| --- |
| license: bsd-3-clause |
| library_name: braindecode |
| pipeline_tag: feature-extraction |
| tags: |
| - eeg |
| - biosignal |
| - pytorch |
| - neuroscience |
| - braindecode |
| - foundation-model |
| - transformer |
| --- |
| |
| # CBraMod |
|
|
| **C**\ riss-\ **C**\ ross **Bra**\ in **Mod**\ el for EEG Decoding from Wang et al. (2025) [cbramod]. |
|
|
| > **Architecture-only repository.** Documents the |
| > `braindecode.models.CBraMod` class. **No pretrained weights are |
| > distributed here.** Instantiate the model and train it on your own |
| > data. |
|
|
| ## Quick start |
|
|
| ```bash |
| pip install braindecode |
| ``` |
|
|
| ```python |
| from braindecode.models import CBraMod |
| |
| model = CBraMod( |
| n_chans=22, |
| sfreq=200, |
| input_window_seconds=4.0, |
| n_outputs=2, |
| ) |
| ``` |
|
|
| The signal-shape arguments above are illustrative defaults — adjust to |
| match your recording. |
|
|
| ## Documentation |
| - Full API reference: <https://braindecode.org/stable/generated/braindecode.models.CBraMod.html> |
| - Interactive browser (live instantiation, parameter counts): |
| <https://huggingface.co/spaces/braindecode/model-explorer> |
| - Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/cbramod.py#L23> |
|
|
|
|
| ## Architecture |
|
|
| 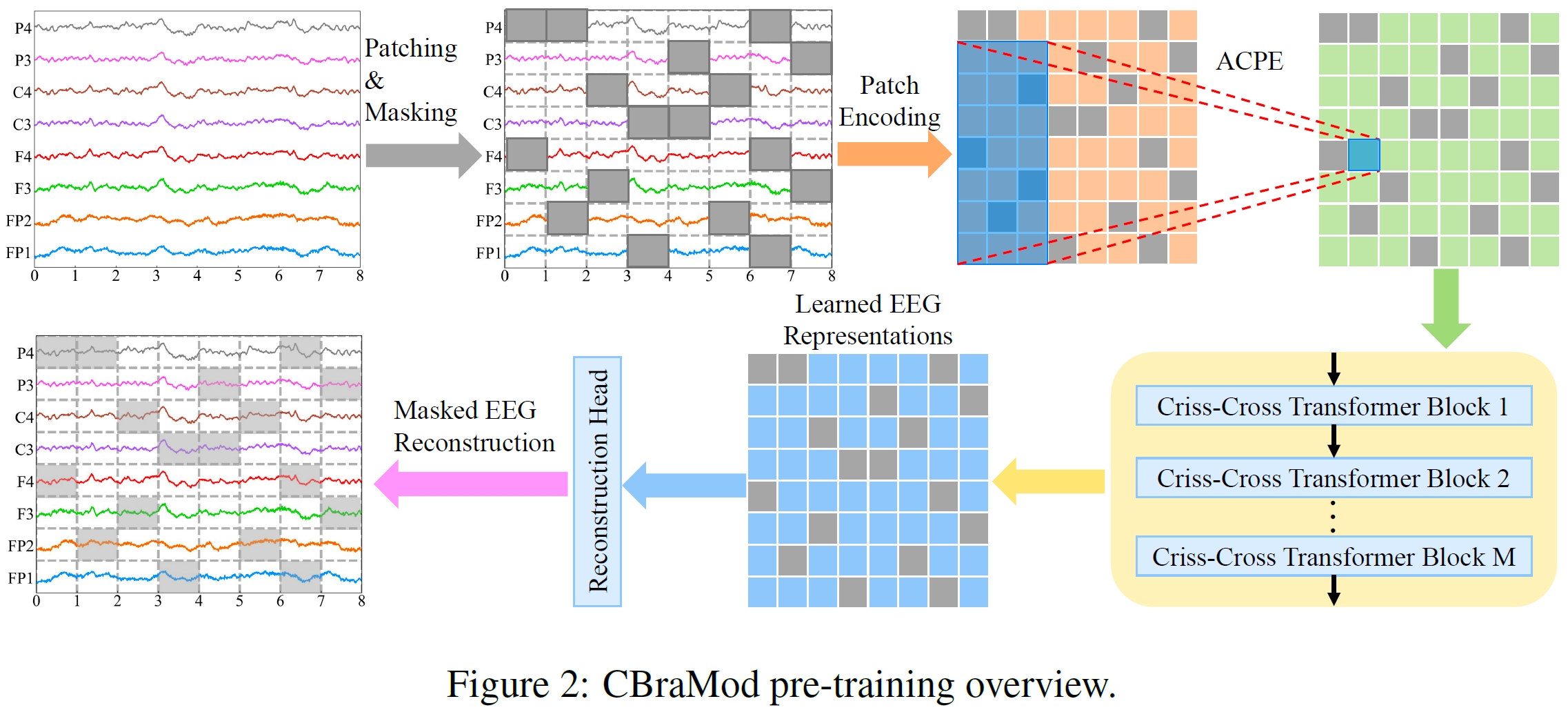 |
|
|
|
|
| ## Parameters |
|
|
| | Parameter | Type | Description | |
| |---|---|---| |
| | `patch_size` | int, default=200 | Temporal patch size in samples (200 samples = 1 second at 200 Hz). | |
| | `dim_feedforward` | int, default=800 | Dimension of the feedforward network in Transformer layers. | |
| | `n_layer` | int, default=12 | Number of Transformer layers. | |
| | `nhead` | int, default=8 | Number of attention heads. | |
| | `activation` | type[nn.Module], default=nn.GELU | Activation function used in Transformer feedforward layers. | |
| | `emb_dim` | int, default=200 | Output embedding dimension. | |
| | `drop_prob` | float, default=0.1 | Dropout probability. | |
| | `return_encoder_output` | bool, default=False | If false (default), the features are flattened and passed through a final linear layer to produce class logits of size `n_outputs`. If True, the model returns the encoder output features. | |
|
|
|
|
| ## References |
|
|
| 1. Wang, J., Zhao, S., Luo, Z., Zhou, Y., Jiang, H., Li, S., Li, T., & Pan, G. (2025). CBraMod: A Criss-Cross Brain Foundation Model for EEG Decoding. In The Thirteenth International Conference on Learning Representations (ICLR 2025). https://arxiv.org/abs/2412.07236 |
|
|
|
|
| ## Citation |
|
|
| Cite the original architecture paper (see *References* above) and braindecode: |
|
|
| ```bibtex |
| @article{aristimunha2025braindecode, |
| title = {Braindecode: a deep learning library for raw electrophysiological data}, |
| author = {Aristimunha, Bruno and others}, |
| journal = {Zenodo}, |
| year = {2025}, |
| doi = {10.5281/zenodo.17699192}, |
| } |
| ``` |
|
|
| ## License |
|
|
| BSD-3-Clause for the model code (matching braindecode). |
| Pretraining-derived weights, if you fine-tune from a checkpoint, |
| inherit the licence of that checkpoint and its training corpus. |
|
|