| --- |
| license: bsd-3-clause |
| library_name: braindecode |
| pipeline_tag: feature-extraction |
| tags: |
| - eeg |
| - biosignal |
| - pytorch |
| - neuroscience |
| - braindecode |
| - convolutional |
| - transformer |
| --- |
| |
| # SSTDPN |
|
|
| SSTDPN from Can Han et al (2025) [Han2025]. |
|
|
| > **Architecture-only repository.** Documents the |
| > `braindecode.models.SSTDPN` class. **No pretrained weights are |
| > distributed here.** Instantiate the model and train it on your own |
| > data. |
|
|
| ## Quick start |
|
|
| ```bash |
| pip install braindecode |
| ``` |
|
|
| ```python |
| from braindecode.models import SSTDPN |
| |
| model = SSTDPN( |
| n_chans=22, |
| sfreq=250, |
| input_window_seconds=4.0, |
| n_outputs=4, |
| ) |
| ``` |
|
|
| The signal-shape arguments above are illustrative defaults — adjust to |
| match your recording. |
|
|
| ## Documentation |
| - Full API reference: <https://braindecode.org/stable/generated/braindecode.models.SSTDPN.html> |
| - Interactive browser (live instantiation, parameter counts): |
| <https://huggingface.co/spaces/braindecode/model-explorer> |
| - Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/sstdpn.py#L17> |
|
|
|
|
| ## Architecture |
|
|
| 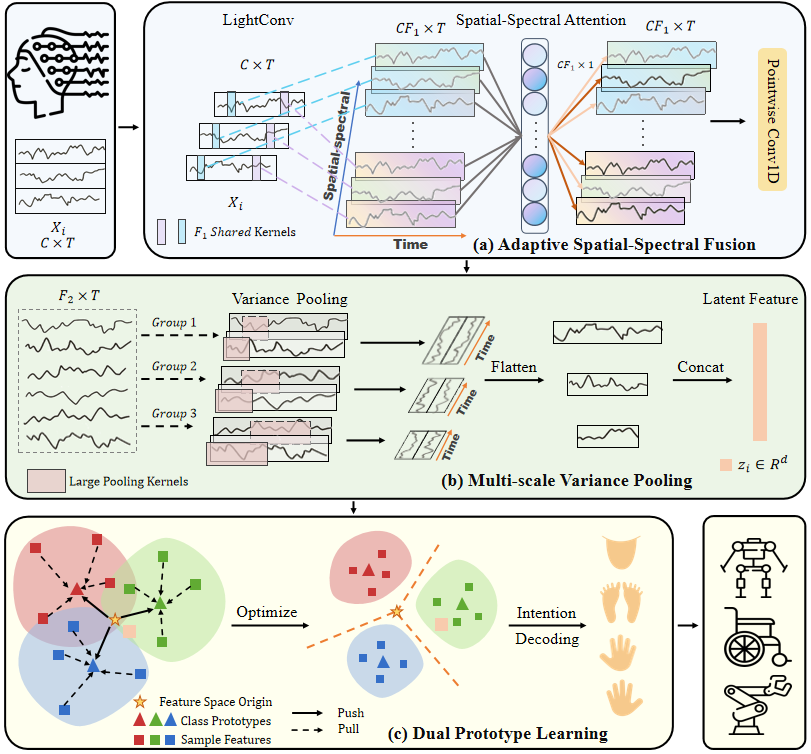 |
|
|
|
|
| ## Parameters |
|
|
| | Parameter | Type | Description | |
| |---|---|---| |
| | `n_spectral_filters_temporal` | int, optional | Number of spectral filters extracted per channel via temporal convolution. These represent the temporal spectral bands (equivalent to :math:`F_1` in the paper). Default is 9. | |
| | `n_fused_filters` | int, optional | Number of output filters after pointwise fusion convolution. These fuse the spectral filters across all channels (equivalent to :math:`F_2` in the paper). Default is 48. | |
| | `temporal_conv_kernel_size` | int, optional | Kernel size for the temporal convolution layer. Controls the receptive field for extracting spectral information. Default is 75 samples. | |
| | `mvp_kernel_sizes` | list[int], optional | Kernel sizes for Multi-scale Variance Pooling (MVP) module. Larger kernels capture long-term temporal dependencies . | |
| | `return_features` | bool, optional | If True, the forward pass returns (features, logits). If False, returns only logits. Default is False. | |
| | `proto_sep_maxnorm` | float, optional | Maximum L2 norm constraint for Inter-class Separation Prototypes during forward pass. This constraint acts as an implicit force to push features away from the origin. Default is 1.0. | |
| | `proto_cpt_std` | float, optional | Standard deviation for Intra-class Compactness Prototype initialization. Default is 0.01. | |
| | `spt_attn_global_context_kernel` | int, optional | Kernel size for global context embedding in Spatial-Spectral Attention module. Default is 250 samples. | |
| | `spt_attn_epsilon` | float, optional | Small epsilon value for numerical stability in Spatial-Spectral Attention. Default is 1e-5. | |
| | `spt_attn_mode` | str, optional | Embedding computation mode for Spatial-Spectral Attention ('var', 'l2', or 'l1'). Default is 'var' (variance-based mean-var operation). | |
| | `activation` | nn.Module, optional | Activation function to apply after the pointwise fusion convolution in :class:`_SSTEncoder`. Should be a PyTorch activation module class. Default is nn.ELU. | |
|
|
|
|
| ## References |
|
|
| 1. Han, C., Liu, C., Wang, J., Wang, Y., Cai, C., & Qian, D. (2025). A spatial–spectral and temporal dual prototype network for motor imagery brain–computer interface. Knowledge-Based Systems, 315, 113315. |
| 2. Han, C., Liu, C., Wang, J., Wang, Y., Cai, C., & Qian, D. (2025). A spatial–spectral and temporal dual prototype network for motor imagery brain–computer interface. Knowledge-Based Systems, 315, 113315. GitHub repository. https://github.com/hancan16/SST-DPN. |
|
|
|
|
| ## Citation |
|
|
| Cite the original architecture paper (see *References* above) and braindecode: |
|
|
| ```bibtex |
| @article{aristimunha2025braindecode, |
| title = {Braindecode: a deep learning library for raw electrophysiological data}, |
| author = {Aristimunha, Bruno and others}, |
| journal = {Zenodo}, |
| year = {2025}, |
| doi = {10.5281/zenodo.17699192}, |
| } |
| ``` |
|
|
| ## License |
|
|
| BSD-3-Clause for the model code (matching braindecode). |
| Pretraining-derived weights, if you fine-tune from a checkpoint, |
| inherit the licence of that checkpoint and its training corpus. |
|
|