers and is able to visually",
"show different types of populations, without making any",
"assumptions of the underlying statistical distribution.",
"The spacing between the different parts of the box help",
"indicate variance, skewness and identify outliers. The",
"thick bar inside the colored box is the median of the",
"observations while the box extends over the Interquartile",
"Range of the observations. The whiskers extend up (down)",
"to +/-1.5xIQR. Unexpected outcomes (e.g. protein coding",
"read count distribution similar to pseudogene read count",
"distribution) indicates poor sample quality.",collapse=" "
),
saturation=paste(
"The read and biotype saturation plots are a set of quality",
"control charts that depict the read count saturation",
"levels at several sequencing depths. Thus, they comprise",
"a means of assessing whether the sequencing depth of an",
"RNA-Seq experiment is sufficient in order to detect the",
"biological features under investigation. These quality",
"control charts are separated in two subgroups: the first",
"subgroup (read saturation per biotype for all samples)",
"is a set of plots, one for each biological feature (e.g.",
"protein coding, pseudogene, lincRNA, etc.), that depict",
"the number of detected features in different sequencing",
"depths and for all samples in the same plot. The second",
"subgroup (read saturation per sample for all biotypes)",
"is a set of plots similar to the above, but with,",
"there is one pair of plots with two panels for each sample,",
"presenting all biological features. The left panel depicts",
"the saturation levels for the less abundatnt features,",
"while the right panel, the saturation for the more abundant",
"features, as placing them all together would make the",
"less abundant features indistinguishable. All the saturation",
"plots should be interpreted as follows: if the read counts",
"for a biotype tend to be saturated, the respective curve",
"should tend to reach a plateau at higher depths. Otherwise,",
"more sequencing is needed for the specific biotype.",
collapse=" "
),
readnoise=paste(
"The read noise plots depict the percentage of biological",
"features detected when subsampling the total number of",
"reads. Very steep curves in read noise plots indicate",
"that although the sequencing depth reaches its maximum,",
"a relatively small percentage of total features is detected,",
"indicating that the level of background noise is relatively",
"high. Less steep RNA composition curves, indicate less noise.",
"When a sample's curve deviate from the rest, it could",
"indicate lower or higher quality, depending on the curves",
"of the rest of the samples.",collapse=" "
),
correl=paste(
"The sample correlation plots depict the accordance among",
"the RNA-Seq samples, as this is manifested through the",
"read counts table used with the metaseqr pipeline, with",
"two representations that both use the correlation matrix",
"(a matrix which depicts all the pairwise correlations",
"between each pair of samples) of the read counts matrix.",
"The first is a correlation clustered heatmap which",
"depicts the correlations among samples as color-scaled",
"image and the hierarchical clustering tree depicts the",
"grouping of the samples according to their correlation.",
"Samples from the same group that are not clustered together",
"provides an indication that there might be a quality",
"problem with the dataset. The second is a 'correlogram'",
"plot, where again the samples are hierarchically clustered",
"and grouped but this time correlations are presented as",
"ellipses inside each cell. Each cell represents a pairwise",
"comparison and each correlation coefficient is represented",
"by an ellipse whose 'diameter', direction and color",
"depict the accordance for that pair of samples. Highly",
"correlated samples are depicted as ellipses with narrow",
"diameter, while poorly correlated samples are depicted",
"as ellipses with wide diameters. Also, highly correlated",
"samples are depicted as ellipses with a left-to-right",
"upwards direction while poorly correlated samples are",
"depicted as ellipses with a right-to-left upwards direction.",
collapse=" "
),
pairwise=paste(
"The pairwise comparison plots are split in three parts:",
"the upper diagonal consists of simple scatterplots for",
"all pairwise sample comparisons, together with their",
"Pearson correlation coefficient. It is a simple measure",
"of between sample correlation using all the available",
"data points instead of only the correlation matrix. The",
"lower diagonal consists of mean-difference plots for all",
"pairwise sample comparisons. A mean-difference plot (or",
"a Bland-Altman plots) is a method of data plotting used",
"in analyzing the agreement between two different",
"assays/variables. In this graphical method the differences",
"(or alternatively the ratios) between the two variables",
"are plotted against the averages of the two. Such a plot",
"is useful, for example, for analyzing data with strong",
"correlation between x and y axes, when the (x,y) dots on",
"the plot are close to the diagonal x=y. In this case, the",
"value of the transformed variable X is about the same as",
"x and y and the variable Y shows the difference between",
"x and y. In both represantations, irregular shapes of the",
"red smoother lines are an indication of poor correlation",
"between samples or of other systematic bias sources,",
"which is usually corrected through data normalization.",
collapse=" "
),
rnacomp=paste(
"The RNA composition plots depict the differences in the",
"distributions of reads in the same biological features",
"across samples. The following is taken from the NOISeq",
"vignette: <em>'...when two samples have different RNA",
"composition, the distribution of sequencing reads across",
"the features is different in such a way that although",
"a feature had the same number of read counts in both",
"samples, it would not mean that it was equally expressed",
"in both... To check if this bias is present in the data,",
"the RNA composition plot and the correponding diagnostic",
"test can be used. In this case, each sample s is compared",
"to the reference sample r (which can be arbitrarily",
"chosen). To do that, M values are computed as",
"log2(counts_sample = counts_reference). If no bias is",
"present, it should be expected that the median of M",
"values for each comparison is 0. Otherwise, it would be",
"indicating that expression levels in one of the samples",
"tend to be higher than in the other, and this could lead",
"to false discoveries when computing differencial expression.",
"Confidence intervals for the M median are also computed by",
"bootstrapping. If value 0 does not fall inside the interval,",
"it means that the deviation of the sample with regard",
"to the reference sample is statistically significant.",
"Therefore, a normalization procedure is required.'</em>",
collapse=" "
),
boxplot=paste(
"The boxplot comprises a means of summarizing the read",
"counts distribution of a sample in the form of a bar",
"with extending lines, as a commonly used way of",
"graphically presenting groups of numerical data. A",
"boxplot also indicates which observations, if any, might",
"be considered outliers and is able to visually show",
"different types of populations, without making any",
"assumptions about the underlying statistical distribution.",
"The spacings between the different parts of the box help",
"indicate variance, skewness and identify outliers. The",
"thick bar inside the colored box is the median of the",
"observations while the box extends over the Interquartile",
"Range of the observations. The whiskers extend up (down)",
"to +/-1.5xIQR. Boxplots at similar levels indicate good",
"quality of the normalization. If boxplots remain at",
"different levels after normalization, maybe another",
"normalization algorithm may have to be examined.",
"The un-normalized boxplots show the need for data",
"normalization in order for the data from different",
"samples to follow the same underlying distribution and",
"statistical testing becoming possible.",collapse=" "
),
gcbias=paste(
"The GC-content bias plot is a quality control chart that",
"shows the possible dependence of the read counts (in log2",
"scale) under a gene to the GC content percentage of that",
"gene. In order for the statistical tests to be able to",
"detect statistical significance which occurs due to real",
"biological effects and not by other systematic biases",
"present in the data (e.g. a possible GC-content bias),",
"the latter should be accounted for by the applied",
"normalization algorithm. Although the tests are performed",
"for each gene across biological conditions one could assume",
"that the GC content does not represent a bias, as it is the",
"same for the tested gene across samples and conditions.",
"However, Risso et al. (2011) showed that the GC-content",
"could have an impact in the statistical testing procedure.",
"The GC-content bias plot depicts the dependence of the",
"read counts to the GC content before and after normalization.",
"The smoothing lines for each sample, should be as 'straight'",
"as possible after normalization. In addition, if the",
"smoothing lines differ significantly between biological",
"conditions, this would constitute a possible quality warning.",
collapse=" "
),
lengthbias=paste(
"The gene/transcript length bias plot is a quality control",
"chart that shows the possible dependence of the read counts",
"(in log2 scale) under a gene to the length that gene (whole",
"gene or sum of exons depending on the analysis). In order",
"for the statistical tests to be able to detect statistical",
"significance which occurs due to real biological effects",
"and not by other systematic biases present in the data",
"(e.g. a possible length bias), the latter should be accounted",
"for by the applied normalization algorithm. Although the",
"tests are performed for each gene across bioogical conditions,",
"one could assume that the gene length does not represent",
"a bias as it's the same for the tested gene across samples",
"and conditions. However, it has been shown in several",
"studies that the gene length could have an impact on the",
"statistical testing procedure. The length bias plot",
"depicts the dependence of the read counts to the",
"gene/transcript length before and after normalization.",
"The smoothing lines for each sample, should be as 'straight'",
"as possible after normalization. In addition, if the",
"smoothing lines differ significantly among biological",
"conditions, this would constitute a possible quality warning.",
collapse=" "
),
meandiff=paste(
"A mean-difference plot (or a Bland-Altman plot) is a",
"method of data plotting used in analyzing the agreement",
"between two different assays/variables. In this graphical",
"method the differences (or alternatively the ratios)",
"between the two variables are plotted against the averages",
"of the two. Such a plot is useful, for example, for analyzing",
"data with strong correlation between x and y axes, when",
"the (x,y) dots on the plot are close to the diagonal x=y.",
"In this case, the value of the transformed variable X is",
"about the same as x and y and the variable Y shows the",
"difference between x and y. When the data cloud in a mean",
"difference plot is centered around the horizontal zero line,",
"this is an indication of good data quality and good",
"normalization results. On the other hand, when the data",
"cloud deviates from the center line or has a 'banana'",
"shape, this constitutes an indication of systematic biases",
"present in the data and that either the chosen normalization",
"algorithm has not worked well, or that data are not",
"normalized. The smoothing curve that traverses the data",
"(red curve) summarizes the above trends.",collapse=" "
),
meanvar=paste(
"The mean-variance plot comprises a graphical means of",
"displaying a possible relationship between the means of",
"gene expression (counts) values and their variances",
"across replicates of the same biological condition. Thus",
"data can be inspected for possible overdispersion (greater",
"variability in a dataset than would be expected based on",
"a given simple statistical model). In such plots for",
"RNA-Seq data, overdispersion is usually manifested as",
"increasing variance with increasing gene expression",
"(counts) and it is summarized through a smoothing curve",
"(red curve). The following is taken from the EDASeq package",
"vignette: '<em>...although the Poisson distribution",
"is a natural and simple way to model count data, it has",
"the limitation of assuming equality of the mean and",
"variance. For this reason, the negative binomial",
"distribution has been proposed as an alternative when the",
"data show over-dispersion...'</em> If overdispersion is",
"not present, the data cloud is expected to be evenly",
"scattered around the smoothing curve.",collapse=" "
),
deheatmap=paste(
"The Differentially Expressed Genes (DEGs) heatmaps depict",
"how well samples from different conditions cluster",
"together according to their expression values after",
"normalization and statistical testing, for each requested",
"statistical contrast. If samples from the same biological",
"condition do not cluster together, this would constitute",
"a warning sign regarding the quality of the samples. In",
"addition, DEG heatmaps provide an initial view of",
"possible clusters of co-expressed genes.",collapse=" "
),
volcano=paste(
"A volcano plot is a scatterplot that is often used when",
"analyzing high-throughput -omics data (e.g. microarray",
"data, RNA-Seq data) to give an overview of interesting",
"genes. The log2 fold change is plotted on the x-axis and",
"the negative log10 p-value is plotted on the y-axis. A",
"volcano plot combines the results of a statistical test",
"(aka, p-values) with the magnitude of the change enabling",
"quick visual identification of those genes that display",
"large-magnitude changes that are also statistically",
"significant. The horizontal dashed line sets the threshold",
"for statistical significance, while the vertical dashed",
"lines set the thresholds for biological significance. It",
"should be noted that the volcano plots become harder to",
"interpret when using more than one statistical algorithm",
"and performing meta-analysis. This happens because the genes",
"that have stronger evidence of being differentially",
"expressed obtain lower p-values while the rest either",
"remain at similar levels or obtain higher p-values.",
"The result is a 'warped' volcano plot, with two",
"main data clouds: one in the upper part of the plot, and",
"one in the lower part of the plot. You can always zoom in",
"when using interacting mode (the default).",collapse=" "
),
biodist=paste(
"The chromosome and biotype distributions bar diagram for",
"Differentially Expressed Genes (DEGs) is split in two",
"panels: i) on the left panel DEGs are distributed per",
"chromosome and the percentage of each chromosome in the",
"genome is presented in grey bars, the percentage of DEGs",
"in each chromosome is presented in red lined bars and the",
"percentage of certain chromosomes in the distribution of",
"DEGs is presented in solid red bars. ii) on the right panel,",
"DEGs are distributed per biotype and the percentage of",
"each biotype in the genome (i.e. in the whole set of",
"features provided, for example, protein coding genes, non",
"coding RNAs or pseudogenes) is presented in grey bars,",
"the percentage of DEGs in each biotype is presented in",
"blue lined bars and the percentage of each biotype in",
"DEGs is presented in solid blue lines. The vertical green",
"line separates the most abundant biotypes (on the left-hand",
"side, corresponding to the left axis scale), from the rest",
"(on the right-hand side, corresponding to the right axis",
"scale). Otherwise, the lower abundance, biotypes would be",
"indistinguishable.",collapse=" "
),
filtered=paste(
"The chromosome and biotype distribution of filtered genes",
"is a quality control chart with two rows and four panels:",
"on the left panel of the first row, the bar chart depicts",
"the numbers of filtered genes per chromosome (actual numbers",
"shown above the bars). On the right panel of the first row,",
"the bar chart depicts the numbers of filtered genes per",
"biotype (actual numbers shown above the bars). On the left",
"panel of the second row, the bar chart depicts the fraction",
"of the filtered genes to the total genes per chromosome",
"(actual percentages shown above the bars). On the right",
"panel of the second row, the bar chart depicts the fraction",
"of the filtered genes to the total genes per biotype",
"(actual percentages shown above the bars). This plot",
"should indicate possible quality problems when for example",
"the filtered genes for a specific chromosome (or the",
"fraction) is extremely higher than the rest. Generally,",
"the fractions per chromosome should be uniform and the",
"fractions per biotype should be proportional to the biotype",
"fraction relative to the genome.",collapse=" "
),
venn=paste(
"The Venn diagrams are an intuitive way of presenting",
"overlaps between lists, based on the overlap of basic",
"geometrical shapes. The numbers of overlapping genes per",
"statistical algorithm are shown in the different areas",
"of the Venn diagrams, one for each contrast.",collapse=" "
)
),
references=list(
filein=list(
sam=paste("<NAME>., <NAME>., <NAME>.,",
"<NAME>., <NAME>., <NAME>. (2010).",
"Repitools: an R package for the analysis of",
"enrichment-based epigenomic data. Bioinformatics",
"26(13), 1662-1663."),
bam=paste("<NAME>., <NAME>., <NAME>.,",
"<NAME>., <NAME>., <NAME>. (2010).",
"Repitools: an R package for the analysis of",
"enrichment-based epigenomic data. Bioinformatics",
"26(13), 1662-1663."),
bed=paste("<NAME>., <NAME>., <NAME>.",
"(2009). rtracklayer: an R package for interfacing",
"with genome browsers. Bioinformatics 25(14),",
"1841-1842.")
),
norm=list(
edaseq=paste("<NAME>., <NAME>., <NAME>.,",
"and <NAME>. (2011). GC-content normalization",
"for RNA-Seq data. BMC Bioinformatics 12, 480."),
deseq=paste("<NAME>., and <NAME>. (2010).",
"Differential expression analysis for sequence",
"count data. Genome Biol 11, R106."),
edger=paste("<NAME>., <NAME>., and",
"<NAME>. (2010). edgeR: a Bioconductor package",
"for differential expression analysis of digital",
"gene expression data. Bioinformatics 26,",
"139-140."),
noiseq=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
nbpseq=paste("<NAME>, <NAME>., <NAME>., and",
"<NAME>. (2011). The NBP Negative Binomial",
"Model for Assessing Differential Gene Expression",
"from RNA-Seq. Statistical Applications in",
"Genetics and Molecular Biology 10(1), 1-28."),
none=NULL
),
stat=list(
deseq=paste("<NAME>., and <NAME>. (2010).",
"Differential expression analysis for sequence",
"count data. Genome Biol 11, R106."),
edger=paste("<NAME>., <NAME>., and",
"<NAME>. (2010). edgeR: a Bioconductor package",
"for differential expression analysis of digital",
"gene expression data. Bioinformatics 26,",
"139-140."),
noiseq=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
limma=paste("<NAME>. (2005). Limma: linear models",
"for microarray data. In Bioinformatics and",
"Computational Biology Solutions using R and",
"Bioconductor, <NAME>., <NAME>., <NAME>., <NAME>., and",
"<NAME>., eds. (New York, Springer), pp. 397-420."),
bayseq=paste("<NAME>., and <NAME>.",
"(2010). baySeq: empirical Bayesian methods for",
"identifying differential expression in sequence",
"count data. BMC Bioinformatics 11, 422."),
nbpseq=paste("<NAME>., <NAME>., and",
"<NAME>. (2011). The NBP Negative Binomial",
"Model for Assessing Differential Gene Expression",
"from RNA-Seq. Statistical Applications in",
"Genetics and Molecular Biology 10(1), 1-28."),
ebseq=paste("<NAME>., <NAME>., <NAME>.,",
"<NAME>., <NAME>., <NAME>., Haag,",
"J.D., <NAME>., <NAME>., and",
"<NAME>. (2013). EBSeq: an empirical",
"Bayes hierarchical model for inference in",
"RNA-seq experiments. Bioinformatics 29, 1035-1043")
),
meta=list(
fisher=paste("<NAME>. (1932). Statistical",
"Methods for Research Workers (Edinburgh, Oliver",
"and Boyd)."),
fperm=paste("<NAME>. (1932). Statistical",
"Methods for Research Workers (Edinburgh, Oliver",
"and Boyd)."),
whitlock=c(
paste("<NAME>. (2005). Combining",
"probability from independent tests:",
"the weighted Z-method is superior to Fisher's",
"approach. J Evol Biol 18, 1368-1373."),
paste("<NAME>., <NAME>., Quackenbush,",
"J., and <NAME>. (2011). survcomp:",
"an R/Bioconductor package for performance",
"assessment and comparison of survival",
"models. Bioinformatics 27, 3206-3208.")
),
weight=paste("<NAME>., <NAME>., Wasserman,",
"L. (2006). False discovery control with p-value",
"weighting. Biometrika 93 (3): 509-524."),
simes=paste("<NAME>. (1986). An improved",
"Bonferroni procedure for multiple tests of",
"significance. Biometrika 73 (3): 751-754."),
none=NULL
),
multiple=list(
BH=paste("<NAME>., and <NAME>. (1995). ",
"Controlling the False Discovery Rate: A Practical",
"and Powerful Approach to Multiple Testing.",
"Journal of the Royal Statistical Society Series",
"B (Methodological) 57, 289-300."),
fdr=paste("<NAME>., and <NAME>. (1995). ",
"Controlling the False Discovery Rate: A Practical",
"and Powerful Approach to Multiple Testing.",
"Journal of the Royal Statistical Society Series",
"B (Methodological) 57, 289-300."),
BY=paste("<NAME>., and <NAME>. (2001). The",
"control of the false discovery rate in multiple",
"testing under dependency. Annals of Statistics",
"26, 1165-1188."),
bonferroni=paste("<NAME>. (1995). Multiple",
"hypothesis testing. Annual Review of",
"Psychology 46, 561-576."),
holm=paste("<NAME>. (1979). A simple sequentially",
"rejective multiple test procedure. Scandinavian",
"Journal of Statistics 6, 65-70."),
hommel=paste("<NAME>. (1988). A stagewise rejective",
"multiple test procedure based on a modified",
"Bonferroni test. Biometrika 75, 383-386."),
hochberg=paste("<NAME>. (1988). A sharper",
"Bonferroni procedure for multiple tests of",
"significance. Biometrika 75, 800-803."),
qvalue=paste("<NAME>., and <NAME>. (2003).",
"Statistical significance for genomewide studies.",
"Proc Natl Acad Sci U S A 100, 9440-9445.")
),
figure=list(
mds=paste("<NAME>., <NAME>., <NAME>.,",
"<NAME>., and <NAME>. (2012). htSeqTools:",
"high-throughput sequencing quality control,",
"processing and visualization in R. Bioinformatics",
"28, 589-590."),
biodetection=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
countsbio=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
saturation=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
readnoise=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
gcbias=paste("<NAME>., <NAME>., <NAME>.,",
"and <NAME>. (2011). GC-content normalization",
"for RNA-Seq data. BMC Bioinformatics 12, 480."),
lengthbias=paste("<NAME>., <NAME>., Sherlock,",
"G., and <NAME>. (2011). GC-content",
"normalization for RNA-Seq data.",
"BMC Bioinformatics 12, 480."),
meandiff=paste("<NAME>., <NAME>., <NAME>.,",
"and <NAME>. (2011). GC-content normalization",
"for RNA-Seq data. BMC Bioinformatics 12, 480."),
meanvar=paste("<NAME>., <NAME>., <NAME>.,",
"and <NAME>. (2011). GC-content normalization",
"for RNA-Seq data. BMC Bioinformatics 12, 480."),
rnacomp=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
biodist=paste("<NAME>., <NAME>.,",
"<NAME>., <NAME>., and <NAME>. (2011).",
"Differential expression in RNA-seq: a matter of",
"depth. Genome Res 21, 2213-2223."),
venn=paste("<NAME>., and <NAME>. (2011).",
"VennDiagram: a package for the generation of",
"highly-customizable Venn and Euler diagrams in R.",
"BMC Bioinformatics 12, 35."),
filtered=NULL
)
)
)
}
)
return(messages)
}
#' Interactive volcano plot helper
#'
#' Creates a list which contains the data series of a scatterplot, to be used for
#' serialization with highcharts JavaScript plotting.
#' framework. Internal use only.
#'
#' @param x The x coordinates (should be a named vector!).
#' @param y The y coordinates.
#' @param a Alternative names for each point.
#' @return A list that is later serialized to JSON.
#' @author <NAME>
make.highcharts.points <- function(x,y,a=NULL) {
if (length(x)>0) {
n <- names(x)
x <- unname(x)
y <- unname(y)
stru <- vector("list",length(x))
if (is.null(a)) {
for (i in 1:length(x))
stru[[i]] <- list(
x=round(x[i],digits=3),
y=round(y[i],digits=3),
name=n[i]
)
}
else {
for (i in 1:length(x))
stru[[i]] <- list(
x=round(x[i],digits=3),
y=round(y[i],digits=3),
name=n[i],
alt_name=a[i]
)
}
}
else
stru <- list(x=NULL,y=NULL,name=NULL,alt_name=NULL)
return(stru)
}
#' Create a class vector
#'
#' Creates a class vector from a sample list. Internal to the \code{stat.*} functions.
#' Mostly internal use.
#'
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @return A vector of condition names.
#' @author <NAME>
#' @export
#' @examples
#' \dont'run{
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' clv <- as.class.vector(sample.list)
#'}
as.class.vector <- function(sample.list) {
classes <- vector("list",length(sample.list))
names(classes) <- names(sample.list)
for (n in names(sample.list))
classes[[n]] <- rep(n,times=length(sample.list[[n]]))
classes <- unlist(classes,use.names=FALSE)
names(classes) <- unlist(sample.list,use.names=FALSE)
return(classes)
}
#' Argument getter
#'
#' Get argument(s) from a list of arguments, e.g. normalization arguments.
#'
#' @param arg.list the initial list of a method's (e.g. normalization) arguments.
#' Can be created with the \code{\link{get.defaults}}
#' function.
#' @param arg.name the argument name inside the argument list to fetch its value.
#' @return The argument sub-list.
#' @author <NAME>
#' @examples
#' \dontrun{
#' norm.list <- get.defaults("normalization","egder")
#' a <- get.arg(norm.list,c("main.method","logratioTrim"))
#'}
get.arg <- function(arg.list,arg.name) {
return(arg.list[arg.name])
}
#' Argument setter
#'
#' Set argument(s) to a list of arguments, e.g. normalization arguments.
#'
#' @param arg.list the initial list of a method's (e.g. normalization) arguments.
#' Can be created with the \code{\link{get.defaults}}
#' function.
#' @param arg.name a named list with names the new arguments to be set, and mebers
#' the values to be set or a vector of argument
#' names. In this case, \code{arg.value} must be supplied.
#' @param arg.value when \code{arg.name} is a vector of argument names, the values
#' corresponding to these arguments.
#' @return the \code{arg.list} with the changed \code{arg.value} for \code{arg.name}.
#' @author <NAME>
#' @examples
#' \dontrun{
#' norm.list <- get.defaults("normalization","egder")
#' set.arg(norm.list,list(main.method="glm",logratioTrim=0.4))
#'}
set.arg <- function(arg.list,arg.name,arg.value=NULL) {
if (is.list(arg.name))
arg.list[names(arg.name)] <- arg.name
else if (is.character(arg.name)) {
tmp <- vector("list",length(arg.name))
names(tmp) <- arg.name
i <- 0
for (n in arg.name) {
i <- i + 1
tmp[[n]] <- arg.value[i]
}
arg.list[arg.name] <- tmp
}
return(arg.list)
}
#' Multiple testing correction helper
#'
#' A wrapper around the \code{\link{p.adjust}} function to include also the qvalue
#' adjustment procedure from the qvalue package.
#' Internal use.
#'
#' @param p a vector of p-values.
#' @param m the adjustment method. See the help of \code{\link{p.adjust}}.
#' @author <NAME>
wp.adjust <- function(p,m) {
if (m=="qvalue")
return(qvalue(p))
else
return(p.adjust(p,method=m))
}
#' List apply helper
#'
#' A wrapper around normal and parallel apply (\code{\link{mclapply}} or parallel
#' package) to avoid excessive coding for control of single or parallel code
#' execution. Internal use.
#'
#' @param m a logical indicating whether to execute in parallel or not.
#' @param ... the rest arguments to \code{\link{lapply}} (or \code{mclapply})
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' multic <- check.parallel(0.8)
#' # Test meaningful only in machines where parallel computation supported
#' if (multic) {
#' system.time(r<-wapply(TRUE,1:10,function(x) runif(1e+6)))
#' system.time(r<-wapply(FALSE,1:10,function(x) runif(1e+6)))
#' }
#'}
wapply <- function(m,...) {
if (m)
return(mclapply(...,mc.cores=getOption("cores"),mc.set.seed=FALSE))
else
return(lapply(...))
}
#' Filtering helper
#'
#' Low score filtering function. Internal use.
#'
#' @param x a data numeric matrix.
#' @param f a threshold.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data("mm9.gene.data",package="metaseqR")
#' counts <- as.matrix(mm9.gene.counts[,9:12])
#' f <- filter.low(counts,median(counts))
#'}
filter.low <- function(x,f) { return(all(x<=f)) }
#' Filtering helper
#'
#' High score filtering function. Internal use.
#'
#' @param x a data numeric matrix.
#' @param f a threshold.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' data("mm9.gene.data",package="metaseqR")
#' counts <- as.matrix(mm9.gene.counts[,9:12])
#' f <- filter.high(counts,median(counts))
#'}
filter.high <- function(x,f) { return(all(x>=f)) }
#' Message displayer
#'
#' Displays a message during execution of the several functions. Internal use.
#'
#' @param ... a vector of elements that compose the display message.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' i <- 1
#' disp("Now running iteration ",i,"...")
#'}
disp <- function(...) {
verbose <- get("VERBOSE",envir=meta.env)
if (!is.null(verbose) && verbose) {
message("\n",...,appendLF=FALSE)
}
logger <- get("LOGGER",envir=meta.env)
levalias <- c("one","two","three","four","five")
if (!is.null(logger)) {
switch(levalias[level(logger)],
one = { debug(logger,paste0(...)) },
two = { info(logger,gsub("\\n","",paste0(...))) },
three = { warn(logger,gsub("\\n","",paste0(...))) },
four = { error(logger,gsub("\\n","",paste0(...))) },
five = { fatal(logger,gsub("\\n","",paste0(...))) }
)
}
}
stopwrap <- function(...,t="fatal") {
logger <- get("LOGGER",envir=meta.env)
if (!is.null(logger)) {
if (t=="fatal")
fatal(logger,gsub("\\n","",paste0(...)))
else
error(logger,gsub("\\n","",paste0(...)))
}
stop(paste0(...))
}
warnwrap <- function(...,now=FALSE) {
logger <- get("LOGGER",envir=meta.env)
if (!is.null(logger))
warn(logger,gsub("\\n","",paste0(...)))
if (now)
warning(paste0(...),call.=FALSE,immediate.=TRUE)
else
warning(paste0(...),call.=FALSE)
}
elap2human <- function(start.time) {
start.time <- as.POSIXct(start.time)
dt <- difftime(Sys.time(),start.time,units="secs")
ndt <- as.numeric(dt)
if (ndt<60)
format(.POSIXct(dt,tz="GMT"),"%S seconds")
else if (ndt>=60 && ndt<3600)
format(.POSIXct(dt,tz="GMT"),"%M minutes %S seconds")
else if (ndt>=3600 && ndt<86400)
format(.POSIXct(dt,tz="GMT"),"%H hours %M minutes %S seconds")
else if (ndt>=86400)
format(.POSIXct(dt,tz="GMT"),"%d days %H hours %M minutes %S seconds")
}
deprecated.warning <- function(func) {
switch(func,
read.targets = {
warnwrap("\"yes\" and \"no\" for read strandedness have been ",
"deprecated. Please use \"forward\", \"forward\" or \"no\". ",
"Replacing \"yes\" with \"forward\"...")
}
)
}
metaseqR.version <- function() {
anchor.version <- "1.5-2"
current.version <- as.character(numeric_version(packageVersion("metaseqR")))
current.version <- gsub("(.*)\\.", "\\1-", current.version)
cmp <- compareVersion(current.version,anchor.version)
return(list(current=current.version,compare=cmp))
}
##' Fixed annotation updater
##'
##' A function to update the fixed annotations contained to avoid downloading every
##' time if it's not embedded. It has no parameters.
##'
##' @return This function does not return anything. It updates the fixed annotation
##' files instead.
##' @note This function cannot be used by users when the package is installed. For
##' this reason it is not exported. If you want to maintain a local copy of the
##' package and update annotation at will, you can download the package source.
## @author <NAME>
##' @examples
##' \dontrun{
##' library(metaseqr)
##' annotations.update()
##'}
#annotations.update <- function() {
# if(!require(biomaRt))
# stopwrap("Bioconductor package biomaRt is required to update annotations!")
# VERBOSE <<- TRUE
# supported.types <- c("gene","exon")
# supported.orgs <- c("hg18","hg19","mm9","mm10","rn5","dm3","danrer7")
# if (exists("ANNOTATION")) {
# for (type in supported.types) {
# for (org in supported.orgs) {
# disp("Downloading and writing ",type,"s for ",org,"...")
# tryCatch({
# tmp <- get.annotation(org,type)
# var.name <- paste(org,type,sep=".")
# assign(var.name,tmp)
# #if (!file.exists(ANNOTATION$ENSEMBL[[toupper(type)]]))
# # dir.create(ANNOTATION$ENSEMBL[[toupper(type)]],recursive=TRUE)
# #gzfh <- gzfile(file.path(ANNOTATION$ENSEMBL[[toupper(type)]],
# # paste(org,".txt.gz",sep="")),"w")
# #write.table(tmp,gzfh,sep="\t",row.names=FALSE,quote=FALSE)
# #close(gzfh)},
# save(list=eval(parse(text="var.name")),file=file.path(ANNOTATION,
# paste(org,type,"rda",sep=".")),compress=TRUE)},
# error=function(e) {
# disp("!!! Probable problem with connection to Biomart...")
# },
# finally=""
# )
# }
# }
# disp("Finished!\n")
# }
# else
# stopwrap("metaseqr environmental variables are not properly set up! ",
# "Annotations cannot be updated...")
#}
##' Fixed annotation reader
##'
##' A function to read fixed annotations from the local repository.
##'
##' @param org one of the supported organisms.
##' @param type \code{"gene"} or \code{"exon"}.
##' @return A data frame with the \code{type} annotation for \code{org}.
##' @author <NAME>
##' @export
##' @examples
##' \dontrun{
##' ann <- read.annotation("hg19","gene")
##'}
#read.annotation <- function(org,type) {
# data(list=paste(org,type,sep="."))
# ann <- eval(parse(text=paste(org,type,sep=".")))
# if (type=="gene")
# rownames(ann) <- ann$gene_id
# else if (type=="exon")
# rownames(ann) <- ann$exon_id
# return(ann)
#}
<file_sep>/man/normalize.edaseq.Rd
\name{normalize.edaseq}
\alias{normalize.edaseq}
\title{Normalization based on the EDASeq package}
\usage{
normalize.edaseq(gene.counts, sample.list,
norm.args = NULL, gene.data = NULL,
output = c("matrix", "native"))
}
\arguments{
\item{gene.counts}{a table where each row represents a
gene and each column a sample. Each cell contains the
read counts for each gene and sample. Such a table can be
produced outside metaseqr and is imported during the
basic metaseqr workflow.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{norm.args}{a list of EDASeq normalization
parameters. See the result of
\code{get.defaults("normalization",} \code{"edaseq")} for
an example and how you can modify it.}
\item{gene.data}{an optional annotation data frame (such
the ones produced by \code{get.annotation}) which
contains the GC content for each gene and from which the
gene lengths can be inferred by chromosome coordinates.}
\item{output}{the class of the output object. It can be
\code{"matrix"} (default) for versatility with other
tools or \code{"native"} for the EDASeq native S4 object
(SeqExpressionSet). In the latter case it should be
handled with suitable EDASeq methods.}
}
\value{
A matrix or a SeqExpressionSet with normalized counts.
}
\description{
This function is a wrapper over EDASeq normalization. It
accepts a matrix of gene counts (e.g. produced by
importing an externally generated table of counts to the
main metaseqr pipeline).
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
diagplot.boxplot(data.matrix,sample.list)
lengths <- round(1000*runif(nrow(data.matrix)))
starts <- round(1000*runif(nrow(data.matrix)))
ends <- starts + lengths
gc=runif(nrow(data.matrix))
gene.data <- data.frame(
chromosome=c(rep("chr1",nrow(data.matrix)/2),
rep("chr2",nrow(data.matrix)/2)),
start=starts,end=ends,gene_id=rownames(data.matrix),gc_content=gc
)
norm.data.matrix <- normalize.edaseq(data.matrix,sample.list,
gene.data=gene.data)
diagplot.boxplot(norm.data.matrix,sample.list)
}
}
\author{
<NAME>
}
<file_sep>/man/read.targets.Rd
\name{read.targets}
\alias{read.targets}
\title{Creates sample list and BAM/BED file list from file}
\usage{
read.targets(input, path = NULL)
}
\arguments{
\item{input}{a tab-delimited file structured as
follows: the first line of the external tab
delimited file should contain column names (names
are not important). The first column MUST contain
UNIQUE sample names. The second column MUST contain
the raw BAM/BED files WITH their full path.
Alternatively, the \code{path} argument should be
provided (see below). The third column MUST contain
the biological condition where each of the samples
in the first column should belong to. There is an
optional fourth column which should contain the
keywords \code{"single"} for single-end reads,
\code{"paired"} for paired-end reads or
\code{"mixed"} for BAM files that contain both
single- and paired-end reads (e.g. after a mapping
procedure with two round of alignment). If this
column is not provided, single-end reads will be
assumed. There is an optional fifth column which
stranded read assignment. It should contain the
keywords \code{"forward"} for a forward (5'->3')
strand library construction protocol, \code{"reverse"}
for a reverse (3'->5') strand library construction
protocol, or \code{"no"} for unstranded/unknown protocol.
If this column is not provided, unstranded reads will
be assumed.}
\item{path}{an optional path where all the BED/BAM
files are placed, to be prepended to the BAM/BED
file names in the targets file.}
}
\value{
A named list with four members. The first member is
a named list whose names are the conditions of the
experiments and its members are the samples belonging
to each condition. The second member is like the
first, but this time the members are named vectors
whose names are the sample names and the vector
elements are full path to BAM/BED files. The third
member is like the second, but instead of filenames
it contains information about single- or paired-end
reads (if available). The fourth member is like the
second, but instead of filenames it contains
information about the strandedness of the reads (if
available). The fifth member is the guessed type
of the input files (SAM/BAM or BED). It will be used
if not given in the main \code{\link{read2count}}
function.
}
\description{
Create the main sample list and determine the BAM/BED
files for each sample from an external file.
}
\examples{
\donttest{
targets <- data.frame(sample=c("C1","C2","T1","T2"),
filename=c("C1_raw.bam","C2_raw.bam","T1_raw.bam","T2_raw.bam"),
condition=c("Control","Control","Treatment","Treatment"))
path <- "/home/chakotay/bam"
write.table(targets,file="~/targets.txt",sep="\\t",row.names=FALSE,
quote=FALSE)
the.list <- read.targets("~/targets.txt",path=path)
sample.list <- the.list$samples
bamfile.list <- the.list$files
}
}
\author{
<NAME>
}
<file_sep>/man/set.arg.Rd
\name{set.arg}
\alias{set.arg}
\title{Argument setter}
\usage{
set.arg(arg.list, arg.name, arg.value = NULL)
}
\arguments{
\item{arg.list}{the initial list of a method's (e.g.
normalization) arguments. Can be created with the
\code{\link{get.defaults}} function.}
\item{arg.name}{a named list with names the new arguments
to be set, and mebers the values to be set or a vector of
argument names. In this case, \code{arg.value} must be
supplied.}
\item{arg.value}{when \code{arg.name} is a vector of
argument names, the values corresponding to these
arguments.}
}
\value{
the \code{arg.list} with the changed \code{arg.value} for
\code{arg.name}.
}
\description{
Set argument(s) to a list of arguments, e.g.
normalization arguments.
}
\examples{
\donttest{
norm.list <- get.defaults("normalization","egder")
set.arg(norm.list,list(main.method="glm",logratioTrim=0.4))
}
}
\author{
<NAME>
}
<file_sep>/man/diagplot.pairs.Rd
\name{diagplot.pairs}
\alias{diagplot.pairs}
\title{Massive X-Y, M-D correlation plots}
\usage{
diagplot.pairs(x, output = "x11", path = NULL, ...)
}
\arguments{
\item{x}{the read counts matrix or data frame.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filename of the pairwise comparisons plot produced if
it's a file.
}
\description{
This function uses the read counts matrix to create
pairwise correlation plots. The upper diagonal of the
final image contains simple scatterplots of each sample
against each other (log2 scale) while the lower diagonal
contains mean-difference plots for the same samples (log2
scale). This type of diagnostic plot may not be
interpretable for more than 10 samples.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
diagplot.pairs(data.matrix)
}
\author{
<NAME>
}
<file_sep>/man/graphics.close.Rd
\name{graphics.close}
\alias{graphics.close}
\title{Close plotting device}
\usage{
graphics.close(o)
}
\arguments{
\item{o}{the plotting device, see main metaseqr function}
}
\description{
Wrapper function to close a plotting device. Internal use
only.
}
\examples{
\donttest{
graphics.close("pdf")
}
}
\author{
<NAME>
}
<file_sep>/man/check.num.args.Rd
\name{check.num.args}
\alias{check.num.args}
\title{Numeric argument validator}
\usage{
check.num.args(arg.name, arg.value, arg.type, arg.bounds,
direction)
}
\arguments{
\item{arg.name}{the name of the argument that is checked
(for display purposes).}
\item{arg.value}{the value(s) of the argument to be
checked.}
\item{arg.type}{either the string \code{"numeric"} to
denote generic double-like R numerics or \code{"integer"}
for integer values.}
\item{arg.bounds}{a numeric or a vector with 2 elements,
restraining \code{arg.value} to be within the bounds
defined by the input vector or e.g. larger (smaller) than
the numeric value. See examples.}
\item{direction}{a string denoting to which direction the
\code{arg.value} should be compared with
\code{arg.bounds}. For example, \code{"both"} should be
given with a two element vector against which,
\code{arg.value} will be checked to see whether it is
smaller than the low boundary or larger than the higher
boundary. In that case, the function will throw an error.
The direction parameter can be one of: \code{"both"}
(described above), \code{"botheq"} (as above, but the
\code{arg.val} is also checked for equality -closed
intervals), \code{"gt"} or \code{"gte"} (check whether
\code{arg.val} is smaller or smaller than or equal to the
first value of \code{arg.bounds}), \code{"lt"} or
\code{"lte"} (check whether \code{arg.val} is larger or
larger than or equal to the first value of
\code{arg.bounds}).}
}
\description{
Checks if one or more given numeric argument(s) satisfy
several rules concerning numeric arguments, e.g. proper
bounds or proper format (e.g. it must be a number and not
a character). Mostly for internal use.
}
\examples{
pcut <- 1.2 # A probability cannot be larger than 1! It will throw an error!
#check.num.args("pcut",pcut,"numeric",c(0,1),"botheq")
pcut <- 0.05 # Pass
check.num.args("pcut",pcut,"numeric",c(0,1),"botheq")
gc.col <- 3.4 # A column in a file cannot be real! It will throw an error!
#check.num.args("gc.col",gc.col,"integer",0,"gt")
gc.col <- 5L # Pass
check.num.args("gc.col",gc.col,"integer",0,"gt")
}
\author{
<NAME>
}
<file_sep>/man/construct.gene.model.Rd
\name{construct.gene.model}
\alias{construct.gene.model}
\title{Assemble a gene model based on exon counts}
\usage{
construct.gene.model(exon.counts, sample.list, gene.data,
multic = FALSE)
}
\arguments{
\item{exon.counts}{the exon counts data frame produced by
reading the exon read counts file.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{gene.data}{an annotation data frame from the same
organism as \code{exon.counts} (such the ones produced by
\code{get.annotation}).}
\item{multic}{a logical value indicating the presence of
multiple cores. Defaults to \code{FALSE}. Do not change
it if you are not sure whether package multicore has been
loaded or not.}
}
\value{
A named list where names represent samples. Each list
member is a also a named list where names correspond to
gene ids and members are named vectors. Each vector is
named according to the exons corresponding to each gene
and contains the read counts for each exon. This
structure is used for exon filtering and assembling final
gene counts in the metaseqr pipeline.
}
\description{
This function assembles gene models (single genes, not
isoforms) based on the input exon read counts file and a
gene annotation data frame, either from an external file
provided by the user, or with the
\code{\link{get.annotation}} function. The
\code{gene.data} argument should have a specific format
and for this reason it's better to use one of the two
aforementioned ways to supply it. This function is
intended mostly for internal use but can be used if the
requirements are met.
}
\examples{
\donttest{
# Takes some time to run...
data("hg19.exon.data",package="metaseqR")
gene.data <- get.annotation("hg19","gene","ensembl")
reduced.gene.data <- reduce.gene.data(hg19.exon.counts,
gene.data)
multic <- check.parallel(0.4)
gene.model <- construct.gene.model(hg19.exon.counts,
sample.list.hg19,gene.data,multic)
}
}
\author{
<NAME>
}
<file_sep>/man/build.export.Rd
\name{build.export}
\alias{build.export}
\title{Results export builder}
\usage{
build.export(gene.data, raw.gene.counts,
norm.gene.counts, flags, sample.list, cnt,
statistics, raw.list, norm.list,
p.mat = matrix(NA, nrow(gene.data), length(statistics)),
adj.p.mat = matrix(NA, nrow(gene.data), length(statistics)),
sum.p = rep(NA, nrow(gene.data)),
adj.sum.p = rep(NA, nrow(gene.data)),
export.what = c("annotation", "p.value", "adj.p.value", "meta.p.value",
"adj.meta.p.value", "fold.change", "stats", "counts","flags"),
export.scale = c("natural", "log2", "log10", "rpgm", "vst"),
export.values = c("raw", "normalized"),
export.stats = c("mean", "median", "sd", "mad", "cv", "rcv"),
log.offset = 1, report = TRUE)
}
\arguments{
\item{gene.data}{an annotation data frame (such the ones
produced by \code{\link{get.annotation}}).}
\item{raw.gene.counts}{a matrix of filtering flags (0,1), created
by the filtering functions.}
\item{norm.gene.counts}{a matrix of normalized gene
counts.}
\item{flags}{a matrix of normalized gene
counts.}
\item{sample.list}{see the documentation of
\code{\link{metaseqr}}.}
\item{cnt}{the statistical contrast for which the export
builder is currently running.}
\item{statistics}{the statistical tests used (see the
documentation of \code{\link{metaseqr}}).}
\item{raw.list}{a list of transformed un-normalized
counts, see the documentation
of \code{\link{make.transformation}}.}
\item{norm.list}{a list of transformed normalized counts,
see the documentation
of \code{\link{make.transformation}}.}
\item{p.mat}{a matrix of p-values, see the documentation
of \code{\link{metaseqr}}.}
\item{adj.p.mat}{a matrix of adjusted p-values, see the
documentation of \code{\link{metaseqr}}.}
\item{sum.p}{a vector of combined p-values, see the
documentation of \code{\link{metaseqr}}.}
\item{adj.sum.p}{a vector of adjusted combined p-values,
see the documentation of \code{\link{metaseqr}}.}
\item{export.what}{see the documentation of
\code{\link{metaseqr}}.}
\item{export.scale}{see the documentation of
\code{\link{metaseqr}}.}
\item{export.values}{see the documentation of
\code{\link{metaseqr}}.}
\item{export.stats}{see the documentation of
\code{\link{metaseqr}}.}
\item{log.offset}{see the documentation of
\code{\link{metaseqr}}.}
\item{report}{see the documentation of
\code{\link{metaseqr}}.}
}
\value{
A list with three members: a data frame to be exported in
a text file, a long string with the result in a html
formatted table (if \code{report=TRUE}) and the column
names of the output data frame.
}
\description{
This function help build the output files of the metaseqr
pipeline based on several elements produced during the
pipeline execution. It is intended for internal use and
not available to the users.
}
\examples{
\dontrun{
# Not yet available
}
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.count.R
#' SAM/BAM/BED file reader helper for the metaseqr pipeline
#'
#' This function is a helper for the \code{metaseqr} pipeline, for reading SAM/BAM
#' or BED files when a read counts file is not available.
#'
#' @param targets a named list, the output of \code{\link{read.targets}}.
#' @param file.type the type of raw input files. It can be \code{"bed"} for BED
#' files or \code{"sam"}, \code{"bam"} for SAM/BAM files. See the same argument
#' in the main \code{\link{metaseqr}} function for the case of auto-guessing.
#' @param annotation see the \code{annotation} argument in the main
#' \code{\link{metaseqr}} function. The \code{"annotation"} parameter here is the
#' result of the same parameter in the main function. See also
#' \code{\link{get.annotation}}.
#' @param has.all.fields a logical variable indicating if all annotation fields
#' used by \code{metaseqr} are available (that is apart from the main chromosome,
#' start, end, unique id and strand columns, if also present are the gene name and
#' biotype columns). The default is \code{FALSE}.
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not.
#' @return A data frame with counts for each sample, ready to be passed to the
#' main \code{\link{metaseqr}} pipeline.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' my.targets <- read.targets("my_mm9_study_bam_files.txt")
#' sample.list <- my.targets$samples
#' file.list <- my.targets$files
#' gene.data <- get.annotation("mm9","gene")
#' r2c <- read2count(files.list=file.list,file.type=my.targets$type,
#' annotation=gene.data)
#' gene.counts <- r2c$counts
#' libsize.list <- r2s$libsize
#'}
read2count <- function(targets,annotation,file.type=targets$type,
trans.level="gene",utr.flank=500,has.all.fields=FALSE,multic=FALSE) {
if (missing(targets))
stopwrap("You must provide the targets argument!")
if (missing(annotation))
stopwrap("You must provide an annotation data frame!")
if (!require(GenomicRanges))
stopwrap("The Bioconductor package GenomicRanges is required to ",
"proceed!")
if (file.type=="bed" && !require(rtracklayer))
stopwrap("The Bioconductor package rtracklayer is required to process ",
"BED files!")
if (file.type %in% c("sam","bam")) {
if (!require(Rsamtools))
stopwrap("The Bioconductor package Rsamtools is required to ",
"process BAM files!")
}
if (!is.list(targets) && file.exists(targets))
targets <- read.targets(targets)
else if (!is.list(targets) && !file.exists(targets))
stopwrap("You must provide a targets list or a valid targets file!")
# Convert annotation to GRanges
disp("Converting annotation to GenomicRanges object...")
if (packageVersion("GenomicRanges")<1.14) { # Classic way
if (has.all.fields)
annotation.gr <- GRanges(
seqnames=Rle(annotation[,1]),
ranges=IRanges(start=annotation[,2],end=annotation[,3]),
strand=Rle(annotation[,6]),
name=as.character(annotation[,4]),
symbol=as.character(annotation[,7]),
biotype=as.character(annotation[,8])
)
else
annotation.gr <- GRanges(
seqnames=Rle(annotation[,1]),
ranges=IRanges(start=annotation[,2],end=annotation[,3]),
strand=Rle(annotation[,6]),
name=as.character(annotation[,4])
)
}
else # Use native method in newer versions of GenomicRanges
annotation.gr <- makeGRangesFromDataFrame(
df=annotation,
keep.extra.columns=TRUE,
seqnames.field="chromosome"
)
# If the count type is "exon", we must reduce the overlapping exons belonging
# to multiple transcripts, so as to avoid inflating the final read count when
# summing all exons
inter.feature <- FALSE
if (length(grep("exon",colnames(annotation)))>0) { # count.type is exon
if (length(grep("MEX",annotation$exon_id[1]))) # Retrieved from previous
merged.annotation <- annotation
else {
if (trans.level=="gene") {
disp("Merging exons to create unique gene models...")
annotation.gr <- reduce.exons(annotation.gr,multic=multic)
#merged.annotation <- as.data.frame(annotation.gr) # Bug?
}
#else if (trans.level=="transcript") {
# disp("Merging exons to create unique gene models...")
# annotation.gr <- reduce.exons.transcript(annotation.gr,
# multic=multic)
#}
merged.annotation <- data.frame(
chromosome=as.character(seqnames(annotation.gr)),
start=start(annotation.gr),
end=end(annotation.gr),
exon_id=if (!is.null(annotation.gr$exon_id))
as.character(annotation.gr$exon_id) else
as.character(annotation.gr$name),
gene_id=if (!is.null(annotation.gr$gene_id))
as.character(annotation.gr$gene_id) else
as.character(annotation.gr$name),
strand=as.character(strand(annotation.gr)),
gene_name=if (!is.null(annotation.gr$gene_name))
as.character(annotation.gr$gene_name) else
if (!is.null(annotation.gr$symbol))
as.character(annotation.gr$name) else NULL,
biotype=if (!is.null(annotation.gr$biotype))
as.character(annotation.gr$biotype) else NULL
)
rownames(merged.annotation) <-
as.character(merged.annotation$exon_id)
}
}
else if (length(grep("transcript",colnames(annotation)))>0) { # count.type may be utr
if (length(grep("MET",annotation$transcript_id[1]))
|| length(grep("MEU",annotation$transcript_id[1]))) # Retrieved from previous
merged.annotation <- annotation
else {
if (trans.level=="gene") {
disp("Merging transcript 3' UTRs to create unique ",
"gene models...")
annotation.gr <-
reduce.transcripts.utr(annotation.gr,multic=multic)
}
if (trans.level=="transcript") {
disp("Merging transcript 3' UTRs to create unique ",
"transcript models...")
annotation.gr <-
reduce.transcripts.utr.transcript(annotation.gr,
multic=multic)
}
if (utr.flank > 0) {
disp("Flanking merged transcript 3' UTRs per ",utr.flank,
"bp...")
w <- width(annotation.gr)
annotation.gr <- promoters(annotation.gr,upstream=utr.flank,
downstream=0)
annotation.gr <- resize(annotation.gr,width=w+2*utr.flank)
}
#merged.annotation <- as.data.frame(annotation.gr) # Bug?
merged.annotation <- data.frame(
chromosome=as.character(seqnames(annotation.gr)),
start=start(annotation.gr),
end=end(annotation.gr),
transcript_id=if (!is.null(annotation.gr$transcript_id))
as.character(annotation.gr$transcript_id) else
as.character(annotation.gr$name),
gene_id=if (!is.null(annotation.gr$gene_id))
as.character(annotation.gr$gene_id) else
as.character(annotation.gr$name),
strand=as.character(strand(annotation.gr)),
gene_name=if (!is.null(annotation.gr$gene_name))
as.character(annotation.gr$gene_name) else
if (!is.null(annotation.gr$symbol))
as.character(annotation.gr$name) else NULL,
biotype=if (!is.null(annotation.gr$biotype))
as.character(annotation.gr$biotype) else NULL
)
rownames(merged.annotation) <-
as.character(merged.annotation$transcript_id)
}
inter.feature = FALSE # Quant-Seq
}
else
merged.annotation <- NULL
# Continue
files.list <- targets$files
sample.names <- unlist(lapply(files.list,names),use.names=FALSE)
sample.files <- unlist(files.list,use.names=FALSE)
names(sample.files) <- sample.names
if (!is.null(targets$paired)) {
paired <- unlist(targets$paired,use.names=FALSE)
names(paired) <- sample.names
}
else
paired <- NULL
if (!is.null(targets$stranded)) {
stranded <- unlist(targets$stranded,use.names=FALSE)
names(stranded) <- sample.names
}
else
stranded <- NULL
counts <- matrix(0,nrow=length(annotation.gr),ncol=length(sample.names))
if (length(grep("exon",colnames(annotation)))>0)
rownames(counts) <- as.character(annotation.gr$exon_id)
else if (length(grep("transcript",colnames(annotation)))>0)
rownames(counts) <- as.character(annotation.gr$transcript_id)
else
rownames(counts) <- as.character(annotation.gr$gene_id)
colnames(counts) <- sample.names
libsize <- vector("list",length(sample.names))
names(libsize) <- sample.names
if (file.type=="bed") {
ret.val <- wapply(multic,sample.names,function(n,sample.files) {
disp("Reading bed file ",basename(sample.files[n]),
" for sample with name ",n,". This might take some time...")
bed <- import.bed(sample.files[n],trackLine=FALSE)
disp(" Checking for chromosomes not present in the annotation...")
bed <- bed[which(!is.na(match(as(seqnames(bed),"character"),
seqlevels(annotation.gr))))]
libsize <- length(bed)
if (length(bed)>0) {
disp(" Counting reads overlapping with given annotation...")
counts <- countOverlaps(annotation.gr,bed)
}
else
warnwrap(paste("No reads left after annotation chromosome ",
"presence check for sample ",n,sep=""))
gc(verbose=FALSE)
return(list(counts=counts,libsize=libsize))
},sample.files)
}
else if (file.type %in% c("sam","bam")) {
if (file.type=="sam") {
for (n in sample.names) {
dest <- file.path(dirname(sample.files[n]),n)
disp("Converting sam file ",basename(sample.files[n]),
" to bam file ",basename(dest),"...")
asBam(file=sample.files[n],destination=dest,overwrite=TRUE)
sample.files[n] <- paste(dest,"bam",sep=".")
}
}
ret.val <- wapply(multic,sample.names,function(n,sample.files,paired,
stranded) {
disp("Reading bam file ",basename(sample.files[n])," for sample ",
"with name ",n,". This might take some time...")
if (!is.null(paired)) {
p <- tolower(paired[n])
if (p=="single") {
singleEnd <- TRUE
fragments <- FALSE
asMates <- FALSE
}
else if (p=="paired") {
singleEnd <- FALSE
fragments <- FALSE
asMates <- TRUE
}
else if (p=="mixed") {
singleEnd <- FALSE
fragments <- TRUE
asMates <- TRUE
}
else {
warnwrap("Information regarding single- or paired-end ",
"reads is not correctly provided! Assuming single...")
singleEnd <- TRUE
fragments <- FALSE
asMates <- FALSE
}
}
else {
singleEnd <- TRUE
fragments <- FALSE
asMates <- FALSE
}
if (!is.null(stranded)) {
s <- tolower(stranded[n])
if (s %in% c("forward","reverse"))
ignore.strand <- FALSE
else if (s=="no")
ignore.strand <- TRUE
else {
warnwrap("Information regarding strandedness of the reads ",
"is not correctly provided! Assuming unstranded...")
ignore.strand <- TRUE
}
}
else
ignore.strand <- TRUE
# Check remoteness
if (length(grep("^(http|ftp)",sample.files[n],perl=TRUE))>=1) {
reads <- as(readGAlignments(file=sample.files[n]),"GRanges")
libsize <- length(reads)
is.remote <- TRUE
}
else {
reads <- BamFile(sample.files[n],asMates=asMates)
libsize <- countBam(reads,
param=ScanBamParam(scanBamFlag(isUnmappedQuery=FALSE)))$records
is.remote <- FALSE
}
if (libsize>0) {
disp(" Counting reads overlapping with given annotation...")
if (singleEnd & !fragments)
disp(" ...for single-end reads...")
else if (!singleEnd & !fragments)
disp(" ...for paired-end reads...")
else if (!singleEnd & fragments)
disp(" ...for mixed single- and paired-end reads...")
if (ignore.strand)
disp(" ...ignoring strandedness...")
else {
disp(" ...assuming ",s," sequenced reads...")
if (s=="reverse")
strand(annotation.gr) <- ifelse(strand(
annotation.gr)=="+","-","+")
}
if (is.remote)
disp(" ...for remote BAM file... might take longer...")
counts <- summarizeOverlaps(annotation.gr,reads,
singleEnd=singleEnd,fragments=fragments,
ignore.strand=ignore.strand,inter.feature=inter.feature)
counts <- assays(counts)$counts
}
else
warnwrap(paste("No reads left after annotation chromosome ",
"presence check for sample ",n,sep=""))
gc(verbose=FALSE)
return(list(counts=counts,libsize=libsize))
},sample.files,paired,stranded)
}
for (i in 1:length(ret.val)) {
counts[,i] <- ret.val[[i]]$counts
libsize[[i]] <- ret.val[[i]]$libsize
}
return(list(counts=counts,libsize=libsize,mergedann=merged.annotation))
}
#' Merges exons to create a unique set of exons for each gene
#'
#' This function uses the \code{"reduce"} function of IRanges to construct virtual
#' unique exons for each gene, so as to avoid inflating the read counts for each
#' gene because of multiple possible transcripts. If the user wants transcripts
#' instead of genes, they should be supplied to the original annotation table.
#'
#' @param gr a GRanges object created from the supplied annotation (see also the
#' \code{\link{read2count}} and \code{\link{get.annotation}} functions.
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not.
#' @return A GRanges object with virtual merged exons for each gene/transcript.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(GenomicRanges)
#' ann <- get.annotation("mm9","exon")
#' gr <- makeGRangesFromDataFrame(
#' df=ann,
#' keep.extra.columns=TRUE,
#' seqnames.field="chromosome"
#' )
#' re <- reduce.exons(gr)
#'}
reduce.exons <- function(gr,multic=FALSE) {
gene <- unique(as.character(gr$gene_id))
if (!is.null(gr$gene_name))
gn <- gr$gene_name
else
gn <- NULL
if (!is.null(gr$biotype))
bt <- gr$biotype
else
bt <- NULL
red.list <- wapply(multic,gene,function(x,a,g,b) {
tmp <- a[a$gene_id==x]
if (!is.null(g))
gena <- as.character(tmp$gene_name[1])
if (!is.null(b))
btty <- as.character(tmp$biotype[1])
merged <- reduce(tmp)
n <- length(merged)
meta <- DataFrame(
exon_id=paste(x,"MEX",1:n,sep="_"),
gene_id=rep(x,n)
)
if (!is.null(g))
meta$gene_name <- rep(gena,n)
if (!is.null(b))
meta$biotype <- rep(btty,n)
mcols(merged) <- meta
return(merged)
},gr,gn,bt)
return(do.call("c",red.list))
}
#' Merges 3' UTR of transcripts to create a unique set of coordinates for each
#' transcript
#'
#' This function uses the \code{"reduce"} function of IRanges to construct virtual
#' unique transcripts for each gene, so as to avoid inflating the read counts for
#' each gene because of multiple possibly overlaping 3' UTR starts/ends when using
#' metaseqR with QuantSeq protocol.
#'
#' @param gr a GRanges object created from the supplied annotation (see also the
#' \code{\link{read2count}} and \code{\link{get.annotation}} functions.
#' @param multic a logical value indicating the presence of multiple cores. Defaults
#' to \code{FALSE}. Do not change it if you are not sure whether package parallel
#' has been loaded or not.
#' @return A GRanges object with virtual merged exons for each gene/transcript.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(GenomicRanges)
#' ann <- get.annotation("mm9","exon")
#' gr <- makeGRangesFromDataFrame(
#' df=ann,
#' keep.extra.columns=TRUE,
#' seqnames.field="chromosome"
#' )
#' re <- reduce.exons(gr)
#'}
reduce.transcripts.utr <- function(gr,multic=FALSE) {
gene <- unique(as.character(gr$gene_id))
if (!is.null(gr$gene_name))
gn <- gr$gene_name
else
gn <- NULL
if (!is.null(gr$biotype))
bt <- gr$biotype
else
bt <- NULL
red.list <- wapply(multic,gene,function(x,a,g,b) {
tmp <- a[a$gene_id==x]
if (!is.null(g))
gena <- as.character(tmp$gene_name[1])
if (!is.null(b))
btty <- as.character(tmp$biotype[1])
merged <- reduce(tmp)
n <- length(merged)
meta <- DataFrame(
transcript_id=paste(x,"MET",1:n,sep="_"),
gene_id=rep(x,n)
)
if (!is.null(g))
meta$gene_name <- rep(gena,n)
if (!is.null(b))
meta$biotype <- rep(btty,n)
mcols(merged) <- meta
return(merged)
},gr,gn,bt)
return(do.call("c",red.list))
}
reduce.transcripts.utr.transcript <- function(gr,multic=FALSE) {
trans <- unique(as.character(gr$transcript_id))
if (!is.null(gr$gene_name))
gn <- gr$gene_name
else
gn <- NULL
if (!is.null(gr$biotype))
bt <- gr$biotype
else
bt <- NULL
red.list <- wapply(multic,trans,function(x,a,g,b) {
tmp <- a[a$transcript_id==x]
if (!is.null(g))
gena <- as.character(tmp$gene_name[1])
if (!is.null(b))
btty <- as.character(tmp$biotype[1])
merged <- reduce(tmp)
n <- length(merged)
meta <- DataFrame(
transcript_id=paste(x,"MEU",1:n,sep="_"),
gene_id=rep(x,n)
)
if (!is.null(g))
meta$gene_name <- rep(gena,n)
if (!is.null(b))
meta$biotype <- rep(btty,n)
mcols(merged) <- meta
return(merged)
},gr,gn,bt)
return(do.call("c",red.list))
}
#' Creates sample list and BAM/BED file list from file
#'
#' Create the main sample list and determine the BAM/BED files for each sample
#' from an external file.
#'
#' @param input a tab-delimited file structured as follows: the first line of the
#' external tab delimited file should contain column names (names are not important).
#' The first column MUST contain UNIQUE sample names. The second column MUST contain
#' the raw BAM/BED files WITH their full path. Alternatively, the \code{path}
#' argument should be provided (see below). The third column MUST contain the
#' biological condition where each of the samples in the first column should belong
#' to. There is an optional fourth column which should contain the keywords
#' \code{"single"} for single-end reads, \code{"paired"} for paired-end reads or
#' \code{"mixed"} for BAM files that contain bith paired- and single-end reads.
#' If this column is not provided, single-end reads will be assumed. There is an
#' optional fifth column which controls stranded read assignment. It should
#' contain the keywords \code{"forward"} for a forward (5'->3') strand library
#' construction protocol, \code{"reverse"} for a reverse (3'->5') strand library
#' construction protocol, or \code{"no"} for unstranded/unknown protocol. If
#' this column is not provided, unstranded reads will be assumed.
#' @param path an optional path where all the BED/BAM files are placed, to be
#' prepended to the BAM/BED file names in the targets file.
#' @return A named list with three members. The first member is a named list whose
#' names are the conditions of the experiments and its members are the samples
#' belonging to each condition. The second member is like the first, but this time
#' the members are named vectors whose names are the sample names and the vector
#' elements are full path to BAM/BED files. The third member is the guessed type
#' of the input files (BAM or BED). It will be used if not given in the main
#' \code{\link{read2count}} function.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' targets <- data.frame(sample=c("C1","C2","T1","T2"),
#' filename=c("C1_raw.bam","C2_raw.bam","T1_raw.bam","T2_raw.bam"),
#' condition=c("Control","Control","Treatment","Treatment"))
#' path <- "/home/chakotay/bam"
#' write.table(targets,file="targets.txt",sep="\t",row.names=F,quote="")
#' the.list <- read.targets("targets.txt",path=path)
#' sample.list <- the.list$samples
#' bamfile.list <- the.list$files
#'}
read.targets <- function(input,path=NULL) {
if (missing(input) || !file.exists(input))
stopwrap("The targets file should be a valid existing text file!")
tab <- read.delim(input,strip.white=TRUE)
samples <- as.character(tab[,1])
conditions <- unique(as.character(tab[,3]))
rawfiles <- as.character(tab[,2])
if (!is.null(path)) {
tmp <- dirname(rawfiles) # Test if there is already a path
if (any(tmp=="."))
rawfiles <- file.path(path,basename(rawfiles))
}
# Check if they exist!!!
for (f in rawfiles) {
if (!file.exists(f))
stopwrap("Raw reads input file ",f," does not exist! Please check!")
}
if (length(samples) != length(unique(samples)))
stopwrap("Sample names must be unique for each sample!")
if (length(rawfiles) != length(unique(rawfiles)))
stopwrap("File names must be unique for each sample!")
sample.list <- vector("list",length(conditions))
names(sample.list) <- conditions
for (n in conditions)
sample.list[[n]] <- samples[which(as.character(tab[,3])==n)]
file.list <- vector("list",length(conditions))
names(file.list) <- conditions
for (n in conditions) {
file.list[[n]] <- rawfiles[which(as.character(tab[,3])==n)]
names(file.list[[n]]) <- samples[which(as.character(tab[,3])==n)]
}
if (ncol(tab)>3) { # Has info about single- or paired-end reads / strand
if (ncol(tab)==4) { # Stranded or paired
whats <- tolower(as.character(tab[,4]))
if (!all(whats %in% c("yes","no","forward","reverse",
"single","paired")))
stopwrap("Unknown options for paired-end reads and/or ",
"strandedness in targets file.")
what <- whats[1]
if (what %in% c("single","paired")) {
has.paired.info <- TRUE
has.stranded.info <- FALSE
}
else {
if (what %in% c("yes","no")) {
deprecated.warning("read.targets")
tmp <- as.character(tab[,4])
tmp[tmp=="yes"] <- "forward"
tab[,4] <- tmp
has.paired.info <- FALSE
has.stranded.info <- TRUE
}
if (what %in% c("forward","reverse","no")) {
has.paired.info <- FALSE
has.stranded.info <- TRUE
}
}
}
if (ncol(tab)==5) { # Both
whats.paired <- tolower(as.character(tab[,4]))
if (!all(whats.paired %in% c("single","paired","mixed")))
stopwrap("Unknown option for type of reads (single, paired, ",
"mixed) in targets file.")
whats.strand <- tolower(as.character(tab[,5]))
if (!all(whats.strand %in% c("yes","no","forward","reverse")))
stopwrap("Unknown option for read strandedness in targets file")
if (any(whats.strand=="yes")) {
deprecated.warning("read.targets")
tmp <- as.character(tab[,5])
tmp[tmp=="yes"] <- "forward"
tab[,5] <- tmp
}
has.paired.info <- TRUE
has.stranded.info <- TRUE
}
if (has.paired.info && !has.stranded.info) {
paired.list <- vector("list",length(conditions))
names(paired.list) <- conditions
for (n in conditions) {
paired.list[[n]] <- character(length(sample.list[[n]]))
names(paired.list[[n]]) <- sample.list[[n]]
for (nn in names(paired.list[[n]]))
paired.list[[n]][nn] <- as.character(tab[which(as.character(
tab[,1])==nn),4])
}
}
else
paired.list <- NULL
if (has.stranded.info && !has.paired.info) {
stranded.list <- vector("list",length(conditions))
names(stranded.list) <- conditions
for (n in conditions) {
stranded.list[[n]] <- character(length(sample.list[[n]]))
names(stranded.list[[n]]) <- sample.list[[n]]
for (nn in names(stranded.list[[n]]))
stranded.list[[n]][nn] <- as.character(tab[which(as.character(
tab[,1])==nn),4])
}
}
else
stranded.list <- NULL
if (has.stranded.info && has.paired.info) {
stranded.list <- vector("list",length(conditions))
names(stranded.list) <- conditions
for (n in conditions) {
stranded.list[[n]] <- character(length(sample.list[[n]]))
names(stranded.list[[n]]) <- sample.list[[n]]
for (nn in names(stranded.list[[n]]))
stranded.list[[n]][nn] <- as.character(tab[which(as.character(
tab[,1])==nn),5])
}
paired.list <- vector("list",length(conditions))
names(paired.list) <- conditions
for (n in conditions) {
paired.list[[n]] <- character(length(sample.list[[n]]))
names(paired.list[[n]]) <- sample.list[[n]]
for (nn in names(paired.list[[n]]))
paired.list[[n]][nn] <- as.character(tab[which(as.character(
tab[,1])==nn),4])
}
}
}
else
paired.list <- stranded.list <- NULL
# Guess file type based on only one of them
tmp <- file.list[[1]][1]
if (length(grep("\\.bam$",tmp,ignore.case=TRUE,perl=TRUE))>0)
type <- "bam"
else if (length(grep("\\.sam$",tmp,ignore.case=TRUE,perl=TRUE))>0)
type <- "sam"
else if (length(grep("\\.bed$",tmp,ignore.case=TRUE,perl=TRUE)>0))
type <- "bed"
else
type <- NULL
return(list(samples=sample.list,files=file.list,paired=paired.list,
stranded=stranded.list,type=type))
}
<file_sep>/man/get.valid.chrs.Rd
\name{get.valid.chrs}
\alias{get.valid.chrs}
\title{Annotation downloader helper}
\usage{
get.valid.chrs(org)
}
\arguments{
\item{org}{the organism for which to return the
chromosomes.}
}
\value{
A character vector of chromosomes.
}
\description{
Returns a vector of chromosomes to maintain after
annotation download. Internal use.
}
\examples{
hg18.chr <- get.valid.chrs("hg18")
}
\author{
<NAME>
}
<file_sep>/man/diagplot.filtered.Rd
\name{diagplot.filtered}
\alias{diagplot.filtered}
\title{Diagnostic plot for filtered genes}
\usage{
diagplot.filtered(x, y, output = "x11", path = NULL, ...)
}
\arguments{
\item{x}{an annotation data frame like the ones produced
by \code{\link{get.annotation}}. \code{x} should be the
filtered annotation according to metaseqR's filters.}
\item{y}{an annotation data frame like the ones produced
by \code{\link{get.annotation}}. \code{y} should contain
the total annotation without the application of any
metaseqr filter.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filenames of the plots produced in a named list with
names the \code{which.plot} argument. If
output=\code{"x11"}, no output filenames are produced.
}
\description{
This function plots a grid of four graphs depicting: in
the first row, the numbers of filtered genes per
chromosome in the first column and per biotype in the
second column. In the second row, the percentages of
filtered genes per chromosome related to the whole genome
in the first columns and per biotype in the second
column.
}
\examples{
y <- get.annotation("mm9","gene")
x <- y[-sample(1:nrow(y),10000),]
diagplot.filtered(x,y)
}
\author{
<NAME>
}
<file_sep>/man/meta.test.Rd
\name{meta.test}
\alias{meta.test}
\title{Meta-analysis using several RNA-Seq statistics}
\usage{
meta.test(cp.list,
meta.p = c("simes", "bonferroni", "fisher", "dperm.min",
"dperm.max", "dperm.weight", "fperm", "whitlock",
"minp", "maxp", "weight", "pandora", "none"), counts,
sample.list, statistics, stat.args, libsize.list,
nperm = 10000, weight = rep(1/length(statistics),
length(statistics)), reprod=TRUE, multic = FALSE)
}
\arguments{
\item{cp.list}{a named list whose names are the contrasts
requested from metaseqr. Each member is a p-value matrix
whose colnames are the names of the statistical tests
applied to the data. See the main \code{\link{metaseqr}}
help page.}
\item{meta.p}{the p-value combination method to use. See
the main \code{\link{metaseqr}} help page.}
\item{counts}{the normalized and possibly filtered read
counts matrix. See the main \code{\link{metaseqr}} help
page.}
\item{sample.list}{the list containing condition names
and the samples under each condition. See the main
\code{\link{metaseqr}} help page.}
\item{statistics}{the statistical algorithms used in
metaseqr. See the main \code{\link{metaseqr}} help page.}
\item{stat.args}{the parameters for each statistical
argument. See the main \code{\link{metaseqr}} help page.}
\item{libsize.list}{a list with library sizes. See the
main \code{\link{metaseqr}} and the \code{stat.*} help
pages.}
\item{nperm}{the number of permutations (Monte Carlo
simulations) to perform.}
\item{weight}{a numeric vector of weights for each
statistical algorithm.}
\item{reprod}{create reproducible permutations when
\code{meta.p="dperm.min"}, \code{meta.p="dperm.max"}
or \code{meta.p="dperm.weight"}. Ideally one would
want to create the same set of indices for a given
dataset so as to create reproducible p-values. If
\code{reprod=TRUE}, a fixed seed is used by
\code{meta.perm} for all the datasets analyzed with
\code{metaseqr}. If \code{reprod=FALSE}, then the
p-values will not be reproducible, although
statistical significance is not expected to change
for a large number of resambling. Finally,
\code{reprod} can be a numeric vector of seeds with
the same length as \code{nperm} so that the user can
supply his/her own seeds.}
\item{multic}{use multiple cores to execute the
premutations. This is an external parameter and implies
the existence of multicore package in the execution
environment. See the main \code{\link{metaseqr}} help
page.}
}
\value{
A named list with combined p-values. The names are the
contrasts and the list members are combined p-value
vectors, one for each contrast.
}
\description{
This function calculates the combined p-values when
multiple statistical algorithms are applied to the input
dataset. It is a helper and it requires very specific
arguments so it should not be used individually
}
\examples{
\donttest{
# Not yet available
}
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.query.R
#' Download annotation from UCSC servers, according to organism and source
#'
#' Directly downloads UCSC and RefSeq annotation files from UCSC servers to be
#' used with metaseqR. This functionality is used when the package RMySQL is not
#' available for some reason, e.g. Windows machines.
#'
#' @param org one of metaseqR supported organisms.
#' @param type either \code{"gene"} or \code{"exon"}.
#' @param refdb one of \code{"ucsc"} or \code{"refseq"} to use the UCSC or RefSeq
#' annotation sources respectively.
#' @return A data frame with annotation elements.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' db.file <- get.ucsc.file("hg18","gene","ucsc")
#'}
get.ucsc.dbl <- function(org,type,refdb="ucsc") {
type <- tolower(type[1])
org <- tolower(org[1])
refdb <- tolower(refdb[1])
check.text.args("type",type,c("gene","exon"))
check.text.args("org",org,c("hg18","hg19","hg38","mm9","mm10","rn5","dm3",
"dm6","danrer7","pantro4","susscr3","tair10"),multiarg=FALSE)
check.text.args("refdb",refdb,c("ucsc","refseq"))
if (!require(RSQLite))
stopwrap("R package RSQLite is required to use annotation from UCSC!")
http.base <- paste("http://hgdownload.soe.ucsc.edu/goldenPath/",
get.ucsc.organism(org),"/database/",sep="")
table.defs <- get.ucsc.tabledef(org,type,refdb,"fields")
file.list <- vector("list",length(table.defs))
names(file.list) <- names(table.defs)
for (n in names(file.list))
file.list[[n]] <- paste(http.base,n,".txt.gz",sep="")
# Fill the fields for each table
drv <- dbDriver("SQLite")
db.tmp <- tempfile()
con <- dbConnect(drv,dbname=db.tmp)
#disp(" Defining tables for temporary SQLite ",refdb," ",org," ",
# type," subset database")
#for (n in names(file.list)) {
# disp(" Creating table ",n,"\n")
# dbSendQuery(con,table.defs[[n]])
#}
disp(" Retrieving tables for temporary SQLite ",refdb," ",org," ",type,
" subset database")
for (n in names(file.list)) {
disp(" Retrieving table ",n)
download.file(file.list[[n]],file.path(tempdir(),
paste(n,".txt.gz",sep="")),quiet=TRUE)
if (.Platform$OS.type == "unix")
system(paste("gzip -df",file.path(tempdir(),
paste(n,".txt.gz",sep=""))))
else
unzip(file.path(tempdir(),paste(n,".txt.gz",sep="")))
sql.df <- read.delim(file.path(tempdir(),paste(n,".txt",sep="")),
row.names=NULL,header=FALSE,strip.white=TRUE)
names(sql.df) <- table.defs[[n]]
dbWriteTable(con,n,sql.df,row.names=FALSE)
}
dbDisconnect(con)
return(db.tmp)
}
#' Get SQLite UCSC table defintions, according to organism and source
#'
#' Creates a list of UCSC Genome Browser database tables and their SQLite
#' definitions with the purpose of creating a temporary SQLite database to be
#' used used with metaseqR. This functionality is used when the package RMySQL
#' is not available for some reason, e.g. Windows machines.
#'
#' @param org one of metaseqR supported organisms.
#' @param type either \code{"gene"} or \code{"exon"}.
#' @param refdb one of \code{"ucsc"} or \code{"refseq"} to use the UCSC or RefSeq
#' annotation sources respectively.
#' @return A list with SQLite table definitions.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' db.tabledefs <- get.ucsc.tables("hg18","gene","ucsc")
#'}
get.ucsc.tabledef <- function(org,type,refdb="ucsc",what="queries") {
type <- tolower(type[1])
org <- tolower(org[1])
refdb <- tolower(refdb[1])
what <- tolower(what[1])
check.text.args("type",type,c("gene","exon"))
check.text.args("org",org,c("hg18","hg19","hg38","mm9","mm10","rn5","dm3",
"dm6","danrer7","pantro4","susscr3","tair10"),multiarg=FALSE)
check.text.args("refdb",refdb,c("ucsc","refseq"))
check.text.args("what",what,c("queries","fields"))
switch(type,
gene = {
switch(refdb,
ucsc = {
switch(org,
hg18 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
hg19 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
hg38 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
mm9 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
mm10 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
rn5 = {
return(list(
mgcGenes=get.ucsc.tbl.tpl("mgcGenes",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm3 = {
return(list(
flyBaseCanonical=
get.ucsc.tbl.tpl("flyBaseCanonical",what),
flyBaseGene=
get.ucsc.tbl.tpl("flyBaseGene",what),
flyBaseToRefSeq=
get.ucsc.tbl.tpl("flyBaseToRefSeq",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm6 = {
# Stub until we find out what will happen with
# Augustus
},
danrer7 = {
return(list(
mgcGenes=get.ucsc.tbl.tpl("mgcGenes",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
pantro4 = {
warnwrap("No UCSC Genome annotation for Pan ",
"troglodytes! Will use RefSeq instead...",
now=TRUE)
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
susscr3 = {
warnwrap("No UCSC Genome annotation for Sus ",
"scrofa! Will use RefSeq instead...",
now=TRUE)
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Borwser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
},
refseq = {
switch(org,
hg18 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what)
))
},
hg19 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
hg38 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what)
))
},
mm9 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
mm10 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
rn5 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm3 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
pantro4 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
susscr3 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Borwser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
}
)
},
exon = {
switch(refdb,
ucsc = {
switch(org,
hg18 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
hg19 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
hg38 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
mm9 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
mm10 = {
return(list(
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownGene=get.ucsc.tbl.tpl("knownGene",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what),
refFlat=get.ucsc.tbl.tpl("refFlat",what)
))
},
rn5 = {
return(list(
mgcGenes=get.ucsc.tbl.tpl("mgcGenes",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm3 = {
return(list(
flyBaseCanonical=
get.ucsc.tbl.tpl("flyBaseCanonical",what),
flyBaseGene=
get.ucsc.tbl.tpl("flyBaseGene",what),
flyBaseToRefSeq=
get.ucsc.tbl.tpl("flyBaseToRefSeq",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(list(
mgcGenes=get.ucsc.tbl.tpl("mgcGenes",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
pantro4 = {
warnwrap("No UCSC Genome annotation for Pan ",
"troglodytes! Will use RefSeq instead...",
now=TRUE)
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
susscr3 = {
warnwrap("No UCSC Genome annotation for Sus ",
"scrofa! Will use RefSeq instead...",
now=TRUE)
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Borwser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
},
refseq = {
switch(org,
hg18 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what)
))
},
hg19 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
hg38 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what)
))
},
mm9 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
mm10 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
knownToRefSeq=
get.ucsc.tbl.tpl("knownToRefSeq",what),
knownCanonical=
get.ucsc.tbl.tpl("knownCanonical",what),
knownToEnsembl=
get.ucsc.tbl.tpl("knownToEnsembl",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
rn5 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm3 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
pantro4 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
susscr3 = {
return(list(
refFlat=get.ucsc.tbl.tpl("refFlat",what),
ensemblToGeneName=
get.ucsc.tbl.tpl("ensemblToGeneName",what),
ensemblSource=
get.ucsc.tbl.tpl("ensemblSource",what)
))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Borwser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
}
)
}
)
}
#' Create SQLite UCSC table template defintions
#'
#' Returns an SQLIte table template defintion, according to UCSC Genome Browser
#' database table schemas. This functionality is used when the package RMySQL
#' is not available for some reason, e.g. Windows machines. Internal use only.
#'
#' @param tab name of UCSC database table.
#' @param what \code{"queries"} for SQLite table definitions or \code{"fields"}
#' for table column names.
#' @return An SQLite table definition.
#' @author <NAME>
#' @examples
#' \dontrun{
#' db.table.tmpl <- get.ucsc.tbl.tpl("knownCanonical")
#'}
get.ucsc.tbl.tpl <- function(tab,what="queries") {
if (what=="queries") {
switch(tab,
knownCanonical = {
return(paste(
"CREATE TABLE",
"`knownCanonical` (",
"`chrom` TEXT NOT NULL DEFAULT '',",
"`chromStart` INTEGER NOT NULL DEFAULT '0',",
"`chromEnd` INTEGER NOT NULL DEFAULT '0',",
"`clusterId` INTEGER NOT NULL DEFAULT '0',",
"`transcript` TEXT NOT NULL DEFAULT '',",
"`protein` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
},
knownGene = {
return(paste(
"CREATE TABLE",
"`knownGene` (",
"`name` TEXT NOT NULL DEFAULT '',",
"`chrom` TEXT NOT NULL DEFAULT '',",
"`strand` TEXT NOT NULL DEFAULT '',",
"`txStart` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`txEnd` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`cdsStart` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`cdsEnd` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`exonCount` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`exonStarts` TEXT NOT NULL,",
"`exonEnds` TEXT NOT NULL,",
"`proteinID` TEXT NOT NULL DEFAULT '',",
"`alignID` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
},
knownToRefSeq = {
return(paste(
"CREATE TABLE",
"`knownToRefSeq` (",
"`name` TEXT NOT NULL DEFAULT '',",
"`value` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
},
refFlat = {
return(paste("CREATE TABLE",
"`refFlat` (",
"`geneName` TEXT NOT NULL,",
"`name` TEXT NOT NULL,",
"`chrom` TEXT NOT NULL,",
"`strand` TEXT NOT NULL,",
"`txStart` UNSIGNED INTEGER NOT NULL,",
"`txEnd` UNSIGNED INTEGER NOT NULL,",
"`cdsStart` UNSIGNED INTEGER NOT NULL,",
"`cdsEnd` UNSIGNED INTEGER NOT NULL,",
"`exonCount` UNSIGNED INTEGER NOT NULL,",
"`exonStarts` TEXT NOT NULL,",
"`exonEnds` TEXT NOT NULL",
")",collapse=" "
))
},
knownToEnsembl = {
return(paste(
"CREATE TABLE",
"`knownToEnsembl` (",
"`name` TEXT NOT NULL DEFAULT '',",
"`value` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
},
ensemblSource = {
return(paste(
"CREATE TABLE",
"`ensemblSource` (",
"`name` TEXT NOT NULL DEFAULT '',",
"`source` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
},
mgcGenes = {
return(paste(
"CREATE TABLE `mgcGenes` (",
"`bin` UNSIGNED INTEGER NOT NULL,",
"`name` TEXT NOT NULL,",
"`chrom` TEXT NOT NULL,",
"`strand` TEXT NOT NULL,",
"`txStart` UNSIGNED INTEGER NOT NULL,",
"`txEnd` UNSIGNED INTEGER NOT NULL,",
"`cdsStart` UNSIGNED INTEGER NOT NULL,",
"`cdsEnd` UNSIGNED INTEGER NOT NULL,",
"`exonCount` UNSIGNED INTEGER NOT NULL,",
"`exonStarts` TEXT NOT NULL,",
"`exonEnds` TEXT NOT NULL,",
"`score` INTEGER DEFAULT NULL,",
"`name2` TEXT NOT NULL,",
"`cdsStartStat` TEXT NOT NULL,",
"`cdsEndStat` TEXT NOT NULL,",
"`exonFrames` TEXT NOT NULL",
")",collapse=" "
))
},
ensemblToGeneName = {
return(paste(
"CREATE TABLE",
"`knownToGeneName` (",
"`name` TEXT NOT NULL,",
"`value` TEXT NOT NULL",
")",collapse=" "
))
},
flyBaseCanonical = {
return(paste(
"CREATE TABLE",
"`flyBaseCanonical` (",
"`chrom` TEXT NOT NULL DEFAULT '',",
"`chromStart` INTEGER NOT NULL DEFAULT '0',",
"`chromEnd` INTEGER NOT NULL DEFAULT '0',",
"`clusterId` INTEGER unsigned NOT NULL DEFAULT '0',",
"`transcript` TEXT NOT NULL DEFAULT '',",
"`protein` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
},
flyBaseGene = {
return(paste(
"CREATE TABLE",
"`flyBaseGene` (",
"`bin` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`name` TEXT NOT NULL DEFAULT '',",
"`chrom` TEXT NOT NULL DEFAULT '',",
"`strand` TEXT NOT NULL DEFAULT '',",
"`txStart` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`txEnd` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`cdsStart` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`cdsEnd` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`exonCount` UNSIGNED INTEGER NOT NULL DEFAULT '0',",
"`exonStarts` TEXT NOT NULL,",
"`exonEnds` TEXT NOT NULL",
")",collapse=" "
))
},
flyBaseToRefSeq = {
return(paste(
"CREATE TABLE",
"`flyBaseToRefSeq` (",
"`name` TEXT NOT NULL DEFAULT '',",
"`value` TEXT NOT NULL DEFAULT ''",
")",collapse=" "
))
}
)
}
else if (what=="fields") {
switch(tab,
knownCanonical = {
return(c("chrom","chromStart","chromEnd","clusterId",
"transcript","protein"))
},
knownGene = {
return(c("name","chrom","strand","txStart","txEnd","cdsStart",
"cdsEnd","exonCount","exonStarts","exonEnds","proteinID",
"alignID"))
},
knownToRefSeq = {
return(c("name","value"))
},
refFlat = {
return(c("geneName","name","chrom","strand","txStart","txEnd",
"cdsStart","cdsEnd","exonCount","exonStarts","exonEnds"))
},
knownToEnsembl = {
return(c("name","value"))
},
ensemblSource = {
return(c("name","source"))
},
mgcGenes = {
return(c("name","chrom","strand","txStart","txEnd","cdsStart",
"cdsEnd","exonCount","exonStarts","exonEnds","score",
"name2","cdsStartStat","cdsEndStat","exonFrames"
))
},
ensemblToGeneName = {
return(c("name","value"))
},
flyBaseCanonical = {
return(c("chrom","chromStart","chromEnd","clusterId",
"transcript","protein"))
},
flyBaseGene = {
return(c("bin","name","chrom","strand","txStart","txEnd",
"cdsStart","cdsEnd","exonCount","exonStarts","exonEnds"))
},
flyBaseToRefSeq = {
return(c("name","value"))
}
)
}
}
#' Return queries for the UCSC Genome Browser database, according to organism and
#' source
#'
#' Returns an SQL query to be used with a connection to the UCSC Genome Browser
#' database and fetch metaseqR supported organism annotations. This query is
#' constructed based on the data source and data type to be returned.
#'
#' @param org one of metaseqR supported organisms.
#' @param type either \code{"gene"} or \code{"exon"}.
#' @param refdb one of \code{"ucsc"} or \code{"refseq"} to use the UCSC or RefSeq
#' annotation sources respectively.
#' @return A valid SQL query.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' db.query <- get.ucsc.query("hg18","gene","ucsc")
#'}
get.ucsc.query <- function(org,type,refdb="ucsc") {
type <- tolower(type[1])
org <- tolower(org[1])
refdb <- tolower(refdb[1])
check.text.args("type",type,c("gene","exon"))
check.text.args("org",org,c("hg18","hg19","hg38","mm9","mm10","rn5","dm3",
"dm6","danrer7","pantro4","susscr3","tair10"),multiarg=FALSE)
check.text.args("refdb",refdb,c("ucsc","refseq"))
switch(type,
gene = {
switch(refdb,
ucsc = {
switch(org,
hg18 = {
return(paste("SELECT knownCanonical.chrom AS ",
"`chromosome`,`chromStart` AS `start`,",
"`chromEnd` AS `end`,`transcript` AS ",
"`gene_id`,0 AS `gc_content`,knownGene.strand ",
"AS `strand`,`geneName` AS `gene_name`,'NA' ",
"AS `biotype` FROM `knownCanonical` INNER ",
"JOIN `knownGene` ON ",
"knownCanonical.transcript=knownGene.name ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"`gene_id` ORDER BY `chromosome`, `start`",
sep=""))
},
hg19 = {
return(paste("SELECT knownCanonical.chrom AS ",
"`chromosome`,`chromStart` AS `start`,",
"`chromEnd` AS `end`,`transcript` AS ",
"`gene_id`,0 AS `gc_content`,",
"knownGene.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`knownCanonical` INNER JOIN `knownGene` ON ",
"knownCanonical.transcript=knownGene.name ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"`gene_id` ORDER BY `chromosome`, `start`",
sep=""))
},
hg38 = {
return(paste("SELECT knownCanonical.chrom AS ",
"`chromosome`,`chromStart` AS `start`,",
"`chromEnd` AS `end`,`transcript` AS ",
"`gene_id`,0 AS `gc_content`,knownGene.strand ",
"AS `strand`,`geneName` AS `gene_name`,'NA' ",
"AS `biotype` FROM `knownCanonical` INNER ",
"JOIN `knownGene` ON ",
"knownCanonical.transcript=knownGene.name ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"`gene_id` ORDER BY `chromosome`, `start`",
sep=""))
# Should be the same as hg19 but is like hg18
},
mm9 = {
return(paste("SELECT knownCanonical.chrom AS ",
"`chromosome`,`chromStart` AS `start`,",
"`chromEnd` AS `end`,`transcript` AS ",
"`gene_id`,0 AS `gc_content`,",
"knownGene.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`knownCanonical` INNER JOIN `knownGene` ON ",
"knownCanonical.transcript=knownGene.name ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"`gene_id` ORDER BY `chromosome`, `start`",
sep=""))
},
mm10 = {
return(paste("SELECT knownCanonical.chrom AS ",
"`chromosome`,`chromStart` AS `start`,",
"`chromEnd` AS `end`,`transcript` AS ",
"`gene_id`,0 AS `gc_content`,",
"knownGene.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`knownCanonical` INNER JOIN `knownGene` ON ",
"knownCanonical.transcript=knownGene.name ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"`gene_id` ORDER BY `chromosome`, `start`",
sep=""))
},
rn5 = {
return(paste("SELECT mgcGenes.chrom AS ",
"`chromosome`,`txStart` AS `start`,`txEnd` ",
"AS `end`,mgcGenes.name AS `gene_id`,0 AS ",
"`gc_content`,mgcGenes.strand AS `strand`,",
"`name2` AS `gene_name`,`source` AS `biotype` ",
"FROM `mgcGenes` INNER JOIN ",
"`ensemblToGeneName` ON ",
"mgcGenes.name2=ensemblToGeneName.value INNER ",
"JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm3 = {
return(paste("SELECT flyBaseCanonical.chrom AS ",
"`chromosome`,`chromStart` AS `start`,",
"`chromEnd` AS `end`,`transcript` AS ",
"`gene_id`,0 AS `gc_content`,",
"flyBaseGene.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`flyBaseCanonical` INNER JOIN `flyBaseGene` ",
"ON flyBaseCanonical.transcript=",
"flyBaseGene.name INNER JOIN ",
"`flyBaseToRefSeq` ON ",
"flyBaseCanonical.transcript=",
"flyBaseToRefSeq.name INNER JOIN `refFlat` ON ",
"flyBaseToRefSeq.value=refFlat.name INNER ",
"JOIN `ensemblToGeneName` ON ",
"ensemblToGeneName.value=refFlat.geneName ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(paste("SELECT mgcGenes.chrom AS ",
"`chromosome`,`txStart` AS `start`,`txEnd` ",
"AS `end`,mgcGenes.name AS `gene_id`,0 AS ",
"`gc_content`,mgcGenes.strand AS `strand`,",
"`name2` AS `gene_name`,`source` AS `biotype` ",
"FROM `mgcGenes` INNER JOIN ",
"`ensemblToGeneName` ON ",
"mgcGenes.name2=ensemblToGeneName.value INNER ",
"JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
pantro4 = {
warnwrap("No UCSC Genome annotation for Pan ",
"troglodytes! Will use RefSeq instead...",
now=TRUE)
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
susscr3 = {
warnwrap("No UCSC Genome annotation for Sus ",
"scrofa! Will use RefSeq instead...",
now=TRUE)
return(paste(
"SELECT refFlat.chrom AS `chromosome`,",
"refFlat.txStart AS `start`, refFlat.txEnd AS ",
"`end`, refFlat.name AS `gene_id`, 0 AS ",
"`gc_content`, refFlat.strand AS `strand`,",
"`geneName` AS `gene_name`, `source` AS ",
"`biotype` FROM `refFlat` INNER JOIN ",
"`ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`,",
"`start`",
sep=""
))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Browser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
},
refseq = {
switch(org,
hg18 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,refFlat.strand ",
"AS `strand`,`geneName` AS `gene_name`,'NA' ",
"AS `biotype` FROM `refFlat` INNER JOIN ",
"`knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"GROUP BY refFlat.name ORDER BY `chromosome`,",
" `start`",
sep=""))
},
hg19 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER ",
"JOIN `knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"GROUP BY refFlat.name ORDER BY `chromosome`,",
" `start`",
sep=""))
},
hg38 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,refFlat.strand ",
"AS `strand`,`geneName` AS `gene_name`,'NA' ",
"AS `biotype` FROM `refFlat` INNER JOIN ",
"`knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"GROUP BY refFlat.name ORDER BY `chromosome`,",
" `start`",
sep=""))
# Should be the same as hg19 but is as hg18
},
mm9 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER ",
"JOIN `knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"GROUP BY refFlat.name ORDER BY `chromosome`,",
" `start`",
sep=""))
},
mm10 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER ",
"JOIN `knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"GROUP BY refFlat.name ORDER BY `chromosome`,",
" `start`",
sep=""))
},
rn5 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm3 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,refFlat.strand ",
"AS `strand`,`geneName` AS `gene_name`,",
"`source` AS `biotype` FROM `refFlat` INNER ",
"JOIN `ensemblToGeneName` ON ",
"ensemblToGeneName.value=refFlat.geneName ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
pantro4 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.txStart AS `start`,",
"refFlat.txEnd AS `end`,refFlat.name AS ",
"`gene_id`,0 AS `gc_content`,",
"refFlat.strand AS `strand`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
susscr3 = {
return(paste(
"SELECT refFlat.chrom AS `chromosome`,",
"refFlat.txStart AS `start`, refFlat.txEnd AS ",
"`end`, refFlat.name AS `gene_id`, 0 AS ",
"`gc_content`, refFlat.strand AS `strand`,",
"`geneName` AS `gene_name`, `source` AS ",
"`biotype` FROM `refFlat` INNER JOIN ",
"`ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`,",
"`start`",
sep=""
))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Browser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
}
)
},
exon = {
switch(refdb,
ucsc = {
switch(org,
hg18 = {
return(paste("SELECT knownGene.chrom AS ",
"`chromosome`,knownGene.exonStarts AS `start`,",
"knownGene.exonEnds AS `end`,knownGene.name ",
"AS `exon_id`,knownGene.strand AS `strand`,",
"`transcript` AS `gene_id`,`geneName` AS ",
"`gene_name`,'NA' AS `biotype` FROM ",
"`knownGene` INNER JOIN `knownCanonical` ON ",
"knownGene.name=knownCanonical.transcript ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"knownGene.name ORDER BY `chromosome`, `start`",
sep=""))
},
hg19 = {
return(paste("SELECT knownGene.chrom AS ",
"`chromosome`,knownGene.exonStarts AS `start`,",
"knownGene.exonEnds AS `end`,knownGene.name ",
"AS `exon_id`,knownGene.strand AS `strand`,",
"`transcript` AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`knownGene` INNER JOIN `knownCanonical` ON ",
"knownGene.name=knownCanonical.transcript ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"knownGene.name ORDER BY `chromosome`, `start`",
sep=""))
},
hg38 = {
return(paste("SELECT knownGene.chrom AS ",
"`chromosome`,knownGene.exonStarts AS `start`,",
"knownGene.exonEnds AS `end`,knownGene.name ",
"AS `exon_id`,knownGene.strand AS `strand`,",
"`transcript` AS `gene_id`,`geneName` AS ",
"`gene_name`,'NA' AS `biotype` FROM ",
"`knownGene` INNER JOIN `knownCanonical` ON ",
"knownGene.name=knownCanonical.transcript ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"knownGene.name ORDER BY `chromosome`, `start`",
sep=""))
# Should be the same as hg19 but is as hg18
},
mm9 = {
return(paste("SELECT knownGene.chrom AS ",
"`chromosome`,knownGene.exonStarts AS `start`,",
"knownGene.exonEnds AS `end`,knownGene.name ",
"AS `exon_id`,knownGene.strand AS `strand`,",
"`transcript` AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`knownGene` INNER JOIN `knownCanonical` ON ",
"knownGene.name=knownCanonical.transcript ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"knownGene.name ORDER BY `chromosome`, `start`",
sep=""))
},
mm10 = {
return(paste("SELECT knownGene.chrom AS ",
"`chromosome`,knownGene.exonStarts AS `start`,",
"knownGene.exonEnds AS `end`,knownGene.name ",
"AS `exon_id`,knownGene.strand AS `strand`,",
"`transcript` AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`knownGene` INNER JOIN `knownCanonical` ON ",
"knownGene.name=knownCanonical.transcript ",
"INNER JOIN `knownToRefSeq` ON ",
"knownCanonical.transcript=knownToRefSeq.name ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"INNER JOIN `refFlat` ON ",
"knownToRefSeq.value=refFlat.name GROUP BY ",
"knownGene.name ORDER BY `chromosome`, `start`",
sep=""))
},
rn5 = {
return(paste("SELECT mgcGenes.chrom AS ",
"`chromosome`,`exonStarts` AS `start`,",
"`exonEnds` AS `end`,mgcGenes.name AS ",
"`exon_id`,mgcGenes.strand AS `strand`,",
"mgcGenes.name AS `gene_id`,`name2` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`mgcGenes` INNER JOIN `ensemblToGeneName` ON ",
"mgcGenes.name2=ensemblToGeneName.value INNER ",
"JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm3 = {
return(paste("SELECT flyBaseCanonical.chrom AS ",
"`chromosome`,flyBaseGene.exonStarts AS ",
"`start`,flyBaseGene.exonEnds AS `end`,",
"`transcript` AS `exon_id`,flyBaseGene.strand ",
"AS `strand`,`transcript` AS `gene_id`,",
"`geneName` AS `gene_name`,`source` AS ",
"`biotype` FROM `flyBaseCanonical` INNER JOIN ",
"`flyBaseGene` ON ",
"flyBaseCanonical.transcript=flyBaseGene.name ",
"INNER JOIN `flyBaseToRefSeq` ON ",
"flyBaseCanonical.transcript=",
"flyBaseToRefSeq.name INNER JOIN `refFlat` ON ",
"flyBaseToRefSeq.value=refFlat.name ",
"INNER JOIN `ensemblToGeneName` ON ",
"ensemblToGeneName.value=refFlat.geneName ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(paste("SELECT mgcGenes.chrom AS ",
"`chromosome`,`exonStarts` AS `start`,",
"`exonEnds` AS `end`,mgcGenes.name AS ",
"`exon_id`,mgcGenes.strand AS `strand`,",
"mgcGenes.name AS `gene_id`,`name2` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`mgcGenes` INNER JOIN `ensemblToGeneName` ON ",
"mgcGenes.name2=ensemblToGeneName.value INNER ",
"JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
pantro4 = {
warnwrap("No UCSC Genome annotation for Pan ",
"troglodytes! Will use RefSeq instead...",
now=TRUE)
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
susscr3 = {
warnwrap("No UCSC Genome annotation for Sus ",
"scrofa! Will use RefSeq instead...",
now=TRUE)
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Browser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
},
refseq = {
switch(org,
hg18 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,'NA' AS `biotype` FROM `refFlat` ",
"INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
hg19 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
hg38 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,'NA' AS `biotype` FROM `refFlat` ",
"INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
# Should be the same as hg19 but is as hg18
},
mm9 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
mm10 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `knownToRefSeq` ON ",
"refFlat.name=knownToRefSeq.value INNER JOIN ",
"`knownCanonical` ON ",
"knownToRefSeq.name=knownCanonical.transcript ",
"INNER JOIN `knownToEnsembl` ON ",
"knownCanonical.transcript=knownToEnsembl.name",
" INNER JOIN `ensemblSource` ON ",
"knownToEnsembl.value=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
rn5 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm3 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"ensemblToGeneName.value=refFlat.geneName ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `gene_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
dm6 = {
# Stub until we find out what is going on with
# Augustus
},
danrer7 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
pantro4 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
susscr3 = {
return(paste("SELECT refFlat.chrom AS ",
"`chromosome`,refFlat.exonStarts AS `start`,",
"refFlat.exonEnds AS `end`,refFlat.name AS ",
"`exon_id`,refFlat.strand AS `strand`,",
"refFlat.name AS `gene_id`,`geneName` AS ",
"`gene_name`,`source` AS `biotype` FROM ",
"`refFlat` INNER JOIN `ensemblToGeneName` ON ",
"refFlat.geneName=ensemblToGeneName.value ",
"INNER JOIN `ensemblSource` ON ",
"ensemblToGeneName.name=ensemblSource.name ",
"GROUP BY `exon_id` ORDER BY `chromosome`, ",
"`start`",
sep=""))
},
tair10 = {
warnwrap("Arabidopsis thaliana genome is not ",
"supported by UCSC Genome Browser database! ",
"Will automatically switch to Ensembl...",
now=TRUE)
return(FALSE)
}
)
}
)
}
)
}
#' Return host, username and password for UCSC Genome Browser database
#'
#' Returns a character vector with a hostname, username and password to connect
#' to the UCSC Genome Browser database to retrieve annotation. Internal use.
#'
#' @return A named character vector.
#' @author <NAME>
#' @examples
#' \dontrun{
#' db.creds <- get.ucsc.credentials()
#'}
get.ucsc.credentials <- function() {
return(c(
host="genome-mysql.cse.ucsc.edu",
user="genome",
password=""
))
}
<file_sep>/man/get.ucsc.tabledef.Rd
\name{get.ucsc.tabledef}
\alias{get.ucsc.tabledef}
\title{Get SQLite UCSC table defintions, according to organism and source}
\usage{
get.ucsc.tabledef(org, type, refdb="ucsc", what="queries")
}
\arguments{
\item{org}{one of metaseqR supported organisms.}
\item{type}{either \code{"gene"} or \code{"exon"}.}
\item{refdb}{one of \code{"ucsc"} or \code{"refseq"}
to use the UCSC or RefSeq annotation sources
respectively.}
\item{what}{either \code{"queries"} for SQLite table definitions
or \code{"fields"} for only a vector of table field names.}
}
\value{
A list with SQLite table definitions.
}
\description{
Creates a list of UCSC Genome Browser database tables and
their SQLite definitions with the purpose of creating a
temporary SQLite database to be used used with metaseqR.
This functionality is used when the package RMySQL is not
available for some reason, e.g. Windows machines.
}
\examples{
\donttest{
db.tabledefs <- get.ucsc.tabledef("hg18","gene","ucsc")
}
}
\author{
<NAME>
}
<file_sep>/man/get.ucsc.tbl.tpl.Rd
\name{get.ucsc.tbl.tpl}
\alias{get.ucsc.tbl.tpl}
\title{Create SQLite UCSC table template defintions}
\usage{
get.ucsc.tbl.tpl(tab, what="queries")
}
\arguments{
\item{tab}{name of UCSC database table.}
\item{what}{\code{"queries"} for SQLite table
definitions or \code{"fields"} for table column names.}
}
\value{
An SQLite table definition.
}
\description{
Returns an SQLIte table template defintion,
according to UCSC Genome Browser database
table schemas. This functionality is used
when the package RMySQL is not available
for some reason, e.g. Windows machines.
Internal use only.
}
\examples{
\donttest{
db.table.tmpl <- get.ucsc.tbl.tpl("knownCanonical")
}
}
\author{
<NAME>
}
<file_sep>/man/diagplot.ftd.Rd
\name{diagplot.ftd}
\alias{diagplot.ftd}
\title{Create False (or True) Positive (or
Negative) curves}
\usage{
diagplot.ftd(truth, p, type = "fpc", N = 2000,
output = "x11", path = NULL, draw = TRUE, ...)
}
\arguments{
\item{truth}{the ground truth differential
expression vector. It should contain only
zero and non-zero elements, with zero denoting
non-differentially expressed genes and non-zero,
differentially expressed genes. Such a vector
can be obtained for example by using the
\code{\link{make.sim.data.sd}} function, which
creates simulated RNA-Seq read counts based on
real data. The elements of \code{truth} MUST
be named (e.g. each gene's name).}
\item{p}{a p-value matrix whose rows correspond
to each element in the \code{truth} vector. If
the matrix has a \code{colnames} attribute, a
legend will be added to the plot using these
names, else a set of column names will be
auto-generated. \code{p} can also be a list or
a data frame. The p-values MUST be named (e.g.
each gene's name).}
\item{type}{what to plot, can be \code{"fpc"}
for False Positive Curves (default),
\code{"tpc"} for True Positive Curves,
\code{"fnc"} for False Negative Curves or
\code{"tnc"} for True Negative Curves.}
\item{N}{create the curves based on the
top (or bottom) \code{N} ranked genes
(default is 2000) to be used with
\code{type="fpc"} or \code{type="tpc"}.}
\item{output}{one or more R plotting device to
direct the plot result to. Supported mechanisms:
\code{"x11"} (default), \code{"png"}, \code{"jpg"},
\code{"bmp"}, \code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{draw}{boolean to determine whether to
plot the curves or just return the calculated
values (in cases where the user wants the
output for later averaging for example).
Defaults to \code{TRUE} (make plots).}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
A named list with two members: the first member
(\code{ftdr}) contains the values used to create
the plot. The second member (\code{path}) contains
the path to the created figure graphic.
}
\description{
This function creates false (or true) discovery
curves using a matrix of p-values (such a matrix
can be derived for example from the result table
of \code{\link{metaseqr}} by subsetting the table
to get the p-values from several algorithms)
given a ground truth vector for differential
expression.
}
\examples{
p1 <- 0.001*matrix(runif(300),100,3)
p2 <- matrix(runif(300),100,3)
p <- rbind(p1,p2)
rownames(p) <- paste("gene",1:200,sep="_")
colnames(p) <- paste("method",1:3,sep="_")
truth <- c(rep(1,40),rep(-1,40),rep(0,20),
rep(1,10),rep(2,10),rep(0,80))
names(truth) <- rownames(p)
ftd.obj <- diagplot.ftd(truth,p,N=100)
}
\author{
<NAME>
}
<file_sep>/man/diagplot.metaseqr.Rd
\name{diagplot.metaseqr}
\alias{diagplot.metaseqr}
\title{Diagnostic plots for the metaseqR package}
\usage{
diagplot.metaseqr(object, sample.list, annotation = NULL,
contrast.list = NULL, p.list = NULL,
thresholds = list(p = 0.05, f = 1),
diagplot.type = c("mds", "biodetection", "countsbio", "saturation",
"readnoise", "rnacomp", "correl", "pairs", "boxplot", "gcbias",
"lengthbias", "meandiff", "meanvar", "deheatmap", "volcano",
"biodist", "filtered", "venn"),
is.norm = FALSE, output = "x11", path = NULL, ...)
}
\arguments{
\item{object}{a matrix or a data frame containing count
data derived before or after the normalization procedure,
filtered or not by the metaseqR's filters and/or p-value.
The object can be fed to any of the
\code{diagplot.metaseqr} plotting systems but not every
plot is meaningful. For example, it's meaningless to
create a \code{"biodist"} plot for a count matrix before
normalization or statistical testing.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{annotation}{a data frame containing annotation
elements for each row in object. Usually, a subset of the
annotation obtained by \code{\link{get.annotation}} or a
subset of possibly embedded annotation with the input
counts table. This parameter is optional and required
only when diagplot.type is any of \code{"biodetection"},
\code{"countsbio"}, \code{"saturation"},
\code{"rnacomp"}, \code{"readnoise"}, \code{"biodist"},
\code{"gcbias"}, \code{"lengthbias"} or
\code{"filtered"}.}
\item{contrast.list}{a named structured list of contrasts
as returned by \code{\link{make.contrast.list}} or just
the vector of contrasts as defined in the main help page
of \code{\link{metaseqr}}. This parameter is optional and
required only when \code{diagplot.type} is any of
\code{"deheatmap"}, \code{"volcano"} or
\code{"biodist"}.}
\item{p.list}{a list of p-values for each contrast as
obtained from any of the \code{stat.*} methods of the
metaseqr package. This parameter is optional and required
only when \code{diagplot.type} is any of
\code{"deheatmap"}, \code{"volcano"} or
\code{"biodist"}.}
\item{thresholds}{a list with the elements \code{"p"} and
\code{"f"} which are the p-value and the fold change
cutoff when \code{diagplot.type="volcano"}.}
\item{diagplot.type}{one or more of the diagnostic plots
supported in metaseqR package. Many of these plots
require the presence of additional package, something
that is checked while running the main metaseqr function.
The supported plots are \code{"mds"},
\code{"biodetection"}, \code{"countsbio"},
\code{"saturation"}, \code{"rnacomp"}, \code{"boxplot"},
\code{"gcbias"}, \code{"lengthbias"}, \code{"meandiff"},
\code{"meanvar"}, \code{"deheatmap"}, \code{"volcano"},
\code{"biodist"}, \code{"filtered"}, \code{"readnoise"},
\code{"venn"}, \code{"correl"}, \code{"pairwise"}. For a
brief description of these plots please see the main
\code{\link{metaseqr}} help page.}
\item{is.norm}{a logical indicating whether object
contains raw or normalized data. It is not essential and
it serves only plot annotation purposes.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"png"},
\code{"jpg"}, \code{"bmp"}, \code{"pdf"}, \code{"ps"} or
\code{"json"}. The latter is currently available for the
creation of interactive volcano plots only when reporting
the output, through the highcharts javascript library.
The default plotting (\code{"x11"}) is not supported due
to instability in certain devices.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
A named list containing the file names of the produced
plots. Each list member is names according to the
selected plotting device and is also a named list, whose
names are the plot types. The final contents are the file
names in case the plots are written to a physical
location (not meaningful for \code{"x11"}).
}
\description{
This is the main function for producing sructured quality
control and informative graphs base on the results of the
various steps of the metaseqR package. The graphs
produced span a variety of issues like good sample
reproducibility (Multi-Dimensional Scaling plot, biotype
detection, heatmaps. diagplot.metaseqr, apart from
implementing certain package-specific plots, is a wrapper
around several diagnostic plots present in other RNA-Seq
analysis packages such as EDASeq and NOISeq.
}
\note{
In order to make the best out of this function, you
should generally provide the annotation argument as most
and also the most informative plots depend on this. If
you don't know what is inside your counts table or how
many annotation elements you can provide by embedding it,
it's always best to set the annotation parameter of the
main metaseqr function to \code{"download"} to use
predefined annotations that work better with the
functions of the whole package.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
contrast <- "A_vs_B"
diagplot.metaseqr(data.matrix,sample.list,diagplot.type=c("mds","boxplot"))
norm.args <- get.defaults("normalization","deseq")
object <- normalize.deseq(data.matrix,sample.list,norm.args)
diagplot.metaseqr(object,sample.list,diagplot.type="boxplot")
## More
#p <- stat.deseq(object,sample.list)
#diagplot.metaseqr(object,sample.list,contrast.list=contrast,p.list=p,
# diagplot.type="volcano")
}
\author{
<NAME>
}
<file_sep>/man/make.stat.Rd
\name{make.stat}
\alias{make.stat}
\title{Calculates several statistices on read counts}
\usage{
make.stat(samples, data.list, stat, export.scale)
}
\arguments{
\item{samples}{a set of samples from the dataset under
processing. They should match sample names from
\code{sample.list}. See also the main help page of
\code{\link{metaseqr}}.}
\item{data.list}{a list containing natural or transformed
data, typically an output
from \code{\link{make.transformation}}.}
\item{stat}{the statistics to calculate. Can be one or
more of \code{"mean"}, \code{"median"}, \code{"sd"},
\code{"mad"}, \code{"cv"}, \code{"rcv"}. See also the
main help page of \code{\link{metaseqr}}.}
\item{export.scale}{the output transformations used as
input also to \code{\link{make.transformation}}.}
}
\value{
A matrix of statistics calculated based on the input
sample names. The different data transformnations are
appended columnwise.
}
\description{
Returns a matrix of statistics calculated for a set of
given samples. Internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
tr <- make.transformation(data.matrix,c("log2","vst"))
st <- make.stat(c("C1","C2"),tr,c("mean","sd"),c("log2","vst"))
}
}
\author{
<NAME>
}
<file_sep>/man/make.path.struct.Rd
\name{make.path.struct}
\alias{make.path.struct}
\title{Project path constructor helper}
\usage{
make.path.struct(main.path)
}
\arguments{
\item{main.path}{The desired project path.}
}
\value{
A named list whose names are the conditions of the
experiments and its members are the samples belonging to
each condition.
}
\description{
Helper for \code{make.project.path}. Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/get.gc.content.Rd
\name{get.gc.content}
\alias{get.gc.content}
\title{Return a named vector of GC-content for each genomic
region}
\usage{
get.gc.content(ann, org)
}
\arguments{
\item{ann}{a data frame which can be converted to a
GRanges object, that means it has at least the
chromosome, start, end fields. Preferably, the output
of \code{link{get.ucsc.annotation}}.}
\item{org}{one of metaseqR supported organisms.}
}
\value{
A named numeric vector.
}
\description{
Returns a named numeric vector (names are the
genomic region names, e.g. genes) given a data
frame which can be converted to a GRanges object
(e.g. it has at least chromosome, start, end
fields). This function works best when the input
annotation data frame has been retrieved using one
of the SQL queries generated from
\code{\link{get.ucsc.query}}, used in
\code{\link{get.ucsc.annotation}}.
}
\examples{
\donttest{
ann <- get.ucsc.annotation("mm9","gene","ucsc")
gc <- get.gc.content(ann,"mm9")
}
}
\author{
<NAME>
}
<file_sep>/man/normalize.edger.Rd
\name{normalize.edger}
\alias{normalize.edger}
\title{Normalization based on the edgeR package}
\usage{
normalize.edger(gene.counts, sample.list,
norm.args = NULL, output = c("matrix", "native"))
}
\arguments{
\item{gene.counts}{a table where each row represents a
gene and each column a sample. Each cell contains the
read counts for each gene and sample. Such a table can be
produced outside metaseqr and is imported during the
basic metaseqr workflow.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{norm.args}{a list of edgeR normalization
parameters. See the result of
\code{get.defaults("normalization",} \code{"edger")} for
an example and how you can modify it.}
\item{output}{the class of the output object. It can be
\code{"matrix"} (default) for versatility with other
tools or \code{"native"} for the edgeR native S4 object
(DGEList). In the latter case it should be handled with
suitable edgeR methods.}
}
\value{
A matrix or a DGEList with normalized counts.
}
\description{
This function is a wrapper over edgeR normalization. It
accepts a matrix of gene counts (e.g. produced by
importing an externally generated table of counts to the
main metaseqr pipeline).
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
diagplot.boxplot(data.matrix,sample.list)
norm.data.matrix <- normalize.edger(data.matrix,sample.list)
diagplot.boxplot(norm.data.matrix,sample.list)
}
}
\author{
<NAME>
}
<file_sep>/man/libsize.list.hg19.Rd
\docType{data}
\name{libsize.list.hg19}
\alias{libsize.list.hg19}
\title{Human RNA-Seq data with three conditions, three samples}
\format{a named \code{list} with library sizes.}
\source{
GEO (http://www.ncbi.nlm.nih.gov/geo/)
}
\description{
The library size list for \code{hg19.exon.counts}. See
the data set description.
}
\author{
<NAME>
}
\keyword{datasets}
<file_sep>/README.md
# metaseqR
An R package for the analysis, meta-analysis and result reporting of RNA-Seq gene expression data
# metaseqR deprecation
**Important**: metaseqR has been deprecated an it no longer exists in the current Bioconductor devel repository. Please switch to [metaseqR2](https://github.com/pmoulos/metaseqR2),
the next generation of the package with many more features, interfaces to more statistical testing and normalization algorithms, 5-50x speed-up and a completely redesigned and
fully interactive report!
<file_sep>/man/diagplot.mds.Rd
\name{diagplot.mds}
\alias{diagplot.mds}
\title{Multi-Dimensinal Scale plots or RNA-Seq samples}
\usage{
diagplot.mds(x, sample.list, method = "spearman",
log.it = TRUE, output = "x11", path = NULL, ...)
}
\arguments{
\item{x}{the count data matrix.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{method}{which correlation method to use. Same as
the method parameter in \code{\link{cor}} function.}
\item{log.it}{whether to log transform the values of x or
not.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"}, \code{"ps"} or \code{"json"}. The latter is
currently available for the creation of interactive
volcano plots only when reporting the output, through the
highcharts javascript library.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filename of the MDS plot produced if it's a file.
}
\description{
Creates a Multi-Dimensional Scale plot for the given
samples based on the count data matrix. MDS plots are
very useful for quality control as you can easily see of
samples of the same groups are clustered together based
on the whole dataset.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
diagplot.mds(data.matrix,sample.list)
}
\author{
<NAME>
}
<file_sep>/man/get.defaults.Rd
\name{get.defaults}
\alias{get.defaults}
\title{Default parameters for several metaseqr functions}
\usage{
get.defaults(what, method = NULL)
}
\arguments{
\item{what}{a keyword determining the procedure for which
to fetch the default settings according to method
parameter. It can be one of \code{"normalization"},
\code{"statistics"}, \code{"gene.filter"},
\code{"exon.filter"} or \code{"biotype.filter"}.}
\item{method}{the supported algorithm included in
metaseqR for which to fetch the default settings. When
\code{what} is \code{"normalization"}, method is one of
\code{"edaseq"}, \code{"deseq"}, \code{"edger"},
\code{"noiseq"} or \code{"nbpseq"}. When \code{what} is
\code{"statistics"}, method is one of \code{"deseq"},
\code{"edger"}, \code{"noiseq"}, \code{"bayseq"},
\code{"limma"} or \code{"nbpseq"}. When \code{method} is
\code{"biotype.filter"}, \code{what} is the input
organism (see the main \code{\link{metaseqr}} help page
for a list of supported organisms).}
}
\value{
A list with default setting that can be used directly in
the call of metaseqr.
}
\description{
This function returns a list with the default settings
for each filtering, statistical and normalization
algorithm included in the metaseqR package. See the
documentation of the main function and the documentation
of each statistical and normalization method for details.
}
\examples{
norm.args.edaseq <- get.defaults("normalization","edaseq")
stat.args.edger <- get.defaults("statistics","edger")
}
\author{
<NAME>
}
<file_sep>/man/make.html.header.Rd
\name{make.html.header}
\alias{make.html.header}
\title{HTML report helper}
\usage{
make.html.header(h)
}
\arguments{
\item{h}{the colnames of a matrix or data frame, usually
as output of \code{\link{make.html.cells}} function.}
}
\value{
A character vector with html formatted header of a
matrix.
}
\description{
Returns a character vector with an html formatted table
head row. Essentially, it collapses the input row to a
single character and puts a <th></th> tag set around. It
is meant to be applied to the output of
\code{\link{make.html.cells}}. Internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
the.cells <- make.html.cells(data.matrix)
the.header <- make.html.header(the.cells[1,])
}
}
\author{
<NAME>
}
<file_sep>/man/diagplot.edaseq.Rd
\name{diagplot.edaseq}
\alias{diagplot.edaseq}
\title{Diagnostic plots based on the EDASeq package}
\usage{
diagplot.edaseq(x, sample.list, covar = NULL,
is.norm = FALSE,
which.plot = c("meanvar", "meandiff", "gcbias", "lengthbias"),
output = "x11", path = NULL, ...)
}
\arguments{
\item{x}{the count data matrix.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{covar}{The covariate to plot counts against.
Usually \code{"gc"} or \code{"length"}.}
\item{is.norm}{a logical indicating whether object
contains raw or normalized data. It is not essential and
it serves only plot annotation purposes.}
\item{which.plot}{the EDASeq package plot to generate. It
can be one or more of \code{"meanvar"},
\code{"meandiff"}, \code{"gcbias"} or
\code{"lengthbias"}. Please refer to the documentation of
the EDASeq package for details on the use of these
plots. The \code{which.plot="lengthbias"} case is
not covered by EDASeq documentation, however it is
similar to the GC-bias plot when the covariate is the
gene length instead of the GC content.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filenames of the plot produced in a named list with
names the which.plot argument. If \code{output="x11"}, no
output filenames are produced.
}
\description{
A wrapper around the plotting functions availale in the
EDASeq normalization Bioconductor package. For analytical
explanation of each plot please see the vignette of the
EDASeq package. It is best to use this function through
the main plotting function
\code{\link{diagplot.metaseqr}}.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
diagplot.edaseq(data.matrix,sample.list,which.plot="meandiff")
}
\author{
<NAME>
}
<file_sep>/man/validate.list.args.Rd
\name{validate.list.args}
\alias{validate.list.args}
\title{Validate list parameters for several metaseqR functions}
\usage{
validate.list.args(what, method = NULL, arg.list)
}
\arguments{
\item{what}{what a keyword determining the procedure
for which to validate arguments. It can be one of
\code{"normalization"}, \code{"statistics"},
\code{"gene.filter"}, \code{"exon.filter"} or
\code{"biotype.filter"}.}
\item{method}{the normalization/statistics/filtering
algorithm included in metaseqR for which to validate
user input. When \code{what} is \code{"normalization"},
method is one of \code{"edaseq"}, \code{"deseq"},
\code{"edger"}, \code{"noiseq"} or \code{"nbpseq"}.
When \code{what} is \code{"statistics"}, method is
one of \code{"deseq"}, \code{"edger"}, \code{"noiseq"},
\code{"bayseq"}, \code{"limma"} or \code{"nbpseq"}.
When \code{method} is \code{"biotype.filter"},
\code{what} is the input organism (see the main
\code{\link{metaseqr}} help page for a list of
supported organisms).}
\item{arg.list}{the user input list of arguments.}
}
\value{
A list with valid arguments to be used as user
input in the algorithms supported by metaseqR.
}
\description{
This function validates the arguments passed
by the user to the normalization, statistics
and filtering algorithms supported by metaseqR.
As these are given into lists and passed to
the algorithms, the list member names must
be valid algorithm arguments for the pipeline
not to crash. This function performs these
checks and ignores any invalid arguments.
}
\examples{
\donttest{
norm.args.edger <- list(method="TMM",refColumn=NULL,
logratioTrim=0.3,sumTrim=0.05,doWeighting=TRUE,
Bcutoff=-1e10,p=0.75)
# Bcutoff does not exist, will throw a warning and ignore it.
norm.args.edger <- validate.list.args("normalization",
"edger",norm.args.edger)
}
}
\author{
<NAME>
}
<file_sep>/man/wp.adjust.Rd
\name{wp.adjust}
\alias{wp.adjust}
\title{Multiple testing correction helper}
\usage{
wp.adjust(p, m)
}
\arguments{
\item{p}{a vector of p-values.}
\item{m}{the adjustment method. See the help of
\code{\link{p.adjust}}.}
}
\description{
A wrapper around the \code{\link{p.adjust}} function to
include also the qvalue adjustment procedure from the
qvalue package. Internal use.
}
\author{
<NAME>
}
<file_sep>/man/diagplot.noiseq.Rd
\name{diagplot.noiseq}
\alias{diagplot.noiseq}
\title{Diagnostic plots based on the NOISeq package}
\usage{
diagplot.noiseq(x, sample.list, covars,
which.plot = c("biodetection", "countsbio", "saturation", "rnacomp",
"readnoise", "biodist"),
output = "x11",
biodist.opts = list(p = NULL, pcut = NULL, name = NULL),
path = NULL, is.norm = FALSE, ...)
}
\arguments{
\item{x}{the count data matrix.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{covars}{a list (whose annotation elements are
ideally a subset of an annotation data frame produced by
\code{\link{get.annotation}}) with the following members:
data (the data matrix), length (gene length), gc (the
gene gc_content), chromosome (a data frame with
chromosome name and co-ordinates), factors (a factor with
the experimental condition names replicated by the number
of samples in each experimental condition) and biotype
(each gene's biotype as depicted in Ensembl-like
annotations).}
\item{which.plot}{the NOISeq package plot to generate. It
can be one or more of \code{"biodetection"},
\code{"countsbio"}, \code{"saturation"},
\code{"rnacomp"}, \code{"readnoise"} or \code{"biodist"}.
Please refer to the documentation of the EDASeq package
for details on the use of these plots. The
\code{which.plot="saturation"} case is modified to be
more informative by producing two kinds of plots. See
\code{\link{diagplot.noiseq.saturation}}.}
\item{biodist.opts}{a list with the following members: p
(a vector of p-values, e.g. the p-values of a contrast),
pcut (a unique number depicting a p-value cutoff,
required for the \code{"biodist"} case), name (a name for
the \code{"biodist"} plot, e.g. the name of the
contrast.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"} or \code{"ps"}.}
\item{path}{the path to create output files.}
\item{is.norm}{a logical indicating whether object
contains raw or normalized data. It is not essential and
it serves only plot annotation purposes.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filenames of the plots produced in a named list with
names the \code{which.plot} argument. If
\code{output="x11"}, no output filenames are produced.
}
\description{
A wrapper around the plotting functions availale in the
NOISeq Bioconductor package. For analytical explanation
of each plot please see the vignette of the NOISeq
package. It is best to use this function through the
main plotting function \code{\link{diagplot.metaseqr}}.
}
\note{
Please note that in case of \code{"biodist"} plots, the
behavior of the function is unstable, mostly due to the
very specific inputs this plotting function accepts in
the NOISeq package. We have tried to predict unstable
behavior and avoid exceptions through the use of tryCatch
but it's still possible that you might run onto an error.
}
\examples{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
lengths <- round(1000*runif(nrow(data.matrix)))
starts <- round(1000*runif(nrow(data.matrix)))
ends <- starts + lengths
covars <- list(
data=data.matrix,
length=lengths,
gc=runif(nrow(data.matrix)),
chromosome=data.frame(
chromosome=c(rep("chr1",nrow(data.matrix)/2),
rep("chr2",nrow(data.matrix)/2)),
start=starts,
end=ends
),
factors=data.frame(class=as.class.vector(sample.list)),
biotype=c(rep("protein_coding",nrow(data.matrix)/2),rep("ncRNA",
nrow(data.matrix)/2))
)
p <- runif(nrow(data.matrix))
diagplot.noiseq(data.matrix,sample.list,covars=covars,
biodist.opts=list(p=p,pcut=0.1,name="A_vs_B"))
}
\author{
<NAME>
}
<file_sep>/R/metaseqR-package.R
#' An R package for the analysis and result reporting of RNA-Seq gene expression
#' data, using multiple statistical algorithms.
#'
#' \tabular{ll}{
#' Package: \tab metaseqR\cr
#' Type: \tab Package\cr
#' Version: \tab 1.9.21\cr
#' Date: \tab 2015-10-08\cr
#' Depends: \tab R (>= 2.13.0), Biobase, BiocGenerics, rjson, biomaRt, utils,
#' knitr, EDASeq, DESeq, edgeR, limma, NOISeq, baySeq, NBPSeq, survcomp,
#' brew, gplots, corrplot, GenomicRanges, Rsamtools, rtracklayer, Repitools,
#' qvalue, vsn, VennDiagram, log4r\cr
#' Encoding: \tab UTF-8\cr
#' License: \tab GPL (>= 3)\cr
#' LazyLoad: \tab yes\cr
#' URL: \tab http://www.fleming.gr\cr
#' }
#'
#' Provides an interface to several normalization and statistical testing packages
#' for RNA-Seq gene expression data. Additionally, it creates several diagnostic
#' plots, performs meta-analysis by combinining the results of several statistical
#' tests and reports the results in an interactive way.
#'
#' @aliases metaseqR-package metaseqR
#' @name metaseqR-package
#' @docType package
#' @title The metaseqR Package
#' @author <NAME> \email{<EMAIL>}
#' @keywords package
NULL
<file_sep>/man/filter.genes.Rd
\name{filter.genes}
\alias{filter.genes}
\title{Filter gene expression based on gene counts}
\usage{
filter.genes(gene.counts, gene.data, gene.filters,
sample.list)
}
\arguments{
\item{gene.counts}{a matrix of gene counts, preferably
after the normalization procedure.}
\item{gene.data}{an annotation data frame usually
obtained with \code{\link{get.annotation}} containing the
unique gene accession identifiers.}
\item{gene.filters}{a named list with gene filters and
their parameters. See the main help page of
\code{\link{metaseqr}} for details.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
}
\value{
a named list with three members. The first member
(\code{result} is a named list whose names are the
gene filter names and its members are the filtered
rownames of \code{gene.data}. The second member
(\code{cutoff} is a named list whose names are the
gene filter names and its members are the cutoff
values corresponding to each filter. The third member
is a matrix of binary flags (0 for non-filtered, 1
for filtered) for each gene. The rownames of the
flag matrix correspond to gene ids.
}
\description{
This function performs the gene expression filtering
based on gene read counts and a set of gene filter rules.
For more details see the main help pages of
\code{\link{metaseqr}}.
}
\examples{
\donttest{
data("mm9.gene.data",package="metaseqR")
gene.counts <- mm9.gene.counts
sample.list <- sample.list.mm9
gene.counts <- normalize.edger(as.matrix(gene.counts[,9:12]),
sample.list)
gene.data <- get.annotation("mm9","gene")
gene.filters <- get.defaults("gene.filter","mm9")
filter.results <- filter.genes(gene.counts,gene.data,
gene.filters,sample.list)
}
}
\author{
<NAME>
}
<file_sep>/man/make.permutation.Rd
\name{make.permutation}
\alias{make.permutation}
\title{Create counts matrix permutations}
\usage{
make.permutation(counts, sample.list, contrast,
repl = FALSE)
}
\arguments{
\item{counts}{the gene read counts matrix.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{contrast}{the contrasts vector. See the main
\code{\link{metaseqr}} help page.}
\item{repl}{the same as the replace argument in
\code{\link{sample}} function.}
}
\value{
A list with three members: the matrix of permuted per
sample read counts, the virtual sample list and the
virtual contrast to be used with the \code{stat.*}
functions.
}
\description{
This function creates a permuted read counts matrix based
on the \code{contrast} argument (to define new virtual
contrasts of the same number) and on the
\code{sample.list} to derive the number of samples for
each virtual condition.It is a helper for the
\code{\link{meta.perm}} function.
}
\examples{
\donttest{
data("mm9.gene.data",package="metaseqR")
per <- make.permutation(mm9.gene.counts,sample.list.mm9,
"e14.5_vs_adult_8_weeks")
}
}
\author{
<NAME>
}
<file_sep>/man/cdplot.Rd
\name{cdplot}
\alias{cdplot}
\title{Old functions from NOISeq}
\usage{
cdplot(dat, samples = NULL, ...)
}
\arguments{
\item{dat}{the returned list from \code{\link{cddat}}.}
\item{samples}{the samples to plot.}
\item{...}{further arguments passed to e.g.
\code{\link{par}}.}
}
\value{
Nothing, it created the old RNA composition plot.
}
\description{
Old functions from NOISeq to create the
\code{"readnoise"} plots. Internal use only.
}
\note{
Adopted from an older version of NOISeq package (author:
<NAME>)
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.stat.R
#' Statistical testing with DESeq
#'
#' This function is a wrapper over DESeq statistical testing. It accepts a matrix
#' of normalized gene counts or an S4 object specific to each normalization
#' algorithm supported by metaseqR.
#'
#' @param object a matrix or an object specific to each normalization algorithm
#' supported by metaseqR, containing normalized counts. Apart from matrix (also
#' for NOISeq), the object can be a SeqExpressionSet (EDASeq), CountDataSet (DESeq)
#' or DGEList (edgeR).
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined
#' in the main help page of \code{\link{metaseqr}}.
#' @param stat.args a list of DESeq statistical algorithm parameters. See the
#' result of \code{get.defaults("statistics",} \code{"deseq")} for an example and
#' how you can modify it. It is not required when the input object is already a
#' CountDataSet from DESeq normalization
#' as the dispersions are already estimated.
#' @return A named list of p-values, whose names are the names of the contrasts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' norm.data.matrix <- normalize.deseq(data.matrix,sample.list)
#' p <- stat.deseq(norm.data.matrix,sample.list,contrast)
#'}
stat.deseq <- function(object,sample.list,contrast.list=NULL,stat.args=NULL) {
#if (is.null(norm.args) && class(object)=="DGEList")
# norm.args <- get.defaults("normalization","edger")
if (is.null(stat.args) && !is(object,"CountDataSet"))
stat.args <- get.defaults("statistics","deseq")
if (is.null(contrast.list))
contrast.list <- make.contrast.list(paste(names(sample.list)[1:2],
collapse="_vs_"),sample.list)
if (!is.list(contrast.list))
contrast.list <- make.contrast.list(contrast.list,sample.list)
classes <- as.class.vector(sample.list)
the.design <- data.frame(condition=classes,row.names=colnames(object))
p <- vector("list",length(contrast.list))
names(p) <- names(contrast.list)
# Check if there is no replication anywhere
if (all(sapply(sample.list,function(x) ifelse(length(x)==1,TRUE,
FALSE)))) {
warnwrap("No replication detected! There is a possibility that ",
"DESeq will fail to estimate dispersions...")
method.disp <- "blind"
sharingMode.disp <- "fit-only"
fitType.disp <- "local"
}
else {
method.disp <- stat.args$method
sharingMode.disp <- stat.args$sharingMode
fitType.disp <- stat.args$fitType
}
switch(class(object)[1],
CountDataSet = { # Has been normalized with DESeq
cds <- object
cds <- estimateDispersions(cds,method=method.disp,
sharingMode=sharingMode.disp,fitType=fitType.disp)
},
DGEList = { # Has been normalized with edgeR
# Trick found at http://cgrlucb.wikispaces.com/edgeR+spring2013
scl <- object$samples$lib.size * object$samples$norm.factors
cds <- newCountDataSet(round(t(t(object$counts)/scl)*mean(scl)),
the.design$condition)
sizeFactors(cds) <- rep(1,ncol(cds))
cds <- estimateDispersions(cds,method=method.disp,
sharingMode=sharingMode.disp)
},
matrix = { # Has been normalized with EDASeq or NOISeq
cds <- newCountDataSet(object,the.design$condition)
sizeFactors(cds) <- rep(1,ncol(cds))
cds <- estimateDispersions(cds,method=method.disp,
sharingMode=sharingMode.disp)
},
list = { # Has been normalized with NBPSeq and main method was "nbpseq"
cds <- newCountDataSet(as.matrix(round(sweep(object$counts,2,
object$norm.factors,"*"))),the.design$condition)
sizeFactors(cds) <- rep(1,ncol(cds))
cds <- estimateDispersions(cds,method=method.disp,
sharingMode=sharingMode.disp)
},
nbp = { # Has been normalized with NBPSeq and main method was "nbsmyth"...
cds <- newCountDataSet(as.matrix(round(object$pseudo.counts)),
the.design$condition)
sizeFactors(cds) <- rep(1,ncol(cds))
cds <- estimateDispersions(cds,method=method.disp,
sharingMode=sharingMode.disp)
}
)
for (con.name in names(contrast.list)) {
disp(" Contrast: ", con.name)
con <- contrast.list[[con.name]]
cons <- unique(unlist(con))
if (length(con)==2) {
res <- nbinomTest(cds,cons[1],cons[2])
p[[con.name]] <- res$pval
}
else {
#cind <- match(cons,the.design$condition)
#if (any(is.na(cind)))
# cind <- cind[which(!is.na(cind))]
cc <- names(unlist(con))
cds.tmp <- cds[,cc]
fit0 <- fitNbinomGLMs(cds.tmp,count~1)
fit1 <- fitNbinomGLMs(cds.tmp,count~condition)
p[[con.name]] <- nbinomGLMTest(fit1,fit0)
}
names(p[[con.name]]) <- rownames(object)
p[[con.name]][which(is.na(p[[con.name]]))] <- 1
}
return(p)
}
#' Statistical testing with edgeR
#'
#' This function is a wrapper over edgeR statistical testing. It accepts a matrix
#' of normalized gene counts or an S4 object specific to each normalization
#' algorithm supported by metaseqR.
#'
#' @param object a matrix or an object specific to each normalization algorithm
#' supported by metaseqr, containing normalized counts. Apart from matrix (also
#' for NOISeq), the object can be a SeqExpressionSet (EDASeq), CountDataSet (DESeq)
#' or DGEList (edgeR).
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined in
#' the main help page of \code{\link{metaseqr}}.
#' @param stat.args a list of edgeR statistical algorithm parameters. See the
#' result of \code{get.defaults("statistics",} \code{"edger")} for an example and
#' how you can modify it.
#' @return A named list of p-values, whose names are the names of the contrasts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' norm.data.matrix <- normalize.edger(data.matrix,sample.list)
#' p <- stat.edger(norm.data.matrix,sample.list,contrast)
#'}
stat.edger <- function(object,sample.list,contrast.list=NULL,stat.args=NULL) {
if (is.null(stat.args))
stat.args <- get.defaults("statistics","edger")
if (is.null(contrast.list))
contrast.list <- make.contrast.list(paste(names(sample.list)[1:2],
collapse="_vs_"),sample.list)
if (!is.list(contrast.list))
contrast.list <- make.contrast.list(contrast.list,sample.list)
classes <- as.class.vector(sample.list)
p <- vector("list",length(contrast.list))
names(p) <- names(contrast.list)
switch(class(object)[1],
CountDataSet = { # Has been normalized with DESeq
dge <- DGEList(counts=counts(object,normalized=TRUE),group=classes)
},
DGEList = { # Has been normalized with edgeR
dge <- object
},
matrix = { # Has been normalized with EDASeq or NOISeq
dge <- DGEList(object,group=classes)
},
list = { # Has been normalized with NBPSeq and main method was "nbpseq"
dge <- DGEList(counts=as.matrix(round(sweep(object$counts,2,
object$norm.factors,"*"))),group=classes)
},
nbp = { # Has been normalized with NBPSeq and main method was "nbsmyth"
dge <- DGEList(counts=as.matrix(round(object$pseudo.counts)),
group=classes)
}
)
# Dispersion estimate step
# Check if there is no replication anywhere
repli = TRUE
if (all(sapply(sample.list,function(x) ifelse(length(x)==1,TRUE,
FALSE)))) {
warnwrap("No replication when testing with edgeR! Consider using ",
"another statistical test or just performing empirical analysis. ",
"Setting to 0.2...")
repli <- FALSE
bcv <- 0.2
}
if (repli) {
if (stat.args$main.method=="classic") {
dge <- estimateCommonDisp(dge,rowsum.filter=stat.args$rowsum.filter)
dge <- estimateTagwiseDisp(dge,prior.df=stat.args$prior.df,
trend=stat.args$trend,span=stat.args$span,
method=stat.args$tag.method,
grid.length=stat.args$grid.length,
grid.range=stat.args$grid.range)
}
else if (stat.args$main.method=="glm") {
design <- model.matrix(~0+classes,data=dge$samples)
dge <- estimateGLMCommonDisp(dge,design=design,
offset=stat.args$offset,
method=stat.args$glm.method,subset=stat.args$subset,
AveLogCPM=stat.args$AveLogCPM)
dge <- estimateGLMTrendedDisp(dge,design=design,
offset=stat.args$offset,
method=stat.args$trend.method,AveLogCPM=stat.args$AveLogCPM)
dge <- estimateGLMTagwiseDisp(dge,design=design,
offset=stat.args$offset,
dispersion=stat.args$dispersion,prior.df=stat.args$prior.df,
span=stat.args$span,AveLogCPM=stat.args$AveLogCPM)
}
}
# Actual statistical test
for (con.name in names(contrast.list))
{
disp(" Contrast: ", con.name)
con <- contrast.list[[con.name]]
if (length(con)==2) {
if (repli) {
if (stat.args$main.method=="classic") {
res <- exactTest(dge,pair=unique(unlist(con)))
}
else if (stat.args$main.method=="glm") {
s <- unlist(con)
us <- unique(s)
ms <- match(names(s),rownames(dge$samples))
if (any(is.na(ms)))
ms <- ms[which(!is.na(ms))]
design <- model.matrix(~0+s,data=dge$samples[ms,])
colnames(design) <- us
#fit <- glmFit(dge[,ms],design=design,
# offset=stat.args$offset,
# weights=stat.args$weights,lib.size=stat.args$lib.size,
# prior.count=stat.args$prior.count,
# start=stat.args$start,method=stat.args$method)
fit <- glmFit(dge[,ms],design=design,
prior.count=stat.args$prior.count,
start=stat.args$start,method=stat.args$method)
co <- makeContrasts(paste(us[2],us[1],sep="-"),
levels=design)
lrt <- glmLRT(fit,contrast=co)
res <- topTags(lrt,n=nrow(dge))
}
}
else {
if (stat.args$main.method=="classic") {
res <- exactTest(dge,pair=unique(unlist(con)),
dispersion=bcv^2)
}
else if (stat.args$main.method=="glm") {
s <- unlist(con)
us <- unique(s)
ms <- match(names(s),rownames(dge$samples))
if (any(is.na(ms)))
ms <- ms[which(!is.na(ms))]
design <- model.matrix(~0+s,data=dge$samples[ms,])
colnames(design) <- us
#fit <- glmFit(dge[,ms],design=design,
# offset=stat.args$offset,weights=stat.args$weights,
# lib.size=stat.args$lib.size,
# prior.count=stat.args$prior.count,
# start=stat.args$start,
# method=stat.args$method,dispersion=bcv^2)
fit <- glmFit(dge[,ms],design=design,dispersion=bcv^2,
prior.count=stat.args$prior.count,start=stat.args$start,
method=stat.args$method)
co <- makeContrasts(paste(us[2],us[1],sep="-"),
levels=design)
lrt <- glmLRT(fit,contrast=co)
res <- topTags(lrt,n=nrow(dge))
}
}
}
else { # GLM only
s <- unlist(con)
us <- unique(s)
#design <- model.matrix(~0+s,data=dge$samples) # Ouch!
ms <- match(names(s),rownames(dge$samples))
if (any(is.na(ms)))
ms <- ms[which(!is.na(ms))]
design <- model.matrix(~s,data=dge$samples[ms,])
if (repli)
#fit <- glmFit(dge[,ms],design=design,offset=stat.args$offset,
# weights=stat.args$weights,lib.size=stat.args$lib.size,
# prior.count=stat.args$prior.count,start=stat.args$start,
# method=stat.args$method)
fit <- glmFit(dge[,ms],design=design,start=stat.args$start,
prior.count=stat.args$prior.count)
else
#fit <- glmFit(dge[,ms],design=design,offset=stat.args$offset,
# weights=stat.args$weights,lib.size=stat.args$lib.size,
# prior.count=stat.args$prior.count,start=stat.args$start,
# method=stat.args$method,dispersion=bcv^2)
# dispersion=bcv^2)
fit <- glmFit(dge[,ms],design=design,dispersion=bcv^2,
prior.count=stat.args$prior.count,start=stat.args$start)
#lrt <- glmLRT(fit,coef=2:ncol(fit$design))
lrt <- glmLRT(fit,coef=2:ncol(fit$design))
res <- topTags(lrt,n=nrow(dge))
}
p[[con.name]] <- res$table[,"PValue"]
names(p[[con.name]]) <- rownames(res$table)
p[[con.name]] <- p[[con.name]][rownames(dge)]
p[[con.name]][which(is.na(p[[con.name]]))] <- 1
}
return(p)
}
#' Statistical testing with limma
#'
#' This function is a wrapper over limma statistical testing. It accepts a matrix
#' of normalized gene counts or an S4 object specific to each normalization
#' algorithm supported by metaseqR.
#'
#' @param object a matrix or an object specific to each normalization algorithm
#' supported by metaseqr, containing normalized counts. Apart from matrix (also
#' for NOISeq), the object can be a SeqExpressionSet (EDASeq), CountDataSet (DESeq)
#' or DGEList (edgeR).
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined in
#' the main help page of \code{\link{metaseqr}}.
#' @param stat.args a list of edgeR statistical algorithm parameters. See the
#' result of \code{get.defaults("statistics",} \code{"limma")} for an example and
#' how you can modify it.
#' @return A named list of p-values, whose names are the names of the contrasts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' norm.data.matrix <- normalize.edger(data.matrix,sample.list)
#' p <- stat.limma(norm.data.matrix,sample.list,contrast)
#'}
stat.limma <- function(object,sample.list,contrast.list=NULL,stat.args=NULL) {
if (is.null(stat.args))
stat.args <- get.defaults("statistics","limma")
if (is.null(contrast.list))
contrast.list <- make.contrast.list(paste(names(sample.list)[1:2],
collapse="_vs_"),sample.list)
if (!is.list(contrast.list))
contrast.list <- make.contrast.list(contrast.list,sample.list)
classes <- as.class.vector(sample.list)
p <- vector("list",length(contrast.list))
names(p) <- names(contrast.list)
switch(class(object)[1],
CountDataSet = { # Has been normalized with DESeq
dge <- DGEList(counts=counts(object,normalized=TRUE),group=classes)
},
DGEList = { # Has been normalized with edgeR
dge <- object
},
matrix = { # Has been normalized with EDASeq or NOISeq
dge <- DGEList(object,group=classes)
},
list = { # Has been normalized with NBPSeq and main method was "nbpseq"
dge <- DGEList(counts=as.matrix(round(sweep(object$counts,2,
object$norm.factors,"*"))),group=classes)
},
nbp = { # Has been normalized with NBPSeq and main method was "nbsmyth"
dge <- DGEList(counts=as.matrix(round(object$pseudo.counts)),
group=classes)
}
)
for (con.name in names(contrast.list))
{
disp(" Contrast: ", con.name)
con <- contrast.list[[con.name]]
s <- unlist(con)
us <- unique(s)
ms <- match(names(s),rownames(dge$samples))
if (any(is.na(ms)))
ms <- ms[which(!is.na(ms))]
if (length(con)==2) {
design <- model.matrix(~0+s,data=dge$samples[ms,])
colnames(design) <- us
vom <- voom(dge[,ms],design,
normalize.method=stat.args$normalize.method)
fit <- lmFit(vom,design)
fit <- eBayes(fit)
co <- makeContrasts(contrasts=paste(us[2],us[1],sep="-"),
levels=design)
fit <- eBayes(contrasts.fit(fit,co))
p[[con.name]] <- fit$p.value[,1]
}
else {
design <- model.matrix(~s,data=dge$samples[ms,])
vom <- voom(dge[,ms],design,
normalize.method=stat.args$normalize.method)
fit <- lmFit(vom,design)
fit <- eBayes(fit)
res <- topTable(fit,coef=2:ncol(fit$design),number=nrow(vom))
p[[con.name]] <- res[,"P.Value"]
names(p[[con.name]]) <- rownames(res)
p[[con.name]] <- p[[con.name]][rownames(dge)]
}
p[[con.name]][which(is.na(p[[con.name]]))] <- 1
}
return(p)
}
#' Statistical testing with NOISeq
#'
#' This function is a wrapper over NOISeq statistical testing. It accepts a matrix
#' of normalized gene counts or an S4 object specific to each normalization
#' algorithm supported by metaseqR.
#'
#' @param object a matrix or an object specific to each normalization algorithm
#' supported by metaseqr, containing normalized counts. Apart from matrix (also
#' for NOISeq), the object can be a SeqExpressionSet (EDASeq), CountDataSet (DESeq)
#' or DGEList (edgeR).
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined in
#' the main help page of \code{\link{metaseqr}}.
#' @param stat.args a list of edgeR statistical algorithm parameters. See the
#' result of \code{get.defaults("statistics",} \code{"noiseq")} for an example
#' and how you can modify it.
#' @param gene.data an optional annotation data frame (such the ones produced by
#' \code{get.annotation} which contains the GC content for each gene and from
#' which the gene lengths can be inferred by chromosome coordinates.
#' @param log.offset a number to be added to each element of data matrix in order
#' to avoid Infinity on log type data transformations.
#' @return A named list of NOISeq q-values, whose names are the names of the
#' contrasts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' lengths <- round(1000*runif(nrow(data.matrix)))
#' starts <- round(1000*runif(nrow(data.matrix)))
#' ends <- starts + lengths
#' gc=runif(nrow(data.matrix)),
#' gene.data <- data.frame(
#' chromosome=c(rep("chr1",nrow(data.matrix)/2),rep("chr2",nrow(data.matrix)/2)),
#' start=starts,end=ends,gene_id=rownames(data.matrix),gc_content=gc
#' )
#' norm.data.matrix <- normalize.noiseq(data.matrix,sample.list,gene.data)
#' p <- stat.noiseq(norm.data.matrix,sample.list,contrast,gene.data=gene.data)
#'}
stat.noiseq <- function(object,sample.list,contrast.list=NULL,stat.args=NULL,
gene.data=NULL,log.offset=1) {
#if (is.null(norm.args) && class(object)=="DGEList")
# norm.args <- get.defaults("normalization","edger")
if (is.null(stat.args))
stat.args <- get.defaults("statistics","noiseq")
if (is.null(contrast.list))
contrast.list <- make.contrast.list(paste(names(sample.list)[1:2],
collapse="_vs_"),sample.list)
if (!is.list(contrast.list))
contrast.list <- make.contrast.list(contrast.list,sample.list)
if (is.null(gene.data)) {
gc.content <- NULL
chromosome <- NULL
biotype <- NULL
gene.length <- NULL
}
else {
gc.content <- gene.data$gc_content
biotype <- as.character(gene.data$biotype)
names(gc.content) <- names(biotype) <- rownames(gene.data)
if (is.null(attr(gene.data,"gene.length")))
gene.length <- NULL
else {
gene.length <- attr(gene.data,"gene.length")
names(gene.length) <- rownames(gene.data)
}
}
classes <- as.class.vector(sample.list)
p <- vector("list",length(contrast.list))
names(p) <- names(contrast.list)
switch(class(object)[1],
CountDataSet = { # Has been normalized with DESeq
ns.obj <- NOISeq::readData(
data=counts(object,normalized=TRUE),
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)
},
DGEList = { # Has been normalized with edgeR
# Trick found at http://cgrlucb.wikispaces.com/edgeR+spring2013
scl <- object$samples$lib.size * object$samples$norm.factors
dm <- round(t(t(object$counts)/scl)*mean(scl))
ns.obj <- NOISeq::readData(
data=dm,
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)
},
ExpressionSet = { # Has been normalized with NOISeq
ns.obj <- object
},
matrix = { # Has been normalized with EDASeq
ns.obj <- NOISeq::readData(
data=object,
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)
},
list = { # Has been normalized with NBPSeq and main method was "nbpseq"
ns.obj <- NOISeq::readData(
data=as.matrix(round(sweep(object$counts,2,
object$norm.factors,"*"))),
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)
},
nbp = { # Has been normalized with NBPSeq and main method was "nbsmyth"
ns.obj <- NOISeq::readData(
data=as.matrix(round(object$pseudo.counts)),
length=gene.length,
gc=gc.content,
chromosome=gene.data[,1:3],
factors=data.frame(class=classes),
biotype=biotype
)
}
)
for (con.name in names(contrast.list)) {
disp(" Contrast: ", con.name)
con <- contrast.list[[con.name]]
if (length(con)==2) {
stat.args$conditions=unique(unlist(con))
if (any(sapply(sample.list,function(x) ifelse(length(x)==1,
TRUE,FALSE))))
# At least one condition does not have replicates
stat.args$replicates <- "no"
if (stat.args$replicates %in% c("technical","no"))
res <- noiseq(ns.obj,k=log.offset,norm="n",
replicates=stat.args$replicates,factor=stat.args$factor,
conditions=stat.args$conditions,pnr=stat.args$pnr,
nss=stat.args$nss,v=stat.args$v,lc=stat.args$lc)
else
res <- noiseqbio(ns.obj,k=log.offset,norm="n",
nclust=stat.args$nclust,factor=stat.args$factor,
lc=stat.args$lc,conditions=stat.args$conditions,
r=stat.args$r,adj=stat.args$adj,a0per=stat.args$a0per,
cpm=stat.args$cpm,
filter=stat.args$filter,depth=stat.args$depth,
cv.cutoff=stat.args$cv.cutoff)
# Beware! This is not the classical p-value!
p[[con.name]] <- 1 - res@results[[1]]$prob
}
else {
warnwrap(paste("NOISeq differential expression algorithm does not ",
"support multi-factor designs (with more than two ",
"conditions to be compared)! Switching to DESeq for this ",
"comparison:",con.name))
M <- assayData(ns.obj)$exprs
cds <- newCountDataSet(round(M),data.frame(condition=unlist(con),
row.names=names(unlist(con))))
sizeFactors(cds) <- rep(1,ncol(cds))
cds <- estimateDispersions(cds,method="blind",
sharingMode="fit-only")
fit0 <- fitNbinomGLMs(cds,count~1)
fit1 <- fitNbinomGLMs(cds,count~condition)
p[[con.name]] <- nbinomGLMTest(fit1,fit0)
}
names(p[[con.name]]) <- rownames(ns.obj)
p[[con.name]][which(is.na(p[[con.name]]))] <- 1
}
return(p)
}
#' Statistical testing with baySeq
#'
#' This function is a wrapper over baySeq statistical testing. It accepts a matrix
#' of normalized gene counts or an S4 object specific to each normalization
#' algorithm supported by metaseqR.
#'
#' @param object a matrix or an object specific to each normalization algorithm
#' supported by metaseqr, containing normalized counts. Apart from matrix (also
#' for NOISeq), the object can be a SeqExpressionSet (EDASeq), CountDataSet
#' (DESeq) or DGEList (edgeR).
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined in
#' the main help page of \code{\link{metaseqr}}.
#' @param stat.args a list of edgeR statistical algorithm parameters. See the
#' result of \code{get.defaults("statistics",} \code{"bayseq")} for an example
#' and how you can modify it.
#' @param libsize.list an optional named list where names represent samples (MUST
#' be the same as the samples in \code{sample.list}) and members are the library
#' sizes (the sequencing depth) for each sample. If not provided, they will be
#' estimated from baySeq.
#' @return A named list of the value 1-likelihood that a gene is differentially
#' expressed, whose names are the names of the contrasts.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' norm.data.matrix <- normalize.edaseq(data.matrix,sample.list,gene.data)
#' p <- stat.bayseq(norm.data.matrix,sample.list,contrast)
#'}
stat.bayseq <- function(object,sample.list,contrast.list=NULL,stat.args=NULL,
libsize.list=NULL) {
if (is.null(stat.args))
stat.args <- get.defaults("statistics","bayseq")
if (is.null(contrast.list))
contrast.list <- make.contrast.list(paste(names(sample.list)[1:2],
collapse="_vs_"),sample.list)
if (!is.list(contrast.list))
contrast.list <- make.contrast.list(contrast.list,sample.list)
classes <- as.class.vector(sample.list)
p <- vector("list",length(contrast.list))
names(p) <- names(contrast.list)
switch(class(object)[1],
CountDataSet = { # Has been normalized with DESeq
bayes.data <- counts(object,normalized=TRUE)
},
DGEList = { # Has been normalized with edgeR
scl <- object$samples$lib.size * object$samples$norm.factors
bayes.data <- round(t(t(object$counts)/scl)*mean(scl))
},
matrix = { # Has been normalized with EDASeq or NOISeq
bayes.data <- object
},
list = {
bayes.data <- as.matrix(round(sweep(object$counts,2,
object$norm.factors,"*")))
},
nbp = {
bayes.data <- as.matrix(round(object$pseudo.counts))
}
)
CD <- new("countData",data=bayes.data,replicates=classes)
if (is.null(libsize.list))
baySeq::libsizes(CD) <- baySeq::getLibsizes(CD)
else
baySeq::libsizes(CD) <- unlist(libsize.list)
for (con.name in names(contrast.list)) {
disp(" Contrast: ", con.name)
con <- contrast.list[[con.name]]
#cd <- CD[,names(unlist(con))]
cd <- CD[,match(names(unlist(con)),colnames(CD@data))]
if (length(con)==2)
baySeq::groups(cd) <- list(NDE=rep(1,length(unlist(con))),
DE=c(rep(1,length(con[[1]])),rep(2,length(con[[2]]))))
else
baySeq::groups(cd) <- list(NDE=rep(1,length(unlist(con))),
DE=unlist(con,use.names=FALSE)) # Maybe this will not work
baySeq::replicates(cd) <- as.factor(classes[names(unlist(con))])
cd <- baySeq::getPriors.NB(cd,samplesize=stat.args$samplesize,
samplingSubset=stat.args$samplingSubset,
equalDispersions=stat.args$equalDispersions,
estimation=stat.args$estimation,zeroML=stat.args$zeroML,
consensus=stat.args$consensus,cl=stat.args$cl)
cd <- baySeq::getLikelihoods(cd,pET=stat.args$pET,
marginalise=stat.args$marginalise,subset=stat.args$subset,
priorSubset=stat.args$priorSubset,bootStraps=stat.args$bootStraps,
conv=stat.args$conv,nullData=stat.args$nullData,
returnAll=stat.args$returnAll,returnPD=stat.args$returnPD,
discardSampling=stat.args$discardSampling,cl=stat.args$cl)
tmp <- baySeq::topCounts(cd,group="DE",number=nrow(cd))
p[[con.name]] <- 1 - as.numeric(tmp[,"Likelihood"])
names(p[[con.name]]) <- rownames(tmp)
p[[con.name]] <- p[[con.name]][rownames(CD@data)]
p[[con.name]][which(is.na(p[[con.name]]))] <- 1
}
return(p)
}
#' Statistical testing with NBPSeq
#'
#' This function is a wrapper over NBPSeq statistical testing. It accepts a matrix
#' of normalized gene counts or an S4 object specific to each normalization
#' algorithm supported by metaseqR.
#'
#' @param object a matrix or an object specific to each normalization algorithm
#' supported by metaseqr, containing normalized counts. Apart from matrix (also
#' for NOISeq), the object can be a SeqExpressionSet (EDASeq), CountDataSet
#' (DESeq), DGEList (edgeR) or list (NBPSeq).
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined in
#' the main help page of \code{\link{metaseqr}}.
#' @param stat.args a list of NBPSeq statistical algorithm parameters. See the
#' result of \code{get.defaults("statistics",} \code{"nbpseq")}
#' for an example and how you can modify it. It is not required when the input
#' object is already a list from NBPSeq normalization as the dispersions are
#' already estimated.
#' @param libsize.list an optional named list where names represent samples
#' (MUST be the same as the samples \code{in sample.list}) and members are the
#' library sizes (the sequencing depth) for each sample. If not provided, the
#' default is the column sums of the \code{gene.counts} matrix.
#' @return A named list of p-values, whose names are the names of the contrasts.
#' @note There is currently a problem with the NBPSeq package and the workflow that
#' is specific to the NBPSeq package. The problem has to do with function exporting
#' as there are certain functions which are not recognized from the package
#' internally. For this reason and until it is fixed, only the Smyth workflow
#' will be available with the NBPSeq package.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' norm.data.matrix <- normalize.nbpseq(data.matrix,sample.list)
#' p <- stat.nbpseq(norm.data.matrix,sample.list,contrast)
#'}
stat.nbpseq <- function(object,sample.list,contrast.list=NULL,stat.args=NULL,
libsize.list=NULL) {
if (is.null(stat.args) && !is(object,"list"))
stat.args <- get.defaults("statistics","nbpseq")
if (is.null(contrast.list))
contrast.list <- make.contrast.list(paste(names(sample.list)[1:2],
collapse="_vs_"),sample.list)
if (!is.list(contrast.list))
contrast.list <- make.contrast.list(contrast.list,sample.list)
classes <- as.class.vector(sample.list)
p <- vector("list",length(contrast.list))
names(p) <- names(contrast.list)
switch(class(object)[1],
CountDataSet = { # Has been normalized with DESeq
counts <- round(counts(object,normalized=TRUE))
if (is.null(libsize.list)) {
libsize.list <- vector("list",length(classes))
names(libsize.list) <- unlist(sample.list,use.names=FALSE)
for (n in names(libsize.list))
libsize.list[[n]] <- sum(counts[,n])
}
lib.sizes <- unlist(libsize.list)
},
DGEList = { # Has been normalized with edgeR
# Trick found at http://cgrlucb.wikispaces.com/edgeR+spring2013
scl <- object$samples$lib.size * object$samples$norm.factors
counts <- round(t(t(object$counts)/scl)*mean(scl))
if (is.null(libsize.list)) {
libsize.list <- vector("list",length(classes))
names(libsize.list) <- unlist(sample.list,use.names=FALSE)
for (n in names(libsize.list))
libsize.list[[n]] <- sum(counts[,n])
}
lib.sizes <- unlist(libsize.list)
},
matrix = { # Has been normalized with EDASeq or NOISeq
counts <- object
if (is.null(libsize.list)) {
libsize.list <- vector("list",length(classes))
names(libsize.list) <- unlist(sample.list,use.names=FALSE)
for (n in names(libsize.list))
libsize.list[[n]] <- sum(counts[,n])
}
lib.sizes <- unlist(libsize.list)
},
list = { # Has been normalized with NBPSeq
object$counts <- as.matrix(object$counts)
nb.data <- object
if (is.null(libsize.list)) {
libsize.list <- vector("list",length(classes))
names(libsize.list) <- unlist(sample.list,use.names=FALSE)
for (n in names(libsize.list))
libsize.list[[n]] <- sum(nb.data$counts[,n])
}
lib.sizes <- unlist(libsize.list)
nb.data$pseudo.lib.sizes=rep(1e+7,dim(object$counts)[2])
},
nbp = { # Same...
object$pseudo.counts <- as.matrix(object$pseudo.counts)
nb.data <- object
if (is.null(libsize.list)) {
libsize.list <- vector("list",length(classes))
names(libsize.list) <- unlist(sample.list,use.names=FALSE)
for (n in names(libsize.list))
libsize.list[[n]] <- sum(nb.data$counts[,n])
}
lib.sizes <- unlist(libsize.list)
}
)
# To avoid repeating the following chunk in the above
if (!is(object,"list") && !is(object,"nbp")) {
#if (stat.args$main.method=="nbpseq") {
# nb.data <- list(
# counts=as.matrix(counts),
# lib.sizes=lib.sizes,
# norm.factors=rep(1,dim(counts)[2]),
# eff.lib.sizes=lib.sizes*rep(1,dim(counts)[2]),
# rel.frequencies=as.matrix(sweep(counts,2,
# lib.sizes*rep(1,dim(counts)[2]),"/")),
# tags=matrix(row.names(counts),dim(counts)[1],1)
# )
#}
#else if (stat.args$main.method=="nbsmyth") {
# nb.data <- new("nbp",list(
# counts=as.matrix(counts),
# lib.sizes=lib.sizes,
# grp.ids=classes,
# eff.lib.sizes=lib.sizes*rep(1,dim(counts)[2]),
# pseudo.counts=as.matrix(counts),
# pseudo.lib.sizes=colSums(as.matrix(counts))*rep(1,dim(counts)[2])
# ))
nb.data <- list(
counts=as.matrix(counts),
lib.sizes=lib.sizes,
grp.ids=classes,
eff.lib.sizes=lib.sizes*rep(1,dim(counts)[2]),
pseudo.counts=as.matrix(counts),
#pseudo.lib.sizes=colSums(as.matrix(counts)) *
# rep(1,dim(counts)[2])
pseudo.lib.sizes=rep(1e+7,dim(counts)[2])
)
class(nb.data) <- "nbp"
#}
}
for (con.name in names(contrast.list)) {
disp(" Contrast: ", con.name)
con <- contrast.list[[con.name]]
cons <- unique(unlist(con))
if (length(con)==2) {
#if (stat.args$main.method=="nbpseq") {
# dispersions <- estimate.dispersion(nb.data,
# model.matrix(~classes),
# method=stat.args$method$nbpseq)
# res <- test.coefficient(nb.data,dispersion=dispersions,
# x=model.matrix(~classes),beta0=c(NA,0),
# tests=stat.args$tests,
# alternative=stat.args$alternative,print.level=1)
# #res <- nb.glm.test(nb.data$counts,x=model.matrix(~classes),
# beta0=c(NA,0),lib.sizes=lib.sizes,
# # dispersion.method=stat.args$method$nbpseq,
# tests=stat.args$tests)
# p[[con.name]] <- res[[stat.args$tests]]$p.values
# #p[[con.name]] <- res$test[[stat.args$tests]]$p.values
#}
#else if (stat.args$main.method=="nbsmyth") {
obj <- suppressWarnings(estimate.disp(nb.data,
model=stat.args$model$nbsmyth,print.level=0))
obj <- exact.nb.test(obj,cons[1],cons[2],print.level=0)
p[[con.name]] <- obj$p.values
#}
}
else {
warnwrap(paste("NBPSeq differential expression algorithm does not ",
"support ANOVA-like designs with more than two conditions to ",
"be compared! Switching to DESeq for this comparison:",
con.name))
cds <- newCountDataSet(nb.data$counts,
data.frame(condition=unlist(con),
row.names=names(unlist(con))))
sizeFactors(cds) <- rep(1,ncol(cds))
cds <- estimateDispersions(cds,method="blind",
sharingMode="fit-only")
fit0 <- fitNbinomGLMs(cds,count~1)
fit1 <- fitNbinomGLMs(cds,count~condition)
p[[con.name]] <- nbinomGLMTest(fit1,fit0)
}
names(p[[con.name]]) <- rownames(nb.data$counts)
p[[con.name]][which(is.na(p[[con.name]]))] <- 1
}
return(p)
}
<file_sep>/man/make.sample.list.Rd
\name{make.sample.list}
\alias{make.sample.list}
\title{Creates sample list from file}
\usage{
make.sample.list(input, type=c("simple","targets"))
}
\arguments{
\item{input}{a tab-delimited file structured as follows:
the first line of the external tab delimited file should
contain column names (names are not important). The first
column MUST contain UNIQUE sample names and the second
column MUST contain the biological condition where each
of the samples in the first column should belong to.}
\item{type}{one of \code{"simple"} or \code{"targets"} to
indicate if the input is a simple two column text file or
the targets file used to launch the main analysis pipeline.}
}
\value{
A named list whose names are the conditions of the
experiments and its members are the samples belonging to
each condition.
}
\description{
Create the main sample list from an external file.
}
\examples{
\donttest{
targets <- data.frame(sample=c("C1","C2","T1","T2"),
condition=c("Control","Control","Treatment","Treatment"))
write.table(targets,file="targets.txt",sep="\\t",row.names=FALSE,
quote="")
sample.list <- make.sample.list("targets.txt")
}
}
\author{
<NAME>
}
<file_sep>/man/diagplot.boxplot.Rd
\name{diagplot.boxplot}
\alias{diagplot.boxplot}
\title{Boxplots wrapper for the metaseqR package}
\usage{
diagplot.boxplot(mat, name = NULL, log.it = "auto",
y.lim = "default", is.norm = FALSE, output = "x11",
path = NULL, alt.names = NULL, ...)
}
\arguments{
\item{mat}{the count data matrix.}
\item{name}{the names of the samples plotted on the
boxdiagplot. If \code{NULL}, the function check the
column names of mat. If they are also \code{NULL}, sample
names are autogenerated. If \code{name="none"}, no sample
names are plotted. If name is a list, it should be the
sample.list argument provided to the manin metaseqr
function. In that case, the boxes are colored per group.}
\item{log.it}{whether to log transform the values of mat
or not. It can be \code{TRUE}, \code{FALSE} or
\code{"auto"} for auto-detection. Auto-detection log
transforms by default so that the boxplots are smooth and
visible.}
\item{y.lim}{custom y-axis limits. Leave the string
\code{"default"} for default behavior.}
\item{is.norm}{a logical indicating whether object
contains raw or normalized data. It is not essential and
it serves only plot annotation purposes.}
\item{output}{one or more R plotting device to direct the
plot result to. Supported mechanisms: \code{"x11"}
(default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
\code{"pdf"}, \code{"ps"} or \code{"json"}. The latter is
currently available for the creation of interactive
volcano plots only when reporting the output, through the
highcharts javascript library (JSON for boxplots not yet
available).}
\item{path}{the path to create output files.}
\item{alt.names}{an optional vector of names, e.g. HUGO
gene symbols, alternative or complementary to the unique
rownames of \code{mat} (which must exist!). It is used only
in JSON output.}
\item{...}{further arguments to be passed to plot
devices, such as parameter from \code{\link{par}}.}
}
\value{
The filename of the boxplot produced if it's a file.
}
\description{
A wrapper over the general boxplot function, suitable for
matrices produced and processed with the metaseqr
package. Intended for internal use but can be easily used
as stand-alone. It can colors boxes based on group
depending on the name argument.
}
\examples{
# Non-normalized boxplot
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
diagplot.boxplot(data.matrix,sample.list)
# Normalized boxplot
norm.args <- get.defaults("normalization","deseq")
object <- normalize.deseq(data.matrix,sample.list,norm.args)
diagplot.boxplot(object,sample.list)
}
\author{
<NAME>
}
<file_sep>/man/log2disp.Rd
\name{log2disp}
\alias{log2disp}
\title{Display value transformation}
\usage{
log2disp(mat, base = 2)
}
\arguments{
\item{mat}{input data matrix}
\item{base}{logarithmic base, 2 or 10}
}
\description{
Logarithmic transformation for display purposes. Internal
use only.
}
\author{
<NAME>
}
<file_sep>/man/make.project.path.Rd
\name{make.project.path}
\alias{make.project.path}
\title{Project path constructor}
\usage{
make.project.path(path, f = NULL)
}
\arguments{
\item{path}{The desired project path. Can be NULL for
auto-generation.}
\item{f}{The input counts table file.}
}
\value{
A list with project path elements.
}
\description{
Create the main metaseqr project path. Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/get.biotypes.Rd
\name{get.biotypes}
\alias{get.biotypes}
\title{Biotype converter}
\usage{
get.biotypes(a)
}
\arguments{
\item{a}{the annotation data frame (output of
\code{\link{get.annotation}}).}
}
\value{
A character vector of biotypes.
}
\description{
Returns biotypes as character vector. Internal use.
}
\examples{
\donttest{
hg18.genes <- get.annotation("hg18","gene")
hg18.bt <- get.biotypes(hg18.genes)
}
}
\author{
<NAME>
}
<file_sep>/man/get.ensembl.annotation.Rd
\name{get.ensembl.annotation}
\alias{get.ensembl.annotation}
\title{Ensembl annotation downloader}
\usage{
get.ensembl.annotation(org, type)
}
\arguments{
\item{org}{the organism for which to download
annotation.}
\item{type}{either \code{"gene"} or \code{"exon"}.}
}
\value{
A data frame with the canonical (not isoforms!) genes or
exons of the requested organism. When
\code{type="genes"}, the data frame has the following
columns: chromosome, start, end, gene_id, gc_content,
strand, gene_name, biotype. When \code{type="exon"} the
data frame has the following columns: chromosome, start,
end, exon_id, gene_id, strand, gene_name, biotype. The
gene_id and exon_id correspond to Ensembl gene and exon
accessions respectively. The gene_name corresponds to
HUGO nomenclature gene names.
}
\description{
This function connects to the EBI's Biomart service using
the package biomaRt and downloads annotation elements
(gene co-ordinates, exon co-ordinates, gene
identifications, biotypes etc.) for each of the supported
organisms. See the help page of \code{\link{metaseqr}}
for a list of supported organisms. The function downloads
annotation for an organism genes or exons.
}
\note{
The data frame that is returned contains only "canonical"
chromosomes for each organism. It does not contain
haplotypes or random locations and does not contain
chromosome M.
}
\examples{
\donttest{
hg19.genes <- get.ensembl.annotation("hg19","gene")
mm9.exons <- get.ensembl.annotation("mm9","exon")
}
}
\author{
<NAME>
}
<file_sep>/R/metaseqr-data.R
#' @docType data
#' @name hg19.exon.counts
#' @title Human RNA-Seq data with three conditions, three samples
#' @description This data set contains RNA-Seq exon read counts for 3 chromosomes.
#' Data are derived from three colon tissue types (normal, paracancerous, cancerous).
#' It contains three coditions (normal, paracancerous, cancerous) with one replicate
#' each (three samples in total). It also contains a predefined \code{sample.list}
#' and \code{libsize.list} named \code{sample.list.hg19} and \code{libsize.list.hg19}.
#' Data were downloaded from GEO (GSE33782) and the corresponding reference is
#' Wu et al., Transcriptome profiling of the cancer, adjacent non-tumor and distant
#' normal tissues from a colorectal cancer patient by deep sequencing. PLoS One
#' 2012, 7(8), e41001.
#' @usage hg19.exon.counts
#' @format a \code{data.frame} with exon read counts and some embedded annotation,
#' one row per exon.
#' @source GEO (http://www.ncbi.nlm.nih.gov/geo/)
#' @author <NAME>
NULL
#' @docType data
#' @name sample.list.hg19
#' @title Human RNA-Seq data with three conditions, three samples
#' @description The sample list for \code{hg19.exon.counts}. See the data set
#' description.
#' @usage sample.list.hg19
#' @format a named \code{list} with condition and sample names.
#' @source GEO (http://www.ncbi.nlm.nih.gov/geo/)
#' @author <NAME>
NULL
#' @docType data
#' @name libsize.list.hg19
#' @title Human RNA-Seq data with three conditions, three samples
#' @description The library size list for \code{hg19.exon.counts}. See the data
#' set description.
#' @usage libsize.list.hg19
#' @format a named \code{list} with library sizes.
#' @source GEO (http://www.ncbi.nlm.nih.gov/geo/)
#' @author <NAME>
NULL
#' @docType data
#' @name mm9.gene.counts
#' @title mouse RNA-Seq data with two conditions, four samples
#' @description This data set contains RNA-Seq gene read counts for 3 chromosomes.
#' The data were downloaded from the ENCODE public repository and are derived
#' from the study of Mortazavi et al., 2008 (<NAME>, <NAME>, <NAME>,
#' <NAME>, <NAME>. Mapping and quantifying mammalian transcriptomes by RNA-Seq.
#' Nat Methods. 2008 Jul;5(7):621-8). In their experiment, the authors studied
#' among others genes expression at two developmental stages of mouse liver cells.
#' It has two conditions-developmental stages (e14.5,
#' adult_8_weeks) and four samples (e14.5_1, e14.5_2, a8w_1, a8w_2). It also
#' contains a predefined \code{sample.list} and \code{libsize.list}
#' named \code{sample.list.mm9} and \code{libsize.list.mm9}.
#' @usage mm9.gene.counts
#' @format a \code{data.frame} with gene read counts and some embedded annotation,
#' one row per gene.
#' @source ENCODE (http://genome.ucsc.edu/encode/)
#' @author <NAME>
NULL
#' @docType data
#' @name sample.list.mm9
#' @title Mouse RNA-Seq data with two conditions, four samples
#' @description The sample list for \code{mm9.gene.counts}. See the data set
#' description.
#' @usage sample.list.mm9
#' @format a named \code{list} with condition and sample names.
#' @source ENCODE (http://genome.ucsc.edu/encode/)
#' @author <NAME>
NULL
#' @docType data
#' @name libsize.list.mm9
#' @title Mouse RNA-Seq data with two conditions, four samples
#' @description The library size list for \code{mm9.gene.counts}. See the data set
#' description.
#' @usage libsize.list.mm9
#' @format a named \code{list} with library sizes.
#' @source ENCODE (http://genome.ucsc.edu/encode/)
#' @author <NAME>
NULL
<file_sep>/man/make.venn.pairs.Rd
\name{make.venn.pairs}
\alias{make.venn.pairs}
\title{Helper for Venn diagrams}
\usage{
make.venn.pairs(algs)
}
\arguments{
\item{algs}{a vector with the names of the sets (up to
length 5, if larger, it will be truncated with a
warning).}
}
\value{
A list with as many pairs as the comparisons to be made
for the construction of the Venn diagram. The pairs are
encoded with the uppercase letters A through E, each one
corresponding to order of the input sets.
}
\description{
This function creates a list of pairwise comparisons to
be performed in order to create an up to 5-way Venn
diagram using the R package VennDiagram. Internal use
mostly.
}
\examples{
\donttest{
sets <- c("apple","pear","banana")
pairs <- make.venn.pairs(sets)
}
}
\author{
<NAME>
}
<file_sep>/man/meta.perm.Rd
\name{meta.perm}
\alias{meta.perm}
\title{Permutation tests for meta-analysis}
\usage{
meta.perm(contrast, counts, sample.list, statistics,
stat.args, libsize.list, nperm = 10000,
weight = rep(1/ncol(counts), ncol(counts)),
select = c("min", "max", "weight"), replace = "auto",
reprod=TRUE, multic = FALSE)
}
\arguments{
\item{contrast}{the contrasts to be tested by each
statistical algorithm. See the main
\code{\link{metaseqr}} help page.}
\item{counts}{a normalized read counts table, one row for
each gene, one column for each sample.}
\item{sample.list}{the list containing condition names
and the samples under each condition. See the main
\code{\link{metaseqr}} help page.}
\item{statistics}{the statistical algorithms used in
metaseqr. See the main \code{\link{metaseqr}} help page.}
\item{stat.args}{the parameters for each statistical
algorithm. See the main \code{\link{metaseqr}} help page.}
\item{libsize.list}{a list with library sizes. See the
main \code{\link{metaseqr}} and the \code{stat.*} help
pages.}
\item{nperm}{the number of permutations (Monte Carlo
simulations) to perform.}
\item{weight}{a numeric vector of weights for each
statistical algorithm.}
\item{select}{how to select the initial vector of
p-values. It can be \code{"min"} to select the minimum
p-value for each gene (more conservative), \code{"max"}
to select the maximum p-value for each gene (less
conservative), \code{"weight"} to apply the weights to
the p-value vector for each gene and derive a weighted
p-value.}
\item{replace}{same as the \code{replace} argument in the
\code{\link{sample}} function. Implies bootstraping or
simple resampling without replacement. It can also be
\code{"auto"}, to determine bootstraping or not with the
following rule: if \code{ncol(counts)<=6}
\code{replace=FALSE else} \code{replace=TRUE}. This
protects from the case of having zero variability across
resampled conditions. In such cases, most statistical
tests would crash.}
\item{reprod}{create reproducible permutations. Ideally
one would want to create the same set of indices for a
given dataset so as to create reproducible p-values. If
\code{reprod=TRUE}, a fixed seed is used by
\code{meta.perm} for all the datasets analyzed
with \code{metaseqr}. If \code{reprod=FALSE}, then the
p-values will not be reproducible, although statistical
significance is not expected to change for a large
number of resambling. Finally, \code{reprod} can be a
numeric vector of seeds with the same length as
\code{nperm} so that the user can supply his/her own
seeds.}
\item{multic}{use multiple cores to execute the
premutations. This is an external parameter and implies
the existence of multicore package in the execution
environment. See the main \code{\link{metaseqr}} help
page.}
}
\value{
A vector of meta p-values
}
\description{
This function performs permutation tests in order to
derive a meta p-value by combining several of the
statistical algorithms of metaseqr. This is probably the
most accurate way of combining multiple statistical
algorithms for RNA-Seq data, as this issue is different
from the classic interpretation of the term
"meta-analysis" which implies the application of the same
statistical test on different datasets treating the same
subject/experiment. For other methods, see also the main
\code{\link{metaseqr}} help page. You should keep in mind
that the permutation procedure can take a long time, even
when executed in parallel.
}
\examples{
\donttest{
# Not yet available
}
}
\author{
<NAME>
}
<file_sep>/man/check.graphics.file.Rd
\name{check.graphics.file}
\alias{check.graphics.file}
\title{Check graphics file}
\usage{
check.graphics.file(o)
}
\arguments{
\item{o}{the plotting device, see main metaseqr function}
}
\description{
Graphics file checker. Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/downsample.counts.Rd
\name{downsample.counts}
\alias{downsample.counts}
\title{Downsample read counts}
\usage{
downsample.counts(counts,seed=42)
}
\arguments{
\item{counts}{the read counts table
which is subjected to downsampling.}
\item{seed}{random seed for reproducible
downsampling.}
}
\value{
The downsampled counts matrix.
}
\description{
This function downsamples the library sizes
of a read counts table to the lowest library
size, according to the methdology used in
(Soneson and Delorenzi, BMC Bioinformatics,
2013).
}
\examples{
## Dowload locally the file "bottomly_count_table.txt" from
## the ReCount database
#download.file(paste("http://bowtie-bio.sourceforge.net/",
# "recount/countTables/bottomly_count_table.txt",sep=""),
# destfile="~/bottomly_count_table.txt")
#M <- as.matrix(read.delim("~/bottomly_count_table.txt",row.names=1))
#D <- downsample.counts(M)
}
\author{
<NAME>
}
<file_sep>/man/combine.weight.Rd
\name{combine.weight}
\alias{combine.weight}
\title{Combine p-values using weights}
\usage{
combine.weight(p, w)
}
\arguments{
\item{p}{a p-value matrix (rows are genes,
columns are statistical tests).}
\item{w}{a weights vector, must sum to 1.}
}
\value{
A vector of combined p-values.
}
\description{
This function combines p-values from the
various statistical tests supported by
metaseqR using p-value weights.
}
\examples{
p <- matrix(runif(300),100,3)
pc <- combine.weight(p,w=c(0.2,0.5,0.3))
}
\author{
<NAME>
}
<file_sep>/R/metaseqr.plot.R
#' Diagnostic plots for the metaseqr package
#'
#' This is the main function for producing sructured quality control and informative
#' graphs base on the results of the various steps of the metaseqR package. The
#' graphs produced span a variety of issues like good sample reproducibility
#' (Multi-Dimensional Scaling plot, biotype detection, heatmaps. diagplot.metaseqr,
#' apart from implementing certain package-specific plots, is a wrapper around
#' several diagnostic plots present in other RNA-Seq analysis packages such as
#' EDASeq and NOISeq.
#'
#' @param object a matrix or a data frame containing count data derived before or
#' after the normalization procedure, filtered or not by the metaseqR's filters
#' and/or p-value. The object can be fed to any of the \code{diagplot.metaseqr}
#' plotting systems but not every plot is meaningful. For example, it's meaningless
#' to create a \code{"biodist"} plot for a count matrix before normalization or
#' statistical testing.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param annotation a data frame containing annotation elements for each row in
#' object. Usually, a subset of the annotation obtained by \code{\link{get.annotation}}
#' or a subset of possibly embedded annotation with the input counts table. This
#' parameter is optional and required only when diagplot.type is any of
#' \code{"biodetection"}, \code{"countsbio"}, \code{"saturation"}, \code{"rnacomp"},
#' \code{"readnoise"}, \code{"biodist"}, \code{"gcbias"}, \code{"lengthbias"} or
#' \code{"filtered"}.
#' @param contrast.list a named structured list of contrasts as returned by
#' \code{\link{make.contrast.list}} or just the vector of contrasts as defined in
#' the main help page of \code{\link{metaseqr}}. This parameter is optional and
#' required only when \code{diagplot.type} is any of \code{"deheatmap"},
#' \code{"volcano"} or \code{"biodist"}.
#' @param p.list a list of p-values for each contrast as obtained from any of the
#' \code{stat.*} methods of the metaseqr package. This parameter is optional and
#' required only when \code{diagplot.type} is any of \code{"deheatmap"},
#' \code{"volcano"} or \code{"biodist"}.
#' @param thresholds a list with the elements \code{"p"} and \code{"f"} which are
#' the p-value and the fold change cutoff when \code{diagplot.type="volcano"}.
#' @param diagplot.type one or more of the diagnostic plots supported in metaseqR
#' package. Many of these plots require the presence of additional package,
#' something that is checked while running the main metaseqr function. The supported
#' plots are \code{"mds"}, \code{"biodetection"}, \code{"countsbio"},
#' \code{"saturation"}, \code{"rnacomp"}, \code{"boxplot"}, \code{"gcbias"},
#' \code{"lengthbias"}, \code{"meandiff"}, \code{"meanvar"}, \code{"deheatmap"},
#' \code{"volcano"}, \code{"biodist"}, \code{"filtered"}, \code{"readnoise"},
#' \code{"venn"}, \code{"correl"}, \code{"pairwise"}. For a brief description of
#' these plots please see the main \code{\link{metaseqr}} help page.
#' @param is.norm a logical indicating whether object contains raw or normalized
#' data. It is not essential and it serves only plot annotation purposes.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"png"}, \code{"jpg"}, \code{"bmp"}, \code{"pdf"},
#' \code{"ps"} or \code{"json"}. The latter is currently available for the creation
#' of interactive volcano plots only when reporting the output, through the
#' highcharts javascript library. The default plotting (\code{"x11"}) is not
#' supported due to instability in certain devices.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return A named list containing the file names of the produced plots. Each list
#' member is names according to the selected plotting device and is also a named
#' list, whose names are the plot types. The final contents are the file names in
#' case the plots are written to a physical location (not meaningful for \code{"x11"}).
#' @note In order to make the best out of this function, you should generally
#' provide the annotation argument as most and also the most informative plots
#' depend on this. If you don't know what is inside your counts table or how many
#' annotation elements you can provide by embedding it, it's always best to set
#' the annotation parameter of the main metaseqr function to \code{"download"} to
#' use predefined annotations that work better with the functions of the whole
#' package.
#' @author <NAME>
#' @export
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' diagplot.metaseqr(data.matrix,sample.list,diagplot.type=c("mds","boxplot"))
#'
#' norm.args <- get.defaults("normalization","deseq")
#' object <- normalize.deseq(data.matrix,sample.list,norm.args)
#' diagplot.metaseqr(object,sample.list,diagplot.type="boxplot")
#'
#' p <- stat.deseq(object)
#' diagplot.metaseqr(object,sample.list,contrast.list=contrast,p.list=p,
#' diagplot.type="volcano")
#'}
diagplot.metaseqr <- function(object,sample.list,annotation=NULL,contrast.list=NULL,
p.list=NULL,thresholds=list(p=0.05,f=1),diagplot.type=c("mds","biodetection",
"countsbio","saturation","readnoise","rnacomp","correl","pairs","boxplot",
"gcbias","lengthbias","meandiff","meanvar","deheatmap","volcano","biodist",
"filtered","venn"),is.norm=FALSE,output="x11",path=NULL,...) {
# annotation should have the format internally created here... This function
# can be used outside so it must be checked at some point...
if (!is.matrix(object) && !is.data.frame(object))
stopwrap("object argument must be a matrix or data frame!")
if (is.null(annotation) && any(diagplot.type %in% c("biodetection",
"countsbio","saturation","rnacomp","readnoise","biodist","gcbias",
"lengthbias","filtered")))
stopwrap("annotation argument is needed when diagplot.type is ",
"\"biodetection\", \"countsbio\",\"saturation\",\"rnacomp\", ",
"\"readnoise\", \"biodist\", \"gcbias\", \"lengthbias\", ",
"\"filtered\" or \"venn\"!")
if (any(diagplot.type %in% c("deheatmap","volcano","biodist","venn"))) {
if (is.null(contrast.list))
stopwrap("contrast.list argument is needed when diagplot.type is ",
"\"deheatmap\",\"volcano\", \"biodist\" or \"venn\"!")
if (is.null(p.list))
stopwrap("The p argument which is a list of p-values for each ",
"contrast is needed when diagplot.type is \"deheatmap\", ",
"\"volcano\", \"biodist\" or \"venn\"!")
if (is.na(thresholds$p) || is.null(thresholds$p) || thresholds$p==1) {
warnwrap(paste("The p-value threshold when diagplot.type is ",
"\"deheatmap\", \"volcano\", \"biodist\" or \"venn\" must allow ",
"the normal plotting of DEG diagnostic plots! Setting to 0.05..."))
thresholds$p <- 0.05
}
}
if (is.null(path)) path <- getwd()
if (is.data.frame(object) && !("filtered" %in% diagplot.type))
object <- as.matrix(object)
if (any(diagplot.type %in% c("biodetection","countsbio","saturation",
"rnacomp","biodist","readnoise")))
covars <- list(
data=object,
length=annotation$end - annotation$start,
gc=annotation$gc_content,
chromosome=annotation[,1:3],
factors=data.frame(class=as.class.vector(sample.list)),
biotype=annotation$biotype,
gene_name=as.character(annotation$gene_name)
)
raw.plots <- c("mds","biodetection","countsbio","saturation","readnoise",
"correl","pairwise")
norm.plots <- c("boxplot","gcbias","lengthbias","meandiff","meanvar",
"rnacomp")
stat.plots <- c("deheatmap","volcano","biodist")
other.plots <- c("filtered")
venn.plots <- c("venn")
files <- list()
for (p in diagplot.type) {
disp(" Plotting ",p,"...")
if (p %in% raw.plots && !is.norm) {
switch(p,
mds = {
files$mds <- diagplot.mds(object,sample.list,output=output,
path=path)
},
biodetection = {
files$biodetection <- diagplot.noiseq(object,sample.list,
covars,which.plot=p,output=output,path=path,...)
},
countsbio = {
files$countsbio <- diagplot.noiseq(object,sample.list,
covars,which.plot=p,output=output,path=path,...)
},
saturation = {
fil <- diagplot.noiseq(object,sample.list,covars,
which.plot=p,output=output,path=path,...)
files$saturation$biotype <- fil[["biotype"]]
files$saturation$sample <- fil[["sample"]]
},
readnoise = {
files$readnoise <- diagplot.noiseq(object,sample.list,
covars,which.plot=p,output=output,path=path,...)
},
correl = {
files$correl$heatmap <- diagplot.cor(object,type="heatmap",
output=output,path=path,...)
files$correl$correlogram <- diagplot.cor(object,
type="correlogram",output=output,path=path,...)
},
pairwise = {
files$pairwise <- diagplot.pairs(object,output=output,
path=path)
}
)
}
if (p %in% norm.plots) {
switch(p,
boxplot = {
files$boxplot <- diagplot.boxplot(object,name=sample.list,
is.norm=is.norm,output=output,path=path,...)
},
gcbias = {
files$gcbias <- diagplot.edaseq(object,sample.list,
covar=annotation$gc_content,is.norm=is.norm,
which.plot=p,output=output,path=path,...)
},
lengthbias = {
files$lengthbias <- diagplot.edaseq(object,sample.list,
covar=annotation$end-annotation$start,is.norm=is.norm,
which.plot=p,output=output,path=path,...)
},
meandiff = {
fil <- diagplot.edaseq(object,sample.list,is.norm=is.norm,
which.plot=p,output=output,path=path,...)
for (n in names(fil)) {
if (!is.null(fil[[n]]))
files$meandiff[[n]] <- unlist(fil[[n]])
}
},
meanvar = {
fil <- diagplot.edaseq(object,sample.list,is.norm=is.norm,
which.plot=p,output=output,path=path,...)
for (n in names(fil)) {
if (!is.null(fil[[n]]))
files$meanvar[[n]] <- unlist(fil[[n]])
}
},
rnacomp = {
files$rnacomp <- diagplot.noiseq(object,sample.list,covars,
which.plot=p,output=output,is.norm=is.norm,path=path,
...)
}
)
}
if (p %in% stat.plots && is.norm) {
for (cnt in names(contrast.list)) {
disp(" Contrast: ",cnt)
samples <- names(unlist(contrast.list[[cnt]]))
mat <- as.matrix(object[,match(samples,colnames(object))])
switch(p,
deheatmap = {
files$deheatmap[[cnt]] <- diagplot.de.heatmap(mat,cnt,
output=output,path=path)
},
volcano = {
fc <- log2(make.fold.change(cnt,sample.list,object,1))
for (contrast in colnames(fc)) {
files$volcano[[contrast]] <- diagplot.volcano(
fc[,contrast],p.list[[cnt]],contrast,
fcut=thresholds$f,pcut=thresholds$p,
output=output,path=path)
}
},
biodist = {
files$biodist[[cnt]] <- diagplot.noiseq(object,
sample.list,covars,which.plot=p,output=output,
biodist.opts=list(p=p.list[[cnt]],
pcut=thresholds$p,name=cnt),path=path,...)
}
)
}
}
if (p %in% other.plots) {
switch(p,
filtered = {
files$filtered <- diagplot.filtered(object,annotation,
output=output,path=path)
}
)
}
if (p %in% venn.plots) {
switch(p,
venn = {
for (cnt in names(contrast.list)) {
disp(" Contrast: ",cnt)
if (!is.null(annotation)) {
alt.names <- as.character(annotation$gene_name)
names(alt.names) <- rownames(annotation)
}
else
alt.names <- NULL
files$venn[[cnt]] <- diagplot.venn(p.list[[cnt]],
pcut=thresholds$p,nam=cnt,output=output,path=path,
alt.names=alt.names)
}
}
)
}
}
return(files)
}
#' Boxplots wrapper for the metaseqR package
#'
#' A wrapper over the general boxplot function, suitable for matrices produced
#' and processed with the metaseqr package. Intended for internal use but can be
#' easily used as stand-alone. It can colors boxes based on group depending on
#' the name argument.
#'
#' @param mat the count data matrix.
#' @param name the names of the samples plotted on the boxplot. If \code{NULL},
#' the function check the column names of mat. If they are also \code{NULL}, sample
#' names are autogenerated. If \code{name="none"}, no sample names are plotted.
#' If name is a list, it should be the sample.list argument provided to the manin
#' metaseqr function. In that case, the boxes are colored per group.
#' @param log.it whether to log transform the values of mat or not. It can be
#' \code{TRUE}, \code{FALSE} or \code{"auto"} for auto-detection. Auto-detection
#' log transforms by default so that the boxplots are smooth and visible.
#' @param y.lim custom y-axis limits. Leave the string \code{"default"} for default
#' behavior.
#' @param is.norm a logical indicating whether object contains raw or normalized
#' data. It is not essential and it serves only plot annotation purposes.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"}, \code{"ps"} or \code{"json"}. The latter is
#' currently available for the creation of interactive volcano plots only when
#' reporting the output, through the highcharts javascript library (JSON for
#' boxplots not yet available).
#' @param path the path to create output files.
#' @param alt.names an optional vector of names, e.g. HUGO gene symbols, alternative
#' or complementary to the unique names of \code{f} or \code{p} (one of them must
#' be named!). It is used only in JSON output.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filename of the boxplot produced if it's a file.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.boxplot(data.matrix,sample.list)
#'
#' norm.args <- get.defaults("normalization","deseq")
#' object <- normalize.deseq(data.matrix,sample.list,norm.args)
#' diagplot.boxplot(object,sample.list)
#'}
diagplot.boxplot <- function(mat,name=NULL,log.it="auto",y.lim="default",
is.norm=FALSE,output="x11",path=NULL,alt.names=NULL,...) {
if (is.null(path)) path <- getwd()
if (is.norm)
status<- "normalized"
else
status<- "raw"
# Need to log?
if (log.it=="auto") {
if (diff(range(mat,na.rm=TRUE))>1000)
mat <- log2disp(mat)
}
else if (log.it=="yes")
mat <- log2disp(mat)
# Define the axis limits based on user input
if (!is.numeric(y.lim) && y.lim=="default") {
min.y <- floor(min(mat))
max.y <- ceiling(max(mat))
}
else if (is.numeric(y.lim)) {
min.y <- y.lim[1]
max.y <- y.lim[2]
}
grouped <- FALSE
if (is.null(name)) {
if (is.null(colnames(mat)))
nams <- paste("Sample",1:ncol(mat),sep=" ")
else
nams <- colnames(mat)
}
else if (length(name)==1 && name=="none")
nams <- rep("",ncol(mat))
else if (is.list(name)) { # Is sample.list
nams <- unlist(name)
grouped <- TRUE
}
cols <- c("red3","green3","blue2","gold","skyblue","orange3","burlywood",
"red","blue","green","orange","darkgrey","green4","black","pink",
"brown","magenta","yellowgreen","pink4","seagreen4","darkcyan")
if (grouped) {
tmp <- as.numeric(factor(as.class.vector(name)))
b.cols <- cols[tmp]
}
else b.cols <- cols
mat.list <- list()
for (i in 1:ncol(mat))
mat.list[[i]] <- mat[,i]
names(mat.list) <- nams
if (output != "json") {
fil <- file.path(path,paste("boxplot_",status,".",output,sep=""))
graphics.open(output,fil)
if (!is.numeric(y.lim) && y.lim=="default")
b <- boxplot(mat.list,names=nams,col=b.cols,las=2,main=paste(
"Boxplot ",status,sep=""),...)
else
b <- boxplot(mat.list,names=nams,col=b.cols,ylim=c(min.y,max.y),
las=2,main=paste("Boxplot ",status,sep=""),...)
graphics.close(output)
}
else {
# Create boxplot object
b <- boxplot(mat.list,plot=FALSE)
colnames(b$stat) <- nams
# Locate the outliers
o.list <- lapply(names(mat.list),function(x,M,b) {
v <- b[,x]
o <- which(M[[x]]<v[1] | M[[x]]>v[5])
if (length(o)>0)
return(M[[x]][o])
else
return(NULL)
},mat.list,b$stat)
# Create output object
obj <- list(
x=NULL,
y=NULL,
plot=b,
samples=name,
ylims=c(min.y,max.y),
xlims=NULL,
status=status,
pcut=NULL,
fcut=NULL,
altnames=alt.names,
user=o.list
)
json <- boxplotToJSON(obj)
fil <- file.path(path,paste("boxplot_",status,".json",sep=""))
disp("Writing ",fil)
write(json,fil)
}
return(fil)
}
#' Multi-Dimensinal Scale plots or RNA-Seq samples
#'
#' Creates a Multi-Dimensional Scale plot for the given samples based on the count
#' data matrix. MDS plots are very useful for quality control as you can easily
#' see of samples of the same groups are clustered together based on the whole
#' dataset.
#'
#' @param x the count data matrix.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param method which correlation method to use. Same as the method parameter in
#' \code{\link{cor}} function.
#' @param log.it whether to log transform the values of x or not.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"}, \code{"ps"} or \code{"json"}. The latter is
#' currently available for the creation of interactive volcano plots only when
#' reporting the output, through the highcharts javascript library.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filename of the MDS plot produced if it's a file.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' diagplot.mds(data.matrix,sample.list)
#'}
diagplot.mds <- function(x,sample.list,method="spearman",log.it=TRUE,
output="x11",path=NULL,...) {
if (is.null(path)) path <- getwd()
classes <- as.factor(as.class.vector(sample.list))
design <- as.numeric(classes)
colspace <- c("red","blue","yellowgreen","orange","aquamarine2",
"pink2","seagreen4","brown","purple","chocolate")
pchspace <- c(20,17,15,16,8,3,2,0,1,4)
if (ncol(x)<3) {
warnwrap("MDS plot cannot be created with less than 3 samples! ",
"Skipping...")
return(NULL)
}
if (log.it)
y <- nat2log(x,base=2)
else
y <- x
d <- as.dist(0.5*(1-cor(y,method=method)))
mds.obj <- cmdscale(d,eig=TRUE,k=2)
xr <- diff(range(min(mds.obj$points[,1]),max(mds.obj$points[,1])))
yr <- diff(range(min(mds.obj$points[,2]),max(mds.obj$points[,2])))
xlim <- c(min(mds.obj$points[,1])-xr/10,max(mds.obj$points[,1])+xr/10)
ylim <- c(min(mds.obj$points[,2])-yr/10,max(mds.obj$points[,2])+yr/10)
if (output!="json") {
fil <- file.path(path,paste("mds.",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=9,height=7)
else
graphics.open(output,fil,width=1024,height=768)
plot(mds.obj$points[,1],mds.obj$points[,2],
col=colspace[1:length(levels(classes))][design],
pch=pchspace[1:length(levels(classes))][design],
xlim=xlim,ylim=ylim,
main="MDS plot",xlab="MDS 1",ylab="MDS 2",
cex=0.9,cex.lab=0.9,cex.axis=0.9,cex.main=0.9)
text(mds.obj$points[,1],mds.obj$points[,2],labels=colnames(x),pos=3,
cex=0.7)
grid()
graphics.close(output)
}
else {
# Create output object
xx <- mds.obj$points[,1]
yy <- mds.obj$points[,2]
names(xx) <- names(yy) <- unlist(sample.list)
obj <- list(
x=xx,
y=yy,
plot=NULL,
samples=sample.list,
ylim=ylim,
xlim=xlim,
status=NULL,
pcut=NULL,
fcut=NULL,
altnames=NULL,
user=NULL
)
json <- mdsToJSON(obj)
fil <- file.path(path,"mds.json")
disp("Writing ",fil)
write(json,fil)
}
return(fil)
}
#' Massive X-Y, M-D correlation plots
#'
#' This function uses the read counts matrix to create pairwise correlation plots.
#' The upper diagonal of the final image contains simple scatterplots of each
#' sample against each other (log2 scale) while the lower diagonal contains
#' mean-difference plots for the same samples (log2 scale). This type of diagnostic
#' plot may not be interpretable for more than 10 samples.
#'
#' @param x the read counts matrix or data frame.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filename of the pairwise comparisons plot produced if it's a file.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' diagplot.pairs(data.matrix)
#'}
diagplot.pairs <- function(x,output="x11",path=NULL,...) {
x <- as.matrix(x)
x <- nat2log(x)
n <- ncol(x)
if (!is.null(colnames(x)))
nams <- colnames(x)
else
nams <- paste("Sample_",1:ncol(x),sep="")
if (!is.null(path))
fil <- file.path(path,paste("correlation_pairs",output,sep="."))
else
fil <- paste("correlation_pairs",output,sep=".")
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=12,height=12)
else {
if (ncol(x)<=5)
graphics.open(output,fil,width=800,height=800,res=100)
else
graphics.open(output,fil,width=1024,height=1024,res=150)
}
# Setup the grid
par(mfrow=c(n,n),mar=c(1,1,1,1),oma=c(1,1,0,0),mgp=c(2,0.5,0),cex.axis=0.6,
cex.lab=0.6)
# Plot
for (i in 1:n)
{
for (j in 1:n)
{
if (i==j)
{
plot(0:10,0:10,type="n",xaxt="n",yaxt="n",xlab="",ylab="") # Diagonal
text(c(3,5,3),c(9.5,5,1),c("X-Y plots",nams[i],"M-D plots"),
cex=c(0.8,1,0.8))
arrows(6,9.5,9.5,9.5,angle=20,length=0.1,lwd=0.8,cex=0.8)
arrows(0.2,3.2,0.2,0.2,angle=20,length=0.1,lwd=0.8,cex=0.8)
}
else if (i<j) # XY plot
{
plot(x[,i],x[,j],pch=20,col="blue",cex=0.4,xlab=nams[i],
ylab=nams[j],...)
lines(lowess(x[,i],x[,j]),col="red")
cc <- paste("cor:",formatC(cor(x[,i],x[,j]),digits=3))
text(3,max(x[,j]-1),labels=cc,cex=0.7,)
#grid()
}
else if (i>j) # MD plot
{
plot((x[,i]+x[,j])/2,x[,j]-x[,i],pch=20,col="blue",cex=0.4,...)
lines(lowess((x[,i]+x[,j])/2,x[,j]-x[,i]),col="red")
#grid()
}
}
}
graphics.close(output)
return(fil)
}
#' Summarized correlation plots
#'
#' This function uses the read counts matrix to create heatmap or correlogram
#' correlation plots.
#'
#' @param mat the read counts matrix or data frame.
#' @param type create heatmap of correlogram plots.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filename of the pairwise comparisons plot produced if it's a file.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' diagplot.cor(data.matrix,type="heatmap")
#' diagplot.cor(data.matrix,type="correlogram")
#'}
diagplot.cor <- function(mat,type=c("heatmap","correlogram"),output="x11",
path=NULL,...) {
x <- as.matrix(mat)
type <- tolower(type[1])
check.text.args("type",type,c("heatmap","correlogram"))
#if (!require(corrplot) && type=="correlogram")
# stop("R package corrplot is required!")
cor.mat <- cor(mat)
if (!is.null(colnames(mat)))
colnames(cor.mat) <- colnames(mat)
if (!is.null(path))
fil <- file.path(path,paste("correlation_",type,".",output,sep=""))
else
fil <- paste("correlation_",type,".",output,sep="")
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=7,height=7)
else
graphics.open(output,fil,width=640,height=640,res=100)
if (type=="correlogram")
corrplot(cor.mat,method="ellipse",order="hclust",...)
else if (type=="heatmap") {
n <- dim(cor.mat)[1]
labs <- matrix(NA,n,n)
for (i in 1:n)
for (j in 1:n)
labs[i,j] <- sprintf("%.2f",cor.mat[i,j])
if (n <= 5)
notecex <- 1.2
else if (n > 5 & n < 10)
notecex <- 0.9
else
notecex <- 0.7
heatmap.2(cor.mat,col=colorRampPalette(c("yellow","grey","blue")),
revC=TRUE,trace="none",symm=TRUE,Colv=TRUE,cellnote=labs,
keysize=1,density.info="density",notecex=notecex,cexCol=0.9,
cexRow=0.9,font.lab=2)
}
graphics.close(output)
return(fil)
}
#' Diagnostic plots based on the EDASeq package
#'
#' A wrapper around the plotting functions availale in the EDASeq normalization
#' Bioconductor package. For analytical explanation of each plot please see the
#' vignette of the EDASeq package. It is best to use this function through the
#' main plotting function \code{\link{diagplot.metaseqr}}.
#'
#' @param x the count data matrix.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param covar The covariate to plot counts against. Usually \code{"gc"} or
#' \code{"length"}.
#' @param is.norm a logical indicating whether object contains raw or normalized
#' data. It is not essential and it serves only plot annotation purposes.
#' @param which.plot the EDASeq package plot to generate. It can be one or more
#' of \code{"meanvar"}, \code{"meandiff"}, \code{"gcbias"} or \code{"lengthbias"}.
#' Please refer to the documentation of the EDASeq package for details on the use
#' of these plots. The \code{which.plot="lengthbias"} case is not covered by
#' EDASeq documentation, however it is similar to the GC-bias plot when the
#' covariate is the gene length instead of the GC content.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filenames of the plot produced in a named list with names the
#' which.plot argument. If \code{output="x11"}, no output filenames are produced.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' gc <- runif(nrow(data.matrix))
#' diagplot.edaseq(data.matrix,sample.list,which.plot="meandiff")
#'}
diagplot.edaseq <- function(x,sample.list,covar=NULL,is.norm=FALSE,
which.plot=c("meanvar","meandiff","gcbias","lengthbias"),output="x11",
path=NULL,...) {
if (is.null(path)) path <- getwd()
check.text.args("which.plot",which.plot,c("meanvar","meandiff","gcbias",
"lengthbias"),multiarg=TRUE)
if (is.null(covar) && which.plot %in% c("gcbias","lengthbias"))
stopwrap("\"covar\" argument is required when \"which.plot\" is ",
"\"gcbias\ or \"lengthbias\"!")
if (is.norm)
status <- "normalized"
else
status <- "raw"
if (is.null(covar)) covar <- rep(NA,nrow(x))
s <- newSeqExpressionSet(x,phenoData=AnnotatedDataFrame(
data.frame(conditions=factor(as.class.vector(sample.list)),
row.names=colnames(x))),featureData=AnnotatedDataFrame(data.frame(
gc=covar,length=covar,row.names=rownames(x))))
switch(which.plot,
meandiff = {
fil <- vector("list",length(sample.list))
names(fil) <- names(sample.list)
for (n in names(sample.list)) {
if (length(sample.list[[n]])==1) {
warnwrap("Cannot create a mean-difference plot with one ",
"sample per condition! Skipping...")
next
}
pair.matrix <- combn(1:length(sample.list[[n]]),2)
fil[[n]] <- vector("list",ncol(pair.matrix))
for (i in 1:ncol(pair.matrix)) {
s1 <- sample.list[[n]][pair.matrix[1,i]]
s2 <- sample.list[[n]][pair.matrix[2,i]]
fil[[n]][[i]] <- file.path(path,paste(which.plot,"_",
status,"_",n,"_",s1,"_",s2,".",output,sep=""))
names(fil[[n]][i]) <- paste(s1,"vs",s2,sep="_")
graphics.open(output,fil[[n]][[i]])
MDPlot(s,y=pair.matrix[,i],main=paste("MD plot for ",n," ",
status," samples ",s1," and ",s2,sep=""),cex.main=0.9)
graphics.close(output)
}
}
},
meanvar = {
fil <- vector("list",length(sample.list))
names(fil) <- names(sample.list)
for (n in names(sample.list)) {
if (length(sample.list[[n]])==1) {
warnwrap("Cannot create a mean-variance plot with one ",
"sample per condition! Skipping...")
next
}
pair.matrix <- combn(1:length(sample.list[[n]]),2)
fil[[n]] <- vector("list",ncol(pair.matrix))
for (i in 1:ncol(pair.matrix)) {
s1 <- sample.list[[n]][pair.matrix[1,i]]
s2 <- sample.list[[n]][pair.matrix[2,i]]
fil[[n]][[i]] <- file.path(path,paste(which.plot,"_",status,
"_",n,"_",s1,"_",s2,".",output,sep=""))
names(fil[[n]][i]) <- paste(s1,"vs",s2,sep="_")
graphics.open(output,fil[[n]][[i]])
suppressWarnings(meanVarPlot(s,main=paste("MV plot for ",n,
" ",status," samples ",s1," and ",s2,sep=""),
cex.main=0.9))
graphics.close(output)
}
}
},
gcbias = {
if (!output=="json") {
fil <- file.path(path,paste(which.plot,"_",status,".",output,
sep=""))
graphics.open(output,fil)
biasPlot(s,"gc",xlim=c(0.1,0.9),log=TRUE,ylim=c(0,15),
main=paste("Expression - GC content ",status,sep=""))
grid()
graphics.close(output)
}
else {
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=sample.list,
ylim=NULL,
xlim=NULL,
status=status,
pcut=NULL,
fcut=NULL,
altnames=NULL,
user=list(counts=x,covar=covar,covarname="GC content")
)
json <- biasPlotToJSON(obj)
fil <- file.path(path,paste(which.plot,"_",status,".json",
sep=""))
disp("Writing ",fil)
write(json,fil)
}
},
lengthbias = {
if (output!="json") {
fil <- file.path(path,paste(which.plot,"_",status,".",output,
sep=""))
graphics.open(output,fil)
biasPlot(s,"length",log=TRUE,ylim=c(0,10),
main=paste("Expression - Gene length ",status,sep=""))
grid()
graphics.close(output)
}
else {
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=sample.list,
ylim=NULL,
xlim=NULL,
status=status,
pcut=NULL,
fcut=NULL,
altnames=NULL,
user=list(counts=x,covar=covar,
covarname="Gene/transcript length")
)
json <- biasPlotToJSON(obj)
fil <- file.path(path,paste(which.plot,"_",status,".json",
sep=""))
disp("Writing ",fil)
write(json,fil)
}
}
)
return(fil)
}
#' Diagnostic plots based on the NOISeq package
#'
#' A wrapper around the plotting functions availale in the NOISeq Bioconductor
#' package. For analytical explanation of each plot please see the vignette of
#' the NOISeq package. It is best to use this function through the main plotting
#' function \code{\link{diagplot.metaseqr}}.
#'
#' @param x the count data matrix.
#' @param sample.list the list containing condition names and the samples under
#' each condition.
#' @param covars a list (whose annotation elements are ideally a subset of an
#' annotation data frame produced by \code{\link{get.annotation}})
#' with the following members: data (the data matrix), length (gene length), gc
#' (the gene gc_content), chromosome (a data frame with chromosome name and
#' co-ordinates), factors (a factor with the experimental condition names
#' replicated by the number of samples in each experimental condition) and biotype
#' (each gene's biotype as depicted in Ensembl-like annotations).
#' @param which.plot the NOISeq package plot to generate. It can be one or more
#' of \code{"biodetection"}, \code{"countsbio"}, \code{"saturation"},
#' \code{"rnacomp"}, \code{"readnoise"} or \code{"biodist"}. Please refer to the
#' documentation of the EDASeq package for details on the use of these plots. The
#' \code{which.plot="saturation"} case is modified to be more informative by
#' producing two kinds of plots. See \code{\link{diagplot.noiseq.saturation}}.
#' @param biodist.opts a list with the following members: p (a vector of p-values,
#' e.g. the p-values of a contrast), pcut (a unique number depicting a p-value
#' cutoff, required for the \code{"biodist"} case), name (a name for the
#' \code{"biodist"} plot, e.g. the name of the contrast.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param is.norm a logical indicating whether object contains raw or normalized
#' data. It is not essential and it serves only plot annotation purposes.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filenames of the plots produced in a named list with names the
#' \code{which.plot} argument. If \code{output="x11"}, no output filenames are
#' produced.
#' @note Please note that in case of \code{"biodist"} plots, the behavior of the
#' function is unstable, mostly due to the very specific inputs this plotting
#' function accepts in the NOISeq package. We have tried to predict unstable
#' behavior and avoid exceptions through the use of tryCatch but it's still
#' possible that you might run onto an error.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' lengths <- round(1000*runif(nrow(data.matrix)))
#' starts <- round(1000*runif(nrow(data.matrix)))
#' ends <- starts + lengths
#' covars <- list(
#' data=data.matrix,
#' length=lengths,
#' gc=runif(nrow(data.matrix)),
#' chromosome=data.frame(
#' chromosome=c(rep("chr1",nrow(data.matrix)/2),rep("chr2",nrow(data.matrix)/2)),
#' start=starts,
#' end=ends
#' ),
#' factors=data.frame(class=as.class.vector(sample.list)),
#' biotype=c(rep("protein_coding",nrow(data.matrix)/2),rep("ncRNA",
#' nrow(data.matrix)/2))
#' )
#' p <- runif(nrow(data.matrix))
#' diagplot.noiseq(data.matrix,sample.list,covars=covars,
#' biodist.opts=list(p=p,pcut=0.1,name="A_vs_B"))
#'}
diagplot.noiseq <- function(x,sample.list,covars,which.plot=c("biodetection",
"countsbio","saturation","rnacomp","readnoise","biodist"),output="x11",
biodist.opts=list(p=NULL,pcut=NULL,name=NULL),path=NULL,is.norm=FALSE,
...) {
if (is.null(path)) path <- getwd()
# covars is a list of gc-content, factors, length, biotype, chromosomes,
# factors, basically copy of the noiseq object
which.plot <- tolower(which.plot[1])
check.text.args("which.plot",which.plot,c("biodetection","countsbio",
"saturation","readnoise","rnacomp","biodist"),multiarg=FALSE)
if (missing(covars))
stopwrap("\"covars\" argument is required with NOISeq specific plots!")
else {
covars$biotype <- as.character(covars$biotype)
names(covars$length) <- names(covars$gc) <-
rownames(covars$chromosome) <- names(covars$biotype) <-
rownames(x)
}
if (which.plot=="biodist") {
if (is.null(biodist.opts$p))
stopwrap("A p-value must be provided for the \"biodist\" plot!")
if (is.null(biodist.opts$pcut) || is.na(biodist.opts$pcut))
biodist.opts$pcut=0.05
}
if (is.norm)
status<- "normalized"
else
status<- "raw"
# All of these plots are NOISeq specific so we need a local NOISeq object
if (any(is.na(unique(covars$biotype))))
covars$biotype=NULL # Otherwise, it will probably crash
local.obj <- NOISeq::readData(
data=x,
length=covars$gene.length,
gc=covars$gc.content,
chromosome=covars$chromosome,
#factors=data.frame(class=covars$factors),
factors=covars$factors,
biotype=covars$biotype
)
switch(which.plot,
biodetection = {
diagplot.data <- NOISeq::dat(local.obj,type=which.plot)
samples <- unlist(sample.list)
if (output!="json") {
fil <- character(length(samples))
names(fil) <- samples
for (i in 1:length(samples)) {
fil[samples[i]] <- file.path(path,paste(which.plot,"_",
samples[i],".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil[samples[i]],width=9,height=7)
else
graphics.open(output,fil[samples[i]],width=1024,height=768)
explo.plot(diagplot.data,samples=i)
graphics.close(output)
}
}
else {
diagplot.data.save = NOISeq::dat2save(diagplot.data)
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=sample.list,
ylims=NULL,
xlims=NULL,
status=status,
pcut=NULL,
fcut=NULL,
altnames=covars$gene_name,
user=list(plotdata=diagplot.data.save,covars=covars)
)
json <- bioDetectionToJSON(obj)
fil <- character(length(samples))
names(fil) <- samples
for (i in 1:length(samples)) {
fil[samples[i]] <- file.path(path,
paste(which.plot,"_",samples[i],".json",sep=""))
disp("Writing ",fil[samples[i]])
write(json[[i]],fil[samples[i]])
}
}
},
countsbio = {
samples <- unlist(sample.list)
if (output!="json") {
diagplot.data <- NOISeq::dat(local.obj,type=which.plot,
factor=NULL)
fil <- character(length(samples))
names(fil) <- samples
for (i in 1:length(samples)) {
fil[samples[i]] <- file.path(path,paste(which.plot,"_",
samples[i],".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil[samples[i]],width=9,height=7)
else
graphics.open(output,fil[samples[i]],width=1024,
height=768)
explo.plot(diagplot.data,samples=i,plottype="boxplot")
graphics.close(output)
}
}
else {
colnames(x) <- unlist(sample.list)
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=sample.list,
ylims=NULL,
xlims=NULL,
status=status,
pcut=NULL,
fcut=NULL,
altnames=covars$gene_name,
user=list(counts=nat2log(x),covars=covars)
)
# Write JSON by sample
fil <- vector("list",2)
names(fil) <- c("sample","biotype")
fil[["sample"]] <- character(length(samples))
names(fil[["sample"]]) <- samples
bts <- unique(as.character(obj$user$covars$biotype))
fil[["biotype"]] <- character(length(bts))
names(fil[["biotype"]]) <- bts
json <- countsBioToJSON(obj,by="sample")
for (i in 1:length(samples)) {
fil[["sample"]][samples[i]] <- file.path(path,
paste(which.plot,"_",samples[i],".json",sep=""))
disp("Writing ",fil[["sample"]][samples[i]])
write(json[[i]],fil[["sample"]][samples[i]])
}
json <- countsBioToJSON(obj,by="biotype")
for (i in 1:length(bts)) {
fil[["biotype"]][bts[i]] <- file.path(path,
paste(which.plot,"_",bts[i],".json",sep=""))
disp("Writing ",fil[["biotype"]][bts[i]])
write(json[[i]],fil[["biotype"]][bts[i]])
}
}
},
saturation = {
# For 10 saturation points
diagplot.data <- NOISeq::dat(local.obj,k=0,ndepth=9,type=which.plot)
d2s <- NOISeq::dat2save(diagplot.data)
if (output != "json")
fil <- diagplot.noiseq.saturation(d2s,output,covars$biotype,
path=path)
else {
samples <- unlist(sample.list)
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=sample.list,
ylims=NULL,
xlims=NULL,
status=status,
pcut=NULL,
fcut=NULL,
altnames=covars$gene_name,
user=list(plotdata=d2s)
)
# Write JSON by sample
fil <- vector("list",2)
names(fil) <- c("sample","biotype")
fil[["sample"]] <- character(length(samples))
names(fil[["sample"]]) <- samples
json <- bioSaturationToJSON(obj,by="sample")
for (i in 1:length(samples)) {
fil[["sample"]][samples[i]] <- file.path(path,
paste(which.plot,"_",samples[i],".json",sep=""))
disp("Writing ",fil[["sample"]][samples[i]])
write(json[[i]],fil[["sample"]][samples[i]])
}
json <- bioSaturationToJSON(obj,by="biotype")
fil[["biotype"]] <- character(length(json))
names(fil[["biotype"]]) <- names(json)
for (n in names(json)) {
fil[["biotype"]][n] <- file.path(path,
paste(which.plot,"_",n,".json",sep=""))
disp("Writing ",fil[["biotype"]][n])
write(json[[n]],fil[["biotype"]][n])
}
}
},
rnacomp = {
if (ncol(local.obj)<3) {
warnwrap("RNA composition plot cannot be created with less ",
"than 3 samples! Skipping...")
return(NULL)
}
if (ncol(local.obj)>12) {
warnwrap("RNA composition plot cannot be created with more ",
"than 12 samples! Skipping...")
return(NULL)
}
diagplot.data <- NOISeq::dat(local.obj,type="cd")
fil <- file.path(path,paste(which.plot,"_",status,".",output,
sep=""))
graphics.open(output,fil)
explo.plot(diagplot.data)
grid()
graphics.close(output)
},
readnoise = {
D <- cddat(local.obj)
if (output!="json") {
fil <- file.path(path,paste(which.plot,".",output,sep=""))
graphics.open(output,fil)
cdplot(D,main="RNA-Seq reads noise")
grid()
graphics.close(output)
}
else {
colnames(D$data2plot)[2:ncol(D$data2plot)] <-
unlist(sample.list)
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=sample.list,
xlim=NULL,
ylim=NULL,
status=NULL,
pcut=NULL,
fcut=NULL,
altnames=NULL,
user=D$data2plot
)
json <- readNoiseToJSON(obj)
fil <- file.path(path,paste(which.plot,".json",sep=""))
disp("Writing ",fil)
write(json,fil)
}
},
biodist = { # We have to fake a noiseq object
p <- biodist.opts$p
if (is.matrix(p)) p <- p[,1]
dummy <- new("Output",
comparison=c("Dummy.1","Dummy.2"),
factor=c("class"),
k=1,
lc=1,
method="n",
replicates="biological",
results=list(
data.frame(
Dummy.1=rep(1,length(p)),
Dummy.2=rep(1,length(p)),
M=rep(1,length(p)),
D=rep(1,length(p)),
prob=as.numeric(p),
ranking=rep(1,length(p)),
Length=rep(1,length(p)),
GC=rep(1,length(p)),
Chrom=as.character(covars$chromosome[,1]),
GeneStart=covars$chromosome[,2],
GeneEnd=covars$chromosome[,3],
Biotype=covars$biotype
)
),
nss=5,
pnr=0.2,
v=0.02
)
if (!is.null(biodist.opts$name))
fil <- file.path(path,paste(which.plot,"_",biodist.opts$name,
".",output,sep=""))
else
fil <- file.path(path,paste(which.plot,".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=10,height=6)
else
graphics.open(output,fil,width=1024,height=640)
tryCatch( # A lot of times, there is a problem with this function
DE.plot(dummy,chromosomes=NULL,q=biodist.opts$pcut,
graphic="distr"),
error=function(e) {
disp(" Known problem with NOISeq and external ",
"p-values detected! Trying to make a plot with ",
"alternative p-values (median of p-value ",
"distribution)...")
fil="error"
tryCatch(
DE.plot(dummy,chromosomes=NULL,
q=quantile(biodist.opts$p,0.5),
graphic="distr"),
error=function(e) {
disp(" Cannot create DEG biotype plot! This ",
"is not related to a problem with the ",
"results. Excluding...")
fil="error"
},
finally=""
)
},
finally=""
)
graphics.close(output)
}
)
return(fil)
}
#' Simpler implementation of saturation plots inspired from NOISeq package
#'
#' Helper function for \code{\link{diagplot.noiseq}} to plot feature detection
#' saturation as presented in the NOISeq package vignette. It has two main outputs:
#' a set of figures, one for each input sample depicting the saturation for each
#' biotype and one single multiplot which depicts the saturation of all samples
#' for each biotype. It expands the saturation plots of NOISeq by allowing more
#' samples to be examined in a simpler way. Don't use this function directly. Use
#' either \code{\link{diagplot.metaseqr}} or \code{\link{diagplot.noiseq}}.
#'
#' @param x the count data matrix.
#' @param o one or more R plotting device to direct the plot result to. Supported
#' mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"}, \code{"bmp"},
#' \code{"pdf"} or \code{"ps"}.
#' @param tb the vector of biotypes, one for each row of x.
#' @param path the path to create output files.
#' @return The filenames of the plots produced in a named list with names the
#' \code{which.plot} argument. If \code{output="x11"}, no output filenames are
#' produced.
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' biotype=c(rep("protein_coding",nrow(data.matrix)/2),rep("ncRNA",
#' nrow(data.matrix)/2))
#' diagplot.noiseq.saturation(data.matrix,"x11",biotype)
#'}
diagplot.noiseq.saturation <- function(x,o,tb,path=NULL) {
if (is.null(path)) path <- getwd()
if (length(unique(tb))==1) {
warnwrap("Saturation plot cannot be created with only one biotype! ",
"Skipping...")
return(NULL)
}
total.biotypes <- table(tb)
the.biotypes <- names(tb)
biotypes <- colnames(x[[1]][,2:ncol(x[[1]])])
colspace <- c("red3","green4","blue2","orange3","burlywood",
"lightpink4","gold","skyblue","red2","green2","firebrick3",
"orange4","yellow4","skyblue3","tan4","gray40",
"brown2","darkgoldenrod","cyan3","coral2","cadetblue",
"bisque3","blueviolet","chocolate3","darkkhaki","dodgerblue")
pchspace <- c(rep(c(15,16,17,18),6),15)
# Plot all biotypes per sample
f.sample <- character(length(names(x)))
names(f.sample) <- names(x)
for (n in names(x)) {
f.sample[n] <- file.path(path,paste("saturation_",n,".",o,sep=""))
if (o %in% c("pdf","ps","x11"))
graphics.open(o,f.sample[n],width=10,height=7)
else
graphics.open(o,f.sample[n],width=1024,height=800)
y <- x[[n]]
sep <- match(c("global","protein_coding"),colnames(y))
yab <- cbind(y[,"depth"],y[,sep])
ynab <- y[,-sep]
colnames(yab)[1] <- colnames(ynab)[1] <- "depth"
xlim <- range(y[,"depth"])
ylim.ab <- range(yab[,2:ncol(yab)])
ylim.nab <- range(ynab[,2:ncol(ynab)])
par(cex.axis=0.9,cex.main=1,cex.lab=0.9,font.lab=2,font.axis=2,pty="m",
lty=2,lwd=1.5,mfrow=c(1,2))
plot.new()
plot.window(xlim,ylim.nab)
axis(1,at=pretty(xlim,10),labels=as.character(pretty(xlim,10)/1e+6))
axis(2,at=pretty(ylim.nab,10))
title(main="Non abundant biotype detection saturation",
xlab="Depth in millions of reads",ylab="Detected features")
co <- 0
for (b in biotypes) {
co <- co + 1
if (b=="global" || b=="protein_coding") {
# Silently do nothing
}
else {
lines(ynab[,"depth"],ynab[,b],col=colspace[co])
points(ynab[,"depth"],ynab[,b],pch=pchspace[co],
col=colspace[co],cex=1)
}
}
grid()
graphics::legend(
x="topleft",legend=colnames(ynab)[2:ncol(ynab)],xjust=1,yjust=0,
box.lty=0,x.intersp=0.5,cex=0.6,text.font=2,
col=colspace[1:(ncol(ynab)-1)],pch=pchspace[1:(ncol(ynab)-1)]
)
plot.new()
plot.window(xlim,ylim.ab)
axis(1,at=pretty(xlim,10),labels=as.character(pretty(xlim,10)/1e+6))
axis(2,at=pretty(ylim.ab,10))
title(main="Abundant biotype detection saturation",
xlab="Depth in millions of reads",ylab="Detected features")
co <- 0
for (b in c("global","protein_coding")) {
co <- co + 1
lines(yab[,"depth"],yab[,b],col=colspace[co])
points(yab[,"depth"],yab[,b],pch=16,col=colspace[co],cex=1.2)
}
grid()
graphics::legend(
x="topleft",legend=c("global","protein_coding"),xjust=1,yjust=0,
box.lty=0,lty=2,x.intersp=0.5,cex=0.7,text.font=2,
col=colspace[1:2],pch=pchspace[1:2]
)
mtext(n,side=3,line=-1.5,outer=TRUE,font=2,cex=1.3)
graphics.close(o)
}
# Plot all samples per biotype
g <- make.grid(length(biotypes))
f.all <- file.path(path,paste("biotype_saturation.",o,sep=""))
if (o %in% c("pdf","ps"))
graphics.open(o,f.all,width=14,height=14)
else
graphics.open(o,f.all,width=1600,height=1600,res=150)
par(cex.axis=0.8,cex.main=0.9,cex.lab=0.8,pty="m",lty=2,lwd=1.5,mfrow=g,
mar=c(3,3,1,1),oma=c(1,1,0,0),mgp=c(2,0.5,0))
for (b in biotypes) {
y <- depth <- vector("list",length(x))
names(y) <- names(depth) <- names(x)
for (n in names(x)) {
y[[n]] <- x[[n]][,b]
depth[[n]] <- x[[n]][,"depth"]
}
y <- do.call("cbind",y)
xlim <- range(do.call("c",depth))
ylim <- range(y)
plot.new()
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,5),labels=as.character(pretty(xlim,5)/1e+6),
line=0.5)
axis(2,at=pretty(ylim,5),line=0.5)
title(main=b,xlab="Depth in millions of reads",
ylab="Detected features")
co <- 0
for (n in colnames(y)) {
co <- co + 1
lines(depth[[n]],y[,n],col=colspace[co])
points(depth[[n]],y[,n],pch=pchspace[co],col=colspace[co])
}
grid()
graphics::legend(
x="bottomright",legend=colnames(y),xjust=1,yjust=0,
box.lty=0,x.intersp=0.5,
col=colspace[1:length(colnames(y))],
pch=pchspace[1:length(colnames(y))]
)
}
graphics.close(o)
return(list(sample=f.sample,biotype=f.all))
}
#' (Interactive) volcano plots of differentially expressed genes
#'
#' This function plots a volcano plot or returns a JSON string which is used to
#' render aninteractive in case of HTML reporting.
#'
#' @param f the fold changes which are to be plotted on the x-axis.
#' @param p the p-values whose -log10 transformation is going to be plotted on
#' the y-axis.
#' @param con an optional string depicting a name (e.g. the contrast name) to
#' appear in the title of the volcano diagplot.
#' @param fcut a fold change cutoff so as to draw two vertical lines indicating
#' the cutoff threshold for biological significance.
#' @param pcut a p-value cutoff so as to draw a horizontal line indicating the
#' cutoff threshold for statistical significance.
#' @param alt.names an optional vector of names, e.g. HUGO gene symbols, alternative
#' or complementary to the unique names of \code{f} or \code{p} (one of them must
#' be named!). It is used only in JSON output.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"}, \code{"ps"} or \code{"json"}. The latter is currently
#' available for the creation of interactive volcano plots only when reporting the
#' output, through the highcharts javascript library.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filenames of the plots produced in a named list with names the
#' \code{which.plot} argument. If \code{output="x11"}, no output filenames are
#' produced.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' M <- norm.edger(data.matrix,sample.list)
#' p <- stat.edger(M,sample.list,contrast)
#' ma <- apply(M[,sample.list$A],1,mean)
#' mb <- apply(M[,sample.list$B],1,mean)
#' f <- log2(ifelse(mb==0,1,mb)/ifelse(ma==0,1,ma))
#' diagplot.volcano(f,p,con=contrast)
#' j <- diagplot.volcano(f,p,con=contrast,output="json")
#'}
diagplot.volcano <- function(f,p,con=NULL,fcut=1,pcut=0.05,alt.names=NULL,
output="x11",path=NULL,...) { # output can be json here...
## Check rjson
#if ("json" %in% output && !require(rjson))
# stopwrap("R package rjson is required to create interactive volcano plot!")
if (is.null(path)) path <- getwd()
if (is.null(con))
con <- conn <- ""
else {
conn <- con
con <- paste("for ",con)
}
fil <- file.path(path,paste("volcano_plot_",conn,".",output,sep=""))
if (output!="json") {
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=8,height=10)
else
graphics.open(output,fil,width=768,height=1024,res=100)
}
rem <- which(is.na(p))
if (length(rem)>0) {
p <- p[-rem]
f <- f[-rem]
if (!is.null(alt.names))
alt.names <- alt.names[-rem]
}
# Fix problem with extremely low p-values, only for display purposes though
p.zero <- which(p==0)
if (length(p.zero)>0)
p[p.zero] <- runif(length(p.zero),0,1e-256)
xlim <- c(-max(abs(f)),max(abs(f)))
ylim <- c(0,ceiling(-log10(min(p))))
up <- which(f>=fcut & p<pcut)
down <- which(f<=-fcut & p<pcut)
u <- union(up,down)
alt.names.neutral <- NULL
if (length(u)>0) {
ff <- f[-u]
pp <- p[-u]
if (!is.null(alt.names))
alt.names.neutral <- alt.names[-u]
}
else {
ff <- f
pp <- p
if (!is.null(alt.names))
alt.names.neutral <- alt.names
}
if (output!="json") {
par(cex.main=1.1,cex.lab=1.1,cex.axis=1.1,font.lab=2,font.axis=2,
pty="m",lwd=1.5)
plot.new()
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,10),labels=as.character(pretty(xlim,10)))
axis(2,at=pretty(ylim,10))
title(paste(main="Volcano plot",con),
xlab="Fold change",ylab="-log10(p-value)")
points(ff,-log10(pp),pch=20,col="blue2",cex=0.9)
points(f[down],-log10(p[down]),pch=20,col="green3",cex=0.9)
points(f[up],-log10(p[up]),pch=20,col="red2",cex=0.9)
abline(h=-log10(pcut),lty=4)
abline(v=-fcut,lty=2)
abline(v=fcut,lty=2)
grid()
graphics::legend(
x="topleft",
legend=c("up-regulated","down-regulated","unregulated",
"p-value threshold","fold change threshold"),
col=c("red2","green3","blue1","black","black"),
pch=c(20,20,20,NA,NA),lty=c(NA,NA,NA,4,2),
xjust=1,yjust=0,box.lty=0,x.intersp=0.5,cex=0.8,text.font=2
)
graphics.close(output)
}
else {
obj <- list(
x=f,
y=p,
plot=NULL,
samples=NULL,
xlim=xlim,
ylim=ylim,
status=NULL,
pcut=pcut,
fcut=fcut,
altnames=alt.names,
user=list(up=up,down=down,unf=ff,unp=pp,ualt=alt.names.neutral,
con=con)
)
#json <- volcanoToJSON(obj)
#fil <- file.path(path,paste("volcano_",con,".json",sep=""))
#write(json,fil)
fil <- volcanoToJSON(obj)
}
return(fil)
}
#' Diagnostic heatmap of differentially expressed genes
#'
#' This function plots a heatmap of the differentially expressed genes produced
#' by the metaseqr workflow, useful for quality control, e.g. whether samples
#' belonging to the same group cluster together.
#'
#' @param x the data matrix to create a heatmap for.
#' @param con an optional string depicting a name (e.g. the contrast name) to
#' appear in the title of the volcano plot.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"}, \code{"ps"}.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filenames of the plots produced in a named list with names the
#' \code{which.plot} argument. If \code{output="x11"}, no output filenames are
#' produced.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' require(DESeq)
#' data.matrix <- counts(makeExampleCountDataSet())
#' sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
#' contrast <- "A_vs_B"
#' M <- norm.edger(data.matrix,sample.list)
#' p <- stat.edger(M,sample.list,contrast)
#' diagplot.de.heatmap(data.matrix[p[[1]]<0.05])
#'}
diagplot.de.heatmap <- function(x,con=NULL,output="x11",path=NULL,...) {
if (is.null(path)) path <- getwd()
if (is.null(con))
con <- conn <- ""
else {
conn <- con
con <- paste("for ",con)
}
y <- nat2log(x,2,1)
# First plot the normal image
fil <- file.path(path,paste("de_heatmap_",conn,".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=10,height=10)
else
graphics.open(output,fil,width=800,height=800)
heatmap.2(y,trace="none",col=bluered(16),labRow="",cexCol=0.9,keysize=1,
font.lab=2,main=paste("DEG heatmap",con),cex.main=0.9)
graphics.close(output)
## Then the "interactive" using sendplot
#xy.labels <- list(normalized_counts=x,log2_normalized_counts=y)
#x.labels <- data.frame(
# label=colnames(x),
# description=paste("Sample",colnames(x))
#)
#y.labels <- data.frame(
# label=rownames(x),
# description=paste("Gene ID:",rownames(x))
#)
#suppressWarnings(heatmap.send(
# y,
# distfun=dist,
# hclustfun=hclust,
# MainColor=bluered(16),
# labRow="",
# labCol=NULL,
# keep.dendro=TRUE,
# x.labels=x.labels,
# y.labels=y.labels,
# xy.labels=xy.labels,
# image.size="2048x4096",
# fname.root=paste("iframe_de_heatmap_",conn,sep=""),
# dir=paste(path,.Platform$file.sep,sep=""),
# header="v3",
# window.size="2048x4192"
#))
return(fil)
}
#' Diagnostic plot for filtered genes
#'
#' This function plots a grid of four graphs depicting: in the first row, the
#' numbers of filtered genes per chromosome in the first column and per biotype
#' in the second column. In the second row, the percentages of filtered genes
#' per chromosome related to the whole genome in the first columns and per biotype
#' in the second column.
#'
#' @param x an annotation data frame like the ones produced by
#' \code{\link{get.annotation}}. \code{x} should be the filtered annotation
#' according to metaseqR's filters.
#' @param y an annotation data frame like the ones produced by
#' \code{\link{get.annotation}}. \code{y} should contain the total annotation
#' without the application of any metaseqr filter.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filenames of the plots produced in a named list with names the
#' \code{which.plot} argument. If output=\code{"x11"}, no output filenames are
#' produced.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' y <- get.annotation("mm9","gene")
#' x <- y[-sample(1:nrow(y),10000),]
#' diagplot.filtered(x,y)
#'}
diagplot.filtered <- function(x,y,output="x11",path=NULL,...) {
if (output !="json") {
if (is.null(path)) path <- getwd()
fil <- file.path(path,paste("filtered_genes.",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=12,height=8)
else
graphics.open(output,fil,width=1200,height=800,res=100)
chr <- table(as.character(x$chromosome))
bt <- table(as.character(x$biotype))
chr.all <- table(as.character(y$chromosome))
bt.all <- table(as.character(y$biotype))
barlab.chr <- as.character(chr)
barlab.bt <- as.character(bt)
per.chr <- chr/chr.all[names(chr)]
per.bt <- bt/bt.all[names(bt)]
# Some bug...
per.chr[per.chr>1] <- 1
per.bt[per.bt>1] <- 1
#
suppressWarnings(per.chr.lab <- paste(formatC(100*per.chr,digits=1,
format="f"),"%",sep=""))
suppressWarnings(per.bt.lab <- paste(formatC(100*per.bt,digits=1,
format="f"),"%",sep=""))
par(mfrow=c(2,2),mar=c(1,4,2,1),oma=c(1,1,1,1))
# Chromosomes
barx.chr <- barplot(chr,space=0.5,
ylim=c(0,max(chr)+ceiling(max(chr)/10)),yaxt="n",xaxt="n",
plot=FALSE)
plot.new()
plot.window(xlim=c(0,ceiling(max(barx.chr))),
ylim=c(0,max(chr)+ceiling(max(chr)/10)),mar=c(1,4,1,1))
axis(2,at=pretty(0:(max(chr)+ceiling(max(chr)/10))),cex.axis=0.9,padj=1,
font=2)
text(x=barx.chr,y=chr,label=barlab.chr,cex=0.7,font=2,col="green3",
adj=c(0.5,-1.3))
title(main="Filtered genes per chromosome",cex.main=1.1)
mtext(side=2,text="Number of genes",line=2,cex=0.9,font=2)
grid()
barplot(chr,space=0.5,ylim=c(0,max(chr)+ceiling(max(chr)/10)),
col="blue3",border="yellow3",yaxt="n",xaxt="n",font=2,add=TRUE)
# Biotypes
barx.bt <- barplot(bt,space=0.5,ylim=c(0,max(bt)+ceiling(max(bt)/10)),
yaxt="n",xaxt="n",plot=FALSE)
plot.new()
plot.window(xlim=c(0,ceiling(max(barx.bt))),
ylim=c(0,max(bt)+ceiling(max(bt)/10)),mar=c(1,4,1,1))
axis(2,at=pretty(0:(max(bt)+ceiling(max(bt)/10))),cex.axis=0.9,padj=1,
font=2)
text(x=barx.bt,y=bt,label=barlab.bt,cex=0.7,font=2,col="blue",
adj=c(0.5,-1.3),xpd=TRUE)
title(main="Filtered genes per biotype",cex.main=1.1)
mtext(side=2,text="Number of genes",line=2,cex=0.9,font=2)
grid()
barplot(bt,space=0.5,ylim=c(0,max(bt)+ceiling(max(bt)/10)),col="red3",
border="yellow3",yaxt="n",xaxt="n",font=2,add=TRUE)
# Chromosome percentage
barx.per.chr <- barplot(per.chr,space=0.5,ylim=c(0,max(per.chr)),
yaxt="n",xaxt="n",plot=FALSE)
plot.new()
par(mar=c(9,4,1,1))
plot.window(xlim=c(0,max(barx.per.chr)),ylim=c(0,max(per.chr)))
#axis(1,at=barx.per.chr,labels=names(per.chr),cex.axis=0.9,font=2,
# tcl=-0.3,col="lightgrey",las=2)
axis(1,at=barx.per.chr,labels=FALSE,tcl=-0.3,col="lightgrey")
axis(2,at=seq(0,max(per.chr),length.out=5),labels=formatC(seq(0,
max(per.chr),length.out=5),digits=2,format="f"),cex.axis=0.9,padj=1,
font=2)
text(barx.per.chr,par("usr")[3]-max(per.chr)/17,labels=names(per.chr),
srt=45,adj=c(1,1.1),xpd=TRUE,cex=0.9,font=2)
text(x=barx.per.chr,y=per.chr,label=per.chr.lab,cex=0.7,font=2,
col="green3",adj=c(0.5,-1.3),xpd=TRUE)
mtext(side=2,text="fraction of total genes",line=2,cex=0.9,font=2)
grid()
barplot(per.chr,space=0.5,ylim=c(0,max(per.chr)),col="blue3",
border="yellow3",yaxt="n",xaxt="n",font=2,add=TRUE)
# Biotype percentage
barx.per.bt <- barplot(per.bt,space=0.5,ylim=c(0,max(per.bt)),yaxt="n",
xaxt="n",plot=FALSE)
plot.new()
par(mar=c(9,4,1,1))
plot.window(xlim=c(0,max(barx.per.bt)),ylim=c(0,max(per.bt)))
#axis(1,at=barx.per.bt,labels=names(per.bt),cex.axis=0.9,font=2,
# tcl=-0.3,col="lightgrey",las=2)
axis(1,at=barx.per.bt,labels=FALSE,tcl=-0.3,col="lightgrey")
axis(2,at=seq(0,max(per.bt),length.out=5),
labels=formatC(seq(0,max(per.bt),length.out=5),digits=2,format="f"),
cex.axis=0.9,padj=1,font=2)
text(barx.per.bt,par("usr")[3]-max(per.bt)/17,
labels=gsub("prime","'",names(per.bt)),srt=45,adj=c(1,1.1),
xpd=TRUE,cex=0.9,font=2)
text(x=barx.per.bt,y=per.bt,label=per.bt.lab,cex=0.7,font=2,col="blue",
adj=c(0.5,-1.3),xpd=TRUE)
mtext(side=2,text="fraction of total genes",line=2,cex=0.9,font=2)
grid()
barplot(per.bt,space=0.5,ylim=c(0,max(per.bt)),col="red3",
border="yellow3",yaxt="n",xaxt="n",font=2,add=TRUE)
graphics.close(output)
}
else {
obj <- list(
x=NULL,
y=NULL,
plot=NULL,
samples=NULL,
xlim=NULL,
ylim=NULL,
status=NULL,
pcut=NULL,
fcut=NULL,
altnames=NULL,
user=list(filtered=x,total=y)
)
fil <- list(chromosome=NULL,biotype=NULL)
json <- filteredToJSON(obj,by="chromosome")
fil$chromosome <- file.path(path,"filtered_genes_chromosome.json")
write(json,fil$chromosome)
json <- filteredToJSON(obj,by="biotype")
fil$biotype <- file.path(path,"filtered_genes_biotype.json")
write(json,fil$biotype)
}
return(fil)
}
#' Venn diagrams when performing meta-analysis
#'
#' This function uses the R package VennDiagram and plots an up to 5-way Venn
#' diagram depicting the common and specific to each statistical algorithm genes,
#' for each contrast. Mostly for internal use because of its main argument which
#' is difficult to construct, but can be used independently if the user grasps
#' the logic.
#'
#' @param pmat a matrix with p-values corresponding to the application of each
#' statistical algorithm. The p-value matrix must have the colnames attribute and
#' the colnames should correspond to the name of the algorithm used to fill the
#' specific column (e.g. if \code{"statistics"=c("deseq","edger","nbpseq")} then
#' \code{colnames(pmat) <-} \code{c("deseq","edger","nbpseq")}.
#' @param fcmat an optional matrix with fold changes corresponding to the application
#' of each statistical algorithm. The fold change matrix must have the colnames
#' attribute and the colnames should correspond to the name of the algorithm used
#' to fill the specific column (see the parameter \code{pmat}).
#' @param pcut a p-value cutoff for statistical significance. Defaults to
#' \code{0.05}.
#' @param fcut if \code{fcmat} is supplied, an absolute fold change cutoff to be
#' applied to \code{fcmat} to determine the differentially expressed genes for
#' each algorithm.
#' @param direction if \code{fcmat} is supplied, a keyword to denote which genes
#' to draw in the Venn diagrams with respect to their direction of regulation. It
#' can be one of \code{"dereg"} for the total of regulated genes, where
#' \code{abs(fcmat[,n])>=fcut} (default), \code{"up"} for the up-regulated genes
#' where \code{fcmat[,n]>=fcut} or \code{"down"} for the up-regulated genes where
#' \code{fcmat[,n]<=-fcut}.
#' @param nam a name to be appended to the output graphics file (if \code{"output"}
#' is not \code{"x11"}).
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param alt.names an optional named vector of names, e.g. HUGO gene symbols,
#' alternative or complementary to the unique gene names which are the rownames
#' of \code{pmat}. The names of the vector must be the rownames of \code{pmat}.
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return The filenames of the plots produced in a named list with names the
#' \code{which.plot} argument. If output=\code{"x11"}, no output filenames are
#' produced. If \code{"path"} is not \code{NULL}, a file with the intersections
#' in the Venn diagrams will be produced and written in \code{"path"}.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p1 <- 0.001*matrix(runif(300),100,3)
#' p2 <- matrix(runif(300),100,3)
#' p <- rbind(p1,p2)
#' rownames(p) <- paste("gene",1:200,sep="_")
#' colnames(p) <- paste("method",1:3,sep="_")
#' venn.contents <- diagplot.venn(p)
#'}
diagplot.venn <- function(pmat,fcmat=NULL,pcut=0.05,fcut=0.5,
direction=c("dereg","up","down"),nam=as.character(round(1000*runif(1))),
output="x11",path=NULL,alt.names=NULL,...) {
check.text.args("direction",direction,c("dereg","up","down"))
if (is.na(pcut) || is.null(pcut) || pcut==1)
warnwrap("Illegal pcut argument! Using the default (0.05)")
algs <- colnames(pmat)
if (is.null(algs))
stopwrap("The p-value matrices must have the colnames attribute ",
"(names of statistical algorithms)!")
if (!is.null(fcmat) && (is.null(colnames(fcmat)) ||
length(intersect(colnames(pmat),colnames(fcmat)))!=length(algs)))
stopwrap("The fold change matrices must have the colnames attribute ",
"(names of statistical algorithms) and must be the same as in the ",
"p-value matrices!")
nalg <- length(algs)
if(nalg>5) {
warnwrap(paste("Cannot create a Venn diagram for more than 5 result ",
"sets! ",nalg,"found, only the first 5 will be used..."))
algs <- algs[1:5]
nalg <- 5
}
lenalias <- c("two","three","four","five")
aliases <- toupper(letters[1:nalg])
names(algs) <- aliases
genes <- rownames(pmat)
pairs <- make.venn.pairs(algs)
areas <- make.venn.areas(length(algs))
counts <- make.venn.counts(length(algs))
# Initially populate the results and counts lists so they can be used to create
# the rest of the intersections
results <- vector("list",nalg+length(pairs))
names(results)[1:nalg] <- aliases
names(results)[(nalg+1):length(results)] <- names(pairs)
if (is.null(fcmat)) {
for (a in aliases) {
results[[a]] <- genes[which(pmat[,algs[a]]<pcut)]
counts[[areas[[a]]]] <- length(results[[a]])
}
}
else {
switch(direction,
dereg = {
for (a in aliases) {
results[[a]] <-
genes[which(pmat[,algs[a]]<pcut & abs(
fcmat[,algs[a]])>=fcut)]
counts[[areas[[a]]]] <- length(results[[a]])
}
},
up = {
for (a in aliases) {
results[[a]] <-
genes[which(pmat[,algs[a]]<pcut &
fcmat[,algs[a]]>=fcut)]
counts[[areas[[a]]]] <- length(results[[a]])
}
},
down = {
for (a in aliases) {
results[[a]] <-
genes[which(pmat[,algs[a]]<pcut &
fcmat[,algs[a]]<=-fcut)]
counts[[areas[[a]]]] <- length(results[[a]])
}
}
)
}
# Now, perform the intersections
for (p in names(pairs)) {
a = pairs[[p]][1]
b = pairs[[p]][2]
results[[p]] <- intersect(results[[a]],results[[b]])
counts[[areas[[p]]]] <- length(results[[p]])
}
# And now, the Venn diagrams must be constructed
color.scheme <- make.venn.colorscheme(length(algs))
fil <- file.path(path,paste("venn_plot_",nam,".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=8,height=8)
else
graphics.open(output,fil,width=800,height=800,res=100)
switch(lenalias[length(algs)-1],
two = {
v <- draw.pairwise.venn(
area1=counts$area1,
area2=counts$area2,
cross.area=counts$cross.area,
category=paste(algs," (",aliases,")",sep=""),
lty="blank",
fill=color.scheme$fill,
cex=1.5,
cat.cex=1.3,
cat.pos=c(0,0),
cat.col=color.scheme$font,
#cat.dist=0.07,
cat.fontfamily=rep("Bookman",2)
)
},
three = {
#overrideTriple <<- TRUE
v <- draw.triple.venn(
area1=counts$area1,
area2=counts$area2,
area3=counts$area3,
n12=counts$n12,
n13=counts$n13,
n23=counts$n23,
n123=counts$n123,
category=paste(algs," (",aliases,")",sep=""),
lty="blank",
fill=color.scheme$fill,
cex=1.5,
cat.cex=1.3,
#cat.pos=c(0,0,180),
cat.col=color.scheme$font,
#cat.dist=0.07,
cat.fontfamily=rep("Bookman",3)
)
},
four = {
v <- draw.quad.venn(
area1=counts$area1,
area2=counts$area2,
area3=counts$area3,
area4=counts$area4,
n12=counts$n12,
n13=counts$n13,
n14=counts$n14,
n23=counts$n23,
n24=counts$n24,
n34=counts$n34,
n123=counts$n123,
n124=counts$n124,
n134=counts$n134,
n234=counts$n234,
n1234=counts$n1234,
category=paste(algs," (",aliases,")",sep=""),
lty="blank",
fill=color.scheme$fill,
cex=1.5,
cat.cex=1.3,
c(0.1,0.1,0.05,0.05),
cat.col=color.scheme$font,
cat.fontfamily=rep("Bookman",4)
)
},
five = {
v <- draw.quintuple.venn(
area1=counts$area1,
area2=counts$area2,
area3=counts$area3,
area4=counts$area4,
area5=counts$area5,
n12=counts$n12,
n13=counts$n13,
n14=counts$n14,
n15=counts$n15,
n23=counts$n23,
n24=counts$n24,
n25=counts$n25,
n34=counts$n34,
n35=counts$n35,
n45=counts$n45,
n123=counts$n123,
n124=counts$n124,
n125=counts$n125,
n134=counts$n134,
n135=counts$n135,
n145=counts$n145,
n234=counts$n234,
n235=counts$n235,
n245=counts$n245,
n345=counts$n345,
n1234=counts$n1234,
n1235=counts$n1235,
n1245=counts$n1245,
n1345=counts$n1345,
n2345=counts$n2345,
n12345=counts$n12345,
category=paste(algs," (",aliases,")",sep=""),
lty="blank",
fill=color.scheme$fill,
cex=1.5,
cat.cex=1.3,
cat.dist=0.1,
cat.col=color.scheme$font,
cat.fontfamily=rep("Bookman",5)
)
}
)
graphics.close(output)
# Now do something with the results
if (!is.null(path)) {
results.ex <- vector("list",length(results))
names(results.ex) <- names(results)
if (!is.null(alt.names)) {
for (n in names(results))
results.ex[[n]] <- alt.names[results[[n]]]
}
else {
for (n in names(results))
results.ex[[n]] <- results[[n]]
}
max.len <- max(sapply(results.ex,length))
for (n in names(results.ex)) {
if (length(results.ex[[n]])<max.len) {
dif <- max.len - length(results.ex[[n]])
results.ex[[n]] <- c(results.ex[[n]],rep(NA,dif))
}
}
results.ex <- do.call("cbind",results.ex)
write.table(results.ex,file=file.path(path,"..","..","lists",
paste0("venn_categories_",nam,".txt")),sep="\t",
row.names=FALSE,quote=FALSE,na="")
}
return(fil)
}
#' Helper for Venn diagrams
#'
#' This function creates a list of pairwise comparisons to be performed in order
#' to create an up to 5-way Venn diagram using the R package VennDiagram. Internal
#' use mostly.
#'
#' @param algs a vector with the names of the sets (up to length 5, if larger, it
#' will be truncated with a warning).
#' @return A list with as many pairs as the comparisons to be made for the
#' construction of the Venn diagram. The pairs are encoded with the uppercase
#' letters A through E, each one corresponding to order of the input sets.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' sets <- c("apple","pear","banana")
#' pairs <- make.venn.pairs(sets)
#'}
make.venn.pairs <- function(algs)
{
lenalias <- c("two","three","four","five")
switch(lenalias[length(algs)-1],
two = {
return(list(
AB=c("A","B")
))
},
three = {
return(list(
AB=c("A","B"),
AC=c("A","C"),
BC=c("B","C"),
ABC=c("AB","C")
))
},
four = {
return(list(
AB=c("A","B"),
AC=c("A","C"),
AD=c("A","D"),
BC=c("B","C"),
BD=c("B","D"),
CD=c("C","D"),
ABC=c("AB","C"),
ABD=c("AB","D"),
ACD=c("AC","D"),
BCD=c("BC","D"),
ABCD=c("ABC","D")
))
},
five = {
return(list(
AB=c("A","B"),
AC=c("A","C"),
AD=c("A","D"),
AE=c("A","E"),
BC=c("B","C"),
BD=c("B","D"),
BE=c("B","E"),
CD=c("C","D"),
CE=c("C","E"),
DE=c("D","E"),
ABC=c("AB","C"),
ABD=c("AB","D"),
ABE=c("AB","E"),
ACD=c("AC","D"),
ACE=c("AC","E"),
ADE=c("AD","E"),
BCD=c("BC","D"),
BCE=c("BC","E"),
BDE=c("BD","E"),
CDE=c("CD","E"),
ABCD=c("ABC","D"),
ABCE=c("ABC","E"),
ABDE=c("ABD","E"),
ACDE=c("ACD","E"),
BCDE=c("BCD","E"),
ABCDE=c("ABCD","E")
))
}
)
}
#' Helper for Venn diagrams
#'
#' This function creates a list with names the arguments of the Venn diagram
#' construction functions of the R package VennDiagram and list members the
#' internal encoding (uppercase letters A to E and combinations among then) used
#' to encode the pairwise comparisons to create the intersections needed for the
#' Venn diagrams. Internal use mostly.
#'
#' @param n the number of the sets used for the Venn diagram.
#' @return A named list, see descritpion.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' sets <- c("apple","pear","banana")
#' pairs <- make.venn.pairs(sets)
#' areas <- make.venn.areas(length(sets))
#'}
make.venn.areas <- function(n)
{
lenalias <- c("two","three","four","five")
switch(lenalias[n-1],
two = {
return(list(
A="area1",
B="area2",
AB="cross.area"
))
},
three = {
return(list(
A="area1",
B="area2",
C="area3",
AB="n12",
AC="n13",
BC="n23",
ABC="n123"
))
},
four = {
return(list(
A="area1",
B="area2",
C="area3",
D="area4",
AB="n12",
AC="n13",
AD="n14",
BC="n23",
BD="n24",
CD="n34",
ABC="n123",
ABD="n124",
ACD="n134",
BCD="n234",
ABCD="n1234"
))
},
five = {
return(list(
A="area1",
B="area2",
C="area3",
D="area4",
E="area5",
AB="n12",
AC="n13",
AD="n14",
AE="n15",
BC="n23",
BD="n24",
BE="n25",
CD="n34",
CE="n35",
DE="n45",
ABC="n123",
ABD="n124",
ABE="n125",
ACD="n134",
ACE="n135",
ADE="n145",
BCD="n234",
BCE="n235",
BDE="n245",
CDE="n345",
ABCD="n1234",
ABCE="n1235",
ABDE="n1245",
ACDE="n1345",
BCDE="n2345",
ABCDE="n12345"
))
}
)
}
#' Helper for Venn diagrams
#'
#' This function creates a list with names the arguments of the Venn diagram
#' construction functions of the R package VennDiagram and list members are
#' initially \code{NULL}. They are filled by the \code{\link{diagplot.venn}}
#' function. Internal use mostly.
#'
#' @param n the number of the sets used for the Venn diagram.
#' @return A named list, see descritpion.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' sets <- c("apple","pear","banana")
#' counts <- make.venn.counts(length(sets))
#'}
make.venn.counts <- function(n)
{
lenalias <- c("two","three","four","five")
switch(lenalias[n-1],
two = {
return(list(
area1=NULL,
area2=NULL,
cross.area=NULL
))
},
three = {
return(list(
area1=NULL,
area2=NULL,
area3=NULL,
n12=NULL,
n13=NULL,
n23=NULL,
n123=NULL
))
},
four = {
return(list(
area1=NULL,
area2=NULL,
area3=NULL,
area4=NULL,
n12=NULL,
n13=NULL,
n14=NULL,
n23=NULL,
n24=NULL,
n34=NULL,
n123=NULL,
n124=NULL,
n134=NULL,
n234=NULL,
n1234=NULL
))
},
five = {
return(list(
area1=NULL,
area2=NULL,
area3=NULL,
area4=NULL,
area5=NULL,
n12=NULL,
n13=NULL,
n14=NULL,
n15=NULL,
n23=NULL,
n24=NULL,
n25=NULL,
n34=NULL,
n35=NULL,
n45=NULL,
n123=NULL,
n124=NULL,
n125=NULL,
n134=NULL,
n135=NULL,
n145=NULL,
n234=NULL,
n235=NULL,
n245=NULL,
n345=NULL,
n1234=NULL,
n1235=NULL,
n1245=NULL,
n1345=NULL,
n2345=NULL,
n12345=NULL
))
}
)
}
#' Helper for Venn diagrams
#'
#' This function returns a list of colorschemes accroding to the number of sets.
#' Internal use.
#'
#' @param n the number of the sets used for the Venn diagram.
#' @return A list with colors for fill and font.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' sets <- c("apple","pear","banana")
#' cs <- make.venn.colorscheme(length(sets))
#'}
make.venn.colorscheme <- function(n) {
lenalias <- c("two","three","four","five")
switch(lenalias[n-1],
two = {
return(list(
fill=c("blue","orange2"),
font=c("darkblue","orange4")
))
},
three = {
return(list(
fill=c("red","green","mediumpurple"),
font=c("darkred","darkgreen","mediumpurple4")
))
},
four = {
return(list(
fill=c("red","green","mediumpurple","orange2"),
font=c("darkred","darkgreen","mediumpurple4","orange4")
))
},
five = {
return(list(
fill=c("red","green","blue","mediumpurple","orange2"),
font=c("darkred","darkgreen","darkblue","mediumpurple4",
"orange4")
))
}
)
}
#' Create basic ROC curves
#'
#' This function creates basic ROC curves using a matrix of p-values (such a matrix
#' can be derived for example from the result table of \code{\link{metaseqr}} by
#' subsetting the table to get the p-values from several algorithms) given a ground
#' truth vector for differential expression and a significance level.
#'
#' @param truth the ground truth differential expression vector. It should contain
#' only zero and non-zero elements, with zero denoting non-differentially expressed
#' genes and non-zero, differentially expressed genes. Such a vector can be obtained
#' for example by using the \code{\link{make.sim.data.sd}} function, which creates
#' simulated RNA-Seq read counts based on real data.
#' @param p a p-value matrix whose rows correspond to each element in the
#' \code{truth} vector. If the matrix has a \code{colnames} attribute, a legend
#' will be added to the plot using these names, else a set of column names will
#' be auto-generated. \code{p} can also be a list or a data frame.
#' @param sig a significance level (0 < \code{sig} <=1).
#' @param x what to plot on x-axis, can be one of \code{"fpr"}, \code{"fnr"},
#' \code{"tpr"}, \code{"tnr"} for False Positive Rate, False Negative Rate, True
#' Positive Rate and True Negative Rate respectively.
#' @param y what to plot on y-axis, same as \code{x} above.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param draw boolean to determine whether to plot the curves or just return the
#' calculated values (in cases where the user wants the output for later averaging
#' for example). Defaults to \code{TRUE} (make plots).
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return A named list with two members. The first member is a list containing
#' the ROC statistics: \code{TP} (True Postives), \code{FP} (False Positives),
#' \code{FN} (False Negatives), \code{TN} (True Negatives), \code{FPR} (False
#' Positive Rate), \code{FNR} (False Negative Rate), \code{TPR} (True Positive
#' Rate), \code{TNR} (True Negative Rate), \code{AUC} (Area Under the Curve). The
#' second is the path to the created figure graphic.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p1 <- 0.001*matrix(runif(300),100,3)
#' p2 <- matrix(runif(300),100,3)
#' p <- rbind(p1,p2)
#' rownames(p) <- paste("gene",1:200,sep="_")
#' colnames(p) <- paste("method",1:3,sep="_")
#' truth <- c(rep(1,40),rep(-1,40),rep(0,20),rep(1,10),rep(2,10),rep(0,80))
#' names(truth) <- rownames(p)
#' roc.obj <- diagplot.roc(truth,p)
#'}
diagplot.roc <- function(truth,p,sig=0.05,x="fpr",y="tpr",output="x11",
path=NULL,draw=TRUE,...) {
check.text.args("x",x,c("fpr","fnr","tpr","tnr","scrx","sens","spec"),
multiarg=FALSE)
check.text.args("y",y,c("fpr","fnr","tpr","tnr","scry","sens","spec"),
multiarg=FALSE)
if (is.list(p))
pmat <- do.call("cbind",p)
else if (is.data.frame(p))
pmat <- as.matrix(p)
else if (is.matrix(p))
pmat <- p
if (is.null(colnames(pmat)))
colnames(pmat) <- paste("p",1:ncol(pmat),sep="_")
ax.name <- list(
tpr="True Positive Rate",
tnr="True Negative Rate",
fpr="False Positive Rate",
fnr="False Negative Rate",
scrx="Ratio of selected",
scry="Normalized TP/(FP+FN)",
sens="Sensitivity",
spec="1 - Specificity"
)
ROC <- vector("list",ncol(pmat))
names(ROC) <- colnames(pmat)
colspace.universe <- c("red","blue","green","orange","darkgrey","green4",
"black","pink","brown","magenta","yellowgreen","pink4","seagreen4",
"darkcyan")
colspace <- colspace.universe[1:ncol(pmat)]
names(colspace) <- colnames(pmat)
eps <- min(pmat[!is.na(pmat) & pmat>0])
for (n in colnames(pmat)) {
disp("Processing ",n)
gg <- which(pmat[,n]<=sig)
psample <- -log10(pmax(pmat[gg,n],eps))
#psample <- pmat[gg,n]
size <- seq(1,length(gg))
cuts <- seq(-log10(sig),max(psample),length.out=length(gg))
#cuts <- seq(min(psample),sig,length.out=length(gg))
local.truth <- truth[gg]
S <- length(size)
TP <- FP <- FN <- TN <- FPR <- FNR <- TPR <- TNR <- SENS <- SPEC <-
SCRX <- SCRY <- numeric(S)
for (i in 1:S) {
TP[i] <- length(which(psample>cuts[i] & local.truth!=0))
FP[i] <- length(which(psample>cuts[i] & local.truth==0))
FN[i] <- length(which(psample<cuts[i] & local.truth!=0))
TN[i] <- length(which(psample<cuts[i] & local.truth==0))
## Alternatives which I keep in the code
#TP[i] <- length(intersect(names(which(psample>cuts[i])),
# names(which(local.truth!=0))))
#FP[i] <- length(intersect(names(which(psample>cuts[i])),
# names(which(local.truth==0))))
#FN[i] <- length(intersect(names(which(psample<cuts[i])),
# names(which(local.truth!=0))))
#TN[i] <- length(intersect(names(which(psample<cuts[i])),
# names(which(local.truth==0))))
#bad <- which(psample<cuts[i])
#good <- which(psample>cuts[i])
#TP[i] <- length(which(local.truth[good]!=0))
#FP[i] <- length(which(local.truth[good]==0))
#TN[i] <- length(which(local.truth[bad]==0))
#FN[i] <- length(which(local.truth[bad]!=0))
SCRX[i] <- i/S
SCRY[i] <- TP[i]/(FN[i]+FP[i])
if (FP[i]+TN[i] == 0)
FPR[i] <- 0
else
FPR[i] <- FP[i]/(FP[i]+TN[i])
FNR[i] <- FN[i]/(TP[i]+FN[i])
TPR[i] <- TP[i]/(TP[i]+FN[i])
if (TN[i]+FP[i] == 0)
TNR[i] <- 0
else
TNR[i] <- TN[i]/(TN[i]+FP[i])
SENS[i] <- TPR[i]
SPEC[i] <- 1 - TNR[i]
}
#if (all(FPR==0))
# FPR[length(FPR)] <- 1
#if (all(TNR==0)) {
# TNR[1] <- 1
# SPEC[i] <- 0
#}
ROC[[n]] <- list(TP=TP,FP=FP,FN=FN,TN=TN,
FPR=FPR,FNR=FNR,TPR=TPR,TNR=TNR,SCRX=SCRX,SCRY=SCRY/max(SCRY),
SENS=SENS,SPEC=SPEC,AUC=NULL)
}
for (n in colnames(pmat)) {
disp("Calculating AUC for ",n)
auc <- 0
for (i in 2:length(ROC[[n]][[toupper(y)]])) {
auc <- auc +
0.5*(ROC[[n]][[toupper(x)]][i]-ROC[[n]][[toupper(x)]][i-1])*
(ROC[[n]][[toupper(y)]][i]+ROC[[n]][[toupper(y)]][i-1])
}
ROC[[n]]$AUC <- abs(auc)
# There are some extreme cases, with the Intersection case for the paper
# where there are no FPs or TNs for a p-value cutoff of 0.2 (which is
# imposed in order to avoid the saturation of the ROC curves). In these
# cases, performance is virtually perfect, and the actual AUC should be
# 1. For these cases, we set it to a value between 0.95 and 0.99 to
# represent a more plausible truth.
if (ROC[[n]]$AUC==0) ROC[[n]]$AUC <- sample(seq(0.95,0.99,by=0.001),1)
}
disp("")
if (draw) {
fil <- file.path(path,paste("ROC",output,sep="."))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=8,height=8)
else
graphics.open(output,fil,width=1024,height=1024,res=100)
xlim <- c(0,1)
ylim <- c(0,1)
par(cex.axis=0.9,cex.main=1,cex.lab=0.9,font.lab=2,font.axis=2,pty="m",
lwd=1.5,lty=1)
plot.new()
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,10))
axis(2,at=pretty(ylim,10))
for (n in names(ROC))
lines(ROC[[n]][[toupper(x)]],ROC[[n]][[toupper(y)]],
col=colspace[n],...)
grid()
title(xlab=ax.name[[x]],ylab=ax.name[[y]])
auc.text <- as.character(sapply(ROC,function(x)
round(x$AUC,digits=3)))
graphics::legend(x="bottomright",col=colspace,lty=1,cex=0.9,
legend=paste(names(ROC)," (AUC = ",auc.text,")",sep=""))
graphics.close(output)
}
else
fil <- NULL
return(list(ROC=ROC,truth=truth,sig.level=sig,x.axis=x,y.axis=y,path=fil))
}
## Create averaged basic ROC curves
##
## This function creates averaged basic ROC curves using a list of objects returned
## from the \code{\link{diagplot.roc}} function.
##
## @param roc.obj a list containing several lists returned from the application
## of \code{\link{diagplot.roc}} function.
## @param output one or more R plotting device to direct the plot result to.
## Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
## \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
## @param path the path to create output files.
## @param ... further arguments to be passed to plot devices, such as parameter
## from \code{\link{par}}.
## @return A named list with two members. The first member is a list containing
## the mean and standard deviation of ROC statistics. The second is the path to
## the created figure graphic.
## @export
## @author <NAME>
## @examples
## \dontrun{
## # Not yet available
##}
#diagplot.avg.roc <- function(roc.obj,output="x11",path=NULL,...) {
# ax.name <- list(
# tpr="True Positive Rate",
# tnr="True Negative Rate",
# fpr="False Positive Rate",
# fnr="False Negative Rate"
# )
# stats <- names(roc.obj[[1]]$ROC)
# x <- toupper(roc.obj[[1]]$x.axis)
# y <- toupper(roc.obj[[1]]$y.axis)
# avg.ROC <- vector("list",length(stats))
# avg.ROC <- lapply(avg.ROC,function(x) {
# return(list(TP=NULL,FP=NULL,FN=NULL,TN=NULL,
# FPR=NULL,FNR=NULL,TPR=NULL,TNR=NULL,AUC=NULL))
# })
# names(avg.ROC) <- stats
# colspace.universe <- c("red","blue","green","orange","darkgrey","green4",
# "black","pink","brown","yellowgreen","magenta","pink2","seagreen4",
# "darkcyan")
# colspace <- colspace.universe[1:length(stats)]
# names(colspace) <- stats
# for (s in stats) {
# disp("Retrieving ",s)
# for (r in names(avg.ROC[[s]])) {
# if (r != "AUC") {
# #avg.ROC[[s]][[r]] <- do.call("cbind",lapply(roc.obj,
# # function(x,s,r) x$ROC[[s]][[r]],s,r))
# lapply(roc.obj,function(x,s,r) print(length(x$ROC[[s]][[r]])),s,r)
# mn <- apply(avg.ROC[[s]][[r]],1,mean)
# st <- apply(avg.ROC[[s]][[r]],1,sd)
# avg.ROC[[s]][[r]] <- list(mean=mn,sd=st)
# }
# }
# }
# disp("")
# means <- do.call("cbind",lapply(avg.ROC,function(x) x$mean))
# stds <- do.call("cbind",lapply(avg.ROC,function(x) x$sd))
# fil <- file.path(path,paste("ROC",output,sep="."))
# if (output %in% c("pdf","ps","x11"))
# graphics.open(output,fil,width=8,height=8)
# else
# graphics.open(output,fil,width=1024,height=1024,res=100)
# xlim <- c(0,1)
# ylim <- c(0,1)
# par(cex.axis=0.9,cex.main=1,cex.lab=0.9,font.lab=2,font.axis=2,pty="m",
# lwd=1.5,lty=1)
# plot.new()
# plot.window(xlim,ylim)
# axis(1,at=pretty(xlim,10))
# axis(2,at=pretty(ylim,10))
# for (n in names(ROC)) {
# lines(ROC[[n]][[x]],ROC[[n]][[y]],col=colspace[n],...)
# }
# grid()
# title(xlab=ax.name[[x]],ylab=ax.name[[y]])
# legend(x="bottomright",legend=names(ROC),col=colspace,lty=1)
# graphics.close(output)
# return(list(ROC=ROC,path=fil))
#}
#' Create False (or True) Positive (or Negative) curves
#'
#' This function creates false (or true) discovery curves using a matrix of
#' p-values (such a matrix can be derived for example from the result table of
#' \code{\link{metaseqr}} by subsetting the table to get the p-values from several
#' algorithms) given a ground truth vector for differential expression.
#'
#' @param truth the ground truth differential expression vector. It should contain
#' only zero and non-zero elements, with zero denoting non-differentially expressed
#' genes and non-zero, differentially expressed genes. Such a vector can be obtained
#' for example by using the \code{\link{make.sim.data.sd}} function, which creates
#' simulated RNA-Seq read counts based on real data. The elements of \code{truth}
#' MUST be named (e.g. each gene's name).
#' @param p a p-value matrix whose rows correspond to each element in the
#' \code{truth} vector. If the matrix has a \code{colnames} attribute, a legend
#' will be added to the plot using these names, else a set of column names will
#' be auto-generated. \code{p} can also be a list or a data frame. The p-values
#' MUST be named (e.g. each gene's name).
#' @param type what to plot, can be \code{"fpc"} for False Positive Curves
#' (default), \code{"tpc"} for True Positive Curves, \code{"fnc"} for False
#' Negative Curves or \code{"tnc"} for True Negative Curves.
#' @param N create the curves based on the top (or bottom) \code{N} ranked genes
#' (default is 2000) to be used with \code{type="fpc"} or \code{type="tpc"}.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param draw boolean to determine whether to plot the curves or just return the
#' calculated values (in cases where the user wants the output for later averaging
#' for example). Defaults to \code{TRUE} (make plots).
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return A named list with two members: the first member (\code{ftdr}) contains
#' the values used to create the plot. The second member (\code{path}) contains
#' the path to the created figure graphic.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p1 <- 0.001*matrix(runif(300),100,3)
#' p2 <- matrix(runif(300),100,3)
#' p <- rbind(p1,p2)
#' rownames(p) <- paste("gene",1:200,sep="_")
#' colnames(p) <- paste("method",1:3,sep="_")
#' truth <- c(rep(1,40),rep(-1,40),rep(0,10),rep(1,10),rep(2,10),rep(0,80))
#' names(truth) <- rownames(p)
#' ftd.obj <- diagplot.ftd(truth,p,N=100)
#'}
diagplot.ftd <- function(truth,p,type="fpc",N=2000,output="x11",path=NULL,
draw=TRUE,...) {
check.text.args("type",type,c("fpc","tpc","fnc","tnc"),multiarg=FALSE)
if (is.list(p))
pmat <- do.call("cbind",p)
else if (is.data.frame(p))
pmat <- as.matrix(p)
else if (is.matrix(p))
pmat <- p
else if (is.numeric(p))
pmat <- as.matrix(p)
if (is.null(colnames(pmat)))
colnames(pmat) <- paste("p",1:ncol(pmat),sep="_")
y.name <- list(
tpc="Number of True Positives",
fpc="Number of False Positives",
tnc="Number of True Negatives",
fnc="Number of False Negatives"
)
ftdr.list <- vector("list",ncol(pmat))
names(ftdr.list) <- colnames(pmat)
colspace.universe <- c("red","blue","green","orange","darkgrey","green4",
"black","pink","brown","magenta","yellowgreen","pink4","seagreen4",
"darkcyan")
colspace <- colspace.universe[1:ncol(pmat)]
names(colspace) <- colnames(pmat)
switch(type,
fpc = {
for (n in colnames(pmat)) {
disp("Processing ",n)
z <- sort(pmat[,n])
for (i in 1:N) {
nn <- length(intersect(names(z[1:i]),
names(which(truth==0))))
if (nn==0)
ftdr.list[[n]][i] <- 1
else
ftdr.list[[n]][i] <- nn
}
}
},
tpc = {
for (n in colnames(pmat)) {
disp("Processing ",n)
z <- sort(pmat[,n])
for (i in 1:N)
ftdr.list[[n]][i] <- length(intersect(names(z[1:i]),
names(which(truth!=0))))
}
},
fnc = {
for (n in colnames(pmat)) {
disp("Processing ",n)
z <- sort(pmat[,n],decreasing=TRUE)
for (i in 1:N) {
nn <- length(intersect(names(z[1:i]),
names(which(truth!=0))))
if (nn==0)
ftdr.list[[n]][i] <- 1
else
ftdr.list[[n]][i] <- nn
}
}
},
tnc = {
for (n in colnames(pmat)) {
disp("Processing ",n)
z <- sort(pmat[,n],decreasing=TRUE)
for (i in 1:N)
ftdr.list[[n]][i] <- length(intersect(names(z[1:i]),
names(which(truth==0))))
}
}
)
disp("")
if (draw) {
fil <- file.path(path,paste("FTDR_",type,".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=8,height=8)
else
graphics.open(output,fil,width=1024,height=1024,res=100)
xlim <- ylim <- c(1,N)
#ylim <- c(1,length(which(truth!=0)))
par(cex.axis=0.9,cex.main=1,cex.lab=0.9,font.lab=2,font.axis=2,pty="m",
lwd=1.5,lty=1)
plot.new()
switch(type,
fpc = {
plot.window(xlim,ylim,log="y")
axis(1,at=pretty(xlim,10))
axis(2)
for (n in names(ftdr.list)) {
lines(ftdr.list[[n]],col=colspace[n],...)
}
grid()
title(main="Selected genes vs False Positives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="topleft",legend=names(ftdr.list),
col=colspace,lty=1)
},
tpc = {
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,10))
axis(2,at=pretty(ylim,10))
for (n in names(ftdr.list)) {
lines(ftdr.list[[n]],col=colspace[n],...)
}
grid()
title(main="Selected genes vs True Positives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="bottomright",legend=names(ftdr.list),
col=colspace,lty=1)
},
fnc = {
plot.window(xlim,ylim,log="y")
axis(1,at=pretty(xlim,10))
axis(2)
for (n in names(ftdr.list)) {
lines(ftdr.list[[n]],col=colspace[n],...)
}
grid()
title(main="Selected genes vs False Negatives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="topleft",legend=names(ftdr.list),
col=colspace,lty=1)
},
tnc = {
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,10))
axis(2,at=pretty(ylim,10))
for (n in names(ftdr.list)) {
lines(ftdr.list[[n]],col=colspace[n],...)
}
grid()
title(main="Selected genes vs True Negatives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="bottomright",legend=names(ftdr.list),
col=colspace,lty=1)
}
)
graphics.close(output)
}
else
fil <- NULL
return(list(ftdr=ftdr.list,truth=truth,type=type,N=N,path=fil))
}
#' Create average False (or True) Discovery curves
#'
#' This function creates false (or true) discovery curves using a list containing
#' several outputs from \code{\link{diagplot.ftd}}.
#'
#' @param ftdr.obj a list with outputs from \code{\link{diagplot.ftd}}.
#' @param output one or more R plotting device to direct the plot result to.
#' Supported mechanisms: \code{"x11"} (default), \code{"png"}, \code{"jpg"},
#' \code{"bmp"}, \code{"pdf"} or \code{"ps"}.
#' @param path the path to create output files.
#' @param draw boolean to determine whether to plot the curves or just return the
#' calculated values (in cases where the user wants the output for later averaging
#' for example). Defaults to \code{TRUE} (make plots).
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @return A named list with two members: the first member (\code{avg.ftdr})
#' contains a list with the means and the standard deviations of the averaged
#' \code{ftdr.obj} and are used to create the plot. The second member (\code{path})
#' contains the path to the created figure graphic.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p11 <- 0.001*matrix(runif(300),100,3)
#' p12 <- matrix(runif(300),100,3)
#' p21 <- 0.001*matrix(runif(300),100,3)
#' p22 <- matrix(runif(300),100,3)
#' p31 <- 0.001*matrix(runif(300),100,3)
#' p32 <- matrix(runif(300),100,3)
#' p1 <- rbind(p11,p21)
#' p2 <- rbind(p12,p22)
#' p3 <- rbind(p31,p32)
#' rownames(p1) <- rownames(p2) <- rownames(p3) <-
#' paste("gene",1:200,sep="_")
#' colnames(p1) <- colnames(p2) <- colnames(p3) <-
#' paste("method",1:3,sep="_")
#' truth <- c(rep(1,40),rep(-1,40),rep(0,20),
#' rep(1,10),rep(2,10),rep(0,80))
#' names(truth) <- rownames(p1)
#' ftd.obj.1 <- diagplot.ftd(truth,p1,N=100,draw=FALSE)
#' ftd.obj.2 <- diagplot.ftd(truth,p2,N=100,draw=FALSE)
#' ftd.obj.3 <- diagplot.ftd(truth,p3,N=100,draw=FALSE)
#' ftd.obj <- list(ftd.obj.1,ftd.obj.2,ftd.obj.3)
#' avg.ftd.obj <- diagplot.avg.ftd(ftd.obj)
#'}
diagplot.avg.ftd <- function(ftdr.obj,output="x11",path=NULL,draw=TRUE,...) {
y.name <- list(
tpc="Number of True Positives",
fpc="Number of False Positives",
tnc="Number of True Negatives",
fnc="Number of False Negatives"
)
stats <- names(ftdr.obj[[1]]$ftdr)
type <- ftdr.obj[[1]]$type
truth <- ftdr.obj[[1]]$truth
N <- ftdr.obj[[1]]$N
avg.ftdr.obj <- vector("list",length(stats))
names(avg.ftdr.obj) <- stats
colspace.universe <- c("red","blue","green","orange","darkgrey","green4",
"black","pink","brown","magenta","yellowgreen","pink4","seagreen4",
"darkcyan")
colspace <- colspace.universe[1:length(stats)]
names(colspace) <- stats
for (s in stats) {
disp("Retrieving ",s)
avg.ftdr.obj[[s]] <- do.call("cbind",lapply(ftdr.obj,
function(x) x$ftdr[[s]]))
}
disp("")
avg.ftdr.obj <- lapply(avg.ftdr.obj,function(x) {
mn <- apply(x,1,mean)
st <- apply(x,1,sd)
return(list(mean=mn,sd=st))
})
means <- do.call("cbind",lapply(avg.ftdr.obj,function(x) x$mean))
stds <- do.call("cbind",lapply(avg.ftdr.obj,function(x) x$sd))
if (draw) {
fil <- file.path(path,paste("AVG_FTDR_",type,".",output,sep=""))
if (output %in% c("pdf","ps","x11"))
graphics.open(output,fil,width=8,height=8)
else
graphics.open(output,fil,width=1024,height=1024,res=100)
xlim <- ylim <- c(1,N)
par(cex.axis=0.9,cex.main=1,cex.lab=0.9,font.lab=2,font.axis=2,pty="m",
lwd=1.5,lty=1)
plot.new()
switch(type,
fpc = {
plot.window(xlim,ylim,log="y")
axis(1,at=pretty(xlim,10))
axis(2)
for (n in colnames(means)) {
lines(means[,n],col=colspace[n],...)
}
grid()
title(main="Selected genes vs False Positives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="topleft",legend=colnames(means),
col=colspace,lty=1)
},
tpc = {
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,10))
axis(2,at=pretty(ylim,10))
for (n in colnames(means)) {
lines(means[,n],col=colspace[n],...)
}
grid()
title(main="Selected genes vs True Positives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="bottomright",legend=colnames(means),
col=colspace,lty=1)
},
fnc = {
plot.window(xlim,ylim,log="y")
axis(1,at=pretty(xlim,10))
axis(2)
for (n in colnames(means)) {
lines(means[,n],col=colspace[n],...)
}
grid()
title(main="Selected genes vs False Negatives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="topleft",legend=colnames(means),
col=colspace,lty=1)
},
tnc = {
plot.window(xlim,ylim)
axis(1,at=pretty(xlim,10))
axis(2,at=pretty(ylim,10))
for (n in colnames(means)) {
lines(means[,n],col=colspace[n],...)
}
grid()
title(main="Selected genes vs True Negatives",
xlab="Number of selected genes",ylab=y.name[[type]])
graphics::legend(x="bottomright",legend=colnames(means),
col=colspace,lty=1)
}
)
graphics.close(output)
}
else
fil <- NULL
return(list(avg.ftdr=list(means=means,stds=stds),path=fil))
}
#' Open plotting device
#'
#' Wrapper function to open a plotting device. Internal use only.
#'
#' @param o the plotting device, see main metaseqr function
#' @param f a filename, if the plotting device requires it (e.g. \code{"pdf"})
#' @param ... further arguments to be passed to plot devices, such as parameter
#' from \code{\link{par}}.
#' @author <NAME>
#' @examples
#' \dontrun{
#' graphics.open("pdf","test.pdf",width=12,height=12)
#'}
graphics.open <- function(o,f,...) {
if(!check.graphics.type(o))
stopwrap("Invalid graphics output type!")
if(check.graphics.file(o) && is.null(f))
stopwrap("Please specify an output file name for your plot")
switch(o,
x11 = { dev.new(...) },
pdf = { pdf(file=f,pointsize=10,...) },
ps = { postscript(file=f,pointsize=10,...) },
png = { png(filename=f,pointsize=12,...) },
jpg = { jpeg(filename=f,pointsize=12,quality=100,...) },
bmp = { bmp(filename=f,pointsize=12,...) },
tiff = { tiff(filename=f,pointsize=12,...) }
)
}
#' Close plotting device
#'
#' Wrapper function to close a plotting device. Internal use only.
#'
#' @param o the plotting device, see main metaseqr function
#' @author <NAME>
#' @examples
#' \dontrun{
#' graphics.close("pdf")
#'}
graphics.close <- function(o) {
if (!is.element(o,c("x11","png","jpg","tiff","bmp","pdf","ps")))
return(FALSE)
if (o!="x11")
dev.off()
}
#' Check plotting device
#'
#' Plotting device checker. Internal use only.
#'
#' @param o the plotting device, see main metaseqr function
#' @author <NAME>
check.graphics.type <- function(o) {
if (!is.element(o,c("x11","png","jpg","tiff","bmp","pdf","ps")))
return(FALSE)
else
return(TRUE)
}
#' Check graphics file
#'
#' Graphics file checker. Internal use only.
#'
#' @param o the plotting device, see main metaseqr function
#' @author <NAME>
check.graphics.file <- function(o) {
if (is.element(o,c("png","jpg","tiff","bmp","pdf","ps")))
return(TRUE)
else
return(FALSE)
}
#' Display value transformation
#'
#' Logarithmic transformation for display purposes. Internal use only.
#'
#' @param mat input data matrix
#' @param base logarithmic base, 2 or 10
#' @author <NAME>
log2disp <- function(mat,base=2) {
mat[mat==0] <- 1
if (base==10)
return(log10(mat))
else
return(log2(mat))
}
#' General value transformation
#'
#' Logarithmic transformation. Internal use only.
#'
#' @param x input data matrix
#' @param base logarithmic base, 2 or 10
#' @param off offset to avoid Infinity
#' @author <NAME>
nat2log <- function(x,base=2,off=1) {
#x[x==0] <- off
x <- x + off
if (base==2)
return(log2(x))
else
return(log10(x))
}
#' Old functions from NOISeq
#'
#' Old functions from NOISeq to create the \code{"readnoise"} plots. Internal use
#' only.
#'
#' @param input input to cddat.
#' @return a list with data to plot.
#' @note Adopted from an older version of NOISeq package (author: <NAME>).
#' @author <NAME>
cddat <- function (input) {
if (inherits(input,"eSet") == FALSE)
stopwrap("The input data must be an eSet object.\n")
if (!is.null(assayData(input)$exprs)) {
if (ncol(assayData(input)$exprs) < 2)
stopwrap("The input object should have at least two samples.\n")
datos <- assayData(input)$exprs
}
else {
if (ncol(assayData(input)$counts) < 2)
stopwrap("The input object should have at least two samples.\n")
datos <- assayData(input)$counts
}
datos <- datos[which(rowSums(datos) > 0),]
nu <- nrow(datos) # number of detected features
qq <- 1:nu
data2plot = data.frame("%features" = 100*qq/nu)
for (i in 1:ncol(datos)) {
acumu <- 100*cumsum(sort(datos[,i],decreasing=TRUE))/sum(datos[,i])
data2plot = cbind(data2plot, acumu)
}
colnames(data2plot)[-1] = colnames(datos)
# Diagnostic test
KSpval = mostres = NULL
for (i in 1:(ncol(datos)-1)) {
for (j in (i+1):ncol(datos)) {
mostres = c(mostres, paste(colnames(datos)[c(i,j)], collapse="_"))
KSpval = c(KSpval, suppressWarnings(ks.test(datos[,i], datos[,j],
alternative = "two.sided"))$"p.value")
}
}
KSpval = p.adjust(KSpval, method = "fdr")
return(list(
"data2plot"=data2plot,
"DiagnosticTest"=data.frame("ComparedSamples"=mostres,"KSpvalue"=KSpval)
))
}
#' Old functions from NOISeq
#'
#' Old functions from NOISeq to create the \code{"readnoise"} plots. Internal use
#' only.
#' @param dat the returned list from \code{\link{cddat}}.
#' @param samples the samples to plot.
#' @param ... further arguments passed to e.g. \code{\link{par}}.
#' @return Nothing, it created the old RNA composition plot.
#' @note Adopted from an older version of NOISeq package (author: <NAME>)
#' @author <NAME>
cdplot <- function (dat,samples=NULL,...) {
dat = dat$data2plot
if (is.null(samples)) samples <- 1:(ncol(dat)-1)
if (is.numeric(samples)) samples = colnames(dat)[samples+1]
colspace <- c("red","blue","yellowgreen","orange","aquamarine2","pink2",
"seagreen4","brown","purple","chocolate","gray10","gray30","darkblue",
"darkgreen","firebrick2","darkorange4","darkorchid","darkcyan","gold4",
"deeppink3")
if (length(samples)>length(colspace))
miscolores <- sample(colspace,length(samples),replace=TRUE)
else
miscolores <- sample(colspace,length(samples))
plot(dat[,1],dat[,samples[1]],xlab="% features",ylab="% reads",type="l",
col=miscolores[1],...)
for (i in 2:length(samples))
lines(dat[,1],dat[,samples[i]],col=miscolores[i])
graphics::legend("bottom",legend=samples,
text.col=miscolores[1:length(samples)],bty="n",lty=1,lwd=2,
col=miscolores[1:length(samples)])
}
<file_sep>/man/check.parallel.Rd
\name{check.parallel}
\alias{check.parallel}
\title{Parallel run validator}
\usage{
check.parallel(rc)
}
\arguments{
\item{rc}{fraction of available cores to use.}
}
\description{
Checks existence of multiple cores and loads multicore
package.
}
\examples{
multic <- check.parallel(0.8)
}
\author{
<NAME>
}
<file_sep>/man/check.libsize.Rd
\name{check.libsize}
\alias{check.libsize}
\title{Library size validator}
\usage{
check.libsize(libsize.list, sample.list)
}
\arguments{
\item{libsize.list}{the samples-names library size list.}
\item{sample.list}{the input sample list.}
}
\description{
Checks the names of the supplied library sizes. Internal
use only.
}
\examples{
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
libsize.list.1 <- list(A1=1e+6,A2=1.1e+6,B1=1.2e+6,
B2=1.3e+6,B3=1.5e+6)
libsize.list.2 <- list(A1=1e+6,A2=1.1e+6,B1=1.2e+6,
B2=1.3e+6)
check.libsize(libsize.list.1,sample.list) # Will work
#check.libsize(libsize.list.2,sample.list) # Will throw error!
}
\author{
<NAME>
}
<file_sep>/man/fisher.method.perm.Rd
\name{fisher.method.perm}
\alias{fisher.method.perm}
\title{
Derive a p-value for a summary statistic of p-values by permutation
}
\description{
Given a set of p-values and a summary statistic S: \deqn{S = -2 \sum
\log p,} a p-value for this statistic can be derived by randomly
generating summary statistics [Rhodes,2002]. Therefore, a p-value is
randomly sampled from each contributing study and a random statistic is
calculated. The fraction of random statistics greater or equal to S
then gives the p-value.
}
\usage{
fisher.method.perm(pvals, p.corr = c("bonferroni", "BH", "none"),
zero.sub = 1e-05, B = 10000, mc.cores = NULL, blinker = 1000)
}
\arguments{
\item{pvals}{
A matrix or data.frame containing the p-values from the single tests
}
\item{p.corr}{
Method for correcting the summary p-values. BH: Benjamini-Hochberg
(default); Bonferroni's method or no ('none') correction are currently
supported.
}
\item{zero.sub}{
Replacement for p-values of 0
}
\item{B}{
Number of random statistics
}
\item{mc.cores}{
Number of cores used for calculating the permutations. If not
\code{NULL} the \code{multicore} package is used for parallelization
with the given number of cores.
}
\item{blinker}{
An indicator that prints "\code{=}" after each \code{blinker} rows of
\code{pvals} in order to follow the progress.
}
}
\details{
At the moment this function only supports situations were all passed
p-values are not \code{NA}. We plan on extending this functionality
in upcoming versions.
For large data sets and/or large \code{B} we strongly recommend using
the \code{mc.cores} option as the calculation will otherwise be
computationally demanding. This will call the \code{mclapply}
function from the \cite{multicore} package, which you will have to
install in that case.
By default a blinker (a small string "=") is shown after each 1000
rows that were computed. This function allows you to assess the
progress of the analysis. If you don't want to see the blinker set
it to \code{NA}.
As \code{log(0)} results in \code{Inf} we replace p-values of 0 by
default with a small float. If you want to keep them as 0 you have
to provide 0 as a parameter in \code{zero.sub}.
Note that only p-values between 0 and 1 are allowed to be passed to
this method.
}
\value{
This method returns a data.frame containing the following columns
\item{S }{The statistic}
\item{num.p }{The number of p-values used to calculate S}
\item{p.value }{The overall p-value}
\item{p.adj}{The adjusted p-value}
}
\note{
This function was copied from the CRAN package MADAM which is no longer
maintained. Recognition goes to the original author(s) below.
}
\references{
<NAME>., (2002). Meta-analysis of microarrays: interstudy
alidation of gene expression profiles reveals pathway dysregulation
in prostate cancer. Cancer research, 62(15), 4427-33.
}
\author{
<NAME> <<EMAIL>>
}
\seealso{
\code{\link{fisher.sum}}, \code{\link{fisher.method}}
}
\examples{
set.seed(123)
pp <- matrix(c(runif(20),c(0.001,0.02,0.03,0.001)), ncol=4)
fisher.method.perm(pp, B=10, blinker=1)
\dontrun{
fisher.method.perm(pp, B=10000, mc.cores=3, blinker=1) #use multicore
}
}
<file_sep>/R/metaseqr.meta.R
#' Meta-analysis using several RNA-Seq statistics
#'
#' This function calculates the combined p-values when multiple statistical algorithms
#' are applied to the input dataset. It is a helper and it requires very specific
#' arguments so it should not be used individually.
#'
#' @param cp.list a named list whose names are the contrasts requested from metaseqR.
#' Each member is a p-value matrix whose colnames are the names of the statistical
#' tests applied to the data. See the main \code{\link{metaseqr}} help page.
#' @param meta.p the p-value combination method to use. See the main
#' \code{\link{metaseqr}} help page.
#' @param counts the normalized and possibly filtered read counts matrix. See the
#' main \code{\link{metaseqr}} help page.
#' @param sample.list the list containing condition names and the samples under
#' each condition. See the main \code{\link{metaseqr}} help page.
#' @param statistics the statistical algorithms used in metaseqr. See the main
#' \code{\link{metaseqr}} help page.
#' @param stat.args the parameters for each statistical argument. See the main
#' \code{\link{metaseqr}} help page.
#' @param libsize.list a list with library sizes. See the main \code{\link{metaseqr}}
#' and the \code{stat.*} help pages.
#' @param nperm the number of permutations (Monte Carlo simulations) to perform.
#' @param weight a numeric vector of weights for each statistical algorithm.
#' @param reprod create reproducible permutations. Ideally one would want to create
#' the same set of indices for a given dataset so as to create reproducible p-values.
#' If \code{reprod=TRUE}, a fixed seed is used by \code{meta.perm} for all the
#' datasets analyzed with \code{metaseqr}. If \code{reprod=FALSE}, then the
#' p-values will not be reproducible, although statistical significance is not
#' expected to change for a large number of resambling. Finally, \code{reprod}
#' can be a numeric vector of seeds with the same length as \code{nperm} so that
#' the user can supply his/her own seeds.
#' @param multic use multiple cores to execute the premutations. This is an
#' external parameter and implies the existence of parallel package in the execution
#' environment. See the main \code{\link{metaseqr}} help page.
#' @return A named list with combined p-values. The names are the contrasts and
#' the list members are combined p-value vectors, one for each contrast.
#' @author <NAME>
#' @examples
#' \dontrun{
#' # This function is not exported
#'}
meta.test <- function(cp.list,meta.p=c("simes","bonferroni","fisher",
"dperm.min","dperm.max","dperm.weight","fperm","whitlock","minp","maxp",
"weight","pandora","none"),counts,sample.list,statistics,stat.args,
libsize.list,nperm=10000,weight=rep(1/length(statistics),
length(statistics)),reprod=TRUE,multic=FALSE) {
check.text.args("meta.p",meta.p,c("simes","bonferroni","fisher","dperm.min",
"dperm.max","dperm.weight","fperm","whitlock","minp","maxp","weight",
"pandora","none"))
contrast <- names(cp.list)
disp("Performing meta-analysis with ",meta.p)
if (meta.p=="pandora")
meta.p <- "weight"
switch(meta.p,
fisher = {
sum.p.list <- wapply(multic,cp.list,function(x) {
tmp <- fisher.method(x,p.corr="none",
zero.sub=.Machine$double.xmin)
rp <- tmp$p.value
names(rp) <- rownames(x)
return(rp)
})
},
fperm = {
sum.p.list <- wapply(multic,cp.list,function(x) {
if (multic)
tmp <- fisher.method.perm(x,p.corr="none",B=nperm,
mc.cores=getOption("cores"),zero.sub=1e-32)
else
tmp <- fisher.method.perm(x,p.corr="none",B=nperm,
zero.sub=.Machine$double.xmin)
return(tmp$p.value)
})
},
whitlock = {
sum.p.list <- wapply(multic,cp.list,function(x)
return(apply(x,1,combine.test,method="z.transform")))
},
simes = {
sum.p.list <- wapply(multic,cp.list,function(x) {
return(apply(x,1,combine.simes))
})
},
bonferroni = {
sum.p.list <- wapply(multic,cp.list,function(x) {
return(apply(x,1,combine.bonferroni))
})
},
minp = {
sum.p.list <- wapply(multic,cp.list,function(x) {
return(apply(x,1,combine.minp))
})
},
maxp = {
sum.p.list <- wapply(multic,cp.list,function(x) {
return(apply(x,1,combine.maxp))
})
},
weight = {
sum.p.list <- wapply(multic,cp.list,function(x) {
return(apply(x,1,combine.weight,weight))
})
},
dperm.min = {
sum.p.list <- vector("list",length(cp.list))
names(sum.p.list) <- names(cp.list)
conl <- as.list(contrast)
names(conl) <- contrast
temp.p.list <- wapply(multic,conl,meta.perm,
counts=counts,sample.list=sample.list,
statistics=statistics,stat.args=stat.args,
libsize.list=libsize.list,
nperm=nperm,weight=weight,
select="min",reprod=reprod,
multic=multic)
original.p.list <- wapply(multic,cp.list,function(x,m,w=NULL) {
x[which(is.na(x))] <- 1
switch(m,
min = {
return(apply(x,1,min))
},
max = {
return(apply(x,1,max))
},
weight = {
return(apply(x,1,function(p,w) return(prod(p^w)),
w))
}
)
},"min")
for (cc in names(original.p.list))
{
pc <- cbind(temp.p.list[[cc]],original.p.list[[cc]])
ly <- ncol(pc)
sum.p.list[[cc]] <- apply(pc,1,function(y,m)
return(length(which(y[1:(m-1)]<y[m]))/(m-1)),ly)
}
#assign("perm.list",temp.p.list,envir=.GlobalEnv)
#assign("o.list",original.p.list,envir=.GlobalEnv)
},
dperm.max = {
sum.p.list <- vector("list",length(cp.list))
names(sum.p.list) <- names(cp.list)
conl <- as.list(contrast)
names(conl) <- contrast
temp.p.list <- wapply(multic,conl,meta.perm,
counts=counts,sample.list=sample.list,
statistics=statistics,stat.args=stat.args,
libsize.list=libsize.list,
nperm=nperm,weight=weight,
select="max",reprod=reprod,
multic=multic)
original.p.list <- wapply(multic,cp.list,function(x,m,w=NULL) {
switch(m,
min = {
return(apply(x,1,min))
},
max = {
return(apply(x,1,max))
},
weight = {
return(apply(x,1,function(p,w) return(prod(p^w)),
w))
}
)
},"max")
for (cc in names(original.p.list))
{
pc <- cbind(temp.p.list[[cc]],original.p.list[[cc]])
ly <- ncol(pc)
sum.p.list[[cc]] <- apply(pc,1,function(y,m)
return(length(which(y[1:(m-1)]<y[m]))/(m-1)),ly)
}
#assign("perm.list",temp.p.list,envir=.GlobalEnv)
#assign("o.list",original.p.list,envir=.GlobalEnv)
},
dperm.weight = {
sum.p.list <- vector("list",length(cp.list))
names(sum.p.list) <- names(cp.list)
conl <- as.list(contrast)
names(conl) <- contrast
temp.p.list <- wapply(multic,conl,meta.perm,
counts=counts,sample.list=sample.list,
statistics=statistics,stat.args=stat.args,
libsize.list=libsize.list,
nperm=nperm,weight=weight,
select="weight",reprod=reprod,
multic=multic)
original.p.list <- wapply(multic,cp.list,function(x,m,w=NULL) {
switch(m,
min = {
return(apply(x,1,min))
},
max = {
return(apply(x,1,max))
},
weight = {
return(apply(x,1,function(p,w) {return(prod(p^w))},
w))
}
)
},"weight",weight)
for (cc in names(original.p.list))
{
pc <- cbind(temp.p.list[[cc]],original.p.list[[cc]])
ly <- ncol(pc)
sum.p.list[[cc]] <- apply(pc,1,function(y,m)
return(length(which(y[1:(m-1)]<y[m]))/(m-1)),ly)
}
#assign("perm.list",temp.p.list,envir=.GlobalEnv)
#assign("o.list",original.p.list,envir=.GlobalEnv)
},
none = {
# A default value must be there to use with volcanos, we say the one
# of the first statistic in order of input
sum.p.list <- wapply(multic,cp.list,function(x) return(x[,1]))
}
)
return(sum.p.list)
}
#' Permutation tests for meta-analysis
#'
#' This function performs permutation tests in order to derive a meta p-value by
#' combining several of the statistical algorithms of metaseqr. This is probably
#' the most accurate way of combining multiple statistical algorithms for RNA-Seq
#' data, as this issue is different from the classic interpretation of the term
#' "meta-analysis" which implies the application of the same statistical test on
#' different datasets treating the same subject/experiment. For other methods, see
#' also the main \code{\link{metaseqr}} help page. You should keep in mind that
#' the permutation procedure can take a long time, even when executed in parallel.
#'
#' @param counts a normalized read counts table, one row for each gene, one column
#' for each sample.
#' @param sample.list the list containing condition names and the samples under
#' each condition. See the main \code{\link{metaseqr}} help page.
#' @param contrast the contrasts to be tested by each statistical algorithm. See
#' the main \code{\link{metaseqr}} help page.
#' @param statistics the statistical algorithms used in metaseqr. See the main
#' \code{\link{metaseqr}} help page.
#' @param stat.args the parameters for each statistical algorithm. See the main
#' \code{\link{metaseqr}} help page.
#' @param libsize.list a list with library sizes. See the main \code{\link{metaseqr}}
#' and the \code{stat.*} help pages.
#' @param nperm the number of permutations (Monte Carlo simulations) to perform.
#' @param weight a numeric vector of weights for each statistical algorithm.
#' @param select how to select the initial vector of p-values. It can be \code{"min"}
#' to select the minimum p-value for each gene (more conservative), \code{"max"}
#' to select the maximum p-value for each gene (less conservative), \code{"weight"}
#' to apply the weights to the p-value vector for each gene and derive a weighted
#' p-value.
#' @param replace same as the \code{replace} argument in the \code{\link{sample}}
#' function. Implies bootstraping or simple resampling without replacement. It can
#' also be \code{"auto"}, to determine bootstraping or not with the following rule:
#' if \code{ncol(counts)<=6} \code{replace=FALSE else} \code{replace=TRUE}. This
#' protects from the case of having zero variability across resampled conditions.
#' In such cases, most statistical tests would crash.
#' @param multic use multiple cores to execute the premutations. This is an
#' external parameter and implies the existence of parallel package in the
#' execution environment. See the main \code{\link{metaseqr}} help page.
#' @param reprod create reproducible permutations. Ideally one would want to
#' create the same set of indices for a given dataset so as to create reproducible
#' p-values. If \code{reprod=TRUE}, a fixed seed is used by \code{meta.perm} for
#' all the datasets analyzed with \code{metaseqr}. If \code{reprod=FALSE}, then
#' the p-values will not be reproducible, although statistical significance is not
#' expected to change for a large number of resambling. Finally, \code{reprod} can
#' be a numeric vector of seeds with the same length as \code{nperm} so that the
#' user can supply his/her own seeds.
#' @return A vector of meta p-values
#' @author <NAME>
#' @examples
#' \dontrun{
#' # This function is not exported
#'}
meta.perm <- function(contrast,counts,sample.list,statistics,stat.args,
libsize.list,nperm=10000,weight=rep(1/ncol(counts),ncol(counts)),
select=c("min","max","weight"),replace="auto",reprod=TRUE,multic=FALSE) {
check.text.args("select",select,c("min","max","weight"))
if (replace=="auto") {
if (ncol(counts)<=6)
replace=FALSE
else
replace=TRUE
}
# We will construct relist in a way so that we can assign seeds for random
# number generation and track progress at the same time
if (is.logical(reprod)) {
relist <- vector("list",nperm)
if (reprod) {
relist <- wapply(multic,seq_along(relist),function(i)
{return(list(seed=i,prog=i))})
}
else
relist <- wapply(multic,seq_along(relist),function(i)
{return(list(seed=round(1e+6*runif(1)),prog=i))})
}
else if (is.numeric(reprod)) {
if (length(reprod) != nperm)
stopwrap("When reprod is numeric, it must have the same length as ",
"nperm!")
relist <- wapply(multic,seq_along(reprod),function(i)
{return(list(seed=reprod[i],prog=i))})
}
else
stopwrap("reprod must be either a logical or a numeric vector!")
disp(" Resampling procedure started...")
# In this case, we must not use wapply as we want to be able to track progress
# through mc.preschedule...
if (multic)
pp <- mclapply(relist,meta.worker,counts,sample.list,contrast,
statistics,replace,stat.args,libsize.list,select,weight,
mc.preschedule=FALSE,mc.cores=getOption("cores"))
else
pp <- lapply(relist,meta.worker,counts,sample.list,contrast,statistics,
replace,stat.args,libsize.list,select,weight)
disp(" Resampling procedure ended...")
return(do.call("cbind",pp))
}
#' Permutation tests helper
#'
#' This function performs the statistical test for each permutation. Internal use
#' only.
#'
#' @param x a virtual list with the random seed and the permutation index.
#' @param co the counts matrix.
#' @param sl the sample list.
#' @param cnt the contrast name.
#' @param s the statistical algorithms.
#' @param sa the parameters for each statistical algorithm.
#' @param ll a list with library sizes.
#' @param r same as the \code{replace} argument in the \code{\link{sample}} function.
#' @param el min, max or weight.
#' @param w the weights when \code{el="weight"}.
#' @return A matrix of p-values.
#' @author <NAME>
#' @examples
#' \dontrun{
#' # This function is not exported
#'}
meta.worker <- function(x,co,sl,cnt,s,r,sa,ll,el,w) {
set.seed(x$seed)
disp(" running permutation #",x$prog)
pl <- make.permutation(co,sl,cnt,r)
ppmat <- matrix(NA,nrow(co),length(s))
colnames(ppmat) <- s
for (alg in s) {
#disp(" running permutation tests with: ",alg)
tcl <- make.contrast.list(pl$contrast,pl$sample.list)
switch(alg,
deseq = {
p.list <- suppressMessages(stat.deseq(pl$counts,pl$sample.list,
tcl,sa[[alg]]))
},
edger = {
p.list <- suppressMessages(stat.edger(pl$counts,pl$sample.list,
tcl,sa[[alg]]))
},
noiseq = {
p.list <- suppressMessages(stat.noiseq(pl$counts,pl$sample.list,
tcl,sa[[alg]]))
},
bayseq = {
p.list <- suppressMessages(stat.bayseq(pl$counts,pl$sample.list,
tcl,sa[[alg]],ll))
},
limma = {
p.list <- suppressMessages(stat.limma(pl$counts,pl$sample.list,
tcl,sa[[alg]]))
},
nbpseq = {
p.list <- suppressMessages(stat.nbpseq(pl$counts,pl$sample.list,
tcl,sa[[alg]],ll))
}
)
ppmat[,alg] <- as.numeric(p.list[[1]])
}
ppmat[which(is.na(ppmat))] <- 1
switch(el,
min = {
p.iter <- apply(ppmat,1,min)
},
max = {
p.iter <- apply(ppmat,1,max)
},
weight = {
p.iter <- apply(ppmat,1,function(p,w) return(prod(p^w)),w)
}
)
return(p.iter)
}
#' Combine p-values with Simes' method
#'
#' This function combines p-values from the various statistical tests supported by
#' metaseqR using the Simes' method (see reference in the main \code{\link{metasqr}}
#' help page or in the vignette).
#'
#' @param p a p-value matrix (rows are genes, columns are statistical tests).
#' @return A vector of combined p-values.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p <- matrix(runif(300),100,3)
#' pc <- combine.simes(p)
#'}
combine.simes <- function(p) {
ze <- which(p==0)
if (length(ze)>0)
p[ze] <- 0.1*min(p[-ze])
m <- length(p)
y <- sort(p)
s <- min(m*(y/(1:m)))
return(min(c(s,1)))
}
#' Combine p-values with Bonferroni's method
#'
#' This function combines p-values from the various statistical tests supported by
#' metaseqR using the Bonferroni's method (see reference in the main
#' \code{\link{metasqr}} help page or in the vignette).
#'
#' @param p a p-value matrix (rows are genes, columns are statistical tests).
#' @return A vector of combined p-values.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p <- matrix(runif(300),100,3)
#' pc <- combine.bonferroni(p)
#'}
combine.bonferroni <- function(p) {
ze <- which(p==0)
if (length(ze)>0)
p[ze] <- 0.1*min(p[-ze])
b <- length(p)*min(p)
return(min(c(1,b)))
}
#' Combine p-values using weights
#'
#' This function combines p-values from the various statistical tests supported by
#' metaseqR using p-value weights.
#'
#' @param p a p-value matrix (rows are genes, columns are statistical tests).
#' @param w a weights vector, must sum to 1.
#' @return A vector of combined p-values.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p <- matrix(runif(300),100,3)
#' pc <- combine.weight(p,w=c(0.2,0.5,0.3))
#'}
combine.weight <- function(p,w) {
ze <- which(p==0)
if (length(ze)>0)
p[ze] <- 0.1*min(p[-ze])
return(prod(p^w))
}
#' Combine p-values using the minimum p-value
#'
#' This function combines p-values from the various statistical tests supported by
#' metaseqR by taking the minimum p-value.
#'
#' @param p a p-value matrix (rows are genes, columns are statistical tests).
#' @return A vector of combined p-values.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p <- matrix(runif(300),100,3)
#' pc <- combine.min(p)
#'}
combine.minp <- function(p) { return(min(p)) }
#' Combine p-values using the maximum p-value
#'
#' This function combines p-values from the various statistical tests supported by
#' metaseqR by taking the maximum p-value.
#'
#' @param p a p-value matrix (rows are genes, columns are statistical tests).
#' @return A vector of combined p-values.
#' @export
#' @author <NAME>
#' @examples
#' \dontrun{
#' p <- matrix(runif(300),100,3)
#' pc <- combine.max(p)
#'}
combine.maxp <- function(p) { return(max(p)) }
# Copied from ex-CRAN package MADAM and exported. The man pages are copied from
# the original package.
fisher.method <- function(pvals,method=c("fisher"),p.corr=c("bonferroni","BH",
"none"),zero.sub=0.00001,na.rm=FALSE,mc.cores=NULL) {
stopifnot(method %in% c("fisher"))
stopifnot(p.corr %in% c("none","bonferroni","BH"))
stopifnot(all(pvals>=0, na.rm=TRUE) & all(pvals<=1, na.rm=TRUE))
stopifnot(zero.sub>=0 & zero.sub<=1 || length(zero.sub)!=1)
if(is.null(dim(pvals)))
stop("pvals must have a dim attribute")
p.corr <- ifelse(length(p.corr)!=1, "BH", p.corr)
##substitute p-values of 0
pvals[pvals == 0] <- zero.sub
if(is.null(mc.cores)) {
fisher.sums <- data.frame(do.call(rbind,apply(pvals,1,fisher.sum,
zero.sub=zero.sub,na.rm=na.rm)))
}
else {
fisher.sums <- parallel::mclapply(1:nrow(pvals), function(i) {
fisher.sum(pvals[i,],zero.sub=zero.sub,na.rm=na.rm)
}, mc.cores=mc.cores)
fisher.sums <- data.frame(do.call(rbind,fisher.sums))
}
rownames(fisher.sums) <- rownames(pvals)
fisher.sums$p.value <- 1-pchisq(fisher.sums$S,df=2*fisher.sums$num.p)
fisher.sums$p.adj <- switch(p.corr,
bonferroni = p.adjust(fisher.sums$p.value,"bonferroni"),
BH = p.adjust(fisher.sums$p.value,"BH"),
none = fisher.sums$p.value
)
return(fisher.sums)
}
# Copied from ex-CRAN package MADAM and exported. The man pages are copied from
# the original package.
fisher.method.perm <- function(pvals,p.corr=c("bonferroni","BH","none"),
zero.sub=0.00001,B=10000,mc.cores=NULL,blinker=1000) {
stopifnot(is.na(blinker) || blinker>0)
stopifnot(p.corr %in% c("none","bonferroni","BH"))
stopifnot(all(pvals>=0,na.rm=TRUE) & all(pvals<=1,na.rm=TRUE))
stopifnot(zero.sub>=0 & zero.sub<=1 || length(zero.sub)!=1)
if(is.null(dim(pvals)))
stop("pvals must have a dim attribute")
p.corr <- ifelse(length(p.corr)!=1,"BH",p.corr)
pvals[pvals==0] <- zero.sub
res.perm <- lapply(1:nrow(pvals),function(i) {
if(!is.na(blinker) & i%%blinker==0)
message("=", appendLF=FALSE)
##which studies contribute to S (don't have a NA in row i)
good.p <- which(!is.na(pvals[i,]))
S.obs= fisher.sum(pvals[i,good.p], na.rm=FALSE)
if(is.null(mc.cores)) {
S.rand <- unlist(lapply(1:B, function(b) {
##get non NA p-values from studies contributing to S
myp <- sapply(good.p, function(pc){
sample(na.exclude(pvals[,pc]),1)
})
fisher.sum(myp)$S
}))
} else {
S.rand <- unlist(parallel::mclapply(1:B, function(b) {
##get non NA p-values from studies contributing to S
myp <- sapply(good.p, function(pc) {
sample(na.exclude(pvals[,pc]),1)
})
fisher.sum(myp)$S
}, mc.cores=mc.cores))
}
p.value <- sum(S.rand>=S.obs$S)/B
data.frame(S=S.obs$S, num.p=S.obs$num.p, p.value=p.value)
})
res.perm <- data.frame(do.call(rbind, res.perm))
if(!is.na(blinker) && blinker>0)
message()
## rownames(res.perm) <- rownames(pvals)
res.perm$p.adj <- switch(p.corr,
bonferroni = p.adjust(res.perm$p.value,"bonferroni"),
BH = p.adjust(res.perm$p.value,"BH"),
none = res.perm$p.value)
return(res.perm)
}
# Copied from ex-CRAN package MADAM and exported. The man pages are copied from
# the original package.
fisher.sum <- function(p,zero.sub=0.00001,na.rm=FALSE) {
if(any(p>1, na.rm=TRUE)||any(p<0, na.rm=TRUE))
stop("You provided bad p-values")
stopifnot(zero.sub>=0 & zero.sub<=1 || length(zero.sub)!=1)
p[p==0] <- zero.sub
if (na.rm)
p <- p[!is.na(p)]
S = -2*sum(log(p))
res <- data.frame(S=S,num.p=length(p))
return(res)
}
<file_sep>/man/make.fold.change.Rd
\name{make.fold.change}
\alias{make.fold.change}
\title{Calculates fold changes}
\usage{
make.fold.change(contrast, sample.list, data.matrix,
log.offset = 1)
}
\arguments{
\item{contrast}{the vector of requested statistical
comparison contrasts.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{data.matrix}{a matrix of gene expression data whose
column names are the same as the sample names included in
the sample list.}
\item{log.offset}{a number to be added to each element of
data matrix in order to avoid Infinity on log type data
transformations.}
}
\value{
A matrix of fold change ratios, treatment to control, as
these are parsed from contrast.
}
\description{
Returns a matrix of fold changes based on the requested
contrast, the list of all samples and the data matrix
which is produced by the metaseqr workflow. For details
on the \code{contrast}, \code{sample.list} and
\code{log.offset} parameters, see the main usage page of
metaseqr. This function is intended mostly for internal
use but can also be used independently.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
fc <- make.fold.change("Control_vs_Treatment",list(Control=c("C1","C2"),
Treatment=c("T1","T2")),data.matrix)
}
}
\author{
<NAME>
}
<file_sep>/man/metaseqr.Rd
\name{metaseqr}
\alias{metaseqr}
\alias{metaseqr.main}
\title{The main metaseqr pipeline}
\usage{
metaseqr(counts, sample.list, exclude.list = NULL,
file.type = c("auto", "sam", "bam", "bed"),
path = NULL, contrast = NULL, libsize.list = NULL,
id.col = 4, gc.col = NA, name.col = NA, bt.col = NA,
annotation = c("download", "embedded"), gene.file = NULL,
org = c("hg18", "hg19", "hg38", "mm9", "mm10", "rn5", "rn6",
"dm3", "dm6", "danrer7", "pantro4", "susscr3", "tair10",
"equcab2","custom"),
refdb = c("ensembl", "ucsc", "refseq"),
trans.level = c("gene", "transcript", "exon"),
count.type = c("gene", "exon","utr"),
utr.flank = 500,
exon.filters = list(min.active.exons = list(exons.per.gene = 5,
min.exons = 2, frac = 1/5)),
gene.filters = list(length = list(length = 500),
avg.reads = list(average.per.bp = 100, quantile = 0.25),
expression = list(median = TRUE, mean = FALSE, quantile = NA,
known = NA, custom = NA),
biotype = get.defaults("biotype.filter", org[1]),
presence = list(frac = 0.25, min.count = 10,
per.condition = FALSE)),
when.apply.filter = c("postnorm", "prenorm"),
normalization = c("deseq", "edaseq", "edger", "noiseq", "nbpseq",
"each", "none"),
norm.args = NULL,
statistics = c("deseq", "edger", "noiseq", "bayseq", "limma",
"nbpseq"),
stat.args = NULL,
adjust.method = sort(c(p.adjust.methods, "qvalue")),
meta.p = if (length(statistics) > 1) c("simes", "bonferroni", "fisher",
"dperm.min", "dperm.max", "dperm.weight", "fperm", "whitlock",
"minp", "maxp", "weight", "pandora", "none") else "none",
weight = rep(1/length(statistics), length(statistics)),
nperm = 10000, reprod=TRUE, pcut = NA, log.offset = 1,
preset = NULL,
qc.plots = c("mds", "biodetection", "countsbio", "saturation",
"readnoise", "filtered", "correl", "pairwise", "boxplot",
"gcbias", "lengthbias", "meandiff", "meanvar", "rnacomp",
"deheatmap", "volcano", "biodist"),
fig.format = c("png", "jpg", "tiff", "bmp", "pdf", "ps"),
out.list = FALSE, export.where = NA,
export.what = c("annotation", "p.value", "adj.p.value",
"meta.p.value", "adj.meta.p.value", "fold.change",
"stats", "counts","flags"),
export.scale = c("natural", "log2", "log10", "vst", "rpgm"),
export.values = c("raw", "normalized"),
export.stats = c("mean", "median", "sd", "mad", "cv",
"rcv"),
export.counts.table = FALSE,
restrict.cores = 0.6, report = TRUE, report.top = 0.1,
report.template = "default", save.gene.model = TRUE,
verbose = TRUE, run.log = TRUE, progress.fun = NULL, ...)
}
\arguments{
\item{counts}{a text tab-delimited file containing gene
or exon counts in one of the following formats: i) the
first column contains unique gene or exon identifiers and
the rest of the columns contain the read counts for each
sample. Thus the first cell of each row is a gene or exon
accession and the rest are integers representing the
counts for that accession. In that case, the
\code{annotation} parameter should strictly be
\code{"download"} or an external file in proper format.
ii) The first n columns should contain gene or exon
annotation elements like chromosomal locations, gene
accessions, exon accessions, GC content etc. In that
case, the \code{annotation} parameter can also be
\code{"embedded"}. The ideal embedded annotation contains
8 columns, chromosome, gene or exon start, gene or exon
end, gene or exon accession, GC-content (fraction or
percentage), strand, HUGO gene symbol and gene biotype
(e.g. "protein_coding" or "ncRNA"). When the
\code{annotation} parameter is "embedded", certain of
these features are mandatory (co-ordinates and
accessions). If they are not present, the pipeline will
not run. If additional elements are not present (e.g. GC
content or biotypes), certain features of metaseqr will
not be available. For example, EDASeq normalization will
not be performed based on a GC content covariate but
based on gene length which is not what the authors of
EDASeq suggest. If biotypes are not present, a lot of
diagnostic plots will not be available. If the HUGO gene
symbols are missing, the final annotation will contain
only gene accessions and thus be less comprehensible.
Generally, it's best to set the \code{annotation}
parameter to \code{"download"} to ensure the most
comprehensible results. Counts can be a data frame
satisfying the above conditions. It is a data frame
by default when \code{read2count} is used. counts can
also be an .RData file (output of \code{\link{save}}
function which contains static input elements (list
containing the gene model (exon counts for each gene
constructed by the \code{\link{construct.gene.model}}
function, gene and exon annotation to avoid
re-downloading and/or gene counts depending on
\code{count.type}). This kind of input facilitates the
re-analysis of the same experiment, using different
filtering, normalization and statistical algorithms.
Finally, counts can be a list representing the gene model
(exon counts for each gene) constructed by the
\code{\link{construct.gene.model}} function (provided for
backwards compatibility). This .RData file can be generated
by setting \code{save.gene.model=TRUE} when performing data
analysis for the first time.}
\item{sample.list}{a list containing condition names and
the samples under each condition. It should have the
format \code{sample.list <-}
\code{list(ConditionA=c("Sample_A1",} \code{"Sample_A2",
"Sample_A3"),} \code{ConditionB=c("Sample_B1",
"Sample_B2"),} \code{ConditionC=c("Sample_C1",
"Sample_C2"))}. The names of the samples in list members
MUST match the column names containing the read counts in
the counts file. If they do not match, the pipeline will
either crash or at best, ignore several of your samples.
Alternative, \code{sample.list} can be a small
tab-delimited file structured as follows: the first line
of the external tab delimited file should contain column
names (names are not important). The first column MUST
contain UNIQUE sample names and the second column MUST
contain the biological condition where each of the
samples in the first column should belong to. In this
case, the function \code{\link{make.sample.list}} is
used. If the \code{counts} argument is missing, the
\code{sample.list} argument MUST be a targets text
tab-delimited file which contains the sample names, the
BAM/BED file names and the biological conditions/groups
for each sample/file. The file should be text
tab-delimited and structured as follows: the first line
of the external tab delimited file should contain column
names (names are not important). The first column MUST
contain UNIQUE sample names. The second column MUST
contain the raw BAM/BED files WITH their full path.
Alternatively, the \code{path} argument should be
provided (see below). The third column MUST contain the
biological condition where each of the samples in the
first column should belong to.}
\item{exclude.list}{ a list of samples to exclude, in the
same (list) format as \code{sample.list} above.}
\item{path}{an optional path where all the BED/BAM files
are placed, to be prepended to the BAM/BED file names in
the targets file. If not given and if the files in the
second column of the targets file do not contain a path
to a directory, the current directory is assumed to be
the BAM/BED file container.}
\item{file.type}{the type of raw input files. It can be
\code{"auto"} for auto-guessing, \code{"bed"} for BED
files, \code{"sam"} for SAM files or \code{"bam"} for BAM
files.}
\item{contrast}{a character vector of contrasts to be
tested in the statistical testing step(s) of the metaseqr
pipeline. Each element of contrast should STRICTLY have the
format "ConditionA_vs_ConditionB_vs_...". A valid example
based on the \code{sample.list} above is \code{contrast
<- c("ConditionA_vs_ConditionB",}
\code{"ConditionA_vs_ConditionC",}
\code{"ConditionA_vs_ConditionB_vs_ConditionC")}. The
first element of pairwise contrasts (e.g. "ConditionA"
above) MUST be the control condition or any reference
that ConditionB is checked against. metaseqr uses this
convention to properly calculate fold changes. If it's
NULL, a contrast between the first two members of the
\code{sample.list} will be auto-generated.}
\item{libsize.list}{an optional named list where names
represent samples (MUST be the same as the samples in
\code{sample.list}) and members are the library sizes
(the sequencing depth) for each sample. For example
\code{libsize.list <- list(Sample_A1=32456913,}
\code{Sample_A2=4346818)}.}
\item{id.col}{an integer denoting the column number in
the file (or data frame) provided with the counts
argument, where the unique gene or exon accessions are.
Default to \code{4} which is the standard feature name
column in a BED file.}
\item{gc.col}{an integer denoting the column number in
the file (or data frame) provided with the \code{counts}
argument, where each gene's GC content is given. If not
provided, GC content normalization provided by EDASeq
will not be available.}
\item{name.col}{an integer denoting the column number in
the file (or data frame) provided with the counts
argument, where the HUGO gene symbols are given. If not
provided, it will not be available when reporting
results. In addition, the \code{"known"} gene filter will
not be available.}
\item{bt.col}{an integer denoting the column number in
the file (or data frame) provided with the counts
argument, where the gene biotypes are given. If not
provided, the \code{"biodetection"}, \code{"countsbio"},
\code{"saturation"}, \code{"filtered"} and
\code{"biodist"} plots will not be available.}
\item{annotation}{instructs metaseqr where to find the
annotation for the given counts file. It can be one of i)
\code{"download"} (default) for automatic downloading of
the annotation for the organism specified by the org
parameter (using biomaRt), ii) \code{"embedded"} if the
annotation elements are embedded in the read counts file
or iv) a file specified by the user which should be as
similar as possible to the \code{"download"} case, in
terms of column structure.}
\item{gene.file}{an external gene annotation file
required when \code{annotation="embedded"},
\code{count.type="exon"} or \code{count.type="utr"}
and \code{org="custom"}. See the result of
\code{\link{get.annotation}} on the format of the
external gene file.}
\item{org}{the supported organisms by metaseqr. These can
be, for human genomes \code{"hg18"}, \code{"hg19"} or
\code{"hg38"}, for mouse genomes \code{"mm9"}, \code{"mm10"},
for rat genomes \code{"rn5"} or \code{"rn6"}, for drosophila
genome \code{"dm3"} or \code{"dm6"}, for zebrafish genome
\code{"danrer7"}, for chimpanzee genome \code{"pantro4"},
for pig genome \code{"susscr3"}, for Arabidopsis
thaliana genome \code{"tair10"} and for Equus caballus genome
\code{"equcab2"}. Finally, \code{"custom"} will instruct
metaseqR to completely ignore the \code{org} argument and
depend solely on annotation file provided by the user.}
\item{refdb}{the reference annotation repository from
which to retrieve annotation elements to use with
metaseqr. It can be one of \code{"ensembl"} (default),
\code{"ucsc"} or \code{"refseq"}.}
\item{trans.level}{perform differential expression
analysis at which transcriptional unit, can be one of
\code{"gene"} (default), \code{"transcript"} for
reporting differential expression at the transcript
level or \code{"exon"} for exon level.}
\item{count.type}{the type of reads inside the counts
file. It can be one of \code{"gene"}, \code{"exon"} or
\code{"utr"} for quant seq (lexogen) protocol. This is
a very important and mandatory parameter as it defines
the course of the workflow.}
\item{utr.flank}{the number of base pairs to flank the
3' UTR of transcripts when analyzing Quant-Seq data.}
\item{exon.filters}{a named list whose names are the
names of the supported exon filters and its members the
filter parameters. See section "Exon filters" below for
details.}
\item{gene.filters}{a named list whose names are the
names of the supported gene filters and its members the
filter parameters. See section "Gene filters" below for
details.}
\item{when.apply.filter}{a character string determining
when to apply the exon and/or gene filters, relative to
normalization. It can be \code{"prenorm"} to apply apply
the filters and exclude genes from further processing
before normalization, or \code{"postnorm"} to apply the
filters after normalization (default). In the case of
\code{when.apply.filter="prenorm"}, a first normalization
round is applied to a copy of the gene counts matrix in
order to derive the proper normalized values that will
constitute the several expression-based filtering
cutoffs.}
\item{normalization}{the normalization algorithm to be
applied on the count data. It can be one of
\code{"edaseq"} for EDASeq normalization, \code{"deseq"}
for the normalization algorithm (individual options
specified by the \code{norm.args} argument) in the DESq
package (default), \code{"edger"} for the
normalization algorithms present in the edgeR package
(specified by the \code{norm.args} argument),
\code{"noiseq"} for the normalization algorithms present
in the NOISeq package (specified by the \code{norm.args}
argument), \code{"nbpseq"} for the normalization
algorithms present in the NBPSeq package (specified by
the \code{norm.args} argument) or \code{"none"} to not
normalize the data (highly unrecommended). It can also
be \code{"each"} where in this case, the normalization
applied will be specific to each statistical test used
(i.e. the normalization method bundled with each package
and used in its examples and documentation). The last
choice is for future use!}
\item{norm.args}{a named list whose names are the names
of the normalization algorithm parameters and its members
parameter values. See section "Normalization parameters"
below for details. Leave \code{NULL} for the defaults of
\code{normalization}. If \code{normalization="each"}, it
must be a named list of lists, where each sub-list
contains normalization parameters specific to each
statistical test to be used. The last choice is for
future use!}
\item{statistics}{one or more statistical analyses to be
performed by the metaseqr pipeline.It can be one or more
of \code{"deseq"} (default) to conduct statistical
test(s) implemented in the DESeq package, \code{"edger"}
to conduct statistical test(s) implemented in the edgeR
package, \code{"limma"} to conduct the RNA-Seq version of
statistical test(s) implemented in the limma package,
\code{"noiseq"} to conduct statistical test(s)
implemented in the NOISeq package, \code{"bayseq"} to
conduct statistical test(s) implemented in the baySeq
package and \code{"nbpseq"} to conduct statistical
test(s) implemented in the NBPSeq package. In any case
individual algorithm parameters are controlled by the
contents of the \code{stat.args} list.}
\item{stat.args}{a named list whose names are the names
of the statistical algorithms used in the pipeline. Each
member is another named list whose names are the
algorithm parameters and its members are the parameter
values. See section "Statistics parameters" below for
details. Leave \code{NULL} for the defaults of
\code{statistics}.}
\item{adjust.method}{the multiple testing p-value
adjustment method. It can be one of
\code{\link{p.adjust.methods}} or \code{"qvalue"} from
the qvalue Bioconductor package. Defaults to \code{"BH"}
for Benjamini-Hochberg correction.}
\item{meta.p}{the meta-analysis method to combine
p-values from multiple statistical tests . It can be
one of \code{"simes"} (default), \code{"bonferroni"},
\code{"minp"}, \code{"maxp"}, \code{"weight"}, \code{"pandora"},
\code{"dperm.min"}, \code{"dperm.max"}, \code{"dperm.weight"},
\code{"fisher"}, \code{"fperm"}, \code{"whitlock"} or
\code{"none"}. For the \code{"fisher"} and \code{"fperm"}
methods, see the documentation of the R package MADAM. For
the \code{"whitlock"} method, see the documentation of the
survcomp Bioconductor package. With the \code{"maxp"}
option, the final p-value is the maximum p-value out of
those returned by each statistical test. This is
equivalent to an "intersection" of the results derived
from each algorithm so as to have a final list with the
common genes returned by all statistical tests. Similarly,
when \code{meta.p="minp"}, is equivalent to a "union" of
the results derived from each algorithm so as to have a
final list with all the genes returned by all statistical
tests. The latter can be used as a very lose statistical
threshold to aggregate results from all methods regardless
of their False Positive Rate. With the \code{"simes"}
option, the method proposed by Simes (<NAME>., 1986)
is used. With the \code{"dperm.min"}, \code{"dperm.max"},
\code{"dperm.weight"} options, a permutation procedure is
initialed, where \code{nperm} permutations are performed
across the samples of the normalized counts matrix,
producing \code{nperm} permuted instances of the initital
dataset. Then, all the chosen statistical tests are
re-executed for each permutation. The final p-value is
the number of times that the p-value of the permuted
datasets is smaller than the original dataset.
The p-value of the original dataset is created based on
the choice of one of \code{dperm.min}, \code{dperm.max}
or \code{dperm.weight} options. In case of
\code{dperm.min}, the intial p-value vector is consists
of the minimum p-value resulted from the applied
statistical tests for each gene. The maximum p-value
is used with the \code{dperm.max} option. With the
\code{dperm.weight} option, the \code{weight}
weighting vector for each statistical test is used to
weight each p-value according to the power of
statistical tests (some might work better for a
specific dataset). Be careful as the permutation
procedure usually requires a lot of time. However,
it should be the most accurate. This method will NOT
work when there are no replicated samples across
biological conditions. In that case, use
\code{meta.p="simes"} instead. Finally, there are the
\code{"minp"}, \code{"maxp"} and \code{"weight"}
options which correspond to the latter three methods
but without permutations. Generally, permutations
would be accurate to use when the experiment includes
>5 samples per condition (or even better 7-10) which
is rather rare in RNA-Seq experiments. Finally,
\code{"pandora"} is the same as \code{"weight"} and is
added to be in accordance with the metaseqR paper.}
\item{weight}{a vector of weights with the same length as
the \code{statistics} vector containing a weight for each
statistical test. It should sum to 1. \strong{Use with
caution with the} \code{dperm.weight} \strong{parameter!
Theoretical background is not yet} \strong{solid and only
experience shows improved results!}}
\item{nperm}{the number of permutations performed to
derive the meta p-value when \code{meta.p="fperm"} or
\code{meta.p="dperm"}. It defaults to 10000.}
\item{reprod}{create reproducible permutations when
\code{meta.p="dperm.min"}, \code{meta.p="dperm.max"}
or \code{meta.p="dperm.weight"}. Ideally one would
want to create the same set of indices for a given
dataset so as to create reproducible p-values. If
\code{reprod=TRUE}, a fixed seed is used by
\code{meta.perm} for all the datasets analyzed with
\code{metaseqr}. If \code{reprod=FALSE}, then the
p-values will not be reproducible, although statistical
significance is not expected to change for a large
number of resambling. Finally, \code{reprod} can be
a numeric vector of seeds with the same length as
\code{nperm} so that the user can supply his/her
own seeds.}
\item{pcut}{a p-value cutoff for exporting differentially
genes, default is to export all the non-filtered genes.}
\item{log.offset}{an offset to be added to values during
logarithmic transformations in order to avoid Infinity
(default is \code{1}).}
\item{preset}{an analysis strictness preset.
\code{preset} can be one of \code{"all.basic"},
\code{"all.normal"}, \code{"all.full"},
\code{"medium.basic"}, \code{"medium.normal"},
\code{"medium.full"}, \code{"strict.basic"},
\code{"strict.normal"} or \code{"strict.full"}, each of
which control the strictness of the analysis and the
amount of data to be exported. For an explanation of the
presets, see the section "Presets" below.}
\item{qc.plots}{a set of diagnostic plots to show/create.
It can be one or more of \code{"mds"},
\code{"biodetection"}, \code{"rnacomp"},
\code{"countsbio"}, \code{"saturation"},
\code{"readnoise"}, \code{"filtered"}, \code{"boxplot"},
\code{"gcbias"}, \code{"lengthbias"}, \code{"meandiff"},
\code{"meanvar"}, \code{"deheatmap"}, \code{"volcano"},
\code{"biodist"}, \code{"venn"}. The \code{"mds"} stands
for Mutlti-Dimensional Scaling and it creates a PCA-like
plot but using the MDS dimensionality reduction instead.
It has been succesfully used for NGS data (e.g. see the
package htSeqTools) and it shows how well samples from
the same condition cluster together. For
\code{"biodetection"}, \code{"countsbio"},
\code{"saturation"}, \code{"rnacomp"},
\code{"readnoise"}, \code{"biodist"} see the vignette of
NOISeq package. The \code{"saturation"} case has been
rewritten in order to display more samples in a more
simple way. See the help page of
\code{\link{diagplot.noiseq.saturation}}. In addition,
the \code{"readnoise"} plots represent an older version
or the RNA composition plot included in older versions of
NOISeq. For \code{"gcbias"}, \code{"lengthbias"},
\code{"meandiff"}, \code{"meanvar"} see the vignette of
EDASeq package. \code{"lenghtbias"} is similar to
\code{"gcbias"} but using the gene length instead of the
GC content as covariate. The \code{"boxplot"} option
draws boxplots of log2 transformed gene counts. The
\code{"filtered"} option draws a 4-panel figure with the
filtered genes per chromosome and per biotype, as
absolute numbers and as fractions of the genome. See also
the help page of \code{\link{diagplot.filtered}}. The
\code{"deheatmap"} option performs hierarchical
clustering and draws a heatmap of differentially
expressed genes. In the context of diagnostic plots, it's
useful to see if samples from the same groups cluster
together after statistical testing. The \code{"volcano"}
option draws a volcano plot for each contrast and if a
report is requested, an interactive volcano plot is
presented in the HTML report. The \code{"venn"} option
will draw an up to 5-way Venn diagram depicting the
common and specific to each statistical algorithm genes
and for each contrast, when meta-analysis is performed.
The \code{"correl"} option creates two correlation
graphs: the first one is a correlation heatmap (a
correlation matrix which depicts all the pairwise
correlations between each pair of samples in the counts
matrix is drawn as a clustered heatmap) and the second
one is a correlogram plot, which summarizes the
correlation matrix in the form of ellipses (for an
explanation please see the vignette/documentation of the
R package corrplot. Set \code{qc.plots=NULL} if you don't
want any diagnostic plots created.}
\item{fig.format}{the format of the output diagnostic
plots. It can be one or more of \code{"png"},
\code{"jpg"}, \code{"tiff"}, \code{"bmp"}, \code{"pdf"},
\code{"ps"}. The native format \code{"x11"} (for direct
display) is not provided as an option as it may not
render the proper display of some diagnostic plots in
some devices.}
\item{out.list}{a logical controlling whether to export a
list with the results in the running environment.}
\item{export.where}{an output directory for the project
results (report, lists, diagnostic plots etc.)}
\item{export.what}{the content of the final lists. It can
be one or more of \code{"annotation"}, to bind the
annoation elements for each gene, \code{"p.value"}, to
bind the p-values of each method, \code{"adj.p.value"},
to bind the multiple testing adjusted p-values,
\code{"meta.p.value"}, to bind the combined p-value from
the meta-analysis, \code{"adj.meta.p.value"}, to bind the
corrected combined p-value from the meta-analysis,
\code{"fold.change"}, to bind the fold changes of each
requested contrast, \code{"stats"}, to bind several
statistics calclulated on raw and normalized counts (see
the \code{export.stats} argument), \code{"counts"}, to
bind the raw and normalized counts for each sample.}
\item{export.scale}{export values from one or more
transformations applied to the data. It can be one or
more of \code{"natural"}, \code{"log2"}, \code{"log10"},
\code{"vst"} (Variance Stabilizing Transormation, see the
documentation of DESeq package) and \code{"rpgm"} which
is ratio of mapped reads per gene model (either the gene
length or the sum of exon lengths, depending on
\code{count.type} argument). Note that this is not RPKM
as reads are already normalized for library size using
one of the supported normalization methods. Also,
\code{"rpgm"} might be misleading when \code{normalization}
is other than \code{"deseq"}.}
\item{export.values}{It can be one or more of
\code{"raw"} to export raw values (counts etc.) and
\code{"normalized"} to export normalized counts.}
\item{export.stats}{calculate and export several
statistics on raw and normalized counts, condition-wise.
It can be one or more of \code{"mean"}, \code{"median"},
\code{"sd"}, \code{"mad"}, \code{"cv"} for the
Coefficient of Variation, \code{"rcv"} for a robust
version of CV where the median and the MAD are used
instead of the mean and the standard deviation.}
\item{export.counts.table}{exports also the calculated
read counts table when input is read from bam files
and exports also the normalized count table in all
cases. Defaults to \code{FALSE}.}
\item{restrict.cores}{in case of parallel execution of
several subfunctions, the fraction of the available cores
to use. In some cases if all available cores are used
(\code{restrict.cores=1} and the system does not have
sufficient RAM, the pipeline running machine might
significantly slow down.}
\item{report}{a logical value controlling whether to
produce a summary report or not. Defaults to
\code{TRUE}.}
\item{report.top}{a fraction of top statistically
significant genes to append to the HTML report. This
helps in keeping the size of the report as small as
possible, as appending the total gene list might
create a huge HTML file. Users can always retrieve
the whole gene lists from the report links. Defaults
to \code{0.1} (top 10% of statistically significant
genes). Set to \code{NA} or \code{NULL} to append all
the statistically significant genes to the HTML report.}
\item{report.template}{an HTML template to use for the
report. Do not change this unless you know what you are
doing.}
\item{save.gene.model}{in case of exon analysis, a list
with exon counts for each gene will be saved to the file
\code{export.where/data/gene_model.RData}. This file can
be used as input to metaseqR for exon count based
analysis, in order to avoid the time consuming step of
assembling the counts for each gene from its exons}
\item{verbose}{print informative messages during
execution? Defaults to \code{TRUE}.}
\item{run.log}{write a log file of the \code{metaseqr}
run using package log4r. Defaults to \code{TRUE}. The
filename will be auto-generated.}
\item{progress.fun}{a function which updates a
\code{Progress} object from shiny. This function must
accept a \code{detail} argument. See
http://shiny.rstudio.com/articles/progress.html}
\item{...}{further arguments that may be passed to
plotting functions, related to \code{\link{par}}.}
}
\value{
If \code{out.list} is \code{TRUE}, a named list whose
length is the same as the number of requested contrasts.
Each list member is named according to the corresponding
contrast and contains a data frame of differentially
expressed genes for that contrast. The contents of the
data frame are defined by the \code{export.what,
export.scale, export.stats, export.values} parameters. If
\code{report} is \code{TRUE}, the output list contains
two main elements. The first is described above (the
analysis results) and the second contains the same
results but in HTML formatted tables.
}
\description{
This function is the main metaseqr workhorse and
implements the main metaseqr workflow which performs data
read, filtering, normalization and statistical selection,
creates diagnostic plots and exports the results and a
report if requested. The metaseqr function is responsible
for assembling all the steps of the metaseqr pipeline
which i) reads the input gene or exon read count table
ii) performs prelimininary filtering of data by removing
chrM and other non-essential information for a typical
differential gene expression analysis as well as a
preliminary expression filtering based on the exon
counts, if an exon read count file is provided. iii)
performs data normalization with one of currently widely
used algorithms, including EDASeq (Risso et al., 2011),
DESeq (<NAME>, 2010), edgeR (Robinson et al.,
2010), NOISeq (Tarazona et al., 2012) or no normalization
iv) performs a second stage of filtering based on the
normalized gene expression according to several gene
filters v) performs statistical testing with one or more
of currently widely used algorithms, including DESeq
(<NAME>, 2010), edgeR (Robinson et al., 2010),
NOISeq (Tarazona et al., 2012), limma (Smyth et al.,
2005) for RNA-Seq data, baySeq (Hardcastle et al., 2012)
vi) in the case of multiple statistical testing
algorithms, performs meta-analysis using one of five
available methods (see the meta.p argument) vii) exports
the resulting differentially expressed gene list in text
tab-delimited format viii) creates a set of diagnostic
plots either available in the aforementioned packages or
metaseqr specific ones and ix) creates a comprehensive
HTML report which summarizes the run information, the
results and the diagnostic plots. Certain diagnostic
plots (e.g. the volcano plot) can be interactive with the
use of the external Highcharts
(http://www.highcharts.com) JavaScript library for
interactive graphs. Although the inputs to the metaseqr
workflow are many, in practice, setting only very few of
them and accepting the defaults as the rest can result in
quite comprehensible results for mainstream organisms
like mouse, human, fly and rat.
}
\note{
Please note that currently only gene and exon annotation
from Ensembl (http://www.ensembl.org), UCSC and RefSeq
are supported. Thus, the unique gene or exon ids in the
counts files should correspond to valid Ensembl, UCSC or
RefSeq gene or exon accessions for the organism of interest.
If you are not sure about the source of your counts file or
do not know how to produce it, it's better to start from the
original BAM/BED files (metaseqr will use the
\code{\link{read2count}} function to create a counts
file). Keep in mind that in the case of BED files, the
performance will be significantly lower and the overall
running time significanlty higher as the R functions
which are used to read BED files to proper structures
(GenomicRanges) and calculate the counts are quite slow.
An alternative way is maybe the easyRNASeq package
(Delhomme et al, 2012). The \code{\link{read2count}}
function does not use this package but rather makes use
of standard Bioconductor functions to handle NGS data. If
you wish to work outside R, you can work with other
popular read counters such as the HTSeq read counter
(http://www-huber.embl.de/users/anders/HTSeq/doc/overview.html).
Please also note that in the current version, the members
of the \code{gene.filters} and \code{exon.filters} lists
are not checked for validity so be careful to supply with
correct names otherwise the pipeline will crash or at the
best case scenario, will ignore the filters. Also note
that when you are supplying metaseqr wtih an exon counts
table, gene annotation is always downloaded so please be
sure to have a working internet connection. In addition
to the above, if you have a multiple core system, be very
careful on how you are using the \code{restrict.cores}
argument and generally how many cores you are using with
scripts purely written in R. The analysis with exon read
data can very easily cause memory problems, so unless you
have more than 64Gb of RAM available, consider setting
restrict.cores to something like 0.2 when working with
exon data. Finally, if you do not wish to download the
same annotation again and again when performing multiple
analyses, it is best to use the
\code{\link{get.annotation}} function to download and
store the resulting data frames in local files and then
use these files with the \code{annotation} option.
Please note that the \strong{meta-analysis} feature
provided by metaseqr does not satisfy the strict definition
of "meta-analysis", which is the combination of multiple
similar datasets under the same statistical methodology.
Instead it is the use of mulitple statistical tests applied
to the same data. For the Simes method, please consult also
"<NAME>. (1986). "An improved Bonferroni procedure
for multiple tests of significance". Biometrika 73 (3):
751–754."
}
\section{Exon filters}{
The exon filters are a set of filters which are applied
after the gene models are assembled from the read counts
of individual exons and before the gene expression is
summarized from the exons belonging to each gene. These
filters can be applied when the input read counts file
contains exon reads. It is not applicable when the input
file already contains gene counts. Such filters can be
for example "accept genes where all the exons contain
more than x reads" or "accept genes where there is read
presence in at least m/n exons, n being the total exons
of the gene". Such filters are NOT meant for detecting
differential splicing as also the whole metaseqr
pipeline, thus they should not be used in that context.
The \code{exon.filters} argument is a named list of
filters, where the names are the filter names and the
members are the filter parameters (named lists with
parameter name, parameter value). See the usage of the
\code{metaseqr} function for an example of how these
lists are structured. The supported exon filters in the
current version are: i) \code{min.active.exons} which
implements a filter for demanding m out of n exons of a
gene to have a certain read presence with parameters
\code{exons.per.gene}, \code{min.exons} and \code{frac}.
The filter is described as follows: if a gene has up to
\code{exons.per.gene} exons, then read presence is
required in at least \code{min.exons} of them, else read
presence is required in a \code{frac} fraction of the
total exons. With the default values, the filter
instructs that if a gene has up to 5 exons, read presence
is required in at least 2, else in at least 20% of the
exons, in order to be accepted. More filters will be
implemented in future versions and users are encouraged
to propose exon filter ideas to the author by mail. See
\code{metaseqr} usage for the defaults. Set
\code{exon.filters=NULL} to not apply any exon filtering.
}
\section{Gene filters}{
The gene filters are a set of filters applied to gene
expression as this is manifested through the read
presence on each gene and are preferably applied after
normalization. These filters can be applied both when the
input file or data frame contains exon read counts and
gene read counts. Such filter can be for example "accept
all genes above a certain count threshold" or "accept all
genes with expression above the median of the normalized
counts distribution" or "accept all with length above a
certain threshold in kb" or "exclude the 'pseudogene'
biotype from further analysis". The supported gene
filters in the current version, which have the same
structure as the exon filters (named list of lists with
filter names, parameter names and parameter arguments)
are: i) \code{length} which implements a length filter
where genes are accepted for further analysis if they are
above \code{length} (its parameter) kb. ii)
\code{avg.reads} which implements a filter where a gene
is accepted for further analysis if it has more average
reads than the \code{quantile} of the average count
distribution per \code{average.per.bp} base pairs. In
summary, the reads of each gene are averaged per
\code{average.per.bp} based on each gene's length (in
case of exons, input the "gene's length" is the sum of
the lengths of exons) and the \code{quantile} quantile of
the average counts distribution is calculated for each
sample. Genes passing the filter should have an average
read count larger than the maximum of the vector of the
quantiles calculated above. iii) \code{expression} which
implements a filter based on the overall expression of a
gene. The parameters of this filter are: \code{median},
where genes below the median of the overall count
distribution are not accepted for further analysis (this
filter has been used to distinguish between "expressed"
and "not expressed" genes in several cases, e.g. (Mokry
et al., NAR, 2011) with a logical as value, \code{mean}
which is the same as \code{median} but using the mean,
\code{quantile} which is the same as the previous two but
using a specific quantile of the total counts
distribution, \code{known}, where in this case, a set of
known not-expressed genes in the system under
investigation are used to estimate an expression cutoff.
This can be quite useful, as the genes are filtered based
on a "true biological" cutoff instead of a statistical
cutoff. The value of this filter is a character vector of
HUGO gene symbols (MUST be contained in the annotation,
thus it's better to use \code{annotation="download"})
whose counts are used to build a "null" expression
distribution. The 90th quantile of this distribution is
then the expression cutoff. This filter can be combined
with any other filter. Be careful with gene names as they
are case sensitive and must match exactly ("Pten" is
different from "PTEN"!). iv) \code{biotype} where in this
case, genes with a certain biotype (MUST be contained in
the annotation, thus it's better to use
\code{annotation="download"}) are excluded from the
analysis. This filter is a named list of logical, where
names are the biotypes in each genome and values are
\code{TRUE} or \code{FALSE}. If the biotype should be
excluded, the value should be \code{TRUE} else
\code{FALSE}. See the result of
\code{get.defaults("biotype.filter","hg19")} for an
example. Finally, in future versions there will be
support for user-defined filters in the form of a
function. v) \code{presence} where in this case, a gene
is further considered for statistical testing if
\code{frac} (x100 for a percentage value) have more
than \code{min.count} reads across all samples
(\code{per.condition=FALSE}) or across the samples
of each condition (\code{per.condition=TRUE}).
}
\section{Normalization parameters}{
The normalization parameters are passed again as a named
list where the names of the members are the normalization
parameter names and the values are the normalization
parameter values. You should check the documentation of
the packages EDASeq, DESeq, edgeR, NOISeq and NBPSeq for
the parameter names and parameter values. There are a few
exceptions in parameter names: in case of
\code{normalization="edaseq"} the only parameter names
are \code{within.which} and \code{between.which},
controlling the withing lane/sample and between
lanes/samples normalization algorithm. In the case
of \code{normalization="nbpseq"}, there is one
additional parameter called \code{main.method} which can
take the calues \code{"nbpseq"} or \code{"nbsmyth"}.
These values correspond to the two different workflows
available in the NBPSeq package. Please, consult the
NBPSeq package documentation for further details. For the
rest of the algorithms, the parameter names are the same
as the names used in the respective packages. For
examples, please use the \code{\link{get.defaults}}
function.
}
\section{Statistics parameters}{
The statistics parameters as passed to statistical
algorithms in metaseqr, exactly with the same way as the
normalization parametes above. In this case, there is one
more layer in list nesting. Thus, \code{stat.args} is a
named list whose names are the names the algorithms used
(see the \code{statistics} parameter). Each member is
another named list,with parameters to be used for each
statistical algorithm. Again, the names of the member
lists are parameter names and the values of the member
lists are parameter values. You should check the
documentations of DESeq, edgeR, NOISeq, baySeq, limma and
NBPSeq for these parameters. There are a few exceptions
in parameter names: In case of \code{statistics="edger"},
apart from the rest of the edgeR statistical testing
arguments, there is the argument \code{main.method} which
can be either \code{"classic"} or \code{"glm"}, again
defining whether the binomial test or GLMs will be used
for statistical testing. For examples, please use the
\code{\link{get.defaults}} function. When
\code{statistics="nbpseq"}, apart from the rest arguments
of the NBPSeq functions \code{estimate.disp} and
\code{estimate.dispersion}, there is the argument
\code{main.method} which can be \code{"nbpseq"} or
\code{"nbsmyth"}. This argument determines the parameters
to be used by the \code{estimate.dispersion} function or
by the \code{estimate.disp} function to estimate RNA-Seq
count dispersions. The difference between the two is that
they constitute different starting points for the two
workflows in the package NBPSeq. The first worklfow (with
\code{main.method="nbpseq"} and the
\code{estimate.dispersion} function is NBPSeq package
specific, while the second (with
\code{main.method="nbsmyth"} and the \code{estimate.disp}
function is similar to the workflow of the edgeR package.
For additional information regarding the statistical
testing in NBPSeq, please consult the documentation of
the NBPSeq package. \strong{Additinally, please note that
there is currently a problem with the NBPSeq package and
the workflow that is specific to the NBPSeq package. The
problem has to do with function exporting as there are
certain functions which are not recognized from the
package internally. For this reason and until it is
fixed, only the Smyth workflow will be available with the
NBPSeq package (thus}
\code{stat.args$main.method="nbpseq"} \strong{ will not
be available)!}
}
\section{Presets}{
The analysis presets are a set of keywords (only one can
be used) that predefine some of the parameters of the
metaseqr pipeline. For the time being they are quite
simple and they control i) the strictness of filtering
and statistical thresholding with three basic levels
("all", "medium", "strict") and ii) the data columns that
are exported, again in three basic ways ("basic",
"normal", "full") controlling the amount of data to be
exported. These keywords can be combined with a dot in
the middle (e.g. \code{"all.basic"} to define an analysis
preset. When using analysis presets, the following
argumentsof metaseqr are overriden: \code{exon.filters},
\code{gene.filters}, \code{pcut}, \code{export.what},
\code{export.scale}, \code{export.values},
\code{exon.stats}. If you want to explicitly control the
above arguments, the \code{preset} argument should be set
to \code{NULL} (default). Following is a synopsis of the
different presets and the values of the arguments they
moderate: \itemize{ \item \code{"all.basic"}: use all
genes (do not filter) and export all genes and basic
annotation and statistics elements. In this case, the
above described arguments become: \itemize{ \item
\code{exon.filters=NULL} \item \code{gene.filters=NULL}
\item \code{pcut=1} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change")} \item
\code{export.scale=c("natural","log2")} \item
\code{export.values=c("normalized")} \item
\code{export.stats=c("mean")} } \item
\code{"all.normal"}: use all genes (do not filter) and
export all genes and normal annotation and statistics
elements. In this case, the above described arguments
become: \itemize{ \item \code{exon.filters=NULL} \item
\code{gene.filters=NULL} \item \code{pcut=1} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change","stats","counts")}
\item \code{export.scale=c("natural","log2")} \item
\code{export.values=c("normalized")} \item
\code{export.stats=c("mean","sd","cv")} } In this case,
the above described arguments become: \itemize{ \item
\code{exon.filters=NULL} \item \code{gene.filters=NULL}
\item \code{pcut=1} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change","stats","counts")}
\item
\code{export.scale=c("natural","log2","log10","vst")}
\item \code{export.values=c("raw","normalized")} \item
\code{export.stats=c("mean","median","sd","mad","cv","rcv")}
} \item \code{"medium.basic"}: apply a medium set of
filters and and export statistically significant genes
and basic annotation and statistics elements. In this
case, the above described arguments become: \itemize{
\item
\code{exon.filters=list(min.active.exons=list(exons.per.gene=5,min.exons=2,frac=1/5))}
\item \code{gene.filters=list(length=list(length=500),}
\code{avg.reads=list(average.per.bp=100,quantile=0.25),}
\code{expression=list(median=TRUE,mean=FALSE,quantile=NA,known=NA,custom=NA),}
\code{biotype=get.defaults("biotype.filter",org[1]))}
\item \code{pcut=0.05} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change")} \item
\code{export.scale=c("natural","log2")} \item
\code{export.values=c("normalized")} \item
\code{export.stats=c("mean")} } \item
\code{"medium.normal"}: apply a medium set of filters and
and export statistically significant genes and normal
annotation and statistics elements. In this case, the
above described arguments become: \itemize{ \item
\code{exon.filters=list(min.active.exons=list(exons.per.gene=5,min.exons=2,frac=1/5))}
\item \code{gene.filters=list(length=list(length=500),}
\code{avg.reads=list(average.per.bp=100,quantile=0.25),}
\code{expression=list(median=TRUE,mean=FALSE,quantile=NA,known=NA,custom=NA),}
\code{biotype=get.defaults("biotype.filter",org[1]))}
\item \code{pcut=0.05} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change","stats","counts")}
\item \code{export.scale=c("natural","log2")} \item
\code{export.values=c("normalized")} \item
\code{export.stats=c("mean","sd","cv")} } and statistics
elements. In this case, the above described arguments
become: \itemize{ \item
\code{exon.filters=list(min.active.exons=list(exons.per.gene=5,min.exons=2,frac=1/5))}
\item \code{gene.filters=list(length=list(length=500),}
\code{avg.reads=list(average.per.bp=100,quantile=0.25),}
\code{expression=list(median=TRUE,mean=FALSE,quantile=NA,known=NA,custom=NA),}
\code{biotype=get.defaults("biotype.filter",org[1]))}
\item \code{pcut=0.05} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change","stats","counts")}
\item
\code{export.scale=c("natural","log2","log10","vst")}
\item \code{export.values=c("raw","normalized")} \item
\code{export.stats=c("mean","median","sd","mad","cv","rcv")}
} \item \code{"strict.basic"}: apply a strict set of
filters and and export statistically significant genes
and basic annotation and statistics elements. In this
case, the above described arguments become: \itemize{
\item
\code{exon.filters=list(min.active.exons=list(exons.per.gene=4,min.exons=2,frac=1/4))}
\item \code{gene.filters=list(length=list(length=750),}
\code{avg.reads=list(average.per.bp=100,quantile=0.5),}
\code{expression=list(median=TRUE,mean=FALSE,quantile=NA,known=NA,custom=NA),}
\code{biotype=get.defaults("biotype.filter",org[1]))}
\item \code{pcut=0.01} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change")} \item
\code{export.scale=c("natural","log2")} \item
\code{export.values=c("normalized")} \item
\code{export.stats=c("mean")} } \item
\code{"strict.normal"}: apply a strict set of filters and
and export statistically significant genes and normal
annotation and statistics elements. In this case, the
above described arguments become: \itemize{ \item
\code{exon.filters=list(min.active.exons=list(exons.per.gene=4,min.exons=2,frac=1/4))}
\item \code{gene.filters=list(length=list(length=750),}
\code{avg.reads=list(average.per.bp=100,quantile=0.5),}
\code{expression=list(median=TRUE,mean=FALSE,quantile=NA,known=NA,custom=NA),}
\code{biotype=get.defaults("biotype.filter",org[1]))}
\item \code{pcut=0.01} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change","stats","counts")}
\item \code{export.scale=c("natural","log2")} \item
\code{export.values=c("normalized")} \item
\code{export.stats=c("mean","sd","cv")} } and statistics
elements. In this case, the above described arguments
become: \itemize{ \item
\code{exon.filters=list(min.active.exons=list(exons.per.gene=4,min.exons=2,frac=1/4))}
\item \code{gene.filters=list(length=list(length=750),}
\code{avg.reads=list(average.per.bp=100,quantile=0.5),}
\code{expression=list(median=TRUE,mean=FALSE,quantile=NA,known=NA,custom=NA),}
\code{biotype=get.defaults("biotype.filter",org[1]))}
\item \code{pcut=0.01} \item
\code{export.what=c("annotation","p.value","adj.p.value","meta.p.value",}
\code{"adj.meta.p.value","fold.change","stats","counts")}
\item
\code{export.scale=c("natural","log2","log10","vst")}
\item \code{export.values=c("raw","normalized")} \item
\code{export.stats=c("mean","median","sd","mad","cv","rcv")}
} }
}
\examples{
\donttest{
# An example pipeline with exon counts
data("hg19.exon.data",package="metaseqR")
metaseqr(
counts=hg19.exon.counts,
sample.list=list(normal="normal",paracancerous="paracancerous",cancerous="cancerous"),
contrast=c("normal_vs_paracancerous","normal_vs_cancerous",
"normal_vs_paracancerous_vs_cancerous"),
libsize.list=libsize.list.hg19,
id.col=4,
annotation="download",
org="hg19",
count.type="exon",
normalization="edaseq",
statistics="deseq",
pcut=0.05,
qc.plots=c("mds", "biodetection", "countsbio", "saturation", "rnacomp",
"boxplot", "gcbias", "lengthbias", "meandiff", "readnoise","meanvar",
"readnoise", "deheatmap", "volcano", "biodist", "filtered"),
fig.format=c("png","pdf"),
export.what=c("annotation","p.value","adj.p.value","fold.change","stats",
"counts"),
export.scale=c("natural","log2","log10","vst"),
export.values=c("raw","normalized"),
export.stats=c("mean","median","sd","mad","cv","rcv"),
restrict.cores=0.8,
gene.filters=list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.25
),
expression=list(
median=TRUE,
mean=FALSE
),
biotype=get.defaults("biotype.filter","hg18")
)
)
# An example pipeline with gene counts
data("mm9.gene.data",package="metaseqR")
result <- metaseqr(
counts=mm9.gene.counts,
sample.list=list(e14.5=c("e14.5_1","e14.5_2"), adult_8_weeks=c("a8w_1","a8w_2")),
contrast=c("e14.5_vs_adult_8_weeks"),
libsize.list=libsize.list.mm9,
annotation="download",
org="mm9",
count.type="gene",
normalization="edger",
statistics=c("deseq","edger","noiseq"),
meta.p="fisher",
pcut=0.05,
fig.format=c("png","pdf"),
export.what=c("annotation","p.value","meta.p.value","adj.meta.p.value",
"fold.change"),
export.scale=c("natural","log2"),
export.values="normalized",
export.stats=c("mean","sd","cv"),
export.where=getwd(),
restrict.cores=0.8,
gene.filters=list(
length=list(
length=500
),
avg.reads=list(
average.per.bp=100,
quantile=0.25
),
expression=list(
median=TRUE,
mean=FALSE,
quantile=NA,
known=NA,
custom=NA
),
biotype=get.defaults("biotype.filter","mm9")
),
out.list=TRUE
)
head(result$data[["e14.5_vs_adult_8_weeks"]])
}
}
\author{
<NAME>
}
<file_sep>/man/load.bs.genome.Rd
\name{load.bs.genome}
\alias{load.bs.genome}
\title{Loads (or downloads) the required BSGenome package}
\usage{
load.bs.genome(org)
}
\arguments{
\item{org}{one of \code{\link{metaseqr}} supported
organisms.}
}
\value{
A proper BSgenome package name.
}
\description{
Retrieves the required BSgenome package when the
annotation source is \code{"ucsc"} or
\code{"refseq"}. These packages are required in order
to estimate the GC-content of the retrieved genes from
UCSC or RefSeq.
}
\examples{
\donttest{
bs.obj <- load.bs.genome("mm9")
}
}
\author{
<NAME>
}
<file_sep>/man/get.ucsc.credentials.Rd
\name{get.ucsc.credentials}
\alias{get.ucsc.credentials}
\title{Return host, username and password for UCSC
Genome Browser database}
\usage{
get.ucsc.credentials()
}
\value{
A named character vector.
}
\description{
Returns a character vector with a hostname, username
and password to connect to the UCSC Genome Browser
database to retrieve annotation. Internal use mostly.
}
\examples{
\donttest{
db.creds <- get.ucsc.credentials()
}
}
\author{
<NAME>
}
<file_sep>/man/sample.list.hg19.Rd
\docType{data}
\name{sample.list.hg19}
\alias{sample.list.hg19}
\title{Human RNA-Seq data with three conditions, three samples}
\format{a named \code{list} with condition and sample names.}
\source{
GEO (http://www.ncbi.nlm.nih.gov/geo/)
}
\description{
The sample list for \code{hg19.exon.counts}. See the data
set description.
}
\author{
<NAME>
}
\keyword{datasets}
<file_sep>/man/check.packages.Rd
\name{check.packages}
\alias{check.packages}
\title{Required packages validator}
\usage{
check.packages(m, p)
}
\arguments{
\item{m}{meta-analysis method.}
\item{p}{QC plot types.}
}
\description{
Checks if all the any required packages, not attached
during installation or loading, are present according
to metaseqR input options. Internal use only.
}
\examples{
check.packages(c("simes","whitlock"),
c("gcbias","correl"))
}
\author{
<NAME>
}
<file_sep>/man/diagplot.noiseq.saturation.Rd
\name{diagplot.noiseq.saturation}
\alias{diagplot.noiseq.saturation}
\title{Simpler implementation of saturation plots inspired from
NOISeq package}
\usage{
diagplot.noiseq.saturation(x, o, tb, path = NULL)
}
\arguments{
\item{x}{the count data matrix.}
\item{o}{one or more R plotting device to direct the plot
result to. Supported mechanisms: \code{"x11"} (default),
\code{"png"}, \code{"jpg"}, \code{"bmp"}, \code{"pdf"} or
\code{"ps"}.}
\item{tb}{the vector of biotypes, one for each row of x.}
\item{path}{the path to create output files.}
}
\value{
The filenames of the plots produced in a named list with
names the \code{which.plot} argument. If
\code{output="x11"}, no output filenames are produced.
}
\description{
Helper function for \code{\link{diagplot.noiseq}} to plot
feature detection saturation as presented in the NOISeq
package vignette. It has two main outputs: a set of
figures, one for each input sample depicting the
saturation for each biotype and one single multiplot
which depicts the saturation of all samples for each
biotype. It expands the saturation plots of NOISeq by
allowing more samples to be examined in a simpler way.
Don't use this function directly. Use either
\code{\link{diagplot.metaseqr}} or
\code{\link{diagplot.noiseq}}.
}
\author{
<NAME>
}
<file_sep>/man/make.export.list.Rd
\name{make.export.list}
\alias{make.export.list}
\title{Intitialize output list}
\usage{
make.export.list(con)
}
\arguments{
\item{con}{The contrasts.}
}
\value{
An empty named list.
}
\description{
Initializes metaseqr R output. Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/validate.alg.args.Rd
\name{validate.alg.args}
\alias{validate.alg.args}
\title{Validate normalization and statistical
algorithm arguments}
\usage{
validate.alg.args(normalization, statistics,
norm.args, stat.args)
}
\arguments{
\item{normalization}{a keyword determining the
normalization strategy to be performed by
metaseqR. See \code{\link{metaseqr}} main help
page for details.}
\item{statistics}{the statistical tests to be
performed by metaseqR. See \code{\link{metaseqr}}
main help page for details.}
\item{norm.args}{the user input list of
normalization arguments. See
\code{\link{metaseqr}} main help page for
details.}
\item{stat.args}{the user input list of
statistical test arguments. See
\code{\link{metaseqr}} main help page for
details.}
}
\value{
A list with two members (\code{norm.args},
\code{stat.args}) with valid arguments to
be used as user input for the algorithms
supported by metaseqR.
}
\description{
This function checks and validates the arguments
passed by the user to the normalization and
statistics algorithms supported by metaseqR. As
these are given into lists and passed to the
algorithms, the list members must be checked
for \code{NULL}, valid names etc. This function
performs these checks and ignores any invalid
arguments.
}
\examples{
\donttest{
normalization <- "edaseq"
statistics <- "edger"
norm.args <- get.defaults("normalization","edaseq")
stat.args <- get.defaults("statistics","deseq")
# Will return as is
val <- validate.alg.args(normalization,statistics,norm.args,stat.args)
val$norm.args
val$stat.args
# but...
stat.args <- c(stat.args,my.irrelevant.arg=999)
val <- validate.alg.args(normalization,statistics,norm.args,stat.args)
# irrelevant argument will be removed
val$norm.args
val$stat.args
}
}
\author{
<NAME>
}
<file_sep>/man/make.venn.areas.Rd
\name{make.venn.areas}
\alias{make.venn.areas}
\title{Helper for Venn diagrams}
\usage{
make.venn.areas(n)
}
\arguments{
\item{n}{the number of the sets used for the Venn
diagram.}
}
\value{
A named list, see descritpion.
}
\description{
This function creates a list with names the arguments of
the Venn diagram construction functions of the R package
VennDiagram and list members the internal encoding
(uppercase letters A to E and combinations among then)
used to encode the pairwise comparisons to create the
intersections needed for the Venn diagrams. Internal use
mostly.
}
\examples{
\donttest{
sets <- c("apple","pear","banana")
pairs <- make.venn.pairs(sets)
areas <- make.venn.areas(length(sets))
}
}
\author{
<NAME>
}
<file_sep>/man/mlfo.Rd
\name{mlfo}
\alias{mlfo}
\title{MLE dispersion estimate}
\usage{
mlfo(phi, y)
}
\arguments{
\item{phi}{the parameter to be optimized.}
\item{y}{count samples used to perform
the optimization.}
}
\value{
The objective function value.
}
\description{
MLE function used to estimate negative
binomial dispersions from real RNA-Seq
data, as in (Soneson and Delorenzi, BMC
Bioinformatics, 2013) and (Robles et al.,
BMC Genomics, 2012). Internal use.
}
\examples{
\donttest{
# Not yet available
}
}
\author{
<NAME>
}
<file_sep>/man/check.graphics.type.Rd
\name{check.graphics.type}
\alias{check.graphics.type}
\title{Check plotting device}
\usage{
check.graphics.type(o)
}
\arguments{
\item{o}{the plotting device, see main metaseqr function}
}
\description{
Plotting device checker. Internal use only.
}
\author{
<NAME>
}
<file_sep>/man/make.html.table.Rd
\name{make.html.table}
\alias{make.html.table}
\title{HTML report helper}
\usage{
make.html.table(b, h = NULL, id = NULL)
}
\arguments{
\item{b}{the table body as produced by
\code{\link{make.html.body}}.}
\item{h}{the table header as produced by
\code{\link{make.html.header}}.}
\item{id}{the table id attribute.}
}
\value{
A fully formatted html table.
}
\description{
Returns a character vector with a fully html formatted
table. Essentially, it binds the outputs of
\code{\link{make.html.cells}},
\code{\link{make.html.rows}},
\code{\link{make.html.header}} and
\code{\link{make.html.body}} to the final table and
optionally assigns an id attribute. The above functions
are meant to format a data table so as it can be rendered
by external tools such as DataTables.js during a report
creation. It is meant for internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
the.cells <- make.html.cells(data.matrix)
the.header <- make.html.header(the.cells[1,])
the.rows <- make.html.rows(the.cells)
the.body <- make.html.body(the.rows)
the.table <- make.html.table(the.body,the.header,id="my_table")
}
}
\author{
<NAME>
}
<file_sep>/man/make.html.rows.Rd
\name{make.html.rows}
\alias{make.html.rows}
\title{HTML report helper}
\usage{
make.html.rows(mat)
}
\arguments{
\item{mat}{the data matrix, usually the output of
\code{\link{make.html.cells}} function.}
}
\value{
A character vector with html formatted rows of a matrix.
}
\description{
Returns a character vector with html formatted rows.
Essentially, it collapses every row of a matrix to a
single character and puts a <tr></tr> tag set around. It
is meant to be applied to the output of
\code{\link{make.html.cells}}. Internal use.
}
\examples{
\donttest{
data.matrix <- round(1000*matrix(runif(400),100,4))
rownames(data.matrix) <- paste("gene_",1:100,sep="")
colnames(data.matrix) <- c("C1","C2","T1","T2")
the.cells <- make.html.cells(data.matrix)
the.rows <- make.html.rows(the.cells)
}
}
\author{
<NAME>
}
<file_sep>/man/normalize.nbpseq.Rd
\name{normalize.nbpseq}
\alias{normalize.nbpseq}
\title{Normalization based on the NBPSeq package}
\usage{
normalize.nbpseq(gene.counts, sample.list,
norm.args = NULL, libsize.list = NULL,
output = c("matrix", "native"))
}
\arguments{
\item{gene.counts}{a table where each row represents a
gene and each column a sample. Each cell contains the
read counts for each gene and sample. Such a table can be
produced outside metaseqr and is imported during the
basic metaseqr workflow.}
\item{sample.list}{the list containing condition names
and the samples under each condition.}
\item{norm.args}{a list of NBPSeq normalization
parameters. See the result of
\code{get.defaults("normalization",} \code{"nbpseq")} for
an example and how you can modify it.}
\item{libsize.list}{an optional named list where names
represent samples (MUST be the same as the samples in
\code{sample.list}) and members are the library sizes
(the sequencing depth) for each sample. If not provided,
the default is the column sums of the \code{gene.counts}
matrix.}
\item{output}{the class of the output object. It can be
\code{"matrix"} (default) for versatility with other
tools or \code{"native"} for the NBPSeq native S4 object
(a specific list). In the latter case it should be
handled with suitable NBPSeq methods.}
}
\value{
A matrix with normalized counts or a list with the
normalized counts and other NBPSeq specific parameters.
}
\description{
This function is a wrapper over DESeq normalization. It
accepts a matrix of gene counts (e.g. produced by
importing an externally generated table of counts to the
main metaseqr pipeline).
}
\examples{
\donttest{
require(DESeq)
data.matrix <- counts(makeExampleCountDataSet())
sample.list <- list(A=c("A1","A2"),B=c("B1","B2","B3"))
diagplot.boxplot(data.matrix,sample.list)
norm.data.matrix <- normalize.nbpseq(data.matrix,sample.list)
diagplot.boxplot(norm.data.matrix,sample.list)
}
}
\author{
<NAME>
}
<file_sep>/man/calc.f1score.Rd
\name{calc.f1score}
\alias{calc.f1score}
\title{Calculate the F1-score}
\usage{
calc.f1score(truth, p, sig = 0.05)
}
\arguments{
\item{truth}{the ground truth differential
expression vector. It should contain only
zero and non-zero elements, with zero denoting
non-differentially expressed genes and non-zero,
differentially expressed genes. Such a vector
can be obtained for example by using the
\code{\link{make.sim.data.sd}} function, which
creates simulated RNA-Seq read counts based on
real data. It MUST be named with gene names,
the same as in \code{p}.}
\item{p}{a p-value matrix whose rows correspond
to each element in the \code{truth} vector. If
the matrix has a \code{colnames} attribute, a
legend will be added to the plot using these
names, else a set of column names will be
auto-generated. \code{p} can also be a list or
a data frame. In any case, each row (or element)
MUST be named with gene names (the same as in
\code{truth}).}
\item{sig}{a significance level (0 < \code{sig}
<=1).}
}
\value{
A named list with two members. The first member
is a data frame with the numbers used to
calculate the TP/(FP+FN) ratio and the second
member is the ratio TP/(FP+FN) for each
statistical test.
}
\description{
This function calculates the F1 score
(2*(precision*recall/precision+racall) or
2*TP/(2*TP+FP+FN) given a matrix of p-values (one
for each statistical test used) and a vector of
ground truth (DE or non-DE). This function serves
as a method evaluation helper.
}
\examples{
p1 <- 0.001*matrix(runif(300),100,3)
p2 <- matrix(runif(300),100,3)
p <- rbind(p1,p2)
rownames(p) <- paste("gene",1:200,sep="_")
colnames(p) <- paste("method",1:3,sep="_")
truth <- c(rep(1,40),rep(-1,40),rep(0,20),rep(1,10),
rep(2,10),rep(0,80))
names(truth) <- rownames(p)
f1 <- calc.f1score(truth,p)
}
\author{
<NAME>
}
<file_sep>/man/get.strict.biofilter.Rd
\name{get.strict.biofilter}
\alias{get.strict.biofilter}
\title{Group together a more strict biotype filter}
\usage{
get.strict.biofilter(org)
}
\arguments{
\item{org}{one of the supported organisms.}
}
\value{
A list of booleans, one for each biotype.
}
\description{
Returns a list with TRUE/FALSE according to the biotypes
that are going to be filtered in a more strict way than
the defaults. This is a helper function for the analysis
presets of metaseqR. Internal use only.
}
\examples{
\donttest{
sf <- get.strict.biofilter("hg18")
}
}
\author{
<NAME>
}
| f06961d1fc7fd9fe4860f75cd76e4269c00d46d7 | [
"Markdown",
"R"
] | 146 | R | pmoulos/metaseqr | 81b42a9120c1830a46e68a50ce9dddb975975edf | 175f4e1f79593697fc98aa8b775cbd8c9d6ff3d7 |
refs/heads/master | <file_sep>public class SomeProgram {
public static void main(String[] args) {
System.out.println("We are learning to use Gitzmiana.");
}
public static void main(String[] args) {
System.out.println("Zarządzanie rakietami.");
} | 66eeb14294eb51c24599ab90b42b765d44206bb5 | [
"Java"
] | 1 | Java | michalowska11/mwo-git | 7728d665e61a51b58c08eb5eaf0d1a498a856b28 | 6877d21cf1a0d6aba1eb30feb9a46d74e4ac5e2c |
refs/heads/master | <file_sep>*Work in progress, adding sweet pics as I go.*
# Features
- Live OpenGL rendering
- Wavefront object loading
# Rendering
- KD tree for bounding boxes
- Cosine importance sampling
# Gallery


<file_sep>R"(
#version 330
in vec2 tex_coord;
uniform sampler2D tex;
void main()
{
gl_FragColor = pow(texture(tex, tex_coord), vec4(1 / 2.2));
}
)"
<file_sep>#include "aabb.hpp"
#include "utils.hpp"
#include <algorithm>
// Modified slabs method from Real Time Rendering ch 16.7.1
bool Box::intersect(const Ray &ray)
{
float t_min = -INF;
float t_max = INF;
vec3f to_p1 = ll - ray.pos;
vec3f to_p2 = ur - ray.pos;
// TODO extra checks.
for (int i=0; i<3; i++) {
float f = ray.dir[i];
float t1 = to_p1[i] / f;
float t2 = to_p2[i] / f;
if (t1 > t2) {
float temp = t1;
t1 = t2;
t2 = temp;
}
t_min = std::max(t1, t_min);
t_max = std::min(t2, t_max);
if (t_min > t_max) return false;
if (t_max < 0) return false;
}
return true;
}
<file_sep>#include <algorithm>
#include <chrono>
#include <deque>
#include <functional>
#include <iostream>
#include <libgen.h>
#include <time.h>
#include <vector>
#include "image.hpp"
#include "model.hpp"
#include "scene.hpp"
Scene::Scene(std::string model_name)
{
std::string err;
std::cout << "Loading model " << model_name << std::endl;
unsigned int flags =
tinyobj::triangulation | tinyobj::calculate_normals;
std::string model_name_copy(model_name);
char* model_name_char = const_cast<char*>(model_name_copy.c_str());
char* obj_dir = dirname(model_name_char);
strcat(obj_dir, "/");
bool success = tinyobj::LoadObj(m_shapes, m_mats, err,
model_name.c_str(), obj_dir, flags);
if (!success) {
std::cout << err << std::endl;
std::cerr << "Model failed to load, Exiting." << std::endl;
std::exit(EXIT_FAILURE);
} else {
std::cout << "Model loaded successfully." << std::endl;
}
std::cout << "Constructing triangles" << std::endl;
std::vector<Triangle> tris;
for (size_t s=0; s < m_shapes.size(); s++) {
Shape shape = m_shapes[s];
tinyobj::mesh_t mesh = shape.mesh;
for (size_t i=0; i < mesh.indices.size(); i += 3) {
unsigned int j1 = mesh.indices[i]*3;
unsigned int j2 = mesh.indices[i+1]*3;
unsigned int j3 = mesh.indices[i+2]*3;
vec3f v1 = to_vec3f(&mesh.positions[j1]);
vec3f v2 = to_vec3f(&mesh.positions[j2]);
vec3f v3 = to_vec3f(&mesh.positions[j3]);
vec3f norm = to_vec3f(&mesh.normals[j1]);
Triangle tri = {
.verts[0] = v1,
.verts[1] = v2,
.verts[2] = v3,
.norm = norm,
.shape_data = &(m_shapes[s]),
.index = i
};
tris.push_back(tri);
}
}
std::cout << "Constructing KdTree" << std::endl;
m_tree = new KdTree(tris);
}
<file_sep>#ifndef VECTOR_H
#define VECTOR_H
#include <eigen3/Eigen/Dense>
#include <vector>
#include <numeric>
typedef Eigen::Vector3f vec3f;
typedef Eigen::Vector3d vec3i;
inline vec3f to_vec3f(float* a)
{
vec3f v(a[0], a[1], a[2]);
return v;
}
inline vec3f unit(const vec3f &v)
{
vec3f vn(v / v.norm());
return vn;
}
inline vec3f vec_average(const std::vector<vec3f> &vecs)
{
vec3f accum(0.0, 0.0, 0.0);
accum = std::accumulate(vecs.begin(), vecs.end(), accum);
return accum / vecs.size();
}
inline vec3f rand_hemisphere_vec(const vec3f &norm)
{
vec3f randy = unit(Eigen::Vector3f::Random());
if (randy.dot(norm) < 0) {
return -randy;
} else {
return randy;
}
}
inline vec3f cos_dist_hemisphere_vec(const vec3f &norm)
{
float theta = 2 * M_PI * ((float) rand() / (float) RAND_MAX);
float radius = sqrtf((float) rand() / (float) RAND_MAX);
float radius_sqrt = sqrtf(radius);
// Generate vector non-parallel to norm.
vec3f base;
do {
base = unit(Eigen::Vector3f::Random());
} while (fabs(base.dot(norm)) < 0.01);
vec3f u = norm.cross(base);
vec3f v = norm.cross(u);
return unit((norm * sqrtf(1 - radius)) + radius_sqrt * (u * cos(theta)) + (v * sin(theta)));
}
#endif
<file_sep>#ifndef UTILS_H
#define UTILS_H
#define INF std::numeric_limits<float>::infinity()
#endif
<file_sep>#include <cstdlib>
#include <fstream>
#include <iostream>
#include <math.h>
#include <stdio.h>
#include <stdlib.h>
#include <string>
#include <time.h>
#include <vector>
#include "frontends.hpp"
#include "image.hpp"
#include "scene.hpp"
// TODO gamma correction / scaling.
void ImageFrontend::render_image(RenderOpts render_opts, Scene &scene,
std::string outfile_path) {
Renderer renderer(render_opts);
renderer.start_render(scene);
std::cout << "{0% " << std::string(render_opts.bar_length - 8, ' ')
<< " 100%}" << std::endl;
std::cout << "{" << std::flush;
auto start = std::chrono::steady_clock::now();
std::vector<vec3f> *pixels = renderer.get_pixels();
const int total_needed = renderer.m_render_opts.image_width
* renderer.m_render_opts.image_height
* renderer.m_render_opts.num_samples;
// Draw a progress bar while rendering.
int dot_delta = total_needed / render_opts.bar_length;
int pixels_done = 0;
int total_dots = 0;
while (renderer.m_pixels_done < total_needed) {
std::this_thread::sleep_for(std::chrono::milliseconds(300));
pixels_done = renderer.m_pixels_done;
int last_dot = total_dots * dot_delta;
if (pixels_done - last_dot >= dot_delta) {
int new_dots = (pixels_done - last_dot) / dot_delta;
total_dots += new_dots;
if (total_dots > render_opts.bar_length) {
new_dots -= total_dots - render_opts.bar_length;
}
std::cout << std::string(new_dots, '.') << std::flush;
}
}
renderer.stop_render(true);
auto end = std::chrono::steady_clock::now();
std::cout << "}" << std::endl;
std::cout << "Traced image in " <<
(std::chrono::duration_cast<std::chrono::milliseconds>
(end - start).count() / 1000.0) << " seconds." << std::endl;
std::cout << "Saving image to " << outfile_path << std::endl;
write_png(outfile_path.c_str(), *pixels,
renderer.m_render_opts.image_width,
renderer.m_render_opts.image_height);
}
void OpenGLFrontend::render_scene(RenderOpts render_opts, Scene &scene) {
Renderer renderer(render_opts);
renderer.start_render(scene);
std::vector<vec3f> *pixels = renderer.get_pixels();
GLvoid* pixel_data = &((*pixels)[0]);
GLFWwindow* win = glfw_setup(render_opts.image_width, render_opts.image_height);
load_program();
GLfloat verts[] =
{
-1.0f, 1.0f,
-1.0f, -1.0f,
1.0f, 1.0f,
1.0f, -1.0f,
};
GLfloat tex_coords[] =
{
0.0f, 0.0f,
0.0f, 1.0f,
1.0f, 0.0f,
1.0f, 1.0f,
};
GLuint vert_vbo = 0;
glGenBuffers(1, &vert_vbo);
glBindBuffer(GL_ARRAY_BUFFER, vert_vbo);
glBufferData(GL_ARRAY_BUFFER, sizeof(verts), verts, GL_STATIC_DRAW);
GLuint tex_vbo = 0;
glGenBuffers(1, &tex_vbo);
glBindBuffer(GL_ARRAY_BUFFER, tex_vbo);
glBufferData(GL_ARRAY_BUFFER, sizeof(tex_coords), tex_coords, GL_STATIC_DRAW);
glTexParameteri(GL_TEXTURE_2D, GL_TEXTURE_MIN_FILTER, GL_LINEAR);
glTexParameteri(GL_TEXTURE_2D, GL_TEXTURE_MAG_FILTER, GL_LINEAR);
glTexParameteri(GL_TEXTURE_2D, GL_TEXTURE_WRAP_S, GL_CLAMP_TO_BORDER);
glTexParameteri(GL_TEXTURE_2D, GL_TEXTURE_WRAP_T, GL_CLAMP_TO_BORDER);
GLuint tex;
glGenTextures(1, &tex);
glActiveTexture(GL_TEXTURE0);
glBindTexture(GL_TEXTURE_2D, tex);
glTexImage2D(GL_TEXTURE_2D, 0, GL_RGB, render_opts.image_width,
render_opts.image_height, 0, GL_RGB, GL_FLOAT, pixel_data);
glGenerateMipmap(GL_TEXTURE_2D);
GLuint vao = 0;
glGenVertexArrays(1, &vao);
glBindVertexArray(vao);
glEnableVertexAttribArray(0);
glBindBuffer(GL_ARRAY_BUFFER, vert_vbo);
glVertexAttribPointer(0, 2, GL_FLOAT, GL_FALSE, 0, (GLvoid*) 0);
glEnableVertexAttribArray(1);
glBindBuffer(GL_ARRAY_BUFFER, tex_vbo);
glVertexAttribPointer(1, 2, GL_FLOAT, GL_FALSE, 0, (GLvoid*) 0);
float pixels_one_frame = render_opts.image_height * render_opts.image_width;
while (!glfwWindowShouldClose(win)) {
std::this_thread::sleep_for(std::chrono::milliseconds(70));
std::cout << "Pixels rendered: " << renderer.m_pixels_done << ", or ~"
<< renderer.m_pixels_done / pixels_one_frame << " samples per pixel"
<< std::endl;
glClear(GL_COLOR_BUFFER_BIT | GL_DEPTH_BUFFER_BIT);
glDrawArrays(GL_TRIANGLE_STRIP, 0, 4);
glfwSwapBuffers(win);
glfwPollEvents();
pixel_data = &((*pixels)[0]);
glBindBuffer(GL_ARRAY_BUFFER, tex_vbo);
glBufferData(GL_ARRAY_BUFFER, sizeof(tex_coords), tex_coords, GL_STATIC_DRAW);
glBindTexture(GL_TEXTURE_2D, tex);
glTexImage2D(GL_TEXTURE_2D, 0, GL_RGB, render_opts.image_width,
render_opts.image_height, 0, GL_RGB, GL_FLOAT, pixel_data);
glGenerateMipmap(GL_TEXTURE_2D);
}
glfwTerminate();
exit(EXIT_SUCCESS);
}
int OpenGLFrontend::load_shader(std::string shader_text, GLenum shader_type)
{
const GLchar* shader_source = shader_text.c_str();
GLuint shader = glCreateShader(shader_type);
glShaderSource(shader, 1, (const GLchar **) &shader_source, NULL);
glCompileShader(shader);
// Shader error reporting
GLint is_compiled = GL_FALSE;
glGetShaderiv(shader, GL_COMPILE_STATUS, &is_compiled);
if(is_compiled == GL_FALSE) {
GLint max_len = 0;
glGetShaderiv(shader, GL_INFO_LOG_LENGTH, &max_len);
GLchar* infoLog = (GLchar*) malloc(max_len * sizeof(GLchar));
glGetShaderInfoLog(shader, max_len, &max_len, infoLog);
printf("%s:%s\n", shader_source, infoLog);
glDeleteShader(shader);
exit(EXIT_FAILURE);
}
return shader;
}
int OpenGLFrontend::load_program()
{
GLuint vs = load_shader(vertex_shader, GL_VERTEX_SHADER);
GLuint fs = load_shader(fragment_shader, GL_FRAGMENT_SHADER);
GLuint shader_program = glCreateProgram();
glAttachShader(shader_program, vs);
glAttachShader(shader_program, fs);
glLinkProgram(shader_program);
glUseProgram(shader_program);
return shader_program;
}
// GLFW Callbacks
void error_callback(int error, const char* description)
{
std::cerr << "Error " << error << ": " << description << std::endl;
}
void key_callback(GLFWwindow* window, int key, int scancode, int action, int mods)
{
if (key == GLFW_KEY_Q) {
std::cout << "Exiting renderer." << std::endl;
std::exit(EXIT_SUCCESS);
}
}
GLFWwindow* OpenGLFrontend::glfw_setup(int win_res_x, int win_res_y)
{
// Windowing setup
if (!glfwInit()) {
std::cerr << "ERROR: couldn't start GLFW3." << std::endl;
exit(EXIT_FAILURE);
}
glfwWindowHint(GLFW_CONTEXT_VERSION_MAJOR, 3);
glfwWindowHint(GLFW_CONTEXT_VERSION_MINOR, 3);
glfwWindowHint(GLFW_OPENGL_PROFILE, GLFW_OPENGL_CORE_PROFILE);
GLFWwindow* win = glfwCreateWindow(win_res_x, win_res_y, "Opengl Demo",
NULL, NULL);
glfwSetErrorCallback(error_callback);
if (!win) {
std::cerr << "ERROR: couldn't open window.\n" << std::endl;
glfwTerminate();
exit(EXIT_FAILURE);
}
glfwMakeContextCurrent(win);
glfwSetKeyCallback(win, key_callback);
glewExperimental = GL_TRUE;
glewInit();
glEnable(GL_DEPTH_TEST);
glDepthFunc(GL_LESS);
return win;
}
<file_sep>#ifndef AABB_H
#define AABB_H
#include "vector.hpp"
#include "ray.hpp"
// Axis aligned bounding box (AABB)
class Box {
public:
vec3f ll, ur;
bool intersect(const Ray &ray);
};
#endif
<file_sep>#ifndef RENDERER_H
#define RENDERER_H
#include <atomic>
#include "camera.hpp"
#include "ray.hpp"
#include "scene.hpp"
#include "vector.hpp"
#include "workqueue.hpp"
typedef struct {
int image_width, image_height; // Image dimensions in pixels
int num_samples; // Number of samples for each pixel
int num_bounces; // Number of bounces for each ray
int num_threads; // Number of threads to use for rendering.
int bar_length; // Number of '.' to show in progress bar.
float fov; // Field of view angle in radians
vec3f cam_eye, cam_up, cam_at;
} RenderOpts;
class Renderer {
private:
std::vector<vec3f> m_pixel_buf; // Internal buffer for RGB pixel data.
std::vector<int> m_sample_counts; // How many times each pixel has been sampled.
float m_frust_top, m_frust_bottom, m_frust_right, m_frust_left;
bool m_render_running = false;
WorkQueue m_pixel_queue; // Consumer queue to read next pixel from.
std::vector<std::thread> m_workers; // Thread pool for parallel pixel rendering.
Camera* m_camera;
/* Cast a ray into the scene and return a color. */
vec3f sample(Scene& scene, Ray ray, int num_bounces);
public:
std::atomic_int m_pixels_done;
RenderOpts m_render_opts;
std::vector<vec3f>* get_pixels() { return &m_pixel_buf; }
void start_render(Scene& scene);
void stop_render(bool wait);
void work_render(Scene& scene);
int get_num_pixels();
void update_frustum_view();
Renderer(RenderOpts render_opts);
};
#endif
<file_sep>project(pathtracer)
cmake_minimum_required(VERSION 0.1)
set(CMAKE_EXPORT_COMPILE_COMMANDS ON)
file(GLOB_RECURSE sources "src/*.cpp" "src/*.hpp")
include_directories("src/lib")
add_library(tiny_obj_loader STATIC "src/lib/tiny_obj_loader.cc" "src/lib/tiny_obj_loader.h")
set(CMAKE_CXX_COMPILER clang++)
set(CMAKE_CXX_FLAGS "${CMAKE_CXX_FLAGS} -std=c++11 -Ofast -Wall -Weffc++ -Wuninitialized")
set(CMAKE_EXE_LINKER_FLAGS "${CMAKE_EXE_LINKER_FLAGS} -lGL -lGLEW -lGLU -lSOIL -lX11 -lXi -lXrandr -lXxf86vm -lglfw -lpng -lprofiler -lpthread")
add_executable(pathtracer ${sources})
target_link_libraries(pathtracer tiny_obj_loader)
<file_sep>#ifndef KDTREE_H
#define KDTREE_H
#include <vector>
#include "aabb.hpp"
#include "triangle.hpp"
class KdTree {
private:
Box m_box;
KdTree *m_left, *m_right;
std::vector<Triangle> m_tris;
public:
KdTree(std::vector<Triangle> &tris, int dim_split = 0,
int max_tris = 3);
TriangleHit hit(const Ray &ray);
};
#endif
<file_sep>#ifndef WORKQUEUE_H
#define WORKQUEUE_H
#include <deque>
#include <thread>
#include <mutex>
/* A simple thread-safe queue for integer operations. */
class WorkQueue {
private:
std::mutex m_mutex;
std::deque<int> m_queue;
public:
WorkQueue() {};
void push_back(int n);
int pop_front();
int size();
};
#endif
<file_sep>#include "camera.hpp"
#include "ray.hpp"
#include "vector.hpp"
Camera::Camera(Ray view, vec3f up_dir)
{
m_pos = view.pos;
m_view = unit(view.dir);
m_right = m_view.cross(unit(up_dir));
m_up = m_right.cross(m_view);
};
<file_sep>/*
* A pathtracer. This is a successor to a raytracer I wrote a few years ago.
* Hopefully I can get some even cooler pictures out of this one. This time
* around I'm using libraries for wavefront .obj loading, .png writing, and
* vector math, whereas before they were hand rolled. Much nicer this way.
*
* Requires: tinyobjloader (included), Eigen, png++, soil,
* OpenGL 3.3 (for live-render), glfw, glew
*
* Building: Only tested on Linux. See Makefile.
*
* Related:
* - Metropolis light transport
* - Volumetric Pathtracing
* - PBRT book
*
* References:
* https://en.wikipedia.org/wiki/Path_tracing
* https://en.wikipedia.org/wiki/Rendering_equation
* http://www.flipcode.com/archives/Raytracing_Topics_Techniques-Part_7_Kd-Trees_and_More_Speed.shtml
* https://blog.frogslayer.com/kd-trees-for-faster-ray-tracing-with-triangles/
* http://www.iquilezles.org/www/articles/simplepathtracing/simplepathtracing.htm
* https://www.cs.cornell.edu/courses/CS6630/
* http://www.kevinbeason.com/smallpt/
* http://raytracey.blogspot.com/2016/01/gpu-path-tracing-tutorial-3-take-your.html
*
* Other projects / libraries:
* https://github.com/embree/embree
*
*
* Author: <NAME>, 2016
*/
// TODO: Moving camera in live render
// Movement speed depends on scene size
// TODO AA
// TODO Russian-roulette path termination
// TODO look at logging libraries
// https://github.com/easylogging/easyloggingpp
// https://github.com/easylogging/easyloggingpp
// TODO: Snapshot picture save from OpenGL frontend.
// OPT: Cache opts
// OPT: opencl?
#include "getopt.h"
#include <cstdlib>
#include <iostream>
#include <sstream>
#include <string>
#include "frontends.hpp"
#include "renderer.hpp"
#include "scene.hpp"
void show_help() {
std::cout << "Usage: ./pathtracer [options]\n"
"Flags: -h: Print this help.\n"
" -i: Input model file.\n"
" -o: Output image file.\n"
" -r: Use real-time OpenGL output.\n"
<< std::endl;
}
int main(int argc, char *argv[]) {
// Default args
bool real_time = false;
int image_width = 512, image_height = 512;
int num_samples = 100;
int num_bounces = 5;
int num_threads = 9;
// Required args
std::string model_name;
std::string output_name;
std::stringstream argparse_errors;
char cli_opt = 0;
while ((cli_opt = getopt(argc, argv, "b:d:hi:o:rs:t:")) != -1) {
switch (cli_opt) {
case 'b':
sscanf(optarg, "%d", &num_bounces);
break;
case 'd':
sscanf(optarg, "%dx%d", &image_width, &image_height);
break;
case 'i':
model_name.assign(optarg);
break;
case 'o':
output_name.assign(optarg);
break;
case 'r':
real_time = true;
break;
case 's':
sscanf(optarg, "%d", &num_samples);
break;
case 't':
sscanf(optarg, "%d", &num_threads);
break;
case 'h':
show_help();
std::exit(EXIT_SUCCESS);
break;
}
}
if (model_name.empty()) {
argparse_errors << "You must specify an input model with -i <model>"
<< std::endl;
}
if (output_name.empty() && !real_time) {
argparse_errors
<< "You must specify an output image with -o <image> or use -r."
<< std::endl;
}
if (!(argparse_errors.rdbuf()->in_avail() == 0)) {
std::cerr << argparse_errors.str() << std::endl;
std::cerr << "Invalid flags given. Use -h for usage info."
<< std::endl;
std::exit(EXIT_FAILURE);
}
// TODO Set through config
vec3f look_eye(0, 1.0, 4.0);
vec3f look_dir(0.0, 0, -1.0);
vec3f up_dir(0, 1.0, 0.0);
// Pathtracer settings
RenderOpts render_opts = {
.image_width = image_width,
.image_height = image_height,
.num_samples = num_samples,
.num_bounces = num_bounces,
.num_threads = num_threads,
.bar_length = 72,
.fov = M_PI / 5.0,
.cam_up = up_dir,
.cam_eye = look_eye,
.cam_at = look_dir,
};
std::cout << "Preprocessing scene" << std::endl;
Scene scene(model_name);
if (real_time) {
std::cout << "Using OpenGL context for rendering." << std::endl;
OpenGLFrontend rendering_frontend;
rendering_frontend.render_scene(render_opts, scene);
} else {
std::cout << "Rendering to PNG image." << std::endl;
ImageFrontend rendering_frontend;
rendering_frontend.render_image(render_opts, scene, output_name);
}
return 0;
}
<file_sep>#include "workqueue.hpp"
void WorkQueue::push_back(int n)
{
std::lock_guard<std::mutex> lock_g(m_mutex);
m_queue.push_back(n);
}
int WorkQueue::pop_front()
{
std::lock_guard<std::mutex> lock_g(m_mutex);
int front = m_queue.front();
m_queue.pop_front();
return front;
}
int WorkQueue::size()
{
std::lock_guard<std::mutex> lock_g(m_mutex);
return m_queue.size();
}
<file_sep>#include <vector>
#include "kdtree.hpp"
// Naive BSP construction (midpoint)
// A tree is constructed with at most max_tris on each leaf.
// OPT: use SAH
// OPT: precalc midpoints
KdTree::KdTree(std::vector<Triangle> &tris, int dim_split, int max_tris)
{
// Leaf creation.
if (tris.size() <= max_tris) {
m_tris = tris; // PROFILE copy
m_left = nullptr;
m_right = nullptr;
return;
}
// Internal node creation.
// Precalculate min/max/average coordinate info.
float midpoint_total = 0.0;
float max_dims[3] = {0.0};
float min_dims[3] = {INF};
for (int i=0; i < tris.size(); i++) {
for (int vert=0; vert < 3; vert++) {
for (int dim=0; dim < 3; dim++) {
float val = tris[i][vert][dim];
if (max_dims[dim] < val) max_dims[dim] = val;
if (min_dims[dim] > val) min_dims[dim] = val;
}
}
midpoint_total += tris[i].midpoint()[dim_split];
}
// Find the average coordinate along dimension dim_split for left / right
// tree separation.
float dim_avg = midpoint_total / tris.size();
// Use max / min dimensions for bounding box creation.
vec3f lower_left = to_vec3f(min_dims);
vec3f upper_right = to_vec3f(max_dims);
m_box.ll = lower_left;
m_box.ur = upper_right;
// Split the tree at the midpoint and construct each half.
std::vector<Triangle> left_tris, right_tris;
for (int i=0; i < tris.size(); i++) {
if (tris[i].midpoint()[dim_split] < dim_avg) {
left_tris.push_back(tris[i]);
} else {
right_tris.push_back(tris[i]);
}
}
int next_dim = (dim_split + 1) % 3;
m_left = new KdTree(left_tris, next_dim);
m_right = new KdTree(right_tris, next_dim);
}
TriangleHit KdTree::hit(const Ray &ray)
{
// At leaf
if (m_left == nullptr && m_right == nullptr) {
TriangleHit hit_data;
float cl_dist = INF;
Triangle *cl_tri = nullptr;
// Find which triangle (if any) our ray hits.
bool hit = false;
for (int t=0; t < m_tris.size(); t++) {
Triangle &tri = m_tris[t];
float dist = tri.intersect(ray);
if (dist < cl_dist && dist != 0) {
hit = true;
cl_dist = dist;
cl_tri = &(m_tris[t]);
}
}
hit_data.tri = cl_tri;
hit_data.dist = cl_dist;
return hit_data;
}
// At interior node
else if (m_box.intersect(ray)) {
TriangleHit left_hit = m_left->hit(ray);
TriangleHit right_hit = m_right->hit(ray);
// We may hit triangles in both trees. In this case, take the
// closer hit.
if (left_hit.dist < right_hit.dist) {
return left_hit;
} else {
return right_hit;
}
}
TriangleHit miss;
return miss;
}
<file_sep>#ifndef IMAGE_H
#define IMAGE_H
#include "vector.hpp"
void write_png(const char* filename, std::vector<vec3f> &pixel_data,
int img_width, int img_height);
#endif
<file_sep>#ifndef CAMERA_H
#define CAMERA_H
#include "ray.hpp"
#include "vector.hpp"
class Camera {
public:
vec3f m_view, m_up, m_right;
vec3f m_pos;
Camera(Ray view, vec3f up);
};
#endif
<file_sep>#ifndef RAY_H
#define RAY_H
#include "vector.hpp"
typedef struct {
vec3f pos, dir;
} Ray;
#endif
<file_sep>#include "triangle.hpp"
#include "ray.hpp"
vec3f& Triangle::operator[] (const int index)
{
return verts[index];
}
vec3f Triangle::midpoint()
{
return (verts[0] + verts[1] + verts[2]) / 3.0;
}
// https://en.wikipedia.org/wiki/Moller-Trumbore_intersection_algorithm
// OPT: precalculate dominant triangle axis.
float Triangle::intersect(const Ray &ray)
{
float EPSILON = 0.0001;
vec3f e1, e2;
vec3f p, q, r;
float det, inv_det, u, v;
float t;
e1 = verts[1] - verts[0];
e2 = verts[2] - verts[0];
p = ray.dir.cross(e2);
det = e1.dot(p);
if (det > -EPSILON && det < EPSILON) return 0;
inv_det = 1.0 / det;
r = ray.pos - verts[0];
u = r.dot(p) * inv_det;
if (u < 0.0 || u > 1.0) return 0;
q = r.cross(e1);
v = ray.dir.dot(q) * inv_det;
if (v < 0.0 || u + v > 1.0) return 0;
t = e2.dot(q) * inv_det;
if (t > EPSILON) {
return t;
} else {
return 0;
}
}
<file_sep>#ifndef MODEL_H
#define MODEL_H
#include "tiny_obj_loader.h"
typedef tinyobj::shape_t Shape;
typedef tinyobj::material_t Material;
#endif
<file_sep>#ifndef SCENE_H
#define SCENE_H
#include <vector>
#include "kdtree.hpp"
#include "model.hpp"
struct Scene {
std::vector<Shape> m_shapes;
std::vector<Material> m_mats;
KdTree* m_tree;
Scene(std::string model_name);
};
#endif
<file_sep>#include <string>
// OpenGL libs
#include <GL/glew.h>
#include <GLFW/glfw3.h>
// Image loading
#include <SOIL/SOIL.h>
// OpenGL Matrix math
#define GLM_FORCE_RADIANS
#define GLM_SWIZZLE GLM_SWIZZLE_XYZW
#include "renderer.hpp"
#include "scene.hpp"
class ImageFrontend {
public:
void render_image(RenderOpts render_opts, Scene &scene, std::string outfile_path);
};
class OpenGLFrontend {
// The shader files contain raw string literals.
std::string vertex_shader =
#include "shader.vert"
;
std::string fragment_shader =
#include "shader.frag"
;
int load_shader(std::string filename, GLenum shader_type);
int load_program();
void update_pixels(float *pixel_data);
GLFWwindow* glfw_setup(int win_res_x, int win_res_y);
public:
void render_scene(RenderOpts render_opts, Scene &scene);
};
void error_callback(int error, const char* description);
<file_sep>#include <algorithm>
#include <chrono>
#include <deque>
#include <functional>
#include <iostream>
#include <libgen.h>
#include <thread>
#include <time.h>
#include <vector>
#include "image.hpp"
#include "model.hpp"
#include "ray.hpp"
#include "renderer.hpp"
#include "scene.hpp"
#include "vector.hpp"
#include "workqueue.hpp"
// DES: Separate renderer class from rendering instance.
Renderer::Renderer(RenderOpts render_opts) : m_render_opts(render_opts) {
Ray look_at = { render_opts.cam_eye, render_opts.cam_at };
m_camera = new Camera(look_at, render_opts.cam_up);
m_pixels_done = 0;
int num_pixels = get_num_pixels();
for (int i=0; i < num_pixels; i++) {
vec3f empty_vec(0.0, 0.0, 0.0);
m_pixel_buf.push_back(empty_vec);
m_sample_counts.push_back(0);
}
update_frustum_view();
// Populate pixel queue with a random ordering of pixels.
std::vector<int> range(num_pixels);
std::iota(range.begin(), range.end(), 0);
std::random_shuffle(range.begin(), range.end());
for (int i=0; i < num_pixels; i++) {
m_pixel_queue.push_back(range[i]);
}
// Seed for places we need random vector directions.
srand(time(NULL));
};
void Renderer::start_render(Scene& scene) {
m_render_running = true;
for (int t=0; t < m_render_opts.num_threads; t++) {
m_workers.push_back(std::thread(&Renderer::work_render, this,
std::ref(scene)));
}
}
void Renderer::stop_render(bool wait=false) {
m_render_running = false;
if (wait) {
for (std::thread &t : m_workers) {
t.join();
}
}
}
int Renderer::get_num_pixels() {
return m_render_opts.image_height * m_render_opts.image_width;
}
vec3f Renderer::sample(Scene& scene, Ray ray, int num_bounces)
{
vec3f accum_radiance(0.0, 0.0, 0.0);
vec3f rem_radiance(1.0, 1.0, 1.0);
for (int b=0; b < num_bounces; b++) {
TriangleHit hit_data = scene.m_tree->hit(ray);
Triangle* tri = hit_data.tri;
float dist = hit_data.dist;
if (tri == nullptr) {
return vec3f(0.0, 0.0, 0.0);
}
tinyobj::mesh_t mesh = tri->shape_data->mesh;
Material mat = scene.m_mats[mesh.material_ids[tri->index / 3]];
// Material properties
vec3f emittance = to_vec3f(mat.emission);
vec3f diffuse = to_vec3f(mat.diffuse);
// vec3f specular = to_vec3f(mat.specular);
vec3f &norm = tri->norm;
// Calculate BRDF
vec3f brdf = 2 * diffuse;
// Reflect in a random direction on the normal's unit hemisphere.
ray.pos = ray.pos + dist * ray.dir;
// ray.dir = rand_hemisphere_vec(norm);
ray.dir = cos_dist_hemisphere_vec(norm);
// Add accumulation of outgoing radiance.
accum_radiance += rem_radiance.cwiseProduct(emittance);
rem_radiance = rem_radiance.cwiseProduct(brdf);
// For specular, reflect perfectly.
// Ray spec_reflect_ray;
// vec3f spec_reflected_amt;
// spec_reflect_ray.pos = reflect_ray.pos;
// spec_reflect_ray.dir = ray.dir + (2 * cos_theta * norm);
// spec_reflected_amt = sample(scene, spec_reflect_ray, bounce + 1,
// max_bounces);
}
return accum_radiance;
}
// OPT: precompute / store ray directions for x, y.
void Renderer::work_render(Scene &scene) {
while (m_render_running) {
int pixel_id = m_pixel_queue.pop_front();
int x = pixel_id % m_render_opts.image_width;
int y = pixel_id / m_render_opts.image_height;
float u = m_frust_left + ((m_frust_right - m_frust_left)
* (x + 0.5) / m_render_opts.image_height);
float v = m_frust_bottom + ((m_frust_top - m_frust_bottom)
* (y + 0.5) / m_render_opts.image_width);
v = -v;
vec3f dir = (u * m_camera->m_right) + (v * m_camera->m_up) + m_camera->m_view;
Ray ray = { m_camera->m_pos, unit(dir) };
vec3f new_sample = sample(scene, ray, m_render_opts.num_bounces);
vec3f prev_sample = m_pixel_buf[pixel_id];
int sample_count = m_sample_counts[pixel_id];
m_pixel_buf[pixel_id] = (new_sample + (prev_sample * sample_count)) / (sample_count + 1);
m_sample_counts[pixel_id] += 1;
m_pixel_queue.push_back(pixel_id);
m_pixels_done++;
}
}
void Renderer::update_frustum_view() {
m_frust_top = tan(m_render_opts.fov / 2);
m_frust_bottom = -m_frust_top;
m_frust_left = -m_frust_top;
m_frust_right = m_frust_top;
}
<file_sep>#ifndef TRIANGLE_H
#define TRIANGLE_H
#include "model.hpp"
#include "ray.hpp"
#include "utils.hpp"
#include "vector.hpp"
// REF: move shape_data / index to shader
typedef struct {
vec3f verts[3];
vec3f norm;
Shape *shape_data;
size_t index;
vec3f& operator[] (const int index);
vec3f midpoint();
float intersect(const Ray &ray);
} Triangle;
typedef struct {
Triangle* tri = nullptr;
float dist = INF;
} TriangleHit;
#endif
<file_sep>#include "png++/png.hpp"
#include "image.hpp"
#include "vector.hpp"
void write_png(const char* filename, std::vector<vec3f> &pixel_data,
int img_width, int img_height)
{
png::image<png::rgb_pixel> image(img_width, img_height);
for (png::uint_32 y = 0; y < image.get_height(); ++y) {
for (png::uint_32 x = 0; x < image.get_width(); ++x) {
int index = (y * image.get_width() + x);
image.set_pixel(x, y, png::rgb_pixel(
(uint8_t) (255 * pixel_data[index].x()),
(uint8_t) (255 * pixel_data[index].y()),
(uint8_t) (255 * pixel_data[index].z())));
}
}
image.write(filename);
}
| 1f741ab0ca6e6a101ea6bf39db0ba5ee2b3f6035 | [
"Markdown",
"CMake",
"JavaScript",
"C++"
] | 26 | Markdown | CLaverdiere/pathtracer | 73d17fe3316068f467869355694afff8e5722c7c | 8c60b2a470b6126ed142265a66784e27c14056ac |
refs/heads/master | <file_sep>---
title: 大前端笔记之03 🔍 CSS 选择器
date: 2020-01-20 16:57:55
abbrlink: 1lc5rbtd
tags: CSS
categories: 大前端
excerpt: 要修改元素的样式,那么首先要将这些元素找出来。CSS 提供了丰富的选择器用来选择想要修饰的元素。
---
# 大前端笔记之03 🔍 CSS 选择器
要修改元素的样式,那么首先要将这些元素找出来。CSS 提供了丰富的选择器用来选择想要修饰的元素。
## 简单选择器
下面是一些最为基础的选择器。
### 元素选择器
元素选择器也称为类型选择器,它通过 HTML 元素的名称选择对应的元素。
```css
div {}
```
### 类选择器
类选择器由一个点`.`后跟元素的类名称构成,类名称对应的是 HTML 元素中`class`属性的值。多个元素可以拥有相同的类名称,而一个元素也可以拥有多个类名称,多个类名称之间在`class`属性中使用空格隔开。
如果类选择器的名称包含多个单词,应该使用短横线命名法。
```html
<button class="btn btn-danger"></button>
```
```css
.btn-danger {}
```
### ID 选择器
ID 选择器是由一个`#`符号后跟元素的 ID 构成的,ID 对应的是 HTML 元素中`id`属性的值。元素的`id`属性在当前文档中必须是唯一的,主要供 JavaScript 脚本使用。
如果 ID 选择器的名称包含多个单词,应该使用驼峰命名法。
```html
<div id="toolBar"></div>
```
```css
#toolBar {}
```
### 通配选择器
通配选择器用一个星号`*`表示,它可以用来选择页面上所有的元素。对于页面元素不多的页面,可以使用它来清除默认样式。但是,该选择器容易意外地将一些元素设置样式,并且在页面元素非常多的情况下可能会产生性能下降问题(虽然以目前的机器性能来说几乎可以忽略不计),所以最好还是不要滥用。
```css
* {
margin: 0;
padding: 0;
}
```
## 组选择器
有时单个选择器可能无法完成比较复杂的选择,因此它们也可以组合使用。
### 交集选择器
将多个选择器直接合并到一起(只要语法不产生歧义即可),可以选择同时符合这些选择器的元素。
```css
div.box {}
```
### 并集选择器
将多个选择器使用逗号`,`隔开,可以同时使用这些选择器选择不同的元素。为了方便阅读,逗号后最好换行再写下一个选择器。
```css
h1,
h2 {}
```
### 后代选择器
将多个选择器之间使用空格隔开,可以选择父元素中所有符合条件的后代元素。
```css
div span {} /* 选择 <div> 中所有的 <span> */
```
### 子选择器
将多个选择器之间使用`>`隔开,可以选择父元素中所有符合条件的子元素。
```css
div > span {} /* 选择 <div> 中的子元素 <span> */
```
### 相邻后续选择器
将多个选择器之间使用`+`隔开,可以选择该元素相邻的**下一个**满足条件的元素,它要求这两个元素必须是同级而且紧邻,中间不能间隔其它元素。
```html
<span>A</span>
<span class="B">B</span>
<span>C</span>
```
```css
.B + span {} /* 选择 C 元素 */
```
### 所有后续选择器
将多个选择器之间使用`~`隔开,可以选择该元素**之后所有**满足条件的元素,只要这些元素是同级即可,中间可以间隔其它元素。
```html
<span>A</span>
<span class="B">B</span>
<em>hi</em>
<span>C</span>
<span>C</span>
<span>C</span>
```
```css
.B ~ span {} /* 选择三个 C 元素 */
```
## 伪类选择器
伪类选择器可以选择文档结构以外的元素,它由一个冒号`:`后接关键字组成。注意,在使用伪选择器(包括下文的伪元素)时,**不要为了美观添加多余的空格**,这会导致触发后代选择器,从而无法得到想要的结果。
### :hover
选择鼠标当前悬停的元素。
```css
div:hover {}
```
### :focus
选择当前获得焦点的元素。
```css
input:focus {}
```
### :empty
选择没有子元素(包括普通文本、空格和换行,但注释除外)的元素,也包括单标签元素。
```css
p:emtpy {}
```
### :only-child
选择没有兄弟元素的元素。
在下面的代码中,由于`<span>`没有同级的元素,因此可以被选中。
```html
<p>
<span></span>
</p>
```
```css
span:only-child {}
```
### :only-of-type
选择没有**相同类型**兄弟元素的元素。
在下面的代码中,虽然`<span>`有兄弟元素,但并不是与之相同类型的`<span>`,因此可以被选中。
```html
<p>
<span></span>
<strong></strong>
</p>
```
```css
span:only-of-type {}
```
### :first-child / :last-child
选择一组兄弟元素中的第一个(或最后一个)。注意,该元素必须是同级元素中的第一个,否则无法被选中。
在下面的代码中,由于`.item`并不是同级的`<li>`中的第一个,所以没有元素被选中。
```html
<ul>
<li>1</li>
<li class="item">2</li>
<li class="item">3</li>
</ul>
```
```css
.item:first-child {}
```
### :first-of-type / :last-of-type
选择一组**同类型**兄弟元素中的第一个(或最后一个)。注意,该元素必须是同级元素中的第一个,否则无法被选中。
在下面的代码中,虽然`<span>`并不是同级元素中的第一个,但它是`<span>`类型中的第一个,所以可以被选中。
```html
<div>
<p>p</p>
<span>span</span>
</div>
```
```css
span:first-of-type {}
```
### :nth-child() / :nth-last-child()
选择一组兄弟元素中的第 n 个(或倒数第 n 个), n 从`1`开始,也就是说第 1 个元素索引为`1`,而不是`0`。注意,该元素必须是同级元素中的第 n 个,否则无法被选中。
其中括号的参数可以为:
| 参数 | 描述 |
| --- | --- |
| 关键字`odd` / `even` | 表示第奇数 / 偶数个元素 |
| 整数值`n` | 表示第`n`个元素,最小值为`1` |
| 表达式`An+B` | 表示第`An+B`个元素,`n`为从`0`开始的自然数,`A`和`B`必须为正整数 |
```css
:nth-child(odd) {} /* 选择第奇数个元素,如第 1、3、5、7... 个 */
:nth-child(even) {} /* 选择第偶数个元素,如第 2、4、6、8... 个 */
:nth-child(3) {} /* 选择第 3 个元素 */
:nth-child(3n+2) {} /* 选择第 3n+2 个元素,如第 2、5、8、11... 个 */
:nth-child(-n+5) /* 选择第 5、4、3、2、1 个元素,即前 5 个元素 */
:nth-child(n+3) /* 选择第 3、4、5、6... 个元素 */
:nth-child(-n+5):nth-child(n+3) /* 选择第 3、4、5 个元素 */
```
### :nth-of-type() / :nth-last-of-type()
选择一组**同类型**兄弟元素中的第 n 个(或倒数第 n 个),与上文用法一样,不再赘述。
### :not()
选择不符合指定选择器的元素。
```css
code:not(.hljs) /* 选择不包含 .hljs 类的所有 <code> 元素 */
```
## 伪元素选择器
伪元素由两个冒号`::`后接关键字组成。
### ::before / ::after
创建一个伪元素,将其作为**子元素**追加到匹配元素内部的最前面(或最后面),默认是**行内元素**。注意,必须使用`content`属性设置伪元素内容才可以使之生效,如果不需要内容,则设置为`content: ""`。
```css
div::after {
content: "";
}
```
注意,`<img>`和一些表单元素为空元素,不能有任何的子元素,因此它们也不能与`::before`或`::after`连用。
### ::selection
改变选中的元素样式。注意,该选择器仅适用于下面的特定样式属性,其它属性无效。
```css
a::selection {
color: green;
background-color: orange;
cursor: pointer;
}
```
## 属性选择器
根据元素的属性选择元素。属性值最好用双引号`""`包裹,并且区分大小写。它的用法包括:
| 选择器 | 描述 |
| --- | --- |
| `[attr]` | 选择所有包含`attr`属性的元素,无论它的值是什么 |
| `[attr="val"]` | 选择所有包含`attr`属性的元素,但是值必须完全是`val` |
| `[attr\|="val"]` | 选择所有包含`attr`属性的元素,但是值必须完全是`val`或者以`val-`开头(注意短横线) |
| `[attr^="val"]` | 选择所有包含`attr`属性的元素,但是值必须以`val`开头 |
| `[attr$="val"]` | 选择所有包含`attr`属性的元素,但是值必须以`val`结尾 |
| `[attr~="val"]` | 选择所有包含`attr`属性的元素,但是值必须包含`val`,且必须与其它值以**空格**隔开,不能连在一起 |
| `[attr*="val"]` | 选择所有包含`attr`属性的元素,但是值必须包含`val`作为子字符串(例如`cat`是`caterpillar`的子字符串) |
```html
<p>0</p>
<p data-fruit="">1</p>
<p data-fruit="apple">2</p>
<p data-fruit="banana apple">3</p>
<p data-fruit="apples">4</p>
<p data-fruit="apple-bad">5</p>
```
```css
[data-fruit] {} /* 选择除 0 号以外的元素 */
[data-fruit = "apple"] {} /* 选择 2 号元素 */
[data-fruit ~= "apple"] {} /* 选择 2、3 号元素 */
[data-fruit |= "apple"] {} /* 选择 2、5 号元素 */
[data-fruit ^= "apple"] {} /* 选择 2、4、5 号元素 */
[data-fruit $= "les"] {} /* 选择 4 号元素 */
[data-fruit *= "apple"] {} /* 选择 2、3、4、5 号元素 */
```<file_sep>---
title: 大前端笔记之21 🍀 Vue.js
date: 2020-02-15 21:15:10
abbrlink: ovrgsm0u
tags: Vue.js
categories: 大前端
excerpt: Vue.js 是目前国内最流行的前端渐进式框架,所谓渐进式指的是,你不需要一开始就学会 Vue 的全部功能特性,就可以将项目一点点引入 Vue。也就是说,它可以作为一个普通的 JavaScript 库用来提交表单、管理 DOM;也可以在大型项目中用来管理路由、实现组件化开发。随着项目的不断发展,以及对于 Vue 了解的不断加深,每个人所使用的侧重点也是不同的。
---
# 大前端笔记之 21 🍀 Vue.js
[Vue.js](https://cn.vuejs.org/v2/guide/installation.html)是目前国内最流行的前端渐进式框架,所谓渐进式指的是,你不需要一开始就学会 Vue 的全部功能特性,就可以将项目一点点引入 Vue。也就是说,它可以作为一个普通的 JavaScript 库用来提交表单、管理 DOM;也可以在大型项目中用来管理路由、实现组件化开发。随着项目的不断发展,以及对于 Vue 了解的不断加深,每个人所使用的侧重点也是不同的。
如果只是在浏览器端简单使用,则可通过 CDN 的方式在页面直接引入,后期则会采用其它的使用方式。
```html
<script src="https://cdn.jsdelivr.net/npm/vue/dist/vue.js"></script>
```
## 基本语法
引入 Vue.js 之后,首先通过`Vue()`实例化一个对象,它的参数也是一个对象,其中的属性值有很多,会在接下来逐一了解到:
```js
const vue = new Vue({
el: '#app', // 挂载数据的元素选择器,表示只有在这个元素下才能使用 Vue 的语法
data: {
// 数据对象,通常从服务器接口获取,这里临时模拟
msg: '你好呀~'
}
})
```
然后在页面上要准备一个对应的`#app`元素。其中可以使用双大括号(插值表达式)语法,将要渲染的数据名称写在里面,Vue 就会自动在实例中寻找相应的数据,将其替换成数据的值。当然,也可以在其中进行一些简单的表达式运算。
```html
<div id="app">
<span>{{ msg }}</span>
<span>{{ 'Hello,' + msg }}</span>
<!-- 拼接了字符串 -->
</div>
```
如果此时通过`vue.msg = '好个毛'`修改`msg`的值,那么会发现页面中的值也发生了变化,这是因为 Vue 默认会一直监听渲染后的数据,只要数据发生改变,那么相应的 DOM 也会被改变。
> 注意,所有的`data`数据可以直接使用实例本身调用,不需要`vue.data.msg`。
### 方法
方法`methods`可以像`data`一样保存数据,区别在于`data`存储的是值,而`methods`存储的是方法。
在下面的代码中,为元素[绑定了事件](#绑定事件),当触发事件时,会调用相应的`show()`方法。
```html
<button @click="show">点我</button>
```
```js
var vue = new Vue({
methods: {
show() {
console.log('hello')
}
}
})
```
如果需要在方法中访问`data`中的值,则需要使用`this`。这里的`this`指代的是实例化的`Vue`对象,可以通过`this.msg`获取到值,因为 Vue 自动在该对象下添加了属性,不需要调用`this.data.msg`。
```js
var vue = new Vue({
data: { msg: '你好呀~' },
methods: {
show() {
console.log(this.msg)
}
}
})
```
### 计算属性
计算属性用来抽取一些复杂的插值表达式计算,可以将这些计算定义 Vue 实例中统一管理,而不是直接写在模板上。
在下面的代码中,需要对书的总价进行计算,并显示在页面上,这时就可以使用计算属性。
```html
<div id="app">
<h1>{{ total }}</h1>
</div>
```
```js
var vue = new Vue({
el: '#app',
data: {
books: [
{ id: 1000, name: '哈利波特', price: 300 },
{ id: 1001, name: '北欧众神', price: 50 },
{ id: 1002, name: 'JOJO的奇妙冒险', price: 450 }
]
},
computed: {
total() {
return this.books.reduce(function(sum, book) {
return sum + book.price
}, 0)
}
}
})
```
可以看到,在使用计算属性时,不需要使用圆括号`()`,直接**当作属性使用**即可。原因是,其实计算属性的值本身也是一个对象,而其中包含两个函数属性,分别为该属性的`get()`和`set()`,因此计算属性的完整写法是:
```js
var vue = new Vue({
computed: {
total: {
get() {}
set() {} // 如果修改了 app.total,就会调用 set() 方法
}
}
})
```
由于`set()`方法实际使用很少,因此 Vue 才提供了这种简写方式,如果传入的不是对象,而是一个函数,那么就相当于调用它的`get()`方法。
### 方法与计算属性的区别
计算属性与`methods`非常相似,但是两者有一定的区别。其中最重要的一点是,计算属性是具有缓存机制的,而方法并没有。
在下面的代码中,分别使用计算属性和方法输出了内容,并对数据做了相同的处理,可以看到,计算属性只调用了一次,而方法调用了两次。其原因是,**只要数据没有发生改变**,那么计算属性就会直接读取缓存中计算好的值,不会多次进行计算;而方法是只要被调用,那么就会重新计算一次,因此如果只是希望获取一个计算后的值,那么应该使用计算属性,可以提高性能。
```html
<p>{{ getSumComputed }}</p>
<p>{{ getSumComputed }}</p>
<p>{{ getSumMethod() }}</p>
<p>{{ getSumMethod() }}</p>
```
```js
var vue = new Vue({
data: { num1: 233 },
computed: {
getSumComputed() {
console.log('计算属性执行了')
return this.num1 + 2
}
},
methods: {
getSumMethod() {
console.log('方法执行了')
return this.num1 + 2
}
}
})
```
### 指令
指令是 Vue 的一种特殊语法,有很多功能都需要指令来实现。它的本质是类似于`data-`的自定义属性,通常以`v-`开头,定义在元素标签上。当指令的值发生改变时,Vue 会自动操作相应的 DOM 元素,而不需要手动处理,即**以数据驱动 DOM**,这是 Vue 的重要理念。
指令的值与插值表达式一样,如果只写名称,没有引号,那么均会当作 Vue 数据处理。同样,也可以对于这些数据进行表达式运算。
### 过滤器
过滤器用于文本格式化显示,它只能使用在**双花括号的插值表达式**和`v-bind`指令中,分为全局和局部两种。
使用`Vue.filter()`方法可以定义一个全局过滤器,注意,全局过滤器只能定义在实例化`Vue()`之前。
| 参数 | 描述 |
| -------- | -------------------------------------------------------------------------- |
| 字符串 | 过滤器名称 |
| 处理函数 | 它的参数为要过滤的数据,即显示数据时,管道符前面的值;返回值为过滤后的数据 |
```js
// 将所有的 Fuck 替换成 F**k
Vue.filter('msgFilter', function(msg) {
return msg.replace(/Fuck/, 'F**k')
})
var vue = new Vue({}) // 然后才能实例化 Vue 对象
```
在`Vue()`实例对象内部也可以定义私有过滤器,表示只有该实例内部才能使用。如果与全局过滤器重名,那么优先调用私有过滤器。
```js
var vue = new Vue({
el: '#app', // 只有在 #app 下的元素才能调用该过滤器
filters: {
// 注意这里的属性名包含 s
msgFilter(msg) {
return msg.replace(/Fuck/, words)
}
}
})
```
在显示数据时,可以在原本数据最后添加一个管道符`|`,然后跟过滤器的名称:
```html
<p>{{ msg | msgFilter }}</p>
```
过滤器也可以传递参数,比如将要替换的固定值`F**k`改成通过参数传入:
```html
<p>{{ msg | msgFilter('F**k') }}</p>
```
```js
Vue.filter('msgFilter', function(msg, words) {
return msg.replace(/Fuck/, words)
})
```
过滤器也可以调用多次,Vue 将从左到右依次将前面的结果交给后面的过滤器。
```html
<p>{{ msg | msgFilter | otherFilter }}</p>
```
### 侦听器
使用`Vue()`实例中的`watch`属性可以定义侦听器,用于监听数据变化。一旦数据发生变化,则立刻触发对应的监听函数。监听函数中还可以传入两个参数,分别表示变化之后和变化之前的值。
```html
<input type="text" v-model="msg" />
```
```js
var vue = new Vue({
data: {
msg: ''
},
watch: {
msg(newVal, oldVal) {
// 当 msg 被修改时,会调用 msg() 侦听器
console.log('msg 发生了变化!')
}
}
})
```
### Vue 实例的生命周期
当使用`new Vue()`创建 Vue 实例的时候,这一条简单的语句会使得 Vue 内部作出非常复杂的操作。这些操作可以分为多个阶段,从而构成了它的生命周期。Vue 为这些阶段提供了对应的生命周期函数(或生命周期钩子),当执行到相应的阶段时,Vue 会自动的试图调用这些函数。
下图主要取自官网,它描述了 Vue 实例的整个生命周期。

这些生命周期函数均定义在实例对象的参数中,与`el`、`data`等属性并列。
```js
var vue = new Vue({
el: '#app',
data: {
msg: '你好呀~'
},
created() {
console.log(this.msg + 'Vue 被创建啦!数据可以使用了!')
}
})
```
## 显示数据
除了使用插值表达式以外,也可以使用一些特定的指令来显示数据。但是相比于插值表达式,指令方式并没有那么灵活,因此除非是特定的需求,否则尽量还是使用插值表达式。
### v-text
将指令的值作为数据名称,以纯文本形式,直接渲染到元素中。它的作用与插值表达式基本一致,但是如果需要元素中的部分内容,则必须使用插值表达式语法。
```html
<div v-text="msg"></div>
<!-- 等价于 -->
<div>{{ msg }}</div>
```
```js
var vue = new Vue({
data: {
msg: '你好呀~'
}
})
```
### v-html
将指令的值作为数据名称,以 HTML 形式,渲染到元素中。也就是说,如果数据内容包含 HTML,则会被正确解析。注意,如果数据是外部传入的,则可能会导致遭受恶意攻击。
```html
<div v-html="msg"></div>
```
```js
var vue = new Vue({
data: {
msg: '<h1>你好呀~</h1>' // 页面会出现一个 <h1> 标题
}
})
```
### v-once
如果不希望 Vue 监听数据的变化,从而改变 DOM,则可以使用`v-once`指令。使用该指令的元素只会被渲染一次,之后就会变成静态,从而提升页面性能。
```html
<div v-once>{{ msg }}</div>
```
```js
vue.msg = '好个毛' // 不会改变页面显示
```
### v-cloak
当页面使用插值表达式时,由于渲染需要时间,此时页面的双大括号可能会直接显示出来,然后才会被替换成真正的内容,从而导致数据出现闪烁。
如果为页面元素添加`v-cloak`指令,并使用 CSS 将其设置为隐藏,那么在内容渲染完成之前,该元素就不会显示。当 Vue 渲染完成后,会自动删除元素上的`v-cloak`样式,从而避免闪烁。
```html
<span v-cloak>{{ msg }}</span>
```
```css
[v-cloak] {
display: none;
}
```
## 绑定属性
使用`v-bind`指令可以绑定元素属性。
```html
<a v-bind:href="url">点我跳转~</a> <img v-bind:src="src + '.png'" />
```
```js
var vue = new Vue({
data: {
url: 'https://www.baidu.com',
src: 'test'
}
})
```
该指令提供了语法糖形式,可以将前面的`v-bind`省略,保留一个冒号`:`即可。
```html
<a :href="url">点我跳转~</a>
```
### class 样式属性
样式属性是一种比较特殊的元素属性,Vue 提供了专门的语法格式来绑定。如果元素同时拥有本身的类与绑定的类,那么 Vue 会将两者合并,而非简单覆盖。
如果传入一个字符串数组,那么表示该元素使用这些样式。其中的样式名必须以引号包裹,否则会被解析为数据名。
```css
.tianyi {
color: #66ccff;
}
.italic {
font-style: italic;
}
.thin {
font-weight: 200;
}
```
```html
<h1 :class="className">注意我要变形了!</h1>
```
```js
var vue = new Vue({
data: {
className: ['tianyi', 'thin', 'italic']
}
})
```
如果数组中的某些元素为对象,那么它表示该类是否起作用。其中对象的属性名为样式名,属性值为布尔值。
```js
var vue = new Vue({
data: {
// 对象的属性名可以省略引号哒!这是 JS 语法!和 Vue 没关系!
className: ['tianyi', 'thin', { italic: true }]
}
})
```
如果直接传入一个对象,则可以对其中的每个属性进行可用性判断。其中对象的属性名为样式名,属性值为布尔值。
```js
var vue = new Vue({
data: {
className: {
tianyi: true,
thin: true,
italic: false
}
}
})
```
### style 样式属性
除了类样式以外,也可以为元素绑定内联样式。注意,对于带有连字符`-`的属性,需要转换成驼峰形式,否则必须使用引号包裹(因为连字符不符合标识符规则)。
如果绑定的值为对象,那么每个键值对表示一条样式规则。其中属性名为样式名称,属性值为样式值。
```html
<h1 :style="s1">注意我要变形了!</h1>
```
```js
var vue = new Vue({
data: {
s1: { fontWeight: '200', color: '#66ccff' }
}
})
```
如果绑定的值为数组,那么可以同时绑定多个样式对象。
```html
<h1 :style="[s1, s2]">注意我要变形了!</h1>
```
```js
var vue = new Vue({
data: {
s1: { fontWeight: '200', color: '#66ccff' },
s2: { fontStyle: 'italic' }
}
})
```
## 绑定事件
使用`v-on`指令可以为元素绑定事件。注意,传入方法时加不加圆括号效果是一样的:
```html
<button v-on:click="show">点我</button>
<!-- 或者 -->
<button v-on:click="show()">点我</button>
```
```js
var vue = new Vue({
methods: {
show() {
alert('hello')
}
}
})
```
该指令同样提供了语法糖,可以将前面的`v-on:`替换成一个`@`。
```html
<button @click="show">点我</button>
```
绑定事件时也可以同时传入参数和事件对象。如果调用时**省略了圆括号**,那么 Vue 会默认传入一个事件对象参数:
```html
<a href="https://www.bilibili.com" @click="show">点我</a>
```
```js
var app = new Vue({
methods: {
// 可以使用原生的事件对象,这里只做演示,使用下面的事件修饰符实现更合理
show(e) {
e.preventDefault()
}
}
})
```
但是,**如果手动添加了圆括号,则必须显式传入一个`$event`对象才能获取到事件对象**,此时也可以传入其它参数作为方法本身的参数:
```html
<button @click="test1(10, $event)">点我</button>
<button @click="test2()">点我</button>
```
```js
var app = new Vue({
methods: {
test1(num, e) {
e // => 原生事件对象
num // => 10
},
test2(e) {
e // => undefined
}
}
})
```
### 事件修饰符
事件修饰符用来简化类似于阻止默认事件、事件冒泡等常见需求,它们均定义在事件名称后,由一个点`.`和修饰符名称组成。注意,事件修饰符是可以串联的,根据串联顺序不同结果也可能会有所不同。
| 事件修饰符 | 描述 |
| ---------- | ---------------------- |
| `.stop` | 阻止元素冒泡 |
| `.prevent` | 阻止元素的默认事件 |
| `.once` | 使绑定的事件仅触发一次 |
```html
<div @click="outer">
<div @click.stop="inner"></div>
</div>
<a @click.prevent="showMsg" href="https://www.baidu.com">点我跳转</a>
<button @click.once="showMsg">点我</button>
```
```js
var vue = new Vue({
methods: {
outer() {
console.log('外部元素事件被触发啦')
}, // => 不会被触发
inner() {
console.log('内部元素事件被触发啦')
}
}
})
```
### 按键修饰符
使用按键修饰符可以监听具体的按键。Vue 将[常用的按键](https://cn.vuejs.org/v2/guide/events.html#按键码)进行了封装,如果需要的按键不在列表中,可以使用`keyCode`码作为按键修饰符名称。
```html
<input type="text" @keypress.enter="show" />
<input type="text" @keypress.65="show" /><!-- 按下 a 触发 -->
```
```js
var vue = new Vue({
methods: {
show() {
alert('hello')
}
}
})
```
使用`Vue.config.keyCodes`属性可以为`keyCode`自定义一个名称。
```js
Vue.config.keyCodes.a = 65
```
```html
<input type="text" @keypress.a="show" />
```
## 条件渲染
使用`v-if`和`v-show`指令可以根据条件来判断某个元素是否被渲染。
两者的区别在于,`v-if`是通过创建或删除 DOM 元素来实现,而`v-show`则是通过`display`属性来实现。因此,如果是需要频繁切换显示隐藏的场景,应该使用`v-show`;如果一个元素经过判断之后自始至终就不需要显示,那么使用`v-if`即可,因为元素根本就不会创建,而不是先创建再隐藏。并且,`v-show`不能用在`<template>`中,也没有多分支判断。
### v-if
控制元素的显示或隐藏。如果指令的值为`true`,则元素正常显示,否则会被隐藏。
```html
<h1 v-if="flag">看不见我!</h1>
```
```js
var vue = new Vue({
data: {
flag: false // 如果为 true,则元素正常显示
}
})
```
与普通的流程控制一样,它也具有多分支的判断功能:
```html
<h1 v-if="age < 18">回家玩去!</h1>
<h1 v-else-if="age >= 18 && age <= 65">可以上网吧了!</h1>
<h1 v-else>退休以后还来网吧??</h1>
```
```js
var app = new Vue({
data: {
age: 18 // => 显示第二个 <h1>可以上网吧了!</h1>
}
})
```
> 对于本例来说,最好还是使用计算属性。但是如果要控制多个元素是否显示,可以将它们放到一个`<template>`模板中,然后为模板添加`v-if`指令,这样其中的元素就会根据条件判断隐藏或显示,但是`<template>`本身不会被渲染。`<template>`模板在后面的[组件](#组件)一节中会讨论到。
#### key 属性
如果将文本框用于判断时,可能会出现一个令人迷惑的现象:虽然文本框组件被切换了,但是用户输入的值却保留着:
```html
<input v-if="login == 'username'" type="text" placeholder="账户名" />
<input v-else-if="login == 'email'" type="text" placeholder="邮箱" />
```
```js
var vue = new Vue({
data: {
login: 'username' // 修改它会导致文本框发生改变,但是输入值却保留了
}
})
```
这是因为 Vue 为了最大化性能采用了虚拟 DOM,在替换元素时 Vue 会对比两个元素之间的差异,由于它们都是文本框元素,因此并不会真正的将前者删除,重新渲染 DOM,而是将它们的属性进行替换,就好像看起来是创建了一个新元素一样,所以之前的输入值才被保留了。
如果希望 Vue 将它们重新创建,可以添加一个`key`属性,如果两个元素的`key`属性不同,则 Vue 会认为它们是两个完全不同的元素。
```html
<input
v-if="login == 'username'"
type="text"
placeholder="账户名"
key="username"
/>
<input
v-else-if="login == 'email'"
type="text"
placeholder="邮箱"
key="email"
/>
```
### v-show
控制元素的显示或隐藏。如果指令的值为`true`,则元素正常显示,否则会被隐藏。
```html
<h1 v-show="flag">隐藏啦!</h1>
```
```js
var vue = new Vue({
data: {
flag: false
}
})
```
## 列表渲染
使用`v-for`可以遍历数组和对象,将其渲染到页面上。
### 遍历数组
在下面的代码中,每次从数组中取出一个数字,并渲染到页面上。
```html
<p v-for="num in nums">{{ num }}</p>
```
```js
const vue = new Vue({
data: {
nums: [3, 8, 12, 22, 28, 31]
}
})
```
除了数组元素本身外,也可以同时获取索引:
```html
<p v-for="(num, index) in nums">当前数字是{{ num }},这是第{{ index }}个数字</p>
```
> 注意,只有使用会修改原数组的方法操作数组时,才会导致数据被同步修改。
### 遍历对象
在下面的代码中,使用`v-for`遍历了`person`对象的每个属性值:
```html
<p v-for="val in person">{{ val }}</p>
```
```js
var vue = new Vue({
data: {
person: { name: '御坂美琴', age: 16 }
}
})
```
当然也可以同时获取对象的属性名,注意其中第一个参数为属性值,第二个参数为属性名:
```html
<p v-for="(val, key) in person">{{ key }} --- {{ val }}</p>
```
### 遍历整数
该指令还可以用来迭代整数。
```html
<p v-for="n in 10">{{ n }}</p>
```
### key 属性
与`v-if`中绑定的`key`属性的原因一样,`v-for`也必须使用`key`属性来标识自己的唯一性,否则当中间插入其它数据时,原本的输入值就会保留下来。
在下面的代码中,使用了`v-for`遍历输出了`persons`数组,并且每条数据前都有一个复选框。
```html
<div v-for="person in persons">
<input type="checkbox" />{{ person.id }} : {{ person.name }}
</div>
```
```js
var vue = new Vue({
data: {
persons: [
{ id: 1, name: '御坂美琴' },
{ id: 2, name: '温蒂' },
{ id: 3, name: 'JOJO' }
]
}
})
```
但是假如先勾选任意一条数据,再添加一条新数据到开头,就会发现复选框选中的数据发生了改变。这是因为 Vue 在虚拟 DOM 中比对两个元素时,发现都是相同的元素,因此为了提高性能,会在末尾创建一个新元素,而之前的元素就仅做属性覆盖。

要解决这个问题,需要为`v-for`绑定一个唯一的`key`属性,就可以使虚拟 DOM 找到对应的元素了。
```html
<div v-for="person in persons" :key="person.id"></div>
```
### 数据同步
由于 JavaScript 的限制,通过索引直接修改数组项、通过对象属性名直接修改值,或者修改数组长度均不会导致数据内容同步显示。
```html
<p v-for="num in nums">{{ num }}</p>
```
```js
var vue = new Vue({
data: {
nums: [10, 20, 30, 40]
}
})
// 输入 this.nums[0] = 233,可以看到数据被修改了,但是页面不会同步
```
为了解决这个问题,Vue 提供了一个内置的方法`Vue.set()`,用来修改数组内容。
| 参数 | 描述 |
| ------------- | ---------------------------- |
| 数组 / 对象 | 要修改的数组或对象名称 |
| 数值 / 字符串 | 要修改的索引位置或对象属性名 |
| 对象 | 修改后的值 |
```js
Vue.set(this.nums, 0, 233)
Vue.set(this.person, 'age', 16)
```
## 绑定表单元素
使用`v-model`指令可以将实例中的数据与视图中的数据进行**双向绑定**,它只能用于表单元素。之前所有的指令,只能实现由数据(Model,即`data`数据)到视图(View,即页面元素)的单向绑定,如果修改了数据,那么视图会随之变化,但是如果修改了页面元素内容,数据是不会发生改变的。而数据双向绑定,则是将两者关联起来,只要一方发生了变化,那么另一方也随之改变。
### 文本框 / 文本域
将文本框或文本域绑定为字符串数据,表示文本框的值与该字符串数据进行双向绑定。注意,**绑定之后会导致控件本身的`value`属性失效**。
```html
<input type="text" v-model="msg" /> <span>{{ msg }}</span>
```
```js
var vue = new Vue({
data: { msg: '你好呀' }
})
```
### 单选框
将单选框绑定为字符串数据,当它的`value`属性值等于该字符串时,那么该单选框会被选中,与此同时,其它绑定相同字符串的单选框则会被取消选中。
```html
<input type="radio" v-model="sex" value="male" />男
<input type="radio" v-model="sex" value="female" />女
```
```js
var vue = new Vue({
data: {
sex: 'male' // 与单选框的值同步改变,如果单选框选中了 女,那么该值也会改成 female
}
})
```
> 如果使用 Ajax 方式提交表单,那么`name`属性其实可以省略,因为`v-model`本身就可以实现互斥功能。当然原生提交还是必须写明`name`属性的,否则无法获取到属性名称。
### 复选框
复选框分为两种情况,首先,如果绑定一个布尔值,那么表示该复选框是否被选中(用于同意协议等场景):
```html
<input type="checkbox" v-model="isAgree" />我已阅读并同意以上扯淡条款
```
```js
var vue = new Vue({
data: {
isAgree: false
}
})
```
如果绑定一个数组,那么表示当数组中包含它们的`value`属性值时,这些复选框会被选中:
```html
<input type="checkbox" v-model="hobbies" value="sing" />唱
<input type="checkbox" v-model="hobbies" value="jump" />跳
<input type="checkbox" v-model="hobbies" value="rap" />Rap
<input type="checkbox" v-model="hobbies" value="basketball" />篮球
```
```js
var vue = new Vue({
data: {
hobbies: ['sing', 'jump']
}
})
```
### 下拉列表
如果绑定的是单个字符串数据,那么当`<option>`的`value`属性值等于该字符串时,该选项被选中。如果`value`属性不存在,则会以`<option>`中的内容作为备选项。
```html
<select v-model="province">
<option value="bj">北京</option>
<option value="sh">上海</option>
<option value="sd">山东</option>
</select>
```
```js
var vue = new Vue({
data: {
province: 'sd'
}
})
```
如果下拉列表允许多选,那么需要绑定一个数组。
```html
<select v-model="province" multiple>
<option value="bj">北京</option>
<option value="sh">上海</option>
<option value="sd">山东</option>
</select>
```
```js
var vue = new Vue({
data: {
province: ['sd', 'bj']
}
})
```
### 表单修饰符
表单修饰符可以对表单数据进行简单处理。
| 表单修饰符 | 描述 |
| ---------- | -------------------------------------------------------------------------------------------------------------------------------------------------------- |
| `.number` | 将数据转换成数值类型 |
| `.trim` | 过滤输入数据的首尾空格 |
| `.lazy` | 默认情况下,`v-model`绑定的元素只要发生`input`事件就会进行数据同步,使用该修饰符可以将其更改为`change`事件,也就是只有当表单元素失去焦点时,数据才会同步 |
```html
<input type="text" v-model.number="num" />
```
```js
var vue = new Vue({
data: { num: 0 }, // => 输入 30
computed: {
sum() {
return this.num + 10
} // => 如果不加修饰符,则结果是 3010,转换成数字后,则结果为 40
}
})
```
## 组件
Vue 可以将页面上的不同部分划分成一个个组件,每个组件实现一个单独的功能块,以方便维护和复用。
### 注册组件
使用`Vue.component()`方法可以注册一个全局组件,与[过滤器](#过滤器)一样,它依然要位于`Vue()`实例化之前。
| 参数 | 描述 |
| ------ | -------------- |
| 字符串 | 组件名称 |
| 对象 | 组件的配置参数 |
在组件对象中使用`template`属性可以为组件定义一个模板,也就是组件的内容。将组件名称作为 HTML 元素定义在页面上,那么 Vue 会使用组件的模板替换掉这个组件元素。
```html
<div id="app">
<my-com></my-com
><!-- <h1>这是一个标题组件啦</h1> -->
</div>
```
```js
Vue.component('my-com', {
template: '<h1>这是一个标题组件啦</h1>'
})
```
> 组件的名称如果包含多个单词,应该以**短横线**的方式命名和引用,因为 HTML 标签是忽略大小写的。如果标签名写作`<myCom>`,那么解析后的名称其实为`mycom`,而 JavaScript 又是一门大小写敏感的语言,它定义的组件名如果是`myCom`,那么很明显是不能对应`mycom`这个名称的。
组件也可以使用`components`属性定义在实例对象内部,作为**局部组件**使用。
```js
var vue = new Vue({
components: {
'my-com': {
// 组件名称,带有短横线的属性名必须加引号包裹
template: '<h1>这是一个标题组件啦</h1>'
}
}
})
```
注意,组件模板**只能有一个根元素**,因此下面的写法是错误的:
```js
Vue.component('my-com', {
template: '<h1>这是一个标题组件啦</h1> <h2>只能有一个根元素!</h2>'
})
```
将 HTML 写在 JavaScript 中会使得代码混乱不堪,因此可以使用`<template>`元素预先定义好模板,然后将`id`(必须是以`#`开头才会被识别为选择器,与`el`属性不同)传入`template`属性中:
```html
<div class="app">
<my-com></my-com>
</div>
<template id="tpl">
<h1>这是一个组件啦</h1>
</template>
```
```js
var vue = new Vue({
components: {
'my-com': {
template: '#tpl'
}
}
})
```
### 组件数据
组件与`Vue()`实例一样,也可以定义属于该组件的`data`数据。但是区别在于,**组件中的数据`data`不是一个对象,而是一个函数**,该函数的返回值才是数据对象。
```js
Vue.component('my-com', {
data() {
return { msg: '这是组件的数据啦' }
}
})
```
之所以这样设计,是为了保证组件在复用时,彼此之间的数据独立。如果`data`是一个对象,那么这些组件会通过引用传递,共享同一个对象。
在下面的代码中模拟了这种情况,组件的`data()`并不是直接返回一个对象,而是返回了一个外部的全局对象。此时所有的组件获得的都是该对象的**引用**,也就是说,它们指向同一块内存空间。因此一旦通过任何一个组件修改了数据,那么所有的组件数据都会被修改。
```html
<div id="app">
<my-com></my-com>
<my-com></my-com>
<my-com></my-com>
</div>
<template id="tpl">
<h1>{{ msg }}</h1>
</template>
```
```js
const obj = { msg: '这是组件的数据啦' }
Vue.component('my-com', {
template: '#tpl',
data() {
return obj
}
})
```
### 组件通信
组件是可以嵌套的,只需要将一个组件(如`child`)放到另一个组件(如`parent`)的`components`属性中,那么`parent`就是`child`的父组件。
> 可以看出,组件与之前一直使用`Vue()`实例非常相似,其实`Vue()`实例也可以看作一个组件,并且是根组件。
```js
var vue = new Vue({
components: {
// Vue 实例的 components 属性
parent: {
// 注册在 Vue 实例中的父组件
template: '<h1>这是父组件啦,里面包含<child></child></h1>',
components: {
// 父组件的 components 属性
child: {
// 注册在 parent 父组件中的子组件
template: '<h3>这是子组件啦</h3>'
}
}
}
}
})
```
注意,**子组件是不能直接使用父组件中的`data`数据的**,但是在开发中,通常都是由父组件统一请求数据(如果由子组件发送,那么发送次数也太频繁了),因此父子组件必须通过一定的方式来交换数据,这称为父子组件之间的通信。
#### 父组件向子组件传递数据
如果父组件需要向子组件传递数据,那么子组件首先要使用`props`属性定义用来保存父组件数据的变量名,该属性默认是一个字符串数组,每个元素为变量名。子组件中也只能使用`props`属性中的变量名,用来显示父组件数据。
```js
var vue = new Vue({
data: {
parentMsg: '父组件的数据'
},
components: {
child: {
template: '<h1>这是子组件啦{{ childMsgProp }}</h1>', // 只能使用 props 中定义的变量名
props: ['childMsgProp'] // 用来保存接收的父组件数据
}
}
})
```
然后,子组件需要绑定一个属性,其中属性名是`props`中用来接收数据的变量名,属性值是父组件的数据名,为要传递的数据。注意,**如果实例中`props`的变量名为驼峰形式,那么绑定元素的属性名必须要转换成连字符形式。**
```html
<div id="app">
<!-- 如果 props 名称是驼峰式,这里的属性名要转换成连字符形式 -->
<child :child-msg-prop="parentMsg"></child>
</div>
```
此外,`props`也可以是一个对象,它的属性名依然是接收数据的变量名称,但是它的值可以是数据类型(`String`、`Number`、`Boolean`、`Array`、`Object`、`Date`、`Function`、`Symbol`)或者另一个对象。如果是单独的数据类型,那么表示该变量必须接收该类型的数据,否则会报错;如果是另一个对象,那么还可以实现更复杂的验证操作:
```js
var vue = new Vue({
components: {
child: {
props: Array, // 类型限制
props: {
// 传入更复杂的验证对象
type: [Number, String], // 类型限制,满足其中之一即可
default: 233, // 默认值
required: true, // 必须通过元素属性传值
validator(value) {
//自定义验证函数
// 这个值必须匹配下列字符串中的一个
return (
['success', 'warning', 'danger'].indexOf(value) !== -1
)
}
}
}
}
})
```
对于对象和数组类型的默认值,其类型也必须是一个工厂函数,将默认值作为对象返回,原因与组件中的`data`必须是函数一样。
```js
var vue = new Vue({
components: {
child: {
props: {
default() {
// 默认是空数组,必须通过函数返回
return []
}
}
}
}
})
```
注意,虽然可以直接使用`props`中的数据,但是与`data`数据不同的是,**`props`的数据是不能修改的**。因此,在下面的代码中,虽然将`props`数据进行了双向绑定,但是程序会报出`Avoid mutating a prop directly...`的错误。
```html
<div id="app" class="container">
<child :child-msg="parentMsg"></child>
</div>
```
```js
const vue = new Vue({
data: { parentMsg: '父组件数据' },
components: {
child: {
// 绑定了 props 数据,程序会报错
template:
'<h1>这是子组件啦 -- {{ childMsg }} <input type="text" v-model="childMsg"></h1>',
props: ['childMsg']
}
}
})
```
根据错误信息提示,为了避免这个错误,应该将`props`数据保存到自己的`data`等属性中,然后绑定`data`中的数据就可以了。
```js
const vue = new Vue({
data: { parentMsg: '父组件数据' },
components: {
child: {
// 绑定了自己的 data 数据
template:
'<h1>这是子组件啦 -- {{ childOwnMsg }} <input type="text" v-model="childOwnMsg"></h1>',
props: ['childMsg'],
data() {
// 将 props 的数据传递给 data
return { childOwnMsg: this.childMsg }
}
}
}
})
```
#### 子组件向父组件传递数据
如果子组件需要向父组件传递数据,那么过程会比较麻烦,因为它们需要通过**子组件的事件绑定父组件的方法作为监听函数**来传递。具体来说,子组件需要设置一个自定义事件,将父组件的方法作为监听函数传入,然后子组件自己触发自定义事件,就相当于调用了父组件的方法。如果父组件方法中设置了参数,那么子组件在调用时就可以通过参数将数据传递过去了。
首先,为子组件绑定一个自定义事件`@child-event`,将父组件的方法`parentMethod()`传入。
> 由于 HTML 不区分大小写,因此自定义事件名称的多个单词建议以**连字符**分隔,切记不能使用驼峰命名法。
```html
<div id="app" class="container">
<child @child-event="parentMethod"></child>
</div>
```
```js
var vue = new Vue({
methods: {
// 父组件方法,用于传递给子组件调用
parentMethod() {
console.log('调用了父组件方法')
}
},
components: {
child: {
data() {
return { childMsg: '子组件的数据' } // 要传递的子组件数据
}
}
}
})
```
然后,需要在子组件中使用`$emit()`方法触发自己绑定的事件,该方法的第一个参数为事件名称,之后若干参数为监听函数中的参数。当然,为了调用`$emit()`,在子组件中再定义一个自己的方法。
```js
var vue = new Vue({
methods: {
parentMethod() {
console.log('调用了父组件方法')
}
},
components: {
child: {
template: '<h1>这是子组件啦{{ childMethod() }}</h1>', // 调用自己的 childMethod() 方法
data() {
return { childMsg: '子组件的数据' }
},
methods: {
childMethod() {
// 触发自己绑定的自定义 @child-event 事件,从而调用了监听函数,也就是父组件的方法 parentMethod()
this.$emit('child-event')
}
}
}
}
})
```
此时可以发现父组件方法`parentMethod()`已经被调用了,因此为了传递数据,只需要在`parentMethod()`中设置参数,在子组件调用时传递就可以了。
```js
var vue = new Vue({
data: {
parentMsg : ''
}
methods: {
parentMethod(msg) {
console.log('调用了父组件方法')
this.parentMsg = msg // 接收子组件调用时传递过来的数据
}
},
components: {
child: {
template: '<h1>这是子组件啦{{ childMethod() }}</h1>',
data() {
return { childMsg: '子组件的数据' }
},
methods: {
childMethod() {
// 调用父组件的方法 parentMethod() 时同时传递子组件数据
this.$emit('child-event', this.childMsg)
}
},
}
}
})
```
#### 父组件访问子组件
有些时候我们不需要组件的数据进行传递,而只是希望从父(子)组件中获取到子(父)组件中数据或方法直接使用。Vue 为此提供了专门的方式用来获取父子组件。
如果希望在父组件中访问子组件,总共有两种方式。第一种是直接在父组件中使用`$children`数组,该数组保存了它所有的子组件,使用索引取出后即可直接访问子组件中的数据。
> 注意,无论是`$children`,还是接下来的`$refs`和`$parent`,它们必须要等到模板渲染完成,也就是生命周期函数`mounted()`之后才能被正确获取,因为这些组件元素是通过 DOM 获取的。否则的话,这些属性的值会为空。
```html
<div id="app">
<child></child>
</div>
```
```js
const vue = new Vue({
el: '#app',
mounted() {
this.$children[0].childMethod() // => 子组件方法被调用了!
},
components: {
child: {
template: '<div></div>',
methods: {
childMethod() {
console.log('子组件方法被调用了!')
}
}
}
}
})
```
但是由于该方式通过索引来获取组件,如果组件顺序发生改变就会影响代码执行,因此通常都会使用下面的第二种方式。
第二种方式需要首先在子组件元素上添加`ref`属性,然后父组件就可以通过`$refs`对象获取到子组件了,该对象保存了所有定义了`ref`属性的元素。
```html
<div id="app">
<child ref="childRef"></child>
</div>
```
```js
const vue = new Vue({
el: '#app',
mounted() {
this.$refs.childRef.childMethod() // => 子组件方法被调用了!
},
components: {
child: {
template: '<div></div>',
methods: {
childMethod() {
console.log('子组件方法被调用了!')
}
}
}
}
})
```
### 插槽
虽然同一个组件可以多次复用,但是这些组件可能会有细小的差别,如果因为这些差别再去定义几乎一致的不同组件,那么就太浪费了。为了解决这一问题,Vue 提供了插槽的概念,可以将组件中有差别的部分定义成一个插槽`<slot>`,然后在调用组件时提供这些差异内容就可以了。
在下面的代码中,我们希望每个组件的最后一个元素有所不同,因此在注册组件时,将最后一个元素的位置设置为插槽。然后在调用组件时,传入想要的元素即可。
```html
<div id="app">
<com><button>将插槽替换成按钮元素</button></com>
<com><strong>将插槽替换成加粗元素</strong></com>
<com><span>将插槽替换成 span 元素</span></com>
</div>
<template id="comTpl">
<div>
<h2>这是一个组件啦</h2>
<slot></slot>
</div>
</template>
```
```js
const vue = new Vue({
components: {
com: {
template: '#comTpl'
}
}
})
```
在插槽中定义元素,可以为插槽设置一个默认值,如果调用组件时没有传入元素,那么会渲染为默认元素。
```html
<div id="app">
<com></com>
<!-- 最后一个元素为 <button>默认的按钮元素</button> -->
</div>
<template id="childTpl">
<div>
<h2>这是一个组件啦</h2>
<slot><button>默认的按钮元素</button></slot>
</div>
</template>
```
#### 具名插槽
一个组件中也可以定义多个插槽,在调用组件时为不同的插槽传入不同的元素。但是要实现这样的功能,需要为插槽起个名字,这样 Vue 才能知道哪些元素对应哪个插槽。这些拥有名字的插槽被称为具名插槽。
在下面的代码中,为导航栏组件定义了三个插槽,并且分别在`<slot>`上使用`name`属性为插槽定义名称:
```html
<template id="comTpl">
<div class="navbar">
<slot name="left"></slot>
<slot name="center"></slot>
<slot name="right"></slot>
</div>
</template>
```
当调用插槽时,需要在元素上使用`v-slot`指令冒号后面的值(该指令的属性值稍后再用)设置其对应的插槽。注意,该指令只能用于组件或`<template>`上。
```html
<div id="app">
<com>
<template v-slot:left><span>左侧内容</span></template>
<template v-slot:center><span>中间内容</span></template>
<template v-slot:right><span>右侧内容</span></template>
</com>
</div>
```
没有`name`名称的插槽,称为匿名插槽。如果调用组件时定义了没有`v-slot`指令的内容,那么就会将其对应为匿名插槽,如同本节开始时一样。
具名插槽同样有语法糖形式,只需将`v-slot:`替换成一个`#`即可:
```html
<div id="app">
<com>
<template #left><span>左侧内容</span></template>
</com>
</div>
```
#### 作用域插槽
在讨论作用域插槽之前,需要先明确一下 Vue 中作用域的问题。首先,在父组件中调用子组件时,子组件是没法获得自己数据的,即便是传入插槽的元素也不行。因为这整个位置都是父组件的作用域,而只有在子组件模板中才是子组件的作用域。
```html
<div id="app">
<com>
<!-- 这里只能获取到父组件的数据,因此 childMsg 为空 -->
<button>{{ childMsg }}</button>
</com>
</div>
<template id="comTpl">
<div>
<slot></slot>
<!-- 这里可以正常获取 childMsg -->
{{ childMsg }}
</div>
</template>
```
```js
const vue = new Vue({
components: {
com: {
template: '#comTpl',
data() {
return { childMsg: '子组件的数据' }
}
}
}
})
```
要解决这个问题,需要先在子组件模板中为插槽`<slot>`绑定一个属性,属性名可以自定义,属性值则是要传递的数据。由于这里是子组件的作用域,因此可以访问到`childMsg`数据。
```html
<template id="comTpl">
<div>
<!-- 子组件模板中可以访问子组件数据,属性名可以自定义 -->
<slot :msg="childMsg"></slot>
</div>
</template>
```
然后,在调用组件时为组件添加`v-slot`指令,它的属性值为一个对象(同样可以自定义名称),包含了所有插槽传递数据的名称(如上文中的`msg`)。通过该对象访问这些名称,就可以获取到子组件中的数据。
```html
<div id="app">
<com v-slot="slotProps">
<button>{{ slotProps.msg }}</button>
<!-- <button>子组件的数据</button> -->
</com>
</div>
```
如果匿名插槽与作用域插槽同时使用,还希望使用语法糖形式,那么必须要添加匿名插槽的名称`default`,不能直接省略。
```html
<div id="app">
<com #default="slotProps">
<button>{{ slotProps.msg }}</button>
</com>
<!-- 错误的写法 -->
<com #="slotProps">
<button>{{ slotProps.msg }}</button>
</com>
</div>
```
> 注意,`v-slot`指令为 2.6.0 版本新增,之前具名插槽`slot`和作用域插槽的`slot-scope`语法已经被废弃。
### 动态组件
如果组件名称是动态的,那么就不能使用组件名称作为标签了。为此,需要使用`<component>`定义一个动态组件,通过它的`is`属性可以使其变成对应名称的组件。这样只需将`is`属性绑定为一个变量数据,就可以切换不同的组件。
```html
<component :is="comName"></component>
```
```js
const vue = new Vue({
data: {
comName: 'login' // 修改 comName 就可以切换渲染的组件了
},
components: {
reg: { template: '<h1>注册组件</h1>' },
login: { template: '<h1>登录组件</h1>' }
}
})
```
不过注意,如果此时为组件添加生命周期函数`created()`和`destroy()`的话,会发现切换组件时它们会一直被创建和销毁。这无疑是比较影响性能的。如果希望它们可以被缓存下来,可以在组件外侧包裹一层`<keep-alive>`。
```html
<keep-alive>
<component :is="comName"></component>
</keep-alive>
```
### 单文件组件
之前我们均是使用`Vue.component()`定义全局组件,或者使用`components`属性定义局部组件,然后将组件相关的数据、模板写到对象属性中。但是随着项目变得复杂,这种方式定义的组件阅读起来会非常的麻烦。
因此,Vue 提供了一种单文件组件的方式来管理组件,可以将一个组件所有的模板、脚本和样式抽离到一个单独的`.vue`文件中管理。不过,此时必须要将组件作为一个模块导入才可以使用,为此,则必须使用[webpack](/posts/1zcig0yl.html)来提供模块化支持。
```html
<!-- app.vue -->
<template>
<div>{{ msg }}</div>
</template>
<script>
export default {
data() {
return { msg: '你好呀' }
}
}
</script>
<style>
div {
background-color: #66ccff;
}
</style>
```
虽然通常情况下我们都是使用[Vue-CLI](#Vue-CLI)来直接搭建带有 webpack 的 Vue 项目,但是也可以安装[Vue Loader](https://vue-loader.vuejs.org/zh/)来手动配置。
```powershell
npm install vue # 安装 vue
npm install webpack webpack-cli --save-dev # 安装 webpack
npm install -D vue-loader vue-template-compiler # 安装 vue-loader
```
> 如果希望组件的`<style>`生效,还必须安装[渲染样式](/posts/1zcig0yl.html#样式文件)的 loader。
```js
/* webpack.config.js */
const { VueLoaderPlugin } = require('vue-loader')
module.exports = {
module: {
rules: [
{
test: /\.vue$/,
loader: 'vue-loader'
},
{
test: /\.css$/,
use: ['style-loader', 'css-loader']
}
]
},
plugins: [
new VueLoaderPlugin() // 请确保引入这个插件!
]
}
```
这样就可以将`.vue`文件直接作为模块导入,并作为组件来使用了:
```js
import Vue from 'vue'
import app from './vue/app.vue' // 导入根组件模块
// CommonJS 导入方式,注意 Vue 对象保存在 default 属性中
// const Vue = require('vue').default
new Vue({
el: '#app',
template: '<app></app>', // 会覆盖 index.html 中的 <div id="app"></div> 作为根组件模板
components: { app } // 根组件
})
```
```html
<body>
<div id="app"></div>
<script src="./dist/main.js"></script>
</body>
```
与之前将根组件数据直接写在`Vue()`实例中不同的是,这里把根组件也作为一个子组件抽离到了外部。此时`index.html`页面上只有一个`#app`元素,而没有其它组件或数据。当 Vue 渲染页面时,会使用`template`属性中的`<app></app>`替换掉页面上的`#app`元素,又因为`<app>`本身是一个组件,因此会找到`app.vue`中的模板内容`<div>{ msg }</div>`来替换掉`<app></app>`。
注意,此时浏览器可能会报`You are using the runtime-only build of Vue...`这样的错误,这是因为 Vue 包含多个不同的版本,而默认的版本是不包含编译器的。因此如果`Vue()`实例中使用了模板,那么 Vue 就无法在运行时将其编译。要修改成完整版,需要在 webpack 中添加一项配置:
```js
module.exports = {
mode: 'development', // 否则 webpack 会隐藏错误信息
resolve: {
alias: {
vue$: 'vue/dist/vue.esm.js'
}
}
}
```
> 如果没有报错信息,但是也没有出现效果,则需要在`webpack.config.js`中配置`mode: 'development'`,否则 webpack 会将错误信息隐藏。
之所以出现这样的情况,是因为 Vue 将版本分为了两种:仅运行时版本(runtime-only)与运行时 + 编译器版本(runtime-only + compiler)。
要了解它们的区别,首先要简单了解 Vue 运行时的工作原理,注意接下来的步骤均是工作在**运行时**,也就是浏览器运行代码时,而非 webpack 编译时。当创建`Vue()`实例时,Vue 首先会将实例中`template`属性的值(也就是模板)进行编译成 AST(Abstract Syntax Tree,抽象语法树),这是一个编译原理的概念,不作展开讨论,有兴趣可以看看[这篇文章](https://segmentfault.com/a/1190000016231512)。
然后 Vue 会将 AST 传入一个渲染函数,通过渲染函数来创建虚拟 DOM。该函数可以通过实例中的`render()`的参数来手动调用,它接收诸如元素名称、元素属性、元素内容等参数,从而根据这些参数创建出虚拟 DOM。最后,Vue 才会将虚拟 DOM 转换成真实 DOM,并根据`el`属性找到页面上对应的元素,然后使用 DOM 的内容将其替换,从而呈现到页面上。
而这两个版本的区别,就在于运行时将模板编译为 AST 这一步,也就是说,runtime-only 版本没有这一步,而是直接从渲染函数开始的。这样做的原因是为了减少打包后的代码体积,但是缺少了编译模板的功能。
既然不使用模板,那么必须另寻他法来解决编译的问题。好在,渲染函数可以直接**传入一个组件**,Vue 可以通过组件生成虚拟 DOM,从而完成剩下的工作。因此,我们可以将之前创建实例的代码稍作修改(这也基本上是脚手架中自动生成的代码了):
```js
new Vue({
el: '#app', // 设置要替换的元素,脚手架中使用了 $mount('#app'),效果相同
// 直接将组件传入,渲染函数可以通过组件生成虚拟 DOM
render: function(h) {
// h 为渲染函数,也可以称为 createElement()
return h(app) // 返回值为虚拟 DOM
}
})
```
这时你可能有一个疑问,组件中不是也有`<template>`模板,那么 Vue 是怎么处理的呢?原因是,这些模板**早在 webpack 编译时就已经被处理成渲染函数了**,也就是说,它们**没有发生在运行时**。在刚才安装 vue-loader 时,我们还安装了另外一个 vue-template-compiler,它的作用就是在编译时将模板处理成渲染函数,这样就不需要运行时再次处理了。并且,可以看到参数中使用了`-D`即开发时依赖,也就证明了它们是没有工作在运行时的。
```powershell
npm install -D vue-loader vue-template-compiler # 安装 vue-loader
```
## Vue-CLI
[Vue-CLI](https://cli.vuejs.org/zh/guide/)(Vue Command-Line Interface,即 Vue 命令行工具,俗称脚手架)是一个用于快速搭建 Vue 项目开发环境的工具,使用它可以运行 Vue 的命令行指令,从而帮助我们快速生成目录结构、配置 webpack 等。
```powershell
npm install -g @vue/cli
```
然后,使用`vue create`命令可以创建一个项目:
```powershell
vue create hello-world
```
此时终端会提示`Please pick a preset`,即要求选择一个预设,选择`Manually select features`即手动配置后,按照顺序可以进行以下配置:
- 选择项目支持的特性,如 Babel、路由、vuex 等
- 对于一些特性是否生成独立的配置文件,还是保存到`package.json`中
- 是否将配置保存为一个预设,这样在创建项目时就会多出一个选项
选择完成后,便会生成一个如下的项目目录:
```powershell
hello-world
├─ .gitignore
├─ README.md
├─ babel.config.js # bebel 的配置文件
├─ node_modules
├─ package-lock.json
├─ package.json
├─ public # 静态资源目录,所有的内容会被原封不动打包到 dist 目录中
│ ├─ favicon.ico
│ └─ index.html # htmlWebpackPlugin 载入的模板,以此创建 index.html 文件
└─ src # 源代码目录
├─ App.vue # 根组件,可以自行修改
├─ assets # 资源文件,如图片、样式文件等
│ └─ logo.png
├─ components # 子组件目录
│ └─ HelloWorld.vue
└─ main.js # 项目入口
```
使用`vue ui`命令可以创建一个图形化界面的项目管理工具,如果希望修改配置、管理依赖等,也可以在这里更改。
## 前端路由
路由(route)在网站开发中指的是 URL 和页面之间的对应关系。在早期的网站开发中,服务器通常需要使用诸如 JSP、PHP 等技术,使用这些技术的页面除了 HTML 代码以外,还需要内嵌一些 Java 或 PHP 等后端代码。当用户请求一个 URL 之后,服务器会根据 URL 找到对应的页面,并通过后端代码查询数据库,然后将拼接好数据的页面返回给用户。可以看到,此时 URL 和页面之间的对应关系是由后端处理的,而这样的方式被称为**后端路由**。并且,由于返回给用户的是渲染之后的静态页面,因此对于 SEO 是比较友好的。但是,这种方式的缺点也很明显,因为页面中 HTML 和后端代码耦合在一起,使得页面维护变得非常麻烦,修改一个页面需要前端和后端人员一起配合才能完成。

后来,随着 Ajax 技术的普及,我们将之前的页面拆分成了两个部分:静态资源(不包含数据的 HTML、CSS、JS)和数据接口。当用户请求一个 URL 之后,依然是从服务器找到对应的 HTML 页面,但是此时的 HTML 页面只是一个骨架,并没有任何实际数据。但是,HTML 中会包含 JS 代码,用来向数据接口发送请求,然后再通过操作 DOM 的方式,将数据拼接到页面上,从而完成页面渲染。当然,为了方便操作 DOM,HTML 通常会包含模板引擎代码,这样页面只需要加载模板引擎,就可以快速地将模板引擎代码替换成获取的数据了。此时前后端责任更加清晰,开发、部署互不影响,而这样的开发模式称为「前后端分离」。

随之而来的下一个阶段则是**单页面应用**(SPA,Single Page Application),与之前的区别在于,它将多个 URL 对应的多套静态资源合并成了一个,也就是说,用户发送任何一个 URL 请求,返回的都只有一套 HTML + CSS + JS。然后,浏览器会监听 URL 变化,由 JavaScript 根据用户请求的 URL 不同,从这些资源中找到对应的部分(组件)进行渲染,但是不需要再向服务器请求静态资源了(接口数据还是得正常请求)。可见,此时的 URL 与页面(组件)的映射关系是由前端 JavaScript 维护的,因此这样的方式被称为**前端路由**。单页面应用的优点在于,它仅在用户首次访问时需要稍长的加载时间,但是之后切换页面不需要再次发送请求,这样增加了用户体验,也减轻了服务器压力。

此时,URL 只是一个页面的标识,并不需要真正发送请求。因此,我们必须通过一些方式使得 URL 发生改变,但是又不发送请求。
第一种方式是修改 URL 中的 hash 部分(`#`之后),然后通过监听`hashchange`事件来重新渲染页面。
第二种方式是通过 HTML5 新增的[History](/posts/gjqbkyw3.html#History)对象中的方法来修改 URL,这些方法也不会导致重新发送请求。但如果用户刷新了页面,那么浏览器还是会将其当作正常的 URL 向服务端发送请求。
而 Vue 通过[Vue-Router](https://router.vuejs.org/zh/installation.html)扩展提供了对前端路由的支持,它的实现原理也是上述两种方式。
### 安装与配置
安装 Vue-Router 可以使用浏览器端和模块两种方式:
```html
<script src="https://cdn.jsdelivr.net/npm/vue/dist/vue.js"></script>
<script src="https://unpkg.com/vue-router/dist/vue-router.js"></script>
```
```powershell
npm install vue-router
```
为了目录更加清晰,我们通常会在`src`目录中新建一个`router`文件夹,再创建一个`index.js`文件(导入`router`文件夹时会自动寻找该文件),将创建路由对象的部分放到里面,而入口文件只需要导入该模块即可。
接下来,需要实例化一个路由对象,并通过`Vue()`实例挂载。如果使用了模块引用,则还需要导入路由模块,并通过`Vue.use()`加载。
```js
/* index.js */
import Vue from 'vue'
import VueRouter from 'vue-router'
import Home from '../components/Home'
import About from '../components/About'
Vue.use(VueRouter) // 模块导入 VueRouter 的话必须加载插件
export default new VueRouter({
// 将模块对象导出,供 main.js 调用
routes: [
// 配置路由,即 URL 和组件的对应关系
]
})
/* main.js */
import router from './router' // 会自动寻找目录下的 index.js 文件
new Vue({
render: h => h(App),
router: router // 挂载路由模块
}).$mount('#app')
```
当引入路由模块后,会发现 URL 自动变成了`http://localhost/#/`的形式,最后一个`/`的后面也就是接下来要自己定义的 Hash 值。
### 配置路由规则
接下来,需要在`VueRouter()`实例中的`routes`属性中配置具体的路由规则。该属性是一个对象数组,每个对象表示一条匹配规则。对象的第一个属性`path`表示匹配的路径,第二个属性`component`对应的组件,它必须为一个**组件对象**。
```js
import VueRouter from 'vue-router'
import Home from '../components/Home'
import About from '../components/About'
export default new VueRouter({
routes: [
{ path: '/home', component: Home },
{ path: '/about', component: About }
]
})
```
最后在页面上使用`<router-view>`作为占位符,一旦找到匹配的路由,就会将该元素替换为对应的组件。这时将 URL 修改成`http://localhost/#/home`就可以跳转到首页组件了。当然让用户修改 URL 肯定是不现实的,为此 Vue 提供了`<router-link>`元素,将要跳转的路由添加到`to`属性中,Vue 会自动将该元素渲染为仅改变 Hash 值的`<a>`标签。
> 注意,`<router-view>`和`<router-link>`本身也是两个全局组件。
```html
<!-- App.vue -->
<template>
<div id="app">
<router-link to="home">首页</router-link>
<router-link to="about">关于</router-link>
<router-view></router-view>
</div>
</template>
```
如果希望`<router-link>`渲染为其它的元素,而非`<a>`元素,那么为其添加一个`tag`属性即可。注意,无论渲染为任何元素,它都是可以点击跳转的。
```html
<!-- 会渲染为一个 <button> 元素 -->
<router-link to="home" tag="button">首页</router-link>
```
### 重定向
将路由规则对象中的`component`替换为`redirect`,可以使该路由重定向到另外的路由:
```js
var myRouter = new VueRouter({
routes: [
// 当访问根路径时,重定向到首页
{ path: '/', redirect: '/home' },
{ path: '/home', component: Home },
{ path: '/about', component: About }
]
})
```
### History 模式
如果要使用 HTML5 中`History`方式来实现前端路由,以去掉 URL 中的`#`,只需设置一个`mode`属性即可:
```js
var myRouter = new VueRouter({
mode: 'history', // 设置为 History 实现前端路由
routes: []
})
```
### 取消历史记录
默认情况下,点击浏览器的后退按钮是可以返回上一条历史记录的,如果希望直接将当前记录替换,而非压入一条新记录,则可以为`<router-link>`添加一个`replace`属性。
```html
<router-link to="home" replace>首页</router-link>
<router-link to="about" replace>关于</router-link>
```
### 当前链接样式
对于当前选中的链接,Vue 会自动默认添加一个`.router-link-active`类,通过它可以设置当前选中链接的样式。如果要修改这个类名,在路由实例中设置`linkActiveClass`属性即可:
```js
var myRouter = new VueRouter({
routes: [],
linkActiveClass: 'active' // 修改默认的选中类
})
```
### 编程式导航
除了使用`<router-link>`创建标签来定义导航链接,也可以借助`Vue`实例内部提供的`$router`调用实例方法,通过编写代码来实现跳转。
在下面的代码中,通过两个按钮的点击事件来实现跳转。
```html
<button @click="goIndex">首页</button> <button @click="goAbout">关于</button>
```
```js
export default {
name: 'App',
methods: {
goIndex() {
this.$router.push('/home').catch(err => {}) // 跳转到 /home
this.$router.replace('/home').catch(err => {}) // 不添加新的历史记录,而是直接替换
this.$router.go(-1) // 跳转到某个历史记录,如果不存在则无效
},
goAbout() {
this.$router.push('/about').catch(err => {})
}
}
}
```
注意对于`push()`和`replace()`两个方法来说,如果当前已经是该组件,那么重复跳转会导致`NavigationDuplicated`错误。好在它们均返回`Promise`对象,使用`catch()`可以捕获异常(它们的第二个和第三个参数其实是跳转成功和失败的回调函数,可以使用`then()`和`catch()`替代)。
### 路由懒加载
随着业务代码越来越多,如果当用户第一次打开时就将所有页面对应的组件全部载入,可能速度会非常慢,影响用户体验。为此 Vue 提供了路由懒加载,它可以将原本打包成一个文件的业务代码分成多个文件,每个路由对应其中一个,当用户真正跳转到这个页面时,对应的组件文件才会被下载。
要实现路由懒加载,只需将原本组件对象的位置,替换成一个箭头函数即可:
```js
/* router/index.js */
const Home = () => import('../components/Home')
const About = () => import('../components/About')
export default new VueRouter({
routes: [
{ path: '/home', component: Home },
{ path: '/about', component: About }
]
})
```
### 嵌套路由
Vue 支持在路由下继续定义子路由。首先,需要在路由规则对象中添加`children`属性,以配置二级路由规则。注意子路由的`path`不要添加`/`,否则表示根路由,会导致路由拼接错误。
```js
/* router/index.js */
const Home = () => import('../components/Home')
const HomeNews = () => import('../components/HomeNews')
const HomeMessage = () => import('../components/HomeMessage')
// 注意两个子路由模块不要忘记定义,这里就不写了
export default new VueRouter({
routes: [
{
path: '/home',
component: Home,
children: [
// 子路由规则,为首页添加了两个路由,新闻和消息
{ path: '', redirect: 'news' }, // 设置重定向,默认显示新闻
{ path: 'news', component: HomeNews }, // path 不要加 /
{ path: 'message', component: HomeMessage }
]
}
]
})
```
然后,在父级路由的模板中设置`<router-link>`和`<router-view>`。注意`<router-link>`里的`to`属性要定义完整的路径,因为它无法识别当前是跟路由还是子路由。
```html
<!-- Home.vue -->
<template>
<div>
<h1>这里是 home 组件啦</h1>
<router-link to="/home/news">新闻</router-link>
<router-link to="/home/message">消息</router-link>
<router-view></router-view>
</div>
</template>
```
### 传递参数
前端路由在跳转到其它页面时也可以传递参数,主要有下面两种方式。
#### 路由参数
第一种方式称为路由参数,将路由规则中的`path`最后添加一个以冒号开头的自定义参数名,那么 Vue 会将这个部分识别为参数。
```js
/* router/index.js */
import User from '../components/User'
export default new VueRouter({
routes: [
// 冒号后面为自定义参数名
{ path: '/user/:userID', component: User }
]
})
```
此时任何以`/user/`开头的路由(如`/user/Leon`、`/user/Claire`)都会被渲染为`User`组件。
```html
<!-- App.vue -->
<router-link to="/user/zhangsan">用户</router-link>
```
如果参数为动态设置,那么需要将`to`绑定一个对象,其中`name`参数为**路径规则的名称**,`params`参数为一个对象,表示要传递的参数。
```html
<!-- App.vue -->
<router-link :to="{ name: 'User', params: { id: 666 }}">用户</router-link>
```
```js
/* /router/index.js */
export default new VueRouter({
routes: [
{ path: '/', redirect: '/home' },
{ path: '/home', component: Home },
{ path: '/about', component: About },
{ path: '/user/:id', component: User, name: 'User' } // 与 router-link 中的 name 对应
]
})
```
然后在组件中使用`$route.params.自定义参数名`来获取路由参数的值:
```html
<!-- User.vue -->
<template>
<div>
<h1>这里是用户组件啦</h1>
<!-- 与 path 中的 id 对应 -->
<h2>用户 ID 为:{{ $route.params.id }}</h2>
</div>
</template>
```
注意,`$route`是**当前活跃的路由对象**,也就是`routes`数组中的路由对象。根据调用它的组件不同,取得的当前路由对象自然也是不同的。
#### 地址栏传参
第二种方式是通过传统的地址栏传参,不过这里不需要手动在路由中拼接`?`部分,而是同样将`to`绑定为一个对象,其中`path`参数依然为之前的路径,`query`参数为一个对象,表示要传递的参数。
```html
<router-link :to="{ path: '/user', query: { id: 233 }}">用户</router-link>
```
然后在组件中使用`$route.query`对象来获取这些参数:
```html
<!-- User.vue -->
<template>
<div>
<h1>这里是用户组件啦</h1>
<h2>用户 ID 为:{{ $route.query.id }}</h2>
</div>
</template>
```
### 导航守卫
导航守卫可以监听并拦截页面跳转,以触发对应的回调函数,类似于中间件。使用`VueRouter()`实例对象的`beforeEach()`方法可以创建一个全局导航守卫,它接收一个函数参数,拥有三个参数分别表示目的路由对象、源路由对象以及用来放行的`next()`函数。
```js
/* /router/index.js */
const vueRouter = new VueRouter({
routes: [
{ path: '/home', component: Home },
{ path: '/about', component: About }
]
})
vueRouter.beforeEach((to, from, next) => {
next() // 拦截之后,必须手动调用 next() 以放行,否则无法正常跳转
})
```
在下面的代码中,通过在路由对象中定义元信息,可以实现跳转后修改当前页面的标题。
```js
const vueRouter = new VueRouter({
routes: [
{ path: '/home', component: Home, meta: { title: '首页' } },
{ path: '/about', component: About, meta: { title: '关于' } }
]
})
vueRouter.beforeEach((to, from, next) => {
document.title = to.meta.title // 修改页面标题
next()
})
```
但是,如果路由对象中包含子路由,那么需要使用`matched`属性获取到所有匹配的路由对象,通过索引`0`取出其中第一个,才能使父路由正确得到`meta`属性。
```js
vueRouter.beforeEach((to, from, next) => {
document.title = to.matched[0].meta.title
next()
})
```
## Vuex
[Vuex](https://vuex.vuejs.org/zh/)是 Vue 提供的一个集中管理组件状态(变量)的工具,通俗来讲,就是将所有组件都需要用到的数据放置到一起统一管理,比如用户登录状态、地理位置等等。
Vuex 可以使用 CDN 直接引入,也可以使用 npm 的方式导入模块。
```powershell
npm install vuex
```
如果采用模块方式,那么与 vue-router 一样,也需要使用`Vue.use()`加载。然后,创建一个`Vuex.Store()`实例对象,并在`Vue()`实例中挂载:
```js
/* 新建 store 文件夹用来保存 Vuex 相关文件,/store/index.js */
import Vuex from 'vuex'
Vue.use(Vuex) // 加载插件
export default new Vuex.Store({
// 相关数据和操作
})
```
```js
/* main.js */
import store from './store'
new Vue({
store,
render: h => h(App)
}).$mount('#app')
```
然后将公共数据放到`Vuex.Store()`的`state`属性中,就可以在任何一个组件通过`$store.state`对象获取了:
```js
/* /store/index.js */
export default new Vuex.Store({
state: {
msg: '这是公共状态啦'
}
})
```
```html
<!-- Home.vue -->
<h2>这是首页组件啦{{ $store.state.msg }}</h2>
```
### mutations
要修改`state`中保存的状态,不能在组件中直接使用`$store.state`来修改。虽然这样做确实可以修改成功,但是会导致 Vue 无法跟踪状态变化。
因此,官方建议在`Vuex.Store()`中的另一个属性`mutations`中定义修改的方法,使用它来修改`state`,而组件中只需要调用`mutations`中的方法即可。这些方法默认会传入一个`state`参数,通过它可以直接获取`state`中的属性。
```js
/* /store/index.js */
export default new Vuex.Store({
state: {
msg: '这是公共状态啦'
},
mutations: {
// 定义方法,供组件调用,并传入参数
change(state, param) {
state.msg = param // 通过 state 直接获取 msg,并进行修改
}
}
})
```
在组件中,则需要通过`$store.commit()`方法调用,其中第一个参数为`mutations`中定义的方法名,第二个参数为本身需要传递的参数,**如果需要传递多个参数,则可以通过对象来传递**,因此它也被称为载荷(Payload)。
```html
<!-- Home.vue -->
<h2>这是首页组件啦{{ $store.state.msg }}</h2>
<button @click="$store.commit('change', '状态被修改了!')">改变状态</button>
```
#### 类型常量
对于`mutations`中的方法名称,官方也称为「类型」。实际开发中,建议类型统一定义为常量,并保存到一个配置文件中,以方便管理。
```js
/* /store/mutations-types.js */
export const CHANGE = 'change'
```
```js
/* /store/index.js */
import { CHANGE } from './mutations-types.js' // 导入常量配置文件
export default new Vuex.Store({
state: { msg: '这是公共状态啦' },
mutations: {
// 以方括号的特殊语法来使用常量
[CHANGE](state, payload) {
state.msg = payload
}
}
})
```
```js
/* Home.vue */
import { CHANGE } from '../store/mutations-types.js'
export default {
methods: {
change() {
// 调用时同样使用常量
this.$store.commit(CHANGE, '新状态')
}
}
}
```
### getters
`getters`类似于之前的计算属性,通过函数的返回值对状态作出进一步处理。与`mutations`一样,它的第一个参数同样为`state`,而它的第二个参数为`getters`本身,可以用来获取其它`getters`。
在下面的代码中,使用`getters`过滤了价格较贵的书籍,并获取了这些书籍的数量,然后在组件中直接调用`getters`获取过滤后的结果:
```js
/* /store/index.js */
export default new Vuex.Store({
state: {
books: [
{ id: 1000, name: '北欧神话', price: 21 },
{ id: 1001, name: 'JOJO的奇妙冒险', price: 199 },
{ id: 1002, name: '从零开始的异世界生活', price: 233 }
]
},
getters: {
expensiveBooks(state) {
// 过滤较贵的书籍
return state.books.filter(el => el.price > 100)
},
expensiveBooksLength(state, getters) {
// 调用过滤后的 expensiveBooks 再获取数量
return getters.expensiveBooks.length
}
}
})
```
```html
<!-- Home.vue -->
<h2>{{ $store.getters.expensiveBooks }}</h2>
<h2>{{ $store.getters.expensiveBooksLength }}</h2>
```
但是与`mutations`不同的一点是,它不能接收参数。因此,如果希望数据是在调用时传入的,则需要将`getters`返回一个函数。
```js
/* /store/index.js */
export default new Vuex.Store({
getters: {
expensiveBooks(state) {
return function(price) {
return state.books.filter(el => el.price > price)
}
}
}
})
```
```html
<!-- Home.vue -->
<h2>{{ $store.getters.expensiveBooks(200) }}</h2>
```
<file_sep>---
title: 大前端笔记之04 📄 CSS 文本
date: 2020-01-20 17:29:32
abbrlink: ckgj1ldm
tags: CSS
categories: 大前端
excerpt: 页面中最常见的内容就是普通文本,CSS 中提供了大量属性对文本进行修饰。
---
# 大前端笔记之04 📄 CSS 文本
页面中最常见的内容就是普通文本,CSS 中提供了大量属性对文本进行修饰。
## 字体
使用`font-family`设置字体。注意,如果字体名称**包含空格或符号**,那么要使用双引号将它包裹起来。如果有多个备选字体,可以使用逗号`,`将它们连接在一起,这样如果前面的字体不可用,那么浏览器会自动使用下一个。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | 浏览器自定义 |
```css
body { font-family: consolas, "微软雅黑"; }
```
如果这些字体都不存在,那么浏览器会使用默认字体。因此最好在字体列表最后添加一个通用字体,这样可以保证至少有类似风格的字体替代。
| 通用字体 | 描述 |
| --- | ---|
| `Serif` | 该系列的字体包含衬线,每个字符的尺寸和宽度不同 |
| `Sans-serif` | 该系列的字体不包含衬线,每个字符的尺寸和宽度不同 |
| `Monospace` | 该系列的字体都是**等宽**的,每个字符在水平上均占据相等的空间,通常用来显示程序代码 |
由于一些系统不支持中文字体名称,因此除了使用对应的英文名称外,还可以使用 Unicode 字符集的方式设置字体,如:
```css
p { font-family: "\5FAE\8F6F\96C5\9ED1"; /* 微软雅黑 这四个字的 Unicode 码点 */ }
```
## 字重
使用`font-weight`设置字重,即文本的粗细。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | 分为`100`~`900`九个不同等级 |
| `normal` | 对应数字值`400` |
| `bold` | 对应数字值`700` |
数字值`100`到`900`并不是标明具体的字重等级,实际上几乎不可能有字体拥有如此多的变体。CSS 规范中仅要求,只要**数字更大的字重不小于前一个数字的字重**就可以。也就是说`200`字重可能比`100`字重更粗,但也有可能一样。
## 字号
使用`fong-size`设置字号,即[行内盒子内容区](/posts/08603cll.html)的高度。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | Chrome 有最小字号`12px`的限制,如果希望文字不显示,可以把字号设置为`0` |
| 百分比值 | 相对于父元素的`font-size`属性 |
| `normal` | 通常相当于`16px` |
## 斜体
使用`font-style`设置斜体。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| `normal` | 不使用斜体 |
| `italic` | 调用该字体的斜体变体,如果不存在强行渲染为倾斜文字 |
| `oblique` | 另一种斜体变体,但几乎没有字体有这种变体,因此没有使用的必要 |
## 行高
使用`line-height`设置行高,即[行内盒子](/posts/08603cll.html)的高度。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | 表示当前元素`font-size`属性的倍数 |
| 长度值 | 可用来单行文本垂直居中 |
| `normal` | 多数情况下相当于数字值`1.2`或`1.3` |
## 字体属性简写
使用`font`简写上述五个属性。它的最后两个值必须是`font-size`和`font-family`,顺序不能打乱,也不能省略。前面的`font-style`和`font-weight`可以互换位置和省略,`line-height`也可以省略。
```css
p {
font:
italic /* font-style */
bold /* font-weight */
26px/1.2 /* font-size/line-height */
"微软雅黑"; /* font-family */
}
```
## 水平对齐
使用`text-align`设置文本(准确说是行内元素)水平方向的对齐方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| **块级元素**,并仅对其中**行内元素**生效 | 可继承 | `left` |
| 可选值 | 描述 |
| --- | --- |
| `left` | 起边对齐 |
| `right` | 终边对齐 |
| `center` | 居中对齐 |
对于默认的书写模式(从左向右的语言)来说,起边指的是左边,终边是右边。如果设置了`writing-mode: vertical-lr`的话,那么`left`指的是顶端对齐,`right`为底端对齐。
## 垂直对齐
使用`verticle-align`设置[行内盒子](/posts/08603cll.html)在行框中的对齐方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| **行内元素** | 不可继承 | `baseline` |
| 可选值 | 描述 |
| --- | --- |
| `baseline` | 基线对齐 |
| `top` | 顶端对齐 |
| `center` | 居中对齐 |
| `bottom` | 底端对齐 |
| `middle` | 中线对齐 |
| 百分比值 | 相对于本元素的行内盒子高度,即`line-height`值 |
## 文字颜色
使用`color`设置文字的颜色。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | 浏览器自定义 |
| 可选值 | 描述 |
| --- | --- |
| 颜色值 | 任何合法的颜色值 |
## 装饰线
使用`text-decoration`设置文本的装饰线。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承(但实际是可以继承) | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 无装饰线,可以用来清除超链接自带的下划线 |
| `underline` | 下划线 |
| `overline` | 上划线 |
| `line-through` | 删除线 |
> 其实该属性还可以设置装饰线的颜色和样式,但是由于装饰线不能调整距离和粗细,因此并不常用。通常可以使用`border`来设置装饰线效果。由于行内元素不会因为`border`而改变高度,这样的方式也不会影响到布局和排版。
## 文本缩进
使用`text-indent`设置段落的首行缩进。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素 | 可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | 将段落设置为`text-indent: 2em`可以达到首行缩进两个汉字的效果 |
| 百分比值 | 相对于[包含块](/posts/vv37590w.html)的`width` |
## 空白处理
使用`white-space`设置空格、换行或者制表符的处理方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `normal` |
| 可选值 | 多余空格字符 | 换行符 | 自动换行 |
| --- | --- | --- | --- |
| `normal` | 合并 | 忽略 | 是 |
| `pre-line` | 合并 | 保留 | 是 |
| `nowrap` | 合并 | 忽略 | 否 |
| `pre` | 保留 | 保留 | 否 |
| `pre-wrap` | 保留 | 保留 | 是 |
## 单词换行处理
使用`word-break`设置文本在**自动换行**时,行尾单词的处理方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 行内元素 | 可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| `normal` | 文本在单词之间(以空格为界)发生换行,但是在中日韩等语言中,每个字符都会被当作一个单词,因此在任何两个字之间都可能换行 |
| `break-all` | 在任意位置换行,因此一个英语单词可能会被分成两行 |
| `keep-all` | 只有在空格出现时才会换行,因此在汉语中只要没有空格出现,那么也不会换行 |
## 文字阴影
使用`text-shadow`设置文字阴影。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 行内元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 没有文字阴影 |
| 长度值 | 两个值分别表示阴影水平和垂直方向的**偏移量**,正的数值表示阴影向右下方移动,负值表示阴影向左上方移动;如果有第三个长度值,则表示阴影的**模糊半径**,该值不能为负 |
| 颜色值 | 必须放在长度值最后,表示阴影的颜色,默认为黑色 |
```css
div { text-shadow: 1px 1px 2px #ccc; }
/* 用逗号分隔,可以同时添加多个阴影效果 */
div { text-shadow: 1px 1px 2px #ccc, -2px -2px 1px blue; }
```
## 文本溢出处理
使用`text-overflow`设置**单行的溢出文本**以`...`来显示,必须要配合`overflow: hidden`和`white-space: nowrap`来使用。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 行内元素 | 不可继承 | `clip` |
| 可选值 | 描述 |
| --- | --- |
| `clip` | 不作任何处理 |
| `ellipsis` | 以`...`显示溢出的文本 |
```css
span {
text-overflow: ellipsis;
overflow: hidden;
white-space: nowrap;
}
```<file_sep>---
title: 大前端支线笔记之01 🧱 CSS 包含块
date: 2020-01-22 6:22:00
abbrlink: vv37590w
tags: CSS
categories:
- 大前端
- 支线
excerpt: 包含块(containing block)是 CSS 布局中比较重要的一个概念,因为一个元素的布局或宽高均是由自身的包含块决定的。在不同的布局方式中,一个元素的包含块也会有所不同。
---
# 大前端支线笔记之01 🧱 CSS 包含块
包含块(containing block)是 CSS 布局中比较重要的一个概念,因为一个元素的布局或宽高均是由自身的包含块决定的。在不同的布局方式中,一个元素的包含块也会有所不同。
## 普通文档流
在普通文档流中,一个元素的包含块是其**最近父元素的内容区**。此时,子元素在**水平方向**上的`margin`、`border`、`padding`和`width`之和必须与包含块的宽度相等。
由于其中的`border`和`padding`只能是固定值,而`margin`和`width`能够设置为`auto`,表示根据情况来调整自身大小,以确保这个等式被满足。
注意,只要`margin`与任何其它属性同时为`auto`,那么`margin`的`auto`会被视为`0`。
首先,当只有一个值为`auto`时,该值会自动调整宽度,以满足等式。
```html
<div>
<p></p>
</div>
```
```css
div { width: 500px; }
p {
margin-left: auto; /* auto 会被计算为 300px */
margin-right: 100px;
width: 100px;
}
```
当两个`margin`的值为`auto`时,那么它们会**等分剩余的距离**以满足等式,从而使该元素在包含块中居中,这是**块级元素水平居中方式**之一。
```css
div { width: 500px; }
p {
margin-left: auto;
margin-right: auto; /* 左右外边距均是( 500-300 )/ 2 = 100px */
width: 300px;
}
```
当三个属性都为固定值,而且加起来也不够包含块的宽度,此时浏览器会**强制将`margin-right`设置为`auto`**,以满足等式。
```css
div { width: 500px; }
p {
margin-left: 100px;
margin-right: 100px; /* 右外边距会被强行设置为 auto,从而被计算为 300px */
width: 100px;
}
```
如果三个固定值加起来超过了包含块宽度,那么子元素就会超出包含块,导致一部分位于包含块外部。
## 定位元素
定位元素的包含块与普通文档流有所不同:
* 绝对定位:为最近的**定位**父元素,直到最外层的`<html>`元素
* 如果父元素是块级盒子,那么包含块是父元素的**内边距**
* 如果父元素是行内盒子,那么包含块是父元素的**内容区**
* 相对定位:为最近的父元素内容区
* 固定定位:为浏览器窗口
与普通文档流一样,定位元素也要满足包含块的等式。但是,普通文档流的等式仅针对水平方向,而定位元素的公式也同时适用于**垂直方向**。
以水平方向为例,定位元素水平方向上的`margin`、`border`、`padding`、`width`、`left`和`right`之和必须与包含块宽度相等。
其中`border`和`Padding`的值只能是固定值,而`margin`、`width`、`left`和`right`都能够设置为`auto`从而根据情况来调整自身大小,以确保这个等式被满足。注意,只要`margin`和任何一个属性都为`auto`,那么`margin`的`auto`会被视为`0`。
当 5 个值(`margin-left`、`margin-right`、`width`、`left`、`right`)中只有一个为`auto`,该值会自动调整宽度,以满足等式。
```css
div {
position: relative;
width: 500px;
}
p {
position: absolute;
left: 0;
right: 0;
margin-left: auto; /* auto 会被计算为 300px */
margin-right: 100px;
width: 100px;
}
```
当 2 个`margin`的值为`auto`,那么它们会**等分剩余的距离**以满足等式,如果要居中元素,最好将对立的偏移量设置为**同一个值**(比如`0`).
```css
p {
position: absolute;
left: 0;
right: 0;
margin-left: auto; /* ( 500-100 ) / 2 = 200px */
margin-right: auto; /* 同上 */
width: 100px;
}
```
当 2 个偏移量为`auto`,那么元素位于**定位之前的位置**。相当于仅仅为一个元素设置了绝对定位,而没有设置偏移量。
当`width`和 1 个偏移量为`auto`,那么`width`会收缩到最小,由偏移量满足公式。
当`width`和 2 个偏移量为`auto`,那么`width`会收缩到最小,元素位于定位之前的位置。
当 5 个属性都为固定值,而且加起来也不够包含块的宽度,`right`会被视为`auto`。<file_sep>---
title: 大前端笔记之18 🎉 JavaScript ES6
date: 2020-02-06 10:24:12
abbrlink: j5h1kgw7
tags: JavaScript
categories: 大前端
excerpt: ECMAScript 6(简称 ES6)是 2015 年发布的最新版本,其中添加了大量的新特性。虽然之后 ECMAScript 版本依然在不断更新,但是新特性并不是很多,因此将 ES6 及之后的版本统称为 ES6。
---
# 大前端笔记之18 🎉 JavaScript ES6
ECMAScript 6(简称 ES6)是 2015 年发布的最新版本,其中添加了大量的新特性。虽然之后 ECMAScript 版本依然在不断更新,但是新特性并不是很多,因此将 ES6 及之后的版本统称为 ES6。
## 变量
新增了用于定义变量的`let`关键字。
```js
let num = 10
```
与之前的`var`相比,它主要有以下几个特点。首先,使用`let`关键字声明的变量具有**块级作用域**。也就是说,使用`let`在花括号内部声明的变量不能在花括号外部访问了,因为花括号拥有了自己的作用域。
```js
if (true) {
var a = 10
let b = 10
}
console.log(a) // => 10
console.log(b) // => b is not defined
```
第二,使用`let`定义的变量声明**不会被提升**。
```js
console.log(a) // => undefined
var a = 2
console.log(b) // => Cannot access 'b' before initialization
let b = 2
```
第三,虽然使用`let`定义的变量没有提升的特性,但是它会与作用域绑定,产生**暂时性死区**。
在下面的代码中,虽然在块级作用域内先调用后定义了`num`,但是在调用时不会去访问全局作用域下的`num`,而是直接报错。这是因为虽然 JavaScript 不会把`let`变量提升,但是依然知道该作用域内有同名变量,而且将其与当前作用域进行了绑定,如果提前调用,就会直接报错。在`let`声明之前无法调用该变量的这段运行过程,称为暂时性死区。
```js
var num = 10
if (true) {
console.log(num) // Cannot access 'num' before initialization
let num = 20
}
```
最后,使用`let`定义的全局变量不能使用`window`访问,因为它被存放到了单独的作用域中。
```js
let a = 10
window.a // => undefined
a // => 10
```
### let 与 for 循环
使用`let`声明`for`循环的迭代变量是非常合适的,因为循环结束后迭代变量就会被销毁,从而释放空间。
```js
for (var i = 0; i < 10; i++) {}
console.log(i) // => 10
for (let i = 0; i < 10; i++) {}
console.log(i) // => i is not defined
```
但是,如果`for`循环中包含异步任务,那么可能会产生令人迷惑的现象。在下面的代码中,分别使用`let`和`var`声明的`for`循环为 3 个按钮绑定了事件,当点击时输出绑定时的`i`,而结果是不同的:
```js
for (var i = 0; i < btns.length; i++) {
btns[i].addEventListener('click', function () {
console.log(i)
})
} // => 3 3 3
for (let i = 0; i < btns.length; i++) { // 3 个按钮
btns[i].addEventListener('click', function () {
console.log(i)
})
} // => 0 1 2
```
[异步任务](#异步编程)是当整段代码执行完成后,才会根据情况(计时器、触发事件)触发的,这里得到的`i`应该是`for`循环执行完毕之后的。可以看到,`var`声明的`for`循环符合预期,当循环结束后,`i`的值会自增为`3`,因此点击任何一个按钮都是输出`3`。
但是,`let`循环的输出结果就有些令人迷惑了,虽然它可以使自己拥有作用域,但是`i`也依然只有一个,为何最后会输出不同的值呢?
原因是,`let`循环在执行过程中,不但生成了自己的作用域,而且会记录本次循环的值。因为`let`可以识别块级作用域(花括号),因此不同块里的同名变量不会发生冲突。也就是说,上面`let`循环的代码可以表示为:
```js
{
let i = 0 // 初始化循环变量,即圆括号中的 i
{
let t = i // 将 i 的值保存下来
btns[t].addEventListener('click', function () { // 实际使用的不是最后的 i,而是本作用域内保存的值
console.log(t)
})
}
i++ // 循环变量自增
// 第二遍循环
{
let t = i
btns[t].addEventListener('click', function () {
console.log(t)
})
}
i++
// 第三遍循环
{
let t = i
btns[t].addEventListener('click', function () {
console.log(t)
})
}
i++
}
```
可以看到,此时执行事件处理函数时,寻找的其实是本作用域中保存的变量,而非外层迭代完毕的`i`。此外,还可以看到,**圆括号与循环体其实分属于两个不同的作用域**,而且,**圆括号作用域是包裹着循环体作用域的**。
而对于`var`循环就简单的多了,因为无论是哪一次循环,最后只能访问全局下共同的`i`:
```js
var i = 0 // 初始化循环变量,即圆括号中的 i
btns[i].addEventListener('click', function () { // 使用的依然是全局作用域下的 i
console.log(i)
})
i++ // 循环变量自增
// 第二遍循环
btns[i].addEventListener('click', function () {
console.log(i)
})
i++
// 第三遍循环
btns[i].addEventListener('click', function () {
console.log(i)
})
i++
```
在下面的代码中,分别在圆括号和循环体中声明了重名的变量`i`,可见它们并不冲突:
```js
for (let i = 0; i < 3; console.log(i++)) {
let i = 'abcd'
console.log(i)
} // => abcd 0 abcd 1 abcd 2
```
其中`abcd`是循环体内的`i`,数字值是圆括号内的`i`,根据上述理解,这段代码可以表示为:
```js
{
let i = 0 // 初始化循环变量,即圆括号中的 i
{
let i = 'abcd' // 循环体内的 i,由于 let 变量可以识别花括号,因此不会与外部冲突
btns[i].addEventListener('click', function () { // 实际使用的是最近的,也就是说本作用域内的 i
console.log(i) // => 'abcd'
})
}
console.log(i++) // 输出外层作用域下的 i,也就是 0,然后自增为 1
// 第二遍循环
{
let i = 'abcd'
btns[i].addEventListener('click', function () {
console.log(i) // => 'abcd'
})
}
console.log(i++) // => 1
// 第三遍循环
{
let i = 'abcd'
btns[i].addEventListener('click', function () {
console.log(i) // => 'abcd'
})
}
console.log(i++) // => 2
}
```
## 常量
新增了用于定义常量的`const`关键字。
```js
const PI = 3.14
```
除了以上`let`关键字的**所有特性之外**,它还具有下面单独的特点。首先,常量在声明的同时**必须赋初始值**,否则会报错。
```js
const PI // Missing initializer in const declaration,声明常量没有初始化
```
然后,常量的最明显特性就是值一旦确定就**不可更改**。注意,对于复杂数据类型来说,这里的值指的是**地址**而非具体的内容。
```js
const PI = 3.14
PI = 100 // Assignment to constant variable,基本数据类型不能修改值
const ARR = [10, 20]
ARR[0] = 5 // => [5, 20],可以修改具体内容
ARR = [5, 20] // Assignment to constant variable,不能直接修改指向的地址
```
## 对象的简洁表示法
如果对象中的属性名和属性值名称相同,那么可以合并成一个。
```js
let age = 18
let person = { age: age } // 第一个 age 是属性名,第二个 age 是变量 18
/* ===== 等同于 =====*/
let age = 18
let person = { age }
```
对象中的函数也可以简写,如:
```js
let person = {
eat: function() { console.log('吃呀吃呀吃~') }
}
/* ===== 等同于 =====*/
let person = {
eat() { console.log('吃呀吃呀吃~') }
}
```
## for of 循环
使用`for of`循环可以更方便地遍历数组。
```js
const nums = [10, 20, 30]
for (let num of nums) {
num
} // => 10 20 30
```
> 注意与`for in`循环区分开,`for in`循环的迭代变量为键,且属于 ES5 规范;而`for of`循环的迭代变量为值,属于 ES6 规范。因此前者主要由于遍历对象,后者主要用于遍历数组。
## 解构赋值
解构赋值是一种为变量赋值的新方式,可以将数组或对象中的元素直接提取出来,放到对应的变量中。
### 数组解构
将变量名称使用方括号`[]`包裹起来,表示数组解构赋值,它的值必须为一个数组。
```js
let [a, b] = [1, 2]
a // => 1
b // => 2
```
如果两侧的值不完全对应,则未赋值的变量为`undefined`,多余的值被忽略。
```js
let [a, b, c] = [1, 2]
c // => undefined
let [a] = [1, 2]
a // => 1
```
### 对象解构
将变量名称使用花括号`{}`包裹起来,表示对象解构赋值,它的值必须为一个对象。如果在对象中找到了与解构语法中的变量名称一致的属性,则将属性的值赋值给该变量。因此,**解构语法中的变量顺序与对象中的属性顺序无关**。
```js
let {name, age} = {name: 'Wendy', age: 13}
name // => Wendy
age // => 13
```
如果需要修改变量名称,则可以使用下面的解构语法:
```js
let {name: myName, age: myAge} = {name: 'Wendy', age: 13}
myName // => Wendy
```
## 箭头函数
箭头函数是函数表达式的一种简化写法,但是由于它的一些特性(主要是下文中箭头函数的`this`),并不是可以无脑的将所有函数表达式替换成箭头函数。
箭头函数的格式为`() => {}`,其中圆括号为形参列表,花括号为函数体。由于箭头函数没有名称,因此可以将它放到一个变量中调用:
```js
let fn = function() {
console.log('hello')
}
// 相当于
let fn = () => { console.log('hello') }
fn() // => 输出 hello
```
前面的圆括号中可以传入形参:
```js
let fn = (num1, num2) => { return num1 + num2 }
fn(10, 20) // => 30
```
如果参数只有一个,那么可以省略圆括号:
```js
let fn = num1 => { return num1 }
fn(10) // => 10
```
如果函数体中**只有一条语句**,那么可以省略花括号和`return`关键字。此时函数会**自动将这一行代码的执行结果作为返回值,无论这行代码是否有返回结果**。因此如果省略了花括号,那么就不能再添加`return`关键字了。
```js
let fn = (num1, num2) => { return num1 + num2 }
// 相当于
let fn = (num1, num2) => num1 + num2
fn(10, 20) // => 30
// 即使这条语句没有返回值,也会自动返回
let fn = () => console.log('hello')
fn() // => 输出 hello,但是返回值为 undefined
```
### 箭头函数中的 this
箭头函数与其它函数最大的一个区别在于,它并没有自己的`this`,而是直接使用外层作用域下的`this`,也就是说箭头函数中的`this`仅与定义位置有关,与何时调用无关。
在下面的代码中,由于`addEventListener()`定义在全局作用域中,因此其中箭头函数的`this`指代的是`window`,而不是按钮元素:
```js
button.addEventListener('click', function() {
console.log(this) // => <button>...</button>
})
button.addEventListener('click', () => {
console.log(this) // => window
})
this // => 与外层作用域中的该 this 相同
```
由于`this`的指向问题,因此如果函数体中包含`this`的话,切记依然要使用普通函数。另外,如果函数包含多条语句,而不是单纯的通过计算返回一个值,也建议使用普通函数,因为这样代码会更加清晰易读。
## 剩余参数
由于箭头函数中并不能使用`arguments`属性,因此 ES6 提供了剩余参数来解决不定参数的问题(函数声明和表达式也可以使用)。在参数列表中定义一个`...参数名`表示剩余参数,为包含所有剩余参数的数组。
在下面的代码中,将所有传入的参数作为剩余参数:
```js
function fn(...args) {
console.log(args)
}
fn(10, 20) // => [10, 20]
fn(10, 20, 30) // => [10, 20, 30]
```
之所以称之为剩余参数,是因为它除了可以接收所有参数外,也可以接收部分参数:
```js
function fn(p1, p2, ...args) {
console.log(args)
}
fn(10, 20, 30) // => [30]
fn(10, 20, 30, 40) // => [30, 40]
```
## 扩展运算符
扩展运算符`...`可以将数组(或者任何实现了**遍历器**的伪数组)拆分成以逗号分隔的**参数序列**。
在下面的代码中,`log()`的输出结果是`1 2 3`,而不是`1,2,3`。这是因为传入`log()`方法的其实是三个参数,相当于`console.log(1, 2, 3)`。由于`log()`方法可以传入多个参数,并以空格分隔后输出,才出现了这样的结果。
```js
let arr = [1, 2, 3]
console.log(...arr) // => 1 2 3
```
因此,可以用它将数组直接拆分为若干参数传递:
```js
let arr = [10, 20]
let fn = (x, y) => x + y
fn(...arr) // => 30
```
或者用来合并数组:
```js
let arr1 = [10, 20]
let arr2 = [30, 40]
// ...arr1 相当于 10, 20 ...arr2 相当于 30, 40
// 将它们放到括号中再加上中间的逗号,就实现了合并
let result = [...arr1, ...arr2] // => [10, 20, 30, 40]
```
由于 DOM 集合中的`HTMLCollection`也实现了遍历器,因此可以使用扩展运算符将其转换成真正的数组:
```js
let divs = document.querySelectorAll('div')
let result = [...divs] // => [div, div, div],真正的 div 元素数组
```
> 转换真正数组也可以使用`Array`对象的`from()`方法。
## 模板字符串
模板字符串是新增的字符串定义方式,允许在字符串中直接使用变量,这些变量会自动替换成相应的值,而不需要通过`+`拼接字符串。模板字符串使用反引号包裹,里面的变量使用`${}`包裹。
```js
let age = 18
let str = `今年 ${age} 岁` // => 今年 18 岁
```
模板字符串中的变量可以使用表达式,也可以直接调用函数。
```js
let age = 18
let str = `今年 ${age / 2} 岁`
function getSum(num1, num2) {
return num1 + num2
}
let str = `数字的和是 ${getAge(10, 20)}`
```
模板字符串会保留所有的空格和换行。
```js
let str = `
<div>
<span></span>
</div>
` // 显示的结果会保留缩进格式
```
## Set 集合
ES6 中新增了类似于数组的结构 Set,但是它的元素值都是唯一的,不会重复。使用构造函数`Set()`可以创建一个 Set 结构,它可以接收任何数组(伪数组)参数,将其转换为`Set`,并自动去除重复元素。
```js
var arr = [10, 10, 30, 50]
new Set(arr) // => Set {10, 30, 50}
```
它包含下列的常用属性和方法:
| 属性或方法 | 描述 |
| --- | --- |
| `size` | 获取集合的长度 |
| `add()` | 向集合中添加一个值 |
| `delete()` | 删除集合中的一个值,如果删除成功则返回`true`,否则返回`false` |
| `has()` | 判断集合中是否包含某个值,如果包含则返回`true`,否则返回`false` |
| `clear()` | 清空一个集合 |
```js
const arr = [10, 20, 30]
new Set(arr).size // => 3
new Set(arr).add(40).add(50) // => Set {10, 20, 30, 40, 50}
new Set(arr).delete(20) // => Set {10, 30}
new Set(arr).has(30) // => true
new Set(arr).clear() // => Set {}
```
## 类
ES6 中提供了类的概念,用于简化之前~~反人类~~的原型写法,属于**原型的语法糖**。
使用`class`关键字可以定义一个类,其中可以包含`constructor()`构造方法、普通方法,最后通过`new`实例化即可:
```js
class Animal {
constructor (age) { this.age = age }
eat() { console.log('吃呀吃呀吃') }
}
var cat = new Animal(3)
cat.age // => 3
cat.eat() // => 吃呀吃呀吃
```
### 类的继承
使用`extends`关键字可以使子类继承父类所有的属性和方法。
```js
class Animal {
eat() { console.log('吃呀吃呀吃') }
}
class Cat extends Animal {
run() { console.log('猫在跑') }
}
let cat = new Cat()
cat.eat() // => 吃呀吃呀吃
```
如果子类有自定义的构造函数,那么必须在其中先使用`super()`调用父类的构造函数,否则会出现语法错误。这是因为 ES6 的继承需要首先创建父类实例,然后再将子类的属性和方法添加到父类实例上,再返回子类实例,因此子类的实例化必须依赖于父类实例。如果子类没有自定义构造函数,那么引擎也会自动添加上。
```js
class Animal {}
class Cat extends Animal {
constructor () {
super() // 必须先手动调用父类构造函数
}
}
```
如果子类中定义了与父类相同的方法,那么会将父类的覆盖。
```js
class Animal {
eat() { console.log('动物在各种吃') }
}
class Cat extends Animal {
eat() { console.log('猫在各种吃') }
}
var cat = new Cat()
cat.eat() // => 猫在各种吃
```
子类中的`super`关键字指向父类本身,也就是子类的原型对象,因此可以直接调用父类的方法:
```js
class Animal {
eat() { console.log('动物在各种吃') }
}
class Cat extends Animal {
eat() { super.eat() } // 使用 super 指向父类
}
var cat = new Cat()
cat.eat() // => 动物在各种吃
```
但是,`super`指向的不是父类的实例对象,因此不能获取父类实例中的属性:
```js
class Animal {
constructor() { this.age = 3 } // this 指向的是实例对象,因此 age 是实例对象中的属性
}
class Cat extends Animal {
print() { console.log(super.age) }
}
var cat = new Cat()
cat.print() // => undefined
```
## 异步编程
JavaScript 是一门**单线程**的语言,也就是说,同一时间只能做一件事情。这意味着,如果程序执行中遇到了耗时很长的任务,那么后面的代码都要等待。于是 JavaScript 使用了一套专门的机制,将任务分成了同步和异步。
其中大部分的代码都是同步任务,它们位于**主线程**依次执行,只有上一个任务执行完毕后才能执行下一个任务。而一些不能立即获得结果的任务(比如定时器函数、网络请求、事件响应、IO 操作等),会被放到**任务队列**(也称消息队列)中。当主线程任务执行完毕后,会从任务队列中读取一条任务继续执行,完成后再次读取任务队列,这样产生的循环称为**事件循环**。
由于异步任务的执行时间未知,没法立即返回结果,因此在 ES5 及之前,实现异步编程的主要方式为回调函数。
在下面的代码中,主线程会先执行第一条语句,当执行到第二条语句时,引擎发现这是一个定时器异步任务,会将其加入任务队列并跳过(即便延迟时间为`0`),然后继续执行第三条语句。之后,主线程任务均执行完毕后,查看任务队列,然后取出定时器函数中的语句执行。
```js
console.log('张三')
setTimeout(() => console.log('王五'), 1000)
console.log('李四')
// => 张三 李四 王五
```
但回调函数带来了新的问题。
在下面的代码中,先延迟`1s`输出`first`,然后再延迟`1s`输出`second`,为了确保这一点,只能将第二次输出的代码放在第一次的回调函数中,以此类推。
```js
setTimeout(function () {
console.log('first')
setTimeout(function () {
console.log('second')
setTimeout(function () {
console.log('third')
}, 1000)
}, 1000)
}, 1000)
```
可想而知,如果要依次输出的内容更多,那么代码就会层层嵌套,完全失去可维护性,这样的情况被称为**回调地狱**。
### Promise
为了解决这一问题,ES6 新增了一个`Promise`对象,该对象类似于一个容器,用来封装异步任务。
异步任务有很多,但是它们都有一个共性:要么执行成功要么失败。因此,要实例化`Promise`对象需要传入一个函数参数,它的两个参数`resolve`和`reject`又是两个函数,分别表示成功和失败这两种状态。异步任务在执行完成后,不再通过回调函数执行具体的功能,而是通过这两个函数将执行完毕后的结果传递到外部,由外部对结果进行处理。
```js
const first = new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第一个定时器啦') // 1 秒过后,通过回调函数执行 resolve() 将结果传递出去,而不是直接输出
}, 1000) // 由于定时器任务没有失败的回调函数,因此不需要调用 reject()
})
```
接下来,`Promise`对象提供了一个`then()`方法,它接收一个函数参数表示成功状态时的处理函数,该函数的形参为`resolve()`传递出来的结果,这样就实现了将异步任务结果分离到回调函数之外来处理。
```js
first.then(function (result) {
console.log(result) // 这是第一个定时器啦
})
```
除此之外,`then()`还可以接收第二个参数作为失败状态时`reject()`的处理函数,用法与`resolve()`一样。捕获错误也可以使用单独的`catch()`方法,由于`then()`会**默认自动**返回调用它的`Promise`对象,因此可以链式编程。
```js
first.then(function (result) {
console.log(result) // 这是第一个定时器啦
}, function (err) {
console.log(err) // 捕获错误,相当于 reject()
})
// 相当于
first.then(function (result) {
console.log(result) // 这是第一个定时器啦
}).catch(function (err) {
console.log(err) // 捕获错误,相当于 reject()
})
```
接下来继续回到之前的问题,发现问题依然没有完全解决。因为一旦调用了`new Promise()`,它内部的异步任务就会立刻被执行,这样的效果就相当于同时定义了三个定时器并执行,没有先后顺序。
因此为了防止它们自动执行,要先将它们放到一个函数中,等需要时手动调用,将实例化的`Promise`对象返回:
```js
function first() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第一个定时器啦')
}, 1000)
})
}
first().then(function (result) {
console.log(result) // 这是第一个定时器啦
})
```
之前提到,默认情况下`then()`会返回调用它的`Promise`对象,但是也可以手动设置它的返回值。因为第二个定时器依赖于第一个定时器执行完毕,所以在第一个定时器中返回第二个定时器的`Promise`对象,就可以实现依次调用的效果了。
```js
function first() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第一个定时器啦')
}, 1000)
})
}
function second() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第二个定时器啦')
}, 1000)
})
}
function third() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第三个定时器啦')
}, 1000)
})
}
first().then(function (result) {
console.log(result) // 这是第一个定时器啦
return second() // 返回第二个 Promise,通过它再去调用第二个 then(),从而执行第二个定时器
}).then(function (result) {
console.log(result) // 这是第二个定时器啦
return third()
}).then(function (result) {
console.log(result) // 这是第三个定时器啦
})
```
由此可见,虽然可以继续将返回`Promise`对象的函数合并封装,但是这个过程还是非常繁琐的。建议直接使用类似于[bluebird.js](/posts/0wuyppfk.html)的第三方库。
### 异步函数
虽然通过`Promise`解决了回调地狱的问题,但是连续的`then()`方法使得代码看起来依然不是很直观。于是 ES7 中又加入了**异步函数**的概念,它可以使异步的代码看起来与同步代码一样,更加直观易读。
普通的函数一旦开始执行,中间是不能暂停的,而异步函数可以中途停止,等待某个任务执行完毕返回结果,再继续执行。
在`function`前添加`async`关键字,可以定义一个异步函数。
```js
async function fn() {}
```
在其中的异步任务前添加`await`关键字,可以告知函数在此处暂停,直到异步任务返回结果。注意,**异步任务必须返回`Promise`对象才能被正确处理**,否则就相当于没有添加`await`。使用了`await`之后,就可以直接通过返回值来获取`Promise`对象中`resolve()`的结果,与同步代码几乎一样。
```js
function first() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第一个定时器啦')
}, 1000)
})
}
function second() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第二个定时器啦')
}, 1000)
})
}
function third() {
return new Promise(function (resolve, reject) {
setTimeout(function () {
resolve('这是第三个定时器啦')
}, 1000)
})
}
async function fn() {
let r1 = await first()
console.log(r1)
let r2 = await second()
console.log(r2)
let r3 = await third()
console.log(r3)
}
fn()
```
如果使用了[bluebird.js](/posts/0wuyppfk.md),那么`delay()`函数本身返回的便是`Promise`对象,因此代码可以进一步简化。
```js
Promise.delay(1000).then(function () {
console.log('这是第一个定时器啦')
}).delay(1000).then(function () {
console.log('这是第二个定时器啦')
}).delay(1000).then(function () {
console.log('这是第三个定时器啦')
})
// 等同于
async function fn() {
await Promise.delay(1000)
console.log('这是第一个定时器啦')
await Promise.delay(1000)
console.log('这是第二个定时器啦')
await Promise.delay(1000)
console.log('这是第三个定时器啦')
}
fn()
```
## 模块化
JavaScript 自从诞生以来,一直没有类似于模块(Module)的体系,它无法将大型程序拆分成多个小文件,再拼装起来。比如在浏览器端,虽然一个页面可以引入多个 JavaScript 文件,但是它们其实会被浏览器合并成一个大文件,因此会出现命名冲突的问题。
而所谓模块化,就是将每个`.js`文件视为一个**模块**,每个模块会形成一个独立的模块作用域,它们彼此之间不能访问,因此不会出现命名冲突等问题。如果模块之间需要互相访问,则只需在一个模块中导入另一个模块即可;如果要访问其它模块中的具体数据,则要在被访问的模块中先导出数据,再从另一个模块中导入。
为了实现模块化,JavaScript 社区提供了一些解决方案,如浏览器端的 AMD(require.js)和 CMD(Sea.js)等,而服务端的 Node.js 也提出了 CommonJS,并沿用至今。
之后,JavaScript 官方在 ES6 中实现了模块化的功能,但是由于历史原因,Node.js 目前还无法支持。而浏览器端的大部分现代浏览器虽然已经实现了该特性,但是考虑到兼容性问题,也无法使用。因此,要使用原生的 ES6 模块化语法,只能通过 babel 等兼容性工具转换后才可以。
首先,在`<script>`中使用`import`关键字可以导入一个模块,并**自动执行其中的代码**。注意,此时的`<script>`标签必须添加`type="module"`属性,否则无法使用`import`语句。
```html
<script type="module">
import './test.js'
</script>
```
或者也可以写在单独的`.js`文件中:
```html
<script type="module" src="index.js"></script>
```
```js
/* index.js */
import './test.js'
```
### 按需导入与导出
模块可以使用`export`关键字根据需要导出指定的数据,数据需要使用花括号包裹,多个数据以逗号分隔:
```js
/* test.js */
let str = 'hello'
let num = 233
export { str, num }
```
导入模块时,要获取这些数据,也需要通过花括号包裹,并且导出和导入时的**数据名称要保持一致**:
```html
<script type="module">
import { str, num } from './test.js'
str // => hello
num // => 233
</script>
```
### 默认导入与导出
但有时在加载第三方模块时,去逐一地了解需要导入哪些数据实在有些困难,因此在导出模块时,可以使用`default`关键字指定**默认导出**的数据。这样在导入模块时,就不需要确切地声明要导入的数据了,只要任意起名即可。并且,此时导入语句中的数据名**不能加大括号**:
```js
/* test.js */
let num = 233
export default num
```
```html
<script type="module">
import something from './test.js' // something 不需要加大括号
something // => 233
</script>
```
默认导出本质上是将`default`后面的值赋值给`default`变量,而导入时`import`后面的内容就是这个`default`变量。因此`export default`后可以跟任意数据类型,但是不能跟一条语句。
也可以同时使用默认和按需两种导入导出方式:
```js
/* test.js */
let num = 233
let str = 'hello'
export { str }
export default { num }
```
```html
<script type="module">
import something, { str } from './test.js' // 使用逗号隔开按需和默认导入导出
something // => { num: 233 }
str // => hello
</script>
```
此外,默认导出的语句在模块中**只能出现一次,不允许多次使用**。<file_sep>---
title: 大前端支线笔记之07 ⚽️ AJAX
date: 2020-02-17 12:00:26
abbrlink: ljghe0it
tags: JavaScript
categories:
- 大前端
- 支线
excerpt: AJAX(Asynchronous Javascript And XML)指的是异步加载 JavaScript 和 XML,也就是使浏览器异步地向服务器发送请求,以获得新的数据,再通过操作 DOM 实现在不刷新页面的情况下展示新的内容。
---
# 大前端支线笔记之07 ⚽️ AJAX
在传统的 Web 应用中,当用户提交表单时,需要向服务端发送一个请求,服务端处理完收到的表单后,再将一个新的网页响应回来。但是这样的做法有些缺点:首先,填写表单的页面和返回的页面大部分内容都是相同的,浪费了传输的带宽;其次,对于用户来说,提交表单之后需要等待页面重新加载,体验并不友好。
为了解决这些问题,有人提出了 AJAX 这一概念。AJAX(Asynchronous Javascript And XML)指的是异步加载 JavaScript 和 XML,也就是使浏览器异步地向服务器发送请求,以获得新的数据,再通过操作 DOM 实现在不刷新页面的情况下展示新的内容。
这一技术目前已被 JavaScript 正式引入,它通常应用在如页面滚动加载数据、局部更新、实时验证用户表单、搜索框信息提示等多种场景。
> AJAX 名称中之所以包含 XML 是因为早期的互联网信息传输中主要以 XML 为内容格式,但是目前 XML 基本已被 JSON 取代。按道理说,这个名称现在也可以改为 AJAJ,但是由于已经大家习惯了 AJAX,因此它暂时还是被保留了下来。
## 原生 AJAX
JavaScript 通过`XMLHttpRequest`对象提供对 Ajax 的支持,因此需要先实例化该对象。
```js
var xhr = new XMLHttpRequest()
```
然后使用该对象的`open()`方法可以初始化一个请求,建立与服务端的连接。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 请求方式 |
| 字符串 | 请求 URL |
```js
xhr.open('GET', 'http://localhost/test')
```
接下来,使用该对象的`send()`方法发送请求。
```js
xhr.send()
```
如果要传递`GET`请求参数,将其拼接到 URL 一并发送即可。
如果要传递`POST`请求参数,需要先了解请求体的几种类型,通过表单提交数据时,该类型通过`enctype`属性设置,否则的话,需要手动设置请求头中的`Content-Type`属性:
* `application/x-www-form-urlencoded`:表单`POST`提交方式的默认类型,如`name=lucy&age=18`
* `multipart/form-data`:二进制类型,上传文件时经常使用
* `application/json`:JSON 格式数据,不支持表单提交,如`{"name": "lucy", "age": 18}`
然后通过`setRequestHeader()`设置请求体类型:
```js
xhr.setRequestHeader('Content-Type', 'application/x-www-form-urlencoded')
```
最后将数据放到`send()`方法中即可:
```js
xhr.send('name=lucy&age=18')
```
## 接收响应
由于 Ajax 请求是异步操作,因此当请求发送成功后会触发`load`事件,此时通过请求对象的`responseText`属性可以获取服务器响应的结果。
```js
xhr.addEventListener('load', function() {
this.responseText // => ok
})
```
### 低版本浏览器
低版本浏览器并不支持`load`事件,因此需要通过监听 Ajax 的状态来实现接收响应。当 Ajax 的状态发生变化时,会触发`onreadystatechange`事件,而当前的状态通过`readyState`属性来保存,它共有 5 个数字值,分别为:
* `0`:`open()`方法还未被调用
* `1`:`open()`方法已经被调用
* `2`:`send()`方法已经被调用,并且获取了响应头的信息
* `3`:`responseText`已经获取了部分响应体的数据
* `4`:整个请求过程完毕
也就是说,当`readyState`属性为`4`时,才可以通过`responseText`属性获取响应体的数据和其它响应报文信息。
```js
xhr.addEventListener('readystatechange', function() {
if (this.readyState == 4) {
this.responseText // 获取响应体的数据
this.status // => 200,HTTP 状态码
this.statusText // => OK,HTTP 状态信息
}
})
```
## FormData
通过`FormData`对象可以模拟一个表单,并通过 Ajax 将其提交到服务器。首先通过表单元素实例化一个`FormData`对象,使用 Ajax 将`FormData`对象发送到服务器。
```html
<form id="myform">
<input type="text" id="username" name="username" value="hello">
<input type="text" id="age" name="age" value="18">
<button type="button" id="submit">提交</button>
</form>
```
```js
$('#submit').click(function () {
var formData = new FormData($('#myform')[0])
var xhr = new XMLHttpRequest()
xhr.open('POST', 'http://localhost/')
xhr.send(formData)
})
```
### 相关方法
此外,它还包括下列的常用方法。
| 方法名 | 描述 |
| ---------- | ------------------------------------------------------------ |
| `get()` | 获取表单属性的值 |
| `set()` | 设置表单属性的值,如果存在同名属性,则覆盖该属性值;如果不存在则会自动创建 |
| `delete()` | 删除表单属性 |
| `append()` | 添加表单属性 |
```js
formData.get('age') // => 18
formData.set('age', 19) // => { username: 'hello', age: '19' }
formData.delete('age') // => { username: 'hello' }
formData.append('sex', 'male') // => { username: 'hello', age: '19', sex: 'male' }
```
## 同源策略
同源策略是浏览器的一种安全策略,所谓同源指的是两个地址之间的**域名**、**协议**和**端口号**完全相同,只要有其中一项不同,那么这两个地址就是不同源的。只有同源的地址之间才能互相发送 Ajax 请求,不同源之间的相互请求被称为**跨域**。
在下面的请求中,从`http://localhost`向`http://test.io`发送请求,浏览器会出现报错信息。
```js
xhr.open('GET', 'http://test.io')
xhr.send()
```
### 跨域资源共享(☢️IE10)
从上面的报错信息中可以看到,请求被 CORS 策略阻止了,它指的是跨域资源共享(Cross-origin resource sharing)。而后面提示中指出,在响应头信息中没有找到`Access-Control-Allow-Origin`字段,也就是说,如果服务端希望接收来自其它域的 Ajax 请求,只需要添加一条响应头信息即可:
```js
// 通过中间件拦截所有请求
app.use((req, res) => {
// * 表示对任何请求地址均采用 CORS,如果指定了域名,那么只有来自该域名的请求能正确接收
res.header('Access-Control-Allow-Origin', '*')
next()
})
```
> 在跨域资源共享出现之前,是通过 JSONP 来实现这一功能的,但是由于代码比较复杂,不再赘述,直接使用 jQuery 中的 JSONP 即可。
>
> 此外,由于服务端没有同源策略限制,可以先由客户端请求同源的服务器,由服务器再发送跨域请求,获取到数据后再返回给客户端。
### 跨域携带 Cookie
默认情况下,跨域请求是不会携带客户端 Cookie 的。要修改这一点,需要客户端和服务端同时进行设置。
```js
// 客户端
xhr.open('GET', 'http://test.io')
xhr.withCredentials = true // 设置跨域时携带 Cookie
xhr.send()
```
```js
// 服务端
res.header('Access-Control-Allow-Credentials', true) // 响应头也要同时设置
```
## $.ajax()
发送 Ajax 请求。它的参数为一个对象,其中包含若干配置属性:
```js
$.ajax({
url: '/add', // 请求地址
type: 'post', // 请求类型
data: { name: '离散懵逼', age: 38 }, // 传递给服务器的参数,会被自动转换成 urlencode 格式
// 设置为 dataType: 'jsonp' 表示发送 JSONP 请求
dataType: 'json', // 要求响应体的数据必须为 JSON 格式,默认为根据响应头的 content-type 自动识别
beforeSend: function() { // 请求发送前的回调函数,可以用来验证数据或者显示 Loading 动画
return false // 如果返回 false 则取消发送当前请求
},
// 根据状态码(是否为 200)判断请求状态
success: function (res) {}, // res 为服务端返回的响应体内容,会根据 dataType 自动完成转换
error: function (xhr) {}, // xhr 为原生的 XMLHttpRequest 对象
})
```
## serialize()
将表单控件中的值转换成 URLencoded 格式的字符串。
```js
$('form').serialize() // => name=Lucy&age=18
```
如果要直接转换成对象,可以使用 jQuery 的第三方插件[serializeJSON](https://github.com/marioizquierdo/jquery.serializeJSON )。
## $.get() / $.post()
快速发送 GET / POST 请求。
```js
$.get('/add', { age: 18 }, function(res) {})
```
## $.ajaxStart() / $.ajaxComplete()
统一设置 Ajax 请求发送之前 / 之后的回调函数,该事件只能绑定到`document`对象上。
```js
$(document).ajaxStart(function () {
// 请求即将发送
})
```
## load()
在 Ajax 请求成功并获取返回数据后,自动调用`html()`将数据插入到匹配的元素中。
```js
$('div').load('backend/test.php', { key: 'value'}, function(res) {});
```
如果请求返回的是一个页面,还可以在 URL 参数后面添加一个空格,然后追加选择器,来获取更加具体的内容。
```js
$('#div-elem').load('test.php #banner');
```
<file_sep>---
title: 📄 art-template
date: 2020-01-10 22:32:43
abbrlink: 3q4y7l2d
tags: 模板引擎
categories:
- 大前端
- 第三方库
excerpt: 施工中...
---
# 📄 art-template
施工中,修改时间
[art-template](http://aui.github.io/art-template/zh-cn/docs/)是一款国内开发的模板引擎,由于有中文文档,并且语法简单容易上手,非常适合初学者使用。
```powershell
npm install art-template
```
```js
const template = require('art-template')
```
### 拼接模板和数据
首先通过`template()`方法将模板和数据拼接起来。
> * 参数①:模板的路径
>* 参数②:要拼接的数据,必须是对象
> * 返回值:拼接后的 HTML 代码
```js
resPage = template('/list.html', { name: '御坂美琴', age: 16 }) // => 我是御坂美琴,今年16岁啦!
```
接下来,在模板中使用特定的语法即可使用传入的数据。
```ejs
我是 {{ name }},今年 {{ age }} 岁啦!
```
## 基本语法
### 输出
使用下面的方式可以直接取出对象或数组中的值进行输出,并且支持简单的表达式运算。注意,模板引擎会自动将 HTML 标记进行转义处理,不会进行解析。
```js
template(tplPath, {
name: '御坂美琴',
age: 16,
hobbies: ['打人', '放电'],
friend: { name: '白井黑子' }
})
```
```
{{ name }} // => 御坂美琴
{{ hobbies[0] }} // => 打人,通常用下面的循环来遍历数组
{{ friend.name }} // => 白井黑子
{{ age > 18 ? '已成年' : '未成年' }} // => 未成年,注意其中显示的值要用单引号包裹
```
### 条件
根据数据可以判断某些 HTML 是否出现。
```
{{if age > 18}} 我要上网吧!{{/if}}
{{if price < 10}}
这个好便宜
{{else if price < 20}}
还可以
{{else}}
太贵了!
{{/if}}
```
### 循环
通过循环可以遍历数组和对象。在循环中使用`$value`和`$index`可以获取当前元素的值和索引。
```
{{ each hobbies }}
当前为 {{ $index }} 号元素,值为{{ $value }}
{{ /each }}
```
如果只需要遍历指定的次数(比如循环输出分页按钮),那么只能使用原始语法:
```ejs
<% for(var i = 0; i < totalPage; i++){ %>
<li class="page-item"><a class="page-link" href="/admin/user/?page={{ i }}"> {{ i }} </a></li>
<% } %>
```
### 子模板
通过子模板可以将页面公共的部分抽离出来,方便统一维护。注意这里的路径使用相对路径即可,它相对的是当前模板,而不是调用它的 JavaScript 文件。
```
{{ include './list.art' }}
```
### 模板继承
通过模板继承可以使多个页面模板继承共同的一个 HTML 骨架,对于每个页面独有的内容,可以在骨架页面中预留位置,子模板只需要设置预留位置的内容即可。
下面的骨架模板中通过双标记`{{block}}`预留了两个位置,为了区分它们,可以在其中自定义一个名称。
```
// 骨架模板 layout.art
<!DOCTYPE html>
<html lang="en">
<head>
<meta charset="UTF-8">
<meta name="viewport" content="width=device-width, initial-scale=1.0">
<meta http-equiv="X-UA-Compatible" content="ie=edge">
{{block 'title'}} {{/block}}
</head>
<body>
{{block 'content'}} {{/block}}
</body>
</html>
```
接下来,在子模板中使用`extend`引入骨架,并设置预留内容。
```
{{extend './layout.art'}}
{{block 'title'}}
<title>首页</title>
{{/block}}
{{block 'content'}}
<p>这里是首页啦</p>
{{/block}}
```
## 配置选项
配置选项通过`template.defaults`对象设置。
### extname
模板默认的后缀名是`.art`,如果传入路径参数时省略了后缀名,那么模板引擎会自动补充。
```js
template.defaults.extname = '.html'
```
### root
模板通常会放到统一的文件夹中,使用该属性可以修改模板的根目录。
```js
template.defaults.root = path.join(__dirname, 'views')
// 引入模板时,可以省略拼接路径和后缀了
template('index', data)
```
## 浏览器环境
前台的模板语法不能直接写在页面上,因为其中的语法部分会被浏览器当做普通文本呈现出来。为了避免这个问题,可以将模板放入`<script>`标签中,它的内容默认会被浏览器隐藏,而且只要将其设置为`type="text/html"`就能将其中的 HTML 元素正常着色。
```html
<script id="tpl-users" type="text/html"></script>
```
然后,依然使用模板引擎的`template()`方法,将模板与数据联系起来。
> * 参数①:模板元素的`id`
>* 参数②:要拼接的数据,必须是对象
> * 返回值:拼接后的 HTML 代码
```js
let html = template('tpl-users', { name: '御坂美琴', age: 16 })
```
接下来,通过模板引擎的语法在模板中解析数据:
```html
<script id="tpl-users" type="text/html">
我是{{ name }},今年{{ age }}岁啦!
</script>
```
最后,将`template()`方法返回的拼接字符串追加到页面元素中即可。
```js
$('.msg').html(html)
```<file_sep>---
title: 大前端笔记之17 🔔 JavaScript 事件
date: 2020-02-05 14:39:44
abbrlink: g6oz01bg
tags: JavaScript
categories: 大前端
excerpt: 事件是程序的各个组成部分之间的一种通信方式,可以认为是元素的一种行为。将事件与监听函数绑定,当事件触发时,那么就会执行对应的监听函数。
---
# 大前端笔记之17 🔔 JavaScript 事件
事件是程序的各个组成部分之间的一种通信方式,可以认为是元素的一种行为。将事件与监听函数绑定,当事件触发时,那么就会执行对应的监听函数。
## 绑定事件
由于元素对象会自动获取所有自带的属性,而事件属性也属于其中之一,因此可以直接通过元素的事件属性可以绑定事件。
```js
btn.onclick = function() { alert('事件似乎被触发了!') }
```
但是,此时每个元素的同一事件只能绑定一个监听函数,如果再次定义了同一事件,那么之前的监听函数会被覆盖,因此不推荐使用。
更多情况下,都会使用元素对象的`addEventListener()`方法绑定事件。而使用该方法绑定的事件,可以使用该元素对象的`removeEventListener()`方法移除,而且它们的参数都是一样的。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 事件名称,注意没有`on`前缀 |
| 函数 | 事件触发时执行的监听函数 |
```js
function sendMsg() { alert('事件似乎被触发了!') }
btn.addEventListener('click', sendMsg)
btn.removeEventListener('click', sendMsg)
```
> IE9 之前有一个类似的绑定事件方法`attachEvent()`,但是已被从规范中移除,不要使用。
## 事件对象
每当事件被触发时,都会产生一个事件对象作为监听函数的参数,这些事件对象均继承自`Event`对象。
```js
// e 为事件对象,可以是任何名称
btn.addEventListener('click', function (e) {})
```
### target
获取触发事件的元素对象。注意,监听函数中的`this`指代**绑定事件**的对象,而`target`获取的是**触发事件**的对象,由于事件具有冒泡特性,因此两者未必是相同的。
在下面的代码中,事件绑定在`<ul>`上,而点击的是`<li>`元素,因此`this`指代的是`<ul>`,而`target`指代的是`<li>`:
```html
<ul>
<li>冥灯龙</li>
<li>灭尽龙</li>
</ul>
```
```js
ul.addEventListener('click', function (e) {
this // => ul
e.target // => li
})
```
### type
获取触发的事件类型。
```js
div.addEventListener('click', function (e) {
e.type // => click
})
```
### preventDefault()
阻止元素自带的默认事件,比如`<a>`元素拥有默认事件`click`,当事件触发时会跳转到指定页面:
```html
<a href="http://www.bilibili.com">哔哩哔哩</a>
```
```js
a.addEventListener('click', function (e) {
e.preventDefault() // 阻止页面跳转
})
```
> 除此之外,还可以在监听函数中加入`return false`语句,但是它不仅会阻止默认事件,还会**阻止事件冒泡**。由于它的双重作用,因此不推荐使用这种方式。
### 鼠标位置
使用事件对象的下列属性可以获取鼠标位置的相关信息,它们的值均**没有单位**,而且为只读。
| 属性 | 描述 |
| --- | --- |
| `offsetX`/`offsetY` | 鼠标相对于**触发事件元素**内边距区域的水平 / 垂直方向坐标 |
| `clientX`/`clientY` | 鼠标相对于**视口**的水平 / 垂直方向坐标 |
| `pageX`/`pageY` | 鼠标相对于**页面文档**的水平 / 垂直方向坐标 |
| `screenX`/`screenY` | 鼠标相对于**设备屏幕**的水平 / 垂直方向坐标 |
```js
document.addEventListener('mousemove', function (e) {
e.pageX
})
```
> 注意在实现跟随鼠标移动效果时,**不要将鼠标完全放在跟随的元素上**,这样可能会导致一些鼠标事件无法正常触发。比如点击事件正好点在跟随的元素上,而不是原本想要点击的页面元素。
### 键盘按键
使用事件对象的`code`和`key`属性可以获取用户按下的键。但是`code`属性 IE 和 Edge 不支持,`key`属性 IE8 不支持,而且它们的返回值也有所不同。
```js
document.onkeydown = function (e) { // 分别输入回车和字母 A
e.code // => Enter 和 keyA
e.key // => Enter 和 a
e.keyCode // => 13 和 65
e.which // => 13 和 65
}
```
> 注意,`keyCode`和`which`也可以实现同样功能,但是已经被废弃,不建议使用。jQuery 中将`which`属性进行了封装,可以直接使用。
### 屏幕触摸
使用事件对象的`targetTouches`属性可以获取触发事件时位于**触发事件元素**上的触摸点,它返回一个`TouchList`集合,通过`length`和方括号`[]`可以获取其中的`Touch`对象以及数量。每个`Touch`对象代表当前手指的触摸点,其中包含[鼠标位置](#鼠标位置)中的全部属性(`offsetX`和`offsetY`除外)。
```js
div.addEventListener('touchstart', function (e) {
e.targetTouches[0].clientX // 获取当前触摸点相对视口的 x 坐标
})
```
使用事件对象的`touches`属性可以获取触发事件时位于**整个屏幕**上的触摸点,而`changedTouches`可以获取触发事件时**触摸状态发生改变**(之前触摸现在松开,或反之)的触摸点。它们的用法与`targetTouches`基本一致。
## 事件传播
如果子元素与父元素重叠在一起,并且都绑定了事件,那么某些操作(比如鼠标点击、移入移出)可能会导致它们的事件同时触发,这个现象称为事件的传播。JavaScript 将事件传播分为三个阶段,使用事件对象的`eventParse`属性可以获取:
* 捕获阶段:从`html`对象传递到目标节点(从顶层传递到底层),`eventParse`值为`1`
* 目标阶段:在目标节点上触发,为嵌套最深的子节点,`eventParse`值为`2`
* 冒泡阶段:从目标节点传递到`html`对象(从底层传递到顶层),`eventParse`值为`3`

默认情况下,事件会绑定在**冒泡阶段**,也就是说事件会从被点击的内层元素开始触发,直到外层元素。
```html
<div id="outer">
<p id="inner"></p>
</div>
```
```js
inner.addEventListener('click', function (e) {
console.log('内层元素被点击啦,当前阶段为' + e.eventPhase)
})
outer.addEventListener('click', function (e) {
console.log('外层元素被点击啦,当前阶段为' + e.eventPhase)
})
// => 内层元素被点击啦,当前阶段为 2
// => 外层元素被点击啦,当前阶段为 3
```
通过`addEventListener()`方法的第三个参数,可以将事件绑定在捕获阶段。该参数为布尔值,将其设置为`true`,表示绑定在捕获阶段。但是通常情况下不需要这么做,使用冒泡阶段即可。
```js
inner.addEventListener('click', function (e) {
console.log('内层元素被点击啦,当前阶段为' + e.eventPhase)
}, true)
outer.addEventListener('click', function (e) {
console.log('外层元素被点击啦,当前阶段为' + e.eventPhase)
}, true)
// => 外层元素被点击啦,当前阶段为 1
// => 内层元素被点击啦,当前阶段为 2
```
### 阻止事件传播
通过事件对象的`stopPropagation()`方法可以阻止事件传播。
```js
inner.addEventListener('click', function (e) {
e.stopPropagation() // 阻止了事件传播,后面的所有阶段不再执行
})
outer.addEventListener('click', function (e) {
console.log('触发不了啦')
})
```
## 常用事件
下面是一些常用的事件。
### 鼠标事件
| 事件 | 描述 |
| --- | --- |
| `click` | 鼠标左键单击时触发,该事件在触发前会首先触发另外两个事件:`mousedown`与`mouseup` |
| `dblclick` | 鼠标左键双击时触发 |
| `mousedown` | 鼠标键按下时触发 |
| `mouseup` | 鼠标键抬起时触发 |
| `mousemove` | 鼠标在元素上移动时触发 |
| `mouseenter` | 鼠标移入元素时触发(不冒泡),并且当进入该元素的子元素时,**不会**导致该事件再次被触发 |
| `mouseover` | 鼠标移入元素时触发(冒泡),并且当进入该元素的子元素时,会导致该事件再次被触发 |
| `mouseleave` | 鼠标移出元素时触发(不冒泡),并且只会在移出该元素本身时触发一次 |
| `mouseout` | 鼠标移出元素时触发(冒泡),并且当移出父元素进入子元素,或者移出子元素时,都会导致该事件再次被触发 |
| `contextmenu` | 点击右键时弹出菜单,为`document`绑定并阻止默认事件可以禁用弹出菜单 |
### 键盘事件
下面三个事件如果同时注册,那么执行顺序从上到下,即`keydown`一定会优先触发,然后为`keypress`,而`keyup`最后触发。
| 事件 | 描述 |
| --- | --- |
| `keydown` | 键盘按键按下时触发,如果不松开按键该事件也会持续触发 |
| `keypress` | 键盘有值按键(即方向键、<kbd>Ctrl</kbd>等功能键除外)按下时触发,如果不松开按键该事件也会持续触发 |
| `keyup` | 键盘按键抬起时触发 |
### 表单事件
| 事件 | 描述 |
| --- | --- |
| `focus` | 表单元素获得焦点时触发 |
| `blur` | 表单元素失去焦点时触发 |
| `change` | 当`<input>`、`<select>`、`<textarea>`元素的值发生改变时触发 |
| `input` | 与`change`类似,区别在于只要是元素的值发生了改变,该事件会连续触发,而`change`不会 |
| `select` | 当文本框和文本域的内容被选中时触发,通过 DOM 元素调用`select()`方法可以主动选中其中的内容 |
| `submit` | 当表单提交时触发,设置`preventDefault()`可以阻止表单默认提交 |
下面是`change`事件详细的触发条件:
- 激活单选框或复选框时触发
- 在`<select>`或日期控件完成选择时触发
- 当文本框或`<textarea>`元素的值发生改变,并且丧失焦点时触发
### 动画事件
| 事件 | 描述 |
| --------------- | -------------------------------------- |
| `transitionend` | 当元素的`transition`动画执行完毕后触发 |
| `animationend` | 当元素的`animation`动画执行完毕后触发 |
### BOM 事件
| 事件 | 描述 |
| --------- | ------- |
| `scroll` | 元素的滚动条发生滚动时触发 |
| `load` | 当页面加载完成时触发,必须绑定给`window`对象 |
| `pageshow` | 当页面加载完成时触发,必须绑定给`window`对象 |
| `DOMContentLoaded` | 当 DOM 元素加载完成时触发,不包括图片、样式等部分,必须绑定给`document`对象 |
| `resize` | 当浏览器窗口发生改变时触发,必须绑定给`window`对象 |
#### 页面加载
`load`和`pageshow`事件都是在页面加载完成后触发,区别在于,火狐浏览器在页面跳转再通过后退按钮返回时,会将页面保存到缓存中,此时`load`事件不会再次触发,而`pageshow`事件则不会有这个问题。
### 触屏事件
| 事件 | 描述 |
| ------------ | ---------------------- |
| `touchstart` | 手指触摸到元素时触发 |
| `touchmove` | 手指在元素上滑动时触发 |
| `touchend` | 手指从元素上松开时触发 |
### 其它事件
| 事件 | 描述 |
| --------- | ------- |
| `selectstart` | 当用户选择页面文字时触发,为`document`绑定并阻止默认事件可以禁止用户选择页面文字 |
<file_sep>---
title: 大前端笔记之06 🏢 CSS 布局
date: 2020-01-23 0:49:46
abbrlink: rbty8hf8
tags: CSS
categories: 大前端
excerpt: 默认情况下,元素都位于普通文档流中,此时元素会按照从上到下、从左到右依次排列。但是通过下面的方式,可以修改元素的排列方式。
---
# 🏢 CSS 布局
默认情况下,元素都位于普通文档流中,此时元素会按照从上到下、从左到右依次排列。但是通过下面的方式,可以修改元素的排列方式。
## 浮动
浮动原本是用来修改文章中图片位置的,但是它经常被误用于布局中,用来使多个元素排列在一行。
使用`float`可以设置元素浮动。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 不浮动 |
| `left` | 左浮动 |
| `right` | 右浮动 |
浮动的元素会部分脱离文档流,具体来说,它有以下几个特征:
**首先,块级元素看不到浮动元素,会沉到浮动元素下层。**

```html
<div class="float-box"></div>
<div class="block-box"></div>
```
```css
.float-box { float: left; }
```
**然后,行内元素或行内块元素可以看到浮动元素,会环绕在浮动元素周围。**

```html
<div class="float-box"></div>
<span class="inline-box">啦啦啦</span>
```
```css
.float-box { float: left; }
```
**但是,浮动元素能看到文档流中的所有元素,避开块级元素排列到它们下方,避开行内元素排列到它们左侧或右侧。**

```html
<div class="block-box"></div>
<div class="float-box"></div>
```
```css
.float-box { float: left; }
```
除了脱离文档流之外,它还有下面一些其它特性:
- 如果没有为元素设置`width`,那么浮动元素会根据内容收缩
- 会自动拥有**行内块元素**的一些特性,可以定义宽度高度,也可以与其它元素排成一行
- `margin`的`auto`值无效
### 清除浮动
使用`clear`可以避免文档流中后面的**块级元素**沉到浮动元素下层,使其排列到浮动元素下方,而不是与它重叠在一起。也就是说,可以让块级元素看到浮动元素。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 不清除浮动 |
| `both` | 避开两侧的浮动元素,排列到它们下方 |

```html
<div class="float-box"></div>
<div class="block-box"></div>
```
```css
.float-box { float: left; }
.block-box { clear: both; }
```
### 高度塌陷
如果一个块级元素没有设置高度,并且只包含浮动元素,那么会导致它无法获得子元素高度,从而高度为`0`,这称为浮动元素引起的高度塌陷。
在下面的代码中,虽然浮动元素自身有高度,但是父元素的高度依然为`0`,导致后面的元素直接将该元素覆盖。
```html
<div class="container">
<div class="float-box"></div>
</div>
```
```css
.float-box {
float: left;
height: 200px;
}
```
使用`clear`可以解决这个问题,只需在父元素中通过伪元素`::after`追加一个清除浮动的**块级元素**即可:
```css
.container::after {
display: block; /* clear 只能应用于块级元素 */
content: ""; /* 伪元素必须有 content 才能生效 */
clear: both;
}
```
除此之外,也可以使用触发父元素[BFC](/posts/9obo9xpo.html)的方式避免高度塌陷。
## 定位
使用`position`设置元素定位。对于绝对定位和固定定位元素来说,它们会变成块级元素,并且即便[包含块](/posts/vv37590w.html)没有定义具体高度,`height: 100%`这样的属性也可以生效。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `static` |
| 可选值 | 描述 |
| --- | --- |
| `static` | 不定位 |
| `relative` | 相对定位,使元素相对于**原本位置**进行偏移,并且之前的位置依然会被**保留** |
| `absolute` | 绝对定位,使元素从文档流中被**完全移除**,它的位置仅受[包含块](/posts/vv37590w.html)以及偏移量影响 |
| `fixed` | 固定定位,与绝对定位的唯一不同是,它的[包含块](/posts/vv37590w.html)是浏览器窗口 |
### 偏移量
使用`top`、`right`、`bottom`、`left`设置定位元素的偏移量,即相对于[包含块](/posts/vv37590w.html)某个方向的距离。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 定位元素 | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 见[包含块](/posts/vv37590w.html)一节 |
| 长度值 | 设置具体的偏移量 |
| 百分比值 | 水平方向相对于[包含块](/posts/vv37590w.html)的`width`,垂直方向相对于[包含块](/posts/vv37590w.html)的`height` |
### 层叠位置
由于定位后的元素可能会相互重叠,使用`z-index`属性可以设置在同一个[层叠上下文](/posts/xsvc751j.html)中哪个元素显示在上层。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 定位元素 | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 如果只有一个[层叠上下文](/posts/xsvc751j.html)的话,相当于`0` |
| 整数值 | 数值越大,元素越靠近上层,可以为负数 |
## 弹性布局
弹性布局是 CSS3 新增的一种非常强大的布局方式,但是由于兼容性问题,目前主要使用在移动端。
使用`display: flex`或者`display: inline-flex`将元素变成**弹性容器**,其中的**直接**子元素则会自动变成**弹性项目**。
```html
<div class="box">
<div class="item">我是项目1号</div>
<span class="item">我是项目2号</span>
<p class="item">我是项目3号</p>
</div>
```
```css
.box { display: flex; }
```
虽然容器中的元素原本不同,但是现在它们都变成了项目,除了元素自带的样式外,它们都是一样的。这时项目的一些属性会失效,比如`float`、`vertical-align`或者`display: block`等。
容器有两根轴,分别为**主轴**和**交叉轴**,其中的项目会依次沿主轴排列,默认**从左到右**。将主轴**顺时针旋转90°**就得到了交叉轴,因此交叉轴默认**从上到下**。
### 主轴方向
使用`flex-direction`设置主轴方向,注意这会导致**交叉轴同时被旋转**。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `row` |
| 可选值 | 描述 |
| --- | --- |
| `row` | 从左到右 |
| `row-reverse` | 从右到左 |
| `column` | 从上到下 |
| `column-reverse` | 从下到上 |

### 主轴对齐
使用`justify-content`设置项目在主轴上的对齐方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `flex-start` |
| 可选值 | 描述 |
| --- | --- |
| `flex-start` | 左对齐(主轴的起点) |
| `flex-end` | 右对齐(主轴的末尾) |
| `center` | 居中对齐 |
| `space-between` | 两端对齐,项目之间的距离相等 |
| `space-around` | 项目两侧的距离相等 |

### 交叉轴对齐
使用`align-items`和`align-self`设置所有项目 / 单个项目在交叉轴上的对齐方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `stretch` |
| 可选值 | 描述 |
| --- | --- |
| `stretch` | 拉伸至容器的高度(交叉轴对应的方向)。如果项目设置了高度,则该属性值无效 |
| `flex-start` | 顶部对齐(交叉轴的起点) |
| `flex-end` | 底部对齐(交叉轴的末尾) |
| `center` | 居中对齐 |
| `baseline` | 基线对齐,以项目内的第一行文本为准 |

### 主轴剩余空间分配
如果项目的宽度总和不能填满主轴的宽度,那么多余的空间保留会被保留,使用`flex-grow`属性可以设置这些**剩余空间**如何分配。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | 表示分配给该项目的剩余空间比例。设置为`0`表示该项目不分配剩余空间,保持原来大小;所有项目均为同一个`>= 1`的数,那么表示所有项目**等分剩余空间**;如果它们不为同一个数,则按照数值的比例进行分配 |

如果项目的宽度总和超过了填满主轴的宽度,那么剩余空间为负,使用`flex-shrink`属性可以设置这些**负的剩余空间**如何分配。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `1` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | 表示分配给该项目的剩余空间比例。设置为`0`表示空间不足时该项目不会缩小自身,如果所有项目均设置为`0`,那么它们会超出容器;所有项目均为同一个`>= 1`的数,那么表示项目以相同比例缩小;如果它们不为同一个数,则按照数值的比例进行分配 |
注意,即使项目设置了宽度`width`,当剩余空间不足时,这些项目也会被缩小。如果不希望它们的宽度被改变,那么将其设置为`flex-shrink: 0`。
### 自动换行
使用`flex-wrap`设置当主轴空间不足时,项目自动换行。该属性的优先级比项目缩小要高,如果一行下不能容纳所有元素,那么会优先换行而不是缩小元素。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `nowrap` |
| 可选值 | 描述 |
| --- | --- |
| `nowrap` | 不换行 |
| `wrap` | 允许项目自动换行 |

### 多行对齐
使用`align-content`设置当发生换行时,行的排列方式。如果容器只有一行,或者不允许自动换行,那么该属性无效。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 弹性容器 | 不可继承 | `stretch` |
| 可选值 | 描述 |
| --- | --- |
| `stretch` | 拉伸到当前行高度最大项目的高度 |
| `flex-start` | 顶端对齐(交叉轴的起点) |
| `flex-end` | 底部对齐(交叉轴的终点) |
| `center` | 居中对齐 |
| `space-between` | 两端对齐,行之间的距离相等 |
| `space-around` | 行两侧的距离相等 |

## 表格布局
表格是 CSS 中类似于传统 HTML 表格的布局方式。使用`display`属性可以将元素设置为表格样式:
- `table`:块级表格,等同于`<table>`标签
- `inline-table`:行内块表格,等同于`<table>`标签,并设置为`inline-block`
- `table-row`:定义行,等同于`<tr>`标签
- `table-cell`:定义单元格,等同于`<td>`或`<th>`标签
### 表格边框模型
使用`border-collapse`设置表格的边框模型。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 表格元素 | 可继承 | `separate` |
| 可选值 | 描述 |
| --- | --- |
| `separate` | 分离边框模型,单元格之间彼此分开,产生一定的缝隙,每个单元格有自己独立的边框 |
| `collapse` | 折叠边框模型,单元格之间的缝隙完全清除,相邻的单元格共用一条边框 |
不同的边框模型会导致表格元素的部分盒模型属性失效:
| | 表格(分离) | 表格(折叠) | 行(分离) | 行(折叠) | 单元格(分离) | 单元格(折叠) |
| --- | --- | --- | --- | --- | --- | --- |
| `margin` | ✔️ | ✔️ | ❌ | ❌ | ❌ | ❌ |
| `border` | ✔️ | ✔️ | ❌ | ✔️ | ✔️ | ✔️ |
| `padding` | ✔️ | ❌ | ❌ | ❌ | ✔️ | ✔️ |
如果使用了折叠边框模型,但是相邻单元格或行的边框不同,那么会采用下面的规则决定边框的优先级:
- 如果边框为`border-style: hidden`,那么优先级最高
- 如果边框都是可见的,那么宽度更大的边框优先
- 如果边框宽度相同,那么按照边框样式排序,优先级由高到低为`double`、`solid`、`dashed`、`dotted`、`ridge`、`outset`、`groove`、`inset`、`none`
- 如果样式和宽度都相同,但颜色不同,那么优先级由高到低为单元格、行、列、表。如果两个元素都是单元格,那么位置更靠近**右下**的优先
### 单元格间距
使用`border-spacing`设置单元格的间距,即单元格边框到表格内边距,以及相邻单元格之间的距离。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 分离边框模型的表格元素 | 可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | 设置具体的距离,如果只有一个值则同时表示水平和垂直方向;如果有两个值,则前者表示水平方向,后者表示垂直方向 |
```css
table {
padding: 20px;
border-spacing: 20px;
}
```

### 空单元格
使用`empty-cell`设置空单元格的处理方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 分离边框模型的表格元素 | 可继承 | `show` |
| 可选值 | 描述 |
| --- | --- |
| `show` | 如果单元格为空,它的边框和背景依然会正常显示 |
| `hide` | 如果单元格为空,隐藏其边框和背景 |
### 宽度计算
使用`table-layout`设置表格宽度的计算方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 表格元素 | 可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 自动宽度布局,表格的宽度根据内容自动计算。此时表格的`width`相当于`min-width`,也就是说如果内容过多的话,浏览器会尽量将内容压缩,并忽略`width` |
| `fixed` | 固定宽度布局,表格的宽度与内容无关。如果表格的`width`为`auto`(默认值),那么浏览器会强制使用自动宽度布局 |
## 多列布局
多栏布局类似于报纸文章的排版,可以将子元素自动划分为多列显示。
### 列的数量
使用`column-count`设置列的数量,并且将元素设置为多列布局容器。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素 | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 由`column-width`来决定 |
| 整数值 | 表示列的数量 |
### 列宽
使用`column-width`设置列的**最小**宽度,并且将元素设置为多列布局容器。注意,该属性与`column-count`可能会产生矛盾,最好不要同时设置。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素 | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 由`column-count`来决定 |
| 长度值 | 表示列的最小宽度 |
### 列间距
使用`column-gap`设置列的间距。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 多列布局容器 | 不可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| `normal` | 相当于`1em` |
| 长度值 | 设置具体的列间距 |

### 列分隔线
使用`column-rule`设置列间隙上的分隔线。该属性是`column-rule-width`、`column-rule-style`和`column-rule-color`三个属性的简写属性,顺序没有要求。它的用法与边框基本一致,不再赘述。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 多列布局容器 | 不可继承 | `normal` |
```css
.container {
column-rule: 5px solid orange;
}
```

### 内容分割
使用`break-inside`阻止内容被截断。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 多列布局容器的**内部元素** | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 可能会将一块内容截断到下一列,以调整合适的高度 |
| `avoid` | 不截断元素 |
```css
.container > div {
break-inside: avoid;
}
```
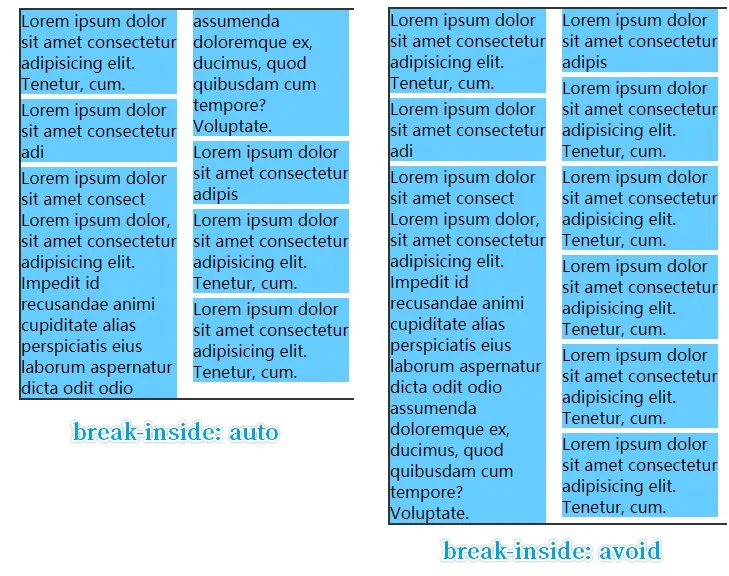<file_sep>---
title: 🐤 bluebird.js
date: 2020-01-10 22:32:43
abbrlink: 0wuyppfk
tags: JavaScript
categories:
- 大前端
- 第三方库
excerpt: 施工中...
---
# 🐤 bluebird.js
施工中,修改时间<file_sep>---
title: 大前端支线笔记之04 🌈 CSS 渐变
date: 2020-01-23 9:15:55
abbrlink: qvto1gxk
tags: CSS
categories:
- 大前端
- 支线
excerpt: 渐变的本质是图片,因此只要是允许使用图片路径<code>url()</code>的属性,都可以使用渐变。CSS 中的渐变分为两种,线性渐变和径向渐变。
---
# 大前端支线笔记之04 🌈 CSS 渐变
渐变的**本质是图片**,因此只要是允许使用图片路径`url()`的属性,都可以使用渐变。CSS 中的渐变分为两种,线性渐变和径向渐变。
## 线性渐变
使用`linear-gradient()`设置线性渐变,它可以包含下面三个参数,参数之间使用逗号隔开。
### 渐变方向
第一个参数是渐变方向,它的可选值为:
| 参数 | 描述 |
| --- | --- |
| `to top`/`to right`/`to bottom`/`to left` | 表示**朝某个方向**渐变 |
| 角度值 | `0deg`表示从元素的正下方开始渐变,然后顺时针旋转,度数随之增加,如`90deg`表示从左到右渐变 |
```css
div { background-image: linear-gradient(to right); }
```
### 色标
第二个参数是色标,分别表示渐变的起始色和终止色。
默认情况下,第一个色标的位置为`0%`,最后一个色标的位置为`100%`,色标所在的位置也就是该颜色(纯色)的位置。如下面左图中,元素的最右侧即`100%`的位置正好是正黄色。如果有多个色标,那么它们会**平均分布**(右图)。
```css
div {
background-image: linear-gradient(90deg, #66ccff, yellow); /* 两个色标 */
background-image: linear-gradient(90deg, #66ccff, yellow, orange); /* 三个色标 */
}
```

如果要修改色标的位置,可以在颜色值后面添加一个长度值或者百分比值(相对于元素大小)。
此时由于渐变并不足以铺满整个元素,因此到`50%`的位置就结束了,**剩余的位置以最后的色标来填充**。如果渐变的距离大于元素,那么多余的渐变也会被直接截掉。
```css
div {
background-image: linear-gradient(90deg, #66ccff, yellow 25%, orange 50%);
}
```

如果两个色标的位置相同,那么它们之间不会有渐变的效果。
```css
div {
background-image: linear-gradient(90deg, #66ccff, yellow 50%, orange 50%);
}
```

### 中色点
在两个色标参数之间还可以插入一个百分比值,作为第三个参数中色点。默认情况下,两种颜色之间的渐变,中间`50%`的位置正好是两个颜色的中点。如果设置了中色点,那么这个中间颜色就会向前 / 向后推移。
```css
div {
background-image: linear-gradient(90deg, #66ccff, 25%, orange);
}
```

## 径向渐变
径向渐变`radial-gradient()`的语法与线性渐变基本一致,但是不需要设置方向,取而代之的是形状、尺寸和位置。
### 形状
径向渐变的形状可选值为圆形`circle`和椭圆`ellipse`,默认与元素的形状一致。如果设置一个正方形的元素为椭圆形状的渐变,那么也不会看出效果,因为即便是椭圆,它的长和宽也是一样的。
```css
div {
width: 200px;
height: 100px;
background-image: radial-gradient(ellipse, orange, #333); /* 左图 */
background-image: radial-gradient(circle, orange, #333); /* 右图 */
}
```

### 尺寸
在形状的后面还可以添加长度值表示渐变的大小。1 个值同时表示水平和垂直方向,只能用于圆形;2 个值则前者表示水平,后者表示垂直方向,只能用于椭圆。
```css
div {
width: 200px;
height: 200px;
background-image: radial-gradient(100px, orange, #333); /* 左图 */
background-image: radial-gradient(50px 200px, orange, #333); /* 右图 */
}
```

### 位置
默认情况下,径向渐变圆心的位置为元素的中心,在尺寸值的后面(如果有的话)添加`at 位置`可以设置圆心的位置。位置参数与背景`background-position`属性的值完全一样,区别在于位置参数的默认值不是`0% 0%`,而是`center`。
```css
div {
width: 200px;
height: 200px;
background-image: radial-gradient(100px at left bottom, orange, #333);
}
```
<file_sep>---
title: 大前端支线笔记之06 😨 Node.js 原生构建网络服务
date: 2020-01-24 0:04:24
abbrlink: tb618gp3
tags: Node.js
categories:
- 大前端
- 支线
excerpt: 施工中
---
# 大前端支线笔记之06 😨 Node.js 原生构建网络服务
> 施工中,修改时间
### 创建服务器
下面的代码可以创建一个最简单的服务器:
```js
const http = require('http') // 引入模块
const app = http.createServer() // 创建服务器
app.listen(3000, function() { // 监听端口
console.log('服务器启动成功,正在监听 3000 端口')
})
```
### 接收请求
当服务器接收的请求时会触发服务器对象的`request`事件,Node.js 也是通过`on()`方法监听事件,该方法定义在`Events`模块中。
```js
app.on('request', (req, res) => { // req 和 res 分别为请求对象与响应对象
// 处理请求
})
```
请求行和请求头可以通过请求对象获取:
```js
app.on('request', (req, res) => {
req.method // 请求方式,值为 'GET' 或 'POST'
req.url // 请求 URL 的 path 部分,如 /index?id=233,使用 URL 模块可以提取出 GET 方式传递的参数
req.headers // 请求头的信息,使用方括号传入键名可以获取对应的值
})
```
请求体(比如`POST`方式传递的参数)的获取比较麻烦,因为它是通过流的方式传递的。
当流接收到新的缓存`Buffer`时会触发`data`事件,但是这时的内容可能并不完全,当流接收到全部内容时,会触发`end`事件。因此最好是在`data`事件中将缓存数组拼接起来,然后在`end`事件中输出:
```js
app.on('request', (req, res) => {
let reqBody = '' // 声明一个变量,表示请求体数据
req.on('data', temp => reqBody += temp) // 拼接获取的缓冲
req.on('end', () => reqBody) // 传输完毕后获取请求体
})
```
### 返回响应
当请求处理完毕后,应当给客户端返回响应,否则客户端会一直处于等待状态(浏览器转圈)。
不过在此之前,可以先通过`writeHead()`方法设置一下响应行和响应头。因为默认情况下,服务器响应的数据为 UTF-8 编码,但是如果没有在响应头中声明,或者在 HTML 文件中使用`<meta charset="UTF-8">`告知浏览器的话,那么浏览器会采用当前操作系统的编码即 GBK 来解析,会出现乱码问题。
> * 参数①:HTTP 状态码
> * 参数②:可选,响应头信息的键值对
> * 返回值:调用它的响应对象,可以链式编程
```js
app.on('request', (req, res) => {
res.writeHead(200, { // 第一个参数为响应行的状态码,第二个参数为响应头的键值对
'content-type': 'text/html;charset=utf8' // 设置响应头中的资源类型和编码
})
})
```
最后,通过`end()`方法为客户端返回响应,同时设置响应体。
> * 参数①:字符串类型的响应体,如果是复杂数据不要忘记转换成 JSON
```js
app.on('request', (req, res) => {
res.end('<h1>hello</h1>') // 响应内容
})
```
### 页面路由
然而此时无论用户访问任何地址,服务器都只能响应相同的内容。如果希望实现用户请求不同路径,返回对应内容,首先要根据请求对象的`url`解析出`pathname`部分(使用 url 模块),然后根据不同的`pathname`返回对应的内容。
下面的代码实现了简单的页面路由,可以看出,**用户请求的路径与实际返回的内容是完全没有关系的**,全凭服务器如何处理。
```js
const url = require('url')
app.on('request', (req, res) => {
// req.url 是包含参数的,如 /index?id=233,使用 parse() 可以提取出 /index 部分
let pathname = url.parse(req.url).pathname
// 根据请求路径返回对应内容
switch (pathname) {
case '/':
case '/index':
res.end('欢迎来到主页')
break
case '/list':
res.end('欢迎来到列表页')
break
default:
res.end('您访问的页面不存在')
break
}
})
```
> *参考资料*
> - 《深入浅出 Node.js》<file_sep>---
title: 🔐 MongoDB
date: 2020-02-09 13:57:38
abbrlink: e6q2910w
tags: MongoDB
categories: 数据库
excerpt: 施工中...
---
# 🔐 MongoDB
> 施工中... 修改时间
MongoDB 与传统的关系型数据库区别在于,它不需要设计表结构,存取数据更加灵活。它保存的不是表,而是一种称为 BSON 的数据结构,它与 JSON 非常相似,但是比 JSON 强大的一点在于,它可以保存二进制数据,其它与 JSON 区别不大。正是由于这一点,MongoDB 的一些概念与关系型数据库有所不同:
| SQL | MongoDB | 描述 |
| --- | --- | --- |
| database | database | 数据库 |
| table | collection | 表 / 集合 |
| row | document | 行 / 文档 |
## 安装与配置
在[官网](https://www.mongodb.com/download-center/community)可以下载社区版(非企业级商用)的 MongoDB,安装比较简单,在自定义安装下可以修改路径,除此之外没有其它配置。安装的过程中会提示安装 MongoDB Compass 图形化界面,但是这个并不好用,建议直接使用 Navicat(最新版本支持 MongoDB)。
安装完成后,将安装路径下的`bin`路径配置到环境变量中(如`C:\MongoDB\Server\bin`),以便命令行调用。
系统服务下的`MongoDB Server`为 MongoDB 的服务,在使用 MongoDB 之前记得先启动它。
开启 CMD 输入`mongo`即可进入 MongoDB 的命令行工具。在命令行工具中输入`show databases`查看所有的数据库,如果能正常显示,则说明连接成功。
安装目录下的`data`文件夹保存了数据库文件,如果要修改保存位置,需要打开`bin\mongod.cfg`,修改其中的`dbPath`一项。该文件是数据库的配置文件,很多选项都可以在这里修改。
## 操作数据库和集合
使用`use 数据库名`可以切换当前使用的数据库。注意,即使这个数据库不存在,也是可以直接切换的。因为 MongoDB 中的**数据库和集合不需要手动创建**,在第一次向集合中插入文档时,对应的数据库和集合会自动创建。
```js
use test
```
使用`show collections`可以查看当前数据库中的所有集合。由于数据库都还没有创建,因此此时的数据库肯定也是空的。
## CRUD
使用`db.集合名`可以调用下列方法,进行数据库的 CRUD 操作。其中`db`指向当前数据库名,类似于`this`;而集合名类似于表名,可以自定义。注意,这些方法仅适用于控制台,而不是供后端语言调用。
### 查询操作符
[官方文档](https://docs.mongodb.com/manual/reference/operator/query/)
```js
// 查询 name 为 'Enter the Gungeon' 的文档
Game.find({ name: 'Enter the Gungeon' })
// 查询 name 为 'Resident Evil' ,且 type 为 'AVG' 的文档
Game.find({ name: 'Resident Evil', type: 'AVG' })
// 查询 age 大于 15 小于 30 的文档
User.find({ age: { $gt: 15, $lt: 30 }})
// 查询 age 不等于 20 的文档
User.find({ age: { $ne: 20 }})
// 查询 age 为 10 或 14 或 20 的文档
User.find({ age: { $in: [10, 14, 20] }})
// 查询全部文档
Game.find()
```
### insert()
向集合中插入若干条文档。
* 参数①:对象或数组,表示插入的数据
```js
// 插入一条文档
db.students.insert({ name:'御坂美琴', age: 16 })
// 插入多条文档
db.students.insert([
{ name: 'JOJO', age: 17 },
{ name: 'Wendy', age: 14 }
])
```
此时使用`show databases`和`show collections`就可以看到自动创建的数据库和集合了。
注意,可以看到每一条文档前自动添加了一个`_id`字段。它用来作为文档的唯一标识,但是其类型为比较特殊的`ObjectId`,该类型并不是普通的字符串,而是一个自定义对象,因此在其它语言查询数据库时,需要额外的处理。
### find() / findOne()
查询集合中的一条 / 多条文档。
* 参数①:对象,表示查询条件
* 返回值:以数组 / 对象形式返回符合条件的文档
```js
// 查询所有文档
db.students.find()
// 查询符合单个条件的文档,注意这里根据 id 查询数据的方式
db.students.find({ _id: ObjectId('5de238bbb52000002c000d74') })
// 查询同时符合多个条件的文档
db.students.find({ name:'JOJO', age: 17 })
// 查询第一条符合条件的文档
db.students.findOne({ age: 17 })
```
如果使用返回的结果继续调用`count()`,可以获取查询的结果数。
```js
db.students.find().count() // => 5
```
### updateOne() / updateMany()
更新文档需要使用**更新操作符**,它用来对本次更新操作进行一些配置,例如可以更新字段名、可以更新字段值或者移除一个字段等。
修改集合中的一条 / 多条文档。
> * 参数①:对象,表示查询条件
> * 参数②:包含更新操作符的对象
```js
// 将第一条 age 为 14 的数据的 age 属性值更新为 23
db.students.updateOne({ age: 14 }, {
$set: { age: 23 } // $set 操作符用来更新属性值
})
```
### deleteOne() / deleteMany()
删除集合中的一条 / 多条文档。
> * 参数①:对象,表示查询条件
```js
// 将 _id 为 5de238bbb52000002c000d74 的文档删除
db.students.deleteOne({ _id: ObjectId('5de238bbb52000002c000d74') })
```<file_sep>---
title: 📄 IEEE 754
date: 2020-01-30 11:49:55
abbrlink: xp50sskp
categories: 杂七杂八
excerpt: IEEE 754 是美国电气电子工程师学会(IEEE)在 1985 年发布的一个技术标准,它定义了浮点型的有限数值、无限数值和特殊值 NaN 的二进制存储格式。
---
# 📄 IEEE 754
IEEE 754 是美国电气电子工程师学会(IEEE)在 1985 年发布的一个技术标准,它定义了浮点型的有限数值、无限数值和特殊值`NaN`的二进制存储格式。
## 存储格式
以一个 64 位的数据类型(比如 Java 中的`double`或者 JavaScript 中的`Number`)为例,它在内存中的存储格式如下:

然后,对于任意一个数值,它都可以换算成下面的格式,以二进制形式存储:
$$ \Large (-1)^s×2^q×c $$
### 符号位
$s$表示符号,直接对应内存中的符号位,只能为`0`或者`1`。可以看到,当符号位为`0`时,该数值为正数($(-1)^0$为`1`),反之则为负数。
### 指数位
$q$表示指数,对应内存中的指数位。通过内存可以看出,它的表示范围最小是全部 11 个指数位取`0`时,最大是全部位取`1`时,即$[0,2047]$($[0,2^{11}-1]$)之间。
但是,指数有正负之分,因此这里设置了一个**偏移量(bias)**,用来移动表示范围。它的计算公式为$bias=2^{k-1}-1$,其中$k$为指数位的位数,是固定值。
- 当数据格式为 64 位时,$k=11$
- 当数据格式为 32 位时,$k=8$
这样将表示范围与偏移量相减,相当于将表示范围的数轴向左移动了一半的距离,从而出现正负指数各一半的情况。此时的$bias=2^{11-1}-1=1023$,因此指数$q$实际的表示范围在$(-1023,1024)$(全`0`和全`1`有另外的作用),即$[-1022,1023]$之间,对应的指数位分别是$00000000001$和$11111111110$。
### 尾数位
$c$表示尾数,对应内存中的小数位,由 1 个隐藏位和 52 个其它位组成。由于指数位只能取整数,因此必须要尾数位才能真正的表示小数。
隐藏位用来表示尾数位是否大于`1`,直接当作十进制即可。也就是说,如果该位为`0`,那么尾数位一定是`0.x`,如果该位为`1`,那么尾数位一定是`1.x`。
其它位则是通过**定点数表示法**转换成十进制小数值,从左到右依次为$2^{-1}$、$2^{-2}$,依次类推。例如,隐藏位为`1`,其它位为`011000...`,转换后为$0×2^{-1}+1×2^{-2}+1×2^{-3}=0.375$,考虑隐藏位之后,尾数位最终为`1.375`。
### 特殊值
到目前为止,还有几个特殊值没法表示。因此使用尾数位结合之前指数位预留的两种情况(全`0`和全`1`),来表示这些值:
- 指数位全是`0`,尾数位全是`0`,表示`±0`(因为符号位还在)
- 指数位全是`0`,尾数位不全是`0`,表示**非规格化的值**,此时指数位的全`0`表示十进制的最小值`-1022`
- 指数位全是`1`,尾数位全是`0`,表示$\pm\infty$
- 指数位全是`1`,尾数位不全是`0`,表示`NaN`
> *参考资料*
> * [Java 与 IEEE754 浅浅谈](https://blog.csdn.net/u011679955/article/details/52750345)
> * [细说 JavaScript 七种数据类型](https://www.cnblogs.com/onepixel/p/5140944.html)
> * [IEEE 754 Calculator](http://weitz.de/ieee/)<file_sep>---
title: 大前端支线笔记之03 💠 CSS 层叠上下文
date: 2020-01-23 7:17:55
abbrlink: xsvc751j
tags: CSS
categories:
- 大前端
- 支线
excerpt: 与 BFC 类似,某些元素也具有层叠上下文(Stacking Context)的特性,在同一个层叠上下文中的元素,会按照下列顺序层叠在一起。
---
# 💠 CSS 层叠上下文
与 BFC 类似,某些元素也具有层叠上下文(Stacking Context)的特性,在同一个层叠上下文中的元素,会按照下列顺序层叠在一起。

> 注意,其中**负数**的`z-index`定位元素甚至比文档流中的元素位置还要低。
要使一个元素拥有层叠上下文特性,只需满足下列条件之一:
* 根元素`<html>`
* `z-index`不为`auto`
* `opcacity`的值小于`1`
* `transform`的值不为`none`
* `perspective`的值不为`none`
在下面的代码中,`.a`元素没有定位,`.a-child`元素使用了绝对定位,并设置了`z-index: 1`,`.b`元素也设置了绝对定位,但是没有设置层叠顺序,因此它们的位置应该是`.a-child`最高,中间是`.b`,最底层是`.a`。(左图)
```html
<div class="a">
<div class="a-child"></div>
</div>
<div class="b"></div>
```
```css
.a-child {
position: absolute;
z-index: 1;
}
.b {
position: absolute;
z-index: 0;
}
```
但是,如果`.a`设置了`transform: scale(1)`,那么它就会创建一个**独立的层叠上下文**。此时,由于`.a`的层级不如`.b`高(`.b`有绝对定位,而`.a`没有定位),那么无论`.a-child`层级有多高,也无法摆脱`.a`的限制。也就是说,`.a-child`设置的`z-index`只能与`.a`的其它子元素相比,而无法与整个文档的元素相比了。(右图)
```css
.a {
transform: scale(1);
}
```
<file_sep>---
title: 大前端笔记之11 😄 JavaScript 基本概念
date: 2020-01-27 10:51:09
abbrlink: pk33pcs4
tags: JavaScript
categories: 大前端
excerpt: JavaScript 的主要作用是赋予页面行为。如今经常用 JavaScript 和 ECMASrcipt 表达相同的含义,但实际上 JavaScript 表示的更加广泛:它由三个部分组成,分别是 ECMAScript、DOM 和 BOM。
---
# 大前端笔记之11 😄 JavaScript 基本概念
JavaScript 的主要作用是赋予页面行为。如今经常用 JavaScript 和 ECMASrcipt 表达相同的含义,但实际上 JavaScript 表示的更加广泛:它由三个部分组成,分别是 ECMAScript、DOM 和 BOM。
ECMAScript 定义的是这门语言的基础,包括语法、类型、语句、关键字、保留字、操作符等,而浏览器只是 ECMAScript 的**宿主环境**之一。宿主环境不仅提供 ECMAScript 的实现,同时也会提供 ECMAScript 的 API,以便**ECMAScript 与宿主环境交互**,其它的宿主环境还有 Node.js、Adobe Flash 等。
到 2019 年为止,它总共经历了 10 个版本,目前最新的是 2019 年 6 月发布的 ECMAScript10。其中变革最大的为 2015 年发布的 ECMAScript6,由于后续版本更新速度加快,新特性并不是很多,因此以 ES6 指代后续所有的新版本。接下来为了方便起见,也通常使用 JavaScript 来泛指 ECMAScript。
DOM(Document Object Model,文档对象模型)是浏览器提供的 API,以便 ECMAScript 与浏览器交互。它将整个页面映射成一个多层节点的结构,其中每个部分都是某种类型的节点,这些节点又包含不同类型的数据,通过 DOM 可以轻松地删除、添加、替换或修改任何节点。
BOM(Browser Object Model,浏览器对象模型)也是浏览器提供的一个 API。通过 BOM 可以控制浏览器视口以外的部分,比如浏览器的窗口和框架等等。
## 加载 JavaScript
使用`<script>`的`src`属性可以引入外部的 JavaScript 文件:
```html
<html>
<head></head>
<body>
<!-- 这里是页面内容 -->
<script src="test.js"></script>
</body>
</html>
```
> 按照传统做法,所有`<script>`元素都应放在`<head>`元素中。但是,这意味着必须等到全部 JavaScript 都被下载、解析和执行完毕后,页面的内容才会呈现,这样用户在打开页面时就会看到长时间的空白。因此通常把 JavaScript 的引用放到`</body>`之前。
也可以在页面中的任何位置添加`<script>`并将代码直接写入其中:
```html
<html>
<head></head>
<body>
<script>
alert('hello')
</script>
</body>
</html>
```
## 语句和表达式
语句是为了执行某个任务而进行的操作,比如赋值、判断等。在 JavaScript 中多数语句以分号`;`结尾,而且**分号可以省略**,省略与否取决于个人习惯。
```js
var a = 3 /* 这是一条语句,执行了将 3 赋值给变量 a 的操作 */
```
> 如果省略了分号,那么切记在`[`、`(`、`+`、`-`和反引号(模板字符串标记)作为首行开头时,在前面添加一个`;`,以防解析出错。
表达式是可以得到计算结果的式子,它一定会返回一个值。也就是说,只要预期为值的地方,都可以替换成表达式。
```js
var a = 1 + 2 /* 将原本的值 3 替换成了表达式 1 + 2 */
```
## 变量
变量是用来保存值的,相当于为值起一个名字,以便识别和重复使用。JavaScript 中的变量是**弱类型**的,任何[数据类型](/posts/p4xy9gkd.html)的值都可以使用`var`关键字声明的变量保存。
```js
var msg = "hello"
```
也可以在一条语句中同时声明多个变量:
```js
var msg = "hello",
num = 233,
isIdiot = true
```
> `var`也可以省略,但是不推荐这样做。因为正常情况下,函数中声明的变量是局部变量。假如省略了`var`关键字,那么它就不再是一个局部变量,而会成为一个**全局变量**,这会导致代码难以阅读和维护。
## 标识符
标识符是变量和函数的名称,它们的名称需要遵循一系列命名规则,不符合规则的标识符会导致程序报错,无法继续执行。
- 由字母、数字、美元符号`$`和下划线`_`组成,不允许以数字开头
- 不能使用[保留字](http://www.ecma-international.org/ecma-262/9.0/index.html#sec-reserved-words)(页面比较大,耐心等待锚点跳转)、`true`、`false`、`null`
- 区分大小写
- 建议使用驼峰命名法
## 注释
注释是 JavaScript 引擎忽略的部分,它们不会被解析,可以用来对程序进行说明。其中单行注释以双斜线`//`开头,多行注释以`/*`开头,以`*/`结尾。
```js
// 这是一条单行注释
/*
这是一条多行注释
*/
```
## 运算符
运算符用来对值进行处理,从而得到新的值。
### 赋值运算符
赋值运算符会将右侧的运算数赋值给左侧的运算数。
| 运算符 | 运算符 | 运算符 |
| --- | --- | --- |
| 赋值`=` | 加赋值`+=` | 减赋值`-=` |
| 乘赋值`*=` | 除赋值`/=` | 模赋值`%=` |
为了简化代码,JavaScript 对一些常用的算数并赋值的运算进行了合并,称为**复合赋值运算符**。
```js
num = num + 2 // 相当于
num += 2
```
赋值运算符可以连续调用,执行顺序是从右向左。但是这样的用法会带来很多意料之外的问题,建议不要使用。
```js
var a = b = c = 2 // 相当于
var a
c = 2
b = c
a = b
```
### 算数运算符
算数运算符用来完成基本的算术运算。
| 运算符 | 运算符 | 运算符 | 运算符 | 运算符
| --- | --- | --- | --- | --- |
| 加法运算符`+` | 减法运算符`-` | 乘法运算符`*` | 除法运算符`/` | 指数运算符`**` |
| 取余运算符`%` | 自增运算符`++` | 自减运算符`--` | 一元正号`+` | 一元负号`-` |
#### 加法运算符
与其它算术运算符有所不同的是,如果其中一个运算数为字符串,那么就会将其它的运算数通过`String()`转换为字符串,表示字符串拼接;否则,就将所有运算数转换为数值,表示数学运算。(见[数据类型](/posts/p4xy9gkd.html)一节)
```js
1 + 2 // => 3
1 + false // => 1
null + false // => 0
'hello' + 1 // => hello1
'hello' + false // => hellofalse
```
#### 自增减运算符
自增减运算符根据相对于运算数的位置,含义有所不同。当它放在运算数左侧时,表示**先自增,后取值**;反之,当它放在运算数右侧时,表示**先取值,后自增**。
```js
var num = 8
console.log(num++) // => 8
var num = 8
console.log(++num) // => 9
```
注意,自增减运算符**只能操作变量**,不能操作值。
### 比较运算符
| 运算符 | 运算符 | 运算符 | 运算符 |
| --- | --- | --- | --- |
| 相等`==` | 不相等`!=` | 严格相等`===` | 严格不相等`!==` |
| 大于`>` | 大于等于`>=` | 小于`<` | 小于等于`<=` |
#### 严格相等
JavaScript 中的相等运算分为普通相等和严格相等两种。(见[数据类型](/posts/p4xy9gkd.html)一节)
普通相等运算的规则如下:
- 如果运算数类型相同,就直接进行比较
- 如果运算数类型不同,则尝试先转换成相同的类型,再进行比较:
- 比较数字和字符串时,会将字符串转换为数字,再比较两个数字
- 比较布尔值和其它类型时,会先将布尔值转换为数字,再比较数字和其它类型
- 比较`null`和`undefined`时,结果是`true`
使用严格相等运算时,只有两个运算数的类型和值都相等,才返回`true`。
```js
99 == '99' // => true
99 === '99' // => false
```
当比较两个对象时,`==`和`===`效果相同,即比较两个引用所指向的对象是否是同一个。如果是同一个对象,就返回`true`,否则返回`false`。
### 逻辑运算符
下面的逻辑运算符会导致运算数自动调用`Boolean()`进行[数据类型](/posts/p4xy9gkd.html)转换。
| 运算符 | 运算符 | 运算符 |
| --- | --- | --- |
| 逻辑与`&&` | 逻辑或`\|\|` | 逻辑非`!` |
#### 短路运算
逻辑与`&&`和逻辑或`||`运算符会从左到右依次判断每个条件,如果左侧的某个条件已经确定了整个运算的结果,那么后面的条件就不再执行(即使发生语法错误也不会报错)。
```js
10 < 5 && Math.sqrt(-1) // false
10 > 5 || Math.sqrt(-1) // true
```
## 流程控制
程序默认都是从上向下依次执行的,使用流程控制语句则可以改变顺序执行结构。流程控制语句总共有两类,分支结构和循环结构。
### 分支结构
分支结构可以使程序根据情况作出判断,从多条路线中选择一条继续执行,而忽略其它路线。
#### if 结构
最简单的`if`结构由`if`关键字、圆括号(表示判断条件)和花括号(表示条件满足时执行的操作)组成。判断条件可以是布尔型的值或者表达式,如果不是,则会自动调用`Boolean()`进行[数据类型](/posts/p4xy9gkd.html)转换。
```js
if (age > 18) {
console.log('可以进网吧啦')
}
```
使用`else`可以表示当条件不满足时,所执行的操作。
```js
if (age > 18) {
console.log('可以进网吧啦')
} else {
console.log('一边玩去!')
}
```
多个`if else`结合在一起使用,可以表示多次判断。注意,JavaScript 中没有`elseif`这个关键词。
```js
if (age < 18) {
console.log('一边玩去!')
} else if (age >= 18 && age < 50) {
console.log('可以进网吧啦')
} else if (age >= 50) {
console.log('啥?你是来找网瘾少年的吗?')
}
```
#### 三元运算符
三元运算符`判断条件 ? 表达式1 : 表达式2 `是`if else`的简写形式,它表示如果`条件表达式`满足,则返回`表达式1`,否则返回`表达式2`。
```js
console.log(age >= 18 ? '可以进网吧啦' : '一边玩去!')
```
#### switch 结构
如果要判断是否为一个具体的值,而非某个范围,那么使用`switch`可以使结构更加清晰。
在下面的代码中,根据`fruit`的值不同从而执行不同的操作。每个可能的值使用`case`表示,冒号后面为满足该值时执行的操作。注意,每个`case`之后都使用了`break`语句,否则当其中一个`case`满足时,会连带后续的所有`case`一并执行,发生所谓的穿透现象。最后,可以使用`default`表示当所有`case`均不满足时执行的操作。此外,如果有多个`case`的值执行相同操作,那么可以连续使用`case`。
```js
var fruit = 'orange'
switch (fruit) {
case '橘子':
case 'orange':
console.log('橘子五毛一斤')
break
case 'banana':
console.log('香蕉一块一斤')
break
default:
console.log('没这东西!')
}
```
注意,JavaScript 中的`switch`结构比较结果采用的是**严格相等**,因此比较时不会发生类型转换。
### 循环结构
循环结构可以使程序根据条件反复执行同一段操作,直到条件不成立为止。
#### while 结构
`while`循环由两部分组成,圆括号中的循环条件和花括号中的循环体。
```js
var i = 1
while (i <= 10) {
console.log('这是第' + i + '次循环啦')
i++ /* 改变循环条件,否则会出现死循环 */
}
```
#### for 结构
如果有明确的循环次数,那么可以使用更加方便的`for`循环。
```js
for (var i = 1; i <= 10; i++) {
/* 执行 10 次相应代码 */
}
```
#### 跳出循环
使用`break`和`continue`语句可以跳出循环。区别在于,`break`会跳出整个循环体,而`continue`是跳出本次循环,继续下一次循环。
```js
for (var i = 1; i <= 5; i++) {
if (i == 3) {
console.log('这个包子有毒!不吃了!吃下一个!')
continue
}
console.log('我正在吃第' + i + '个包子。')
}
```
```js
for (var i = 1; i <= 5; i++) {
if (i == 3) {
console.log('这个包子有毒!去拉肚子了!剩下的不吃了!')
break
}
console.log('我正在吃第' + i + '个包子。')
}
```<file_sep>---
title: 大前端笔记之07 🎨 CSS 背景
date: 2020-01-23 0:53:12
abbrlink: 86mehcq2
tags: CSS
categories: 大前端
excerpt: 元素可以通过下面的属性设置背景颜色或图片。
---
# 大前端笔记之07 🎨 CSS 背景
元素可以通过下面的属性设置背景颜色或图片。
## 背景颜色
使用`background-color`设置元素的背景颜色。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `transparent` |
| 可选值 | 描述 |
| --- | --- |
| `transparent` | 透明背景 |
| 颜色值 | 设置具体的背景颜色 |
## 背景图片
使用`background-image`设置元素的背景图片。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 没有背景图片 |
| `url()` | 图片的 URL |
| `linear-gradient()` | 设置[渐变](/posts/qvto1gxk.html)背景 |
如果同时设置了背景颜色和图片,那么**图片永远在最上层**,只有图片没有覆盖的区域才会显示为背景色。
如果要同时添加多个背景图片,只要将路径使用逗号隔开即可。此时**最左边的图片层叠优先级最高**,向右依次递减:
```css
div { background-image: url("bg01.png"), url("bg02.gif"); }
```
## 背景平铺
使用`background-repeat`设置背景的平铺方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `repeat` |
下面的四个值可以出现最多两次,以空格隔开,分别设置水平和垂直方向的平铺方式。如果省略了其中一个,表示该值同时作用于两个方向。
| 可选值 | 描述 |
| --- | --- |
| `repeat` | 在水平和垂直方向平铺 |
| `no-repeat` | 不平铺 |
| `space` | 根据元素的宽高自动调整背景平铺的数量,图片不会被裁切或压缩,多余的空间会被平分为图片的间隔。例如,背景图片宽为`300px`,元素宽度为`800px`,那么会显示两张图片,其余的`200px`作为背景的间隔 |
| `round` | 根据元素的宽高自动调整背景平铺的数量,图片会被压缩,因此不会产生多余空间。例如,背景图片宽为`300px`,元素宽度为`800px`,那么会显示三张图片,每张图片宽度调整为`266.67px` |
下面的两个值只能出现一次,表示仅在某个方向平铺,而另外一个方向不平铺。
| 可选值 | 描述 |
| --- | --- |
| `repeat-x` | 仅在水平方向平铺 |
| `repeat-y` | 仅在垂直方向平铺 |
```css
div {
background-repeat: repeat no-repeat;
/* 相当于 */
background-repeat: repeat-x;
}
```
## 背景位置
使用`background-position`设置背景在元素中的位置。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0% 0%` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | 表示以元素**左上角**为起始点的偏移量,可以为负值 |
| `top`/`right`/`bottom`/`left`/`center` | 表示背景紧靠该方向 |
| 百分比值 | 相对的值为**元素的宽高 - 背景的宽高**,因此左上角为`0% 0%`,右下角为`100% 100%`,居中则是`50% 50%` |
上述可选值根据数量不同,表示的含义也不同:
| 值的数量 | 描述 |
| --- | --- |
| 1 个长度值 | 该值表示水平方向,垂直方向为`center` |
| 2 个长度值 | 前者表示水平方向,后者表示垂直方向 |
| 1 个关键字 | 表示背景紧靠该方向,另外一个方向为`center` |
| 2 个关键字 | 分别表示水平和垂直方向,顺序并不重要,只要方向不冲突即可 |
| 1 个关键字 + 1 个长度值 | 前者表示水平方向,后者表示垂直方向。如果前面的值不为水平方向关键字,后面的值不为垂直方向关键字,那么该值无效 |
| 4 个值 | 1 个关键字和 1 个长度表示**与这个方向的距离** |
```css
div {
background-position: 10px; /* 距离左侧 10px,垂直方向居中 */
background-position: 10px 20px; /* 距离左侧 10px,距离上方 20px */
background-position: bottom; /* 垂直方向位于底部,水平方向居中 */
background-position: bottom left; /* 位于左下 */
background-position: left 10px; /* 水平方向位于左侧,垂直方向距离上方 10px */
background-position: right 10px bottom 5px; /* 距离右侧 10px,距离底部 15px */
}
```
## 背景固定
使用`background-attachment`设置背景相对于浏览器窗口定位。此时只有当元素位于背景的位置时,背景才能显示出来。即使两个背景的位置重合也没有关系,只要元素覆盖到自己的背景之上,那么背景就会显示出来。
注意,该属性会导致`background-position`相对于浏览器窗口设置位置,而非元素本身。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `scroll` |
| 可选值 | 描述 |
| --- | --- |
| `scroll` | 背景跟随元素滚动 |
| `fixed` | 背景固定,将元素背景以浏览器窗口作为参照物,而不是元素 |
## 背景区域
使用`background-clip`设置背景的**显示区域**。如果一个元素包含`padding`,而显示区域设置为`content-box`,那么背景会显示不全。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `border-box` |
| 可选值 | 描述 |
| --- | --- |
| `border-box` | 背景显示在`border`、`padding`和`content`区域 |
| `padding-box` | 背景显示在`padding`和`content`区域 |
| `content-box` | 背景显示在`content`区域 |
使用`background-origin`设置背景的**定位区域**,它可以修改图片渲染的**起始点**,但是不代表其它部分不渲染,只是起始点变了而已。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `padding-box` |
| 可选值 | 描述 |
| --- | --- |
| `padding-box` | 从`padding`区域开始绘制 |
| `border-box` | 从`border`区域开始绘制 |
| `content-box` | 从`content`区域开始绘制 |

## 背景尺寸
使用`background-size`设置背景尺寸。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `auto auto` |
下面的值可以出现一至两次,以空格隔开,分别设置水平和垂直方向。如果只有一个值,那么它表示水平方向,垂直方向自动设置为`auto`。
| 可选值 | 描述 |
| --- | --- |
| `auto` | 使用图片本身的尺寸,如果其中一个方向尺寸被改变了,那么`auto`表示等比例缩放 |
| 长度值 | 设置固定的背景尺寸 |
| 百分比值 | 相对于`background-origin`区域的宽度或高度 |
| `cover` | 表示背景图片在**保持比例**的前提下填充满元素,超出的部分不会显示 |
| `contain` | 表示背景图片在**保持比例**的前提下完整的显示,多余的部分以空白填充 |
## 背景属性简写
使用`background`简写上述所有属性。将这些值以空格隔开,它们的顺序并不重要,也可以随意省略。不过注意`background-position`必须与`background-size`用`/`隔开。
```css
div {
background:
orange /* background-color */
url("img.png") /* background-image */
no-repeat /* background-repeat */
fixed /* background-attachment */
center center / 50% /* background-position / background-size */
content-box /* background-origin */
content-box; /* background-clip */
}
```<file_sep>---
title: 大前端笔记之20 ⚙️ Node.js
date: 2020-02-09 13:57:38
abbrlink: o48l9v2o
tags: Node.js
categories: 大前端
excerpt: Node.js 是一个类似于 Chrome V8 引擎的 JavaScript 运行环境,它使得 JavaScript 可以工作在服务器端,实现操作文件、构建网络服务等功能。
---
# 大前端笔记之20 ⚙️ Node.js
Node.js 是一个类似于 Chrome V8 引擎的 JavaScript 运行环境,它使得 JavaScript 可以工作在服务器端,实现操作文件、构建网络服务等功能。
## 安装与配置
首先在[官网](https://nodejs.org/en/download/)选择对应的版本下载安装。然后打开 CMD 输入`node -v`命令,如果安装成功,则会显示当前 Node.js 的版本号。
> Node.js 安装版会自动配置环境变量,如果没有,则需要自行配置。
新建一个 JavaScript 文件(如`app.js`),在其中写入 JavaScript 代码。保存后,在 CMD 输入`node app.js`,即可执行该文件。可见,虽然现在的宿主环境由之前的浏览器变成了 Node.js,但是 ECMAScript 的基本语法与之前是完全一样的,只不过缺少了 DOM 和 BOM 等浏览器提供的接口,取而代之的是 Node.js 提供的服务器相关接口。
## 模块化
虽然 [ES6](/posts/j5h1kgw7.html#模块化) 已经提出了模块化相关的功能,但是 Node.js 在创建早期就已经实现了自己特有的模块化方式 CommonJS,并且由于历史原因,即使是最新版本(v12.14.1)的 Node.js 也无法原生支持 ES6 模块化的语法,必须通过 Babel 转换后才可以使用,因此这里依然需要使用 CommonJS。
与 ES6 模块化一样,每个单独的 JavaScript 文件被称为一个模块。
### 导入模块
首先,使用`require()`方法可以导入一个模块并**自动执行其中的代码**。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 模块路径或名称 |
| 返回值 | 描述 |
| --- | --- |
| 对象 | `module.exports`对象,用于接收模块导出的变量,见下文 |
根据传入的参数不同,查找模块的方式也略有不同。
如果传入的是一个路径,则:
- 如果是完整路径,则直接查找该文件
- 如果省略了后缀名,那么会先查找同名的`.js`文件,再查找同名文件夹
- 如果找到了文件夹,则找其中的`index.js`文件
- 如果没有找到,则找其中的`package.json`文件,寻找`main`属性配置的入口文件
- 如果没有找到配置文件或入口文件,则报错
如果传入的是一个名称,那么引擎会认为这是一个系统模块或第三方模块,并在当前的`node_modules`文件夹中先查找同名文件,再查找同名文件夹,之后的步骤与上面一样。
因此,如果要加载的是当前路径下的模块,必须要传入`./`表示当前路径,否则会被识别成模块名。
```js
// hello.js
require('./includes.js') // 加载自定义模块
```
### 导出属性和方法
模块之间通过`module.exports`对象进行信息传递,它默认是一个空对象`{}`,可以为其添加属性和方法,如果只有一个要导出的值,也可以直接将其覆盖。
```js
// includes.js
const msg = '这是要传递的信息啦'
module.exports = msg // 覆盖了 module.exports 对象
module.exports.msg = msg // 或者将变量添加到 module.exports 对象中
```
通过`require()`方法的返回值可以获取`module.exports`对象,从而调用其中的属性或方法:
```js
// hello.js
var includes = require('./includes') // 引入模块
// 如果是直接导出的,那么该对象就是导出的值
includes // => '这是要传递的信息啦'
// 如果属性和方法在对象中,那么也要先取出才能访问
includes // => { msg: '这是要传递的信息啦' }
includes.msg // => '这是要传递的信息啦'
```
## 系统模块
Node.js 将很多系统功能也封装成了单独的模块,需要时再加载即可。
### 全局属性和方法
与浏览器环境不同,Node.js 中的全局对象不再是`window`,而是`global`。它也包含类似于定时器的全局方法,注意不要与`window`混淆。
#### __dirname
当前模块的物理路径。注意它有单独的实现方式,并不是位于`global`中,但是可以在全局任意位置直接调用。
```js
// 当前文件路径为 D:\htdocs\test.js
__dirname // => D:\htdocs
```
### Path
`Path`模块提供了与路径相关的功能。
```js
const path = require('path')
```
#### join()
拼接路径。将需要拼接的部分作为参数传入,它会自动忽略掉每个部分两端的路径分隔符,并使用**当前系统环境的分隔符**将其拼接起来。
| 参数 | 描述 |
| --- | --- |
| 字符串若干 | 要拼接的文件夹、文件名或路径 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 拼接后的路径 |
```js
path.join(__dirname, 'dist', 'index.html') // => D:\htdocs\dist\index.html
```
#### resolve()
将若干路径片段拼接成绝对路径。
| 参数 | 描述 |
| --- | --- |
| 字符串若干 | 要拼接的路径片段 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 拼接后的绝对路径 |
给出的路径参数会从右向左解析并拼接,一旦发现了绝对路径(开头为`/`),则停止解析。
```js
path.resolve('foo', '/src', 'dist') // => D:\src\dist
```
如果没有发现绝对路径,那么以当前工作目录(命令行的执行位置)作为绝对路径拼接。
```js
// 当前命令行指向 D:\WorkSpace\test
path.resolve('dist') // => D:\WorkSpace\test\dist
```
### URL
`URL`模块提供了与 URL 相关的功能。
```js
const url = require('url')
```
#### parse()
解析一个 URL。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要解析的 URL |
| 布尔值 | 默认为`false`,表示以字符串获取`query`部分。如果为`true`则以对象方式获取 |
| 返回值 | 描述 |
| --- | --- |
| 对象 | 解析后的对象 |
```js
let myURL = url.parse('https://user:pass@sub.example.com:8080/p/a/t/h?id=233#hash')
let port = myURL.port // => 8080
// { protocol: 'https:',
// slashes: true,
// auth: 'user:pass',
// host: 'sub.example.com:8080',
// port: '8080',
// hostname: 'sub.example.com',
// hash: '#hash',
// search: '?id=233',
// query: 'id=233',
// pathname: '/p/a/t/h',
// path: '/p/a/t/h?id=233',
// href: 'https://user:pass@sub.example.com:8080/p/a/t/h?id=233#hash' }
let myURL = url.parse('https://sub.example.com:8080?id=233', true)
myURL.query // => {id: 233}
```
### File System
`fs`模块提供了与文件操作相关的功能。
```js
const fs = require('fs')
```
#### readFile()
读取文件的内容。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 文件路径 |
| 字符串 | 可选,文件的编码类型,默认为当前系统的编码,可以设置为`utf8` |
| 函数 | 回调函数,该函数包含两个参数,第一个参数表示错误对象,如果没有错误则为`null`,第二个参数表示读取的文件内容,如果发生错误则为`null` |
| 返回值 | 描述 |
| --- | --- |
| 对象 | 解析后的对象 |
```js
fs.readFile('./test.txt', 'utf8', function(err, content) {
if (err) {
console.log(err) // 如果出现错误,则输出错误信息
} else {
console.log(content) // 否则输出文件内容
}
})
```
#### writeFile()
向文件写入内容。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 文件路径,如果文件不存在则自动创建 |
| 字符串 | 要写入的文件内容 |
| 函数 | 回调函数,该函数包含一个参数表示错误对象,如果没有错误则为`null` |
| 返回值 | 描述 |
| --- | --- |
| 对象 | 解析后的对象 |
```js
fs.writeFile('./test.txt', '这里是文件内容啦', function(err) {
if (err) {
console.log(err) // 如果出现错误,则输出错误信息
} else {
console.log('文件写入成功!')
}
})
```
### Util
`util`模块提供了各种常用的功能。
```js
const util = require('util')
```
#### promisify()
将 Node.js 中原本的异步任务改造成返回`Promise`对象的函数。
| 参数 | 描述 |
| --- | --- |
| 函数 | 普通的异步函数 |
| 返回值 | 描述 |
| --- | --- |
| 函数 | 包装后的新函数,返回`Promise`对象 |
```js
const readFile = require('util').promisify(fs.readFile) // 包装后的异步任务
readFile().then() // 调用后返回 Promise 对象
```
## npm
Node.js 也有很多其它开发者提供的第三方功能,由于这些功能通常由多个模块组成,因此将它们统称为包。Node.js 内置了一个称为 [npm](https://www.npmjs.com/)(Node Package Manager)的包管理工具,使用它输入命令后可以在公共平台上自动下载安装这些第三方包。
使用`npm install`命令(可以简写为`i`)可以在**命令行的当前路径**安装对应的包。如果在后面添加`-g`参数,则表示全局安装,也就是在任何项目下都可以使用该包。
```powershell
npm install 包名称
npm i 包名称
npm i 包名称 -g
```
在当前项目安装的包,会保存到项目根目录下的`node_modules`文件夹下,在下载包时该文件夹会自动生成,无需手动创建。
模块分为**项目依赖**和**开发依赖**。项目依赖表示开发和实际环境都需要使用的模块(比如 jQuery),这也是上述默认的安装方式;而开发依赖仅需要在开发时使用,与实际运行无关(比如之后的 Gulp 等自动化构建工具),需要在安装命令后添加`--save-dev`参数。
```powershell
npm i 包名称 --save-dev
```
使用`npm uninstall`命令可以卸载已经安装的包。
```powershell
npm uninstall 包名称
```
### 配置文件
但是如果需要将项目拷贝给其它人时,单凭这一个文件夹很难判断出项目依赖的模块和对应的版本,并且由于包文件很多,也会影响拷贝的速度。
因此,使用`npm init`命令可以生成一个配置文件`package.json`,它保存了项目的版本号、依赖模块等信息。
```powershell
npm init # 按回车表示使用默认值
npm init -y # 全部使用默认值
```
项目运行时依赖的模块会保存在配置文件的`dependencies`属性中,而开发依赖的模块保存到`devDependencies`属性中。当模块下载或更新后,这些属性自动也会同步更新。
要拷贝项目时,只需要将源代码和该文件复制到新的工作路径,然后使用`npm install`命令就会自动根据配置文件下载对应的模块。
```powershell
npm install
npm i # 简写形式
npm i --production # 实际部署项目时使用,只下载项目依赖模块,不下载开发依赖模块
```
### 常用的全局包
下面安装几个常用的全局包。它们均是方便开发的命令行工具,因此使用全局安装。
#### nodemon
nodemon 可以监控文件保存状态,并自动重新运行文件。
```powershell
npm i nodemon -g
```
安装成功后,不需要使用`node`,而是使用`nodemon`执行当前目录下的文件,可以看到命令行窗口多出了一些`[nodemon]`开头的信息,而且当执行完毕后,并没有将控制台释放给系统,依然是等待的状态。如果修改了文件,可以看到控制台中自动输出了更改后的信息。
如果要断开链接,可以连续按下两次<kbd>CTRL + C</kbd>。
#### nrm
nrm 可以切换默认的下载节点。由于 npm 默认的下载站点在国外,因此下载速度可能会比较慢。
```powershell
npm i nrm -g
```
安装完成后,使用`nrm ls`命令可以列出所有的下载节点,前面的`*`表示当前正在使用的节点:
然后,使用`nrm use taobao`将下载节点切换为国内的淘宝。此时继续使用`npm`下载包时,会自动使用国内的镜像站,速度应该有明显提升。
### npx
npm 从 5.2 版本开始提供了`npx`命令,使用该命令可以方便地调用项目内部安装的模块。例如,在项目中安装了本地(非全局)的 webpack,如果需要在命令行中使用 webpack,那么必须要手动在`node_modules`中找到 webpack 的可执行文件,因为**只要是在终端直接运行的命令,它会去全局环境中寻找**。由于这里使用了局部安装,因此直接使用`webpack`命令自然是找不到的。
```powershell
./node_modules/.bin/webpack
```
> `.bin`是`node_modules`目录下的一个文件夹,里面保存着模块的可执行文件,不是什么特殊的指令~
如果使用 npx,那么只需要在项目中调用如下命令即可。它的原理非常简单,也就是自动到项目的`node_modules/.bin`以及环境变量中寻找命令是否存在。
```powershell
npx webpack
```
> *参考资料*
>
> [npx 使用教程](https://www.ruanyifeng.com/blog/2019/02/npx.html)
### npm scripts
npm 允许在项目中`package.json`中,使用`scripts`属性定义脚本命令。例如,我们需要将`less`目录中的`test.less`编译为`css`目录中的`test.css`文件:
```powershell
lessc ./less/test.less ./css/test.css
```
但是这样写太麻烦了,于是可以在`package.json`中添加这样一个属性:
```json
// ...
"scripts": {
"test": "echo \"Error: no test specified\" && exit 1",
"less": "lessc ./less/test.less ./css/test.css"
},
// ...
```
这样,只需要使用`npm run 属性名`命令就可以执行这段脚本:
```powershell
npm run less
```
注意,与在终端中直接运行命令不同的是,npm scripts 会**优先从本地寻找该命令**,如果找不到才会去全局环境中寻找。所以,除了使用`npx webpack`可以运行本地 webpack 以外,也可以在 npm scripts 中先定义好运行 webpack 的脚本,然后使用`npm run build`即可运行本地的 webpack。
```json
"scripts": {
"build": "webpack"
},
```
> *参考资料*
>
> [npm scripts 使用指南](https://www.ruanyifeng.com/blog/2016/10/npm_scripts.html)
## 构建网络服务
Node.js 不需要像 PHP 一样依赖于 Apache 等服务器软件,它内置了多个模块如`Net`、`Dgram`、`HTTP`和`HTTPS`分别来处理 TCP、UDP、HTTP 和 HTTPS 的网络请求。
由于诸如 Express 这样的主流框架已经对这些功能进行了高级封装,实际应用中也几乎不可能自己去实现一个服务器,所以如果希望了解原理,请参考[另一篇文章](/posts/tb618gp3.html),本文直接使用[Express](https://expressjs.com/)构建网络服务。
```powershell
npm i express
```
### 创建服务器
使用下面的代码可以创建一个最简单的服务器,Express 会帮你自动设置 MIME 类型、编码和状态码(见[HTTP 协议](/posts/dzta2ga7.html#HTTP)一节)。对于没有配置的路径,也会自动返回`404`状态码和错误页面。
```js
const express = require('express') // 引入模块
const app = express() // 创建服务器对象
const port = 3000 // 端口号
// 监听端口
app.listen(port, function() {
console.log(`服务器启动成功,正在监听 ${port} 端口`)
})
```
### 页面路由
创建服务器后,需要根据用户请求的路径来返回相应的内容,这被称之为页面路由(Routing)。这里的内容通常来说是一个 HTML 页面,但是也可以是一个普通字符串、一段 JSON 等等。
使用`METHOD()`方法设置当客户端以特定方式(如`GET`、`POST`等)请求特定路径时所返回的内容。这里的`METHOD`需要替换成具体的方式,其详细列表见[官网](https://expressjs.com/en/4x/api.html#app.METHOD)。
| 参数 | 描述 |
| --- | --- |
| 字符串或正则表达式 | 要匹配的请求路径 |
| 函数 | 匹配后的处理函数,该函数包含两个参数,分别为请求对象和响应对象 |
匹配成功后,可以使用响应对象的`send()`方法返回响应内容,否则客户端会一直停留在等待状态。
| 参数 | 描述 |
| --- | --- |
| 字符串 / 数组 / 对象 | 当参数为字符串时,返回的`Content-Type`为`text/html`,否则为`application/json` |
```js
app.get('/', function(req, res) {
res.send('欢迎来到首页!')
})
app.post('/login', function(req, res) {
res.send('登录成功!')
})
app.post(/a/, function(req, res) { // 正则表达式,表示任何包含字符 a 的路径
res.send('嗯?')
})
```
#### 页面重定向
通过`res`对象的`redirect()`方法可以实现页面重定向,Express 会自动返回`302`状态码,并使客户端跳转到重定向后的页面。
```js
res.redirect('/admin/login')
```
#### 模块化路由
> 模块化路由需要使用中间件,可以先跳过本节,看完下文的中间件之后再回来。
如果一个网站页面比较多,那么将所有的路由写到`app.js`中会使其难以维护。为此 Express 提供了模块化路由,通过它可以将一级路径与路由对象绑定,然后在路由对象中继续定义二级路由,相当于创建了一个小型的`app`对象。这样的方式可以将二级路由放到单独的模块中,并返回路由对象,在`app.js`中只需要引入模块并获取路由对象,然后将其绑定即可。
使用`express`对象的`Router()`方法可以创建一个路由对象,然后使用`use()`中间件将路由对象与一级路径匹配,就表示这个一级路径下的所有请求都交给该路由对象处理,然后再通过路由对象创建二级路由即可。
```js
// blog.js 单独的路由模块
const blog = express.Router() // 创建路由对象
blog.get('/list', (req, res) => { // blog 下的二级路由,匹配 /blog/list
res.send('欢迎来到博客列表页')
})
module.exports = blog
```
```js
// app.js 主模块
const blog = require('./blog.js')
app.use('/blog', blog) // 将所有 /blog 交给路由对象处理
```
注意,在其它路由模块中不需要重复创建`app`对象,使用请求对象`req.app`也可以获取到`app`对象。
### 接收参数
#### GET 请求参数
使用请求对象的`query`属性可以获取`GET`请求参数。
```js
// localhost/index?name=daisy&age=18
app.get('/index', function(req, res) {
req.query // => {name: daisy, age: 18}
})
```
#### POST 请求参数
要获取 POST 请求参数,需要先安装一个 Express 的扩展模块[express-formidable](https://www.npmjs.com/package/express-formidable)。然后使用中间件(见下文)拦截所有请求:
```
npm install express-formidable
```
```js
const formidable = require('express-formidable')
// 中间件:获取并处理请求参数
app.use(formidable())
app.post('/add', function(req, res) {
req.fields // POST 请求参数
})
```
#### 路由参数
路由参数是另一种传递 GET 参数的方式,它使参数看起来像是路由的一部分,而不是问号引导的形式。目前主流的前端框架也都实现了这种传递参数的方式,使路径看起来更加的友好。
要使用路由参数,需要在路由后面使用`/:参数`作为占位符,当服务器接收到这样的请求时,就会将占位符的部分作为参数了。注意,路由参数的获取需要使用`req.params`属性,而不是普通`GET`请求方式的`req.query`。
```js
// 当请求路径为 localhost/user/add/233,那么 233 就会被识别为 id 参数
// 注意,如果是路径为 localhost/user/add,那么不会匹配到该路由
app.get('/add/:id', function(req, res) {
req.params // => { id: 233 }
})
```
### 中间件
中间件指的是位于接收请求和发送响应之间的处理模块,它们通常不会用来处理请求和响应。可以理解成污水到最终排放之间,需要经过层层过滤和处理,这个过程就需要若干个中间件来完成。
使用`use()`可以创建一个中间件,拦截服务器接收的请求。注意,中间件会拦截所有以某路径**开头**的路由,而不仅仅是当前指定的路由。如配置的拦截参数是`/test`,那么它会拦截所有`/test/a`、`/test/a/b`等所有子路由。
| 参数 | 描述 |
| --- | --- |
| 字符串或正则表达式 | 要拦截的请求路径 |
| 函数 | 匹配后的处理函数,该函数包含三个参数,分别为请求对象、响应对象和函数`next()` |
一旦中间件拦截成功,那么默认不会执行下面匹配的其它中间件。如果需要继续匹配,那么需要手动调用`next()`函数。
```js
app.use('/test', function (req, res) { // 拦截了 /test 的请求
console.log('被拦截了!')
// next() // 除非调用 next()
})
app.get('/test', function (req, res) { // 虽然这个路由也匹配,但是不会执行
res.send('hello')
})
```
反之,如果将上述两个函数调换位置,那么中间件也永远不会被执行,因为路由函数`get()`表示该路由已经处理完毕,也没有`next()`函数可供调用。
### 静态资源
但是很多情况下,并不是所有的请求都需要额外处理,比如客户端请求一个 CSS / JS 文件、一张图片等等,如果要将这些请求都手动进行路由,那么实在有些过于繁琐了。而 Express 提供了`static()`方法可以直接访问这些**静态资源**。我们通常将静态资源目录命名为`public`,将其与根目录拼接即可获得物理路径。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 静态资源目录的路径 |
这是一个中间件函数,它需要放到`use()`中才可以正确执行。
```js
app.use(express.static(path.join(__dirname, 'public'))) // 托管的目录相当于 D:\WorkSpace\Test\public
```
它会自动根据静态资源目录下的文件生成路由。例如,现在的项目目录结构如下,其中`public`是放置静态资源的目录。那么当配置静态资源托管之后,Express 会自动拦截如`/300.png`、`/css/a.css`、`/index.html`这些路由,并自动返回静态资源的目录下对应的文件。
```
Test
├─ app.js
├─ node_modules
├─ public
│ ├─ 300.png
│ ├─ css
│ │ └─ layout.css
└─ └─ index.html
```
因此如果需要链接一个 CSS 文件,那么可以写作:
```html
<link rel="stylesheet" href="/css/layout.css">
<!-- 自动返回 /public/css/layout.css 文件 -->
```
### 模板引擎
实际当中,我们很少将内容完全固定的页面直接返回,而通常是先从数据库查询出需要的数据,然后填充到页面结构中,再将整个页面响应给客户端。与 PHP、JSP 等语言不同,Node.js 并没有这类将前端页面和后端语言结合的语法,因此需要借助模板引擎来实现这一功能。
也就是说,我们不再写直接交给客户端的 HTML 页面,而是按照模板引擎的语法写出页面,将其交给模板引擎。模板引擎会根据其中的代码进行数据拼接等操作,然后将其处理成最终的页面响应给客户端。
或者,如果你采用了「前后端分离」的开发模式(前端只通过接口获取 JSON 数据,而不是直接获取渲染后的页面),那么也可以将模板引擎部署在浏览器端。否则必须通过 JavaScript 动态创建元素,将数据存放在元素中,再追加到页面上。而这样并不利于文档结构的维护,因此同样需要借助模板引擎拼接 HTML 和数据。
模板引擎有很多种,比如[EJS](https://ejs.co/)、[Jade](http://www.nooong.com/docs/jade_chinese.htm)等等,它们的用法大同小异,这里使用的是国内制作的[art-template](/posts/3q4y7l2d.html)。
```powershell
npm install express-art-template art-template
```
使用下面的代码在 Express 中部署 art-template 模板引擎。这些参数均为 Express 中[要求配置的参数](http://expressjs.com/en/5x/api.html#app.set),并不属于 art-template 模板引擎,因此其它模板引擎配置的方式也基本一样。
```js
// 根据要渲染的文件后缀名(也就是下面 render() 文件的后缀名),来引入相应的模板引擎,必须设置
app.engine('html', require('express-art-template'))
// 设置模板的存放目录,默认值为 process.cwd() + '/views',process.cwd() 为当前 Node.js 的进程工作目录,也就是项目根目录下的 views 文件夹
app.set('views', path.join(__dirname, 'views'))
// 设置要渲染的文件后缀名省略时,默认补充的后缀
app.set('view engine', 'html')
// 使用 render() 方法响应模板内容,后面传入数据参数
// 如果 views 下面还有子目录 home,那么要使用 res.render('home/index.html'),不要在 home 前添加斜杠了
app.get('/', (req, res) => {
res.render('index.html', { name: 'aui', age: 18 }) // 这里使用了后缀名,如果省略,则根据 view engine 的配置自动补充
})
```
对于每个模板都需要使用的公共数据,可以将其配置到`app.locals`对象中,这样每个模板都能直接获取到该数据。
```js
app.locals.user = { name: '啦啦啦', age: 18 } // 在模板中,直接使用`user`即可获取到该对象
```
### Session
在我们浏览网站时,经常会见到这样一些功能:如果用户登录成功了,那么在某段时间内,用户不再需要重复登录。或者,用户只有在登录之后才能访问特定的页面。
这些需求看似常见,然而并不容易实现。因为 **HTTP 协议是无状态的**,当客户端与服务器的一次请求结束后,服务器就不再认识客户端了。也就是说,虽然用户本次登录成功了,但是服务器无法知道本次与下次登录的是不是同一个用户,只能当作一个新用户来要求其重新登录。或者,如果用户直接访问登录后的页面,那么服务器也无法判断这个用户是否已经登录成功。
> 为了理解这个问题,可以看生活中的一个例子:有一家烤鸭店搞活动,只要一个顾客购买 5 只烤鸭,就可以免费再获得 1 只。但是客人这么多,烤鸭店要如何记住每一个客人呢?店铺可以制作一些带有卡号的会员卡,然后在店里准备一个记录本,将顾客的会员卡号与购买的烤鸭数记录下来。这样,当顾客购买烤鸭前,先出示一下会员卡,这样店铺就知道顾客之前购买过几只烤鸭了,从而判断是否要赠送烤鸭。
其实解决方案与上述例子很相似,为了使服务器(烤鸭店)记住客户端(顾客),那么也要准备一个记录本和会员卡,记录本是服务器的 session 对象,以 sessionID 识别用户,它可以位于服务器内存、文件、数据库,甚至专门的服务器集群中;而会员卡是客户端的本地存储 cookie。当用户首次登录(购物)后,服务器会生成一个随机的 sessionID(卡号),记录到客户端的 cookie 中(会员卡)。
当客户端再次发送 HTTP 请求时,会将本地**同域**的 cookie 携带在请求头中,一同发送到服务器(出示会员卡)。于是服务器可以检查 session 中(记录本)是否包含之前生成的 sessionID(卡号),如果找到了,则说明用户之前登录成功过(购买过烤鸭)。
> session 的本义是「会话」,而无论是服务端的 session 对象,还是 cookie 中记录的 sessionID,都是 session 的**具体实现**。session 的实现并不一定需要依赖于 cookie,只不过 cookie 是目前最简单有效的实现方式之一。
在 Node.js 中要实现服务端 session,需要安装扩展包[express-session](https://www.npmjs.com/package/express-session)。
```
npm install express-session
```
```js
const session = require('express-session')
```
`session`的构造函数是一个中间件函数,它可以传入一个对象,进行初始化配置:
```js
app.use(session({
secret: 'open-sesame', // 增加生成 sessionID 的强度,防止被篡改,类似于盐值
resave: false, // cookie 过期之前,客户端再次访问是否重复保存 session,建议 false
saveUninitialized: false // 是否保存未添加实际内容的 session,建议 false,
cookie: {
maxAge: 24 * 60 * 60 * 1000, // 默认关闭浏览器就会清除,因此要设置 cookie 过期时间,单位为毫秒,这里表示的是一天
}
}))
```
当客户端登录成功后,可以在 session 中记录一个变量(如登录的用户名),如果设置了`saveUninitialized: false`,那么此时服务器会将同时生成的 sessionID 保存到客户端的 cookie 中:
```js
app.post('/login', function (req, res) {
if (isValid) { // 如果登录成功
req.session.username = result.username // 在 session 中记录用户名,或者其它值
// 跳转到登录后页面
}
})
```
之后在其它页面只需要先验证该变量是否存在,然后根据情况返回不同页面即可。
```js
app.get('/user', function (req, res) {
req.session.username ? res.render('admin/user.html') : res.render('admin/login.html')
})
```
使用响应对象的`clearCookie()`可以清除当前客户端保存的 cookie,该方法要求传入一个 cookie 名称。
```js
app.get('/', function (req, res) {
res.clearCookie('connect.sid') // 清除客户端保存的 sessionID
})
```
## 数据库
Node.js 与其它后端语言一样,都可以与数据库进行交互。由于 MongoDB 是直接操作 JSON 数据,因此与 Node.js 结合使用会更加方便。
MongoDB 本身提供了连接 Node.js 的一系列接口,但是原生代码一向比较复杂。因此与 Express 一样,这里直接使用第三方模块[Mongoose](https://mongoosejs.com/) 操作 MongoDB,对于 MongoDB 本身的安装与使用,请查看[另一篇文章](/posts/e6q2910w.html)。
```powershell
npm install mongoose
```
```js
const mongoose = require('mongoose')
```
### 连接数据库
首先,使用`mongoose`对象的`connect()`方法连接数据库。由于 MongoDB 不需要手动创建数据库,因此这里的数据库名称可以随意填写,之后创建文档后会自动创建。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 数据库地址 |
| 对象 | 配置参数 |
```js
// 参数均是清除一些警告信息,需要的时候加上即可
mongoose.connect('mongodb://localhost/test', { useNewUrlParser: true, useUnifiedTopology: true })
// 如果数据库有密码
mongoose.connect('mongodb://admin:123456@localhost/test', { useNewUrlParser: true, useUnifiedTopology: true })
```
### Schema
由于 MongoDB 本身对存入的数据没有任何约束,因此即便传入错误的数据(甚至是某些字段没有传入),MongoDB 也都会将它们保存起来。为了解决这一问题,Mongoose 引入了规则(Schema)这一概念,它类似于表结构,规定了一个集合(表)的字段类型。
使用`mongoose.Schema()`构造函数可以创建一个规则,它的参数为一个对象,其属性为各个字段的名称,而属性值为该字段的类型。字段的类型有很多,而且针对于不同的类型,还可以进一步作出约束。
常用的字段类型如下:
- `String`:字符串,如果传入了一个非字符串值,那么 Mongoose 会自动调用`toString()`方法尝试自动转换
- `Number`:数值,如果传入了一个非数值,那么 Mongoose 会尝试自动转换
- `null`、`undefined`不会被转换
- `NaN`、没有`valueOf()`的数组或对象会导致转换错误
- `Date`:日期
- `Boolean`:布尔值,除了下列值以外,其它任何值都会导致转换错误
- 被转换成`true`的值:`'true'`、`1`、`'1'`、`yes`
- 被转换成`false`的值:`'false'`、`0`、`'0'`、`no`
- `ObjectId`:由于 MongoDB 中的`id`并不是一个简单的字符串,因此 Mongoose 提供了这样一个单独的类型
```js
// 创建了一个 游戏 的规则
const GameSchema = new mongoose.Schema({
id: mongoose.ObjectId, // ID
name: String, // 游戏名称
type: String, // 游戏类型
isPublished: Boolean, // 是否发售
publishDate: Date, // 发售日期
endings: Number // 结局数量
})
```
如果需要更详细的约束,那么属性的值可以为一个对象。这里只列出较为常见的约束,全部类型详见[官方文档](https://mongoosejs.com/docs/schematypes.html)。
- 所有的类型都可以包含的约束:
- `required`:布尔值,非空约束。要求该字段必须提供,且值不能为`null`、`undefined`、`''`
- `default`:任意类型,如果**插入文档**(更新文档无效)时该字段为`undefined`(对`null`或`''`无效),所使用的默认值
- `validate`:对象,自定义验证规则,包含`validator`和`message`两个属性。前者为一个函数,其返回值为验证规则;后者为字符串,表示自定义的错误信息
- `get`:函数,获取数据时,对数据进行自定义处理
- `set`:函数,传入数据时,对数据进行自定义处理
- 只有字符串类型可以包含的约束:
- `lowercase`:布尔值,是否调用`toLowerCase()`将传入值转换成小写
- `uppercase`:布尔值,是否调用`toUppercase()`将传入值转换成大写
- `trim`:布尔值,是否调用`trim()`去除传入值两端的空格
- `match`:正则表达式,验证传入值是否匹配
- `enum`:数组,验证传入值是否为其中之一
- `minlength`:数值,验证传入值的长度是否符合条件
- `maxlength`:数值,验证传入值的长度是否符合条件
- 只有数值类型可以包含的约束:
- `min`:数值,验证传入值的长度是否符合条件
- `max`:数值,验证传入值的长度是否符合条件
- `enum`:数组,验证传入值是否为其中之一,对于`null`和`undefined`无效
```js
const GameSchema = new mongoose.Schema({
name: {
type: String,
required: true, // 不能为空
default: 'Resident Evil', // 默认值
minlength: 2, // 最小字符数,汉字也算 1 个字符
maxlength: 50, // 最大字符数
trim: true, // 自动去除字符串两端空格
},
type: {
type: String,
enum: ['ACT', 'RPG'] // 只能是这两个值之一
}
endings: {
validate: { // 自定义验证规则,v 表示要验证的值
validator: v => { return v > 3 }, // 如果传入的值大于 3,表示验证通过
message: '什么破游戏, 连 3 个结局都没有!' // 验证失败时的错误信息
},
get: v => Math.round(v), // 获取或存入数据时,对其进行处理
set: v => Math.round(v),
min: 3, // 最小值
max: 10, // 最大值
}
})
```
上述`required`、`max`、`min`、`enum`、`match`、`minlength`和`maxlength`会调用内置的验证器,将这些值放到数组中,并传入第二个字符串元素,可以自定义这些验证器的错误信息。
```js
const GameSchema = new mongoose.Schema({
endings: {
max: [99, '最大值不能超过99啦']
enum: [[22, 33], '只能是 22 或者 33']
}
})
```
对于不符合验证规则的数据,Mongoose 会抛出一个错误。在下面增删查改的方法中可以使用`Promise`的`catch()`,或者在回调函数中获取这个错误对象。使用`错误对象.errors.字段名.message`可以获取到错误提示信息。
```js
const GameSchema = new mongoose.Schema({
endings: {
validate: {
validator: v => v === 233, // 只能传入 233
message: '传的什么鬼东西!'
}
}
})
GameModel.create({endings: 666}).catch(function (err) { // err 为错误对象
err.errors.endings.message
})
```
### Model
将定义的规则传入`model()`方法可以创建一个模型(Model),模型是 Mongoose 特有的概念,可以理解成集合(表)。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要操作的集合(表)名称,这里建议采用**首字母大写的单数形式**,然后 Mongoose 会将其转换成**全部小写的复数形式**,与数据库中的集合对应 |
| `Schema`对象 | 之前创建的包含集合规则的对象 |
| 返回值 | 描述 |
| --- | --- |
| `Model`对象 | 模型对象,用来对数据库进行操作 |
```js
const GameModel = mongoose.model('Game', GameSchema) // 集合名称为 Game,则数据库实际操作的集合名称为 games
```
然后,使用模型就可以调用一系列增删查改方法对数据库进行基本操作了。
#### create()
插入若干条文档。
| 参数 | 描述 |
| --- | --- |
| 若干对象 / 数组 | 要添加的数据 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;之后若干参数为插入的数据,包括自动生成的`_id` |
| 返回值 | 描述 |
| --- | --- |
| `Promise`对象 | 可以取代回调函数 |
```js
// 参数是若干个单独的对象
GameModel.create(
{ name: 'Resident Evil', isPublish: true, endings: 1 },
{ name: 'Enter the Gungeon', isPublish: true, endings: 8 },
function(err, p1, p2) {
err // 错误对象
p1 // { _id: 5e4687310a6da6283498057d, name: 'Resident Evil', isPublish: true, endings: 1 }
p2 // { _id: 5e4687310a6da6283498057d, name: 'Enter the Gungeon', isPublish: true, endings: 8 }
}
)
// 参数是一个对象数组
GameModel.create([
{ name: 'Resident Evil', isPublish: true, endings: 1 },
{ name: 'Enter the Gungeon', isPublish: true, endings: 8 }],
).then(function(p1, p2) {})
.catch(function(err) {})
```
#### find()
根据条件查询文档。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示查询全部 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为查询结果,无论结果数据是多条、单条还是没有,均为**对象数组** |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
// 查询 name 为 'Enter the Gungeon' 的文档
GameModel.find({ name: 'Enter the Gungeon' })
// 查询全部文档
GameModel.find()
```
注意,MongoDB 中的`_id`在 JavaScript 中会表现为`Object`类型,**并不是字符串**,因此在使用时可以直接使用 Mongoose 自动生成的`id`属性来获取字符串类型的`_id`,而不需要手动去调用`_id.toString()`。
#### findOne()
根据条件查询一条文档。如果有多个符合条件的文档,只返回其中的第一条。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示查询全部 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为查询结果,如果查询到文档则返回单个对象,如果没有找到文档则返回`null` |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
// 查询第一条文档
GameModel.findOne({ name: 'Enter the Gungeon' })
```
#### findById()
根据文档`_id`查询一条文档,与`findOne({ _id: id })`效果相同,如果要实现这一功能,建议使用该方法。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 文档的`_id` |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为查询结果,如果查询到文档则返回单个对象,如果没有找到文档则返回`null` |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
GameModel.findById('5db6cbb42b004b2840128f79')
```
#### countDocuments()
获取查询的文档数量。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示查询全部 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为符合条件的文档数量 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
GameModel.countDocuments({ ending: 3 }, function(err, result) {
result // 结局数为 3 的游戏数量
})
```
#### updateOne()
根据条件修改一条文档。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示不修改 |
| 对象 | 要修改的值组成的对象 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为包含修改条数的结果对象 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
// 将 id 为 5d9f6b147589b70e201eca72 的文档的 name 修改为 'Biohazard'
GameModel.updateOne({ _id: '5d9f6b147589b70e201eca72' }, { name: 'Biohazard' })
```
#### updateMany()
根据条件修改多条文档。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示不修改 |
| 对象 | 要修改的值组成的对象 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为包含修改条数的结果对象 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
// 将所有 type 为 'AVG' 的文档的 name 修改为 'Biohazard'
GameModel.updateMany({ type: 'AVG' }, { name: 'Biohazard' })
```
#### deleteOne()
删除符合条件的第一条文档。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示删除全部文档的第一条 |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为包含删除条数的结果对象 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
// 删除 endings 为 3 的第一条文档
GameModel.deleteOne({ endings: 3 })
```
#### deleteMany()
删除符合条件的多条文档。
| 参数 | 描述 |
| --- | --- |
| 对象 | 可选,表示[查询条件](/posts/e6q2910w.html#查询操作符),如果省略则表示**不删除** |
| 函数 | 可选,回调函数,可以使用`Promise`替代。第一个参数为错误对象,如果没有错误则为`null`;第二个参数为包含删除条数的结果对象 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | [查询结果对象](#Query) |
```js
// 删除 endings 为 3 的所有文档
GameModel.deleteMany({ endings: 3 })
```
### Query
`Query`对象是 Mongoose 提供的一个查询结果对象,其中包含了一些处理查询结果的常用方法,并且封装了`then()`和`catch()`,可以当作`Promise`来使用,但是它并不是真正的`Promise`。
#### select()
设置查询的字段。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要查询的字段。多个字段以空格隔开,字段名前加`-`表示不查询该字段 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | 调用它的`Query`对象 |
```js
// 仅查询所有文档的 name 和 type 字段
GameModel.find().select('name type').then()
// 不查询所有文档的 name 字段
GameModel.find().select('-name').then()
```
#### sort()
对查询结果排序,默认为升序。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要排序的字段,字段名前加`-`表示降序 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | 调用它的`Query`对象 |
```js
// 将查询结果按照结局数 由小到大 排序
GameModel.find().sort('endings')
// 将查询结果按照结局数 由大到小 排序
GameModel.find().sort('-endings')
```
#### skip()
跳过若干条文档。
| 参数 | 描述 |
| --- | --- |
| 数值 | 跳过的文档数量 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | 调用它的`Query`对象 |
```js
// { _id: 5e4685ffd96f66282c100c61, endings: 3 },
// { _id: 5e468608bc19a50e64ae6263, endings: 233 },
// { _id: 5e46860c31150e222cd388f1, endings: 0 }
GameModel.find().skip(1) // 跳过第一条,即没有 endings: 3 这一条
```
#### limit()
限制返回的文档数。
| 参数 | 描述 |
| --- | --- |
| 数值 | 限制查询的文档数量 |
| 返回值 | 描述 |
| --- | --- |
| `Query`对象 | 调用它的`Query`对象 |
```js
GameModel.find().limit(1) // 只查询 1 条文档,相当于 findOne()
```
<file_sep>---
title: 计算机网络笔记之02 🔌 物理层
date: 2020-01-19 10:16:58
abbrlink: 5yfyor2a
tags: 计算机网络
categories: 计算机网络
excerpt: 物理层是五层模型中的最底层,它主要考虑计算机在传输媒体(如光纤、双绞线等)上如何传输比特流,但传输媒体本身不在物理层的考虑范围。它的作用是尽可能屏蔽掉传输媒体和通信手段的差异,使上面的数据链路层感受不到这些差异。
---
# 计算机网络笔记之02 🔌 物理层
物理层是五层模型中的最底层,它主要考虑计算机在传输媒体(如光纤、双绞线等)上如何传输比特流,但传输媒体本身不在物理层的考虑范围。它的作用是尽可能屏蔽掉传输媒体和通信手段的差异,使上面的数据链路层感受不到这些差异。
物理层定义了与传输媒体接口有关的特性:
- 机械特性:接口形状、尺寸、引脚数目等
- 电气特性:电压范围
- 功能特性:不同电压表示的含义,如`-5V`表示`0`,`+5V`表示`1`
- 过程特性:定义各条物理线路的工作规程和时序关系
之所以规定这些特性,是为了使不同厂家生产的设备能够遵循统一的标准,从而互相协调工作,就好像任何国内的家用电器都可以接到任何的插排一样。
## 数据通信的基础知识
通信的目的是传递消息(如文字、图像、视频等)。数据是运送消息的载体,通常是有意义的符号序列。信号则是数据的电气或电磁表现,它可以分为**模拟信号和数字信号**(统称为电信号)。
模拟信号是连续的波形,而数字信号是离散的取值。通常来说,数字信号用来表示二进制只有两种状态,分别表示`0`和`1`(如`-5V`表示`0`,`+5V`表示`1`),这每一种状态称为数字信号的**码元**。

> 一个码元并非只能表示`0`或`1`,根据调制和解码方式不同,可以使用一个码元表示多个 bit,如`0V`表示`000`,`1V`表示`001`,...,`7V`表示`111`。
由于用户到公用电话网之间距离通常较远,由于成本和传播速率的问题(大概),因此这一段距离需要使用调制解调器(modem,也就是说平时所说的猫)将计算机的数字信号转换成模拟信号传输,到达公用电话网之后再重新转换成数字信号。
> 目前家庭用户大多已经接入光纤,所使用的「光猫」可以将数字信号转换成光信号,从而使用光纤传输。
### 信号调制
信号的调制大致分为两种,一种是仅对信号的波形进行转换,使其与信道特性相适应,转换后的信号依然是数字信号,这种方式称为**编码**(也称为基带调制)。另一种是将数字信号转换成模拟信号,这种方式则称为**带通调制**。
常用的编码方式有下面四种:
- 不归零制:正电平表示`1`,负电平表示`0`
- 归零制:正脉冲表示`1`,负脉冲表示`0`,每个比特位在结束时均要回归零电平
- 曼彻斯特:高电平到低电平表示`1`,低电平到高电平表示`0`,反之也是可以的
- 差分曼彻斯特:在每个比特位均有跳变,如果下一位与上一位跳变相同表示`0`,不同则表示`1`(第一位表示`0`还是`1`可以提前约定)

<file_sep>---
title: 📘 大前端笔记之 Part1 HTML
date: 2020-01-10 22:32:43
abbrlink: andb1byi
tags: HTML
categories: 大前端
excerpt: HTML 是用来标识互联网传输数据语义的一门语言。浏览器根据不同的 HTML 标签将数据进行解析,才能呈现出具有含义的页面。
---
# 📘 大前端笔记之 Part1 HTML
<abbr title="Hyper Text Markup Language,超文本标记语言">HTML</abbr>是用来标识互联网传输数据语义的一门语言,浏览器根据不同的 HTML 标签将数据进行解析,才能呈现出具有含义的页面。
HTML 标签也称 HTML 元素,它由标签名称和其中内容组成。标签名外侧由尖括号`<>`包裹,并且**不区分大小写**(这一点很重要,因为之后会用到自定义标签),因此如果有多个单词会使用**短横线式命名法**。
> 短横线式命名法(kebab-case)指的是以短横线分隔多个单词(如`my-name`),其中 kebab 的含义是「烤肉串」,可见还是比较形象的。与此对应的还有驼峰式命名法(camel-case),以大写字母作为单词分隔(如`myName`或者`MyName`),其中小写字母开头称为小驼峰式,而大写字母开头则称为大驼峰式。
HTML 元素分为单标签元素和双标签元素,前者只需要一个标签,而没有内容。后者需要开始标签和结束标签成对出现,结束标签的名称前会有一个`/`作为标识。

HTML 元素上可以定义属性,用来对元素作出进一步的解释说明,它由属性名和属性值组成,两者使用`=`连接,属性值使用**双引号**包裹。与标签名称一样,属性也可以自定义,并且**属性名同样不区分大小写**,因此如果有多个单词也应该使用短横线式命名法。

## 基本元素
这些元素是 HTML 文档的基本结构,几乎在任何一个文档中都必须存在。
### HTML5 规范
使用`<!DOCTYPE html>`元素表示当前文档遵循 HTML5 规范,以便浏览器按照最新的规范来解析。该元素必须位于 HTML 文档的最顶部。
```html
<!DOCTYPE html>
```
### 根元素
使用`<html>`表示 HTML 文档的顶级元素,也称为根元素,所有其他元素必须是此元素的后代。
如果使用 VS Code 等编辑器快速生成页面模板时,会发现该元素带有`lang`属性,它可以设置文档的语种,以便于浏览器针对页面进行翻译或者搜索引擎检索。对于中文页面来说,该属性可以设置为`zh`(谷歌)、`zh-CN`(淘宝)、`zh-Hans`(哔哩哔哩)或者直接不设置(京东、百度)。这些差异是由于规范中将中文根据方言(如普通话、粤语、闽南语等)分成了非常多的子语种,但是浏览器并没有完全实现,从而导致的问题。
```html
<html lang="zh-Hans"></html>
```
### 文档头部
使用`<head>`表示[文档元数据](#文档元数据)的容器,可以包含一些特殊的标签用来对文档解释说明(比如标题、编码等),它们不会显示在页面上。
```html
<html>
<head></head>
</html>
```
### 文档主体
使用`<body>`表示文档的主体,大部分元素都位于该元素内,用来呈现具体的内容。
```html
<html>
<body></body>
</html>
```
## 文档元数据
文档元数据指的是并不会被解析在页面上的内容,用来设置文档的一些配置信息,它们通常出现在`<head>`标签中。
### 页面标题
使用`<title>`定义页面的标题,也就是浏览器标签页或者标题栏上的文字。
```html
<title>这是一个神奇的网页</title>
```
### 文档编码
使用`<meta>`的`charset`属性设置文档编码,也就是说,告诉浏览器当前页面应该以什么[字符集](/posts/1x90rjzf.html)解析,通常为`UTF-8`。
它的作用仅仅是告诉浏览器应该以什么字符集来解析,而不是设置文档本身的字符集。如果要设置文档本身的字符集,以 VS Code 为例,在页面的右下角即可修改。因此,只有这两者对应起来才能够正确显示页面文字,否则会出现乱码。
```html
<meta charset="UTF-8" />
```
## 文本内容
接下来的标签均会被直接渲染在页面上,通常放在`<body>`中。
### 标题和段落
使用`<h1>`到`<h6>`定义不同级别的标题,其中`<h1>`为最大的标题,`<h6>`为最小的标题。注意,`<h1>`在一篇文档通常只能出现一次,然后根据结构依次递进,不要因为字体大小或样式滥用标题。
```html
<h1>这是一个一级标题哒~</h1>
```
使用`<p>`定义一个段落。
```html
<p>这是一个段落哒~</p>
```
### 强调和删除
使用`<strong>`或`<em>`将需要强调的内容包裹起来,前者默认为**加粗样式**,后者为*倾斜样式*。
```html
<strong>这是强调内容啦</strong>
```
使用`<del>`将需要删除的内容包裹起来,默认为~~删除线样式~~。
```html
<del>这是删除内容啦</del>
```
### 换行
使用`<br>`定义一个换行符。HTML 会默认将文档中的回车以及多个空格,解析为**一个空格**。
### 超链接
使用`<a>`定义一个超链接,用来在不同页面之间跳转。
| 属性 | 描述 |
| -------- | ----------------------------------------------------------------- |
| `href` | 跳转的 URL |
| `target` | 新页面的打开方式,`_self`表示当前页面打开,`_blank`表示新页面打开 |
```html
<!-- 在当前页打开 -->
<a href="http://www.muyumiao.com" target="_self">木鱼喵</a>
<!-- 在新页面打开 -->
<a href="http://www.muyumiao.com" target="_blank">木鱼喵</a>
```
## 页面划分
将页面划分成不同的区域,可以使功能相似的内容归在一起,从而使页面具有清晰的结构。页面划分的元素功能基本一致,但是它们具有不同的语义,应该根据情况选择使用。
| 标签 | 描述 |
| ----------- | ------------------------------------------------------------------------------- |
| `<header>` | 页面的头部,通常包含章节的标题、Logo、搜索框等 |
| `<main>` | 页面的主体,只能在页面出现一次,用于替代`<div id="main">`或`<div id="content">` |
| `<nav>` | 页面的导航,通常包含多个超链接,用于跳转到网站的其它页面 |
| `<section>` | 页面的区块,可以将多个有联系的内容放在一个区块中 |
| `<aside>` | 与页面内容几乎无关的部分,比如侧边栏等 |
| `<article>` | 文章内容区域,比如论坛的帖子、新闻、博客或者用户提交的评论 |
| `<footer>` | 页面的页脚,通常包含章节作者、版权数据、联系方式等 |
| `<div>` | 无语义块级元素,在以上元素均不符合时再考虑使用 |
| `<span>` | 无语义行内元素,在以上元素均不符合时再考虑使用 |
## 表格
使用`<table>`可以定义一个数据表格,其中包含`<tr>`标签作为表格的行,而`<tr>`中又包含若干个`<th>`或`<td>`作为表头或普通单元格。
```html
<table>
<tr>
<th>姓名</th>
<th>年龄</th>
</tr>
<tr>
<td>御坂美琴</td>
<td>16</td>
</tr>
</table>
```
如果表格比较复杂,可以使用`<thead>`、`<tbody>`、`<tfoot>`将`<tr>`包裹起来,划分表格的区域,方便对不同的区域进行样式控制。
```html
<table>
<thead>
<tr>
<th>姓名</th>
<th>年龄</th>
</tr>
</thead>
<tbody>
<tr>
<td>御坂美琴</td>
<td>16</td>
</tr>
</tbody>
</table>
```
在`<th>`或`<td>`中使用`colspan`或`rowspan`属性可以实现相邻单元格跨列或跨行合并。
```html
<table>
<tr>
<!-- 跨列合并,因此 标题 独占一行 -->
<th colspan="2">标题</th>
</tr>
<tr>
<td>御坂美琴</td>
<td>16</td>
</tr>
</table>
```
## 列表
使用`<ul>`和`<ol>`定义一个无序 / 有序列表,在其中使用`<li>`定义列表项。注意,`<ul>`和`<ol>`中只允许包含`<li>`,其它任何元素都是不合法的。
```html
<ul>
<li>苹果</li>
<li>香蕉</li>
<li>橘子</li>
</ul>
```
使用`<dl>`可以定义一个描述列表。它包含两个部分,分别为一个`<dt>`作为描述主题,以及若干个`<dd>`作为描述内容。
```html
<dl>
<dt>帮助中心</dt>
<dd>账户管理</dd>
<dd>入坑指南</dd>
<dd>订单操作</dd>
</dl>
```
## 表单
使用`<form>`创建一个表单区域,表单中可以包含下列各种交互控件,用来接收用户输入并发送到服务端,然后刷新当前页面以呈现服务端返回的新页面。不过需要注意的是,向服务端发送数据远不止提交表单一种方式,因为提交表单需要刷新页面,用户体验并不是特别好。而且其中的控件并不是依赖于表单才能使用,它们只是供用户输入数据,具体的提交方式还要看客户端如何处理。
| 属性 | 描述 | 取值 |
| ------------ | -------------------------------------------------- | ------------- |
| `action` | 发送请求的服务端 URL | |
| `enctype` | 设置请求体的编码方式,在之后文件上传的部分才会用到 | |
| `novalidate` | 布尔属性,禁止浏览器自动验证表单 | |
| `method` | 发送表单数据的方式 | `get`或`post` |
### 单行输入控件
使用`<input>`元素创建单行输入控件。它们中的大部分使用`value`属性来保存数据,使用`name`属性值来命名,以便服务器区分。使用`type`属性可以设置控件的类型,根据其属性值的不同,它的功能也有所不同:
| 属性值 | 描述 |
| ---------- | ----------------------------------------------------------------------- |
| `text` | 文本框 |
| `password` | 密码框 |
| `button` | 普通按钮,通过`value`属性设置按钮上的文字 |
| `submit` | 提交按钮,通过`value`属性设置按钮上的文字 |
| `radio` | 单选框,每组单选框的`name`属性值必须相同 |
| `checkbox` | 复选框,每组复选框的`name`属性值必须相同 |
| `file` | 文件上传控件 |
| `email` | 【☢️HTML5】邮箱地址框,浏览器会自动进行验证 |
| `url` | 【☢️HTML5】网址框,浏览器会自动进行验证 |
| `tel` | 【☢️HTML5】电话号码框 |
| `search` | 【☢️HTML5】搜索框 |
| `number` | 【☢️HTML5】数字框,在移动端可以唤醒数字键盘,浏览器会自动进行验证 |
| `range` | 【☢️HTML5】滑动数字框,通过`min`、`max`属性设置最值、`step`属性设置步长 |
| `color` | 【☢️HTML5】颜色选择框 |
| `date` | 【☢️HTML5】日期选择框,在移动端可以唤醒日期选择控件 |
除了上述属性之外,它还可能包含下列的其它属性:
| 属性 | 描述 |
| -------------- | ------------------------------------------------------------------------------------------ |
| `checked` | 布尔属性,设置默认选中,只适用于单选框或复选框 |
| `readonly` | 布尔属性,设置只读 |
| `placeholder` | 在文本输入框生成一段提示信息 |
| `autofocus` | 布尔属性,当页面加载后自动获得焦点。但是该属性拥有兼容性问题,即使是现代浏览器也可能不支持 |
| `autocomplete` | 可选值为`on`或`off`,自动填充之前提交过的表单数据,建议关闭 |
| `multiple` | 布尔属性,使文件上传控件可以一次选择多个文件 |
### 下拉菜单
通过`<select>`可以创建一个下拉菜单,其中的`<option>`用来设置选项。每个`<option>`的`value`属性值为提交的数据,如果`value`为空,则提交`<option>`包裹的内容。
不过,由于`<select>`表单不可能使用 CSS 在不同浏览器中获得统一的样式,因此建议使用第三方库中对应的插件,或者通过其它元素自己实现一个下拉菜单。
### 按钮
通过`<button>`也可以创建一个按钮,如同`<input type="button">`一样,不过它是一个双标签,包裹的内容为按钮的文字。根据它的`type`属性值不同,按钮的作用也有所不同:
| 取值 | 描述 |
| -------- | -------------------- |
| `submit` | 默认值,提交表单按钮 |
| `reset` | 重置表单按钮 |
| `button` | 普通按钮 |
注意,它的`type`属性默认值是`submit`,也就是说,如果它在表单中的话,即便没有设置`type`属性,点击后也会直接提交表单。
### 文本域
通过`<textarea>`可以创建一个文本域,用于输入多行文本。标签包裹的内容为提交的数据,`name`属性值为文本域名称,以便服务器区分。
为文本域添加如下的 CSS 代码,可以禁止用户自由缩放。
```css
textarea {
resize: none;
}
```
### 标签
通过`<label>`可以为表单控件设置一个标签。如果点击`<label>`内的文本,浏览器就会自动将焦点设置到`<label>`绑定的表单控件上。
它只有一个`for`属性,在其中传入元素的`id`属性值,就可以将标签与表单元素绑定。或者直接将控件放到标签元素中,也可以实现同样效果。
```html
<label for="username">姓名:</label>
<input type="text" id="username" />
<label>姓名:<input type="text"/></label>
```
## 多媒体
### 图片
使用`<img>`定义一张图片。
| 属性 | 描述 |
| ------- | -------------------------- |
| `src` | 图片的 URL |
| `alt` | 当图片加载失败时的替代文本 |
| `title` | 当鼠标悬停时的显示文本 |
```html
<img src="test.jpg" alt="这是一张神奇的图片" />
```
### 音频
使用`<audio>`可以定义一个音频,但是目前由于版权问题,浏览器支持的格式各不相同。
| 属性 | 描述 |
| ---------- | ---------------------------------------------------------------------- |
| `src` | 音频的 URL |
| `autoplay` | 布尔属性,音频在就绪后立刻自动播放,2018 年 1 月 Chrome 浏览器将其禁用 |
| `controls` | 布尔属性,显示播放控件,但各个浏览器样式不同 |
| `loop` | 布尔属性,开启洗脑循环 |
为了解决这个问题,可以在一个`<audio>`中嵌套多个`<source>`标签,用来加载不同格式的音频。浏览器会从最上面的开始尝试加载,如果遇到不支持的格式,则自动尝试下一个。
```html
<audio autoplay controls loop>
<source src="Level5.mp3" type="audio/mpeg" />
<source src="Level5.ogg" type="audio/ogg" />
</audio>
```
### 视频
使用`<video>`定义一个视频,目前多数浏览器均支持`mp4`格式。它的属性如下:
| 属性 | 描述 |
| ---------- | --------------------------------------------------------------------------------------- |
| `src` | 视频的 URL |
| `autoplay` | 布尔属性,视频在就绪后立刻自动播放,2018 年 1 月 Chrome 浏览器将其禁用,除非设置`muted` |
| `muted` | 布尔属性,静音 |
| `controls` | 布尔属性,显示播放控件,但各个浏览器样式不同 |
| `loop` | 布尔属性,开启洗脑循环 |
```html
<video src="极乐净土.mp4" autoplay controls loop></video>
```
> 也可以直接将视频传到第三方视频网站(优酷、腾讯等),然后获取视频的分享地址,缺点是有广告植入,优点是节省自己服务器的空间。
<file_sep>---
title: 大前端支线笔记之05 💨 CSS 逐帧动画
date: 2020-01-24 0:04:24
abbrlink: m69g0i43
tags: CSS
categories:
- 大前端
- 支线
excerpt: 过渡和动画默认的调速函数都是贝塞尔曲线,它会在动画的关键帧之间进行插值运算,使得关键帧之间变化的非常平滑和自然。但是,如果要实现类似于翻书的走马灯效果,那么中间的过渡反而是多余的。
---
# 大前端支线笔记之05 💨 CSS 逐帧动画
过渡和动画默认的调速函数都是贝塞尔曲线,它会在动画的关键帧之间进行插值运算,使得关键帧之间变化的非常平滑和自然。但是,如果要实现类似于翻书的走马灯效果,那么中间的过渡反而是多余的。
> 使用 GIF 图片也是一种选择,但是 GIF 颜色十分单一,并且不具备半透明效果,同时也不方便修改动画速度等参数。
使用调速函数(`transition-timing-function`或`animation-timing-function`)的`steps()`函数可以使动画逐帧显示,其中的整数值参数表示将**两个关键帧之间的部分**分成几份,而不是整段动画,这一点对于关键帧动画来说尤其需要注意。
首先,使用逐帧动画实现进度条填充的效果,为了方便理解,将其简化成两个阶段,分别为阶段 1(`0px ~ 250px`)和阶段 2(`250px ~ 500px`)。

```html
<div class="container">
<div class="bar"></div>
</div>
```
```css
.container { width: 500px; }
.bar {
width: 0%; /* 为了还原现象,需要手动设置进度条的初始值为 0,否则它的默认值为 100% */
animation: progress 2s steps(2); /* 分为两个阶段,因此设置参数为 2 */
}
@keyframes progress { /* 定义两个关键帧,将这两个帧之间的动画分为两份 */
0% { width: 0%; }
100% { width: 100%; }
}
```
但是,出现的效果可能与想象中有所不同:

这是因为,虽然动画阶段有两个,但是分隔后的动画却包含三个关键帧,分别为`0%`的`0px`,`50%`的`250px`和`100%`的`500px`。而`steps()`只能呈现其中的两个,也就是说**第一帧和最后一帧不能同时出现**。
因此要么只能看到从宽度`0px`到`250px`,然后动画结束回到初始状态`0px`(前提是`animation-fill-mode`为默认值);要么只能看到从宽度`250px`到宽度`500px`,然后动画结束变为`0px`。这两种方式分别对应`steps()`第二个参数的`end`(默认)和`start`值(注意顺序正好是相反的)。
```css
.bar { animation: progress 2s steps(2, start); }
```
如果要显示类似进度条填充的效果,那么需要将进度条的初始宽度设置为`500px`(或者默认的`auto`、`100%`),这样虽然动画只能显示宽度从`0`到`250px`,但是由于动画结束时会恢复到初始状态,也就是宽度为`500px`,看起来就构成了一段完整的动画。
```css
.bar {
width: 100%; /* 用于动画回到初始状态 */
animation: progress 2s steps(2);
}
```
或者,也可以使用`animation-fill-mode: forwards`设置动画结束时维持在最后一帧(即默认时`@keyframes`中`100%`的状态),而不回到初始状态。由于最后一帧定义的为`width: 100%`,因此也可以实现同样效果。
```css
.bar {
animation: progress 2s steps(2) forwards; /* 维持在 @keyframes 的最后一帧 */
}
```
但是,如果动画属性使用了`infinite`值,那么情况就会变得更麻烦。比如将之前的两种解决方式添加`infinite`:
```css
.bar { animation: progress 2s steps(2) infinite forwards; }
```

此时发现效果又没有出现,这是因为,添加了`infinite`的动画效果没有**完成期**,与`forwards`根本就没有关系,而且由于它在不断的循环**执行期**,也没法回到动画结束时的初始状态。
为了方便理解,再次将效果简化,只需要`0% → 100% → 0%`这样一个循环的过程。

根据之前的分析,这次只有两个关键帧,因此设置为`steps(1)`即可。但是这样由于最后一帧不会出现,那么进度条永远为第一帧的白色,不会变成蓝色。
为了解决这个问题,这次需要定义三个关键帧,由于第三个关键帧没法显示,因此将第二个和第三个关键帧设置为同一个状态。为了使时间均匀分布,第二个关键帧的位置需要选在`50%`。此时`100%`的关键帧可以省略,因为它默认就与前一帧相同。
```css
@keyframes progress {
0% { width: 0%; }
50% { width: 100%; }
100% { width: 100%; } /* 可以省略 */
}
```<file_sep>---
title: 大前端笔记之09 👀 CSS 其它属性
date: 2020-01-26 8:47:14
abbrlink: lay8q60u
tags: CSS
categories: 大前端
excerpt: 由于一些属性不便于分类,因此在本节讨论。
---
# 大前端笔记之09 👀 CSS 其它属性
由于一些属性不便于分类,因此在本节讨论。
## 元素阴影
使用`box-shadow`为元素添加阴影。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 行内元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 没有元素阴影 |
| 长度值 | 两个值分别表示阴影水平和垂直方向的**偏移量**,正的数值表示阴影向右下方移动,负值表示阴影向左上方移动;如果有第三个长度值,则表示阴影的**模糊半径**,该值不能为负;如果有第四个长度值,则表示阴影的**扩大或缩小**,该效果发生在模糊之后 |
| 颜色值 | 必须放在长度值最后,表示阴影的颜色,默认为黑色 |
| `inset` | 表示阴影会生成在元素**内部**,它不能出现在长度值中间,其它位置均可 |
```css
div { box-shadow: 2px 2px 2px 2px #000 inset; }
/* 用逗号分隔,可以同时添加多个阴影效果 */
div { box-shadow: 1px 1px 2px #ccc, -2px -2px 1px blue; }
```
## 可见性
使用`visibility`设置元素是否可见。注意,设置不可见的元素依然会**影响布局**,相当于设置`opacity`为`0`,如果希望元素彻底消失则应该使用`display: none`。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `visible` |
| 可选值 | 描述 |
| --- | --- |
| `visible` | 元素可见 |
| `hidden` | 元素不可见 |
## 不透明度
使用`opacity`设置元素的**不透明度**。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | `1` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | `1`表示完全不透明,`0`表示完全透明,可以是`0`到`1`之间的任意数字值 |
## 溢出
使用`overflow`设置元素内容超出自身宽高时的处理方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素和行内替换元素 | 不可继承 | `visible` |
| 可选值 | 描述 |
| --- | --- |
| `visible` | 溢出部分会超出元素本身显示 |
| `hidden` | 溢出部分会被隐藏,且无法通过正常方式查看 |
| `scroll` | 总是生成滚动条,无论内容是否溢出 |
| `auto` | 如果内容溢出则生成滚动条,否则不生成 |
## 指针
使用`cursor`设置鼠标指针在元素上的样式,全部的可选值见[MDN](https://developer.mozilla.org/zh-CN/docs/Web/CSS/cursor)。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | 根据元素决定 |
## 点击触摸
使用`pointer-events`使元素的**鼠标和触摸事件**失效,就好像是「透明」一样,可以用来阻止移动端的**图片点击预览**。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 事件正常触发 |
| `none` | 使鼠标和触摸事件失效 |
例如在超链接上覆盖一个半透明的元素,默认情况下,下面的超链接由于被覆盖,是不可以点击的。但是如果为半透明元素设置`pointer-events: none`,那么超链接就可以正常点击了。

```html
<div class="box box-auto">
<a href="https://www.google.com">这个链接点不到啦</a>
<div class="mask"></div>
</div>
<div class="box box-none">
<a href="https://www.google.com">这个链接可以点</a>
<div class="mask"></div>
</div>
```
```css
.box {
position: relative;
width: 100px;
height: 100px;
}
.mask {
position: absolute;
width: 100%;
height: 100%;
}
.box-none > .mask { pointer-events: none; }
```
## 列表
使用`list-style-type`可以设置列表的标记样式,全部可选值见[MDN](https://developer.mozilla.org/zh-CN/docs/Web/CSS/list-style-type)。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 列表项`<li>` | 可继承 | `none` |
使用`list-style-position`设置文本在发生换行时,标记在内容中的位置。由于该属性可操作性很低,因此如果使用复杂的列表标记,均是通过伪元素添加背景图片的方式来设置。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 列表项`<li>` | 可继承 | `outside` |
| 可选值 | 描述 |
| --- | --- |
| `outside` | 位于内容之外 |
| `inside` | 位于内容之内 |
## 清除点击高亮
使用☢️`-webkit-tap-highlight-color`可以清除移动端链接和按钮在点击时的高亮效果。
```css
a, button {
-webkit-tap-highlight-color: transparent;
}
```
## 清除 iOS 控件的默认样式
使用☢️`-webkit-appearance`清除 iOS 中默认的表单控件样式(文本框自带内阴影、按钮自带圆角等),否则没法设置其它样式。注意,该属性会使 PC 端的控件消失:
```css
button {
-webkit-appearance: none;
}
```
## 禁止长按弹出菜单
使用☢️`-webkit-touch-callout`禁止在移动端长按链接和图片时弹出的选择菜单。
```css
a, img {
-webkit-touch-callout: none;
}
```<file_sep>---
title: 大前端笔记之15 ❄️ JavaScript DOM
date: 2020-02-02 16:00:46
abbrlink: y57hcuma
tags: JavaScript
categories: 大前端
excerpt: DOM 对象分为不同的种类,包括整个文档、HTML 元素或者元素内的文本。这些对象都继承自一个统一的<code>Node</code>接口,因此它们有许多共同的方法和属性,并统一称为节点。
---
# 大前端笔记之15 ❄️ JavaScript DOM
浏览器将页面上的元素映射成一个树状的结构,树上的每一个元素称为 DOM 对象(或者叫**节点**)。DOM 对象分为不同的种类,它们都统一继承自`Node`对象(通过原型链),因此有许多共同的方法和属性:
- `Document`:整个文档
- `Element`:HTML 元素,根据不同的元素又分为`HTMLDivElement`、`HTMLHeadingElement`等等
- `Text`:元素内的文本
由于节点的继承关系比较复杂,可以随时参考[这里](http://w3help.org/zh-cn/causes/SD9024)。
使用`window.document`属性可以获取文档对象(`window`可以省略)。而其它节点则可以使用本文下面的各种方式获取。
## 操作元素
与元素相关的属性和方法大部分位于`document`(直接获取)或`Element`(根据元素上下级关系获取)中。
### body / documentElement
获取`<body>`/`<html>`元素对象。
```js
document.body // => <body>
document.documentElement // => <html>
```
### getElementById()
根据元素的`id`值获取元素。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 元素的`id`值,区分大小写 |
| 返回值 | 描述 |
| --- | --- |
| 元素对象 | 相应的元素对象,如果有多个元素有相同`id`,则只返回其中第一个 |
```html
<button id="btn">这是按钮啦</button>
```
```js
document.getElementById('btn')
```
### getElementsByName()
根据元素的`name`值获取元素。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 元素的`name`值 |
| 返回值 | 描述 |
| --- | --- |
| `NodeList`集合 | 包含符合条件元素的伪数组 |
```html
<input type='text' name="username">
```
```js
document.getElementsByName('username')[0]
```
### getElementsByTagName()
根据元素的标签名获取元素对象。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 元素的标签名 |
| 返回值 | 描述 |
| --- | --- |
| `HTMLCollection`集合 | 包含符合条件元素的伪数组 |
```js
document.getElementsByTagName('p')[0]
```
元素对象`Element`也定义了该方法,可以返回元素节点的后代中符合条件的元素。
```js
p.getElementsByTagName('span')
```
### getElementsByClassName()
根据元素的类名获取元素对象。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 元素的类名,多个类使用空格隔开即可,顺序并不重要 |
| 返回值 | 描述 |
| --- | --- |
| `HTMLCollection`集合 | 包含符合条件元素的伪数组 |
```js
document.getElementsByClassName('foo bar') // 返回同时具有 foo 和 bar 两个类的元素
```
元素对象`Element`也定义了该方法,可以返回元素节点的后代中符合条件的元素。
```js
p.getElementsByClassName('foo') // 获取 p 元素下具有 foo 类的元素
```
### querySelector() / querySelectorAll()
根据 CSS 选择器获取匹配的第一个 / 全部对象。
| 参数 | 描述 |
| --- | --- |
| 字符串 | CSS 选择器 |
| 返回值 | 描述 |
| --- | --- |
| 元素对象 / `NodeList`集合 | 符合条件的元素 / 包含符合条件元素的伪数组 |
```js
document.querySelector('.btn') // 获取第一个拥有 btn 类的元素
document.querySelectorAll('.btn') // 获取全部拥有 btn 类的元素
```
### firstElementChild / lastElementChild / children
获取该元素的第一个 / 最后一个 / 所有子元素。
| 返回值 | 描述 |
| --- | --- |
| 元素对象 / `HTMLCollection`集合 | 符合条件的元素 / 包含符合条件元素的伪数组 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var ul = document.querySelector('#list')
ul.children.length // => 2
ul.firstElementChild // => <li>冥灯龙</li>
ul.lastElementChild // => <li>炎妃龙</li>
```
### previousElementSibling / nextElementSibling
获取该元素的前一个 / 后一个兄弟元素。
| 返回值 | 描述 |
| --- | --- |
| 元素对象 | 符合条件的元素 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var first = document.querySelector('#first')
first.nextElementSibling // => <li id="second">炎妃龙</li>
```
### parentElement
获取该元素的父元素。
| 返回值 | 描述 |
| --- | --- |
| 元素对象 | 符合条件的元素 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var first = document.querySelector('#first')
first.parentElement // => <ul id="list">...</ul>
```
### createElement()
创建元素对象,但是创建后的元素需要使用`Node`中的相关方法插入到文档中才有效果。
| 参数 | 描述 |
| --- | --- |
| 字符串 | HTML 标签名 |
| 返回值 | 描述 |
| --- | --- |
| 元素对象 | 创建的元素对象 |
```js
var newDiv = document.createElement('div')
```
## 操作节点
节点相关的属性和方法大部分位于`Node`中,由于元素对象`Element`也继承了`Node`,因此有些方法看起来依然是通过元素对象调用的。
### nodeType / nodeName
获取节点的类型和名称。
```js
document.nodeType // => 9
document.nodeName // => #document
```
### firstChild / lastChild / childNodes
获取该节点的第一个 / 最后一个 / 所有子节点。
| 返回值 | 描述 |
| --- | --- |
| 节点对象 / `NodeList`集合 | 符合条件的节点 / 包含符合条件节点的伪数组 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var ul = document.querySelector('#list')
ul.childNodes.length // => 5
ul.firstChild.nodeName // => #text,因为是换行,而不是 <li>
```
### previousSibling / nextSibling
获取该节点的前一个 / 后一个兄弟节点。
| 返回值 | 描述 |
| --- | --- |
| 节点对象 | 符合条件的节点 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var first = document.querySelector('#first')
first.nextSibling.nodeName // => #text,是一个换行,而不是 <li>
```
### parentNode
获取该节点的父节点。
| 返回值 | 描述 |
| --- | --- |
| 节点对象 | 符合条件的节点 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var first = document.querySelector('#first')
first.parentNode // => <ul id="list">...</ul>
```
### cloneNode()
复制节点。与`createElement()`一样,复制后的节点需要使用`Node`中的相关方法插入到文档中才有效果。
| 参数 | 描述 |
| --- | --- |
| 布尔值 | 可选。默认值`false`为浅拷贝,表示仅复制该节点本身,不包括后代节点;`true`为深拷贝,表示连同后代节点一同复制 |
| 返回值 | 描述 |
| --- | --- |
| 节点对象 | 复制的节点 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
</ul>
```
```js
var list = document.querySelector('#list')
var newList = list.cloneNode(true) // 如果为 false,则不会拷贝 <li id="first">冥灯龙</li> 这个子节点
```
### appendChild()
将一个节点插入到父节点的最后,作为其最后一个子节点。如果要插入的节点是页面上已经存在的,那么该方法会**移动**该节点,而不是复制。
| 参数 | 描述 |
| --- | --- |
| 节点对象 | 要插入的节点 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
</ul>
```
```js
var list = document.querySelector('#list')
var first = document.querySelector('#first')
var newItem = first.cloneNode(true) // 必须设置为 true,否则无法克隆 冥灯龙 这个文本节点
list.appendChild(newItem)
// <ul id="list">
// <li id="first">冥灯龙</li>
// <li id="first">冥灯龙</li>
// </ul>
```
### insertBefore()
将一个节点插入到父节点的内部,作为子节点。
| 参数 | 描述 |
| --- | --- |
| 节点对象 | 要插入的节点 |
| 节点对象 | 父节点的一个内部节点,新节点将插入到该内部节点之前 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var list = document.querySelector('#list')
var newItem = document.createElement('li') // 创建新元素
var second = document.querySelector('#second')
list.insertBefore(newItem, second) // 将新元素插入到 second 之前
// <ul id="list">
// <li id="first">冥灯龙</li>
// <li></li>
// <li id="second">炎妃龙</li>
// </ul>
```
### removeChild()
移除当前节点中的一个子节点。
| 参数 | 描述 |
| --- | --- |
| 节点对象 | 要移除的节点 |
| 返回值 | 描述 |
| --- | --- |
| 节点对象 | 已移除的节点 |
```html
<ul id="list">
<li id="first">冥灯龙</li>
<li id="second">炎妃龙</li>
</ul>
```
```js
var list = document.querySelector('#list')
var second = document.querySelector('#second')
list.removeChild(second)
// <ul id="list">
// <li id="first">冥灯龙</li>
// </ul>
```
## 集合
为了保存多个元素 / 节点对象,DOM 提供了两个伪数组对象作为集合,它们与`Node`平级,是两个独立于`Node`的特殊对象。
### HTMLCollection
保存多个元素对象。
```js
p.children.length // HTMLCollection 的长度
```
### NodeList
保存多个节点对象。
```js
p.childNodes.length // NodeList 的长度
```
## 操作文本
文本对象`Text`本身也是节点之一,但是由于直接使用它的可能性不大,因此通常都是用元素对象`Element`的相关属性获取文本内容即可。
### textContent / innerText
获取元素内部的所有文本内容,也可以用来修改元素内容。它们会忽略内容中的 HTML 标签,返回只包含文字部分的内容,将 HTML 输出到页面也只会显示代码,不会被解析。
```html
<span id="msg">这里是<strong>文本内容</strong>啦</span>
```
```js
var msg = document.querySelector('msg')
msg.textContent // => 这里是文本内容啦
msg.innerText // => 这里是文本内容啦
```
### innerHTML
获取元素内部的所有 HTML 代码,也可以用来修改元素内容。它不会忽略内容中的 HTML 标签,将 HTML 输出到页面会将其解析。
```html
<span id="msg">这里是<strong>文本内容</strong>啦</span>
```
```js
var msg = document.querySelector('msg')
msg.innerHTML // => 这里是<strong>文本内容</strong>啦
```
## 操作属性
DOM 会自动为每个元素添加其标签上的属性(自定义属性除外),通过它们可以直接获取元素的属性值。
```html
<a id="test"></a>
```
```js
a.id // => test
a.href = 'http://www.example.com' // 修改属性值
```
对于布尔型的属性,也应该使用布尔型的值:
```js
input.disable = true
radio.checked = true
```
### className
获取元素的完整类名称。
```html
<div class="one two three"></div>
```
```js
div.className // => 'one two three'
```
### classList
不过完整的类属性字符串很难操作,因此 DOM 提供了`classList`可以获取一个类属性对象,它包含下列属性和方法:
| 名称 | 描述 |
| --- | --- |
| `length` | 获取类的数量,只读 |
| `add()` | 增加类 |
| `remove()` | 移除类 |
| `contains()` | 检查当前对象是否包含某个类 |
| `toggle()` | 如果某个类存在,则移除该类,否则添加该类 |
| `item()` | 返回指定索引位置的类,注意**没有方括号**形式 |
```js
div.classList // => { 0: "one", 1: "two", 2: "three", length: 3 }
div.classList.add('btn')
div.classList.add('btn', 'danger')
div.classList.remove('btn')
div.classList.toggle('btn')
div.classList.contains('btn')
div.classList.item(0) // 没有 div.classList[0] 这样的格式
```
### 自定义属性
使用 HTML 标准提供的`data-`前缀形式可以添加自定义属性:
```js
<div id="person" data-score="30"></div>
```
然后,通过元素对象的`dataset`属性,可以**读写**标签的所有`data-`属性:
```js
person.dataset.score // => 30
```
注意,`data-`后面的属性名不要使用大写字母,因为 HTML 不区分大小写。比如`data-helloWorld`应该写成`data-hello-world`。当它转成`dataset`的属性名时,会自动将短横线后面的字母转为大写,并移除短横线,反之也是一样。
```html
<div id="person" data-max-score="30"></div>
```
```js
person.dataset.maxScore // => 30
```
## 操作样式
元素样式相关的属性和方法均位于`Element`元素对象中。
### style
设置元素的**行内样式**,但不能操作定义在**样式表中的规则**。对于包含连字符的属性名称,需要变成驼峰式写法。
```js
div.style.width = '100px';
div.style.backgroundColor = 'orange';
```
### getComputedStyle()
获取元素计算后的最终样式。但是该对象获取的属性值是**只读**的,因此它只能获取样式,要设置样式只能使用`style`属性。
```js
getComputedStyle(div).width
```
### 元素大小
使用下列属性可以获取元素大小相关的信息,它们均**不包含单位**。
| 属性名 | 描述 |
| --- | --- |
| `scrollWidth` | **只读**,获取元素的内容宽度,包括溢出容器的部分。如果内容没有占满容器,那么与`clientWidth`一致 |
| `scrollHeight` | **只读**,获取元素的内容高度,包括溢出容器的部分。如果内容没有占满容器,那么与`clientHeight`一致 |
| `offsetWidth` | **只读**,获取元素以`border`为界的宽度 |
| `offsetHeight` | **只读**,获取元素以`border`为界的高度 |
| `clientWidth` | 只适用于**块级元素**,获取元素以`padding`为界的宽度 |
| `clientHeight` | 只适用于**块级元素**,获取元素以`padding`为界的高度 |
| `clientLeft` | 只适用于**块级元素**,获取元素`border-left`的大小 |
| `clientTop` | 只适用于**块级元素**,获取元素`border-top`的大小 |
```js
div.offsetWidth // => 300
```
### 滚动距离
使用下列属性可以获取元素滚动距离相关的信息,它们均**不包含单位**。
| 属性名 | 描述 |
| --- | --- |
| `scrollLeft` | **可读写**,获取元素内容水平方向滚动出去的距离,如果元素没有滚动条,那么该值为`0` |
| `scrollTop` | **可读写**,获取元素内容垂直方向滚动出去的距离,如果元素没有滚动条,那么该值为`0` |
| `pageXOffset` | **只读**,获取页面的水平滚动距离 |
| `pageYOffset` | **只读**,获取页面的垂直滚动距离 |
> 如果使用`scrollTop`获取整个页面滚动的话,在 Chrome 和 IE 中,必须使用`document.documentElement.scrollTop`才能正确获取,其它方式该值均为`0`。
```js
document.addEventListener('scroll', function() {
window.pageYoffset
document.documentElement.scrollTop
})
```
### 位置
使用下列属性可以获取元素位置相关的信息,它们均**不包含单位**。
| 属性名 | 描述 |
| --- | --- |
| `offsetParent` | **只读**,获取最靠近当前元素的,且 CSS 的`position`属性不等于`static`的父元素,直到`<body>`元素 |
| `offsetLeft` | **只读**,获取元素相对于`offsetParent`元素左侧的偏移距离 |
| `offsetTop` | **只读**,获取元素相对于`offsetParent`元素上方的偏移距离 |
> 注意,如果元素中包含未设置宽高的图片,那么由于代码执行时图片可能还没加载,从而导致无法正确获取元素距离顶部的高度
## 其它
除了上述通用属性和方法以外,有些特定元素还定义了独有的属性和方法。
### submit()
使用`HTMLFormElement`的`submit()`方法可以直接提交表单。
```js
form.submit()
```
### focus()
使用`HTMLElement`的`focus()`方法可以使元素获得焦点(通常用在表单控件元素上)。
```js
input.focus()
```<file_sep>---
title: 计算机网络笔记之01 😄 概述
date: 2020-01-15 15:12:48
abbrlink: gv4bco96
tags: 计算机网络
categories: 计算机网络
excerpt: 在当今社会,人们熟悉的网络有三种:电信网络、有线电视网络和计算机网络,随着技术发展,这三种网络正逐步在高层业务层面融合为一体,称为「三网融合」,本文主要讨论的是其中的计算机网络。
---
# 计算机网络笔记之01 😄 概述
网络的概念非常宽泛,比如电信网络、有线电视网络、交通网络、神经系统网络等等,而这里的计算机网络是通信技术和计算机技术相结合的产物。具体来说,计算机网络指的是一个将分散的、具体独立功能的计算机系统(如个人电脑、移动设备等),通过通信设备(如交换机和路由器)与线路(如双绞线、光纤、无线信号等)连接起来,由功能完善的软件(操作系统)实现资源共享和信息传递的系统。
总的来说,计算机网络是互连的(一定有通信线路相连)、自治的(无主从关系)计算机集合。
## 性能指标
要评价一个计算机网络的性能,主要通过下面七个性能指标。
### 速率
速率指的是**理想状态下**主机在网络上传输数据的速度,单位为`bit/s`(可简写为`b/s`或`bps`)。因为传输的数据均是`0`或`1`的比特位,因此速率也称为比特率。
当速度更快时,可以在`b/s`前加上单位,如`Kb/s`、`Mb/s`。注意,它们之间的换算关系**与内存中**的换算有所不同:
- $1Kb/s$ = $10^3b/s$
- $1Mb/s$ = $10^6b/s$
- $1Gb/s$ = $10^9b/s$
### 带宽
带宽指的是网络所能传输的最高速度,单位同样为`bit/s`。它原本是通信中表示频带宽度的术语,如果频谱超过了最大频带,那么就会出现问题,但是在计算机网络中并不存在这样的问题,因此基本可以与速率等同。实际当中更多使用的也是带宽这一概念,是评测一个网络质量的重要指标之一。
平时购买网络设备时,所标识的千兆、百兆端口指的均是带宽;在系统中查看当前网卡的速度,也同样指的是带宽。

> 运营商提供的宽带业务通常会标明宽带速率是`200M`,全称应该是`200Mbit/s`。但是在实际使用时,最高下载速度只有`20MB/s`。这是因为运营商是以网络传输中的`bit`为计量单位,但是在计算机中则是以字节`Byte`(`1Byte`=`8bit`)为计量单位,因此通常将宽带运营商的数据除以 8 才是理论上最高的速度(考虑到各种其它因素,差不多除以 10 才是真正能达到的最高速度)。
### 吞吐量
吞吐量指的是单位时间内通过某个网络(或信道、接口)的数据量,即网络当前的实际速度,单位同样为`bit/s`。也就是说,虽然网络可以承载的最大速度(带宽)可能为`1Gbps`,但实际速度可能只有`200Mbps`。
### 时延
时延指的是数据从网络的一端到另一端所需要的时间,也称为延迟,是评测一个网络质量的重要指标之一。
时延由发送时延、传播时延、处理时延和排队时延四部分组成,其中最重要的是发送时延和传播时延。
$$ 时延 = \textbf{发送时延} + \textbf{传播时延} + 处理时延 + 排队时延 $$
#### 发送时延
发送时延指的是设备从发送数据的第一个 bit 起,到最后一个 bit 离开设备为止,所需要的时间。由于发送时延仅存在于网络适配器(如路由器、交换机)中,因此发送时延仅与数据帧长度以及设备的发送速率(即带宽)有关。
$$ 发送时延 = \frac{数据帧长度}{发送速率} $$

> 发送时延也称为传输时延,但是这个名词非常容易与下面的传播时延搞混,因此不建议使用。
#### 传播时延
传播时延指的是数据在信道中传播所需要的时间。由于信道介质的传播速率基本是固定的(铜线约为$2.3 × 10^5 km/s$,光纤约为$2.0 × 10^5 km/s$),因此影响传播时延的主要因素就是信道的长度,也就是通讯双方的距离。
$$ 传播时延 = \frac{信道长度}{传播速率} $$

> 光纤的传播速率其实比铜线要低,但是由于光纤的发送时延小,因此通常使用光纤。
#### 排队时延和处理时延
排队时延指的是通信量较大时,数据在路由器等候的时间(比如在机场排队等待安检的时间)。处理时延指的是主机或服务器收到数据帧后处理如差错校验、提出数据部分、查找路由的时间(比如在安检时被检查的时间)。这两种时延在网络正常情况下所占的比重比较小,而且情况复杂不容易计算,因此不作重点。
### 时延带宽积
将传播时延和带宽相乘,就得到了另外一个概念,时延带宽积。它表示信道可以同时容纳多少个 bit,可以理解成信道的体积。由于传播时延越大,意味着信道越长;而带宽越大,说明信道越宽(从数据角度,而非实际的通信线路),因此时延带宽积也称为以比特为单位的信道长度。
### 往返时间
往返时间(RTT)指的是数据双向交互一次所需要的时间。假设 A 与 B 通信,A 向 B 发送了数据,而且需要等待 B 接收后返回确认信息后,A 才能发送下一个数据,那么这之间就会产生 A 的等待时间,也就是往返时间。往返时间包含**2 倍的传播时延**、末端处理时间(如排队时延和处理时延),以及 B 在接收数据后的处理时间,是一个判断网络性能的综合指标。
> 使用命令行的`ping`命令查看的便是往返时间。
### 利用率
利用率指的是信道或网络的数据通过率。就好比是现实中的公路一样,利用率并不是越高越好,当利用率到达一定程度后,就会出现排队、延迟等网络性能问题(堵车),但是利用率太低,又会造成资源浪费。
## 网络体系结构
要保证数据的正确传输,需要非常多的协调工作,例如检查通信线路是否连通、如何寻找目的主机、检查对方主机是否做好接收数据的准备、如何解决可能出现的差错和意外等等问题。为了解决这些问题,早在计算机网络诞生初期就诞生了「分层」的概念,这样可以将庞大复杂的问题转化为若干较小的局部问题,便于研究和处理。
在分层结构中,有一些概念需要提前了解:
- 实体:任何可以发送或接收信息的软硬件,发送端和接收端同一层的实体称为对等实体
- 协议:**对等实体**之间进行通信的规则(因此不同层之间是没有协议一说的),也就是说双方必须按照一定约定来通信,才能保证数据可以被正确理解和传送。具体来说,它包括:
- 语法:规定了「如何讲」,即数据格式、编码方式等
- 语义:规定了「讲什么」,即控制信息的含义和功能
- 时序:规定了各种操作的顺序
- 接口:也称为访问服务点SAP,是上下层之间交互的入口
- 服务:下层为上层通过接口提供的功能调用,即**下层为上层提供服务**
在网络体系结构中,实际要传递的数据称为**服务数据单元 SDU**(Service Data Unit)。在数据经过每一层时,该层都需要为数据加上一些控制信息,这些控制信息称为**协议控制信息 PCI**(Protocol Control Information)。SDU 和 PCI 合并到一起,就会作为对等层次之间传递的数据,称为**协议数据单元 PDU**(Protocol Data Unit)。
### OSI 参考模型
国际标准化组织 ISO 在 1984 年提出了 OSI 参考模型(开放系统互连参考模型),但是由于缺乏实际经验、设计复杂、标准指定时间过长等原因,该模型只有理论上的指导作用,没有产生任何的商用产品。
它将网络分为了七层,从上到下依次为应用层、表示层、会话层、传输层、网络层、数据链路层、物理层。当一个主机需要向另外一台主机发送数据时,数据会由应用层开始,依次打上本层的头部控制信息,然后交给下一层,直到数据链路层为止(数据链路层除了添加头部信息,还需要添加尾部信息)。而物理层则会将数据链路层的信息转换成比特流进行传输。
当经过中间路由时,路由会将其拆开以寻找传递的下一站。注意,这些通信子网的设备(交换机、路由器等)只拥有低三层的功能,而没有资源子网设备(PC 终端、手机等)高三层的功能。
到达目的主机后,它会将数据层层拆分,直到最后的数据。这一系列过程就好比寄快递一样,发送方需要将货物包装、塞上填充物、放到箱子中、贴上快递单,而经过层层转交之后,到达收件人手中,再将箱子打开、取出填充物、打开包装,最后获得货物。而这些中间步骤均是为了使货物能够安全、准确的到达,虽然与货物本身没有关系,但也是必不可少的。

### TCP/IP 体系结构
目前计算机网络中实际使用的分层模型是 TCP/IP 体系结构,它是一个由很多协议组成的协议栈,由于其中的 TCP 和 IP 协议最为常用,才由此得名。
TCP/IP 体系结构将网络分为了四层,从上到下依次为应用层、传输层、网络层、网络接口层。但是它的网络接口层几乎没有任何实质内容,为了讲解方便,这里借用 OSI 中的物理层和数据链路层,将它们对应到网络接口层中。因此在理论上我们将网络划分为五层,但实际使用中依然是 TCP/IP 的四层。

<file_sep>---
title: 大前端笔记之23 🐈 Git
date: 2020-02-21 21:17:12
abbrlink: wmjtf4cp
tags:
categories: 大前端
excerpt: Git 是目前最主流的分布式版本管理系统,它可以将项目多个不同的版本状态保存起来,以便随时取回之前的版本进行重写或查阅。
---
# 大前端笔记之23 🐈 Git
Git 是目前最主流的分布式版本管理系统,它可以将项目多个不同的版本状态保存起来,以便随时取回之前的版本进行重写或查阅。
既然有分布式,那么与之相对的便是集中式,比如 SVN。集中式版本管理是将版本库保存在一个统一的中央服务器,其它用户都需要通过网络连接中央服务器,如果要修改项目,则需要先从中央服务器获取到版本库的文件,修改完成后再提交回去。而分布式版本管理则是将版本库保存到用户本机上,如果有需要,则只需两个人同步,就能获取到对方更新后的版本了。
## 安装与配置
首先在[官网](https://git-scm.com/downloads )下载对应版本的 Git,根据提示安装即可。
打开任意文件夹,右键选择`Git Bash Here`,打开 Git 的命令行窗口,输入`git --version`,如果出现版本号信息(如`git version 2.24.0.windows.2`),则说明安装成功。
在第一次使用 Git 之前,需要先配置提交人的姓名和邮箱。其中的`--global`表示全局配置,即本地所有的仓库都采用该配置。
```powershell
git config --global user.name 小鸡米花 # 配置姓名
git config --global user.email <EMAIL> # 配置邮箱
git config --list # 查看所有配置信息
```
## 提交修改
在使用 Git 之前,需要先明确几个概念。首先,用户编辑文件的项目目录称为**工作区**,也就是电脑上能直接看到的目录。当文件发生修改后,需要先将文件添加到**暂存区**,如果有多个文件就可以添加多次,它类似于购物车,当你选好所有的商品后一并付款,这样可以将多个文件的变更统一作为一个版本修改。当本次暂存区的所有文件修改完毕后,将它们一并提交到**仓库**,进行持久化保存。
### 初始化仓库
在项目文件夹下使用`git init`命令初始化仓库,此时项目文件夹会出现一个`.git`的隐藏文件夹。它是当前项目的版本管理仓库,暂存区和仓库都保存在这里。
### 添加到暂存区
在项目文件夹下随便新建一个文本文件`index.html`,然后使用命令`git add index.html`告诉 Git 跟踪该文件,也就是将该文件添加到暂存区。
### 提交到仓库
使用`git commit -m 提交内容`将跟踪的文件提交到仓库保存,也就是将暂存区的文件提交到仓库。Git 会提示提交的文件数量,新增和删除了多少行数据。其中`-m`之后表示这次提交的内容,**必须写**清楚以便之后查阅。否则会弹出一个文件要求输入提交内容,关闭文件之后才会继续提交操作。
> 注意,版本管理工具不能跟踪二进制文件内容的变化,只能跟踪它们的大小或日期。
### 查看仓库状态
然后对文件进行一些修改,使用`git status`命令,可以查看当前仓库的状态。其中第一行括号提示文件没有被跟踪,以红色的文件名标识;第二行括号提示`index.html`文件发生了修改,但是还没有被提交。
接下来继续使用`git add index.html`和`git commit -m "第一次修改"`提交修改,再使用`git status`命令,便提示没有需要提交的内容了。
### 比较差异
使用`git diff`命令可以查看工作区文件与暂存区文件有哪些不同。使用`git diff --cached`命令可以查看暂存区文件与仓库文件(最近的一次提交)有哪些不同。
## 时光机穿梭
### 查看提交记录
使用`git log`命令可以查看所有的仓库提交记录,每条记录以一长串的字符作为本次提交的`id`,下面是提交的作者和时间。如果感觉信息太乱,可以使用`git log --pretty=oneline`命令隐藏作者和时间,使其显示在一行。
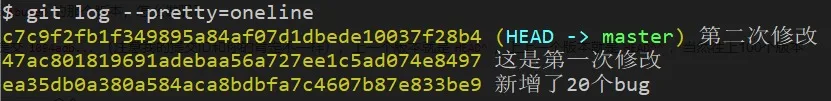
第一条记录后面括号中的`HEAD`表示这是最近的一次提交(后面的`master`表示主分支,见下文)。
> 如果记录显示后控制台不能操作,按下<kbd>Q</kbd>即可。
### 回退版本
如果希望回退到上一个的版本,使用`git reset --hard HEAD`命令,其中`HEAD`表示上一个版本,`HEAD^`表示上上个版本。当版本回退之后,使用`git log`命令就无法再查看最新的版本了(比如`c7c9f2`开头的这条就消失了),如果要回到这条版本,则需要使用`git reset --hard c7c9f2`命令,后面的编号为记录编号,不需要写全,通常取前 6 位即可。
但是,如果关闭了控制台,那么新版本编号就看不到了,此时需要使用`git reflog`命令查看指令输入的历史记录,从中就可以找到对应的编号了。
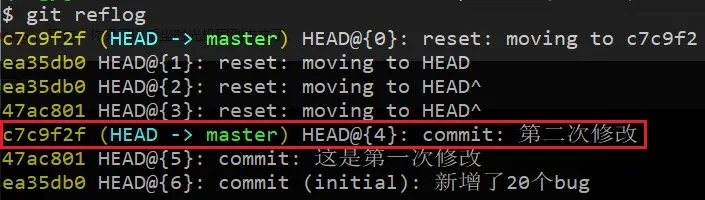
### 撤销修改
如果修改了文件内容,还没有提交到暂存区,那么使用`git checkout -- 文件名`可以撤销工作区的修改。
如果修改了文件内容,而且提交到暂存区了,那么使用`git reset HEAD 文件名`可以将暂存区恢复为最近的一次提交,这样就回到了第一步。
如果修改了文件内容,而且不但提交到了暂存区,也提交到了本地仓库,那么直接回退版本。
如果修改了文件内容,而且还提交到了远程仓库(见下文),那么就等着 GG 吧。
### 删除文件
如果要删除工作区的文件,那么可以使用`git rm 文件名`命令,该命令会自动将删除操作同步到暂存区。如果已经先手动删除了文件,那么也需要使用该命令将操作同步。接下来继续使用`git commit`命令提交修改到仓库,那么该文件就会从仓库中被删除。
如果要恢复被删除的文件,与上文撤销修改其实是一样的。
### 忽略文件
实际开发中由于一次添加的文件比较多,如果手动去挨个添加会比较繁琐。Git 提供了忽略文件清单的功能,在项目文件夹下新建一个后缀为`.gitignore`的文件(不需要文件名),在其中写明要忽略的文件或文件夹(如果省略了后缀名,那么 Git 会将其识别为文件夹),那么 Git 就不会跟踪这些文件了。
```powershell
# .gitignore
test.html
public/test_images
```
接下来,只需要使用`git add .`命令就可以将所有文件添加到暂存区了。
## 分支
之前的操作都是在一条时间线上完成的,这条时间线被称为**主分支**(master)。分支可以使不同的工作分开进行,例如开发分支、功能分支、Bug 修复分支等。
严格说来,`HEAD`直接指向的并不是最近的一次提交,而是指向的`master`,而`master`指向的才是最近一次提交。
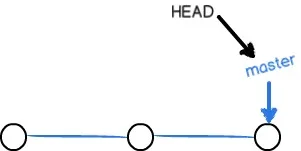
当新的分支创建后,Git 会创建一个新的指针,指向当前最近一次提交,并将`HEAD`指向该分支指针。
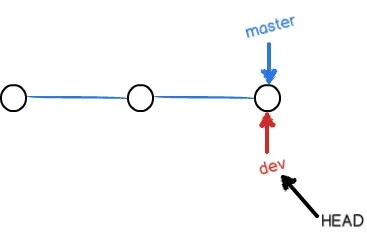
接下来,对工作区的操作就会针对该`dev`分支了,当再次提交新版本时,`master`依然指向原来的版本,而`dev`则指向了新提交的版本。
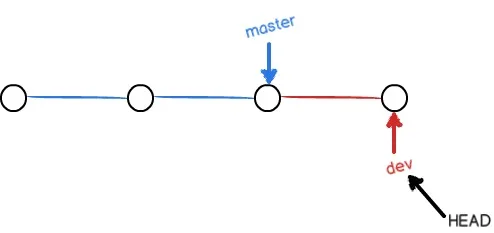
如果`dev`的工作完成了,就可以将该分支合并到主分支上,所谓合并,其实将`master`重新指向最新版本,并将`HEAD`指向它。
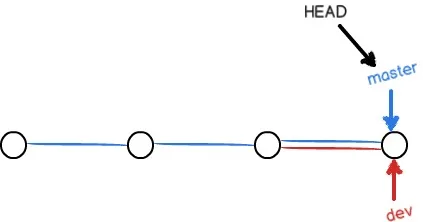
最后,只需要删除掉`dev`指针,那么就只剩下一条主分支了。
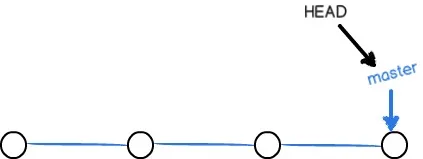
使用`git branch`命令可以查看目前所有的分支。分支前的`*`和绿色字体表示这是当前分支。
使用`git branch dev`命令可以基于当前分支创建一个`dev`分支。
使用`git checkout dev`命令或者`git switch dev`可以切换到`dev`分支。
上述两条命令可以简写成`git checkout -b dev`或者`git switch -c dev`一条。
> 由于`checkout`命令既用于撤销修改,又用于切换分支,容易令人迷惑,因此新版本中添加了`switch`命令用来切换分支。
使用`git merge dev`命令可以将当前分支与`dev`分支合并。
使用`git branch -d dev`命令可以将`dev`分支删除,不过要求该分支必须被合并,防止误删。使用`git branch -D dev`命令可以强制删除分支。
## GitHub
上述的所有操作都是在本地仓库完成的,如果需要与其他人协作,那么就需要一个公共区域供大家下载和推送自己本地的代码。GitHub 是目前世界上最流行的远程仓库和多人协作开发平台,开发者 A 可以将本地的仓库推送到 GitHub 上的远程仓库,然后开发者 B 就可以将其拉取下来,在本地开发完成后再推送到远程仓库,这样就可以完成协作了。
在 GitHub 上注册账号后,需要先创建一个远程仓库(repository)。点击导航栏右上角的加号,选择`New repository`即可打开创建新仓库的页面。在页面中填写项目名称`test`,选择默认的`public`即公开权限,点击下面的绿色按钮创建即可。
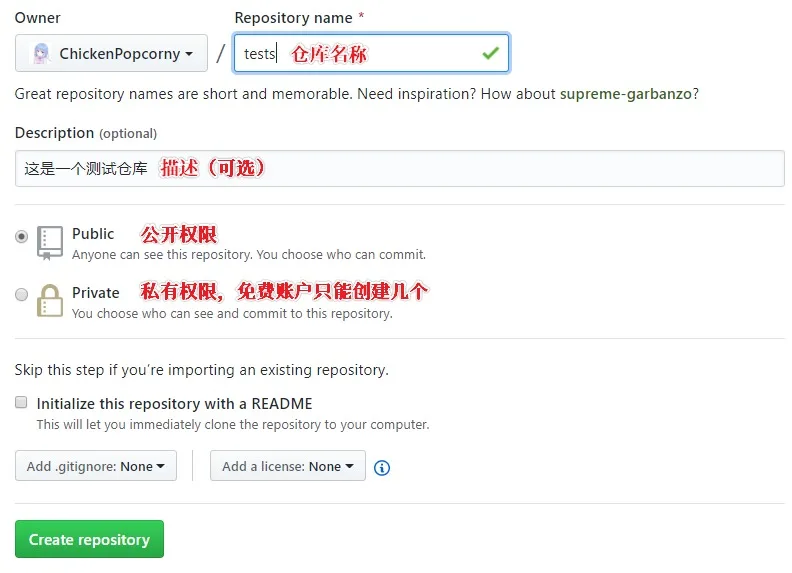
### 推送本地仓库
项目管理者将仓库创建完成后,进入该仓库页面。首先,使用`git push 远程仓库地址 分支名称`命令(如`git push https://github.com/ChickenPopcorny/test.git master`)将本地仓库推送到远程仓库。这时会弹出一个窗口,提示输入该账号的用户名和密码(之后系统会将其保存起来,不需要再次输入)。推送完成后,在远程仓库页面刷新,就可以看到推送的文件了。
但是每次粘贴地址比较麻烦,因此可以先使用`git remote add origin https://github.com/ChickenPopcorny/test.git`命令为远程地址起一个别名`origin`(自定义的,可以修改),然后再使用`git push origin master`命令就可以了。
如果使用`git push -u origin master`命令,那么表示将当前远程地址和分支作为临时的默认推送地址,那么之后只需要使用`git push`即可。
### 克隆远程仓库
其它合作开发者使用`git clone https://github.com/ChickenPopcorny/test.git`命令可以将远程仓库克隆到本地仓库。克隆命令不需要权限,因为仓库的权限是公共的。
### 邀请合作开发者
合作开发者克隆仓库后,并不能直接使用`push`命令将修改推送到远程仓库,因为没有权限。管理者需要在项目的设置页面添加想要邀请的合作开发者,而且合作开发者需要接受邀请后,才能使用`push`命令推送修改。
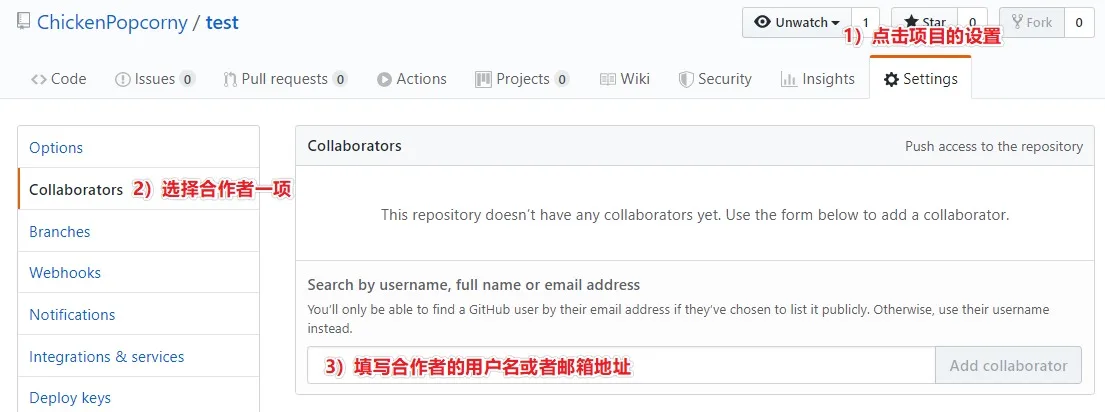
### 拉取最新版本
合作开发者提交版本后,管理者可以通过`git pull 远程仓库地址 分支名称`命令将最新版本拉取到本地仓库并自动合并。与克隆命令不同的是,克隆命令是在本地仓库不存在的情况下使用,而后续只需要使用拉取命令即可。而且,如果远程仓库版本高于本地,那么本地是不能推送的,必须先使用`pull`命令获取最新版本,才能进行推送。<file_sep>---
title: 大前端笔记之10 📱 CSS 移动端
date: 2020-01-26 9:15:55
abbrlink: x3i0w12c
tags: CSS
categories: 大前端
excerpt: 随着移动设备的普及,Web 开发已经不仅仅是 PC 端的问题了,因此 CSS 提出了很多概念来适应不同的移动设备。
---
# 大前端笔记之10 📱 CSS 移动端
随着移动设备的普及,Web 开发已经不仅仅是 PC 端的问题了,因此 CSS 提出了很多概念来适应不同的移动设备。
## 基本概念
### 英寸
英寸(inch)一般用来描述屏幕的物理大小,如电脑显示器的`17`、`22`,手机显示器的`4.8`、`5.7`等使用的单位都是英寸,而且它们指的是屏幕对角线的长度。

> inch 在荷兰语中的本意是大拇指,指的是大拇指关节处的宽度,即 1 英寸 = 2.54 厘米。
### 物理像素
物理像素是设备屏幕的真实物理单元,可以简单理解为一组三色的 LED 灯就构成了一个物理像素,如果把屏幕锯掉一半,那么物理像素也就只有之前的一半了。
例如苹果官网上对于各型号手机**像素分辨率**的说明,指的就是物理像素。

### PPI
但是物理像素越高,并不代表屏幕越清晰,因为还得看这些像素放在多大的地方,也就是屏幕的尺寸。于是,只有通过 PPI(Pixel Per Inch,像素每英寸)才可以描述屏幕的清晰度,PPI 越高,屏幕才越清晰(即使屏幕可能会更小),反之则越模糊。
下面是苹果官网对于 XS Max 和 SE 两款设备的说明,可见前者的屏幕是要比后者更加清晰的。
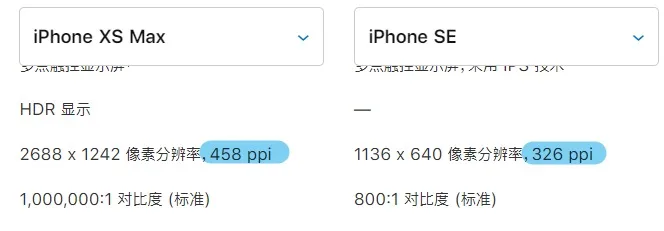
PPI 的计算方式也比较简单,只需要将对角线的物理像素数除以对角线的英寸数即可:
$$ PPI=\frac{\sqrt{水平物理像素^2+垂直物理像素^2}}{对角线英寸} $$
如 iPhone SE 的像素分辨率为`1136 × 640`,屏幕大小为`4`英寸,那么它的 PPI 为:
$$ PPI=\frac{\sqrt{1136^2+640^2}}{4}=\frac{1303.88}{4} \approx 326 $$
### 设备独立像素
上面所有的概念都是以物理像素为基础的,但是随着智能手机的发展,人们开始使用手机来阅读文章或图片。如果一段同样的内容,在分辨率`320 × 480`的手机上大小算作正常的话,那么在分辨率为`640 × 960`的手机来看,则会比前者小了一半,原因就是后者的物理像素更多,渲染同样大小的内容只需要更小的空间。可以想象按这样的情况发展,那么更大分辨率的设备就几乎不可能看清上面的内容了。
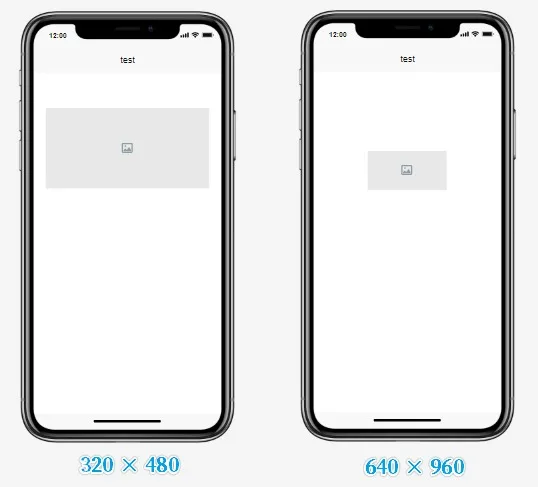
各个移动设备厂商不可能将这个问题置之不理。乔布斯在 2010 年 iPhone4 发布会上首次提出了 Retina Display(视网膜屏幕)的概念,完美解决了这一问题。

其大致原理是,通过算法将之前的 1 个物理像素通过 4 个物理像素来渲染(指平面上,而非单个方向)。于是原本正常大小的内容不会被缩小,而是变得更加精细了。
也就是说,在上文的例子中,假如内容宽度为`300px`,那么左侧手机依然使用`300px`渲染它,而右侧的手机则会使用`600px`来渲染。于是,这里提出了一个新的概念设备独立像素(Device Independent Pixels,简称 DIP)来描述内容的宽度。由此可见,在 CSS 中使用的单位`px`指的**并不是物理像素,而是设备独立像素**。
### DPR
为了定义这种渲染的比例,又规定了在水平方向或垂直方向上,**物理像素与设备独立像素的比值,称为 DPR(Device Pixel Ratio)**。
如上文中 iPhone4 在水平或垂直方向上将 1 个物理像素通过 2 个物理像素来渲染,因此它的 DPR 就是`2 / 1 = 2`。通过在 Chrome 开发工具中选择不同的设备,然后使用 JavaScript 提供的`window.devicePixelRatio`可以直接查看当前设备的 DPR。
并且,设备名称右侧的分辨率也是以**设备独立像素**为单位的,因此将这个分辨率乘以该设备的 DPR 得到的就是设备的物理像素。以 iPhone6 为例,它的 DPR 为 2,设备独立像素分辨率为`667 × 375`,将其乘以 2 得到的就是物理分辨率`1334 × 750`。
> 注意,有的设备如 iPhone 6 / 7 / 8 Plus 计算的结果并不准确,这是由于苹果通过另外的技术将更多的设备独立像素塞进了物理像素中,不必在意。
后来,各个 Android 厂商也纷纷通过另外的技术实现了类似于苹果的视网膜屏幕,开发了 DPR 高于 1 的设备。
### 多倍图
DPR 还带来了另外一个问题,虽然文字或者图形是通过 CSS 渲染出来的,可以任意放大或缩小,但是如果是固定像素的图片被放大了,那么就会出现模糊的情况。
为了解决这个问题,通常会将原本`50 × 50`图片,先放大一倍也就是变成`100 × 100`,然后在 CSS 中使用`width`或`background-size`将其缩小到`50 × 50`。这样原本不需要缩放的 PC 端依然按照原样显示,而移动端虽然会将其放大,但是图片本身就是`100 × 100`的大小,也不会变的模糊。
```css
img { /* 原图大小 100,实际显示 50 */
width: 50px;
}
div { /* 原背景大小 100,实际显示 50 */
background-size: 50px;
}
```
对于 @2x 的精灵图来说,除了要缩小宽高,定位的位置同样也要缩小一半。
```css
/* 精灵图原大小为 400 × 400,原位置在 -166px 0 */
div {
background: url(sprite.png) no-repeat -83px 0 / 200px;
}
```
### 视口
视口指的是浏览器窗口上能看到页面内容的区域。在移动设备上,由于设备屏幕通常比较窄,因此会使用一种**虚拟视口**(virtual viewport)。它通常比屏幕尺寸要宽,然后将渲染的页面放到虚拟视口中,经过收缩之后显示到屏幕上,从而使用户可以一次性看到整个页面。例如,移动设备的屏幕宽度为`375px`,虚拟视口宽度为`980px`,页面宽度也是`980px`,那么页面会正好完整的显示在`375px`大小的屏幕内。
```css
div {
border: 2px solid #000; /* 为了看清元素的边界,加了 2px 边框 */
width: 976px;
}
```

> 虚拟视口的宽度根据不同的移动设备会有所不同,Safari 通常为 980px,Opera 为 850px,Android 的 WebKit 为 800px,而 IE 为 974px。
但是这样也存在弊端,`16px`的文字在`980px`的区域中,显示在手机屏幕这么小的范围,虽然不会出现滚动条,但是文字会变得非常小,根本没法看清,用户打开每个页面都必须手动缩放到合适的大小才能查看。并且,这样的方式也没法使用媒体查询技术,因为对于页面来说,它认为所有的设备都是`980px`,没法针对更小的宽度作出更好的响应。
为了解决这个问题,Safari 提供了一个视口标签,使页面开发者可以根据需要自己设定虚拟视口的宽度。
```html
<meta name="viewport" content="width=1200">
```
此时会发现之前`980px`的元素并不能占满整个窗口了。

但是,由于移动设备屏幕大小不一,这样的固定视口宽度并没有很大用处。因此,通常会将视口宽度设置成与设备宽度相同。该标签除了宽度以外,还可以设置一些其它的参数:
```html
<meta name="viewport" content="width=device-width, initial-scale=1.0, user-scalable=no, maximum-scale=1.0, minimum-scale=1.0">
```
| 参数 | 描述 |
| --- | --- |
| `width=device-width` | 将视口宽度设置为设备宽度 |
| `initial-scale=1.0` | 设置初始的缩放比率为`1`,即没有缩放 |
| `maximum-scale=1.0` | 设置最大的缩放比率为`1` |
| `minimum-scale=1.0` | 设置最小的缩放比率为`1` |
| `user-scalable` | 不允许用户缩放页面,但是在 iOS10 之后的 Safari 中无效 |
不过这样的弊端也是有的,如果是 PC 端固定宽度的页面,放在较小宽度的视口内,那么就只能显示其中的一部分,用户只能通过进度条才能看到页面其它内容。于是,为了达到理想效果,还需要结合自适应布局、响应式布局、媒体查询甚至针对不同大小重新设计页面布局才可以达到完美的效果。
> *参考资料*
>
> * [Using the viewport meta tag to control layout on mobile browsers](https://developer.mozilla.org/en-US/docs/Mozilla/Mobile/Viewport_meta_tag)
## 媒体查询
媒体查询可以使浏览器根据不同的设备类型(比如打印机、屏幕)或者参数(视口宽度)使用不同的样式。
在样式表中通过`@media`可以使用媒体查询:
```css
@media screen and (max-width: 500px) {}
```
或者在`<link>`和`<style>`中使用`media`属性:
```html
<link rel="stylesheet" href="500.css" media="screen and (max-width: 500px)">
<style media="screen and (max-width: 400px)"></style>
```
媒体查询的语法由媒体类型和媒体特性构成,之间使用逻辑运算符相连。
| 媒体查询语法 | 描述 |
| --- | --- |
| 媒体类型 | 即设备的类型,包括全部`all`、打印机`print`和屏幕`screen`,如果省略则默认为`all` |
| 媒体特性 | 必须由**圆括号**`()`包裹,包括`width`和`height`,前面可以加大于等于`min-`或小于等于`max-` |
| 运算符 | 包括逻辑且`and`、逻辑非`not`和逻辑或`,` |
注意,媒体查询的样式也是具有层叠性的,例如:
```css
@media (min-width: 400px) {
body { background-color: #333; }
}
@media (min-width: 300px) {
body { background-color: #66ccff; }
}
```
在上面的代码中,由于后者的区间`>=300`包含了前者的区间`>=400`,因此在进行判断时,后者的样式总是会生效。如果要实现`>=400`显示灰色,`300 ~ 400`之间显示蓝色的效果的话,那么必须将两者的顺序调换过来。
## 自适应布局
多数情况下,网页的设计稿只有一份(一般以`750px`为准),因此最简单直接的方案就是在不同的屏幕大小下,都以同样的方式**等比缩放**显示,因此在大屏下看到的文字和元素也更大,这样的方式称为**自适应布局**。不过,虽然在宽度上可以借助百分比单位、弹性布局等多种方式来实现自适应效果,但是高度和文字大小等部分却并没有这么简单。
以京东移动端为例,可以看到同一个底部链接在 iPhone5 和 iPhone6P 下的高度是不同的。

### rem 实现方式
最初实现自适应布局是通过相对长度单位`rem`,它相对的是根元素`<html>`的`font-size`。例如,`<html>`的`font-size`为`14px`,那么`2rem`就是`28px`。与之类似的`em`是相对于本元素的`font-size`(包括从父元素继承的)。
然后将各个元素的大小都设置为`rem`,再使用媒体查询动态调整`<html>`的`font-size`,就能使相关的元素随之调整。
```css
@media (max-width: 414px) { html { font-size: 41.4px } }
@media (min-width: 320px) { html { font-size: 32px } }
div {
font-size: .5rem;
height: 1rem;
line-height: 1rem;
}
```
为了实现等比缩放,那么必须规定一个统一的**缩放比率**,才能确保不同尺寸下缩放效果相同。例如,假如`750px`设计稿中的`font-size`为`75px`,即缩小十分之一,那么在`320px`下的`font-size`则必须为`32px`,才是同样缩小十分之一。因此,将需要适应的尺寸乘以该比例,得出的就是当前尺寸下的`font-size`。
```css
@media (max-width: 750px) {
html { font-size: 75px; /* 750px / 10 */ }
}
@media (max-width: 320px) {
html { font-size: 32px; /* 320px / 10 */ }
}
```
> 常见的尺寸有`320px`、`360px`、`375px`、`384px`、`400px`、`414px`、`424px`、`480px`、`540px`、`720px`、`750px`,更高尺寸的屏幕以`750px`即设计稿原始大小为准即可。
接下来,假如一个元素在`750px`设计稿中的高度为`100px`,由于此时`1rem = 75px`,那么它的高度`100px`换算成`rem`应该是:
```less
div { height: 1.33rem; /* 100 / 75 */ }
```
如果屏幕大小缩小一半为`375px`,那么计算此时的高度为`1.33rem = 1.33 × 37.5px = 49.8px`,基本为原大小的一半,可见结果是正确的。
> 淘宝基于这种原理开发了一款插件[flexible.js](https://github.com/amfe/lib-flexible),可以节省手写媒体查询的步骤。但是由于`rem`的局限性以及`vw`兼容性的提高,这款插件已经停止开发。
### vw 实现方式
相对长度单位`vw`与`rem`类似,但是它是根据视口宽度来计算的,表示**视口宽度除以`100`**。比如视口的宽度是`750px`,那么`1vw`就相当于`7.5px`。
它的实际使用方式同样与`rem`类似,以大多数设计稿的尺寸`750px`为例,`1vw`就相当于`7.5px`。因此,一个高度为`100px` 的元素,换算后的结果为:
```less
div { height: 13.33vw; /* 100 / 7.5 */ }
```
可以看到这种方式相当于省略了媒体查询,将单位直接与视口挂钩,使用起来更加方便,完全可以取代`rem`方式。此外,如果觉得手动计算比较麻烦,可以使用插件,比如 VSCode 的 px2vw。
### 响应式布局
虽然在不同的屏幕大小上直接等比缩放比较方便,但是这样的方式其实有些简单粗暴了。更加理想的是,**屏幕越大看到的内容应该越多,而不是将字变得更大**。
因此,针对不同的屏幕大小,可以给出多份设计稿(通常包含 PC、手机端、平板三种尺寸)然后开发人员借助媒体查询,可以使元素在不同设备上呈现特定的效果(比如在移动端隐藏侧边栏、改变导航栏布局等),这样的方式称为响应式布局。
下面是 Bootstrap 中给出的屏幕尺寸,可以作为参考:
| 设备 | 尺寸 | 容器宽度 |
| --- | --- | --- |
| 手机 | `< 768px` | `100%` |
| 平板 | `768px ~ 991px` | `750px` |
| 小型桌面显示器 | `992px ~ 1199px` | `970px` |
| 大型桌面显示器 | `≥ 1200px` | `1170px` |
```css
.container {
width: 1280px; /* 默认的 PC 端 */
}
@media (max-width: 1199px) { /* 小型显示器 */
.container { width: 970px; }
}
@media (max-width: 991px) { /* 平板 */
.container { width: 750px; }
}
@media (max-width: 767px) { /* 手机 */
.container { width: 100%; }
}
```<file_sep>---
title: 大前端笔记之08 💨 CSS 动画
date: 2020-01-23 23:27:28
abbrlink: sa91bwec
tags: CSS
categories: 大前端
excerpt: CSS3 中提供了原生的动画效果,相比于之前使用 JavaScript 操作 DOM 实现的动画效果,它的性能更高。
---
# 大前端笔记之08 💨 CSS 动画
CSS3 中提供了原生的动画效果,相比于之前使用 JavaScript 操作 DOM 实现的动画效果,它的性能更高。
## 过渡
过渡可以使元素从一种状态平滑的过渡到另外一种状态。
### 过渡属性
首先,在要过渡的元素上添加`transition-property`属性,指明要过渡的属性。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `all` |
| 可选值 | 描述 |
| --- | --- |
| `all` | 表示所有可过渡的属性(并不是所有的属性都可以被过渡) |
| 属性名 | 要过渡的属性,如果有多个属性以逗号分隔 |
```css
div { transition-property: width, height; }
```
### 过渡时间
使用`transition-duration`设置过渡的持续时间。**该属性不能省略**,因为它的默认值为`0s`,即没有过渡效果。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0s` |
| 可选值 | 描述 |
| --- | --- |
| 时间值 | 表示过渡的持续时间 |
```css
div { transition-duration: .3s; }
```
### 调速函数
使用`transition-timing-function`设置过渡的调速函数。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `ease` |
| 可选值 | 描述 |
| --- | --- |
| `ease` | 中间速度快,开始和结束速度慢 |
| `ease-in-out` | 中间速度慢,开始和结束速度快 |
| `ease-in` | 开始时速度慢,然后逐渐加速 |
| `ease-out` | 开始时速度快,然后逐渐减速 |
| `linear` | 匀速 |
| `steps()` | 将平滑的动画切割成[逐帧动画](/posts/m69g0i43.html)显示 |
```css
div { transition-timing-function: linear; }
```
### 延迟时间
使用`transition-delay`设置过渡的延迟时间。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0s` |
| 可选值 | 描述 |
| --- | --- |
| 时间值 | 表示过渡的延迟时间,`0s`表示立刻开始 |
### 过渡属性简写
使用`transition`简写上述所有属性。除了`transition-duration`之外,其它值均可以被省略,而且顺序也不重要。不过,如果有两个时间值,则前者表示持续时间,后者表示延迟时间;如果只有一个时间值,那么它表示持续时间。如果要同时定义多个不同的过渡属性,那么使用逗号将其隔开。
```css
div {
transition:
width 3s ease 0.5s,
height 1s ease 0s;
}
```
## 关键帧动画
过渡只能实现两种状态之间的变化,如果需要更复杂的动画效果,CSS 还提供了更强大的关键帧动画。
> 与过渡一样,关键帧动画也可以使用`animation-duration`、`animation-timing-function`和`animation-delay`设置动画的持续时间,调速函数和延迟时间。这些属性的用法与可选值与过渡完全一样,不再赘述。
### 关键帧
要使用关键帧动画,首先需要使用`@keyframes`定义关键帧,它由关键帧名称以及关键帧块组成。在关键帧块中可以设置动画过程中某个时间节点的属性值,它们的值可以是:
| 时间节点 | 描述 |
| --- | --- |
| 百分比值 | 设置动画具体的时间节点,`0%`为动画开始节点,`100%`为动画结束节点 |
| 关键字 | `from`相当于`0%`、`to`相当于`100%` |
```css
@keyframes change-color {
from { background-color: orange; }
to { background-color: green; }
}
@keyframes change-shape {
0% { width: 50px }
50% { width: 100px; }
100% { width: 200px; }
}
```
如果省略了`100%`的关键帧,那么它会与之前最近的关键帧相同。
### 动画名称
使用`animation-name`将关键帧绑定到某个元素上,就可以使其拥有关键帧的动画效果,该属性不能省略。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| 关键帧名称 | 要绑定到元素上的关键帧 |
```css
div {
animation-name: change-color;
}
```
### 播放次数
使用`animation-iteration-count`设置动画的播放次数,具体来说,它指的是动画**执行期**的循环次数(见下文)。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `1` |
| 可选值 | 描述 |
| --- | --- |
| 整数值 | 设置具体的播放次数 |
| `infinite` | 表示循环播放 |
### 动画方向
使用`animation-direction`设置动画的播放方向。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `normal` |
| 可选值 | 描述 |
| --- | --- |
| `normal` | 正常播放 |
| `reverse` | 逆向播放 |
| `alternate` | 动画正常播放完成后,第二遍重新逆向播放 |
| `alternate-reverse` | 动画逆向播放完成后,第二遍重新正常播放 |
注意,逆向播放指的是第二遍播放动画时再逆向播放一遍,也就是说动画播放次数**至少要两次**才可以看到效果。以`0%`为动画第一帧,`100%`为动画最后一帧为例,通过下表来表示动画播放的方向:
| 属性 | 第一遍动画 | 第二遍动画 |
| --- | --- | --- |
| `normal` | 0% → 100% | 0% → 100% |
| `reverse` | 100% → 0% | 100% → 0% |
| `alternate` | 0% → 100% | 100% → 0% |
| `alternate-reverse` | 100% → 0% | 0% → 100% |
### 结束状态
使用`animation-fill-mode`设置动画结束时的状态。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 等待期为初始状态,完成期为初始状态 |
| `backwards` | 等待期为第一帧,完成期为初始状态 |
| `forwards` | 等待期为初始状态,完成期为最后一帧 |
| `both` | 等待期为第一帧,完成期为最后一帧 |
该属性的可选值理解起来比较复杂,需要展开讨论。
首先,动画分为**初始状态、等待期、执行期、完成期**四个阶段。
**初始状态**指的是元素没有设置`animation`时处于的状态,也就是与动画效果无关的状态。只有在动画第一遍执行之前才会出现。
**等待期**是`animation-delay`设置的延迟时间,这段时间内元素的状态会受到该属性影响。
- 如果是`none`或`forwards`,那么等待期为初始状态
- 如果是`both`或`backwards`,那么等待期为第一帧
这里的第一帧根据`animationo-direction`会有所不同:
- 如果是`normal`或`alternate`,第一帧是`0%`
- 如果是`reverse`或`alternate-reverse`,第一帧是`100%`
**执行期**指的是延迟结束后动画的第一帧直到最后一帧。这里的最后一帧根据动画方向`animationo-direction`和播放次数`animation-iteration-count`也有所不同(第一帧与之相反即可):
- 如果动画方向是`normal`,那么最后一帧为`100%`,与播放次数无关
- 如果动画方向是`reverse`,那么最后一帧为`0%`,与播放次数无关
- 如果动画方向是`alternate`,那么
- 播放次数为单数,最后一帧为`100%`
- 播放次数为双数,最后一帧为`0%`
- 如果动画方向是`alternate-reverse`,与`alternate`正好相反
**完成期**指的是动画执行完最后一帧后,元素的状态。注意,`infinite`的动画只有第一遍。

```html
<div class="box">
<div class="bar"></div>
</div>
```
```css
.bar { width: 0; /* 初始状态 */ }
@keyframes progress {
0% { width: 30%; } /* 第一帧 */
100% { width: 100%; } /* 最后一帧 */
}
```
> *参考资料*
> - [如何理解animation-fill-mode及其使用?- Druidiurd 的回答](https://segmentfault.com/q/1010000003867335)
### 播放状态
使用`animation-play-state`设置动画的播放状态。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `running` |
| 可选值 | 描述 |
| --- | --- |
| `running` | 正常播放 |
| `paused` | 暂停播放 |
```css
div:hover { animation-play-state: paused; }
```
### 动画属性简写
使用`animation`简写上述除了`animation-play-state`以外所有的属性。除了`animation-name`和`animation-duration`之外,其它值均可以被省略,而且顺序也不重要。不过,如果有两个时间值,则前者表示持续时间,后者表示延迟时间;如果只有一个时间值,那么它表示持续时间。
```css
div { animation: change-color 3s linear 0s infinite alternate forwards; }
```
## 变形
使用`transform`属性可以使元素在二维或三维空间内变形。注意,元素在变形后所占的**实际位置**均与变形前一样,不会影响其它元素的位置。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素和行内替换元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 没有变形 |
该属性的其它可选值为一个函数列表,包含下面的各种变形函数。要同时使用多个函数,只要将它们用**空格**隔开即可。浏览器会按照顺序从第一个函数开始执行,直到最后一个结束。因此,**即便是相同的函数,但顺序不同,也会导致不同的结果**。
注意,在关键帧动画中使用变形时,每个关键帧都是**相对于元素的初始状态**,而非变形后的状态。
### 移动
使用下列函数可以使元素沿着某条轴进行移动。
| 函数 | 描述 |
| --- | --- |
| `translateX()` | 水平方向移动,参数为长度值或百分比值(相对于自身边框区域宽度),正数表示向右移动,负数表示向左移动 |
| `translateY()` | 垂直方向移动,参数为长度值或百分比值(相对于自身边框区域高度),正数表示向下移动,负数表示向上移动 |
| `translate()` | 同时设置水平和垂直方向移动,如果省略了一个参数,那么它表示水平方向 |
| `translateZ()` | 前后方向移动,参数**只能为长度值**,正数表示向屏幕前方移动,负数表示向屏幕后方移动 |
| `translate3d()` | 同时设置水平、垂直和前后方向移动,三个参数均不能省略 |
```css
div { transform: translateX(100px) }; /* 向右 100px */
div { transform: translateY(100px) }; /* 向下 100px */
div { transform: translate(30px, 30px) }; /* 向右、向下 30px */
div { transform: translateZ(100px) }; /* 向前 100px */
div { transform: translate3d(10px, 30px, 50px) }; /* 向右 10px、向下 30px、向前 50px */
```
由于移动的百分比值是相对于元素自身的`border`区域计算,根据这个特性,也可以使**块级元素水平垂直居中**。首先通过绝对定位将子元素的左上角移动到中心位置,但是不同于之前使用负外边距的作法,而是使用`translate()`将其向回移动自身宽高的一半,这样的好处是不需要知道子元素的宽高,也不需要进行重新计算。
```css
.father {
position: relative;
width: 600px;
height: 600px;
}
.child {
position: absolute;
left: 50%;
top: 50%;
transform: translate(-50%, -50%);
}
```
### 缩放
使用下列函数可以将元素缩放。它们的参数均为无单位数值,而且必须是正数,表示原来大小的倍数:
| 函数 | 描述 |
| --- | --- |
| `scaleX()`/`scaleY()`/`scaleZ()` | 设置水平 / 垂直 / 前后的缩放倍数 |
| `scale()` | 同时设置水平和垂直方向的缩放倍数,如果省略了一个参数,那么它同时表示水平和垂直方向 |
| `scale3d()` | 同时设置水平、垂直和前后方向缩放,三个参数均不能省略 |
```css
div { transform: scale(2); } /* 元素会被放大两倍 */
```
注意,如果设置了元素阴影,那么**阴影也会被同时放大**。如果不希望改变阴影的效果,那么直接修改宽高即可。

### 倾斜
使用下列函数可以将元素倾斜,它们的参数均为角度值:
| 函数 | 描述 |
| --- | --- |
| `skewX()`/`skewY()` | 沿 x 轴 / y 轴倾斜 |

### 旋转
使用下列函数可以将元素绕某个轴旋转,它们的参数均为角度值,正负均可。
| 函数 | 描述 |
| --- | --- |
| `rotateX()`/`rotateY()`/`rotateZ()` | 分别设置绕 x 轴 / y 轴 / z 轴旋转 |
| `rotate()` | 设置 2D 旋转,类似于`rotateZ()` |

注意,旋转会导致元素的**坐标轴同时旋转**,因此如果移动和旋转同时使用时,切记**先移动,再旋转**,否则元素会朝旋转后的方向移动。
```css
div { transform: translateX(100px) rotateZ(90deg); }
```
> 要记住`rotateX()`和`rotateY()`旋转的方向,可以将左手大拇指指向 x 轴或 y 轴的正方向(右和下),然后手指弯曲的方向就是正角度值旋转的方向。
### 透视
如果在 3D 空间内变形元素,那么必须使用透视才能使其表现出效果。浏览器通过透视值可以模拟出图像的近大远小,从而呈现出立体感。
使用`perspective`属性设置透视值,即**人眼距离屏幕的距离**。一般来说,将透视值固定为`500px`,然后调整元素的 3D 位置即可。注意,该元素必须设置给 3D 变形元素的**父元素**。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 变形元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 没有透视效果 |
| 长度值 | 设置具体的透视值 |
```css
body { perspective: 500px; }
div { transform: rotateX(45deg); }
```
此外,也可以使用函数`perspective()`来设置透视,此时该函数必须放在 3D 变形函数之前才能生效。
```css
div { transform: perspective(500px) rotateX(45deg); }
```
### 保留 3D 效果
使用`transform-style`属性保留元素 3D 变形后的效果,而不是与其它元素合并为一个平面。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 变形元素 | 不可继承 | `flat` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 不保留 3D 效果 |
| `preserve-3d` | 保留 3D 效果 |
以 x 轴的旋转为例,当使用`rotateX()`和`perspective()`创建出一个具有透视效果的 3D 旋转后,如果再将父元素进行 3D 旋转,会发现两个元素已经合并到了一个平面,所谓 3D 旋转只是一个假象:

要保留 3D 效果,则要在**父元素**中设置`transform-style: preserve-3d`,表示保留变形后的 3D 效果。

```html
<div class="outer">
<div class="inner"></div>
</div>
```
```css
.outer {
transform-style: preserve-3d; /* 保留变形后的 3D 效果 */
transform: perspective(500px) rotateY(60deg);
}
.inner {
transform: perspective(500px) rotateX(45deg);
}
```
### 变形中心点
使用`transform-origin`设置变形中心点的位置。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 变形元素 | 不可继承 | `50% 50%` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | 基于左上角的偏移量,`0 0`表示左上角 |
| 百分比值 | |
| `left`/`right`/`top`/`bottom`/`center` | 表示中心点紧贴该方向 |
上述值可以出现一至两次,一个值表示同时表示水平和垂直方向,两个值则前者表示水平方向,后者表示垂直方向。
下图以旋转为例,分别表示参照点为元素中心,以及右下角`right bottom`的情况:
<file_sep>---
title: 大前端笔记之16 ⏰ JavaScript BOM
date: 2020-02-05 14:00:41
abbrlink: gjqbkyw3
tags: JavaScript
categories: 大前端
excerpt: BOM(Browser Object Model,浏览器对象模型)也是浏览器提供的一个 API。通过 BOM 可以控制浏览器视口以外的部分,比如浏览器的窗口和框架等等。
---
# 大前端笔记之16 ⏰ JavaScript BOM
BOM(Browser Object Model,浏览器对象模型)也是浏览器提供的一个 API。通过 BOM 可以控制浏览器视口以外的部分,比如浏览器的窗口和框架等等。
下面的大部分属性和方法均属于`window`对象,它指代当前的浏览器窗口,是当前页面中的顶层对象。调用它的属性和方法均可以省略`window`,比如可以写`alert()`而不是`window.alert()`。
## 弹框
下列原生的浏览器弹框样式无法修改,并且在对话框弹出期间,浏览器窗口处于冻结状态,用户无法进行其它操作,因此不推荐使用。
### alert()
弹出普通提示框。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 对话框内容 |
```js
alert('烦人的对话框')
```
### prompt()
弹出输入对话框,用来获取用户输入。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 对话框内容 |
| 字符串 | 可选,输入框的默认值 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 / `null` | 用户输入的内容,如果用户点击取消则返回`null` |
```js
var result = prompt('请输出您的年龄', 38)
```
### confirm()
弹出判断对话框,用来获取用户输入。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 对话框内容 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 用户点击确定返回`true`,否则返回`false` |
```js
var result = confirm('您确定要离开吗?')
```
## 定时器
定时器可以用来设置某段代码在特定时间之后 / 每隔特定时间执行。
### setTimeout() / setInterval()
创建定时器。区别在于前者是延迟一段时间执行,后者是每隔一段时间执行。
| 参数 | 描述 |
| --- | --- |
| 函数 | 要定时执行的函数 |
| 整数值 | 延迟时间 / 间隔时间,单位为毫秒 |
| 若干对象 | 执行函数的参数 |
| 返回值 | 描述 |
| --- | --- |
| 整数值 | 定时器的编号 |
```js
var id1 = setTimeout(function(){
console.log('延迟一秒后输出!')
}, 1000)
var id2 = setInterval(function(){
console.log('每隔一秒输出!')
}, 1000)
setTimeout(function(msg) { // 1 秒后输出 hello
console.log(msg)
}, 1000, 'hello')
```
### clearTimeout() / clearInterval()
清除定时器。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 定时器的编号 |
```js
var id = setTimeout(function(){
console.log('延迟一秒后输出!')
}, 1000)
clearTimeout(id)
```
## Location
使用`window.location`属性可以获取`Location`对象,它包含了与 URL 相关的属性和方法。
### 相关属性
下列属性都是**可读写**的,因此修改`href`属性的值会导致页面发生跳转。
```js
// 当前 URL 为:http://www.example.com:4097/path/a.html?x=111#part1
location.href // 完整 URL:"http://www.example.com:4097/path/a.html?x=111#part1"
location.protocol // 协议:"http:"
location.host // 主机:"www.example.com:4097"
location.hostname // 主机名称:"www.example.com"
location.port // 端口号:"4097"
location.pathname // 路径:"/path/a.html"
location.search // 提交参数:"?x=111"
location.hash // 哈希值:"#part1"
```
### replace() / assign()
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要跳转的页面地址 |
使当前页面跳转到新的地址。它们的区别在于,`replace()`方法会删除历史记录中之前的地址,也就是说当页面跳转后,浏览器的后退按钮就无法使用了。而通过`assign()`和`href`跳转的页面依然可以后退。
```js
location.replace('http://www.example.com')
location.assign('http://www.example.com')
location.href = 'http://www.example.com'
```
### reload()
刷新当前页面,相当于浏览器的刷新按钮。
```js
location.reload()
```
## History
使用`window.history`属性可以获取`History`对象,它包含了与历史记录相关的属性和方法。注意,调用这些方法修改页面地址不会导致向服务器发送请求,因此它们也是[前端路由](/posts/ovrgsm0u.html#前端路由)的一种实现方式。
### back() / forward()
跳转到上一个 / 下一个浏览的页面,相当于浏览器的后退 / 前进按钮。如果不存在上一个或下一个页面,那么该方法无效。
```js
history.back()
history.forward()
```
### go()
以当前网址为基准,跳转到前 n 个 / 后 n 个页面。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 负值表示后退,正值表示前进,`0`相当于刷新页面 |
```js
history.go(-1) // 相当于 back()
history.go(1) // 相当于 forward()
history.go(0) // 相当于 刷新页面
```
### pushState()
历史记录本质是一个栈结构,当前的 URL 就是栈顶的值。使用`pushState()`可以向栈中压入一条记录,使当前的 URL 发生改变。
```js
// 使 URL 由 http://localhost:8080/ 变成 http://localhost:8080/home,前两个参数固定为空即可,基本没有作用
history.pushState({}, '', 'home')
```
### replaceState()
与`pushState()`类似,但是它是直接替换栈顶的值,因此虽然当前的 URL 也会发生改变,但是前进后退按钮是不可用的,因为栈中并没有添加新的历史记录。
```js
// 使 URL 由 http://localhost:8080/home 变成 http://localhost:8080/about
history.replaceState({}, '', 'about')
```
## Navigator
使用`window.navigator`属性可以获取`Navigator`对象,它包含了与用户环境相关的属性和方法。
```js
navigator.userAgent // 浏览器的厂商和版本
navigator.platform // 用户的操作系统信息
```
下面的代码可以简单判断用户的设备:
```js
if((navigator.userAgent.match(/(phone|pad|pod|iPhone|iPod|ios|iPad|Android|Mobile|BlackBerry|IEMobile|MQQBrowser|JUC|Fennec|wOSBrowser|BrowserNG|WebOS|Symbian|Windows Phone)/i))) {
/* 手机端 */
} else {
/* PC 端 */
}
```
## 本地存储
### Cookie
使用`document`对象的`cookie`属性获取`Cookie`对象,包含了本域的所有 Cookie。它的值为字符串,多个 Cookie 以分号隔开,必须手动拆分才能获取每一个 Cookie 的值。
```js
var cookies = document.cookie // => 'foo=bar;baz=bar'
for (var i = 0; i < cookies.length; i++) {
cookies[i]
}
// => foo=bar
// => baz=bar
```
> 由于原生操作的复杂性,可以使用第三方插件来操作 Cookie,比如 [JavaScript Cookie](https://github.com/js-cookie/js-cookie) 。
### Web Storage
Web Storage 是 HTML5 新增的本地存储 API,同样使用键值对来保存数据。相比于传统的 Cookie,它的存储空间更大,操作也更加方便。使用`window`对象的`sessionStorage`或`localStorage`属性可以获取相应的对象,它们的方法和属性都是相同的,区别在于前者存储的数据在浏览器关闭后就会清空,使用相对较少;后者存储的数据则会一直存在。
它们的主要方法如下:
| 方法 | 描述 |
| --- | --- |
| `setItem()` | 存储一条数据,它的两个参数为**字符串**(不然会类型转换)。对于一些复杂的数据格式,应该转换为 JSON 后再保存 |
| `getItem()` | 根据键获取对应数据 |
| `removeItem()` | 根据键删除对应数据 |
| `clear()` | 清除当前域下的所有数据 |
```js
sessionStorage.setItem('bgcolor', 'red')
localStorage.setItem('name', 'Saber')
localStorage.getItem('name') // => Saber
localStorage.removeItem('name')
localStorage.clear()
```
## Selection
使用`window`对象的`getSelection()`方法可以获取`Selection`对象,它包含了与用户选中文本相关的属性和方法。
### removeAllRanges()
禁止用户选择页面文本。
```js
getSelection().removeAllRanges()
```<file_sep>---
title: 大前端笔记之02 😄 CSS 基本概念
date: 2020-01-20 16:38:18
abbrlink: vayido08
tags: CSS
categories: 大前端
excerpt: CSS(Cascading Style Sheets,层叠样式表)是一种样式表语言,可以用来对 HTML 文档进行装饰。
---
# 大前端笔记之02 😄 CSS 基本概念
CSS(Cascading Style Sheets,层叠样式表)是一种样式表语言,可以用来对 HTML 文档进行装饰。
## 基础语法
CSS 的语法非常简单,每个 CSS 文档由若干条规则组成,每条规则以选择器开头,然后紧跟一个声明块`{}`。声明块中包含若干条声明,每条声明需要以分号结尾。每条声明又由两部分组成,分别为属性名和属性值,以`:`隔开。
```css
div { /* 选择器 */
font-size: 16px; /* 声明 */
color: #66ccff;
}
```
## 引入 CSS
要在 HTML 文档中加载创建好的 CSS 共有以下三种方式。
第一种是直接在元素上使用`style`属性,将声明作为属性值写在里面,那么这些声明就会应用在该元素上,这样的方式称为**内联样式**。但是这样的方式不方便 CSS 规则的重用,修改起来也非常麻烦,因此并不常用。
```html
<div style="color: #66ccff;"></div>
```
第二种方式是在`<head>`中使用`<style>`元素,然后将规则写在里面。但是如果 CSS 规则比较多,那么就会使得 HTML 页面变得很长,可读性变差。
```html
<head>
<style>
div { color: #66ccff; }
</style>
</head>
```
第三种是将规则定义在一个单独的 CSS 文件中,然后在`<head>`中使用`<link>`元素将文件引入。这样的方式符合结构与样式分离的原则,最为常用。
| 属性 | 描述 |
| --- | --- |
| `rel` | 通常为固定值`stylesheet`,表示引入的外部文件是样式表,不可省略 |
| `href` | 样式表的 URL |
```html
<head>
<link rel="stylesheet" href="test.css">
</head>
```
```css
/* CSS 文件 test.css */
div { color: #66ccff; }
```
## 元素分类
CSS 将元素分成了不同的种类,有些属性只能适用于特定种类的元素。注意,HTML 也规定了块级元素和行内元素,例如行内元素`<span>`只能包含其它行内元素,但是不能包裹`<p>`这样的块级元素。但是 CSS 定义的块级元素和行内元素只是让元素拥有特定的样式表现,并且可以通过`display`属性修改。然而这并不能修改其本身的定义,也就是说,在任何情况下让`<span>`包裹`<p>`都是不合法的。
CSS 中块级元素与行内元素的主要区别在于:
| | 块级元素 | 行内元素 |
| --- | --- | --- |
| 宽高 | 通过`width`和`height`控制 | 只能通过`line-height`控制高度 |
| 边距和边框 | 四个方向均有效果 | 只有**水平**方向生效,垂直方向虽然可以添加,但是**不会影响布局** |
| 排列方式 | 独占一行 | 相互紧靠排列 |
| 默认宽度 | 与父元素的宽度相等 | 根据内容收缩 |
| 嵌套 | 可以包裹块级元素(`<p>`除外)或行内元素 | 只能包裹行内元素(`<a>`除外)|
除此之外还有一种比较特殊的行内块元素,它既可以像行内元素与其它元素排在一行,也可以设置宽高。
元素还分为替换元素和非替换元素两种。替换元素是指其内容可以被替换的元素,这些内容并不会在文档中直接表现出来,如`<img>`元素以及各类表单控件元素。替换元素的一个明显特征就是**自带宽度和高度**,除此之外,它与**行内块元素**的表现效果基本是一样的。
## 特性
CSS 本身有一些语法上的特性,了解这些特性可以更好地理解某些情况下最终渲染出的效果。
### 继承性
CSS 属性可以继承,也就是说,一些样式不仅会应用于选择的元素上,还会影响它的后代元素。注意,有些属性(如盒模型属性)没有继承性。
在下面的代码中,`<h1>`和`<em>`中的文本都会变成灰色,因为`<em>`从`<h1>`继承了`color`属性。
```html
<h1>Meerkat<em>Central</em></h1>
```
```css
h1 { color: gray; }
```
### 优先级
如果有多个样式适用于同一个元素,那么浏览器会计算每个样式的优先级,优先级最高的样式最终会被应用到元素上,其它的样式则会被忽略。
```css
/* 标题的实际颜色为 橙色 */
body h1 { color: orange; }
h1 { color: red; }
```
优先级由[选择器](/posts/1lc5rbtd.html)决定,可以使用一个五位数`00000`(不能进位)来表示优先级。具体规则如下:
- 行内样式,加`1000`分
- 选择器每包含一个`#id`,加`100`分
- 选择器每包含一个`.class`、`[attr]`或者`:pseudo-class`,加`10`分
- 选择器每包含一个`元素名`或者`::pseudo-element`,加`1`分
- 通配选择器`*`加`0`分
- 选择器组合没有特定性
- 继承的属性没有特定性
- 带有`!important`的规则,加`10000`分(继承的`!important`也没有特定性)
注意,优先级为`0`依然比没有优先级要高。在下面的代码中,`<strong>`会从`<p>`继承红色,并且通配符又设置该元素为灰色。那么由于通配符的优先级为`0`,而继承的红色没有优先级,因此`<strong>`最终会显示为灰色。
```html
<p>看我<strong>变色</strong>啦~</p>
```
```css
* { color: gray; }
p { color: red; }
```
如果两个规则的优先级相同,那么后引入的样式表中的规则更重要,如果两个规则在同一张样式表中,那么靠后的规则更重要。
## 浏览器渲染引擎
有些属性前可能会带有`-webkit-`这样的前缀,这是因为一些属性处于实验性状态或者是该浏览器专有,使用前缀标明之后,就可以防止与其它浏览器的同属性发生冲突。
下面是目前常见浏览器的引擎和前缀:
| 浏览器 | 引擎 | CSS 前缀 |
| --- | --- | --- |
| IE | Trident | `-ms-` |
| Edge | ~~EdgeHTML~~ → Blink | `-ms-`和`-webkit-`(Blink) |
| Firefox | Gecko | `-moz-` |
| Opera | ~~Presto~~ → Blink | `-o-`(Presto)和`-webkit-`(Blink)|
| Safari | Webkit | `-webkit-` |
| Chrome | ~~Webkit~~ → Blink | `-webkit-` |
> 注意,这只是浏览器样式的渲染引擎,对于解析 JavaScript 来说,使用的通常是 V8 引擎。<file_sep>---
title: 大前端笔记之05 📦 CSS 盒模型
date: 2020-01-22 1:08:01
abbrlink: 08603cll
tags: CSS
categories: 大前端
excerpt: 每个元素都会生成一个矩形的元素盒子,从外到内依次为外边距、边框、内边距和内容区。
---
# 大前端笔记之05 📦 CSS 盒模型
每个元素都会生成一个矩形的元素盒子,从外到内依次为外边距、边框、内边距和内容区。
## 外边距
使用`margin`设置元素的外边距,即元素之间的距离。该属性是`margin-top`、`margin-right`、`margin-bottom`和`margin-left`四个属性的简写属性。可以单独使用它们设置某个方向的外边距,或者使用`margin`同时设置不同的方向。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 见[包含块](/posts/vv37590w.html)一节 |
| 长度值 | 设置具体的外边距 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「上,右,下,左」 |
| 3 | 分别对应「上,左右,下」 |
| 2 | 分别对应「上下,左右」 |
| 1 | 同时设置四个方向 |
### 外边距合并
如果同一个[BFC](/posts/9obo9xpo.html)中的上下外边距紧挨在一起的话,那么会产生合并的现象,相当于它们共同拥有一个外边距。
合并的规则如下:如果都是正值,则保留较大的一个;如果是一正一负,则保留它们的和;如果都是负值,则保留绝对值较大的一个。
```css
div:first-child { /* 上面的 <div> */
margin-bottom: 50px;
}
div:last-child { /* 下面的 <div> */
margin-top: 30px;
}
/* 它们实际的距离为 50px */
```
另外,父元素和子元素在**上方**的外边距也会发生合并,合并规则同上,但是合并后的外边距会交给**父元素**来表现。
在下面的代码中,一个`<div>`元素中包含一个子元素`<p>`,此时`<div>`不会出现在浏览器最顶端,而是被子元素的`margin-top`连带着一起向下移动了`100px`,父子元素之间则没有任何距离。
```css
div { height: 400px; }
p { height: 200px; margin-top: 100px; }
```
使用下面的方法可以避免这种情况:
- 父元素触发[BFC](/posts/9obo9xpo.html)
- 父元素对应方向的`border`或`padding`不为`0`
- 父子元素之间有行内元素分隔
## 内边距
使用`padding`设置元素的内边距,即元素内容与边框之间的距离。该属性是`padding-top`、`padding-right`、`padding-bottom`和`padding-left`四个属性的简写属性。可以单独使用它们设置某个方向的内边距,或者使用`padding`同时设置不同的方向。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | 设置具体的内边距 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「上,右,下,左」 |
| 3 | 分别对应「上,左右,下」 |
| 2 | 分别对应「上下,左右」 |
| 1 | 同时设置四个方向 |
## 边框
使用`border`设置元素的边框。该属性是`border-width`、`border-style`和`border-color`三个属性的简写属性,顺序没有要求。
### 边框宽度
使用`border-width`设置边框的宽度,该属性是`border-top-width`、`border-right-width`、`border-bottom-width`和`border-left-width`四个属性的简写属性。可以单独使用它们设置某个方向的边框宽度,或者使用`border-width`同时设置不同的方向。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `medium` |
| 可选值 | 描述 |
| --- | --- |
| `medium` | 浏览器自定义 |
| 长度值 | 设置具体的边框宽度 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「上,右,下,左」 |
| 3 | 分别对应「上,左右,下」 |
| 2 | 分别对应「上下,左右」 |
| 1 | 同时设置四个方向 |
### 边框样式
通过`border-style`可以设置边框的样式,该属性是`border-top-style`、`border-right-style`、`border-bottom-style`和`border-left-style`四个属性的简写属性。可以单独使用它们设置某个方向的边框样式,或者使用`border-style`同时设置不同的方向。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 无边框样式,由于该值是默认值,因此**如果没有这个属性,边框就不会存在** |
| `solid` | 实线 |
| `dotted` | 点线 |
| `dashed` | 破折线 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「上,右,下,左」 |
| 3 | 分别对应「上,左右,下」 |
| 2 | 分别对应「上下,左右」 |
| 1 | 同时设置四个方向 |
### 边框颜色
使用`border-color`设置边框的颜色,该属性是`border-top-color`、`border-right-color`、`border-bottom-color`和`border-left-color`四个属性的简写属性。可以单独使用它们设置某个方向的边框颜色,或者使用`border-color`同时设置不同的方向。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | 当前元素的`color`属性值 |
| 可选值 | 描述 |
| --- | --- |
| 颜色值 | 任何合法的颜色值 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「上,右,下,左」 |
| 3 | 分别对应「上,左右,下」 |
| 2 | 分别对应「上下,左右」 |
| 1 | 同时设置四个方向 |
### 边框方向
除上述方式之外,还可以使用`border-top`、`border-bottom`、`border-left`和`border-right`来简写对应方向的**宽度、样式和颜色**值。这些属性要求值的顺序严格,但是可以省略其中的任意值。
```css
div { border-top: 1px solid #ccc; }
```
### 边框圆角
使用`border-radius`设置边框圆角。注意,该属性不会改变元素盒子实际的形状。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| 长度值 | 设置具体的圆角大小 |
| 百分比值 | 相对于对应方向的`border`长度 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「左上,右上,右下,左下」 |
| 3 | 分别对应「左上,右上左下,右下」 |
| 2 | 分别对应「左上右下,右上左下」 |
| 1 | 同时设置四个方向 |
圆角的大小本质是一个圆或者椭圆的半径,使用它的四分之一来定义边框的弧度。比如:
```css
div { border-radius: 10px; }
```

### 边框图片
边框图片用于设置更复杂的边框。注意,必须将`border-style`设置成一个不为`none`的值,否则边框图片无法显示。
#### 图片路径
使用`border-image-source`设置图片的路径,不能省略。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `none` |
| 可选值 | 描述 |
| --- | --- |
| `none` | 没有边框图片 |
| `url()` | 图片的 URL |
#### 裁切位置
使用`border-image-slice`设置图片的裁切位置。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `100%` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | **不能加单位**,默认为像素 |
| 百分比值 | 相对于图片的宽高 |
该属性根据数量不同分别表示不同的方向:
| 值的数量 | 描述 |
| --- | --- |
| 4 | 分别对应「上,右,下,左」 |
| 3 | 分别对应「上,左右,下」 |
| 2 | 分别对应「上下,左右」 |
| 1 | 同时设置四个方向 |
该属性的作用原理是,浏览器会在边框图片上放置 4 条切割线,每条线到图片边缘的距离就是`border-image-slice`的值。
例如,背景图片为`500 × 500`大小,那么将 4 个值设置为`167`或`33.3%`正好可以将其等分为 9 份,从而形成一个九宫格。
```css
div {
border-image-slice: 167; /* 左图 */
border-image-slice: 10% 20% 30% 50%; /* 右图 */
}
```

然后,九宫格的四个角会分别作为边框的四角,而上下左右四个部分会作为边框的四边,按照一定的方式平铺:

因此,如果将 4 个值全部设置为`50%`,那么图片只能被等分为 4 份,只有 4 个角才有图案,分别为图片的四分之一;如果将 4 个值全部设置为`100%`,那么图片只有 1 份,该图片会被完整的显示在 4 个角上,四边依然没有图案。

#### 边的平铺方式
使用`border-image-repeat`设置四边的平铺方式。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `stretch` |
| 可选值 | 描述 |
| --- | --- |
| `stretch` | 拉伸 |
| `repeat` | 将边框图片放在边框的中央,然后向两侧重复,边缘的部分可能会被截断 |
| `round` | 改变边框图片的尺寸,使其正好放置在边框之内 |

#### 边框图片宽度
使用`border-image-width`设置边框图片的宽度。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `1` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | 表示`border-width`的倍数 |
| 长度值 | 设置具体的边框图片宽度 |
该属性的原理与`border-image-slice`相似,区别在于`border-image-width`是在元素(边框区域)上绘制 4 条切割线,然后将切割出来的部分作为边框图片的绘制区域。

上面的两个元素均设置了`30px`的`border-width`、`200px`的宽高,区别在于左图单独设置了`20px`的`border-image-width`。可见默认情况下,边框图片绘制在**元素的内部**,不会导致元素扩大。而且,虽然图片边框没有实际边框大,但是**只要是图片边框存在,那么实际边框就不会显示,就好像是透明了一样(因为大小还在)**,因此被元素的背景色填充。
> 按照《CSS 权威指南》中的说法,`border-width`不会影响边框图片的显示,只会影响元素的大小。但是在 Chrome 下,如果没有设置`border-width`,那么 Chrome 会默认为其添加`3px`的大小。如果直接将其设置为`0`,那么边框图片会**无法显示**。
#### 绘制位置
默认情况下,边框图片绘制在元素的内部,不会导致元素扩大。使用`border-image-outset`属性可以设置边框图片向元素外推移。注意,该属性可能会导致边框图片与其它元素重叠,或者被浏览器边界截断。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 所有元素 | 不可继承 | `0` |
| 可选值 | 描述 |
| --- | --- |
| 数字值 | 表示`border-width`的倍数 |
| 长度值 | 设置具体向外推移的距离 |

#### 简写属性
使用`border-image`可以简写上述所有属性,其中图片路径`border-image-source`和平铺方式`border-image-repeat`可以写在任意位置,但是`border-image-slice`、`border-image-width`和`border-image-outset`必须按照顺序写在一起,并使用`/`分隔。其中除了图片路径外,其它的值都可以省略。
```css
div { border-image: url("a.jpg") 33.3% / 20px / 30px round; }
```
## 宽高
使用`width`可以设置元素**内容区(默认)**的宽度。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素和行内替换元素 | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 见[包含块](/posts/vv37590w.html)一节 |
| 长度值 | 设置具体的宽度 |
| 百分比值 | 相对于[包含块](/posts/vv37590w.html)的`width` |
使用`height`设置元素**内容区(默认)**的高度。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素和行内替换元素 | 不可继承 | `auto` |
| 可选值 | 描述 |
| --- | --- |
| `auto` | 表示高度正好容纳它的子元素 |
| 长度值 | 设置具体的高度 |
| 百分比值 | 相对于[包含块](/posts/vv37590w.html)的`height`,如果包含块没有一个**具体**的`height`,则相当于设置为`auto` |
### 修改作用区域
使用`box-sizing`改变宽高的作用区域。默认情况下,`width`和`height`作用于内容区,添加额外的内边距或边框会导致元素实际会大于设置的`width`或`height`。
| 适用于 | 继承性 | 默认值 |
| --- | --- | --- |
| 块级元素和行内替换元素 | 不可继承 | `content-box` |
| 可选值 | 描述 |
| --- | --- |
| `content-box` | 表示宽高作用于内容区 |
| `border-box` | 表示宽高作用于边框区域 |
### 最大 / 最小宽高
使用`min-width`、`max-width`、`min-height`和`max-height`四个属性可以限制元素的宽度和高度,它们的用法与`width`和`height`基本一致。
## 轮廓
使用`outline`设置元素的轮廓。它显示在边框和外边距之间,但是**不占据任何空间**。而且,轮廓**没有简写属性**,因此只能将四个方向的轮廓设置为同一个样式或者不设置。
将文本框设置为`outline: none`可以清除其默认的蓝色轮廓效果。
## 行内盒模型
对于行内元素来说,它们在普通盒模型中的内容区比块级元素更加复杂。
首先,设计师在 EM Square 中设计出每个字符,不同字体的字符在 EM Square 中的位置可能会有所不同(例如微软雅黑会偏下),有些装饰笔画甚至会超出 EM Square 。多个字符的 EM Square 排列在一起会构成行内盒模型的**内容区**(与普通盒模型中内容区没关系),使用`font-size`可以控制它的高度。
然后,每个内容区外会包裹一个**行内盒子**,使用`line-height`可以控制它的高度,默认情况下内容区和行内盒子的高度应该是一样的。而且,只有`line-height`才能控制行内盒子的实际高度,如果行内盒子高度小于内容区,那么虽然内容可以正常显示,但是上下两行会重叠在一起。
但是,由于有些字体的个别部分可能会超出内容区,因此浏览器为了使行内盒子能容纳下每个字符而不至于发生重叠,通常默认将行内盒子的高度也就是`line-height`的值设置的比内容区稍大一点,通常为`1.2`,即内容区高度的`1.2`倍。因此,行内盒子和内容区之间会产生一定的距离,称为**行间距**。然后浏览器会将行间距等分成两份,分别在内容区的上下各添加一份。由于行间距是等分的,因此**内容区总是在行内盒子的中间位置**,通过这个特性,可以实现单行文本的垂直居中。

最后,一行中的所有行内盒子又会被一个更大的盒子包裹,称为**行框**。行框的高度由其内部所有的行内盒子决定,浏览器会计算每个行内盒子的高度,并且采用它们的**最高点和最低点**作为行框的上下边缘。使用`vertical-align`可以控制行内盒子在行框中的位置。
### 单行文本垂直居中
假如一个**块级元素**中只包含单行的文本,那么可以将文本的行高设置成与块级元素的高度一致。由于行间距的等分特性,那么文字正好可以在块级元素内垂直居中。当然,如果文本超过了一行,这样的方式就行不通了。
```html
<div>我垂直居中啦~</div>
```
```css
div {
height: 300px;
line-height: 300px;
}
```
### 去除图片下方缝隙
行内元素默认的基线对齐会导致图片下方出现缝隙,这是因为浏览器会在**每个行框的开头**添加一个**没有宽度的、永远透明的空白字符**,规范中称之为**支柱**(strut)。由于支柱的基线下方有一定距离,而图片的基线是下边缘,因此为了基线对齐,图片需要向上移动一小段距离,导致出现了缝隙。

解决这个问题的方法有很多,比较常见的是将图片的`vertical-align`改为`middle`或`top`,或者将图片设置为块级元素。<file_sep>---
title: 📜 正则表达式
date: 2020-02-02 14:17:22
abbrlink: 9d8oxogs
categories: 杂七杂八
excerpt: 正则表达式是由特定字符组成的规则字符串,通过各个语言中给出的相关方法,可以判断字符串是否符合正则表达式的规则,也可以将字符串中符合规则的部分提取出来,或者修改为其它值。通常用于验证表单数据、和谐某些关键字等。
---
# 📜 正则表达式
正则表达式是由特定字符组成的**规则字符串**,通过各个语言中给出的相关方法,可以判断字符串是否符合正则表达式的规则,也可以将字符串中符合规则的部分提取出来,或者修改为其它值。通常用于验证表单数据、和谐某些关键字等。
由于在不同的语言中,规则的语法都是相同的,因此这里仅给出规则,并以 JavaScript 为例:
## 特殊字符
### 字面量字符
字面量字符指的是表示字面本身含义的字符。注意,只要是内容中包含该字面量,即视为匹配,与位置无关。
```js
/dog/.test('good dog') // => true
```
### 边界符
边界符`^`表示内容的开始位置,`$`表示内容的结束位置。
```js
/^cat/.test('cat is cute') // => true,表示 cat 必须出现在内容开始的位置
/^cat/.test('The cat is mine') // => false
/dog$/.test('dog is cute') // => false,表示 dog 必须出现在内容结束的位置
/dog$/.test('This is my dog') // => true
```
如果规则同时使用了两个边界符,也就是既满足起点,又满足终点,那么表示该字符串必须完全符合规则,而不是仅仅包含就可以。
```js
/^cat$/.test('cat') // 只有这种情况才是 true
```
### 字符类
字符类表示有一系列字符可供选择,只要匹配其中一个就可以了,注意**一个字符类仅表示一个字符**。所有可供选择的字符都放在方括号内,比如`[xyz]` 表示只要字符串中有`x`、`y`、`z`中任何一个即可匹配。
```js
/[abc]/.test('helloworld') // => false,不包含 a、b、c 三个字母中任意一个
/[abc]/.test('apple') // => true,因为包含 a
```
### 连字符
对于连续序列的字符,可以使用连字符`-`提供简写形式,表示字符的范围。比如,`[abc]`可以写作`[a-c]`,`[0123456789]`可以写作`[0-9]`,所有大写字母可以写作`[A-Z]`。
```js
/[a-z]/.test('b') // => true
/[a-z0-9A-Z]/.test('1') // => true
```
注意,连字符只有在字符类中,且两侧均有字符时,才表示连字符。其他情况下,它依然表示`-`本身,不需要转义。
对于一些常用的字符类,还提供了简写模式:
| 字符类简写 | 描述 |
| --- | --- |
| `\d` | 表示`0 ~ 9`之间的任一数字,相当于`[0-9]` |
| `\D` | 表示所有`0 ~ 9`以外的字符,相当于`[^0-9]` |
| `\w` | 表示任意的字母、数字和下划线,相当于`[A-Za-z0-9_]` |
| `\W` | 表示除所有字母、数字和下划线以外的字符,相当于`[^A-Za-z0-9_]` |
| `\s` | 匹配空格(包括换行符、制表符、空格符等),相当于`[ \t\r\n\v\f]` |
| `\S` | 匹配所有非空格的字符,相当于`[^ \t\r\n\v\f]` |
### 脱字符
如果方括号内的第一个字符是`^`,则表示除了字符类之中的字符,其他字符都可以匹配。比如,`[^xyz]`表示除了`x`、`y`、`z`之外都可以匹配。
```js
/[^abc]/.test('hello world') // => true
/[^abc]/.test('bbc') // => false
```
注意,脱字符只有在**整个正则表达式的开头**或者**字符类的开头**才有特殊含义,否则就是字面含义。
### 量词符
量词符用来限定某个字符出现的次数。
| 量词符 | 描述 |
| --- | --- |
| `?` | 表示 0 次或 1 次 |
| `*` | 表示 0 次或多次,即任意次 |
| `+` | 表示 1 次或多次 |
```js
/sto?p/.test('stop') // => true
/sto?p/.test('stp') // => true
/sto*p/.test('stoooop') // => true
/sto+p/.test('stoooop') // => true
/sto+p/.test('stp') // => false
```
除了上述固定数量的量词符外,使用`{}`还可以限定任意数量:
| 量词符 | 描述 |
| --- | --- |
| `{n}` | 表示正好`n`次 |
| `{n,}` | 表示至少`n`次 |
| `{n,m}` | 表示至少`n`次,至多`m`次(均包括) |
```js
/lo{2}k/.test('look') // => true
/lo{2,5}k/.test('looook') // => true
```
### 分组
使用圆括号`()`可以将字符分组,将这些字符视为一个整体。
```js
/stop{2}/.test('stopp') // => true,表示字符 p 必须出现 2 次
/(stop){2}/.test('stopp') // => false,表示单词 stop 必须出现 2 次
```
### 选择符
选择符`|`表示或者关系,它会将前后的正则自动视为一个整体,不需要添加圆括号。
```js
/^cat$|^dog$/.test('cat') // => true,只要正好是 cat 或 dog 都可以
/^cat|dog$/.test('cattt') // => true,只要是 cat 开头或 dog 结尾都可以
```
### 点字符
点字符`.`匹配除了回车`\r`和换行`\n`以外的所有字符。
```js
/c.t/.test('cat') // => true
/c.t/.test('cut') // => true
```
### 转义字符
对于有特殊含义的元字符,如果要匹配它们本身,需要在前面添加转义字符`\`。
```js
/1\+1/.test('1+1') // => true
```
## 常用的正则表达式
```js
/^1[3-9]\d{9}$/ // 手机号码
/^[1-9]\d{4,}$/ // QQ号码
/^\w+([-+.]\w+)*@\w+([-.]\w+)*\.\w+([-.]\w+)*$/ // 邮箱
/^[\u4e00-\u9fa5]{2}$/ // 2 个汉字,根据情况修改数量即可
/^\w{6,16}$/ // 6 ~ 16 位密码,只能包含数字、字母、下划线
```<file_sep>---
title: 计算机网络笔记之06 🚛 传输层
date: 2020-01-19 10:16:58
abbrlink: 3z5qc3ni
tags: 计算机网络
categories: 计算机网络
excerpt: 施工中
---
# 计算机网络笔记之06 🚛 传输层
> 施工中... 不要忘记修改时间
## 端口
服务器上通常有很多程序同时运行,因此找到服务器之后,还需要找到接收请求的程序。计算机的每个程序会对应一个端口,而服务器用来接收 HTTP 请求的端口通常为`80`(HTTPS 为`443`)。接下来,为了保证连接的稳定,浏览器也会在客户端随机选择一个端口,与服务器的`80`端口建立一条通道。这一过程称为三次握手,是由 TCP 协议完成的。它的作用就是保证接下来传输的数据能准确无误的送达,而不关心数据的内容。<file_sep>---
title: 👩💻 字符集和编码
date: 2020-01-13 06:02:32
abbrlink: 1x90rjzf
categories: 杂七杂八
excerpt: 计算机本身无法识别字符,因此通过字符集将每个字符表示成一个十进制数字,相当于为每个字符起一个编号(学名为码点,Code Point)。
---
# 👩💻 字符集和编码
计算机本身无法识别字符,因此通过字符集将每个字符表示成一个十进制数字,相当于为每个字符起一个编号(学名为码点,Code Point)。
最早的字符集为美国采用的 ASCII,包含了英文字母、数字和简单符号在内的 127 个字符。但随着计算机在全世界逐渐普及,其它国家的文字就无法使用 ASCII 字符集表示了。因此各个国家纷纷建立了自己的字符集,如中国的 GB2312(后来进化为字库更加庞大的 GBK)、台湾的 BIG5 等。
> 微软在 Windows 中将这些各国家自己的字符集统称为 ANSI 字符集,然后根据用户地区来识别究竟应该使用具体哪一种本地的字符集。
但是,如此多的字符集带来了新的问题,因为同一个字符在某个字符集可能是某个编号,在另外一个字符集又成为了其它编号,这使得本应正常显示的内容,在另一套字符集中成为了乱码,从而阻碍了不同地区之间信息的交互。
之后,为了解决字符集的不统一性,Unicode 字符集出现了。它包括了世界上所有的文字和符号,这样大家都使用这一套字符集,就不会出现乱码问题了。
然而,要在一个字符集中保存这么多字符,那么每个字符会占据更多的空间。比如英文字母本身在 ASCII 字符集下只需要 1 个字节就可以表示,现在则需要 3 个字节。如果只把英文字符作为 1 个字节存储,当这些数据保存到计算机后,它并不知道一次读取多少个 bit 才算是一个字符。例如`abc`三个字母在一起,写作十六进制是`009700980099`,如果一次读取两个字节则是`0097_0098_0099`,但是一次读取 3 个字节就变成了`009700_980099`,就只有两个字符了,而且表示的字符也是不正确的。
因此为了解决这一问题,又诞生了 UTF-8 编码方式,用来告诉计算机究竟多少字节表示一个字符。具体来说,它使用 1 ~ 4 个字节表示一个字符,根据不同符号而变化长度:
- 对于单字节的符号,字节的第一位设为`0`,后面 7 位为这个符号的 Unicode 码,因此对于英语字母,UTF-8 编码和 ASCII 码是相同的。也就是说,只要计算机读到一个字节的第一位是`0`,那么就将其当作一个字符来读取。
- 对于`n`字节的符号(`n > 1`),第一个字节的前`n`位都设为`1`,第`n + 1`位设为`0`,后面字节的前两位一律设为`10`。剩下的没有提及的二进制位,全部为这个符号的 Unicode 码。也就是说,某个字节开头有多少个`1`,就表示包括该字节在内之后的多少个字节为一个字符。
如下表中展示了 Unicode 字符集与对应的 UTF-8 编码方式:
| Unicode | UTF-8 |
| --- | --- |
| `0000 0000`~`0000 007F` | `0xxxxxxx` |
| `0000 0080`~`0000 07FF` | `110xxxxx 10xxxxxx` |
| `0000 0800`~`0000 FFFF` | `1110xxxx 10xxxxxx 10xxxxxx` |
| `0001 0000`~`0010 FFFF` | `11110xxx 10xxxxxx 10xxxxxx 10xxxxxx` |
以汉字「严」为例,它的 Unicode 字符集码点为`4E25`,根据上表,发现`4E25`位于第三行范围内。因此将`4E25`转换为二进制的,**从低位开始**依次填充到相应的 UTF-8 格式内,不足的高位处补`0`,从而得到「严」的 UTF-8 编码。由于汉字均位于第三行范围之内,所以汉字的 UTF-8 编码会占 3 个字节。

此时,当计算机读取到`11100100`这第一个字节后,就知道包括该字节在内的连续三个字节为一个字符,根据格式去掉冗余位之后,就能解析出该字符的二进制为`0100 111000 100101`,从而在 Unicode 字符集找到该码点对应的字符「严」了。<file_sep>---
title: 大前端笔记之22 📦 webpack
date: 2020-02-20 16:21:01
abbrlink: 1zcig0yl
tags: webpack
categories: 大前端
excerpt: webpack 是一个前端项目的自动化构建工具,更确切地来说,它是一个模块打包器。模块化可以使得越来越复杂的前端项目中的文件依赖变得清晰,更容易管理。但是由于浏览器的兼容性问题,ES6、CommonJS 等模块化语法并不能得到很好的支持。而 webpack 提供了各种模块化方式的支持,可以将这些包含模块化语法的代码进行打包合并,转换成浏览器能够运行的代码。可见,虽然它也具有类似于 Gulp 提供诸如代码转换(ES6 → ES5、Less → CSS )、代码压缩等功能,但是提供模块化支持,才是它更强大的地方。
---
# 大前端笔记之22 📦 webpack
[webpack](https://www.webpackjs.com/concepts)是一个前端项目的自动化构建工具,更确切地来说,它是一个模块打包器。模块化可以使得越来越复杂的前端项目中的文件依赖变得清晰,更容易管理。但是由于浏览器的兼容性问题,ES6、CommonJS 等模块化语法并不能得到很好的支持。而 webpack 提供了各种模块化方式的支持,可以将这些包含模块化语法的代码进行打包合并,转换成浏览器能够运行的代码。可见,虽然它也具有类似于 Gulp 提供诸如代码转换(ES6 → ES5、Less → CSS )、代码压缩等功能,但是提供模块化支持,才是它更强大的地方。
## 基本概念
webpack 是一个基于 Node.js 的工具,需要使用 npm 安装。由于它仅仅是一个开发工具,因此将其作为开发依赖即可:
```powershell
npm i webpack webpack-cli --save-dev
```
接下来在项目目录中新建一个`src`目录,用来存放源代码文件,等待 webpack 处理。
```powershell
test
├─ node_modules # 模块文件夹
├─ package-lock.json
├─ package.json # 使用 npm init 生成的
├─ index.html # 用于引入打包后的 js 文件,临时放到这里作为测试
└─ src # 存放源代码文件
└─ index.js # 随便写了一行 console.log() 代码
```
接下来使用[npx](/posts/o48l9v2o.html#npx)执行 webpack 进行项目打包。
```powershell
npx webpack
```
此时会发现项目下多出了一个`dist`文件夹,它用来存放处理后的代码,当前包含了打包后的`main.js`文件。打开该文件发现,虽然源文件中只有一行`console.log()`,但是打包后的文件是经过压缩后的一串复杂代码,这是因为 webpack 提供了模块化支持。
然后在`index.html`引入刚生成的`main.js`,可以发现源文件`index.js`中的代码被正确执行了。
如果要修改 webpack 的配置,需要在项目根目录下新建一个`webpack.config.js`文件,该文件是一个模块,且**只能使用 CommonJS 语法**,因此配置对象需要通过模块化导出。
```js
/* webpack.config.js */
module.exports = {
// 配置参数
}
```
## loader
webpack 除了可以管理 JavaScript 文件以外,也可以管理诸如 CSS、图片等各种静态资源,也就是说,可以使用模块化语法直接导入这些其它类型的资源文件,而不仅仅是 JavaScript 文件。不过,要管理这些资源,需要首先安装对应的 loader,不同的资源有不同的 loader,对应的配置方式也不太相同。
### 样式文件
管理 CSS 样式文件需要使用[css-loader](https://www.webpackjs.com/loaders/css-loader/)和[style-loader](https://www.webpackjs.com/loaders/style-loader/)两个 loader,前者负责导入 CSS 文件并提供返回的 CSS 代码,后者负责将 CSS 代码嵌入到页面的 DOM 使其生效。因此你只需在页面中引用一个输出的 JavaScript 文件,就可以使页面拥有样式了!
```powershell
npm install style-loader css-loader --save-dev
```
```js
module.exports = {
module: {
rules: [{
test: /\.css$/,
use: ['style-loader', 'css-loader']
}]
}
}
```
> 注意配置文件中加载 loader 的顺序,是**从右向左**执行的,因此`css-loader`必须要写在后面(先执行)才能正确运行。
如果还需要管理 Less 文件,则需要使用[less-loader](https://www.webpackjs.com/loaders/less-loader/)和 Less 本身。并且由于 Less 最终依然会转换为 CSS,因此也需要依赖`css-loader`和`style-loader`。
```powershell
npm install --save-dev less-loader less
```
```js
module.exports = {
module: {
rules: [{
test: /\.less$/,
use: ['style-loader', 'css-loader', 'less-loader']
}]
}
}
```
> 注意,虽然这里的规则中也使用了`css-loader`和`less-loader`,但是如果希望正确处理 CSS 文件的话,依然需要之前 CSS 的规则,这里的规则仅仅用于管理 Less 文件,不会管理 CSS 文件。
### 图片文件
当其它文件中引入图片文件时(比如 CSS 文件中通过`url()`设置了背景图片),那么需要使用[file-loader](https://www.webpackjs.com/loaders/file-loader/)来管理图片文件。
```powershell
npm install --save-dev file-loader
```
```js
module.exports = {
module: {
rules: [{
test: /\.(png|jpg|gif)$/,
use: ['file-loader']
}]
}
}
```
可以看到`dist`目录中生成了一个随机文件名的图片文件,不过由于此时`index.html`文件的位置问题导致图片路径不正确,因此图片无法正常显示,之后处理 HTML 文件时就会解决这个问题。
除此之外,还可以使用[url-loader](https://www.webpackjs.com/loaders/url-loader/)将比较小的图片转换成 Base64。当图片的体积小于`limit`(单位为字节)时,webpack 会将图片转换为 Base64。如果大于`limit`,则会自动调用 file-loader。
> 注意!一旦配置了 url-loader,虽然 file-loader 必须安装,但是**千万不要配置**!否则会由于冲突导致打包结果不正确!
```powershell
npm install --save-dev url-loader
```
```js
module.exports = {
module: {
rules: [{
test: /\.(png|jpg|gif)$/,
use: [
{
loader: 'url-loader',
options: {
limit: 8192 // 相当于 8KB
}
}
]
}]
}
}
```
### Babel
使用[babel-loader](https://www.webpackjs.com/loaders/babel-loader/)可以将 ES6 语法转换成兼容性比较好的 ES5 语法。
```powershell
npm install -D babel-loader @babel/core @babel/preset-env
```
```js
module.exports = {
module: {
rules: [{
test: /\.m?js$/,
exclude: /(node_modules|bower_components)/,
use: {
loader: 'babel-loader',
options: {
presets: ['@babel/preset-env']
}
}
}]
}
}
```
## 管理输出
### 入口
使用`entry`设置项目的起点,默认为`./src/index.js`。
```js
module.exports = {
entry: './src/main.js' // 修改为 main.js 文件
}
```
### 出口
使用`output`设置编译后的文件名称和保存位置,默认为`./dist/main.js`。该属性是一个对象,它包含`path`和`filename`两个属性,前者用来设置保存位置,后者设置文件名称。注意,其中`path`必须为**绝对路径**,因此需要引入`path`模块拼接路径。
```js
const path = require('path')
module.exports = {
output: { // 将打包后的文件修改为 项目目录下的 output/bundle.js
path: path.join(__dirname, 'output'),
filename: 'bundle.js'
}
}
```
## 插件
插件可以对 webpack 进行功能上的扩充,比如代码压缩、添加版权信息等。与 loader 不同的是,loader 用于将特定类型的文件进行转换,而插件则是将转换后的代码作出进一步处理。
### 自动生成 HTML 文件
使用[HtmlWebpackPlugin](https://github.com/jantimon/html-webpack-plugin#configuration)可以在发布目录中自动生成一个 HTML 文件,并引入包含打包后`.js`文件。
```powershell
npm i --save-dev html-webpack-plugin
```
```js
const HtmlWebpackPlugin = require('html-webpack-plugin')
module.exports = {
plugins: [
new HtmlWebpackPlugin()
]
}
```
调用插件时可以传入一个对象作为配置参数,比如希望以自定义的模板来创建 HTML,可以使用[template 属性](https://github.com/jantimon/html-webpack-plugin/blob/master/docs/template-option.md)。它依然会自动在最后添加打包文件的引用,不需要手动添加。
```js
module.exports = {
plugins: [
new HtmlWebpackPlugin({
template: './index.html' // 相对于 webpack.config.js 文件所在目录
})
]
}
```
### 代码压缩
使用[UglifyjsWebpackPlugin](https://www.webpackjs.com/plugins/uglifyjs-webpack-plugin/)可以对代码进行压缩。
> 在 webpack4 中,如果配置了`mode: 'production'`(默认)选项,那么 webpack 会自动进行压缩,这里等用到时再研究。
## 其它功能
### 本地服务器
使用[webpack-dev-server](https://www.webpackjs.com/configuration/dev-server/)可以快速构建一个基于 Node.js 的本地服务器,它内置了 [Express 框架](/posts/o48l9v2o.html#构建网络服务),可以用于本地调试。
```powershell
npm install webpack-dev-server --save-dev
```
```js
module.exports = {
devServer: {
contentBase: path.join(__dirname, "dist"), // 根路径,通常配置为部署目录 dist
port: 9000 // 端口,默认为 8080
}
}
```
然后运行项目本地的`webpack-server-dev`启动服务器,使用 npx 或者 npm scripts 均可。
```powershell
npx webpack-server-dev
```
> 也可以直接使用 VSCode 中的 live-server 扩展。
### 分离配置文件
使用[webpack-merge](https://github.com/survivejs/webpack-merge)可以将配置文件分离成开发环境与生产环境,然后根据环境再合并为一个统一的配置文件。
<file_sep>---
title: 大前端笔记之19 😄 jQuery
date: 2020-02-08 11:14:12
abbrlink: xw3mvken
tags: JavaScript
categories: 大前端
excerpt: jQuery 是最常用的 JavaScript 的第三方库,它封装了复杂的原生 DOM、事件等操作,并且提供了一些简单的动画效果。
---
# 大前端笔记之19 😄 jQuery
[jQuery](https://jquery.com/)是最常用的 JavaScript 的第三方库,它封装了复杂的原生 DOM、事件等操作,并且提供了一些简单的动画效果。
## 基本概念
使用`$()`并传入字符串形式的 CSS 选择器即可获取一个 jQuery 对象,其中的`$`是`jQuery`的简写。注意,DOM 对象是原生的,而 jQuery 对象是将 DOM 对象封装后产生的,它们两者的属性和方法不能混用。
```js
$('.nav-item') // => jQuery 对象
```
要将 jQuery 对象转换成 DOM 对象,可以通过 jQuery 对象的索引取出 DOM 对象。这是因为 jQuery 的对象是将 DOM 对象封装在一个**伪数组**中:
```js
$('#footer')[0] // DOM 对象
```
或者使用 jQuery 对象的`get()`方法:
```js
$('#footer').get(0) // DOM 对象
```
要将 DOM 对象转换成 jQuery 对象,那么直接将它放到`$()`中就可以了:
```js
var footer = document.querySelector('footer') // DOM 对象
$(footer) // jQuery 对象
```
由于 jQuery 对象是一个伪数组,因此不能通过`null`判断要获取的 jQuery 对象是否存在。因为即便是没有找到对应的 DOM 对象,这个 jQuery 对象是肯定会存在的,所以应该通过**数组长度**来判断:
```js
var $footer = $("#footer")
if ($footer == null) {} // 错误
if ($footer.length == 0) {} // 正确
```
jQuery 对象拥有**隐式迭代**的特性,也就是说如果同时获取到多个页面元素,那么 jQuery 会自动在内部完成遍历,无需手动取出单个对象进行操作。
```js
$('.nav-item').css('background', '#66ccff') // 页面有多个 .nav-item 元素,会全部修改背景色
```
## 操作元素
jQuery 最主要的特性之一就是简化了 DOM 操作。
### children() / find()
获取该元素的子元素 / 后代元素。
| 参数 | 描述 |
| --- | --- |
| 选择器 | 对元素进一步过滤 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('div').children('p')
$('div').find('p')
```
### parent() / parents()
获取该元素的父元素 / 祖先元素。
| 参数 | 描述 |
| --- | --- |
| 选择器 | 对元素进一步过滤 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('span').parent('p')
$('span').parents('p')
```
### siblings()
获取该元素的兄弟元素,但不包括本身。
| 参数 | 描述 |
| --- | --- |
| 选择器 | 对元素进一步过滤 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('li').siblings()
```
### next() / nextAll()
获取该元素的下一个 / 后续全部兄弟元素。
| 参数 | 描述 |
| --- | --- |
| 选择器 | 对元素进一步过滤 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('#header').next()
$('#header').nextAll()
```
### prev() / prevAll()
获取该元素的上一个 / 之前全部兄弟元素。
| 参数 | 描述 |
| --- | --- |
| 选择器 | 对元素进一步过滤 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('#header').prev()
$('#header').prevAll()
```
### first() / last()
获取多个元素中的第一个 / 最后一个元素。
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('li').first() // 获取第一个 <li>
$('li').last() // 获取最后一个 <li>
```
### eq()
获取多个元素中的第 n 个元素。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 元素索引,从`0`开始计数,如果为负数则表示从后开始,`-1`表示最后一个元素 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('li').eq(0) // 获取第一个 <li>
```
### slice()
根据索引获取多个元素中之间的元素。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 起始索引(包括),如果为负数则表示从后开始,`-1`表示最后一个元素 |
| 整数值 | 结束索引(不包括),如果为负数则表示从后开始,`-1`表示最后一个元素;如果省略则一直到最后一个元素 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 符合条件的对象 |
```js
$('li').slice(0, 2) // 获取 第一个、第二个 <li>
```
### index()
获取元素相对于**兄弟元素**的索引位置,如果要查找的元素不止一个,那么以其中第一个为准。
| 返回值 | 描述 |
| --- | --- |
| 整数值 | 元素相对于兄弟元素的索引位置 |
```html
<ul>
<li></li>
<li></li>
<li class="target"></li>
</ul>
```
```js
$('.target').index() // => 2
```
### each()
遍历多个元素。
| 参数 | 描述 |
| --- | --- |
| 函数 | 迭代函数,注意其中的`this`表示当前 DOM 对象,而不是 jQuery 对象。该函数提供两个参数,第一个参数表示当前索引,第二个参数表示当前的 DOM 对象 |
```js
$('div').each(function (index, el) {
el // 当前 DOM 元素,与 this 相同
})
```
### append() / prepend()
将元素(参数,子元素)添加到另一个元素(调用者,父元素)的末尾 / 开头。
| 参数 | 描述 |
| --- | --- |
| DOM 对象 / jQuery 对象 / HTML 字符串 | 要添加的子元素,如果元素在页面上已经存在,那么该方法会**移动**该元素,而非复制 |
```html
<ul>
<li>咩咩子</li>
</ul>
```
```js
$('ul').append('<li>炎王喵</li>')
// <ul>
// <li>咩咩子</li>
// <li>炎王喵</li>
// </ul>
```
### after() / before()
将元素(参数)添加到另一个元素(调用者)之后 / 之前,作为其兄弟元素。
| 参数 | 描述 |
| --- | --- |
| DOM 对象 / jQuery 对象 / HTML 字符串 | 要添加的子元素,如果元素在页面上已经存在,那么该方法会**移动**该元素,而非复制 |
```html
<div class="target">炎王喵</div>
```
```js
$('.target').after('<div>咩咩子</div>')
// <div class="target">炎王喵</div>
// <div>咩咩子</div>
```
### clone()
复制元素,包括其中的子元素和文本。
| 参数 | 描述 |
| --- | --- |
| 布尔值 | 默认值为`false`,如果为`true`,则表示将元素绑定的事件一同复制 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 复制的元素 |
```js
var newLi = $('li').clone(true) // 同时复制事件
```
### wrap()
在指定元素外部包裹一个元素。
| 参数 | 描述 |
| --- | --- |
| DOM 对象 / jQuery 对象 / HTML 字符串 | 用来包裹在外层的元素 |
```js
$('p').wrap("<div></div>") // 在所有 <p> 外层包裹一个`<div>`
```
### remove()
删除元素,包括子元素和绑定的事件。
| 参数 | 描述 |
| --- | --- |
| 选择器 | 对元素进一步过滤 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 被删除的元素 |
```js
$('li').remove('.first') // 删除包含 first 类的 <li>
```
### empty()
清空元素,包括子元素和文本内容。
```js
$('ul').empty() // 清空 <ul>,但 <ul> 本身还在
```
### 元素过滤
在上述的选择器参数中,还可以继续使用下列过滤器对元素过滤。其中一些过滤器与上述某些方法功能相同,此时为了操作方便,应该优先考虑使用方法。
#### 基础过滤
索引都是从`0`开始的,因此第一个元素是索引为`0`的元素。
| 选择器 | 描述 |
| --- | --- |
| `:first` | 选择第一个元素 |
| `:last` | 选择最后一个元素 |
| `:not(selector)` | 选择不满足给定条件的元素 |
| `:even` | 选择索引为偶数的元素(索引从`0`开始) |
| `:odd` | 选择索引为奇数的元素(索引从`0`开始) |
| `:eq(index)` | 选择等于该索引的元素 |
| `:gt(index)` | 选择**大于**该索引的元素 |
| `:lt(index)` | 选择**小于**该索引的元素 |
| `:header` | 选择所有的标题元素 |
| `:focus` | 选择当前获得焦点的元素 |
```js
$('p').first() // 等价于
$('p:first')
```
#### 内容过滤
| 选择器 | 描述 |
| --- | --- |
| `:contains('text')` | 选择包含指定文本的元素,参数的引号可选 |
| `:empty` | 选择所有没有子元素的元素(包括文本节点) |
| `:parent` | 选择所有包含子元素的元素(包括文本节点) |
#### 可见性过滤
| 选择器 | 描述 |
| --- | --- |
| `:hidden` | 选择所有隐藏的元素 |
| `:visible` | 选择所有可见的元素 |
隐藏元素具体指的是:
- `display: none`的元素
- `type`属性为`hidden`的表单元素
- `width`和`height`都为`0`的元素
也就是说,如果一个元素没有占据布局空间,那么就被认为是隐藏的。注意,`visibility: hidden`或`opacity: 0`的元素被认为是可见的,因为它们依然占据布局空间。
#### 子元素过滤
| 选择器 | 描述 |
| --- | --- |
| `:nth-child()` | 选择第 n 个子元素 |
| `:first-child` | 选择第一个子元素 |
| `:last-child` | 选择最后一个子元素 |
注意,由于`:nth-child()`是基于 CSS 规范的选择器,因此与`:eq()`不同,它的索引是从`1`开始的。
此外,`:first-child`可以选择所有匹配的父元素下的所有子元素,而`:first`只能选择一个。例如:
```html
<ul>
<li>咩咩子</li>
<li>炎王喵</li>
<li>麒麟</li>
</ul>
<ul>
<li>钢龙</li>
<li>尸套龙</li>
<li>贝西摩斯</li>
</ul>
```
```js
$('ul li:first') // 只能选择咩咩子
$('ul li:first-child') // 选择咩咩子和钢龙
```
#### 表单元素过滤
| 选择器 | 描述 |
| --- | --- |
| `:input` | 选择所有的`<input>`、`<textarea>`、`<select>`和`<button>`元素 |
| `:text` | 选择所有`type="text"`或者没有设置`type`属性的`<input>`元素 |
| `:password` | 选择所有的密码框元素 |
| `:radio` | 选择所有的单选框元素 |
| `:checkbox` | 选择所有的复选框元素 |
| `:submit` | 选择所有的提交按钮(包括表单中没有设置`type`属性的`<button>`元素) |
| `:reset` | 选择所有的重置按钮 |
| `:button` | 选择所有的按钮元素 |
| `:file` | 选择所有的上传域 |
| `:checked` | 选择所有被选中的单选框和复选框 |
| `:selected` | 选择所有被选中的下拉列表项 |
| `:enabled` | 选择所有未被禁用的元素 |
| `:disabled` | 选择所有被禁用的元素 |
## 操作文本
### html() / text()
获取或设置单个元素的文本内容。后者会忽略内容中的 HTML 标签,返回只包含文字部分的内容。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 可选,如果为空则表示获取值,否则为设置值 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 / 字符串 | 如果是设置内容,则返回调用方法的元素;如果是获取内容,则返回内容的字符串 |
```html
<div class="container">
<div class="box">Demonstration Box</div>
</div>
```
```js
$('.container').html() // => <div class="box">Demonstration Box</div>
$('.container').text() // => Demonstration Box
```
## 操作属性
### val()
设置或者获取表单元素的`value`属性值。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 可选,如果为空则表示获取值,否则为设置值 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 / 字符串 | 如果是设置内容,则返回调用方法的元素;如果是获取内容,则返回内容的字符串 |
```js
$('input').val('我是谁?我在哪?') // 设置 <input> 元素的 value 属性值
$('input').val() // => 我是谁?我在哪?
```
### attr() / prop()
获取或设置元素的属性。它们的区别在于,`attr()`只能获取 HTML 标签上的属性,因此,对于当前不存在的属性,`attr()`会返回`undefined`,这会导致一些**布尔属性**如`checked`、`selected`或`disabled`无法正确的返回`false`值。而`prop()`获取的是元素 DOM 对象的属性,比 HTML 元素标签上写明的属性要多很多,因此可以正确获取布尔属性的值。
| 参数 | 描述 |
| --- | --- |
| 单个字符串 | 单个字符串为属性名,表示获取该属性 |
| 两个字符串 | 两个字符串分别为属性名和属性值,表示设置该属性 |
| 单个对象 | 同时设置多个属性 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 / 字符串 | 如果是设置内容,则返回调用方法的元素;如果是获取内容,则返回内容的字符串 |
```js
$('img').attr('src') // 获取单个属性
$('img').attr('src', '1.jpg') // 设置单个属性
$('img').attr({ // 设置多个属性
src: '1.jpg',
alt: '嗯,图片没显示出来'
})
```
### removeAttr()
移除元素的属性。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要移除的属性名 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$('img').removeAttr('src') // 移除 src 属性
```
### 类属性
jQuery 提供了下列方法用来简化类属性操作。如果需要同时操作多个类,可以使用空格将其隔开:
| 方法 | 描述 |
| --- | --- |
| `addClass()` | 添加类 |
| `removeClass()` | 删除类 |
| `hasClass()` | 判断该对象是否包含某个类 |
| `toggleClass()` | 如果某个类存在,则移除该类,否则添加该类 |
```js
$('button').addClass('btn-danger btn').removeClass('btn-warning')
$('button').hasClass('btn-danger btn')
$('button').toggleClass('btn-danger btn')
```
### 自定义属性
使用`data()`获取或设置元素的自定义属性,也可以获取元素上的`data-`属性,不需要添加前缀。注意,虽然通过该方法可以获取`data-`属性,但是该方法设置的属性却不会呈现到元素的`data-`属性上。
```js
$('div').data('id', 1) // 设置自定义属性
$('div').data('id') // => 1,获取自定义属性
```
## 操作样式
### css()
获取或设置元素的 CSS 样式。注意,该方法不支持简写属性,如`margin`、`border`等,表示属性值的参数如果为数字值,也可以不使用字符串。
| 参数 | 描述 |
| --- | --- |
| 单个字符串 | 单个字符串为样式名,表示获取该样式 |
| 两个字符串 | 两个字符串分别为样式名和样式值,表示设置该样式 |
| 单个对象 | 同时设置多个样式 |
| 单个数组 | 同时获取多个样式 |
```js
$('div').css('backgroundColor') // 获取单个样式
$('div').css(['width', 'height']) // => { width: "300px", height: "300px" },获取多个样式
$('div').css('width', '500px') // 设置单个样式
$('div').css({ // 设置多个样式
width: 500, // 可以是数字值或字符串
height: '500px'
})
```
> 不同于原生的样式操作,这里的样式名称也可以是连字符形式,也就是说`css('font-size')`和`css('fontSize')`都是合法的。
### 元素大小
使用 jQuery 对象的下列方法可以获取与元素大小相关的信息,它们均**不包含单位**。
| 方法 | 描述 |
| --- | --- |
| `width()` | 获取元素内容区的宽度 |
| `height()` | 获取元素内容区的高度 |
| `innerWidth()` | 获取元素`padding`区域的宽度 |
| `innerHeight()` | 获取元素`padding`区域的高度 |
| `outerWidth()` | 获取元素`border`区域的宽度,如果传入参数`true`,则表示`margin`区域 |
| `outerHeight()` | 获取元素`border`区域的高度,如果传入参数`true`,则表示`margin`区域 |
对于`width()`和`height()`来说,即便设置了`box-sizing: border-box`,那么返回的值是`width`依然是内容区的大小。
```css
div {
box-sizing: border-box;
width: 400px;
padding: 50px;
}
```
```js
$('div').width() // => 300
```
### 滚动距离
使用 jQuery 对象的下列方法可以获取或设置与元素滚动距离相关的信息,它们均**不包含单位**。
| 方法 | 描述 |
| --- | --- |
| `scrollLeft()` | 获取或设置元素水平方向滚动出去的距离 |
| `scrollTop()` | 获取或设置元素垂直方向滚动出去的距离 |
```js
// 获取页面的垂直滚动距离
$(document).scroll(function () { // 监听 window 和 document 对象均可
$(document).scrollTop() // 获取 window 和 document 对象均可
})
// 设置页面的滚动距离
$(document).scrollTop(500)
```
### 位置
使用 jQuery 对象的下列方法可以获取与元素位置相关的信息,它们均**不包含单位**。
| 方法 | 描述 |
| --- | --- |
| `offset()` | 获取元素相对于**文档**的位置,该方法以对象形式返回坐标,包含`left`和`top`两个属性 |
| `position()` | 获取元素相对于**最近且定位的父元素**的位置,该方法以对象形式返回坐标,包含`left`和`top`两个属性,对应原生的`offsetLeft`和`offsetTop`两个属性 |
```js
$('div').offset().top
$('div').position().top
```
## 动画
### fadeIn() / fadeOut() / fadeToggle()
使元素淡入显示 / 淡出隐藏 / 切换淡入淡出。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 可选,动画持续时间,单位为毫秒,默认为`400` |
| 字符串 | 可选,调速函数,默认为`swing`,可修改为`linear` |
| 函数 | 动画结束后的回调函数 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$('div').fadeIn(300, 'linear', function() {
console.log('动画完成!')
})
```
### fadeTo()
使元素淡入 / 淡出到指定的透明度。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 可选,动画持续时间,单位为毫秒,默认为`400` |
| 数值 | 目标不透明度 |
| 字符串 | 可选,调速函数,默认为`swing`,可修改为`linear` |
| 函数 | 动画结束后的回调函数 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
### slideDown() / slideUp() / slideToggle()
使元素滑动显示 / 滑动隐藏 / 切换滑动显示与隐藏。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 可选,动画持续时间,单位为毫秒,默认为`400` |
| 字符串 | 可选,调速函数,默认为`swing`,可修改为`linear` |
| 函数 | 动画结束后的回调函数 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$('div').slideDown(300, 'linear', function() {
console.log('动画完成!')
})
```
### animate()
自定义动画。该方法默认不支持颜色的动画效果,除非使用插件。
| 参数 | 描述 |
| --- | --- |
| 对象 | 包含 CSS 属性值的对象,表示动画的结束状态,属性值可以为**相对值**,比如`-=`或者`+=`,表示基于之前动画的结果进行新的动画 |
| 整数值 | 可选,动画持续时间,单位为毫秒,默认为`400` |
| 字符串 | 可选,调速函数,默认为`swing`,可修改为`linear` |
| 函数 | 动画结束后的回调函数 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$div.animate({
left: 1000,
width: 100
}, 1000, 'linear', function () {})
```
一些非 CSS 属性如`scrollTop`、`scrollLeft`也可用于动画。比如下面的代码可以使页面逐渐滚动到顶部:
```js
// 通过 html 还是 body 元素有待根据浏览器测试
$('html, body').animate({
scrollTop: 0
}, 1000)
```
### stop()
jQuery 会将动画效果存储到一个队列当中依次执行。但是这样会带来一个问题,如果连续触发动画,由于上一步的动画还没有完成,新的动画又被触发,因此会出现点击与动画不一致的情况。
要解决这个问题,可以在调用动画函数前添加`stop()`函数,使元素停止当前正在进行的动画,这样就能在每次执行新的动画前,结束上次动画的效果。
| 参数 | 描述 |
| --- | --- |
| 布尔值 | 是否清空整个动画序列,而不仅仅是停止**当前**动画 |
| 布尔值 | 是否立即完成当前动画,而不仅仅是**停止**动画 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$div.stop(true, true).slideToggle()
```
### delay()
延迟下一个动画的执行。
| 参数 | 描述 |
| --- | --- |
| 整数值 | 延迟时间,单位为毫秒 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$div.fadeIn().delay(200).fadeOut()
```
## 事件
### on() / one()
为元素绑定一个或多个事件。区别在于后者当**每个元素**上触发**同一个事件**后,它绑定的事件就会被自动解除。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 事件名称,多个事件以空格隔开 |
| 选择器 | 表示事件委托的子元素(见下文) |
| 函数 | 事件监听函数,该函数提供一个参数表示事件对象 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$btn.on('click mouseover', function (e) { // 为元素绑定多个事件
console.log('事件被触发!')
})
```
通常情况下事件会绑定到要触发的元素上,但这样有一个问题,页面加载完成后新添加的元素是不会有事件的。
因此 jQuery 提供了一种**委托事件**的机制,将事件绑定到已经存在的父元素上,然后利用事件冒泡把事件交给子元素,这样无论子元素绑定事件时是否存在,都可以从父元素获取到相应的事件。
要使用事件委托,需要在`on()`方法中传入一个选择器参数,表示要绑定的子元素,再通过**父元素**调用`on()`方法即可:
```js
$('ul').on('click', 'li', function () {
console.log('事件被触发啦')
})
```
此外,对于一些常用的事件,jQuery 也提供了单独的方法进行绑定,相当于通用事件的快捷版本。
```js
$btn.on('click', function () {}) // 等价于
$btn.click(function () {})
```
### off()
移除元素的事件。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 事件名称,多个事件以空格隔开,如果为空则移除元素上的所有的事件 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$btn.off('click mouseover') // 移除对应的事件
$btn.off() // 移除所有的事件
$btn.off('click', 'li') // 移除事件委托
```
### trigger() / triggerHandler()
触发元素的事件。区别在于,后者不会触发元素的默认行为(比如文本框的`focus`会自动定位光标),并且不会将事件冒泡。
```js
$btn.trigger('click')
```
此外,如果事件方法没有传入事件监听函数,那么相当于触发该元素的对应事件,为`trigger()`的快捷版本。
```js
$btn.click()
```
### hover()
同时设置鼠标进入和离开的事件,相当于同时绑定了`mouseenter`和`mouseleave`两个事件。
| 参数 | 描述 |
| --- | --- |
| 一个函数 | 事件监听函数,鼠标进入和离开时触发 |
| 两个函数 | 事件监听函数,前者鼠标进入时触发,后者鼠标离开时触发 |
| 返回值 | 描述 |
| --- | --- |
| jQuery 对象 | 调用方法的元素 |
```js
$('div').hover(function () {})
```
## 扩展方法
除了上述的特性以外,jQuery 也提供了一些方法用来扩展原生的对象操作。它们均是通过`jQuery`对象也就是`$`调用。
### extend()
将多个对象合并成一个。如果目标对象与源对象属性出现重复,那么目标对象中的属性会被覆盖。该方法会修改保留的对象。
| 参数 | 描述 |
| --- | --- |
| 布尔值 | 可选,表示深拷贝 |
| 对象 | 合并后保留的对象 |
| 若干对象 | 合并后被删除的对象 |
| 返回值 | 描述 |
| --- | --- |
| 对象 | 合并后保留的对象 |
```js
var target = {id: 1}
var source = {id: 233, name: 'JOJO'}
$.extend(target, source) // => {id: 233, name: "JOJO"}
target // => {id: 233, name: "JOJO"}
```
如果在目标对象参数前添加一个布尔值参数,则表示深拷贝。即如果源对象中包含复杂数据类型(如数组、对象等),那么采用递归方式复制对象内容,而不是简单的复制地址。
```js
var target = {}
var source = {
address: { city: 'NY' }
}
// 默认浅拷贝,target 和 source 中的 address 对象都指向一个值
$.extend(target, source)
target.address.city = 'Tokyo'
source.address.city // => Tokyo
// 深拷贝,target 中的 address 对象为新的内容
$.extend(true, target, source)
target.address.city = 'Tokyo'
source.address.city // => NY
```
### each()
遍历对象。与 jQuery 元素的`each()`方法不同的是,`each()`仅能遍历 jQuery 对象,而`$.each()`还可以遍历数组或者 JavaScript 的键值对对象。
| 参数 | 描述 |
| --- | --- |
| 函数 | 迭代函数,该函数提供两个参数,第一个参数表示当前索引,第二个参数表示当前元素 |
```js
var arr = ['red', 'blue', 'green']
$.each(arr, function (index, el) {
index + ' ' + el
})
// 0 red
// 1 blue
// 2 green
var arr = {name: 'dante', age: 18}
$.each(arr, function (index, el) {
index + ' ' + el
})
// name dante
// age 18
```<file_sep>---
title: 大前端笔记之13 😍 JavaScript 面向对象
date: 2020-02-01 9:23:53
abbrlink: vmir6875
tags: JavaScript
categories: 大前端
excerpt: 虽然 JavaScript 对于面向对象的实现与其它语言有所不同,但是它同样是一门面向对象的语言。
---
# 大前端笔记之13 😍 JavaScript 面向对象
虽然 JavaScript 对于面向对象的实现与其它语言有所不同,但是它同样是一门面向对象的语言。
## 构造函数
JavaScript 在 ES6 之前没有类的概念,只能通过构造函数作为对象的「模板」。构造函数本质也只是一个普通函数,但是使用`new`调用后,就可以实例化一个对象。
```js
function Person(name, age) { // 习惯上构造函数首字母大写
this.name = name
this.age = age
this.eat = function() { console.log('吃呀吃') }
}
var person = new Person('Claire', 19)
person.name // => Claire
```
构造函数中使用了`this`关键字,这里的`this`指代的是该构造函数创建的**实例对象**。也就是说,构造函数创建的对象是`person`,因此`this.name`指的是实例对象`person`的属性,第二个`name`则是传递的参数,只用来接收值。
## this 关键字
`this`在不同场合所指向的对象会有所不同,但是无论如何,它总是会指向一个对象。
### 构造函数中的 this
正如上文所说,构造函数中的`this`指向其创建的实例对象。
```js
function Person(name) {
this.name = name
}
var person = new Person('Claire')
person.name // => Claire
```
### 全局作用域中的 this
全局作用域中的`this`指代全局对象`Window`(仅限于浏览器作为宿主环境时)。
```js
console.log(this) // Window
```
### 普通函数中的 this
普通函数中的`this`指代**当前调用函数的对象**,与函数的定义位置、调用位置都没有关系,只与函数的**调用者**有关。如果函数没有调用者,则`this`同样指代全局对象`Window`。
在下面的代码中,由于`foo()`前没有具体的调用对象,因此函数中的`this`指代的是`Window`,从而找到全局作用域中`num`的值为`666`。
```js
function foo() {
var num = 233
console.log(this.num)
}
var num = 666
foo() // => 666
```
即便是嵌套的函数也是一样:
```js
function outer() {
var num = 233
function inner() { console.log(this.num) }
inner() // 没有调用者,依然是全局对象
}
var a = 666
outer() // => 666
```
如果函数是通过对象调用的,那么在该函数中,`this`指代的就是调用函数的对象。例如:
```js
function foo() { console.log(this.num) }
var obj = { num: 233, fn: foo }
var num = 666
obj.fn() // => 233
```
即便函数发生了传递,那么依然也要看**最终调用**时前面是否有对象。因为 ECMAScript 中的函数传递,本质上传递的只是函数的代码。
```js
function foo() { console.log(this.num) }
var obj = { num: 233, fn: foo }
var num = 666
var newFn = obj.fn
newFn() // => 666,因为最终调用时前面没有对象
```
### 定时器函数中的 this
定时器中的`this`指代的是全局对象`Window`。(见[BOM](/posts/ybdcxino.html#定时器)一节)
```js
setInterval(function () {
this // => window
}, 1000)
```
### 事件监听函数中的 this
事件监听函数中的`this`指代的是触发事件的对象。(见[事件](/posts/uxfasi54.html)一节)
```js
btn.addEventListener('click', function () {
this // => <button id="btn">触发事件</button>
})
```
### 改变 this 指向
正如上文所说,随着场景不同`this`的指向也会不同,但是通过一些方式可以修改特定场景下`this`的指向。
#### call()
使用函数调用`call()`,并传入一个对象作为参数,会导致函数中的`this`指向该对象参数:
```js
var obj = { age: 18 }
function say() { console.log(this.age) }
say() // => undefined,this 指向 Window,但是 Window 中没有 age 属性
say.call(obj) // => 18,this 指向 obj
```
如果没有传参数或者传入`null`,那么相当于直接调用函数,不会改变`this`的指向。
```js
function say() { console.log(this) }
say() // => Window
say.call() // => Window
```
如果传入两个以上的参数,则后面的参数表示函数本身的参数。
```js
function getSum(num1, num2) {
return num1 + num2
}
getSum.call(null, 10, 20) // => 30
```
#### apply()
它的用法与`call()`几乎一致,区别在于`apply()`的函数参数,需要以数组的方式传入。
```js
getSum.apply(obj, [10]) // => 参数必须是数组
```
利用这一特性,可以借助`Math.max()`方法获取数组中的最大值,因为它的参数是若干数值,而使用`apply()`可以将一个数组作为一系列数值参数传入:
```js
Math.max.apply(null, [10, 20, 30]) // 不需要改变 this 指向,传入 null 即可
```
#### bind()
该方法会改变`this`的指向,但是不会调用函数,而是返回一个改变了`this`指向的新函数。
```js
function fn() { console.log(this) }
var obj = {}
var newFn = fn.bind(obj)
newFn() // => obj
```
## 原型
虽然 JavaScript 提供的构造函数可以作为模板,用来创建多个实例对象。但是,由同一个构造函数创建的多个实例,即使它们的方法是相同的,也会各自开辟空间,这样会造成系统资源的浪费。
在下面的代码中,虽然`cat`和`dog`的`run()`方法内容完全相同,但是指向的对象却不相同。
```js
function Animal() {
this.run = function () { console.log('各种跑~') }
}
var cat = new Animal()
var dog = new Animal()
cat.run == dog.run // => false
```
因此,JavaScript 引入了原型(prototype)的概念。原型的**本质是一个对象**,通过构造函数的属性`prototype`,或者实例对象中的`__proto__`属性(该属性为浏览器使用,不推荐程序员调用)可以获取。当实例对象在本身找不到要调用的方法时,会通过`__proto__`属性获取原型对象,再从原型对象中继续寻找。于是,将相同的方法定义到原型对象中,就可以达到**共享资源,节省空间**的目的。

因此上面的示例可修改为:
```js
function Animal() {}
// 将 run() 方法添加到原型对象中
Animal.prototype.run = function () { console.log('各种跑~') }
var cat = new Animal()
var dog = new Animal()
cat.run == dog.run // => true
```
### 原型链
其实原型对象中也有`__proto__`属性,该属性同样指向另一个原型对象,它们形成了一个链条,称为原型链。
当调用对象的某个属性时,会先从实例对象本身寻找,如果不存在则去它的原型对象中找,如果依然不存在则去它的原型对象的原型对象中找。链条的顶端是`Object`构造函数的原型对象`Object.prototype`,该对象不再有原型,其`__proto__`属性的值为`null`。
<file_sep>---
title: 大前端笔记之12 📝 JavaScript 数据类型
date: 2020-01-30 10:59:41
abbrlink: p4xy9gkd
tags: JavaScript
categories: 大前端
excerpt: JavaScript 中的值分为六种数据类型,其中除了<code>Object</code>之外均为基本类型,而<code>Object</code>又包含三种类型,狭义的对象、数组和函数。
---
# 大前端笔记之12 📝 JavaScript 数据类型
JavaScript 中的值分为六种数据类型,其中除了`Object`之外均为基本类型,而`Object`又包含三种类型,狭义的对象、数组和函数。
## 基本数据类型
基本数据类型包括`Undefined`、`Null`、`Boolean`、`String`和`Number`。
> 虽然`Boolean`、`String`和`Number`是基本数据类型,但是它们均有对应的**包装类**对象。当它们调用方法时,这些值会被临时转换为包装类对象,调用结束后就会被立刻销毁。
### Undefined 和 Null
JavaScript 中这两个类型均表示「空」,它们都只有一个值,即小写的`undefined`和`null`。
对于`undefined`来说,它不能手动定义,只会在下列情况下出现:
- 声明了变量,但没有赋值,该变量为`undefined`
- 数组中的空元素(修改数组长度、跨索引添加元素等)为`undefined`
- 调用函数时,没有传入实参,那么形参为`undefined`
- 访问**对象**中未声明的属性,该属性为`undefined`。如果访问一个未声明的变量,那么会报错
- 函数没有返回值,则返回`undefined`
对于`null`来说,如果一个对象类型的变量没有指向任何一个实际对象,那么该变量的值就是`null`。
### Boolean
布尔类型`Boolean`只有`true`和`false`两个值。
### String
字符串类型`String`使用单引号`''`或双引号`""`均可创建。不过由于 JSON 格式内部只能使用双引号,因此建议使用单引号创建字符串,否则就要使用转义字符`\`。
```js
var str = 'helloworld'
```
使用`length`属性可以获取字符串长度。
```js
'helloworld'.length // => 10
```
### Number
数值类型`Number`采用[IEEE 754](/posts/xp50sskp.html)标准,不区分整数值和小数值,所有数值都是以 64 位浮点数存储,即便是整数也是如此。
全局属性`NaN`表示非数字(Not a Number),当某些计算失败时(如负数开偶次方),返回的结果可能为`NaN`。它并不是独立的数据类型,而是`Number`类型的一个特殊值。
此外,`NaN`不等于任何值,包括**它本身**。由于这个特性,JavaScript 提供了全局方法`isNaN()`用来判断一个值是否为`NaN`。
```js
isNaN(NaN) // => true
```
## 数组
数组以`Array`内置对象表示。它可以同时存放不同类型的数据,但通常没有必要,因为这样无法对数据进行统一的处理。数组的本质其实是有序的键值对,但是它的特殊之处在于,它的键是有序的数字,而非具体的名称。
### 创建数组
要创建数组,主要通过字面量方括号`[]`语法。
```js
var arr = ['a', 'b', 'c']
```
> 通过`Array`对象的构造函数`Array()`也可以创建数组,但是根据传入的参数不同,其行为并不一致,因此并不推荐。
### 访问数组
使用`数组名[索引]`的方式可以获取数组中对应索引的元素,因此通过`for`循环可以遍历数组。
```js
var arr = ['a', 'b', 'c']
arr[1] // => b
for (var i = 0; i < arr.length; i++) {
arr[i]
}
```
> 除了使用`for`循环以外,也可以使用数组的[相关方法]((/posts/i82icjg7.html#Array))或者[`for of`循环](/posts/j5h1kgw7.html#for-of-循环)遍历。
### 数组长度
使用`length`属性获取数组的长度。JavaScript 中的数组长度是**动态**和**可写**的,因此将数组长度设置为`0`可以清空数组。
```js
var arr = ['a', 'b', 'c']
arr.length // => 3
arr.length = 2 // 数组 arr 会变为 ['a', 'b']
```
## 函数
JavaScript 中的函数以`Function`内置对象表示。
### 创建函数
第一种方式称为**函数声明**:
```js
function sendMsg() { console.log('S.H.E.I.L.D') }
```
第二种方式称为**函数表达式**,函数表达式大多是没有函数名称的,这样的函数也被称为**匿名函数**:
```js
var print = function() { console.log('S.H.E.I.L.D') }
```
JavaScript 会根据代码本行的开头是否为`function`来区分函数声明和函数表达式。
> 由于函数也是对象,其实也可以通过`new Function()`构造函数的方式来创建,但这种方式并不直观,因此没有人使用。
### 调用函数
在函数声明的名称,或者保存函数表达式的变量后面添加一对圆括号`()`表示调用该函数。
```js
function sendMsg() { console.log('S.H.E.I.L.D') }
sendMsg() // 调用函数
```
### 参数
调用函数的同时也可以传入参数。在 JavaScript 中即使在创建函数时定义了形参,但调用时并**不要求**提供所有的参数,省略的参数的值为`undefined`,多余的参数会被忽略。
```js
function print(a, b){
a // => 10
b // => undefined
}
print(10) // 只传入了一个参数
```
由于参数的数量不确定,因此`Function`对象中提供了`arguments`伪数组,用来在函数体内部读取所有参数。
```js
function foo() {
arguments[0] // => 10
arguments[1] // => 20
arguments.length // => 2
}
foo(10, 20)
```
### 返回值
使用`return`语句设置函数的返回值,函数在执行时遇到`return`语句会立刻终止。如果没有返回值,或者`return`后面没有内容,那么返回`undefined`。
```js
function getSum(a, b) { return a + b }
```
### 函数是一等公民
如果一个函数可以存储在变量或数据结构中,并且可以进行引用传递,那么称这样的函数为**一等公民**,表示它和其它类型享有同样的待遇。因此,JavaScript 中的函数可以直接赋值给一个变量,也可以作为函数的结果返回。也就是说,函数声明的名称本质就是一个**保存了函数代码的变量**。
在下面的代码中,将函数的代码传递给了其它变量,通过该变量依然可以调用函数:
```js
function print() { console.log('我是一等公民啦!') }
var other = print
other() // => 我是一等公民啦!
```
### 立即执行函数
如果希望函数声明后立刻调用,而不是单独写一条调用语句,或许会尝试下面的写法:
```js
function sendMsg() { console.log('S.H.E.I.L.D') }() // => SyntaxError: Unexpected token (
```
但是这样是错误的,因为代码的首行开头是`function`,编译器会认为这是一个函数声明,不能以圆括号结尾。那么解决的办法就是不要让`function`出现在行首,让引擎将其理解成一个表达式。下面是两种比较常见的写法:
```js
(function() { // 由于已经成为了表达式,函数名称可以省略
console.log('S.H.E.I.L.D')
})();
// 或者
(function() {
console.log('S.H.E.I.L.D')
}());
```
这样引擎就会将其作为一个表达式来执行,它的最大作用是**创建一个独自的作用域,避免命名发生冲突**。
> 如果采用了无分号的代码风格,那么立即执行函数的前面应当添加一个分号`;(function(){})()`,防止解析出错。
### 作用域
作用域指的是变量所在的范围,它会把自己范围内的变量收集起来并统一管理。作用域分为两种(ES6 中还新增了块级作用域):
- 全局作用域:在整个运行过程中一直存在的作用域。JavaScript 会将一个页面中的全部`<script>`标签和单独的`.js`文件都合并成一个整体,因此即便是**多个文件也是共享一个全局作用域**。在全局作用域中声明的变量称为**全局变量**,在程序任何位置都可以访问。
- 局部作用域(函数作用域):每声明一个函数,就会在**函数内部**创建一个新的局部作用域。在局部作用域中声明的变量称为**局部变量**,只能在该函数内部访问。此外,JavaScript 采用的为**词法作用域**,也就是说**作用域由函数声明**时的位置决定,与调用的位置无关。
```js
/* 全局作用域 */
var num = 10
console.log(num) // => 10
function foo() {
/* 局部作用域 */
var num = 20
console.log(num) // => 20
}
```
#### 作用域链
当函数发生嵌套时,其对应的局部作用域也会嵌套。因此作用域除了保存范围内的变量之外,还会**保存外部的作用域**。因此,在当前的作用域中无法找到某个变量时,就会在嵌套的上一级作用域中继续查找该变量,直到全局作用域为止。这一连串的作用域被称为**作用域链**。
```js
/* 全局作用域 */
var num = 10
function outer() {
/* outer 的局部作用域 */
var num = 20
function inner() {
/* inner 的局部作用域 */
console.log(num) // => 20,inner 作用域内没有 num 变量,因此去上一级 outer的作用域中找
}
}
```
#### 提升
JavaScript 在所有代码执行前会首先收集所有**变量声明**和**函数声明**(不包括函数表达式),然后将它们提升到**所在作用域**的开头。例如:
```js
console.log(a) // => undefined
var a = 2
// 等价于
var a
console.log(a)
a = 2
```
函数声明也会被提升:
```js
fun() // => 我被提升啦
function fun() { console.log('我被提升啦') }
```
### 闭包
当内部函数通过作用域链访问了外部函数的变量,则称外部函数为一个闭包,通过 Chrome 设置断点可以查看:
```js
function father() {
var num = 1
function son() { console.log(num) } // 访问了外部函数的 num 变量
son()
}
father()
```

不过这样的闭包并没有意义,它的最主要作用在于**延长变量的生命周期**。在下面的代码中,每次调用`father()`函数,`num`的值都是`1`,因为当调用结束后,`father()`的作用域就被销毁了。
```js
function father() {
var num = 1
function son() { console.log(num++) }
son()
}
father() // => 1
father() // => 1
```
但是,如果将内部函数作为返回值,传递到外部,再由外部调用,结果就不一样了:
```js
function father() {
var num = 1
function son() { console.log(num++) }
return son
}
var fn = father()
fn() // => 1
fn() // => 2
```
这是因为,虽然`father()`调用结束了,但是它其中的函数被传递到了外部,由于内部函数依然在使用,所以`father()`的作用域也被保存了下来。
> 可以这么理解:我叫独孤求败(内部函数),我在一个山洞(闭包)里,里面有世界上最好的剑法和武器(内部变量)。我学习了里面的剑法,拿走了最好的剑,离开了这里(闭包返回了函数)。我来到这个江湖,快意恩仇,但是从来没有人知道我这把剑,和我这一身的武功的来历(外部无法访问闭包内部的变量,只有内部函数才可以)。
## 对象
这里的对象指的是狭义的对象,它的本质是无序的键值对。
### 创建对象
使用字面量花括号`{}`语法创建一个对象。对象的属性名称可以加引号,也可以不加,通常建议**不加引号**。但是,如果属性名称不符合标识符规则(如包含短横线`-`或空格、以数字开头等),那么必须加引号。
```js
var person = {
name : 'Claire',
age : 19,
eat: function() { console.log('吃呀吃') }
}
```
> 通过`Object`对象的构造函数`Object()`也可以创建对象,但是在这种方式下只能通过单独的赋值语句为对象添加属性,因此不推荐使用。
### 访问属性
使用`.`运算符获取对象的属性:
```js
var person = { age: 18 }
person.age // => 18
```
除此之外,也可以使用方括号`[]`语法,这种方式可以使用变量作为属性名:
```js
var person = { age: 18 }
var attr = 'age'
person['age'] // => 18
person[attr] // => 18,此时的 attr 是一个变量,而非属性名
```
由于对象为键值对,因此没法使用普通`for`循环来遍历对象,取而代之的是`for in`循环。
```js
var person = { name: 'Claire', age: 19 }
for (var key in person) { // 设置一个临时变量 key,命名随意
key // 遍历键
person[key] // 用方括号语法遍历值
}
```
### 判断属性存在
使用`in`可以判断对象中是否存在某个属性。
```js
var person = { name: 'Claire', age: 19 }
'name' in person // => true
```
## 自动类型转换
在某些特殊情况下,JavaScript 中的数据类型会发生自动类型转换。
### 转换为布尔型
在某些情况下(比如将值用于`if`条件判断),JavaScript 会自动调用构造函数`Boolean()`将一些值转换为`Boolean`类型。转换规则如下:
| 数据类型 | 结果 |
| --- | --- |
| `Undefined` | 转换为`false` |
| `Null` | 转换为`false` |
| `Number` | `0`或者`NaN`,转换为`false`,其它转换为`true` |
| `String` | 空字符串`''`转换为`false`,其它转换为`true` |
| `Object` | 转换为`true` |
### 转换为字符串
在某些情况下(比如使用加法运算符`+`将其它数据类型与字符串相连),JavaScript 会自动调用构造函数`String()`将一些值转换为`String`类型。而转换的规则与使用这些值手动调用[toString()](/posts/i82icjg7.html)方法是完全一样的。
### 转换为数字型
在某些情况下,JavaScript 会自动调用构造函数`Number()`将一些值转换为`Number`类型。转换规则如下:
| 数据类型 | 结果 |
| --- | --- |
| `Undefined` | 转换为`NaN` |
| `Null` | 转换为`0` |
| `Boolean` | `true`转换为`1`,`false`转换为`0` |
| `String` | 见下文 |
| `Object` | 见下文 |
使用`Number()`将`String`类型转换成`Number`,规则如下:
- 如果字符串为合法数字,那么将其转换成十进制数值,例如`'123'`转为`123`,`'023'`转为`23`、`'1.2'`转为`1.2`
- 如果字符串为空,则转换成`0`
- 除以上情况外,均转换成`NaN`
使用`Number()`将`Object`类型转换成`Number`,规则如下:
- 首先调用对象的`valueOf()`方法,然后按照上述规则进行转换
- 如果上一步结果为`NaN`,那么调用对象的`toString()`方法转换成字符串,然后再按照上述规则进行转换
可以看到这个过程比较复杂,因此通常会手动调用`Number`对象的相关方法来进行数字转换(见[标准库](/posts/i82icjg7.html#Number)一节)。
## 判断数据类型
使用下面的方式可以判断一个值的数据类型。
### typeof
使用`typeof`运算符获取一个值的数据类型。其中的函数并非数据类型,但有特殊返回值。
| 数据类型 | 返回值 |
| --- | --- |
| `Undefined` | `undefined` |
| `Null` | `object` |
| `Boolean` | `boolean` |
| `String` | `string` |
| `Number` | `number` |
| `Object` | `object` |
| 函数 | `function` |
```js
typeof 'abc' // => string
typeof 123 // => number
```
### instanceof
虽然`typeof`可以检测基本类型,但是对于引用类型,无论什么类型的对象都会返回`object`。因此要判断一个对象是不是某个类型的实例时,可以使用`instanceof`,它根据原型链来识别该实例是否属于某个对象。
```js
var arr = [1, 2, 3]
arr instanceof Array // => true
```
### toString()
由于任何对象调用[toString()](/posts/i82icjg7.html)方法默认会返回表示该对象类型的字符串,因此可以利用这一特性判断对象的数据类型。<file_sep>---
title: 计算机网络笔记之07 💻 应用层
date: 2020-01-19 10:16:58
abbrlink: dzta2ga7
tags: 计算机网络
categories: 计算机网络
excerpt: 应用层主要是由若干相互独立的服务和协议所构成的,每个服务或协议可以协同工作从而为计算机上的应用程序提供网络服务。
---
# 计算机网络笔记之07 💻 应用层
> 施工中...修改时间
应用层主要是由若干相互独立的服务和协议所构成的,每个服务或协议可以协同工作从而为计算机上的应用程序提供网络服务。
## DNS
用户(客户端)与互联网上的其它主机(服务器)通信时,必须知道对方的 IP 地址,但是 IP 地址由纯数字组成,非常难以记忆。因此为了既方便计算机处理,又方便用户识别记忆,域名系统 DNS(Domain Name System)应运而生。它可以将一个域名解析成一段 IP 地址,用户在使用域名访问服务器之前,需要先访问 DNS 服务器,从中获得该主机域名对应的 IP 地址,然后再使用 IP 地址访问指定的服务器。
当然,为了提高访问速度,减少网络资源浪费,系统会将访问过的网站 DNS 存放到系统缓存中(在命令行输入`ipconfig /displaydns`可以查看)。然后,用户也可以在本地的 hosts 文件中(`c:\Windows\System32\drivers\etc\hosts`)配置某个域名对应的实际 IP 地址。如果上述两个步骤均没有找到,那么系统才会访问 DNS 服务器以获取域名对应的 IP 地址。
### 域名
早期的互联网其实就是以一个简短的单词作为 IP 地址的名称的,但是随着网站越来越多,一个单独的名称已经不能满足需求了。于是互联网就采用了目前这种树状的具有层级的命名方法,其中的每一层称为一个**域**,一个完整的域名由多个层级即多个域组成,因此才称为域名(Domain Name)。
顶级域名大部分由国家或特定机构持有,之后也开放了普通公司机构的申请权利,但是费用极高(几十万美元)。常见的`.com`、`.cn`、`.edu`、`.org`等均是顶级域名。
二级域名位于顶级域名的下一层,个人和公司均可以申请,价格也非常便宜(指没有被注册过的),我们平时需要付费购买的基本限于二级域名。如`google.com`中的`google`就是二级域名。
由此继续向左延伸,也可以继续有三级域名、四级域名等,但是域名层级越深,记忆难度越大,使用的人也相对较少。
每个域名均是独一无二的,因此个人在申请域名之前,需要查询该域名是否已经被人注册,如果是他人已经注册的域名,也可以通过协商的方式进行购买并转让。
### 记录类型
购买域名之后,还需要为域名配置解析,也就是在 DNS 服务器登记。记录的类型有很多种,包括:
- `A`记录:最常见的记录类型(A 表示 address),根据域名返回对应的 IPv4 地址
- `CNAME`记录:别名记录,当查询该域名(如`testa.com`)时,返回另一个域名(如`testb.com`),再通过`testb.com`查询对应的`A`记录,从而获得对应的 IP 地址。注意,使用`CNAME`记录配置的记录值(别名)必须在本域名之下有`A`记录,不能解析到其他人的域名
- `AAAA`记录:根据域名返回对应的 IPv6 地址
如果你申请了一个域名如`muyumiao.com`,想要用户使用`muyumiao.com`和`www.muyumiao.com`两种方式访问,那么需要配置`A`记录中对应的主机记录,也就是主机名称。第一种方式主机名为空,第二种方式主机名为`www`,因此对应的主机记录分别为`@`(表示主机名为空)和`www`。而`A`记录的记录值填写的是主机的 IP 地址。最下方的 TTL 表示该记录的缓存时间,也就是多长时间之后,系统会再次去 DNS 服务器获取 IP 地址,而不是本地缓存。

## URL
URL(Uniform Resource Locator,统一资源定位符)类似于现实中的居住地址,可以用来定位互联网上某个资源的位置。它的具体格式如下:

> 严格来说,URL 属于 URI(Uniform Resource Identifier,统一资源标识符)的一种,表示通过地址来描述一个资源。但是,要描述资源不仅只有地址一种方式,也可以通过名称来描述。这种描述方式称为 URN(Uniform Resource Name,统一资源名称),比如常见的书籍国际标准书号 ISBN 就是 URN 的一种。
>
> 总的来说,URI 包括了 URL 和 URN 两种具体描述方式,只是在 Web 中 URL 更加常见。有些文档或书籍中也会以 URI 来表述,但是其含义与 URL 其实是一致的。
**协议**标明了通信双方需要共同遵循的格式约定,这里主要使用的是应用层的 HTTP、HTTPS 或 FTP 协议。
**服务器地址**包括了主机名称和域名,已经在上文 DNS 中详细讨论过。
**端口号**
#### 路径
这一段通常表示资源在服务器的物理路径,但是,返回的内容其实全凭服务器如何解析,因为本质上来说,它只是一段普通的字符串而已。这种根据路径来选择资源的过程,称为**页面的路由**。
路径分为绝对路径和相对路径,在一些 HTML 标签中,如果请求资源与当前资源的服务器、端口等均相同,那么这些部分是可以省略的。如果路径以`/`开头,那么表示从服务器的根路径开始定位;否则的话,则是以当前资源的路径开始定位。
对于相对路径,`./`表示的是当前路径,多数情况下与省略不写是一样的,而`../`表示上一级路径。
```html
<!-- 当前资源为 http://www.test.com/public/index.html -->
<!-- 绝对路径,相当于 http://www.test.com/test.html -->
<a href="/test.html">
<!-- 相对路径,相当于 http://www.test.com/public/test.html -->
<a href="test.html">
```
#### 查询参数
客户端发起请求的时候同时也可以携带参数,其中一种方式就是通过 URL 的`?`部分传递。这些参数以`=`分隔,前面为参数名,后面为参数值。多对参数以`&`分隔。
#### 哈希值
哈希可以用来进一步定位页面上的某一个部分,只需要在元素上添加`id`属性,使哈希与该属性值一致即可。注意,**如果只有哈希部分发生了改变,那么浏览器是不会再次向服务器发起请求的**,这是之后前端路由技术的原理。
## HTTP
HTTP 协议是互联网通信主要使用的协议,通过它传递的数据称为报文。
### 报文
此时浏览器会将客户端的信息打成一个包(称为请求报文),把它通过通道发送到服务器端。但是,服务器必须知道这个包的格式才能拆包,从中提取出想要的信息。因此,双方会以约定好的格式来发送内容,这一格式就是 **HTTP 协议**。
请求报文和接下来的响应报文必须按照一定的格式来保存,这样双方才能从中解析出需要的数据。
请求报文分为三个部分,由上到下为**请求行**、**请求头**与**请求体**,其中请求头包含了客户端的额外信息(浏览器版本、Cookie 等),请求体为发送到服务端的参数(比如注册信息、商品的当前页数等)。具体格式如下:
服务器上通常都会安装类似于 Apache 的服务器软件,用来处理来自浏览器的请求。由于 HTTP 协议是通过`80`端口传输的,因此默认情况下 Apache 会监听服务器的`80`端口,一旦发现了请求,则会根据请求文件的扩展名找到对应的 MIME Type(比如 Apache 的`mime.types`文件),判断用户请求的是静态资源(图片、CSS / JS 文件等)还是动态页面。如果是静态资源,那么直接找到相应文件;如果是动态页面,则需要调用该类型文件的处理工具(比如 PHP),将其转换成浏览器可以识别的 HTML。
请求方法分为`GET`和`POST`两种,`GET`通常使用地址栏后`?`部分传递参数,如`http://127.0.0.1:3000?id=233&page=1`,其中`id=233&page=1`就是参数。而`POST`会将请求参数放在请求体中,打开浏览器控制台工具可以查看:
然后,服务器也会将返回的数据打成一个包(称为响应报文),同样包含了服务器端的信息、返回的页面内容,并通过`80`端口返回。
响应报文同样分为三个部分,由上到下为**响应行**、**响应头**与**响应体**,其中响应头包含了服务端的额外信息,请求体通常为返回的页面内容。具体格式如下:
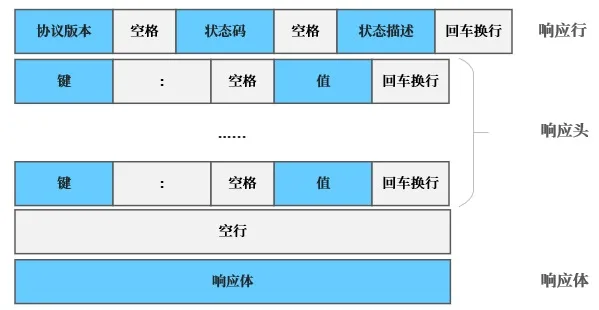
其中常见的状态码有(位于响应行):
* `200`:请求成功
* `404`:资源没有找到
* `500`:服务器端有语法错误
* `400`:客户端请求有语法错误
常见的 MIME 类型有(位于响应头的`content-type`键值对):
* HTML 文件:`text/html`
* CSS 文件:`text/css`
* JavaScript 文件:`application/javascript`
* 图片文件:`image/jpeg`
* JSON:`application/json`
打开浏览器,从开发者工具找到`Network`一项,刷新一个页面,就可以从中看到请求报文和响应报文的信息。
### Cookie
Cookie 是浏览器保存数据的一种方式,数据以键值对形式存储,可以供客户端或服务端调用。在 Chrome 控制台点击`Application`标签可以查看本地所有的 Cookie,可以看到,Cookie 是按照域名分类的,也就是说,一个域名下的网页无法访问其它域名下的 Cookie。并且,当用户提交表单时,浏览器会自动将本域的 Cookie 提交到服务端。
<file_sep>// 为标签追加字体图标
$('.tag-list-item').prepend('<span class="iconfont icon-biaoqian"></span>')
// 回到顶部的猫
$('.back-to-top').click(function () {
$('html, body').animate({
scrollTop: 0
}, 500)
})
// 监听 回到顶部的猫 是否显示
$(document).scroll(function () {
if ($(document).scrollTop() > 50) {
$('.back-to-top').addClass('show')
} else {
$('.back-to-top').removeClass('show')
}
})
// 在表格外包裹一层,用来生成滚动条
$('.post-content > table').wrap('<div class="table-container"></div>')
// Tocbot 实例化,用来快速生成文档目录
tocbot.init({
tocSelector: '.toc', // 渲染目录的容器
contentSelector: '.post-content', // 标题所在的容器
headingSelector: 'h2, h3', // 要渲染的目录层级,
headingsOffset: -50, // 微调目录高亮的显示位置(偏移量),防止点击的链接和高亮的链接不一致
})
// 初始化代码高亮 highlight.js
hljs.initHighlightingOnLoad()<file_sep>---
title: 大前端支线笔记之02 💠 CSS BFC
date: 2020-01-22 23:20:26
abbrlink: 9obo9xpo
tags: CSS
categories:
- 大前端
- 支线
excerpt: BFC(Block Formatting Context,块级格式化上下文)指的是一个元素的特性,拥有该特性的元素可以生成一个独立的空间,无论其内部的元素如何改变,都不可能脱离出这个空间。
---
# 大前端支线笔记之02 💠CSS BFC
BFC(Block Formatting Context,块级格式化上下文)指的是一个元素的特性,拥有该特性的元素可以生成一个独立的空间,无论其内部的元素如何改变,都不可能脱离出这个空间。例如,整个文档的根元素`<html>`就具有 BFC 特性,因此所有的元素都不可能从中脱离出去。
要使一个元素拥有 BFC 特性,只需满足下列条件之一:
* 根元素`<html>`
* 浮动元素,即`float`的值不是`none`
* 定位元素,即`position`的值不是`static`或`relative`
* 行内块元素,即`display`的值是`inline-block`
* 弹性元素,即`display`的值是`flex`或`inline-flex`
* `overflow`值不是`visible`的元素
当一个元素满足以上条件之一,那么就会触发该元素的 BFC。
在下面的代码中,容器`.container`仅包含了一个浮动元素和一些文本内容,可以看到容器的高度仅受文本影响,浮动的元素完全超出了容器(左图)。
```html
<div class="container">
<div class="float-box"></div>
这是容器的内容啦
</div>
```
```css
.container { width: 300px; }
.float-box {
float: left;
width: 100px;
height: 100px;
}
```
但是如果为容器添加了`overflow: hidden`,即触发了容器的 BFC 之后,那么此时它的子元素是无论如何也不会跑到父元素之外的,因此容器的高度被撑大了。
```css
.container {
overflow: hidden;
width: 300px;
}
```
<file_sep>---
title: 大前端笔记之14 🔌 JavaScript 标准库
date: 2020-02-01 11:24:15
abbrlink: i82icjg7
tags: JavaScript
categories: 大前端
excerpt: 本节均是一些 JavaScript 提供的常用 API,需要时查询即可。
---
# 大前端笔记之14 🔌 JavaScript 标准库
本节均是一些 JavaScript 提供的常用 API,需要时查询即可。
## Array
### indexOf() / lastIndexOf()
获取元素在数组中首次出现的索引位置。
| 参数 | 描述 |
| --- | --- |
| 对象 | 要查找的元素 |
| 数值 | 可选,表示从该索引位置(包括)向后查找 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 元素在数组中首次出现的索引位置,如果不存在返回`-1` |
```js
var array = [10, 20, 30, 20]
array.indexOf(20) // => 1
array.indexOf(233) // 元素不存在,返回 -1
// 第二个参数表示从该索引位置(包括)向后查找
array.indexOf(20, 2) // => 3,因为是从 30 开始查找的
```
与之类似的`lastIndexOf()`则是从后开始查找,注意它返回的索引是不会改变的,依然是从左开始计数。
```js
var array = [10, 20, 30, 20]
array.lastIndexOf(20) // => 3
```
### unshift() / push()
在数组开头 / 末尾添加若干元素,会修改原数组。
| 参数 | 描述 |
| --- | --- |
| 若干对象 | 要添加的元素 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 数组修改后的长度 |
```js
var array = [10]
array.unshift(6, 8) // => 3
// array => [6, 8, 10]
array.push(20, 30) // => 3
// array => [10, 20, 30]
```
### shift() / pop()
删除数组第一个 / 最后一个元素,会修改原数组。
| 返回值 | 描述 |
| --- | --- |
| 对象 | 删除的元素 |
```js
var array = [10, 20, 30]
array.shift() // => 10
// array => [20, 30]
array.pop() // => 30
// array => [10, 20]
```
### splice()
删除从指定索引开始的若干元素(并添加新元素),会修改原数组。
| 参数 | 描述 |
| --- | --- |
| 数值 | 起始索引(包括) |
| 数值 | 可选,要删除的元素数量,如果省略则删除起始索引之后的所有元素 |
| 若干对象 | 可选,删除元素后在起始索引处添加的新元素,如果省略则仅删除元素 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 删除的元素构成的数组,如果没有则返回空数组 |
```js
var array = [10, 20, 30, 40, 50]
array.splice(1, 2) // => [20, 30]
// array => [10, 40, 50]
array.splice(1, 2, 233) // => [20, 30]
// array => [10, 233, 40, 50]
```
### slice()
截取数组的一部分并返回,不会修改原数组。
| 参数 | 描述 |
| --- | --- |
| 数值 | 起始索引(包括),如果为负数则表示从后开始计数,最后一个元素的索引为`-1`,依次向前类推;如果省略则默认为`0` |
| 数值 | 结束索引(不包括),如果为负数则表示从后开始计数,最后一个元素的索引为`-1`,依次向前类推;如果省略则截取到数组最后 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 截取的数组 |
```js
var array = [10, 20, 30, 40]
array.slice(1) // => [20, 30, 40]
array.slice(1, 3) // => [20, 30]
array.slice(-2) // => [30, 40]
array.slice(-3, -1) // => [20, 30]
```
### reverse()
将数组中的元素倒序排列,会修改原数组。
| 返回值 | 描述 |
| --- | --- |
| 数组 | 倒序的数组 |
```js
var array = [10, 20, 30, 40]
array.reverse() // => [40, 30, 20, 10]
```
### sort()
将数组按照一定规则排序,会修改原数组。注意,其默认的排序规则是 Unicode 码点,因此对于数值来说,它并不会比较数值的大小。
| 参数 | 描述 |
| --- | --- |
| 函数 | 表示自定义排序规则。其两个参数表示数组的任意两个元素,通过比较后的返回值,可以决定排序规则。该函数写法比较固定,记住即可不必深究原理 |
```js
;['d', 'c', 'b', 'a'].sort() // => ['a', 'b', 'c', 'd']
;[13, 1, 77, 7].sort() // => [1, 13, 7, 77]
// 表示升序排列,return b - a 则表示降序
function rule(a, b) { return a - b }
;[10, 1000, 100].sort(rule) // => array: [10, 100, 1000]
```
### concat()
将一个数组(参数)追加到另一个数组(调用者)的后面,不会修改原数组。
| 参数 | 描述 |
| --- | --- |
| 数组 | 要追加的数组 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 合并后的新数组 |
```js
;[10, 20].concat([30, 40]) // => [10, 20, 30, 40]
```
### forEach()
遍历数组。
| 参数 | 描述 |
| --- | --- |
| 函数 | 表示处理函数,它有三个参数,第一个表示当前元素,第二个表示当前索引,第三个表示遍历的数组本身 |
```js
var arr = [10, 20, 30]
arr.forEach(function (value, index, array) {
value // 当前元素
index // 当前元素的索引
array // 数组对象
})
```
### find() / findIndex()
根据条件查找第一个匹配的元素 / 元素索引并返回:
| 参数 | 描述 |
| --- | --- |
| 函数 | 表示处理函数,它有三个参数,第一个表示当前元素,第二个表示当前索引,第三个表示遍历的数组本身,返回值为判断条件 |
| 返回值 | 描述 |
| --- | --- |
| 对象 / 数值 | 匹配的元素 / 元素索引 |
```js
var data = [
{ id: 1, name: 'Wendy'},
{ id: 2, name: 'Lucy'}
]
data.find(function (el, index, array) {
return el.id == 1
}) // => { id: 1, name: 'Wendy'}
data.findIndex(function (el, index, array) {
return el.id == 1
}) // => 0
```
### filter()
根据条件过滤数组中的所有元素,将满足条件的元素放入一个新数组,不会影响原数组。
| 参数 | 描述 |
| --- | --- |
| 函数 | 表示处理函数,它有三个参数,第一个表示当前元素,第二个表示当前索引,第三个表示遍历的数组本身,返回值为判断条件 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 满足条件元素组成的新数组 |
```js
var arr = [10, 5, 3, 0, 233, 666, 0]
arr.filter(function (value, index, array) {
return value != 0
}) // => [10, 5, 3, 233, 666]
```
### some()
查找数组中是否有满足条件的元素,只要找到了符合条件的元素就会**立刻终止遍历**,不会继续判断后续的元素。
| 参数 | 描述 |
| --- | --- |
| 函数 | 表示处理函数,它有三个参数,第一个表示当前元素,第二个表示当前索引,第三个表示遍历的数组本身,返回值为判断条件 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果数组中包含满足条件的元素返回`true`,否则返回`false` |
```js
var arr = [10, 50, 100]
arr.some(function (value, index, array) {
return value > 50
}) // => true
```
### includes()
查找数组中是否包含指定的元素。
| 参数 | 描述 |
| --- | --- |
| 对象 | 要查找的元素 |
| 数值 | 可选,表示从该索引位置(包括)向后查找 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果数组中包含指定元素返回`true`,否则返回`false` |
```js
;[10, 50, 100].includes(10) // => true
```
### join()
以特定字符作为分隔符,将数组元素转换成一个字符串。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 表示分隔符,如果省略则默认为逗号,与直接使用数组调用`toString()`方法的结果相同 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 拼接后的字符串 |
```js
var array = [10, 20, 30]
array.join() // => '10,20,30'
array.join('|') // => '10|20|30'
```
### reduce()
对数组中的某些数据进行累计,返回一个累计后的值。
| 参数 | 描述 |
| --- | --- |
| 函数 | 迭代函数,包含最多以下四个参数: |
| 子参数① | 累计的结果值 |
| 子参数② | 当前处理的数组元素 |
| 子参数③ | 可选,起始索引,如果提供了初始值,那么默认为`0`,否则为`1` |
| 子参数④ | 可选,调用该方法的数组 |
| 对象 | 可选,迭代的初始值,默认为数组的第一个元素 |
| 返回值 | 描述 |
| --- | --- |
| 对象 | 累计之后的结果 |
该方法可以依次取出数组元素,累计到结果值(子参数①)上。迭代函数的返回值表示本次的计算结果,迭代函数每次执行时,结果值均为上一次迭代的结果,通过与当前数组元素进行运算(不一定是加法,任何运算都可以),从而实现累计。
在下面的代码中,迭代函数首先取出数组的第一个元素`10`作为`sum`,取出第二个元素`20`作为`current`,然后将累加后的结果`30`作为返回值,保存到`sum`;第二次迭代时,取出上次的`sum`即`30`,取出第三个元素`40`作为`current`,将累加后的结果`70`作为返回值`sum`;此时迭代结束,将最后的`sum`即`70`作为方法的返回值。
```js
[10, 20, 40].reduce(function (sum, current) {
return sum + current
}) // => 70
```
如果给出迭代函数之外的第二个参数,表示迭代的初始值。此时迭代函数会将该初始值作为`sum`,而不是数组的第一个元素;第一次迭代时也会变成取出数组的第一个元素。
```js
[10, 20, 40].reduce(function (sum, current) {
return sum + current
}, 0) // => 70
```
两种方式的计算结果是一样的,区别在于,如果给出了初始值,那么迭代函数会多执行一次(因为第一次迭代只能取出第一个元素,否则可以一次取出两个)。不过还是建议总是给出初始值,因为如果传入了空数组,第一种方式会因为找不到初始值而报错。
```js
[].reduce(function (sum, current) {
return sum + current
}, 0) // => 0
[].reduce(function (sum, current) {
return sum + current
}) // => TypeError: Reduce of empty array with no initial value
```
下面的代码可以累加对象数组中的值:
```js
let books = [
{ id: 1000, name: '哈利波特', price: 300 },
{ id: 1001, name: '北欧众神', price: 50 },
{ id: 1002, name: 'JOJO的奇妙冒险', price: 450 },
]
books.reduce(function(totalPrice, book){
return totalPrice + book.price
}, 0) // => 800
```
### map()
将数组的每个元素调用处理函数之后,返回一个新数组。该方法不会修改原数组。
| 参数 | 描述 |
| --- | --- |
| 函数 | 迭代函数,包含最多以下三个参数: |
| 子参数① | 当前数组元素 |
| 子参数② | 可选,当前索引 |
| 子参数③ | 可选,调用该方法的数组 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 处理后的新数组 |
```js
[10, 20, 30].map(function (current, index, array) {
return current * 2
}) // => [20, 40, 60]
```
### from()
静态方法,将一个伪数组或者类似数组的对象(只要包含`length`属性)转换为真正的数组。
| 参数 | 描述 |
| --- | --- |
| 对象 | 要转换的对象 |
| 函数 | 可选,表示处理函数,可以对当前遍历的元素进行操作,它的参数为当前元素,返回值为要执行的操作 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 转换后的数组 |
```js
var obj = { 0: 'hello', 1: 'world', length: 2 }
Array.from(obj) // => ['hello', 'world']
Array.from(obj, function (value) {
return value += 'a'
}) // => ['helloa', 'worlda']
```
### isArray()
静态方法,判断一个值是否为数组。
| 参数 | 描述 |
| --- | --- |
| 对象 | 要判断的值 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果是数组则返回`true`,否则返回`false` |
```js
Array.isArray([1, 2, 3]) // => true
```
## String
### indexOf() / lastIndexOf()
获取一个字符串(参数)中的**第一个字符**在另一个字符串(调用者)中首次出现的位置。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要查找的字符串 |
| 数值 | 可选,表示从该索引位置(包括)向后查找 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 参数在字符串中首次出现的索引位置,如果不存在返回`-1` |
```js
var str = 'helloworld'
str.indexOf('o') // => 4
str.indexOf('world') // => 5
str.indexOf('Daisy') // => -1
str.indexOf('') // => 0
str.indexOf('o', 5) // => 6
```
与之类似的`lastIndexOf()`则是从后开始查找,注意它返回的索引是不会改变的,依然是从左开始计数。
```js
str.lastIndexOf('o') // => 6
str.indexOf('world') // => 5
```
### charAt() / charCodeAt()
获取字符串中某个位置的字符 / 字符编码。
| 参数 | 描述 |
| --- | --- |
| 数值 | 要查找的索引位置 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 / 数值 | 该索引位置的字符 / 字符编码 |
```js
var str = 'helloworld'
str.charAt(5) // => 'w'
str.charCodeAt(5) // => 119
```
> `charAt()`与字符串直接加方括号`[]`的效果相同,如`str.charAt(5)`等价于`str[5]`。
### substring() / slice()
截取字符串,不会修改原字符串。
| 参数 | 描述 |
| --- | --- |
| 数值 | 起始索引(包括) |
| 数值 | 结束索引(不包括),如果省略则截取到最后 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 截取后的字符串 |
```js
var str = 'helloworld'
str.substring(5, 7) // => wo
str.substring(5) // => world
str.substring(-3, -1) // => rl
```
> 与之类似的`substr()`已经被废弃,不要使用。
### replace()
替换字符串的一部分,不会修改原字符串。注意,该方法只能替换第一个匹配(除非使用全局匹配的正则)的内容。
| 参数 | 描述 |
| --- | --- |
| 字符串或正则表达式 | 被替换的旧内容 |
| 字符串 | 新内容 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 替换后的字符串 |
```js
var str = 'hello123world123'
str.replace('o', 'x') // => hellx123world123,将第一个 o 替换成 x
str.replace(/123/, 'x') // => helloxworld123,将第一个 123 替换成 x
str.replace(/123/g, 'x') // => helloxworldx,开启了全局匹配
```
### repeat()
复制字符串,不会修改原字符串。
| 参数 | 描述 |
| --- | --- |
| 数值 | 复制的次数 |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 复制后的字符串 |
```js
var str = 'hello'
str.repeat(2) // => hellohello
```
### trim()
去除字符串两端的空格,不会修改原字符串。
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 去除两端空格后的字符串 |
```js
' hello world '.trim() // => 'hello world'
```
### startsWith() / endsWith()
判断字符串是否以指定内容开头 / 结尾。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要判断的内容 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果以指定内容开头 / 结尾返回`true`,否则返回`false` |
```js
var str = 'hello'
str.startsWith('he') // => true
str.endsWith('lo') // => true
```
### includes()
查找字符串中是否包含指定内容。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要查找的内容 |
| 数值 | 可选,表示从该索引位置(包括)向后查找 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果字符串中包含指定子串返回`true`,否则返回`false` |
```js
'helloworld'.includes('hello') // => true
'helloworld'.includes('') // => true,注意,空字符串也视为包含
```
### padStart() / padEnd()
当字符串长度不足时,以指定内容填充到字符串开头 / 末尾。
| 参数 | 描述 |
| --- | --- |
| 数值 | 目标长度 |
| 字符串 | 要填充的内容,如果填充后超过了目标长度,那么截断右侧多出的部分,默认值为`''` |
| 返回值 | 描述 |
| --- | --- |
| 字符串 | 填充后的新字符串 |
```js
'hello'.padStart(10) // => ' hello'
'hello'.padStart(10, 'a') // => 'aaaaahello'
'hello'.padStart(10, '1234567') // => '12345hello'
```
### split()
以某个字符作为分隔符,将字符串拆分为数组。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 表示分隔符,如果省略则相当于不拆分,如果为空字符串则拆分成单个字符 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 拆分后的数组 |
```js
var str = 'a|b|c'
str.split() // => ["a|b|c"]
str.split('') // => ["a", "|", "b", "|", "c"]
str.split('|') // => ["a", "b", "c"]
```
## Number
### toFixed()
设置小数的保留位数。
| 参数 | 描述 |
| --- | --- |
| 数值 | 要保留的位数,如果不传参数则**四舍五入**去掉小数位 |
| 返回值 | 描述 |
| --- | --- |
| **字符串** | 转换后的数值,注意它的类型 |
```js
var num = 9.876
num.toFixed() // => 10
num.toFixed(2) // => 9.88
num.toFixed(4) // => 9.8760
```
### parseInt() / parseFloat()
静态方法。将字符串转为整数(舍去小数位)/ 浮点数。它们会**依次**将每个字符转换为数字,直到遇到不能转为数字的字符,就不再继续,返回已经转换的部分。如果第一个字符就不能转化为数字(包括空字符串),返回`NaN`。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要转换的值,如果不是字符串,那么会先调用`toString()`将该值转换成字符串 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 转换后的数值 |
```js
Number.parseInt('15px') // =>15
Number.parseInt('abc') // => NaN
```
> ES5 这两个方法位于`window`对象中,ES6 将其移动到了`Number`对象中。
### isInteger()
判断一个值是否为整数。
| 参数 | 描述 |
| --- | --- |
| 对象 | 要判断的值 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果是整数则返回`true`,否则返回`false` |
```js
Number.isInteger('a') // => false
```
## Math
### abs()
静态方法。获取绝对值。
| 参数 | 描述 |
| --- | --- |
| 数值 | 要转换的值 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 转换后的绝对值 |
```js
Math.abs(-1) // => 1
```
### max() / min()
静态方法。获取参数列表中的最大值 / 最小值。
| 参数 | 描述 |
| --- | --- |
| 若干数值 | 要比较的值 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 参数列表的最大值 / 最小值 |
```js
Math.max(1, 5, 3) // => 5
Math.min(-1, -10) // => -1
```
### floor() / ceil() / round()
将数值向下 / 向上 / 四舍五入取整。
| 参数 | 描述 |
| --- | --- |
| 数值 | 要取整的值 |
| 返回值 | 描述 |
| --- | --- |
| 数值 | 取整后的值 |
```js
Math.floor(3.2) // 3
Math.floor(-3.2) // -4
Math.ceil(3.2) // 4
Math.ceil(-3.2) // -3
Math.round(0.4) // 0
Math.round(0.5) // 1
Math.round(-1.5) // -1,注意负数时,也是向大数取整,因此 -2 -1.5 -1,取更大的 -1
Math.round(-1.6) // -2
```
### random()
获取一个$[0, 1)$之间的伪随机数。
| 返回值 | 描述 |
| --- | --- |
| 数值 | 随机数 |
下面的函数可以返回一个$[min, max)$之间的**随机数**:
```js
function getRandom(min, max) {
return Math.random() * (max - min) + min
}
getRandom(1, 6)
```
下面的函数可以返回一个$[min, max]$之间的**随机整数**:
```js
function getRandomInt(min, max) {
return Math.floor(Math.random() * (max - min + 1)) + min
}
getRandomInt(1, 6)
```
## Date
使用构造函数`Date()`可以实例化一个日期对象:
```js
new Date() // 创建一个当前时间的日期对象
new Date('1949-10-1 12:00:23') // 创建一个对应时间的日期对象
```
日期对象覆盖了`toString()`方法,因此直接`console.log()`会打印一长串的时间信息。
```js
console.log(new Date()) // => Sun Dec 22 2019 09:54:21 GMT+0800 (中国标准时间)
```
但是这样的格式基本没法直接使用,因此还需要使用日期对象中的一系列方法来获取想要的数据:
```js
var date = new Date('1949-10-1 12:00:23')
date.getFullYear() // => 1949,获取年份
date.getMonth() // => 9,注意月份是从 0 开始的,因此要得到正确的月份,请手动 +1
date.getDate() // => 1,获取天
date.getDay() // => 6,获取星期,注意周日为 0
date.getHours() // => 12,获取小时数
date.getMinutes() // => 0,获取分钟数
date.getSeconds() // => 23,获取秒数
date.toLocaleString() // => 1949/10/1 下午12:00:23,获取本地时间格式
```
### 时间戳
JavaScript 的日期基于 Unix 时间戳(从世界协调时 1970 年 1 月 1 日开始经过的毫秒数),通过日期对象的下列方法可以获取它的时间戳:
```js
var date = new Date('1949-10-1 12:00:23')
date.getTime() // => -639086377000
date.valueOf() // => -639086377000
```
或者在实例化日期对象时使用一元运算符`+`将其转为`Number`型,这样也可以直接使该对象自动调用`valueOf()`方法,从而获取时间戳:
```js
var timeStamp = +new Date('1949-10-1 12:00:23') // => -639086377000
```
如果只是获取当前时间的时间戳,使用`Date`对象的静态方法`now()`即可:
```js
var timeStamp = Date.now();
```
## RegExp
这里仅讨论正则表达式`RegExp`对象的相关方法,在这里查看[正则表达式规则](/posts/9d8oxogs.html)。
使用两条斜杠`//`包裹创建一个正则表达式对象。
```js
var regex = /hello/
```
### test()
验证字符串是否符合特定的正则表达式规则。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要验证的字符串 |
| 返回值 | 描述 |
| --- | --- |
| 布尔值 | 如果符合规则返回`true`,否则返回`false` |
```js
/hello/.test('helloworld') // => true
```
### exec()
获取匹配结果。
| 参数 | 描述 |
| --- | --- |
| 字符串 | 要验证的字符串 |
| 返回值 | 描述 |
| --- | --- |
| 数组 / `null` | 如果匹配则以数组索引`0`返回匹配结果,否则返回`null` |
如果正则表达式包含分组,则返回的数组会包括多个元素,其中索引`0`依然是整个匹配结果,而索引`1`之后则是每个单独分组匹配的结果。
```js
var regex = /(\w+)@(\w+).(\w+)/
var arr = regex.exec('<EMAIL>')
// arr[0] => <EMAIL>
// arr[1] => qingyibest
// arr[2] => 163
// arr[3] => com
```
此外,该数组还拥有两个属性,其中`index`表示成功匹配的第一个索引位置,`input`表示原字符串。
```js
var arr = /e/.exec('hello')
// index => 1
// input => hello
```
注意,如果字符串中有多个子字符串匹配,那么首次调用该方法只会返回第一个,要获取全部的结果,则需要使用循环遍历,且必须开启全局匹配。
```js
var str = '中国移动:10086;中国电信:10000;中国联通:10010'
var regex = /[0-9]{5}/g
var arr1 = regex.exec(str) // arr1[0] => 10086
var arr2 = regex.exec(str) // arr2[0] => 10000
var arr3 = regex.exec(str) // arr3[0] => 10010
```
### 修饰符
修饰符类似于正则对象的配置参数,它们定义在正则对象的`/`之后,且可以同时定义多个。
修饰符`g`表示全局匹配。默认情况下,当第一次找到符合规则的部分后,就不再继续向下匹配。开启全局匹配后表示查找整个字符串中全部符合规则的部分,主要用于搜索和替换。
```js
'helloworld'.replace(/o/, 'x') // => hellxworld
'helloworld'.replace(/o/g, 'x') // => hellxwxrld
```
修饰符`i`表示忽略大小写。
```js
;/abc/.test('ABC') // => false
;/abc/i.test('ABC') // => true
```
## JSON
JSON(JavaScript Object Notation,JavaScript 对象表示法)是现在非常流行的一种数据格式,它取代了之前常用的 XML,用来传输数据。它的主要格式规则如下:
- 有大括号`{}`和方括号`[]`两种语法,大括号`{}`用来描述不同类型的无序键值对集合,方括号`[]`用来描述相同类型的有序数据集合
- 键名必须用**双引号**包裹
- 字符串必须用**双引号**包裹,不能用单引号
- 引用类型的值只能是数组或对象,不能是函数、日期等
- 基本类型的值只能是字符串、数值、布尔值或`null`
JavaScript 提供了内置对象`JSON`来操作 JSON 数据,它除了调用以下两个静态方法以外本身并没有其它作用。
```js
// 下面是一些合法的 JSON
["one", "two", "three"]
{"one": 1, "two": 2, "three": 3}
{"names": ["张三", "李四"]}
[{ "name": "张三"}, {"name": "李四"}]
```
### stringify()
将 JavaScript 对象转换为 JSON 字符串。
```js
JSON.stringify({name: "Lucy", weight: 15}) // => {"name":"Lucy","weight":15}
JSON.stringify(['a', 'b', 'c']) // => ["a","b","c"]
```
### parse()
将 JSON 字符串转换为 JavaScript 对象。
```js
JSON.parse('{"name": "Lucy", "age": 18}') // => 对象 {name: 'Lucy', age: 18}
JSON.parse('["a", "b", "c"]') // => 数组 ['a', 'b', 'c']
```
## Object
### toString()
获取表示该对象类型的字符串。不过,由于 JavaScript 内置的多个对象(如数组、字符串、函数、日期等)都使用了自己的`toString()`将其覆盖,因此使用这些对象调用`toString()`时,会输出不一样的内容。
| 数据类型或对象 | 返回值 |
| --- | --- |
| `Undefined` | `'undefined'` |
| `Null` | `'null'` |
| `Boolean` | `'true'`或者`'false'` |
| `Number` | 数值对应的字符串形式 |
| `Object` | `[object Object]` |
| 数组 | 以逗号分隔的元素,如`[1, 2, 3]`返回`1,2,3` |
| 函数 | 函数源代码 |
| 日期 | 如`Mon Sep 16 2019 17:01:43 GMT+0800 (中国标准时间)` |
由于任何对象调用`toString()`方法默认会返回表示该对象类型的字符串,因此可以利用这一特性判断对象的数据类型。不过因为某些对象的`toString()`方法已经被覆盖,因此要通过`Object`来调用该方法,然后将要判断的对象作为`call()`的参数传进去。
```js
var arr = [1, 2, 3]
Object.prototype.toString.call(arr) // => [object Array]
```
常见对象的返回值如下,其它对象均返回`[object Object]`:
| 对象 | 返回值 | 对象 | 返回值 | 对象 | 返回值 |
| --- | --- | --- | --- | --- | --- |
| `Number` | `[object Number]` | `String` | `[object String]` | `Boolean` | `[object Boolean]` |
| `Undefined` | `[object Undefined]` | `Null` | `[object Null]` | `Array` | `[object Array]`|
| `Arguments` | `[object Arguments]` | `Function` | `[object Function]` | `Date` | `[object Date]` |
### keys()
获取一个对象的属性名,以数组形式返回。
| 参数 | 描述 |
| --- | --- |
| 对象 | 要获取属性的对象 |
| 返回值 | 描述 |
| --- | --- |
| 数组 | 对象的属性名组成的数组 |
```js
var phone = { id: 1, pname: '小米', price: 3999 }
Object.keys(obj) // => [id, pname, price]
```
## Console
### log()
在控制台输出信息。
```js
console.log('hello')
```
注意,Chrome 浏览器在输出数组或对象类型时默认是折叠的,详细数据需要展开才能显示。但是,当展开数据时,浏览器会重新获取一次数据。也就是说,展开时看到的数据**不一定**是当时代码执行到`console.log()`处的结果。
### dir()
在控制台输出对象的所有属性和值。这种方式可以更加直观的看到对象的各种信息。
```js
console.log({f1: 'foo', f2: 'bar'}) // => {f1: "foo", f2: "bar"}
``` | 195536556dd045067102d160be69cb6ece47a8fd | [
"Markdown",
"JavaScript"
] | 41 | Markdown | ChickenPopcorny/chicken-blog | 36e7348e30863d079a3af383bd7a1f14c2be4597 | 011c7745a706af0a0ce04da9cfcfc47e5fd52dca |
refs/heads/master | <file_sep>/* This program uses code from "Algorithms in C, Third Edition,"
* by <NAME>, Addison-Wesley, 1998.
*
*/
// Modified and added functions by <NAME>
#include <stdio.h>
#include <stdlib.h>
#include "list_hw.h"
struct node_struct
{
Item item;
link next;
};
struct list_struct
{
link first;
int length;
};
/* Builds and returns a list with integers read from standard input.
* Note that it assumes the Item is int.
* Notice that all the list access is done through it's interface (functions).
*/
list buildListOfInts()
{
list the_list = newList();
link current_link = NULL;
while (1) {
Item number;
printf("please enter an integer: ");
int items_read = scanf("%d", &number);
if (items_read != 1) {
break;
}
/* allocate memory for the next link */
link next_item = newLink(number, NULL);
insertLink(the_list, current_link, next_item);
current_link = next_item;
}
return the_list;
}
list arrayToList(int arr[], int sz)
{
int i;
link p, new_p;
list A = newList();
p = newLink(arr[0], NULL);
insertLink(A, NULL, p);
for (i = 1; i < sz; i++) {
new_p = newLink(arr[i], NULL);
insertLink(A, p, new_p);
p = new_p;
}
return A;
}
// ------------- link functions
/* Creates a new link, that contains the value specified in the argument,
* and that points to next_in.
*/
link newLink(Item value, link next_in)
{
link result = (link) malloc(sizeof (struct node_struct));
result->item = value;
result->next = next_in;
return result;
}
void destroyLink(link the_link)
{
if (the_link != NULL) {
free(the_link);
}
}
Item getLinkItem(link the_link)
{
if (the_link != NULL) {
return the_link->item;
}
else {
printf("\n getLinkItem: Empty link. Returned -1.");
return -1; // NOTE: This line can be improved.
}
}
link getLinkNext(link the_link)
{
if (the_link != NULL) {
return the_link->next;
}
else {
return NULL;
}
}
void setLinkItem(link the_link, Item value)
{
if (the_link != NULL) {
the_link->item = value;
}
}
void setLinkNext(link the_link, link next)
{
if (the_link != NULL) {
the_link->next = next;
}
}
/* -------- LIST functions */
int listIsNULL(list the_list)
{
if (the_list == NULL) {
printf("\nThe list pointer is NULL)\n");
return 1;
}
else {
return 0;
}
}
/* Creates and returns an empty list. */
list newList()
{
list result = (list) malloc(sizeof (*result));
result->first = NULL;
result->length = 0;
return result;
}
/* Deallocates memory for all nodes in the list and the list object itself. */
void destroyList(list the_list)
{
if (listIsNULL(the_list) == 1) {
return;
}
link current = the_list->first;
while (1) {
if (current == NULL) {
break;
}
link next = current->next;
destroyLink(current);
current = next;
}
free(the_list);
}
link getFirst(list the_list)
{
if (listIsNULL(the_list) == 1) {
return NULL;
}
return the_list->first;
}
int getLength(list the_list)
{
if (listIsNULL(the_list) == 1) {
return -1;
}
return the_list->length;
}
/* Sets the data to empty (first is null, length is 0,...)
* It does NOT free any memory.
*/
void setEmpty(list the_list)
{
the_list->first = NULL;
the_list->length = 0;
}
/* Inserts new_link to the specified list, at the position right after
* the link called "previous". */
void insertLink(list my_list, link previous, link new_link)
{
if (listIsNULL(my_list) == 1) {
return;
}
/* We need a special case when we want to insert to the beginning.
* In that case, the previous link is NULL. */
if (previous == NULL) {
new_link->next = my_list->first;
my_list->first = new_link;
}
else {
new_link->next = previous->next;
previous->next = new_link;
}
my_list->length += 1;
}
/* Inserts new_link to the beginning of the list. */
void insertAtBeginning(list my_list, link new_link)
{
setLinkNext(new_link, getFirst(my_list)); // replaces: new_link->next = my_list->first;
my_list->first = new_link;
my_list->length += 1;
}
/* Removes from the list and returns the link coming AFTER link x.
* This function DOES NOT remove link x itself.
*/
link removeNext(list my_list, link x)
{
if (listIsNULL(my_list) == 1) {
return NULL;
}
link temp;
if (x == NULL) // try to delete the first node
{
temp = my_list->first;
if (my_list->first != NULL) // There is at least one node in the list.
{
my_list->first = my_list->first->next;
my_list->length -= 1;
}
}
else {
temp = x->next;
x->next = x->next->next; // IS THIS CODE SAFE? JUSTIFY YOUR ANSWER.
my_list->length -= 1;
}
return temp;
}
/* Removes from the list and returns the link at the beginning of the list.
It does not free the memory for that node. */
link removeFirst(list my_list)
{
if (listIsNULL(my_list) == 1) {
return NULL;
}
link first_node = my_list->first;
if (first_node != NULL) {
my_list->first = first_node->next;
my_list->length -= 1;
}
return first_node;
}
void printList(list my_list)
{
if (listIsNULL(my_list) == 1) {
return;
}
int i = 0;
int n = my_list->length;
link curr;
printf("\n List length = %d\n List items:\n", my_list->length);
for (i = 0, curr = my_list->first; (curr != NULL) && (i < n); curr = curr->next) {
printf("item %d: %d\n", i, curr->item);
i++;
}
printf("\n");
}
void printListHoriz(list my_list)
{
if (listIsNULL(my_list) == 1) {
return;
}
int i = 0;
int n = my_list->length;
link curr;
printf("\n List length = %d\n List items:\n", my_list->length);
for (i = 0, curr = my_list->first; (curr != NULL) && (i < n); curr = curr->next) {
printf("%5d ", curr->item);
i++;
}
printf("\n");
}
void printListHorizPointer(list my_list)
{
if (listIsNULL(my_list) == 1) {
return;
}
int i = 0;
int n = my_list->length;
link curr;
printf("\n List length = %d\nList items: ", my_list->length);
for (i = 0, curr = my_list->first; (curr != NULL) && (i < n); curr = curr->next) {
printf("%-11d ", curr->item);
i++;
}
printf("\n");
printf("List pointers: ");
for (i = 0, curr = my_list->first; (curr != NULL) && (i < n); curr = curr->next) {
printf("%p ", curr);
i++;
}
printf("\n");
}
int computeLength(list my_list)
{
if (listIsNULL(my_list) == 1) {
return -1;
}
int counter = 0;
link i;
for (i = my_list->first; i != NULL; i = i->next) {
counter++;
}
return counter;
}
void setFirst(list my_list, link new_link) // Method to set the first address to the list.
{
my_list->first = new_link;
}
void myDestroyList(list the_list) //Method used to delete the list when memory errors occur of some inputs
{
if (listIsNULL(the_list) == 1) {
return;
}
int listLength = getLength(the_list);
int i;
link current = the_list->first;
link nextLink;
for (i = 0; i < listLength; i++) {
if (current == NULL) {
i = listLength;
}
nextLink = getLinkNext(current);
destroyLink(current);
current = nextLink;
}
free(the_list);
the_list = NULL;
}
/*----------------------------------------------------------------
New functions.
Finish their implementation below.
If you need to use helper functions you can define them and use them in this file.
***You should not change the behavior of any of the functions defined above.
Instead, write a new function with the behavior that you want.***
*/
// Implementations provided by <NAME>
void deleteOccurrences(list A, Item V)
{
if (A == NULL) { //Check if the list is NULL
printf("List A is NULL\n");
return;
}
int ListLength = getLength(A);
if (ListLength == 0) { //Check if the list has zero length
printf("List A is empty \n");
}
else {
int i;
link current = getFirst(A);
link previous = NULL;
int ListLength = getLength(A); //Get list Length
for (i = 0; i < ListLength; i++) {
Item TestElement = getLinkItem(current); //Get the First Item from the List A
link nextLinkNew = getLinkNext(current);
if (TestElement == V) {
link temp; // If the Item is equal to item to be searched
if (i == 0) {
temp = removeFirst(A); //Remove the first link
destroyLink(temp);
}
else {
temp = removeNext(A, previous); //else removes the next link of previous
destroyLink(temp);
}
current = nextLinkNew;
}
else {
previous = current;
current = nextLinkNew;
}
}
}
return;
}
list sublist(list A, list pos_list)
{
if (A == NULL) {
printf("list A is NULL\n"); //Check if the list A is NULL
return NULL;
}
int listLength = getLength(A);
if (listLength == 0) {
printf("List A is empty\n"); //Check if the list A has zero length
return NULL;
}
if (pos_list == NULL) {
printf("List A is NULL \n"); //Check if the pos_list is NULL
return NULL;
}
if (pos_list->length == 0) { //Check if the list pos_list has zero length
printf("List A is empty \n");
return NULL;
}
else {
int i;
list subList = newList(); //Create new sublist to store the values returned from the function
link sublistLink = NULL;
link newLinkToAdd;
link startLink = getFirst(A); //Get first link of list A
link currentLink, nextLink;
link previousLinkAddedToSublist;
link currentLinkForPosList = getFirst(pos_list); //Get first link of pos_list
int posListLength = getLength(pos_list); //Get length of pos_list
int count = 0;
Item itemAtRequiredIndex;
Item itemToGet = getLinkItem(currentLinkForPosList); //Get first item of Pos_list
currentLink = startLink;
for (i = 0; i < posListLength; i++) {
if (itemToGet > listLength) { //Handled for Index out of bound test case
destroyList(subList);
return NULL;
}
while (count != itemToGet) {
nextLink = getLinkNext(currentLink); //get the value from pos_list
count++;
currentLink = nextLink;
}
newLinkToAdd = newLink(getLinkItem(currentLink), NULL); //if lengthof sublist ==0 Insert the value returned from the list A based on the pos_list value to the new sublist
if (getLength(subList) == 0) {
insertAtBeginning(subList, newLinkToAdd);
}
else {
insertLink(subList, previousLinkAddedToSublist, newLinkToAdd); //else insert to next node
}
previousLinkAddedToSublist = newLinkToAdd;
currentLinkForPosList = getLinkNext(currentLinkForPosList);
if (currentLinkForPosList != NULL) {
itemToGet = getLinkItem(currentLinkForPosList);
}
count = 0;
currentLink = startLink;
}
return subList;
}
}
void moveAllMaxAtEnd(list A)
{
int listLength = getLength(A);
if (A == NULL) { //Check if the list A is NULL
printf("List A is NULL \n");
return;
}
else if (listLength == 0) {
printf("List A is empty \n"); //Check if List length is empty
}
else if (listLength > 1) {
int i, j;
int listLength = getLength(A);
link current = getFirst(A); //Get the first link of a list A
Item testItem;
Item newTestItem;
int maxValue = 0; //Assign maxvalue=0
link previous = NULL;
int countOfOccurences = 0; //assign countOfOccurences = 0
for (i = 0; i < listLength; i++) {
Item testItem = getLinkItem(current); //For i = 0 to listlength,compare if testitem is greater than maxvalue
if (testItem > maxValue) {
maxValue = testItem; //if testitem>maxvalue then maxvalue is asigned a new value of testItem
}
link nextLink = getLinkNext(current); //Do the above action for all the items
current = nextLink;
}
current = getFirst(A);
for (i = 0; i < listLength; i++) { //Checks for all the iterations og listlength if testitem==maxvalue
Item testItem = getLinkItem(current);
if (testItem == maxValue) {
countOfOccurences = countOfOccurences + 1; //Increment countOfOccurences for all the items if testitem==maxvalue
}
link nextLink = getLinkNext(current);
current = nextLink;
}
if (countOfOccurences < listLength) { //If countOfOccurences is less than Listlength then remove the links
link linksRemoved[countOfOccurences];
link nextCurrent = getFirst(A);
int arrayPointer = 0;
for (i = 0; i < listLength; i++) {
Item testItem = getLinkItem(nextCurrent);
link nextCurrentLink = getLinkNext(nextCurrent); //remove the max element from the list to be added at the end
if (testItem == maxValue) {
if (previous == NULL) {
setFirst(A, nextCurrentLink);
}
else if (nextCurrentLink != NULL) {
setLinkNext(previous, nextCurrentLink);
}
linksRemoved[arrayPointer] = nextCurrent;
arrayPointer++;
}
previous = nextCurrent;
nextCurrent = nextCurrentLink;
}
for (i = 0; i < countOfOccurences; i++) {
link maxLink = linksRemoved[i]; //add the removed items to the end of the list
setLinkNext(maxLink, NULL);
setLinkNext(previous, maxLink);
previous = maxLink;
}
}
}
}
// swaps the first and the third nodes
void swapFirstThird(list A)
{
int listLength = getLength(A); //Get List length
link previous = NULL;
if (A == NULL) {
printf("List A is Null \n"); //Check if A is NULL
}
else if (listLength == 0) { //Check if list length is zero
printf("List A is empty\n");
}
else if (listLength > 2) {
link current = getFirst(A);
link secondLink = getLinkNext(current);
link thirdLink = getLinkNext(secondLink);
link fourthLink = getLinkNext(thirdLink); //If list length is greater than 2 swap first and third
setLinkNext(current, fourthLink);
setLinkNext(secondLink, current);
setLinkNext(thirdLink, secondLink);
setFirst(A, thirdLink);
}
else if (listLength == 2) {
link current = getFirst(A);
link temp = NULL;
link secondLink = getLinkNext(current); //If list length is equal to 2 swap first and second
temp = current;
A->first = secondLink;
link newFirst = getFirst(A);
setLinkNext(newFirst, temp);
}
}
//
// Set-up and call your testing functions from this function.
int run_student_tests()
{
list A;
list pos_list;
list result;
printf("\n Running function run_student_test.\n ");
printf("\n\n--------------------------------------\n");
/**************************************************TEST CASES FOR DELETE OCCURENCE FUNCTION*******************************************************/
printf("\n Running TestCases For:deleteOccurrences(list A, int V)");
printf("\n*****************************************\n");
/*********************TestCase:1************************************************/
A = NULL;
printf("Test case 1: List A is NULL \n");
deleteOccurrences(A, 20);
destroyList(A);
printf("\n");
/**********************TestCase:2*********************************************/
printf("Test case 2 : List A is empty \n");
A = newList();
deleteOccurrences(A, 20);
destroyList(A);
printf("\n");
/*************************TestCase:3****************************************************/
printf("Test case 3 : Normal data V is in A \n");
int del_arr[] = {15, 100, 7, 5, 100, 7, 30};
A = arrayToList(del_arr, 7);
deleteOccurrences(A, 7);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*************************TestCase:4****************************************************/
printf("Test case 4 : V does not occur in A\n");
int del_arr1[] = {15, 100, 7, 5};
A = arrayToList(del_arr1, 4);
deleteOccurrences(A, 9);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*************************TestCase:5****************************************************/
printf("Test case 5 : Repeated consecutive occurrences\n");
int del_arr2[] = {15, 7, 7, 5};
A = arrayToList(del_arr2, 4);
deleteOccurrences(A, 7);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*************************TestCase:6****************************************************/
printf("Test case 6 : A has one item and that is V\n");
int del_arr3[] = {7};
A = arrayToList(del_arr3, 1);
deleteOccurrences(A, 7);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*************************TestCase:7****************************************************/
printf("Test case 7 : A has only items with Value V in it\n");
int del_arr4[] = {7, 7, 7};
A = arrayToList(del_arr4, 3);
deleteOccurrences(A, 7);
printListHorizPointer(A);
destroyList(A);
printf("\n******************************************************************************\n");
/**************************************************TEST CASES FOR SUBLIST FUNCTION*******************************************************/
printf("\n Running TestCases For:sublist(list A, list pos_list)");
printf("\n*****************************************\n");
/*********************TestCase:1************************************************/
A = NULL;
printf("Test case 1: List A is NULL \n");
int pos_arr[] = {6};
pos_list = arrayToList(pos_arr, 1);
result = sublist(A, pos_list);
printListHorizPointer(result);
destroyList(A);
destroyList(pos_list);
destroyList(result);
printf("\n");
/**********************TestCase:2*********************************************/
printf("Test case 2 : List A is empty \n");
A = newList();
int pos_arr1[] = {5};
pos_list = arrayToList(pos_arr1, 1);
result = sublist(A, pos_list);
printListHorizPointer(result);
destroyList(A);
destroyList(pos_list);
destroyList(result);
printf("\n");
/*********************TestCase:3************************************************/
printf("Test case 3: Index out of bound\n");
int sub_arr2[] = {10, 10, 40, 20};
int pos_arr2[] = {3, 50, 3};
pos_list = arrayToList(pos_arr2, 3);
A = arrayToList(sub_arr2, 4);
result = sublist(A, pos_list);
printListHorizPointer(result);
myDestroyList(pos_list);
destroyList(A);
myDestroyList(result);
printf("\n");
/*********************TestCase:4************************************************/
printf("Test case 4: A is not modified by sublist");
int sub_arr3[] = {15, 100, 7, 5, 100};
int pos_arr3[] = {3, 0, 2};
pos_list = arrayToList(pos_arr3, 3);
A = arrayToList(sub_arr3, 5);
result = sublist(A, pos_list);
printListHorizPointer(A);
destroyList(pos_list);
destroyList(A);
destroyList(result);
printf("\n");
/*********************TestCase:5************************************************/
printf("Test case 5: Normal data");
int sub_arr4[] = {15, 100, 7, 5, 100, 7, 30};
int pos_arr4[] = {3, 0, 6, 4};
pos_list = arrayToList(pos_arr4, 4);
A = arrayToList(sub_arr4, 7);
result = sublist(A, pos_list);
printListHorizPointer(result);
destroyList(A);
destroyList(pos_list);
destroyList(result);
printf("\n");
/*********************TestCase:6************************************************/
printf("Test case 6: Repeated positions");
int sub_arr5[] = {5};
int pos_arr5[] = {0, 0, 0};
pos_list = arrayToList(pos_arr5, 3);
A = arrayToList(sub_arr5, 1);
result = sublist(A, pos_list);
printListHorizPointer(result);
destroyList(A);
destroyList(pos_list);
destroyList(result);
printf("\n");
/*********************TestCase:7***********************************************/
pos_list = NULL;
printf("Test case 7: List pos_list is NULL \n");
int sub_arr6[] = {6, 7};
A = arrayToList(sub_arr6, 2);
result = sublist(A, pos_list);
printListHorizPointer(result);
destroyList(A);
destroyList(pos_list);
destroyList(result);
printf("\n");
/**********************TestCase:8*********************************************/
printf("Test case 8 : List pos_list is empty \n");
pos_list = newList();
int sub_arr7[] = {8, 9, 10};
A = arrayToList(sub_arr7, 3);
result = sublist(A, pos_list);
printListHorizPointer(result);
destroyList(A);
destroyList(pos_list);
destroyList(result);
printf("\n*******************************************************************\n");
/**************************************************TEST CASES FOR SWAP FIRST AND THIRD*******************************************************/
printf("\n Running TestCases For:swapFirstThird(list A)");
printf("\n*****************************************\n");
/*********************TestCase:1************************************************/
A = NULL;
printf("Test case 1: List A is NULL \n");
swapFirstThird(A);
destroyList(A);
printf("\n");
/**********************TestCase:2*********************************************/
printf("Test case 2 : List A is empty \n");
A = newList();
swapFirstThird(A);
destroyList(A);
printf("\n");
/*************************TestCase:3****************************************************/
printf("Test case 3 : List A has one element");
int swap_arr[] = {15};
A = arrayToList(swap_arr, 1);
swapFirstThird(A);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*************************TestCase:4****************************************************/
printf("Test case 4 : List A has two elements");
int swap_arr1[] = {15, 30};
A = arrayToList(swap_arr1, 2);
swapFirstThird(A);
printListHorizPointer(A);
myDestroyList(A);
printf("\n");
/*************************TestCase:5****************************************************/
printf("Test case 5 : List A has three elements");
int swap_arr2[] = {15, 30, 20};
A = arrayToList(swap_arr2, 3);
swapFirstThird(A);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*************************TestCase:6****************************************************/
printf("Test case 6 : List A has more than three elements");
int swap_arr3[] = {15, 30, 20, 40, 50, 16};
A = arrayToList(swap_arr3, 6);
swapFirstThird(A);
printListHorizPointer(A);
destroyList(A);
printf("\n*******************************************************************************/");
/**************************************************TEST CASES FOR MOVE ALL MAX TO END*******************************************************/
printf("\n Running TestCases For:moveAllMaxAtEnd(list A)");
printf("\n*****************************************\n");
/*********************TestCase:1************************************************/
A = NULL;
printf("Test case 1: List A is NULL \n");
moveAllMaxAtEnd(A);
destroyList(A);
printf("\n");
/**********************TestCase:2*********************************************/
printf("Test case 2 : List A is empty \n");
A = newList();
moveAllMaxAtEnd(A);
destroyList(A);
printf("\n");
/*********************TestCase:3************************************************/
printf("Test case 3: Normal data \n");
int max_arr1[] = {15, 100, 5, 100, 30};
A = arrayToList(max_arr1, 5);
moveAllMaxAtEnd(A);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*********************TestCase:4************************************************/
printf("Test case 4: A has one item\n");
int max_arr2[] = {7};
A = arrayToList(max_arr2, 1);
moveAllMaxAtEnd(A);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*********************TestCase:5************************************************/
printf("Test case 5: A has only one item of same value in it\n");
int max_arr3[] = {7, 7, 7};
A = arrayToList(max_arr3, 3);
moveAllMaxAtEnd(A);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*********************TestCase:6************************************************/
printf("Test case 6: Max is on the first position\n");
int max_arr4[] = {100, 7, 20};
A = arrayToList(max_arr4, 3);
moveAllMaxAtEnd(A);
printListHorizPointer(A);
destroyList(A);
printf("\n");
/*********************TestCase:7************************************************/
printf("Test case 7: Max is on the last position\n");
int max_arr5[] = {10, 7, 200};
A = arrayToList(max_arr5, 3);
moveAllMaxAtEnd(A);
printListHorizPointer(A);
myDestroyList(A);
printf("\n************************************************************************\n");
}
<file_sep>/****************************************INSTRUCTIONS************************************************************/
/* Compiling instructions:
*
* connect to omega server with putty by providing host name and username
*
* copy search.c to omega server and execute the following command in SSH to compile
*
* gcc -o search search.c
*
* run the search.c by following command
*
* ./search
*
*
*/
// Created by :<NAME>
/**************************************************************************************************************/
#include <stdio.h>
#include <stdlib.h>
#include<math.h>
#include<time.h>
#include<unistd.h>
#include<string.h>
/*********************************PROGRAM FUNCTIONALITY********************************************************/
/*
1. Code provides 3 options for the user to enter 0-exit ,1-to create and save random data for search,2-load the data ,sort and run searches
2. When user selects 0 code terminates.
3. When user enters 1 he is asked to enter numbers of elements , elements to be searched within a particular range and a filename.
4. Code generates random numbers for N and S
5. Codes reads and stores the data
6. Runs selection sort on N and prints the sorted array
7. Runs the binary and interpolation search on S values
8. Compare the results of binary and interpolation search
*/
/***************************************************************************************************************/
/***********************************STRUCTURE DEFINITION********************************************************/
struct binresult
{
int bin_index;
int bin_iterations;
};
struct interresult
{
int inter_index;
int inter_iterations;
};
struct searchResults
{
int index;
int value;
int interpolationIndex;
int binaryIndex;
int interpolationIteration;
int binaryInteration;
};
/*****************************************************************************************************************/
/**************************************FUNCTION DECLARATION*******************************************************/
void selection(int A[], int M);
void printArray(int arr[], int size);
struct binresult binarysearch(int A[], int N, int v);
struct interresult interpolatedsearch(int A[], int N, int v);
/****************************************************************************************************************/
int main(void)
{
/****************************************VARIABLE DECLARATION****************************************************/
int N; //To store the number of elements
int S; //To store elements to be searched
int start_value; //Start value of the range
int end_value; //End value of the range
int temp; //temporary variable to store random numbers generated for N and S values
int i, j, k, l; //Index used in For loop
int x; //Element to be searched in binary search
int y; //Element to be searched in interpolation search
char filename[100]; //Name of the file where data will be saved
int exitflag = 1; //Used for While loop
int choice; //To store the user entered choice
FILE *fptr; //Declare pointer of a type file
/***************************************************************************************************************/
while (exitflag) {
/****************************************CHOICE ENTERED BY THE USER************************************/
printf("Choose from the menu option below\n");
printf("0-exit\n");
printf("1-create and save random data for search.\n");
printf("2-load data from file,sort array and run searches.\n");
printf("Enter your choice:\n");
scanf("%d", &choice);
switch (choice) {
/*****************************************WHEN USER SELECTS CHOICE '0'********************************************/
case 0:
{
printf("See You In Class");
exit(1);
break;
}
/****************************************CASE 1: CREATE AND SAVE RANDOM DATA FOR SEARCHES***************************/
case 1:
{
/***************************************READ DATA FROM CONSOLE STORE IT IN A VARIABLE********************************/
printf("Enter N S start_value end_value filename : ");
scanf("%d", &N);
scanf("%d", &S);
scanf("%d", &start_value);
scanf("%d", &end_value);
scanf("%s", filename);
/*******************************************SETTING BOUNDARY LIMIT FOR START_VALUE AND END_VALUE(start_value should be less than end_value)***********************/
while (start_value > end_value) {
printf("Enter N S start_value end_value filename : ");
scanf("%d", &N);
scanf("%d", &S);
scanf("%d", &start_value);
scanf("%d", &end_value);
scanf("%s", filename);
}
/**************************************************STORE THE VALUES ENTERED BY THE USER INTO A FILE**********************************************************************************/
fptr = fopen(filename, "w");
fprintf(fptr, "%d %d %d %d\n", N, S, start_value, end_value);
fclose(fptr);
/**************************************************GENERATE RANDOM NUMBERS FOR N AND STORE IT IN A FILE**********************************************************************/
int arrayOfrandomNumbers[N];
srand((unsigned int) time(NULL));
for (i = 0; i <= N - 1; i++) {
temp = (rand() % (end_value - start_value + 1)) + start_value;
arrayOfrandomNumbers[i] = temp;
}
for (i = 0; i <= N - 1; i++) {
fptr = fopen(filename, "a");
fprintf(fptr, "%d ", arrayOfrandomNumbers[i]);
fclose(fptr);
}
fptr = fopen(filename, "a");
fprintf(fptr, "\n");
fclose(fptr);
/******************************************************GENERATE RANDOM NUMBERS FOR S AND STORE IT IN A FILE****************************************************************/
int arrayOfNumbersToSearch[S];
srand((unsigned int) time(NULL));
for (j = 0; j <= S - 1; j++) {
temp = (rand() % (end_value - start_value + 1)) + start_value;
arrayOfNumbersToSearch[j] = temp;
}
for (i = 0; i <= S - 1; i++) {
fptr = fopen(filename, "a");
fprintf(fptr, "%d ", arrayOfNumbersToSearch[i]);
fclose(fptr);
}
break;
}
/*******************************************************END OF CASE 1*******************************************************************************************************/
/*****************************************************CASE 2 :LOAD DATA FROM FILE,SORT ARRAY AND RUN SEARCHES***************************************************************/
case 2:
{
FILE *fptrToOpen; //Declare pointer of a type file
int verboseFlag; // Used for while loop
int option; // To store user entered option for verbose and non verbose output
float a, b; //To store the averages of loop iterations
float binarySum = 0.0, interpolationSum = 0.0; //To store the sum of loop iterations
/*******************************************************USER SELECTS THE FILENAME TO READ AND OPTION 1 OR 2 FOR VERBOSE AND NON-VERBOSE****************************************/
printf("Enter: filename, mode(1-verbose, 2-not verbose:");
scanf("%s", filename);
scanf("%d", &option);
verboseFlag = (option == 1 || option == 2);
while (!verboseFlag) {
printf("\nEnter the format for the output(1- for verbose , 2- for non-verbose):");
scanf("%d", &option);
verboseFlag = (option == 1 || option == 2);
}
/****************************************************WHEN USER ENTERS WRONG FILENAME*******************************************************************************************/
fptrToOpen = fopen(filename, "r");
if (fptrToOpen == NULL) {
printf("\n ERROR:File could not be opened \n\n");
break;
}
/**************************************************READ AND STORE DATA FROM A FILE AND STORE IT IN AN ARRAY*********************************************************************/
fscanf(fptrToOpen, "%d", &N);
fscanf(fptrToOpen, "%d", &S);
fscanf(fptrToOpen, "%d", &start_value);
fscanf(fptrToOpen, "%d", &end_value);
int randomNumberArrayFromFile[N], searchNumbersArrayFromFile[S];
struct searchResults sr[S];
for (i = 0; i < N; i++) {
fscanf(fptrToOpen, "%d", &temp);
randomNumberArrayFromFile[i] = temp;
}
for (j = 0; j < S; j++) {
fscanf(fptrToOpen, "%d", &temp);
searchNumbersArrayFromFile[j] = temp;
}
/*************************************************FUNCTION CALL :SELECTION SORT,PRINT ARRAY,BINARY SEARCH AND INTERPOLATION SEARCH************************************************/
selection(randomNumberArrayFromFile, N);
printf("\nSorted array: ");
printArray(randomNumberArrayFromFile, N);
// //call binary search
for (k = 0; k < S; k++) {
x = searchNumbersArrayFromFile[k];
struct binresult binresultValue = binarysearch(randomNumberArrayFromFile, N, x);
struct interresult interresultValue = interpolatedsearch(randomNumberArrayFromFile, N, x);
sr[k].index = k;
sr[k].value = searchNumbersArrayFromFile[k];
sr[k].interpolationIndex = interresultValue.inter_index;
sr[k].binaryIndex = binresultValue.bin_index;
sr[k].interpolationIteration = interresultValue.inter_iterations;
sr[k].binaryInteration = binresultValue.bin_iterations;
binarySum = binarySum + sr[k].binaryInteration;
interpolationSum = interpolationSum + sr[k].interpolationIteration;
}
/***************************************************PERFORM AVERAGES OF NUMBER OF ITERATIONS OF BINARY AND INTERPOLATION SEARCH******************************************************************/
a = binarySum / S;
b = interpolationSum / S;
/****************************************************PRINT RESULT FOR VERBOSE OUTPUT WITH FORMATTED TABLE****************************************************************************************/
if (option == 1) {
printf("\n| | | found at index | repetitions |\n");
printf("| i| value| interp| binary| interp| binary|\n");
for (l = 0; l < S; l++) {
printf("|%8d|%8d|%8d|%8d|%8d|%8d|\n", sr[l].index, sr[l].value,
sr[l].interpolationIndex, sr[l].binaryIndex,
sr[l].interpolationIteration, sr[l].binaryInteration);
}
printf("| avg| | | |%8.2f|%8.2f|\n", b, a);
}
/****************************************************PRINT RESULT FOR NON-VERBOSE OUTPUT WITH FORMATTED TABLE*********************************************************************************/
else {
printf("\n| | | found at index | repetitions |\n");
printf("| i| value| interp| binary| interp| binary|\n");
printf("| avg| | | |%8.2f|%8.2f|\n", b, a);
}
break;
}
}
}
}
/**************************************************************END OF CASE 2********************************************************************************************/
/**************************************************************FUNCTION:SELECTION SORT**********************************************************************************/
void selection(int A[], int M)
{
int i, j, temp;
for (j = 0; j < M; j++) {
int min_idx = j;
for (i = j + 1; i < M; i++)
if (A[i] < A[min_idx])
min_idx = i;
temp = A[min_idx];
A[min_idx] = A[j];
A[j] = temp;
}
}
/***************************************************************FUNCTION:TO PRINT SORTED ARRAY************************************************************************************/
void printArray(int arr[], int size)
{
int i;
for (i = 0; i < size; i++) {
printf("%d ", arr[i]);
}
printf("\n");
}
/***************************************************************BINARY SEARCH***********************************************************************************/
struct binresult binarysearch(int A[], int N, int v)
{
struct binresult r;
int left, right, count = 0;
left = 0;
right = N - 1;
while (left <= right) {
int m = (left + right) / 2;
count++;
if (v == A[m]) {
r.bin_index = m;
r.bin_iterations = count;
return r;
}
if (v < A[m])
right = m - 1;
else
left = m + 1;
}
r.bin_index = -1;
r.bin_iterations = count;
return r;
}
/******************************************************************INTERPOLATION SEARCH**********************************************************************************/
struct interresult interpolatedsearch(int A[], int N, int v)
{
struct interresult interPolatedSearchResult;
int left, right, count = 0;
left = 0;
right = N - 1;
while (left <= right && v >= A[left] && v <= A[right]) {
int m = left + ((v - A[left])*(right - left) / (A[right] - A[left]));
count++;
if (v == A[m]) {
interPolatedSearchResult.inter_index = m;
interPolatedSearchResult.inter_iterations = count;
return interPolatedSearchResult;
}
if (v < A[m])
right = m - 1;
else
left = m + 1;
}
interPolatedSearchResult.inter_index = -1;
interPolatedSearchResult.inter_iterations = count;
return interPolatedSearchResult;
}
<file_sep>// Implementation provided by <NAME>
#include <stdio.h>
#include <stdlib.h>
#include <stdbool.h>
#include <string.h>
/******************************************Structure definition**************************************************************/
struct item
{
char englishWord[100];
char spanishWord[1000];
};
/****************************************Function: Quadratic Probing*********************************************************/
int QuadraticProbing(int hashKey, int i, int M)
{
return (hashKey + (3 * i)+(5 * i * i)) % M;
}
/****************************************Function:Calculate hash value******************************************************/
int hash(char *v, int M)
{
int h = 0, a = 127;
for (; *v != '\0'; v++)
h = (a * h + *v) % M;
return h;
}
/***************************************Structure(Copy english word to hash table)********************************************/
struct item copyStructItem(struct item source, struct item destination)
{
strncpy(destination.englishWord, source.englishWord, strlen(source.englishWord));
strncpy(destination.spanishWord, source.spanishWord, strlen(source.spanishWord));
return destination;
}
/***************************************Structure(Append spanish word)**********************************************************/
struct item appendSpanishWord(struct item source, struct item destination)
{
strncat(destination.spanishWord, ";", 1);
strncat(destination.spanishWord, source.spanishWord, strlen(source.spanishWord));
return destination;
}
/***************************************unction:To print message************************************************************************************/
void printInsertMessage(struct item source)
{
printf("\tWill insert pair [%s,%s]\n", source.englishWord, source.spanishWord);
}
/*********************Function:To check if the position in the table is empty*******************************************************************/
bool isPositionEmpty(struct item destination)
{
int test = strcmp(destination.englishWord, " ");
int test1 = strcmp(destination.englishWord, "DELETED");
return (test == 0) && (test1 != 0);
}
/***************************************Function:Check if word is deleted************************************************************************************/
bool isWordDeleted(struct item destination)
{
int test = strcmp(destination.englishWord, "DELETED");
return (test == 0);
}
/***************************************Function:To check if new word is present************************************************************************************/
bool isNewWordAlreadyPresent(struct item destination, struct item source)
{
int test = (strcmp(destination.englishWord, source.englishWord));
int test1 = strcmp(destination.englishWord, "DELETED");
return (test == 0) && (test1 != 0);
}
/***************************************Function:check if word is present************************************************************************************/
bool isSearchWordPresent(struct item destination, char* source)
{
int test = (strcmp(destination.englishWord, source));
int test1 = strcmp(destination.englishWord, "DELETED");
return (test == 0)&& (test1 != 0);
}
/***************************************Function:Calculate hash table result************************************************************************************/
void calculateAndPrintHashTableResults(int keyCount[])
{
int i;
float totalProbesDictionary = 0;
float totalKeysDictionary = 0;
int maxProbes = 0;
for (i = 1; i <= 100; i++) {
totalProbesDictionary = totalProbesDictionary + (i * keyCount[i]);
totalKeysDictionary = totalKeysDictionary + keyCount[i];
if (keyCount[i] != 0) {
maxProbes = i;
}
}
//***************************************Print hash table************************************************************************/
printf("\n\n");
printf("Hash Table\n");
printf("\taverage number of probes:\t\t %.2f\n", (totalProbesDictionary / totalKeysDictionary));
printf("\tmax_run of probes:\t\t\t %d\n", maxProbes);
printf("\ttotal PROBES (for %f items):\t%.2f\n", totalKeysDictionary, totalProbesDictionary);
printf("\titems NOT hashed (out of %f):\t 0\n", totalKeysDictionary);
}
//***************************************Function: Print dictionary**************************************************************/
void printDictionary(struct item dictionary[], int size)
{
int i;
for (i = 0; i < size; i++) {
printf("%s-%s\n", dictionary[i].englishWord, dictionary[i].spanishWord);
}
}
void printDataItem(struct item a)
{
printf("%s-%s", a.englishWord, a.spanishWord);
}
/*****************************************Function:To split english and spanish word********************************************/
struct item splitWord(char str[], int lengthstr)
{
struct item dictionaryitems;
int x, y;
int i = 0;
while (str[i] != '\t') {
i++;
}
x = i;
y = i + 1;
for (i = 0; i < x; i++) {
dictionaryitems.englishWord[i] = str[i];
}
dictionaryitems.englishWord[i] = '\0';
for (i = 0; i < (lengthstr - 1 - y); i++) {
dictionaryitems.spanishWord[i] = str[y + i];
}
dictionaryitems.spanishWord[i] = '\0';
return dictionaryitems;
}
//***************************************Function: To search english word**************************************************************/
int search(char english[], struct item dictionary[], int size)
{
int probes = 1;
int key = hash(english, size);
bool wordFound = false;
bool repeat = true;
if (isSearchWordPresent(dictionary[key], english)) {
printf("\t%d probes\n", probes);
printf("\tTranslation: %s\n", dictionary[key].spanishWord);
wordFound = true;
}
else if (isWordDeleted(dictionary[key])) {
int i = 0;
while (repeat) {
probes = probes + 1;
int quadraticKey = QuadraticProbing(key, i, size);
if (isSearchWordPresent(dictionary[quadraticKey], english)) {
printf("\t%d probes\n", probes);
printf("\tTranslation: %s\n", dictionary[quadraticKey].spanishWord);
wordFound = true;
repeat = false;
break;
}
else if (!isWordDeleted(dictionary[quadraticKey])) {
repeat = false;
break;
}
else {
i++;
}
}
}
else {
int i = 0;
while (i < size) {
probes = probes + 1;
int quadraticKey = QuadraticProbing(key, i, size);
if (isSearchWordPresent(dictionary[quadraticKey], english)) {
printf("\t%d probes\n", probes);
printf("\tTranslation: %s\n", dictionary[quadraticKey].spanishWord);
i = size;
wordFound = true;
break;
}
else {
i++;
}
}
}
if (!wordFound) {
printf("\t%d probes\n", probes);
printf("\tNOT found\n");
}
return probes;
}
//***************************************Function:To insert a english word and its translations**********************************/
struct item* insert(struct item dictionary[], struct item data, int size, bool addToProbes, int *keyCount, int *insertProbes)
{
int probes = 1;
int key = hash(data.englishWord, size);
if (isPositionEmpty(dictionary[key])) {
dictionary[key] = copyStructItem(data, dictionary[key]);
}
else if (isNewWordAlreadyPresent(dictionary[key], data)) {
dictionary[key] = appendSpanishWord(data, dictionary[key]);
}
else {
int i = 0;
while (i < size) {
probes = probes + 1;
int quadraticKey = QuadraticProbing(key, i, size);
if (isPositionEmpty(dictionary[quadraticKey])) {
dictionary[quadraticKey] = copyStructItem(data, dictionary[quadraticKey]);
break;
}
if (isNewWordAlreadyPresent(dictionary[quadraticKey], data)) {
dictionary[quadraticKey] = appendSpanishWord(data, dictionary[quadraticKey]);
break;
}
else {
i++;
}
}
}
if (addToProbes) {
*insertProbes = *insertProbes + probes;
printf("\t%d probes\n", probes);
}
else {
keyCount[probes]++;
}
return dictionary;
}
//***************************************Function:To delete key value pair**************************************************************/
int delete(char english[], struct item dictionary[], int size)
{
int probes = 1;
int key = hash(english, size);
bool wordFound = false;
bool repeat = true;
if (isSearchWordPresent(dictionary[key], english)) {
strncpy(dictionary[key].englishWord, "DELETED", 100);
strncpy(dictionary[key].spanishWord, "DELETED", 1000);
printf("\t%d probes\n", probes);
printf("\tItem was deleted\n");
wordFound = true;
}
else if (isWordDeleted(dictionary[key])) {
int i = 0;
while (repeat) {
probes = probes + 1;
int quadraticKey = QuadraticProbing(key, i, size);
if (isSearchWordPresent(dictionary[quadraticKey], english)) {
strncpy(dictionary[quadraticKey].englishWord, "DELETED", 100);
strncpy(dictionary[quadraticKey].spanishWord, "DELETED", 1000);
wordFound = true;
repeat = false;
break;
}
else if (!isWordDeleted(dictionary[quadraticKey])) {
repeat = false;
break;
}
else {
i++;
}
}
}
else {
int i = 0;
while (i < size) {
probes = probes + 1;
int quadraticKey = QuadraticProbing(key, i, size);
if (isSearchWordPresent(dictionary[quadraticKey], english)) {
strncpy(dictionary[quadraticKey].englishWord, "DELETED", 100);
strncpy(dictionary[quadraticKey].spanishWord, "DELETED", 1000);
printf("\t%d probes\n", probes);
printf("\tItem was deleted\n");
wordFound = true;
i = size;
}
else {
i++;
}
}
}
if (!wordFound) {
printf("\t%d probes\n", probes);
printf("\tItem not found => no deletion.\n");
}
return probes;
}
int main()
{
char filename[100];
FILE *test;
char choice[50];
char str[500];
char c;
int size = 20123;
int count = 0;
char file[100];
char english[100];
char spanish[100];
char result[1000];
struct item temp;
int keyCount[101] = {0};
int totalProbes = 0;
int totalOperations = 0;
int probes = 1;
int i;
int key;
//****************************************Create a empty hash table***********************************************************************************/
struct item *dictionary = (struct item*) malloc(size * sizeof (struct item));
for (i = 0; i < size; i++) {
strncpy(dictionary[i].englishWord, " ", 100);
strncpy(dictionary[i].spanishWord, " ", 1000);
}
printf("Enter the filename with the dictionary data (include the extension e.g. Spanish.txt):");
scanf("%s", &filename);
test = fopen(filename, "r");
for (c = getc(test); c != EOF; c = getc(test)) {
if (c == '\n') { // Increment count if this character is newline
count = count + 1;
}
}
i = 1;
fclose(test);
int j = 1;
test = fopen(filename, "r");
while (j <= count) {
fgets(str, 250, test);
int lengthstr = strlen(str);
temp = splitWord(str, lengthstr);
dictionary = insert(dictionary, temp, size, false, &keyCount[0], &totalProbes);
j++;
}
fclose(test);
/****************************************Print hash table results*************************************************************/
calculateAndPrintHashTableResults(keyCount);
/*****************************************Print the table with count of keys**************************************************/
printf("\n\n");
printf("Probes|Count of keys\n");
printf("-------------\n");
for (i = 1; i <= 100; i++) {
printf("%6d|%6d\n", i, keyCount[i]);
printf("-------------\n");
}
printf("\n");
printf("Enter words to look-up. Enter -1 to stop.");
printf("\n");
/******************************************Perform read operations*************************************************************/
while (1) {
totalOperations = totalOperations + 1;
printf("READ op:");
scanf("%s", &choice);
printf("%s", choice);
if (strcasecmp(choice, "-1") == 0) {
return 0;
}
int value2 = strcmp(choice, "q");
if (value2 == 0) {
printf("\n\n\n");
printf("Average probes per operation: %f", (float) totalProbes / (float) totalOperations);
printf("\n\n\n");
free(dictionary);
exit(0);
}
int value = strcmp(choice, "s");
if (value == 0) {
printf(" query:");
scanf("%s", &english);
printf("%s", english);
printf("\n");
probes = search(english, dictionary, size);
totalProbes = totalProbes + probes;
}
int value1 = strcmp(choice, "d");
if (value1 == 0) {
printf(" query:");
scanf("%s", &english);
printf("%s", english);
printf("\n");
probes = delete(english, dictionary, size);
totalProbes = totalProbes + probes;
}
int value3 = strcmp(choice, "i");
if (value3 == 0) {
printf(" query:");
scanf("%s", temp.englishWord);
printf("%s", temp.englishWord);
scanf("%s", temp.spanishWord);
printf("\n");
insert(dictionary, temp, size, true, &keyCount[0], &totalProbes);
printInsertMessage(temp);
}
}
}
<file_sep>// Implementation provided by <NAME>
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include<strings.h>
/********************************************Function:To find minimum between 3 numbers**********************************************************************************/
int findMinimum(int a, int b, int c)
{
if (a <= b && a <= c) {
return a;
}
if (b <= a && b <= c) {
return b;
}
return c;
}
/*********************************************Function: To calculate edit distance*************************************************************************************************/
int editDistance(char str1[], char str2[], int length_str1, int length_str2)
{
int i, j;
int editDistance[length_str1 + 1][length_str2 + 1];
for (i = 0; i <= length_str1; i++) {
for (j = 0; j <= length_str2; j++) {
if (i == 0 && j == 0) { //When i =0 and j=0
editDistance[0][0] = 0;
}
else if(i == 0) //When i=0
{
editDistance[0][j] = editDistance[i][j-1]+1;
}
else if (j == 0) //When j=0
{
editDistance[i][0] = editDistance[i-1][j]+1;
}
else {
if (str1[i - 1] == str2[j - 1]) { //when str1[i-1]=str2[i-1]
editDistance[i][j] = findMinimum(editDistance[i - 1][j - 1], editDistance[i][j - 1] + 1, editDistance[i - 1][j] + 1); //Find minimum
}
else {
editDistance[i][j] = findMinimum(editDistance[i - 1][j - 1] + 1, editDistance[i][j - 1] + 1, editDistance[i - 1][j] + 1); //Find minimum
}
}
}
}
/*******************************************************PRINT THE OUTPUT IN TABLE FORM*********************************************************************************/
printf("\n");
int m;
for (i = 0; i <= length_str2 + 1; i++) {
if (i < 2) {
printf(" |");
}
else {
printf("%3c|", str2[i - 2]);
}
}
printf("\n");
for (m = 0; m < (length_str2 + 2)*4; m++) {
printf("-");
}
printf("\n");
for (i = 0; i <= length_str1; i++) {
for (j = 0; j <= length_str2 + 1; j++) {
if (j == 0 && i == 0) {
printf(" |");
}
else if (j == 0) {
printf("%3c|", str1[i - 1]);
}
else {
printf("%3d|", editDistance[i][j - 1]);
}
}
printf("\n");
for (m = 0; m < (length_str2 + 2)*4; m++) {
printf("-");
}
printf("\n");
}
printf("edit distance: %d", editDistance[length_str1][length_str2]);
printf("\n");
printf("--------------------------------------------------------------------------------------------------------");
printf("\n\n");
}
/***************************************************MAIN FUNCTION****************************************************************************/
int main()
{
char str1[100]; //Variable declaration
char str2[100];
printf("\n");
printf("Enter two words separated by a space (e.g.: cat someone).Stop with: -1 -1 : ");
printf("\n");
while (1) {
printf("First : ");
scanf("%s", str1);
printf("%s", str1);
printf("\n");
printf("Second :");
scanf("%s", str2);
printf("%s", str2);
printf("\n");
if ((strcmp("-1", str1)) == 0) {
break;
}
if((strcmp("-1", str2)) == 0){
break;
}
int length_str1 = strlen(str1);
int length_str2 = strlen(str2);
int value=editDistance(str1, str2, length_str1, length_str2); //Call edit distance function
}
return 0;
}
<file_sep>// Implementation provided by <NAME>
#include <stdio.h>
#include <stdbool.h>
#include <stdlib.h>
#include <stdlib.h>
#include <string.h>
#include<strings.h>
/****************************Substring Function declaration****************************************************/
char * substring(char s[], int p, int l);
/*****************************Function:To test elements in increasing order or not****************************/
int increasing(int * A, int N) {
if (N <= 1)
return 1;
else if (A[N - 1] >= A[N - 2]) {
return increasing(A, N - 1);
}
return 0;
}
/*****************************Function min_rec_1:To find minimum in an array**********************************/
int min_rec_1(int * A, int N, int * rec_ct, int * base_ct,int *max_depth) {
*max_depth=*max_depth+1;
if (N == 1) //base case
{
* base_ct = * base_ct + 1;
return A[0];
}
* rec_ct = * rec_ct + 1;
int temp = min_rec_1(A, N - 1, & ( * rec_ct), & ( * base_ct),&(*max_depth));
if (temp < A[N - 1])
return temp;
else
return A[N - 1];
}
/*****************************Function min_rec_2:To find minimum in an array**********************************/
int min_rec_2(int * A, int N, int * rec_ct, int * base_ct, int *max_depth) {
if (N == 1) //base case
{
* base_ct = * base_ct + 1;
return A[0];
}
if (min_rec_2(A, N - 1, & ( * rec_ct), & ( * base_ct), & ( *max_depth))< A[N - 1]) {
* rec_ct = * rec_ct + 1;
*max_depth=*max_depth+1;
return min_rec_2(A, N - 1, & ( * rec_ct), & ( * base_ct), & ( *max_depth));
} else{
*rec_ct = * rec_ct + 1;
}
return A[N - 1];
}
/*****************************Function min_rec_tail:To find minimum in an array**********************************/
int min_rec_tail(int * A, int N, int * rec_ct, int * base_ct,int *max_depth) {
*max_depth=*max_depth+1;
if (N == 0) {
* base_ct = * base_ct + 1;
return A[0];
}
* rec_ct = * rec_ct + 1;
N--;
return min_rec_tail(A + (A[0] > A[N]), N, & ( * rec_ct), & ( * base_ct),&(*max_depth));
}
/*****************************Function substring:To generate substring from a string*******************************/
char * substring(char s[], int p, int l) {
int substringLength = l - p;
int i;
char * newSubstring = malloc((substringLength+1) *sizeof(char));
int count = 0;
for (i = p; i < l; i++) {
newSubstring[count] = s[i];
count = count + 1;
}
newSubstring[count] = '\0';
return newSubstring;
free(newSubstring);
}
/*****************************Function checkPalindrome :To Test if the string is palindrome or not****************/
int checkPalindrome(char * Arr, int l, int r) {
while (l < r)
{
if (Arr[l] != Arr[r]) //compare characters
return 0; //return 0 if its not a palindrome
l++;
r--;
}
return 1; ////return 1 if its a palindrome
}
/*****************************Function PalindromeDecomp :Recursive function for palindrome decomposition****************/
void PalindromeDecomp(int stringlength, int pos, char * myString, char testArray[100][100], int * count) {
int i = pos;
int z;
if (pos >= stringlength) { //if the index is greater than string length print the array
for (z = 0; z < stringlength; z++) {
if (strlen(testArray[z]) != 0) {
printf("%s,", testArray[z]);
}
}
( * count) ++; //generate the count of substrings generated
printf("\n");
return;
}
for (i = pos; i < stringlength; i++) {
if (checkPalindrome(myString, pos, i)) { //Function call to check palindrome
int newSubsLength = i - pos + 1;
char * newSubstring;
newSubstring = substring(myString, pos, i + 1); //if entered string is palindrome then call a substring function
strcpy(testArray[pos], newSubstring); //copy the substring generated to test array
PalindromeDecomp(stringlength, i + 1, myString, testArray, count); //palindrome decomposition recursive call
strcpy(testArray[pos], "");
free(newSubstring);
}
}
}
/*********************Function palindromeDecompositionWrapper:Wrapper function to support Function PalindromeDecomp****************/
void palindromeDecompositionWrapper(char st[]) {
int stringlength = strlen(st); //calculate string length
char *myString = malloc((stringlength + 1) * sizeof(char));
strcpy(myString, st); //copy entered string to a new variable
printf("\n");
int count = 0;
char testArray[100][100] ={0};
PalindromeDecomp(stringlength, 0, myString, testArray, & count); //Function call
printf("%d", count);
free(myString);
}
/*********************************************************************************************************************************/
int main() {
/*******************************************Variable declaration*****************************************************************/
int N; //To enter array size
int i; //variable for the loops
char st[100]; //declare a string
/*******************************************************************************************************************************/
/**************************values to be entered to test if array is increasing or decreasing**************************************/
while (1) {
printf("Enter size of array.(Stop with -1)"); //Enter array size
scanf("%d", &N);
if (N == -1) { //when user enters -1 program terminates and ask user to enter values for next function
break;
}
printf("Enter array elements separated by spaces:"); //Enter array elements and store it in an array
int a[N];
for (i = 0; i < N; ++i) {
scanf("%d", &a[i]);
}
int increasing_result = increasing(a, N); //Function call
if (increasing_result == 1)
printf("increasing:1\n");
else
printf("increasing:0\n");
}
/***********************************values to be entered for Function:to find minimum in an array****************************************/
while (1) {
printf("Enter size of array(Stop with -1.)\n"); //Enter array size
scanf("%d", &N);
if (N == -1) {
break; //when user enters -1 program terminates and ask user to enter values for next function
}
printf("Enter array elements separated by spaces\n");
int a[N]; //Enter array elements and store it in an array
for (i = 0; i < N; ++i) {
scanf("%d", &a[i]);
}
int rec_ct = 0;
int base_ct = 0;
int max_depth=0;
int minimum2 = min_rec_1(a, N, &rec_ct, &base_ct,&max_depth);
printf("\n");
printf("smallest_1:\tsmallest = %d, base_ct = %d, rec_ct = %d, max_depth = %d\n",minimum2,base_ct,rec_ct,max_depth); //Function call
rec_ct = 0;
base_ct = 0;
max_depth=0;
int minimum1 = min_rec_2(a, N, &rec_ct, &base_ct, &max_depth);
printf("smallest_2:\tsmallest = %d, base_ct = %d, rec_ct = %d, max_depth = %d\n",minimum1,base_ct,rec_ct,max_depth); //Function call
rec_ct = 0;
base_ct = 0;
max_depth=0;
int minimum3 = min_rec_tail(a, N, &rec_ct, &base_ct,&max_depth);
printf("smallest_tail:\tsmallest = %d, base_ct = %d, rec_ct = %d, max_depth = %d\n",minimum3,base_ct,rec_ct,max_depth); //Function call
}
/*******************************Values to be Entered for function to decompose the strings**********************************************/
int count = 0;
printf("Enter strings. Stop with -1."); //Enter strings
while (1) {
printf("\ns =");
scanf("%s", & st);
if (strcasecmp(st, "-1") == 0) { //When user enters -1 terminates the program
return 0;
}
printf("\n");
palindromeDecompositionWrapper(st); //palindromeDecompositionWrapper Function call
printf("\n");
}
return 0;
}
<file_sep>// Implementation provided by <NAME>
#include <stdio.h>
#include <stdlib.h>
#include <stdbool.h>
/***********************************Function:Depth First Search************************************************************************/
void depthFirstSearch(int G[10][10], int* nodeIncluded, int node, int size, char name[][200], bool print) {
int j;
if(print)
printf("%s\n",name[node]);
nodeIncluded[node] = 1;
for(j=0; j<size; j++) {
if(nodeIncluded[j]==0 && G[node][j] == 1) {
depthFirstSearch(G,nodeIncluded,j,size, name, print);
}
}
}
/***********************************Main Function**************************************************************************************/
int main() {
int N; //Declare Number of vertices
int i; //Variable for loops
int j; //Variables for loops
int numberOfEdges = 0;
bool flag = true;
printf("\n");
printf("Program output: ");
printf("\n");
printf("\n");
printf("Enter number of vertices, N:");
scanf("%d", &N);
if(N==-1)
{
return 0;
}
/************************************Variable declaration******************************************************************************************/
char name[N][200];
char name1[N][200];
char name2[N][200];
char inputName1[200];
char inputName2[200];
int resultMatrix[N][N];
for (i = 0; i < N; i++) {
printf("Enter name1: ");
scanf("%s", &name[i]);
if (strcasecmp(name[i], "-1") == 0) {
return 0;
}
}
/*************************************Print Matrix********************************************************************************************/
for(i=0; i<N; i++)
{
for(j=0; j<N; j++)
{
resultMatrix[i][j] = 0;
}
}
i = 0;
while (flag) {
printf("Enter name1 name2:");
scanf("%s", &name1[i]);
scanf("%s", &name2[i]);
if (strcasecmp(name1[i], "-1") == 0 && strcasecmp(name2[i], "-1") == 0) {
flag = false;
numberOfEdges = i;
}
i++;
}
printf("\nGRAPH:\nN = %d\n",N);
for (i = 0; i < N; i++) {
printf("%d-%s\n",i, name[i]);
}
for(i=0; i<numberOfEdges; i++) {
int indexOfFirstNode = 0;
int indexOfSecondNode = 0;
for(j=0; j<N; j++) {
if(strcasecmp(name1[i], name[j]) == 0) {
indexOfFirstNode = j;
}
}
for(j=0; j<N; j++) {
if(strcasecmp(name2[i], name[j]) == 0) {
indexOfSecondNode = j;
}
}
resultMatrix[indexOfFirstNode][indexOfSecondNode] = 1;
resultMatrix[indexOfSecondNode][indexOfFirstNode] = 1;
}
printf("\n");
for(i=0; i<N; i++)
{
for(j=0; j<N; j++)
{
printf("%d ",resultMatrix[i][j]);
}
printf("\n");
}
/**************************************Implementing DFS To form Groups*********************************************************************************************/
int nodeIncluded[N];
for(i=0; i<N; i++) {
nodeIncluded[i] = 0;
}
int groupNumber = 1;
int x = 0;
for(i=0; i<N; i++) {
if(nodeIncluded[i] == 0) {
x = x+1;
depthFirstSearch(resultMatrix,nodeIncluded,i,N, name, false);
}
}
printf("Number of groups: %d\n",x);
for(i=0; i<N; i++) {
nodeIncluded[i] = 0;
}
x = 1;
for(i=0; i<N; i++) {
if(nodeIncluded[i] == 0) {
printf("Group: %d\n",x );
x = x+1;
depthFirstSearch(resultMatrix,nodeIncluded,i,N, name, true);
printf("\n");
}
}
return 0;
}
<file_sep>// Implementation provided by <NAME>
#include <stdio.h>
#include <stdlib.h>
/*******************************************RECURSIVE FUNCTION******************************************************************/
int foo(int N) {
if (N <= 1) return 5;
int res1 = 3 * foo(N / 2);
int res2 = foo(N - 1);
if (res1 >= res2)
return res1;
else
return res2;
}
/*******************************************ITERATIVE FUNCTION******************************************************************/
int foo_iterative(int N) {
int i;
int res1;
int res2;
int itr[N];
itr[0] = 5;
itr[1] = 5;
if(N<=1){
return 5;
}
for(i=2;i<=N;i++){
res1 = 3* itr[i/2];
res2 = itr[i-1];
if(res1>= res2){
itr[i]= res1;
}
else{
itr[i]= res2;
}
}
return itr[N];
}
/*******************************************WRAPPER FUNCTION FOR MEMOIZED CODE***************************************************/
int foo_wrapper(int N) {
int i;
int solution[N + 1];
solution[0] = 5;
solution[1] = 5;
for (i = 2; i < N + 1; i++) {
solution[i] = -1;
}
int max_depth = 2;
return foo_memoized(N, solution, max_depth);
}
/**********************************************MEMOIZED FUNCTION**********************************************************************************/
int foo_memoized(int N, int * Solution, int max_depth) {
int i;
max_depth = max_depth + 3;
for (i = 0; i < max_depth; i++) {
printf(" ");
}
printf("N = %d\n\n", N);
if (Solution[N] != -1) {
max_depth = max_depth - 3;
return Solution[N];
}
int res1 = 3 * foo_memoized(N / 2, Solution, max_depth);
int res2 = foo_memoized(N - 1, Solution, max_depth);
if (res1 >= res2) {
max_depth = max_depth - 3;
Solution[N] = res1;
return res1;
} else {
max_depth = max_depth - 3;
Solution[N] = res2;
return res2;
}
}
/**************************************************MAIN FUNCTION*****************************************************************************/
int main() {
int N;
int result_rec;
int result_iter;
int result_wrap;
int max_depth=0;
while (1) {
printf("Enter N:");
scanf("%d", & N);
if (N == -1) {
break;
}
//call recursive function and print the result
int result_rec = foo(N);
printf("Result of Recursive function for N=%d is:%d\n\n",N,result_rec);
//call iterative function and print the result
int result_iter = foo_iterative(N);
printf("Result of Iterative function for N=%d is:%d\n\n", N,result_iter);
//call wrapper function and print the memoized
int result_wrap = foo_wrapper(N);
printf("Result of memoized function for N=%d is:%d\n\n",N,result_wrap);
}
}
| 1f6cc7e90270bd466f08f3dcb87b287efe4e1066 | [
"C"
] | 7 | C | Iam-El/Algorithm-and-Datastructures | 875719d13c473575a5dba3deb7f54af1f3472956 | 9af6411f607f8e4126d33ca8f7b1d389e3a61d09 |
refs/heads/master | <file_sep>'use strict';
const mongoose = require('mongoose');
mongoose.set('useFindAndModify', false);
const alertSchema = new mongoose.Schema({
search: { type: String, required: true },
email: { type: String, required: true },
time: { type: Number, required: true },
createdAt: { type: Date, default: Date.now }
});
const AlertModel = mongoose.model('alert', alertSchema);
module.exports = AlertModel;<file_sep>'use strict';
const VError = require('verror');
const AlertModel = require('./model');
/**
* Get all alerts
* @param {*} req
* @param {*} res
* @param {*} next
*/
async function getAlert(req, res, next) {
try {
const alerts = await AlertModel.find({}, {__v: 0 }).exec();
res.status(200).json(alerts);
} catch (err) {
next(new VError(err, 'Failed to get the alert'));
}
}
/**
* Create an alert
* @param {*} req
* @param {*} res
* @param {*} next **
*/
async function createAlert(req, res, next) {
try {
const payload = req.body;
const Alert = new AlertModel(payload);
const newAlert = await Alert.save();
return res.status(200).json(newAlert);
} catch (err) {
next(new VError(err, 'Failed to create alert'));
}
}
/**
* Get all alerts
* @param {*} req
* @param {*} res
* @param {*} next
*/
async function deleteAlert(req, res, next) {
try {
AlertModel.deleteOne({_id: req.body.id}).exec();
res.status(200).end();
} catch (err) {
next(new VError(err, 'Failed to get the alert'));
}
}
module.exports = {
getAlert,
createAlert,
deleteAlert
};
<file_sep>'use strict';
/**
* It returns decorated request object fot better logging
* @param {Express.Request} req
* @returns
*/
function getCommonRequestDetails(req) {
return {
method: req.method,
status: req.status,
headers: req.headers,
url: req.url,
body: req.body,
params: req.params,
query: req.query
};
}
module.exports = {
getCommonRequestDetails
};<file_sep>'use strict';
const { getCommonRequestDetails } = require('../utils');
/**
* Handles all errors from the requests
* @param {HttpException} err
* @param {Express.Request} req
* @param {Express.Response} res
* @param {Function} next
*/
// eslint-disable-next-line no-unused-vars
function errorHandler(err, req, res, next) {
const { args, name, status, message, stack } = err;
console.error(
{
request: getCommonRequestDetails(req),
error: {
name,
message,
args,
stack
}
},
`Failed to process the request to ${req.method} ${req.url}`
);
res.status(status || 500).json({
message
});
}
function loggingHandler(req, res, next) {
console.log(getCommonRequestDetails(req), `Received request to ${req.method} ${req.url}`);
next();
}
module.exports = {
errorHandler,
loggingHandler
};
<file_sep># RankMyApp challenge
## How to run locally
1. Clone this repository
`$ git clone https://github.com/PatrickRNG/RankMyApp_challenge`
2. Configure environment variables - [configurations](#Configurations)
3. If you already have Docker installed, run:
`$ docker-compose up --build`
4. Go to http://localhost:3001/
## Configurations
Create a ".env" file in the root of the /server folder. Inside set the below env variables. (For testing purposes there is already a .env.example in this repository (server folder), just copy it).
**The environment variables**
1. PORT -> Port for the web server
2. MONGO_URI -> Mongo URI for MongoDB configuration
3. CLIENT_ID -> ClientID for the smtp configuration
4. CLIENT_SECRET -> Secret of your Google Oauth
5. REDIRECT_URL -> Configuration for Google Oauth
6. REFRESH_TOKEN -> Configuration for Google Oauth
## TO DO / Improvements
1. More tests!
2. Better way to control every E-mail sent by each alarm
3. More environment variables configurations
**PS**
The ClientId, ClientSecret and other Google/Auth information in here are just for testing purposes, specificaly for this test/challenge. You should generate your own SMTP and Google Oauth info.
<file_sep>import { createContext } from 'react';
const AlertContext = createContext({});
export const AlertProvider = AlertContext.Provider;
export const AlertConsumer = AlertContext.Consumer;
export default AlertContext;<file_sep>import React from 'react';
import { Layout } from 'antd';
import styled from 'styled-components';
const { Header : AntHeader } = Layout;
const H1 = styled.h1`
color: #ccc;
`;
function Header() {
return (
<AntHeader>
<H1>Ebay Alerter</H1>
</AntHeader>
);
}
export default Header;
<file_sep>const API_URL = 'http://localhost:3000';
const ALERT_URL = `${API_URL}/alerts`;
/**
* GET alerts
*/
async function getAlerts() {
const alerts = await fetch(ALERT_URL);
return alerts.json();
}
/**
* POST create an alert
* @param {*} payload
*/
async function createAlert(payload) {
const newAlert = await fetch(ALERT_URL, {
method: 'post',
body: JSON.stringify(payload),
headers: { 'Content-Type': 'application/json' }
});
return newAlert.json();
}
/**
* DELETE alert
*/
async function deleteAlert(id) {
fetch(ALERT_URL, {
method: 'delete',
body: JSON.stringify(id),
headers: { 'Content-Type': 'application/json' }
});
}
/**
* GET Ebay products by keyword
* @param {*} keyword
*/
async function getEbayProductByKeyword(keyword) {
const ebayURL = `${API_URL}/products?search=${keyword}`;
const products = await fetch(ebayURL);
return products.json();
}
/**
* GET Send aE-mail
* @param {*} payload
*/
function sendEmail(payload) {
const emailURL = `${API_URL}/email`;
fetch(emailURL, {
method: 'post',
body: JSON.stringify(payload),
headers: { 'Content-Type': 'application/json' }
});
}
export {
getAlerts,
createAlert,
deleteAlert,
getEbayProductByKeyword,
sendEmail
};
<file_sep>'use strict';
const express = require('express');
const bodyParser = require('body-parser');
const cors = require('cors');
const helmet = require('helmet');
const VError = require('verror');
const HttpServer = require('./http-server');
const config = require('./config');
const { errorHandler, loggingHandler } = require('../middleware/handler');
const routes = require('../routes/index');
class Express {
constructor() {
this.app = express;
this.port = config.PORT;
}
/**
* Initialize express server
*/
async init() {
this.app = express();
this.app.use(cors());
this.app.use(helmet());
this.app.use(bodyParser.json());
this.app.use(loggingHandler);
this.app.use(routes);
this.app.use(errorHandler);
await this.registerHttpServer();
}
/**
* Register the http server
*/
async registerHttpServer() {
this.httpServer = new HttpServer(this.app, this.port);
this.httpServer.on('error', err => {
return new VError(err, 'Error to initialize the server');
});
this.httpServer.on('info', msg => {
console.log(msg);
});
this.httpServer.init();
}
}
module.exports = Express;<file_sep>/* eslint-disable no-undef */
const fetch = require('node-fetch');
const chai = require('chai');
const expect = chai.expect;
describe('Email API test', () => {
const EMAIL_API = 'http://localhost:3000/email';
it('should send an E-mail', async () => {
const emailPayload = {
from: '<EMAIL>',
to: '<EMAIL>',
subject: 'Ebay Alert',
text: 'Ebay test'
};
const response = await fetch(EMAIL_API, {
method: 'post',
body: JSON.stringify(emailPayload),
headers: { 'Content-Type': 'application/json' }
});
expect(response.status).to.equal(200);
});
});
<file_sep>PORT=3000
MONGO_URI=mongodb://mongo/alerts
CLIENT_ID=441362831599-1sliu0ecmo4rbc4m7a895o1ji6a63f69.apps.googleusercontent.com
CLIENT_SECRET=<KEY>
REDIRECT_URL=https://developers.google.com/oauthplayground
REFRESH_TOKEN=<KEY><file_sep>import React, { useState, useEffect } from 'react';
import { Layout } from 'antd';
import styled from 'styled-components';
import { AlertProvider } from '../contexts/AlertContext';
import {
getAlerts,
createAlert,
deleteAlert as deleteAlertApi,
getEbayProductByKeyword,
sendEmail
} from '../api';
import { Search, Header, Footer, Alerts } from '../components';
const { Content } = Layout;
const Paper = styled.div`
background: #fff;
padding: 24px;
min-height: 280;
`;
const StyledHeader = styled.h2`
margin-top: 30px;
`;
function App() {
const [alert, setAlert] = useState({
search: '',
email: '',
time: 10
});
const [alerts, setAlerts] = useState([]);
const [intervals, setIntervals] = useState([]);
/**
* Get all alerts
*/
async function getAllAlerts() {
const alerts = await getAlerts();
alerts.reverse();
setAlerts(alerts);
return alerts;
}
/**
* Create an alert
* @param {*} payload
*/
async function addAlert(payload) {
const newAlert = await createAlert(payload);
getAllAlerts();
return newAlert;
}
/**
* Delete an alert by id
* @param {*} id
*/
async function deleteAlert(id) {
deleteAlertApi({ id });
const filteredAlerts = alerts.filter(alert => alert._id !== id);
setAlerts(filteredAlerts);
}
/**
* Send the alert E-mail
* @param {Array} payload
*/
async function sendAlertEmail(payload) {
const products = await getEbayProductByKeyword(payload.search);
if (products.length) {
const lastProducts = getLastItems(products);
const sortedProducts = sortByPrice(lastProducts);
const text = createEmailBody(sortedProducts);
const emailPayload = {
from: '<EMAIL>',
to: payload.email,
subject: `Ebay Alert | ${payload.search}`,
text
};
sendEmail(emailPayload);
}
}
/**
* Create the email body
* @param {Array} products
*/
function createEmailBody(products) {
let email = '';
for (let product of products) {
console.log(product);
const price = product.sellingStatus[0].currentPrice[0];
email += `
Product: ${product.title[0]}
Price: ${price['@currencyId']} ${price.__value__}
----------
`;
}
return email;
}
/**
* Sort array by price
* @param {Array} products
*/
const sortByPrice = products =>
products.sort((a, b) => {
const price1 = a.sellingStatus[0].currentPrice[0].__value__;
const price2 = b.sellingStatus[0].currentPrice[0].__value__;
return price1 - price2;
});
/**
* Get the last 3 items from an array
* @param {Array} products
*/
const getLastItems = products => products.slice(0, 3);
/**
* Call a function with intervals for each item of an array
* @param {Array} arr
* @param {Function} fn
* @param {Number} time
*/
function interval(arr, fn, time) {
let intervalsArr = [];
for (const interval of intervals) {
clearInterval(interval);
}
for (const item of arr) {
const minutes = (item.time || time) * 60 * 1000;
const interval = setInterval(() => {
fn(item);
}, minutes);
intervalsArr.push(interval);
setIntervals(intervalsArr);
}
}
useEffect(() => {
getAllAlerts();
}, []);
useEffect(() => {
if (alerts) {
interval(alerts, sendAlertEmail);
// timeFunction();
}
}, [alerts]);
return (
<AlertProvider value={{ alert, setAlert, alerts }}>
<div className="App">
<Layout className="layout">
<Header />
<Content style={{ padding: '0 50px' }}>
<StyledHeader>Create an alert</StyledHeader>
<Paper>
<Search addAlert={addAlert} />
</Paper>
<Alerts deleteAlert={deleteAlert} />
</Content>
<Footer />
</Layout>
</div>
</AlertProvider>
);
}
export default App;
<file_sep>import React, { useContext } from 'react';
import PropTypes from 'prop-types';
import styled from 'styled-components';
import AlertContext from '../../contexts/AlertContext';
import { Card as AntCard } from 'antd';
const StyledHeader = styled.h2`
margin-top: 30px;
`;
const Wrapper = styled.div`
display: flex;
flex-wrap: wrap;
`;
const Card = styled(AntCard)`
position: relative;
width: 350px;
margin: 6px;
& .ant-card-body {
display: flex;
justify-content: space-between;
}
`;
const Left = styled.div`
word-break: break-word;
width: 75%;
`;
const P = styled.p`
margin: 0;
`;
const Time = styled.div`
font-size: 20px;
font-weight: 500;
text-align: center;
`;
const Minute = styled.div`
font-size: 14px;
`;
const CloseButton = styled.span`
position: absolute;
right: 13px;
top: 6px;
font-size: 18px;
cursor: pointer;
color: #aaa;
&:hover {
color: #888;
}
`;
function Alerts({deleteAlert}) {
const { alerts } = useContext(AlertContext);
return (
<>
<StyledHeader>Alerts</StyledHeader>
<Wrapper>
{alerts.map((alert, i) => (
<Card key={i}>
<Left>
<b>{alert.search}</b>
<P>{alert.email}</P>
</Left>
<div>
<Time>
{alert.time}
<br />
<Minute>minutes</Minute>
</Time>
</div>
<CloseButton onClick={() => deleteAlert(alert._id)}>×</CloseButton>
</Card>
))}
</Wrapper>
</>
);
}
Alerts.propTypes = {
deleteAlert: PropTypes.func
};
export default Alerts;
<file_sep>'use strict';
const VError = require('verror');
const fetch = require('node-fetch');
const EBAY_URL =
'https://svcs.ebay.com/services/search/FindingService/v1?OPERATION-NAME=findItemsByKeywords&SERVICE-VERSION=1.0.0&SECURITY-APPNAME=PatrickP-RankMyAp-PRD-33882c1d0-3c03000f&RESPONSE-DATA-FORMAT=JSON&REST-PAYLOAD';
/**
* Get Ebay products by keywords
* @param {*} req
* @param {*} res
* @param {*} next
*/
async function getEbayProductByKeyword(req, res, next) {
try {
const ebayURL = `${EBAY_URL}&keywords=${req.query.search}`;
const response = await fetch(ebayURL);
const jsonResponse = await response.json();
const products = jsonResponse.findItemsByKeywordsResponse[0].searchResult[0].item;
res.status(200).json(products);
} catch (err) {
next(new VError(err, 'Failed to get the products'));
}
}
module.exports = {
getEbayProductByKeyword
};
<file_sep>'use strict';
const VError = require('verror');
const Email = require('../../infra/email');
/**
* Send an E-mail
* @param {*} req
* @param {*} res
* @param {*} next
*/
async function sendEmail(req, res, next) {
try {
const email = new Email('<EMAIL>');
const transport = email.configEmail();
const { from, to, subject, text } = req.body;
const mailOptions = {
from, // Sender address
to, // List of recipients
subject, // Subject line
text // Plain text body
};
transport.sendMail(mailOptions, function(err, response) {
err ? console.log(err) : console.log(response);
transport.close();
});
res.status(200).end();
} catch (err) {
next(new VError(err, 'Failed to send the E-mail'));
}
}
module.exports = {
sendEmail
};
<file_sep>import Header from './Header';
import Footer from './Footer';
import Search from './Search';
import Alerts from './Alerts';
export {
Header,
Footer,
Search,
Alerts
};<file_sep>const fetch = require('node-fetch')
const chai = require('chai');
const expect = chai.expect;
describe('Ebay API test', () => {
const EBAY_API = 'http://localhost:3000/products';
it('should return the ebay products correctly', async () => {
const response = await fetch(EBAY_API);
const products = await response.json();
expect(response.status).to.equal(200);
expect(products).to.be.a('array');
});
});<file_sep>'use strict';
const Express = require('./infra/express');
const Database = require('./infra/database');
const config = require('./infra/config');
const express = new Express();
const mongoOpts = {
useCreateIndex: true,
useUnifiedTopology: true
};
const database = new Database(config.MONGO_URI, mongoOpts);
database.init();
database.on('info', msg => console.log(msg));
database.on('error', msg => console.log(msg));
express.init();<file_sep>'use strict';
const { Router } = require('express');
const alertController = require('../api/alert/controller');
const ebayController = require('../api/ebay/controller');
const emailController = require('../api/email/controller');
const router = Router();
router.get('/alerts', alertController.getAlert);
router.post('/alerts', alertController.createAlert);
router.delete('/alerts', alertController.deleteAlert);
router.get('/products', ebayController.getEbayProductByKeyword);
router.post('/email', emailController.sendEmail);
module.exports = router;<file_sep>'use strict';
const nodemailer = require('nodemailer');
const { google } = require('googleapis');
const VError = require('verror');
const config = require('./config');
class Email {
constructor(userEmail) {
this.userEmail = userEmail;
this.clientId = config.CLIENT_ID;
this.clientSecret = config.CLIENT_SECRET;
this.redirectURL = config.REDIRECT_URL;
this.refreshToken = config.REFRESH_TOKEN;
}
/**
* Configure nodemailer with smtp
*/
configEmail() {
try {
const OAuth2 = google.auth.OAuth2;
const oauth2Client = new OAuth2(
this.clientId,
this.clientSecret,
this.redirectURL
);
oauth2Client.setCredentials({
refresh_token: this.refreshToken
});
const accessToken = oauth2Client.getAccessToken();
const transport = this.creteSmtpTransport(accessToken);
return transport;
} catch (err) {
throw new VError(err, 'Failed to configure the E-mail');
}
}
creteSmtpTransport(accessToken) {
try {
return nodemailer.createTransport({
service: 'gmail',
auth: {
type: 'OAuth2',
user: this.userEmail,
clientId: this.clientId,
clientSecret: this.clientSecret,
refreshToken: this.refreshToken,
accessToken
}
});
} catch(err) {
throw new VError(err, 'Failed to configure smtp');
}
}
}
module.exports = Email;
| 93c505fe709b53dd6ffbaa929651903596a9832f | [
"JavaScript",
"Markdown",
"Shell"
] | 20 | JavaScript | PatrickRNG/RankMyApp_challenge | 066a090c1b0031a32eb761dbcdd43ad92670ee35 | 5f761e74053a14c5c1475f97d695005b292863b8 |
refs/heads/master | <repo_name>gfall94/PyTrinamicMicro<file_sep>/PyTrinamicMicro/platforms/motionpy/examples/modules/max/max14912pmb.py
'''
Example using the MAX14912PMB.
This script switches all the outputs to high and then back to low.
Created on 15.02.2021
@author: JH
'''
from pyb import Pin
from PyTrinamicMicro.platforms.motionpy.modules.max.max14912 import MAX14912
import time
import struct
import logging
logger = logging.getLogger(__name__)
logger.info("MAX14912PMB example running")
module = MAX14912()
while(True):
logger.info("Switching everything to HIGH")
for y in range(0, 8):
module.set_output(y,1)
time.sleep(0.5)
logger.info("Switching everything to LOW")
for y in range(0, 8):
module.set_output(y,0)
time.sleep(0.5)
<file_sep>/PyTrinamicMicro/platforms/motionpy/examples/modules/max/max14914pmb_output.py
'''
Example using the MAX14914PMB.
This scripts toggles output of MAX14914.
Created on 5.03.2021
@author: JH
'''
from pyb import Pin
from PyTrinamicMicro.platforms.motionpy.modules.max.max14914 import MAX14914
import time
import logging
logger = logging.getLogger(__name__)
logger.info("MAX14914PMB Output example running")
module = MAX14914()
module.setIOMode(0)
description = """\nThis scripts toggles output of MAX14914.\n """
print(description)
while(True):
state = 0
module.setDO(state)
faults = module.getFault()
OV_VDD = module.getOV_VDD()
for cursor in '|/-\\':
text = cursor+" Output state: " + str(state) + ";Fault state: " + str(faults) +"; OV_VDD: " + str(OV_VDD)
print(text, end='\r')
time.sleep(0.2)
state = 1
module.setDO(state)
faults = module.getFault()
OV_VDD = module.getOV_VDD()
for cursor in '|/-\\':
text = cursor+" Output state: " + str(state) + ";Fault state: " + str(faults) +"; OV_VDD: " + str(OV_VDD)
print(text, end='\r')
time.sleep(0.2)
<file_sep>/PyTrinamicMicro/platforms/motionpy/examples/modules/max/max22190pmb.py
'''
This file implements an example for using the MAX22190PMB in SPI Mode 1.
It displays the input read and wire break readout in the terminal.
For further details on MAX22190PMB refer to the data sheet.
Created on 25.02.2021
@author: JH
'''
from pyb import Pin
from PyTrinamicMicro.platforms.motionpy.modules.max.max22190 import MAX22190
import time
import logging
logger = logging.getLogger(__name__)
logger.info("MAX22190PMB example running")
module = MAX22190()
description = """\nThis scripts displays the digital channel inputs as well as the wire break detection states.
Channel input => 0: Channel is driven low; 1: Channel is driven high.
Wire break detection=> 0: wire break condition detected; 1: no wire break condition detected.\n """
print(description)
legend = "Channel nr.: " + "12345678"+ "; Channel nr.: " + "12345678"
print(legend)
while(True):
for cursor in '|/-\\':
input_states = module.get_digital_input_states()
wire_breaks = module.get_wire_break_states()
text = cursor+" IO states: " + ''.join(str(e) for e in input_states)+ "; Wire Break states: " + ''.join(str(e) for e in wire_breaks)
print(text, end='\r')
time.sleep(0.1)
<file_sep>/PyTrinamicMicro/platforms/motionpy/examples/modules/max/max14914pmb_input.py
'''
Example using the MAX14914PMB.
This scripts displays the readout of the max14914 in digital input mode.
Created on 5.03.2021
@author: JH
'''
from pyb import Pin
from PyTrinamicMicro.platforms.motionpy.modules.max.max14914 import MAX14914
import time
import logging
logger = logging.getLogger(__name__)
logger.info("MAX14914PMB Input example running")
module = MAX14914()
module.setIOMode(1)
description = """\nThis scripts displays the readout of the max14914 in digital input mode.\n """
print(description)
while(True):
for cursor in '|/-\\':
state = module.getDIDO_LVL()
faults = module.getFault()
OV_VDD = module.getOV_VDD()
text = cursor+" Input state: " + str(state) + ";Fault state: " + str(faults) +"; OV_VDD: " + str(OV_VDD)
print(text, end='\r')
time.sleep(0.1)
4 | 58ce54411518a1ec8f540e6e3a93508a8109717d | [
"Python"
] | 4 | Python | gfall94/PyTrinamicMicro | 62da5c980c9f3b2ee75b37b0de0f843f6f9b0a68 | 0e89464d957f4cd9311484b488b995dfece65b9f |
refs/heads/master | <file_sep>package org.example;
import com.google.protobuf.ByteString;
import io.grpc.*;
import java.io.File;
import java.io.FileInputStream;
import com.google.flatbuffers.FlatBufferBuilder;
import com.google.protobuf.LiteralByteString;
import io.grpc.stub.StreamObserver;
import org.example.ProtoFileTransfer.TransferMsgProto;
import org.example.ProtoFileTransfer.TransferReplyProto;
import org.example.TransferMsg;
import org.example.TransferReply;
public class Client {
private FileTransferGrpc.FileTransferStub asyncStubFlat;
private ProtobufServiceGrpc.ProtobufServiceStub asyncStubProto;
final long[] recv = new long[1];
int partId = 0;
public Client(Channel channel){
asyncStubFlat = FileTransferGrpc.newStub(channel);
asyncStubProto = ProtobufServiceGrpc.newStub(channel);
recv[0] = 0;
}
public void execFlatClient() throws Exception{
System.out.println("Starting streaming with Flatbuffers");
StreamObserver<TransferMsg> requestObserver = asyncStubFlat.sendData(new StreamObserver<TransferReply>(){
@Override
public void onNext(TransferReply msg) {
recv[0]++;
}
@Override
public void onError(Throwable t) {
Status status = Status.fromThrowable(t);
System.out.println(status);
System.out.println("Finished streaming with errors");
}
@Override
public void onCompleted() {
System.out.println("Finished streaming");
}
});
try{
int i = 0;
byte[] chunk = new byte[64 * 1024];
int chunkLen = 0;
while(i < 1000) {
File file = new File("./bigtext.txt");
FileInputStream is = new FileInputStream(file);
while ((chunkLen = is.read(chunk)) != -1) {
partId++;
FlatBufferBuilder builder = new FlatBufferBuilder();
int dataOff = TransferMsg.createDataVector(builder, chunk);
int off = TransferMsg.createTransferMsg(builder, partId, dataOff);
builder.finish(off);
TransferMsg msg = TransferMsg.getRootAsTransferMsg(builder.dataBuffer());
requestObserver.onNext(msg);
}
i++;
}
} catch (Exception e){
System.out.println(e);
}
requestObserver.onCompleted();
Thread.sleep(1000*100);
if(recv[0] == partId){
System.out.println("Transfer Successfull....");
} else{
System.out.println("Some error occurred...");
}
}
public void execProto() throws Exception{
System.out.println("Starting streaming with Protobuffers");
StreamObserver<TransferMsgProto> requestObserver = asyncStubProto.sendData(new StreamObserver<TransferReplyProto>(){
@Override
public void onNext(TransferReplyProto msg) {
recv[0]++;
}
@Override
public void onError(Throwable t) {
Status status = Status.fromThrowable(t);
System.out.println(status);
System.out.println("Finished streaming with errors");
}
@Override
public void onCompleted() {
System.out.println("Finished streaming");
}
});
try{
int i = 0;
byte[] chunk = new byte[64 * 1024];
int chunkLen = 0;
while(i < 1000) {
File file = new File("../bigtext.txt");
FileInputStream is = new FileInputStream(file);
while ((chunkLen = is.read(chunk)) != -1) {
partId++;
TransferMsgProto msg = TransferMsgProto.newBuilder().setPartId(partId).setData(ByteString.copyFrom(chunk)).build();
requestObserver.onNext(msg);
}
i++;
}
} catch (Exception e){
System.out.println(e);
}
requestObserver.onCompleted();
Thread.sleep(1000*100);
if(recv[0] == partId){
System.out.println("Transfer Successfull....");
} else{
System.out.println("Some error occurred...");
}
}
public static void main(String[] args) throws Exception{
String target = "localhost:50051";
ManagedChannel channel = ManagedChannelBuilder.forTarget(target).usePlaintext().build();
Client c = new Client(channel);
if(args.length == 0){
c.execFlatClient();
} else{
c.execProto();
}
}
}
// Refractor Required....<file_sep><?xml version="1.0" encoding="UTF-8"?>
<project xmlns="http://maven.apache.org/POM/4.0.0"
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
xsi:schemaLocation="http://maven.apache.org/POM/4.0.0 http://maven.apache.org/xsd/maven-4.0.0.xsd">
<modelVersion>4.0.0</modelVersion>
<groupId>org.example</groupId>
<artifactId>grpc-flatbuffers</artifactId>
<version>1.0-SNAPSHOT</version>
<properties>
<maven-compiler-plugin.version>3.7.0</maven-compiler-plugin.version>
<javac.version>1.8</javac.version>
<protobuf-maven-plugin.version>0.5.1</protobuf-maven-plugin.version>
</properties>
<dependencies>
<dependency>
<groupId>javax.annotation</groupId>
<artifactId>javax.annotation-api</artifactId>
<version>1.3.2</version>
</dependency>
<dependency>
<groupId>com.google.flatbuffers</groupId>
<artifactId>flatbuffers-java</artifactId>
<version>1.11.0</version>
</dependency>
<dependency>
<groupId>com.google.flatbuffers</groupId>
<artifactId>flatbuffers-java-grpc</artifactId>
<version>1.11.0</version>
</dependency>
<dependency>
<groupId>io.grpc</groupId>
<artifactId>grpc-netty</artifactId>
<version>1.29.0</version>
</dependency>
<dependency>
<groupId>io.grpc</groupId>
<artifactId>grpc-protobuf</artifactId>
<version>1.29.0</version>
</dependency>
<dependency>
<groupId>io.grpc</groupId>
<artifactId>grpc-stub</artifactId>
<version>1.29.0</version>
</dependency>
<dependency>
<groupId>io.grpc</groupId>
<artifactId>grpc-api</artifactId>
<version>1.29.0</version>
</dependency>
</dependencies>
<build>
<extensions>
<extension>
<groupId>kr.motd.maven</groupId>
<artifactId>os-maven-plugin</artifactId>
<version>1.5.0.Final</version>
</extension>
</extensions>
<plugins>
<plugin>
<groupId>org.apache.maven.plugins</groupId>
<artifactId>maven-compiler-plugin</artifactId>
<version>${maven-compiler-plugin.version}</version>
<configuration>
<source>${javac.version}</source>
<target>${javac.version}</target>
<fork>true</fork>
<meminitial>512m</meminitial>
<maxmem>2048m</maxmem>
<showDeprecation>false</showDeprecation>
<useIncrementalCompilation>false</useIncrementalCompilation>
<compilerArgs>
<arg>-Xlint:all,-options,-path</arg>
</compilerArgs>
</configuration>
</plugin>
<plugin>
<groupId>org.xolstice.maven.plugins</groupId>
<artifactId>protobuf-maven-plugin</artifactId>
<version>${protobuf-maven-plugin.version}</version>
<configuration>
<protocArtifact>com.google.protobuf:protoc:3.4.0:exe:${os.detected.classifier}</protocArtifact>
<pluginId>grpc-java</pluginId>
<pluginArtifact>io.grpc:protoc-gen-grpc-java:1.7.0:exe:${os.detected.classifier}</pluginArtifact>
</configuration>
<executions>
<execution>
<goals>
<goal>compile</goal>
<goal>compile-custom</goal>
</goals>
</execution>
</executions>
</plugin>
</plugins>
</build>
</project><file_sep>//Generated by flatc compiler (version 1.11.0)
//If you make any local changes, they will be lost
//source: FileTransfer.fbs
package org.example;
import com.google.flatbuffers.grpc.FlatbuffersUtils;
import java.nio.ByteBuffer;
import static io.grpc.MethodDescriptor.generateFullMethodName;
import static io.grpc.stub.ClientCalls.asyncBidiStreamingCall;
import static io.grpc.stub.ClientCalls.asyncClientStreamingCall;
import static io.grpc.stub.ClientCalls.asyncServerStreamingCall;
import static io.grpc.stub.ClientCalls.asyncUnaryCall;
import static io.grpc.stub.ClientCalls.blockingServerStreamingCall;
import static io.grpc.stub.ClientCalls.blockingUnaryCall;
import static io.grpc.stub.ClientCalls.futureUnaryCall;
import static io.grpc.stub.ServerCalls.asyncBidiStreamingCall;
import static io.grpc.stub.ServerCalls.asyncClientStreamingCall;
import static io.grpc.stub.ServerCalls.asyncServerStreamingCall;
import static io.grpc.stub.ServerCalls.asyncUnaryCall;
import static io.grpc.stub.ServerCalls.asyncUnimplementedStreamingCall;
import static io.grpc.stub.ServerCalls.asyncUnimplementedUnaryCall;
/**
*/
@javax.annotation.Generated(
value = "by gRPC proto compiler",
comments = "Source: FileTransfer.fbs")
public final class FileTransferGrpc {
private FileTransferGrpc() {}
public static final String SERVICE_NAME = "org.example.FileTransfer";
// Static method descriptors that strictly reflect the proto.
@io.grpc.ExperimentalApi("https://github.com/grpc/grpc-java/issues/1901")
@java.lang.Deprecated // Use {@link #getSendDataMethod()} instead.
public static final io.grpc.MethodDescriptor<org.example.TransferMsg,
org.example.TransferReply> METHOD_SEND_DATA = getSendDataMethod();
private static volatile io.grpc.MethodDescriptor<org.example.TransferMsg,
org.example.TransferReply> getSendDataMethod;
private static volatile FlatbuffersUtils.FBExtactor<org.example.TransferMsg> extractorOfTransferMsg;
private static FlatbuffersUtils.FBExtactor<org.example.TransferMsg> getExtractorOfTransferMsg() {
if (extractorOfTransferMsg != null) return extractorOfTransferMsg;
synchronized (FileTransferGrpc.class) {
if (extractorOfTransferMsg != null) return extractorOfTransferMsg;
extractorOfTransferMsg = new FlatbuffersUtils.FBExtactor<org.example.TransferMsg>() {
public org.example.TransferMsg extract (ByteBuffer buffer) {
return org.example.TransferMsg.getRootAsTransferMsg(buffer);
}
};
return extractorOfTransferMsg;
}
}
private static volatile FlatbuffersUtils.FBExtactor<org.example.TransferReply> extractorOfTransferReply;
private static FlatbuffersUtils.FBExtactor<org.example.TransferReply> getExtractorOfTransferReply() {
if (extractorOfTransferReply != null) return extractorOfTransferReply;
synchronized (FileTransferGrpc.class) {
if (extractorOfTransferReply != null) return extractorOfTransferReply;
extractorOfTransferReply = new FlatbuffersUtils.FBExtactor<org.example.TransferReply>() {
public org.example.TransferReply extract (ByteBuffer buffer) {
return org.example.TransferReply.getRootAsTransferReply(buffer);
}
};
return extractorOfTransferReply;
}
}
@io.grpc.ExperimentalApi("https://github.com/grpc/grpc-java/issues/1901")
public static io.grpc.MethodDescriptor<org.example.TransferMsg,
org.example.TransferReply> getSendDataMethod() {
io.grpc.MethodDescriptor<org.example.TransferMsg, org.example.TransferReply> getSendDataMethod;
if ((getSendDataMethod = FileTransferGrpc.getSendDataMethod) == null) {
synchronized (FileTransferGrpc.class) {
if ((getSendDataMethod = FileTransferGrpc.getSendDataMethod) == null) {
FileTransferGrpc.getSendDataMethod = getSendDataMethod =
io.grpc.MethodDescriptor.<org.example.TransferMsg, org.example.TransferReply>newBuilder()
.setType(io.grpc.MethodDescriptor.MethodType.BIDI_STREAMING)
.setFullMethodName(generateFullMethodName(
"org.example.FileTransfer", "SendData"))
.setSampledToLocalTracing(true)
.setRequestMarshaller(FlatbuffersUtils.marshaller(
org.example.TransferMsg.class, getExtractorOfTransferMsg()))
.setResponseMarshaller(FlatbuffersUtils.marshaller(
org.example.TransferReply.class, getExtractorOfTransferReply()))
.setSchemaDescriptor(null)
.build();
}
}
}
return getSendDataMethod;
}
/**
* Creates a new async stub that supports all call types for the service
*/
public static FileTransferStub newStub(io.grpc.Channel channel) {
return new FileTransferStub(channel);
}
/**
* Creates a new blocking-style stub that supports unary and streaming output calls on the service
*/
public static FileTransferBlockingStub newBlockingStub(
io.grpc.Channel channel) {
return new FileTransferBlockingStub(channel);
}
/**
* Creates a new ListenableFuture-style stub that supports unary calls on the service
*/
public static FileTransferFutureStub newFutureStub(
io.grpc.Channel channel) {
return new FileTransferFutureStub(channel);
}
/**
*/
public static abstract class FileTransferImplBase implements io.grpc.BindableService {
/**
*/
public io.grpc.stub.StreamObserver<org.example.TransferMsg> sendData(
io.grpc.stub.StreamObserver<org.example.TransferReply> responseObserver) {
return asyncUnimplementedStreamingCall(getSendDataMethod(), responseObserver);
}
@java.lang.Override public final io.grpc.ServerServiceDefinition bindService() {
return io.grpc.ServerServiceDefinition.builder(getServiceDescriptor())
.addMethod(
getSendDataMethod(),
asyncBidiStreamingCall(
new MethodHandlers<
org.example.TransferMsg,
org.example.TransferReply>(
this, METHODID_SEND_DATA)))
.build();
}
}
/**
*/
public static final class FileTransferStub extends io.grpc.stub.AbstractStub<FileTransferStub> {
private FileTransferStub(io.grpc.Channel channel) {
super(channel);
}
private FileTransferStub(io.grpc.Channel channel,
io.grpc.CallOptions callOptions) {
super(channel, callOptions);
}
@java.lang.Override
protected FileTransferStub build(io.grpc.Channel channel,
io.grpc.CallOptions callOptions) {
return new FileTransferStub(channel, callOptions);
}
/**
*/
public io.grpc.stub.StreamObserver<org.example.TransferMsg> sendData(
io.grpc.stub.StreamObserver<org.example.TransferReply> responseObserver) {
return asyncBidiStreamingCall(
getChannel().newCall(getSendDataMethod(), getCallOptions()), responseObserver);
}
}
/**
*/
public static final class FileTransferBlockingStub extends io.grpc.stub.AbstractStub<FileTransferBlockingStub> {
private FileTransferBlockingStub(io.grpc.Channel channel) {
super(channel);
}
private FileTransferBlockingStub(io.grpc.Channel channel,
io.grpc.CallOptions callOptions) {
super(channel, callOptions);
}
@java.lang.Override
protected FileTransferBlockingStub build(io.grpc.Channel channel,
io.grpc.CallOptions callOptions) {
return new FileTransferBlockingStub(channel, callOptions);
}
}
/**
*/
public static final class FileTransferFutureStub extends io.grpc.stub.AbstractStub<FileTransferFutureStub> {
private FileTransferFutureStub(io.grpc.Channel channel) {
super(channel);
}
private FileTransferFutureStub(io.grpc.Channel channel,
io.grpc.CallOptions callOptions) {
super(channel, callOptions);
}
@java.lang.Override
protected FileTransferFutureStub build(io.grpc.Channel channel,
io.grpc.CallOptions callOptions) {
return new FileTransferFutureStub(channel, callOptions);
}
}
private static final int METHODID_SEND_DATA = 0;
private static final class MethodHandlers<Req, Resp> implements
io.grpc.stub.ServerCalls.UnaryMethod<Req, Resp>,
io.grpc.stub.ServerCalls.ServerStreamingMethod<Req, Resp>,
io.grpc.stub.ServerCalls.ClientStreamingMethod<Req, Resp>,
io.grpc.stub.ServerCalls.BidiStreamingMethod<Req, Resp> {
private final FileTransferImplBase serviceImpl;
private final int methodId;
MethodHandlers(FileTransferImplBase serviceImpl, int methodId) {
this.serviceImpl = serviceImpl;
this.methodId = methodId;
}
@java.lang.Override
@java.lang.SuppressWarnings("unchecked")
public void invoke(Req request, io.grpc.stub.StreamObserver<Resp> responseObserver) {
switch (methodId) {
default:
throw new AssertionError();
}
}
@java.lang.Override
@java.lang.SuppressWarnings("unchecked")
public io.grpc.stub.StreamObserver<Req> invoke(
io.grpc.stub.StreamObserver<Resp> responseObserver) {
switch (methodId) {
case METHODID_SEND_DATA:
return (io.grpc.stub.StreamObserver<Req>) serviceImpl.sendData(
(io.grpc.stub.StreamObserver<org.example.TransferReply>) responseObserver);
default:
throw new AssertionError();
}
}
}
private static volatile io.grpc.ServiceDescriptor serviceDescriptor;
public static io.grpc.ServiceDescriptor getServiceDescriptor() {
io.grpc.ServiceDescriptor result = serviceDescriptor;
if (result == null) {
synchronized (FileTransferGrpc.class) {
result = serviceDescriptor;
if (result == null) {
serviceDescriptor = result = io.grpc.ServiceDescriptor.newBuilder(SERVICE_NAME)
.setSchemaDescriptor(null)
.addMethod(getSendDataMethod())
.build();
}
}
}
return result;
}
}
| ce74d7a23caa1a6766246249c7d49f98cad2a77c | [
"Java",
"Maven POM"
] | 3 | Java | anshkhannasbu/grpc-flatbuffers | 1c72356490b7959c0e423b2a5ec05a292f7a902c | 7b829c04005e7c3e569c9d0ee9ccd3e26c080209 |
refs/heads/master | <file_sep>它们 ByteBuffer 分配内存的两种方式。HeapByteBuffer 顾名思义其内存空间在 JVM 的 heap(堆)上分配,可以看做是 jdk 对于 byte[] 数组的封装;
而 DirectByteBuffer 则直接利用了系统接口进行内存申请,其内存分配在c heap 中,这样就减少了内存之间的拷贝操作,如此一来,在使用 DirectByteBuffer 时,
系统就可以直接从内存将数据写入到 Channel 中,而无需进行 Java 堆的内存申请,复制等操作,提高了性能。既然如此,为什么不直接使用 DirectByteBuffer,
还要来个 HeapByteBuffer?原因在于, DirectByteBuffer 是通过full gc来回收内存的,DirectByteBuffer会自己检测情况而调用 system.gc(),
但是如果参数中使用了 DisableExplicitGC 那么就无法回收该快内存了,-XX:+DisableExplicitGC标志自动将 System.gc() 调用转换成一个空操作,
就是应用中调用 System.gc() 会变成一个空操作,那么如果设置了就需要我们手动来回收内存了,所以DirectByteBuffer使用起来相对于完全托管于 java 内存管理的Heap ByteBuffer 来说更复杂一些,
如果用不好可能会引起OOM。Direct ByteBuffer 的内存大小受 -XX:MaxDirectMemorySize JVM 参数控制(默认大小64M),
在 DirectByteBuffer 申请内存空间达到该设置大小后,会触发 Full GC。
<file_sep>package test;
import org.apache.hadoop.conf.Configuration;
import org.apache.hadoop.ipc.RPC;
import proto.DfsService;
import proto.DfsServiceImpl;
import java.io.IOException;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/6/28 22:39
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public class DfsServiceServer {
public static void main(String[] args) throws IOException {
DfsServiceImpl dfsService = new DfsServiceImpl();
// 创建一个RPC builder
RPC.Builder builder = new RPC.Builder(new Configuration());
//指定RPC Server的参数
builder.setBindAddress("localhost");
builder.setPort(7788);
//将自己的程序部署到server上
builder.setProtocol(DfsService.class);
builder.setInstance(dfsService);
//创建Server
RPC.Server server = builder.build();
//启动服务
server.start();
System.out.println("server is start");
}
}
<file_sep># 序列化
## 1. 为什么需要序列化
1. 网络传输的数据必须是二进制数据
2. java中的对象(网络中交互数据), 是不能直接在网络中传输
所以我们需要将需要在网络传输的对象转化为可传输的二进制, 并且这种转换算法是可逆的
**java中的序列化** 就是将一个Java对象 **"转换"** 为一串连续的bytes字节(字节数组或流),以便写入文件或通过Socket发送出去, 这一串字节包括了对象的数据(成员变量), 以及对象及其成员变量的类型信息(元信息)等,
**java的反序列化** 正好相反可以将从文件或Socket中读取到的这一串字节还原为一个几乎一模一样的Java对象

**总结来说**, 序列化就是将对象转换成二进制数据的过程,而反序列就是反过来将二进制转换
为对象的过程
## 2. 序列化在RPC中的应用
### 2.1 RPC通信流程图

不妨借用个例子帮助你理解,比如发快递,我们要发一个需要自行组装的物件。发件人发之
前,会把物件拆开装箱,这就好比序列化;这时候快递员来了,不能磕碰呀,那就要打包,
这就好比将序列化后的数据进行编码,封装成一个固定格式的协议;过了两天,收件人收到
包裹了,就会拆箱将物件拼接好,这就好比是协议解码和反序列化。
## 3. 有哪些常用的序列化框架
- JDK 原生序列化
- JSON
- Protobuf
- Thrift
- Hessian
- Kryo
> 为方便集成多种序列化, 实现多种序列化组件的插拔和替换,将序列化及反序列化抽象为[service.ObjectSerializer](src/main/java/service/ObjectSerializer.java)接口
### 3.1 JDK 原生序列化
JDK 自带的序列化机制对使用者而言是非常简单的。序列化具体的实现是由 **ObjectOutputStream** 完成的,而反序列化的具体实现是由 **ObjectInputStream** 完成的
```java
@Data
public class Student implements Serializable {
/**
* serialVersionUID
* 反序列化时,会根据serialVersionUID这个常量值来判断反序列得到的类信息是否与原来一致
* 如果我们不指定serialVersionUID值,序列化将会把当前类的hashCode值赋给它
*
*/
private static final long serialVersionUID = 3L;
private int no;
private String name;
public static void main(String[] args) throws IOException, ClassNotFoundExc
String home = System.getProperty("user.home");
String basePath = home + "/Desktop";
FileOutputStream fos = new FileOutputStream(basePath + "student.dat");
Student student = new Student();
student.setNo(100);
student.setName("TEST_STUDENT");
ObjectOutputStream oos = new ObjectOutputStream(fos);
oos.writeObject(student);
oos.flush();
oos.close();
FileInputStream fis = new FileInputStream(basePath + "student.dat");
ObjectInputStream ois = new ObjectInputStream(fis);
Student deStudent = (Student) ois.readObject();
ois.close();
System.out.println(deStudent);
}
}
```
**JDK序列化的过程:**

序列化过程就是在读取对象数据的时候,不断加入一些特殊分隔符,这些特殊分隔符用于在反序列化过程中截断用。
- 头部数据用来声明序列化协议、序列化版本,用于高低版本向后兼容
- 对象数据主要包括类名、签名、属性名、属性类型及属性值,当然还有开头结尾等数据,除了属性值属于真正的对象值,其他都是为了反序列化用的元数据
- 存在对象引用、继承的情况下,就是递归遍历“写对象”逻辑
**封装JDK序列化实现代码:**
[JdkSerialaizerImpl](./src/main/java/impl/JdkSerialaizerImpl.java)
**测试**

### 3.2 JSON
JSON 是典型的 Key-Value 方式,没有数据类型,是一种文本型序列化框架
JSON使用广泛,无论WEB,存储,还是基于HTTP协议的RPC框架通信, 都会选择JSON格式
使用JSON进行序列化有两个问题需要注意:
- JSON 进行序列化的额外空间开销比较大,对于大数据量服务这意味着需要巨大的内存和磁盘开销;
- JSON 没有类型,但像 Java 这种强类型语言,需要通过反射统一解决,所以性能不会太好。
**所以如果 RPC 框架选用 JSON 序列化,服务提供者与服务调用者之间传输的数据量要相对
较小,否则将严重影响性能。**
```java
public byte[] serialize(Object obj) throws IOException {
return JSON.toJSONBytes(obj);
}
public <T> T deserialize(byte[] bytes, Class<T> clazz) throws IOException, ClassNotFoundException {
return JSON.parseObject(bytes, clazz);
}
```
### 3.3 Hessian序列化
Hessian 是动态类型、二进制、紧凑的,并且**可跨语言移植**的一种序列化框架
Hessian序列化是Hessian组件中的一个模块, 而Hessian组件是基于HTTP协议的一个跨语言的轻量级,二进制Web服务解决方案。Hessian 协议要比 JDK、JSON 更加紧凑,性能上要比 JDK、JSON 序列化高效很多,而且生成的字节数也更小。
相对于 JDK、JSON,由于 Hessian 更加高效,生成的字节数更小,有非常好的兼容性和稳定性,所以 Hessian 更加适合作为 RPC 框架远程通信的序列化协议。
**缺点:**
- 对序列化支持的不够全面, 在特殊情况下序列化会失败
- 反序列化得到的新对象可能与原来的对象存在细微差异
官方版本对Java里面的一些常见对象的类型不支持如:
Linked 系列,LinkedHashMap、LinkedHashSet 等,但是可以通过扩展CollectionDeserializer 类修复;
Locale 类,可以通过扩展 ContextSerializerFactory 类修复;
Byte/Short 反序列化的时候变成 Integer。
### 3.4 protobuf
Protobuf 是 Google 公司内部的混合语言数据标准,是一种轻便、高效的结构化数据存储
格式,可以用于结构化数据序列化,支持 Java、Python、C++、Go 等语言。Protobuf
使用的时候需要定义 IDL(Interface description language),然后使用不同语言的 IDL
编译器,生成序列化工具类,它的优点是:
- 序列化后体积相比 JSON、Hessian 小很多;
- IDL 能清晰地描述语义,所以足以帮助并保证应用程序之间的类型不会丢失,无需类似XML 解析器;
- 序列化反序列化速度很快,不需要通过反射获取类型;
- 消息格式升级和兼容性不错,可以做到向后兼容。
缺点:
- 要为传输的对象编写专门的结构化文件并编译
Protobuf 非常高效,但是对于具有反射和动态能力的语言来说,这样用起来很费劲,这一
点就不如 Hessian,比如用 Java 的话,这个预编译过程不是必须的,可以考虑使用**Protostuff**。
Protostuff 不需要依赖 IDL 文件,可以直接对 Java 领域对象进行反 / 序列化操作,在效率
上跟 Protobuf 差不多,生成的二进制格式和 Protobuf 是完全相同的,可以说是一个 Java
版本的 Protobuf 序列化框架
### 总结
实际上任何一种序列化框架, 核心思想就是设计一种序列化协议, 将对象的类型, 属性类型, 属性值 一一按照固定的格式写到二进制字节流中来完成序列化, 再按照固定的格式一一读出对象的类型,属性类型,属性值,通过这些信息重新创建出一个新的对象, 来完成反序列化
## 4. RPC框架中如何选择序列化
序列化框架的选择需要综合考虑一下因素:

综合上面几个参考因素,现在我们再来总结一下这几个序列化协议。
我们首选的还是**Hessian** 与 **Protobuf**,因为他们在性能、时间开销、空间开销、通用性、兼容性和安全性上,都满足了我们的要求。其中 Hessian 在使用上更加方便,在对象的兼容性上更好;Protobuf 则更加高效,通用性上更有优势。
### 4.1注意事项:
在使用 RPC 框架的过程中,我们构造入参、返回值对象,主要记住以下几点:
- 对象要尽量简单,没有太多的依赖关系,属性不要太多,尽量高内聚
- 入参对象与返回值对象体积不要太大,更不要传太大的集合;
- 尽量使用简单的、常用的、开发语言原生的对象,尤其是集合类;
- 对象不要有复杂的继承关系,最好不要有父子类的情况。
实际上,虽然 RPC 框架可以让我们发起全程调用就像调用本地,但在 RPC 框架的传输过程中,入参与返回值的根本作用就是用来传递信息的,为了提高 RPC 调用整体的性能和稳定性,我们的入参与返回值对象要构造得尽量简单,这很重要。<file_sep>
> Netty封装了JDK的NIO,使用简单健壮, 它是一个异步事件驱动的网络应用框架, 用于快速开发可维护的高性能服务器和客户端
## 1. 为什么使用Netty,而不使用JDK原生的NIO呢?
- 使用 JDK 自带的NIO需要了解太多的概念,编程复杂,一不小心 bug 横飞
- Netty 底层 IO 模型随意切换,而这一切只需要做微小的改动,改改参数,Netty可以直接从 NIO 模型变身为 IO 模型
- Netty 自带的拆包解包,异常检测等机制让你从NIO的繁重细节中脱离出来,让你只需要关心业务逻辑
- Netty 解决了 JDK 的很多包括空轮询在内的 Bug
- Netty 底层对线程,selector 做了很多细小的优化,精心设计的 reactor 线程模型做到非常高效的并发处理
- 自带各种协议栈让你处理任何一种通用协议都几乎不用亲自动手
- Netty 社区活跃,遇到问题随时邮件列表或者 issue
- Netty 已经历各大 RPC 框架,消息中间件,分布式通信中间件线上的广泛验证,健壮性无比强大
<file_sep>package proxy;
import net.sf.cglib.proxy.Enhancer;
/**
* <p>Title: TestCglib</p>
* <p>github </p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 18:59
*/
public class TestCglib {
public static void main(String[] args) {
CglibMethodInterceptor cglibMethodInterceptor = new CglibMethodInterceptor();
// 创建加强器,用来创建动态代理类
Enhancer eh = new Enhancer();
// 为代理类指定需要代理的类,也即是父类
eh.setSuperclass(HelloServiceImpl.class);
// 设置方法拦截器回调引用,对于代理类上所有方法的调用,都会调用CallBack,而Callback则需要实现intercept() 方法进行拦截
eh.setCallback(cglibMethodInterceptor);
HelloServiceImpl cglibProxy = (HelloServiceImpl)eh.create();
// cglib 创建对象消耗时间比较多 > JDk代理方式, 但处理的性能>JDK代理方式
// 一次创建 多次使用的对象 推荐使用cglib
cglibProxy.say();
}
}
<file_sep>
本专栏为分布式算法专题
> 分布式系统开发核心,在于如何选择或设计合适的算法, 解决一致性和可用性相关的问题
目前很多公司的开源产品, 开源的其实只是单机版本,如果要使用集群功能, 要么自研,要么购买企业版
而企业版一般都蛮贵的
比如InfluxDB,分布式企业版License授权费一年1.5万美刀,相比于单机版本,企业版本的技术壁垒又是什么呢?
其实它的护城河就是**以分布式算法为核心的分布式集群能力**
分布式系统的价值和意义很大, 我们需要准确的理解分布式算法, 这样才能开发出稳定的分布式系统
<file_sep>https://github.com/lightningMan/flash-netty<file_sep>package com.pushkin;
import org.springframework.boot.SpringApplication;
import org.springframework.boot.autoconfigure.SpringBootApplication;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/4/19 13:09
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
@SpringBootApplication
public class Start {
public static void main(String[] args) {
SpringApplication.run(Start.class, args);
}
}
<file_sep>
异步使用子线程实现, 回调就是调用另一类去调用自己(通过回调接口, 自己得实现这个接口)
异步回调, 就是使用子线程去调用另一个类去调用自己的回调方法
> 哪边想要回调,哪边就实现回调接口, 回调其实就是方法里传接口==>接口编程而已
## 1. 异步化
一般我们用多线程优化性能,其实不过就是将串行操作变成并行操作。
如果仔细观察, 你还会发现在串行转换成并行的过程中, 一定会涉及到异步化。
> 例: 为提升性能将以下串行任务变为并行任务
```java
// 串行
JobA();
JobB();
/** 使用两个子线程去执行,实现并行。
* 主线程无需等待JobA或者JobB的执行结果 = JobA与JobB两个操作已经被异步化
*/
new Thread(()->JobA()).start();
new Thread(()->JobB()).start();
```
**异步化**,是并行方案得以实施的基础,更深入地讲其实就是: **利用多线程优化性能这个核心方案得以实施的基础**
目前JDK1.8提供了CompletableFuture 来支持异步编程(JDK1.8之前的Future没有实现真正的异步回调,
get()获取结果,该方法会阻塞当前线程,非阻塞的方式isDone(),需要主线程循环询问子线程是否完成)
## 2. 回调
>核心:接口作为方法参数,其实际传入引用指向的是实现类
### 2.1 什么是回调
软件模块之间总是存在着一定的接口,从调用方式上,可以把他们分为三类:同步调用、回调和异步调用。
>在计算机程序设计中,回调函数,或简称回调,是指通过函数参数传递到其它代码的某一块可执行代码的引用。
>这一设计允许了底层代码调用在高层定义的子程序
具体说来:就是A类中调用B类中的某个方法C,然后B类中反过来调用A类中的方法D,D这个方法就叫回调方法,
### 2.2 三种方式的回调
回调是一种特殊的调用, 有3种方式的回调:
- 同步回调,即阻塞,单向。
- 回调,即双向(类似自行车的两个齿轮)。
- 异步调用,即通过异步消息进行通知。
#### 2.2.1 同步调用
一种阻塞式调用,调用方要等待对方执行完毕才返回,它是一种单向调用
```python
a() {
print "a";
}
b() {
a();
print "b";
}
```
#### 2.2.2 回调
一种双向调用模式,也就是说,被调用方在接口被调用时也会调用对方的接口;
代码模拟经典场景: 客户端向服务端发送请求, 服务端处理完请求然后进行响应
[见代码](./src/main/java/CH01/回调/SYNC_CS/CallBack.java)
*结果测试方法:*
```java
// one-by-one式请求, 服务器方同步式处理响应
client.request("仿真运行请求1");
System.out.println("--------------over-----------------");
client.request("仿真运行请求2");
System.out.println("--------------over-----------------");
client.request("仿真运行请求3");
System.out.println("--------------over-----------------");
```
*运行情况:*
```java
1. 客户端发起请求: 仿真运行请求1
2. 服务端: 服务端接收到客户端请求消息:仿真运行请求1
3. 服务端处理逻辑ing....
4. 服务端: 数据处理成功, 状态码 200, 返回结果
5. 客户端: 接受到服务端响应的消息状态码 200, 响应结果内容*****
--------------over-----------------
1. 客户端发起请求: 仿真运行请求2
2. 服务端: 服务端接收到客户端请求消息:仿真运行请求2
3. 服务端处理逻辑ing....
4. 服务端: 数据处理成功, 状态码 200, 返回结果
5. 客户端: 接受到服务端响应的消息状态码 200, 响应结果内容*****
--------------over-----------------
1. 客户端发起请求: 仿真运行请求3
2. 服务端: 服务端接收到客户端请求消息:仿真运行请求3
3. 服务端处理逻辑ing....
4. 服务端: 数据处理成功, 状态码 200, 返回结果
5. 客户端: 接受到服务端响应的消息状态码 200, 响应结果内容*****
--------------over-----------------
```
#### 2.2.3 异步回调
一种类似消息或事件的机制,接口的服务在收到某种讯息或发生某种事件时,会主动通知客户方(即调用客户方的接口)
>异步回调核心还是的配合多线程+回调实现
2.2.2 是同步的One-By-One, 客户端向服务器发起请求,服务器给出响应,客户端得到响应之后继续下一个请求
但实际在使用这种CS场景中,更多的是异步的场景,即客户端更多是的并发的发起不同的请求, 而服务端并不是同步的处理请求,而是多个线程去处理以提高响应效率
为实现这种场景, 代码如下: [见异步回调实现代码](./src/main/java/CH01/回调/ASYNC_CS/CallBack.java)
*测试方法:*
```java
// 主线程异步发送3个请求
client.request("仿真运行请求1");
client.request("仿真运行请求2");
client.request("仿真运行请求3");
```
*运行情况:*
```
1. 客户端发起请求(异步方式): 仿真运行请求1
1. 客户端发起请求(异步方式): 仿真运行请求2
1. 客户端发起请求(异步方式): 仿真运行请求3
2. 服务端: 服务端接收到客户端请求消息:仿真运行请求1 当前线程: Thread-0
2. 服务端: 服务端接收到客户端请求消息:仿真运行请求2 当前线程: Thread-1
3. 服务端处理逻辑ing.... 当前线程:Thread-1
2. 服务端: 服务端接收到客户端请求消息:仿真运行请求3 当前线程: Thread-2
3. 服务端处理逻辑ing.... 当前线程:Thread-0
3. 服务端处理逻辑ing.... 当前线程:Thread-2
4. 服务端: 数据处理成功, 状态码 200, 返回结果
4. 服务端: 数据处理成功, 状态码 200, 返回结果
5. 客户端: 接受到服务端响应的消息状态码 200, 响应结果内容***** 当前线程: Thread-2
4. 服务端: 数据处理成功, 状态码 200, 返回结果
5. 客户端: 接受到服务端响应的消息状态码 200, 响应结果内容***** 当前线程: Thread-1
5. 客户端: 接受到服务端响应的消息状态码 200, 响应结果内容***** 当前线程: Thread-0
```
<file_sep>package test;
import org.apache.commons.lang.StringUtils;
/**
* <p>Title: Test</p>
* <p>Company:浩鲸云计算科技股份有限公司 </p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-07-22 16:58
*/
public class Test {
public static void main(String[] args) {
// StringBuilder evtIdList = new StringBuilder();
// System.out.println(StringUtils.isEmpty(evtIdList.toString()));
System.out.println(10/(double)3);
}
}
<file_sep>package CH01.回调.ASYNC_CS;
/**
* <p>Title: Client</p>
*
* <p>Description:
* 描述:异步回调
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-15 08:14
*/
public class Client implements CallBack {
private Server server;
public Client(Server server) {
this.server = server;
}
public void request(final String requestHead){
System.out.println("1. 客户端发起请求(异步方式): "+requestHead);
System.out.println();
new Thread(() -> server.requestHandler(Client.this, requestHead
+" 当前线程: "+Thread.currentThread().getName())).start();
}
@Override
public void call(String response) {
System.out.println("5. 客户端: 接受到服务端响应的消息" + response+" 当前线程: "+Thread.currentThread().getName());
}
}
<file_sep>
## IO
> 在计算机系统中I/O就是输入(Input)和输出(Output)的意思,针对不同的操作对象,可以划分为磁盘I/O模型,网络I/O模型,
>内存映射I/O, Direct I/O、数据库I/O等,只要具有输入输出类型的交互系统都可以认为是I/O系统,也可以说I/O是整个操作系统数据交换
>与人机交互的通道,这个概念与选用的开发语言没有关系,是一个通用的概念
在如今的系统中I/O却拥有很重要的位置,现在系统都有可能处理大量文件,大量数据库操作,而这些操作都依赖于系统的I/O性能,
也就造成了现在系统的瓶颈往往都是由于I/O性能造成的。因此,为了解决磁盘I/O性能慢的问题,系统架构中添加了缓存来提高响应速度;
或者有些高端服务器从硬件级入手,使用了固态硬盘(SSD)来替换传统机械硬盘;在大数据方面,Spark越来越多的承担了实时性计算任务,
而传统的Hadoop体系则大多应用在了离线计算与大量数据存储的场景,这也是由于磁盘I/O性能远不如内存I/O性能而造成的格局(Spark更多的使用了内存,
而MapReduece更多的使用了磁盘)。因此,**一个系统的优化空间,往往都在低效率的I/O环节上**,很少看到一个系统CPU、内存的性能是其整个系统
的瓶颈。也正因为如此,Java在I/O上也一直在做持续的优化,从**JDK 1.4**开始便引入了NIO模型,大大的提高了以往BIO模型下的操作效率。
## IO 模型
- BIO(Blocking I/O):
同步阻塞I/O模式,数据的读取写入必须阻塞在一个线程内等待其完成。这里使用那个经典的烧开水例子,这里假设一个烧开水的场景,
有一排水壶在烧开水,BIO的工作模式就是, 叫一个线程停留在一个水壶那,直到这个水壶烧开,才去处理下一个水壶。
但是实际上线程在等待水壶烧开的时间段什么都没有做。
- NIO(New I/O):
同时支持阻塞与非阻塞模式,但这里我们以其同步非阻塞I/O模式来说明,那么什么叫做同步非阻塞?如果还拿烧开水来说,**NIO的做法是叫一个线程
不断的轮询每个水壶的状态**,看看是否有水壶的状态发生了改变,从而进行下一步的操作。
- AIO(Asynchronous I/O):
异步非阻塞I/O模型。异步非阻塞与同步非阻塞的区别在哪里?异步非阻塞无需一个线程去轮询所有IO操作的状态改变,在相应的状态改变后,
系统会通知对应的线程来处理。对应到烧开水中就是,为每个水壶上面装了一个开关,水烧开之后,水壶会自动通知我水烧开了。
进程中的IO调用步骤大致可以分为以下四步:
1. 进程向OS操作系统请求数据 ;
2. 操作系统把外部数据加载到内核的缓冲区中;
3. 操作系统把内核的缓冲区拷贝到进程的缓冲区 ;
4. 进程获得数据完成自己的功能 ;
当操作系统在把外部数据放到进程缓冲区的这段时间(即上述的第二,三步)[read/write],如果应用进程是挂起等待的,那么就是同步IO,反之,就是异步IO,也就是AIO 。
### BIO
> 阻塞; 面向流
典型应用: apache tomcat
场景: 适用于连接数小(并发量要求不高),且一次发送大量数据的场景,这种方式对于服务器资源要求比较高,并发局限于应用中。
### NIO (New I/O) 同步非阻塞I/O
> 非阻塞; 面向缓存区(通道Channel); IO多路复用
典型应用: Nginx, Netty, Jetty, Mina, Zookeeper
场景: 适用于处理连接数特别多(高并发场景),但是连接比较短(轻操作)的场景,服务器需要支持**超大量的长时间连接**。比如1万+个连接,并且客户端
**不会频繁地发送太多数据**
关于NIO,国内有很多技术博客将英文翻译成No-Blocking I/O,非阻塞I/O模型 ,当然这样就与BIO形成了鲜明的特性对比。
NIO本身是**基于事件驱动**的思想来实现的,其目的就是解决BIO的大并发问题,在BIO模型中,如果需要并发处理多个I/O请求,那就需要多线程来支持,
NIO使用了**多路复用器机制**,以socket使用来说,**多路复用器通过不断轮询各个连接的状态**,只有在socket有流可读或者可写时,应用程序才需要去处理它,
在线程的使用上,就不需要一个连接就必须使用一个处理线程了,而是只是**有效请求**时(确实需要进行I/O处理时),才会使用一个线程去处理,
这样就避免了BIO模型下大量线程处于阻塞等待状态的情景。
相对于BIO的流,NIO抽象出了新的通道(Channel)作为输入输出的通道,并且提供了缓存(Buffer)的支持,在进行读操作时,需要使用Buffer分配空间,
然后将数据从Channel中读入Buffer中,对于Channel的写操作,也需要现将数据写入Buffer,然后将Buffer写入Channel中。
如下NIO对文件的拷贝
```java
public static void copyFile (String srcFileName, String dstFileName) throws IOException {
FileInputStream fis = new FileInputStream(srcFileName);
FileOutputStream fos = new FileOutputStream(dstFileName);
FileChannel readChannel = fis.getChannel();
FileChannel writeChannel = fos.getChannel();
ByteBuffer buffer = ByteBuffer.allocate(1024);
while (true) {
buffer.clear();
if ( readChannel.read(buffer) == -1 ){
break;
}
// 将当前位置设置为limit, 然后再将缓存区游标置于0, 0-limit写入Channel
buffer.flip();
writeChannel.write(buffer);
}
fis.close();
fos.close();
}
```
通过比较New IO的使用方式我们可以发现,新的IO操作不再面向 Stream来进行操作了,改为了通道Channel,并且使用了更加灵活的缓存区类Buffer,
Buffer只是缓存区定义接口, 根据需要,我们可以选择对应类型的缓存区实现类。
在java NIO编程中,我们需要理解以下3个对象Channel、Buffer和Selector。
- Channel
首先说一下Channel,国内大多翻译成“通道”。Channel和IO中的Stream(流)是差不多一个等级的。只不过Stream是单向的,譬如:InputStream, OutputStream。
而Channel是双向的,既可以用来进行读操作,又可以用来进行写操作,NIO中的Channel的主要实现有:FileChannel、DatagramChannel、SocketChannel、ServerSocketChannel;
通过看名字就可以猜出个所以然来:分别可以对应文件IO、UDP和TCP(Server和Client)。
- Buffer
NIO中的关键Buffer实现有:ByteBuffer、CharBuffer、DoubleBuffer、 FloatBuffer、IntBuffer、 LongBuffer,、ShortBuffer,
分别对应基本数据类型: byte、char、double、 float、int、 long、 short。当然NIO中还有MappedByteBuffer, HeapByteBuffer, DirectByteBuffer等这里先不具体陈述其用法细节。
[说一下 DirectByteBuffer 与 HeapByteBuffer 的区别?](./DirectByteBuffer与HeapByteBuffer的区别.md)
- Selector
Selector 是NIO相对于BIO实现多路复用的基础,Selector 运行单线程处理多个 Channel,如果你的应用打开了多个通道,但每个连接的流量都很低,
使用 Selector 就会很方便。例如在一个聊天服务器中。要使用 Selector , 得向 Selector 注册 Channel,然后调用它的 select() 方法。
这个方法会一直阻塞到某个注册的通道有事件就绪。一旦这个方法返回,线程就可以处理这些事件,事件的例子有如新的连接进来、数据接收等。



这里我们再来看一个NIO模型下的TCP服务器的实现,我们可以看到Selector 正是NIO模型下 TCP Server 实现IO复用的关键,请仔细理解下段代码while循环中的逻辑,见下图:

### AIO
> 异步; 多线程
从编程模式上来看AIO相对于NIO的区别在于,**NIO需要使用者线程不停的轮询IO对象,来确定是否有数据准备好可以读了,而AIO则是在数据准备好之后,才会通知数据使用者**,
这样使用者就不需要不停地轮询了。当然AIO的异步特性并不是Java实现的伪异步,而是使用了系统底层API的支持,在Unix系统下,采用了epoll IO模型,
而windows便是使用了IOCP模型。关于Java AIO,本篇只做一个抛砖引玉的介绍,如果你在实际工作中用到了,那么可以参考Netty在高并发下使用AIO的相关技术。
## 总结
BIO、NIO、AIO适用场景分析:
BIO方式适用于连接数目比较小且固定的架构,这种方式对服务器资源要求比较高
NIO方式适用于连接数目多且连接比较短的架构,可充分利用服务器资源
AIO方式使用于连接数目多且连接比较长的架构,充分调用OS参与并发操作
>总 结:
>IO实质上与线程没有太多的关系,但是不同的IO模型改变了应用程序使用线程的方式,NIO与AIO的出现解决了很多BIO无法解决的并发问题,当然任何技术抛开适用场景都是耍流氓,复杂的技术往往是为了解决简单技术无法解决的问题而设计的,在系统开发中能用常规技术解决的问题,绝不用复杂技术,否则大大增加系统代码的维护难度,学习IT技术不是为了炫技,而是要实实在在解决问题。
---
*本文参考博文地址:*
*https://juejin.im/entry/598da7d16fb9a03c42431ed3*
*https://www.cnblogs.com/binghuaZhang/p/11042835.html*
---
*blog 插眼:*
*https://www.jianshu.com/p/cde27461c226*
*https://juejin.im/book/5b4bc28bf265da0f60130116/section/5b6a1a9cf265da0f87595521*<file_sep>package proxy;
/**
* <p>Title: HelloStaticProxy</p>
* <p>github </p>
* <p>Description:
* 描述:代理类
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 18:34
*/
public class HelloStaticProxy implements HelloService{
HelloServiceImpl realHello;
public HelloStaticProxy(HelloServiceImpl realHello) {
this.realHello = realHello;
}
@Override
public void say() {
System.out.println("手动实现代理类: ");
System.out.println("执行前操作-----------");
realHello.say();
System.out.println("执行后操作-----------");
}
@Override
public void play() {
System.out.println("手动实现代理类: ");
System.out.println("执行前操作-----------");
realHello.play();
System.out.println("执行后操作-----------");
}
}
<file_sep>
## 1. 简介
Master-Worker模式是常用的并行设计模式
核心思想:系统有两种进程协议工作
- Master进程: 负责接收与分配任务
- Worker进程: 负责处理子任务
当Worker进程将子任务处理完后,将结果返回给Master进程,由Master进程归纳和汇总, 从而得到系统结果

## 2. 优点
- 任务分解, 并行化, 提升性能
- 异步
<file_sep>
## 代理模式
>将一个对象的直接访问,变为访问这个对象的代理对象,通过代理对象间接的访问原本的对象
**为什么需要代理呢?**
因为一个良好的设计不应该被轻易地修改,这正是开闭原则的体现: 一个良好的设计应该对修改关闭, 对扩展开放。
而代理正是为了扩展类的而存在的, 其可以控制对现有类服务的访问。
## 类的加载
- 静态代理的代理类在运行前(编译期间)就生成了代理类, 不是编译器生成的代理类,而是手动创建的类
- 动态代理的代理类并不是在java代码中定义好的,而是在运行时(运行期动态创建)根据我们在java代码中的"指示"动态生成的
ClassLoader, 可以在运行期动态地载入字节码, 从而装载定义在其中的类信息(Class对象)

## 远程调用的魔法
在项目中,当我们要使用 RPC 的时候,我们一般的做法是先找服务提供方要接口,通过Maven 或者其他的工具把接口
依赖到我们项目中。我们在编写业务逻辑的时候,如果要调用提供方的接口,我们就只需要通过依赖注入的方式把接口
注入到项目中就行了,然后在代码里面直接调用接口的方法 。
我们都知道,接口里并不会包含真实的业务逻辑,业务逻辑都在服务提供方应用里,但我们
通过调用接口方法,确实拿到了想要的结果,是不是感觉有点神奇呢?想一下,在 RPC 里
面,我们是怎么完成这个魔术的。
**核心技术就是动态代理**, RPC 会自动给接口生成一个代理类,当我
们在项目中注入接口的时候,运行过程中实际绑定的是这个接口生成的代理类。这样在接口
方法被调用的时候,它实际上是被生成代理类拦截到了,这样我们就可以在生成的代理类里
面,加入远程调用逻辑。
通过这种“偷梁换柱”的手法,就可以帮用户屏蔽远程调用的细节,实现像调用本地一样地
调用远程的体验,整体流程如下图所示:

## 实现
### 1. 静态代理
在编译期就生成代理类,实现简单,但不灵活
### 2. 动态代理
增加程序的灵活性 (其主要原理是利用动态生成与装载字节码的技术,为指定的接口动态创建代理类)
#### 2.1 基于JDK实现动态代理(基于接口实现)
- 关键接口: InvocationHandler
- 关键类 : Proxy.newProxyInstance
InvocationHandler这个接口是被动态代理类回调的接口,我们增加的针对委托类的统一处理逻辑都增加到invoke 方法里面
JDK动态代理的代理类字节码在创建时,需要实现业务实现类所实现的接口作为参数。如果业务实现类是没有实现接口而是直接定义业务方法的话,
就无法使用JDK动态代理了。(JDK动态代理重要特点是代理接口) 并且,如果业务实现类中新增了接口中没有的方法,
这些方法是无法被代理的(因为无法被调用)。
动态代理只能对接口产生代理,不能对类产生代理
#### 2.2 基于CGLIB实现动态代理(基于继承实现)
- 关键接口: MethodInterceptor
Cglib是针对类来实现代理的,他的原理是对代理的目标类生成一个子类,并覆盖其中方法实现增强,因为底层是基于创建被代理类的一个子类,所以它避免了JDK动态代理类的缺陷。
但因为采用的是继承,所以不能对final修饰的类进行代理。final修饰的类不可继承。
CGLIB创建的动态代理对象在性能方面比JDK创建的动态代理对象要高很多,但创建代理对象所花费的时间却比JDK动态代理多得多,
**因此,对于一次创建而多次使用的对象,用CGLIB更合适,反之,使用JDK的动态代理更合适**
<file_sep>
单例方式(根据创建实例的时期):
- 饿汉模式: 在类的成员初始化创建 (创建时期不优雅, 内存空间浪费等问题)
- 懒汉模式: 在类的方法中创建 (要考虑 安全性; 有序性等问题)
饿汉优化 --> 延迟加载(懒加载)
懒汉优化 --> 双重校验 + 可见volatile
饿汉本身就是线程安全, 饿汉式第一次调用更快, 懒汉式第一次调用需要实例化该对象,
懒汉式双重null的检查,确保了只有第一次调用单例的时候才会去做同步,这样也是线程安全的,同时避免了每次都做同步的性能损耗
<file_sep>package com.pushkin.nettyim.serialize;
import com.pushkin.nettyim.serialize.impl.JSONSerializer;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 14:33
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public interface Serializer {
/**
* json 序列化 [默认]
*/
byte JSON_SERIALIZER = 1;
Serializer DEFAULT = new JSONSerializer();
/**
* 序列化算法
*/
byte getSerializerAlogrithm();
/**
* java 对象转换成二进制
*/
byte[] serialize(Object object);
/**
* 二进制转换成 java 对象
*/
<T> T deserialize(Class<T> clazz, byte[] bytes);
}
<file_sep>package CH00.单例案例;
/**
* <p>Title: Singleton3</p>
* <p>Description:
* 描述:双重校验 + 可见volatile
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-22 09:57
*/
public final class Singleton3 {
/** 可见volatile */
private static volatile Singleton3 instance = null;
private Singleton3() {
}
/** 双重校验 */
public static Singleton3 getInstance() {
if (instance == null) {
synchronized (Singleton3.class) {
if (instance == null) {
instance = new Singleton3();
}
}
}
return instance;
}
}
<file_sep>package com.pushkin.nettyim.protocol.response;
import com.pushkin.nettyim.protocol.Packet;
import static com.pushkin.nettyim.protocol.command.Command.LOGIN_RESPONSE;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 16:36
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public class LoginResponsePacket extends Packet {
private boolean success;
private String reason;
@Override
public Byte getCommand() {
return LOGIN_RESPONSE;
}
public boolean isSuccess() {
return success;
}
public void setSuccess(boolean success) {
this.success = success;
}
public String getReason() {
return reason;
}
public void setReason(String reason) {
this.reason = reason;
}
}
<file_sep>import impl.JdkSerialaizerImpl;
import impl.JsonSerialaizerImpl;
import service.ObjectSerializer;
import java.io.IOException;
/**
* <p>Title: TestSerialaizer</p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-09 14:14
*/
public class TestSerialaizer {
public static void main(String[] args) throws IOException, ClassNotFoundException {
TestStudent t1 = new TestStudent(110, "student1");
// JDK序列化
JdkSerialaizerImpl jdkSerialaizer = new JdkSerialaizerImpl();
// Json序列化
ObjectSerializer jsonSerialaizer = new JsonSerialaizerImpl();
// 测试JDK自带的序列化
testSerialaizer(t1, jdkSerialaizer);
// 测试JSON序列化
// testSerialaizer(t1, jsonSerialaizer);
}
static void testSerialaizer(Object obj, ObjectSerializer objectSerializer) throws IOException, ClassNotFoundException {
// 序列化
byte[] serialize = objectSerializer.serialize(obj);
// 反序列化
TestStudent t3 = objectSerializer.deserialize(serialize, new TestStudent().getClass());
System.out.println(t3.getNo());
System.out.println(t3.getName());
}
}
<file_sep># 异步编程之CompletableFuture
在我们的开发过程中,经常遇到一些复杂的逻辑,需要分解成多个任务job去处理,常规的
one-by-one这种串行编码的方式虽然简单, 但性能方面不尽人意, 一般的优化可以
利用提升系统资源使用率, 使用多线程的方式去实现一些job的并行化, 但job之间很多时候
是相互依赖耦合,job之间需要进行数据的交互, 那这种场景又如何实现呢?
答案是: 异步编程
>串行任务(同步)实现并行化(异步化)本质还是用多线程+回调实现。
JDK1.5提供了Future和Callable的实现,去实现这种需求场景
<file_sep>package proxy;
import proxy.util.MonitorUtil;
import java.lang.reflect.InvocationHandler;
import java.lang.reflect.Method;
import java.util.Arrays;
/**
* <p>Title: HelloinvocationHandler</p>
* <p>github </p>
* <p>Description:
* 描述:
* </p>
* 实现InvocationHandler接口,这个类中持有一个被代理对象(委托类)的实例target。
* 该类别JDK Proxy类回调
*
* InvocationHandler 接口中有一个invoke方法,当一个代理实例的方法被调用时,
* 代理方法将被编码并分发到 InvocationHandler接口的invoke方法执行。
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 18:42
*/
public class HelloInvocationHandler<T> implements InvocationHandler {
/**
* 被代理对象引用,invoke 方法里面method 需要使用这个被代理对象
*/
private T target;
public HelloInvocationHandler(T target) {
this.target = target;
}
/**
* InvocationHandler这个接口 是被动态代理类回调的接口,
* 我们所有需要增加的针对委托类的统一处理逻辑都增加到invoke 方法里面
* 在调用委托类接口方法之前或之后 结束战斗。
*/
@Override
public Object invoke(Object proxy, Method method, Object[] args) throws Throwable {
System.out.println("被动态代理类回调执行, 代理类 proxyClass ="+proxy.getClass()+" 方法名: " + method.getName() + "方法. 方法返回类型:"+method.getReturnType()
+" 接口方法入参数组: "+(args ==null ? "null" : Arrays.toString(args)));
System.out.println("执行前操作-----------");
MonitorUtil.start();
// Thread.sleep(1000);
/** 调用被代理的真实对象*/
Object result = method.invoke(target, args);
System.out.println("执行后操作-----------");
MonitorUtil.finish(method.getName());
return result;
}
}<file_sep>package proxy;
/**
* <p>Title: TestStatic</p>
* <p>github </p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 18:44
*/
public class TestStatic {
public static void main(String[] args) {
HelloStaticProxy helloStaticProxy = new HelloStaticProxy(new HelloServiceImpl());
helloStaticProxy.say();
}
}
<file_sep>package CH01.回调.ASYNC_CS;
/**
* <p>Title: Test</p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-15 08:14
*/
public class Test {
public static void main(String[] args) {
Server server = new Server();
Client client = new Client(server);
// 主线程异步发送3个请求
client.request("仿真运行请求1");
client.request("仿真运行请求2");
client.request("仿真运行请求3");
}
}
<file_sep># 1. 如何实现并发, 并发模型

<file_sep>package com.pushkin.register.registry;
import com.pushkin.register.constants.Constants;
import org.apache.curator.framework.CuratorFramework;
import org.apache.curator.framework.CuratorFrameworkFactory;
import org.apache.curator.retry.RetryNTimes;
import org.apache.zookeeper.CreateMode;
import org.springframework.stereotype.Service;
import java.util.List;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 21:51
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
@Service
public class IRegistryCenterImpl implements IRegistryCenter {
public CuratorFramework curatorFramework = null;
/**
* 连接zookeeper
*
* @param zkAddress zkAddress
* @return CuratorFramework
*/
public CuratorFramework getCuratorFramework(String zkAddress) {
if (curatorFramework == null) {
synchronized (this) {
if (curatorFramework == null) {
curatorFramework = CuratorFrameworkFactory.builder()
.connectString(zkAddress)
.connectionTimeoutMs(10000)
.retryPolicy(new RetryNTimes(3, 2000))
.build();
curatorFramework.start();
try {
curatorFramework.blockUntilConnected();
} catch (InterruptedException e) {
e.printStackTrace();
}
System.out.println("连接zookeeper成功");
}
}
}
return curatorFramework;
}
@Override
public void register(String zkAddress, String serviceName, String serviceAddress) {
curatorFramework = getCuratorFramework(zkAddress);
// zk节点: /registry/xxxx-provider/ip:9090
try {
// 创建根节点
if (curatorFramework.checkExists().forPath(Constants.REGISTRY) == null) {
curatorFramework.create()
// 持久节点
.withMode(CreateMode.PERSISTENT)
.forPath(Constants.REGISTRY);
}
// 创建服务节点
String serviceNode = Constants.REGISTRY + "/" + serviceName + "/" + serviceAddress;
curatorFramework.create()
.creatingParentsIfNeeded()
// 临时节点
.withMode(CreateMode.EPHEMERAL)
.forPath(serviceNode);
System.out.println("服务注册完毕...."+serviceName);
} catch (Exception e) {
e.printStackTrace();
}
}
/**
* 服务发现
*
* @param zkAddress zkAddress
* @param serviceName serviceName
* @return
*/
@Override
public List<String> discoveryService(String zkAddress, String serviceName) {
curatorFramework = getCuratorFramework(zkAddress);
// zk节点: /registry/xxxx-provider/ip:9090
String serviceNode = Constants.REGISTRY + "/" + serviceName;
try {
// 服务提供者可能是集群提供多个
return curatorFramework.getChildren().forPath(serviceNode);
} catch (Exception e) {
e.printStackTrace();
}
return null;
}
}
<file_sep>package lockandconfition;
import java.util.concurrent.locks.Condition;
import java.util.concurrent.locks.Lock;
import java.util.concurrent.locks.ReentrantLock;
/**
* <p>Title: BlockedQueue</p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-28 20:51
*/
public class BlockedQueue {
// notFull 不满锁, 如果队列已满, 就释放锁, 等到队列不满(信号) [出队的话 就不满]; 入队 通知notEmpty线程不用阻塞 可以取数(信号)了
// notEmpty 不空锁, 如果队列已空, 就释放锁, 等到队列不空(信号) [入队的话 就不空]; 出队 通知notFull线程不用阻塞 可以装数(信号)了
// 核心就是: 阻塞(释放锁) + 通知信号
// 用同一把锁, 锁同一个不共享变量 的不同操作!
final ReentrantLock lock = new ReentrantLock();
final Condition notFull = lock.newCondition();
final Condition notEmpty = lock.newCondition();
final Object[] items = new Object[100];
int putptr, takeptr, count;
public void put(Object x) throws InterruptedException {
lock.lock();
try {
// 当数组满了
while (count == items.length) {
// 释放锁,等待
notFull.await();
}
// 放入数据
items[putptr] = x;
// 如果到最后一个位置了,下标从头开始,防止下标越界
if (++putptr == items.length) {
// 从头开始
putptr = 0;
}
// 对 count ++ 加加
++count;
// 通知 take 线程,可以取数据了,不必继续阻塞
notEmpty.signal();
} finally {
lock.unlock();
}
}
public Object take() throws InterruptedException {
lock.lock();
try {
// 如果数组没有数据,则等待
while (count == 0) {
notEmpty.await();
}
// 取数据
Object x = items[takeptr];
// 如果到数组尽头了,就从头开始
if (++takeptr == items.length) {
// 从头开始
takeptr = 0;
}
// 将数量减1
--count;
// 通知阻塞的 put 线程可以装填数据了
notFull.signal();
return x;
}
finally {
lock.unlock();
}
}
public static void main(String[] args) throws InterruptedException {
BlockedQueue blockedQueue = new BlockedQueue();
for (int j = 0; j < 10; j++) {
new Thread(()->{
for (int i = 0; i < 10; i++) {
try {
blockedQueue.put(i);
System.out.println(Thread.currentThread().getName()+"入队: "+i);
}
catch (InterruptedException e) {
e.printStackTrace();
}
}
}).start();
new Thread(()-> {
for (int i = 0; i < 10; i++) {
try {
Object take = blockedQueue.take();
System.out.println(Thread.currentThread().getName()+"出队: "+take);
}
catch (InterruptedException e) {
e.printStackTrace();
}
}
}).start();
}
}
}
<file_sep>import java.io.Serializable;
/**
* <p>Title: TestStudent</p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-09 14:13
*/
public class TestStudent implements Serializable {
/**
* serialVersionUID
* 反序列化时,会根据serialVersionUID这个常量值来判断反序列得到的类信息是否与原来一致
* 如果我们不指定serialVersionUID值,序列化将会把当前类的hashCode值赋给它
*
*/
private int no;
private String name;
public int getNo() {
return no;
}
public void setNo(int no) {
this.no = no;
}
public String getName() {
return name;
}
public void setName(String name) {
this.name = name;
}
public TestStudent(int no, String name) {
this.no = no;
this.name = name;
}
public TestStudent() {
}
@Override
public String toString() {
return "TestStudent{" +
"no=" + no +
", name='" + name + '\'' +
'}';
}
}
<file_sep>https://www.acfun.cn/v/ac13837146_3
pom:
还得配置
<dependency>
<groupId>org.springframework.boot</groupId>
<artifactId>spring-boot-configuration-processor</artifactId>
</dependency>
否则com.pushkin.register.autoconfig.RegistryAutoConfiguration会报spring Boot configuration annotation processor not configured
注意不要忘记此文件
META-INF/spring.factories
指定自动配置的类
# Auto Configure
org.springframework.boot.autoconfigure.EnableAutoConfiguration=com.pushkin.register.autoconfig.RegistryAutoConfiguration
1. 添加注册中心的依赖
2. 提供者项目配置文件:
server.port=9100
spring.application.name=spring-cloud-service-provider
spring.registry.zookeeper.zkAddress=127.0.0.1:2181
spring.registry.zookeeper.server=true<file_sep>package com.pushkin.register.constants;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 22:02
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public class Constants {
public static String REGISTRY = "/registry";
}
<file_sep>package com.pushkin.register.registry;
import org.springframework.stereotype.Component;
import java.util.List;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 21:44
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
@Component
public interface IRegistryCenter{
/**
* 注册服务
*
* @param zkAddress zkAddress
* @param serviceName 服务名称
* @param serviceAddress 服务地址
*/
void register(String zkAddress, String serviceName, String serviceAddress);
/**
* 服务发现.
*
* @param zkAddress zkAddress
* @param serviceName serviceName
* @return serviceList
*/
List<String> discoveryService(String zkAddress, String serviceName);
}
<file_sep>package proxy;
import java.lang.reflect.Proxy;
/**
* <p>Title: TestJdk</p>
* <p>github </p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 18:44
*/
public class TestJdk {
public static void main(String[] args) {
System.out.println("-------------------第一种创建代理类方法--------------");
System.out.println("-------------------通过Proxy.newProxyInstance 方法 获取代理对象--------------");
// 1. 委托类
HelloServiceImpl helloService = new HelloServiceImpl();
// 2. 委托类的统一处理逻辑Handler (回调InvocationHandler接口的invoke方法执行)
HelloInvocationHandler<HelloServiceImpl> helloServiceHelloInvocationHandler = new HelloInvocationHandler<>(helloService);
// 3. 创建一个代理对象 [动态创建- 代码运行期创建代理]
HelloService jdkProxy = (HelloService)Proxy.newProxyInstance(HelloService.class.getClassLoader(),
new Class<?>[]{HelloService.class}, helloServiceHelloInvocationHandler);
// 4. 调用
jdkProxy.say();
}
}
<file_sep>package lockandconfition;
/**
* <p>Title: test06</p>
* <p>Company:浩鲸云计算科技股份有限公司 </p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-07-30 21:05
*/
public class Test06 {
public Object lock = new Object();
public static void main(String[] args) {
Test06 test06 = new Test06();
// 创建线程对象
Thread t = new Thread(new MyRun06(test06));
t.setName("t");
t.start();
// try {
// // 问题:这行代码会让t线程睡眠5秒钟吗?
// t.sleep(5 * 1000);
// } catch (InterruptedException e) {
// e.printStackTrace();
// }
System.out.println("hello world");
}
public void doSleep() throws InterruptedException {
synchronized (lock) {
System.out.println("开始sleep");
Thread.sleep(10000);
System.out.println("结束sleep");
}
}
}
class MyRun06 implements Runnable {
private Test06 t;
public MyRun06(Test06 t) {
this.t = t;
}
@Override
public void run() {
try {
t.doSleep();
}
catch (InterruptedException e) {
e.printStackTrace();
}
for (int i = 0; i < 100; i++) {
System.out.println(Thread.currentThread().getName() + "-->" + i);
}
}
}
<file_sep>package test;
/**
* <p>Title: CallBack</p>
*
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 22:40
*/
public interface CallBack {
public void solve(String result);
}
<file_sep>package com.pushkin.nettyim.server;
import com.pushkin.nettyim.protocol.Packet;
import com.pushkin.nettyim.protocol.PacketCodeC;
import com.pushkin.nettyim.protocol.request.LoginRequestPacket;
import com.pushkin.nettyim.protocol.response.LoginResponsePacket;
import io.netty.buffer.ByteBuf;
import io.netty.channel.ChannelHandlerContext;
import io.netty.channel.ChannelInboundHandlerAdapter;
import java.util.Date;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 15:28
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public class ServerHandler extends ChannelInboundHandlerAdapter {
@Override
public void channelRead(ChannelHandlerContext ctx, Object msg) {
System.out.println(new Date() + ": 客户端开始登录……");
ByteBuf requestByteBuf = (ByteBuf) msg;
Packet packet = PacketCodeC.INSTANCE.decode(requestByteBuf);
if (packet instanceof LoginRequestPacket) {
// 登录流程
LoginRequestPacket loginRequestPacket = (LoginRequestPacket) packet;
LoginResponsePacket loginResponsePacket = new LoginResponsePacket();
loginResponsePacket.setVersion(packet.getVersion());
if (valid(loginRequestPacket)) {
loginResponsePacket.setSuccess(true);
System.out.println(new Date() + ": 登录成功!");
} else {
loginResponsePacket.setReason("账号密码校验失败");
loginResponsePacket.setSuccess(false);
System.out.println(new Date() + ": 登录失败!");
}
// 登录响应
ByteBuf responseByteBuf = PacketCodeC.INSTANCE.encode(ctx.alloc(), loginResponsePacket);
ctx.channel().writeAndFlush(responseByteBuf);
}
}
private boolean valid(LoginRequestPacket loginRequestPacket) {
String username = loginRequestPacket.getUsername();
System.out.println("账号认证成功: "+username);
return true;
}
}
<file_sep># Lock和Condition
编程领域两大核心问题:
- 互斥: 即同一时刻只允许一个线程访问共享资源
- 同步: 即线程之间如何通信、协作
>互斥;同步这两个问题, 管程都能解决, JDK并发包通过Lock和Condition两个接口来实现管程
>其中Lock用于解决互斥问题, Condition用于解决同步问题
## 1.Lock轮子的意义
java语言提的synchronized也是管程的实现,既然java已经从语言层面已经实现了管程,为什么还要在SDK里提供另外一种实现呢?
JDK1.5 synchronized性能不如LOCK
JDK1.6 synchronized性能优于LOCK
Lock有别与synchronized隐式锁的三个特性:
- 响应中断
- 支持超时
- 非阻塞获取锁
### 1.1 死锁问题_破坏不可抢占条件方案
这个方案synchronized没有办法解决, 因为其在申请资源的时候, 如果申请不到, 线程直接进入阻塞状态了,
什么都干不了,也释放不了线程已经占有的资源
> 对于"不可抢占"这个条件, 占有部分资源的线程进一步申请其他资源时, 如果申请不到, 可以主动释放它占有的资源
这样抢占这个条件就破坏了
### 1.2 设计互斥锁
如何重新设计一把互斥锁去解决这个问题呢? 可以有三种方案
#### 1.2.1 能够响应中断
synchronized 的问题是,持有锁 A 后,如果尝试获取锁 B 失败,那么
线程就进入阻塞状态,一旦发生死锁,就没有任何机会来唤醒阻塞的线程。但如果阻塞
状态的线程能够响应中断信号,也就是说当我们给阻塞的线程发送中断信号的时候,能
够唤醒它,那它就有机会释放曾经持有的锁 A。这样就破坏了不可抢占条件了。
#### 1.2.2 支持超时
如果线程在一段时间之内没有获取到锁,不是进入阻塞状态,而是返回一个
错误,那这个线程也有机会释放曾经持有的锁。这样也能破坏不可抢占条件。
#### 1.2.3 非阻塞地获取锁
如果尝试获取锁失败,并不进入阻塞状态,而是直接返回,那这个线
程也有机会释放曾经持有的锁。这样也能破坏不可抢占条件。
这三种方案可以全面弥补 synchronized 的问题
这三个方案就是“重复造LOCK轮子”的主要原因,体现在 API 上,就是 Lock 接口的三个方法。详情如
下:
// 支持中断的 API
void lockInterruptibly()
throws InterruptedException;
// 支持超时的 API
boolean tryLock(long time, TimeUnit unit)
throws InterruptedException;
// 支持非阻塞获取锁的 API
boolean tryLock();
## 2.如何保证可见性
Java SDK 里面 Lock 的使用,有一个经典的范例,就是try{}finally{},需要重点关注
的是在 finally 里面释放锁。这个范例无需多解释,你看一下下面的代码就明白了。但是有
一点需要解释一下,那就是可见性是怎么保证的。你已经知道 Java 里多线程的可见性是通
过 Happens-Before 规则保证的,而 synchronized 之所以能够保证可见性,也是因为有
一条 synchronized 相关的规则:synchronized 的解锁 Happens-Before 于后续对这个锁
的加锁。那 Java SDK 里面 Lock 靠什么保证可见性呢?例如在下面的代码中,线程 T1 对
value 进行了 +=1 操作,那后续的线程 T2 能够看到 value 的正确结果吗?
class X {
private final Lock rtl = new ReentrantLock();
int value;
public void addOne() {
// 获取锁
rtl.lock();
try {
value+=1;
} finally {
// 保证锁能释放
rtl.unlock();
}
}
}
答案必须是肯定的。Java SDK 里面锁的实现非常复杂,这里我就不展开细说了,但是原理
还是需要简单介绍一下:它是利用了 volatile 相关的 Happens-Before 规则。Java SDK
里面的 ReentrantLock,内部持有一个 volatile 的成员变量 state,获取锁的时候,会读写
state 的值;解锁的时候,也会读写 state 的值(简化后的代码如下面所示)。也就是说,
在执行 value+=1 之前,程序先读写了一次 volatile 变量 state,在执行 value+=1 之
后,又读写了一次 volatile 变量 state。根据相关的 Happens-Before 规则:
1. **顺序性规则**:对于线程 T1,value+=1 Happens-Before 释放锁的操作 unlock();
2. **volatile 变量规则**:由于 state = 1 会先读取 state,所以线程 T1 的 unlock() 操作
Happens-Before 线程 T2 的 lock() 操作;
3. **传递性规则**:线程 T1 的 value+=1 Happens-Before 线程 T2 的 lock() 操作。
class SampleLock {
volatile int state;
// 加锁
lock() {
// 省略代码无数
state = 1;
}
// 解锁
unlock() {
// 省略代码无数
state = 0;
}
}
所以说,后续线程 T2 能够看到 value 的正确结果
## 3. 可重入锁
> 所谓可重入锁,顾名思义,指的是线程可以重复获取同一把锁
例如下面代码中,当线程 T1 执行到 ① 处时,已经获取到了锁 rtl ,当在
① 处调用 get() 方法时,会在 ② 再次对锁 rtl 执行加锁操作。此时,如果锁 rtl 是可重入
的,那么线程 T1 可以再次加锁成功;如果锁 rtl 是不可重入的,那么线程 T1 此时会被阻
塞。
class X {
private final Lock rtl = new ReentrantLock();
int value;
public int get() {
// 获取锁
rtl.lock(); ②
try {
return value;
} finally {
// 保证锁能释放
rtl.unlock();
}
}
public void addOne() {
// 获取锁
rtl.lock();
try {
value = 1 + get(); ①
} finally {
// 保证锁能释放
rtl.unlock();
}
}
}
除了可重入锁,可能你还听说过可重入函数,可重入函数怎么理解呢?指的是线程可以重复
调用?显然不是,所谓可重入函数,指的是多个线程可以同时调用该函数,每个线程都能得
到正确结果;同时在一个线程内支持线程切换,无论被切换多少次,结果都是正确的。多线
程可以同时执行,还支持线程切换,这意味着什么呢?线程安全啊。所以,可重入函数是线
程安全的。
## 4. 公平锁与非公平锁
在使用 ReentrantLock 的时候,你会发现 ReentrantLock 这个类有两个构造函数,一个是
无参构造函数,一个是传入 fair 参数的构造函数。fair 参数代表的是锁的公平策略,如果传
入 true 就表示需要构造一个公平锁,反之则表示要构造一个非公平锁。
// 无参构造函数:默认非公平锁
public ReentrantLock() {
sync = new NonfairSync();
}
// 根据公平策略参数创建锁
public ReentrantLock(boolean fair){
sync = fair ? new FairSync()
: new NonfairSync();
}
入口等待队列,锁都对应着一个等待队列,如果一个线程没有获得锁,就会进入等待队列,当有线程释放锁的时候,就需
要从等待队列中唤醒一个等待的线程。如果是公平锁,唤醒的策略就是谁等待的时间长,就
唤醒谁,很公平;如果是非公平锁,则不提供这个公平保证,有可能等待时间短的线程反而
先被唤醒。
## 5. 用锁的最佳实践
你已经知道,用锁虽然能解决很多并发问题,但是风险也是挺高的。可能会导致死锁,也可
能影响性能。这方面有是否有相关的最佳实践呢?有,还很多。但是我觉得最值得推荐的是
并发大师 <NAME>《Java 并发编程:设计原则与模式》一书中,推荐的三个用锁的最佳
实践,它们分别是:
>1. 永远只在更新对象的成员变量时加锁
>2. 永远只在访问可变的成员变量时加锁
>3. 永远不在调用其他对象的方法时加锁
这三条规则,前两条估计你一定会认同,最后一条你可能会觉得过于严苛。但是我还是倾向
于你去遵守,因为调用其他对象的方法,实在是太不安全了,也许“其他”方法里面有线程
sleep() 的调用,也可能会有奇慢无比的 I/O 操作,这些都会严重影响性能。更可怕的
是,“其他”类的方法可能也会加锁,然后双重加锁就可能导致死锁。
## 6. 思考题:
你已经知道 tryLock() 支持非阻塞方式获取锁,下面这段关于转账的程序就使用到了
tryLock(),你来看看,它是否存在死锁问题呢?
```java
class Account {
private int balance;
private final Lock lock = new ReentrantLock();
// 转账
void transfer(Account tar, int amt){
while (true) {
if(this.lock.tryLock()) {
try {
if (tar.lock.tryLock()) {
try {
this.balance -= amt;
tar.balance += amt;
} finally {
tar.lock.unlock();
}
}//if
} finally {
this.lock.unlock();
}
}//if
}//while
}//transfer
}
```<file_sep>package impl;
import com.alibaba.fastjson.JSON;
import service.ObjectSerializer;
import java.io.IOException;
import java.io.InputStream;
import java.io.OutputStream;
/**
* <p>Title: JsonSerialaizerImpl</p>
* <p>github </p>
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-09 14:59
*/
public class JsonSerialaizerImpl implements ObjectSerializer {
public byte[] serialize(Object obj) throws IOException {
return JSON.toJSONBytes(obj);
}
public void serialize(Object obj, OutputStream os) throws IOException {
}
public <T> T deserialize(byte[] bytes, Class<T> clazz) throws IOException, ClassNotFoundException {
return JSON.parseObject(bytes, clazz);
}
public <T> T deserialize(InputStream is, Class<T> clazz) throws IOException, ClassNotFoundException {
return null;
}
public Object deserialize(InputStream is) throws IOException, ClassNotFoundException {
return null;
}
public Object deserialize(byte[] bytes) throws IOException, ClassNotFoundException {
return null;
}
public <T> T deserialize(byte[] b, T co) throws IOException, ClassNotFoundException {
return null;
}
}
<file_sep>
[对象如何在网络中传输](./对象如何在网络中传输.md)
本节主要介绍了RPC框架中常见的几个序列化框架, 以及一些实现<file_sep>package proxy.util;
/**
* <p>Title: MonitorUtil</p>
* <p>github </p>
* <p>Description:
* 描述:方法用时监控类
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-13 15:31
*/
public class MonitorUtil {
private static ThreadLocal<Long> tl = new ThreadLocal<>();
public static void start() {
tl.set(System.currentTimeMillis());
}
/**
* 结束时打印耗时
*
* @param methodName 方法名
*/
public static void finish(String methodName) {
long finishTime = System.currentTimeMillis();
System.out.println(methodName + "方法执行耗时" + (finishTime - tl.get()) + "ms");
}
}
<file_sep>package com.pushkin.controller;
import com.pushkin.websocket.WebSocketPushHandler;
import org.springframework.web.bind.annotation.PostMapping;
import org.springframework.web.bind.annotation.RequestMapping;
import org.springframework.web.bind.annotation.RestController;
import org.springframework.web.socket.TextMessage;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/4/19 13:39
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
@RestController
@RequestMapping("/msg")
public class MsgController {
/**
* 功能描述:向全体广播消息
* @param: [msg] 消息内容
* @return: boolean
* @auther: lzy
* @date: 2019/8/14 16:10
*/
@PostMapping("/sendMsg")
public boolean sendMsg(String msg){
System.out.println("全体广播消息 ["+msg+"]");
TextMessage textMessage = new TextMessage(msg);
try{
WebSocketPushHandler.sendMessagesToUsers(textMessage);
}catch (Exception e){
return false;
}
return true;
}
/**
* 功能描述:向指定用户发送消息
* @param msg 消息内容
* @param userId 用户编号
* @return: boolean
* @auther: lzy
* @date: 2019/8/14 16:13
*/
@PostMapping("/sendMsgByUser")
public boolean sendMsgByUser(String msg,String userId){
System.out.println("向 "+userId+" 发送消息,消息内容为:"+msg);
TextMessage textMessage = new TextMessage(msg);
try{
WebSocketPushHandler.sendMessageToUser(userId,textMessage);
}catch (Exception e){
return false;
}
return true;
}
}
<file_sep>package com.pushkin.nettyim.serialize;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 14:34
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public interface SerializerAlgorithm {
/**
* json 序列化标识
*/
byte JSON = 1;
}
<file_sep>package com.pushkin.demo.netty;
import io.netty.buffer.ByteBuf;
import io.netty.channel.ChannelHandlerContext;
import io.netty.channel.ChannelInboundHandlerAdapter;
import java.nio.charset.Charset;
/**
* @author pushkin
* @version v1.0.0
* @date 2020/5/24 10:59
* <p>
* Modification History:
* Date Author Version Description
* ------------------------------------------------------------
*/
public class FirstServerHandler extends ChannelInboundHandlerAdapter {
@Override
public void channelRead(ChannelHandlerContext ctx, Object msg) {
System.out.println(msg);
System.out.println("收到客户端心跳监测, 开始响应客户端");
ByteBuf byteBuf = getByteBuf(ctx, "你好 客户端, xxx服务运行正常....");
ctx.channel().writeAndFlush(byteBuf);
}
private ByteBuf getByteBuf(ChannelHandlerContext ctx, String inputStr) {
// 1. 获取二进制抽象 ByteBuf
ByteBuf buffer = ctx.alloc().buffer();
// 2. 准备数据
byte[] bytes = inputStr.getBytes(Charset.forName("UTF-8"));
buffer.writeBytes(bytes);
return buffer;
}
}
<file_sep>package CH01.回调.SYNC_CS;
/**
* <p>Title: test</p>
*
* <p>Description:
* 描述:
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-15 07:55
*/
public class Test {
public static void main(String[] args) {
Server server = new Server();
Client client = new Client(server);
// one-by-one式请求, 服务器方同步式处理响应
client.request("仿真运行请求1");
System.out.println("--------------over-----------------");
client.request("仿真运行请求2");
System.out.println("--------------over-----------------");
client.request("仿真运行请求3");
System.out.println("--------------over-----------------");
}
}
<file_sep>package CH00.单例案例;
/**
* <p>Title: Singleton4</p>
* <p>Description: 静态内部类实现
* 描述:实现延迟加载
* <p>
* <p>
* 这种方式同样利用了类加载机制来保证只创建一个instance实例。它与饿汉模式一样,也是利用了类加载机制,因此不存在多线程并发的问题。
* 不一样的是,它是在内部类里面去创建对象实例。这样的话,只要应用中不使用内部类,JVM就不会去加载这个单例类,也就不会创建单例对象,
* 从而实现懒汉式的延迟加载。也就是说这种方式可以同时保证延迟加载和线程安全。
*
*
*
* </p>
*
* @author jinpu.shi
* @version v1.0.0
* @since 2020-04-22 09:52
*/
public class Singleton4 {
/**
* 外部类加载的时候, 静态内部类并不会被加载进去的
* 只有在 Singleton4.newInstance 才会被加载
*
*/
private static class SingletonHolder {
private static Singleton4 instance = new Singleton4();
}
private Singleton4() {
}
public static Singleton4 newInstance() {
return SingletonHolder.instance;
}
}
<file_sep>“有时候,我认为我永远不会离开道拉多雷斯大街了。一旦写下这句话。它对于我来说,就如同永恒的谶言。” - 费尔南多·佩索阿
<file_sep>
>并发问题解决的核心技术就是使用"管程"
# 1. 什么是管程 (Monitor)
java1.5 之前提供唯一并发原语就是管程
java1.5 之后提供的SDK并发包 也是以管程技术为基础的
本质: "管程"和OS的"信号量"是等价的,所谓等价指的是用管程能够实现信号量,也能用信号量实现管程
不知道你是否曾思考过这个问题:为什么 Java 在 1.5 之前仅仅提供了 synchronized 关键字及 wait()、notify()、notifyAll() 这三个看似从天而降的方法?在刚接触 Java 的时候,
我以为它会提供信号量这种编程原语,因为操作系统原理课程告诉我,用信号量能解决所有并发问题,结果我发现不是。后来我找到了原因:Java 采用的是管程技术,
synchronized 关键字及 wait()、notify()、notifyAll() 这三个方法都是管程的组成部分。而管程和信号量是等价的,所谓等价指的是用管程能够实现信号量,也能用信号量实现管程。
但是管程更容易使用,所以 Java 选择了管程。
管程,对应的英文是 Monitor,很多 Java 领域的同学都喜欢将其翻译成“监视器”,这是直译。操作系统领域一般都翻译成“管程”,这个是意译,而我自己也更倾向于使用“管程”。
所谓管程,指的是管理共享变量以及对共享变量的操作过程,让他们支持并发。翻译为 Java 领域的语言,就是管理类的成员变量和成员方法,让这个类是线程安全的。那管程是怎么管的呢?
# 2. 管程模型
- Hasen
- Hoare
- MESA (目前广泛使用, java观察实现也是基于此模型)
在并发编程领域,有两大核心问题:一个是**互斥**,即同一时刻只允许一个线程访问共享资源;
另一个是**同步**,即线程之间如何通信、协作。这两大问题,管程都是能够解决的。
本文重点介绍MESA模型
## 2.1 如何解决互斥问题
管程解决互斥问题的思路很简单,就是将共享变量及其对共享变量的操作统一封装起来。
在下图中,管程 X 将共享变量 queue 这个队列和相关的操作入队 enq()、出队 deq()
都封装起来了;线程 A 和线程 B 如果想访问共享变量 queue,只能通过调用管程提供的 enq()、deq() 方法来实现;
enq()、deq() 保证互斥性,只允许一个线程进入管程。不知你有没有发现,管程模型和面向对象高度契合的。
估计这也是 Java 选择管程的原因吧。而我在前面章节介绍的互斥锁用法,其背后的模型其实就是它。

## 2.2 如何解决同步问题
MESA 模型处理:
在管程模型里,共享变量和对共享变量的操作是被封装起来的,图中最外层的框就代表封装的意思。框的上面只有一个入口,
并且在入口旁边还有一个入口等待队列。当多个线程同时试图进入管程内部时,只允许一个线程进入,其他线程则在入口等待队列中等待。这个过程类似就医流程的分诊,
只允许一个患者就诊,其他患者都在门口等待。
管程里还引入了条件变量的概念,而且每个条件变量都对应有一个等待队列,如下图,条件变量 A 和条件变量 B 分别都有自己的等待队列。

那条件变量和等待队列的作用是什么呢?其实就是解决线程同步问题。你也可以结合上面提到的入队出队例子加深一下理解。
假设有个线程 T1 执行出队操作,不过需要注意的是执行出队操作,有个前提条件,就是队列不能是空的,而队列不空这个前提条件就是管程里的条件变量。 如果线程 T1 进入管程后恰好发现队列是空的,那怎么办呢?等待啊,去哪里等呢?就去条件变量对应的等待队列里面等。此时线程 T1 就去“队列不空”这个条件变量的等待队列中等待。这个过程类似于大夫发现你要去验个血,于是给你开了个验血的单子,你呢就去验血的队伍里排队。线程 T1 进入条件变量的等待队列后,是允许其他线程进入管程的。这和你去验血的时候,医生可以给其他患者诊治,道理都是一样的。
再假设之后另外一个线程 T2 执行入队操作,入队操作执行成功之后,“队列不空”这个条件对于线程 T1 来说已经满足了,此时线程 T2 要通知 T1,告诉它需要的条件已经满足了。当线程 T1 得到通知后,会从等待队列里面出来,但是出来之后不是马上执行,而是重新进入到入口等待队列里面。这个过程类似你验血完,回来找大夫,需要重新分诊。
条件变量及其等待队列我们讲清楚了,下面再说说 wait()、notify()、notifyAll() 这三个操作。前面提到线程 T1 发现“队列不空”这个条件不满足,需要进到对应的等待队列里等待。这个过程就是通过调用 wait() 来实现的。如果我们用对象 A 代表“队列不空”这个条件,那么线程 T1 需要调用 A.wait()。同理当“队列不空”这个条件满足时,线程 T2 需要调用 A.notify() 来通知 A 等待队列中的一个线程,此时这个队列里面只有线程 T1。至于 notifyAll() 这个方法,它可以通知等待队列中的所有线程。
这里我还是来一段代码再次说明一下吧。下面的代码实现的是一个阻塞队列,阻塞队列有两个操作分别是入队和出队,这两个方法都是先获取互斥锁,类比管程模型中的入口。
1. 对于入队操作,如果队列已满,就需要等待直到队列不满,所以这里用了notFull.await();。
2. 对于出队操作,如果队列为空,就需要等待直到队列不空,所以就用了notEmpty.await();。
3. 如果入队成功,那么队列就不空了,就需要通知条件变量:队列不空notEmpty对应的等待队列。
4. 如果出队成功,那就队列就不满了,就需要通知条件变量:队列不满notFull对应的等待队列。
```java
// 条件变量及其等待队列
// await() 和前面我们提到的 wait() 语义是一样的;signal() 和前面我们提到的 notify() 语义是一样的。
public class BlockedQueue<T>{
final Lock lock =
new ReentrantLock();
// 条件变量:队列不满
final Condition notFull =
lock.newCondition();
// 条件变量:队列不空
final Condition notEmpty =
lock.newCondition();
// 入队
void enq(T x) {
lock.lock();
try {
while (队列已满){
// 等待队列不满
notFull.await();
}
// 省略入队操作...
// 入队后, 通知可出队
notEmpty.signal();
}finally {
lock.unlock();
}
}
// 出队
void deq(){
lock.lock();
try {
while (队列已空){
// 等待队列不空
notEmpty.await();
}
// 省略出队操作...
// 出队后,通知可入队
notFull.signal();
}finally {
lock.unlock();
}
}
}
```
**await() 和前面我们提到的 wait() 语义是一样的;signal() 和前面我们提到的 notify() 语义是一样的。**
## 2.3 wait() 的正确姿势
但是有一点,需要再次提醒,对于 MESA 管程来说,有一个编程范式,就是需要在一个 while 循环里面调用 wait()。
**这个是 MESA 管程特有的**
```java
while(条件不满足) {
wait();
```
Hasen 模型、Hoare 模型和 MESA 模型的一个核心区别就是当条件满足后,如何通知相关线程。
管程要求同一时刻只允许一个线程执行,那当线程 T2 的操作使线程 T1 等待的条件满足时,T1 和 T2 究竟谁可以执行呢?
1. Hasen 模型里面,要求 notify() 放在代码的最后,这样 T2 通知完 T1 后,T2 就结束了,然后 T1 再执行,这样就能保证同一时刻只有一个线程执行。
2. Hoare 模型里面,T2 通知完 T1 后,T2 阻塞,T1 马上执行;等 T1 执行完,再唤醒 T2,也能保证同一时刻只有一个线程执行。但是相比 Hasen 模型,T2 多了一次阻塞唤醒操作。
3. MESA 管程里面,T2 通知完 T1 后,T2 还是会接着执行,T1 并不立即执行,仅仅是从条件变量的等待队列进到入口等待队列里面。这样做的好处是 notify() 不用放到代码的最后,T2
也没有多余的阻塞唤醒操作。但是也有个副作用,就是当 T1 再次执行的时候,可能曾经满足的条件,现在已经不满足了,所以需要以循环方式检验条件变量。
## 2.4 notify() 何时可以使用
**除非经过深思熟虑,否则尽量使用 notifyAll()**
notify()需要满足以下三个条件:
1. 所有等待线程拥有相同的等待条件;
2. 所有等待线程被唤醒后,执行相同的操作;
3. 只需要唤醒一个线程。
比如上面阻塞队列的例子中,对于“队列不满”这个条件变量,其阻塞队列里的线程都是在等待“队列不满”这个条件,反映在代码里就是下面这 3 行代码。对所有等待线程来说,都是执行这 3 行代码,重点是 while 里面的等待条件是完全相同的。
```java
while (队列已满){
// 等待队列不满
notFull.await();
}
```
所有等待线程被唤醒后执行的操作也是相同的,都是下面这几行:
```java
// 省略入队操作...
// 入队后, 通知可出队
notEmpty.signal();
```
同时也满足第 3 条,只需要唤醒一个线程。所以上面阻塞队列的代码,使用 signal() 是可以的。
# 3.总结
Java 参考了 MESA 模型,语言内置的管程(synchronized)对 MESA 模型进行了精简。MESA 模型中,条件变量可以有多个,Java 语言内置的管程里只有一个条件变量。具体如下图所示。

Java 内置的管程方案(synchronized)使用简单,synchronized 关键字修饰的代码块,在编译期会自动生成相关加锁和解锁的代码,但是仅支持一个条件变量;而 Java SDK 并发包实现的管程支持多个条件变量,不过并发包里的锁,需要开发人员自己进行加锁和解锁操作。
并发编程里两大核心问题——互斥和同步,都可以由管程来帮你解决。学好管程,理论上所有的并发问题你都可以解决,并且很多并发工具类底层都是管程实现的,所以学好管程,就是相当于掌握了一把并发编程的万能钥匙。<file_sep>
## 并发编程
> 并发编程核心矛盾一直存在, 本质在于CPU, 内存, IO设备的速度差异
为合理利用CPU的高性能, 平衡这三者的速度差异, 计算机系统, 编译程序等都做出了贡献,主要体现为:
1. CPU 增加了缓存,以均衡与内存的速度差异;
2. 操作系统增加了进程、线程,以分时复用 CPU,进而均衡 CPU 与 I/O 设备的速度差
异;
3. 编译程序优化指令执行次序,使得缓存能够得到更加合理地利用
但由此也带来很多并发的问题,要解决这些问题, 我们可以从如下几点考虑解决
- 可见性
- 原子性
- 有序性
## 1 可见性
> 一个线程对共享变量的修改, 另一个线程能够立刻看到, 我们称为可见性
可见性主要问题,来自于当前多核时代,每个CPU都有其自己的缓存, 与内存的数据一致性的问题
多个线程在不同的CPU上运行时,这些线程操作的是不同的CPU缓存
## 2 原子性
> 我们把一个或者多个操作在 CPU 执行的过程中不被中断的特性称为原子性
这块主要的问题,主要来自于操作系统CPU多**线程切换**所带来的。
早期的操作系统基于进程来调度 CPU,不同进程间是不共享内存空间的,所以进程要做任
务切换就要切换内存映射地址,而一个进程创建的所有线程,都是共享一个内存空间的,所
以线程做任务切换成本就很低了。现代的操作系统都基于更轻量的线程来调度,现在我们提
到的“任务切换”都是指“线程切换”
java 并发程序都是基于多线程的,自然也会涉及到任务切换,也许你想不到,任务切换竟
然也是并发编程里诡异 Bug 的源头之一。任务切换的时机大多数是在时间片结束的时候,
我们现在基本都使用高级语言编程,高级语言里一条语句往往需要多条 CPU 指令完成,例
如上面代码中的count += 1,至少需要三条 CPU 指令。
指令 1:首先,需要把变量 count 从内存加载到 CPU 的寄存器;
指令 2:之后,在寄存器中执行 +1 操作;
指令 3:最后,将结果写入内存(缓存机制导致可能写入的是 CPU 缓存而不是内存)
操作系统做任务切换,可以发生在任何一条CPU 指令执行完,是的,是 CPU 指令,而不
是高级语言里的一条语句。对于上面的三条指令来说,我们假设 count=0,如果线程 A 在
指令 1 执行完后做线程切换,线程 A 和线程 B 按照下图的序列执行,那么我们会发现两个
线程都执行了 count+=1 的操作,但是得到的结果不是我们期望的 2,而是 1。
我们潜意识里面觉得 count+=1 这个操作是一个不可分割的整体,就像一个原子一样,线
程的切换可以发生在 count+=1 之前,也可以发生在 count+=1 之后,但就是不会发生在
中间。**我们把一个或者多个操作在 CPU 执行的过程中不被中断的特性称为原子性**。CPU
能保证的原子操作是 CPU 指令级别的,而不是高级语言的操作符,这是违背我们直觉的地
方。因此,很多时候我们需要在高级语言层面保证操作的原子性。
### 2.1 如何解决原子性问题?
原子性问题的源头就是**线程切换**,而操作系统做线程切换时依赖CPU中断的,所以禁止CPU发生中断就能够禁止线程切换
单核时代(同一时刻只有一个线程),此方案可行,但并不适合多核场景(同一时刻多个线程同时运行,且执行在不同的CPU上)
**同一时刻只有一个线程执行**才能真正意义上保证原子性, 这个条件我们可以称之为**互斥**,保证对共享变量的修改时互斥的,
那无论单核还是多核场景都可以解决原子性问题
#### 2.1.1 简易锁模型
> 1. 加锁操作 lock(); 2.临界区:代码逻辑; 3. 解锁操作: unlock()
**临界区:** 我们把一段需要互斥执行的代码称之为临界区
如厕的例子其实就是个简易锁模型,这个事就是临界区,但很容易让我们忽略两个非常重要的点:
- 我们锁的是什么?
- 我们保护的又是什么?
#### 2.1.2 改进后的锁模型
我们知道在现实世界里,锁和锁要保护的资源是有对应关系的,比如你用你家的锁保护你家
的东西,我用我家的锁保护我家的东西。在并发编程世界里,锁和资源也应该有这个关系,
但这个关系在我们上面的模型中是没有体现的,所以我们需要完善一下我们的模型。

首先,我们要把临界区要保护的资源标注出来,如图中临界区里增加了一个元素:受保护的
资源 R;其次,我们要保护资源 R 就得为它创建一把锁 LR;最后,针对这把锁 LR,我们
还需在进出临界区时添上加锁操作和解锁操作。另外,在锁 LR 和受保护资源之间,我特地
用一条线做了关联,这个关联关系非常重要。很多并发 Bug 的出现都是因为把它忽略了,
然后就出现了类似锁自家门来保护他家资产的事情,这样的 Bug 非常不好诊断,因为潜意
识里我们认为已经正确加锁了。
### 2.2 java提供的锁技术: Synchronized
锁是一种通用的技术方案,java提供Synchronized关键字,就是锁的一种实现。
Synchronized关键字可以用来修饰方法, 也可以用来修饰代码块, 使用示例:
```java
class X {
// 修饰非静态方法
synchronized void foo() {
// 临界区
}
// 修饰静态方法
synchronized static void bar() {
// 临界区
}
// 修饰代码块
Object obj = new Object();
void baz() {
synchronized(obj) {
// 临界区
}
}
}
```
Java 编译器会在 synchronized 修饰的方法或代码块前后自动加上加锁 lock() 和解
锁 unlock(),这样做的好处就是加锁 lock() 和解锁 unlock() 一定是成对出现的
>加锁本质就是在锁对象的对象头中写入当前线程id。
>sync锁的对象monitor指针指向一个ObjectMonitor对象,所有线程加入他的entrylist里
面,去cas抢锁,更改state加1拿锁,执行完代码,释放锁state减1,和aqs机制差不多,
只是所有线程不阻塞,cas抢锁,没有队列,属于非公平锁。
wait的时候,线程进waitset休眠,等待notify唤醒
那么synchronized里的加锁lock()与解锁unlock()锁定的对象在哪呢?
上面的代码我们
看到只有修饰代码块的时候,锁定了一个 obj 对象,那修饰方法的时候锁定的是什么呢?
这个也是 Java 的一条隐式规则:
>当修饰静态方法的时候,锁定的是当前类的 Class 对象
>当修饰非静态方法的时候,锁定的是当前实例对象 this。
```java
class X {
// 修饰静态方法
synchronized(X.class) static void bar() {
// 临界区
}
}
class X {
// 修饰非静态方法
synchronized(this) void foo() {
// 临界区
}
}
```
#### 2.2.1 使用 synchronized实现读写互斥锁
```java
class SafeCalc {
long value = 0L;
// get()方法也要synchronized
synchronized long get() {
return value;
}
// addOne synchronized
synchronized void addOne() {
value += 1;
}
}
```
get() 方法和 addOne() 方
法都需要访问 value 这个受保护的资源,这个资源用 this 这把锁来保护。线程要进入临界
区 get() 和 addOne(),必须先获得 this 这把锁,这样 get() 和 addOne() 也是互斥的。

这个模型更像现实世界里面球赛门票的管理,一个座位只允许一个人使用,这个座位就
是“受保护资源”,球场的入口就是 Java 类里的方法,而门票就是用来保护资源
的“锁”,Java 里的检票工作是由 synchronized 解决的。
### 2.3 锁和受保护资源的关系
我们前面提到,受保护资源和锁之间的关联关系非常重要,他们的关系是怎样的呢?一个合
理的关系是:受保护资源和锁之间的关联关系是 N:1 的关系。还拿前面球赛门票的管理来
类比,就是一个座位,我们只能用一张票来保护,如果多发了重复的票,那就要打架了。现
实世界里,我们可以用多把锁来保护同一个资源,但在并发领域是不行的,并发领域的锁和
现实世界的锁不是完全匹配的。不过倒是可以用同一把锁来保护多个资源,这个对应到现实
世界就是我们所谓的“包场”了。
上面那个例子我稍作改动,把 value 改成静态变量,把 addOne() 方法改成静态方法,此
时 get() 方法和 addOne() 方法是否存在并发问题呢?
```java
class SafeCalc {
static long value = 0L;
// get()方法也要synchronized
synchronized long get() {
return value;
}
// addOne synchronized
synchronized static void addOne() {
value += 1;
}
}
```
如果你仔细观察,就会发现改动后的代码是用两个锁保护一个资源。这个受保护的资源就是
静态变量 value,两个锁分别是 this 和 SafeCalc.class。我们可以用下面这幅图来形象描述
这个关系。由于临界区 get() 和 addOne() 是用两个锁保护的,因此这两个临界区没有互斥
关系,临界区 addOne() 对 value 的修改对临界区 get() 也没有可见性保证,这就导致并
发问题了。

### 总结:
互斥锁,在并发领域的知名度极高,只要有了并发问题,大家首先容易想到的就是加锁,因
为大家都知道,加锁能够保证执行临界区代码的互斥性。这样理解虽然正确,但是却不能够
指导你真正用好互斥锁。临界区的代码是操作受保护资源的路径,类似于球场的入口,入口
一定要检票,也就是要加锁,但不是随便一把锁都能有效。所以必须深入分析锁定的对象和
受保护资源的关系,综合考虑受保护资源的访问路径,多方面考量才能用好互斥锁。
synchronized 是 Java 在语言层面提供的互斥原语,其实 Java 里面还有很多其他类型的
锁,但作为互斥锁,原理都是相通的:**锁,一定有一个要锁定的对象,至于这个锁定的对象
要保护的资源以及在哪里加锁 / 解锁,就属于设计层面的事情了**
## 3 有序性
源代码顺序:源代码中所指定的内存访问操作顺序。
程序顺序:我们可以理解为编译得到机器码或者解释执行的字节码(之后把两者统称为字节码)所指定的内存访问顺序。
执行顺序:内存访问在指定处理器上的实际执行顺序。
感知顺序:给定处理器感知到其他处理器内存访问的顺序。
> 指令重排序: 程序顺序与源代码顺序不一致 或者 执行顺序与程序顺序 不一致
> 存储子系统重排序: 表现在感知顺序与执行顺序不一样
### 3.1 编译优化带来的有序性问题
Java平台包括两种编译器:
静态编译器(javac)和动态编译器(jit:just in time)。
静态编译器是将.java文件编译成.class文件(二进制文件),之后便可以解释执行。动态编译器是将.class文件编译成机器码,之后再由jvm运行。
jit主要是做性能上面的优化,如热点代码编译成本地代码,加速调用。
| e2546dc70fc03ec9276dc3fb2387938305e89641 | [
"Markdown",
"Java"
] | 47 | Markdown | Shkin1/D-NOTEBOOK | 0dbaaac6b0a3157e86fe1bea6781e57a8165a047 | 00c8d7c940a2427c918ba5833c8f13ab8c36175e |
refs/heads/master | <repo_name>kupchenko/spring-boot-notes-demo<file_sep>/README.md
# Notes management system
### Getting started
### Setting up UI:
* Install dependencies UI part with ```npm i```
* UI Prod-Build ```npm run build:prod```
* Move output from ```ui/dist``` folder to ```src/main/resources/static```
### Run the application:
* Start the application by executing:
```
./gradlew clean bootRun
```
* DEV: Start the application by executing:
```
./gradlew bootRun --args='--spring.profiles.active=dev'
```
### Build service docker image:
To build run:
```
docker build -t notes:1.0-notes-service -f src/main/docker/Dockerfile .
```
To run:
```
docker run -p 8080:8080 notes:1.0-notes-service
```
### Build ui docker image:
To build run:
```
docker build -t notes:1.0-notes-frontend -f ui/src/docker/Dockerfile .
```
To run:
```
docker run -p 80:80 notes:1.0-notes-frontend
```<file_sep>/src/test/resources/db/schema.sql
DROP TABLE notes IF EXISTS;
CREATE TABLE notes
(
id integer identity primary key,
title varchar(128) not null,
content varchar(512),
owner varchar(128) not null,
created_ts datetime not null,
updated_ts datetime not null
);<file_sep>/src/main/resources/db/test-data.sql
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (0, 'My first note', 'Here is i can white my notes. This text is long enough to show shorter.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (1, 'My second note', 'Some content here. Adding long text. Test test test test.', 'user', '2014-02-15', '2014-02-15');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (2, 'My third note', 'Another content here. Another text is presented here.', 'user1', '2014-02-13', '2014-02-13');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (3, 'My fourth note', 'Some tfgyhu jihugyfdf lhbdf sdhgf shdgf jdshgfjhsdgf jhsdgfjsdgf iuytwegifuyg isdugfiusydgfiuywegifuygw', 'user1', '2014-02-16', '2014-02-17');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (4, 'My fifth note', 'Fifth Some content here. Adding long text. Test test test test.', 'user1', '2014-02-18', '2014-02-18');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (5, '1My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-19', '2014-02-19');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (6, '2My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (7, '3My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (8, '4My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (9, '5My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (10, '6My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (11, '7My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (12, '8My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (13, '9My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (14, '10My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (15, '11My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (16, '12My sixth note', 'Sixth Another content here. Another text is presented here.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (17, 'Note with non-existing user', 'Some content here. Another text is presented here.', -1, '2014-02-14', '2014-02-14');<file_sep>/settings.gradle
rootProject.name = 'notes-service'
<file_sep>/src/test/resources/db/data.sql
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (0, 'My first note', 'Here is i can white my notes. This text is long enough to show shorter.', 'user1', '2014-02-14', '2014-02-14');
INSERT INTO notes(id, title, content, owner, created_ts, updated_ts)
VALUES (1, 'My test note', 'Some content here. Adding long text. Test test test test.', 'user', '2014-02-15', '2014-02-15');<file_sep>/ui/src/js/service/api.service.js
import ExceptionHandlerService from './exception-handler-service';
import NotificationService from "./notification-service";
import {HTTP_METHOD_GET, HTTP_METHOD_POST, HTTP_METHOD_PUT} from "../utils/request-method";
import {APPLICATION_JSON_VALUE, AUTH_TOKEN} from "../utils/request-header";
import appConfig from "../config/config-app";
export default class ApiService {
static fetch(url, payload = {}) {
const options = {
method: HTTP_METHOD_GET,
headers: {
'Authorization': ApiService.getToken(AUTH_TOKEN),
'Content-Type': APPLICATION_JSON_VALUE
},
};
return fetch(ApiService.buildUri(url, payload), options)
.then((response) => {
return ApiService.parseResponse(response, HTTP_METHOD_GET);
});
}
static async update(url, payload) {
NotificationService.loading('Updating ...', url);
const options = {
method: HTTP_METHOD_PUT,
headers: {
'Authorization': ApiService.getToken(AUTH_TOKEN),
'Content-Type': APPLICATION_JSON_VALUE
},
body: JSON.stringify(payload),
};
return fetch(this.getFullUrl(url), options)
.then((response) => {
return ApiService.parseResponse(response, HTTP_METHOD_PUT);
});
}
static async create(url, payload) {
NotificationService.loading('Creating ...', url);
const options = {
method: HTTP_METHOD_POST,
headers: {
'Authorization': ApiService.getToken(AUTH_TOKEN),
'Content-Type': APPLICATION_JSON_VALUE
},
body: JSON.stringify(payload),
};
return fetch(this.getFullUrl(url), options)
.then((response) => {
return ApiService.parseResponse(response, HTTP_METHOD_POST);
});
}
static async retrieveToken(code) {
const username = appConfig.APP_OAUTH2_USERNAME;
const password = <PASSWORD>;
let formdata = new FormData();
let headers = new Headers();
formdata.append('grant_type', 'authorization_code');
formdata.append('code', code);
headers.append('Authorization', 'Basic ' + Buffer.from(username + ":" + password).toString('base64'));
const options = {
method: HTTP_METHOD_POST,
headers: headers,
body: formdata
};
return fetch(appConfig.API_URL_BASE + '/oauth/token', options)
.then((response) => response.json());
}
static buildUri(url, params = {}) {
const fullUrl = this.getFullUrl(url);
if (!params || Object.entries(params).length === 0) return fullUrl;
const uri = fullUrl.indexOf('?') === -1 ? `${fullUrl}?` : fullUrl;
return uri + ApiService.buildParams(params);
}
static getFullUrl(url) {
return appConfig.API_URL_BASE + appConfig.API_CONTEXT_PATH + url;
}
static buildParams(params) {
let paramsArray = ['text=' + params.text, 'page=' + params.page, 'rows=' + params.rows];
return paramsArray.join('&');
}
static parseResponse(response, requestType) {
console.log(`Parsing response with status: ${response.status}`);
ExceptionHandlerService.catchApiErrors(response, requestType);
if (response.status === 204) return null;
return response.json();
}
static getToken(tokenName) {
return localStorage.getItem(tokenName);
}
}
<file_sep>/src/main/java/me/kupchenko/controller/SwaggerController.java
package me.kupchenko.controller;
import org.springframework.stereotype.Controller;
import org.springframework.web.bind.annotation.GetMapping;
import static me.kupchenko.util.Constant.Endpoint.SWAGGER_URL;
@Controller
public class SwaggerController {
@GetMapping({"/swagger"})
public String swaggerForward() {
return "redirect:" + SWAGGER_URL;
}
}
<file_sep>/src/main/java/me/kupchenko/controller/AdminNoteController.java
package me.kupchenko.controller;
import lombok.AllArgsConstructor;
import lombok.extern.slf4j.Slf4j;
import me.kupchenko.dto.AdminNotesResponseDto;
import me.kupchenko.dto.ExtendedNoteDto;
import me.kupchenko.dto.NotesSearchDto;
import me.kupchenko.service.NoteService;
import org.springframework.web.bind.annotation.GetMapping;
import org.springframework.web.bind.annotation.PathVariable;
import org.springframework.web.bind.annotation.RequestMapping;
import org.springframework.web.bind.annotation.RestController;
import javax.validation.Valid;
import static org.springframework.http.MediaType.APPLICATION_JSON_VALUE;
@Slf4j
@RestController
@AllArgsConstructor
@RequestMapping("/admin/notes")
public class AdminNoteController {
private NoteService noteService;
@GetMapping("/{noteId:[0-9]+}")
public ExtendedNoteDto getNote(@PathVariable Long noteId) {
return noteService.getNote(noteId);
}
@GetMapping(value = "/user/{userId:[0-9]+}", produces = APPLICATION_JSON_VALUE)
public AdminNotesResponseDto searchUserNotes(@PathVariable String userId,
@Valid NotesSearchDto searchDto) {
return noteService.searchAdminUserNotes(userId, searchDto);
}
}
<file_sep>/ui/src/docker/Dockerfile
FROM nginx
COPY ui/dist /data/html
COPY nginx.conf /etc/nginx/nginx.conf
<file_sep>/ui/src/js/actions/note-create.js
import {
C_NOTE_CREATE_FAILURE,
C_NOTE_CREATE_IS_IN_PROGRESS,
C_NOTE_CREATE_MODAL_HIDE,
C_NOTE_CREATE_MODAL_SHOW,
C_NOTE_CREATE_SUCCESS
} from "./action-type";
import {actionDoNotesSearch} from "./notes-search";
import {actionRestoreCreateValues} from "./note-creating";
import ApiService from "../service/api.service";
export const actionNoteCreateInProgress = (bool) => ({
type: C_NOTE_CREATE_IS_IN_PROGRESS,
isLoading: bool
});
export const actionNoteCreateSuccess = (note) => ({
type: C_NOTE_CREATE_SUCCESS,
note
});
export const actionNoteCreateFailure = () => ({
type: C_NOTE_CREATE_FAILURE
});
export const actionShowNoteCreateModal = () => ({
type: C_NOTE_CREATE_MODAL_SHOW
});
export const actionHideNoteCreateModal = () => ({
type: C_NOTE_CREATE_MODAL_HIDE
});
export const actionDoNoteCreate = (newTitle, newContent) => {
return (dispatch) => {
dispatch(actionNoteCreateInProgress(true));
ApiService.create(`/notes`, {
'userId': 0,
'title': newTitle,
'content': newContent
}).then(() => {
dispatch(actionNoteCreateSuccess());
dispatch(actionRestoreCreateValues());
dispatch(actionDoNotesSearch());
}).catch(() => {
dispatch(actionNoteCreateFailure())
});
}
};<file_sep>/ui/src/js/component/notes/create/note-create-modal-body.js
import {Input} from 'antd';
import React from "react";
import {connect} from "react-redux";
import {actionDoUpdateNoteContent, actionDoUpdateNoteTitle} from "../../../actions/note-creating";
class NoteCreateModalBody extends React.Component {
constructor(props) {
super(props);
}
handleContentChange = (e) => {
this.props.actionDoUpdateNoteContent(e.target.value);
};
handleTitleChange = (e) => {
this.props.actionDoUpdateNoteTitle(e.target.value);
};
render() {
const {TextArea} = Input;
const {title, content} = this.props.noteCreating;
return (
<div>
<Input onChange={this.handleTitleChange}
value={title}
allowClear
placeholder="Title"
/>
<TextArea rows={7}
value={content}
onChange={this.handleContentChange}
allowClear
placeholder="Note content"
style={{
paddingTop: '20px'
}}
/>
</div>
);
}
}
const mapDispatchToProps = (dispatch) => {
return {
actionDoUpdateNoteTitle: (newTitle) => dispatch(actionDoUpdateNoteTitle(newTitle)),
actionDoUpdateNoteContent: (newContent) => dispatch(actionDoUpdateNoteContent(newContent)),
};
};
const mapStateToProps = (state) => ({
noteCreating: state.noteCreating
});
export default connect(mapStateToProps, mapDispatchToProps)(NoteCreateModalBody);<file_sep>/ui/src/js/component/notes/content/note-container.js
import React from 'react';
import {connect} from 'react-redux';
import NoteContainerHeader from "./note-container-header";
import Spinner from "../../common/spinner";
import NoteContainerContent from "./note-container-content";
class NoteContainer extends React.Component {
constructor(props) {
super(props);
}
render() {
let {note, isLoading, hasErrors} = this.props.noteFetch;
if (isLoading) {
return (<Spinner/>);
}
if (!note || hasErrors) {
return '';
}
return (
<div>
<NoteContainerHeader/>
<hr/>
<NoteContainerContent/>
</div>
)
}
}
const mapStateToProps = (state) => ({
noteFetch: state.noteFetch
});
export default connect(mapStateToProps)(NoteContainer);<file_sep>/ui/src/js/component/notes/list/notes-list-container.js
import React, {PureComponent} from 'react';
import NotesListContainerSearch from "./notes-list-container-search";
import NotesPagination from "../pagination/notes-pagination";
import NotesListContainerContent from "./notes-list-container-body";
class NotesListContainer extends PureComponent {
constructor(props) {
super(props);
}
render() {
return (
<div>
<NotesListContainerSearch/>
<NotesListContainerContent/>
<NotesPagination/>
</div>
)
}
}
export default NotesListContainer;<file_sep>/src/main/java/me/kupchenko/client/UserClient.java
package me.kupchenko.client;
import me.kupchenko.auth.service.controller.UserController;
import org.springframework.cloud.openfeign.FeignClient;
@FeignClient("users")
public interface UserClient extends UserController {
}
<file_sep>/ui/src/js/component/notes/create/note-create-modal.js
import {Button, Modal} from 'antd';
import React from "react";
import {actionHideNoteCreateModal} from "../../../actions/note-create";
import {connect} from "react-redux";
import NoteCreateModalBody from "./note-create-modal-body";
import NoteCreateModalSubmit from "./note-create-modal-submit";
class NoteCreateModal extends React.Component {
constructor(props) {
super(props);
}
hideModal = () => {
this.props.actionHideNoteCreateModal();
};
render() {
const {isLoading, modalVisible} = this.props.noteCreate;
return (
<Modal
title="Creating a new note"
visible={modalVisible}
confirmLoading={isLoading}
onCancel={this.hideModal}
footer={[
<Button key="back" onClick={this.hideModal}>
Cancel
</Button>,
<NoteCreateModalSubmit key="submit" isLoading={isLoading}/>
]}
>
<NoteCreateModalBody/>
</Modal>
);
}
}
const mapDispatchToProps = (dispatch) => {
return {
actionHideNoteCreateModal: () => dispatch(actionHideNoteCreateModal())
};
};
const mapStateToProps = (state) => ({
noteCreate: state.noteCreate
});
export default connect(mapStateToProps, mapDispatchToProps)(NoteCreateModal);<file_sep>/ui/src/js/actions/note-select.js
import {
C_NOTE_FETCH_IS_LOADING,
C_NOTE_FETCH_LOAD_FAILURE,
C_NOTE_FETCH_LOAD_SUCCESS,
C_NOTE_FETCH_LOAD_SUCCESS_EMPTY,
C_NOTE_SELECT
} from "./action-type";
import ApiService from "../service/api.service";
export const actionNoteFetchIsLoading = (bool) => ({
type: C_NOTE_FETCH_IS_LOADING,
isLoading: bool
});
export const actionNoteFetchSuccess = (response) => ({
type: C_NOTE_FETCH_LOAD_SUCCESS,
response
});
export const actionNoteFetchSuccessEmpty = () => ({
type: C_NOTE_FETCH_LOAD_SUCCESS_EMPTY
});
export const actionNoteFetchFailure = (errors) => ({
type: C_NOTE_FETCH_LOAD_FAILURE,
errors
});
export const actionNotesSelect = (id) => ({
type: C_NOTE_SELECT,
id
});
export const actionDoNoteFetch = (id) => {
return (dispatch) => {
dispatch(actionNoteFetchIsLoading(true));
ApiService.fetch(`/notes/${id}`).then((json) => {
dispatch(actionNoteFetchSuccess(json));
}).catch(() => {
dispatch(actionNoteFetchFailure())
});
}
};
export const actionDoNoteFetchWithSelect = (id) => {
return (dispatch) => {
dispatch(actionDoNoteFetch(id));
dispatch(actionNotesSelect(id))
}
};
<file_sep>/ui/src/js/reducers/index.js
import {combineReducers} from 'redux';
import {notesSearchReducer} from "./notes-search-reducer";
import {noteFetchReducer} from "./note-fetch-reducer";
import {noteCreateReducer} from "./note-create-reducer";
import {noteUpdateReducer} from "./note-update-reducer";
import {noteEditingReducer} from "./note-editing-reducer";
import {notesSelectReducer} from "./notes-select-reducer";
import {noteCreatingReducer} from "./note-creating-reducer";
export default combineReducers({
notesSearch: notesSearchReducer,
noteFetch: noteFetchReducer,
noteCreate: noteCreateReducer,
noteCreating: noteCreatingReducer,
noteUpdate: noteUpdateReducer,
noteEditing: noteEditingReducer,
selectedNote: notesSelectReducer
});<file_sep>/ui/src/js/component/common/header.js
import React, {PureComponent} from 'react';
import {Layout, Menu} from "antd";
import {actionShowNoteCreateModal} from "../../actions/note-create";
import {connect} from "react-redux";
class NoteHeader extends PureComponent {
constructor(props) {
super(props);
}
showModal = () => {
this.props.actionShowNoteCreateModal();
};
render() {
const {Header} = Layout;
return (
<Header className="header">
<Menu
theme="dark"
mode="horizontal"
style={{
lineHeight: '64px',
}}
selectable={false}
>
<Menu.Item onClick={this.showModal}>Create note</Menu.Item>
</Menu>
</Header>
)
}
}
const mapDispatchToProps = (dispatch) => {
return {
actionShowNoteCreateModal: () => dispatch(actionShowNoteCreateModal())
};
};
export default connect(null, mapDispatchToProps)(NoteHeader);<file_sep>/ui/src/js/component/notes/content/note-container-title.js
import React from 'react';
import {connect} from 'react-redux';
import {Input} from "antd";
import {
actionDisableTitleEditing,
actionDoUpdateNoteTitle,
actionEnableTitleEditing
} from "../../../actions/note-editing";
class NoteContainerTitle extends React.PureComponent {
constructor(props) {
super(props);
}
handleTitleChange = (e) => {
this.props.actionDoUpdateNoteTitle(e.target.value);
};
editTitle = () => {
this.props.actionEnableTitleEditing()
};
disableTitleEditing = () => {
this.props.actionDisableTitleEditing()
};
render() {
const {titleEditable} = this.props;
const title = (this.props.title) ? this.props.title : this.props.note.title;
if (!titleEditable) {
return (
<h1 onClick={this.editTitle}>
{title}
</h1>
);
}
return (
<Input
size="large"
value={title}
onChange={this.handleTitleChange}
onBlur={this.disableTitleEditing}
autoFocus={true}
style={{
maxWidth: '300px'
}}
/>
);
}
}
const mapDispatchToProps = (dispatch) => {
return {
actionDoUpdateNoteTitle: (newTitle) => dispatch(actionDoUpdateNoteTitle(newTitle)),
actionEnableTitleEditing: () => dispatch(actionEnableTitleEditing()),
actionDisableTitleEditing: () => dispatch(actionDisableTitleEditing())
};
};
const mapStateToProps = (state) => ({
titleEditable: state.noteEditing.titleEditable,
title: state.noteEditing.title,
note: state.noteFetch.note
});
export default connect(mapStateToProps, mapDispatchToProps)(NoteContainerTitle);<file_sep>/src/test/java/me/kupchenko/TestUtils.java
package me.kupchenko;
import org.apache.commons.io.IOUtils;
import java.io.IOException;
import java.io.InputStream;
import java.nio.charset.StandardCharsets;
public class TestUtils {
public static <T> String getResourceFileAsString(Class<T> clazz, String filePath) throws IOException {
InputStream resourceAsStream = clazz.getClassLoader().getResourceAsStream(filePath);
return IOUtils.toString(resourceAsStream, StandardCharsets.UTF_8.name());
}
}
<file_sep>/ui/src/js/reducers/note-create-reducer.js
import {
C_NOTE_CREATE_FAILURE,
C_NOTE_CREATE_IS_IN_PROGRESS,
C_NOTE_CREATE_MODAL_HIDE,
C_NOTE_CREATE_MODAL_SHOW,
C_NOTE_CREATE_SUCCESS
} from "../actions/action-type";
const noteCreateInitialState = {
isLoading: false,
hasErrors: false,
isSuccess: false,
errors: null,
note: null,
modalVisible: false
};
export const noteCreateReducer = (state = noteCreateInitialState, action) => {
switch (action.type) {
case C_NOTE_CREATE_IS_IN_PROGRESS: {
return {...state, isSuccess: false, isLoading: action.isLoading, hasErrors: false};
}
case C_NOTE_CREATE_MODAL_SHOW: {
return {
...state,
isLoading: false,
hasErrors: false,
isSuccess: false,
errors: null,
note: null,
modalVisible: true
};
}
case C_NOTE_CREATE_MODAL_HIDE: {
return {
...state,
modalVisible: false,
note: {}
};
}
case C_NOTE_CREATE_SUCCESS: {
return {
...state,
isSuccess: true,
isLoading: false,
hasErrors: false,
modalVisible: false,
note: action.response
};
}
case C_NOTE_CREATE_FAILURE: {
return {...state, isSuccess: false, isLoading: false, hasErrors: true, errors: action.errors};
}
default:
return state
}
};
<file_sep>/src/main/java/me/kupchenko/dto/NotesSearchDto.java
package me.kupchenko.dto;
import lombok.AllArgsConstructor;
import lombok.Data;
import lombok.NoArgsConstructor;
import javax.validation.constraints.Max;
import javax.validation.constraints.Min;
import static me.kupchenko.util.Constant.DEFAULT_ROWS_NUMBER;
import static me.kupchenko.util.Constant.DEFAULT_START_PAGE;
@Data
@AllArgsConstructor
@NoArgsConstructor
public class NotesSearchDto {
private String text = "";
@Min(value = 0, message = "Page number cannot be less then 0")
private Integer page = DEFAULT_START_PAGE;
@Min(value = 1, message = "Number of rows cannot be less then 1")
@Max(value = 100, message = "Number of rows cannot be more then 100")
private Integer rows = DEFAULT_ROWS_NUMBER;
}
<file_sep>/ui/src/js/component/notes/list/notes-list-container-search.js
import React, {PureComponent} from 'react';
import {actionDoNotesSearch, actionUpdateSearchQuery} from "../../../actions/notes-search";
import {connect} from 'react-redux';
import {Input} from "antd";
class NotesListContainerSearch extends PureComponent {
constructor(props) {
super(props);
}
searchNotes = (text) => {
this.props.actionDoNotesSearch(text);
};
updateQuery = (e) => {
this.props.actionUpdateSearchQuery(e.target.value);
};
render() {
const {isLoading} = this.props;
const {Search} = Input;
const query = this.props.notesSearch.query;
return (
<Search
className="search-input"
placeholder="Input search text"
enterButton
size="large"
loading={isLoading}
value={query}
onChange={this.updateQuery}
onSearch={this.searchNotes}
/>
)
}
}
const mapDispatchToProps = (dispatch) => {
return {
actionDoNotesSearch: (e) => dispatch(actionDoNotesSearch(e)),
actionUpdateSearchQuery: (e) => dispatch(actionUpdateSearchQuery(e))
};
};
const mapStateToProps = (state) => ({
notesSearch: state.notesSearch
});
export default connect(mapStateToProps, mapDispatchToProps)(NotesListContainerSearch);<file_sep>/src/main/java/me/kupchenko/config/ResourceServerSecurityConfig.java
package me.kupchenko.config;
import org.springframework.beans.factory.annotation.Value;
import org.springframework.context.annotation.Bean;
import org.springframework.context.annotation.Configuration;
import org.springframework.security.config.annotation.web.builders.HttpSecurity;
import org.springframework.security.config.annotation.web.configuration.EnableWebSecurity;
import org.springframework.security.config.annotation.web.configuration.WebSecurityConfigurerAdapter;
import org.springframework.security.config.annotation.web.configurers.oauth2.server.resource.OAuth2ResourceServerConfigurer;
import org.springframework.security.config.http.SessionCreationPolicy;
import org.springframework.security.oauth2.core.OAuth2TokenValidatorResult;
import org.springframework.security.oauth2.jwt.JwtDecoder;
import org.springframework.security.oauth2.jwt.NimbusJwtDecoder;
import java.security.interfaces.RSAPublicKey;
@Configuration
@EnableWebSecurity
public class ResourceServerSecurityConfig extends WebSecurityConfigurerAdapter {
@Value("${key.location}")
private RSAPublicKey publicKey;
@Override
protected void configure(HttpSecurity http) throws Exception {
http
.authorizeRequests(authorizeRequests ->
authorizeRequests.anyRequest().authenticated()
)
.oauth2ResourceServer(OAuth2ResourceServerConfigurer::jwt)
.sessionManagement(configure -> configure.sessionCreationPolicy(SessionCreationPolicy.STATELESS));
}
@Bean
public JwtDecoder jwtDecoder() {
NimbusJwtDecoder jwtDecoder = NimbusJwtDecoder.withPublicKey(publicKey).build();
jwtDecoder.setJwtValidator(token -> OAuth2TokenValidatorResult.success());
return jwtDecoder;
}
}
<file_sep>/src/main/java/me/kupchenko/dto/AdminNotesResponseDto.java
package me.kupchenko.dto;
import lombok.AllArgsConstructor;
import lombok.Builder;
import lombok.Data;
import lombok.NoArgsConstructor;
import me.kupchenko.auth.service.dto.UserDto;
import java.util.List;
@Data
@Builder
@AllArgsConstructor
@NoArgsConstructor
public class AdminNotesResponseDto {
private UserDto user;
private List<NoteDto> notes;
private ResponsePagination pagination;
}
<file_sep>/build.gradle
plugins {
id 'java'
id 'org.springframework.boot' version '2.2.6.RELEASE'
id 'io.spring.dependency-management' version '1.0.8.RELEASE'
id 'org.hidetake.swagger.generator' version '2.18.1'
}
group = 'me.kupchenko'
version = '0.0.1-SNAPSHOT'
sourceCompatibility = 11
configurations {
compileOnly {
extendsFrom annotationProcessor
}
}
repositories {
mavenLocal()
mavenCentral()
}
dependencies {
implementation 'org.springframework.boot:spring-boot-starter-web'
implementation 'org.springframework.boot:spring-boot-starter-data-jpa'
implementation 'org.springframework.cloud:spring-cloud-starter-openfeign:2.2.1.RELEASE'
implementation 'org.springframework.boot:spring-boot-starter-security'
implementation 'org.springframework.security:spring-security-oauth2-resource-server'
implementation 'org.springframework.security:spring-security-oauth2-jose'
implementation 'mysql:mysql-connector-java'
implementation 'org.hsqldb:hsqldb'
implementation 'me.kupchenko:auth-service-api:1.0'
implementation 'org.mapstruct:mapstruct:1.2.0.Final'
annotationProcessor 'org.mapstruct:mapstruct-processor:1.2.0.Final'
implementation 'io.github.openfeign:feign-okhttp:10.8'
implementation("io.github.openfeign:feign-jackson:10.1.0")
implementation("io.github.openfeign:feign-slf4j:10.1.0")
compileOnly 'org.projectlombok:lombok'
annotationProcessor 'org.projectlombok:lombok'
swaggerUI 'org.webjars:swagger-ui:3.22.1'
testAnnotationProcessor 'org.projectlombok:lombok:1.18.10'
testImplementation 'org.projectlombok:lombok:1.18.10'
testImplementation 'org.springframework.boot:spring-boot-starter-test'
testImplementation 'org.springframework.security:spring-security-test'
}
swaggerSources {
nms {
inputFile = file('src/main/resources/api.spec.yaml')
}
}
task swagger(type: Copy) {
dependsOn generateSwaggerUI
from "$buildDir/swagger-ui-nms"
into "$buildDir/resources/main/static/swagger-ui"
doLast {
delete "$buildDir/swagger-ui-nms"
}
}
processResources.dependsOn swagger
test {
useJUnitPlatform()
}
<file_sep>/ui/src/js/component/notes/pagination/notes-pagination.js
import React from 'react';
import {connect} from "react-redux";
import {actionDoNotesSearch} from "../../../actions/notes-search";
import {Pagination} from "antd";
class NotesPagination extends React.Component {
constructor(props) {
super(props);
}
handlePageChange = (page) => {
const {query} = this.props;
const pageIndex = page - 1;
this.props.actionDoNotesSearch(query, pageIndex)
};
render() {
if (!this.props.pagination) {
return '';
}
const {numFound, page, rows} = this.props.pagination;
const pageIndex = page + 1;
return (
<Pagination
defaultCurrent={pageIndex}
total={numFound}
pageSize={rows}
onChange={this.handlePageChange}
className="pagination"
style={{
paddingTop: 10,
textAlign: 'center'
}}
/>
)
}
}
const mapDispatchToProps = (dispatch) => {
return {
actionDoNotesSearch: (content, page) => dispatch(actionDoNotesSearch(content, page)),
};
};
const mapStateToProps = (state) => ({
query: state.notesSearch.query,
pagination: state.notesSearch.pagination
});
export default connect(mapStateToProps, mapDispatchToProps)(NotesPagination);<file_sep>/src/main/docker/Dockerfile
FROM openjdk:11
WORKDIR /app
ADD build/libs/notes-app-demo*.jar service.jar
ENTRYPOINT ["java", "-jar", "service.jar"]<file_sep>/ui/src/js/reducers/notes-select-reducer.js
import {C_NOTE_SELECT,} from "../actions/action-type";
const notesSelectInitialState = {
id: 0
};
export const notesSelectReducer = (state = notesSelectInitialState, action) => {
switch (action.type) {
case C_NOTE_SELECT: {
return {...state, id: action.id};
}
default:
return state
}
};
<file_sep>/ui/src/js/component/notes/notes-page.js
import React, {PureComponent} from 'react';
import NotesListContainer from "./list/notes-list-container";
import NoteContent from "./content/note-container";
import {Layout} from "antd";
class NotesPage extends PureComponent {
constructor(props) {
super(props);
}
render() {
const {Content, Sider} = Layout;
return (
<Layout style={{marginTop: 5}}>
<Sider width={300} style={{
background: '#fff',
height: '100vh',
padding: 5,
}}>
<NotesListContainer/>
</Sider>
<Layout style={{
padding: '0px 5px 0px',
height: '100vh'
}}>
<Content
style={{
background: '#fff',
padding: 24,
}}
>
<NoteContent/>
</Content>
</Layout>
</Layout>
)
}
}
export default NotesPage;<file_sep>/ui/src/js/actions/note-creating.js
import {
C_NOTE_CREATING_NEW_CONTENT_VALUE,
C_NOTE_CREATING_NEW_TITLE_VALUE,
C_NOTE_CREATING_NEW_VALUES_RESTORE
} from "./action-type";
export const actionDoUpdateNoteTitle = (title) => ({
type: C_NOTE_CREATING_NEW_TITLE_VALUE,
title
});
export const actionDoUpdateNoteContent = (content) => ({
type: C_NOTE_CREATING_NEW_CONTENT_VALUE,
content
});
export const actionRestoreCreateValues = () => ({
type: C_NOTE_CREATING_NEW_VALUES_RESTORE
});<file_sep>/ui/src/js/reducers/note-fetch-reducer.js
import {
C_NOTE_FETCH_IS_LOADING,
C_NOTE_FETCH_LOAD_FAILURE,
C_NOTE_FETCH_LOAD_SUCCESS,
C_NOTE_FETCH_LOAD_SUCCESS_EMPTY,
C_NOTE_UPDATE_IS_IN_PROGRESS
} from "../actions/action-type";
const noteFetchInitialState = {
isLoading: false,
isUpdateInProgress: false,
hasErrors: false,
isSuccess: false,
errors: null,
note: null
};
export const noteFetchReducer = (state = noteFetchInitialState, action) => {
switch (action.type) {
case C_NOTE_FETCH_IS_LOADING: {
return {...state, isLoading: action.isLoading, hasErrors: false};
}
case C_NOTE_FETCH_LOAD_SUCCESS: {
return {
...state,
isSuccess: true,
isUpdateInProgress: false,
isLoading: false,
hasErrors: false,
note: action.response
};
}
case C_NOTE_FETCH_LOAD_SUCCESS_EMPTY: {
return {...state, isSuccess: true, isLoading: false, hasErrors: false, note: null};
}
case C_NOTE_FETCH_LOAD_FAILURE: {
return {...state, isSuccess: false, isLoading: false, hasErrors: true, errors: action.errors};
}
case C_NOTE_UPDATE_IS_IN_PROGRESS: {
return {
...state,
isSuccess: true,
isUpdateInProgress: true,
isLoading: false,
hasErrors: false,
note: state.note
};
}
default:
return state
}
};
<file_sep>/src/main/java/me/kupchenko/dto/CreateNoteDto.java
package me.kupchenko.dto;
import lombok.AllArgsConstructor;
import lombok.Data;
import lombok.NoArgsConstructor;
import javax.validation.constraints.NotEmpty;
@Data
@NoArgsConstructor
@AllArgsConstructor
public class CreateNoteDto {
@NotEmpty
private String title;
private String content;
private String owner;
}
<file_sep>/ui/src/js/utils/response-status.js
export const INTERNAL_SERVER_ERROR = 500;
export const BAD_REQUEST = 400;
export const NOT_FOUND = 404;
export const UNAUTHORIZED = 401;
export const OK = 200;
<file_sep>/ui/src/js/component/auth/success/index.js
import React from 'react';
import queryString from 'query-string'
import {Spin} from "antd";
import {connect} from "react-redux";
import {retrieveToken} from "../../../actions/login";
class AuthSuccess extends React.Component {
constructor(props) {
super(props);
}
componentDidMount() {
const {code} = queryString.parse(this.props.location.search);
console.log("The code is " + code);
this.props.retrieveToken(code)
}
render() {
console.log("Rendering login");
return (
<div style={{
paddingTop: 300,
margin: 'auto',
textAlign: 'center'
}}>
<Spin size="large"/>
</div>
)
}
}
const mapDispatchToProps = (dispatch) => {
return {
retrieveToken: (code) => dispatch(retrieveToken(code))
};
};
export default connect(null, mapDispatchToProps)(AuthSuccess);<file_sep>/ui/src/js/index.js
import "@babel/polyfill";
import React from "react";
import ReactDOM from "react-dom";
import {Provider} from "react-redux";
import configureStore from "./store/configure-store";
import Notes from "./component/notes";
import 'antd/dist/antd.css';
import '@styles/custom.min.css'
import 'ant-design-pro/dist/ant-design-pro.css';
import AuthSuccess from "./component/auth/success";
import {BrowserRouter, Route} from 'react-router-dom'
const store = configureStore();
ReactDOM.render(
<Provider store={store}>
<BrowserRouter>
<Route exact path="/" component={Notes}/>
<Route exact path="/notes" component={Notes}/>
<Route path="/auth/success" component={AuthSuccess}/>
</BrowserRouter>
</Provider>,
document.getElementById('container')
);<file_sep>/src/main/java/me/kupchenko/controller/NoteController.java
package me.kupchenko.controller;
import lombok.AllArgsConstructor;
import lombok.extern.slf4j.Slf4j;
import me.kupchenko.dto.CreateNoteDto;
import me.kupchenko.dto.ExtendedNoteDto;
import me.kupchenko.dto.NoteDto;
import me.kupchenko.dto.NotesResponseDto;
import me.kupchenko.dto.NotesSearchDto;
import me.kupchenko.service.NoteService;
import org.springframework.security.oauth2.server.resource.authentication.JwtAuthenticationToken;
import org.springframework.web.bind.annotation.DeleteMapping;
import org.springframework.web.bind.annotation.GetMapping;
import org.springframework.web.bind.annotation.PatchMapping;
import org.springframework.web.bind.annotation.PathVariable;
import org.springframework.web.bind.annotation.PostMapping;
import org.springframework.web.bind.annotation.PutMapping;
import org.springframework.web.bind.annotation.RequestBody;
import org.springframework.web.bind.annotation.RequestMapping;
import org.springframework.web.bind.annotation.RestController;
import javax.validation.Valid;
import static org.springframework.http.MediaType.APPLICATION_JSON_VALUE;
@Slf4j
@RestController
@AllArgsConstructor
@RequestMapping("/notes")
public class NoteController {
private NoteService noteService;
@GetMapping(value = "/user", produces = APPLICATION_JSON_VALUE)
public NotesResponseDto searchUserNotes(JwtAuthenticationToken auth,
@Valid NotesSearchDto searchDto) {
log.info("Searching notes for user {} by search criteria: {}", auth.getName(), searchDto);
return noteService.searchUserNotes(auth.getName(), searchDto);
}
@GetMapping("/{id:[0-9]+}")
public ExtendedNoteDto getNote(JwtAuthenticationToken auth, @PathVariable Long id) {
return noteService.getNote(auth.getName(), id);
}
@PostMapping(consumes = APPLICATION_JSON_VALUE, produces = APPLICATION_JSON_VALUE)
public NoteDto createNote(JwtAuthenticationToken auth, @Valid @RequestBody CreateNoteDto noteDto) {
noteDto.setOwner(auth.getName());
return noteService.createNote(noteDto);
}
@PutMapping(value = "/{id:[0-9]+}", consumes = APPLICATION_JSON_VALUE)
public NoteDto replaceNote(JwtAuthenticationToken auth, @PathVariable Long id, @Valid @RequestBody NoteDto noteDto) {
noteDto.setId(id);
log.info("Updating note: {}", noteDto);
return noteService.replaceNote(auth.getName(), noteDto);
}
@PatchMapping("/{id:[0-9]+}")
public NoteDto updateNote(@PathVariable Long id, NoteDto noteDto) {
return noteService.updateNote(id, noteDto);
}
@DeleteMapping("/{id:[0-9]+}")
public void deleteNote(JwtAuthenticationToken auth, @PathVariable Long id) {
noteService.deleteNote(auth.getName(), id);
}
}
<file_sep>/ui/src/js/reducers/notes-search-reducer.js
import {
C_NOTES_SEARCH_IS_LOADING,
C_NOTES_SEARCH_LOAD_FAILURE,
C_NOTES_SEARCH_LOAD_SUCCESS,
C_NOTES_SEARCH_UPDATE_QUERY
} from "../actions/action-type";
const notesSearchInitialState = {
isLoading: false,
hasErrors: false,
isSuccess: false,
errors: null,
query: '',
notes: [],
pagination: null
};
export const notesSearchReducer = (state = notesSearchInitialState, action) => {
switch (action.type) {
case C_NOTES_SEARCH_IS_LOADING:
return {
...state,
isLoading: action.isLoading,
hasErrors: false
};
case C_NOTES_SEARCH_LOAD_SUCCESS: {
return {
...state,
isSuccess: true,
isLoading: false,
hasErrors: false,
notes: action.response.notes,
pagination: action.response.pagination,
query: action.searchQuery
};
}
case C_NOTES_SEARCH_LOAD_FAILURE: {
return {
...state,
isSuccess: false,
isLoading: false,
hasErrors: true,
errors: action.errors,
query: action.searchQuery
};
}
case C_NOTES_SEARCH_UPDATE_QUERY: {
return {
...state,
query: action.searchQuery
};
}
default:
return state
}
};<file_sep>/ui/src/js/reducers/note-update-reducer.js
import {C_NOTE_UPDATE_FAILURE, C_NOTE_UPDATE_IS_IN_PROGRESS, C_NOTE_UPDATE_SUCCESS} from "../actions/action-type";
const noteUpdateInitialState = {
isLoading: false,
isUpdateInProgress: false,
hasErrors: false,
isSuccess: false,
errors: null
};
export const noteUpdateReducer = (state = noteUpdateInitialState, action) => {
switch (action.type) {
case C_NOTE_UPDATE_SUCCESS: {
return {...state, isSuccess: true, isLoading: false, hasErrors: false};
}
case C_NOTE_UPDATE_FAILURE: {
return {...state, isSuccess: false, isLoading: false, hasErrors: true, errors: action.errors};
}
case C_NOTE_UPDATE_IS_IN_PROGRESS: {
return {...state, isSuccess: false, isLoading: true, hasErrors: false};
}
default:
return state
}
};
<file_sep>/ui/src/js/reducers/note-creating-reducer.js
import {
C_NOTE_CREATING_NEW_CONTENT_VALUE,
C_NOTE_CREATING_NEW_TITLE_VALUE,
C_NOTE_CREATING_NEW_VALUES_RESTORE
} from "../actions/action-type";
const noteCreatingState = {
title: '',
content: ''
};
export const noteCreatingReducer = (state = noteCreatingState, action) => {
switch (action.type) {
case C_NOTE_CREATING_NEW_TITLE_VALUE:
return {...state, title: action.title};
case C_NOTE_CREATING_NEW_CONTENT_VALUE:
return {...state, content: action.content};
case C_NOTE_CREATING_NEW_VALUES_RESTORE:
return noteCreatingState;
default:
return state
}
}; | 1a2d70167bbf3b1a895780cb3c435a3f5948dd00 | [
"SQL",
"Markdown",
"JavaScript",
"Gradle",
"Java",
"Dockerfile"
] | 40 | Markdown | kupchenko/spring-boot-notes-demo | 1b2529658de5616980f288c7d06e50c72a7c1a76 | 2786931cbec73bf30eb49942c10848689b2f4831 |
refs/heads/main | <repo_name>chiranthancv95/EpAI2.0_CapStone<file_sep>/conftest.py
'''
This is the default script for pytest module to enable the arguments passing to the testing script which test_capstone.py script.
'''
import pytest
def pytest_addoption(parser):
'''
Defines the alternate to parser.add_arguments of parser_args to set the arguments taken from user
'''
parser.addoption('--filename', action="store",type=str, default='/home/cv/workspace2/EpAI2.0_CapStone/file2.csv', help="defines the csv file to be processed")
parser.addoption('--sleep', action="store",type=str, default='1', help="defines the gap between mails sent, default is set to 30secs")
parser.addoption('--path', action="store",type=str, default='/home/cv/workspace2/EpAI2.0_CapStone/certificates', help="defines the path where the certificates are to be stored")
parser.addoption('--certificate_file',action="store", type=str, default="/home/cv/workspace2/EpAI2.0_CapStone/certificate.jpg", help="defines the path where the certificate template is stored for loading")
parser.addoption('--single_mode',action="store",type=str,default='1', help="If single_mode is set to 1, then we ask name, course name, score and total from the user, else 2 is for sending multiple")
parser.addoption('--name',type=str, action="store",help="Specify name to be written on certificate")
parser.addoption('--course_name', action="store",type=str, default="EPAI 2.0 Course", help="Specify the course name to be written on the certificate")
parser.addoption('--score',type=str, action="store",help="Specify the score to be written on the certificate")
parser.addoption('--total',type=str, action="store",default='100', help="Specify the total score to be written on the certificate")
parser.addoption('--email', type=str, action="store",help="Specify the email to be sent with the certificate")
parser.addoption('--sender_email', action="store",type=str, default='chirant<EMAIL>', help="defines the email through which certificates are sent out")
parser.addoption('--passwords', type=str, help="Defines the password for th emailer's account")
@pytest.fixture
def params(request):
'''
Defines all the arguments taken to respective attributes of params dictionary.
'''
params = {}
params['filename'] = request.config.getoption('--filename')
params['sleep'] = request.config.getoption('--sleep')
params['path'] = request.config.getoption('--path')
params['certificate_file'] = request.config.getoption('--certificate_file')
params['single_mode'] = request.config.getoption('--single_mode')
params['name'] = request.config.getoption('--name')
params['course_name'] = request.config.getoption('--course_name')
params['score'] = request.config.getoption('--score')
params['total'] = request.config.getoption('--total')
params['email'] = request.config.getoption('--email')
params['sender_email'] = request.config.getoption('--sender_email')
params['passwords'] = request.config.getoption('--passwords')
if params['name'] is None or params['course_name'] is None or params['score'] is None or params['total'] is None or params['email'] is None or params['sender_email'] is None:
pytest.skip()
return params<file_sep>/utils_package/parser_args.py
import os
import argparse
# required defines a mandatory argument
# default defines a default value if not specified
parser = argparse.ArgumentParser()
parser.add_argument('-f', '--filename', type=str, default='/home/cv/workspace2/EpAI2.0_CapStone/file2.csv', help="defines the csv file to be processed")
parser.add_argument('-s', '--sleep', type=str, default='1', help="defines the gap between mails sent, default is set to 30secs")
parser.add_argument('-p', '--path', type=str, default='/home/cv/workspace2/EpAI2.0_CapStone/certificates', help="defines the path where the certificates are to be stored")
parser.add_argument('-c', '--certificate_file', type=str, default="/home/cv/workspace2/EpAI2.0_CapStone/certificate.jpg", help="defines the path where the certificate template is stored for loading")
parser.add_argument('-sm','--single_mode',type=str,default='1', help="If single_mode is set to 1, then we ask name, course name, score and total from the user, else 2 is for sending multiple")
parser.add_argument('-na','--name',type=str, help="Specify name to be written on certificate")
parser.add_argument('-cn','--course_name', type=str, default="EPAI 2.0 Course", help="Specify the course name to be written on the certificate")
parser.add_argument('-sc','--score',type=str, help="Specify the score to be written on the certificate")
parser.add_argument('-tot','--total',type=str, default='100', help="Specify the total score to be written on the certificate")
parser.add_argument('-em', '--email', type=str, help="Specify the email to be sent with the certificate")
parser.add_argument('-se_em', '--sender_email', type=str, default='<EMAIL>', help="defines the email through which certificates are sent out")
parser.add_argument('-pa','--passwords', type=str, help="Defines the password for th emailer's account")
args = parser.parse_args()
sleep_timer = args.sleep
path = args.path
filename = args.filename
certificate_file = args.certificate_file
single_mode = args.single_mode
name =args.name
course_name = args.course_name
score = args.score
total = args.total
email = args.email
sender_email = args.sender_email
passwords = args.passwords
<file_sep>/utils_package/send_mail_smtp.py
'''
This code is to send mails using smtplib library.
'''
import os
import img2pdf
from PIL import Image
import base64
from utils_package.parser_args import filename, path, certificate_file, sleep_timer,name, score, total, email, single_mode, course_name, sender_email, passwords
from utils_package.decorators import *
import smtplib
from email.mime.multipart import MIMEMultipart
from email.mime.text import MIMEText
from email.mime.base import MIMEBase
from email import encoders
from email_validator import validate_email, EmailNotValidError
import time
def check_valid_email(email):
'''
This method checks for the valid email address and raises an error if not.
'''
try:
# Validate.
valid = validate_email(email)
print("valid email")
# Update with the normalized form.
email = valid.email
return True
except EmailNotValidError as e:
# email is not valid, exception message is human-readable
print(str(e))
return False
@timeit
@func_name
@is_connected
@logged
#@authenticatedOrNot
def mailer_smtp(info):
'''
This method is the main method for sending mails to the candidates to whom the user needs to send emails with the attached certitficates.
'''
#Waiting to send mail
print(f"Waiting for {sleep_timer} seconds to send next mail")
time.sleep(int(sleep_timer))
a=info.split(',')
index,name,score,mailid=a[0],a[1],a[2],a[3]
total=str(100)
try:
total = a[4]
except Exception as e:
pass
name = name.title()
name="".join(name.split())
fromaddr = sender_email
toaddr = mailid
# instance of MIMEMultipart
msg = MIMEMultipart()
# storing the senders email address
msg['From'] = fromaddr
# storing the receivers email address
msg['To'] = toaddr
# storing the subject
msg['Subject'] = "Certificate of Course Completion_new7"
# string to store the body of the mail
#body = "Body_of_the_mail"
body=(f'Dear {name},\nCongratulations! You have cleared the EPAI 2.0 Course with {score} marks out of {total} marks!\n We are excited to share the attached Award of Excellence for your performance!\n Regards')
content_str = body
# attach the body with the msg instance
msg.attach(MIMEText(body, 'plain'))
# open the file to be sent
filename = name+'_certificate.pdf'
attachment = open(os.path.join(path, name+'_certificate.pdf'), 'rb')
# instance of MIMEBase and named as p
p = MIMEBase('application', 'octet-stream')
# To change the payload into encoded form
p.set_payload((attachment).read())
# encode into base64
encoders.encode_base64(p)
p.add_header('Content-Disposition', "attachment; filename= %s" % filename)
# attach the instance 'p' to instance 'msg'
msg.attach(p)
# creates SMTP session
s = smtplib.SMTP('smtp.gmail.com', 587)
s.ehlo()
# start TLS for security
s.starttls()
# Authentication
s.login(fromaddr, passwords)
# Converts the Multipart msg into a string
text = msg.as_string()
if check_valid_email(toaddr) == True:
# sending the mail
s.sendmail(fromaddr, toaddr, text)
# terminating the session
s.quit()
return content_str<file_sep>/main.py
'''
This is the main script that runs the app.
As mentioned in the readme, there are two modes available to the user.
If selected '1', then it enables bulk mailing.
If selected '2' or any other number, then it enables single mailing option to the user.
'''
import os
import argparse
from utils_package.certificate_creator import print_certificate
from utils_package.decorators import timeit,func_name
from utils_package.iterator_class import FileIter
#from utils_package.send_mail import mailer
from utils_package.send_mail_smtp import mailer_smtp
from utils_package.parser_args import filename, path, sleep_timer,name, score, total, email, single_mode,passwords
if single_mode == '1':
try:
items = FileIter(filename)
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
print(next(csv_info_iterator))
cert_map_object = map(print_certificate,csv_info_iterator)
print(list(cert_map_object))
except IndexError as error:
#runs the sending email part since the IndexError is a known error which occurs during the map call
print(error)
try:
# Sending the mails now
mail_items = FileIter(filename)
mail_info= "".join(next(mail_items))
mail_info_list = mail_info.split('\n')
mail_info_iterator = iter(mail_info_list)
print(next(mail_info_iterator))
mail_map_object = map(mailer_smtp,mail_info_iterator)
print(list(mail_map_object))
except IndexError as error:
print(error)
else:
index = '1'
info_obj = str(index)+','+ str(name) + ','+ str(score) +' ,'+ str(email)+','+str(total)
#info_obj = [str(index,name,score,email)]
#index,name,score,mail=a[0],a[1],a[2],a[3]
#info_obj.append(index,name,score,email)
print_certificate(info_obj)
mailer_smtp(info_obj)
print("Finished sending all mails with certificates to the qualified candidates")
<file_sep>/requirements.txt
opencv-python
pytest==6.2.1
email_validator==1.1.1
urllib3==1.25.9
tcp-latency==0.0.10
pyspeedtest==1.2.7
speedtest-cli==2.1.3
pillow==7.2.0
img2pdf==0.4.0
sendgrid==6.7.0
<file_sep>/utils_package/iterator_class.py
'''
This is the custom iterator class file where the data is read from csv file using as a generator.
'''
import cv2
import csv
class FileIter:
'''
This custom iterator yields a generator object of the csv file given by user.
'''
def __init__(self, filename):
file = open(filename)
self.reader = file.read()
def __iter__(self):
return self
def __next__(self):
yield self.reader<file_sep>/utils_package/__init__.py
from . import decorators
from . import parser_args
from . import certificate_creator
from . import iterator_class
from . import send_mail_smtp
# print("In __init_.py script")
<file_sep>/test_capstone.py
import importlib
import os
import sys
import argparse
from email_validator import validate_email, EmailNotValidError
import re
import urllib.request
import csv
from datetime import datetime
from glob import glob
import os.path
from os import path
import shutil
import pytest
import inspect
import main
from utils_package import decorators
from utils_package import send_mail_smtp
from tcp_latency import measure_latency
import pyspeedtest
import speedtest
import utils_package
from utils_package import iterator_class
from utils_package.certificate_creator import print_certificate
from utils_package.iterator_class import FileIter
from utils_package.send_mail_smtp import mailer_smtp
import time
import urllib.request
from functools import wraps
def test_params(params):
print(params)
def test_connected(params):
if urllib.request.urlopen('http://google.com'):
# connect to the host -- tells us if the host is actually
# reachable
print ("Connected to internet")
else:
print ("Not connected to internet")
README_CONTENT_CHECK_FOR = ['iterator_class', 'iterator_class', 'send_mail', 'send_mail_smtp', 'decorators','certificate_creator','conftest','test_capstone']
def test_readme_exists():
assert os.path.isfile("README.md"), "README file missing!"
def test_readme_contents():
readme = open("README.md", "r", encoding="utf-8")
readme_words = readme.read().split()
readme.close()
assert len(readme_words) >= 500, "Make your README interesting! Add atleast 500 words"
def test_readme_proper_description():
READMELOOKSGOOD = True
f = open("README.md", "r", encoding="utf-8")
content = f.read()
f.close()
for c in README_CONTENT_CHECK_FOR:
if c not in content:
READMELOOKSGOOD = False
pass
else:
print(c)
assert READMELOOKSGOOD == True, "You have not described all the functions well in your README file"
def test_readme_file_for_formatting():
f = open("README.md", "r", encoding="utf-8")
content = f.read()
f.close()
assert content.count("#") >= 5
def test_indentations():
''' Returns pass if used four spaces for each level of syntactically \
significant indenting.'''
lines = inspect.getsource(main)
spaces = re.findall('\n +.', lines)
for space in spaces:
assert len(space) % 4 == 2, "Your script contains misplaced indentations"
assert len(re.sub(r'[^ ]', '', space)) % 4 == 0, "Your code indentation does not follow PEP8 guidelines"
def test_function_name_had_cap_letter():
functions = inspect.getmembers(main, inspect.isfunction)
for function in functions:
assert len(re.findall('([A-Z])', function[0])) == 0, "You have used Capital letter(s) in your function names"
#Authenticate
def set_password(password=None):
'''Sets the default password if no values are supplied
:args password: str
:returns inner: closure function
'''
def inner():
nonlocal password
if password == None:
password = '<PASSWORD>'
return hash(password)
return inner
def authenticate(fn):
'''Decorator to authenticate before accessing any functions'''
def check_creds(user_password, in_password, *args, **kwargs):
if user_password() != hash(in_password):
print('Password Mismatch')
else:
print('user Authenticated')
print(f"Function {fn.__name__} is called")
return fn(*args)
return check_creds
@authenticate
def add_auth(*args):
return sum(args)
def test_authentication():
'''Checks the authentication by setting a password and calling the function'''
user_password = set_password()
assert add_auth(user_password, '<PASSWORD>', 1,2) == 3
#assert add_auth(user_password, '<PASSWORD>', 1,2) == None
#email = "<EMAIL>"
def test_valid_email(params):
try:
# Validate.
valid = validate_email(params['email'])
print("valid email")
# Update with the normalized form.
email = valid.email
except EmailNotValidError as e:
# email is not valid, exception message is human-readable
print(str(e))
regex = "^[a-z0-9]+[\._]?[a-z0-9]+[@]\w+[.]\w{2,3}$"
def test_emailid(params):
if(re.search(regex,params['email'])):
print("Valid Email")
else:
print("Invalid Email")
def test_internet_connection():
if urllib.request.urlopen('http://google.com'):
# connect to the host -- tells us if the host is actually
# reachable
print("Connected to internet")
else:
print("Not connected to internet")
# This test has been commented since I'm getting the error:smtplib.SMTPSenderRefused, since gmail allows sending only upto 100 mails at a time.
# Link - https://support.google.com/a/answer/166852?hl=en
# However it works properly when tested locally.
# def test_send_1000_mails(params):
# count = 0
# try:
# items = FileIter(params['filename'])
# csv_info= "".join(next(items))
# csv_info_list = csv_info.split('\n')
# csv_info_iterator = iter(csv_info_list)
# next(csv_info_iterator)
# cert_map_object = map(print_certificate,csv_info_iterator)
# list(cert_map_object)
# except IndexError as error:
# #runs the sending email part since the IndexError is a known error which occurs during the map call
# print(error)
# while count != 10:
# try:
# mail_items = FileIter(params['filename'])
# mail_info= "".join(next(mail_items))
# mail_info_list = mail_info.split('\n')
# mail_info_iterator = iter(mail_info_list)
# l = len(mail_info_list)
# next(mail_info_iterator)
# mail_map_object = map(mailer_smtp,mail_info_iterator)
# list(mail_map_object)
# except IndexError as error:
# print(error)
# count += l
# print("Sent above 1000 mails for testing purposes")
def test_import_datetime_module():
package_name = 'datetime'
if package_name in sys.modules:
print(f'{package_name} is a imported successfully')
def test_is_a_package():
package_name = 'utils_package'
if package_name in sys.modules:
print(f'{package_name} is a imported successfully')
def is_char(s_obj):
a=s_obj.split(',')
name=a[1]
if name.isalpha():
pass
#print("It's all letters here")
else:
print("Some intruder other than character")
def test_char_only(params):
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
next(csv_info_iterator)
test_char_obj = map(is_char,csv_info_iterator)
def is_first_letter_capitalized(s_obj):
a=s_obj.split(',')
name=a[1]
first_let = name.split(' ')[0]
#second_let = name.split(' ')[1]
if first_let.isupper():
pass
#print("First letter is capital here")
else:
print("First letter is not capital here")
def test_char_first_letter_capitalized(params):
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
next(csv_info_iterator)
test_char_obj = map(is_first_letter_capitalized,csv_info_iterator)
def is_second_letter_capitalized(s_obj):
a=s_obj.split(',')
name=a[1]
second_let = name.split(' ')[1]
#second_let = name.split(' ')[1]
if second_let.isupper():
pass
#print("First letter is capital here")
else:
print("Second letter is not capital here")
def test_char_second_letter_capitalized(params):
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
next(csv_info_iterator)
test_char_obj = map(is_second_letter_capitalized,csv_info_iterator)
def is_numeric(s_obj):
a=s_obj.split(',')
name=a[1]
if not name.isnumeric():
pass
#print("It's not all letters here")
else:
print("Some intruder other than character")
def test_not_numeric_only(params):
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
next(csv_info_iterator)
test_char_obj = map(is_numeric,csv_info_iterator)
def test_certificate_file(params):
os.path.exists(params['certificate_file'])
def test_git_folder(params):
os.path.exists(".git")
def test_csv_header(params):
with open(params['filename'], 'r') as csvfile:
sample = csvfile.read()
has_header = csv.Sniffer().has_header(sample)
csvfile.close()
def test_csv_format(params):
with open(params['filename'], 'r') as csvfile:
sample = csvfile.read()
assert (len(sample.split('\n')[0].split(','))) == 4, "Not enough data or not in correct format"
assert (len(sample.split('\n')[1].split(','))) == 4, "Not enough data or not in correct format"
csvfile.close()
def test_csv_file_empty(params):
with open('file1.csv', 'r') as csvfile:
csv_dict = [row for row in csv.DictReader(csvfile)]
if len(csv_dict) == 0:
print('csv file is empty')
csvfile.close()
def test_csv_file_extension(params):
assert params['filename'][-4:] == '.csv', "Not a csv file"
def test_certificate_extension(params):
assert params['filename'][-4:] == '.csv', "Not a csv file"
def test_datetime_format(params):
#date_object = datetime(2021, 4, 24)
#print(date_string)
#date_str = date_object.strftime("%d")+"th " +date_object.strftime("%B")+" "+date_object.strftime("%Y")
info_obj = str('index')+','+ str('name') + ','+ str('score') +' ,'+ str('email')+','+str('total')
date_str=print_certificate(info_obj)
assert date_str == '24th April 2021', "Date not in the right format"
def test_certificate_folder(params):
assert path.isdir(params['path']) == True, "Folder to save is not a directory"
def test_pdf_saving_working(params):
try:
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
next(csv_info_iterator)
cert_map_object = map(print_certificate,csv_info_iterator)
list(cert_map_object)
except IndexError as error:
#runs the sending email part since the IndexError is a known error which occurs during the map call
pass
#print(error)
#pdf_glob = [x for x in os.listdir() if x.endswith(".txt")]
pdf_glob = glob(params['path']+"/*.pdf")
assert len(pdf_glob) != 0, "Pdfs are not generated as required"
# Remove the specified
# file path
try:
shutil.rmtree(params['path'])
except OSError as e:
print ("Error: %s - %s." % (e.filename, e.strerror))
#print("File path can not be removed")
def test_img_saving_working(params):
try:
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
next(csv_info_iterator)
cert_map_object = map(print_certificate,csv_info_iterator)
list(cert_map_object)
except IndexError as error:
#runs the sending email part since the IndexError is a known error which occurs during the map call
pass
#print(error)
img_glob = glob(params['path']+"/*.img")
assert img_glob != 0, "Images are not generated as required"
# Remove the specified
# file path
try:
shutil.rmtree(params['path'])
except OSError as e:
print ("Error: %s - %s." % (e.filename, e.strerror))
def test_import_external_packages():
package_list = ['pytest','datetime','email_validator','PIL','img2pdf','smtplib','csv']
for package_name in package_list:
if package_name in sys.modules:
print(f'{package_name} external package is imported successfully')
def test_no_for_loop():
# checks whether for loop is used anywhere in the code
lines = inspect.getsource(main)
#spaces = re.findall(lines)
for_loops = re.findall('.*for.*', lines)
assert len(for_loops) == 0, "Your script contains for loops in the code"
def test_no_while_loop():
# checks whether for loop is used anywhere in the code
lines = inspect.getsource(main)
#spaces = re.findall(lines)
while_loops = re.findall('.*while.*', lines)
assert len(while_loops) == 0, "Your script contains while loops in the code"
def test_no_list_comprehension():
# checks whether for loop is used anywhere in the code
lines = inspect.getsource(main)
#spaces = re.findall(lines)
list_comprehension = re.findall('.[*].*', lines)
assert len(list_comprehension) == 0, "Your script contains list_comprehension in the code. Invalid"
def test_number_of_decorators():
# checks whether for loop is used anywhere in the code
lines = inspect.getsource(decorators)
#spaces = re.findall(lines)
number_of_decorators = re.findall('.@.*', lines)
assert len(number_of_decorators) >= 0, "Your script does not contain enough decorators in the code"
def test_sleep_applied():
lines = inspect.getsource(send_mail_smtp)
#spaces = re.findall(lines)
sleep_applied = re.findall('.sleep.*', lines)
assert len(sleep_applied) >= 0, "Your script contains sleep in the code"
def test_requirements_txt_exists():
assert os.path.isfile("requirements.txt"), "README file missing!"
def test_iter_class_an_iterable(params):
#iter, any_check = tee(FileIter(params['filename']))
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
csv_info_iterator = iter(csv_info_list)
assert next(csv_info_iterator) is not None, "Not an iterator"
def test_python_version():
assert sys.version_info >= (3, 4), "Python version not satisfied. Please upgrade Python version"
def test_yield_exists():
lines = inspect.getsource(iterator_class)
#spaces = re.findall(lines)
yield_exists = re.findall('.yield.*', lines)
assert len(yield_exists) >= 0, "Your script does not contain yield in the code"
def test_csv_load(params):
items = FileIter(params['filename'])
csv_info= "".join(next(items))
csv_info_list = csv_info.split('\n')
assert len(csv_info_list) >0, "csv file not loaded properly or empty file given"
def test_certificate_names(params):
info_obj = str('index')+','+ str('Name') + ','+ str('score') +' ,'+ str('email')+','+str('total')
date_str=print_certificate(info_obj)
assert os.path.exists(os.path.join(params['path'], 'Name'+'_certificate.pdf')) == True, "The name of the certificate file and name of candidate not matching"
def test_score_above_threshold(params):
assert int(params['score']) >= 70, "The score is not eligible for certificate"
def test_score_is_int(params):
assert isinstance(int(params['score']), int), 'Score parameter of wrong type!'
def test_total_is_int(params):
assert isinstance(int(params['total']), int), 'Score parameter of wrong type!'
def test_score_is_not_decimal(params):
assert not isinstance(int(params['score']), float), 'Score parameter is not of decimal type!'
def test_total_is_not_decimal(params):
assert not isinstance(int(params['total']), float), 'Total parameter is not of decimal type!'
def test_latency():
latency = int(measure_latency(host='google.com')[0])
assert (latency) <= 1000, "The latency is too high"
print("Measured Latency: ", latency)
def test_ping():
test = pyspeedtest.SpeedTest("www.google.com")
ping = int(test.ping())
assert (ping) <= 1000, "The ping is too high"
print("ping: ",ping)
def test_download_speed():
test = pyspeedtest.SpeedTest("www.google.com")
download_speed = int(test.download())
assert (download_speed) > 1, "The download speed is too slow"
print("download_speed: ", download_speed)
def test_upload_speed():
st = speedtest.Speedtest()
upload_speed = int(st.upload())
assert (upload_speed) > 1, "The upload speed is too slow"
print("upload_speed: ",upload_speed)
def test_doc_string(params):
assert main.__doc__ is not None, "Please include a doc string to the code"
<file_sep>/utils_package/certificate_creator.py
'''
This script generates the certificates of the candidates in a specific format and stores them.
'''
import cv2
import img2pdf
from PIL import Image
from datetime import datetime
import os
from utils_package.parser_args import filename, path, certificate_file, sleep_timer,name, score, total, email, single_mode, course_name, passwords
from utils_package.decorators import *
@timeit
@func_name
def print_certificate(info_obj):
'''
This function helps in gathering information from the csv file and aprropriately generating the certificates of the candidates.
Then stores it as .jpg file extension and later converted to .pdf file which can then be exported with the mail.
'''
a=info_obj.split(',')
index,name,score,mail=a[0],a[1],a[2],a[3]
# converting name to standard capitalized format
name_on_certificate = name.title()
name="".join(name_on_certificate.split())
img=cv2.imread(certificate_file)
# font
font = cv2.FONT_HERSHEY_SIMPLEX
# fontScale
fontScale = 1
color = (0, 0, 0)
# Line thickness of 2 px
thickness = 2
#datetime(year, month, day)
date_object = datetime(2021, 4, 24)
#print(date_string)
date_str = date_object.strftime("%d")+"th " +date_object.strftime("%B")+" "+date_object.strftime("%Y")
#positions
course_position = (370,299)
image = cv2.putText(img, course_name, course_position, font,
fontScale, color, thickness, cv2.LINE_AA)
name_position = (400,440)
image = cv2.putText(img, name_on_certificate, name_position, font,
fontScale, color, thickness, cv2.LINE_AA)
date_position = (200,525)
image = cv2.putText(img, date_str, date_position, font,
0.7, color, 2, cv2.LINE_AA)
signature_position = (630,525)
image = cv2.putText(img, '<NAME>', signature_position, font,
0.7, color, thickness, cv2.LINE_AA)
# Create the directory only if it does not exist already
try:
os.mkdir(path)
except OSError as error:
print(error)
print(os.path.join(path, name+'_certificate.jpg'))
cv2.imwrite(os.path.join(path, name+'_certificate.jpg'), img)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
im_pil = Image.fromarray(img)
pdf_bytes = img2pdf.convert(os.path.join(path, name+'_certificate.jpg'))
file1 = open(os.path.join(path, name+'_certificate.pdf'), "wb")
# writing pdf files with chunks
file1.write(pdf_bytes)
# closing pdf file
file1.close()
print("Printing: ", index,name,score,mail)
return date_str<file_sep>/utils_package/decorators.py
'''
This script contains all the decorators used in other programs.
'''
import time
import urllib.request
from functools import wraps
from datetime import datetime, timezone
def timeit(f):
'''
This decorator prints the time that the code too execute a certain task.
'''
def timed(*args, **kw):
ts = time.time()
result = f(*args, **kw)
te = time.time()
print('took: %2.4f sec' % \
(te-ts))
return result
return timed
def func_name(f):
'''
This decorator prints out the function name and other parameters
'''
def func_n(*args, **kw):
result = f(*args, **kw)
print('func:%r args:[%r, %r] '% \
(f.__name__, args, kw,))
return result
return func_n
def logged(fn):
'''
This is a logger decorator which logs all the function parameters with time duration.
'''
@wraps(fn)
def inner(*args, **kwargs):
run_dt = datetime.now(timezone.utc)
result = fn(*args, **kwargs)
print('{0}: called {1}'.format(fn.__name__, run_dt))
return result
return inner
def is_connected(fn):
'''
This decorator checkcs whether the internet connection is present or not.
'''
@wraps(fn)
def connected(*args, **kwargs):
result = fn(*args, **kwargs)
if urllib.request.urlopen('http://google.com'):
# connect to the host -- tells us if the host is actually
# reachable
print ("Connected to internet")
else:
print ("Not connected to internet")
return connected
def authenticatedOrNot(auth):
'''
This decorator ensures that the function or user is authenticated to run the program.
'''
def dec(fn):
def inner(*args, **kwargs):
return fn(*args, **kwargs)
return inner
if (auth):
return dec
else:
print("You are not authenticated")
<file_sep>/README.md
# EpAI CapStone Project
# Mailer App
## Problem Statement
You are going to build a Python app that can:<br>
read a CSV file that has NAME, SCORE and EMAIL (10+, use the fact that <EMAIL> and <EMAIL> are the same email ids, so
you can use your own for testing)<br>
loads this image (Links to an external site.) (you can use OpenCV, PIL, or anything else you want)<br>
copies each name from the CSV file and adds it to the image (again use OpenCV and font support there to write it on the image) along with other details.<br> The certificate should finally read:<br>
Award of Excellence for "COURSE NAME" Awarded to "NAME". Date 24th April 2021. Signature "FIXED NAME". You must make sure that the items are placed exactly at the location they should be and with the appropriate font size. <br>
learn how to send emails using Python using this tutorial (Links to an external site.). <br>
Then compose an email that:<br>
will send the final certificate to the email address<br>
reads another file where this email content is written:<br>
Dear NAME,<br>
Congratulations! You have cleared the COURSE NAME with SCORE marks out of TOTAL marks!<br>
We are excited to share the attached Award of Excellence for your performance!<br>
Regards<br>
These are the things that are expected:<br>
your code is well documented<br>
your code has at least 50 test cases that test various problems that might be there. Some tests that are expected:<br>
regex check for emails<br>
can handle 1000+ emails without getting crashed <br>
check if names have only characters<br>
checks if internet connection exists<br>
checks if all required (including external) packages are installed<br>
NAME, COURSE NAME, SCORE, TOTAL are variables and can be changed while calling your Python App<br>
you are using at least 2 different custom modules<br>
you are using at least 1 or more packages<br>
you are NOT using while or for loop anywhere<br>
you are use generators and NOT list comprehension anywhere<br>
you are using your OWN Iterator class for creating your name, email and scores database.<br>
you are using:<br>
named tuples<br>
Python's datetime module<br>
at least 2 decorators (eg, a decorator that "knows" if internet connection is there, etc).<br>
the certificates you attach have a proper name (not image1, etc)<br>
all certificates are finally stored in a folder<br>
checks internet connection before doing anything<br>
checks if the external packages that are required (like PIL, OpenCV are already installed)<br>
checks if the certificate already exists in the folder, and if yes, then do not create it again<br>
emails must be sent out at a gap of 30-60 seconds (selectable while running the app)<br>
your code automatically converts names to their "proper form", e.g. <NAME> to <NAME><br>
your app can be called from the command line while providing the variable details as well as the CSV file location<br>
your app first runs "some of the tests" before actually sending emails<br>
**In your final submission, you need to send emails to yourself 10+ times, then add the 10+ certificates in the folder on Github along with the code, as well as the screenshot of your Gmail box showing these 10+ email along with the time stamp<br>**
## Introduction
This is a repo which helps the user to generate certificates with the candidate's name, marks scored, total marks, course name, date issued, signature of the instructor. <br>
The user can then send the certificates to the candidates through mail.<br>
The user is given option whether to use a csv file with format of 'name, score, total' to send mails in bulk.<br>
Otherwise, the user can send a single mail using the command line arguments.<br>
The usage and other instructions are provided below.<br>
## Installation and Requirements
opencv-python
pytest
email_validator
urllib3
tcp-latency
pyspeedtest
speedtest-cli
pillow
img2pdf
sendgrid
Note - All these can be installed using the requirements.txt<br>
Command -
'''shell
pip install -r requirements.txt
'''
## Usage
Two Modes are provided for the user which can be used on the need -
1. Bulk Mode<br>
2. Single Mode<br>
1. Bulk Mode -
Using this mode, the user can send multiple mails to the candidates whose information is given in the form of a csv file.<br>
For using this mode, please give a csv file which is in the format of 'name, score, email'<br>
Command -
```shell
python main.py --filename <file-path-for-csv> --sleep <sleep-timer-in-int> --path <path-to-store-certificates> --certificate_file <path-for-certificate-file> --single_mode <bulk-mode-or-single-mode-selector> --sender_email <mailer_address> --password <<PASSWORD>>
```
Sample Command -
```shell
python main.py --filename /home/cv/workspace2/EpAI2.0_CapStone/file2.csv --sleep 0 --path /home/cv/workspace2/EpAI2.0_CapStone/certificates --certificate_file /home/cv/workspace2/EpAI2.0_CapStone/certificate.jpg --single_mode 1
```
2. Single Mode -
Using this mode, the user can send a single mail to the candidate whose information is to be given as a command line argument instead of a csv file <br>
Command -
```shell
python main.py --filename <file-path-for-csv> --sleep <sleep-timer-in-int> --path <path-to-store-certificates> --certificate_file <path-for-certificate-file> --single_mode <bulk-mode-or-single-mode-selector> --name <string> --course_name <string> --score <int> --total <int> --email <email_address> --sender_email <mailer_address> --password <<PASSWORD>>
```
Sample Command -
```shell
python main.py --filename /home/cv/workspace2/EpAI2.0_CapStone/file2.csv --sleep 0 --path /home/cv/workspace2/EpAI2.0_CapStone/certificates --certificate_file /home/cv/workspace2/EpAI2.0_CapStone/certificate.jpg --single_mode 2 --name chiru --course_name new1111 --score 80 --total 101 --email <EMAIL> --sender_email <EMAIL> --password <PASSWORD>
```
## Command-line-arguments
**--filename**
defines the csv file to be processed
**--sleep**
defines the gap between mails sent, default is set to 30secs
**--path**
defines the path where the certificates are to be stored
**--certificate_file**
defines the path where the certificate template is stored for loading
**--single_mode**
If single_mode is set to 1, then we ask name, course name, score and total from the user, else 2 is for sending multiple
**--name**
Specify name to be written on certificate
**--course_name**
Specify the course name to be written on the certificate
**--score**
Specify the score to be written on the certificate
**--total**
Specify the total score to be written on the certificate
**--email**
Specify the email to be sent with the certificate
**--sender_email**
defines the email through which certificates are sent out
**--password**
Defines the password for th emailer's account
## Testing of Code
The code contains around 50 test codes to ensure the standard quality of the code.<br>
The codes are written using the pytest module.<br>
The file named "test_capstone.py" contains all the test cases.<br>
The user can test out the code using the following commands for both the modes.<br>
Command -
Pytest Command for multiple -
```shell
pytest -s test_capstone.py --filename /home/cv/workspace2/EpAI2.0_CapStone/file2.csv --sleep 0 --path /home/cv/workspace2/EpAI2.0_CapStone/certificates --certificate_file /home/cv/workspace2/EpAI2.0_CapStone/certificate.jpg --single_mode 1 --name chiru --course_name new1111 --score 80 --total 101 --email <EMAIL> --sender_email <EMAIL> --password <PASSWORD>
```
Pytest Command for single mail -
```shell
pytest -s test_capstone.py --filename /home/cv/workspace2/EpAI2.0_CapStone/file2.csv --sleep 0 --path /home/cv/workspace2/EpAI2.0_CapStone/certificates --certificate_file /home/cv/workspace2/EpAI2.0_CapStone/certificate.jpg --single_mode 2 --name chiru --course_name new1111 --score 80 --total 101 --email <EMAIL> --sender_email <EMAIL> --password <PASSWORD>
```
## Module Explanation
### iterator_class
This module is to create a generator using custom iterator class.
### parser_args
This module is used to get all the command-line arguments from the user.
### send_mail
This module is used to send mails to candidates using a third-party offering from sendgrid.
### send_mail_smtp
This module is used to send mails with certificates to candidates using the python's built-in library.(It's freeeee)
### decorators
This module is used to create the decorators used for better functioning of other modules.
### certificate_creator
This module is used to create certificates with the given information from the user in both .jpg and .pdf file which can be attached while sending the mail.
### conftest
This module is used to fix the arguments for the testing of code using pytest.
### test_capstone
This module is the one which contains all the test cases for testing purposes.
**Note - Apart from conftest, test_capstone are contained in the utils_package which is a custom package.**
| a9d3bdbe9d2cab671f67ce962cad70b818a98b8b | [
"Markdown",
"Python",
"Text"
] | 11 | Python | chiranthancv95/EpAI2.0_CapStone | a50bbe9f65e091766e245a50a32a1379b75122af | 5e42ef148f92eaaf608fa881a39cca258ca7035b |
refs/heads/master | <file_sep>import com.google.api.ads.common.lib.auth.OfflineCredentials;
import com.google.api.ads.common.lib.auth.OfflineCredentials.Api;
import com.google.api.ads.dfp.axis.factory.DfpServices;
import com.google.api.ads.dfp.axis.utils.v201708.StatementBuilder;
import com.google.api.ads.dfp.axis.v201708.Company;
import com.google.api.ads.dfp.axis.v201708.CompanyPage;
import com.google.api.ads.dfp.axis.v201708.CompanyServiceInterface;
import com.google.api.ads.dfp.axis.v201708.CompanyType;
import com.google.api.ads.dfp.lib.client.DfpSession;
import com.google.api.client.auth.oauth2.Credential;
public class TestDfpConnection {
public static void runExample(DfpServices dfpServices, DfpSession session) throws Exception {
CompanyServiceInterface companyService =
dfpServices.get(session, CompanyServiceInterface.class);
// Create a statement to select companies.
StatementBuilder statementBuilder = new StatementBuilder()
.where("type = :type")
.orderBy("id ASC")
.limit(StatementBuilder.SUGGESTED_PAGE_LIMIT)
.withBindVariableValue("type", CompanyType.ADVERTISER.toString());
// Retrieve a small amount of companies at a time, paging through
// until all companies have been retrieved.
int totalResultSetSize = 0;
do {
CompanyPage page =
companyService.getCompaniesByStatement(statementBuilder.toStatement());
if (page.getResults() != null) {
// Print out some information for each company.
totalResultSetSize = page.getTotalResultSetSize();
int i = page.getStartIndex();
for (Company company : page.getResults()) {
System.out.printf(
"%d) Company with ID %d, name '%s', and type '%s' was found.%n",
i++,
company.getId(),
company.getName(),
company.getType()
);
}
}
statementBuilder.increaseOffsetBy(StatementBuilder.SUGGESTED_PAGE_LIMIT);
} while (statementBuilder.getOffset() < totalResultSetSize);
System.out.printf("Number of results found: %d%n", totalResultSetSize);
}
public static void main(String[] args) throws Exception {
// Generate a refreshable OAuth2 credential for authentication.
Credential oAuth2Credential = new OfflineCredentials.Builder()
.forApi(Api.DFP)
.fromFile()
.build()
.generateCredential();
// Construct an API session configured from a properties file and the OAuth2
// credentials above.
DfpSession session = new DfpSession.Builder()
.fromFile()
.withOAuth2Credential(oAuth2Credential)
.build();
DfpServices dfpServices = new DfpServices();
runExample(dfpServices, session);
}
}
<file_sep>package com.admaru.service;
import java.util.Timer;
import org.apache.log4j.Logger;
public class BatchMon {
Timer timer;
private final Logger logger = Logger.getLogger(BatchMon.class);
public BatchMon() {
System.out.println("Service start");
try {
timer = new Timer();
timer.schedule(new TransTask(), 1000 * 5, 1000 * 60 * 60 * 2);// 1시간 간격
} catch (Exception e) {
System.out.println(e.getMessage());
e.printStackTrace();
System.exit(-1);
}
}
public static void main(String[] args) {
BatchMon batchmon = new BatchMon();
}
}
//TODO 시작일, 종료일 넣으면 일별로 엑셀다운받아서 처리하고 다음날짜로 넘어가서 엑셀다운받고 처리하도록 수정
| f4ced0ada723cc3fcb03ecee70c06b5367b1194f | [
"Java"
] | 2 | Java | m4292007/dfp | 49f31b46e6792d05224bf5e488167a875668fa54 | cdf1175de3adc23d95c1f2c81414fd1893842300 |
refs/heads/master | <file_sep>using System;
using System.Collections.Generic;
using System.ComponentModel;
using System.Data;
using System.Diagnostics;
using System.Drawing;
using System.IO;
using System.Linq;
using System.Text;
using System.Threading;
using System.Threading.Tasks;
using System.Windows.Forms;
using FindAssocExe;
using System.Drawing.Imaging;
namespace Graico
{
public partial class Form1 : Form
{
private readonly AsyncGraphicRead AsyncGraphReader = new ();
private List<string> FileList = null;
private string nowFile;
private bool zipFile;
private Form2 ProgressForm = new();
private string ZipFileName;
private readonly List<PictureBox> PicBoxList = new();
private readonly List<Image> PictureList = new();
private readonly PictureBox pictureBox = new();
private readonly Stopwatch KeyTimeSW = new();
private bool exec;
private Point mouseDownLocation;
private bool VolBtnNext;
private readonly List<string> graphicFileExt = new() { ".jpg", ".jpe", ".jpeg", ".gif", ".bmp", ".png", ".tif", ".tiff", ".avif" };
public Form1()
{
InitializeComponent();
MouseDown += PicBox_MouseDown;
MouseUp += PicBox_MouseUp;
PreviewKeyDown += Form1_PreviewKeyDown;
}
private void Form1_Load(object sender, EventArgs e)
{
var cmdLine = System.Environment.GetCommandLineArgs();
bool exeExec = false;
cmdLine.ForEach(async cmd =>
{
int jumpNo = 0;
string exeName = cmd.ToLower();
if (exeName.EndsWith("graico.exe") ||
exeName.EndsWith("graico.vshost.exe")) // VS Debug用
{
exeExec = true;
}
else if (exeExec)
{
string ext = Path.GetExtension(cmd).ToLower();
if ((ext == ".zip" || ext == ".cbz") ||
graphicFileExt.Contains(ext) &&
File.Exists(cmd))
{
await OpenGrapicFile(cmd);
if (FileList == null || FileList.Count <= 0)
{
// 既定の方法で開く
var proc = System.Diagnostics.Process.Start(cmd);
var procName = Path.GetFileName(Path.GetExtension(cmd).FindAssociatedExecutable()).ToLower();
var myProcName = Path.GetFileName(System.Environment.GetCommandLineArgs()[0]).ToLower();
if (procName == myProcName)
{
proc.Kill();
}
Close();
return;
}
else if (ext != ".zip" && ext != ".cbz")
{
jumpNo = FileList.FindIndex(fileName => fileName == cmd);
}
}
else if (cmd.IsNumeric())
{
string cmdNo = cmd;
jumpNo = Convert.ToInt32(cmdNo);
}
if (FileList != null && FileList.Count > 0)
await SetJumpImage(jumpNo);
}
});
}
public async Task OpenGrapicFile(string file)
{
if (ProgressForm.IsDisposed)
{
ProgressForm = new Form2();
}
ProgressForm.Show();
// 指定されたファイルを元にZIPアーカイブ内/フォルダ内ファイル一覧を取得
FileList = await AsyncGraphReader.GetFileListAsync(file);
if (FileList != null && FileList.Count > 0)
{
ZipFileName = file;
Debug.WriteLine("File List Count=" + FileList.Count);
string ext = Path.GetExtension(file).ToLower();
if (ext == ".zip" || ext == ".cbz")
{
zipFile = true;
nowFile = FileList[0];
}
else
{
zipFile = false;
nowFile = file;
}
}
ProgressForm.Close();
}
/// <summary>
/// 画面からPictureBoxを削除し、メモリも解放する
/// </summary>
/// <param name="picBoxList">PictureBoxリスト</param>
private void PicBoxListClear(List<PictureBox> picBoxList)
{
if (picBoxList.Count > 0)
{
picBoxList.ForEach(pic =>
{
pic.Image.Dispose();
pic.Image = null;
this.Controls.Remove(pic);
});
picBoxList.Clear();
}
}
/// <summary>
/// PictureBoxプロパティ設定
/// </summary>
/// <param name="picBox">設定するPictureBox</param>
/// <param name="img">PictureBoxに設定するイメージ</param>
/// <param name="yPos">PictureBox表示開始縦位置(横位置は0固定)</param>
private void SetPictureBoxProperty(PictureBox picBox, Image img, int yPos)
{
picBox.Location = new Point(0, yPos);
picBox.Image = img;
picBox.Size = new Size(img.Width, img.Height);
picBox.SizeMode = PictureBoxSizeMode.CenterImage;
picBox.Enabled = true;
picBox.Visible = true;
picBox.ContextMenuStrip = contextMenuStrip1;
int index = FileList.FindIndex(file => file == nowFile) + 1;
picBox.DoubleClick += async (sender, e) => await GetNextImage(index);
picBox.MouseDown += PicBox_MouseDown;
picBox.MouseUp += PicBox_MouseUp;
}
private void PicBox_MouseDown(object sender, MouseEventArgs e)
{
if (e.Button == MouseButtons.Right)
{
//this.OnMouseUp(e);
return;
}
mouseDownLocation = e.Location;
Debug.WriteLine("Mouse Down Event. Location=" + mouseDownLocation.ToString());
}
private async void PicBox_MouseUp(object sender, MouseEventArgs e)
{
if (e.Button == MouseButtons.Right)
{
//this.OnMouseUp(e);
return;
}
Debug.WriteLine("Mouse Up Event. Location=" + e.Location.ToString());
int mouseMoveX = e.X - mouseDownLocation.X;
//int mouseMoveY = e.Y - mouseDownLocation.Y;
//int maxVertScroll = VerticalScroll.Maximum - VerticalScroll.LargeChange;
//int maxHoriScroll = HorizontalScroll.Maximum - HorizontalScroll.LargeChange;
//bool scroll = false;
//try
//{
// if (VerticalScroll.Value < maxVertScroll && mouseMoveY != 0)
// {
// VerticalScroll.Value += mouseMoveY;
// scroll = true;
// }
// if (HorizontalScroll.Value < maxHoriScroll && mouseMoveX != 0)
// {
// HorizontalScroll.Value += mouseMoveX;
// scroll = true;
// }
// if (scroll)
// {
// return;
// }
//}
//catch {}
// 横方向に移動なし、または、移動距離が短い
if (mouseMoveX == 0 || Math.Abs(mouseMoveX) < 100)
{
return;
}
if (mouseMoveX > 0)
{
await MouseSwipe(1);
}
else
{
await MouseSwipe(-1);
}
}
/// <summary>
/// イメージリスト画像フォーム貼り付け
/// </summary>
/// <param name="imgList">フォーム貼り付けるイメージリスト</param>
private void PicBoxSetImage(List<Image> imgList)
{
int yPos = 0;
try
{
if (imgList.Count <= 0) return;
Size imgSize = new(imgList[0].Size.Width, 0);
imgList.ForEach(img =>
{
var picBox = new PictureBox();
SetPictureBoxProperty(picBox, img, yPos);
imgSize.Height += img.Height;
this.Controls.Add(picBox);
PicBoxList.Add(picBox);
yPos += img.Height;
});
}
catch (Exception ex)
{
MessageBox.Show(ex.Message);
}
}
/// <summary>
/// 次の画像読み込みと関連する処理の実施
/// </summary>
/// <param name="index">表示インデックス</param>
/// <returns>戻り値なし(待機可能にするための指定->voidにすると待機なしになる)</returns>
private async Task GetNextImage(int index)
{
try
{
if (ProgressForm.IsDisposed)
{
ProgressForm = new Form2();
}
ProgressForm.Show();
PicBoxListClear(PicBoxList);
ImageListClear(PictureList);
nowFile = await AsyncGraphReader.GetIndexImage(PictureList, FileList, index, zipFile);
if (PictureList.Count > 0)
{
PicBoxSetImage(PictureList);
int newIndex = FileList.FindIndex(file => file == nowFile) + 1;
SetMainTitle(newIndex);
}
ProgressForm.Close();
}
catch (Exception ex)
{
MessageBox.Show(ex.Message);
}
}
private async void OpenToolStripMenuItem_Click(object sender, EventArgs e)
{
var openFile = new OpenFileDialog
{
Filter = "Graphic Files|*.jpg;*.jpeg;*.bmp;*.gif;*.png;*.zip;*.cbz"
};
if (openFile.ShowDialog() == DialogResult.OK)
{
string file = openFile.FileName;
await OpenGrapicFile(file);
if (FileList.Count <= 0)
{
MessageBox.Show("対応の画像ファイルが存在しません(または、未対応のファイルです):" + file);
}
if (FileList != null && FileList.Count > 0)
await SetJumpImage(0);
}
}
private void CloseToolStripMenuItem_Click(object sender, EventArgs e)
{
Close();
}
private async void Form1_PreviewKeyDown(object sender, PreviewKeyDownEventArgs e)
{
if ((e.KeyCode == Keys.Right || e.KeyCode == Keys.Enter) ||
(e.KeyCode == Keys.Left || e.KeyCode == Keys.Back))
{
bool viewOk;
if (KeyTimeSW.IsRunning)
{
Debug.WriteLine("Time = " + KeyTimeSW.ElapsedMilliseconds + "ms");
if (KeyTimeSW.ElapsedMilliseconds > 200)
{
viewOk = true;
KeyTimeSW.Restart();
}
else
{
viewOk = false;
}
}
else
{
KeyTimeSW.Start();
viewOk = true;
}
//Debug.WriteLine("View=" + (viewOk ? "OK" : "NG"));
if (!viewOk)
{
return;
}
}
int addIndex = 0;
if (VolBtnNext && e.KeyCode == Keys.VolumeDown)
{
addIndex = 1;
await SetPicBoxSizeMode(addIndex);
e.IsInputKey = false;
}
else if (VolBtnNext && e.KeyCode == Keys.VolumeUp)
{
addIndex = -1;
await SetPicBoxSizeMode(addIndex);
e.IsInputKey = false;
}
else if (e.KeyCode == Keys.Right || e.KeyCode == Keys.Enter)
{
Debug.WriteLine("============= Rigth or Enter Input! =============");
addIndex = 1;
//await SetPicBoxSizeMode(addIndex);
}
else if (e.KeyCode == Keys.Left || e.KeyCode == Keys.Back)
{
Debug.WriteLine("============= Left or Back Input! =============");
addIndex = -1;
}
if ((e.KeyCode == Keys.Right || e.KeyCode == Keys.Enter) ||
(e.KeyCode == Keys.Left || e.KeyCode == Keys.Back))
{
//var findIndex = FileList.FindIndex(file => file == nowFile) + addIndex + 1;
//if (findIndex <= 0)
//{
// findIndex = FileList.Count;
//}
//else if (findIndex > FileList.Count)
//{
// findIndex = 1;
//}
//await GetNextImage(-1);
//await SetJump(-1);
//Debug.WriteLine("KeyPreview index=" + findIndex);
//await SetPicBoxSizeMode(findIndex);
await SetPicBoxSizeMode(addIndex);
}
else if (e.KeyCode == Keys.Escape)
{
Close();
}
else if (e.KeyCode == Keys.Space)
{
int newX = (ClientSize.Width / 2) - (contextMenuStrip1.Width / 2);
int newY = (ClientSize.Height / 2) - (contextMenuStrip1.Height / 2);
contextMenuStrip1.Show(this, new Point(newX, newY));
}
}
private void SetMainTitle(int index)
{
Text = Path.GetFileName(ZipFileName) + " - " + nowFile + " (" + (index + 1) + "/" + FileList.Count + ")";
}
private static void ImageListClear(List<Image> ImageList)
{
ImageList.ForEach(img => img.Dispose());
ImageList.Clear();
}
public async Task SetJumpImage(int index)
{
if (ProgressForm.IsDisposed)
{
ProgressForm = new Form2();
}
ProgressForm.Show();
ImageListClear(PictureList);
nowFile = await AsyncGraphReader.GetIndexImage(PictureList, FileList, index, zipFile);
if (nowFile == null)
{
return;
}
if (PictureList.Count > 0)
{
PicBoxListClear(PicBoxList);
}
PicBoxSetImage(PictureList);
Debug.WriteLine("SetJumpImage indx = " + index + " Now file = " + nowFile);
SetMainTitle(index);
ProgressForm.Close();
}
private async void JumpToolStripMenuItem_Click(object sender, EventArgs e)
{
int index = FileList.FindIndex(file => file == nowFile);
var jumpInputForm = new Form3();
jumpInputForm.setNumericUpDownMaxValue(FileList.Count);
jumpInputForm.setNumericUpDownValue(index + 1);
if (jumpInputForm.ShowDialog() == DialogResult.OK)
{
index = jumpInputForm.getNumericUpDownValue();
//await SetJumpImage(index-1);
await SetPicBoxSizeMode(index - 1, false);
}
}
private async void NextToolStripMenuItem_Click(object sender, EventArgs e)
{
//int index = FileList.FindIndex(file => file == nowFile) + 1;
//await GetNextImage(index);
await SetPicBoxSizeMode(1);
}
private async void BeforeToolStripMenuItem_Click(object sender, EventArgs e)
{
//int index = FileList.FindIndex(file => file == nowFile) - 1;
//await GetNextImage(index);
await SetPicBoxSizeMode(-1);
}
private void SaveAsToolStripMenuItem_Click(object sender, EventArgs e)
{
if (string.IsNullOrEmpty(nowFile))
{
return;
}
var saveDialog = new SaveFileDialog
{
Filter = "Graphic Files|*.jpg;*.jpeg;*.bmp;*.gif;*.png|JPEG|*.jpg|PNG|*.png|Bitmap|*.bmp"
};
if (saveDialog.ShowDialog() == DialogResult.OK)
{
ImageFormat imgform = null;
string file = saveDialog.FileName;
string ext = Path.GetExtension(file).ToLower();
if (ext == ".jpg" || ext == "*.jpe" || ext == ".jpeg")
{
imgform = ImageFormat.Jpeg;
}
else if (ext == ".bmp")
{
imgform = ImageFormat.Bmp;
}
else if (ext == ".png")
{
imgform = ImageFormat.Png;
}
Image.FromFile(nowFile).Save(file, imgform);
}
}
private async Task SetJump(int indexAdd = 0)
{
await SetJumpImage(NextFileIndex(indexAdd));
}
private int NextFileIndex(int index)
{
var findIndex = FileList.FindIndex(file => file == nowFile) + index;
if (findIndex < 0)
{
findIndex = FileList.Count - 1;
}
else if (findIndex >= FileList.Count)
{
findIndex = 0;
}
return findIndex;
}
private async Task SetPicBoxSizeMode(int index, bool next = true)
{
if (ProgressForm.IsDisposed)
{
ProgressForm = new Form2();
}
ProgressForm.Show();
// ズームモード
// サイズが小さくなるので、分割なしにする
//var phMen = new Microsoft.VisualBasic.Devices.ComputerInfo().AvailablePhysicalMemory;
if ((System.Environment.Is64BitOperatingSystem
// && phMen >= (1L * 1024L * 1024L * 1024L)
) ||
(zoomToolStripMenuItem.Checked || wideFitZoomToolStripMenuItem.Checked))
{
var findIndex = FileList.FindIndex(file => file == nowFile);
if (next)
{
findIndex = NextFileIndex(index);
}
// PictureBoxリストクリア
// まずはコントロールコレクションから
ControlsPicBoxClear(PicBoxList);
// リスト自体の削除
PicBoxListClear(PicBoxList);
// イメージリストも削除
ImageListClear(PictureList);
// ここから、新たに読み込みと追加
//nowFile = await AsyncGraphReader.GetIndexImage(PictureList, FileList, findIndex, zipFile, 0);
//SetMainTitle(findIndex);
//PicBoxSetImage(PictureList);
//nowFile = FileList[index - 1];
//SetPictureBoxProperty(pictureBox, PictureList[0], 0);
//pictureBox = PicBoxList[0];
AsyncGraphReader.GetPictureSize(FileList, FileList[findIndex], out Size newSize, zipFile);
if (zoomToolStripMenuItem.Checked)
{
GetScreenFitSize(ref newSize);
var newImage = await AsyncGraphReader.GetZoomImageFromFile(FileList, FileList[findIndex], newSize, zipFile);
nowFile = FileList[findIndex];
SetMainTitle(findIndex);
pictureBox.Image = newImage;
if (zoomToolStripMenuItem.Checked)
{
pictureBox.Size = ClientSize;
pictureBox.SizeMode = PictureBoxSizeMode.Zoom;
}
else
{
pictureBox.Size = newImage.Size;
}
pictureBox.Location = new Point(0, 0);
pictureBox.MouseDown += PicBox_MouseDown;
pictureBox.MouseUp += PicBox_MouseUp;
Controls.Add(pictureBox);
pictureBox.Refresh();
}
else if (wideFitZoomToolStripMenuItem.Checked)
{
GetScreenWideFitSize(ref newSize);
var newImage = await AsyncGraphReader.GetZoomImageFromFile(FileList, FileList[findIndex], newSize, zipFile);
nowFile = FileList[findIndex];
SetMainTitle(findIndex);
AutoScrollPosition = new Point(0, 0);
pictureBox.Image = newImage;
pictureBox.Size = newImage.Size;
pictureBox.Location = new Point(0, 0);
pictureBox.MouseDown += PicBox_MouseDown;
pictureBox.MouseUp += PicBox_MouseUp;
Controls.Add(pictureBox);
pictureBox.Refresh();
}
else
{
if (pictureBox.Image != null)
{
pictureBox.Image.Dispose();
pictureBox.Image = null;
}
if (Controls.Contains(pictureBox))
Controls.Remove(pictureBox);
if (next)
{
await SetJump(index);
}
else
{
await SetJumpImage(index);
}
}
ProgressForm.Close();
}
}
/// <summary>
/// 画面のコントロールコレクションからPicBoxListのアイテムを削除する
/// </summary>
/// <param name="PicBoxList">削除対象のリスト</param>
private void ControlsPicBoxClear(List<PictureBox> PicBoxList)
{
PicBoxList.ForEach(picBox =>
{
if (Controls.Contains(picBox))
{
Debug.WriteLine("Remove PictureBox=" + picBox.Size.ToString());
Controls.Remove(picBox);
}
});
}
private async void ZoomToolStripMenuItem_Click(object sender, EventArgs e)
{
if (zoomToolStripMenuItem.Checked && wideFitZoomToolStripMenuItem.Checked)
{
wideFitZoomToolStripMenuItem.Checked = false;
}
var findIndex = FileList.FindIndex(file => file == nowFile);
if (findIndex < 0)
{
findIndex = FileList.Count - 1;
}
else if (findIndex > FileList.Count)
{
findIndex = 1;
}
await SetPicBoxSizeMode(findIndex, false);
}
private static void GetScreenFitSize(ref Size imageSize)
{
// スクリーンサイズの取得
Rectangle Rect = Screen.GetWorkingArea(new Point(0, 0));
// スクリーンサイズの幅と高さ
int screenX = Rect.Size.Width;
int screenY = Rect.Size.Height;
// 画像の幅と高さ
int imageX = imageSize.Width;
int imageY = imageSize.Height;
// 画像の比率に沿った幅と高さ計算
int RX = imageX * screenY / imageY;
int RY = imageY * screenX / imageX;
int newX;
int newY;
if ((RX < screenX) && (RY > screenY))
{
newX = RX;
newY = screenY;
}
else
{
newX = screenX;
newY = RY;
}
imageSize.Width = newX;
imageSize.Height = newY;
}
private static void GetScreenWideFitSize(ref Size imageSize)
{
// スクリーンサイズの取得
Rectangle Rect = Screen.GetWorkingArea(new Point(0, 0));
// スクリーンサイズの幅と高さ
int screenX = Rect.Size.Width;
//int screenY = Rect.Size.Height;
// 画像の幅と高さ
int imageX = imageSize.Width;
int imageY = imageSize.Height;
// 画像の比率に沿った幅と高さ計算
//int RX = imageX * screenY / imageY;
int RY = imageY * screenX / imageX;
// 横幅はスクリーンサイズ固定
int newY = RY;
int newX = screenX;
imageSize.Width = newX;
imageSize.Height = newY;
}
private async Task MouseSwipe(int indexAdd)
{
// 2重起動防止(1回の処理中に同イベントが発生しても何もしない)
if (exec)
{
return;
}
exec = true;
await SetPicBoxSizeMode(indexAdd);
exec = false;
}
private void Form1_Scroll(object sender, ScrollEventArgs e)
{
Debug.WriteLine("" + e.ScrollOrientation);
Debug.WriteLine("Scroll=" + e.NewValue);
Debug.WriteLine("VScroll=" + VerticalScroll.Value);
Debug.WriteLine("HScroll=" + HorizontalScroll.Value);
Debug.WriteLine("VLarge Scroll=" + VerticalScroll.LargeChange);
Debug.WriteLine("HLarge Scroll=" + HorizontalScroll.LargeChange);
Debug.WriteLine("VSmall Scroll=" + VerticalScroll.SmallChange);
Debug.WriteLine("HSmall Scroll=" + HorizontalScroll.SmallChange);
Debug.WriteLine("VerticalScroll.Maximum=" + VerticalScroll.Maximum);
Debug.WriteLine("HorizontalScroll.Maximum=" + HorizontalScroll.Maximum);
}
private void FullscreenToolStripMenuItem_Click(object sender, EventArgs e)
{
if (fullscreenToolStripMenuItem.Checked)
{
WindowState = FormWindowState.Normal;
FormBorderStyle = FormBorderStyle.None;
TopMost = true;
WindowState = FormWindowState.Maximized;
}
else
{
WindowState = FormWindowState.Maximized;
FormBorderStyle = FormBorderStyle.Sizable;
TopMost = false;
}
}
private void CopyToolStripMenuItem_Click(object sender, EventArgs e)
{
Clipboard.SetImage(pictureBox.Image);
}
private void VolButtonEnableToolStripMenuItem_Click(object sender, EventArgs e)
{
if (volButtonEnableToolStripMenuItem.Checked)
{
VolBtnNext = true;
}
else
{
VolBtnNext = false;
}
}
private void Form1_KeyDown(object sender, KeyEventArgs e)
{
if (e.KeyCode == Keys.Up)
{
this.AutoScrollPosition = new Point(this.AutoScrollPosition.X, -this.AutoScrollPosition.Y - 10);
}
else if (e.KeyCode == Keys.Down)
{
this.AutoScrollPosition = new Point(this.AutoScrollPosition.X, -this.AutoScrollPosition.Y + 10);
}
else if (e.KeyCode == Keys.PageUp)
{
this.AutoScrollPosition = new Point(this.AutoScrollPosition.X, -this.AutoScrollPosition.Y - 100);
}
else if (e.KeyCode == Keys.PageDown)
{
this.AutoScrollPosition = new Point(this.AutoScrollPosition.X, -this.AutoScrollPosition.Y + 100);
}
else if (e.KeyCode == Keys.Home)
{
this.AutoScrollPosition = new Point(this.AutoScrollPosition.X, 0);
}
else if (e.KeyCode == Keys.End)
{
this.AutoScrollPosition = new Point(this.AutoScrollPosition.X, 30000);
}
}
private async void WideFitZoomToolStripMenuItem_Click(object sender, EventArgs e)
{
if (zoomToolStripMenuItem.Checked && wideFitZoomToolStripMenuItem.Checked)
{
zoomToolStripMenuItem.Checked = false;
}
var findIndex = FileList.FindIndex(file => file == nowFile);
if (findIndex < 0)
{
findIndex = FileList.Count - 1;
}
else if (findIndex > FileList.Count)
{
findIndex = 1;
}
await SetPicBoxSizeMode(findIndex, false);
}
}
}
<file_sep>using System;
using System.Collections.Generic;
using System.Linq;
using System.Text;
using System.Threading.Tasks;
namespace Graico
{
public static class Utility
{
public static bool IsNumeric(this string str)
{
return Int32.TryParse(str, out _);
}
}
/// <summary>
/// for文、foreach文をなくすための、標準で存在しない拡張メソッド
/// </summary>
public static class EnumerableExtentions
{
/// <summary>
/// index付、ForEach拡張メソッド
/// </summary>
/// <typeparam name="T"></typeparam>
/// <param name="source"></param>
/// <param name="action"></param>
public static void ForEach<T>(this IEnumerable<T> source, Action<T, int> action)
{
int index = 0;
foreach (var x in source)
action(x, index++);
}
/// <summary>
/// indexなし、ForEach拡張メソッド
/// </summary>
/// <typeparam name="T"></typeparam>
/// <param name="source"></param>
/// <param name="action"></param>
public static void ForEach<T>(this IEnumerable<T> source, Action<T> action)
{
foreach (var x in source)
action(x);
}
}
}
<file_sep>using System;
using System.Collections.Generic;
using System.ComponentModel;
using System.Data;
using System.Drawing;
using System.Linq;
using System.Text;
using System.Threading.Tasks;
using System.Windows.Forms;
namespace Graico
{
public partial class Form3 : Form
{
public void setNumericUpDownMaxValue(int max)
{
numericUpDown1.Maximum = max;
label1.Text = "/" + max;
}
public int getNumericUpDownValue()
{
return Convert.ToInt32(numericUpDown1.Value);
}
public void setNumericUpDownValue(int val)
{
numericUpDown1.Value = val;
}
public Form3()
{
InitializeComponent();
numericUpDown1.Accelerations.Add(new NumericUpDownAcceleration(5, 10));
}
}
}
<file_sep>using System;
using System.Collections;
using System.Collections.Generic;
using System.Linq;
using System.Text;
using System.Text.RegularExpressions;
using System.Threading.Tasks;
namespace Graico
{
public class StringComparer : IComparer<string>
{
public static bool NumCheck = true;
private static string _numRegex = @"^(.*?)([0-9]+).*?$";
private static Regex regex = new Regex(_numRegex);
//xがyより小さいときはマイナスの数、大きいときはプラスの数、
//同じときは0を返す
public int Compare(string a, string b)
{
string aorg = a;
string borg = b;
// 何もしなくても等しかったら0
if (a == b)
{
return 0;
}
// 数字部分切り出し保存用
long? ai = null;
long? bi = null;
// 数字チェックするなら
if (NumCheck)
{
// 正規表現で切り出す
Match matchCol = regex.Match(a);
// マッチしたら
if (matchCol.Success)
{
// 数字の前までの文字列と
a = matchCol.Groups[1].Value;
// 数字に分ける
ai = Convert.ToInt64(matchCol.Groups[2].Value);
}
// 正規表現
matchCol = regex.Match(b);
// マッチ
if (matchCol.Success)
{
// 文字列
b = matchCol.Groups[1].Value;
// 数字
bi = Convert.ToInt64(matchCol.Groups[2].Value);
}
}
// 文字列の比較
int t = string.Compare(a, b);
// 等しければ
if (NumCheck && t == 0)
{
//
if (ai == null && bi != null)
{
t = -1;
}
else if (ai != null && bi == null)
{
t = 1;
}
else if (ai == null && bi == null)
{
t = string.Compare(aorg, borg);
}
else
{
t = (int)(ai - bi);
if (t == 0)
{
t = string.Compare(aorg, borg);
}
}
}
return t;
}
}
}
<file_sep>using System;
using System.Runtime.InteropServices;
using System.Text;
namespace FindAssocExe
{
public static class FindAssocExe
{
/// <summary>
/// 指定された拡張子に関連付けられた実行ファイルのパスを取得する。
/// </summary>
/// <param name="extName">".txt"などの拡張子。</param>
/// <returns>見つかった時は、実行ファイルのパス。
/// 見つからなかった時は、空の文字列。</returns>
/// <example>
/// 拡張子".txt"に関連付けられた実行ファイルのパスを取得する例
/// <code>
/// string exePath = FindAssociatedExecutable(".txt");
/// </code>
/// </example>
public static string FindAssociatedExecutable(this string extName)
{
//pszOutのサイズを取得する
uint pcchOut = 0;
//ASSOCF_INIT_IGNOREUNKNOWNで関連付けられていないものを無視
//ASSOCF_VERIFYを付けると検証を行うが、パフォーマンスは落ちる
AssocQueryString(AssocF.Init_IgnoreUnknown, AssocStr.Executable,
extName, null, null, ref pcchOut);
if (pcchOut == 0)
{
return string.Empty;
}
//結果を受け取るためのStringBuilderオブジェクトを作成する
StringBuilder pszOut = new StringBuilder((int)pcchOut);
//関連付けられた実行ファイルのパスを取得する
AssocQueryString(AssocF.Init_IgnoreUnknown, AssocStr.Executable,
extName, null, pszOut, ref pcchOut);
//結果を返す
return pszOut.ToString();
}
[DllImport("Shlwapi.dll",
SetLastError = true,
CharSet = CharSet.Auto)]
private static extern uint AssocQueryString(AssocF flags,
AssocStr str,
string pszAssoc,
string pszExtra,
[Out] StringBuilder pszOut,
[In][Out] ref uint pcchOut);
[Flags]
private enum AssocF
{
None = 0,
Init_NoRemapCLSID = 0x1,
Init_ByExeName = 0x2,
Open_ByExeName = 0x2,
Init_DefaultToStar = 0x4,
Init_DefaultToFolder = 0x8,
NoUserSettings = 0x10,
NoTruncate = 0x20,
Verify = 0x40,
RemapRunDll = 0x80,
NoFixUps = 0x100,
IgnoreBaseClass = 0x200,
Init_IgnoreUnknown = 0x400,
Init_FixedProgId = 0x800,
IsProtocol = 0x1000,
InitForFile = 0x2000,
}
private enum AssocStr
{
Command = 1,
Executable,
FriendlyDocName,
FriendlyAppName,
NoOpen,
ShellNewValue,
DDECommand,
DDEIfExec,
DDEApplication,
DDETopic,
InfoTip,
QuickTip,
TileInfo,
ContentType,
DefaultIcon,
ShellExtension,
DropTarget,
DelegateExecute,
SupportedUriProtocols,
Max,
}
}
}<file_sep>using System;
using System.Collections.Generic;
using System.Diagnostics;
using System.Drawing;
using System.Drawing.Drawing2D;
using System.IO;
using System.IO.Compression;
using System.Linq;
using System.Text;
using System.Threading.Tasks;
namespace Graico
{
public class AsyncGraphicRead
{
public string FileName { get; set; }
private List<ZipArchiveEntry> ZipArcEntryList = null;
private readonly Object thisLock = new();
private readonly string[] graphicFileExt = { ".jpg", ".jpe", ".jpeg", ".gif", ".bmp", ".png", ".tif", ".tiff", ".avif" };
/// <summary>
/// 非同期画像ファイル読み込み
/// </summary>
/// <param name="fileName">読み込む画像ファイル名</param>
/// <param name="progress">進行状況の更新プロバイダー</param>
/// <returns></returns>
//private async Task<Image>
// GetImageFileAsync(
// string fileName,
// IProgress<int> progress)
//{
// var readList = new List<byte>();
// using (var fs = File.Open(fileName, FileMode.Open, FileAccess.Read, FileShare.Read))
// {
// int readsize = Convert.ToInt32(fs.Length / 100);
// if (readsize <= 100)
// {
// readsize = 100;
// }
// foreach (int i in Enumerable.Range(0, 100))
// {
// var readBuff = new byte[readsize];
// var readSize = await fs.ReadAsync(readBuff, 0, (int)readBuff.Length);
// if (readSize <= 0)
// {
// break;
// }
// Array.Resize(ref readBuff, readSize);
// readList.AddRange(readBuff);
// progress.Report(i + 1);
// //Debug.WriteLine((i + 1) + "%");
// }
// }
// //var imgconv = new ImageConverter();
// //return (imgconv.ConvertFrom(readList.ToArray()) as Image);
// var imgconv = new ImageConverter();
// var img = imgconv.ConvertFrom(readList.ToArray()) as Image;
// readList.Clear();
// readList = null;
// return (img);
//}
private async Task GetZipGraphicImage(
List<Image> imgList,
int newIndex,
int DivSize = 4096)
{
if (ZipArcEntryList.Count < (newIndex))
{
newIndex = 0;
}
else if (newIndex < 0)
{
newIndex = ZipArcEntryList.Count - 1;
}
await Task.Run(() =>
{
// ロックをかけないとデータ破壊の恐れあり
// 付けないとヘッダが壊れているという例外が発生することがある
lock (thisLock)
{
Debug.WriteLine("ZIP Arc index=" + newIndex);
var ZipArc = ZipArcEntryList[newIndex];
Debug.WriteLine("Zip new Index File=" + ZipArc.FullName);
using var zipStream = ZipArc.Open();
try
{
//Image Img = Image.FromStream(zipStream);
//Debug.WriteLine("New Index=" + newIndex + " FileName = " + ZipArcEntryList[newIndex].FullName);
GetDivideImageFormStream(zipStream, imgList, DivSize);
return;
}
catch (Exception ex)
{
Debug.WriteLine(ex.Message);
return;
}
}
});
return;
}
/// <summary>
/// ファイル一覧取得
/// </summary>
/// <remarks>
/// ZIPファイルやディレクトリから、グラフィックファイル一覧を作成する
/// </remarks>
/// <param name="fileName">ZIPファイル名/基となる画像ファイル</param>
public async Task<List<string>> GetFileListAsync(
string fileName)
{
var extens = Path.GetExtension(fileName).ToLower();
if (extens == ".zip" || extens == ".cbz")
{
var zipStream = File.OpenRead(fileName);
var ret = await Task.Run(() =>
{
try
{
var arc = new ZipArchive(zipStream, ZipArchiveMode.Read);
ZipArcEntryList = arc.Entries
.Where(x =>
{
var ext = Path.GetExtension(x.Name).ToLower();
if (graphicFileExt.Contains(ext))
{
return true;
}
return false;
})
//.OrderBy(x => x.Name, new NaturalComparer())
.OrderBy(x => x.Name, new StrNatComparer())
.ToList();
}
catch(Exception ex)
{
Debug.WriteLine(ex.Message);
return null;
}
return ZipArcEntryList.Select(x => x.FullName).ToList();
});
return ret;
}
else
{
// 検索ディレクトリ=カレントディレクトリ
string currDir = Path.GetDirectoryName(fileName);
// EnumerateFilesは単一のパターンのみしか指定できないため、
// とりあえず全部取得し、Whereで絞り込む
string searchPattern = "*";
// ファイルリストの取得
// LINQでファイルの絞り込み(LINQなら複雑な処理を記述しなくて済む)
// さらに、別途ソートしていたのをLINQでナチュラルソートするように変更
// List化も別途から一緒にするように変更した
var ret = await Task.Run(() =>
{
var rtc = Directory.EnumerateFiles(currDir, searchPattern)
.Where(file =>
{
// 拡張子の取得
string ext = Path.GetExtension(file).ToLower();
if (graphicFileExt.Contains(ext))
{
return true;
}
return false;
//}).OrderBy(x => x, new NaturalComparer()).ToList();
}).OrderBy(x => x, new StrNatComparer()).ToList();
return rtc;
});
return ret;
}
}
/// <summary>
/// ファイルリストから指定のファイルを見つけ出し、
/// addIndex加算した次のファイル(マイナスなら前)を
/// 返す
/// </summary>
/// <param name="FileList"></param>
/// <param name="fileName"></param>
/// <param name="addIndex"></param>
/// <returns></returns>
private string GetNextFile(List<string> FileList, string fileName, int addIndex)
{
var findIndex = FileList.FindIndex(file => file == fileName);
if (findIndex < 0)
{
return null;
}
int index = findIndex + addIndex;
if (index >= FileList.Count)
{
index = 0;
}
else if (index < 0)
{
index = FileList.Count - 1;
}
string getFile = string.Empty;
while (index < FileList.Count)
{
getFile = FileList[index];
var ext = Path.GetExtension(fileName).ToLower();
//if (ext != ".jpg" && ext != ".jpeg" && ext != ".png" &&
// ext != ".bmp" && ext != ".gif")
if (!graphicFileExt.Contains(ext))
{
int newIndex = index + addIndex;
if (newIndex >= 0 && newIndex < FileList.Count)
{
index += addIndex;
}
else if (newIndex < 0)
{
index = FileList.Count - 1;
continue;
}
else if (newIndex > FileList.Count)
{
index = 0;
continue;
}
}
else
{
break;
}
}
return getFile;
}
private async Task<Image> GetNextImage(
List<string> FileList,
string fileName,
int add,
bool zipFile)
{
var imgList = new List<Image>();
FileName = GetNextFile(FileList, fileName, add);
if (zipFile)
{
int index = FileList.FindIndex(file => file == FileName);
await GetZipGraphicImage(imgList, index, 0);
}
else
{
//return await GetImageFileAsync(FileName, new Progress<int>(prog => { }));
GetDivideImageFormFile(FileName, imgList, 0);
}
return imgList[0];
}
public async Task<string> GetIndexImage(
List<Image> imgList,
List<string> FileList,
//string fileName,
int index,
bool zipFile,
int divSize = 4096)
{
if (index < 0 || index >= FileList.Count)
{
Debug.WriteLine("Index Under/Over Error! index=" + index);
return null;
}
Debug.WriteLine("Index Image=" + index);
FileName = FileList[index];
Debug.WriteLine("Get Index Image file=" + FileName);
if (zipFile)
{
await GetZipGraphicImage(imgList, index, divSize);
}
else
{
//return await GetImageFileAsync(FileName, new Progress<int>(prog => { }));
GetDivideImageFormFile(FileName, imgList, divSize);
}
return FileName;
}
//private async Task GetNextImage(
// List<Image> imgList,
// List<string> FileList,
// string fileName,
// int add,
// bool zipFile)
//{
// FileName = GetNextFile(FileList, fileName, add);
// Debug.WriteLine("Get Next Image file=" + FileName);
// if (zipFile)
// {
// int index = FileList.FindIndex(file => file == FileName);
// await GetZipGraphicImage(imgList, index);
// return;
// }
// else
// {
// //return await GetImageFileAsync(FileName, new Progress<int>(prog => { }));
// GetDivideImageFormFile(FileName, imgList);
// }
//}
public static void GetDivideImage(
Image img,
List<Image> divImg,
int DivSize = 4096)
{
int height = img.Height;
// 分割の必要なしなら、読み込んだイメージをリストに追加して返す
if (height <= DivSize || DivSize <= 0)
{
divImg.Add(img);
return;
}
// 分割数計算
int divNo = 0;
try
{
divNo = height / DivSize + 1;
}
catch (DivideByZeroException)
{
divNo = 1;
}
var enumRange = Enumerable.Range(0, divNo);
int heightPos = 0;
// 最後は別途計算する
int lastSize = height - (DivSize * (divNo - 1));
// 読み込んだ画像を縦分割する
enumRange.ForEach(i =>
{
int ySize = DivSize;
if (i == enumRange.Last())
{
ySize = lastSize;
}
if (i == enumRange.Last() && lastSize <= 0)
{
}
else
{
//画像ファイルのImageオブジェクトを作成する
var image = new Bitmap(img.Width, ySize);
//ImageオブジェクトのGraphicsオブジェクトを作成する
Graphics g = Graphics.FromImage(image);
//切り取る部分の範囲を決定する
Rectangle srcRect = new(0, heightPos, img.Width, ySize);
//描画する部分の範囲を決定する
Rectangle desRect = new(0, 0, img.Width, srcRect.Height);
//画像の一部を描画する
g.DrawImage(img, desRect, srcRect, GraphicsUnit.Pixel);
//Graphicsオブジェクトのリソースを解放する
g.Dispose();
heightPos += ySize;
divImg.Add(image);
}
});
img.Dispose();
img = null;
}
/// <summary>
/// 画素数が大きいサイズの画像を縦分割して読み込む
/// </summary>
/// <param name="fileName">読み込む画像ファイル名</param>
/// <param name="divImg">読み込んだ画像イメージリスト</param>
/// <param name="DivSize">分割する縦サイズ</param>
public static void GetDivideImageFormFile(
string fileName,
List<Image> divImg,
int DivSize = 4096)
{
// まず全部を読み込む(これでメモリ不足になるなら、別の手を考える必要あり)
var img = Image.FromFile(fileName);
GetDivideImage(img, divImg, DivSize);
}
/// <summary>
/// 画素数が大きいサイズの画像を縦分割して読み込む
/// </summary>
/// <param name="fileStream">読み込む画像ファイル名</param>
/// <param name="divImg">読み込んだ画像イメージリスト</param>
/// <param name="DivSize">分割する縦サイズ</param>
public static void GetDivideImageFormStream(
Stream fileStream,
List<Image> divImg,
int DivSize = 4096)
{
// まず全部を読み込む(これでメモリ不足になるなら、別の手を考える必要あり)
var img = Image.FromStream(fileStream);
GetDivideImage(img, divImg, DivSize);
}
public async Task<Image> GetZoomImage(List<string> fileList, string imageFile, Size newSize, bool zip)
{
//描画先とするImageオブジェクトを作成する
Bitmap canvas = new(newSize.Width, newSize.Height);
//ImageオブジェクトのGraphicsオブジェクトを作成する
Graphics g = null;
await Task.Run(() => g = Graphics.FromImage(canvas));
//画像ファイルを読み込んで、Imageオブジェクトとして取得する
Image img = await GetNextImage(fileList, imageFile, 0, zip);
//画像のサイズを2倍にしてcanvasに描画する
g.DrawImage(img, 0, 0, newSize.Width, newSize.Height);
//Imageオブジェクトのリソースを解放する
img.Dispose();
//Graphicsオブジェクトのリソースを解放する
g.Dispose();
return canvas;
}
public static void GetImageSize(Stream fs, out Size imgSize)
{
using Image img = Image.FromStream(fs, false, false);
Console.WriteLine(img.Width + " x " + img.Height);
imgSize = new Size
{
Width = img.Size.Width,
Height = img.Size.Height
};
img.Dispose();
}
public void GetZipInImageSize(int index, out Size imgSize)
{
var ZipArc = ZipArcEntryList[index];
using var zipStream = ZipArc.Open();
GetImageSize(zipStream, out imgSize);
}
public static void GetFileImageSize(string fileName, out Size imgSize)
{
using FileStream fs = File.OpenRead(fileName);
GetImageSize(fs, out imgSize);
}
public void GetPictureSize(List<string> FileList, string fileName, out Size imgSize, bool zip)
{
if (zip)
{
int index = FileList.FindIndex(file => file == fileName);
GetZipInImageSize(index, out imgSize);
}
else
{
GetFileImageSize(fileName, out imgSize);
}
}
public static Image GetZoomImageFromStream(Stream fs, Size imgSize)
{
using Image img = Image.FromStream(fs, false, false);
//描画先とするImageオブジェクトを作成する
Bitmap canvas = new(imgSize.Width, imgSize.Height);
//ImageオブジェクトのGraphicsオブジェクトを作成する
Graphics g = Graphics.FromImage(canvas);
g.InterpolationMode =
InterpolationMode.HighQualityBicubic;
//画像のサイズを2倍にしてcanvasに描画する
g.DrawImage(img, 0, 0, imgSize.Width, imgSize.Height);
//Imageオブジェクトのリソースを解放する
img.Dispose();
//Graphicsオブジェクトのリソースを解放する
g.Dispose();
return canvas;
}
public async Task<Image> GetZoomImageFromFile(List<string> FileList, string fileName, Size imgSize, bool zip)
{
Image img = null;
if (zip)
{
int index = FileList.FindIndex(file => file == fileName);
var ZipArc = ZipArcEntryList[index];
using var zipStream = ZipArc.Open();
await Task.Run(() => img = GetZoomImageFromStream(zipStream, imgSize));
}
else
{
using FileStream fs = File.OpenRead(fileName);
await Task.Run(() => img = GetZoomImageFromStream(fs, imgSize));
}
return img;
}
}
}
| 99189009eddd13480ee220939cda29afb5cfa3ea | [
"C#"
] | 6 | C# | ShiraiShiika/Graico | 8ffc8e2c569a0281fbb3ae7545540551af13f53f | 0f3ba23ac9d5bd0dfd48b5dfd421443d97365f04 |
refs/heads/master | <file_sep>#!/bin/bash
#===========================================================================
# Title : ctuppi.sh
# Description : Script for setting up my dev environment
# Author : aristaako
# Version : 2.3
# Notes : Check readme.md for commands cheatsheet
# Usage : Just run the thing and hope for the best. See below
# for further instructions
#===========================================================================
VERSION=2.3
CTUPPIID=ctuppi023000
LOCK=/tmp/$CTUPPIID.lock
DEFAULT_DISTRO=ubuntu
USER_DISTRO=
VIRTUALIZED=false
RED='\033[0;31m'
GREEN='\033[0;32m'
NO_COLOR='\033[0m'
declare -a OPERATION_STARTED=()
declare -a FAILED=()
declare -a OPERATION_LIST=()
showHelp() {
cat <<-END
Usage: ./ctuppi.sh
(to run Ctuppi)
or ./ctuppi.sh [options]
(to run Ctuppi options)
where options include:
-h | --help print help message to output stream
-v | --version print Ctuppi version information
END
}
showVersion() {
echo "Ctuppi $VERSION"
}
inquire_virtualbox() {
while true; do
read -p "Is this a virtualized environment? (y/n) " yn
case $yn in
[Yy]* ) echo "Roger that."; VIRTUALIZED=true; break;;
[Nn]* ) echo "Okay."; break;;
* ) echo "Please answer yes or no.";;
esac
done
}
is_distro_debian() {
while true; do
read -p "Are you using debian based distro? (y/n) " yn
case $yn in
[Yy]* ) echo "Great."; USER_DISTRO=debian; break;;
[Nn]* ) echo "Sadness. Ctuppi does not support your distro."; exit;;
* ) echo "Please answer yes or no.";;
esac
done
}
inquire_distro() {
while true; do
read -p "Are you using ubuntu based distro? (y/n) " yn
case $yn in
[Yy]* ) echo "Okay."; USER_DISTRO=ubuntu; break;;
[Nn]* ) is_distro_debian; break;;
* ) echo "Please answer yes or no.";;
esac
done
}
update_apt_packages() {
sudo apt update -y -q
}
apt_install() {
display_name=$1
[[ ! -z "$2" ]] && package_name=$2 || package_name=$display_name
echo "Installing $display_name"
operation="Installation: $package_name"
OPERATION_STARTED+=( "$operation" )
sudo apt install $package_name -y -q || FAILED+=( "$operation" )
}
apt_remove() {
display_name=$1
[[ ! -z "$2" ]] && package_name=$2 || package_name=$display_name
echo "Removing $display_name"
operation="Remove: $package_name"
OPERATION_STARTED+=( "$operation" )
if [ "$(which $package_name 2> /dev/null)" != "" ]; then
sudo apt remove $package_name -y -q || FAILED+=( "$operation" )
else
echo "$package_name not found"
fi
}
install_konsole() {
apt_install "konsole"
}
install_nerd_font() {
echo "Installing FiraCode Nerd Font"
OPERATION_STARTED+=( "Installation: FiraCode Nerd Font" )
wget https://github.com/ryanoasis/nerd-fonts/releases/download/v2.1.0/FiraCode.zip
unzip FiraCode.zip -d FiraCode
mkdir ~/.local/share/fonts/
cp -r FiraCode/* ~/.local/share/fonts/
rm FiraCode.zip
rm -rf FiraCode
fc-cache
}
copy_konsole_profile() {
echo "Copying konsole profile"
mkdir -p ~/.local/share/konsole
cp files/konsole.profile ~/.local/share/konsole
}
copy_bash_configs() {
echo "Copying bash configs to user root"
cp files/bash_aliases ~/.bash_aliases
cp files/bashrc ~/.bashrc
mkdir -p ~/opt/git-prompt
cp files/git-prompt.sh ~/opt/git-prompt/git-prompt.sh
}
set_username_to_bash_configs() {
echo "Setting current user [$USER] for the bash config paths"
sed -i "s/_username_/$USER/" ~/.bash_aliases
}
install_git() {
apt_install "git"
apt_install "git-cola"
apt_install "kdiff3"
}
configure_git() {
git_username=$(git config user.name)
git_useremail=$(git config user.email)
if [[ -z "$git_username" ]]; then
echo "Configuring git: git username not set"
username=
while [[ $username = "" ]]; do
read -p "Enter git username: " username
done
git config --global user.name "$username"
fi
if [[ -z "$git_useremail" ]]; then
echo "Configuring git: git email not set"
email=
while [[ $email = "" ]]; do
read -p "Enter git email: " email
done
git config --global user.email "$email"
fi
echo "Configuring git: merge tool kdiff3"
git config --global merge.tool kdiff3
}
install_utils() {
apt_install "curl"
echo "Installing ripgrep"
OPERATION_STARTED+=( "Installation: ripgrep" )
curl -LO https://github.com/BurntSushi/ripgrep/releases/download/12.1.1/ripgrep_12.1.1_amd64.deb
sudo dpkg -i "ripgrep_12.1.1_amd64.deb"
echo "Removing ripgrep deb"
OPERATION_STARTED+=( "Remove: ripgrep_12.1.1_amd64.deb" )
rm "ripgrep_12.1.1_amd64.deb"
apt_install "python-pip"
apt_install "python3"
apt_install "python3-pip"
apt_install "sqlite3" "libsqlite3-dev"
apt_install "ruby"
echo "Installing mailcatcher"
OPERATION_STARTED+=( "Installation: mailcatcher" )
sudo gem install mailcatcher
apt_install "gedit"
wget -qO - https://packagecloud.io/AtomEditor/atom/gpgkey | sudo apt-key add
sudo sh -c 'echo "deb [arch=amd64] https://packagecloud.io/AtomEditor/atom/any/ any main" > /etc/apt/sources.list.d/atom.list'
update_apt_packages
apt_install "atom"
apt_install "tree"
echo "Installing SDKMAN!"
OPERATION_STARTED+=( "Installation: SDKMAN!" )
curl -s "https://get.sdkman.io" | bash
source "/home/$USER/.sdkman/bin/sdkman-init.sh"
apt_install "Silver Searcher" "silversearcher-ag"
apt_install "ssh" "openssh-server"
if [ "$VIRTUALIZED" == "false" ]; then
apt_install "solaar for Logitech bluetooth devices" "solaar"
fi
}
install_tmux() {
apt_install "tmux"
echo "Copying tmux configs to user root"
cp files/tmux.conf ~/.tmux.conf
cp files/splitter ~/.tmux/splitter
cp files/tmux-status.sh ~/.tmux/tmux-status.sh
echo "Downloading tpm"
OPERATION_STARTED+=( "Installation: tpm" )
git clone https://github.com/tmux-plugins/tpm ~/.tmux/plugins/tpm
echo "Downloading tmux reset"
OPERATION_STARTED+=( "Installation: tmux reset" )
curl -Lo ~/.tmux/reset --create-dirs \
https://raw.githubusercontent.com/hallazzang/tmux-reset/master/tmux-reset
}
install_nvm() {
echo "Installing nvm"
OPERATION_STARTED+=( "Installation: nvm" )
curl -o- https://raw.githubusercontent.com/nvm-sh/nvm/v0.39.1/install.sh | bash
refresh_bashrc
}
install_node() {
echo "Installing node"
OPERATION_STARTED+=( "Installation: node" )
nvm install node
}
install_java() {
apt_install "default java jdk" "default-jdk"
}
install_docker() {
echo "Preparing to install docker"
echo "First cleaning older versions"
apt_remove "docker"
apt_remove "docker-engine"
apt_remove "docker.io"
apt_remove "containerd"
apt_remove "runc"
echo "Installing packages to allow apt to use repositories over HTTPS"
apt_install " apt-transport-https" "apt-transport-https"
apt_install " ca-certificates" "ca-certificates"
apt_install " gnupg-agent" "gnupg-agent"
apt_install " software-properties-common" "software-properties-common"
if [ "$DEFAULT_DISTRO" == "$USER_DISTRO" ]; then
echo "Adding Docker's official GPG key"
curl -fsSL "https://download.docker.com/linux/$DEFAULT_DISTRO/gpg" | sudo apt-key add -
echo "Setting up docker stable repository"
sudo add-apt-repository \
"deb [arch=amd64] https://download.docker.com/linux/$DEFAULT_DISTRO \
$(lsb_release -cs) \
stable" -y
update_apt_packages
echo "Installing docker"
apt_install " docker-ce" "docker-ce"
apt_install " docker-ce-cli" "docker-ce-cli"
apt_install " containerd.io" "containerd.io"
else
echo "Downloading docker "
sudo curl -L "https://download.docker.com/linux/$USER_DISTRO/dists/$(lsb_release -cs)/pool/stable/amd64/docker-ce_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb" -o "~/Downloads/docker-ce_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb"
echo "Downloading docker cli"
sudo curl -L "https://download.docker.com/linux/$USER_DISTRO/dists/$(lsb_release -cs)/pool/stable/amd64/docker-ce-cli_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb" -o "~/Downloads/docker-ce-cli_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb"
echo "Downloading containerd.io"
sudo curl -L "https://download.docker.com/linux/$USER_DISTRO/dists/$(lsb_release -cs)/pool/stable/amd64/containerd.io_1.3.7-1_amd64.deb" -o "~/Downloads/containerd.io_1.3.7-1_amd64.deb"
echo "Installing containerd.io"
sudo dpkg -i "~/Downloads/containerd.io_1.3.7-1_amd64.deb"
echo "Installing docker-ce-cli"
sudo dpkg -i "~/Downloads/docker-ce-cli_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb"
echo "Installing docker-ce"
sudo dpkg -i "~/Downloads/docker-ce_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb"
echo "Removing downloaded docker debs"
rm "~/Downloads/containerd.io_1.3.7-1_amd64.deb"
rm "~/Downloads/docker-ce-cli_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb"
rm "~/Downloads/docker-ce_19.03.9~3-0~$USER_DISTRO-$(lsb_release -cs)_amd64.deb"
fi
sudo usermod -aG docker "$USER"
echo "Installing docker-compose with pip3"
OPERATION_STARTED+=( "Installation: docker-compose" )
pip3 install --user docker-compose
}
install_maven() {
apt_install "maven"
}
install_gradle() {
echo "Installing gradle"
OPERATION_STARTED+=( "Installation: gradle" )
sdk install gradle 6.6.1
}
install_aws() {
apt_install "AWS" "awscli"
}
install_clojure() {
apt_install "readline wrapper" "rlwrap"
curl -O https://download.clojure.org/install/linux-install-1.10.1.727.sh
chmod +x linux-install-1.10.1.727.sh
sudo ./linux-install-1.10.1.727.sh
echo "Removing clojure installation script"
rm linux-install-1.10.1.727.sh
echo "Install leiningen"
OPERATION_STARTED+=( "Installation: Leiningen" )
curl -o ~/bin/lein --create-dirs https://raw.githubusercontent.com/technomancy/leiningen/stable/bin/lein
chmod a+x ~/bin/lein
PATH=$PATH:~/bin
lein
}
install_emacs() {
apt_install "Emacs" "emacs"
echo "Installing emacs live"
OPERATION_STARTED+=( "Installation: Emacs live" )
yes '' |bash <(curl -fksSL https://raw.github.com/overtone/emacs-live/master/installer/install-emacs-live.sh)
echo "Installing flowapack"
OPERATION_STARTED+=( "Installation: flowa-pack for Emacs" )
git clone https://github.com/flowa/flowa-pack.git ~/.flowa-pack
mv ~/.flowa-pack/.emacs-live.el ~/.emacs-live.el
cd ~/.flowa-pack
git submodule init
git submodule update
cd ~/.flowa-pack/lib/helm
make
cd ~/
echo "Setting emacs as the default editor"
echo 'export EDITOR=~/bin/emacsnw' >> ~/.bashrc
}
install_vscode() {
wget -qO- https://packages.microsoft.com/keys/microsoft.asc | gpg --dearmor > packages.microsoft.gpg
sudo install -o root -g root -m 644 packages.microsoft.gpg /etc/apt/trusted.gpg.d/
sudo sh -c 'echo "deb [arch=amd64 signed-by=/etc/apt/trusted.gpg.d/packages.microsoft.gpg] https://packages.microsoft.com/repos/vscode stable main" > /etc/apt/sources.list.d/vscode.list'
update_apt_packages
apt_install "Visual Studio Code" "code"
}
install_brave() {
curl -s https://brave-browser-apt-release.s3.brave.com/brave-core.asc | sudo apt-key --keyring /etc/apt/trusted.gpg.d/brave-browser-release.gpg add -
echo "deb [arch=amd64] https://brave-browser-apt-release.s3.brave.com/ stable main" | sudo tee /etc/apt/sources.list.d/brave-browser-release.list
update_apt_packages
apt_install "Brave browser" "brave-browser"
}
install_mpv() {
if [ "$DEFAULT_DISTRO" == "$USER_DISTRO" ]; then
echo "Adding repository for mpv"
sudo add-apt-repository ppa:mc3man/mpv-tests -y
fi
apt_install "mpv"
}
refresh_bashrc() {
echo "Refreshing .bashrc"
source ~/.bashrc
}
display_greetings() {
echo "Thank you for using Ctuppi!"
echo "Install tmux plugins by opening tmux and pressing PREFIX + I (capital i)."
echo "Over and out."
}
create_operation_list() {
longest_operation_name_length=0
for item in "${OPERATION_STARTED[@]}"
do
item_length=${#item}
if [ "$item_length" -gt "$longest_operation_name_length" ]; then
longest_operation_name_length=$item_length
fi
done
for value in "${OPERATION_STARTED[@]}"
do
value_length=${#value}
value_shorter_than_longest=$((longest_operation_name_length-value_length))
spaces=
for i in $(seq 1 $value_shorter_than_longest); do spaces+=" "; done
operation_row=
if [[ ! " ${FAILED[*]} " =~ " ${value} " ]]; then
INSTALLED+=( "$value" )
operation_row="$spaces$value $GREEN OK $NO_COLOR"
OPERATION_LIST+=( "$operation_row" )
else
operation_row="$spaces$value $RED NOK $NO_COLOR"
OPERATION_LIST+=( "$operation_row" )
fi
done
}
print_operation_list() {
echo ""
for item in "${OPERATION_LIST[@]}"
do
echo -e "$item"
done
echo ""
}
setup_environment() {
echo "Setting up dev environment"
inquire_virtualbox
inquire_distro
update_apt_packages
install_konsole
install_nerd_font
copy_konsole_profile
copy_bash_configs
set_username_to_bash_configs
install_git
configure_git
install_utils
install_tmux
install_nvm
refresh_bashrc
install_node
install_java
install_docker
install_maven
install_gradle
install_aws
install_clojure
install_emacs
install_vscode
install_brave
install_mpv
refresh_bashrc
create_operation_list
print_operation_list
display_greetings
}
invalid() {
echo "Invalid argument $1"
echo "Use argument -h or --help for instructions."
}
while getopts ":hv-:" opt; do
case $opt in
h)
showHelp
exit 0 ;;
v)
showVersion
exit 0 ;;
-)
case "$OPTARG" in
version)
showVersion
exit 0 ;;
help)
showHelp
exit 0 ;;
*)
invalid "--${OPTARG}"
exit 0 ;;
esac ;;
*)
invalid "-$OPTARG"
exit 0 ;;
esac
done
cleanup() {
rm -f $LOCK
}
startup() {
if [ -f "$LOCK" ]; then
echo "Ctuppi already running"
exit
fi
touch $LOCK
trap cleanup EXIT
}
startup
setup_environment<file_sep>#aliakset
alias aliakset='less /home/_username_/.bash_aliases'
alias reload='. ~/.bashrc'
alias forget='unalias -a'
alias gs='git status'
alias GS='git status'
alias gitco='git checkout'
alias rbmaster='git pull --rebase origin master:master'
alias rbmain='git pull --rebase origin main:main'
alias rbdev='git pull --rebase origin dev:dev'
alias rbdevelop='git pull --rebase origin develop:develop'
alias eiku='git reset HEAD~'
alias eikumerge='git reset --soft HEAD@{1}'
alias eikueka='git update-ref -d HEAD'
alias kokis='git-cola'
alias apdate='sudo apt-get update'
alias doi='docker image'
alias doc='docker container'
alias g='cd /home/_username_/git/'
alias sudona='sudo $(history -p !!)'
#Docker/Leiningen/etc. service starting
alias docsta='sudo service docker start'
alias docsto='sudo service docker stop'
alias sohva='sudo docker start couchdb'
alias haamu='lein doo phantom'
alias viikuna='rlwrap lein figwheel'
alias figge='rlwrap lein do clean, figwheel'
alias leet='lein test'
alias leinstall='lein install'
alias jousi='sudo mvn spring-boot:run'
alias mcatch='mailcatcher --http-ip=0.0.0.0'
alias ..='cd ..'
alias ..2='cd ../..'
alias ..3='cd ../../..'
alias ..4='cd ../../../..'
alias ..5='cd ../../../../..'
alias ..6='cd ../../../../../..'
alias ..7='cd ../../../../../../..'
alias ..8='cd ../../../../../../../..'
alias lst='ls --full-time'
alias whereami='pwd'
alias syslogaa='tail -f /var/log/syslog'
alias aallot='mpv https://stream.bauermedia.fi/auranaallot/auranaallot_64.aac'
alias puhe='mpv http://mediau.yle.fi/liveradiopuhe'
alias ylex='mpv http://mediau.yle.fi/liveylex'
function rgf {
rg -g "*$1*" --files ;
}
function cdl { cd $1; ls; }
function cdll { cd $1; ls -l; }
function cd3 { cd $1; tree; }
function cdg {
kohde=$(find . -name $1 |head -1 |grep -Eo "/[ /._A-Za-z0-9\-]+/")
kohde=$(sed 's/^.//' <<< ${kohde})
cd ${kohde}
}<file_sep>#!/bin/bash
CURRENT_PATH=$1
SHOW_FOLDER=$2
DEFAULT_COLOR='colour34'
TEXT_COLOR='colour240'
BRANCH_COLOR_NOTHING='colour240'
BRANCH_COLOR_UNTRACKED='colour166'
BRANCH_COLOR_COMMITED='colour28'
BRANCH_COLOR_CHANGES='colour196'
BRANCH_COLOR_CONFLICTS='colour196'
BRANCH_ICON=""
BRANCH_ICON_CONFLICTS="✖"
FOLDER_ICON=""
FOLDER_ICON_COLOR='colour243'
FILE_ICON=""
FILE_ICON_COLOR='colour243'
SYMLINK_ICON=""
SYMLINK_ICON_COLOR='colour243'
SPLITTER_ICON=""
SPLITTER_ICON_COLOR='colour243'
get_git() {
local is_git
is_git="$(cd $CURRENT_PATH; git rev-parse --is-inside-work-tree 2> /dev/null)"
echo "$is_git"
}
get_git_status() {
local git_status
git_status="$(cd $CURRENT_PATH; git status 2> /dev/null)"
echo "$git_status"
}
get_branch_icon() {
local git_status
git_status=$(cd $CURRENT_PATH; git status 2> /dev/null)
case "${git_status}" in
*"nothing to commit"*)
echo "#[fg=$BRANCH_COLOR_NOTHING]$BRANCH_ICON" #colour240
;;
*"Changes not staged"*)
echo "#[fg=$BRANCH_COLOR_CHANGES]$BRANCH_ICON" #colour196
;;
*"nothing added to commit but untracked"*)
echo "#[fg=$BRANCH_COLOR_UNTRACKED]$BRANCH_ICON" #colour166
;;
*"conflicts"*)
echo "#[fg=$BRANCH_COLOR_CONFLICTS]$BRANCH_ICON_CONFLICTS" #✖colour196
;;
*"Changes to be committed"*)
echo "#[fg=$BRANCH_COLOR_COMMITED]$BRANCH_ICON" #colour28
;;
esac
}
get_branch_name() {
local branch_name
branch_name="$(cd $CURRENT_PATH; git rev-parse --abbrev-ref HEAD)"
echo $branch_name
}
get_folder_status() {
local folder_count="$(cd $CURRENT_PATH; find . -mindepth 1 -maxdepth 1 -type d | wc -l)"
local file_count="$(cd $CURRENT_PATH; find . -mindepth 1 -maxdepth 1 -type f | wc -l)"
local symlink_count="$(cd $CURRENT_PATH; find . -mindepth 1 -maxdepth 1 -type l | wc -l)"
folder="#[fg=$FOLDER_ICON_COLOR]$FOLDER_ICON #[fg=$TEXT_COLOR]${folder_count}"
file="#[fg=$FILE_ICON_COLOR]$FILE_ICON #[fg=$TEXT_COLOR]${file_count}"
symlink="#[fg=$SYMLINK_ICON_COLOR]$SYMLINK_ICON #[fg=$TEXT_COLOR]${symlink_count}"
splitter="#[fg=$SPLITTER_ICON_COLOR]$SPLITTER_ICON#[fg=$TEXT_COLOR]"
folder_status="$folder $file $symlink $splitter"
echo $folder_status
}
set_status_bar() {
local git_status
local branch_name
local branch_icon
local branch_status
git_status=$(get_git_status)
branch_name="$(get_branch_name)"
branch_icon="$(get_branch_icon)"
splitter="#[fg=$SPLITTER_ICON_COLOR]$SPLITTER_ICON#[fg=$TEXT_COLOR]"
branch_status="$branch_icon#[fg=$DEFAULT_COLOR] ${branch_name} $splitter"
echo $branch_status
}
folder_status=""
if [ "$SHOW_FOLDER" = "true" ]; then
folder_status=$(get_folder_status)
fi
IS_GIT=$(get_git)
status=""
if [ $IS_GIT = true ]; then
status=$(set_status_bar)
echo $folder_status $status
else
echo $folder_status
fi
<file_sep>#!/bin/bash
#===========================================================================
# Title : bunsen.sh
# Description : Script for copying bunsenlabs configs
# Author : aristaako
# Version : 1.0
# Usage : Run the script. It will copy the files. Yes?
#===========================================================================
VERSION=1.0
BUNSENID=bunsen010000
LOCK=/tmp/$BUNSENID.lock
BUNSENWEATHER_SCRIPT_FILE=$HOME/.config/conky/scripts/bunsenweather.sh
showHelp() {
cat <<-END
Usage: ./bunsen.sh
(to run Bunsen config copy script)
or ./bunsen.sh [options]
(to run Bunsen config copy script options)
where options include:
-h | --help print help message to output stream
-v | --version print Bunsen config copy script version information
END
}
showVersion() {
echo "Bunsen config copy script $VERSION"
}
copy_bunsenlabs_configs() {
echo "Copying bunsenlabs configs"
cp conky.conf ~/.config/conky/conky.conf
cp prepend.csv ~/.config/jgmenu/prepend.csv
}
set_bunsenweather_apikey() {
read -p "Type your OpenWeatherMap API key: " apikey
sed -i "s/api=$/api=$apikey/" $BUNSENWEATHER_SCRIPT_FILE
}
bunsenweather_apikey_setup() {
while true; do
read -p "Would you like to give OpenWeatherMap API key to bunsenweather? (y/n) " yn
case $yn in
[Yy]* ) set_bunsenweather_apikey; break;;
[Nn]* ) break;;
* ) echo "Please answer yes or no.";;
esac
done
}
setup_bunsenlabs() {
echo "Setting up bunsenlabs"
copy_bunsenlabs_configs
bunsenweather_apikey_setup
}
invalid() {
echo "Invalid argument $1"
echo "Use argument -h or --help for instructions."
}
while getopts ":hv-:" opt; do
case $opt in
h)
showHelp
exit 0 ;;
v)
showVersion
exit 0 ;;
-)
case "$OPTARG" in
version)
showVersion
exit 0 ;;
help)
showHelp
exit 0 ;;
*)
invalid "--${OPTARG}"
exit 0 ;;
esac ;;
*)
invalid "-$OPTARG"
exit 0 ;;
esac
done
cleanup() {
rm -f $LOCK
}
startup() {
if [ -f "$LOCK" ]; then
echo "Bunsen config copying already running"
exit
fi
touch $LOCK
trap cleanup EXIT
}
startup
setup_bunsenlabs
<file_sep># Ctuppi
Script for setting up my personal dev environment.
## Setup info
* Konsole
* Git
* Curl
* Ripgrep
* Pip
* Python 3 & Pip3
* SQLite3
* Ruby
* Mailcatcher
* Gedit
* Atom
* Tree
* SDKMAN!
* Silver Searcher
* OpenSSH Server
* Solaar
* Tmux
* Nvm
* Node
* Java (current default jdk)
* Docker
* Maven
* Gradle
* AWS Command Line Interface
* Clojure
* Emacs
* Visual Studio Code
* Brave
* mpv
## Todo
* Install everything silently
* Improve emacs cheatsheet
* Install mpv for debian
* Ensure Clojure has compatible Java version (8 or 11)
* List failed installations after ctuppi
## Thanks
First of all, I would like to thank all those who have made my dev environment possible. I thank you all Tmux-plugin creating gurus and Flowa for the great emacs live pack.
## Cheatsheet
Because sometimes I forget some of the rarely used commands
### Tmux
#### Common
```
prefix = Ctrl-b
```
```
Reload tmux conf: prefix + r
```
```
Refresh tmux client: prefix + R
```
```
Split window horizontally: prefix + Shift-2
Split window vertically: prefix + Shift-5
```
```
Move cursor to pane above: Alt-↑
Move cursor to pane below: Alt-↓
Move cursor to pane on the left: Alt-←
Move cursor to pane on the right: Alt-→
```
```
Open tmux window 10: prefix + F1
Open tmux window 11: prefix + F2
Open tmux window 12: prefix + F3
Open tmux window 13: prefix + F4
Open tmux window 14: prefix + F5
Open tmux window 15: prefix + F6
Open tmux window 16: prefix + F7
Open tmux window 17: prefix + F8
Open tmux window 18: prefix + F9
Open tmux window 19: prefix + F10
Open tmux window 20: prefix + F11
Open tmux window 21: prefix + F12
```
```
Close pane: prefix + x
```
#### Splitter
```
Split top left pane into 4 panes: prefix + j
```
#### TPM - Tmux plugin manager
```
Install plugins: prefix + I
Update plugins: prefix + u
Uninstall plugins: prefix + Alt-u (Uninstalls plugins not in the list of plugins in .tmux.conf)
```
#### Tmux resurrect
```
Save session: prefix + Ctrl-s
Restore session: prefix + Ctrl-r
```
#### Tmux yank aka copy to clipboard
Normal mode
```
Copy text from command line: prefix + y
Copy current pane's working directory: prefix + Y
```
Copy mode
```
Copy selection: y
Copy selection and paste into command line: Y
```
#### Tmux sidebar
```
Open sidebar: prefix + Tab
Open sidebar and focus: prefix + Backspace
```
#### Tmux open
Copy mode
```
Open selection with default program: o
Open selection with $EDITOR: Ctrl-o
Search selection from a search engine (default= Google): Shift-s
```
Change default search engine
```
Add line to .tmux.conf:
set -g @open-S 'https://www.duckduckgo.com/'
```
### Emacs
```
Split window vertically: Ctrl-x 2
Split window horizontally: Ctrl-x 3
```
```
Revert buffer: Ctrl-c x
```
```
Cancel command: Ctrl-g
```
```
Find and open file into current window: Ctrl-x Ctrl-f
```
```
Find and open file into another window: Ctrl-x 4 f
Find and open file into another frame: Ctrl-x 5 f
```
```
Search text forwards: Ctrl-s
Search text backwards: Ctrl-r
```
```
Kill whitespace: Alt-\ (Alt-AltGr-+)
```
```
Undo: Ctrl-_ (Ctrl-Shift--)
Undo tree: Ctrl-x u
```
```
Close window: Ctrl-x 0
```
```
Arrange windows evenly: Ctrl-x +
```
## Author
<NAME> - [aristaako](https://github.com/aristaako) | adf3313ac4bad64a2c03d97e35a710cb68218614 | [
"Markdown",
"Shell"
] | 5 | Shell | aristaako/ctuppi | 73e2b17db30ad93fcb0ebfff592c3e2075fa3e45 | d67e4c241bcfb7f9a8b9c759e0fa71d8f9933e44 |
refs/heads/master | <file_sep>#include <vector>
int fib(int n) {
static vector<int> memo;
if (memo.size() == 0) {
memo.push_back(1);
memo.push_back(1);
}
if (n >= memo.size()) {
memo.push_back(fib(n-2) + fib(n-1));
}
return memo[n];
}
int climbStairs(int n) {
return fib(n);
}
<file_sep>#include <stdio.h>
int main (int argc, char * argv[]) {
int t;
scanf("%d", &t);
getchar();
while (t--) {
int count = 0;
char c;
while ((c = getchar()) != '\n') {
switch(c) {
case 'B':
count += 2;
break;
case 'A':
case 'D':
case 'O':
case 'P':
case 'Q':
case 'R':
count += 1;
break;
default:
break;
}
}
printf("%d\n", count);
}
}
<file_sep>#include <stdio.h>
// return number of bits used in bin representation, up to msot significant 1
int numBits(int a) {
int bits = 0;
if (a == 0) {
return 0;
}
for (int i = 0; i < 32; i++) {
int test = 1 << i;
if (test > a) {
break;
}
bits++;
}
return bits;
}
// return number of bits at front that are the same
int sameBitsFront(int a, int b) {
int lenA = numBits(a);
int lenB = numBits(b);
int sameBits = 0;
// extract first bit of both while both are equal
do {
int chkA = a & 1 << (lenA - 1);
int chkB = b & 1 << (lenB - 1);
if ((chkA == 0 && chkB != 0) || (chkB == 0 && chkA != 0)) {
break;
}
sameBits++;
lenA--;
lenB--;
} while(lenA && lenB);
return sameBits;
}
int main(int argc, char *argv[]) {
int n;
scanf("%d", &n);
while (n--) {
unsigned int a, b;
scanf("%d %d", &a, &b);
int front = sameBitsFront(a, b);
printf("%d\n", numBits(a) - front + numBits(b) - front);
}
}
<file_sep>#include <stdio.h>
int abs(int a) {
return a < 0 ? -a: a;
}
int main(int argc, char *argv[]) {
int n;
scanf("%d", &n);
int lead = 0;
int maxLead = -1;
int winner;
for (int i = 0; i < n; i++) {
int a, b;
scanf("%d %d", &a, &b);
lead = lead + a - b;
if (abs(lead) > maxLead) {
maxLead = abs(lead);
if (lead < 0) {
winner = 2;
} else {
winner = 1;
}
}
}
printf("%d %d\n", winner, abs(maxLead));
}
<file_sep>#include <stdio.h>
char adds(char a, char b) {
if (a <= b) {
return b - a;
} else {
return 26 - a + b;
}
}
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
char c = getchar();
while (t--) {
int total = 0;
int wordLen = 0;
do {
char last = c;
c = getchar();
if (c == '\n' || c == EOF)
break;
wordLen++;
if (last == '\n') {
total += 2; // load and print first char
} else {
total += 1 + adds(last, c); // inc * adds(last, c) and print this char
}
} while (c != '\n' && c != EOF);
if (total > 11 * wordLen) {
printf("NO\n");
} else {
printf("YES\n");
}
}
}
<file_sep>#include <stdio.h>
int main (int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int n, c;
scanf("%d %d", &n, &c);
int total = 0;
int temp;
for (int i = 0; i < n; i++) {
scanf("%d", &temp);
total += temp;
}
if (total <= c) {
printf("Yes\n");
} else {
printf("No\n");
}
}
}
<file_sep>#include <stdio.h>
bool isLetter(char c) {
return (c >= 65 && c <= 90) || (c >= 97 && c <= 122);
}
bool isCap(char c) {
return c < 97;
}
bool isSpace(char c) {
return c == 95;
}
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
char alpha[26];
getchar();
for (int i = 0; i < 26; i++) {
alpha[i] = getchar();
}
while (t--) {
char sentence[101];
scanf("%s", &sentence);
for (int i = 0; sentence[i] != 0; i++) {
char *curr = &sentence[i];
if (isSpace(*curr)) {
*curr = ' ';
} else if (isLetter(*curr)) {
if (isCap(*curr)) {
sentence[i] = alpha[*curr - 65] - 32;
} else {
sentence[i] = alpha[*curr - 97];
}
}
}
printf("%s\n", sentence);
}
}
<file_sep>#include <stdio.h>
#include <math.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int p, s;
float b, v;
scanf("%d %d", &p, &s);
b = (p - (float)sqrt((float)(p * p - 24 * s))) / 12;
v = b * (s / 2 - p * b / 4 + b * b);
printf("%.2f\n",v);
}
return 0;
}
<file_sep>#include <stdio.h>
#include <math.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
unsigned long long n;
scanf("%llu", &n);
unsigned long long fac = 5, count = 0;
while (fac <= n) {
count += n / fac;
fac *= 5;
}
printf("%llu\n", count);
}
}
<file_sep>#include <stdio.h>
#include <string>
#include <vector>
#include <iostream>
using namespace std;
class chunk {
public:
int freq;
char val;
chunk(int a, char b) :
freq(a),
val(b)
{}
};
string chunksToStr(vector<chunk> vals) {
string final = "";
for (int i = 0; i < vals.size(); i++) {
final += (char)(vals[i].freq + '0');
final += vals[i].val;
}
return final;
}
vector<chunk> strToChunks(string prev) {
vector<chunk> parse;
int count = 1;
char curr = prev[0];
for (int i = 1; i <= prev.size(); i++) {
if (i == prev.size()) {
parse.push_back(chunk(count, curr));
} else if (prev[i] != curr) {
parse.push_back(chunk(count, curr));
curr = prev[i];
count = 1;
} else {
count++;
continue;
}
}
return parse;
}
string countAndSay(int n) {
if (n == 1) {
return "1";
} else {
string prev = countAndSay(n - 1);
vector<chunk> parse = strToChunks(prev);
return chunksToStr(parse);
}
}
int main(int argc, char *argv[]) {
for (int i = 1; i < 6; i++) {
cout << countAndSay(i) << endl;
}
}
<file_sep>#!/usr/bin/bash
lynx http://www.codechef.com/problems/easy -dump > temp
sed '/\[1\]CodeChef/,/^References$/ d' temp > page
mv page temp
sed '1,60d' temp > page
mv page temp
sed '/users\|submit\|status/ d' temp > page
mv page temp
sed '/javascript/,/http:\/\/www.codechef.com\/goforgold\n[1-9][0-9]*\. http:\/\/www.codechef.com\/goforgold/ d' temp > page
mv page temp
sed 's/ *[1-9][0-9]*\. http:\/\/www.codechef.com\/problems\///' temp > page
for prb in $( cat page ); do
echo "downloading: "$prb
lynx http://www.codechef.com/problems/$prb -dump > $prb
echo "sedding"
mv $prb temp
sed '1,68d' temp > $prb
mv $prb temp
sed 's/\[[1-9][0-9]*\]//' temp > $prb
mv $prb temp
sed '/___*/,/http:\/\/www.codechef.com\/goforgold\n[1-9][0-9]*\. http:\/\/www.codechef.com\/goforgold/ d' temp > $prb
done
rm temp
<file_sep>#include <stdio.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
// use divide and conquer!
}
}
<file_sep>#include <stdio.h>
int main(int argc, char *argv[]) {
int a, b;
scanf("%d %d", &a, &b);
int diff = a - b;
if (diff % 10 == 9) {
diff -= 1;
} else {
diff += 1;
}
printf("%d\n", diff);
}
<file_sep>#include <stdio.h>
#include <vector>
#include <algorithm>
using namespace std;
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int n;
scanf("%d", &n);
int temp;
vector<int> pies, racks;
for (char i = 0; i < n; i++) {
scanf("%d", &temp);
pies.push_back(temp);
}
sort(pies.begin(), pies.end());
for (char i = 0; i < n; i++) {
scanf("%d", &temp);
racks.push_back(temp);
}
sort(racks.begin(), racks.end());
int i = 0, j = 0;
while (i < n && j < n) {
if (pies[i] <= racks[j]) {
i++;
}
j++;
}
printf("%d\n", i);
}
}
<file_sep>#include <stdio.h>
#include <math.h>
#define BIGINT_SIZE 200
class bigInt {
char digits[BIGINT_SIZE];
int len;
bigInt() {
len = 0;
}
template <type T> bigInt(T a) {
int bytes = sizeof(a);
long long max = pow((double)2, (double)bytes);
int len = ceil(log10(max));
}
void add(bigInt a) {
}
void sub(bigInt a) {
}
void mul(int a) {
}
void div(int a) {
}
};
int main(int argc, char *argv[]) {
}
<file_sep>#include <stdio.h>
#include <deque>
#include <algorithm>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int n;
scanf("%d", &n);
int temp;
std::deque<int> vals;
for (int i = 0; i < n; i++) {
scanf("%d", &temp);
vals.push_back(temp);
}
std::sort(vals.begin(), vals.end());
for (int i = 0; i < n; i++) {
printf("%d ", i % 2 == 0 ? vals.front() : vals.back());
i % 2 == 0 ? vals.pop_front() : vals.pop_back();
}
printf("\n");
}
}
<file_sep>#include <stdio.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int n;
scanf("%d", &n);
int songs[n];
for (int i = 0; i < n; i++) {
scanf("%d", &songs[i]);
}
int k;
scanf("%d", &k);
int smallerCount = 0;
for (int i = 0; i < n; i++) {
if (i != k - 1 && songs[i] < songs[k - 1]) {
smallerCount++;
}
}
printf("%d\n", smallerCount + 1);
}
}
<file_sep>#include <stdio.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int n;
scanf("%d", &n);
int nums[n];
int i;
for (i = 0; i < n; i++) {
scanf("%d", &nums[i]);
}
int inv = 0, loc = 0;
for (i = 0; i < n; i++) {
int j;
for (j = 1; j < n - i; j++) {
if (nums[j + i] < nums[i]) {
inv++;
if (j == 1) {
loc++;
}
}
}
}
if (inv ^ loc) {
printf("%s", "NO\n");
} else {
printf("%s", "YES\n");
}
}
}
<file_sep>#include <stdio.h>
#define START 2048
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int p, total = 0, div = START;
scanf("%d", &p);
while (div) {
total += p / div;
p %= div;
div /= 2;
}
printf("%d\n", total);
}
}
<file_sep>#include <stdio.h>
int main(int argc, char *argv[]) {
int n;
scanf("%d", &n);
while(n--) {
int lines;
scanf("%d", &lines);
int arr[lines][lines];
for (int i = 0; i < lines; i++) {
for (int j = 0; j <= i; j++) {
scanf("%d", &arr[i][j]);
}
}
for (int i = lines - 2; i >= 0; i--) {
for (int j = 0; j <= i; j++) {
arr[i][j] += arr[i + 1][j] > arr[i + 1][j + 1] ? arr[i + 1][j] : arr[i + 1][j + 1];
}
}
printf("%d\n", arr[0][0]);
}
}
<file_sep>#include <stdio.h>
#include <math.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
int t1, t2, t3, t4;
scanf("%d %d %d %d", &t1, &t2, &t3, &t4);
}
}
<file_sep>#include <stdio.h>
#include <math.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while (t--) {
float b, ls;
scanf("%f %f", &b, &ls);
printf("%f %f\n", sqrt(pow(ls, 2) - pow(b, 2)), sqrt(pow(ls, 2) + pow(b, 2)));
}
}
<file_sep>#include <stdio.h>
int main(int argc, char *argv[]) {
int t;
scanf("%d", &t);
while(t--) {
int n, a, b;
scanf("%d %d %d", &n, &a, &b);
int bitCount = 0;
for (int i = 0; i < n; i++) {
bitCount += (a & 1) + (b & 1);
a = a >> 1;
b = b >> 1;
}
int finalBits = bitCount == n ? n : bitCount > n ? 2 * n - bitCount : bitCount;
int ans = 0;
for (int i = 0; i < n; i++) {
if (i < finalBits) {
ans = ans | 1;
}
if (i != n - 1) {
ans = ans << 1;
}
}
printf("%d\n", ans);
}
}
<file_sep>#include <stdio.h>
#include <cctype>
#include <string>
#include <iostream>
using namespace std;
string reverseWords(string st) {
bool isword = 0;
int i = 0;
string buff = "";
string res = "";
for (int i = 0; i < st.size(); i++) {
if (isword) {
if (isspace(st[i])) {
isword = 0;
res = buff + res;
buff = "";
} else {
buff += st[i];
}
} else {
if (isspace(st[i])) {
continue;
} else {
isword = 1;
buff += st[i];
if (res.size() != 0)
res = " " + res;
}
}
}
return res;
}
int main() {
string test = " this is a string with leading and trailing spaces ";
cout << "|" << reverseWords(test) << "|";
}
<file_sep>#include <stdio.h>
#include <iostream>
#include <vector>
#include <algorithm>
int main() {
int t, n;
std::vector<int> vec;
scanf("%d", &t);
while (t--) {
scanf("%d", &n);
vec.push_back(n);
}
std::sort(vec.begin(), vec.end());
for (int i = 0; i < vec.size(); i++)
printf("%d\n", vec[i]);
}
| 6550b65311e13a84242cffa9fedd461c4fbe143e | [
"C++",
"Shell"
] | 25 | C++ | szhongren/probs | 5ce1008dd91be6bdc01a4b7d145c3a75051d9e68 | 3da25a6c5bd9ddae34b41f3bc9eb5dc62849f8b0 |
refs/heads/master | <repo_name>Frankie937/Python_basic<file_sep>/python기본_11.py
# #**11-1 모듈(쉽게 설명하면, 필요한 것끼리 부품처럼 만들어진 파일이라고 보면 됨.
# # 예를 들어, 자동차를 타다가 타이어가 마모가 되면 타이어만 교체하면 되는 것 처럼
# # 소프트웨어도 부품만 교체하거나 추가할 수 있게 만들면 유지보수도 쉽고, 코드의 재사용도 수월해지는 장점이 있음
# # 이런식으로 딱 필요한 것들끼리 부품처럼 잘 만드는 것을 '모듈화'라고 함
# # 파이썬에서는 함수 정의나 클래스 등의 파이썬 문장들을 담고 있는 파일을 모듈이라고 함 모듈은 확장자가 '.py'임)
# # 모듈은 내가 그 모듈을 쓰려는 파일과 같은 경로에 있거나 혹은 파이썬 라이브러리들이 모여있는 폴더에 있어야 사용 가능함!
# # # 방법1
# # import theater_module
# # theater_module.price(3)
# # theater_module.price_morning(4)
# # theater_module.price_soldier(5)
# # 방법2
# import theater_module as mv # 모듈명이 길 때, as를 활용
# mv.price(3)
# mv.price_morning(4)
# mv.price_soldier(5)
# # 방법3
# from theater_module import * # 이렇게 쓰면, 모듈 내의 함수를 바로 호출 가능
# price(3)
# price_morning(4)
# price_soldier(5)
# # 방법4
# from theater_module import price, price_morning # 필요한 함수만 불러올 수 있음
# price(3)
# price_morning(4)
# # 방법5
# from theater_module import price_soldier as price # 불러온 함수에도 as 활용가능
# price(5)
# #**11-2 패키지(모듈들을 모아놓은 집합)
# # 하나의 디렉토리에 여러 모듈 파일들을 갖다 놓은 것을 패키지라고 쉽게 이해하면 됨
# import travel.thailand # 주의할 점: import를 쓸 때 맨뒤에는 모듈이나 패키지만 가능! (클래스나 함수는 import를 직접 바로 할 수 없음!)
# trip_to = travel.thailand.ThailandPackage()
# trip_to.detail()
# # from import 구문에서는 모듈, 패키지, 클래스, 함수 모두 import 할 수 있음!
# from travel.thailand import ThailandPackage
# trip_to = ThailandPackage()
# trip_to.detail()
# from travel import vietnam
# trip_to = vietnam.VietnamPackage()
# trip_to.detail()
# #**11-3 __all__
# from travel import *
# trip_to = vietnam.VietnamPackage() # -> 오류 발생!! (__init__파일에 아무것도 없을 시)
# # travel 폴더 안에 __init__파일에 __all__ =[ ]을 정의해주지 않고 from travel import * 을 하게 되면 오류 발생
# # 그 이유는, import *은 travel 패키지 않에 모든 것을 갖고오겠다는 것인데 실제 사용할 때에는 개발자가 그 공개 범위를 설정해줘야 함!
# # 패키지 안에 포함된 것들 중에서 import되기 원하는 것만 공개하고 원하지 않는 것을 비공개로 설정할 수 있다는 의미
# #**11-4 모듈직접실행
# # 실제로 패키지나 모듈을 만들 때, 잘 동작하는지 테스트를 해봐야 함(if __name__ == "__main__" 구문 활용)
# # 모듈 내에서 실행되는 건지 외부에서 가져와서 실행하는 건지 구분해서 필요한 코드를 작성할 수 있음
# # thailand 모듈에 예제 있음(줄 5번 부터 11번까지)
# #**11-5 패키지, 모듈 위치 확인 방법
# import inspect
# import random
# print(inspect.getfile(random)) # 랜덤이라는 모듈이 어느 위치에 있는지 파일 정보를 알려주는 것
# from travel import *
# print(inspect.getfile(thailand))
# #**11-6 pip install (pip로 패키지 설치하기)
# # 지금 이미 수많은 패키지들이 존재하고, 지금도 누군가가 패키지를 새롭게 개발하고 있음
# # 그러므로, 파이썬은 새로운 코드를 무조건 다 작성하는 것보다 이미 잘 만들어진 패키지를 필요한 곳에 가져다 쓰는 것도 굉장히 중요함!!
# # pypi 검색(구글링) https://pypi.org/
# # beautifulsoup4 검색(웹스크래핑에 대한 유명한 패키지)
# # <terminal 에 작성하는 부분>
# # pip install beautifulsoup4 -> beautifulsoup4 패키지가 설치됨
# # pip list -> 현재 설치되어 있는 패키지들이 어떤 것이 있는 지 볼 수 있음
# # pip show beautifulsoup4 -> beautifulsoup4 패키지에 대한 정보를 알려줌
# # pip install --upgrade beautifulsoup4 -> 설치되어있는 패키지가 새로운 버전이 나와서 업그레이드가 필요할 때
# # pip uninstall beautifulsoup4 -> 패키지 삭제할 때
# #**11-7 내장함수(내장되어 있기 때문에 따로 import 할 필요 없이 바로 사용가능한 함수)
# # input : 사용자 입력을 받는 함수
# language = input("무슨 언어를 좋아하세요?")
# print("{0}은 아주 좋은 언어입니다.!".format(language))
# # dir : 어떤 객체를 넘겨줬을 때 그 객체가 어떤 변수와 함수를 가지고 있는지 표시
# print(dir())
# import random # 외장 함수
# print(dir()) # random이 추가되어 있음
# import pickle
# print(dir()) random, pickle이 추가되어 있음
# print(dir(random)) # random 모듈 내에서 쓸 수 있는 모든 것들이 나옴
# # 내장함수-list
# lst = [1, 2, 3]
# print(dir(lst)) # list 함수 내에서 쓸 수 있는 모든 것들이 나옴
# # 내장함수-문자열 str 함수
# name = "Jim" # 문자열 함수 내에서 쓸 수 있는 모든 것들이 나옴
# print(dir(name))
# # 더 많은 내장함수를 찾으려면,
# # 구글에 'list of python builtins'로 검색하면 'https://docs.python.org/ko/3/library/functions.html'사이트 클릭해서 확인하면 됨
# #**11-8 외장함수(직접 import 해서 사용해야 하는 함수)
# # 구글에 'list of python modules'로 검색하면 'https://docs.python.org/3/py-modindex.html' 외장함수 목록 확인 가능
# # glob : 경로 내의 폴더/ 파일 목록 조회 (윈도우 dir 과 똑같음)
# import glob
# print(glob.glob("*.py")) # 확장자가 py인 모든 파일에 대해서 알려줘라는 것
# # os : 운영체제에서 제공하는 기본 기능
# import os
# print(os.getcwd()) # 현재 디렉토리를 표시해달라는 의미
# folder = "sample_dir"
# if os.path.exists(folder): # sample_dir라는 폴더가 있으면 이 구문을 타라는 것
# print("이지 존재하는 폴더입니다.")
# os.rmdir(folder)
# print(folder, "폴더를 삭제하였습니다.")
# else:
# os.makedirs(folder) # 폴더 생성
# print(folder, "폴더를 생성하였습니다." )
# print(os.listdir()) # os.listdir ->glob과 비슷하게 쓸 수 있음
# # time : 시간 관련 함수들을 제공하는 외장함수
# import time
# print(time.localtime())
# print(time.strftime("%Y-%m-%d %H:%M:%S"))
# import datetime
# print("오늘 날짜는 ", datetime.date.today())
# # timedelta : 두 날짜 사이의 간격
# today = datetime.date.today() # 오늘 날짜 저장
# td = datetime.timedelta(days=100) # 100일 저장
# print("우리가 만난지 100일은", today + td) # 오늘부터 100일 후
# # 퀴즈 10
# # --내가 한 것--
# import byme
# byme.sign("김민서", "<EMAIL>")
# # --나도코딩--
# import byme
# byme.sign()
<file_sep>/python기본_8.py
# #**8-1표준 입출력
# print("Python","Java") # Python Java 출력
# print("Python"+"Java") # PythonJava 출력
# print("Python","Java",sep=",") # Python,Java 출력
# print("Python","Java",sep=" vs ") # Python vs Java 출력
# import sys
# print("Python","Java", file=sys.stdout) # 보이는 결과는 차이없어보이지만, stdout은 표준출력으로 찍히는 것이고, stderr은 표준에러로 처리되는 것
# print("Python","Java", file=sys.stderr) # 즉, stdout은 크게 신경쓸 필요 없는데 stderr는 확인을 해서 프로그램 코드를 수정하든지 해야 되는 것
# # 시험성적 출력
# scores = {"수학":0, "영어":50, "코딩":100}
# for subject, score in scores.items():
# print(subject, score)
# print(subject.ljust(8), str(score).rjust(4), sep=":") # 보기좋게 왼쪽 정렬, 오른쪽정렬(괄호숫자는 공간확보길이)
# # 은행 대기순번표처럼 출력-001, 002, 003... 같이 나타내려면 'zfill' 함수 활용!
# for num in range(1,21):
# print("대기번호 : " + str(num).zfill(3))
# # 표준 입력
# answer = input("아무 값이나 입력하세요 : ")
# print(type(answer)) # <calss 'str'> 이라고 나옴! 즉, input(사용자입력)을 통해서 값을 받게되면 항상 문자열 형태로 저장됨!!꼭 기억!!
# #**8-2 다양한 출력 포맷
# # 빈 자리는 빈공간으로 두고, 오른쪽 정렬을 하되, 총 10자리 공간을 확보
# print("{0: >10}".format(500)) # 500 나옴
# # 양수일 땐 +로 표시, 음수일 땐 -로 표시
# print("{0: >+10}".format(500)) # +500 나옴
# # 왼쪽 정렬하고, 빈칸으로 _로 채움
# print("{0:_<10}".format(500))
# # 3자리 마다 콤마를 찍어주기
# print("{0:,}".format(100000000000))
# # 3자리 마다 콤마를 찍어주기, +-부호도 붙이기
# print("{0:+,}".format(100000000000))
# # 3자리 마다 콤마를 찍어주기, +-부호도 붙이기, 30자리수, 빈자리는 ^, 왼쪽정렬
# print("{0:^<+30,}".format(100000000000))
# # 소수점 출력
# print("{0:f}".format(5/3))
# # 소수점 특정 자리수 까지만 표시_소수점 3째 자리에서 반올림하는 예시
# print("{0:.2f}".format(5/3))
# #**8-3 파일 입출력(파이썬을 통해 파일을 불러올 수 있고, 안에 있는 내용을 쓸 수도 있음)
# # 파일 열어서 쓰기
# score_file = open("score.txt", "w", encoding="utf8") # open(파일명, 용도(w-쓰기), encoding 써주는게 좋음)
# print("수학 : 0", file=score_file)
# print("영어 : 50", file=score_file)
# score_file.close() # 파일은 항상 열어주면 닫아줘야 함!
# score_file = open("score.txt", "a", encoding="utf8") # "W"로 하면 덮어쓰기가 되버림! 그래서 "a"(append의미)를 써서 내용 더해주기
# score_file.write("과학 : 80")
# score_file.write("\n코딩 : 100") # print는 저절로 줄바꿈이 되지만, 파일에서 write 함수는 줄바꿈이 안되기 때문에 '\n'을 사용!
# score_file.close()
# # 파일 읽어오기_전체
# score_file = open("score.txt", "r", encoding="utf8")
# print(score_file.read()) # print를 써야 읽는 내용이 출력되어 볼 수 있음
# score_file.close()
# # 파일 읽어오기_한 줄씩 읽기, 한 줄 읽고 커서는 다음 줄로 이동->한줄씩 띄어져서 출력되는데 바로 밑으로 오게 하고 싶으면 print 구문에서 ','하고 'end=""' 쓰면 됨)
# score_file = open("score.txt", "r", encoding="utf8")
# print(score_file.readline())
# print(score_file.readline())
# print(score_file.readline())
# print(score_file.readline())
# score_file.close()
# # 파일이 몇 줄인지 모를 경우(다른사람의 파일인 경우 몇줄인지 모르기 때문)
# score_file = open("score.txt", "r", encoding="utf8")
# while True:
# line = score_file.readline()
# if not line:
# break
# print(line, end="") #한줄 바로 뒤에 오려고 'end=""'입력
# score_file.close()
# # list 형태로 저장
# score_file = open("score.txt", "r", encoding="utf8")
# lines = score_file.readlines() # 'readlines' 모든 라인을 갖고와서 list 형태로 저장
# for line in lines:
# print(line, end="")
# score_file.close()
# #**8-4 pickle (프로그램상에서 사용하고 있는 데이터를 파일형태로 저장을 해주는 유용한 라이브러리***)
# #피클_쓰기(우리가 가지고 있는 데이터를 피클을 이용하여 파일에 저장)
# import pickle
# profile_file = open("profile.pickle", "wb") #pickle은 encoding 할 필요 없음, "wb" w는 쓰기, b는 바이널을 의미 피클을 쓰기 위해서는 항상 바이널타입을 정의를 해줘야 함!
# profile = {"이름":"박명수", "나이":30, "취미": ["축구", "골프", "코딩"]}
# print(profile) # 생략해도 무관
# pickle.dump(profile, profile_file) # profile에 있는 정보를 profile_file 에 저장
# profile_file.close()
# #피클_불러와서 데이터 읽기(파일에 있는 내용을 load를 통해 불러와서 변수에 저장을 해서 계속 쓸 수 있도록 하는)
# profile_file = open("profile.pickle", "rb")
# profile = pickle.load(profile_file) # load함수로 file에 있는 정보를 profile에 불러오기
# print(profile)
# profile_file.close()
# #**8-5 with (이전보다 좀더 수월하게 파일을 읽고 쓸 수 있음-매번 close할 필요 없음)
# import pickle
# with open("profile.pickle", "rb") as profile_file: # 파일을 열어서 profile_file 변수로 저장을 하고,
# print(pickle.load(profile_file)) # (변수에 저장된) 파일의 내용을 load를 통해서 불러와서 출력을 해주는 것
# # pickle 사용하지 않고 일반적인 파일을 쓰는 경우
# with open("study.txt", "w", encoding="utf8") as study_file:
# study_file.write("파이썬을 열심히 공부하고 있어요")
# # pickle 사용하지 않고 일반적인 파일을 읽는 경우
# with open("study.txt", "r", encoding="utf8") as study_file:
# print(study_file.read())
# # 퀴즈7
# #--내가 푼 것--
# for week in range(1,51):
# with open("{0}주차.txt".format(week), "w", encoding="utf8") as report_file:
# report_file.write("-{0} 주차 주간보고-\n부서 : \n이름 : \n업무 요약 : ".format(week))
# #--나도코딩--
# for i in range(1,51):
# with open(str(i)+"주차.txt", "w", encoding="utf8") as report_file:
# report_file.write("-{0} 주차 주간보고-".format(i))
# report_file.write("\n부서 :")
# report_file.write("\n이름 :")
# report_file.write("\n업무 요약 :")<file_sep>/python기본_1-5.py
# print("안녕")
# hobby= "산책"
# print(3==3)
# cabinet = {"A-3" : "유재석", "b-3":"김태호"}
# print(cabinet)
# print(cabinet.keys())
# print(cabinet.values())
#숫자처리함수
print(abs(-5))
print(round(3.67))
print(max(5,12,85))
from math import * #math 라는 라이브러리에서 불러온다는 의미
print(floor(3.65)) #내림
print(ceil(3.14)) #올림
print(sqrt(16)) #제곱근
#랜덤함수
from random import *
print(random()) #0.0 ~ 1.0 미만의 임의의 값 출력
print(random()*10) #0.0 ~ 10.0미만의 임의의 값 출력
print(int(random()*10)) #0 ~ 10 미만의 임임의 값(정수) 출력
print(randrange(0,10)) #0 ~ 10 미만의 임의의 값(정수) 출력
#로또 관련 코드 써보기 (1부터 45 이하의 임의의 값(정수) 출력)
print(int(random()*45)+1)
print(randrange(1,46))
print(randint(1,45)) # 양쪽 값 모두 포함
#퀴즈_2
date = randint(4,28)
print("오프라인 스터디 모임 날짜는 매월"+str(date)+"일로 선정되었습니다.")
sentence= """나는 28세 여성입니다.
안녕하세요 반갑습니다."""
print(sentence)
python = "Python is Amazing"
print(python.find("java")) # find 에서는 없는 값 찾을 때 '-1' 출력
#print(python.index("java")) # index에서는 없는 값 찾을 때 오류남
print("Reddd Apple\rPine")
print("Red\tApple")
#퀴즈_3
url = "http://naver.com"
my_str = url.replace("http://", "")
my_str = my_str[:my_str.index(".")]
password = my_str[:3]+str(len(my_str))+str(my_str.count("e"))+"!"
print("{0}의 비밀번호는 {1} 입니다." .format(url, password))
#퀴즈_4
from random import *
id = [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20]
shuffle(id)
chicken = sample(id, 1)
id_b = set(id)-set(chicken)
id_c = list(id_b)
shuffle(id_c)
coffee = sample(id_c, 3)
print("""---당첨자 발표---
치킨 당첨자 : {0}
커피 당첨자 : {1}
--축하합니다--""" .format(chicken, coffee))
#퀴즈_4 정답(나도코딩)
users = range(1,21) #1부터 20까지 숫자 생성
#print(type(users)) #-> users 타입이 range로 나옴
users = list(users)
shuffle(users)
winners = sample(users, 4)
print("""---당첨자 발표---
치킨 당첨자 : {0}
커피 당첨자 : {1}
--축하합니다--""" .format(winners[0], winners[1:]))<file_sep>/python기본_6-7.py
# # if문 (굉장히 많이 쓰임)
# weather = input("오늘 날씨는 어때요?")
# if weather == "비" or weather == "눈":
# print("우산을 챙기세요")
# elif weather == "미세먼지":
# print("마스크를 챙기세요")
# else :
# print("준비물 필요 없어요치")
# temp = int(input("오늘 기온은 어때요?"))
# if 30 <= temp:
# print("너무 더워요. 나가지 마세요")
# elif 10 <= temp and temp <30:
# print("괜찮은 날씨에요")
# elif 0 <= temp < 10:
# print("쌀쌀하니 외투를 챙기세요")
# else:
# print("너무 추워요. 나가지 마세요.")
# # for(반복문)
# for waiting_no in [0, 1, 2, 3, 4]:
# print("대기번호 : {0}" .format(waiting_no))
# for waiting_no in range(5): #또는 range(0,5) 해도 똑같은 값(0~4까지_5직전) 나옴
# print("대기번호 : {0}" .format(waiting_no))
# starbucks = ["아이언맨", "토르", "아이엠그루트"]
# for customer in starbucks:
# print ("{0}, 커피가 준비되었습니다.".format(customer))
# # while(반복문) while 뒤에 오는 조건이 성립될 때까지 반복함
# customer = "토르"
# index = 5
# while index >=1:
# print("{0}, 커피가 준비되었습니다. {1}번 남았어요.".format(customer, index))
# index -= 1
# if index == 0:
# print("폐기처분되었습니다.")
# # 무한루프에 빠지는 경우 -> ctrl+C 누르면 강제 종료됨 !
# # customer2 ="아이언맨"
# # index = 1
# # while True:
# # print("{0}, 커피가 준비되었습니다. 호출: {1}회".format(customer2, index))
# # index += 1
##퀴즈 5
# from random import *
# cnt = 0 # 총 탑승승객 수
# for customer in range(1,51): # 1 ~ 50 이라는 수 (승객)
# time = randrange(5,51) # 5분 ~ 50분 소요시간
# if 5<= time <=15: # 5분~ 15분 이내의 손님(매칭성공), 탑승 승객 수 증가 처리
# print("[O] {0}번째 손님 (소요시간 : {1}분".format(customer, time))
# cnt += 1
# else: # 매칭 실패한 경우
# print("[ ] {0}번째 손님 (소요시간 : {1}분".format(customer, time))
# print("총 탑승 승객: {0} 분" .format(cnt))
# #7. 함수 (7-2 전달값, 반환값 )
# def deposit(balance, money): # 입금
# print("입금이 완료되었습니다. 잔액은 {0}원 입니다.".format(balance+money))
# return balance+money
# def withdraw(balance, money): #출금
# if balance >= money: #잔액이 출금보다 많으면
# print("출금이 완료되었습니다. 잔액은 {0}원 입니다.".format(balance-money))
# return balance-money
# else:
# print("출금이 완료되지 않았습니다. 잔액은 {0}원 입니다.".format(balance))
# return balance
# def withdraw_night(balance, money): #저녁에 출금
# commission = 100
# return commission, balance-money-commission
# balance = 0
# balance = deposit(balance, 2000)
# balance = withdraw(balance, 500)
# commission, balance = withdraw_night(balance, 500)
# print("수수료는 {0}원 이고, 잔액은 {1}원 입니다.".format(commission, balance))
# #7-3 기본값
# def profile(name, age, main_lang) :
# print("이름: {0}\t나이: {1}\t주 사용 언어 : {2}" \
# .format(name, age, main_lang)) # 코드가 길면 \+엔터누르면 줄바꿈이 됨(같은 줄이라는 의미는 있으면서)
# profile("유재석", 20, "파이썬")
# profile("김태호",25, "자바")
# #만약 유재석, 김태호가 같은나이이고, 같은 언어이면 계속 반복적으로 적을 필요 없음
# #그러므로, 기본값을 넣어줌
# def profile(name, age=17, main_lang="파이썬"):
# print("이름: {0}\t나이: {1}\t주 사용 언어 : {2}" \
# .format(name, age, main_lang))
# profile("유재석")
# profile("김태호")
# #7-4 키워드값으로 함수 호출 가능
# def profile(name, age, main_lang) :
# print(name, age, main_lang)
# profile(name="유재석", main_lang="파이썬", age=20)
# profile(main_lang="자바", age=25, name="김태호")
# #7-5 가변인자
# def profile(name, age, *language): # *language 같은 변수가 가변인자(서로 다른 갯수의 값을 넣어줄 때 유용)
# print("이름 : {0}\t나이 : {1}\t".format(name, age), end=" ")
# for lang in language:
# print(lang, end=" ")
# print()
# profile("유재석", 20, "Python", "Java", "C", "C++", "C#", "JavaScript")
# profile("김태호", 25, "Kotlin", "Swift")
# #7-6 지역변수와 전역변수
# #지역변수: 함수 내에서 쓸 수 있는 것(함수 호출될 때 만들어졌다가 호출이 끝나면 사라지는 것)
# #전역변수: 모든 공간에서 프로그램 내에서 어디서든지 부를 수 있는 함수
# #일반적으로 전역변수 많이 쓰면 코드관리도 어려워지기 때문에 권장되는 방법은 아님
# # 지역변수
# gun = 10
# def checkpoint(soldiers): # 경계근무나가는 군인
# gun = 20
# gun = gun-soldiers
# print("[함수 내] 남은 총 : {0}".format(gun))
# print("전체 총 : {0}".format(gun)) # 10이 나옴
# checkpoint(2) # 18이 나옴
# print("남은 총 : {0}".format(gun)) # 10이 나옴
# # '전역변수'를 이용하면 다르게 나옴-'global' 이용!
# gun = 10
# def checkpoint(soldiers): # 경계근무나가는 군인
# global gun # 전역공간에 있는 gun 사용 (즉, 함수 밖에있는 'gun=10'이라는 변수를 'checkpoint'함수 내에서 쓰겠다는 의미!)
# gun = gun-soldiers
# print("[함수 내] 남은 총 : {0}".format(gun))
# print("전체 총 : {0}".format(gun)) # 10이 나옴
# checkpoint(2) # 8이 나옴
# print("남은 총 : {0}".format(gun)) # 8이 나옴
# 그러나 일반적으로 전역변수를 많이 쓰면 코드가 어려워져서.. 가급적이면 함수의 전달값으로 던져서 계산하고 반환값을 받아서 사용함!
# #(일반적임)전달값으로 던져서 계산하고 반환값으로 받아서 사용하는 방법
# gun = 10
# def checkpoint_ret(gun, soldiers):
# gun = gun-soldiers
# print("[함수 내] 남은 총 : {0}".format(gun))
# return gun
# print("전체 총 : {0}".format(gun)) # 10이 나옴
# gun = checkpoint_ret(gun,2) # 8이 나옴
# print("남은 총 : {0}".format(gun)) # 8이 나옴
# #퀴즈 6
# #--내가 푼 것 --
# def std_weight(height, gender) :
# if gender == "male":
# print ("키 {0}cm 남자의 표준 체중은 {1} kg 입니다.".format(height*100, round(height*height*22, 2)))
# else:
# print("키 {0}cm 여자의 표준 체중은 {1} kg 입니다.".format(height*100, round(height*height*21, 2)))
# std_weight(1.75, "male")
# #--나도코딩 정답--
# def std_weight(height, gender): # 키 m 단위(실수), 성별 "남자"/"여자"
# if gender == "남자":
# return height*height*22
# else:
# return height*height*21
# height = 175 # cm 단위
# gender = "남자"
# weight = round(std_weight(height/100, gender), 2)
# print("키 {0}cm {1}의 표준 체중은 {2}kg 입니다.".format(height, gender, weight))
<file_sep>/python기본_10.py
# #**10-1 예외처리 (어떤 에러가 발생했을 때 그에 대해서 처리를 해주는 것)
# try:
# # 나누기전용 계산기 프로그램
# print("나누기 전용 계산기입니다.")
# nums =[]
# nums.append(int(input("첫 번째 숫자를 입력하세요 : ")))
# nums.append(int(input("두 번째 숫자를 입력하세요 : ")))
# nums.append(int(nums[0]/nums[1]))
# print("{0} / {1} = {2}".format(nums[0], nums[1], nums[2]))
# except ValueError: #숫자가 아닌 '삼' 이런식으로 넣을 경우
# print("에러! 잘못된 값을 입력하였습니다.")
# except ZeroDivisionError as err: # 나누는 값을 0을 넣을 경우(나누기에서 0으로 나눌 수 없음)
# print(err)
# except Exception as err: # 위에 2개 에러 말고 다른 에러들 발생할 경우 ('err'은 무슨 에러인지 알려주는 것)
# print("알 수 없는 에러가 발생하였습니다.")
# print(err)
# #**10-2 에러 발생시키기(의도적으로 에러를 발생시키는 것)
# #한 자리 숫자 나누기 전용 계산기
# try:
# print("한 자리 숫자 나누기 전용 계산기입니다.")
# num1 = int(input("첫 번째 숫자를 입력하세요 : "))
# num2 = int(input("두 번째 숫자를 입력하세요 : "))
# if num1 >= 10 or num2 >= 10:
# raise ValueError # 에러를 내가 원하는 조건에 해당하는 경우, 'raise'를 사용하여 의도적으로 만들 수 있음
# print("{0} / {1} = {2}".format(num1, num2, int(num1/ num2)))
# except ValueError:
# print("잘못된 값을 입력하였습니다. 한 자리 숫자만 입력하세요.")
# #**10-3 사용자 정의 예외처리(파이썬에서 정의하는 에러(ValueError, ZeroDivisionError 등)가 아닌 사용자가 직접 만든 에러를 처리하는 것)
# class BigNumberError(Exception):
# def __init__(self, msg):
# self.msg = msg
# def __str__(self):
# return self.msg
# try:
# print("한 자리 숫자 나누기 전용 계산기입니다.")
# num1 = int(input("첫 번째 숫자를 입력하세요 : "))
# num2 = int(input("두 번째 숫자를 입력하세요 : "))
# if num1 >= 10 or num2 >= 10:
# raise BigNumberError("입력값 : {0}, {1}".format(num1, num2)) # 사용자가 클래스를 사용하여 만든 에러
# print("{0} / {1} = {2}".format(num1, num2, int(num1/ num2)))
# except ValueError:
# print("잘못된 값을 입력하였습니다. 한 자리 숫자만 입력하세요.")
# except BigNumberError as err:
# print("에러가 발생하였습니다. 한 자리 숫자만 입력하세요.")
# print(err)
# finally:
# print("계산기를 이용해주셔서 감사합니다.")
# #**10-4 finally(예외처리 구문에서 정상적으로 수행이 되건 오류가 발생하건 상관없이 무조건 실행되는 구문)
# #항상 try 구문 내에서 맨 마지막에 작성
# #**퀴즈 9
# #--내가 푼 것
# class SoldOutError(Exception):
# def __init__(self, msg):
# self.msg = msg
# def __str__(self):
# return self.msg
# chicken = 10
# waiting = 1 # 홀 안에는 현재 만석, 대기번호 1부터 시작
# while(True):
# try:
# print("[남은 치킨 : {0}]".format(chicken))
# order = int(input("치킨 몇 마리 주문하시겠습니까?"))
# if order > chicken: # 남은 치킨보다 주문량이 많을 때
# print("재료가 부족합니다.")
# elif order <= 0:
# raise ValueError
# else:
# print("[대기번호 {0}] {1} 마리 주문이 완료되었습니다.".format(waiting, order))
# waiting += 1
# chicken -= order
# if chicken <= 0 :
# raise SoldOutError("재고가 소진되어 더 이상 주문을 받지 않습니다.")
# except ValueError:
# print("잘못된 값을 입력하였습니다.")
# except SoldOutError as err:
# print(err)
# break # while문 탈출(프로그램 종료)
# # --나도코딩--
# class SoldOutError(Exception):
# pass
# chicken = 10
# waiting = 1 # 홀 안에는 현재 만석, 대기번호 1부터 시작
# while(True):
# try:
# print("[남은 치킨 : {0}]".format(chicken))
# order = int(input("치킨 몇 마리 주문하시겠습니까?"))
# if order > chicken: # 남은 치킨보다 주문량이 많을 때
# print("재료가 부족합니다.")
# elif order <= 0:
# raise ValueError
# else:
# print("[대기번호 {0}] {1} 마리 주문이 완료되었습니다.".format(waiting, order))
# waiting += 1
# chicken -= order
# if chicken <= 0 :
# raise SoldOutError
# except ValueError:
# print("잘못된 값을 입력하였습니다.")
# except SoldOutError as err:
# print("재고가 소진되어 더 이상 주문을 받지 않습니다.")
# break # while문 탈출(프로그램 종료)<file_sep>/byme.py
# --내가 한 것---
# def sign(name, email):
# print("이 프로그램은 {0}에 의해 만들어졌습니다. \n유튜브 : http://youtube.com \n이메일 : {1}".format(name, email))
# --나도코딩 정답--
def sign():
print("이 프로그램은 나도코딩에 의해 만들어졌습니다.")
print("유튜브 : http://youtube.com")
print("이메일 : <EMAIL>")<file_sep>/python기본_12퀴즈.py
# #기본편 다 들은 후 푸는 퀴즈 강의
# #퀴즈1
# #--내가 푼것--
# names = ["유튜버1", "유튜버2", "유튜버3", "유튜버4"]
# for name in names:
# with open(str(name)+".txt", "w", encoding="utf-8") as f:
# f.write("안녕하세요? {0}님.\n\n(주)나도출판 편집자 나코입니다.\
# \n현재 저희 출판사는 파이썬에 관한 주제로 책 출간을 기획 중입니다.\
# \n{0}님의 유튜브 영상을 보고 연락을 드리게 되었습니다.\
# \n자세한 내용은 첨부드리는 제안서를 확인 부탁드리며, 긍정적인 회신 기다리겠습니다.\
# \n\n좋은하루 보내세요^^\n감사합니다.\
# \n\n- 나코드림.".format(name))
# #--나도코딩--
# names = ["아이어맨TV", "<NAME>", "토르코딩", "헐크에러"]
# for name in names:
# with open("{}.txt".format(name), "w", encoding="utf-8") as email_file:
# email_file.write(f"""
# 안녕하세요? {name}님.
# (주)나도출판 편집자 나코입니다.
# 현재 저희 출판사는 파이썬에 관한 주제로 책 출간을 기획 중입니다.
# {name}님의 유튜브 영상을 보고 연락을 드리게 되었습니다.
# 자세한 내용은 첨부드리는 제안서를 확인 부탁드리며, 긍정적인 회신 기다리겠습니다.
# 좋은하루 보내세요^^
# 감사합니다.
# - 나코드림.
# """)
# # 위에 wirte 부분에서 들여쓰기를 못해서 코드가 예쁘게 안보임("""사용할 때 들여쓰기하면 만드는 파일에도 그대로 적용되서). contents 를 사용하기!
# names = ["아이어맨TV", "<NAME>", "토르코딩", "헐크에러"]
# for name in names:
# with open("{}.txt".format(name), "w", encoding="utf-8") as email_file:
# contents = (f"안녕하세요? {name}님\n\n"
# "(주)나도출판 편집자 나코입니다.\n"
# "현재 저희 출판사는 파이썬에 관한 주제로 책 출간을 기획 중입니다.\n"
# f"{name}님의 유튜브 영상을 보고 연락을 드리게 되었습니다.\n"
# "자세한 내용은 첨부드리는 제안서를 확인 부탁드리며, 긍정적인 회신 기다리겠습니다.\n\n"
# "좋은하루 보내세요^^\n"
# "감사합니다.\n\n"
# "- 나코드림." )
# email_file.write(contents)
#퀴즈2<file_sep>/python기본_9_1-9.py
# #**9-1 클래스(어렵지만 python에서 굉장히 중요한 부분!)
# # 비유를 하자면, '붕어빵 틀'이라고 생각하면 됨(그 틀에 재료를 넣으면 틀은 1개인데 붕어빵은 무한대로 만들 수 있음)
# # 일반적인 설명으로는, 서로 연관이 있는 변수와 함수의 집합정도로 이해하면 될 것 같음
# class Unit:
# def __init__(self, name, hp, damage):
# self.name = name
# self.hp = hp
# self.damage = damage
# print("{0} 유닛이 생성 되었습니다.".format(self.name))
# print("체력 {0}, 공격력 {1}".format(self.hp, self.damage))
# marine1 = Unit("마린", 40, 5)
# marine2 = Unit("마린", 50, 5)
# tank = Unit("탱크", 150, 35)
# #**9-2 __init__(파이썬에서 쓰이는 생성자 즉, 마린이나 탱크같은 객체가 만들어질 때 자동으로 호출되는 부분)
# # 마린과 탱크는 Unit 클래스의 인스턴스라고 표현함
# # 객체(마린, 탱크같은)가 생성될 때에는 기본적으로 __init__함수에 정의된 갯수와 동일하게 해야 함(self 제외하고)
# # 예를 들어, tank2 = Unit("탱크2", 120) 이렇게만 넣을 경우 오류 발생
# #**9-3 멤버변수
# # 클래스 내에서 정의된 변수, 그 변수를 가지고 외부에서 실제로 쓸 수 있는 것 ex)self.name, self.hp 이런 변수
# wraith1 = Unit("레이스", 80, 5)
# print("유닛 이름 : {0}, 공격력 : {1}".format(wraith1.name, wraith1.damage)) # 멤버변수를 외부에서 wraith1.name 이렇게 쓸 수 있음
# # 마인드 컨트롤 : 상대방 유닛을 내 것으로 만드는 것(빼앗음)_스타크래프트 설명
# wraith2 = Unit("빼<NAME>", 80, 5)
# wraith2.clocking = True # 클래스 외부에서 clocking이라는 변수를 추가로 할당한 것(파이썬은 어떤 객체에 추가로 변수를 외부에서 만들어서 쓸 수 있음)
# if wraith2.clocking == True: # 외부에서 추가 할당한 변수는 그 할당을 한 객체에서만 적용됨!(ex-wraith1에서는 clocking을 할당하지 않은 채로 쓰면 오류남)
# print("{0} 는 현재 클로킹 상태입니다. ".format(wraith2.name))
# #**9-4 메소드 (클래스에서 멤버변수 이외의 함수들)
# class AttackUnit:
# def __init__(self, name, hp, damage):
# self.name = name
# self.hp = hp
# self.damage = damage
# def attack(self, location):
# print("{0} : {1} 방향으로 적군을 공격합니다. [공격력 {2}"\
# .format(self.name, location, self.damage)) #self.name, self.damage같은 'self.~~' 변수는 클래스 자기자신에 있는 멤버변수의 값을 출력, location은 그냥 전닯받은 값을 출력
# def damaged(self, damage):
# print("{0} : {1} 데미지를 입었습니다.".format(self.name, damage))
# self.hp -= damage
# print("{0} : 현재 체력은 {1} 입니다.".format(self.name, self.hp))
# if self.hp <= 0:
# print("{0} : 파괴되었습니다.".format(self.name))
# firebat1 = AttackUnit("파이어뱃", 50, 16) # 파이어뱃 : 공격유닛, 화염방사기_스타크래프트 설명
# firebat1.attack("5시")
# # 공격 2번 받는다고 가정
# firebat1.damaged(25)
# firebat1.damaged(25)
# #**9-5 상속(상속받는 유닛의 멤버변수와 메소드를 그대로 사용할 수 있게 됨)
# # 일반유닛
# # class Unit:
# # def __init__(self, name, hp):
# # self.name = name
# # self.hp = hp
# # 공격유닛
# # class AttackUnit(Unit): # 상속받고 싶은 클래스를 ()괄호 안에 넣어줌 (AttackUnit클래스가 Unit클래스를 상속받음_AttackUnit클래스:자식/Unit크랠스:부모)
# # def __init__(self, name, hp, damage):
# # Unit.__init__(self, name, hp) # 상속받는 클래스의 멤버변수 가져올 때 (Unit에서 만들어진 생성자를 호출)
# # self.damage = damage
# #**9-6 다중상속
# # 날 수 있는 기능을 가진 클래스
# class Flyable:
# def __init__(self, flying_speed):
# self.flying_speed = flying_speed
# def fly(self, name, location):
# print("{0} : {1} 방향으로 날아갑니다. [속도 {2}]".format(name, location, self.flying_speed))
# # 공중공격유닛_다중상속 예시
# class FlyableAttackUnit(AttackUnit, Flyable): # 이 클래스는 두 개의 클래스를 상속받아 초기화해준 것뿐임(두 클래스의 멤버변수, 메소드 다 받음)
# def __init__(self, name, hp, damage, flying_speed):
# AttackUnit.__init__(self, name, hp, damage)
# Flyable.__init__(self, flying_speed)
# valkyrie = FlyableAttackUnit("발키리", 200, 6, 5) # 발키리 : 공중공격유닛, 한번에 14발 미사일 발사_스타크래프트 설명
# valkyrie.fly(valkyrie.name, "3시")
# #**9-7 메소드 오버라이딩
# # -> 부모클래스에서 정의한 메소드말고 자식클래스에서 정의한 메소드를 쓰고 싶을 때 메소드를 새롭게 정의해서 사용하는 것
# class Unit:
# def __init__(self, name, hp, speed):
# self.name = name
# self.hp = hp
# self.speed = speed
# def move(self, location):
# print("[지상 유닛 이동]")
# print("{0} : {1} 방향으로 이동합니다. [속도 {2}]".format(self.name, location, self.speed))
# class AttackUnit(Unit):
# def __init__(self, name, hp, speed, damage):
# Unit.__init__(self, name, hp, speed)
# self.damage = damage
# def attack(self, location):
# print("{0} : {1} 방향으로 적군을 공격합니다. [공격력 {2}]".format(self.name, location, self.damage))
# def damaged(self, damage):
# print("{0} : {1} 데미지를 입었습니다.".format(self.name, damage))
# self.hp -= damage
# print("{0} : 현재 체력은 {1} 입니다.".format(self.name, self.hp))
# if self.hp <= 0:
# print("{0} : 파괴되었습니다.".format(self.name))
# class Flyable:
# def __init__(self, flying_speed):
# self.flying_speed = flying_speed
# def fly(self, name, location):
# print("{0} : {1} 방향으로 날아갑니다. [속도 {2}]".format(name, location, self.flying_speed))
# class FlyableAttackUnit(AttackUnit, Flyable):
# def __init__(self, name, hp, damage, flying_speed):
# AttackUnit.__init__(self, name, hp, 0, damage) # 지상 speed 0으로 처리
# Flyable.__init__(self, flying_speed)
# def move(self, location): # move 재정의 (메소드 오버라이딩)_상속받고 있는 AttackUnit의 move함수(Unit에게 상속받은)를 쓰지 않고 다시 재정의해서 씀
# print("[공중 유닛 이동]")
# self.fly(self.name, location)
# vulture = AttackUnit("벌쳐", 80, 10, 20) # 벌쳐 : 지상유닛, 기동성이 좋음_스타크래프트 설명
# battlecruiser = FlyableAttackUnit("배틀크루저", 500, 25, 3) # 배틀크루저 : 공중유닛, 체력과 공격력 굉장히 좋음_스타크래프트 설명
# # 아래와 같이, 벌쳐와 배틀크루저가 지상유닛인지 공격유닛인지 항상 확인해가면서 함수(move/fly)를 구별해서 써줘야 함;; 되게 귀찮음..
# vulture.move("11시")
# # battlecruiser.fly(battlecruiser.name,"9시")
# # 그러므로, '메소드 오버라이딩'을 써서 똑같이 move함수만 쓰면 지상유닛인 경우에는 이동을 하고, 공중유닛인 경우에는 날아갈 수 있도록 처리하겠음
# battlecruiser.move("9시")
# #**9-8 pass (아무것도 안하고 그냥 넘어가는 의미, 정의O, 오류X)
# class BuildingUnit(Unit):
# def __init__(self, name, hp, locaiton):
# pass
# supply_depot = BuildingUnit("서플라이디폿", 500, "7시") # 서플라이 디폿 : 건물, 1개 건물 = 8 유닛 _스타크래프트 설명
# # pass 예제2
# def game_start():
# print("[알림] 새로운 게임을 시작합니다.")
# def game_over():
# pass
# game_start()
# game_over()
# #**9-9 super
# class BuildingUnit(Unit):
# def __init__(self, name, hp, locaiton):
# # Unit.__init__(self, name, hp, 0)
# super().__init__(name, hp, 0) # 상속받는 클래스의 멤버변수 가져올 때(생성자 호출할 때), 이렇게 super로도 사용 가능
# self.location = location
# # 다중상속일 때 super의 문제점
# class Unit:
# def __init__(self):
# print("Unit 생성자")
# class Flyable:
# def __init__(self):
# print("Flyable 생성자")
# class FlyableUnit(Flyable, Unit):
# def __init__(self):
# super().__init__() # 다중상속 받을 경우, super는 처음 오는 클래스인 'Flyable'클래스의 __init__함수가 호출됨
# dropship = FlyableUnit() # 'Flyable 생성자'만 출력됨 | 5ba9c291ae17342b9d86b80d77613fd2b16a1e4e | [
"Python"
] | 8 | Python | Frankie937/Python_basic | d2fb5cc5ae254b10425874f6191204732644627b | 60aa070de66bb9cc74867d0a7304eb9c5e94e861 |
refs/heads/master | <repo_name>mohamed0328/PieShop-Asp.net-Core-2.2<file_sep>/BethanysPieShop/Iconfigurstion.cs
namespace BethanysPieShop
{
public class IConfigurstion
{
}
} | 420d1e423edac3c0926aaf04144d96487db8644c | [
"C#"
] | 1 | C# | mohamed0328/PieShop-Asp.net-Core-2.2 | fbed67400b056f31551875b23a38164b74608055 | f620c8acfe043907aeb901527a6b885444ae3e7c |
refs/heads/master | <repo_name>snehasingi/ReactAssignments<file_sep>/src/Components/TSkills.js
import React,{Component} from 'react'
class TSkills extends Component{
state = {
emplist : this.props.employeeDetails
}
componentDidMount = () => {
this.setState({emplist : this.props.employeeDetails});
}
render(){
console.log(this.props.employeeDetails);
this.state.emplist = this.props.employeeDetails.map((row) => {
return(
<tr key={row.id}>
<td>{row.name}</td>
<td>{row.skill}</td>
{/* <td><button onClick={this.props.deleteItem(row.id)}>Delete</button></td> */}
</tr>
)
});
return <tbody>{this.state.emplist}</tbody>;
}
}
export default TSkills;<file_sep>/src/Components/Assignment1.js
import React,{Component} from 'react'
import ReactDOM from 'react-dom'
import EmployeeSkills from './EmployeeSkills'
import { States, CounterFunctionalComponent } from './States';
import StaticClock from './JavaScriptFunctions/StaticClock'
import DynamicClock from './JavaScriptFunctions/DynamicClock'
import Timer from './JavaScriptFunctions/Timer'
import {ShowAddition} from './ShowAddition'
class Assignment1 extends React.Component{
render(){
return(
<React.Fragment>
<h1 align="center"> Assignment - 1</h1>
<table align="center">
<tbody>
<tr>
<td><h3>1. Extend the Dynamic Clock example using a start and stop button which helps us control the clock </h3></td><td align="right"><DynamicClock date={new Date()}/></td>
</tr><br></br>
<tr>
<td><h3>2. Create 2 text boxes and set initial values. As soon as there is a change in the text values , calculate the sum of the values</h3></td><td align="right"><ShowAddition/></td>
</tr><br></br><br></br>
<tr>
<td><h3>3. Add a new Skill and Employee in the project to include new skills and Employees dynamically through form elements </h3></td><td align="right"><EmployeeSkills/></td>
</tr>
</tbody>
</table>
</React.Fragment>
)
}
}
export default Assignment1;<file_sep>/src/Components/JavaScriptFunctions/StaticClock.js
import React from 'react';
class StaticClock extends React.Component{
render(){
return(
<React.Fragment>
<h3> Static Clock : {this.props.date.toLocaleTimeString()}</h3>
</React.Fragment>
)
}
}
export default StaticClock;<file_sep>/src/Components/EmployeeSkills.js
import React, {Component} from 'react'
import THeader from './THeader'
import TSkills from './TSkills'
import './Css/header.css'
var employee = [
{
id : 1,
name: "Sneha",
skill: "ReactJs"
},
{
id : 2,
name: "Ram",
skill: "Selenium"
}
]
class EmployeeSkills extends Component{
state = {
employeeName : "",
skillName : "",
id : 2,
}
handleChange = (event) =>{
switch(event.target.name){
case "employeeName" : this.setState({employeeName : event.target.value});break;
case "skill" : this.setState({skillName : event.target.value});break;
}
}
addSkill = () =>{
//employee.push({name: this.state.employeeName, skill: this.state.skillName});
this.setState({id : (this.state.id++)})
//console.log(this.state.id);
alert(this.state.id);
console.log()
employee = [...employee,{id: this.state.id, name: this.state.employeeName, skill: this.state.skillName}];
console.log(employee);
}
deleteItem = (index) =>{
alert("deleted item at" + index);
//this.state.emplist.actionItems.splice(index,1);
}
render(){
return(
<React.Fragment>
<input
type={Text}
value={this.state.employeeName}
name="employeeName"
onChange = {this.handleChange}>
</input>
<input
type={Text}
value={this.state.skillName}
name="skill"
onChange = {this.handleChange}>
</input>
<button onClick={this.addSkill}> Add Skill </button>
<table border="1">
<THeader/>
<TSkills employeeDetails = {employee} />
</table>
</React.Fragment>
)
}
}
export default EmployeeSkills;<file_sep>/src/Components/THeader.js
import React,{Component} from 'react'
import ReactDOM from 'react-dom'
class THeader extends React.Component{
render(){
return(
<React.Fragment>
<th><td>Employee Name</td></th>
<th><td>Skills</td></th>
</React.Fragment>
)
}
}
export default THeader;<file_sep>/src/Components/JavaScriptFunctions/DynamicClock.js
import React,{Component} from 'react'
class DynamicClock extends React.Component {
constructor(props) {
super(props);
this.state = {date: new Date()};
}
startClock = () =>{
this.date = setInterval(() => this.tick(),1000);
}
stopClock = () =>{
clearInterval(this.date);
}
componentWillUnmount() {
clearInterval(this.date);
}
tick() {
this.setState({
date: new Date()
});
}
render() {
return (
<div>
<h3> Dynamic Clock : {this.state.date.toLocaleTimeString()}</h3>
<button name="start" onClick={this.startClock}> Start </button>
<button name="stop" onClick={this.stopClock}> Stop </button>
</div>
);
}
}
export default DynamicClock; | efa7fbd5609e818471e51d0a2b2fb82bd5d5240d | [
"JavaScript"
] | 6 | JavaScript | snehasingi/ReactAssignments | 76d78c5f48447d4ef3ce3e2880edc8545b32afaa | 70cc1e22f40adb516eed92b815457bd54ddd75aa |
refs/heads/master | <file_sep># -*- encoding: utf-8 -*-
module XmppServer
#drb interface
class Pusher
def push(username, content)
m = XmppServer::M.new(username, content)
XmppServer::QUEUE.push m
return nil #from : http://ruby-china.org/topics/5550
end
end
end
<file_sep>require 'spec_helper'
describe "XmppServer.push" do
it "should push message to queue" do
XmppServer::Pusher.new().push("tester1", "hello world")
m = XmppServer::QUEUE.pop
m.username.should == "tester1"
m.content.should == "hello world"
end
it "push data to queue" do
3.times do |i|
m = XmppServer::M.new("tester002", "hello #{i}")
XmppServer::QUEUE.push(m)
sleep 1
p XmppServer::QUEUE
end
end
end
<file_sep># -*- encoding: utf-8 -*-
namespace :xmppserver do
desc "启动xmppserver-drb服务器"
task :start do
puts "start xmpp drb server"
path = File.expand_path("../../../bin",__FILE__)
file_path = File.join(path, "xmppserver")
system("#{file_path}")
end
end
<file_sep># -*- encoding: utf-8 -*-
module XmppServer
class Config
@@server = "weishanke.com"
@@username = "tester001"
@@password = "<PASSWORD>"
class << self
def server; @@server ;end
def username; @@username; end
def password; @@password; end
def server=(server)
@@server = server
end
def username=(username)
@@username = username
end
def password=(<PASSWORD>)
@@password = <PASSWORD>
end
def config
yield self
end
end
end
end
<file_sep>require 'spec_helper'
describe "push messgae to client" do
it "should push message" do
jid = "#{XmppServer::Config.username}@#{XmppServer::Config::server}/pusher"
jid.should == "<EMAIL>/pusher"
password = XmppServer::Config.password
client = XmppServer::Client.new(jid, password)
m = XmppServer::M.new("tester002", "hello,man")
client.connect
client.auth
client.push m
end
it "default should could push message" do
m = XmppServer::M.new("tester002", "hello, world. default push")
client = XmppServer::Client.get_default_client
client.push m
end
end
<file_sep>require 'xmpp_server'
require 'rails'
module XmppServer
class Railtie < Rails::Railtie
rake_tasks do
load File.join(File.expand_path("../../tasks", __FILE__), "xmpp_server.rake")
end
end
end
<file_sep>require 'rubygems'
require 'bundler/setup'
Bundler.require(:default, :development)
RSpec.configure do |config|
config.mock_with :rspec
config.before(:each) do
#pending
end
end
<file_sep>## xmpp_server
给xmpp 服务器push 消息的中转站
###实例
* 如何启动服务
```shell
./bin/xmppserver
```
rake启动
```ruby
rake xmppserver:start
```
* 客户端code
```ruby
require 'drb/drb'
SERVER_URI="druby://localhost:8787"
xmppserver = DRbObject.new_with_uri(SERVER_URI)
3.times do |i|
xmppserver.push("tester002", "testdddeeee....")
sleep 1
end
```
* 修改服务器地址,用户名和密码
```ruby
XmppServer::Config.config do |s|
s.username = "zhangsan"
s.password = "<PASSWORD>"
s.server = "test.com"
end
```
Rails中,可以新建`config/initializers/xmppserver.rb`
* Rails项目中的使用
新建`config/initializers/xmppserver.rb`文件
```ruby
require 'drb/drb'
SERVER_URI="druby://localhost:8787"
XMPPSERVER = DRbObject.new_with_uri(SERVER_URI)
XmppServer::Config.config do |s|
s.username = "zhangsan"
s.password = "<PASSWORD>"
s.server = "test.com"
end
```
<file_sep># -*- encoding: utf-8 -*-
require "bundler/gem_tasks"
desc "Run all tests by default"
task :default do
system("rspec spec --color")
end
namespace :xmppserver do
desc "启动xmppserver-drb服务器"
task :start do
puts "start xmpp drb server"
path = File.expand_path("../bin",__FILE__)
file_path = File.join(path, "xmppserver")
system("#{file_path}")
end
end
<file_sep>#!/usr/bin/env ruby
XMPP_PATH = File.expand_path("../../lib", __FILE__)
$:.unshift(XMPP_PATH)
require "drb/drb"
require "xmpp_server"
DRB_URI = "druby://localhost:8787"
client = XmppServer::Client.get_default_client
consumer = Thread.new {
loop {
m = XmppServer::QUEUE.pop
client.push m
}
}
FRONT_OBJECT = XmppServer::Pusher.new
DRb.start_service(DRB_URI, FRONT_OBJECT)
DRb.thread.join
<file_sep># -*- encoding: utf-8 -*-
module XmppServer
QUEUE = Queue.new
#message node
class M
attr_accessor :username, :content
def initialize(username, content)
@username = username
@content = content
end
end
end
<file_sep>require "xmpp_server/version"
require "xmpp4r/client"
require "xmpp_server/config"
require "xmpp_server/queue"
require "xmpp_server/push"
require "xmpp_server/railtie" if defined?(Rails)
module XmppServer
def self.config
yield self
end
#xmpp client to push connect and push message to xmpp server
class Client
attr_accessor :jid, :password
def initialize(jid, password)
@jid = jid
@password = <PASSWORD>
@client = Jabber::Client.new @jid
end
def connect
@client.connect
end
def auth
@client.auth @password
end
def push(m)
to = "#{m.username}@#{XmppServer::Config.server}/nickname"
msg = Jabber::Message::new(to, m.content).set_type(:normal).set_id(1)
@client.send msg
end
class << self
def get_default_client
jid = "#{XmppServer::Config.username}@#{XmppServer::Config::server}/pusher"
password = XmppServer::Config.password
client = XmppServer::Client.new(jid, password)
client.connect
client.auth
client
end
end
end
end
<file_sep>require "spec_helper"
require 'drb/drb'
describe "xmppserver bin" do
it "should push message to drb server" do
SERVER_URI="druby://localhost:8787"
DRb.start_service
xmppserver = DRbObject.new_with_uri(SERVER_URI)
3.times do |i|
xmppserver.push("tester002", "testdddeeee....")
puts i
sleep 1
end
end
end
<file_sep>require 'drb/drb'
SERVER_URI="druby://localhost:8787"
DRb.start_service
xmppserver = DRbObject.new_with_uri(SERVER_URI)
3.times do |i|
xmppserver.push("tester002", "testdddeeee....")
puts i
sleep 1
end
| 4b8b711690008e83959f2ffc3640d208edd349ad | [
"Markdown",
"Ruby"
] | 14 | Ruby | chucai/xmpp_server | 699d398c24a840b687c63148aefe9b469a8ae20c | bd80ed739d414743f82b84a72210d96e25b756a9 |
refs/heads/master | <repo_name>chrispicato/TickrTaker<file_sep>/app/components/cart.jsx
import React, {Component} from 'react';
export default class Cart extends Component {
constructor(props) {
super(props)
this.state = {
sum: 0
}
}
componentDidMount () {
var context = this;
setTimeout(function () {
context.props.winningBids.forEach(function (item) {
var sum = parseInt(context.state.sum) + parseInt(item.highestBid);
var decimalSum = sum.toFixed(2);
context.setState({
sum: decimalSum
});
});
console.log(context.state.sum);
}, 4000);
// this.setState({
// sum: sum
// });
}
render () {
return (
<div>
<h5>Subtotal: $</h5><h5 id="subtotal">{this.state.sum}</h5>
</div>
);
}
}<file_sep>/app/components/payment.jsx
import React, {Component} from 'react';
import Cart from './cart.jsx';
export default class Payment extends Component {
constructor(props) {
super(props);
}
componentDidMount () {
Stripe.setPublishableKey('<KEY>');
}
render () {
return (
<div className="payment">
<h5 className="payment-header">Payment Info</h5>
<form onChange={this.props.paymentChange} onSubmit={this.props.submitPayment} method="POST" id="payment-form">
<div className="payment-div">
<label>
<span className="payment-span">Name On Card</span>
<input className="payment-input" type="text" size="20" data-stripe="number" />
</label>
</div>
<div className="payment-div">
<label>
<span className="payment-span">Card Number</span>
<input className="payment-input" type="text" size="20" id="number" data-stripe="number" />
</label>
</div>
<div className="payment-div">
<label>
<span className="payment-span">Expiration (MM/YY)</span>
<input className="payment-input" type="text" size="2" id="exp_month" data-stripe="exp_month" />
</label>
<span className="payment-span"> / </span>
<input className="payment-input" type="text" size="2" id="exp_year" data-stripe="exp_year" />
</div>
<div className="payment-div">
<label>
<span className="payment-span">CVC</span>
<input className="payment-input" type="text" size="4" id="cvc" data-stripe="cvc" />
</label>
</div>
<div className="col-xs-12 payment-container">
<Cart winningBids={this.props.winningBids} submitPayment={this.props.submitPayment} />
</div>
<input type="submit" className="submit" value="Submit Payment" id="form-submit-disable" />
<span className="payment-span payment-errors"></span>
</form>
</div>
);
}
} | b84bc664254557bffc1cfc0bdaeed11d7fccd0c9 | [
"JavaScript"
] | 2 | JavaScript | chrispicato/TickrTaker | 0b28fa904ed5bd6c88c448d19aedd7a02a21783d | 6767a09a954940c4301b4688c20b967aaf9cfc75 |
refs/heads/master | <repo_name>leoliuzhou/SinaVideoSdkDemo<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoPlayButton.java
package com.sina.sinavideo.sdk.widgets;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.util.Log;
import android.view.KeyEvent;
import android.view.View;
import android.view.ViewGroup;
import android.widget.ImageView;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideControllerListener;
import com.sina.sinavideo.sdk.data.VDVideoInfo;
import com.sina.sinavideo.sdk.dlna.DLNAController;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.utils.VDPlayPauseHelper;
import com.sina.sinavideo.sdk.R;
/**
* 播放\暂停视频按钮
*
* @author liuqun
*/
public final class VDVideoPlayButton extends ImageView implements VDBaseWidget,
VDVideoViewListeners.OnPlayVideoListener, OnShowHideControllerListener {
private Context mContext;
private int mPlayRes = R.drawable.play_ctrl_pause;
private int mPauseRes = R.drawable.play_ctrl_play;
private VDPlayPauseHelper mVDPlayPauseHelper;
public VDVideoPlayButton(Context context) {
super(context);
mContext = context;
VDLog.d("VDVideoPlayButton", "context ctt=" + context);
setImageResource(mPauseRes);
registerListeners();// 注册事件监听
init();
}
public VDVideoPlayButton(Context context, AttributeSet attrs) {
super(context, attrs);
mContext = context;
VDLog.d("VDVideoPlayButton", "context ctt=" + context);
TypedArray typedArr = context.obtainStyledAttributes(attrs,
R.styleable.VDVideoPlayButton);
if (typedArr != null) {
for (int i = 0; i < typedArr.getIndexCount(); i++) {
int resID = -1;
if (typedArr.getIndex(i) == R.styleable.VDVideoPlayButton_pausedRes) {
resID = typedArr.getResourceId(
R.styleable.VDVideoPlayButton_pausedRes, -1);
if (resID != -1) {
mPauseRes = resID;
}
} else if (typedArr.getIndex(i) == R.styleable.VDVideoPlayButton_playingRes) {
resID = typedArr.getResourceId(
R.styleable.VDVideoPlayButton_playingRes, -1);
if (resID != -1) {
mPlayRes = resID;
}
}
// switch (typedArr.getIndex(i)) {
// case R.styleable.VDVideoPlayButton_pausedRes :
// resID =
// typedArr.getResourceId(R.styleable.VDVideoPlayButton_pausedRes,
// -1);
// if (resID != -1) {
// mPauseRes = resID;
// }
// break;
// case R.styleable.VDVideoPlayButton_playingRes :
// resID =
// typedArr.getResourceId(R.styleable.VDVideoPlayButton_playingRes,
// -1);
// if (resID != -1) {
// mPlayRes = resID;
// }
// break;
// }
}
typedArr.recycle();
}
init();
registerListeners();// 注册事件监听
VDVideoViewController.getInstance(context).addOnPlayVideoListener(this);
}
private void init() {
mVDPlayPauseHelper = new VDPlayPauseHelper(getContext());
setOnFocusChangeListener(new OnFocusChangeListener() {
@Override
public void onFocusChange(View v, boolean hasFocus) {
if (hasFocus) {
VDVideoViewController controller = VDVideoViewController
.getInstance(VDVideoPlayButton.this.getContext());
if (controller != null)
controller.notifyShowControllerBar(true);
}
}
});
}
@Override
public void reset() {
// if (VDVideoViewController.getInstance().mVDPlayerInfo.mPlayStatus ==
// VDPlayerInfo.PLAYER_ISPLAYING) {
// setSelected(false);
// } else {
// setSelected(true);
// }
onPlayStateChanged();
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.addOnPlayVideoListener(this);
}
@Override
public void hide() {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.removeOnPlayVideoListener(this);
}
/**
* 为自己注册事件
*/
private void registerListeners() {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null)
controller.addOnShowHideControllerListener(this);
// click事件
this.setOnClickListener(new OnClickListener() {
@Override
public void onClick(View v) {
if (DLNAController.mIsDLNA) {
DLNAController.getInstance(mContext).onClickPlay();
return;
}
doClick();
}
});
}
private void doClick() {
mVDPlayPauseHelper.doClick();
}
@Override
public void onVideoInfo(VDVideoInfo info) {
}
@Override
public void onShowLoading(boolean show) {
}
@Override
public void onVideoPrepared(boolean prepare) {
}
@Override
public void onPlayStateChanged() {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null && controller.mVDPlayerInfo.mIsPlaying) {
setImageResource(mPlayRes);
} else {
setImageResource(mPauseRes);
}
}
@Override
public boolean onKeyDown(int keyCode, KeyEvent event) {
VDLog.e("VDVideoPlayButton", " onKeyDown ");
// if(keyCode == KeyEvent.KEYCODE_ENTER || keyCode ==
// KeyEvent.KEYCODE_DPAD_CENTER){
// doClick();
// return true;
// }
return super.onKeyDown(keyCode, event);
}
@Override
public boolean onKeyUp(int keyCode, KeyEvent event) {
VDLog.e("VDVideoPlayButton", " onKeyUp ");
if (keyCode == KeyEvent.KEYCODE_ENTER
|| keyCode == KeyEvent.KEYCODE_DPAD_CENTER) {
doClick();
return true;
}
if (keyCode == KeyEvent.KEYCODE_DPAD_RIGHT) {
VDLog.e("VDVideoPlayButton",
" onKeyUp KEYCODE_DPAD_RIGHT 111111111");
int id = getNextFocusRightId();
View v = ((ViewGroup) getParent()).findViewById(id);
if (v != null) {
VDLog.e("VDVideoPlayButton", " onKeyUp KEYCODE_DPAD_RIGHT");
v.requestFocus();
}
return true;
}
return super.onKeyUp(keyCode, event);
}
@Override
public void doNotHideControllerBar() {
}
@Override
public void hideControllerBar(long delay) {
}
@Override
public void onPostHide() {
Log.i("VDVideoPlayButton", "onPostHide key--> 失去焦点");
clearFocus();
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null && controller.getVideoView() != null) {
((View) controller.getVideoView()).requestFocus();
}
}
@Override
public void onPostShow() {
postDelayed(new Runnable() {
@Override
public void run() {
// setFocusable(true);
requestFocus();
Log.i("VDVideoPlayButton", "key onPostShow --> " + isFocused());
}
}, 50);
}
@Override
public void onPreHide() {
}
@Override
public void showControllerBar(boolean delayHide) {
}
@Override
public void onPreShow() {
}
}
<file_sep>/SinaVideoSdk/settings.gradle
include ':app', ':sinavideo_playercore', ':sinavideosdk2'
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/playlist/VDVideoPlayListContainer.java
package com.sina.sinavideo.sdk.widgets.playlist;
import android.content.Context;
import android.util.AttributeSet;
import android.view.View;
import android.view.animation.Animation;
import android.view.animation.AnimationUtils;
import android.widget.LinearLayout;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoListVisibleChangeListener;
import com.sina.sinavideo.sdk.widgets.VDBaseWidget;
import com.sina.sinavideo.sdk.R;
/**
* 播放列表弹出界面,容器部分
*
* @author liuqun
*/
public class VDVideoPlayListContainer extends LinearLayout implements VDBaseWidget, OnVideoListVisibleChangeListener {
public interface OnPlayListItemClick {
public void onItemClick(int position);
}
private Animation mShowAnim;
private Animation mHideAnim;
private Context mContext = null;
/**
* 容器的适配器,因为不知道当前适配的界面,所以,ViewHolder没法弄了,短列表姑且认为没问题吧
*
* @author sunxiao
*
*/
public VDVideoPlayListContainer(Context context, AttributeSet attrs) {
super(context, attrs);
// TODO Auto-generated constructor stub
init(context, attrs);
}
private void init(Context context, AttributeSet attrs) {
mContext = context;
setOnClickListener(new OnClickListener() {
@Override
public void onClick(View v) {
// setVisibility(GONE);
removeCallbacks(mHideAction);
post(mHideAction);
}
});
// 如果有自定义属性,加载
// int itemID = -1;
// TypedArray typedArr = context.obtainStyledAttributes(attrs,
// R.styleable.VDVideoPlaylistContainer);
// for (int i = 0; i < typedArr.getIndexCount(); i++) {
// switch (typedArr.getIndex(i)) {
// default :
// break;
// case R.styleable.VDVideoPlaylistContainer_listItem :
// itemID =
// typedArr.getResourceId(R.styleable.VDVideoPlaylistContainer_listItem,
// -1);
// break;
// }
// }
// typedArr.recycle();
// 设置adapter
// VDVideoViewController.getInstance().addOnVideoListListener(this);
VDVideoViewController controller = VDVideoViewController.getInstance(context);
if(null!=controller)controller.addOnVideoListVisibleChangeListener(this);
mShowAnim = AnimationUtils.loadAnimation(mContext, R.anim.video_list_from_right_in);
mShowAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
setVisibility(View.VISIBLE);
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
clearAnimation();
}
});
mHideAnim = AnimationUtils.loadAnimation(mContext, R.anim.video_list_fade_from_right);
mHideAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
setVisibility(View.GONE);
clearAnimation();
//VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
//if(null!=controller)controller.notifyShowBottomControllerBar();
}
});
}
@Override
public void reset() {
}
@Override
public void hide() {
setVisibility(GONE);
}
@Override
public void toogle() {
if (getAnimation() != null) {
return;
}
if (getVisibility() == VISIBLE) {
removeCallbacks(mHideAction);
hidePlayList();
// TODO 什么意思??
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if(null!=controller)controller.notifyShowBottomControllerBar();
} else {
setVisibility(VISIBLE);
showPlayList();
}
}
public Runnable mHideAction = new Runnable() {
@Override
public void run() {
if (getVisibility() == VISIBLE && getAnimation() == null) {
// setVisibility(GONE);
//VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
//if(null!=controller)controller.notifyHideControllerBar(0);
startAnimation(mHideAnim);
}
}
};
@Override
public void showPlayList() {
startAnimation(mShowAnim);
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if(null!=controller)controller.notifyNotHideControllerBar();
postDelayed(mHideAction, VDVideoViewController.DEFAULT_DELAY);
// VDVideoViewController.getInstance().notifyHideBottomControllerBar();
}
@Override
public void hidePlayList() {
removeCallbacks(mHideAction);
if (getVisibility() == VISIBLE) {
// VDVideoViewController.getInstance().notifyHideControllerBar(VDVideoViewController.DEFAULT_DELAY);
// startAnimation(mHideAnim);
post(mHideAction);
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if(null!=controller)controller.notifyHideTopControllerBar();
if(null!=controller)controller.notifyHideBottomControllerBar();
// setVisibility(GONE);
}
}
@Override
public void removeAndHideDelay() {
removeCallbacks(mHideAction);
postDelayed(mHideAction, VDVideoViewController.DEFAULT_DELAY);
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoLoadingPercentView.java
package com.sina.sinavideo.sdk.widgets;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnBufferingUpdateListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnLoadingListener;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.R;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.View;
import android.widget.TextView;
public final class VDVideoLoadingPercentView extends TextView implements
VDBaseWidget, OnLoadingListener, OnBufferingUpdateListener {
private String mLoadingText = "正在缓冲%d%%";
private String mPreLoadingText = "开始加载";
private Context mContext = null;
private boolean mIsVisible = false;
public VDVideoLoadingPercentView(Context context) {
super(context);
// TODO Auto-generated constructor stub
init(context);
}
public VDVideoLoadingPercentView(Context context, AttributeSet attrs) {
super(context, attrs);
// TODO Auto-generated constructor stub
TypedArray typedArr = context.obtainStyledAttributes(attrs,
R.styleable.VDVideoLoadingPercentView);
if (typedArr != null) {
for (int i = 0; i < typedArr.getIndexCount(); i++) {
if (typedArr.getIndex(i) == R.styleable.VDVideoLoadingPercentView_loadingText) {
mLoadingText = typedArr
.getString(R.styleable.VDVideoLoadingPercentView_loadingText);
} else if (typedArr.getIndex(i) == R.styleable.VDVideoLoadingPercentView_preLoadingText) {
mPreLoadingText = typedArr
.getString(R.styleable.VDVideoLoadingPercentView_preLoadingText);
}
}
typedArr.recycle();
}
init(context);
}
private void init(Context context) {
mContext = context;
registerListener();
}
private void registerListener() {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller != null) {
controller.addOnBufferingUpdateListener(this);
controller.addOnLoadingListener(this);
}
}
@Override
public void reset() {
// TODO Auto-generated method stub
registerListener();
}
@Override
public void hide() {
// TODO Auto-generated method stub
}
@Override
public void showLoading() {
// TODO Auto-generated method stub
setText(mPreLoadingText);
setVisibility(View.VISIBLE);
mIsVisible = true;
}
@Override
public void hideLoading() {
// TODO Auto-generated method stub
setVisibility(View.GONE);
mIsVisible = false;
}
@Override
public void onBufferingUpdate(int percent) {
// TODO Auto-generated method stub
if (mIsVisible) {
try {
String formatTxt = String.format(mLoadingText, percent);
setText(formatTxt);
} catch (Exception ex) {
VDLog.e(VIEW_LOG_TAG, ex.getMessage());
ex.printStackTrace();
}
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoADSoundButton.java
package com.sina.sinavideo.sdk.widgets;
import com.sina.sinavideo.sdk.R;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSoundChangedListener;
import com.sina.sinavideo.sdk.utils.VDPlayerSoundManager;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.View;
import android.widget.ImageButton;
/**
* 广告用的音量按钮,静音开关方式,点击后或者开启或者关闭声音
*
* @author sunxiao
*
*/
public final class VDVideoADSoundButton extends ImageButton implements
VDBaseWidget, OnSoundChangedListener {
private int mResID = -1;
private int mSilentResID = -1;
private Context mContext = null;
public VDVideoADSoundButton(Context context) {
super(context);
// TODO Auto-generated constructor stub
registerListeners();
init(context);
}
public VDVideoADSoundButton(Context context, AttributeSet attrs) {
super(context, attrs);
// TODO Auto-generated constructor stub
// 支持自定义背景
TypedArray typedArr = context.obtainStyledAttributes(attrs,
new int[] { android.R.attr.background });
if (typedArr != null) {
mResID = typedArr.getResourceId(0, -1);
if (mResID == -1) {
mResID = R.drawable.play_ctrl_volume;
}
typedArr.recycle();
}
// 静音部分的定义
typedArr = context.obtainStyledAttributes(attrs,
R.styleable.VDVideoADSoundButton);
mSilentResID = R.drawable.ad_silent_selcetor;
if (typedArr != null) {
for (int i = 0; i < typedArr.getIndexCount(); i++) {
if (typedArr.getIndex(i) == R.styleable.VDVideoADSoundButton_adSoundSeekSilent) {
// 自定义背景的时候,换静音按钮
mSilentResID = typedArr.getResourceId(i, -1);
}
}
typedArr.recycle();
}
registerListeners();
init(context);
}
private void init(Context context) {
mContext = context;
changeBackground(VDPlayerSoundManager.isMuted(context));
}
private void changeBackground(boolean isSilent) {
if (isSilent)
setBackgroundResource(mSilentResID);
else
setBackgroundResource(mResID);
}
/**
* 为自己注册事件
*/
private void registerListeners() {
// click事件
this.setOnClickListener(new OnClickListener() {
@Override
public void onClick(View v) {
// 先测试添加于此
boolean isMuted = VDPlayerSoundManager.isMuted(mContext);
isMuted = !isMuted;
VDPlayerSoundManager.setMute(mContext, isMuted, false);
changeBackground(isMuted);
}
});
}
@Override
public void reset() {
// TODO Auto-generated method stub
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.addOnSoundChangedListener(this);
}
@Override
public void hide() {
// TODO Auto-generated method stub
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.removeOnSoundChangedListener(this);
}
@Override
public void onSoundChanged(int currVolume) {
// TODO Auto-generated method stub
int currSoundNum = VDPlayerSoundManager.getCurrSoundVolume(mContext);
if (currSoundNum > 0) {
changeBackground(false);
} else {
changeBackground(true);
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoSoundSeekButton.java
package com.sina.sinavideo.sdk.widgets;
import android.app.Activity;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.View;
import android.widget.ImageButton;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSoundChangedListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSoundVisibleListener;
import com.sina.sinavideo.sdk.utils.VDPlayerSoundManager;
import com.sina.sinavideo.sdk.R;
/**
* 音量控制按钮部分,点击打开音量控制拖拉条
*
* @author seven
*/
public final class VDVideoSoundSeekButton extends ImageButton implements
VDBaseWidget, OnSoundChangedListener, OnSoundVisibleListener {
private Context mContext = null;
private int mContainerID = -1;
// 普通背景
private int mResID = -1;
// 静音情况下的背景
private int mSilentResID = -1;
public VDVideoSoundSeekButton(Context context, AttributeSet attrs) {
super(context, attrs);
// TODO Auto-generated constructor stub
mContext = context;
// 支持自定义背景
TypedArray typedArr = context.obtainStyledAttributes(attrs,
new int[] { android.R.attr.background });
mResID = R.drawable.play_ctrl_volume;
if (typedArr != null) {
mResID = typedArr.getResourceId(0, -1);
if (mResID == -1) {
mResID = R.drawable.play_ctrl_volume;
}
typedArr.recycle();
}
setBackgroundResource(mResID);
// 指定相应的音量控制容器类
typedArr = context.obtainStyledAttributes(attrs,
R.styleable.VDVideoSoundSeekButton);
mSilentResID = R.drawable.ad_silent_selcetor;
if (typedArr != null) {
for (int i = 0; i < typedArr.getIndexCount(); i++) {
if (typedArr.getIndex(i) == R.styleable.VDVideoSoundSeekButton_soundSeekContainer) {
mContainerID = typedArr.getResourceId(i, -1);
} else if (typedArr.getIndex(i) == R.styleable.VDVideoSoundSeekButton_soundSeekSilent) {
// 自定义背景的时候,换静音按钮
mSilentResID = typedArr.getResourceId(i, -1);
}
}
typedArr.recycle();
}
registerListeners();
}
@Override
public void reset() {
// TODO Auto-generated method stub
VDVideoViewController.getInstance(this.getContext())
.addOnSoundChangedListener(this);
}
@Override
public void hide() {
// TODO Auto-generated method stub
// VDVideoViewController.getInstance().removeOnSoundChangedListener(this);
}
/**
* 为自己注册事件
*/
private void registerListeners() {
// click事件
this.setOnClickListener(new OnClickListener() {
@Override
public void onClick(View v) {
// 先测试添加于此
if (mContainerID == -1) {
return;
}
View container = ((Activity) mContext)
.findViewById(mContainerID);
if (container != null) {
if (container.getVisibility() == View.GONE) {
// container.setVisibility(View.VISIBLE);
VDVideoViewController controller = VDVideoViewController
.getInstance(VDVideoSoundSeekButton.this
.getContext());
if (controller != null)
controller.notifySoundSeekBarVisible(true);
} else {
// container.setVisibility(View.GONE);
VDVideoViewController controller = VDVideoViewController
.getInstance(VDVideoSoundSeekButton.this
.getContext());
if (controller != null)
controller.notifySoundSeekBarVisible(false);
}
}
}
});
}
@Override
public void onSoundChanged(int soundIndex) {
if (soundIndex <= 0) {
setBackgroundResource(mSilentResID);
} else {
setBackgroundResource(mResID);
}
}
@Override
public void onSoundVisible(boolean isVisible) {
int soundIndex = VDPlayerSoundManager.getCurrSoundVolume(mContext);
if (soundIndex <= 0) {
setBackgroundResource(mSilentResID);
} else {
setBackgroundResource(mResID);
}
if (isVisible) {
setPressed(true);
} else {
setPressed(false);
}
}
@Override
public void onSoundSeekBarVisible(boolean isVisible) {
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoADTicker.java
package com.sina.sinavideo.sdk.widgets;
import java.util.ArrayList;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoInsertADListener;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.R;
import android.annotation.SuppressLint;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.View;
import android.widget.ImageView;
import android.widget.LinearLayout;
import android.widget.TextView;
/**
* 视频插入广告的时钟跳数widget,规则如下:<br>
* 1、当时钟跳数为0,则不管当前广告是播放完毕,则直接进入整片<br>
* 2、当前广告因为编码、网络等原因导致无法播放,不进跳数过程,按照错误处理,直接跳入正片
*
* @author sunxiao
*/
public final class VDVideoADTicker extends LinearLayout implements
VDBaseWidget, OnVideoInsertADListener {
private ArrayList<Integer> mTickerImgList = new ArrayList<Integer>();
private ImageView mImg1 = null;
private ImageView mImg2 = null;
private TextView mTV1 = null;
private TextView mTV2 = null;
// 字体默认为白色
private int mTVTextColor = 0xFFFFFF;
// 字体大小默认为12sp
private float mTVTextSize = 12;
@SuppressLint("nouse")
private Context mContext = null;
private final String TAG = "VDVideoADTicker";
private void refreshCurrSecNum(final int tickerNum) throws Exception {
int units = 9;
int decade = 9;
if (tickerNum < 100) {
units = tickerNum % 10;
decade = tickerNum / 10;
}
// TODO Auto-generated method stub
if (mTickerImgList == null || mTickerImgList.size() == 0) {
mTV1.setText(decade + "");
mTV2.setText(units + "");
} else {
mImg1.setImageDrawable(getResources().getDrawable(
mTickerImgList.get(decade)));
mImg2.setImageDrawable(getResources().getDrawable(
mTickerImgList.get(units)));
}
}
public VDVideoADTicker(Context context) {
super(context);
// TODO Auto-generated constructor stub
mContext = context;
initLayout(context);
}
public VDVideoADTicker(Context context, AttributeSet attrs) {
super(context, attrs);
mContext = context;
// TODO Auto-generated constructor stub
int tickerImgListRes = -1;
TypedArray typedArr = context.obtainStyledAttributes(attrs,
R.styleable.VDVideoADTicker);
if (typedArr != null) {
for (int i = 0; i < typedArr.getIndexCount(); i++) {
if (typedArr.getIndex(i) == R.styleable.VDVideoADTicker_tickerImgList) {
int resID = typedArr.getResourceId(
R.styleable.VDVideoADTicker_tickerImgList, -1);
if (resID != -1) {
tickerImgListRes = resID;
}
} else if (typedArr.getIndex(i) == R.styleable.VDVideoADTicker_tickerTextColor) {
int txtColor = typedArr.getColor(
R.styleable.VDVideoADTicker_tickerTextColor, -1);
if (txtColor != -1) {
mTVTextColor = txtColor;
}
} else if (typedArr.getIndex(i) == R.styleable.VDVideoADTicker_tickerTextSize) {
float txtSize = typedArr.getDimension(
R.styleable.VDVideoADTicker_tickerTextSize, -1);
if (txtSize != -1) {
mTVTextSize = txtSize;
}
}
}
}
mTickerImgList.clear();
if (tickerImgListRes != -1) {
// 判断是否是一个array,如果不是,报错
String type = getResources().getResourceTypeName(tickerImgListRes);
if (type != null && type.equals("array")) {
TypedArray imgList = getResources().obtainTypedArray(
tickerImgListRes);
if (imgList == null || imgList.length() != 10) {
imgList.recycle();
throw new IllegalArgumentException("数字图片数组必须为10个");
}
for (int i = 0; i < imgList.length(); i++) {
mTickerImgList.add(imgList.getResourceId(i, -1));
}
imgList.recycle();
}
}
typedArr.recycle();
initLayout(context);
}
private void initLayout(Context context) {
if (mImg1 == null)
mImg1 = new ImageView(context);
if (mImg2 == null)
mImg2 = new ImageView(context);
if (mTV1 == null) {
mTV1 = new TextView(context);
mTV1.setTextColor(mTVTextColor);
mTV1.setTextSize(mTVTextSize);
}
if (mTV2 == null) {
mTV2 = new TextView(context);
mTV2.setTextColor(mTVTextColor);
mTV2.setTextSize(mTVTextSize);
}
// 设置下层的两个图片的属性
LayoutParams params1 = new LayoutParams(
LayoutParams.WRAP_CONTENT, LayoutParams.WRAP_CONTENT);
LayoutParams params2 = new LayoutParams(
LayoutParams.WRAP_CONTENT, LayoutParams.WRAP_CONTENT);
if (mTickerImgList == null || mTickerImgList.size() == 0) {
addView(mTV1, params1);
addView(mTV2, params2);
} else {
addView(mImg1, params1);
addView(mImg2, params2);
}
}
@Override
public void reset() {
// TODO Auto-generated method stub
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.addOnVideoInsertADListener(this);
// 需要同步一下,当前的时间值,可能会有变化
}
@Override
public void hide() {
// TODO Auto-generated method stub
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.removeOnVideoInsertADListener(this);
}
@Override
public void onVideoInsertADBegin() {
// TODO Auto-generated method stub
setVisibility(View.VISIBLE);
try {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller == null) {
return;
}
refreshCurrSecNum(controller.getADTickerSec());
} catch (Exception ex) {
VDLog.e(TAG, ex.getMessage());
}
}
@Override
public void onVideoInsertADTicker() {
// TODO Auto-generated method stub
try {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller == null) {
return;
}
refreshCurrSecNum(controller.refreshADTickerSec());
} catch (Exception ex) {
VDLog.e(TAG, ex.getMessage());
}
}
@Override
public void onVideoInsertADEnd() {
// TODO Auto-generated method stub
setVisibility(View.GONE);
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoFullScreenButton.java
/**
* 全屏按钮部分,点击后,横屏显示
*
* @author sunxiao
*/
package com.sina.sinavideo.sdk.widgets;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.View;
import android.widget.ImageButton;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.utils.VDVideoFullModeController;
import com.sina.sinavideo.sdk.R;
public final class VDVideoFullScreenButton extends ImageButton implements
VDBaseWidget {
public VDVideoFullScreenButton(Context context) {
super(context);
init();
}
public VDVideoFullScreenButton(Context context, AttributeSet attrs) {
super(context, attrs);
// 判断是否有新定义的背景图片,没有则使用sina视频的默认图片
TypedArray typedArr = context.obtainStyledAttributes(attrs,
new int[] { android.R.attr.background });
if (typedArr != null) {
int resouceID = typedArr.getResourceId(0, -1);
if (resouceID == -1) {
setBackgroundResource(R.drawable.play_ctrl_fullscreen);
}
typedArr.recycle();
} else {
setBackgroundResource(R.drawable.play_ctrl_fullscreen);
}
init();
}
private void init() {
registerListeners();
}
private void registerListeners() {
setOnClickListener(new OnClickListener() {
@Override
public void onClick(View v) {
if (!VDVideoFullModeController.getInstance().getIsFullScreen()) {
VDVideoViewController controller = VDVideoViewController
.getInstance(VDVideoFullScreenButton.this
.getContext());
if (null != controller)
controller.setIsFullScreen(true);
}
}
});
}
@Override
public void reset() {
}
@Override
public void hide() {
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoResolutionList.java
package com.sina.sinavideo.sdk.widgets;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.MotionEvent;
import android.view.View;
import android.widget.LinearLayout;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnResolutionContainerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnScreenTouchListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideControllerListener;
import com.sina.sinavideo.sdk.R;
/**
* 清晰度选择菜单
*
* @author liuqun
*
*/
public final class VDVideoResolutionList extends LinearLayout implements
VDBaseWidget, OnShowHideControllerListener, OnScreenTouchListener,
OnResolutionContainerListener {
String tag = "VDVideoResolutionList";
public VDVideoResolutionList(Context context) {
super(context);
init();
}
public VDVideoResolutionList(Context context, AttributeSet attrs) {
super(context, attrs);
init();
TypedArray a = context.obtainStyledAttributes(attrs,
R.styleable.ResolutionBackGround);
a.recycle();
}
private void init() {
setBackgroundResource(R.drawable.definition_select_bg);
}
@Override
protected void onFinishInflate() {
super.onFinishInflate();
}
@Override
public void reset() {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null) {
// controller.addOnResolutionListener(this);
controller.addOnResolutionContainerListener(this);
controller.addOnScreenTouchListener(this);
controller.addOnShowHideControllerListener(this);
}
}
@Override
public void hide() {
this.setVisibility(View.GONE);
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null) {
// controller.removeOnResolutionListener(this);
controller.removeOnResolutionContainerListener(this);
controller.removeOnScreenTouchListener(this);
controller.removeOnShowHideControllerListener(this);
}
}
// public void setResolutionList(VDResolutionData resolutionData) {
// Log.i(tag, "setResolutionList --> " + resolutionData.toString());
// if (resolutionData != null) {
//
// }
// }
@Override
public void onSingleTouch(MotionEvent ev) {
setVisibility(GONE);
}
// @Override
// public void onResolutionSelect(String resolution) {
// Log.i(tag, "onResolutionSelect -> " + resolution);
// if (resolution != null) {
//
// }
// }
// @Override
// public void onParseResolution(VDResolutionData list) {
// setResolutionList(list);
// }
// @Override
// public void hideResolution() {
// setVisibility(GONE);
// }
public void focusFirstView() {
// mResolutionButtonBase.requestFocus();
}
@Override
public void doNotHideControllerBar() {
}
@Override
public void hideControllerBar(long delay) {
}
@Override
public void showControllerBar(boolean delayHide) {
}
@Override
public void onPostHide() {
setVisibility(View.GONE);
}
@Override
public void onPostShow() {
}
@Override
public void onPreHide() {
setVisibility(View.GONE);
}
@Override
public void onPreShow() {
// TODO Auto-generated method stub
}
@Override
public void onResolutionContainerVisible(boolean isVisible) {
// TODO Auto-generated method stub
if (isVisible) {
setVisibility(View.VISIBLE);
} else {
setVisibility(View.GONE);
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/VDVideoViewListeners.java
package com.sina.sinavideo.sdk;
import java.util.HashSet;
import java.util.Set;
import android.content.Context;
import android.graphics.PointF;
import android.media.TimedText;
import android.os.Handler;
import android.util.Log;
import android.view.MotionEvent;
import com.sina.sinavideo.coreplayer.ISinaVideoView;
import com.sina.sinavideo.sdk.data.VDPlayerInfo;
import com.sina.sinavideo.sdk.data.VDResolutionData;
import com.sina.sinavideo.sdk.data.VDVideoInfo;
import com.sina.sinavideo.sdk.data.VDVideoListInfo;
import com.sina.sinavideo.sdk.dlna.DLNAController;
import com.sina.sinavideo.sdk.utils.VDApplication;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.utils.VDPlayerSoundManager;
import com.sina.sinavideo.sdk.utils.VDVideoFullModeController;
/**
* 管理控制类中所有回调消息
*
* @author haifeng9
*
*/
public class VDVideoViewListeners {
private final static String TAG = "VDVideoViewListeners";
private Context mContext;
// 日志相关
// private LogPushManager mLogPushManager;
/**
* 所有的事件的定义
*/
public interface OnBufferingUpdateListener {
abstract void onBufferingUpdate(int percent);
}
public interface OnCompletionListener {
abstract void onCompletion();
}
public interface OnErrorListener {
abstract boolean onError(int what, int extra);
}
public interface OnPauseListener {
public void onPause();
}
public interface OnInfoListener {
abstract boolean onInfo(int what, int extra);
}
public interface OnPreparedListener {
abstract void onPrepared();
}
public interface OnSeekCompleteListener {
abstract void onSeekComplete();
}
public interface OnVideoSizeChangedListener {
abstract void onVideoSizeChanged(int width, int height);
}
public interface OnTimedTextListener {
public void onTimedText(TimedText text);
}
public interface OnProgressUpdateListener {
public void onProgressUpdate(long current, long duration);
public void onDragProgess(long progress, long duration);
}
public interface OnVideoOpenedListener {
public void onVideoOpened();
}
public interface OnVideoUIRefreshListener {
public void onVideoUIRefresh();
}
public interface OnPlayVideoListener {
public void onVideoInfo(VDVideoInfo info);
public void onShowLoading(boolean show);
public void onVideoPrepared(boolean prepare);
public void onPlayStateChanged();
}
// public interface OnLiveVideoListener {
//
// /**
// * 如果有分辨率的信息,就有list,如果没有list为null
// */
// public void onResolutionInfo(VDResolutionData list);
//
// public void onResolutionIndex(int index);
// }
/**
* 全屏事件,转屏时候,会发送此通知
*
* @author sunxiao
*
*/
public interface OnFullScreenListener {
public void onFullScreen(boolean isFullScreen, boolean isFromHand);
}
/**
* 音量改变后的回调
*
* @author liuqun1
*
*/
public interface OnSoundChangedListener {
public void onSoundChanged(int currVolume);
}
/**
* 点击按钮,播放或暂停操作
*
* @author liuqun
*
*/
public interface OnClickPlayListener {
public void onClickPlay();
}
/**
* 音量调节空间显示
*
* @author sunxiao
*
*/
public interface OnSoundVisibleListener {
public void onSoundVisible(boolean isVisible);
public void onSoundSeekBarVisible(boolean isVisible);
}
/**
* 清晰度控件显示
*
* @author GengHongchao
*
*/
public interface OnVMSResolutionListener {
public void onVMSResolutionContainerVisible(boolean isVisible);
public void onVMSResolutionChanged();
}
public enum eSingleTouchListener {
eTouchListenerSingleTouchStart, eTouchListenerSingleTouch, eTouchListenerSingleTouchEnd,
}
public enum eDoubleTouchListener {
eTouchListenerDoubleTouchStart, eTouchListenerDoubleTouch, eTouchListenerDoubleTouchEnd,
}
public enum eVerticalScrollTouchListener {
eTouchListenerVerticalScrollStart, eTouchListenerVerticalScroll, eTouchListenerVerticalScrollSound, eTouchListenerVerticalScrollLighting, eTouchListenerVerticalScrollEnd,
}
public enum eHorizonScrollTouchListener {
eTouchListenerHorizonScrollStart, eTouchListenerHorizonScroll, eTouchListenerHorizonScrollEnd,
}
/**
* 触屏相关的消息
*
* @author sunxiao
*
*/
public interface OnScreenTouchListener {
/**
* 单击,隐藏、显示
*
* @param ev
*/
public void onSingleTouch(MotionEvent ev);
}
/**
* 亮度调节
*
* @author sunxiao
*
*/
public interface OnLightingChangeListener {
public void onLightingChange(float curr);
}
/**
* 亮度调节显示
*
* @author sunxiao
*
*/
public interface OnLightingVisibleListener {
public void onLightingVisible(boolean isVisible);
}
/**
* 清晰度解析完回调,以及选择清晰度的回调
*
* @author liuqun1
*
*/
// public interface OnResolutionListener {
//
// void onResolutionSelect(String resolution);
//
// void onParseResolution(VDResolutionData list);
//
// void hideResolution();
// }
/**
* 清晰度改变
*
* @author sunxiao
*
*/
public interface OnResolutionListener {
/**
* 清晰度按钮点击后回传
*
* @param tag
*/
public void onResolutionChanged(String tag);
/**
* 清晰度解析完毕时候回传
*
* @param list
*/
public void onResolutionParsed(VDResolutionData list);
}
/**
* 清晰度按钮容器之类的隐藏显示
*
* @author alexsun
*
*/
public interface OnResolutionContainerListener {
public void onResolutionContainerVisible(boolean isVisible);
}
/**
* 在遥控器等TV项目中,在进入container焦点的时候,触发的第一个清晰度按钮
*
* @author alexsun
*
*/
public interface OnResolutionListButtonListener {
public void onResolutionListButtonFocusFirst();
}
/**
* 清晰度选择按钮,显示或隐藏清晰度列表
*
* @author liuqun1
*
*/
// public interface OnResolutionVisibleChangeListener {
//
// void toogle();
// }
/**
* 提示通知
*
* @author liuqun1
*
*/
public interface OnTipListener {
void onTip(String tip);
void onTip(int tipResId);
void hideTip();
}
/**
* 加载进度条回调
*
* @author liuqun1
*
*/
public interface OnLoadingListener {
void showLoading();
void hideLoading();
}
/**
* 新手引导页部分
*
* @author sunxiao
*
*/
public interface OnVideoGuideTipsListener {
public void onVisible(boolean isVisible);
}
/**
* 静帧广告部分,弄起来很麻烦,可能有问题 [点击暂停的时候,出现此东东]
*
* @author sunxiao
*
*/
public interface OnVideoFrameADListener {
/**
* 广告显示开始
*/
public void onVideoFrameADBegin();
/**
* 广告播放时长结束
*/
public void onVideoFrameADEnd();
}
public interface OnVideoInsertADListener {
/**
* 插入式广告开始
*/
public void onVideoInsertADBegin();
/**
* 插入式广告的跳秒信号
*/
public void onVideoInsertADTicker();
/**
* 插入式广告结束
*/
public void onVideoInsertADEnd();
}
/**
* 视频列表通知
*
* @author liuqun1
*
*/
public interface OnVideoListListener {
// /**
// * @deprecated 不再使用,用这个:onVideoList(VDVideoListInfo infoList)
// * @param videoList
// */
// void onVideoList(ArrayList<VDVideoInfo> videoList);
public void onVideoList(VDVideoListInfo infoList);
}
/**
* 视频列表切换显示隐藏通知
*
* @author liuqun1
*
*/
public interface OnVideoListVisibleChangeListener {
void toogle();
void showPlayList();
void hidePlayList();
void removeAndHideDelay();
}
/**
* 更多操作面板切换显示隐藏通知
*
* @author liuqun1
*
*/
public interface OnMoreOprationVisibleChangeListener {
void showPanel();
void hidePanel();
void removeAndHideDelay();
}
/**
* 双击播放/暂停动画view的通知
*
* @author liuqun1
*
*/
public interface OnVideoDoubleTapListener {
void onDoubleTouch();
}
/**
* 解码器部分的触发
*
* @author sunxiao
*
*/
public interface OnDecodingTypeListener {
public void onChange(boolean isFFMpeg);
}
/**
* 屏幕方向发生改变
*
* @author liuqun
*
*/
public interface OnScreenOrientationChangeListener {
void onScreenOrientationVertical();
void onScreenOrientationHorizontal();
}
/**
* 显示或隐藏控制栏
*
* @author liuqun
*
*/
public interface OnShowHideControllerListener {
void doNotHideControllerBar();
void hideControllerBar(long delay);
void showControllerBar(boolean delayHide);
void onPostHide();
void onPostShow();
void onPreHide();
void onPreShow();
}
/**
* 显示或隐藏底部控制栏
*
* @author liuqun
*
*/
public interface OnShowHideBottomControllerListener {
void hideBottomControllerBar();
void showBottomControllerBar();
}
/**
* 显示或者隐藏顶部的控制栏
*
* @author alexsun
*
*/
public interface OnShowHideTopContainerListener {
public void hideTopControllerBar();
public void showTopControllerBar();
}
/**
* 显示或者隐藏前贴片部分控制容器的事件
*
* @author alexsun
*
*/
public interface OnShowHideADContainerListener {
public void hideADContainerBar();
public void showADContainerBar();
}
/**
* 手势调整进度
*
* @author liuqun
*
*/
public interface OnProgressViewVisibleListener {
public void onProgressVisible(boolean isVisible);
}
/**
* 方向键改变进度
*
* @author liuqun
*
*/
public interface OnKeyChangeProgressListener {
public void onKeyDown(boolean keyLeft);
}
/**
* TV 键盘事件
*
* @author liuqun
*
*/
public interface OnKeyEventListener {
/**
* 单击,隐藏、显示
*/
public void onKeyEvent();
/**
* 单击,隐藏、显示
*/
public void onKeyLeftRight();
}
/**
* 音量设置的回调,用于DLNA
*
* @author liuqun1
*
*/
public interface OnSetSoundListener {
public void onSetCurVolume(int currVolume);
public void onSetMaxVolume(int maxVolume);
}
/**
* VDVideoView注册DLNAListener
*
* @author liuqun1
*
*/
public interface OnRegisterDLNAListener {
public void register();
}
/**
* DLNA显示/隐藏Listener
*
* @author liuqun1
*
*/
public interface OnDLNALinearLayoutListener {
public void setLayoutVisiable(boolean visiable);
}
/**
* 横竖屏切换Listener
*
* @author liuqun1
*
*/
public interface OnScreenOrientationSwitchListener {
public void onScreenOrientationSwitch(boolean fullScreen);
}
/**
* 点击错误重试按钮Listener
*
* @author liuqun1
*
*/
public interface OnClickRetryListener {
public void onClickRetry();
}
/**
* 所有的事件保存在此
*/
private Set<OnBufferingUpdateListener> mOnBufferingUpdateListener = new HashSet<OnBufferingUpdateListener>();
private Set<OnCompletionListener> mOnCompletionListener = new HashSet<OnCompletionListener>();
private Set<OnErrorListener> mOnErrorListener = new HashSet<OnErrorListener>();
private Set<OnErrorListener> mOnRetryErrorListener = new HashSet<OnErrorListener>();
private Set<OnInfoListener> mOnInfoListener = new HashSet<OnInfoListener>();
private Set<OnPreparedListener> mOnPreparedListener = new HashSet<OnPreparedListener>();
private Set<OnSeekCompleteListener> mOnSeekCompleteListener = new HashSet<OnSeekCompleteListener>();
private Set<OnVideoSizeChangedListener> mOnVideoSizeChangedListener = new HashSet<OnVideoSizeChangedListener>();
private Set<OnVideoOpenedListener> mOnVideoOpenedListener = new HashSet<OnVideoOpenedListener>();
private Set<OnTimedTextListener> mOnTimedTextListener = new HashSet<OnTimedTextListener>();
private Set<OnProgressUpdateListener> mOnProgressUpdateListener = new HashSet<OnProgressUpdateListener>();
private Set<OnPlayVideoListener> mOnPlayVideoListener = new HashSet<OnPlayVideoListener>();
// private Set<OnLiveVideoListener> mOnLiveVideoListener = new
// HashSet<OnLiveVideoListener>();
private Set<OnFullScreenListener> mOnFullScreenListener = new HashSet<OnFullScreenListener>();
private Set<OnSoundChangedListener> mOnSoundChangedListener = new HashSet<OnSoundChangedListener>();
private Set<OnScreenTouchListener> mOnScreenTouchListener = new HashSet<OnScreenTouchListener>();
private Set<OnResolutionListener> mOnResolutionListener = new HashSet<OnResolutionListener>();
private Set<OnResolutionContainerListener> mOnResolutionContainerListener = new HashSet<OnResolutionContainerListener>();
private Set<OnResolutionListButtonListener> mOnResolutionListButtonListener = new HashSet<OnResolutionListButtonListener>();
private Set<OnTipListener> mOnTipListener = new HashSet<OnTipListener>();
private Set<OnLightingChangeListener> mOnLightingChangeListener = new HashSet<OnLightingChangeListener>();
private Set<OnSoundVisibleListener> mOnSoundVisibleListener = new HashSet<OnSoundVisibleListener>();
private Set<OnVMSResolutionListener> mOnVMSResolutionListener = new HashSet<OnVMSResolutionListener>();
private Set<OnLightingVisibleListener> mOnLightingVisibleListener = new HashSet<OnLightingVisibleListener>();
private Set<OnLoadingListener> mOnLoadingListener = new HashSet<OnLoadingListener>();
private Set<OnVideoGuideTipsListener> mOnVideoGuideTipsListener = new HashSet<OnVideoGuideTipsListener>();
private Set<OnPauseListener> mOnPauseListener = new HashSet<OnPauseListener>();
private Set<OnVideoFrameADListener> mOnVideoFrameADListener = new HashSet<OnVideoFrameADListener>();
private Set<OnVideoInsertADListener> mOnVideoInsertADListener = new HashSet<OnVideoInsertADListener>();
private Set<OnVideoListListener> mOnVideoListListener = new HashSet<OnVideoListListener>();
private Set<OnVideoListVisibleChangeListener> mOnVideoListVisibleChangeListener = new HashSet<OnVideoListVisibleChangeListener>();
private Set<OnMoreOprationVisibleChangeListener> mOnMoreOprationVisibleChangeListener = new HashSet<OnMoreOprationVisibleChangeListener>();
// private Set<OnResolutionVisibleChangeListener>
// mOnResolutionVisibleChangeListener = new
// HashSet<VDVideoViewListeners.OnResolutionVisibleChangeListener>();
private Set<OnVideoDoubleTapListener> mOnVideoDoubleTapListener = new HashSet<OnVideoDoubleTapListener>();
private Set<OnDecodingTypeListener> mOnDecodingTypeListener = new HashSet<OnDecodingTypeListener>();
private Set<OnScreenOrientationChangeListener> mOnScreenOrientationChangeListener = new HashSet<OnScreenOrientationChangeListener>();
private Set<OnShowHideControllerListener> mOnShowHideControllerListener = new HashSet<OnShowHideControllerListener>();
private Set<OnShowHideBottomControllerListener> mOnShowHideBottomControllerListener = new HashSet<OnShowHideBottomControllerListener>();
private Set<OnShowHideTopContainerListener> mOnShowHideTopContainerListener = new HashSet<OnShowHideTopContainerListener>();
private Set<OnProgressViewVisibleListener> mOnProgressViewVisibleListener = new HashSet<OnProgressViewVisibleListener>();
private Set<OnClickPlayListener> mOnClickPlayListener = new HashSet<OnClickPlayListener>();
private Set<OnKeyChangeProgressListener> mOnKeyChangeProgressListener = new HashSet<OnKeyChangeProgressListener>();
private Set<OnKeyEventListener> mOnKeyEventListener = new HashSet<OnKeyEventListener>();
private Set<OnSetSoundListener> mOnSetSoundListener = new HashSet<OnSetSoundListener>();
private Set<OnRegisterDLNAListener> mOnRegisterDLNAListener = new HashSet<OnRegisterDLNAListener>();
private Set<OnDLNALinearLayoutListener> mOnDLNALinearLayoutListener = new HashSet<OnDLNALinearLayoutListener>();
private Set<OnScreenOrientationSwitchListener> mOnScreenOrientationSwitchListener = new HashSet<OnScreenOrientationSwitchListener>();
private Set<OnClickRetryListener> mOnClickRetryListener = new HashSet<OnClickRetryListener>();
private Set<OnVideoUIRefreshListener> mOnVideoUIRefreshListener = new HashSet<OnVideoUIRefreshListener>();
private Set<OnShowHideADContainerListener> mOnShowHideADContainerListener = new HashSet<OnShowHideADContainerListener>();
/**
* 通过主线程回调消息到界面
*/
private Handler mMainHandler;
public VDVideoViewListeners(Context context) {
mMainHandler = new Handler();
mContext = context;
}
public void addOnVideoUIRefreshListener(OnVideoUIRefreshListener l) {
mOnVideoUIRefreshListener.add(l);
}
public void removeOnVideoUIRefreshListener(OnVideoUIRefreshListener l) {
mOnVideoUIRefreshListener.remove(l);
}
void addOnBufferingUpdateListener(OnBufferingUpdateListener l) {
mOnBufferingUpdateListener.add(l);
}
void removeOnBufferingUpdateListener(OnBufferingUpdateListener l) {
mOnBufferingUpdateListener.remove(l);
}
void addOnCompletionListener(OnCompletionListener l) {
mOnCompletionListener.add(l);
}
void removeOnCompletionListener(OnCompletionListener l) {
mOnCompletionListener.remove(l);
}
void addOnErrorListener(OnErrorListener l) {
mOnErrorListener.add(l);
}
void removeOnErrorListener(OnErrorListener l) {
mOnErrorListener.remove(l);
}
void addOnRetryErrorListener(OnErrorListener l) {
mOnRetryErrorListener.add(l);
}
void removeOnRetryErrorListener(OnErrorListener l) {
mOnRetryErrorListener.remove(l);
}
void addOnInfoListener(OnInfoListener l) {
mOnInfoListener.add(l);
}
void removeOnInfoListener(OnInfoListener l) {
mOnInfoListener.remove(l);
}
void addOnPreparedListener(OnPreparedListener l) {
mOnPreparedListener.add(l);
}
void removeOnPreparedListener(OnPreparedListener l) {
mOnPreparedListener.remove(l);
}
void addOnSeekCompleteListener(OnSeekCompleteListener l) {
mOnSeekCompleteListener.add(l);
}
void removeOnSeekCompleteListener(OnSeekCompleteListener l) {
mOnSeekCompleteListener.remove(l);
}
void addOnVideoSizeChangedListener(OnVideoSizeChangedListener l) {
mOnVideoSizeChangedListener.add(l);
}
void removeOnVideoSizeChangedListener(OnVideoSizeChangedListener l) {
mOnVideoSizeChangedListener.remove(l);
}
void addOnVideoOpenedListener(OnVideoOpenedListener l) {
mOnVideoOpenedListener.add(l);
}
void removeOnVideoOpenedListener(OnVideoOpenedListener l) {
mOnVideoOpenedListener.remove(l);
}
public void addOnProgressUpdateListener(OnProgressUpdateListener l) {
mOnProgressUpdateListener.add(l);
}
public void removeOnProgressUpdateListener(OnProgressUpdateListener l) {
mOnProgressUpdateListener.remove(l);
}
public void addOnTimedTextListener(OnTimedTextListener l) {
mOnTimedTextListener.add(l);
}
public void removeOnTimedTextListener(OnTimedTextListener l) {
mOnTimedTextListener.remove(l);
}
public void addOnPlayVideoListener(OnPlayVideoListener l) {
mOnPlayVideoListener.add(l);
}
public void removeOnPlayVideoListener(OnPlayVideoListener l) {
mOnPlayVideoListener.remove(l);
}
// public void addOnLiveVideoListener(OnLiveVideoListener l) {
// mOnLiveVideoListener.add(l);
// }
//
// public void removeOnLiveVideoListener(OnLiveVideoListener l) {
// mOnLiveVideoListener.remove(l);
// }
public void addOnFullScreenListener(OnFullScreenListener l) {
mOnFullScreenListener.add(l);
}
public void removeOnFullScreenListener(OnFullScreenListener l) {
mOnFullScreenListener.remove(l);
}
public void addOnSoundChangedListener(OnSoundChangedListener l) {
mOnSoundChangedListener.add(l);
}
public void removeOnSoundChangedListener(OnSoundChangedListener l) {
mOnSoundChangedListener.remove(l);
}
public void addOnScreenTouchListener(OnScreenTouchListener l) {
mOnScreenTouchListener.add(l);
}
public void removeOnScreenTouchListener(OnScreenTouchListener l) {
mOnScreenTouchListener.remove(l);
}
public void addOnResolutionListener(OnResolutionListener l) {
mOnResolutionListener.add(l);
}
public void removeOnResolutionListener(OnResolutionListener l) {
mOnResolutionListener.remove(l);
}
public void addOnResolutionContainerListener(OnResolutionContainerListener l) {
mOnResolutionContainerListener.add(l);
}
public void removeOnResolutionContainerListener(
OnResolutionContainerListener l) {
mOnResolutionContainerListener.remove(l);
}
public void addOnResolutionListButtonListener(
OnResolutionListButtonListener l) {
mOnResolutionListButtonListener.add(l);
}
public void removeOnResolutionListButtonListener(
OnResolutionListButtonListener l) {
mOnResolutionListButtonListener.remove(l);
}
public void addOnTipListener(OnTipListener l) {
mOnTipListener.add(l);
}
public void removeOnTipListener(OnTipListener l) {
mOnTipListener.remove(l);
}
public void addOnLightingChangeListener(OnLightingChangeListener l) {
mOnLightingChangeListener.add(l);
}
public void removeOnLightingChangeListener(OnLightingChangeListener l) {
mOnLightingChangeListener.remove(l);
}
public void addOnLightingVisibleListener(OnLightingVisibleListener l) {
mOnLightingVisibleListener.add(l);
}
public void removeOnLightingVisibleListener(OnLightingVisibleListener l) {
mOnLightingVisibleListener.remove(l);
}
public void addOnSoundVisibleListener(OnSoundVisibleListener l) {
mOnSoundVisibleListener.add(l);
}
public void removeOnSoundVisibleListener(OnSoundVisibleListener l) {
mOnSoundVisibleListener.remove(l);
}
public void addOnVMSResolutionListener(OnVMSResolutionListener l) {
mOnVMSResolutionListener.add(l);
}
public void removeOnVMSResolutionListener(OnVMSResolutionListener l) {
mOnVMSResolutionListener.remove(l);
}
public void addOnLoadingListener(OnLoadingListener l) {
mOnLoadingListener.add(l);
}
public void removeOnLoadingListener(OnLoadingListener l) {
mOnLoadingListener.remove(l);
}
public void addOnVideoGuideTipsListener(OnVideoGuideTipsListener l) {
mOnVideoGuideTipsListener.add(l);
}
public void removeOnVideoGuideTipsListener(OnVideoGuideTipsListener l) {
mOnVideoGuideTipsListener.remove(l);
}
public void addOnPauseListener(OnPauseListener l) {
mOnPauseListener.add(l);
}
public void removeOnPauseListener(OnPauseListener l) {
mOnPauseListener.remove(l);
}
public void addOnVideoADListener(OnVideoFrameADListener l) {
mOnVideoFrameADListener.add(l);
}
public void removeOnVideoAdListener(OnVideoFrameADListener l) {
mOnVideoFrameADListener.remove(l);
}
public void addOnVideoInsertADListener(OnVideoInsertADListener l) {
mOnVideoInsertADListener.add(l);
}
public void removeOnVideoInsertADListener(OnVideoInsertADListener l) {
mOnVideoInsertADListener.remove(l);
}
public void addOnVideoListListener(OnVideoListListener l) {
mOnVideoListListener.add(l);
}
public void removeOnVideoListListener(OnVideoListListener l) {
mOnVideoListListener.remove(l);
}
public void addOnVideoListVisibleChangeListener(
OnVideoListVisibleChangeListener l) {
mOnVideoListVisibleChangeListener.add(l);
}
public void removeOnVideoListVisibleChangeListener(
OnVideoListVisibleChangeListener l) {
mOnVideoListVisibleChangeListener.remove(l);
}
public void addOnMoreOprationVisibleChangeListener(
OnMoreOprationVisibleChangeListener l) {
mOnMoreOprationVisibleChangeListener.add(l);
}
public void removeOnMoreOprationVisibleChangeListener(
OnMoreOprationVisibleChangeListener l) {
mOnMoreOprationVisibleChangeListener.remove(l);
}
// public void
// addOnResolutionVisibleChangeListener(OnResolutionVisibleChangeListener l)
// {
// mOnResolutionVisibleChangeListener.add(l);
// }
//
// public void
// removeOnResolutionVisibleChangeListener(OnResolutionVisibleChangeListener
// l) {
// mOnResolutionVisibleChangeListener.remove(l);
// }
public void addOnVideoDoubleTapListener(OnVideoDoubleTapListener l) {
mOnVideoDoubleTapListener.add(l);
}
public void removeOnVideoDoubleTapListener(OnVideoDoubleTapListener l) {
mOnVideoDoubleTapListener.remove(l);
}
public void addOnDecodingTypeListener(OnDecodingTypeListener l) {
mOnDecodingTypeListener.add(l);
}
public void removeOnDecodingTypeListener(OnDecodingTypeListener l) {
mOnDecodingTypeListener.remove(l);
}
public void addOnScreenOrientationChangeListener(
OnScreenOrientationChangeListener l) {
mOnScreenOrientationChangeListener.add(l);
}
public void removeOnScreenOrientationChangeListener(
OnScreenOrientationChangeListener l) {
mOnScreenOrientationChangeListener.remove(l);
}
public void addOnShowHideControllerListener(OnShowHideControllerListener l) {
mOnShowHideControllerListener.add(l);
}
public void removeOnShowHideControllerListener(
OnShowHideControllerListener l) {
mOnShowHideControllerListener.remove(l);
}
public void addOnShowHideBottomControllerListener(
OnShowHideBottomControllerListener l) {
mOnShowHideBottomControllerListener.add(l);
}
public void removeOnShowHideBottomControllerListener(
OnShowHideBottomControllerListener l) {
mOnShowHideBottomControllerListener.remove(l);
}
public void addOnShowHideTopContainerListener(
OnShowHideTopContainerListener l) {
mOnShowHideTopContainerListener.add(l);
}
public void removeOnShowHideTopContainerListener(
OnShowHideTopContainerListener l) {
mOnShowHideTopContainerListener.remove(l);
}
public void addOnProgressViewVisibleListener(OnProgressViewVisibleListener l) {
mOnProgressViewVisibleListener.add(l);
}
public void removeOnProgressViewVisibleListener(
OnProgressViewVisibleListener l) {
mOnProgressViewVisibleListener.remove(l);
}
public void addOnClickPlayListener(OnClickPlayListener l) {
mOnClickPlayListener.add(l);
}
public void removeOnClickPlayListener(OnClickPlayListener l) {
mOnClickPlayListener.remove(l);
}
public void addOnKeyChangeProgressListener(OnKeyChangeProgressListener l) {
mOnKeyChangeProgressListener.add(l);
}
public void removeOnKeyChangeProgressListener(OnKeyChangeProgressListener l) {
mOnKeyChangeProgressListener.remove(l);
}
public void addOnKeyEventListener(OnKeyEventListener l) {
mOnKeyEventListener.add(l);
}
public void removeOnKeyEventListener(OnKeyEventListener l) {
mOnKeyEventListener.remove(l);
}
public void addOnSetSoundListener(OnSetSoundListener l) {
mOnSetSoundListener.add(l);
}
public void removeOnSetSoundListener(OnSetSoundListener l) {
mOnSetSoundListener.remove(l);
}
public void addOnRegisterDLNAListener(OnRegisterDLNAListener l) {
mOnRegisterDLNAListener.add(l);
}
public void removeOnRegisterDLNAListener(OnRegisterDLNAListener l) {
mOnRegisterDLNAListener.remove(l);
}
public void addOnDLNALinearLayoutListener(OnDLNALinearLayoutListener l) {
mOnDLNALinearLayoutListener.add(l);
}
public void removeOnDLNALinearLayoutListener(OnDLNALinearLayoutListener l) {
mOnDLNALinearLayoutListener.remove(l);
}
public void addOnScreenOrientationSwitchListener(
OnScreenOrientationSwitchListener l) {
mOnScreenOrientationSwitchListener.add(l);
}
public void removeOnScreenOrientationSwitchListener(
OnScreenOrientationSwitchListener l) {
mOnScreenOrientationSwitchListener.remove(l);
}
public void addOnClickRetryListener(OnClickRetryListener l) {
mOnClickRetryListener.add(l);
}
public void removeOnClickRetryListener(OnClickRetryListener l) {
mOnClickRetryListener.remove(l);
}
public void addOnShowHideADContainerListener(OnShowHideADContainerListener l) {
mOnShowHideADContainerListener.add(l);
}
public void removeOnShowHideADContainerListener(
OnShowHideADContainerListener l) {
mOnShowHideADContainerListener.remove(l);
}
public void notifyVideoUIRefreshListener() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoUIRefreshListener listener : mOnVideoUIRefreshListener) {
listener.onVideoUIRefresh();
}
}
});
}
public void notifyClickRetry() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnClickRetryListener listener : mOnClickRetryListener) {
listener.onClickRetry();
}
}
});
}
void notifyBufferingUpdate(final int percent) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnBufferingUpdateListener listener : mOnBufferingUpdateListener) {
listener.onBufferingUpdate(percent);
}
}
});
}
void notifyCompletion() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnCompletionListener listener : mOnCompletionListener) {
listener.onCompletion();
}
}
});
}
void notifyError(final int what, final int extra) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnErrorListener listener : mOnErrorListener) {
listener.onError(what, extra);
}
}
});
}
void notifyRetryError(final int what, final int extra) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnErrorListener listener : mOnRetryErrorListener) {
listener.onError(what, extra);
}
}
});
}
void notifyInfo(final int what, final int extra) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnInfoListener listener : mOnInfoListener) {
listener.onInfo(what, extra);
}
}
});
}
void notifyPrepared() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnPreparedListener listener : mOnPreparedListener) {
listener.onPrepared();
}
}
});
}
void notifySeekComplete() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnSeekCompleteListener listener : mOnSeekCompleteListener) {
listener.onSeekComplete();
}
}
});
}
void notifyVideoSizeChanged(final int width, final int height) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoSizeChangedListener listener : mOnVideoSizeChangedListener) {
listener.onVideoSizeChanged(width, height);
}
}
});
}
void notifyTimedText(final TimedText text) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnTimedTextListener listener : mOnTimedTextListener) {
listener.onTimedText(text);
}
}
});
}
void notifyProgressUpdate(final long current, final long duration) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnProgressUpdateListener listener : mOnProgressUpdateListener) {
listener.onProgressUpdate(current, duration);
}
}
});
}
void notifyVideoOpened() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoOpenedListener listener : mOnVideoOpenedListener) {
listener.onVideoOpened();
}
}
});
}
void notifyVideoInfo(final VDVideoInfo info) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnPlayVideoListener listener : mOnPlayVideoListener) {
listener.onVideoInfo(info);
}
}
});
}
void notifyShowLoading(final boolean show) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnPlayVideoListener listener : mOnPlayVideoListener) {
listener.onShowLoading(show);
}
}
});
}
void notifyVideoPrepared(final boolean prepare) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnPlayVideoListener listener : mOnPlayVideoListener) {
listener.onVideoPrepared(prepare);
}
}
});
}
void notifyPlayStateChanged() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnPlayVideoListener listener : mOnPlayVideoListener) {
listener.onPlayStateChanged();
}
}
});
}
void notifyDragTo(final long position, final long duration) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnProgressUpdateListener listener : mOnProgressUpdateListener) {
listener.onDragProgess(position, duration);
}
}
});
}
// void notifyResolutionInfo(final VDResolutionData list) {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnLiveVideoListener listener : mOnLiveVideoListener) {
// listener.onResolutionInfo(list);
// }
// }
// });
// }
//
// void notifyResolutionIndex(final int index) {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnLiveVideoListener listener : mOnLiveVideoListener) {
// listener.onResolutionIndex(index);
// }
// }
// });
// }
void notifyFullScreen(final boolean isFullScreen, final boolean isFromHand) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnFullScreenListener listener : mOnFullScreenListener) {
listener.onFullScreen(isFullScreen, isFromHand);
}
}
});
}
void notifySoundChanged(final int curr) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnSoundChangedListener listener : mOnSoundChangedListener) {
listener.onSoundChanged(curr);
}
}
});
}
void notifyScreenSingleTouch(final MotionEvent ev,
final eSingleTouchListener flag) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
switch (flag) {
case eTouchListenerSingleTouchStart:
break;
case eTouchListenerSingleTouch:
for (OnScreenTouchListener listener : mOnScreenTouchListener) {
listener.onSingleTouch(ev);
}
break;
case eTouchListenerSingleTouchEnd:
break;
default:
break;
}
}
});
}
void notifyKeyEvent() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnKeyEventListener listener : mOnKeyEventListener) {
listener.onKeyEvent();
}
}
});
}
void notifyKeyLeftRightEvent() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnKeyEventListener listener : mOnKeyEventListener) {
listener.onKeyLeftRight();
}
}
});
}
void notifyScreenDoubleTouch(final MotionEvent ev,
final eDoubleTouchListener flag) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
switch (flag) {
case eTouchListenerDoubleTouchStart:
break;
case eTouchListenerDoubleTouch:
Log.i("VDVideoDoubleTapPlayView",
"eTouchListenerDoubleTouch");
notifyDoubleTouch();
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller != null) {
if (controller.getIsPlaying()) {
controller.pause();
} else {
controller.resume();
controller.start();
}
}
break;
case eTouchListenerDoubleTouchEnd:
break;
default:
break;
}
}
});
}
/**
* 使用坐标点方式得到[0-1]的滑动取值范围
*
* @param point1
* @param point2
* @return
*/
private float getCurrTimeFromEvent(final PointF point1, final PointF point2) {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller == null) {
return 0;
}
DLNAController dlnaController = DLNAController.getInstance(controller
.getContext());
int width = controller.getScreen()[0];
int distance = (int) (point2.x - point1.x);
long current = 0;
long duration = 0;
if (DLNAController.mIsDLNA) {
current = DLNAController.mTmpPosition;
duration = dlnaController.mDuration;
} else {
current = controller.mVDPlayerInfo.mCurrent;
duration = controller.mVDPlayerInfo.mDuration;
}
// LogS.i(TAG, "getCurrTimeFromEvent point1.x:" + point1.x +
// ", point2.x:" + point2.x + ",distance:" + distance);
float rate = DLNAController.mIsDLNA ? dlnaController.mProgressRate
: controller.mProgressRate;
float ret = ((float) current / duration)
+ ((float) distance / (float) width) * rate;
VDLog.e(TAG, "current : " + current + ",duration : " + duration
+ " , distance : " + distance + " , ret = " + ret);
if (ret < 0) {
ret = 0;
} else if (ret > 1) {
ret = 1;
}
return ret;
}
private float mTmpStreamLevel;
private int mScreenHeight;
/**
* 使用坐标点方式得到[0-1]的音量取值范围
*
* @param point1
* @param point2
* @return
*/
private float getCurrSoundFromEvent(final PointF point1,
final PointF point2, float distansY) {
// float y1 = point1.y;
// float y2 = point2.y;
// float distance = y1 - y2;
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller == null) {
return 0;
}
int maxVolume;
int currVolume;
if (DLNAController.mIsDLNA) {
maxVolume = DLNAController.getInstance(controller.getContext())
.getVolumeMax();
currVolume = DLNAController.getInstance(controller.getContext()).mVolume;
} else {
// Context context = VDApplication.getInstance().getContext();
maxVolume = VDPlayerSoundManager.getMaxSoundVolume(controller
.getContext());
currVolume = VDPlayerSoundManager.getCurrSoundVolume(controller
.getContext());
}
// float degressVolume = (float) currVolume / maxVolume;
float degree = (float) distansY / mScreenHeight;
// float currVolumef = currVolume;
// currVolumef += degree*maxVolume;
// currVolumef = currVolume/maxVolume;
Log.i("getCurrSoundFromEvent", "fromDownY = " + distansY
+ " , currVolume = " + currVolume + " , degree = "
+ degree + " , tmp_stream_level = " + mTmpStreamLevel
+ " , maxVolume = " + maxVolume);
mTmpStreamLevel += (degree * maxVolume);
// float ret = (currVolume+degree*maxVolume);
Log.d("getCurrSoundFromEvent", "tmp_stream_level = " + mTmpStreamLevel
+ " , add = " + (degree * maxVolume));
// Log.i("getCurrSoundFromEvent", "fromDownY = " + distansY +
// " , ret = " + ret + " , degree = " + degree + " , y1 = " + y1 +
// " , cur = " + (currVolume+degree*maxVolume));
// float ret = degressVolume + degree * 3;
if (mTmpStreamLevel < 0) {
mTmpStreamLevel = 0;
} else if (mTmpStreamLevel > maxVolume) {
mTmpStreamLevel = maxVolume;
}
return mTmpStreamLevel;
}
/**
* 使用坐标点方式得到[0-1]的亮度取值范围
*
* @param point1
* @param point2
* @return
*/
private float getCurrLightingFromEvent(final PointF point1,
final PointF point2) {
float ret;
float y1 = point1.y;
float y2 = point2.y;
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller == null) {
ret = 1.0f;
return ret;
}
int height = controller.getScreen()[1];
float distance = y1 - y2;
float degree = distance / height;
float curNum = controller.getCurrLightingSetting();
ret = curNum + degree;
if (ret >= 1.0) {
ret = 1.0f;
} else if (ret <= 0.1) { // 不能小于10%亮度
ret = 0.1f;
}
return ret;
}
private boolean getIsRight(final PointF point) {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (null == controller) {
return false;
}
int width = controller.getScreen()[0];
boolean isRight = false;
if (point.x > ((float) width / 2)) {
// 在右边屏幕位置
isRight = true;
}
return isRight;
}
void notifyScreenVerticalScrollTouch(final PointF point1,
final PointF point2, final PointF beginPoint,
final eVerticalScrollTouchListener flag, final float distansY) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
float curr = -1;
boolean isSoundVisible = false;
if (flag == eVerticalScrollTouchListener.eTouchListenerVerticalScrollEnd) {
curr = 0;
} else if (flag == eVerticalScrollTouchListener.eTouchListenerVerticalScrollSound
|| getIsRight(beginPoint)) {
// 调节音量
isSoundVisible = true;
if (point1 != null && point2 != null) {
curr = getCurrSoundFromEvent(beginPoint, point2,
distansY);
}
} else if (flag == eVerticalScrollTouchListener.eTouchListenerVerticalScrollLighting
|| !getIsRight(beginPoint)) {
// 调节亮度··
isSoundVisible = false;
if (point1 != null && point2 != null) {
curr = getCurrLightingFromEvent(point1, point2);
VDLog.e("xxoo", curr + "");
}
}
if (curr == -1) {
return;
}
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
switch (flag) {
case eTouchListenerVerticalScrollStart:
if (!VDVideoFullModeController.getInstance()
.getIsFullScreen()) {
isSoundVisible = true;
}
if (isSoundVisible) {
notifySoundVisible(true);
if (DLNAController.mIsDLNA) {
if (controller != null)
mTmpStreamLevel = DLNAController
.getInstance(controller.getContext()).mVolume;
} else {
Context context = VDApplication.getInstance()
.getContext();
mTmpStreamLevel = VDPlayerSoundManager
.getCurrSoundVolume(context);
}
VDLog.e("getCurrSoundFromEvent", "tmp_stream_level = "
+ mTmpStreamLevel);
} else {
notifyLightingVisible(true);
}
if (controller != null)
mScreenHeight = controller.getScreen()[1];
break;
case eTouchListenerVerticalScrollLighting:
if (controller != null)
controller.dragLightingTo(curr, true);
break;
case eTouchListenerVerticalScrollSound:
if (controller != null)
controller.dragSoundSeekTo((int) curr);
break;
case eTouchListenerVerticalScroll:
// 判断是右边还是左边?
if (getIsRight(beginPoint)) {
VDLog.e(TAG,
"notifyScreenVerticalScrollTouch,eTouchListenerVerticalScroll,dragSoundSeekTo curr : "
+ curr);
if (controller != null)
controller.dragSoundSeekTo((int) curr);
} else {
VDLog.e(TAG,
"notifyScreenVerticalScrollTouch,eTouchListenerVerticalScroll,dragLightingTo");
if (controller != null)
controller.dragLightingTo(curr, true);
}
break;
case eTouchListenerVerticalScrollEnd:
notifySoundVisible(false);
notifyLightingVisible(false);
break;
default:
break;
}
}
});
}
void notifyScreenHorizonScrollTouch(final PointF point1,
final PointF point2, final PointF beginPoint,
final eHorizonScrollTouchListener flag) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// for (OnScreenTouchListener listener : mOnScreenTouchListener)
// {
// }
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
VDVideoInfo videoInfo = null;
if (controller != null) {
VDPlayerInfo playerInfo = controller.getPlayerInfo();
if (!playerInfo.isCanScroll()) {
return;
}
videoInfo = controller.getCurrentVideo();
}
if (videoInfo == null) {
return;
}
switch (flag) {
case eTouchListenerHorizonScrollStart:
if (!videoInfo.mIsLive) {
notifyProgressViewVisible(true);
if (DLNAController.mIsDLNA) {
DLNAController dlnaController = DLNAController
.getInstance(controller.getContext());
DLNAController.mTmpPosition = dlnaController.mPosition;
} else {
controller.mVDPlayerInfo.mCurrent = controller
.getCurrentPosition();
controller.setProgressRate();
controller.pause();
}
}
break;
case eTouchListenerHorizonScroll:
if (!videoInfo.mIsLive) {
float curr = getCurrTimeFromEvent(point1, point2);
controller.dragProgressTo(curr);
}
break;
case eTouchListenerHorizonScrollEnd:
if (!videoInfo.mIsLive) {
notifyProgressViewVisible(false);
controller.dragProgressTo(
getCurrTimeFromEvent(point1, point2), true,
true);
if (DLNAController.mIsDLNA) {
} else {
controller.resume(false);
ISinaVideoView vv = controller.getVideoView();
if (!vv.isPlaying()) {
controller.start();
}
}
notifyPlayStateChanged();
}
break;
default:
break;
}
}
});
}
public void notifyLightingSetting(final float curr) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnLightingChangeListener listener : mOnLightingChangeListener) {
listener.onLightingChange(curr);
}
}
});
}
/**
* 清晰度改变通知
*
* @param tag
*/
public void notifyResolutionChanged(final String tag) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnResolutionListener listener : mOnResolutionListener) {
listener.onResolutionChanged(tag);
}
}
});
}
/**
* 清晰度解析完毕的时候回传
*
* @param list
*/
public void notifyResolutionParsed(final VDResolutionData list) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnResolutionListener listener : mOnResolutionListener) {
listener.onResolutionParsed(list);
}
}
});
}
/**
* 通知清晰度选择
*
* @param mCurResolution
*/
// void notifyResolutionSelect(final String mCurResolution) {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnResolutionListener listener : mOnResolutionListener) {
// listener.onResolutionSelect(mCurResolution);
// }
// }
// });
// }
/**
* 通知清晰度解析完成
*
* @param list
*/
// void notifyResolutionParsed(final VDResolutionData list) {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnResolutionListener listener : mOnResolutionListener) {
// listener.onParseResolution(list);
// }
// }
// });
// }
/**
* 隐藏清晰度按钮
*
*/
// void notifyHideResolution() {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnResolutionListener listener : mOnResolutionListener) {
// listener.hideResolution();
// }
// }
// });
// }
public void notifyResolutionVisible(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnResolutionContainerListener listener : mOnResolutionContainerListener) {
listener.onResolutionContainerVisible(isVisible);
}
}
});
}
public void notifyResolutionListButtonFirstFocus() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnResolutionListButtonListener l : mOnResolutionListButtonListener) {
l.onResolutionListButtonFocusFirst();
}
}
});
}
void notifyTip(final String tip) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnTipListener listener : mOnTipListener) {
listener.onTip(tip);
}
}
});
}
void notifyTip(final int tip) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnTipListener listener : mOnTipListener) {
listener.onTip(tip);
}
}
});
}
void notifyHideTip() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnTipListener listener : mOnTipListener) {
listener.hideTip();
}
}
});
}
void notifyLightingVisible(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnLightingVisibleListener listener : mOnLightingVisibleListener) {
listener.onLightingVisible(isVisible);
}
}
});
}
void notifySoundVisible(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnSoundVisibleListener listener : mOnSoundVisibleListener) {
listener.onSoundVisible(isVisible);
}
}
});
}
void notifySoundSeekBarVisible(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnSoundVisibleListener listener : mOnSoundVisibleListener) {
listener.onSoundSeekBarVisible(isVisible);
}
}
});
}
void notifyVMSResolutionContainerVisible(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVMSResolutionListener listener : mOnVMSResolutionListener) {
listener.onVMSResolutionContainerVisible(isVisible);
}
}
});
}
void notifyVMSResolutionChanged() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVMSResolutionListener listener : mOnVMSResolutionListener) {
listener.onVMSResolutionChanged();
}
}
});
}
void notifyShowLoading() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnLoadingListener listener : mOnLoadingListener) {
listener.showLoading();
}
}
});
}
void notifyHideLoading() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnLoadingListener listener : mOnLoadingListener) {
listener.hideLoading();
}
}
});
}
void notifyGuideTips(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoGuideTipsListener listener : mOnVideoGuideTipsListener) {
listener.onVisible(isVisible);
}
}
});
}
void notifyPlayOrPause() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnClickPlayListener listener : mOnClickPlayListener) {
listener.onClickPlay();
}
}
});
}
void notifyPause() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnPauseListener listener : mOnPauseListener) {
listener.onPause();
}
}
});
}
void notifyVideoInsertADBegin() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoInsertADListener listener : mOnVideoInsertADListener) {
listener.onVideoInsertADBegin();
}
}
});
}
void notifyVideoInsertADTicker() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoInsertADListener listener : mOnVideoInsertADListener) {
listener.onVideoInsertADTicker();
}
}
});
}
void notifyVideoInsertADEnd() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoInsertADListener listener : mOnVideoInsertADListener) {
listener.onVideoInsertADEnd();
}
}
});
}
void notifyVideoFrameADBegin() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoFrameADListener listener : mOnVideoFrameADListener) {
listener.onVideoFrameADBegin();
}
}
});
}
void notifyVideoFrameADEnd() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoFrameADListener listener : mOnVideoFrameADListener) {
listener.onVideoFrameADEnd();
}
}
});
}
// /**
// * @deprecated 不再用了,用这个:notifyVideoList(final VDVideoListInfo videoList)
// * @param videoList
// */
// void notifyVideoList(final ArrayList<VDVideoInfo> videoList) {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnVideoListListener listener : mOnVideoListListener) {
// listener.onVideoList(videoList);
// }
// }
// });
// }
void notifyVideoList(final VDVideoListInfo videoList) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoListListener listener : mOnVideoListListener) {
listener.onVideoList(videoList);
}
}
});
}
void notifyVideoListVisibelChange() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoListVisibleChangeListener listener : mOnVideoListVisibleChangeListener) {
listener.toogle();
}
}
});
}
void notifyShowVideoList() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoListVisibleChangeListener listener : mOnVideoListVisibleChangeListener) {
listener.showPlayList();
}
}
});
}
void notifyHideVideoList() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoListVisibleChangeListener listener : mOnVideoListVisibleChangeListener) {
listener.hidePlayList();
}
}
});
}
void removeAndHideDelayVideoList() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoListVisibleChangeListener listener : mOnVideoListVisibleChangeListener) {
listener.removeAndHideDelay();
}
}
});
}
void notifyShowMoreOprationPanel() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnMoreOprationVisibleChangeListener listener : mOnMoreOprationVisibleChangeListener) {
listener.showPanel();
}
}
});
}
void notifyHideMoreOprationPanel() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnMoreOprationVisibleChangeListener listener : mOnMoreOprationVisibleChangeListener) {
listener.hidePanel();
}
}
});
}
void notifyRemoveAndHideDelayMoreOprationPanel() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnMoreOprationVisibleChangeListener listener : mOnMoreOprationVisibleChangeListener) {
listener.removeAndHideDelay();
}
}
});
}
// void notifyResolutionListVisibelChange() {
// mMainHandler.post(new Runnable() {
//
// @Override
// public void run() {
// for (OnResolutionVisibleChangeListener listener :
// mOnResolutionVisibleChangeListener) {
// listener.toogle();
// }
// }
// });
// }
void notifyDoubleTouch() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnVideoDoubleTapListener listener : mOnVideoDoubleTapListener) {
listener.onDoubleTouch();
}
}
});
}
public void notifyDecodingTypeChange(final boolean isFFMpeg) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnDecodingTypeListener l : mOnDecodingTypeListener) {
l.onChange(isFFMpeg);
}
}
});
}
public void notifyScreenOrientationChange(final boolean vertical) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnScreenOrientationChangeListener l : mOnScreenOrientationChangeListener) {
VDLog.e(VDVideoFullModeController.TAG,
": OnScreenOrientationChange");
if (vertical) {
l.onScreenOrientationVertical();
} else {
l.onScreenOrientationHorizontal();
}
}
}
});
}
public void notifyNotHideControllerBar() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.doNotHideControllerBar();
}
}
});
}
public void notifyHideBottomControllerBar() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideBottomControllerListener l : mOnShowHideBottomControllerListener) {
l.hideBottomControllerBar();
}
}
});
}
public void notifyShowBottomControllerBar() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideBottomControllerListener l : mOnShowHideBottomControllerListener) {
l.showBottomControllerBar();
}
}
});
}
public void notifyHideTopControllerBar() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnShowHideTopContainerListener l : mOnShowHideTopContainerListener) {
l.hideTopControllerBar();
}
}
});
}
public void notifyShowTopControllerBar() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnShowHideTopContainerListener l : mOnShowHideTopContainerListener) {
l.showTopControllerBar();
}
}
});
}
public void notifyControllerBarPreHide() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.onPreHide();
}
}
});
}
public void notifyControllerBarPreShow() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.onPreShow();
}
}
});
}
public void notifyControllerBarPostHide() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.onPostHide();
}
}
});
}
public void notifyControllerBarPostShow() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.onPostShow();
}
}
});
}
public void notifyKeyChangeProgress(final boolean keyLeft) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
VDVideoInfo videoInfo = null;
if (controller != null)
videoInfo = controller.getCurrentVideo();
if (videoInfo != null && !videoInfo.mIsLive) {
notifyProgressViewVisible(false);
long current = controller.mVDPlayerInfo.mCurrent;
long duration = controller.mVDPlayerInfo.mDuration;
current += keyLeft ? (-7000) : 7000;
float ret = ((float) current / duration);// ((float)
// distance /
// width) *
// controller.mProgressRate;
VDLog.e(TAG, "current : " + current + ",duration : "
+ duration + " , ret = " + ret);
if (ret < 0) {
ret = 0;
} else if (ret > 1) {
ret = 1;
}
controller.dragProgressTo(ret, true, false);
controller.resume(false);
ISinaVideoView vv = controller.getVideoView();
if (!vv.isPlaying()) {
controller.start();
}
notifyPlayStateChanged();
notifyPlayStateChanged();
}
for (OnKeyChangeProgressListener l : mOnKeyChangeProgressListener) {
l.onKeyDown(keyLeft);
}
}
});
}
public void notifyHideControllerBar(final long delay) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.hideControllerBar(delay);
}
}
});
}
public void notifyShowControllerBar(final boolean delayHide) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnShowHideControllerListener l : mOnShowHideControllerListener) {
l.showControllerBar(delayHide);
}
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller != null)
controller.hideStatusBar(!delayHide);
}
});
}
void notifyProgressViewVisible(final boolean isVisible) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnProgressViewVisibleListener listener : mOnProgressViewVisibleListener) {
listener.onProgressVisible(isVisible);
}
}
});
}
// DLNA
public void notifySetCurVolume(final int currVolume) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnSetSoundListener l : mOnSetSoundListener) {
l.onSetCurVolume(currVolume);
}
}
});
}
public void notifySetMaxVolume(final int maxVolume) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnSetSoundListener l : mOnSetSoundListener) {
l.onSetMaxVolume(maxVolume);
}
}
});
}
public void notifyRegisterDLNAListener() {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnRegisterDLNAListener l : mOnRegisterDLNAListener) {
l.register();
}
}
});
}
void notifySetDLNALayoutVisible(final boolean visiable) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnDLNALinearLayoutListener listener : mOnDLNALinearLayoutListener) {
listener.setLayoutVisiable(visiable);
}
}
});
}
void notifyScreenOrientationSwitch(final boolean fullScreen) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
for (OnScreenOrientationSwitchListener listener : mOnScreenOrientationSwitchListener) {
listener.onScreenOrientationSwitch(fullScreen);
}
}
});
}
public void notifyOnShowHideADContainer(final boolean isShow) {
mMainHandler.post(new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
for (OnShowHideADContainerListener l : mOnShowHideADContainerListener) {
if (isShow) {
l.showADContainerBar();
} else {
l.hideADContainerBar();
}
}
}
});
}
public void clear() {
mOnBufferingUpdateListener.clear();
mOnCompletionListener.clear();
mOnErrorListener.clear();
mOnRetryErrorListener.clear();
mOnInfoListener.clear();
mOnPreparedListener.clear();
mOnSeekCompleteListener.clear();
mOnVideoSizeChangedListener.clear();
mOnVideoOpenedListener.clear();
mOnTimedTextListener.clear();
mOnProgressUpdateListener.clear();
mOnPlayVideoListener.clear();
// mOnLiveVideoListener.clear();
mOnFullScreenListener.clear();
mOnSoundChangedListener.clear();
mOnLightingChangeListener.clear();
mOnScreenTouchListener.clear();
mOnResolutionListener.clear();
mOnResolutionContainerListener.clear();
mOnTipListener.clear();
mOnLightingVisibleListener.clear();
mOnLoadingListener.clear();
mOnSoundVisibleListener.clear();
mOnVMSResolutionListener.clear();
mOnVideoGuideTipsListener.clear();
mOnPauseListener.clear();
mOnVideoFrameADListener.clear();
mOnVideoInsertADListener.clear();
mOnVideoListListener.clear();
mOnVideoListVisibleChangeListener.clear();
// mOnResolutionVisibleChangeListener.clear();
mOnVideoDoubleTapListener.clear();
mOnDecodingTypeListener.clear();
mOnScreenOrientationChangeListener.clear();
mOnShowHideControllerListener.clear();
mOnSetSoundListener.clear();
mOnDLNALinearLayoutListener.clear();
mOnScreenOrientationSwitchListener.clear();
mOnClickRetryListener.clear();
mOnVideoUIRefreshListener.clear();
mOnShowHideADContainerListener.clear();
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/data/VDResolutionData.java
package com.sina.sinavideo.sdk.data;
import java.util.ArrayList;
import java.util.HashMap;
import java.util.Iterator;
import java.util.LinkedHashMap;
import java.util.List;
import java.util.Map;
/**
* 清晰度的类 TODO 这个类,后期要在SDK中再放一份。
*
* @author alexsun
*
*/
public class VDResolutionData {
/** 清晰度:流畅 */
public static final String TYPE_DEFINITION_CIF = "cif";
/** 清晰度:标清 */
public static final String TYPE_DEFINITION_SD = "sd";
/** 清晰度:高清 */
public static final String TYPE_DEFINITION_HD = "hd";
/** 清晰度:超清 */
public static final String TYPE_DEFINITION_FHD = "fhd";
/**
* 3D,这个一般都没有用到
*/
public static final String TYPE_DEFINITION_3D = "3d";
public static LinkedHashMap<String, String> mDefDesc = new LinkedHashMap<String, String>();
static {
mDefDesc.put(TYPE_DEFINITION_CIF, "流畅");
mDefDesc.put(TYPE_DEFINITION_SD, "标清");
mDefDesc.put(TYPE_DEFINITION_HD, "高清");
mDefDesc.put(TYPE_DEFINITION_FHD, "超清");
mDefDesc.put(TYPE_DEFINITION_3D, "3D");
}
public static String getDefDescTagWithIndex(int index) {
Iterator<Map.Entry<String, String>> iter = mDefDesc.entrySet()
.iterator();
int step = 0;
String ret = null;
while (iter.hasNext()) {
Map.Entry<String, String> entry = (Map.Entry<String, String>) iter
.next();
if (step == index) {
ret = entry.getKey();
}
step++;
}
return ret;
}
public static String getDefDescTextWithTag(String tag) {
return mDefDesc.get(tag);
}
public static List<String> getDefDescList() {
ArrayList<String> retList = new ArrayList<String>();
Iterator<Map.Entry<String, String>> iter = mDefDesc.entrySet()
.iterator();
while (iter.hasNext()) {
Map.Entry<String, String> entry = (Map.Entry<String, String>) iter
.next();
retList.add(entry.getValue());
}
return retList;
}
public static List<String> getDefDescTagList() {
ArrayList<String> retList = new ArrayList<String>();
Iterator<Map.Entry<String, String>> iter = mDefDesc.entrySet()
.iterator();
while (iter.hasNext()) {
Map.Entry<String, String> entry = (Map.Entry<String, String>) iter
.next();
retList.add(entry.getKey());
}
return retList;
}
public static int getDefDescIndexWithTag(String tag) {
int ret = -1;
Iterator<Map.Entry<String, String>> iter = mDefDesc.entrySet()
.iterator();
int count = 0;
while (iter.hasNext()) {
Map.Entry<String, String> entry = (Map.Entry<String, String>) iter
.next();
if (entry.getKey().equals(tag)) {
ret = count;
}
count++;
}
return ret;
}
private List<VDResolution> mResolutionList = new ArrayList<VDResolution>();
public VDResolution getFirstResolution() {
if (mResolutionList != null && mResolutionList.size() != 0) {
return mResolutionList.get(0);
}
return null;
}
public List<String> getUrlList() {
if (!mResolutionList.isEmpty()) {
ArrayList<String> retUrlList = new ArrayList<String>();
for (VDResolution resolution : mResolutionList) {
retUrlList.add(resolution.mUrl);
}
return retUrlList;
}
return null;
}
public List<String> getTagList() {
if (!mResolutionList.isEmpty()) {
ArrayList<String> retTagList = new ArrayList<String>();
for (VDResolution resolution : mResolutionList) {
retTagList.add(resolution.mTag);
}
return retTagList;
}
return null;
}
/**
* 深度copy,效率方面似乎有优化余地,要保持次序
*
* @param src
*/
public void deepCopy(VDResolutionData src) {
if (!src.getResolutionList().isEmpty()) {
mResolutionList.clear();
for (VDResolution value : src.mResolutionList) {
mResolutionList.add(value);
}
}
}
public int getResolutionSize() {
return mResolutionList.size();
}
public List<VDResolution> getResolutionList() {
return mResolutionList;
}
public VDResolution getResolutionWithTag(String tag) {
for (VDResolution value : mResolutionList) {
if (value.mTag.equals(tag)) {
return value;
}
}
return null;
}
/**
* 这个函数名字有问题,实际上是返回tag:url的组合
*
* @return
*/
public HashMap<String, String> getResolution() {
HashMap<String, String> retMap = null;
if (mResolutionList != null) {
for (VDResolution value : mResolutionList) {
if (retMap == null) {
retMap = new HashMap<String, String>();
}
retMap.put(value.mTag, value.mUrl);
}
}
return retMap;
}
public boolean isContainTag(String tag) {
if (mResolutionList != null) {
for (VDResolution value : mResolutionList) {
if (value.mTag.equals(tag)) {
return true;
}
}
}
return false;
}
public void addResolution(VDResolution resolution) {
if (!isContainTag(resolution.mTag)) {
mResolutionList.add(resolution);
}
}
/**
* @deprecated 不再使用
* @param tag
* @param resolution
*/
public void addResolution(String tag, VDResolution resolution) {
if (!isContainTag(tag)) {
mResolutionList.add(resolution);
}
}
@Override
public String toString() {
// TODO Auto-generated method stub
StringBuilder sb = new StringBuilder();
for (VDResolution value : mResolutionList) {
sb.append(value.toString() + "\n");
sb.append("-------------\n");
}
return sb.toString();
}
/**
* 清晰度封装
*
* @author alexsun
*
*/
public static class VDResolution {
public VDResolution() {
super();
}
public VDResolution(String tag, String url, int programID, int bandwidth) {
super();
mTag = tag;
mUrl = url;
mProgramID = programID;
mBandWidth = bandwidth;
}
private String mTag = "";
private String mUrl = "";
private int mProgramID = 0;
private int mBandWidth = 0;
public void setTag(String tag) {
mTag = tag;
}
public void setUrl(String url) {
mUrl = url;
}
public void setProgramID(int programID) {
mProgramID = programID;
}
public void setBandWidth(int bandwidth) {
mBandWidth = bandwidth;
}
public String getTag() {
return mTag;
}
public String getUrl() {
return mUrl;
}
public int getProgramID() {
return mProgramID;
}
public int getBandWidth() {
return mBandWidth;
}
public void deepCopy(VDResolution resolution) {
mUrl = new String(resolution.getUrl());
mTag = new String(resolution.getTag());
mProgramID = resolution.getProgramID();
mBandWidth = resolution.getBandWidth();
}
@Override
public String toString() {
// TODO Auto-generated method stub
StringBuilder sb = new StringBuilder();
sb.append("mTag" + mTag + "\n");
sb.append("mUrl" + mUrl + "\n");
sb.append("mProgramID" + mProgramID + "\n");
sb.append("mBandWidth" + mBandWidth + "\n");
return sb.toString();
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/container/VDVideoControlTopContainer.java
package com.sina.sinavideo.sdk.container;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.view.MotionEvent;
import android.view.View;
import android.view.animation.Animation;
import android.view.animation.AnimationUtils;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnKeyEventListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnScreenTouchListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideControllerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideTopContainerListener;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.utils.VDUtility;
import com.sina.sinavideo.sdk.utils.VDVideoScreenOrientation;
import com.sina.sinavideo.sdk.widgets.VDBaseWidget;
import com.sina.sinavideo.sdk.R;
/**
* 控制条容器类,单击隐藏等操作
*
* @author liuqun
*/
public class VDVideoControlTopContainer extends VDVideoControlContainer implements VDBaseWidget, OnScreenTouchListener,
OnShowHideControllerListener, OnKeyEventListener, OnShowHideTopContainerListener {
private int mStateBarHeight = 0;
private boolean mUseStatusBar = true;
private Animation mShowAnim;
private Animation mHideAnim;
// private boolean isAnimating;
public VDVideoControlTopContainer(Context context, AttributeSet attrs) {
super(context, attrs);
init(context);
TypedArray a = context.obtainStyledAttributes(attrs, R.styleable.VDVideoControlTopContainer);
mUseStatusBar = a.getBoolean(R.styleable.VDVideoControlTopContainer_useStatusBar, true);
a.recycle();
if (!mUseStatusBar) {
mStateBarHeight = 0;
}
VDVideoViewController controller = VDVideoViewController.getInstance(context);
if (controller != null)
controller.addOnShowHideControllerListener(this);
}
public VDVideoControlTopContainer(Context context) {
super(context);
init(context);
}
@Override
public boolean onTouchEvent(MotionEvent event) {
// TODO Auto-generated method stub
return true;
}
private void init(Context ctt) {
VDLog.d("VDVideoControlTopContainer", "context ctt=" + ctt);
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (controller != null)
controller.addOnKeyEventListener(this);
mShowAnim = AnimationUtils.loadAnimation(ctt, R.anim.up_to_down_translate);
mShowAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
setVisibility(View.VISIBLE);
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
clearAnimation();
}
});
mHideAnim = AnimationUtils.loadAnimation(ctt, R.anim.down_to_up_translate2);
mHideAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
setVisibility(View.GONE);
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
clearAnimation();
}
});
mStateBarHeight = VDVideoScreenOrientation.getStatusBarHeight(ctt);
}
@Override
public void hide() {
super.hide();
removeCallbacks(hideRun);
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (controller != null)
controller.removeOnShowHideControllerListener(this);
if (controller != null)
controller.removeOnShowHideTopControllerListener(this);
}
@Override
public void reset() {
super.reset();
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (controller != null)
controller.addOnShowHideControllerListener(this);
if (controller != null)
controller.addOnShowHideTopControllerListener(this);
}
@Override
public void onSingleTouch(MotionEvent ev) {
// Log.i("VDVideoControlTopContainer",
// "VDVideoControlTopContainer onSingleTouch " + getVisibility());
if (getVisibility() == VISIBLE) {
removeCallbacks(hideRun);
hideController();
} else {
removeCallbacks(hideRun);
showController();
postDelayed(hideRun, VDVideoViewController.DEFAULT_DELAY);
}
}
@Override
public void doNotHideControllerBar() {
removeCallbacks(hideRun);
}
@Override
public void hideControllerBar(long delay) {
hideControllerBarWithDelay(delay);
}
@Override
public void onPostHide() {
// TODO Auto-generated method stub
}
@Override
public void onPostShow() {
// TODO Auto-generated method stub
}
@Override
public void onPreHide() {
// TODO Auto-generated method stub
}
@Override
public void onKeyEvent() {
if (getVisibility() == VISIBLE) {
removeCallbacks(hideRun);
hideController();
} else {
removeCallbacks(hideRun);
showController();
// postDelayed(hideRun, VDVideoViewController.DEFAULT_DELAY);
}
}
@Override
public void onKeyLeftRight() {
// TODO Auto-generated method stub
}
@Override
public void showControllerBar(boolean delayHide) {
showControllerBarWithDelay(delayHide);
}
@Override
public void onPreShow() {
// TODO Auto-generated method stub
}
@Override
public void hideTopControllerBar() {
// TODO Auto-generated method stub
hideController();
}
@Override
public void showTopControllerBar() {
// TODO Auto-generated method stub
showController();
}
/**
* 隐藏当前容器的私有方法,集中调用这个方法来实现<br />
* TODO 后期考虑将此方法重构,放入统一的helper类中
*
* @param delay 延迟隐藏的毫秒数,立即执行,填写0
*/
private void hideControllerBarWithDelay(long delay) {
removeCallbacks(hideRun);
if (delay <= 0) {
hideController();
} else {
postDelayed(hideRun, delay);
}
}
/**
* 显示当前容器的私有方法,集中调用这个方法来实现<br />
* TODO 后期考虑将此方法重构,放入统一的helper类中
*
* @param delay 是否需要自动隐藏
*/
private void showControllerBarWithDelay(boolean delayHide) {
showController();
removeCallbacks(hideRun);
if (delayHide) {
postDelayed(hideRun, VDVideoViewController.DEFAULT_DELAY);
}
}
private void hideController() {
if (getVisibility() != VISIBLE || getAnimation() != null) {
return;
}
MarginLayoutParams lp = (MarginLayoutParams) getLayoutParams();
if (lp != null) {
lp.topMargin = 0;
}
// setVisibility(GONE);
// TODO 控制栏部分,挪到外部去处理
if (mUseStatusBar) {
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (null != controller)
controller.hideStatusBar(true);
}
startAnimation(mHideAnim);
if (lp != null) {
setLayoutParams(lp);
}
}
private void showController() {
if (getVisibility() == VISIBLE) {
return;
}
MarginLayoutParams lp = (MarginLayoutParams) getLayoutParams();
if (lp != null && !(VDUtility.isMeizu() || VDUtility.isSamsungNoteII())) {
lp.topMargin = mStateBarHeight;
}
// setVisibility(VISIBLE);
startAnimation(mShowAnim);
// TODO 控制栏部分,挪到外部去处理
if (mUseStatusBar) {
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (null != controller)
controller.hideStatusBar(false);
}
if (lp != null) {
VDLog.i("VDVideoControlTopContainer", "padTop = " + mStateBarHeight);
setLayoutParams(lp);
}
}
private Runnable hideRun = new Runnable() {
@Override
public void run() {
hideController();
}
};
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoSoundHorizontalSeekBar.java
package com.sina.sinavideo.sdk.widgets;
import android.content.Context;
import android.content.res.TypedArray;
import android.graphics.drawable.Drawable;
import android.util.AttributeSet;
import android.widget.SeekBar;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSetSoundListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSoundChangedListener;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.utils.VDPlayerSoundManager;
import com.sina.sinavideo.sdk.R;
/**
* 视频音量拖动条
*
* @author seven
*
*/
public final class VDVideoSoundHorizontalSeekBar extends SeekBar implements
VDBaseWidget, OnSoundChangedListener, SeekBar.OnSeekBarChangeListener,
OnSetSoundListener {
private boolean mIsDragging = false;
private Context mContext = null;
private final static String TAG = "VDVideoSoundSeekBar";
public VDVideoSoundHorizontalSeekBar(Context context) {
super(context);
init(context, null);
}
public VDVideoSoundHorizontalSeekBar(Context context, AttributeSet attrs) {
super(context, attrs);
init(context, attrs);
}
private void initVolume(Context context) {
int maxVolume = VDPlayerSoundManager.getMaxSoundVolume(context);
int progress = VDPlayerSoundManager.getCurrSoundVolume(context);
setMax(maxVolume);
setProgress(progress);
VDLog.e(TAG, "max:" + maxVolume + ",progress:" + progress);
}
private void init(Context context, AttributeSet attrs) {
mContext = context;
// 滑动条样式
TypedArray typedArr = context.obtainStyledAttributes(attrs, new int[] {
android.R.attr.progressDrawable, android.R.attr.thumb });
if (typedArr != null) {
Drawable drawable = typedArr.getDrawable(0);
if (drawable == null) {
setProgressDrawable(context.getResources().getDrawable(
R.drawable.play_soundseekbar_background));
}
} else {
setProgressDrawable(context.getResources().getDrawable(
R.drawable.play_soundseekbar_background));
}
// 焦点样式
if (typedArr != null) {
Drawable drawable = typedArr.getDrawable(1);
if (drawable == null) {
setThumb(context.getResources().getDrawable(
R.drawable.play_ctrl_sound_ball));
}
} else {
setThumb(context.getResources().getDrawable(
R.drawable.play_ctrl_sound_ball));
}
if (typedArr != null) {
typedArr.recycle();
}
registerListener();
}
private void registerListener() {
setOnSeekBarChangeListener(this);
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null)
controller.addOnSoundChangedListener(this);
if (controller != null)
controller.addOnSetSoundListener(this);
// VDVideoViewController.getInstance().addOnSoundVisibleListener(this);
}
@Override
public void reset() {
// VDVideoViewController.getInstance().addOnSoundChangedListener(this);
initVolume(mContext);
}
@Override
public void hide() {
// VDVideoViewController.getInstance().removeOnSoundChangedListener(this);
}
@Override
public void onSoundChanged(int currVolume) {
// LogS.e(TAG, "onSoundChanged currVolume:" + currVolume +
// ",mIsDragging : " + mIsDragging);
if (!mIsDragging) {
setProgress(currVolume);
onSizeChanged(getWidth(), getHeight(), 0, 0);
}
}
@Override
public void onProgressChanged(SeekBar seekBar, int progress,
boolean fromUser) {
// VDLog.e(TAG, "onProgressChanged progress:" + progress +
// ",mIsDragging : " + mIsDragging);
VDPlayerSoundManager.dragSoundSeekTo(mContext, progress, mIsDragging);
if (mIsDragging) {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null)
controller.notifyRemoveAndHideDelayMoreOprationPanel();
}
}
@Override
public void onStartTrackingTouch(SeekBar arg0) {
mIsDragging = true;
// VDVideoViewController.getInstance().notifySoundSeekBarVisible(true);
}
@Override
public void onStopTrackingTouch(SeekBar arg0) {
mIsDragging = false;
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.notifyRemoveAndHideDelayMoreOprationPanel();
}
// @Override
// public void onSoundSeekBarVisible(boolean isVisible) {
// setProgress(getProgress());
// onSizeChanged(getWidth(), getHeight(), 0, 0);
// }
@Override
public void onSetCurVolume(int currVolume) {
// LogS.e(TAG, "DLNA onSetCurVolume currVolume:" + currVolume );
setProgress(currVolume);
onSizeChanged(getWidth(), getHeight(), 0, 0);
}
@Override
public void onSetMaxVolume(int maxVolume) {
// LogS.e(TAG, "DLNA onSetMaxVolume maxVolume:" + maxVolume );
if (getMax() != maxVolume) {
setMax(maxVolume);
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/container/VDVideoControlBottomContainer.java
package com.sina.sinavideo.sdk.container;
import android.content.Context;
import android.util.AttributeSet;
import android.view.MotionEvent;
import android.view.View;
import android.view.animation.Animation;
import android.view.animation.AnimationUtils;
import com.sina.sinavideo.sdk.R;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnScreenTouchListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideBottomControllerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideControllerListener;
import com.sina.sinavideo.sdk.widgets.VDBaseWidget;
/**
* 控制条容器类,单击隐藏等操作
*
* @author liuqun
*/
public class VDVideoControlBottomContainer extends VDVideoControlContainer implements VDBaseWidget,
OnScreenTouchListener, OnShowHideControllerListener, OnShowHideBottomControllerListener {
private Animation mShowAnim;
private Animation mHideAnim;
// private boolean isAnimating;
public VDVideoControlBottomContainer(Context context, AttributeSet attrs) {
super(context, attrs);
init(context);
}
public VDVideoControlBottomContainer(Context context) {
super(context);
init(context);
}
private void init(Context ctt) {
// VDVideoViewController.getInstance().addOnShowHideControllerListener(this);
mShowAnim = AnimationUtils.loadAnimation(ctt, R.anim.down_to_up_translate);
mShowAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
VDVideoViewController controller = VDVideoViewController.getInstance(VDVideoControlBottomContainer.this.getContext());
if (null != controller)
controller.setBottomPannelHiding(false);
setVisibility(View.VISIBLE);
if (null != controller)
controller.notifyHideVideoList();
if (null != controller)
controller.notifyHideMoreOprationPanel();
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
VDVideoViewController controller = VDVideoViewController.getInstance(VDVideoControlBottomContainer.this.getContext());
if (null != controller)
controller.notifyControllerBarPostShow();
clearAnimation();
}
});
mHideAnim = AnimationUtils.loadAnimation(ctt, R.anim.up_to_down_translate2);
mHideAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
VDVideoViewController controller = VDVideoViewController.getInstance(VDVideoControlBottomContainer.this.getContext());
if (controller != null)
controller.setBottomPannelHiding(true);
setVisibility(GONE);
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
VDVideoViewController controller = VDVideoViewController.getInstance(VDVideoControlBottomContainer.this.getContext());
if (controller != null)
controller.notifyControllerBarPostHide();
clearAnimation();
// isAnimating = false;
}
});
}
@Override
public boolean onTouchEvent(MotionEvent event) {
// TODO Auto-generated method stub
return true;
}
@Override
public void onSingleTouch(MotionEvent ev) {
if (getVisibility() == VISIBLE) {
hideControllerBarWithDelay(0);
} else {
showControllerBarWithDelay(true);
}
}
@Override
public void doNotHideControllerBar() {
removeCallbacks(hideRun);
}
@Override
public void hideControllerBar(long delay) {
hideControllerBarWithDelay(delay);
}
@Override
public void onPostHide() {
// TODO Auto-generated method stub
}
@Override
public void onPostShow() {
// TODO Auto-generated method stub
}
@Override
public void onPreHide() {
// TODO Auto-generated method stub
}
@Override
public void showControllerBar(boolean delayHide) {
showControllerBarWithDelay(delayHide);
}
@Override
public void onPreShow() {
// TODO Auto-generated method stub
}
@Override
public void reset() {
// Exception e = new Exception("this is a log");
// e.printStackTrace();
super.reset();
removeCallbacks(hideRun);
setVisibility(GONE);
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (controller != null)
controller.addOnShowHideControllerListener(this);
if (controller != null)
controller.addOnShowHideBottomControllerListener(this);
}
@Override
public void hide() {
super.hide();
removeCallbacks(hideRun);
setVisibility(GONE);
VDVideoViewController controller = VDVideoViewController.getInstance(this.getContext());
if (controller != null)
controller.removeOnShowHideControllerListener(this);
if (controller != null)
controller.removeOnShowHideBottomControllerListener(this);
}
@Override
public void hideBottomControllerBar() {
if (getVisibility() == VISIBLE && getAnimation() == null) {
startAnimation(mHideAnim);
}
}
@Override
public void showBottomControllerBar() {
if (getVisibility() != VISIBLE && getAnimation() == null) {
startAnimation(mShowAnim);
}
}
/**
* 显示当前容器的私有方法,集中调用这个方法来实现<br />
* #NOTE 后期考虑将此方法重构,放入统一的helper类中
*
* @param
*/
private void showControllerBarWithDelay(boolean delayHide) {
if (getVisibility() != VISIBLE) {
setVisibility(View.VISIBLE);
startAnimation(mShowAnim);
}
removeCallbacks(hideRun);
if (delayHide) {
postDelayed(hideRun, VDVideoViewController.DEFAULT_DELAY);
}
}
/**
* 隐藏当前容器的私有方法,集中调用这个方法来实现<br />
* #NOTE 后期考虑将此方法重构,放入统一的helper类中
*
* @param delay 延迟隐藏的毫秒数,立即执行,填写0
*/
private void hideControllerBarWithDelay(long delay) {
removeCallbacks(hideRun);
postDelayed(hideRun, delay);
}
private Runnable hideRun = new Runnable() {
@Override
public void run() {
if (getVisibility() == VISIBLE && getAnimation() == null) {
startAnimation(mHideAnim);
VDVideoViewController controller = VDVideoViewController.getInstance(VDVideoControlBottomContainer.this.getContext());
if (null != controller)
controller.notifyControllerBarPreHide();
}
}
};
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/VDVideoViewController.java
package com.sina.sinavideo.sdk;
import java.io.File;
import java.lang.ref.WeakReference;
import java.util.HashMap;
import java.util.List;
import java.util.Map;
import android.app.Activity;
import android.content.Context;
import android.content.IntentFilter;
import android.graphics.PointF;
import android.media.TimedText;
import android.net.ConnectivityManager;
import android.os.Environment;
import android.os.Handler;
import android.os.Looper;
import android.os.Message;
import android.provider.Settings;
import android.util.DisplayMetrics;
import android.view.KeyEvent;
import android.view.MotionEvent;
import android.view.View;
import android.view.Window;
import android.view.WindowManager;
import com.sina.sinavideo.coreplayer.ISinaVideoView;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnBufferingUpdateListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnCompletionListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnErrorListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnInfoListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnPreparedListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnSeekCompleteListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnTimedTextListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnVideoOpenedListener;
import com.sina.sinavideo.coreplayer.splayer.MediaPlayer.OnVideoSizeChangedListener;
import com.sina.sinavideo.coreplayer.splayer.VideoView;
import com.sina.sinavideo.coreplayer.splayer.VideoViewHard;
import com.sina.sinavideo.sdk.VDVideoConfig.eVDDecodingType;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnClickPlayListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnClickRetryListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnDLNALinearLayoutListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnDecodingTypeListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnKeyChangeProgressListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnKeyEventListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnLightingVisibleListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnLoadingListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnMoreOprationVisibleChangeListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnProgressViewVisibleListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnRegisterDLNAListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnResolutionListButtonListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnScreenOrientationChangeListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnScreenOrientationSwitchListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSetSoundListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideADContainerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideBottomControllerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideControllerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnShowHideTopContainerListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnSoundVisibleListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnTipListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVMSResolutionListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoDoubleTapListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoFrameADListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoGuideTipsListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoInsertADListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoListListener;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnVideoListVisibleChangeListener;
import com.sina.sinavideo.sdk.data.VDPlayerErrorInfo;
import com.sina.sinavideo.sdk.data.VDPlayerInfo;
import com.sina.sinavideo.sdk.data.VDResolutionData;
import com.sina.sinavideo.sdk.data.VDVideoInfo;
import com.sina.sinavideo.sdk.data.VDVideoListInfo;
import com.sina.sinavideo.sdk.dlna.DLNAController;
import com.sina.sinavideo.sdk.utils.VDApplication;
import com.sina.sinavideo.sdk.utils.VDLog;
import com.sina.sinavideo.sdk.utils.VDNetworkBroadcastReceiver;
import com.sina.sinavideo.sdk.utils.VDPlayerLightingManager;
import com.sina.sinavideo.sdk.utils.VDPlayerSoundManager;
import com.sina.sinavideo.sdk.utils.VDResolutionManager;
import com.sina.sinavideo.sdk.utils.VDSDKConfig;
import com.sina.sinavideo.sdk.utils.VDSharedPreferencesUtil;
import com.sina.sinavideo.sdk.utils.VDUtility;
import com.sina.sinavideo.sdk.utils.VDVideoFullModeController;
import com.sina.sinavideo.sdk.utils.VDVideoScreenOrientation;
import com.sina.sinavideo.sdk.utils.m3u8.M3u8ContentParser;
import com.sina.sinavideo.sdk.utils.m3u8.M3u8ContentParser.M3u8ParserListener;
/**
* 控制类,使用引用方式调用CorePlayer中的MediaController<br>
* 注:VDVideoviewController不是传统意义上的Videoviewcontroller<br>
* VDVideoViewController更像一个使用EventBus模式处理的消息泵<br>
* 理论上,所有的控制都通过controller来进行<br>
* 所有调用返回的数据都通过VDVideoViewListeners的内部接口来返回
*
* @hide
* @author sunxiao
*/
public class VDVideoViewController implements OnVideoOpenedListener,
OnVideoSizeChangedListener, OnTimedTextListener,
OnBufferingUpdateListener, OnCompletionListener, OnErrorListener,
OnInfoListener, OnPreparedListener, OnSeekCompleteListener {
public static final int MESSAGE_UPDATE_PROGRESS = 0;
public static int DEFAULT_DELAY = 5000;
/**
* 播放列表
*/
public VDVideoListInfo mVDVideoListInfo = new VDVideoListInfo();
/**
* 播放器状态信息
*/
public VDPlayerInfo mVDPlayerInfo = new VDPlayerInfo();
/**
* Core层的VideoView
*/
private ISinaVideoView mVideoView = null;
/**
* 外界的事件处理类
*/
private VDVideoViewListeners mListeners = null;
/**
* 是否有声音提示组件
*/
public boolean mIsHasSoundWidget = false;
/**
* 是否底部控制区执行隐藏动画状态
*/
private boolean mIsBottomPannelHiding = false;
/**
* 当前播放进度(100%)
*/
public float mProgressRate;
/**
* 网络监控
*/
public VDNetworkBroadcastReceiver mReciever = new VDNetworkBroadcastReceiver();
/**
* layer数据
*/
public VDVideoViewLayerContextData mLayerContextData = null;
// 解析m3u8类
private M3u8ContentParser mParser = null;
/**
* 得到外部事件处理
*
* @return
*/
public VDVideoExtListeners getExtListener() {
return mExtListeners;
}
private VDVideoExtListeners mExtListeners = null;
private static Map<Context, VDVideoViewController> mControllers = new HashMap<Context, VDVideoViewController>();
private final static String TAG = "VDVideoViewController";
private Context mContext = null;
private boolean mIsUpdateProgress = false;
private int mRetryTimes = 0;
private boolean mIsPlayed = false; // 是否播放过
private NetChangeListener mNetChangeListener;
// 广告部分变量
/**
* 广告跳秒的变量
*/
private int mADTickerSecNum = -1000;
private int[] mTmpArr = new int[] { 0, 0 };
// ------------- end 日志
/**
* 静帧广告控制,使用或方式进行,对应于attrs.xml-VDVideoAdContainer-adConfig
*/
public boolean mADIsFromStart = false;
public int mADConfigEnum = 0;
private Handler mInsertADHandler = new Handler();
private Runnable mInsertADRunnable = new Runnable() {
@Override
public void run() {
// TODO Auto-generated method stub
try {
if (getCurrentVideo().mIsInsertAD) {
if (getADTickerSec() > 0) {
notifyInsertADTicker();
mInsertADHandler.postDelayed(mInsertADRunnable, 1000);
} else if (getADTickerSec() == 0) {
if (mVDVideoListInfo != null
&& mVDVideoListInfo.isInsertADEnd()) {
mExtListeners
.notifyInsertADEnd(VDPlayerErrorInfo.MEDIA_INSERTAD_ERROR_STEPOUT);
}
playNext();
}
}
} catch (Exception ex) {
}
}
};
private int mWhereTopause = 0;
private Handler mMainHander = new Handler();
private Runnable mSoundDisapperRunnable = new Runnable() {
@Override
public void run() {
notifySoundSeekBarVisible(false);
mListeners.notifySoundVisible(false);
}
};
/**
* 事件泵
*/
private Handler mMessageHandler = new Handler(new Handler.Callback() {
@Override
public boolean handleMessage(Message msg) {
switch (msg.what) {
case MESSAGE_UPDATE_PROGRESS:
if (mVideoView == null || getCurrentVideo() == null) {
return false;
}
VDVideoInfo info = getCurrentVideo();
// 伪直播做点播时候需要seekto到上一次停止时间,所以不能获取真正的当前时间,一般会是0.
// mNeedSeekTo在非直播且清晰度发生切换时标记
long position = 0;
// if (info != null && info.mNeedSeekTo &&
// info.mVideoPosition >= 0) {
// position = info.mVideoPosition;
// } else {
position = mVideoView.getCurrentPosition();
// }
long duration = mVideoView.getDuration();
if (info != null) {
info.mVideoPosition = position;
info.mVideoDuration = duration;
if (!info.mIsLive) {
mListeners.notifyProgressUpdate(position, duration);
}
info.mVideoPosition = position;
}
if (mIsUpdateProgress) {
long delay = 1000 - (position % 1000);
if (delay < 50) {
delay += 1000;
}
mMessageHandler.sendEmptyMessageDelayed(
MESSAGE_UPDATE_PROGRESS, delay);
}
break;
}
return false;
}
});
public void setLayerContextData(VDVideoViewLayerContextData layerContextData) {
mLayerContextData = layerContextData;
}
public VDVideoViewLayerContextData getLayerContextData() {
return mLayerContextData;
}
public void setScreenOrientationPause(boolean isPause) {
if (isPause) {
mWhereTopause = 2;
} else {
mWhereTopause = 0;
}
}
public void setSeekPause(boolean isPause) {
if (isPause) {
mWhereTopause = 1;
} else {
mWhereTopause = 0;
}
}
public void setBeginPause(boolean isPause) {
if (isPause) {
mWhereTopause = 4;
} else {
mWhereTopause = 0;
}
}
/**
* 设置当前底部控制区处于执行隐藏动画状态
*
* @param anim
*/
public void setBottomPannelHiding(boolean anim) {
mIsBottomPannelHiding = anim;
}
/**
* 判定当前底部控制区是否处于执行隐藏动画状态,底部控制区执行隐藏动画屏蔽控制区功能按钮点击操作
*
* @return
*/
public boolean isBottomPannelHiding() {
return mIsBottomPannelHiding;
}
public VDVideoViewController(Context context) {
mContext = context;
mListeners = new VDVideoViewListeners(context);
mExtListeners = new VDVideoExtListeners(context);
}
// public LogPushManager getLogPushManager(){
// return mLogPushManager;
// }
/**
* 截获音量键后触发
*
* @param keyCode
*/
private void handleVolumeKey(int keyCode) {
int curr = VDPlayerSoundManager.getCurrSoundVolume(mContext);
int max = VDPlayerSoundManager.getMaxSoundVolume(mContext);
// float currFloat = (float) curr / max;
if (keyCode == KeyEvent.KEYCODE_VOLUME_DOWN) {
// currFloat -= 0.1f;
curr--;
} else if (keyCode == KeyEvent.KEYCODE_VOLUME_UP) {
// currFloat += 0.1f;
curr++;
}
// if (currFloat < 0) {
// currFloat = 0;
// } else if (currFloat > 1) {
// currFloat = 1.f;
// }
if (curr < 0) {
curr = 0;
} else if (curr > max) {
curr = max;
}
// notifySoundSeekBarVisible(true);
mListeners.notifySoundVisible(true);
// TODO DLNA
notifySetMaxVolume(VDPlayerSoundManager.getMaxSoundVolume(mContext));
notifySetCurVolume(VDPlayerSoundManager.getCurrSoundVolume(mContext));
mMainHander.removeCallbacks(mSoundDisapperRunnable);
mMainHander.postDelayed(mSoundDisapperRunnable, 1000);
dragSoundSeekTo(curr);
}
/**
* 用来提供给activity捕获key的时候使用
*
* @param event
* @return
*/
public boolean onKeyEvent(KeyEvent event) {
boolean flag = false;
if (event.getKeyCode() == KeyEvent.KEYCODE_VOLUME_DOWN
|| event.getKeyCode() == KeyEvent.KEYCODE_VOLUME_UP) {
if (event.getAction() == KeyEvent.ACTION_DOWN) {
// VDVideoViewController.getInstance().notifySoundVisible(true);
handleVolumeKey(event.getKeyCode());
VDLog.e(TAG,
"dispatchKeyEvent截获KEYCODE_VOLUME_DOWN|KEYCODE_VOLUME_UP");
}
if (mIsHasSoundWidget) {
flag = true;
}
} else if (event.getKeyCode() == KeyEvent.KEYCODE_BACK
&& mLayerContextData.getLayerType() != VDVideoViewLayerContextData.LAYER_COMPLEX_NOVERTICAL) {
if (!VDVideoFullModeController.getInstance().getIsPortrait()) {
setIsFullScreen(false);
flag = true;
}
}
return flag;
}
/**
* 设置清晰度
*
* @deprecated 不允许从外部直接设置清晰度
* @param resolution
*/
// public void setCurResolution(String resolution) {
// mCurResolution = resolution;
// }
/**
* 设置是否锁定屏幕方向-不转屏
*
* @param isLockScreen
*/
public void setIsLockScreen(boolean isLockScreen) {
VDVideoFullModeController.getInstance().setFullLock();
}
/**
* 设置横屏,手动控制调用这里
*/
public void setIsFullScreen(boolean isFullScreen) {
if (mContext == null) {
VDLog.e(VDVideoFullModeController.TAG,
"controller---setIsFullScreen---mContext--return null");
return;
}
if (VDVideoFullModeController.getInstance().getIsFullScreen() == isFullScreen) {
return;
}
if (mLayerContextData.getLayerType() == VDVideoViewLayerContextData.LAYER_COMPLEX_NOVERTICAL) {
if (!isFullScreen) {
// 只横不竖的时候,且转到竖屏,那么直接关闭当前activity
((Activity) mContext).finish();
} else {
// 转到横屏,什么都不做
}
return;
}
VDVideoFullModeController.getInstance().setIsManual(true);
VDVideoFullModeController.getInstance().mInHandNum = 0;
// 发送转屏指令
if (isFullScreen) {
VDVideoScreenOrientation.setLandscape(mContext);
} else {
VDVideoScreenOrientation.setPortrait(mContext);
}
}
/**
* 返回VideoView的引用,用来装载
*
* @return
*/
public ISinaVideoView getVideoView() {
return mVideoView;
}
/**
* 获得播放器当前状态
*
* @return
*/
public int getPlayerStatus() {
return mVDPlayerInfo.mPlayStatus;
}
/**
* 当前是否播放状态
*
* @return
*/
public boolean getIsPlaying() {
if (mVideoView == null) {
return false;
}
// NOTE: 后期修改为mPlayerstatus方式
return mVideoView.isPlaying();
}
/**
* 增加一个重置首屏GuideTips的地方
*
* @param context
* @param isFirst
* 为true,则重置为首次打开,false则为已经打开过了
*/
public void setFirstFullScreen(Context context, boolean isFirst) {
VDSharedPreferencesUtil.setFirstFullScreen(context, isFirst);
}
/**
* 单例启动方法,只能在主线程中调用
*
* @return
*/
public static void register(Context context,
VDVideoViewController controller) {
if (Looper.myLooper() != Looper.getMainLooper()) {
throw new IllegalStateException(TAG + " create not in main thread.");
}
mControllers.put(context, controller);
}
public static void unRegister(Context context) {
if (Looper.myLooper() != Looper.getMainLooper()) {
throw new IllegalStateException(TAG + " create not in main thread.");
}
mControllers.remove(context);
}
public static VDVideoViewController getInstance(Context context) {
if (Looper.myLooper() != Looper.getMainLooper()) {
throw new IllegalStateException(TAG + " create not in main thread.");
}
// if (mVDMediaController == null) {
// mVDMediaController = new VDVideoViewController();
// }
return mControllers.get(context);
}
/**
* 设置上下文<br>
* 注:用activity来填充
*
* @param context
*/
public void setContext(Context context) {
if (context == null) {
return;
}
mContext = context;
VDApplication.getInstance().setContext(context);
}
/**
* 得到上下文<br>
* 注:可直接转为容器的activity
*
* @return
*/
public Context getContext() {
return mContext;
}
/**
* 创建方法,每次切换视频,重新建立一个新的
*
* @param context
* 上下文
*/
public static ISinaVideoView create(Context context) {
if (Looper.myLooper() != Looper.getMainLooper()) {
throw new IllegalStateException(TAG + " create not in main thread.");
}
if (context == null) {
return null;
}
VDVideoViewController controller = VDVideoViewController
.getInstance(context);
if (controller == null) {
controller = new VDVideoViewController(context);
VDVideoViewController.register(context, controller);
}
controller.mContext = context;
// 监听网络变化
final IntentFilter intentFilter = new IntentFilter();
intentFilter.addAction(ConnectivityManager.CONNECTIVITY_ACTION);
context.registerReceiver(controller.mReciever, intentFilter);
if (controller.mNetChangeListener == null) {
controller.mNetChangeListener = new NetChangeListener(context);
controller.mReciever.addListener(controller.mNetChangeListener);
}
// 初始化转屏部分
VDVideoFullModeController.getInstance().init(context);
if (VDVideoConfig.mDecodingType == eVDDecodingType.eVDDecodingTypeSoft) {
controller.mVideoView = new VideoView(context);
} else if (VDVideoConfig.mDecodingType == eVDDecodingType.eVDDecodingTypeHardWare) {
controller.mVideoView = new VideoViewHard(context);
} else {
if (VDSharedPreferencesUtil.isDecodingTypeFFMpeg(context)) {
controller.mVideoView = new VideoView(context);
} else {
controller.mVideoView = new VideoViewHard(context);
}
}
// 事件监听
controller.mVideoView.setOnCompletionListener(controller);
controller.mVideoView.setOnPreparedListener(controller);
controller.mVideoView.setOnInfoListener(controller);
controller.mVideoView.setOnErrorListener(controller);
controller.mVideoView.setOnSeekCompleteListener(controller);
controller.mVideoView.setOnBufferingUpdateListener(controller);
// 对于guidetips做一下初始化
if (VDSharedPreferencesUtil.isFirstFullScreen(context)) {
// 是首次打开
controller.notifyGuideTips(true);
VDSharedPreferencesUtil.setFirstFullScreen(context, false);
} else {
controller.notifyGuideTips(false);
}
return controller.mVideoView;
}
public VDVideoListInfo getVideoList() {
return mVDVideoListInfo;
}
// --------控制方法区begin-----------//
/**
* 设置播放列表
*
* @param infoList
*/
public void setVideoList(VDVideoListInfo infoList) {
if (infoList == null) {
return;
}
mVDVideoListInfo = infoList;
if (mListeners == null) {
mListeners = new VDVideoViewListeners(mContext);
}
mListeners.notifyVideoList(infoList);
}
/**
* 获得当前广告的秒数
*
* @return
*/
public int getADTickerSec() {
return getCurrentVideo().mInsertADSecNum;
}
/**
* 返回处理过的广告跳秒数
*
* @return
*/
public synchronized int refreshADTickerSec() {
if (mADTickerSecNum > 0) {
mADTickerSecNum--;
}
if (getCurrentVideo().mInsertADSecNum > 0) {
getCurrentVideo().mInsertADSecNum--;
}
return mADTickerSecNum;
}
/**
* VDVideoListInfo的controller包装版本<br />
* 尽量使用VDVideoView来调用
*
* @param insertADList
* @param currInfo
*/
public int refreshInsertADList(List<VDVideoInfo> insertADList,
VDVideoInfo currInfo) {
return mVDVideoListInfo.refreshInsertADList(insertADList, currInfo);
}
private boolean playVideoOnInfoKey(int index) {
if (mVideoView == null || mVDVideoListInfo == null || index < 0
|| index >= mVDVideoListInfo.getVideoListSize()) {
return false;
}
stop();
notifyVideoUIRefresh();
boolean same = mVDVideoListInfo.mIndex == index;
mVDVideoListInfo.mIndex = index;
// 设置广告的跳秒数
try {
if (!mVDVideoListInfo.isCanPlay()) {
}
} catch (IllegalArgumentException ex) {
VDLog.e(TAG, ex.getMessage());
return false;
}
VDVideoInfo currInfo = mVDVideoListInfo.getCurrInfo();
if (currInfo == null) {
throw new IllegalArgumentException("url为null");
}
if (mVDVideoListInfo.getVideoInfo(index).mIsInsertAD
&& mVDVideoListInfo.mInsertADSecNum == 0
&& mVDVideoListInfo.getADNum() > 1
&& mVDVideoListInfo.mIsSetInsertADTime) {
// 如果是多流多广告类型,那么就需要手动设置广告时间,如果为0,那么要提示错误。
throw new IllegalArgumentException(
"如果未设置mIsSetInsertADTime的多流多广告类型,那么就需要手动设置广告时间,不能为0。");
}
if (mVDVideoListInfo.mInsertADType != VDVideoListInfo.INSERTAD_TYPE_NONE) {
if (mVDVideoListInfo.mInsertADSecNum != 0) {
mADTickerSecNum = mVDVideoListInfo.mInsertADSecNum;
notifyInsertAD(true);
}
}
// 开始具体的播放逻辑
final VDVideoInfo videoInfo = mVDVideoListInfo
.getVideoInfo(mVDVideoListInfo.mIndex);
mListeners.notifyVideoInfo(videoInfo);
VIDEO_TIME_OUT = videoInfo.mIsLive ? VIDEO_LIVE_TIME_OUT
: VIDEO_NORMAL_TIME_OUT;
notifyShowLoading();
if (videoInfo.mIsLive || videoInfo.isM3u8()) {
playLiveVideo(videoInfo);
} else {
playOnDemandVideo(videoInfo);
if (!same) {
mRetryTimes = 0;
}
}
notifyClickRetry();
return true;
}
/**
* 播放视频
*
* @param index
* 正片列表数组中的下标
* @return
*/
public boolean playVideo(int index) {
index = mVDVideoListInfo.getCurrKeyFromRealInfo(index);
// 日志上报的start部分,从这儿开始
return playVideoOnInfoKey(index);
}
private void startM3u8ContentParser(VDVideoInfo info) {
mParser = new M3u8ContentParser(new MyM3u8ParserListener(info),
info.mVideoId, mContext);
mParser.startParserM3u8(info.mPlayUrl);
}
/**
* 播放直播视频
*
* @param info
*/
private void playLiveVideo(VDVideoInfo info) {
if (!info.mIsParsed) {
startM3u8ContentParser(info);
} else {
String url = null;
mListeners.notifyResolutionChanged(VDResolutionManager.getInstance(
mContext).getCurrResolutionTag());
setVideoPath(url);
mListeners.notifyShowLoading(true);
mListeners.notifyVideoPrepared(false);
}
}
/**
* 播放点播视频
*
* @param videoInfo
*/
public void playOnDemandVideo(VDVideoInfo videoInfo) {
if (DLNAController.mIsDLNA) {
DLNAController.getInstance(mContext).open(videoInfo.mRedirectUrl);
} else {
playVideoWithInfo(videoInfo);
}
}
/**
* 播放点播视频
*
* @param videoInfo
*/
public void playVideoWithInfo(VDVideoInfo videoInfo) {
if (VDUtility.isLocalUrl(videoInfo.mPlayUrl)) {
videoInfo.mRedirectUrl = videoInfo.mPlayUrl;
}
String url = videoInfo.mRedirectUrl == null ? videoInfo.mPlayUrl
: videoInfo.mRedirectUrl;
setVideoPath(url);
mListeners.notifyShowLoading(true);
mListeners.notifyVideoPrepared(false);
mIsPlayed = false;
notifyClickRetry();
}
/**
* 设置视频地址
*
* @param url
*/
public void setVideoPath(String url) {
getCurrentVideo().mIsParsed = true;
if (url != null) {
mVideoView.setVideoPath(url);
}
}
private void updatePlayState() {
if (mVDPlayerInfo != null && mVideoView != null) {
mVDPlayerInfo.mPlayStatus = mVideoView.isPlaying() ? VDPlayerInfo.PLAYER_STARTED
: VDPlayerInfo.PLAYER_PAUSE;
mVDPlayerInfo.mIsPlaying = mVideoView.isPlaying();
}
if (mListeners != null) {
mListeners.notifyPlayStateChanged();
}
}
/**
* 通知清晰度容器是否显示
*
* @param isVisible
*/
public void notifyResolutionContainerVisible(boolean isVisible) {
if (mListeners != null) {
mListeners.notifyResolutionVisible(isVisible);
}
}
/**
* 清晰度变更
*
* @param tag
*/
public void notifyResolutionChanged(String tag) {
if (mListeners != null) {
mListeners.notifyResolutionChanged(tag);
}
}
/**
* 清晰度解析完毕后回传
*
* @param resolutionData
*/
public void notifyResolutionParsed(VDResolutionData resolutionData) {
if (mListeners != null) {
mListeners.notifyResolutionParsed(resolutionData);
}
}
/**
* 状态变更入口
*/
public void notifyPlayStateChanged() {
if (mListeners != null) {
mListeners.notifyPlayStateChanged();
}
}
/**
* 表明当前是否处在播放视频状态中
*
* @return
*/
public boolean isInsertAD() {
try {
return getCurrentVideo().mIsInsertAD;
} catch (Exception ex) {
}
return false;
}
/**
* 当前视频信息
*/
public VDVideoInfo getCurrentVideo() {
try {
return mVDVideoListInfo.getVideoInfo(mVDVideoListInfo.mIndex);
} catch (Exception ex) {
VDLog.e(TAG, ex.getMessage());
}
return null;
}
/**
* 当前视频清晰度集合对应信息
*
* @return
*/
public HashMap<String, String> getCurrVMSResolutionInfo() {
if (mVDVideoListInfo.mIndex >= 0
&& mVDVideoListInfo.mIndex < mVDVideoListInfo
.getVideoListSize()) {
return mVDVideoListInfo.getVideoInfo(mVDVideoListInfo.mIndex)
.getVMSDefinitionInfo();
}
return null;
}
/**
* 得到播放列表数量
*
* @return
*/
public int getVideoInfoNum() {
try {
return mVDVideoListInfo.getRealVideoListSize();
} catch (Exception ex) {
return 0;
}
}
/**
* 得到播放器信息【重构的时候,看是否去掉,这个结构可以放到controller里面】
*
* @return
*/
public VDPlayerInfo getPlayerInfo() {
return mVDPlayerInfo;
}
/**
* 得到当前进度
*
* @return
*/
public long getCurrentPosition() {
if (mVideoView != null) {
return mVideoView.getCurrentPosition();
}
return 0;
}
/**
* 开始
*/
public void start() {
if (mVideoView != null) {
mVideoView.start();
updatePlayState();
mListeners.notifyVideoFrameADEnd();
}
}
/**
* 是否可以显示广告
*
* @return
*/
public boolean isCanShowFrameAD() {
// 1、用户点击暂停
if (mWhereTopause == 0 && ((mADConfigEnum & 1) == 1))
return true;
// 2、用户滑动视频引起的暂停
if (mWhereTopause == 1 && ((mADConfigEnum & 2) == 2))
return true;
// 3、转屏引起的暂停
if (mWhereTopause == 2 && ((mADConfigEnum & 4) == 4))
return true;
// 4、视频开始的时候,仅限于正片
if (mWhereTopause == 4 && ((mADConfigEnum & 8) == 8))
return true;
return false;
}
/**
* 暂停
*/
public void pause() {
if (mVideoView != null) {
mVideoView.pause();
}
stopUpdateMessage();
mTimeOutHandler.removeMessages(NET_TIME_OUT);
mTimeOutHandler.removeMessages(CHECK_LIVE_TIME_OUT);
updatePlayState();
pauseInsertAD();
// if (isCanShowFrameAD()) {
// mListeners.notifyVideoFrameADBegin();
// }
}
private void pauseInsertAD() {
try {
if (getCurrentVideo().mIsInsertAD) {
mInsertADHandler.removeCallbacks(mInsertADRunnable);
}
} catch (Exception ex) {
VDLog.e(TAG, ex.getMessage());
}
}
private void resumeInsertAD() {
try {
if (getCurrentVideo().mIsInsertAD) {
mInsertADHandler.postDelayed(mInsertADRunnable, 1000);
}
} catch (Exception ex) {
VDLog.e(TAG, ex.getMessage());
}
}
/**
* 继续
*/
public void resume() {
resume(true);
}
/**
* 继续
*
* @param updateUI
* 是否更新当前UI
*/
public void resume(boolean updateUI) {
if (mVideoView != null) {
mVideoView.resume();
}
if (updateUI) {
startUpdateMessage();
}
updatePlayState();
VDVideoFullModeController.getInstance().enableSensor(true);
resumeInsertAD();
mListeners.notifyVideoFrameADEnd();
mVDPlayerInfo.mPlayStatus = VDPlayerInfo.PLAYER_RESUME;
}
/**
* 进度条拖动
*/
public void dragProgressTo(float curr) {
dragProgressTo(curr, false, false);
}
/**
* 进度条拖动
*
* @param curr
* 当前
* @param seek
* 拖动到
* @param isgesture
* TODO 此处需要修改
*/
public void dragProgressTo(float curr, boolean seek, boolean isgesture) {
long duration = 0;
if (DLNAController.mIsDLNA) {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller != null) {
duration = DLNAController.getInstance(controller.getContext()).mDuration;
}
} else {
duration = mVDPlayerInfo.mDuration;
}
long seekTo = (long) (curr * duration);
VDLog.i("VDVideoViewListeners", "seek to : curr = " + curr + " , "
+ seekTo);
if (seek) {
if (DLNAController.mIsDLNA) {
DLNAController.getInstance(mContext).seek(seekTo);
} else {
mVideoView.seekTo(seekTo);
}
}
if (DLNAController.mIsDLNA) {
DLNAController.mTmpPosition = seekTo;
} else {
mVDPlayerInfo.mCurrent = seekTo;
}
mListeners.notifyDragTo(seekTo, duration);
}
/**
* 屏幕宽高
*
* @return
*/
public int[] getScreen() {
if (mContext == null) {
return mTmpArr;
}
WindowManager wm = (WindowManager) mContext
.getSystemService(Context.WINDOW_SERVICE);
if (VDUtility.getSDKInt() > 13) {
}
DisplayMetrics metrics = new DisplayMetrics();
wm.getDefaultDisplay().getMetrics(metrics);
int[] rect = { metrics.widthPixels, metrics.heightPixels };
return rect;
}
public boolean isPlaying() {
return mVideoView.isPlaying();
}
/**
* 调节音量
*
* @param currVolume
* 音量值
*/
public void dragSoundSeekTo(int currVolume) {
VDPlayerSoundManager.dragSoundSeekTo(mContext, currVolume, true);
}
/**
* 调整系统亮度到
*
* @param curr
*/
public void dragLightingTo(float curr, boolean notify) {
VDPlayerLightingManager.getInstance().dragLightingTo(mContext, curr,
notify);
mVDPlayerInfo.mCurLighting = curr;
Settings.System.putInt(this.mContext.getContentResolver(),
Settings.System.SCREEN_BRIGHTNESS, (int) (curr * 255));
if (notify) {
mListeners.notifyLightingSetting(curr);
}
}
/**
* 得到当前亮度
*
* @return
*/
public float getCurrLightingSetting() {
// if (mCurrLightingNum < 0 && mContext != null) {
// mCurrLightingNum =
// Settings.System.getInt(this.mContext.getContentResolver(),
// Settings.System.SCREEN_BRIGHTNESS, 125);
// mCurrLightingNum = mCurrLightingNum / 255;
// }
// return mCurrLightingNum;
return VDPlayerLightingManager.getInstance().getCurrLightingSetting(
mContext);
}
/**
* 得到系统当前是否是自动调节亮度
*
* @return
*/
public boolean getIsAutoLightingSetting() {
// boolean ret = false;
// try {
// ret = Settings.System.getInt(mContext.getContentResolver(),
// Settings.System.SCREEN_BRIGHTNESS_MODE) ==
// Settings.System.SCREEN_BRIGHTNESS_MODE_AUTOMATIC;
// } catch (Exception ex) {
// ex.printStackTrace();
// VDLog.e(TAG, ex.getMessage());
// }
// return ret;
return VDPlayerLightingManager.getInstance().getIsAutoLightingSetting(
mContext);
}
/**
* 设置是否是自动亮度调节
*
* @param isAutoLighting
*/
public void setAutoLighting(boolean isAutoLighting) {
VDPlayerLightingManager.getInstance().setAutoLighting(mContext,
isAutoLighting);
}
/**
* 结束
*/
public void stop() {
if (mParser != null) {
mParser.cancelParserM3U8();
}
mTimeOutHandler.removeMessages(CHECK_LIVE_TIME_OUT);
mTimeOutHandler.removeMessages(NET_TIME_OUT);
mInsertADHandler.removeCallbacks(mInsertADRunnable);
if (mVideoView != null) {
mVideoView.stopPlayback();
mVideoView.setVideoURI(null);
}
updatePlayState();
mVDPlayerInfo.mPlayStatus = VDPlayerInfo.PLAYER_FINISHING;
}
/**
* 播放上一个
*/
public void playPre() {
stop();
int currIndex = mVDVideoListInfo.mIndex - 1;
if (currIndex < 0) {
currIndex = getVideoInfoNum() - 1;
}
playVideoOnInfoKey(currIndex);
notifyVideoUIRefresh();
}
/**
* 播放下一个视频
*/
public void playNext() {
stop();
if (getCurrentVideo().mIsInsertAD) {
mVDVideoListInfo.mInsertADSecNum = 0;
mADTickerSecNum = 0;
}
int currIndex = mVDVideoListInfo.mIndex + 1;
if (currIndex >= mVDVideoListInfo.getVideoListSize()) {
currIndex = 0;
}
playVideoOnInfoKey(currIndex);
notifyVideoUIRefresh();
}
/**
* 拖动,已毫秒为单位
*
* @param mSec
*/
public void seekTo(long mSec) {
mVideoView.seekTo((int) mSec);
}
/**
* 重新启动播放器,以匹配使用软硬件解码方式
*
* @param sec
* 播放进度值
*/
public void reset(long sec) {
VDVideoInfo videoInfo = getCurrentVideo();
videoInfo.mVideoPosition = sec;
getExtListener().notifySwitchPlayerListener(videoInfo,
mVDVideoListInfo.mIndex);
getExtListener().notifyPlayerChange(mVDVideoListInfo.mIndex, sec);
}
/**
* 调整当前视频的分辨率
*
* @param sec
* 播放进度值
* @param resolutionTag
* 新分辨率对应的数值
*/
public void changeResolution(long sec, String resolutionTag) {
VDResolutionManager.getInstance(mContext).setResolutionTag(
resolutionTag);
mListeners.notifyVMSResolutionChanged();
VDVideoInfo videoInfo = getCurrentVideo();
videoInfo.mPlayUrl = videoInfo.getVMSDefinitionInfo()
.get(resolutionTag);
videoInfo.mVideoPosition = sec;
// NOTE 这个接口用得有问题,准备清理掉
getExtListener().notifySwitchPlayerListener(videoInfo,
mVDVideoListInfo.mIndex);
getExtListener().notifyPlayerChange(mVDVideoListInfo.mIndex, sec);
}
/**
* 处理资源清理,切换时候必须调用
*/
public void release() {
if (mReciever != null && mContext != null) {
mContext.unregisterReceiver(mReciever);
}
stop();
mVideoView = null;
mListeners.clear();
mExtListeners.clear();
VDPlayerSoundManager.clear();
mVDPlayerInfo.init();
VDVideoFullModeController.getInstance().release();
}
/**
* 轻度清理资源,在重新加载视频资源时调用
*/
public void release2() {
if (mReciever != null && mContext != null) {
mContext.unregisterReceiver(mReciever);
}
stop();
mVideoView = null;
mVDPlayerInfo.init();
mContext = null;
}
/**
* 开始更新播放进度
*/
private void startUpdateMessage() {
if (!DLNAController.mIsDLNA) {
mIsUpdateProgress = true;
mMessageHandler.removeMessages(MESSAGE_UPDATE_PROGRESS);
mMessageHandler.sendEmptyMessage(MESSAGE_UPDATE_PROGRESS);
}
}
/**
* 停止更新播放进度
*/
private void stopUpdateMessage() {
mIsUpdateProgress = false;
mMessageHandler.removeMessages(MESSAGE_UPDATE_PROGRESS);
}
/**
* 通知播放暂停
*/
public void notifyPlayOrPause() {
mListeners.notifyPlayOrPause();
}
/**
* 通知屏幕单击
*
* @param ev
* 点击事件对象
* @param flag
* 点击的状态
*/
public void touchScreenSingleEvent(MotionEvent ev,
VDVideoViewListeners.eSingleTouchListener flag) {
mListeners.notifyScreenSingleTouch(ev, flag);
}
/**
* 通知按键事件
*/
public void notifyKeyEvent() {
mListeners.notifyKeyEvent();
}
/**
* 通知导航键左右事件
*/
public void notifyKeyLeftRightEvent() {
mListeners.notifyKeyLeftRightEvent();
}
/**
* 通知屏幕双击
*
* @param ev
* 双击事件对象
* @param flag
* 双击的状态
*/
public void touchScreenDoubleEvent(MotionEvent ev,
VDVideoViewListeners.eDoubleTouchListener flag) {
mListeners.notifyScreenDoubleTouch(ev, flag);
}
/**
* 通知屏幕横向滑动
*
* @param point1
* 手指接触屏幕的最开始坐标点
* @param point2
* 滑动后坐标点
* @param beginPoint
* 滑动前坐标点
* @param flag
* 滑动的状态
*/
public void touchScreenHorizonScrollEvent(final PointF point1,
final PointF point2, final PointF beginPoint,
VDVideoViewListeners.eHorizonScrollTouchListener flag) {
mListeners.notifyScreenHorizonScrollTouch(point1, point2, beginPoint,
flag);
}
/**
* 设置进度显示比例
*/
public void setProgressRate() {
if (mVDPlayerInfo.mDuration <= 0) {
return;
}
boolean isPortrait = VDVideoFullModeController.getInstance()
.getIsPortrait();
long duration = mVDPlayerInfo.mDuration;
if (duration < 10 * 60 * 1000) {
if (isPortrait) {
mProgressRate = 60 * 1000f / duration;
} else {
mProgressRate = 90 * 1000f / duration;
}
} else if (duration < 20 * 60 * 1000) {
if (isPortrait) {
mProgressRate = 2 * 60 * 1000f / duration;
} else {
mProgressRate = 150 * 1000f / duration;
}
} else {
if (isPortrait) {
mProgressRate = 5 * 60 * 1000f / duration;
} else {
mProgressRate = 460 * 1000f / duration;
}
}
}
/**
* 通知屏幕横向滑动
*
* @param point1
* 手指接触屏幕的最开始坐标点
* @param point2
* 滑动后坐标点
* @param beginPoint
* 滑动前坐标点
* @param flag
* 滑动的状态
* @param distansY
* 纵向滑动的距离
*/
public void touchScreenVerticalScrollEvent(final PointF point1,
final PointF point2, final PointF beginPoint,
final VDVideoViewListeners.eVerticalScrollTouchListener flag,
float distansY) {
mListeners.notifyScreenVerticalScrollTouch(point1, point2, beginPoint,
flag, distansY);
}
/**
* 设置清晰度,更新清晰度UI
*
* @param resolution
* 对应数值
*/
// public void setResolution(String resolution) {
// this.mCurResolution = resolution;
// mListeners.notifyResolutionSelect(resolution);
// }
/**
* 通知音量图标是否可见
*
* @param isVisible
* true 可见;false 不可见
*/
public void notifySoundSeekBarVisible(boolean isVisible) {
mListeners.notifySoundSeekBarVisible(isVisible);
}
/**
* 通知清晰度选择窗口是否可见
*
* @param isVisible
* true 可见;false 不可见
*/
public void notifyDefinitionContainerVisible(boolean isVisible) {
mListeners.notifyVMSResolutionContainerVisible(isVisible);
}
/**
* 横竖屏转换时候,会调用此方法
*
* @param isFullScreen
* true 全屏;false 非全屏
* @param isFromHand
* true 手动操作;false 非手动操作
*/
public void notifyFullScreen(final boolean isFullScreen,
final boolean isFromHand) {
mListeners.notifyFullScreen(isFullScreen, isFromHand);
}
/**
* guideTips通知
*
* @param isVisible
* true 可见;false 不可见
*/
public void notifyGuideTips(boolean isVisible) {
mListeners.notifyGuideTips(isVisible);
}
/**
* 通知显示加载页面
*/
public void notifyShowLoading() {
mListeners.notifyShowLoading();
}
/**
* 通知隐藏加载页面
*/
public void notifyHideLoading() {
mListeners.notifyHideLoading();
}
/**
* 通知显示或隐藏相关视频列表
*/
public void notifyToogleVideoList() {
mListeners.notifyVideoListVisibelChange();
}
/**
* 通知显示视频列表
*/
public void notifyShowVideoList() {
mListeners.notifyShowVideoList();
}
/**
* 通知隐藏视频列表
*/
public void notifyHideVideoList() {
mListeners.notifyHideVideoList();
}
/**
* 显示静帧广告
*/
public void notifyVideoFrameADBegin() {
if (mListeners != null) {
mListeners.notifyVideoFrameADBegin();
}
}
public void notifyRemoveAndHideDelayVideoList() {
mListeners.removeAndHideDelayVideoList();
}
public void notifyShowMoreOprationPanel() {
mListeners.notifyShowMoreOprationPanel();
}
public void notifyHideMoreOprationPanel() {
mListeners.notifyHideMoreOprationPanel();
}
public void notifyRemoveAndHideDelayMoreOprationPanel() {
mListeners.notifyRemoveAndHideDelayMoreOprationPanel();
}
/**
* 通知屏幕旋转
*
* @param vertical
* true 垂直;false 水平
*/
public void notifyScreenOrientationChange(boolean vertical) {
mListeners.notifyScreenOrientationChange(vertical);
}
/**
* 通知隐藏底部控制区
*/
public void notifyHideBottomControllerBar() {
mListeners.notifyHideBottomControllerBar();
}
/**
* 通知显示底部控制区
*/
public void notifyShowBottomControllerBar() {
mListeners.notifyShowBottomControllerBar();
}
/**
* 通知隐藏顶部控制区
*/
public void notifyHideTopControllerBar() {
mListeners.notifyHideTopControllerBar();
}
/**
* 通知显示顶部控制区
*/
public void notifyShowTopControllerBar() {
mListeners.notifyShowTopControllerBar();
}
/**
* 通知不隐藏控制区
*/
public void notifyNotHideControllerBar() {
mListeners.notifyNotHideControllerBar();
}
/**
* 通知隐藏控制区
*
* @param delay
* 延时时间
*/
public void notifyHideControllerBar(long delay) {
mListeners.notifyHideControllerBar(delay);
}
/**
* 通知显示控制区
*
* @param delayHide
* 延时时间
*/
public void notifyShowControllerBar(boolean delayHide) {
mListeners.notifyShowControllerBar(delayHide);
}
/**
* 通知控制区需提前隐藏内容
*/
public void notifyControllerBarPreHide() {
mListeners.notifyControllerBarPreHide();
}
/**
* 通知控制区需提前显示内容
*/
public void notifyControllerBarPreShow() {
mListeners.notifyControllerBarPreShow();
}
/**
* 通知控制区需延时隐藏内容
*/
public void notifyControllerBarPostHide() {
mListeners.notifyControllerBarPostHide();
}
/**
* 通知控制区需延时显示内容
*/
public void notifyControllerBarPostShow() {
mListeners.notifyControllerBarPostShow();
}
/**
* 通知TV中导航键中左键右键的操作
*
* @param keyLeft
* true 表示导航左键;false 表示导航右键
*/
public void notifyKeyChangeProgress(boolean keyLeft) {
mListeners.notifyKeyChangeProgress(keyLeft);
}
/**
* 通知视频播放进度
*
* @param current
* 当前播放时间
* @param duration
* 总时长
*/
public void notifyProgressUpdate(long current, long duration) {
mListeners.notifyProgressUpdate(current, duration);
}
public void notifySoundChanged(final int curr) {
mListeners.notifySoundChanged(curr);
}
public void notifyLightingSetting(final float curr) {
mListeners.notifyLightingSetting(curr);
}
public void notifyOnShowHideADContainer(final boolean isShow) {
mListeners.notifyOnShowHideADContainer(isShow);
}
public void notifyResolutionListButtonFocusFirst() {
mListeners.notifyResolutionListButtonFirstFocus();
}
// --------控制方法区end-----------//
// --------回调方法区begin-----------//
/**
* 添加缓冲更新回调
*
* @param l
* 回调接口
*/
public void addOnBufferingUpdateListener(
VDVideoViewListeners.OnBufferingUpdateListener l) {
mListeners.addOnBufferingUpdateListener(l);
}
/**
* 移除缓冲更新回调
*
* @param l
* 回调接口
*/
public void removeOnBufferingUpdateListener(
VDVideoViewListeners.OnBufferingUpdateListener l) {
mListeners.removeOnBufferingUpdateListener(l);
}
/**
* 添加加载完成回调
*
* @param l
* 回调接口
*/
public void addOnCompletionListener(
VDVideoViewListeners.OnCompletionListener l) {
mListeners.addOnCompletionListener(l);
}
/**
* 移除加载完成回调
*
* @param l
* 回调接口
*/
public void removeOnCompletionListener(
VDVideoViewListeners.OnCompletionListener l) {
mListeners.removeOnCompletionListener(l);
}
/**
* 添加错误回调
*
* @param l
* 回调接口
*/
public void addOnErrorListener(VDVideoViewListeners.OnErrorListener l) {
mListeners.addOnErrorListener(l);
}
/**
* 移除错误回调
*
* @param l
* 回调接口
*/
public void removeOnErrorListener(VDVideoViewListeners.OnErrorListener l) {
mListeners.removeOnErrorListener(l);
}
/**
* 添加重试错误回调
*
* @param l
* 回调接口
*/
public void addOnRetryErrorListener(VDVideoViewListeners.OnErrorListener l) {
mListeners.addOnRetryErrorListener(l);
}
/**
* 移除重试错误回调
*
* @param l
* 回调接口
*/
public void removeOnRetryErrorListener(
VDVideoViewListeners.OnErrorListener l) {
mListeners.removeOnRetryErrorListener(l);
}
/**
* 添加信息回调
*
* @param l
* 回调接口
*/
public void addOnInfoListener(VDVideoViewListeners.OnInfoListener l) {
mListeners.addOnInfoListener(l);
}
/**
* 移除信息回调
*
* @param l
* 回调接口
*/
public void removeOnInfoListener(VDVideoViewListeners.OnInfoListener l) {
mListeners.removeOnInfoListener(l);
}
/**
* 添加界面刷新回调
*
* @param l
* 回调接口
*/
public void addOnVideoUIRefreshListener(
VDVideoViewListeners.OnVideoUIRefreshListener l) {
mListeners.addOnVideoUIRefreshListener(l);
}
/**
* 移除界面刷新回调
*
* @param l
* 回调接口
*/
public void removeOnVideoUIRefreshListener(
VDVideoViewListeners.OnVideoUIRefreshListener l) {
mListeners.removeOnVideoUIRefreshListener(l);
}
/**
* 添加视频准备就绪回调
*
* @param l
* 回调接口
*/
public void addOnPreparedListener(VDVideoViewListeners.OnPreparedListener l) {
mListeners.addOnPreparedListener(l);
}
/**
* 移除视频准备就绪回调
*
* @param l
* 回调接口
*/
public void removeOnPreparedListener(
VDVideoViewListeners.OnPreparedListener l) {
mListeners.removeOnPreparedListener(l);
}
/**
* 添加调节视频进度完毕回调
*
* @param l
* 回调接口
*/
public void addOnSeekCompleteListener(
VDVideoViewListeners.OnSeekCompleteListener l) {
mListeners.addOnSeekCompleteListener(l);
}
/**
* 移除添加视频进度完毕回调
*
* @param l
* 回调接口
*/
public void removeOnSeekCompleteListener(
VDVideoViewListeners.OnSeekCompleteListener l) {
mListeners.removeOnSeekCompleteListener(l);
}
/**
* 添加视频大小改变回调
*
* @param l
* 回调接口
*/
public void addOnVideoSizeChangedListener(
VDVideoViewListeners.OnVideoSizeChangedListener l) {
mListeners.addOnVideoSizeChangedListener(l);
}
/**
* 移除视频大小改变回调
*
* @param l
* 回调接口
*/
public void removeOnVideoSizeChangedListener(
VDVideoViewListeners.OnVideoSizeChangedListener l) {
mListeners.removeOnVideoSizeChangedListener(l);
}
/**
* 添加打开视频回调
*
* @param l
* 回调接口
*/
public void addOnVideoOpenedListener(
VDVideoViewListeners.OnVideoOpenedListener l) {
mListeners.addOnVideoOpenedListener(l);
}
/**
* 移除打开视频回调
*
* @param l
* 回调接口
*/
public void removeOnVideoOpenedListener(
VDVideoViewListeners.OnVideoOpenedListener l) {
mListeners.removeOnVideoOpenedListener(l);
}
/**
* 添加视频进度更新回调
*
* @param l
* 回调接口
*/
public void addOnProgressUpdateListener(
VDVideoViewListeners.OnProgressUpdateListener l) {
mListeners.addOnProgressUpdateListener(l);
}
/**
* 移除视频进度更新回调
*
* @param l
* 回调接口
*/
public void removeOnProgressUpdateListener(
VDVideoViewListeners.OnProgressUpdateListener l) {
mListeners.removeOnProgressUpdateListener(l);
}
/**
* 添加时间信息更新回调
*
* @param l
* 回调接口
*/
public void addOnTimedTextListener(
VDVideoViewListeners.OnTimedTextListener l) {
mListeners.addOnTimedTextListener(l);
}
/**
* 移除时间信息更新回调
*
* @param l
* 回调接口
*/
public void removeOnTimedTextListener(
VDVideoViewListeners.OnTimedTextListener l) {
mListeners.removeOnTimedTextListener(l);
}
/**
* @param l
*/
public void addOnPlayVideoListener(
VDVideoViewListeners.OnPlayVideoListener l) {
mListeners.addOnPlayVideoListener(l);
}
public void removeOnPlayVideoListener(
VDVideoViewListeners.OnPlayVideoListener l) {
mListeners.removeOnPlayVideoListener(l);
}
// public void
// addOnLiveVideoListener(VDVideoViewListeners.OnLiveVideoListener l) {
// mListeners.addOnLiveVideoListener(l);
// }
//
// public void
// removeOnLiveVideoListener(VDVideoViewListeners.OnLiveVideoListener l) {
// mListeners.removeOnLiveVideoListener(l);
// }
/**
* 添加全屏回调
*
* @param l
* 回调接口
*/
public void addOnFullScreenListener(
VDVideoViewListeners.OnFullScreenListener l) {
mListeners.addOnFullScreenListener(l);
}
/**
* 移除全屏回调
*
* @param l
* 回调接口
*/
public void removeOnFullScreenListener(
VDVideoViewListeners.OnFullScreenListener l) {
mListeners.removeOnFullScreenListener(l);
}
/**
* 添加音量回调
*
* @param l
* 回调接口
*/
public void addOnSoundChangedListener(
VDVideoViewListeners.OnSoundChangedListener l) {
mListeners.addOnSoundChangedListener(l);
}
/**
* 移除音量回调
*
* @param l
* 回调接口
*/
public void removeOnSoundChangedListener(
VDVideoViewListeners.OnSoundChangedListener l) {
mListeners.removeOnSoundChangedListener(l);
}
/**
* 添加触屏回调
*
* @param l
* 回调接口
*/
public void addOnScreenTouchListener(
VDVideoViewListeners.OnScreenTouchListener l) {
mListeners.addOnScreenTouchListener(l);
}
/**
* 移除触屏回调
*
* @param l
* 回调接口
*/
public void removeOnScreenTouchListener(
VDVideoViewListeners.OnScreenTouchListener l) {
mListeners.removeOnScreenTouchListener(l);
}
/**
* 添加清晰度选择回调
*
* @param l
* 回调接口
*/
public void addOnResolutionListener(
VDVideoViewListeners.OnResolutionListener l) {
mListeners.addOnResolutionListener(l);
}
/**
* 移除清晰度选择回调
*
* @param l
* 回调接口
*/
public void removeOnResolutionContainerListener(
VDVideoViewListeners.OnResolutionContainerListener l) {
mListeners.removeOnResolutionContainerListener(l);
}
/**
* 添加清晰度容器回调
*
* @param l
* 回调接口
*/
public void addOnResolutionContainerListener(
VDVideoViewListeners.OnResolutionContainerListener l) {
mListeners.addOnResolutionContainerListener(l);
}
/**
* 移除清晰度容器回调
*
* @param l
* 回调接口
*/
public void removeOnResolutionListener(
VDVideoViewListeners.OnResolutionListener l) {
mListeners.removeOnResolutionListener(l);
}
/**
* 添加调节亮度消息
*
* @param l
* 回调接口
*/
public void addOnLightingChangeListener(
VDVideoViewListeners.OnLightingChangeListener l) {
mListeners.addOnLightingChangeListener(l);
}
/**
* 移除调节亮度消息
*
* @param l
* 回调接口
*/
public void removeOnLightingChangeListener(
VDVideoViewListeners.OnLightingChangeListener l) {
mListeners.removeOnLightingChangeListener(l);
}
/**
* 添加tip通知
*
* @param l
* 回调接口
*/
public void addOnTipListener(OnTipListener l) {
mListeners.addOnTipListener(l);
}
/**
* 移除tip通知
*
* @param l
* 回调接口
*/
public void removeOnTipListener(OnTipListener l) {
mListeners.removeOnTipListener(l);
}
/**
* 添加亮度调节回调
*
* @param l
* 回调接口
*/
public void addOnLightingVisibleListener(OnLightingVisibleListener l) {
mListeners.addOnLightingVisibleListener(l);
}
/**
* 移除亮度添加回调
*
* @param l
* 回调接口
*/
public void removeOnLightingVisibleListener(OnLightingVisibleListener l) {
mListeners.removeOnLightingVisibleListener(l);
}
/**
* 添加进度条控件显示回调
*
* @param l
* 回调接口
*/
public void addOnProgressViewVisibleListener(OnProgressViewVisibleListener l) {
mListeners.addOnProgressViewVisibleListener(l);
}
/**
* 移除进度条控件显示回调
*
* @param l
* 回调接口
*/
public void removeOnProgressViewVisibleListener(
OnProgressViewVisibleListener l) {
mListeners.removeOnProgressViewVisibleListener(l);
}
/**
* 添加声音控件显示回调
*
* @param l
* 回调接口
*/
public void addOnSoundVisibleListener(OnSoundVisibleListener l) {
mListeners.addOnSoundVisibleListener(l);
}
/**
* 移除声音控件显示回调
*
* @param l
* 回调接口
*/
public void removeOnSoundVisibleListener(OnSoundVisibleListener l) {
mListeners.removeOnSoundVisibleListener(l);
}
/**
* 添加清晰度控件回调
*
* @param l
* 回调接口
*/
public void addOnVMSResolutionListener(OnVMSResolutionListener l) {
mListeners.addOnVMSResolutionListener(l);
}
/**
* 移除清晰度控件回调
*
* @param l
* 回调接口
*/
public void removeOnVMSResolutionListener(OnVMSResolutionListener l) {
mListeners.removeOnVMSResolutionListener(l);
}
/**
* 添加加载中回调
*
* @param l
* 回调接口
*/
public void addOnLoadingListener(OnLoadingListener l) {
mListeners.addOnLoadingListener(l);
}
/**
* 移除加载中回调
*
* @param l
* 回调接口
*/
public void removeOnLoadingListener(OnLoadingListener l) {
mListeners.removeOnLoadingListener(l);
}
/**
* 添加视频引导页回调
*
* @param l
* 回调接口
*/
public void addOnVideoGuideTipsListener(OnVideoGuideTipsListener l) {
mListeners.addOnVideoGuideTipsListener(l);
}
/**
* 移除视频引导页回调
*
* @param l
* 回调接口
*/
public void removeOnVideoGuideTipsListener(OnVideoGuideTipsListener l) {
mListeners.removeOnVideoGuideTipsListener(l);
}
/**
* 添加插入视频广告回调
*
* @param l
*/
public void addOnVideoInsertADListener(OnVideoInsertADListener l) {
mListeners.addOnVideoInsertADListener(l);
}
/**
* 移除插入视频广告回调
*
* @param l
*/
public void removeOnVideoInsertADListener(OnVideoInsertADListener l) {
mListeners.removeOnVideoInsertADListener(l);
}
/**
* 添加帧间视频广告回调
*
* @param l
* 回调接口
*/
public void addOnVideoFrameADListener(OnVideoFrameADListener l) {
mListeners.addOnVideoADListener(l);
}
/**
* 移除帧间视频广告回调
*
* @param l
* 回调接口
*/
public void removeOnVideoFrameADListener(OnVideoFrameADListener l) {
mListeners.removeOnVideoAdListener(l);
}
/**
* 添加视频列表回调
*
* @param l
* 回调接口
*/
public void addOnVideoListListener(OnVideoListListener l) {
mListeners.addOnVideoListListener(l);
}
/**
* 移除视频列表回调
*
* @param l
* 回调接口
*/
public void removeOnVideoListListener(OnVideoListListener l) {
mListeners.removeOnVideoListListener(l);
}
/**
* 添加视频列表可见状态改变回调
*
* @param l
* 回调接口
*/
public void addOnVideoListVisibleChangeListener(
OnVideoListVisibleChangeListener l) {
mListeners.addOnVideoListVisibleChangeListener(l);
}
/**
* 移除视频列表可见状态改变回调
*
* @param l
* 回调接口
*/
public void removeOnVideoListVisibleChangeListener(
OnVideoListVisibleChangeListener l) {
mListeners.removeOnVideoListVisibleChangeListener(l);
}
public void addOnMoreOprationVisibleChangeListener(
OnMoreOprationVisibleChangeListener l) {
mListeners.addOnMoreOprationVisibleChangeListener(l);
}
public void removeOnMoreOprationVisibleChangeListener(
OnMoreOprationVisibleChangeListener l) {
mListeners.removeOnMoreOprationVisibleChangeListener(l);
}
// public void
// addOnResolutionVisibleChangeListener(OnResolutionVisibleChangeListener l)
// {
// mListeners.addOnResolutionVisibleChangeListener(l);
// }
//
// public void
// removeOnResolutionVisibleChangeListener(OnResolutionVisibleChangeListener
// l) {
// mListeners.removeOnResolutionVisibleChangeListener(l);
// }
/**
* 添加双击回调
*
* @param l
* 回调接口
*/
public void addOnVideoDoubleTapListener(OnVideoDoubleTapListener l) {
mListeners.addOnVideoDoubleTapListener(l);
}
/**
* 移除双加回调
*
* @param l
* 回调接口
*/
public void removeOnVideoDoubleTapListener(OnVideoDoubleTapListener l) {
mListeners.removeOnVideoDoubleTapListener(l);
}
/**
* 添加解码类型改变回调
*
* @param l
* 回调接口
*/
public void addOnDecodingTypeListener(OnDecodingTypeListener l) {
mListeners.addOnDecodingTypeListener(l);
}
/**
* 移除解码类型改变回调
*
* @param l
* 回调接口
*/
public void removeOnDecodingTypeListener(OnDecodingTypeListener l) {
mListeners.removeOnDecodingTypeListener(l);
}
/**
* 添加屏幕方向改变回调
*
* @param l
* 回调接口
*/
public void addOnScreenOrientationChangeListener(
OnScreenOrientationChangeListener l) {
mListeners.addOnScreenOrientationChangeListener(l);
}
/**
* 移除屏幕方向改变回调
*
* @param l
* 回调接口
*/
public void removeOnScreenOrientationChangeListener(
OnScreenOrientationChangeListener l) {
mListeners.removeOnScreenOrientationChangeListener(l);
}
/**
* 添加控制区显示状态改变回调
*
* @param l
* 回调接口
*/
public void addOnShowHideControllerListener(OnShowHideControllerListener l) {
mListeners.addOnShowHideControllerListener(l);
}
/**
* 移除控制区显示状态改变回调
*
* @param l
* 回调接口
*/
public void removeOnShowHideControllerListener(
OnShowHideControllerListener l) {
mListeners.removeOnShowHideControllerListener(l);
}
/**
* 添加底部控制区显示状态改变回调
*
* @param l
* 回调接口
*/
public void addOnShowHideBottomControllerListener(
OnShowHideBottomControllerListener l) {
mListeners.addOnShowHideBottomControllerListener(l);
}
/**
* 移除底部控制区显示状态改变回调
*
* @param l
* 回调接口
*/
public void removeOnShowHideTopControllerListener(
OnShowHideTopContainerListener l) {
mListeners.removeOnShowHideTopContainerListener(l);
}
/**
* 添加底部控制区显示状态改变回调
*
* @param l
* 回调接口
*/
public void addOnShowHideTopControllerListener(
OnShowHideTopContainerListener l) {
mListeners.addOnShowHideTopContainerListener(l);
}
/**
* 移除底部控制区显示状态改变回调
*
* @param l
* 回调接口
*/
public void removeOnShowHideBottomControllerListener(
OnShowHideBottomControllerListener l) {
mListeners.removeOnShowHideBottomControllerListener(l);
}
/**
* 添加点击播放按钮回调
*
* @param l
* 回调接口
*/
public void addOnClickPlayListener(OnClickPlayListener l) {
mListeners.addOnClickPlayListener(l);
}
/**
* 移除点击播放按钮回调
*
* @param l
* 回调接口
*/
public void removeOnClickPlayListener(OnClickPlayListener l) {
mListeners.removeOnClickPlayListener(l);
}
/**
* 添加导航键改变播放进度回调
*
* @param l
* 回调接口
*/
public void addOnKeyChangeProgressListener(OnKeyChangeProgressListener l) {
mListeners.addOnKeyChangeProgressListener(l);
}
/**
* 移除导航键改变播放进度回调
*
* @param l
* 回调接口
*/
public void removeOnKeyChangeProgressListener(OnKeyChangeProgressListener l) {
mListeners.removeOnKeyChangeProgressListener(l);
}
/**
* 添加按键事件回调
*
* @param l
* 回调接口
*/
public void addOnKeyEventListener(OnKeyEventListener l) {
mListeners.addOnKeyEventListener(l);
}
/**
* 移除按键事件回调
*
* @param l
* 回调接口
*/
public void removeOnKeyEventListener(OnKeyEventListener l) {
mListeners.removeOnKeyEventListener(l);
}
/**
* 添加DLNA布局回调
*
* @param l
* 回调接口
*/
public void addOnDLNALinearLayoutListener(OnDLNALinearLayoutListener l) {
mListeners.addOnDLNALinearLayoutListener(l);
}
/**
* 移除DLNA布局回调
*
* @param l
* 回调接口
*/
public void removeOnDLNALinearLayoutListener(OnDLNALinearLayoutListener l) {
mListeners.removeOnDLNALinearLayoutListener(l);
}
/**
* 添加设置音量回调
*
* @param l
* 回调接口
*/
public void addOnSetSoundListener(OnSetSoundListener l) {
mListeners.addOnSetSoundListener(l);
}
/**
* 移除设置音量回调
*
* @param l
* 回调接口
*/
public void removeOnSetSoundListener(OnSetSoundListener l) {
mListeners.removeOnSetSoundListener(l);
}
/**
* 添加注册DLNA回调
*
* @param l
* 回调接口
*/
public void addOnRegisterDLNAListener(OnRegisterDLNAListener l) {
mListeners.addOnRegisterDLNAListener(l);
}
/**
* 移除注册DLNA回调
*
* @param l
* 回调接口
*/
public void removeOnRegisterDLNAListener(OnRegisterDLNAListener l) {
mListeners.removeOnRegisterDLNAListener(l);
}
/**
* 添加切换屏幕方向回调
*
* @param l
* 回调接口
*/
public void addOnScreenOrientationSwitchListener(
OnScreenOrientationSwitchListener l) {
mListeners.addOnScreenOrientationSwitchListener(l);
}
/**
* 移除切换屏幕方向回调
*
* @param l
* 回调接口
*/
public void removeOnScreenOrientationSwitchListener(
OnScreenOrientationSwitchListener l) {
mListeners.removeOnScreenOrientationSwitchListener(l);
}
/**
* 添加点击重试回调
*
* @param l
* 回调接口
*/
public void addOnClickRetryListener(OnClickRetryListener l) {
mListeners.addOnClickRetryListener(l);
}
/**
* 移除贴片广告的容器显示隐藏类
*
* @param l
* 回调接口
*/
public void removeOnShowHideADContainerListener(
OnShowHideADContainerListener l) {
mListeners.removeOnShowHideADContainerListener(l);
}
/**
* 添加贴片广告的容器显示隐藏类
*
* @param l
* 回调接口
*/
public void addOnShowHideADContainerListener(OnShowHideADContainerListener l) {
mListeners.addOnShowHideADContainerListener(l);
}
/**
* 移除点击重试回调
*
* @param l
* 回调接口
*/
public void removeOnClickRetryListener(OnClickRetryListener l) {
mListeners.removeOnClickRetryListener(l);
}
public void addOnResolutionListButtonListener(
OnResolutionListButtonListener l) {
mListeners.addOnResolutionListButtonListener(l);
}
public void removeOnResolutionListButtonListener(
OnResolutionListButtonListener l) {
mListeners.removeOnResolutionListButtonListener(l);
}
// --------回调方法区end-----------//
/**
* 发送页面刷新通知,尤其用于广告层里面
*/
public void notifyVideoUIRefresh() {
mListeners.notifyVideoUIRefreshListener();
}
/**
* 点击重试通知
*/
public void notifyClickRetry() {
mListeners.notifyClickRetry();
}
/**
* 屏幕方向改变通知
*
* @param fullScreen
* true 全屏;false 非全屏
*/
public void notifyScreenOrientationSwitch(boolean fullScreen) {
mListeners.notifyScreenOrientationSwitch(fullScreen);
}
/**
* DLNA视图可见通知
*
* @param visiable
* true 可见;false 不可见
*/
public void notifySetDLNALayoutVisible(boolean visiable) {
mListeners.notifySetDLNALayoutVisible(visiable);
}
/**
* DLNA注册通知
*/
public void notifyRegisterDLNAListener() {
mListeners.notifyRegisterDLNAListener();
}
/**
* 设置音量通知
*
* @param currVolume
* 当前音量
*/
public void notifySetCurVolume(int currVolume) {
mListeners.notifySetCurVolume(currVolume);
}
/**
* 设置最大音量通知
*
* @param maxVolume
* 最大音量
*/
public void notifySetMaxVolume(int maxVolume) {
mListeners.notifySetMaxVolume(maxVolume);
}
/**
* 通知前贴片广告的开始与结束
*
* @param isBegin
*/
public void notifyInsertAD(boolean isBegin) {
if (isBegin) {
mListeners.notifyVideoInsertADBegin();
} else {
mListeners.notifyVideoInsertADEnd();
}
}
/**
* 前贴片跳秒
*/
public void notifyInsertADTicker() {
mListeners.notifyVideoInsertADTicker();
}
/**
* 浮框通知
*
* @param tip
*/
public void notifyTip(String tip) {
mListeners.notifyTip(tip);
}
/**
* 隐藏浮框通知
*/
public void notifyHideTip() {
mListeners.notifyHideTip();
}
@Override
public void onVideoOpened(MediaPlayer mp) {
mListeners.notifyVideoOpened();
}
@Override
public void onVideoSizeChanged(MediaPlayer mp, int width, int height) {
mListeners.notifyVideoSizeChanged(width, height);
}
@Override
public void onTimedText(MediaPlayer mp, TimedText text) {
mListeners.notifyTimedText(text);
}
@Override
public void onPrepared(MediaPlayer mp) {
mVDPlayerInfo.mPlayStatus = VDPlayerInfo.PLAYER_PREPARED;
if (mVideoView == null) {
return;
}
mVDPlayerInfo.mDuration = mVideoView.getDuration();
VDVideoViewController controller = VDVideoViewController
.getInstance(this.mContext);
if (controller != null)
controller.notifyShowControllerBar(true);
if (controller != null)
controller.notifyOnShowHideADContainer(true);
startUpdateMessage();
VDVideoInfo info = getCurrentVideo();
if (info != null) {
info.mVideoDuration = mp.getDuration();
// 补全当前广告部分的时长,只要秒就可以了
if (info.mIsInsertAD) {
// 广告的 buffer size 应当很小
// mp.setBufferSize(1000); //有影响反而卡顿了
mADIsFromStart = true;
int duration = (int) Math
.floor((double) mp.getDuration() / (1000));
if (!mVDVideoListInfo.mIsSetInsertADTime) {
// 只有单广告流的时候,才重置当前时间,如果是多流情况,需要手动设置
mVDVideoListInfo.mInsertADSecNum = duration;
mADTickerSecNum = mVDVideoListInfo.mInsertADSecNum;
}
getCurrentVideo().mInsertADSecNum = duration;
notifyInsertAD(true);
} else {
// 正片部分,将音量调回正常来
boolean isCanNormalSound = false;
int key = mVDVideoListInfo.getVideoInfoKey(info) - 1;
try {
if (key >= 0
&& mVDVideoListInfo.getVideoInfo(key).mIsInsertAD) {
isCanNormalSound = true;
}
} catch (Exception ex) {
VDLog.e(TAG, ex.getMessage());
}
if (VDPlayerSoundManager.isMuted(mContext) && isCanNormalSound) {
VDPlayerSoundManager.setMute(mContext, false, false);
}
}
// 帧间广告部分
if (isCanShowFrameAD()) {
mExtListeners.notifyFrameADListenerPrepared();
}
}
mListeners.notifyPrepared();
mListeners.notifyVideoPrepared(true);
mExtListeners.notifyPreparedListener();
mVideoView.start();
updatePlayState();
mIsPlayed = false;
if (info != null && info.mIsLive
&& VDVideoInfo.SOURCE_TYPE_FAKE_LIVE.equals(info.mSourceType)) {
// m3u8伪直播充当真直播时,变换清晰度时,需跳转到服务器端的当前直播位置。
long seekTo = info.getSeekTo();
if (seekTo > mp.getDuration()) {
seekTo = 0L;
}
seekTo(seekTo);
} else if (info != null && info.mNeedSeekTo) {
info.mNeedSeekTo = false;
seekTo(info.mVideoPosition);
}
}
@Override
public boolean onInfo(MediaPlayer mp, int what, int extra) {
mListeners.notifyInfo(what, extra);
if (getCurrentVideo() != null) {
mExtListeners.notifyInfoListener(getCurrentVideo(), what);
}
VDLog.i("demo", "onInfo --> what :" + what + " , extra : " + extra);
if (what == MediaPlayer.MEDIA_INFO_BUFFERING_START) { // MediaPlayer暂停播放等待缓冲更多数据。
// NOTE: video playing quickly bug if pause added here ,by lyh
notifyShowLoading();
VDLog.i("demo", "onInfo --> MEDIA_INFO_BUFFERING_START");
mIsPlayed = true;
mTimeOutHandler.removeMessages(NET_TIME_OUT);
if (VDApplication.getInstance().isNetworkConnected()) {// 有网
mTimeOutHandler.sendEmptyMessageDelayed(NET_TIME_OUT,
VIDEO_TIME_OUT);
} else {// 无网
}
} else if (what == MediaPlayer.MEDIA_INFO_BUFFERING_END) { // MediaPlayer在缓冲完后继续播放。
VDLog.i("demo", "onInfo --> MEDIA_INFO_BUFFERING_END");
mVDPlayerInfo.mPlayStatus = VDPlayerInfo.PLAYER_STARTED;
mTimeOutHandler.removeMessages(NET_TIME_OUT);
notifyHideLoading();
mListeners.notifyHideLoading();
mListeners.notifyHideTip();
// 将前贴片广告的启动点放置到这儿,首帧部分,以规避踹你的计时问题
if (getCurrentVideo().mIsInsertAD && mADIsFromStart) {
mInsertADHandler.postDelayed(mInsertADRunnable, 1000);
mADIsFromStart = false;
}
} else if (what == MediaPlayer.MEDIA_INFO_VIDEO_TRACK_LAGGING) { // 视频过于复杂,无法解码:不能快速解码帧。此时可能只能正常播放音频。
} else if (what == MediaPlayer.MEDIA_ERROR_UNKNOWN) { // 播放错误,未知错误。
}
return false;
}
@Override
public boolean onError(MediaPlayer mp, int what, int extra) {
VDLog.e(TAG, "onError:errorCode1:" + what + ",errorCodeExtra:" + extra);
if (getCurrentVideo() != null) {
if (getCurrentVideo().mIsInsertAD) {
mInsertADHandler.removeCallbacks(mInsertADRunnable);
}
}
mVDPlayerInfo.mPlayStatus = VDPlayerInfo.PLAYER_ERROR;
if (!mIsPlayed) {
mRetryTimes++;
if (mRetryTimes > VDSDKConfig.getInstance().getRetryTime()) {// 重试两次
mRetryTimes = 0;
if (getCurrentVideo() != null) {
// TODO 这个有点问题,只能记录最后一次的
if (mVDVideoListInfo.isInsertADEnd()) {
mExtListeners
.notifyInsertADEnd(VDPlayerErrorInfo.MEDIA_INSERTAD_ERROR_UNKNOWN);
}
if (getCurrentVideo().mIsInsertAD) {
playNext();
} else {
mListeners.notifyRetryError(what, extra);
}
mExtListeners.ontifyErrorListener(getCurrentVideo(), what,
extra);
}
} else {
mMessageHandler.postDelayed(new Runnable() {
@Override
public void run() {
playVideoOnInfoKey(mVDVideoListInfo.mIndex);
VDLog.e(TAG, "avformat_open_input 视频错误2秒重试");
}
}, 2000);
}
} else {
mListeners.notifyError(what, extra);
if (getCurrentVideo() != null) {
mExtListeners.ontifyErrorListener(getCurrentVideo(), what,
extra);
}
mRetryTimes = 0;
}
return false;
}
@Override
public void onCompletion(MediaPlayer mp) {
mVDPlayerInfo.mPlayStatus = VDPlayerInfo.PLAYER_FINISHED;
if (mVDVideoListInfo != null && mVDVideoListInfo.isInsertADEnd()) {
mExtListeners
.notifyInsertADEnd(VDPlayerErrorInfo.MEDIA_INSERTAD_SUCCESS);
}
VDVideoViewController controller = VDVideoViewController
.getInstance(this.mContext);
VDPlayerInfo playerInfo = null;
if (controller != null)
playerInfo = controller.getPlayerInfo();
if (playerInfo != null) {
playerInfo.mCurrent = playerInfo.mDuration;
}
if (!getCurrentVideo().mIsInsertAD) {
mListeners.notifyCompletion();
mExtListeners.notifyCompletionListener(getCurrentVideo(), 0);
} else {
notifyInsertAD(false);
if (mVDVideoListInfo.getADNumOfRemain() > 0) {
// 当前流中还有广告,那么暂停一下,继续开始
mInsertADHandler.removeCallbacks(mInsertADRunnable);
}
}
VDVideoInfo info = getCurrentVideo();
if (info != null) {
info.mVideoPosition = 0;
}
// mVDPlayerInfo.mDuration = 0;
long position = mVideoView.getCurrentPosition();
long duration = mVideoView.getDuration();
mListeners.notifyProgressUpdate(position, duration);
stopUpdateMessage();
updatePlayState();
if (getCurrentVideo().mIsInsertAD) {
playNext();
}
}
@Override
public void onBufferingUpdate(MediaPlayer mp, int percent) {
VDLog.d("onBufferingUpdate", "percent:" + percent);
mListeners.notifyBufferingUpdate(percent);
if (mLastPercent != percent) {
mLastPercentTime = System.currentTimeMillis();
mLastPercent = percent;
mTimeOutHandler.removeMessages(CHECK_LIVE_TIME_OUT);
} else {
try {
if (getCurrentVideo().mIsLive && mVideoView.isBuffering()) {
mTimeOutHandler.sendEmptyMessageDelayed(
CHECK_LIVE_TIME_OUT, CHECK_LIVE_TIME_GAP);
}
} catch (IllegalStateException e) {
e.printStackTrace();
}
}
}
/**
* m3u8解析回调类
*
* @author sina
*/
class MyM3u8ParserListener implements M3u8ParserListener {
private VDVideoInfo mVideoInfo;
public MyM3u8ParserListener(VDVideoInfo _vi) {
mVideoInfo = _vi;
}
@Override
public void onParcelResult(String url, VDResolutionData resolutionData) {
if (resolutionData != null) {
VDResolutionManager resolutionManager = VDResolutionManager
.getInstance(mContext);
if (resolutionManager != null) {
notifyResolutionParsed(resolutionData);
resolutionManager.setResolutionData(resolutionData);
String tag = resolutionManager.getCurrResolutionTag();
setVideoPath(mVideoInfo.getVideoUrl(tag));
if (mListeners != null) {
mListeners.notifyResolutionParsed(resolutionData);
}
} else {
setVideoPath(mVideoInfo
.getVideoUrl(VDResolutionData.TYPE_DEFINITION_SD));
}
} else {
if (mVideoView != null) {
setVideoPath(mVideoInfo.mPlayUrl);
}
}
if (mVideoView != null) {
setVideoViewVisible(View.VISIBLE);
}
notifyShowLoading();
}
@Override
public void onError(int error_msg) {
mListeners.notifyRetryError(0, 0);
if (getCurrentVideo() != null) {
mExtListeners.ontifyErrorListener(getCurrentVideo(),
VDPlayerErrorInfo.MEDIA_ERROR_WHAT_M3U8_PARSER,
error_msg);
}
}
@Override
public void updateVideoPlayUrl(String playUrl) {
}
@Override
public void updateVideoID(String videoId) {
// @sunxiao1
// 看了一下M3u8ParserListener的实现,发现,updateVideoID在构造后直接丢回来,那么这个操作有毛用?
// VDSDKLogData.getInstance().mLogVideoInfo.mVideoId = videoId;
}
}
/**
* 设置视频控件是否可见
*
* @param isVisible
* eg:VISIBLE, INVISIBLE, or GONE.
*/
private void setVideoViewVisible(int isVisible) {
if (mVideoView != null) {
if (mVideoView instanceof VideoViewHard) {
((VideoViewHard) mVideoView).setVisibility(isVisible);
} else {
((VideoView) mVideoView).setVisibility(isVisible);
}
}
}
private static long VIDEO_TIME_OUT = 30 * 1000;
private static final long VIDEO_NORMAL_TIME_OUT = 30 * 1000;
private static final long VIDEO_LIVE_TIME_OUT = 40 * 1000;
private static final long CHECK_LIVE_TIME_GAP = 30 * 1000;
private static final int NET_TIME_OUT = 1;
private static final int CHECK_LIVE_TIME_OUT = 3;
private int mBufferPercent;
private int mLastPercent;
private long mLastPercentTime;
/**
* 处理时间设置的Handler
*/
private static class TimeOutHandler extends Handler {
WeakReference<VDVideoViewController> controller = null;
public TimeOutHandler(VDVideoViewController instance) {
super(Looper.getMainLooper());
controller = new WeakReference<VDVideoViewController>(instance);
}
@Override
public void handleMessage(Message msg) {
// TODO Auto-generated method stub
if (controller == null) {
return;
}
switch (msg.what) {
case NET_TIME_OUT:
int curPercent = controller.get().mVideoView
.getBufferPercentage();
long now = System.currentTimeMillis();
if (Math.abs(curPercent - controller.get().mBufferPercent) < 1
|| (Math.abs(curPercent - controller.get().mLastPercent) < 1 && (now
- controller.get().mLastPercentTime >= VIDEO_TIME_OUT))) {
controller.get().mListeners
.notifyTip(R.string.net_exp_check_and_retry);
} else if (curPercent < 100 /* && !mVideoView.isPlaying() */
&& controller.get().mVideoView.isBuffering()) {
if (controller.get().getCurrentVideo().mIsLive) {
controller.get().mListeners
.notifyTip(R.string.net_no_good_retry);
VDResolutionManager resolutionManager = VDResolutionManager
.getInstance(controller.get().mContext);
if (resolutionManager != null) {
controller
.get()
.setVideoPath(
controller
.get()
.getCurrentVideo()
.getVideoUrl(
resolutionManager
.getCurrResolutionTag()));
}
} else {
controller.get().mListeners
.notifyTip(R.string.your_net_is_no_good);
sendEmptyMessageDelayed(NET_TIME_OUT, VIDEO_TIME_OUT);
}
controller.get().mBufferPercent = controller.get().mVideoView
.getBufferPercentage();
}
break;
case CHECK_LIVE_TIME_OUT:
controller.get().mListeners
.notifyTip(R.string.net_exp_check_and_retry);
break;
default:
break;
}
}
}
private TimeOutHandler mTimeOutHandler = new TimeOutHandler(this);
/**
* 隐藏状态栏
*
* @param flag
* true :隐藏 ;false: 不隐藏
*/
public void hideStatusBar(boolean flag) {
if (mContext == null) {
return;
}
if (VDUtility.isMeizu() || VDUtility.isSamsungNoteII()) {
return;
}
Window window = ((Activity) mContext).getWindow();
WindowManager.LayoutParams layoutparams = window
.getAttributes();
if (flag)
layoutparams.flags = WindowManager.LayoutParams.FLAG_FULLSCREEN
| layoutparams.flags;
else
layoutparams.flags = ~WindowManager.LayoutParams.FLAG_FULLSCREEN
& layoutparams.flags;
window.setAttributes(layoutparams);
}
@Override
public void onSeekComplete(MediaPlayer mp) {
startUpdateMessage();
}
public void onResume() {
resume();
}
public void onPause() {
pause();
notifyHideControllerBar(0);
}
public void onStart() {
if (getCurrentVideo() != null) {
if (getCurrentVideo().mIsInsertAD) {
start();
}
}
VDVideoFullModeController.getInstance().enableSensor(true);
}
public void onStartWithVideoResume() {
start();
VDVideoFullModeController.getInstance().enableSensor(true);
}
public void onStop() {
VDVideoFullModeController.getInstance().enableSensor(false);
}
public static class NetChangeListener
implements
VDNetworkBroadcastReceiver.NetworkNotifyListener {
private Context mContext = null;
public NetChangeListener(Context context) {
mContext = context;
}
@Override
public void wifiConnected() {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller == null) {
return;
}
}
@Override
public void mobileConnected() {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller == null) {
return;
}
}
@Override
public void nothingConnected() {
VDVideoViewController controller = VDVideoViewController
.getInstance(mContext);
if (controller == null) {
return;
}
File file = Environment.getExternalStorageDirectory();
VDVideoInfo mVideoInfo = controller.getCurrentVideo();
if (mVideoInfo != null && mVideoInfo.mPlayUrl != null
&& VDUtility.isSdcardReady() && file != null
&& mVideoInfo.mPlayUrl.startsWith(file.getAbsolutePath())) {
return;
}
controller.notifyTip("网络异常,请您检查网络后重试");
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/playlist/VDVideoPlaylistTextView.java
package com.sina.sinavideo.sdk.widgets.playlist;
import android.content.Context;
import android.content.res.TypedArray;
import android.graphics.Color;
import android.util.AttributeSet;
import android.widget.TextView;
import com.sina.sinavideo.sdk.data.VDVideoInfo;
import com.sina.sinavideo.sdk.widgets.VDBaseWidget;
import com.sina.sinavideo.sdk.R;
/**
* 全屏播放时相关视频列表中数据列中的文字内容
* @author liuqun
*/
public class VDVideoPlaylistTextView extends TextView implements VDBaseWidget, VDVideoPlaylistBase {
protected VDVideoInfo mInfo = null;
protected int mVideoInfoIndex = -1;
private int mCurPlayColor;
private int mNoPlayColor;
public VDVideoPlaylistTextView(Context context, AttributeSet attrs) {
super(context, attrs);
TypedArray a = context.obtainStyledAttributes(attrs, R.styleable.ResolutionBackGround);
int otherTextColor = Color.parseColor("#d1d1d1");
int curBgColor = Color.parseColor("#0078db");
mNoPlayColor = a.getColor(R.styleable.PlayListTextViewColor_NoPlayColor, otherTextColor);
mCurPlayColor = a.getColor(R.styleable.PlayListTextViewColor_CurPlayColor, curBgColor);
a.recycle();
}
@Override
public void reset() {
}
@Override
public void hide() {
}
@Override
public void setData(VDVideoInfo info) {
mInfo = info;
setText(info.mTitle);
}
@Override
public void setVideoInfo(int infoIndex, int curPlayIndex) {
mVideoInfoIndex = infoIndex;
if (mVideoInfoIndex == curPlayIndex) {
setTextColor(mCurPlayColor);
} else {
setTextColor(mNoPlayColor);
}
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/widgets/VDVideoPlaySeekBar.java
package com.sina.sinavideo.sdk.widgets;
import android.content.Context;
import android.content.res.TypedArray;
import android.util.AttributeSet;
import android.widget.SeekBar;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners;
import com.sina.sinavideo.sdk.data.VDPlayerInfo;
import com.sina.sinavideo.sdk.data.VDVideoInfo;
import com.sina.sinavideo.sdk.dlna.DLNAController;
import com.sina.sinavideo.sdk.R;
/**
* 视频进度拖动条
*
* @author liuqun
*
*/
public final class VDVideoPlaySeekBar extends SeekBar implements VDBaseWidget,
VDVideoViewListeners.OnProgressUpdateListener,
SeekBar.OnSeekBarChangeListener,
VDVideoViewListeners.OnBufferingUpdateListener {
private long mDuration;
public VDVideoPlaySeekBar(Context context) {
super(context);
setProgressDrawable(getResources().getDrawable(
R.drawable.play_seekbar_background));
setThumb(getResources().getDrawable(R.drawable.play_ctrl_sound_ball));
init();
}
public VDVideoPlaySeekBar(Context context, AttributeSet attrs) {
super(context, attrs);
TypedArray typedArr = context.obtainStyledAttributes(attrs,
new int[] { android.R.attr.progressDrawable });
if (typedArr != null) {
int resouceID = typedArr.getResourceId(0, -1);
if (resouceID == -1) {
setProgressDrawable(getResources().getDrawable(
R.drawable.play_seekbar_background));
setThumb(getResources().getDrawable(
R.drawable.play_ctrl_sound_ball));
}
typedArr.recycle();
} else {
setProgressDrawable(getResources().getDrawable(
R.drawable.play_seekbar_background));
setThumb(getResources()
.getDrawable(R.drawable.play_ctrl_sound_ball));
}
init();
}
private void init() {
setOnSeekBarChangeListener(this);
}
@Override
public void reset() {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null == controller)
return;
VDVideoInfo info = controller.getCurrentVideo();
if (info != null) {
onProgressUpdate(info.mVideoPosition, info.mVideoDuration);
}
controller.addOnProgressUpdateListener(this);
controller.addOnBufferingUpdateListener(this);
}
@Override
public void hide() {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (null != controller)
controller.removeOnProgressUpdateListener(this);
if (null != controller)
controller.removeOnBufferingUpdateListener(this);
}
@Override
public void onProgressUpdate(long current, long duration) {
if (duration > 0) {
mDuration = duration;
if (getMax() != duration && duration > 0) {
setMax((int) duration);
}
setProgress((int) current);
}
}
@Override
public void onProgressChanged(SeekBar seekBar, int progress,
boolean fromUser) {
if (fromUser) {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null)
controller.dragProgressTo((float) progress / mDuration);
if (controller != null)
controller
.notifyHideControllerBar(VDVideoViewController.DEFAULT_DELAY);
}
}
@Override
public void onStartTrackingTouch(SeekBar seekBar) {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller == null) {
return;
}
if (!DLNAController.mIsDLNA) {
VDPlayerInfo playInfo = controller.getPlayerInfo();
if (playInfo != null) {
}
}
controller.notifyHideControllerBar(VDVideoViewController.DEFAULT_DELAY);
}
@Override
public void onStopTrackingTouch(SeekBar seekBar) {
VDVideoViewController controller = VDVideoViewController
.getInstance(this.getContext());
if (controller != null)
controller
.notifyHideControllerBar(VDVideoViewController.DEFAULT_DELAY);
if (DLNAController.mIsDLNA) {
DLNAController.getInstance(getContext()).seek(getProgress());
} else {
if (controller != null)
controller.seekTo(getProgress());
if (controller != null)
controller.start();
}
}
@Override
public void onDragProgess(long progress, long duration) {
setProgress((int) progress);
}
@Override
public void onBufferingUpdate(int percent) {
setSecondaryProgress(percent * (getMax() / 100));
}
}
<file_sep>/SinaVideoSdk/sinavideosdk2/src/main/java/com/sina/sinavideo/sdk/container/VDVideoOprationpanelContainer.java
package com.sina.sinavideo.sdk.container;
/**
* 操作面板弹出界面,容器部分
*
* @author liuqun
*/
import android.content.Context;
import android.util.AttributeSet;
import android.view.View;
import android.view.animation.Animation;
import android.view.animation.AnimationUtils;
import android.widget.LinearLayout;
import com.sina.sinavideo.sdk.VDVideoViewController;
import com.sina.sinavideo.sdk.VDVideoViewListeners.OnMoreOprationVisibleChangeListener;
import com.sina.sinavideo.sdk.widgets.VDBaseWidget;
import com.sina.sinavideo.sdk.R;
public class VDVideoOprationpanelContainer extends LinearLayout
implements
VDBaseWidget,
OnMoreOprationVisibleChangeListener {
public interface OnPlayListItemClick {
public void onItemClick(int position);
}
private Animation mShowAnim;
private Animation mHideAnim;
private Context mContext = null;
/**
* 容器的适配器,因为不知道当前适配的界面,所以,ViewHolder没法弄了,短列表姑且认为没问题吧
*
* @author liuqun
*
*/
public VDVideoOprationpanelContainer(Context context, AttributeSet attrs) {
super(context, attrs);
init(context, attrs);
}
private void init(Context context, AttributeSet attrs) {
mContext = context;
setOnClickListener(mClickListener);
VDVideoViewController controller = VDVideoViewController.getInstance(context);
if(controller!=null)controller.addOnMoreOprationVisibleChangeListener(this);
mShowAnim = AnimationUtils.loadAnimation(mContext, R.anim.video_list_from_right_in);
mShowAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
setVisibility(View.VISIBLE);
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
clearAnimation();
}
});
mHideAnim = AnimationUtils.loadAnimation(mContext, R.anim.video_list_fade_from_right);
mHideAnim.setAnimationListener(new Animation.AnimationListener() {
@Override
public void onAnimationStart(Animation animation) {
}
@Override
public void onAnimationRepeat(Animation animation) {
}
@Override
public void onAnimationEnd(Animation animation) {
setVisibility(View.GONE);
clearAnimation();
}
});
}
@Override
protected void onFinishInflate() {
super.onFinishInflate();
getChildAt(0).setOnClickListener(null);
}
private OnClickListener mClickListener = new OnClickListener() {
@Override
public void onClick(View v) {
removeCallbacks(mHideAction);
post(mHideAction);
}
};
@Override
public void reset() {
}
@Override
public void hide() {
setVisibility(GONE);
}
public Runnable mHideAction = new Runnable() {
@Override
public void run() {
if (getVisibility() == VISIBLE && getAnimation() == null) {
startAnimation(mHideAnim);
}
}
};
@Override
public void showPanel() {
startAnimation(mShowAnim);
removeCallbacks(mHideAction);
postDelayed(mHideAction, VDVideoViewController.DEFAULT_DELAY);
}
@Override
public void hidePanel() {
removeCallbacks(mHideAction);
post(mHideAction);
}
@Override
public void removeAndHideDelay() {
removeCallbacks(mHideAction);
postDelayed(mHideAction, VDVideoViewController.DEFAULT_DELAY);
}
}
| 5f621a8abad937374021dc975ea54dd6d40f3d47 | [
"Java",
"Gradle"
] | 18 | Java | leoliuzhou/SinaVideoSdkDemo | cf2f04af95845dbd336bc37c1753d1f9567cac59 | 71ed9436b27e6d8416006e08f8f1e01e72ce5fb4 |
refs/heads/master | <file_sep>source 'https://github.com/CocoaPods/Specs.git'
use_frameworks!
target 'SideMenu_Example', :exclusive => true do
pod "SideMenu", :path => "../"
end
target 'SideMenu_Tests', :exclusive => true do
pod "SideMenu", :path => "../"
end
| d71b2e2d4097970ef32890236d68d13c21dd3832 | [
"Ruby"
] | 1 | Ruby | JeffGuKang/SideMenu | c1de417520114653e7d6a3106ebea4c545c2afc1 | 8f8ea9fbc5fea6891f8722bbd78b00ffbb653fd7 |
refs/heads/master | <file_sep>//
// TIMETABLEViewController.swift
// MSRIT CONNECT
//
// Created by Apple on 05/02/20.
// Copyright © 2020 KARTLA. All rights reserved.
//
import UIKit
class TIMETABLEViewController: UIViewController ,UIPopoverPresentationControllerDelegate{
@IBOutlet weak var button: UIButton!
override func viewDidLoad() {
super.viewDidLoad()
}
func prepareForSegue(segue : UIStoryboardSegue , sender : AnyObject?) {
if segue.identifier == "showPopover"
{
let popoverViewController = segue.destination
popoverViewController.popoverPresentationController?.delegate = self
}
}
func adaptivePresentationStyleForPresentationController( controller: UIPresentationController) -> UIModalPresentationStyle {
return UIModalPresentationStyle.none
}
override func didReceiveMemoryWarning() {
super.didReceiveMemoryWarning()
}
}
<file_sep>//
// ViewController.swift
// MSRIT CONNECT
//
// Created by Apple on 04/02/20.
// Copyright © 2020 KARTLA. All rights reserved.
//
import UIKit
class ViewController: UIViewController {
override func viewDidLoad() {
super.viewDidLoad()
}
}
| bc292e6865e55a2b3a1ab9432d54d1bc573e2e58 | [
"Swift"
] | 2 | Swift | sjain117/msrit-connect1 | fc4aec6b7df8167b5556faab2225858c893e210d | 18d1f6c1751cad72c21925495e2595ec2d0c130e |
refs/heads/master | <file_sep>import express from "express"
import bodyParser from "body-parser"
import {MongoClient} from "mongodb"
import path from 'path'
const app = express()
app.use(express.static(path.join(__dirname,'/build')))
app.use(bodyParser.json())
const dbURL = 'mongodb://localhost:27017/'
const setDB = async (operations) => {
try{
const client = await MongoClient.connect(dbURL,{useUnifiedTopology:true})
const db = client.db('my-blog')
await operations(db)
client.close()
} catch(error){
res.status(500).json({message:'Error connecting to database', error})
}
}
app.get('/api/article/:name',async (req,res)=>{
setDB(async (db)=>{
const articleName = req.params.name
const articleInfo = await db.collection('articles').findOne({name:articleName})
res.status(200).json(articleInfo)
})
})
app.post('/api/article/:name/upvote', async (req , res)=>{
setDB(async(db)=>{
const articleName = req.params.name
const articleInfo = await db.collection('articles').findOne({name:articleName})
await db.collection('articles').updateOne({name:articleName},{
'$set': {
upvotes:articleInfo.upvotes + 1
}
})
res.status(200).json(articleInfo)
})
})
app.post('/api/article/:name/addcomment',async (req,res)=>{
setDB(async(db)=>{
const {username,comments} = req.body
const articleName = req.params.name
const articleInfo = await db.collection('articles').findOne({name:articleName})
await db.collection('articles').updateOne({name:articleName},{
'$set':{
comments:articleInfo.comments.concat({username,comments})
}
})
const updatedArticleInfo = await db.collection('articles').findOne({name:articleName})
res.status(200).json(updatedArticleInfo)
})
})
app.get('*',(req,res)=>{
res.sendFile(path.join(__dirname + '/build/index.html'))
})
app.listen(8000,()=>console.log("server listening")) | 236a60763ef983c89acb0c5b6f9c996bf96bda57 | [
"JavaScript"
] | 1 | JavaScript | AbhishaShah/my-blog | ec19a4426ae6815a44e4d53fc6fb55edb324681e | 95f2d46abcc48faa3e98719595b0dd9f32bfe71d |
refs/heads/master | <file_sep>import statistics as es
hitoria_n1 = float(input('Digite a primeira nota: '))
hitoria_n2 = float(input('Digite a segunda nota: '))
lista_notas = [hitoria_n1,hitoria_n2]
media = es.mean(lista_notas)
if media >= 7:
print('aprovado')
elif media >= 5 and media < 7:
print('recuperação')
else: print ('reprovado')
| 084656dccc5e78fe59acf5a9b1232d674a54cedf | [
"Python"
] | 1 | Python | AbnerAbreu/Media-Aritimetica | aa9d0aca052446e618674c21ba75607394ba1ee5 | 2ef0ccab7ec1bdb1fa6cc94aebc1c1dcac7b9d82 |
refs/heads/master | <repo_name>easonlin404/limit<file_sep>/example/example.go
package main
import (
"github.com/easonlin404/limit"
"github.com/gin-gonic/gin"
)
func main() {
r := gin.Default()
r.Use(limit.Limit(200)) // limit the number of current requests
r.GET("/", func(c *gin.Context) {
// your code
})
r.Run()
}
<file_sep>/limit.go
package limit
import (
"github.com/gin-gonic/gin"
"log"
"net/http"
)
//Limit function is gin middleware to limit current requests
func Limit(max int) gin.HandlerFunc {
if max <= 0 {
log.Panic("max must be more than 0")
}
sema := make(chan struct{}, max)
return func(c *gin.Context) {
var called, fulled bool
defer func() {
if called == false && fulled == false {
<-sema
}
if r := recover(); r != nil { // We don't handle panic
panic(r)
}
}()
select {
case sema <- struct{}{}:
c.Next()
called = true
<-sema
default:
fulled = true
c.Status(http.StatusBadGateway)
}
}
}
<file_sep>/limit_test.go
package limit
import (
"github.com/gin-gonic/gin"
"github.com/stretchr/testify/assert"
"net/http/httptest"
"sync"
"testing"
"time"
)
func TestParam(t *testing.T) {
assert.Panics(t, func() {
Limit(0)
})
}
func TestLimit(t *testing.T) {
gin.SetMode(gin.TestMode)
router := gin.New()
router.Use(Limit(1))
router.GET("/", func(*gin.Context) {
time.Sleep(500 * time.Microsecond)
})
w := performRequest("GET", "/", router)
assert.Equal(t, 200, w.Code)
}
func TestHandlerPanic(t *testing.T) {
assert.Panics(t, func() {
gin.SetMode(gin.TestMode)
router := gin.New()
router.Use(Limit(1))
router.GET("/err", func(*gin.Context) {
panic("err")
})
performRequest("GET", "/err", router)
})
}
func TestFulled(t *testing.T) {
const max = 5
attempts := 1000
var failed int
var wg sync.WaitGroup
gin.SetMode(gin.TestMode)
router := gin.New()
router.Use(Limit(max))
router.GET("/", func(*gin.Context) {
time.Sleep(5 * time.Microsecond)
})
for i := 0; i < attempts; i++ {
wg.Add(1)
go func() {
defer wg.Done()
w := performRequest("GET", "/", router)
if w.Code == 502 {
failed++
}
}()
}
wg.Wait()
// We expect some Gets to fail
assert.True(t, failed > 0)
}
func performRequest(method, target string, router *gin.Engine) *httptest.ResponseRecorder {
r := httptest.NewRequest(method, target, nil)
w := httptest.NewRecorder()
router.ServeHTTP(w, r)
return w
}
<file_sep>/README.md
# Limit Gin's middleware
[](https://travis-ci.org/easonlin404/limit)
[](https://codecov.io/gh/easonlin404/limit)
[](https://goreportcard.com/report/github.com/easonlin404/limit)
[](https://godoc.org/github.com/easonlin404/limit)
Gin middleware to limit the number of current requests.
## Usage
### Start using it
Download and install it:
```sh
$ go get github.com/easonlin404/limit
```
Import it in your code:
```go
import "github.com/easonlin404/limit"
```
### Canonical example:
```go
package main
import (
"github.com/easonlin404/limit"
"github.com/gin-gonic/gin"
)
func main() {
r := gin.Default()
r.Use(limit.Limit(200)) // limit the number of current requests
r.GET("/", func(c *gin.Context) {
// your code
})
r.Run()
}
``` | a53a83c269a2c0e4ce95f051ab5ccf7b477dd227 | [
"Markdown",
"Go"
] | 4 | Go | easonlin404/limit | 3d7b09b656e270910ab648115ee537628574de87 | 569c40adf1759327a4291ec7c2066c27f3245a26 |
refs/heads/master | <file_sep>
$(window).on('load', function(){
$('#loader-container').fadeOut();
$('#show-after-window-load').fadeIn();
})
<file_sep>// Initialize Firebase
var config = {
apiKey: "<KEY>",
authDomain: "flashy-84f3e.firebaseapp.com",
databaseURL: "https://flashy-84f3e.firebaseio.com",
projectId: "flashy-84f3e",
storageBucket: "flashy-84f3e.appspot.com",
messagingSenderId: "1035470518401"
};
firebase.initializeApp(config);
// redirect user
firebase.auth().onAuthStateChanged(user => {
if (user) {
//do nothing
}
if (!user) {
window.location = "/nl-test";
}
});
//global variables
const artistInput = document.getElementById("artist");
const selectedArtistInput = document.getElementById("selectedArtist");
//global variable for database ref
const artistRef = firebase.database().ref("artists");
const lyricsRef = firebase.database().ref("/pending_songs");
//listen submit
document.getElementById("lyricsForm").addEventListener("submit", submitForm);
//submit function
function submitForm(e) {
e.preventDefault();
const name = getInput("songTitle");
const albumName = getInput("album");
const artistName = getInput("artist");
let artistInfoArr = artistInfo(artistName);
let artistId = '';
if (artistInfoArr[0] == undefined) {
artistId = 'N/A';
}
else {
artistId = artistInfoArr[0];
}
let artistImageUrl = '';
if (artistInfoArr[1] == undefined) {
artistImageUrl = 'https://firebasestorage.googleapis.com/v0/b/flashy-84f3e.appspot.com/o/artists%2Fnepali_lyrics.png?alt=media&token=<PASSWORD>';
} else {
artistImageUrl = artistInfoArr[1];
}
const artistName2 = selectedArtistInput.textContent;
const genre = getInput("genre");
const movieName = getInput("movieName");
const youTubeVideoId = `${getInput("movieLink")}`;
let contentInput = getInput("lyrics");
const content = parseToHtml(contentInput);
const hasCords = checkBox("cords");
const dateCreated = Math.floor(Date.now());
//get user info
firebase.auth().onAuthStateChanged(currentUser => {
if (currentUser) {
//User is there!
const addedById = currentUser.uid;
const addedByName = currentUser.displayName;
let arg = {
name,
albumName,
artistName,
artistName2,
genre,
movieName,
youTubeVideoId,
content,
hasCords,
addedById,
addedByName,
dateCreated,
artistId,
artistImageUrl
};
//call saveLyrics function
saveLyrics(arg);
}
});
//reset form after submit
document.getElementById("lyricsForm").reset();
//alert user after submission
alert("Thank you for submitting lyrics!");
if ((selectedArtistInput.style.display = "block")) {
selectedArtistInput.style.display = "none";
}
location.reload();
}
//get form values function
function getInput(id) {
return document.getElementById(id).value;
}
//checkbox value function
function checkBox(id) {
if (document.getElementById(id).checked) {
return true;
} else return false;
}
//add artist function
function addArtist() {
if (document.getElementById("artist").value === "") {
alert("Add artist-1 name!");
} else {
const selectedArtist = getInput("artist");
selectedArtistInput.innerHTML =
selectedArtist +
`<a id='closeBtn' onClick='removeArtist()'><i class='fas fa-window-close'></i></a>`;
selectedArtistInput.style.display = "block";
document.getElementById("artist").value = "";
artistRef
.orderByChild("name")
.equalTo(selectedArtist)
.on("value", snapshot => {
snapshot.forEach(data => {
const artistID = data.key;
const imageURL = data.child("imageUrl").val();
artist1ID = artistID;
artist1Image = imageURL;
});
});
}
}
//ArtistInfo Function
function artistInfo(artistName) {
let artistArr = [];
artistRef
.orderByChild("name")
.equalTo(artistName)
.on("value", snapshot => {
snapshot.forEach(data => {
artistArr.push(data.key);
artistArr.push(data.child("imageUrl").val());
});
});
if (artistArr === []) {
return artistName;
}
return artistArr;
}
//Artist options
artistRef.orderByValue().on("value", snapshot => {
snapshot.forEach(data => {
let artist = data.child("name").val();
document.getElementById(
"artistList"
).innerHTML += `<option value='${artist}'></option>`;
});
});
//remove artist function
function removeArtist() {
selectedArtistInput.textContent = "";
selectedArtistInput.style.display = "none";
}
// saveLyrics function
function saveLyrics(arg) {
let lyrics = {
name: arg.name,
albumName: arg.albumName,
artistName: arg.artistName,
genre: arg.genre,
movieName: arg.movieName,
youTubeVideoId: arg.youTubeVideoId,
content: arg.content,
hasCords: arg.hasCords,
addedById: arg.addedById,
addedByName: arg.addedByName,
dateCreated: arg.dateCreated,
artistId: arg.artistId,
artistImageUrl: arg.artistImageUrl,
likeCount: 0,
viewCount: 0,
favouriteCount: 0
};
if (arg.artistName2 !== "") {
lyrics["artistName2"] = arg.artistName2;
}
lyricsRef.push(lyrics);
}
//preview Lyrics
function previewLyrics() {
const name = getInput("songTitle");
const lyrics = getInput('lyrics');
const genre = getInput('genre');
const parsedLyrics = parseToHtml(lyrics);
const artistName = getInput("artist");
let artistInfoArr = artistInfo(artistName);
let artistImage = '';
const youtubeLink = getInput('movieLink');
firebase.auth().onAuthStateChanged(currentUser => {
if (currentUser) {
const userName = currentUser.displayName;
if (artistInfoArr[1] == undefined) {
artistImage = 'https://firebasestorage.googleapis.com/v0/b/flashy-84f3e.appspot.com/o/artists%2Fnepali_lyrics.png?alt=media&token=<PASSWORD>';
}
else {
artistImage = artistInfoArr[1];
}
if ((name && lyrics && artistName) == '') {
alert('Please fill the fields first!');
}
else {
document.getElementById('previewContainer').innerHTML = `
<div class="modal" id="myModal">
<div class="modal-dialog">
<div class="modal-content">
<div class="modal-header">
<h5 class="modal-title">${name}</h5>
<button class="close" data-dismiss="modal">×</button>
</div>
<div class="modal-body">
<div class="row">
<img id="artistImage" style="width:40px; height: 40px; border-radius:50%; margin-left: 10px; margin-right: 8px; margin-bottom:5px;" src="${artistImage}"/>
<font color='#341f97' id="artistNameModal">${artistName}</font>
</div>
<hr/>
<div class="mb-4">Genre: <font color='#341f97'>${genre}</font></div>
<hr/>
<div>Contributed By: <font color='#341f97'>${userName}</font></div>
<hr/>
<div id="lyricsPreviewContainer" class="unselectable">
${parsedLyrics}
</div>
<div id="youtubeVideo" data-toggle="tooltip" data-placement="top" title="Click to toggle video" onclick="playVideo()"><img src="images/if_youtube_317714.png" alt="youtube"/></div>
</div>
<div id="videoContainer">
<div class="embed-responsive embed-responsive-16by9 videos-container">
<iframe class="embed-responsive-item" src="https://youtube.com/embed/${youtubeLink}" allowfullscreen></iframe>
</div>
</div>
<div class="modal-footer">
<button type="submit" class="btn btn-primary">Submit Lyrics</button>
<button class="btn btn-secondary" data-dismiss="modal">Back</button>
</div>
</div>
</div>
</div>
`
}
if (youtubeLink == '') {
document.getElementById('youtubeVideo').style.display = 'none';
}
}
else {
alert('Please Log in!');
}
})
}
function playVideo() {
if (document.getElementById('videoContainer').style.display != 'block') {
document.getElementById('videoContainer').style.display = 'block';
}
else {
document.getElementById('videoContainer').style.display = 'none';
}
}
//test for primary arstist
//youtube link id only DONE!
//add new artist
document.getElementById('newArtistForm').addEventListener('submit', addNewArtist)
function addNewArtist(){
const newArtistName = getInput('newArtistName');
const newArtistImgUrl = getInput('newArtistImageUrl');
const newArtistInfo = {
name: newArtistName,
imageUrl: newArtistImgUrl,
viewCount: 0
}
artistRef.push(newArtistInfo);
}
| 373783e6fbb854b918909faeca5fc08a2a7ace16 | [
"JavaScript"
] | 2 | JavaScript | vijaylama/nl-test | f1100e1d52de31c21454eaf4257b7f84257891df | 64e158a9449c021214d18f4e7e0e8d43f91abac5 |
refs/heads/master | <repo_name>tato-rj/thesciencebreaker<file_sep>/app/Providers/AppServiceProvider.php
<?php
namespace App\Providers;
use App\Category;
use App\Article;
use App\Highlight;
use App\Tag;
use Illuminate\Support\Facades\Blade;
use Illuminate\Support\Facades\App;
use Illuminate\Support\Facades\Lang;
use Illuminate\Support\ServiceProvider;
use Illuminate\Support\Facades\Schema;
class AppServiceProvider extends ServiceProvider
{
/**
* Bootstrap any application services.
*
* @return void
*/
public function boot()
{
Schema::defaultStringLength(191);
\View::composer('*', function($view) {
$categories = cache()->rememberForever('tsb_categories', function() {
return Category::orderBy('name')->get();
});
$view->with('categories', $categories);
});
\View::composer('components/partials/side_bars/suggestions', function($view) {
$view->with('picks', Article::published()->editorPicks()->get());
$view->with('popular', Article::published()->popular(6)->get());
$view->with('topics', Tag::orderBy('articles_count', 'DESC')->take(25)->get());
});
Blade::if('only', function ($group) {
return \Staff::check(auth()->user()->email)->role($group);
});
\Blade::include('components.snippets.datepicker');
}
/**
* Register any application services.
*
* @return void
*/
public function register()
{
}
}
<file_sep>/app/Resources/AuthorResources.php
<?php
namespace App\Resources;
use App\ArticleAuthor;
class AuthorResources extends Resources
{
public function fullName()
{
return $this->model->first_name.' '.$this->model->last_name;
}
public function orderIn($article)
{
return ArticleAuthor::where('article_id', $article->id)->where('author_id', $this->model->id)->pluck('relevance_order')->first();
}
// public static function generateSlugs()
// {
// foreach (self::all() as $author) {
// $author->update([
// 'slug' => str_slug($author->first_name.' '.$author->last_name)
// ]);
// }
// }
}
<file_sep>/resources/lang/en/menu.php
<?php
return [
'home' => 'HOME',
'presentation' => [
'title' => 'PRESENTATION',
'about' => 'ABOUT',
'mission' => 'MISSION',
'team' => 'THE TEAM',
'breakers' => 'BREAKERS COMMUNITY',
'partners' => 'OUR PARTNERS'
],
'breaks' => 'SUBJECTS',
'for_breakers' => [
'title' => 'FOR AUTHORS',
'information' => 'INFORMATION FOR AUTHORS',
'revops' => 'REVIEW OPERATIONS',
'available' => 'AVAILABLE ARTICLES',
'guidelines' => 'WRITING GUIDELINES'
],
'contact' => [
'title' => 'CONTACT',
'submit' => 'SUBMIT YOUR ARTICLE',
'inquiry' => 'PROPOSE A TOPIC',
'question' => 'ASK A QUESTION'
],
'privacy' => 'PRIVACY POLICY',
'partner' => 'partner with',
'search' => 'Search for'
];<file_sep>/resources/lang/en/contact.php
<?php
return [
'inquiry' => [
'description' => 'If a scientific news attracted your attention and you wish to know more about it, directly from the scientists involved - let us know: we\'ll Break about it!',
'notify' => 'We will notify you when the Break will be published'
],
'form' => [
'first_name' => '<NAME>',
'last_name' => '<NAME>',
'institution_email' => [
'label' => 'Institution E-mail',
'note' => 'Please use the official email provided by your research institute'
],
'institute' => 'Research institute, Department, Unit...',
'original_article' => 'Original article title & reference',
'i_am_a' => [
'label' => 'I am a...',
'student' => 'PhD student',
'post_doc' => 'Postdoctoral Research fellow',
'assistant' => 'Research assistant',
'lecturer' => 'Lecturer',
'professor' => 'Professor',
'other' => 'Other',
'your_position' => 'Your position here'
],
'where_did_you_hear' => [
'title' => 'Where did you hear about this scientific news?',
'internet' => 'On the internet',
'journal' => 'In a newspaper/magazine',
'tv' => 'TV / Radio / Other',
'message' => 'Message (optional)',
'article_title' => 'Article title',
'author_name' => '<NAME>',
'url' => 'Article url',
'mag_title' => 'Newspaper/magazine title',
'mag_number' => 'Newspaper/magazine number',
'date' => 'Publication date',
'page' => 'The article is on page',
'describe' => 'Describe the news/subject'
],
'upload' => [
'title' => 'Manuscript upload',
'note' => [
'p1' => 'Please make sure that you read and respected the',
'link' => 'guidelines for authors',
'p2' => '! If not, your manuscript will not be eligible for publication.'
],
'file_types' => 'Upload only <strong>.doc, .docx or .odt</strong> files. Files exceeding 3 MB will not be uploaded.'
],
'your_message' => 'Your message',
'description' => 'Short description (max 400 characters)',
'add_message' => 'Add your message here and please include full information for additional Breakers, if any. Thank you!',
'newsletter' => 'Join the newsletter',
'send' => 'Send'
],
'map' => [
'title' => 'How to find us',
'address' => [
'p1' => 'TheScienceBreaker',
'p2' => 'Université de Genève',
'p3' => 'Faculté des sciences',
'p4' => '30, <NAME><br>1211 Genève 4'
]
]
];<file_sep>/tests/Feature/AdminSelectionsTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
use App\Article;
use App\Highlight;
class AdminSelectionsTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_manager_can_update_the_editors_picks()
{
$this->signIn();
$old_pick = factory('App\Article')->create(['editor_pick' => 1]);
$new_pick = factory('App\Article')->create(['editor_pick' => 0]);
$this->patch('/admin/editor-picks/'.$old_pick->id, [
'pick' => $new_pick->id
])->assertSessionHas('db_feedback');
$this->assertTrue(Article::find($old_pick->id)->editor_pick == 0);
$this->assertTrue(Article::find($new_pick->id)->editor_pick == 1);
}
/** @test */
public function a_manager_can_update_the_highlights()
{
$this->signIn();
$highlight = factory('App\Highlight')->create();
$new_article = factory('App\Article')->create();
$this->patch('/admin/highlights/'.$highlight->id, [
'article_id' => $new_article->id
])->assertSessionHas('db_feedback');
$this->assertTrue(Highlight::find($highlight->id)->article_id == $new_article->id);
}
}<file_sep>/tests/utilities/functions.php
<?php
function create($class, $attributes = [])
{
return factory($class)->create($attributes);
}
function check($class, $array)
{
foreach ($array as $route) {
$class->get($route)->assertSuccessful();
}
}
function createHighlights()
{
factory('App\Highlight', 10)->create();
}
function localize($lang)
{
\Session::put('lang', $lang);
}<file_sep>/app/Manager/Traits/ManagerScopeQueries.php
<?php
namespace App\Manager\Traits;
trait ManagerScopeQueries
{
public function scopeEditors($query)
{
return $query->where('is_editor', 1);
}
}
<file_sep>/app/Http/Controllers/ContactsController.php
<?php
namespace App\Http\Controllers;
use Illuminate\Http\Request;
use Carbon\Carbon;
use App\Mail\MailFactory;
use App\Subscription;
use App\Http\Controllers\Validators\ValidateBreakInquiry;
use App\Http\Controllers\Validators\ValidateQuestion;
use App\Http\Controllers\Validators\ValidateBreakSubmission;
class ContactsController extends Controller
{
public function __construct()
{
// $this->middleware('throttle:4');
}
public function question(Request $request)
{
if (app()->environment() != 'testing') {
if (Carbon::parse($request->time)->addSeconds(10)->gt(Carbon::now()) || ! empty($request->my_name))
return response('Humans only please.', 403);
}
ValidateQuestion::createCheck($request);
MailFactory::question($request);
if ($request->subscribe_me) Subscription::createOrIgnore($request->email);
return redirect()->back()->with('contact', 'Your message has been sent, thank you for your contact!');
}
public function inquiry(Request $request)
{
if (app()->environment() != 'testing') {
if (Carbon::parse($request->time)->addSeconds(10)->gt(Carbon::now()) || ! empty($request->my_name))
return response('Humans only please.', 403);
}
ValidateBreakInquiry::createCheck($request);
MailFactory::breakInquiry($request);
if ($request->subscribe_me) Subscription::createOrIgnore($request->email);
return redirect()->back()->with('contact', 'Your inquiry has been sent');
}
public function submit(Request $request)
{
if (app()->environment() != 'testing') {
if (Carbon::parse($request->time)->addSeconds(10)->gt(Carbon::now()) || ! empty($request->my_name))
return response('Humans only please.', 403);
}
// return $request->file('file');
ValidateBreakSubmission::createCheck($request);
MailFactory::submit($request);
if ($request->subscribe_me) Subscription::createOrIgnore($request->institution_email);
return redirect()->back()->with('contact', 'Your Break has been submitted');
}
}
<file_sep>/database/migrations/2017_10_10_212456_create_authors_table.php
<?php
use Illuminate\Support\Facades\Schema;
use Illuminate\Database\Schema\Blueprint;
use Illuminate\Database\Migrations\Migration;
class CreateAuthorsTable extends Migration
{
/**
* Run the migrations.
*
* @return void
*/
public function up()
{
Schema::create('authors', function (Blueprint $table) {
$table->increments('id');
// GIVENNAME
$table->string('first_name');
// FAMILYNAME
$table->string('last_name');
$table->string('slug')->slug();
$table->string('email')->unique();
// BIOGRAPHY
$table->string('position')->nullable();
// AFFILIATION
$table->string('research_institute')->nullable();
// IGNORE
$table->string('field_research')->nullable();
// IGNORE
$table->string('general_comments')->nullable();
$table->index(['first_name', 'last_name']);
$table->timestamps();
});
}
/**
* Reverse the migrations.
*
* @return void
*/
public function down()
{
Schema::dropIfExists('authors');
}
}
<file_sep>/resources/lang/en/team.php
<?php
return [
'categories' => [
'core' => 'Core Team',
'board' => 'Advisory Board'
],
'roles' => [
'chief' => 'Editor in Chief',
'managing_editor' => 'Managing editor',
'inhouse_editor' => 'In-house Scientific editor',
'scientific_editors' => 'Scientific editors',
'review_ops' => 'Review operations',
'community_management' => 'Community management',
'comm_officer' => 'Comm officer',
'alumni' => 'Alumni',
],
];
<file_sep>/tests/Unit/ArticlesTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
use Illuminate\Http\UploadedFile;
use Illuminate\Support\Facades\Storage;
class ArticlesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function an_article_belongs_to_a_category()
{
$category = $this->category;
$article = $this->article;
$this->assertEquals($category->name, $article->category->name);
}
/** @test */
public function an_article_can_have_may_tags()
{
$tag = $this->tag;
$second_tag = factory('App\Tag')->create();
$article = $this->article;
$article->tags()->attach($tag);
$article->tags()->attach($second_tag);
$this->assertEquals(2, count($article->tags));
}
/** @test */
public function an_article_has_an_editor()
{
$editor = $this->editor;
$article = $this->article;
$this->assertEquals($this->editor->first_name, $this->article->editor->first_name);
}
/** @test */
public function articles_can_have_one_or_more_authors()
{
$article = $this->article;
$second_author = factory('App\Author')->create();
factory('App\ArticleAuthor')->create([
'article_id' => $article->id,
'author_id' => $second_author->id
]);
$this->assertEquals(2, count($article->authors));
}
/** @test */
public function welcome_page_shows_latest_articles()
{
factory('App\Highlight', 10)->create();
$this->get('/')->assertSee($this->article->title);
}
/** @test */
public function article_page_shows_the_description()
{
factory('App\Highlight', 10)->create();
$this->get($this->article->paths()->route())->assertSee($this->article->description);
}
/** @test */
public function guests_can_read_an_article()
{
factory('App\Highlight', 10)->create();
$this->get($this->article->paths()->route())->assertSee($this->article->title);
}
/** @test */
public function guests_can_view_tags_on_the_article_page()
{
factory('App\Article', 10)->create();
$article = $this->article;
$article->tags()->attach($this->tag);
$this->get($article->paths()->route())->assertSee($article->tags->first()->name);
}
/** @test */
public function a_guest_see_the_most_popular_articles_on_the_side_bar()
{
$popular_article = factory('App\Article')->create([
'views' => 10
]);
$this->get($this->article->paths()->route())->assertSee($popular_article->title);
}
/** @test */
public function a_guest_see_the_cover_image_if_it_exists()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$slug = str_slug($title);
$category = $this->category;
$article = $this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => $category->id,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1],
'image' => UploadedFile::fake()->create('image.jpeg', 200)
]);
$this->get("breaks/$category->slug/$slug")->assertSee("$slug.jpeg");
}
/** @test */
public function an_article_keeps_count_of_viewers()
{
$this->assertEquals(0, $this->article->views);
$this->article->increment('views');
$this->assertEquals(1, $this->article->views);
}
/** @test */
public function articles_have_a_volume_number()
{
$this->assertEquals('4', $this->article->volume);
}
/** @test */
public function articles_have_an_issue_number()
{
$this->assertEquals('3', $this->article->issue);
}
}
<file_sep>/resources/assets/js/contactInputs.js
(function($) {
"use strict";
$('input[name="news_from"]').on('change', function() {
var $section = $(this).attr('id');
$('#options').find('input, textarea').hide();
$('.'+$section).fadeIn();
});
})(jQuery);
<file_sep>/resources/lang/fr/menu.php
<?php
return [
'home' => 'ACCUEIL',
'presentation' => [
'title' => 'PRÉSENTATION',
'about' => 'QUI SOMMES-NOUS?',
'mission' => 'MISSION',
'team' => 'L\'ÉQUIPE',
'breakers' => 'LA COMMUNAUTÉ DES BREAKERS',
'partners' => 'LES PARTENAIRES'
],
'breaks' => 'THÉMATIQUE',
'for_breakers' => [
'title' => 'POUR BREAKERS',
'information' => 'INFORMATION POUR LES AUTEURS',
'revops' => 'REVIEW OPERATIONS',
'available' => 'ARTICLES DISPONIBLES',
'guidelines' => 'WRITING GUIDELINES'
],
'contact' => [
'title' => 'CONTACT',
'submit' => 'SOUMETTRE VOTRE BREAK',
'inquiry' => 'DEMANDE DE BREAK',
'question' => 'POSER UNE QUESTION'
],
'privacy' => 'POLITIQUE DE CONFIDENTIALITÉ',
'partner' => 'partenaire avec',
'search' => 'Rechercher'
];
<file_sep>/app/Mail/Contact/ContactFeedback.php
<?php
namespace App\Mail\Contact;
use Illuminate\Bus\Queueable;
use Illuminate\Mail\Mailable;
use Illuminate\Queue\SerializesModels;
use Illuminate\Contracts\Queue\ShouldQueue;
class ContactFeedback extends Mailable
{
use Queueable, SerializesModels;
public $request;
public $message;
public function __construct($request, $message)
{
$this->request = $request;
$this->message = $message;
}
public function build()
{
return $this->markdown('emails/contact/contact_feedback')->subject($this->message['subject']);
}
}
<file_sep>/resources/lang/en/available_articles.php
<?php
return [
'description' => [
'p1' => 'Here below, a list of articles that were object of a Break-request. Don’t hesitate to',
'get_in_touch' => 'get in touch',
'p2' => 'if you wish to draft a Break-manuscript!'
]
];<file_sep>/app/Http/Controllers/IssuesController.php
<?php
namespace App\Http\Controllers;
use App\{Article, Tag};
use Illuminate\Http\Request;
class IssuesController extends Controller
{
public function index()
{
$archives = Article::selectRaw('year(published_at) AS year, issue, volume, count(*) as count')
->published()
->groupBy('year', 'issue', 'volume')
->orderBy('year', 'DESC')
->orderBy('issue', 'DESC')
->get();
$archives = $archives->groupBy('year');
return view('pages/archives/index', compact(['archives']));
}
public function special()
{
$special = Tag::orderBy('articles_count', 'DESC')->take(10)->get();//->where('articles_count', '>=', 15)->all();
return view('pages/archives/special', compact(['special']));
}
public function show($volume, $issue, Request $request)
{
$sort = ($request->sort) ? $request->sort : 'published_at';
$order = ($sort == 'title') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 5;
$articles = Article::where('volume', $volume)->published()->where('issue', $issue)->orderBy($sort, $order)->paginate($show);
return view('pages/archives/issue', compact(['articles', 'volume', 'issue']));
}
}
<file_sep>/resources/lang/fr/team.php
<?php
return [
'categories' => [
'core' => 'Equipe centrale',
'board' => 'Comité consultatif'
],
'roles' => [
'chief' => 'Rédacteur en chef',
'managing_editor' => 'Rédacteur adjoint',
'scientific_editors' => 'Editeurs scientifiquess',
'inhouse_editor' => 'In-house Scientific editor',
'review_ops' => 'Activités de relecture',
'community_management' => 'Gestion de la communité',
'comm_officer' => 'Chargée de communication',
'alumni' => 'Alumni',
],
];
<file_sep>/resources/lang/fr/categories.php
<?php
return [
'health-physiology' => 'Santé & Physiologie',
'neurobiology' => 'Neurobiologie',
'earth-space' => 'Terre & Espace',
'evolution-behaviour' => 'Evolution & Comportement',
'plant-biology' => 'Biologie Végétale',
'microbiology' => 'Microbiologie',
'maths-physics-chemistry' => 'Maths, Physique & Chimie',
'psychology' => 'Psychologie'
];<file_sep>/database/seeds/ManagerSeeder.php
<?php
use Illuminate\Database\Seeder;
use App\Manager;
class ManagerSeeder extends Seeder
{
/**
* Run the database seeds.
*
* @return void
*/
public function run()
{
return Manager::create([
'first_name' => 'Bart',
'last_name' => 'Simpson',
'slug' => 'bart-simpson',
'email' => '<EMAIL>',
'division_id' => 1,
'research_institute' => 'Springfield',
'image_path' => 'storage/managers/avatars/bart-simpson/bart-simpson.png',
'is_editor' => true
]);
}
}
<file_sep>/app/Http/Controllers/ManagersController.php
<?php
namespace App\Http\Controllers;
use App\{Manager, Author, Division, User};
use Illuminate\Http\Request;
use Illuminate\Support\Facades\Input;
use Illuminate\Support\Facades\File;
use App\Http\Requests\ManagerRequest;
class ManagersController extends Controller
{
public function __construct()
{
$this->middleware('auth', ['except' => ['index', 'show']]);
}
public function index()
{
$founders = Manager::where(['division_id' => 1, 'is_alumni' => false])->get();
$managing_editors = Manager::where(['division_id' => 5, 'is_alumni' => false])->get();
$inhouse_editors = Manager::where(['division_id' => 6, 'is_alumni' => false])->get();
$editors = Manager::where(['division_id' => 2, 'is_alumni' => false])->get();
$comm_officers = Manager::where(['division_id' => 3, 'is_alumni' => false])->get();
$advisors = Manager::where(['division_id' => 4, 'is_alumni' => false])->get();
$alumni = Manager::where(['is_alumni' => true])->get();
$breakers = Author::orderBy('first_name')->paginate(10);
$paginated = Input::get('page');
return view('pages.presentation.team', compact('founders', 'editors', 'managing_editors', 'inhouse_editors', 'comm_officers', 'advisors', 'breakers', 'paginated', 'alumni'));
}
public function admins()
{
$admins = User::whereNotIn('last_name', ['Caine', 'Villar'])->get();
return view('admin/pages/managers/permissions', compact('admins'));
}
public function permissions(User $user)
{
$user->is_authorized = ! $user->is_authorized;
$user->save();
return redirect()->back()->with('db_feedback', $user->full_name.' has been successfully authorized!');
}
// CREATE
public function create()
{
$divisions = Division::all();
return view('admin/pages/managers/add', compact('divisions'));
}
public function store(Request $request)
{
ManagerRequest::get()->save();
return redirect()->back()->with('db_feedback', $request->first_name.' '.$request->last_name.' has been successfully added to the team!');
}
// READ
public function show(Manager $member)
{
return view('pages/manager', compact('member'));
}
// UPDATE
public function selectEdit()
{
$managers = Manager::orderBy('first_name')->get();
return view('admin/pages/managers/selectEdit', compact(['managers']));
}
public function edit(Manager $manager)
{
$divisions = Division::all();
$managers = Manager::orderBy('first_name')->get();
return view('admin/pages/managers/edit', compact(['manager', 'divisions', 'managers']));
}
public function update(Request $request, Manager $manager)
{
ManagerRequest::get()->update($manager);
return redirect("admin/managers/$manager->slug/edit")->with('db_feedback', $manager->first_name.'\'s profile has been updated');
}
// DELETE
public function selectDelete()
{
$managers = Manager::orderBy('first_name')->get();
return view('admin/pages/managers/selectDelete', compact(['managers']));
}
public function destroy(Manager $manager)
{
File::deleteDirectory("storage/app/managers/avatars/$manager->slug");
$manager->delete();
return redirect()->back()->with('db_feedback', 'The team member has been removed from the database');
}
}
<file_sep>/app/Mail/Welcome.php
<?php
namespace App\Mail;
use App\Author;
use App\Category;
use Illuminate\Bus\Queueable;
use Illuminate\Mail\Mailable;
use Illuminate\Queue\SerializesModels;
use Illuminate\Contracts\Queue\ShouldQueue;
class Welcome extends Mailable
{
use Queueable, SerializesModels;
public $breaker;
// public $categories;
public function __construct(Author $breaker)
{
$this->breaker = $breaker;
// $this->categories = Category::all();
}
/**
* Build the message.
*
* @return $this
*/
public function build()
{
return $this->markdown('emails/welcome')->subject('Welcome to TheScienceBreaker!');
}
}
<file_sep>/app/Resources/ManagerResources.php
<?php
namespace App\Resources;
use App\ArticleAuthor;
use App\Article;
class ManagerResources extends Resources
{
public function fullName()
{
$title = ($this->model->title) ? $this->model->title.' ' : null;
return $title.$this->model->first_name.' '.$this->model->last_name;
}
public function editedArticles()
{
return Article::where('editor_id', $this->model->id)->published()->orderBy('created_at', 'DESC')->paginate(6);
}
public function editedArticlesCount()
{
return Article::where('editor_id', $this->model->id)->published()->count();
}
// public static function generateSlugs()
// {
// foreach (self::all() as $author) {
// $author->update([
// 'slug' => str_slug($author->first_name.' '.$author->last_name)
// ]);
// }
// }
}
<file_sep>/tests/Feature/ContactPageTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Tests\AppAssertions;
use Tests\TestingEmailsListener;
use Tests\MailManagement;
use Illuminate\Http\UploadedFile;
use Illuminate\Support\Facades\Storage;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class ContactPageTest extends TestCase
{
use DatabaseMigrations;
use MailManagement;
use AppAssertions;
// CONTACT PAGE INTERACTIONS
/** @test */
public function a_guest_can_ask_a_question_through_the_contact_page()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $faker->safeEmail,
'message' => $faker->paragraph
];
$this->post('/contact/ask-a-question', $request)->assertSessionHas('contact');
$this->seeEmailWasSent()->seeEmailSubjectIs('New Contact')->seeEmailContains($request['first_name']);
}
/** @test */
public function a_guest_can_send_a_break_inquiry_through_the_contact_page()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $faker->safeEmail,
'news_from' => $faker->word,
'article_title' => $faker->sentence,
'author_name' => $faker->name,
'article_url' => $faker->url,
'message' => $faker->paragraph
];
$this->post('/contact/break-inquiry', $request)->assertSessionHas('contact');
$this->seeEmailWasSent()->seeEmailSubjectIs('Break Inquiry')->seeEmailContains($request['first_name']);
}
/** @test */
public function a_guest_can_submit_a_new_break()
{
// Storage::fake('test-folder');
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'institution_email' => $faker->safeEmail,
'field_research' => $faker->word,
'research_institute' => $faker->word,
'original_article' => $faker->url,
'position' => $faker->word,
'file' => $file = UploadedFile::fake()->create('document.doc', 20),
'description' => $faker->sentence,
'message' => $faker->paragraph
];
$this->post('/contact/submit-a-break', $request)->assertSessionHas('contact');
Storage::assertExists('uploaded-breaks/'.$request['last_name'].'_'.$request['first_name'].'_break_v1.doc');
$this->seeEmailWasSent()->seeEmailsSent(2);
$this->seeEmailTo(config('app.email'))
->seeEmailSubjectIs('New Break Submission')
->seeEmailContains($request['first_name']);
}
}
<file_sep>/resources/lang/fr/contact.php
<?php
return [
'inquiry' => [
'description' => 'Si une nouvelle scientifique a attiré votre attention et que vous souhaitez en savoir plus à ce sujet, directement auprès des scientifiques impliqués, n\'hésitez pas à nous prévenir: nous préparerons une Break à ce sujet!',
'notify' => 'Nous vous tiendrons informé quand la break sera publiée'
],
'form' => [
'first_name' => 'Prénom',
'last_name' => '<NAME>',
'institution_email' => [
'label' => 'Adresse email liée à votre institution de rattachement',
'note' => 'Veuillez utiliser l\'adresse email officielle fournie par votre institut de recherche'
],
'institute' => 'Institut de recherche, département, unité',
'original_article' => 'Titre original de l\'article et référence',
'i_am_a' => [
'label' => 'Je suis un',
'student' => 'Étudiant en thèse',
'post_doc' => 'Post-doctorant',
'assistant' => 'Assistant de recherche',
'lecturer' => 'Maître de conférence',
'professor' => 'Professeur',
'other' => 'Autre',
'your_position' => 'Votre position ici'
],
'where_did_you_hear' => [
'title' => 'Ou avez-vous entendu parler de cette découverte scientifique?',
'internet' => 'Sur internet',
'journal' => 'Dans un journal ou un magazine',
'tv' => 'Télévision/radio/autre',
'message' => 'Message (facultatif)',
'article_title' => 'Le titre de l\'article',
'author_name' => 'Nom de l\'auteur',
'url' => 'URL de l\'article',
'mag_title' => 'Titre de journal / magazine',
'mag_number' => 'Numéro de journal / magazine',
'date' => 'Date de publication',
'page' => 'L\'article est sur la page',
'describe' => 'Décrivez les nouvelles / sujet'
],
'upload' => [
'title' => 'Téléchargement du manuscrit',
'note' => [
'p1' => 'Veuillez vous assurer que vous avez lu et respecté les consignes à',
'link' => 'l\'attention des auteurs',
'p2' => ', ou votre manuscrit ne pourra pas être publiée'
],
'file_types' => 'Ne téléchargez que des fichiers <strong>.doc</strong>, <strong>.docx</strong> ou <strong>.odt</strong>. Les fichiers dépassant 3MB ne pourront pas être téléchargés'
],
'your_message' => 'Votre message',
'description' => 'Courte description (400 caractères maximum)',
'add_message' => 'Entrer votre message ici en incluant toute information potentielle à l\'attention de Breakers supplémentaires. Merci!',
'newsletter' => 'Parcourir',
'send' => 'Envoyer'
],
'map' => [
'title' => 'Ou nous trouver',
'address' => [
'p1' => 'TheScienceBreaker',
'p2' => 'Université de Genève',
'p3' => 'Faculté des sciences',
'p4' => '30, Quai Ernest-Ansermet<br>1211 Genève 4'
]
]
];<file_sep>/app/Mail/Contact/BreakInquiry.php
<?php
namespace App\Mail\Contact;
use Illuminate\Bus\Queueable;
use Illuminate\Mail\Mailable;
use Illuminate\Queue\SerializesModels;
use Illuminate\Contracts\Queue\ShouldQueue;
class BreakInquiry extends Mailable
{
use Queueable, SerializesModels;
public $request;
public function __construct($request)
{
$this->request = $request;
}
public function build()
{
return $this->markdown('emails/contact/break_inquiry');
}
}
<file_sep>/app/Resources/CategoryPaths.php
<?php
namespace App\Resources;
class CategoryPaths extends Resources
{
public function icon()
{
return "/images/categories-icons/{$this->model->slug}.svg";
}
public function route()
{
return "/breaks/{$this->model->slug}";
}
}
<file_sep>/app/Http/Controllers/Validators/Validator.php
<?php
namespace App\Http\Controllers\Validators;
interface Validator
{
public static function createCheck($request);
public static function editCheck($request);
}<file_sep>/resources/assets/js/overlay.js
$(window).on('load', function(){
setTimeout(function(){
$('#overlay img').fadeOut(function(){
$('#overlay').fadeOut();
});
}, 500);
});<file_sep>/routes/web.php
<?php
Auth::routes();
Route::get('/json/{id}', 'ArticlesController@showData');
// Route::get('/fix', 'ArticlesController@fix');
// Route::get('/generate-issues', 'ArticlesController@generateIssues');
// Route::get('/generate-volumes', 'ArticlesController@generateVolumes');
Route::get('/issues', 'ArticlesController@issues');
// Welcome page
Route::get('/', 'ArticlesController@index')->name('home');
// Breaks
Route::get('/breaks/{category}/{article}', 'ArticlesController@show');
// Issues
Route::get('/content/volume/{volume}/issue/{issue}', 'IssuesController@show');
Route::get('/special-issues', 'IssuesController@special');
// Archives
Route::get('/archives', 'IssuesController@index');
// Breakers
Route::get('/breakers/{author}', 'AuthorsController@show');
// Managers
Route::get('/core-team/{member}', 'ManagersController@show');
// Categories
Route::get('/breaks/{category}', 'CategoryController@show');
// Tags
Route::get('/tags/{tag}', 'TagsController@show');
// Search
Route::get('/search', 'SearchController@index');
Route::post('/search/breakers', 'SearchController@authors');
Route::post('/search/breaks', 'SearchController@articles');
Route::get('subscription/form', 'SubscriptionsController@form')->name('subscription.form');
// Unsubscribe
Route::get('/unsubscribe', function() {
return view('pages.unsubscribe');
});
Route::delete('/unsubscribe', 'SubscriptionsController@unsubscribe');
// RSS Feed
Route::get('/services/feed', 'FeedController@index');
// Localization
Route::post('/language', 'LanguageController@set');
/*
*
* Presentation Pages
*
*/
// About
Route::get('/about', function() {
return view('pages.presentation.about');
});
// Mission
Route::get('/mission', function() {
return view('pages.presentation.mission');
});
// Team
Route::get('/the-team', 'ManagersController@index');
// Breakers
Route::get('/breakers', 'AuthorsController@index');
/*
*
* For Breakers Pages
*
*/
// Information
Route::get('/information', function() {
return view('pages.for_breakers.information');
});
Route::get('/writing-guidelines', function() {
return view('pages.for_breakers.guidelines');
});
// FAQ
Route::get('/review-operations', function() {
return view('pages.for_breakers.faq');
});
// Available Articles
Route::get('/available-articles', 'AvailableArticlesController@index');
/*
*
* Contact page
*
*/
Route::get('/contact/ask-a-question', function() {
return view('pages.contact.question');
});
Route::get('/contact/break-inquiry', function() {
return view('pages.contact.inquiry');
});
Route::get('/contact/submit-your-break', function() {
return redirect()->away('https://oap.unige.ch/journals/tsb/submissions');
// return view('pages.contact.submit');
});
Route::post('/contact/ask-a-question', 'ContactsController@question');
Route::post('/contact/break-inquiry', 'ContactsController@inquiry');
Route::post('/contact/submit-a-break', 'ContactsController@submit');
/*
*
* App
*
*/
Route::get('/app/breaks', 'AppController@breaks');
Route::get('/app/picks', 'AppController@picks');
Route::get('/app/suggestions', 'AppController@suggestions');
Route::get('/app/highlights', 'AppController@highlights');
Route::get('/app/popular', 'AppController@popular');
Route::get('/app/latest', 'AppController@latest');
Route::get('/app/home', 'AppController@home');
Route::get('/app/break/disqus', 'AppController@disqus');
Route::post('/app/breaks/views', 'AppController@incrementViews');
/*
*
* Admin
*
*/
Route::get('/admin/dashboard', 'AdminController@index');
Route::get('/admin/download', 'AdminController@download')->name('admin.download');
Route::get('/admin/graphs', 'AdminController@graphs');
// Breaks routes
Route::get('/admin/breaks/add', 'ArticlesController@create');
Route::get('/admin/breaks/edit', 'ArticlesController@edit');
Route::get('/admin/breaks/{article}/edit', 'ArticlesController@edit');
Route::get('/admin/breaks/delete', 'ArticlesController@selectDelete');
Route::get('/admin/preview-doi', 'ArticlesController@previewDOI');
Route::post('/admin/breaks', 'ArticlesController@store');
Route::post('/admin/breaks/{article}/breakers-order', 'ArticlesController@authorsOrder');
Route::patch('/admin/breaks/{article}', 'ArticlesController@update');
Route::delete('/admin/breaks/{article}', 'ArticlesController@destroy');
Route::delete('/admin/breaks/images/{article}', 'ArticlesController@destroyImage');
Route::get('/admin/breaks/xml', 'ArticlesController@showXml');
Route::post('/admin/breaks/xml', 'ArticlesController@uploadXml')->name('xml');
// Breakers routes
Route::get('/admin/breakers/add', 'AuthorsController@create');
Route::get('/admin/breakers/edit', 'AuthorsController@edit');
Route::get('/admin/breakers/{author}/edit', 'AuthorsController@edit');
Route::get('/admin/breakers/delete', 'AuthorsController@selectDelete');
Route::post('admin/breakers', 'AuthorsController@store');
Route::patch('/admin/breakers/{author}', 'AuthorsController@update');
Route::delete('/admin/breakers/{author}', 'AuthorsController@destroy');
// Managers routes
Route::get('/admin/managers/permissions', 'ManagersController@admins');
Route::patch('/admin/managers/permissions/{user}', 'ManagersController@permissions');
Route::get('/admin/managers/add', 'ManagersController@create');
Route::get('/admin/managers/edit', 'ManagersController@selectEdit');
Route::get('/admin/managers/{manager}/edit', 'ManagersController@edit');
Route::get('/admin/managers/delete', 'ManagersController@selectDelete');
Route::post('admin/managers', 'ManagersController@store');
Route::patch('/admin/managers/{manager}', 'ManagersController@update');
Route::delete('/admin/managers/{manager}', 'ManagersController@destroy');
// Available Articles routes
Route::get('/admin/available-articles', 'AvailableArticlesController@create');
Route::post('admin/available-articles', 'AvailableArticlesController@store');
Route::patch('/admin/available-articles/{availableArticle}', 'AvailableArticlesController@update');
Route::delete('/admin/available-articles/destroy-multiple', 'AvailableArticlesController@destroyMultiple');
Route::delete('/admin/available-articles/{availableArticle}', 'AvailableArticlesController@destroy');
// Editor Picks routes
Route::get('/admin/editor-picks', 'EditorPicksController@edit');
Route::patch('/admin/editor-picks/{pick}', 'EditorPicksController@update');
// Highlights routes
Route::get('/admin/highlights', 'HighlightsController@edit');
Route::patch('/admin/highlights/{highlight}', 'HighlightsController@update');
// Subscription routes
Route::get('/admin/subscriptions', 'SubscriptionsController@index');
Route::post('admin/subscriptions', 'SubscriptionsController@store');
Route::delete('/admin/subscriptions/{email}', 'SubscriptionsController@destroy');
// Tags
Route::get('/admin/tags', 'TagsController@index');
Route::post('/admin/breaks/{article}/tags', 'ArticlesController@setTags');
Route::post('/admin/tags', 'TagsController@store');
Route::patch('/admin/tags/{tag}', 'TagsController@update');
Route::delete('/admin/tags/{tag}', 'TagsController@destroy');<file_sep>/app/Http/Controllers/AvailableArticlesController.php
<?php
namespace App\Http\Controllers;
use App\AvailableArticle;
use App\Category;
use App\Http\Controllers\Validators\ValidateAvailableArticle;
use Illuminate\Http\Request;
class AvailableArticlesController extends Controller
{
public function __construct()
{
$this->middleware('auth', ['except' => ['index']]);
}
// CREATE
public function create(Request $request)
{
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'category_id') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 5;
$articles = AvailableArticle::orderBy($sort, $order)->paginate($show);
return view('admin.pages.available_articles', compact('articles'));
}
public function store(Request $request)
{
ValidateAvailableArticle::createCheck($request);
AvailableArticle::create($request->except('page'));
return redirect()->back()->with('db_feedback', 'The Article is now available');
}
// READ
public function index()
{
$categories = Category::with('available_articles')->get();
// return $categories;
return view('pages.for_breakers.available_articles', compact('categories'));
}
// UPDATE
public function update(Request $request, AvailableArticle $availableArticle)
{
ValidateAvailableArticle::editCheck($request);
$availableArticle->update($request->except('page'));
return redirect()->back()->with('db_feedback', 'The available article has been updated');
}
// DELETE
public function destroyMultiple(Request $request)
{
$articles = json_decode($request->articles);
foreach ($articles as $article) {
AvailableArticle::find($article)->delete();
}
return redirect()->back()->with('db_feedback', 'The articles has been removed from the database');
}
// DELETE
public function destroy(AvailableArticle $availableArticle)
{
$availableArticle->delete();
return redirect()->back()->with('db_feedback', 'The article has been removed from the database');
}
}
<file_sep>/tests/Feature/AdminManagersTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
use App\Article;
class AdminManagersTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_manager_can_add_a_new_team_member()
{
$this->signIn();
$this->post('/admin/managers', [
'first_name' => 'Bart',
'last_name' => 'Simpson',
'email' => '<EMAIL>',
'division_id' => 2,
'position' => 'Professor',
'biography' => 'Bart\'s biography goes here.',
'research_institute' => 'SU',
'is_editor' => 0
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('managers', [
'first_name' => 'Bart'
]);
}
/** @test */
public function a_manager_can_remove_a_team_member()
{
$this->signIn();
$manager = $this->manager;
$this->delete('/admin/managers/'.$manager->slug)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('managers', [
'id' => $manager->id
]);
}
/** @test */
public function a_manager_can_view_a_page_to_edit_a_team_member()
{
$this->signIn();
$this->get('/admin/managers/'.$this->manager->slug.'/edit')->assertSee($this->manager->first_name);
}
/** @test */
public function a_manager_can_edit_a_team_member()
{
$this->signIn();
$manager = $this->manager;
$this->patch('/admin/managers/'.$manager->slug, [
'first_name' => 'Lisa',
'last_name' => $manager->last_name,
'email' => $manager->email,
'division_id' => $manager->division_id,
'position' => $manager->position,
'biography' => $manager->biography,
'research_institute' => $manager->research_institute,
'is_editor' => $manager->is_editor
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('managers', [
'first_name' => 'Lisa'
])->assertDatabaseMissing('managers', [
'first_name' => $manager->first_name
]);
}
/** @test */
public function a_manager_can_authorize_a_new_admin()
{
$this->signIn();
$manager = $this->manager;
$admin = create('App\User');
$this->assertFalse($admin->is_authorized);
$this->patch('/admin/managers/permissions/'.$admin->id)->assertSessionHas('db_feedback');
$this->assertTrue($admin->fresh()->is_authorized);
}
/** @test */
public function a_manager_can_unauthorize_a_new_admin()
{
$this->signIn();
$manager = $this->manager;
$admin = create('App\User', ['is_authorized' => 1]);
$this->assertTrue($admin->is_authorized);
$this->patch('/admin/managers/permissions/'.$admin->id)->assertSessionHas('db_feedback');
$this->assertFalse($admin->fresh()->is_authorized);
}
}<file_sep>/tests/MailManagement.php
<?php
namespace Tests;
use Mail;
use Swift_Message;
trait MailManagement
{
protected $emails = [];
public function setUp()
{
parent::setUp();
Mail::getSwiftMailer()->registerPlugin(new TestingEmailsListener($this));
}
public function addEmail(Swift_Message $email)
{
$this->emails[] = $email;
}
}<file_sep>/tests/Unit/TagsTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class TagsTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_tag_can_have_many_articles()
{
$tag = $this->tag;
$article = $this->article;
$second_article = factory('App\Article')->create();
$article->tags()->attach($tag);
$second_article->tags()->attach($tag);
$this->assertEquals(2, count($tag->articles));
}
/** @test */
public function a_manager_can_create_new_tags()
{
$this->post('/admin/tags', [
'tag' => 'new tag'
]);
$this->assertDatabaseHas('tags', [
'name' => 'new tag'
]);
}
/** @test */
public function a_manager_can_remove_tags()
{
$tag = $this->tag;
$this->delete("/admin/tags/$tag->name");
$this->assertDatabaseMissing('tags', [
'name' => $tag->name
]);
}
/** @test */
public function a_manager_can_edit_tags()
{
$tag = $this->tag;
$old_name = $tag->name;
$this->patch("/admin/tags/$tag->name", [
'tag' => 'newname'
]);
$this->assertDatabaseMissing('tags', [
'name' => $old_name
]);
$this->assertDatabaseHas('tags', [
'name' => 'newname'
]);
}
/** @test */
public function a_removed_tag_also_removes_its_relationships_with_articles()
{
$tag = $this->tag;
$article = $this->article;
$article->tags()->attach($tag);
$this->assertDatabaseHas('article_tag', [
'article_id' => $article->id
]);
$this->delete("/admin/tags/$tag->name");
$this->assertDatabaseMissing('article_tag', [
'article_id' => $article->id
]);
}
/** @test */
public function a_guest_can_view_all_breaks_from_a_tag()
{
$tag = $this->tag;
$article = $this->article;
$article->tags()->attach($tag);
$this->get("/tags/$tag->name")->assertSee($article->title);
}
}
<file_sep>/tests/Unit/ManagersTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class ManagersTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function managers_have_their_own_page()
{
$manager = $this->manager;
$this->get("/core-team/$manager->slug")->assertSee($manager->first_name);
}
}
<file_sep>/app/Division.php
<?php
namespace App;
use App\TheScienceBreaker;
class Division extends TheScienceBreaker
{
//
}
<file_sep>/app/ArticleAuthor.php
<?php
namespace App;
use App\TheScienceBreaker;
class ArticleAuthor extends TheScienceBreaker
{
protected $table = 'article_author';
}
<file_sep>/bower_components/gb-jquery-popover/src/jquery-popover.js
;(function ( $, window, document, undefined ) {
'use strict';
var old = $.fn.popover;
var Popover = function(el, options) {
this.$el = $(el);
this.options = this.getOptions(options);
this.$wrapper = this.$el.parents('.popover-wrapper').eq(0);
this.$body = this.$wrapper.find('.popover-body');
this.listenEvents();
return this;
};
Popover.DEFAULTS = {
trigger: 'click' // click | hover
};
Popover.prototype.getDefaults = function() {
return Popover.DEFAULTS;
};
Popover.prototype.getOptions = function (options) {
options = $.extend({}, this.getDefaults(), this.$el.data(), options);
return options;
};
Popover.prototype.listenEvents = function (options) {
var this_ = this;
var $el = this.$el;
// click outside to close modal
$(document).click(function(e) {
if ( !this.$wrapper.hasClass('open') ){
return;
}
var shouldClose = (
!$.contains( this.$wrapper[0], e.target )
&& this.$wrapper[0] !== e.target
)
if ( shouldClose ) {
this.close();
}
}.bind(this));
// click, hover, or focus based triggers
var trigger = this.options.trigger;
if ( trigger === 'click' ) {
$el.on('click', function(e) {
e.preventDefault();
this_.toggle();
});
} else if ( trigger === 'hover' ) {
$el.on('mouseenter', function(e) {
e.preventDefault();
this_.open();
});
$el.on('mouseleave', function(e) {
e.preventDefault();
this_.close();
});
$el.on('click', function(e) {
e.preventDefault();
this_.toggle();
});
} else if ( trigger === 'focus' ) {
}
// Listen to close buttons
this.$wrapper.find('[data-toggle-role="close"]').on('click', function(e) {
e.preventDefault();
this_.close();
});
return this;
};
Popover.prototype.open = function() {
if (this.$wrapper) this.$wrapper.addClass('open');
return this;
};
Popover.prototype.close = function() {
if (this.$wrapper) this.$wrapper.removeClass('open');
return this;
};
Popover.prototype.toggle = function() {
if (this.$wrapper) this.$wrapper.toggleClass('open');
return this;
};
// PLUGIN DEFINITION
var Plugin = function( options ){
return this.each(function() {
var $this = $(this);
var data = $this.data('gb.popover');
if (!data) {
data = new Popover(this, options);
$this.data('gb.popover', data);
}
});
};
$.fn.popover = Plugin;
$.fn.popover.Constructor = Popover;
// NO CONFLICT
$.fn.popover.noConflict = function() {
$.fn.popover = old;
return this;
};
})( jQuery, window, document );
<file_sep>/app/Http/Controllers/HighlightsController.php
<?php
namespace App\Http\Controllers;
use App\Category;
use App\Highlight;
use Illuminate\Http\Request;
class HighlightsController extends Controller
{
public function __construct()
{
$this->middleware('auth', ['except' => ['index']]);
}
// READ
public function index()
{
//
}
// UPDATE
public function edit()
{
$breaksByCategory = Category::with(['articles' => function($query) {
return $query->orderBy('created_at', 'DESC');
}])->get();
$highlights = Highlight::orderBy('relevance_order')->get();
return view('admin/pages/highlights', compact(['highlights', 'breaksByCategory']));
}
public function update(Request $request, Highlight $highlight)
{
$highlight->update(['article_id' => $request->article_id]);
return redirect()->back()->with('db_feedback', 'The new highlight has been selected');
}
}
<file_sep>/tests/Unit/LocalizationTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
use Illuminate\Http\UploadedFile;
use Illuminate\Support\Facades\Storage;
class LocalizationTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function website_shows_user_preferred_language()
{
createHighlights();
localize('fr');
$this->get('/')->assertSee('Pourquoi TheScienceBreaker?');
}
/** @test */
public function an_article_shows_default_english_title_if_translation_is_empty()
{
createHighlights();
localize('fr');
$article = $this->article;
// At this point there is no french title
$this->get($article->paths()->route())->assertSee($article->title);
$article->title_fr = 'Title in french';
$article->save();
// Now there is a title in french
$this->get($article->paths()->route())->assertSee($article->title_fr);
}
/** @test */
public function an_article_shows_default_english_description_if_translation_is_empty()
{
createHighlights();
localize('fr');
$article = $this->article;
// At this point there is no french description
$this->get($article->paths()->route())->assertSee($article->description);
$article->description_fr = 'Description in french';
$article->save();
// Now there is a title in french
$this->get($article->paths()->route())->assertSee($article->description_fr);
}
/** @test */
public function an_article_shows_default_english_content_if_translation_is_empty()
{
createHighlights();
localize('fr');
$article = $this->article;
// At this point there is no french content
$this->get($article->paths()->route())->assertSee($article->content);
$article->content_fr = 'Content in french';
$article->save();
// Now there is a title in french
$this->get($article->paths()->route())->assertSee($article->content_fr);
}
}
<file_sep>/app/Http/Controllers/AuthorsController.php
<?php
namespace App\Http\Controllers;
use App\Author;
use App\Mail\MailFactory;
use Illuminate\Http\Request;
use App\Http\Requests\AuthorRequest;
class AuthorsController extends Controller
{
public function __construct()
{
$this->middleware('auth', ['except' => ['index', 'show']]);
}
public function index(Request $request)
{
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'first_name' || $sort == 'last_name') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 5;
$breakers = Author::orderBy($sort, $order)->paginate($show);
return view('pages/presentation/breakers', compact('breakers'));
}
// CREATE
public function create()
{
return view('admin/pages/breakers/add');
}
public function store(Request $request)
{
if (Author::where('slug', str_slug($request->first_name . ' ' . $request->last_name))->exists())
return back()->withErrors(['This author already exists']);
$breaker = AuthorRequest::get()->save();
// MailFactory::sendWelcomeEmail($breaker);
return redirect()->back()->with('db_feedback', 'The Breaker '.$request->first_name.' '.$request->last_name.' has been successfully added!');
}
// READ
public function show(Author $author)
{
return view('pages/author', compact('author'));
}
// UPDATE
public function edit(Author $author = null)
{
$breakers = Author::orderBy('first_name')->get();
return view('admin/pages/breakers/edit', compact(['author', 'breakers']));
}
public function update(Request $request, Author $author)
{
AuthorRequest::get()->update($author);
return redirect('/admin/breakers/edit')->with('db_feedback', $author->first_name.'\'s profile has been updated');
}
// DELETE
public function selectDelete()
{
$breakers = Author::orderBy('first_name')->get();
return view('admin/pages/breakers/selectDelete', compact(['breakers']));
}
public function destroy(Author $author)
{
$author->articles()->detach();
$author->delete();
return redirect()->back()->with('db_feedback', 'The Breaker has been removed from the database');
}
}
<file_sep>/app/Manager.php
<?php
namespace App;
use App\TheScienceBreaker;
use App\Manager\Traits\ManagerScopeQueries;
use App\Resources\ManagerPaths;
use App\Resources\ManagerResources;
class Manager extends TheScienceBreaker
{
use ManagerScopeQueries;
protected $casts = ['is_alumni' => 'bool'];
public function getRouteKeyName()
{
return 'slug';
}
public function paths()
{
return new ManagerPaths($this);
}
public function resources()
{
return new ManagerResources($this);
}
public function division()
{
return $this->belongsTo('App\Division');
}
}
<file_sep>/app/Http/Controllers/TagsController.php
<?php
namespace App\Http\Controllers;
use App\Tag;
use Illuminate\Http\Request;
class TagsController extends Controller
{
// CREATE
public function store(Request $request)
{
$request->validate(['tag' => 'required|unique:tags,name']);
$new_tag = Tag::create(['name' => $request->tag]);
if ($request->ajax()) {
return $new_tag->id;
}
return redirect()->back()->with('db_feedback', 'The tag has been created');
}
// READ
public function index(Request $request)
{
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'name') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 20;
$tags = Tag::orderBy($sort, $order)->paginate($show);
return view('admin/pages/tags', compact('tags'));
}
public function show(Tag $tag, Request $request)
{
$input = $request->for;
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'title') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 5;
$articles = $tag->articles()->orderBy($sort, $order)->paginate($show);
return view('pages.tag', compact(['tag', 'articles']));
}
// UPDATE
public function update(Request $request, Tag $tag)
{
$request->validate(['tag' => 'required|unique:tags,name']);
$tag->update(['name' => $request->tag]);
return redirect()->back()->with('db_feedback', 'The tag has been updated');
}
// DELETE
public function destroy(Tag $tag)
{
$tag->articles()->detach();
$tag->delete();
return redirect()->back()->with('db_feedback', 'The tag has been removed');
}
}
<file_sep>/bower_components/gb-jquery-popover/gulpfile.js
var gulp = require('gulp');
var less = require('gulp-less');
var minifyCSS = require('gulp-minify-css');
var uglify = require('gulp-uglify');
var ver = require('gulp-ver');
var path = require('path');
gulp.task('less', function() {
return gulp.src('src/**/*.less')
.pipe(less({
paths: [ path.join(__dirname, 'less', 'includes') ]
}))
.pipe(minifyCSS())
.pipe(ver())
.pipe(gulp.dest('dist'));
});
gulp.task('compress', function() {
return gulp.src('src/**/*.js')
.pipe(uglify())
.pipe(ver())
.pipe(gulp.dest('dist'));
});
gulp.task('default', [ 'less', 'compress' ]);
<file_sep>/bower_components/gb-jquery-popover/README.md
# jquery-popover
> jquery popover plugin
## Install
```shell
bower install gb-jquery-popover
```
In your html, reference the latest versioned files from `/dist`.
```html
<head>
<link rel="stylesheet" href="dist/jquery-popover-x.y.z.css">
</head>
<body>
<script src="dist/jquery-popover-x.y.z.js"></script>
</body>
```
## Usage
Add popover markup
```html
<span class="popover-wrapper">
<a href="#" data-role="popover" data-target="example-popover">Open</a>
<div class="popover-modal example-popover">
<div class="popover-body">
<!-- contents here -->
</div>
</div>
</span>
```
and enable them with jQuery
```js
$('[data-role="popover"]').popover();
```
[Demo](http://goodybag.github.io/jquery-popover/)
## Build
The default task simply minifies the source files
```shell
gulp
```
While developing, we highly recommend live reloading via browsersync. As you
change files in `/src`, browsersync will automatically inject and reload
the page for you.
Also, make sure you are on the `gh-pages` branch because
`master` does not contain the index.html which serves as a dev playground.
```shell
git checkout gh-pages
gulp serve
```
## Test
```
npm test
```
## License
MIT
---
Made with :heart: by [Goodybag](http://goodybag.com)
<file_sep>/database/seeds/DivisionSeeder.php
<?php
use Illuminate\Database\Seeder;
use App\Division;
class DivisionSeeder extends Seeder
{
/**
* Run the database seeds.
*
* @return void
*/
public function run()
{
Division::create([
'name' => 'Editor'
]);
}
}
<file_sep>/app/Manager/Staff.php
<?php
namespace App\Manager;
class Staff
{
private $email;
protected $managers = ['<EMAIL>', '<EMAIL>'];
public static function check($email)
{
$staff = new self;
$staff->email = $email;
return $staff;
}
public function role($role)
{
return in_array($this->email, $this->$role);
}
}
<file_sep>/app/Http/Controllers/Validators/ValidateAvailableArticle.php
<?php
namespace App\Http\Controllers\Validators;
class ValidateAvailableArticle implements Validator
{
public static function createCheck($request)
{
return $request->validate([
'article' => 'required|unique:available_articles',
'category_id' => 'required'
]);
}
public static function editCheck($request)
{
return $request->validate([
'article' => 'required',
'category_id' => 'required'
]);
}
}<file_sep>/app/Category.php
<?php
namespace App;
use App\TheScienceBreaker;
use App\Resources\CategoryPaths;
class Category extends TheScienceBreaker
{
protected $withCount = ['articles'];
public function getRouteKeyName()
{
return 'slug';
}
public function paths()
{
return new CategoryPaths($this);
}
public function articles()
{
return $this->hasMany(Article::class)->orderBy('created_at', 'DESC');
}
public function available_articles()
{
return $this->hasMany(AvailableArticle::class);
}
public function scopeByName($query, $name)
{
return $query->whereRaw('lower(name) like (?)', strtolower($name));
}
}
<file_sep>/public/js/avatars.js
$('#tab-bar a').click(function (e) {
e.preventDefault()
$(this).tab('show')
});
$('.avatar').on('click', function(event) {
event.stopPropagation();
resetAvatars();
$(this).toggleClass('border-color-green').children('.about').toggle();
});
$(window).click(function() {
resetAvatars();
});
function resetAvatars ($elements) {
$('.avatars .avatar').removeClass('border-color-green').children('.about').hide();
}<file_sep>/app/Resources/AuthorPaths.php
<?php
namespace App\Resources;
class AuthorPaths extends Resources
{
public function route()
{
return "/breakers/".$this->model->slug;
}
}
<file_sep>/app/Tag.php
<?php
namespace App;
use App\TheScienceBreaker;
class Tag extends TheScienceBreaker
{
protected $withCount = ['articles'];
public function getRouteKeyName()
{
return 'name';
}
public function articles()
{
return $this->belongsToMany(Article::class)->published();
}
public function path()
{
return "/tags/$this->name";
}
public static function list()
{
$tagsList = '';
foreach (self::pluck('name') as $tag) {
$tagsList .= $tag.', ';
}
$tagsList = substr($tagsList, 0, -2);
return $tagsList;
}
}
<file_sep>/app/Manager/Traits/ArticleScopeQueries.php
<?php
namespace App\Manager\Traits;
trait ArticleScopeQueries
{
public function scopeExcept($query, $model)
{
return $query->where('id', '!=', $model->id);
}
public function scopeRecent($query, $number)
{
return $query->with(['category', 'authors'])->orderBy('published_at', 'desc')->take($number);
}
public function scopeSimilar($query)
{
return $this->category->articles()->except($this)->with(['category', 'authors'])->orderBy('id', 'desc')->take(5);
}
public function scopePopular($query, $number)
{
return $query->with(['category', 'authors'])->orderBy('views', 'desc')->take($number);
}
public function scopeEditorPicks($query)
{
return $query->with(['category', 'authors'])->where('editor_pick', 1);
}
public function scopeByTitle($query, $title)
{
return $query->whereRaw('lower(title) like (?)', strtolower($title));
}
public function scopePicks($query)
{
// return $query->where('editor_pick', 1)->orderBy('title')->get();
}
public function scopeSearch($query, $word)
{
return $query
->where('title', 'LIKE', "%$word%")
->orWhere('content', 'LIKE', "%$word%")
->orWhereHas('authors', function($query) use ($word) {
$query->where('first_name', 'LIKE', "%$word%")->orWhere('last_name', 'LIKE', "%$word%");
})->orWhereHas('editor', function($query) use ($word) {
$query->where('first_name', 'LIKE', "%$word%")->orWhere('last_name', 'LIKE', "%$word%");
})->orWhereHas('category', function($query) use ($word) {
$query->where('name', 'LIKE', "%$word%");
})->orWhereHas('tags', function($query) use ($word) {
$query->where('name', 'LIKE', "%$word%");
});
}
// public function scopeRecords($query, $length)
// {
// return $query->selectRaw('year(created_at) year, monthname(created_at) month, count(*) published')
// ->whereRaw('created_at >= DATE_ADD(LAST_DAY(DATE_SUB(NOW(), INTERVAL '.$length.')), INTERVAL 1 DAY) and created_at <= NOW()')
// ->groupBy('year', 'month')
// ->orderByRaw('min(created_at) asc')
// ->get();
// }
}
<file_sep>/resources/lang/en/global.php
<?php
return [
'logo_name' => 'logo_en',
'sub_title' => 'Science meets Society',
'latest' => 'Latest',
'picks' => 'Editor\'s picks',
'popular' => 'Trending now',
'topics' => 'Popular topics',
'in' => 'in',
'by' => 'by',
'published' => 'published on',
'views' => 'views',
'read' => 'read',
'reading_time' => 'Reading time',
'at' => 'at',
'click_to_read_more' => 'click to read more',
'search' => 'Search here',
'articles' => 'articles',
'sort_bar' => [
'show' => 'show',
'sort_by' => 'sort by',
'showing' => 'showing',
'of' => 'of',
'all' => 'all',
'date' => 'newest',
'breaks_num' => 'number of breaks',
'first_name' => '<NAME>',
'last_name' => '<NAME>',
'popular' => 'most popular',
'title' => 'title',
'reading_time' => 'reading time'
]
];<file_sep>/app/Http/Requests/FileUpload.php
<?php
namespace App\Http\Requests;
use Illuminate\Support\Facades\File;
use Illuminate\Support\Facades\Storage;
use Illuminate\Http\Request;
class FileUpload {
protected $file;
protected $filename;
protected $path;
public function __construct($file)
{
$this->file = $file;
}
public function name($name)
{
$ext = $this->file->extension();
$this->filename = "$name.$ext";
return $this;
}
public function path($path)
{
$this->path = $path;
return $this;
}
public function save()
{
$filepath = $this->path.$this->filename;
Storage::put('public' . $filepath, File::get($this->file));
return "storage" . $filepath;
}
public function replace()
{
File::cleanDirectory("storage/app$this->path");
$this->save();
}
}
<file_sep>/app/Http/Requests/Form.php
<?php
namespace App\Http\Requests;
use App\Article;
use Carbon\Carbon;
use Illuminate\Http\Request;
use Illuminate\Foundation\Validation\ValidatesRequests;
abstract class Form
{
use ValidatesRequests;
protected $request;
protected $slug;
public function __construct(Request $request = null)
{
$this->request = $request ?: request();
}
abstract protected function rules();
public static function get()
{
return new static;
}
public function isValid()
{
try {
$this->created_at = Carbon::parse("$this->created_at 00:00:00")->format('Y-m-d H:i:s');
} catch(\Exception $e) {
return redirect()->back()->withErrors(['Check the date format! It must be M/D/Y']);
}
$this->validate($this->request, $this->rules());
return true;
}
public function save()
{
if ($this->isValid()) {
return $this->create();
}
return false;
}
public function update($model)
{
if ($this->isValid()) {
return $this->edit($model);
}
return false;
}
public function saveFile()
{
$file = null;
if ($this->request->file('pdf')) {
$file = (new FileUpload($this->request->file('pdf')))->name($this->slug)->path("/breaks/")->save();
}
if ($this->request->file('image')) {
$file = (new FileUpload($this->request->file('image')))->name($this->slug)->path("/breaks/images/$this->slug/")->save();
}
if ($this->request->file('avatar')) {
$file = (new FileUpload($this->request->file('avatar')))->name($this->slug)->path("/managers/avatars/$this->slug/")->save();
}
return $file;
}
public function __get($property)
{
if ($this->request->has($property)) {
return $this->request->input($property);
}
return null;
}
public function slug($input)
{
$this->slug = str_slug($input);
}
}
<file_sep>/app/Subscription.php
<?php
namespace App;
use App\TheScienceBreaker;
use Illuminate\Database\QueryException;
class Subscription extends TheScienceBreaker
{
public static function createOrIgnore($email)
{
try
{
self::create([
'email' => $email
]);
} catch (QueryException $e) {}
}
}
<file_sep>/app/Http/Controllers/Validators/ValidateBreakInquiry.php
<?php
namespace App\Http\Controllers\Validators;
use App\Rules\Recaptcha;
class ValidateBreakInquiry implements Validator
{
public static function createCheck($request) {
return $request->validate([
'first_name' => 'required|min:2',
'last_name' => 'required|min:2',
'email' => 'required|email',
'g-recaptcha-response' => ['sometimes', new Recaptcha]
]);
}
public static function editCheck($request) {}
}
<file_sep>/app/Http/Controllers/EditorPicksController.php
<?php
namespace App\Http\Controllers;
use App\Article;
use App\Category;
use Illuminate\Http\Request;
class EditorPicksController extends Controller
{
public function __construct()
{
$this->middleware('auth', ['except' => ['index']]);
}
// READ
public function index()
{
//
}
// UPDATE
public function edit()
{
$breaksByCategory = Category::with(['articles' => function($query) {
return $query->orderBy('created_at', 'DESC');
}])->get();
$picks = Article::picks();
return view('admin/pages/picks', compact(['picks', 'breaksByCategory']));
}
public function update(Request $request, $id)
{
$old_pick = Article::find($id);
$new_pick = Article::find($request->pick);
$old_pick->update(['editor_pick' => 0]);
$new_pick->update(['editor_pick' => 1]);
return redirect()->back()->with('db_feedback', 'The new pick has been selected');
}
}
<file_sep>/tests/Unit/AvailableArticlesTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class AvailableArticlesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function guests_can_see_available_articles()
{
$available_article = factory('App\AvailableArticle')->create();
$this->get('/available-articles')->assertSee($available_article->article);
}
}<file_sep>/app/Resources/ArticlePaths.php
<?php
namespace App\Resources;
use Illuminate\Support\Facades\File;
class ArticlePaths extends Resources
{
public function image()
{
// $file = 'breaks/images/'.$this->model->slug.'/'.$this->model->slug.'.jpeg';
// if (\Storage::exists($file))
// return asset(\Storage::url($file));
return asset($this->model->image_path);
// return "images/no-image.png";
}
public function route()
{
return "/breaks/{$this->model->category->slug}/{$this->model->slug}";
}
public function pdf()
{
return "storage/breaks/".$this->model->slug.".pdf";
}
}
<file_sep>/tests/Feature/AdminSubscriptionsTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
use App\Subscription;
class AdminSubscriptionsTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_manager_can_see_all_subscriptions_on_the_admin_page()
{
$this->signIn();
$this->get('/admin/subscriptions')->assertSee($this->subscription->email);
}
/** @test */
public function a_manager_can_add_a_new_subscription()
{
$this->signIn();
$this->post('/admin/subscriptions', [
'subscription' => '<EMAIL>'
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('subscriptions', [
'email' => '<EMAIL>'
]);
}
/** @test */
public function a_manager_can_remove_a_subscription()
{
$this->signIn();
$email = $this->subscription;
$this->delete('/admin/subscriptions/'.$email->id)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('subscriptions', [
'email' => $email->email
]);
}
}<file_sep>/resources/lang/en/footer.php
<?php
return [
'follow' => 'FOLLOW US',
'subscribe' => [
'title' => 'SUBSCRIBE',
'text' => 'Stay up-to-date with the latest published Breaks!',
'input' => 'Enter email',
'note' => 'We\'ll never share your email with anyone else.',
'button' => 'Subscribe me'
],
'copyright' => 'All rights reserved'
];<file_sep>/resources/lang/en/welcome.php
<?php
return [
'description' => [
'why-title' => 'Editorial Mission',
'why-text' => 'As Online, Open Access, and Outreach Journal, we promote the democratization of scientific literature to foster dialogues and interest over the most recent scientific advances. Discover our <a href="/mission">mission</a>.',
'what-title' => 'Journal content',
'what-text' => 'We publish short lay-summaries ("<i>breaks</i>") of scientific research. Our authors are scientists involved in the field of the summarized research. Our readers are academics and laypeople likewise. <a href="/about">Learn more</a>.',
],
'partners' => 'Collaborations',
'categories' => 'Subjects',
'app' => 'We launched our iOS app, <strong>check it out</strong>!',
'highlights' => 'Highlights',
];
<file_sep>/app/Mail/Contact/Submit.php
<?php
namespace App\Mail\Contact;
use App\Author;
use Illuminate\Bus\Queueable;
use Illuminate\Mail\Mailable;
use Illuminate\Queue\SerializesModels;
use Illuminate\Contracts\Queue\ShouldQueue;
class Submit extends Mailable
{
use Queueable, SerializesModels;
public $request;
public $file;
public function __construct($request, $file)
{
$this->request = $request;
$this->file = $file;
}
public function build()
{
return $this->markdown('emails/contact/submit')->subject('New Break Submission');
}
}
<file_sep>/app/Resources/Resources.php
<?php
namespace App\Resources;
use App\TheScienceBreaker;
abstract class Resources
{
protected $model;
public function __construct(TheScienceBreaker $model)
{
$this->model = $model;
}
}
<file_sep>/app/Http/Controllers/AdminController.php
<?php
namespace App\Http\Controllers;
use App\Article;
use App\Author;
use App\AvailableArticle;
use App\Subscription;
use Maatwebsite\Excel\Facades\Excel;
use Illuminate\Http\Request;
class AdminController extends Controller
{
public function __construct()
{
$this->middleware('auth');
}
public function index()
{
$total_views = Article::sum('views');
$breaks_count = Article::count();
$authors_count = Author::count();
$available_count = AvailableArticle::count();
$subscription_count = Subscription::count();
$breaks_views_excel = Excel::create('breaks_views', function($breaks_views_excel) {
$breaks_views_excel->sheet('Breaks_views', function($sheet) {
$sheet->fromModel(Article::setEagerLoads([])->selectRaw('DATE_FORMAT(created_at,"%d/%m/%Y") as Date, title as Title, views as Views')->orderBy('views', 'DESC')->get(), null, 'A1', true);
});
})->store('xls', storage_path('app/public/breaks/excel'));
$breakers_excel = Excel::create('breakers_emails', function($breakers_excel) {
$breakers_excel->sheet('Breakers', function($sheet) {
$sheet->fromModel(Author::setEagerLoads([])->select('first_name as Name', 'last_name as Surname', 'position as Position', 'email')->orderBy('first_name')->get(), null, 'A1', true);
});
})->store('xls', storage_path('app/public/breakers/excel'));
return view('admin/pages/dashboard', compact(['breaks_count', 'authors_count', 'available_count', 'subscription_count', 'total_views']));
}
public function download(Request $request)
{
return \Storage::disk('public')->download($request->path);
}
}
<file_sep>/resources/lang/fr/global.php
<?php
return [
'logo_name' => 'logo_fr',
'sub_title' => 'La Science rencontre la Société',
'latest' => 'Dernières Breaks publiées',
'picks' => 'Sélection du rédacteur en chef',
'popular' => 'Tendance actuelle',
'topics' => 'Sujets populaires',
'in' => 'dans',
'by' => 'par',
'published' => 'publié sur',
'views' => 'vues',
'read' => 'lire',
'reading_time' => 'Temps de lecture',
'at' => 'à',
'click_to_read_more' => 'cliquez pour lire plus',
'search' => 'Rechercher',
'articles' => 'articles',
'sort_bar' => [
'show' => 'afficher',
'sort_by' => 'trier par',
'showing' => 'affiche',
'of' => 'sur',
'all' => 'tout',
'date' => 'date',
'breaks_num' => 'nombre of breaks',
'first_name' => 'prénom',
'last_name' => '<NAME>',
'popular' => 'le plus populaire',
'title' => 'titre',
'reading_time' => 'temps de lecture'
]
];
<file_sep>/app/Http/Requests/ManagerRequest.php
<?php
namespace App\Http\Requests;
use App\Manager;
/*
|--------------------------------------------------------------------------
| VALIDATION
|--------------------------------------------------------------------------
|
| This class performs validaton according to the given rules.
|
*/
class ManagerRequest extends Form
{
public function rules()
{
return [
'title' => 'max:6',
'first_name' => 'required|min:2',
'last_name' => 'required|min:2',
'email' => 'required|email',
'division_id' => 'required',
'avatar' => 'mimes:jpg,jpeg,png,svg|max:6000'
];
}
public function create()
{
$this->slug("$this->first_name $this->last_name");
$avatar = $this->saveFile();
return Manager::create([
'title' => $this->title,
'first_name' => $this->first_name,
'last_name' => $this->last_name,
'slug' => $this->slug,
'email' => $this->email,
'division_id' => $this->division_id,
'position' => $this->position,
'biography' => $this->biography,
'research_institute' => $this->research_institute,
'is_editor' => $this->is_editor,
'image_path' => $avatar,
'is_alumni' => $this->is_alumni ? 1 : 0
]);
}
public function edit(Manager $manager)
{
$this->slug("$this->first_name $this->last_name");
$avatar = $this->saveFile();
$manager->update([
'title' => $this->title,
'first_name' => $this->first_name,
'last_name' => $this->last_name,
'slug' => $this->slug,
'email' => $this->email,
'division_id' => $this->division_id,
'position' => $this->position,
'biography' => $this->biography,
'research_institute' => $this->research_institute,
'is_editor' => $this->is_editor,
'image_path' => $avatar ?? $manager->image_path,
'is_alumni' => $this->is_alumni ? 1 : 0
]);
}
}
<file_sep>/TODO.txt
SHOW CONTACT FORMS AFTER PAGE LOADS
FIX ARCHIVES: THE LAST ONE IS NOT SHOWING WELL<file_sep>/app/Mail/BreakerNewBreak.php
<?php
namespace App\Mail;
use App\Article;
use Illuminate\Bus\Queueable;
use Illuminate\Mail\Mailable;
use Illuminate\Queue\SerializesModels;
use Illuminate\Contracts\Queue\ShouldQueue;
class BreakerNewBreak extends Mailable
{
use Queueable, SerializesModels;
public $user;
public $break;
public function __construct($user, Article $break)
{
$this->user = $user;
$this->break = $break;
}
public function build()
{
return $this->markdown('emails/breaker_new_break')->subject('Break published');
}
}
<file_sep>/database/seeds/AuthorSeeder.php
<?php
use Illuminate\Database\Seeder;
use App\Author;
class AuthorSeeder extends Seeder
{
/**
* Run the database seeds.
*
* @return void
*/
public function run()
{
Author::create([
'first_name' => 'Brian',
'last_name' => 'Griffin',
'slug' => 'brian-griffin',
'email' => '<EMAIL>',
'position' => 'Student',
'research_institute' => 'Quahog',
]);
}
}
<file_sep>/app/Article.php
<?php
namespace App;
use App\TheScienceBreaker;
use App\Manager\Traits\ArticleScopeQueries;
use App\Resources\ArticlePaths;
use App\Resources\ArticleResources;
class Article extends TheScienceBreaker
{
use ArticleScopeQueries;
protected $dates = ['published_at'];
// protected $with = ['authors', 'editor', 'category', 'tags'];
public function getRouteKeyName()
{
return 'slug';
}
public function resources()
{
return new ArticleResources($this);
}
public function paths()
{
return new ArticlePaths($this);
}
public function category()
{
return $this->belongsTo('App\Category');
}
public function editor()
{
return $this->belongsTo('App\Manager');
}
public function authors()
{
return $this->belongsToMany('App\Author')->withPivot('relevance_order')->orderBy('relevance_order')->withTimestamps();
}
public function tags()
{
return $this->belongsToMany('App\Tag');
}
public function scopePublished($query)
{
return $query->where('published_at', '<=', now());
}
public static function currentIssuePath()
{
$lastBreak = Article::published()->orderBy('published_at', 'DESC')->first();
return $lastBreak ?
"content/volume/$lastBreak->volume/issue/$lastBreak->issue"
: null;
}
public static function hours()
{
$formatter = function ($time) {
return date('G:i', $time);
};
$halfHourSteps = range(0, 47*1800, 1800);
return array_map($formatter, $halfHourSteps);
}
}
<file_sep>/app/Http/Controllers/CategoryController.php
<?php
namespace App\Http\Controllers;
use App\{Category, Article};
use Illuminate\Http\Request;
class CategoryController extends Controller
{
public function index()
{
//
}
public function create()
{
//
}
public function store(Request $request)
{
//
}
public function show(Category $category, Request $request)
{
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'title') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 5;
$articles = Article::where('category_id', $category->id)->published()->orderBy($sort, $order)->paginate($show);
return view('pages.category', compact(['articles', 'category']));
}
public function edit(Category $category)
{
//
}
public function update(Request $request, Category $category)
{
//
}
public function destroy(Category $category)
{
//
}
}
<file_sep>/app/Resources/ManagerPaths.php
<?php
namespace App\Resources;
use Illuminate\Support\Facades\File;
class ManagerPaths extends Resources
{
public function route()
{
return "/core-team/".$this->model->slug;
}
public function avatar()
{
return asset($this->model->image_path);
// if (File::exists("storage/app/managers/avatars/".$this->model->slug)) {
// if (count(File::allFiles("storage/app/managers/avatars/".$this->model->slug))) {
// return File::allFiles("storage/app/managers/avatars/".$this->model->slug)[0];
// }
// }
// return "images/missing-avatar.svg";
}
}
<file_sep>/tests/AppAssertions.php
<?php
namespace Tests;
trait AppAssertions
{
protected function seeEmailWasSent()
{
$this->assertNotEmpty($this->emails, 'No emails were sent.');
return $this;
}
protected function seeEmailsSent($count)
{
$emailsSent = count($this->emails);
$this->assertCount(
$count, $this->emails,
"Expected $count emails to have been sent. Number of sent emails: $emailsSent."
);
return $this;
}
protected function seeEmailWasNotSent()
{
$this->assertEmpty($this->emails, 'You did send emails! Did not expect that.');
}
protected function seeEmailTo($recipient)
{
$hasEmail = false;
foreach ($this->emails as $email) {
if (array_key_exists($recipient, $email->getTo())) {
$hasEmail = true;
}
}
$this->assertTrue($hasEmail, "$recipient did not receive an email.");
return $this;
}
protected function seeEmailFrom($sender)
{
$sentEmail = false;
foreach ($this->emails as $email) {
if (array_key_exists($recipient, $email->getFrom())) {
$sentEmail = true;
}
}
$this->assertTrue($sentEmail, "$sender did not send an email.");
return $this;
}
public function seeEmailContains($excerpt)
{
$contains = false;
foreach ($this->emails as $email) {
if (strpos($email->getBody(), $excerpt)) {
$contains = true;
}
}
$this->assertTrue($contains, "No email containing the text has been found.");
return $this;
}
protected function seeEmailSubjectIs($subject)
{
$hasSubject = false;
foreach ($this->emails as $email) {
if ($subject == $email->getSubject()) {
$hasSubject = true;
}
}
$this->assertTrue($hasSubject, "An email with the subject \"$subject\" does not exist.");
return $this;
}
protected function getEmail(Swift_Message $message = null)
{
$this->seeEmailWasSent();
return $message ?: end($this->emails);
}
}<file_sep>/tests/Feature/AdminAvailableArticlesTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
use App\Article;
class AdminAvailableArticlesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_manager_can_add_available_articles()
{
$this->signIn();
$this->post('/admin/available-articles', [
'article' => 'Article Here',
'category_id' => 1
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('available_articles', [
'article' => 'Article Here'
]);
}
/** @test */
public function a_manager_can_remove_an_available_article()
{
$this->signIn();
$article = $this->available_article;
$this->delete('/admin/available-articles/'.$article->id)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('available_articles', [
'id' => $article->id
]);
}
/** @test */
public function a_manager_can_edit_an_available_article()
{
$this->signIn();
$article = $this->available_article;
$this->patch('/admin/available-articles/'.$article->id, [
'article' => 'This is my new article',
'category_id' => $article->category_id
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('available_articles', [
'article' => 'This is my new article'
])->assertDatabaseMissing('available_articles', [
'article' => $article->article
]);
}
}<file_sep>/database/factories/UserFactory.php
<?php
use Faker\Generator as Faker;
/*
|--------------------------------------------------------------------------
| Model Factories
|--------------------------------------------------------------------------
|
| This directory should contain each of the model factory definitions for
| your application. Factories provide a convenient way to generate new
| model instances for testing / seeding your application's database.
|
*/
$factory->define(App\User::class, function (Faker $faker) {
static $password;
$first_name = $faker->firstName;
$last_name = $faker->lastName;
return [
'first_name' => $first_name,
'last_name' => $last_name,
'slug' => str_slug($first_name.' '.$last_name),
'email' => $faker->unique()->safeEmail,
'is_authorized' => 0,
'password' => $password ?: $password = bcrypt('<PASSWORD>'),
'remember_token' => str_random(10),
];
});
$factory->define(App\Article::class, function (Faker $faker) {
$title = $faker->sentence;
return [
'title' => $title,
'slug' => str_slug($title),
'description' => $faker->sentence,
'image_caption' => $faker->sentence,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => 'https://doi.org/10.25250/thescbr.brk001',
'volume' => (new App\Article)->resources()->generateVolume(),
'issue' => (new App\Article)->resources()->generateIssue(),
'editor_pick' => $faker->boolean($chanceOfGettingTrue = 50),
'views' => 0
];
});
$factory->define(App\Author::class, function (Faker $faker) {
$first_name = $faker->firstName;
$last_name = $faker->lastName;
return [
'first_name' => $first_name,
'last_name' => $last_name,
'slug' => str_slug($first_name.' '.$last_name),
'email' =>$faker->unique()->safeEmail,
'position' => $faker->word,
'research_institute' => $faker->sentence,
'field_research' => $faker->word,
'general_comments' => $faker->sentence
];
});
$factory->define(App\ArticleAuthor::class, function (Faker $faker) {
return [
'article_id' => function() {
return factory('App\Article')->create()->id;
},
'author_id' => function() {
return factory('App\Author')->create()->id;
}
];
});
$factory->define(App\Manager::class, function (Faker $faker) {
$first_name = $faker->firstName;
$last_name = $faker->lastName;
return [
'title' => $faker->title,
'first_name' => $first_name,
'last_name' => $last_name,
'slug' => str_slug($first_name.' '.$last_name),
'email' =>$faker->unique()->safeEmail,
'division_id' => '1',
'position' => $faker->word,
'biography' => $faker->paragraph,
'research_institute' => $faker->sentence,
'is_editor' => $faker->boolean($chanceOfGettingTrue = 50)
];
});
$factory->define(App\Category::class, function (Faker $faker) {
$name = $faker->word;
return [
'name' => $name,
'slug' => str_slug($name)
];
});
$factory->define(App\AvailableArticle::class, function (Faker $faker) {
return [
'article' => $faker->sentence,
'category_id' => function() {
return factory('App\Category')->create()->id;
}
];
});
$factory->define(App\Subscription::class, function (Faker $faker) {
return [
'email' => $faker->unique()->safeEmail
];
});
$factory->define(App\Tag::class, function (Faker $faker) {
return [
'name' => $faker->unique()->word
];
});
$factory->define(App\Highlight::class, function (Faker $faker) {
return [
'article_id' => function() {
return factory('App\Article')->create()->id;
}
];
});<file_sep>/tests/Feature/AdminBreakersTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class AdminBreakersTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_manager_can_add_a_new_breaker()
{
$this->signIn();
$this->post('/admin/breakers', [
'first_name' => 'John',
'last_name' => 'Doe',
'email' => '<EMAIL>',
'position' => 'Professor',
'research_institute' => 'Yale',
'field_research' => 'Neurobiology',
'general_comments' => 'John is a new breaker'
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('authors', [
'first_name' => 'John'
]);
}
/** @test */
public function a_manager_can_remove_a_breaker()
{
$this->signIn();
$author = $this->author;
$this->delete('/admin/breakers/'.$author->slug)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('authors', [
'id' => $author->id
]);
}
/** @test */
public function removing_a_breaker_also_removes_its_relationships()
{
$this->signIn();
$author = $this->author;
$break_id = $author->articles[0]->id;
$this->delete('/admin/breakers/'.$author->slug)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('article_author', [
'article_id' => $break_id
]);
}
/** @test */
public function a_manager_can_edit_a_breaker()
{
$this->signIn();
$author = $this->author;
$this->patch('/admin/breakers/'.$author->slug, [
'first_name' => 'Melissa',
'last_name' => $author->last_name,
'email' =>$author->email,
'position' => $author->position,
'research_institute' => $author->research_institute,
'field_research' => $author->field_research,
'general_comments' => $author->general_comments
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('authors', [
'first_name' => 'Melissa'
])->assertDatabaseMissing('authors', [
'first_name' => $author->first_name
]);
}
}<file_sep>/tests/Feature/AdminArticlesTest.php
<?php
namespace Tests\Feature;
use App\ArticleAuthor;
use Tests\TestCase;
use Illuminate\Http\UploadedFile;
use Illuminate\Support\Facades\Storage;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class AdminArticlesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_manager_can_add_a_new_article()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1]
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('articles', [
'title' => $title
]);
}
/** @test */
public function a_manager_can_upload_a_pdf_when_adding_a_new_article()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1],
'pdf' => $file = UploadedFile::fake()->create('document.pdf', 20)
]);
Storage::assertExists('breaks/'.str_slug($title).'.pdf');
}
/** @test */
public function a_manager_can_upload_a_cover_image_when_adding_a_new_article()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$slug = str_slug($title);
$this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1],
'image' => UploadedFile::fake()->create('image.jpeg', 200)
]);
Storage::assertExists('breaks/images/'.$slug.'/'.$slug.'.jpeg');
}
/** @test */
public function a_manager_can_remove_an_article()
{
$this->signIn();
$article = $this->article;
$this->delete('/admin/breaks/'.$article->slug)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('articles', [
'id' => $article->id
]);
}
/** @test */
public function removing_an_article_also_removes_its_relationships()
{
$this->signIn();
$article = $this->article;
$author_id = $article->authors[0]->id;
$this->delete('/admin/breaks/'.$article->slug)->assertSessionHas('db_feedback');
$this->assertDatabaseMissing('article_author', [
'author_id' => $author_id
]);
}
/** @test */
public function a_pdf_is_removed_along_with_a_removed_article()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$article = $this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1],
'pdf' => $file = UploadedFile::fake()->create('document.pdf', 20)
]);
Storage::assertExists('breaks/'.str_slug($title).'.pdf');
$this->delete('/admin/breaks/'.str_slug($title));
Storage::assertMissing('breaks/'.str_slug($title).'.pdf');
}
/** @test */
public function a_cover_image_is_removed_along_with_a_removed_article()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$slug = str_slug($title);
$article = $this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1],
'image' => $file = UploadedFile::fake()->create('image.jpeg', 20)
]);
Storage::assertExists('breaks/images/'.$slug.'/'.$slug.'.jpeg');
$this->delete('/admin/breaks/'.$slug);
Storage::assertMissing('breaks/images/'.$slug.'/'.$slug.'.jpeg');
}
/** @test */
public function a_manager_can_remove_a_cover_image()
{
$this->signIn();
$faker = \Faker\Factory::create();
$title = $faker->sentence;
$slug = str_slug($title);
$article = $this->post('/admin/breaks', [
'title' => $title,
'content' => $faker->paragraph,
'reading_time' => $faker->randomDigitNotNull,
'original_article' => $faker->sentence,
'category_id' => 1,
'editor_id' => 1,
'doi' => $faker->url,
'editor_pick' => '0',
'authors' => [1],
'image' => $file = UploadedFile::fake()->create('image.jpeg', 20)
]);
Storage::assertExists('breaks/images/'.$slug.'/'.$slug.'.jpeg');
$this->delete('/admin/breaks/images/'.$slug);
Storage::assertMissing('breaks/images/'.$slug.'/'.$slug.'.jpeg');
}
/** @test */
public function a_manager_can_edit_an_article()
{
$this->signIn();
$article = $this->article;
$new_author = factory('App\Author')->create();
$this->patch('/admin/breaks/'.$article->slug, [
'title' => 'New title',
'content' => $article->content,
'reading_time' =>$article->reading_time,
'original_article' => $article->original_article,
'category_id' => $article->category_id,
'editor_id' => $article->editor_id,
'doi' => $article->doi,
'editor_pick' => $article->editor_pick,
'authors' => [1, $new_author->id]
])->assertSessionHas('db_feedback');
$this->assertDatabaseHas('articles', [
'title' => 'New title'
])->assertDatabaseMissing('articles', [
'title' => $article->title
]);
}
/** @test */
public function a_manager_can_reorder_the_authors_of_an_article()
{
$this->signIn();
$article = $this->article;
$new_author = factory('App\Author')->create();
$article->authors()->save($new_author);
$this->assertArraySubset(ArticleAuthor::where('article_id', $article->id)->pluck('relevance_order'), [0,0]);
$this->post('/admin/breaks/'.$article->slug.'/breakers-order', [
'order' => [0,1],
'is_original_author' => [1,0]
]);
$this->assertArraySubset(ArticleAuthor::where('article_id', $article->id)->pluck('relevance_order'), [1,0]);
}
/** @test */
public function a_manager_can_select_if_the_breaks_author_is_also_an_author_of_the_originial_paper()
{
$this->signIn();
$article = $this->article;
$new_author = factory('App\Author')->create();
$article->authors()->save($new_author);
$this->post('/admin/breaks/'.$article->slug.'/breakers-order', [
'order' => [1,2],
'is_original_author' => [true,false]
]);
$this->assertArraySubset(ArticleAuthor::where('article_id', $article->id)->pluck('is_original_author'), [1,0]);
}
/** @test */
public function a_manager_can_upload_a_pdf_when_editing_an_article()
{
$this->signIn();
$article = $this->article;
$this->patch('/admin/breaks/'.$article->slug, [
'title' => $article->title,
'content' => $article->content,
'reading_time' =>$article->reading_time,
'original_article' => $article->original_article,
'category_id' => $article->category_id,
'editor_id' => $article->editor_id,
'doi' => $article->doi,
'editor_pick' => $article->editor_pick,
'authors' => [1],
'pdf' => $file = UploadedFile::fake()->create('document.pdf', 20)
]);
Storage::assertExists('breaks/'.str_slug($article->title).'.pdf');
}
/** @test */
public function a_manager_can_upload_a_cover_image_when_editing_an_article()
{
$this->signIn();
$article = $this->article;
$slug = str_slug($article->title);
$this->patch('/admin/breaks/'.$article->slug, [
'title' => $article->title,
'content' => $article->content,
'reading_time' =>$article->reading_time,
'original_article' => $article->original_article,
'category_id' => $article->category_id,
'editor_id' => $article->editor_id,
'doi' => $article->doi,
'editor_pick' => $article->editor_pick,
'authors' => [1],
'image' => $file = UploadedFile::fake()->create('image.jpeg', 200)
]);
Storage::assertExists('breaks/images/'.$slug.'/'.$slug.'.jpeg');
}
/** @test */
public function a_manager_can_change_the_date_a_break_was_created()
{
$this->signIn();
$article = $this->article;
$old_date = $article->created_at;
$this->patch('/admin/breaks/'.$article->slug, [
'title' => $article->title,
'content' => $article->content,
'reading_time' =>$article->reading_time,
'original_article' => $article->original_article,
'category_id' => $article->category_id,
'editor_id' => $article->editor_id,
'doi' => $article->doi,
'editor_pick' => $article->editor_pick,
'authors' => [1],
'created_at' => '06/23/1984'
]);
$this->assertNotEquals($old_date, $article->fresh()->created_at);
}
}<file_sep>/database/seeds/CategorySeeder.php
<?php
use Illuminate\Database\Seeder;
use App\Category;
class CategorySeeder extends Seeder
{
/**
* Run the database seeds.
*
* @return void
*/
public function run()
{
Category::create(['name' => 'Health & Physiology', 'slug' => 'health-physiology']);
Category::create(['name' => 'Neurobiology', 'slug' => 'neurobiology']);
Category::create(['name' => 'Earth & Space', 'slug' => 'earth-space']);
Category::create(['name' => 'Evolution & Behaviour', 'slug' => 'evolution-behaviour']);
Category::create(['name' => 'Plant Biology', 'slug' => 'plant-biology']);
Category::create(['name' => 'Microbiology', 'slug' => 'microbiology']);
Category::create(['name' => 'Maths, Physics & Chemistry', 'slug' => 'maths-physics-chemistry']);
Category::create(['name' => 'Psychology', 'slug' => 'psychology']);
}
}
<file_sep>/app/Xml/Validator.php
<?php
namespace App\Xml;
use Illuminate\Validation\ValidationException;
use App\{Category, Article};
class Validator
{
function __construct(array $request)
{
$this->request = $request;
}
public function breakers()
{
$breakers = $this->request['authors']['author'] ?? null;
if (! $breakers)
throw ValidationException::withMessages(['authors' => 'The author is missing']);
$data = [];
$keys = array_keys($breakers);
if ($keys[0] === 0) {
foreach ($breakers as $breaker) {
$info = [
'first_name' => $breaker['givenname'] ?? null,
'last_name' => $breaker['familyname'] ?? null,
'email' => $breaker['email'] ?? null,
'position' => $breaker['biography'] ?? null,
// 'position' => $breaker['biography'] ? preg_replace('/\s/', ' ', strip_tags($breaker['biography'])) : null,
'research_institute' => $breaker['affiliation'] ?? null,
];
$this->sanitize($info, $except = ['research_institute']);
array_push($data, $info);
}
} else {
$info = [
'first_name' => $breakers['givenname'] ?? null,
'last_name' => $breakers['familyname'] ?? null,
'email' => $breakers['email'] ?? null,
'position' => $breakers['biography'] ?? null,
// 'position' => $breaker['biography'] ? preg_replace('/\s/', ' ', strip_tags($breaker['biography'])) : null,
'research_institute' => $breakers['affiliation'] ?? null,
];
$this->sanitize($info, $except = ['research_institute']);
array_push($data, $info);
}
return $data;
}
public function break()
{
$data = [
'title' => $this->request['title'] ?? null,
'description' => $this->request['abstract'] ?? null,
'cover_image' => $this->request['covers']['cover']['cover_image'] ?? null,
'reading_time' => $this->request['subjects']['subject'] ?? null,
'original_article' => $this->request['citations']['citation'] ?? null,
'category_id' => Category::byName($this->request['@attributes']['section_ref'] ?? null)->first()->id ?? null,
'doi' => $this->request['id'][1] ?? null,
];
$this->sanitize($data);
if (Article::where('title', 'LIKE', '%'.$data['title'].'%')->exists())
throw ValidationException::withMessages(['break' => 'This break already exists']);
return $data;
}
public function keywords()
{
return $this->request['keywords']['keyword'] ?? [];
}
public function sanitize($data, $except = [])
{
foreach ($data as $field => $value) {
if (! in_array($field, $except) && ! $value)
throw ValidationException::withMessages([$field => 'The '. $field .' is missing']);
}
}
}<file_sep>/resources/lang/en/search.php
<?php
return [
'searching_for' => 'Searching for',
'we_found' => 'We found',
'results' => 'results'
];<file_sep>/tests/Unit/AuthorsTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class AuthorsTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function authors_can_have_more_than_one_article()
{
$second_article = factory('App\Article')->create();
$author = $this->author;
factory('App\ArticleAuthor')->create([
'article_id' => $second_article->id,
'author_id' => $author->id
]);
$this->assertEquals(2, count($author->articles));
}
/** @test */
public function authors_have_their_own_page()
{
$author = $this->author;
$this->get($author->paths()->route())->assertSee($author->first_name);
}
/** @test */
public function authors_know_if_they_are_the_original_authors()
{
$this->signIn();
$article = $this->article;
$new_author = factory('App\Author')->create();
$article->authors()->save($new_author);
$this->post('/admin/breaks/'.$article->slug.'/breakers-order', [
'order' => [1,2],
'is_original_author' => [0,1]
]);
$this->assertEquals(0, $this->author->isOriginalAuthorOf($article->id));
$this->assertEquals(1, $new_author->isOriginalAuthorOf($article->id));
}
}
<file_sep>/tests/Feature/SearchTest.php
<?php
namespace Tests\Feature;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class SearchTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_guest_can_find_an_article_searching_for_a_title()
{
$article = $this->article;
$title = explode(' ',trim($article->title));
$word = $title[0];
$this->get("/search?for=$word")->assertSee($article->title);
}
/** @test */
public function a_guest_can_find_an_article_searching_for_a_keyword()
{
$article = $this->article;
$content = explode(' ',trim($article->content));
$word = $content[0];
$this->get("/search?for=$word")->assertSee($article->title);
}
/** @test */
public function a_guest_can_find_an_article_searching_for_the_authors_name()
{
$article = $this->article;
$author = explode(' ',trim($article->authors()->first()->first_name));
$name = $author[0];
$this->get("/search?for=$name")->assertSee($article->title);
}
/** @test */
public function a_guest_can_find_an_article_searching_for_the_editors_name()
{
$article = $this->article;
$editor = explode(' ',trim($article->editor->first_name));
$name = $editor[0];
$this->get("/search?for=$name")->assertSee($article->title);
}
/** @test */
public function a_guest_can_find_an_article_searching_for_a_category()
{
$article = $this->article;
$category = explode(' ',trim($article->category->name));
$category = $category[0];
$this->get("/search?for=$category")->assertSee($article->title);
}
/** @test */
public function a_guest_can_find_an_article_searching_for_a_tag()
{
$article = $this->article;
$tag = $this->tag;
$article->tags()->attach($tag);
$tag = $tag->name;
$this->get("/search?for=$tag")->assertSee($article->title);
}
}<file_sep>/resources/lang/en/categories.php
<?php
return [
'health-physiology' => 'Health & Physiology',
'neurobiology' => 'Neurobiology',
'earth-space' => 'Earth & Space',
'evolution-behaviour' => 'Evolution & Behaviour',
'plant-biology' => 'Plant Biology',
'microbiology' => 'Microbiology',
'maths-physics-chemistry' => 'Maths, Physics & Chemistry',
'psychology' => 'Psychology'
];<file_sep>/tests/TestCase.php
<?php
namespace Tests;
use App\Exceptions\Handler;
use Illuminate\Contracts\Debug\ExceptionHandler;
use Illuminate\Foundation\Testing\TestCase as BaseTestCase;
abstract class TestCase extends BaseTestCase
{
use CreatesApplication;
public function setUp()
{
parent::setUp();
$this->disableExceptionHandling();
$this->manager = factory('App\Manager')->create();
$this->author = factory('App\Author')->create();
$this->editor = factory('App\Manager')->create([
'is_editor' => 1
]);
$this->category = factory('App\Category')->create();
$this->article = factory('App\Article')->create([
'editor_id' => $this->editor->id,
'category_id' => $this->category->id
]);
$this->article_author = factory('App\ArticleAuthor')->create([
'article_id' => $this->article->id,
'author_id' => $this->author->id
]);
$this->available_article = factory('App\AvailableArticle')->create();
$this->subscription = factory('App\Subscription')->create();
$this->tag = factory('App\Tag')->create();
}
protected function signIn($user = null)
{
$user = $user ?: create('App\User', ['is_authorized' => 1]);
$this->actingAs($user);
return $this;
}
protected function disableExceptionHandling()
{
$this->oldExceptionHandler = $this->app->make(ExceptionHandler::class);
$this->app->instance(ExceptionHandler::class, new class extends Handler {
public function __construct(){}
public function report(\Exception $e) {}
public function render($request, \Exception $e) {
throw $e;
}
});
}
protected function withExceptionHandling()
{
$this->app->instance(ExceptionHandler::class, $this->oldExceptionHandler);
return $this;
}
}
<file_sep>/app/Author.php
<?php
namespace App;
use App\TheScienceBreaker;
use App\Resources\AuthorPaths;
use App\Resources\AuthorResources;
class Author extends TheScienceBreaker
{
protected $withCount = ['articles'];
public function getRouteKeyName()
{
return 'slug';
}
public function resources()
{
return new AuthorResources($this);
}
public function paths()
{
return new AuthorPaths($this);
}
public function articles()
{
return $this->belongsToMany('App\Article')->withTimestamps();
}
public function isOriginalAuthorOf($articleId)
{
return ArticleAuthor::where([
'article_id' => $articleId,
'author_id' => $this->id,
])->value('is_original_author');
}
}
<file_sep>/resources/lang/fr/search.php
<?php
return [
'searching_for' => 'À la recherche de',
'we_found' => 'Nous avons trouvé',
'results' => 'résultats'
];<file_sep>/app/Http/Middleware/Localization.php
<?php
namespace App\Http\Middleware;
use Closure, Session;
class Localization
{
/**
* The availables languages.
*
* @array $languages
*/
protected $languages = ['en'];
/**
* Handle an incoming request.
*
* @param \Illuminate\Http\Request $request
* @param \Closure $next
* @return mixed
*/
public function handle($request, Closure $next)
{
$browserLang = substr($request->server('HTTP_ACCEPT_LANGUAGE'), 0, 2);
if (!in_array($browserLang, $this->languages)) {
$browserLang = 'en';
}
$appLang = $browserLang;
if (Session::has('lang')) {
$appLang = Session::get('lang');
}
app()->setLocale($appLang);
return $next($request);
}
}
<file_sep>/app/Http/Requests/AuthorRequest.php
<?php
namespace App\Http\Requests;
use App\Author;
/*
|--------------------------------------------------------------------------
| VALIDATION
|--------------------------------------------------------------------------
|
| This class performs validaton according to the given rules.
|
*/
class AuthorRequest extends Form
{
public function rules()
{
return [
'first_name' => 'required|min:2',
'last_name' => 'required|min:2',
'email' => 'email',
'position' => 'required',
'research_institute' => 'required'
];
}
public function create()
{
$this->slug("$this->first_name $this->last_name");
return Author::create([
'first_name' => $this->first_name,
'last_name' => $this->last_name,
'slug' => $this->slug,
'email' => $this->email,
'position' => $this->position,
'research_institute' => $this->research_institute,
'field_research' => $this->field_research,
'general_comments' => $this->general_comments
]);
}
public function edit(Author $author)
{
$this->slug("$this->first_name $this->last_name");
$author->update([
'first_name' => $this->first_name,
'last_name' => $this->last_name,
'slug' => $this->slug,
'email' => $this->email,
'position' => $this->position,
'research_institute' => $this->research_institute,
'field_research' => $this->field_research,
'general_comments' => $this->general_comments
]);
}
}
<file_sep>/app/Http/Controllers/FeedController.php
<?php
namespace App\Http\Controllers;
use App\Tag;
use App\Article;
use Illuminate\Support\Facades\Response;
class FeedController extends Controller
{
public function index()
{
$tags = Tag::all();
$articles = Article::orderBy('created_at', 'DESC')->get();
$content = view('feed', compact(['articles', 'tags']));
return Response::make($content, '200')->header('Content-Type', 'text/xml');
}
}
<file_sep>/app/Highlight.php
<?php
namespace App;
use App\TheScienceBreaker;
class Highlight extends TheScienceBreaker
{
public function article()
{
return $this->belongsTo(Article::class)->with(['category', 'authors'])->published();
}
}
<file_sep>/app/Http/Requests/ArticleRequest.php
<?php
namespace App\Http\Requests;
use App\Article;
use Carbon\Carbon;
/*
|--------------------------------------------------------------------------
| VALIDATION
|--------------------------------------------------------------------------
|
| This class performs validaton according to the given rules.
|
*/
class ArticleRequest extends Form
{
public function rules()
{
return [
'title' => 'required|max:255',
'title_fr' => 'max:255',
'description' => 'max:500',
'description_fr' => 'max:500',
'content' => 'required',
'reading_time' => 'required',
'original_article' => 'required',
'category_id' => 'required',
'editor_id' => 'required',
'pdf' => 'mimes:pdf',
'image' => 'mimes:jpg,jpeg,png,svg|max:800',
'image_caption' => 'max:255',
'image_credits' => 'max:144'
];
}
public function create()
{
$this->slug($this->title);
$article = Article::create([
'title' => $this->title,
'title_fr' => $this->title_fr,
'slug' => $this->slug,
'description' => $this->description,
'description_fr' => $this->description_fr,
'image_path' => $this->saveFile(),
'image_caption' => $this->image_caption,
'image_credits' => $this->image_credits,
'content' => $this->content,
'content_fr' => $this->content_fr,
'reading_time' => $this->reading_time,
'original_article' => $this->original_article,
'category_id' => $this->category_id,
'editor_id' => $this->editor_id,
'doi' => (new Article)->resources()->createDoi(),
'issue' => (new Article)->resources()->generateIssue(),
'volume' => (new Article)->resources()->generateVolume(),
'editor_pick' => $this->editor_pick ?? false,
'published_at' => $this->published_at ? Carbon::parse($this->published_at . $this->published_at_time) : null
]);
foreach ($this->authors as $author) {
$article->authors()->attach($author);
}
$article->tags()->attach($this->tags);
return $article;
}
public function edit(Article $article)
{
$this->slug($this->title);
$article->update([
'title' => $this->title,
'title_fr' => $this->title_fr,
'description' => $this->description,
'description_fr' => $this->description_fr,
'image_caption' => $this->image_caption,
'image_credits' => $this->image_credits,
'content' => $this->content,
'content_fr' => $this->content_fr,
'reading_time' => $this->reading_time,
'original_article' => $this->original_article,
'doi' => $this->doi,
'category_id' => $this->category_id,
'editor_id' => $this->editor_id,
'editor_pick' => $this->editor_pick ?? false,
'created_at' => $this->created_at,
'published_at' => $this->published_at ? Carbon::parse($this->published_at . $this->published_at_time) : null
]);
// $path = $this->saveFile();
// if ($path)
// $article->update(['image_path' => $path]);
if ($this->update_url)
$article->update(['slug' => $this->slug]);
$article->authors()->sync($this->authors);
}
}
<file_sep>/resources/lang/fr/footer.php
<?php
return [
'follow' => 'SUIVEZ-NOUS',
'subscribe' => [
'title' => 'SOUSCRIRE',
'text' => 'Abonnez-vous pour être tenus au courant des dernières Breaks publiées!',
'input' => 'Entrez votre adresse e-mail ici',
'note' => 'Nous ne diffuserons jamais votre adresse email',
'button' => 'Inscrivez-moi'
],
'copyright' => 'Tous droits réservés',
];<file_sep>/app/Resources/ArticleResources.php
<?php
namespace App\Resources;
use Illuminate\Support\Collection;
use App\{Article, Tag};
use App;
class ArticleResources extends Resources
{
public function tagsIds()
{
return $this->model->tags->pluck('id')->toArray();
}
public function tagsList()
{
$tagsList = '';
foreach ($this->model->tags as $tag) {
$tagsList .= $tag->name.', ';
}
$tagsList = substr($tagsList, 0, -2);
return $tagsList;
}
public function authorsIds()
{
return $this->model->authors->pluck('id')->toArray();
}
public function authorsList()
{
$namesList = '';
foreach ($this->model->authors as $author) {
$namesList .= $author->resources()->fullName().', ';
}
$namesList = substr($namesList, 0, -2);
return $namesList;
}
public function preview()
{
$pieces = explode(" ", strip_tags($this->model->content, '<br>'));
return implode(" ", array_splice($pieces, 0, 45));
}
public function nextRead()
{
$collection = $this->model->tags()->inRandomOrder()->first() ?? $this->model->category;
$results = $collection->articles()->with(['category', 'authors'])->where('slug', '!=', $this->model->slug);
if ($results->exists())
return $results->first();
return $this->model->category->articles()->with(['category', 'authors'])->where('slug', '!=', $this->model->slug)->first();
}
public function suggestions()
{
$collection = Tag::with('articles')->whereIn('id', $this->model->tags->pluck('id'))
->get()
->pluck('articles')
->flatten()
->unique('id')
->where('id', '!=', $this->model->id)
->take(5);
return $collection->isEmpty() ?
Article::with(['category', 'authors'])->published()->except($this->model)->inRandomOrder()->take(5)->get() :
$collection;
}
public function createDoi()
{
$doi_base = "https://doi.org/10.25250/thescbr.brk";
$last_doi = $this->model::orderBy('id', 'desc')->first()->doi;
$current_number = (int)substr($last_doi, -3);
$current_number+=1;
$doi_number = str_pad($current_number, 3, '0', STR_PAD_LEFT);
return $doi_base.$doi_number;
}
public function localize($input)
{
$column = $input.'_'.App::getLocale();
$request = ($this->model->$column) ? $this->model->$column : $this->model->$input;
return $request;
}
public function generateVolume($published_at = null)
{
$date = $published_at ?? \Carbon\Carbon::now();
return $date->year - 2014;
}
public function generateIssue($published_at = null)
{
$date = $published_at ?? \Carbon\Carbon::now();
$issue = 0;
if ($date->month >= 1 && $date->month <= 3) {
$issue += 1;
} else if ($date->month >= 4 && $date->month <= 6) {
$issue += 2;
} else if ($date->month >= 7 && $date->month <= 9) {
$issue += 3;
} else if ($date->month >= 10 && $date->month <= 12) {
$issue += 4;
}
return $issue;
}
// public static function highlights()
// {
// return self::where('highlight', 1)->orderBy('title')->get();
// }
// public static function last()
// {
// return self::orderBy('id', 'desc')->first();
// }
// public static function random()
// {
// return self::inRandomOrder()->first();
// }
// public static function generateSlugs()
// {
// foreach ($this->model::all() as $model) {
// $model->update([
// 'slug' => str_slug($model->title)
// ]);
// }
// }
}
<file_sep>/app/Http/Controllers/AppController.php
<?php
namespace App\Http\Controllers;
use App\Article;
use App\Highlight;
use Illuminate\Support\Facades\DB;
use Illuminate\Support\Facades\File;
use Illuminate\Http\Request;
class AppController extends Controller
{
public function breaks()
{
$articles = Article::orderBy('created_at', 'DESC')->get();
foreach ($articles as $article) {
$article->doi = "https://thesciencebreaker.org/breaks/".$article->category->slug."/$article->slug";
}
$this->addImage($articles);
return $articles;
}
public function incrementViews(Request $request)
{
$break = Article::findOrFail($request->id)->increment('views');
return $break;
}
public function disqus(Request $request)
{
$article = Article::where('id', $request->id)->get();
return view('pages/app/disqus', compact(['article']));
}
public function home()
{
$results = [];
$results['a_latest'] = $this->latest();
$results['b_highlights'] = $this->highlights();
$results['c_popular'] = $this->popular();
return $results;
}
public function highlights()
{
$articles = [];
$highlights = Highlight::orderBy('relevance_order')->get();
foreach ($highlights as $highlight) {
array_push($articles, $highlight->article);
}
$this->addImage($articles);
return $articles;
}
public function popular()
{
$articles = Article::popular(7)->get();
$this->addImage($articles);
return $articles;
}
public function latest()
{
$articles = Article::recent(7)->get();
$this->addImage($articles);
return $articles;
}
public function picks()
{
$articles = Article::editorPicks()->get();
$this->addImage($articles);
return $articles;
}
public function suggestions(Request $request)
{
$article = Article::find($request->id);
if (!isset($article)) {
return null;
}
$articles = $article->resources()->suggestions();
$this->addImage($articles);
return $articles;
}
public function addImage($collection)
{
foreach ($collection as $break) {
$break->image_path = $break->paths()->image();
}
}
}
<file_sep>/app/Http/Controllers/ArticlesController.php
<?php
namespace App\Http\Controllers;
use App\{Article, Highlight, Author, ArticleAuthor, Category, Manager, Tag};
use Illuminate\Support\Facades\Storage;
use Illuminate\Http\Request;
use Illuminate\Support\Facades\File;
use App\Http\Requests\ArticleRequest;
use Illuminate\Support\Facades\Lang;
use App\Xml\Generator;
use Carbon\Carbon;
class ArticlesController extends Controller
{
public function __construct()
{
$this->middleware('auth', ['except' => ['index', 'show', 'fix', 'showData']]);
}
public function fix()
{
return now()->timestamp;
}
public function showXml()
{
$editors = Manager::editors()->get();
return view('admin/pages/breaks/xml', compact('editors'));
}
public function showData(Request $request, $id)
{
if ($request->id === '$2y$10$P75HCBUqXIO65SvfjKQ4vOg0vCzAiTAmWrZ2YtGhGMTIafOsfIo4e')
return Article::where('id', $id)->with(['authors'])->first();
abort(404);
}
public function uploadXml(Request $request)
{
$request->validate([
'xml' => 'required|file|mimes:xml',
'content' => 'required',
'editor_id' => 'required',
'image_caption' => 'max:255',
'image_credits' => 'max:144'
]);
$file = $request->xml;
$generator = (new Generator($request->xml));
$break = $generator->createBreak([
'editor_id' => $request->editor_id,
'image_caption' => $request->image_caption,
'image_credits' => $request->image_credits,
'content' => $request->content,
'published_at' => $request->published_at ? Carbon::parse($request->published_at . $request->published_at_time) : null
]);
foreach ($generator->createBreakers() as $breaker) {
$break->authors()->attach($breaker);
}
$break->tags()->attach($generator->createKeywords());
return redirect()->back()->with('db_feedback', 'The Break has been created');
}
public function index()
{
$popular = Article::published()->where('published_at', '>=', now()->subMonths(3))->popular(6)->get();
$topics = Tag::orderBy('articles_count', 'DESC')->take(25)->get();
$highlights = Highlight::with('article')->orderBy('relevance_order')->take(4)->get();
$latest_articles = Article::published()->recent(6)->get();
return view('pages.welcome', compact(['highlights', 'latest_articles', 'topics', 'popular']));
}
public function previewDOI()
{
return (new Article)->resources()->createDoi();
}
// CREATE
public function create()
{
$editors = Manager::editors()->get();
$tags = Tag::orderBy('name')->get();
$authors = Author::orderBy('first_name')->get();
return view('admin/pages/breaks/add', compact(['editors', 'authors', 'tags']));
}
public function store(Request $request)
{
ArticleRequest::get()->save();
return redirect()->back()->with('db_feedback', 'The Break has been successfully added!');
}
// READ
public function show(Category $category, Article $article)
{
if (! $article->published_at)
abort(404);
$more_like = $article->resources()->suggestions();
$next_read = $more_like->pop();
$more_from = $article->similar()->published()->get();
$article->increment('views');
return view('pages.article', compact(['article', 'more_from', 'more_like', 'next_read']));
}
// UPDATE
public function edit(Article $article = null)
{
$authors = Author::orderBy('first_name')->get();
$breaks = Article::orderBy('created_at')->get();
// return $breaks;
$tags = Tag::orderBy('name')->get();
$editors = Manager::editors()->get();
return view('admin/pages/breaks/edit', compact(['editors', 'article', 'authors', 'tags', 'breaks']));
}
public function update(Request $request, Article $article)
{
ArticleRequest::get()->update($article);
if ($request->update_url) {
return redirect("/admin/breaks/{$article->slug}/edit")->with('db_feedback', 'The Break has been updated');
}
return redirect()->back()->with('db_feedback', 'The Break has been updated');
}
public function setTags(Request $request, Article $article)
{
$article->tags()->sync($request->tags);
}
public function authorsOrder(Request $request, Article $article)
{
foreach ($request->order as $index => $author) {
ArticleAuthor::where([
'article_id' => $article->id,
'author_id' => $author
])->update([
'relevance_order' => $index,
'is_original_author' => $request->is_original_author[$index]
]);
}
}
// DELETE
public function selectDelete()
{
$breaksByCategory = Category::with(['articles' => function($query) {
return $query->orderBy('created_at', 'DESC');
}])->get();
return view('admin/pages/breaks/selectDelete', compact(['breaksByCategory']));
}
public function destroy(Article $article)
{
$article->authors()->detach();
Storage::delete("breaks/$article->slug.pdf");
File::deleteDirectory("storage/app/breaks/images/$article->slug");
$article->delete();
return redirect()->back()->with('db_feedback', 'The Break has been deleted');
}
public function destroyImage(Article $article)
{
$deleted = File::deleteDirectory("storage/app/breaks/images/$article->slug");
return redirect()->back()->with('db_feedback', 'The image has been deleted');
}
public function generateIssues()
{
$articles = Article::all();
$articles->each(function($article) {
$article->issue = $article->resources()->generateIssue($article->published_at);
$article->save();
});
return Article::all();
}
public function generateVolumes()
{
$articles = Article::all();
$articles->each(function($article) {
$article->volume = $article->resources()->generateVolume($article->published_at);
$article->save();
});
return Article::all();
}
public function issues()
{
$results = Article::selectRaw('year(published_at) AS year, SUBSTRING(issue, 5) AS issue, volume, count(*) as count')
->groupBy('year', 'issue', 'volume')
->orderBy('year', 'DESC')
->orderBy('issue', 'DESC')
->get();
// $issues = $results->groupBy('volume');
return $results;
return view('issues', compact('issues'));
}
}
<file_sep>/app/Console/Commands/SaveImagesPaths.php
<?php
namespace App\Console\Commands;
use Illuminate\Console\Command;
class SaveImagesPaths extends Command
{
/**
* The name and signature of the console command.
*
* @var string
*/
protected $signature = 'temp:save-paths';
/**
* The console command description.
*
* @var string
*/
protected $description = 'Save all image paths to database';
/**
* Create a new command instance.
*
* @return void
*/
public function __construct()
{
parent::__construct();
}
/**
* Execute the console command.
*
* @return mixed
*/
public function handle()
{
foreach (\App\Article::all() as $article) {
$article->update(['image_path' => $article->paths()->image()]);
}
}
}
<file_sep>/resources/lang/fr/welcome.php
<?php
return [
'description' => [
'why-title' => 'Mission éditoriale',
'why-text' => 'En tant que journal en ligne, open-access et outreach, nous encourageons la démocratisation de la littérature scientifique afin de favoriser les dialogues et l\'intérêt pour les avancées scientifiques les plus récentes. Découvrez notre <a href="https://www.thesciencebreaker.org/mission" target="_blank" rel="noopener">mission</a>.',
'what-title' => 'Contenu du journal',
'what-text' => 'Nous publions de brefs résumés vulgarisés ("<i>breaks</i>") de la recherche scientifique. Nos auteurs sont des scientifiques impliqués dans le domaine de la recherche résumée. Nos lecteurs sont des universitaires et grand public également. <a href="https://www.thesciencebreaker.org/about" target="_blank" rel="noopener">En savoir plus</a>'
],
'partners' => 'Collaborations',
'categories' => 'Breaks par thématique',
'app' => 'Nous avons lancé notre application iOS, <strong>téléchargez maintenant</strong>!',
'highlights' => 'A la une'
];
<file_sep>/app/Xml/Generator.php
<?php
namespace App\Xml;
use App\Xml\Validator;
use App\{Article, Tag, Author};
class Generator
{
protected $publication;
function __construct($file)
{
$path = \Storage::putFileAs('xml', $file, 'file.xml');
$xmlObject = simplexml_load_string(\Storage::get($path));
$json = json_encode($xmlObject);
$xmlData = json_decode($json, true);
\Storage::delete($path);
$validator = (new Validator($xmlData['publication']));
$this->validatedBreak = $validator->break();
$this->validatedBreakers = $validator->breakers();
$this->validatedKeywords = $validator->keywords();
}
public function createBreak($attributes)
{
$data = [
'title' => $this->validatedBreak['title'],
'slug' => str_slug($this->validatedBreak['title']),
'description' => strip_tags($this->validatedBreak['description']),
'image_path' => 'https://oap.unige.ch/journals/public/journals/8/' . $this->validatedBreak['cover_image'],
'reading_time' => floatval($this->validatedBreak['reading_time']),
'original_article' => $this->validatedBreak['original_article'],
'category_id' => $this->validatedBreak['category_id'],
'doi' => 'https://doi.org/' . $this->validatedBreak['doi'],
'issue' => (new Article)->resources()->generateIssue(),
'volume' => (new Article)->resources()->generateVolume(),
];
return Article::create(array_merge($data, $attributes));
}
public function createBreakers()
{
$breakers = collect();
foreach ($this->validatedBreakers as $breaker) {
$breakers->push(Author::firstOrCreate([
'first_name' => $breaker['first_name'],
'last_name' => $breaker['last_name']
], [
'slug' => str_slug($breaker['first_name'].' '.$breaker['last_name']),
'email' => $breaker['email'],
'position' => strip_tags($breaker['position']),
'research_institute' => strip_tags($breaker['research_institute']),
]));
}
return $breakers;
}
public function createKeywords()
{
$tags = collect();
foreach ($this->validatedKeywords as $keyword) {
$tags->push(Tag::firstOrCreate(['name' => $keyword])->id);
}
return $tags;
}
}<file_sep>/app/Suggestion.php
<?php
namespace App;
use App\Article;
class Suggestion {
public static function byTag($article)
{
$tag = $article->tags()->having('articles_count', '>', 1)->inRandomOrder()->first();
if (is_null($tag)) {
return Article::inRandomOrder()->take(4)->get();
}
return $tag->articles()->where('slug', '!=', $article->slug)->take(4)->get();
}
public static function one($article)
{
$suggestion = $article->tags()->having('articles_count', '>', 1)->inRandomOrder()->first();
if (is_null($suggestion)) {
$suggestion = $article->category;
}
$article = $suggestion->articles()->inRandomOrder()->whereNotIn('id', [$article->id])->first();
return $article;
}
}<file_sep>/app/AvailableArticle.php
<?php
namespace App;
use App\TheScienceBreaker;
class AvailableArticle extends TheScienceBreaker
{
protected $table = 'available_articles';
protected $with = ['category'];
public function category()
{
return $this->belongsTo('App\Category');
}
}
<file_sep>/resources/lang/fr/available_articles.php
<?php
return [
'description' => [
'p1' => 'Ci-dessous la liste des articles pour lesquels une Break a été demandée. N\'hésitez pas à',
'get_in_touch' => 'nous contacter',
'p2' => 'si vous souhaitez soumettre votre manuscrit!'
]
];<file_sep>/app/Mail/MailFactory.php
<?php
namespace App\Mail;
use Mail;
use Illuminate\Http\Request;
use App\Mail\BreakerNewBreak;
use App\Mail\EditorNewBreak;
use App\Mail\Welcome;
use App\Mail\Contact\BreakInquiry;
use App\Mail\Contact\Question;
use App\Mail\Contact\Submit;
use App\Mail\Contact\ContactFeedback;
use App\Manager;
use App\Author;
use App\Article;
use Carbon\Carbon;
use Illuminate\Support\Facades\Storage;
use Illuminate\Support\Facades\File;
use Illuminate\Queue\Queue;
class MailFactory
{
public static function sendWelcomeEmail(Author $breaker)
{
Mail::to($breaker->email)->send(new Welcome($breaker));
}
public static function sendNotificationsTo($authors, $editor_id, Article $break)
{
$editor = Manager::find($editor_id);
foreach ($authors as $author) {
$breaker = Author::find($author);
Mail::to($breaker->email)->send(new BreakerNewBreak($breaker, $break));
}
Mail::to($editor->email)->send(new EditorNewBreak($editor, $break));
}
// Contact Page
public static function question(Request $request)
{
$message = [
'subject' => 'Your message to TheScienceBreaker',
'body' => 'Thank you for your contact! We have received your message and will get back to you shortly.'
];
Mail::to(config('app.email'))->send(new Question($request->only(['first_name', 'last_name', 'email', 'message'])));
Mail::to($request->email)->send(new ContactFeedback($request, $message));
}
public static function breakInquiry(Request $request)
{
$message = [
'subject' => 'Your Break inquiry',
'body' => 'We have received your Break inquiry. We will get back to you shortly!'
];
Mail::to(config('app.email'))->send(new BreakInquiry($request->except(['_token', 'time', 'subscribe_me'])));
Mail::to($request->email)->send(new ContactFeedback($request, $message));
}
public static function submit(Request $request)
{
$message = [
'subject' => 'Your Break has been submitted!',
'body' => 'Thank you for submitting your Break! We will review your article and you will be notified once the Break is published.'
];
$file = self::saveFile($request);
Mail::to(config('app.email'))->send(new Submit($request->except(['_token', 'time', 'subscribe_me', 'file']), $file));
Mail::to($request->institution_email)->send(new ContactFeedback($request, $message));
}
protected static function saveFile(Request $request)
{
$file = $request->file('file');
$ext = $file->extension();
$name = $request->last_name.'_'.$request->first_name.'_break_v1';
$filename = "/uploaded-breaks/$name.$ext";
Storage::put('public' . $filename, File::get($file));
return "storage" . $filename;
}
}<file_sep>/database/migrations/2017_10_10_215210_create_managers_table.php
<?php
use Illuminate\Support\Facades\Schema;
use Illuminate\Database\Schema\Blueprint;
use Illuminate\Database\Migrations\Migration;
class CreateManagersTable extends Migration
{
/**
* Run the migrations.
*
* @return void
*/
public function up()
{
Schema::create('managers', function (Blueprint $table) {
$table->increments('id');
$table->string('title')->nullable();
$table->string('first_name');
$table->string('last_name');
$table->string('slug')->slug();
$table->string('email')->unique();
$table->unsignedInteger('division_id');
$table->string('position')->nullable();
$table->text('biography')->nullable();
$table->string('research_institute');
$table->string('image_path')->nullable();
$table->boolean('is_editor');
$table->boolean('is_alumni')->default(false);
$table->index(['first_name', 'last_name']);
$table->timestamps();
});
}
/**
* Reverse the migrations.
*
* @return void
*/
public function down()
{
Schema::dropIfExists('managers');
}
}
<file_sep>/public/js/stickyMenu.js
(function($) {
"use strict";
// Set up sticky menu
$('nav').addClass('original').clone().removeClass('mt-3 original').appendTo('header').addClass('pl-4 pr-4 clone');
$('.clone').find('.logo').show();
var sideBar = $('#side-bar');
var navHeight = $('nav').outerHeight();
var headerHeight = $('header').outerHeight();
var dist = $('main').offset().top;
$(window).scroll(function () {
var scroll = $(this).scrollTop();
if (scroll < dist) {
//Sticky menu
$('.original').css('visibility', 'visible');
$('.clone').fadeOut(100);
// Scroll top button
$('.scroll-to-top').fadeOut();
} else if (scroll > dist) {
//Sticky menu
$('.original').css('visibility', 'hidden');
$('.clone').fadeIn(200);
// Scroll top button
$('.scroll-to-top').fadeIn();
}
});
$(document).on('click', 'a.scroll-to-top', function(event) {
var $anchor = $(this);
// Scroll with ease
$('html, body').stop().animate({
scrollTop: ($($anchor.attr('href')).offset().top)
}, 1000, 'easeInOutExpo');
event.preventDefault();
});
})(jQuery);<file_sep>/app/Http/Controllers/SubscriptionsController.php
<?php
namespace App\Http\Controllers;
use Carbon\Carbon;
use App\Subscription;
use Illuminate\Http\Request;
use Maatwebsite\Excel\Facades\Excel;
class SubscriptionsController extends Controller
{
public function __construct()
{
$this->middleware('throttle:2')->only('store');
$this->middleware('auth', ['except' => ['store', 'unsubscribe', 'form']]);
}
// CREATE
public function store(Request $request)
{
if (app()->environment() != 'testing') {
if (Carbon::parse($request->time)->addSeconds(5)->gt(Carbon::now()) || ! empty($request->my_name))
return response('Humans only please.', 403);
}
$request->validate([
'subscription' => 'required|email|unique:subscriptions,email'
]);
Subscription::create(['email' => $request->subscription]);
return redirect()->back()->with('db_feedback', 'The email has been subscribed')->with('subscription', 'Thank you for subscribing!');
}
public function form()
{
return view('components.forms.subscription')->render();
}
// READ
public function index(Request $request)
{
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'email') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 20;
$subscriptions = Subscription::orderBy($sort, $order)->paginate($show);
$subscriptions_count = Subscription::count();
$excel = Excel::create('subscriptions', function($excel) {
$excel->sheet('Subscriptions', function($sheet) {
$sheet->fromModel(Subscription::select('email as Email', 'created_at as Date')->orderBy('created_at')->get(), null, 'A1', true);
});
})->store('xls', storage_path('app/subscriptions/excel'));
$csv = Excel::create('subscriptions', function($excel) use ($subscriptions) {
$excel->sheet('Subscriptions', function($sheet) use ($subscriptions) {
$sheet->fromModel(Subscription::select('email')->orderBy('created_at')->get(), null, 'A1', true);
});
})->store('csv', storage_path('app/subscriptions/csv'));
return view('admin/pages/subscriptions', compact(['subscriptions', 'subscriptions_count']));
}
// DELETE
public function destroy(Subscription $email)
{
$email->delete();
return redirect()->back()->with('db_feedback', 'The email has been removed');
}
public function unsubscribe(Request $request)
{
$subscription = Subscription::where('email', $request->email);
if ($subscription->exists()) {
$subscription->delete();
return back()->with('success', 'You email has been successfully removed');
} else {
return back()->with('error', 'This email is not on our list');
}
}
}
<file_sep>/tests/Unit/CategoriesTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class CategoriesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function a_category_can_have_many_articles()
{
$category = $this->category;
$article = $this->article;
$second_article = factory('App\Article')->create([
'category_id' => $category->id
]);
$this->assertEquals(2, count($category->articles));
}
/** @test */
public function guests_can_browse_through_all_breaks_from_a_category()
{
$category = $this->category;
$article_one = $this->article;
$article_two = factory('App\Article')->create([
'category_id' => $category->id
]);
$article_three = factory('App\Article')->create([
'category_id' => $category->id
]);
$this->get("{$category->paths()->route()}")
->assertSee($article_one->title)
->assertSee($article_two->title)
->assertSee($article_three->title);
}
}
<file_sep>/resources/assets/js/app.js
require('./bootstrap');
$.ajaxSetup({
headers: {
'X-CSRF-TOKEN': $('meta[name="csrf-token"]').attr('content')
}
});
require('./alertBox');
require('./overlay');
require('./contactInputs');
if( /Android|webOS|iPhone|iPad|iPod|BlackBerry|IEMobile|Opera Mini/i.test(navigator.userAgent) ) {
$('.dropdown').click(function() {
$clicked_menu = $(this).find('.dropdown-menu');
$('.dropdown-menu').not($clicked_menu).slideUp('fast');
$clicked_menu.slideToggle('fast');
})
}
$('form').submit(function(){
$(this).find(':input[type=submit]').prop('disabled', true).val('Working on it...');
});
<file_sep>/tests/Unit/SubscriptionsTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Tests\AppAssertions;
use Tests\TestingEmailsListener;
use Tests\MailManagement;
use Carbon\Carbon;
use Illuminate\Http\UploadedFile;
use Illuminate\Support\Facades\Storage;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class SubscriptionsTest extends TestCase
{
use DatabaseMigrations;
use MailManagement;
use AppAssertions;
/** @test */
public function a_guest_can_be_subscribed_when_submitting_a_question()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $faker->safeEmail,
'message' => $faker->paragraph,
'subscribe_me' => 'on'
];
$this->post('/contact/ask-a-question', $request);
$this->assertDatabaseHas('subscriptions', [
'email' => $request['email']
]);
}
/** @test */
public function a_guest_can_be_subscribed_when_inquiring_about_a_break()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $faker->safeEmail,
'news_from' => $faker->word,
'article_title' => $faker->sentence,
'author_name' => $faker->name,
'article_url' => $faker->url,
'message' => $faker->paragraph,
'subscribe_me' => 'on'
];
$this->post('/contact/break-inquiry', $request);
$this->assertDatabaseHas('subscriptions', [
'email' => $request['email']
]);
}
/** @test */
public function a_guest_can_be_subscribed_when_submitting_a_break()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'institution_email' => $faker->safeEmail,
'field_research' => $faker->word,
'research_institute' => $faker->word,
'original_article' => $faker->url,
'position' => $faker->word,
'file' => $file = UploadedFile::fake()->create('document.doc', 20),
'description' => $faker->sentence,
'message' => $faker->paragraph,
'subscribe_me' => 'on'
];
$this->post('/contact/submit-a-break', $request);
$this->assertDatabaseHas('subscriptions', [
'email' => $request['institution_email']
]);
}
/** @test */
public function new_subscriptions_are_ignored_if_duplicated()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $this->subscription->email,
'message' => $faker->paragraph,
'subscribe_me' => 'on'
];
$this->post('/contact/ask-a-question', $request);
$this->assertDatabaseHas('subscriptions', [
'email' => $request['email']
]);
}
/** @test */
public function a_guest_can_unsubscribe()
{
$email = $this->subscription->email;
$this->delete("/unsubscribe", [
'email' => $email
]);
$this->assertDatabaseMissing('subscriptions', [
'email' => $email
]);
}
}
<file_sep>/tests/Unit/GuestRoutesTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class GuestRoutesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function all_web_get_requests_work()
{
$routes = [
'/',
'/breaks/'.$this->category->slug.'/'.$this->article->slug,
'/breakers/'.$this->author->slug,
'/core-team/'.$this->manager->slug,
'/breaks/'.$this->category->slug,
'/tags/'.$this->tag->name,
'/search',
'/unsubscribe',
'/services/feed',
'/about',
'/mission',
'/the-team',
'/breakers',
'/partners',
'/information',
'/review-operations',
'/available-articles',
'/contact/ask-a-question',
'/contact/break-inquiry',
'/contact/submit-your-break',
'/contact/ask-a-question',
'/contact/break-inquiry',
'/contact/submit-your-break'
];
factory('App\Highlight', 10)->create();
check($this, $routes);
}
/** @test */
public function app_routes_work()
{
$routes = [
'/app/breaks',
'/app/picks',
'/app/suggestions',
'/app/highlights',
'/app/latest',
'/app/popular'
];
check($this, $routes);
}
}
<file_sep>/app/Http/Controllers/SearchController.php
<?php
namespace App\Http\Controllers;
use Illuminate\Http\Request;
use App\Article;
use App\Author;
class SearchController extends Controller
{
public function index(Request $request)
{
$input = $request->for;
$sort = ($request->sort) ? $request->sort : 'created_at';
$order = ($sort == 'title') ? 'ASC' : 'DESC';
$show = ($request->show) ? $request->show : 5;
$articles = Article::search($input)->orderBy($sort, $order)->published()->paginate($show);
return view("pages/search", compact(['articles', 'input']));
}
public function authors(Request $request)
{
if ($request->input != '') {
$results = Author::where('first_name', 'LIKE', '%'.$request->input.'%')
->orWhere('last_name', 'LIKE', '%'.$request->input.'%')
->take(10)
->get();
foreach ($results as $result) {
$result->url = $result->paths()->route();
$result->admin = '/admin/breakers/'.$result->slug.'/edit';
}
return $results;
}
}
public function articles(Request $request)
{
if ($request->input != '') {
$results = Article::where('title', 'LIKE', '%'.$request->input.'%')
->published()
->take(10)
->get();
foreach ($results as $result) {
$result->url = $result->paths()->route();
$result->admin = '/admin/breaks/'.$result->slug.'/edit';
}
return $results;
}
}
}
<file_sep>/database/migrations/2017_10_10_203620_create_articles_table.php
<?php
use Illuminate\Support\Facades\Schema;
use Illuminate\Database\Schema\Blueprint;
use Illuminate\Database\Migrations\Migration;
class CreateArticlesTable extends Migration
{
/**
* Run the migrations.
*
* @return void
*/
public function up()
{
Schema::create('articles', function (Blueprint $table) {
$table->increments('id');
// OK
$table->string('title')->unique();
$table->string('title_fr')->unique()->nullable();
$table->string('slug')->unique();
// ABSTRACT
$table->text('description')->nullable();
$table->text('description_fr')->nullable();
// COVERS - COVER - COVER_IMAGE
// https://oap.unige.ch/journals/public/journals/8/
$table->text('image_path')->nullable();
// MISSING (ADD ON THE BACKEND)
$table->string('image_caption', 255)->nullable();
$table->string('image_credits', 144)->nullable();
// MISSING (ADD ON THE BACKEND)
$table->text('content');
$table->text('content_fr')->nullable();
// ADD LATER (missing from xml)
$table->float('reading_time');
// CITATIONS - CITATION
$table->text('original_article');
// section_ref
$table->unsignedInteger('category_id');
// MISSING (ADD ON THE BACKEND)
$table->unsignedInteger('editor_id');
// issue_identification - VOLUME - NUMBER
$table->string('volume')->nullable();
$table->string('issue')->nullable();
// DOI (ADD https://doi.org/)
$table->string('doi');
$table->boolean('editor_pick')->default(0);
$table->boolean('highlight')->default(0);
$table->integer('views')->default(0);
// STAY AS IS
$table->timestamp('published_at')->nullable();
$table->timestamps();
// AUTOMATIC DOWNLOAD AND SAVE PDF FROM RICK'S API
});
}
// ADD DOI EDIT ON WEBSITE
/**
* Reverse the migrations.
*
* @return void
*/
public function down()
{
Schema::dropIfExists('articles');
}
}
<file_sep>/tests/Unit/FeedbacksTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Tests\AppAssertions;
use Tests\TestingEmailsListener;
use Tests\MailManagement;
use Carbon\Carbon;
use Illuminate\Http\UploadedFile;
use Illuminate\Support\Facades\Storage;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class FeedbacksTest extends TestCase
{
use DatabaseMigrations;
use MailManagement;
use AppAssertions;
/** @test */
public function a_new_breaker_receives_an_email_upon_registration()
{
$this->signIn();
$faker = \Faker\Factory::create();
$first_name = $faker->firstName;
$email = $faker->safeEmail;
$this->post('/admin/breakers', [
'first_name' => $first_name,
'last_name' => $faker->lastName,
'email' => $email,
'position' => $faker->word,
'research_institute' => $faker->word,
'field_research' => $faker->word,
'general_comments' => $faker->paragraph
]);
$this->seeEmailWasSent()->seeEmailTo($email)->seeEmailSubjectIs('Welcome to TheScienceBreaker!')->seeEmailContains("Hello $first_name");
}
/** @test */
public function a_guest_receives_an_email_when_asking_a_question()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $faker->safeEmail,
'message' => $faker->paragraph
];
$this->post('/contact/ask-a-question', $request)->assertSessionHas('contact');
$this->seeEmailWasSent()->seeEmailsSent(2);
$this->seeEmailTo($request['email'])
->seeEmailSubjectIs('Your message to TheScienceBreaker')
->seeEmailContains('Thank you for your contact');
}
/** @test */
public function a_guest_receives_an_email_when_inquiring_for_a_new_break()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'email' => $faker->safeEmail,
'news_from' => $faker->word,
'article_title' => $faker->sentence,
'author_name' => $faker->name,
'article_url' => $faker->url,
'message' => $faker->paragraph
];
$this->post('/contact/break-inquiry', $request)->assertSessionHas('contact');
$this->seeEmailWasSent()->seeEmailsSent(2);
$this->seeEmailTo($request['email'])
->seeEmailSubjectIs('Your Break inquiry')
->seeEmailContains('We have received your Break inquiry');
}
/** @test */
public function a_guest_receives_an_email_when_submitting_a_new_break()
{
$faker = \Faker\Factory::create();
$request = [
'first_name' => $faker->firstName,
'last_name' => $faker->lastName,
'institution_email' => $faker->safeEmail,
'field_research' => $faker->word,
'research_institute' => $faker->word,
'original_article' => $faker->url,
'position' => $faker->word,
'file' => $file = UploadedFile::fake()->create('document.doc', 20),
'description' => $faker->sentence,
'message' => $faker->paragraph
];
$this->post('/contact/submit-a-break', $request)->assertSessionHas('contact');
$this->seeEmailWasSent()->seeEmailsSent(2);
$this->seeEmailTo($request['institution_email'])
->seeEmailSubjectIs('Your Break has been submitted!')
->seeEmailContains('Thank you for submitting your Break');
}
// /** @test */
// public function breakers_and_the_editor_receive_an_email_when_their_new_break_is_published()
// {
// $this->signIn();
// $faker = \Faker\Factory::create();
// $breaker_one = factory('App\Author')->create();
// $breaker_two = factory('App\Author')->create();
// $editor = factory('App\Manager')->create([
// 'is_editor' => 1
// ]);
// $this->post('/admin/breaks', [
// 'title' => $faker->sentence,
// 'content' => '<p>'.$faker->paragraph.'</p>',
// 'authors' => [
// $breaker_one->id,
// $breaker_two->id
// ],
// 'reading_time' => '3.5',
// 'original_article' => $faker->url,
// 'category_id' => '1',
// 'editor_id' => $editor->id,
// 'editor_pick' => '0'
// ]);
// $this->seeEmailWasSent();
// $this->seeEmailTo($breaker_one->email)->seeEmailSubjectIs('Break published')->seeEmailContains("Congratulations $breaker_one->first_name");
// $this->seeEmailTo($breaker_two->email)->seeEmailSubjectIs('Break published')->seeEmailContains("Congratulations $breaker_two->first_name");
// $this->seeEmailTo($editor->email)->seeEmailContains("$editor->first_name");
// }
}
<file_sep>/tests/Unit/UserRoutesTest.php
<?php
namespace Tests\Unit;
use Tests\TestCase;
use Illuminate\Foundation\Testing\RefreshDatabase;
use Illuminate\Foundation\Testing\DatabaseMigrations;
class UserRoutesTest extends TestCase
{
use DatabaseMigrations;
/** @test */
public function all_web_get_requests_work()
{
$this->signIn();
$routes = [
'/admin/dashboard',
'/admin/breaks/add',
'/admin/breaks/edit',
'/admin/breaks/'.$this->article->slug.'/edit',
'/admin/breaks/delete',
'/admin/breakers/add',
'/admin/breakers/edit',
'/admin/breakers/'.$this->author->slug.'/edit',
'/admin/breakers/delete',
'/admin/managers/add',
'/admin/managers/edit',
'/admin/managers/'.$this->manager->slug.'/edit',
'/admin/managers/delete',
'/admin/available-articles',
'/admin/editor-picks',
'/admin/highlights',
'/admin/subscriptions',
'/admin/tags'
];
factory('App\Highlight', 10)->create();
check($this, $routes);
}
}
| 70257a968b836d232f6d5da267cd2a7716ff213b | [
"JavaScript",
"Text",
"Markdown",
"PHP"
] | 115 | PHP | tato-rj/thesciencebreaker | 05589cb4a27d4cd07fc04f08f13ab1479c48aae1 | f929b7f8cbb2d5d5f85bae84b28d02d6deb38496 |
refs/heads/master | <file_sep>function law_id(key_){
var searchArray = location.search.split('&');//获取URL的search部分并且以"&"为界限分割
var key = {};
var value = {};
for(var i=0;i < searchArray.length;i++)
{
if (searchArray[i].split('=')[0] ==key_)
{
console.log(searchArray[i].split('=')[1]);
return searchArray[i].split('=')[1]
}
}
}//获取用户的ID
<file_sep>
(function(){
document.addEventListener('DOMContentLoaded', init, false);
document.addEventListener('DOMContentLoaded', init1, false);
document.addEventListener('DOMContentLoaded', init2, false);
$(".certification_btn").click(function(){
certi_img();
});
$(".picker_btn").click(function(){
console.log($(this).val())
});
var base_1,base_2,base_3;
var yulan1,yulan2,yulan3;
function certi_img(){
var reg_start = new RegExp('=', 'g');
var reg_slash = new RegExp(';', 'g');
var date11 = $(".zhiyeren_").val();//获取当前的时间戳
var timestamp2 = Date.parse(new Date(date11));//转换时间戳
timestamp2 = timestamp2 / 1000;//计算时间戳
//var data = ("#picker-name").
var pic1 = $(".base_img0").text().replace(reg_start, '*').replace(reg_slash, '@');//个人信息
var pic2 = $(".base_img1").text().replace(reg_start, '*').replace(reg_slash, '@');//执业信息
var pic3 = $(".base_img2").text().replace(reg_start, '*').replace(reg_slash, '@');//年检认证
var cnt_name1 = $(".cnt_name1").text();//个人信息图片名字
var cnt_name2 = $(".cnt_name2").text();//执业图片名字
var cnt_name3 = $(".cnt_name3").text();//年检图片名字
if (pic1 != "" && pic2 != "" && pic3 != ""){
//var pic1 = "{\"file_type\":\"" + "img/jpeg" + "\",\"file_name\":\"" + cnt_name1 + "\",\"file_data\":\"" + base_img0 + "\"}";
//var pic2 = "{\"file_type\":\"" + "img/jpeg" + "\",\"file_name\":\"" + cnt_name2 + "\",\"file_data\":\"" + base_img1 + "\"}";
//var pic3 = "{\"file_type\":\"" + "img/jpeg" + "\",\"file_name\":\"" + cnt_name3 + "\",\"file_data\":\"" + base_img2 + "\"}";
/* var pic1 = String({"file_type": "img/jpeg","file_name": cnt_name1,"file_data": base_img0})
var pic2 = {"file_type": "img/jpeg","file_name": cnt_name2,"file_data": base_img1}
var pic3 = {"file_type": "img/jpeg","file_name": cnt_name3,"file_data": base_img2}*/
$.ajax({
type: "POST", //用POST方式传输
dataType: "JSON", //数据格式:JSON
url: "/lawyerhomepage/lawyer_cert?id=" + value_id,//value_id, //目标地址
data:"form_data_type=" + "lawyer_cert" + "&pic1=" + pic1 + "&pic2=" + pic2 + "&pic3=" + pic3 + "&cert_time=" + timestamp2,
beforeSend: function(request) {
request.setRequestHeader("X-CSRFToken", getcookie('csrftoken'));
},
error: function (XMLHttpRequest, textStatus, errorThrown) { },
success: function (result){
var result_str = lawyerhomepage(result);
if ( result.data.url != undefined && result.data.url != "" && result.data.url != "."){
$("#prompt").text(result_str).css({padding:".12rem .2rem .12rem .2rem"}).fadeToggle(2000,function(){
$("#prompt").text(result_str).fadeToggle(2000);
window.location.href = result.data.url;
});
}else {
$("#prompt").text(result_str).css({padding:".12rem .2rem .12rem .2rem"}).fadeToggle(2000);
}
}
});
}else {
console.log(2)
}
};
function init() {
var u = new UploadPic();
u.init({
input: document.querySelector('.input'),
callback: function (base64) {
var html = '';
html = '<div class="itm"><div class="tit">图片名称:</div><div class="cnt_name1">' + this.fileName + '</div></div>'
/*+ '<div class="itm"><div class="tit">原始大小:</div><div class="cnt">' + (this.fileSize / 1024).toFixed(2) + 'KB<\/div><\/div>'
+ '<div class="itm"><div class="tit">编码大小:</div><div class="cnt">' + (base64.length / 1024).toFixed(2) + 'KB<\/div><\/div>'
+ '<div class="itm"><div class="tit">原始尺寸:</div><div class="cnt">' + this.tw + 'px * ' + this.th + 'px<\/div><\/div>'
+ '<div class="itm"><div class="tit">编码尺寸:</div><div class="cnt">' + this.sw + 'px * ' + this.sh + 'px<\/div><\/div>'*/
+ '<div class="itm">' + '<div class="cnt_img"><img src="' + base64 + '"><\/div><\/div>'
+ '<div class="line_aa"></div>'
+'<p class="yulan1 yulan_style">预览</p>'
+ '<div class="itm_bt641"><div class="tit">Base64编码:</div><div class="cnt"><textarea class="base_img0">' + base64 + '<\/textarea><\/div><\/div>';
document.querySelector('.imgzip').innerHTML = html;
$(".liezi_img1").css({display:"none"});
base_1 = base64;//赋值
yulan1 = $(".yulan1");
yulan1.click(function(){yilan_click(base_1);});
//yilan_click(base64);//预览
}
});
}//个人信息
function init1() {
var input1 = document.getElementsByClassName("input1")[0];
var imgzip1 = document.getElementsByClassName("imgzip1")[0];
var u = new UploadPic();
u.init({
input: input1,
callback: function (base641) {
var html = '';
html = '<div class="itm"><div class="tit">图片名称:</div><div class="cnt_name2">' + this.fileName + '</div></div>'
/*+ '<div class="itm"><div class="tit">原始大小:</div><div class="cnt">' + (this.fileSize / 1024).toFixed(2) + 'KB<\/div><\/div>'
+ '<div class="itm"><div class="tit">编码大小:</div><div class="cnt">' + (base64.length / 1024).toFixed(2) + 'KB<\/div><\/div>'
+ '<div class="itm"><div class="tit">原始尺寸:</div><div class="cnt">' + this.tw + 'px * ' + this.th + 'px<\/div><\/div>'
+ '<div class="itm"><div class="tit">编码尺寸:</div><div class="cnt">' + this.sw + 'px * ' + this.sh + 'px<\/div><\/div>'*/
+ '<div class="itm">' + '<div class="cnt_img"><img src="' + base641 + '"><\/div><\/div>'
+ '<div class="line_aa"></div>'
+'<p class="yulan2 yulan_style">预览</p>'
+ '<div class="itm_bt641"><div class="tit">Base64编码:</div><div class="cnt"><textarea class="base_img1">' + base641 + '<\/textarea><\/div><\/div>';
imgzip1.innerHTML = html;
$(".liezi_img2").css({display:"none"});
base_2 = base641;//赋值
yulan2 = $(".yulan2");
yulan2.click(function(){yilan_click(base_2);});
}
});
}//执业认证
function init2() {
var input2 = document.getElementsByClassName("input2")[0];
var imgzip2 = document.getElementsByClassName("imgzip2")[0];
var u = new UploadPic();
u.init({
input: input2,
callback: function (base642) {
var html = '';
html = '<div class="itm"><div class="tit">图片名称:</div><div class="cnt_name3">' + this.fileName + '</div></div>'
/*+ '<div class="itm"><div class="tit">原始大小:</div><div class="cnt">' + (this.fileSize / 1024).toFixed(2) + 'KB<\/div><\/div>'
+ '<div class="itm"><div class="tit">编码大小:</div><div class="cnt">' + (base64.length / 1024).toFixed(2) + 'KB<\/div><\/div>'
+ '<div class="itm"><div class="tit">原始尺寸:</div><div class="cnt">' + this.tw + 'px * ' + this.th + 'px<\/div><\/div>'
+ '<div class="itm"><div class="tit">编码尺寸:</div><div class="cnt">' + this.sw + 'px * ' + this.sh + 'px<\/div><\/div>'*/
+ '<div class="itm">' + '<div class="cnt_img"><img src="' + base642 + '"><\/div><\/div>'
+ '<div class="line_aa"></div>'
+'<p class="yulan3 yulan_style">预览</p>'
+ '<div class="itm_bt641"><div class="tit">Base64编码:</div><div class="cnt"><textarea class="base_img2">' + base642 + '<\/textarea><\/div><\/div>';
imgzip2.innerHTML = html;
$(".liezi_img3").css({display:"none"});
base_3 = base642;//赋值
yulan3 = $(".yulan3");
yulan3.click(function(){yilan_click(base_3);});
}
});
}//执业认证
//***********************************************************
function UploadPic() {
this.sw = 0;
this.sh = 0;
this.tw = 0;
this.th = 0;
this.scale = 0;
this.maxWidth = 0;
this.maxHeight = 0;
this.maxSize = 0;
this.fileSize = 0;
this.fileDate = null;
this.fileType = '';
this.fileName = '';
this.input = null;
this.canvas = null;
this.mime = {};
this.type = '';
this.callback = function () {};
this.loading = function () {};
}
UploadPic.prototype.init = function (options) {
this.maxWidth = options.maxWidth || 800;
this.maxHeight = options.maxHeight || 600;
this.maxSize = options.maxSize || 3 * 1024 * 1024;
this.input = options.input;
this.mime = {'png': 'image/png', 'jpg': 'image/jpeg', 'jpeg': 'image/jpeg', 'bmp': 'image/bmp'};
this.callback = options.callback || function () {};
this.loading = options.loading || function () {};
this._addEvent();
};
/**
* @description 绑定事件
* @param {Object} elm 元素
* @param {Function} fn 绑定函数
*/
UploadPic.prototype._addEvent = function () {
var _this = this;
function tmpSelectFile(ev) {
_this._handelSelectFile(ev);
}
this.input.addEventListener('change', tmpSelectFile, false);
};
/**
* @description 绑定事件
* @param {Object} elm 元素
* @param {Function} fn 绑定函数
*/
UploadPic.prototype._handelSelectFile = function (ev) {
var file = ev.target.files[0];
this.type = file.type;
// 如果没有文件类型,则通过后缀名判断(解决微信及360浏览器无法获取图片类型问题)
if (!this.type) {
this.type = this.mime[file.name.match(/\.([^\.]+)$/i)[1]];
}
if (!/image.(png|jpg|jpeg|bmp)/.test(this.type)) {
alert('选择的文件类型不是图片');
return;
}
/* if (file.size > this.maxSize) {
alert('选择文件大于' + this.maxSize / 1024 / 1024 + 'M,请重新选择');
return;
}*/
this.fileName = file.name;
this.fileSize = file.size;
this.fileType = this.type;
this.fileDate = file.lastModifiedDate;
this._readImage(file);
};
/**
* @description 读取图片文件
* @param {Object} image 图片文件
*/
UploadPic.prototype._readImage = function (file) {
var _this = this;
function tmpCreateImage(uri) {
_this._createImage(uri);
}
this.loading();
this._getURI(file, tmpCreateImage);
};
/**
* @description 通过文件获得URI
* @param {Object} file 文件
* @param {Function} callback 回调函数,返回文件对应URI
* return {Bool} 返回false
*/
UploadPic.prototype._getURI = function (file, callback) {
var reader = new FileReader();
var _this = this;
function tmpLoad() {
// 头不带图片格式,需填写格式
var re = /^data:base64,/;
var ret = this.result + '';
if (re.test(ret)) ret = ret.replace(re, 'data:' + _this.mime[_this.fileType] + ';base64,');
callback && callback(ret);
}
reader.onload = tmpLoad;
reader.readAsDataURL(file);
return false;
};
/**
* @description 创建图片
* @param {Object} image 图片文件
*/
UploadPic.prototype._createImage = function (uri) {
var img = new Image();
var _this = this;
function tmpLoad() {
_this._drawImage(this);
}
img.onload = tmpLoad;
img.src = uri;
};
/**
* @description 创建Canvas将图片画至其中,并获得压缩后的文件
* @param {Object} img 图片文件
* @param {Number} width 图片最大宽度
* @param {Number} height 图片最大高度
* @param {Function} callback 回调函数,参数为图片base64编码
* return {Object} 返回压缩后的图片
*/
UploadPic.prototype._drawImage = function (img, callback) {
this.sw = img.width;
this.sh = img.height;
this.tw = img.width;
this.th = img.height;
this.scale = (this.tw / this.th).toFixed(2);
if (this.sw > this.maxWidth) {
this.sw = this.maxWidth;
this.sh = Math.round(this.sw / this.scale);
}
if (this.sh > this.maxHeight) {
this.sh = this.maxHeight;
this.sw = Math.round(this.sh * this.scale);
}
this.canvas = document.createElement('canvas');
var ctx = this.canvas.getContext('2d');
this.canvas.width = this.sw;
this.canvas.height = this.sh;
ctx.drawImage(img, 0, 0, img.width, img.height, 0, 0, this.sw, this.sh);
this.callback(this.canvas.toDataURL(this.type));
ctx.clearRect(0, 0, this.tw, this.th);
this.canvas.width = 0;
this.canvas.height = 0;
this.canvas = null;
};
function yilan_click(base64_img){
//$(".yulan").click(function(){
$(".main_enlarge").css({display:"block"});
$(".grenxingxi").attr({src:base64_img});
//})
}
$(".main_enlarge").click(function(){
main_click();
});
function main_click(){
$(".main_enlarge").css({display:"none"});
}
$(".shili1 ").click(function(){
$(".main_enlarge").css({display:"block"});
$(".grenxingxi").attr({src:"img/Column/shili1.png",alt:"个人信息"});
});
$(".shili2 ").click(function(){
$(".main_enlarge").css({display:"block"});
$(".grenxingxi").attr({src:"img/Column/shili2.png",alt:"执业信息"});
});
$(".shili3 ").click(function(){
$(".main_enlarge").css({display:"block"});
$(".grenxingxi").attr({src:"img/Column/shili3.png",alt:"年检认证"});
});
$(".loading1111").fadeOut();//loding
}());//loading
| ed4572be5d9cc15720343019610ea40044510398 | [
"JavaScript"
] | 2 | JavaScript | lw664256297/tupianshangchuan | 7e4cfb58c8535db62cdd123ecc7d42b880ea55e1 | 934aa14d52f38a5aba09e349fa09d4543da8554e |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.