metadata
license: bsd-3-clause
library_name: braindecode
pipeline_tag: feature-extraction
tags:
- eeg
- biosignal
- pytorch
- neuroscience
- braindecode
- foundation-model
- convolutional
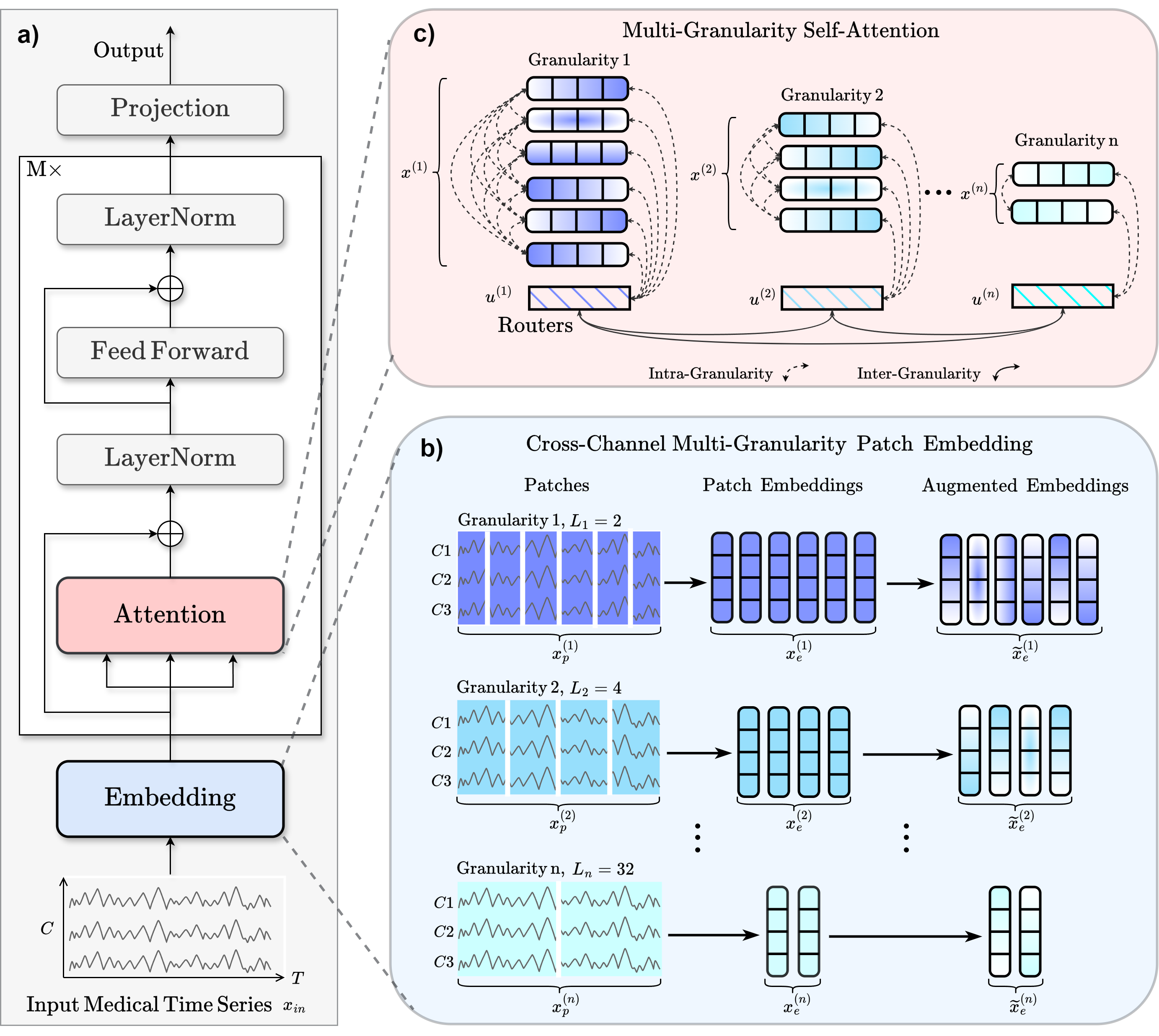
MEDFormer
Medformer from Wang et al (2024) [Medformer2024].
Architecture-only repository. Documents the
braindecode.models.MEDFormerclass. No pretrained weights are distributed here. Instantiate the model and train it on your own data.
Quick start
pip install braindecode
from braindecode.models import MEDFormer
model = MEDFormer(
n_chans=22,
sfreq=250,
input_window_seconds=4.0,
n_outputs=4,
)
The signal-shape arguments above are illustrative defaults — adjust to match your recording.
Documentation
- Full API reference: https://braindecode.org/stable/generated/braindecode.models.MEDFormer.html
- Interactive browser (live instantiation, parameter counts): https://huggingface.co/spaces/braindecode/model-explorer
- Source on GitHub: https://github.com/braindecode/braindecode/blob/master/braindecode/models/medformer.py#L20
Architecture
Parameters
| Parameter | Type | Description |
|---|---|---|
patch_len_list |
list of int, optional | Patch lengths for multi-granularity patching; each entry selects a temporal scale. The default is [14, 44, 45]. |
embed_dim |
int, optional | Embedding dimensionality. The default is 128. |
num_heads |
int, optional | Number of attention heads, which must divide :attr:d_model. The default is 8. |
drop_prob |
float, optional | Dropout probability. The default is 0.1. |
no_inter_attn |
bool, optional | If True, disables inter-granularity attention. The default is False. |
num_layers |
int, optional | Number of encoder layers. The default is 6. |
dim_feedforward |
int, optional | Feedforward dimensionality. The default is 256. |
activation_trans |
nn.Module, optional | Activation module used in transformer encoder layers. The default is :class:nn.ReLU. |
single_channel |
bool, optional | If True, processes each channel independently, increasing capacity and cost. The default is False. |
output_attention |
bool, optional | If True, returns attention weights for interpretability. The default is True. |
activation_class |
nn.Module, optional | Activation used in the final classification layer. The default is :class:nn.GELU. |
References
- Wang, Y., Huang, N., Li, T., Yan, Y., & Zhang, X. (2024). Medformer: A Multi-Granularity Patching Transformer for Medical Time-Series Classification. In A. Globerson, L. Mackey, D. Belgrave, A. Fan, U. Paquet, J. Tomczak, & C. Zhang (Eds.), Advances in Neural Information Processing Systems (Vol. 37, pp. 36314-36341). doi:10.52202/079017-1145.
Citation
Cite the original architecture paper (see References above) and braindecode:
@article{aristimunha2025braindecode,
title = {Braindecode: a deep learning library for raw electrophysiological data},
author = {Aristimunha, Bruno and others},
journal = {Zenodo},
year = {2025},
doi = {10.5281/zenodo.17699192},
}
License
BSD-3-Clause for the model code (matching braindecode). Pretraining-derived weights, if you fine-tune from a checkpoint, inherit the licence of that checkpoint and its training corpus.