File size: 6,074 Bytes
a84bd58 6d1fcc2 a84bd58 6d1fcc2 a84bd58 6d1fcc2 a84bd58 6d1fcc2 a84bd58 6d1fcc2 a84bd58 6d1fcc2 a84bd58 6d1fcc2 a84bd58 | 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 | ---
license: bsd-3-clause
library_name: braindecode
pipeline_tag: feature-extraction
tags:
- eeg
- biosignal
- pytorch
- neuroscience
- braindecode
- convolutional
- transformer
---
# MetaNeuromotorHand
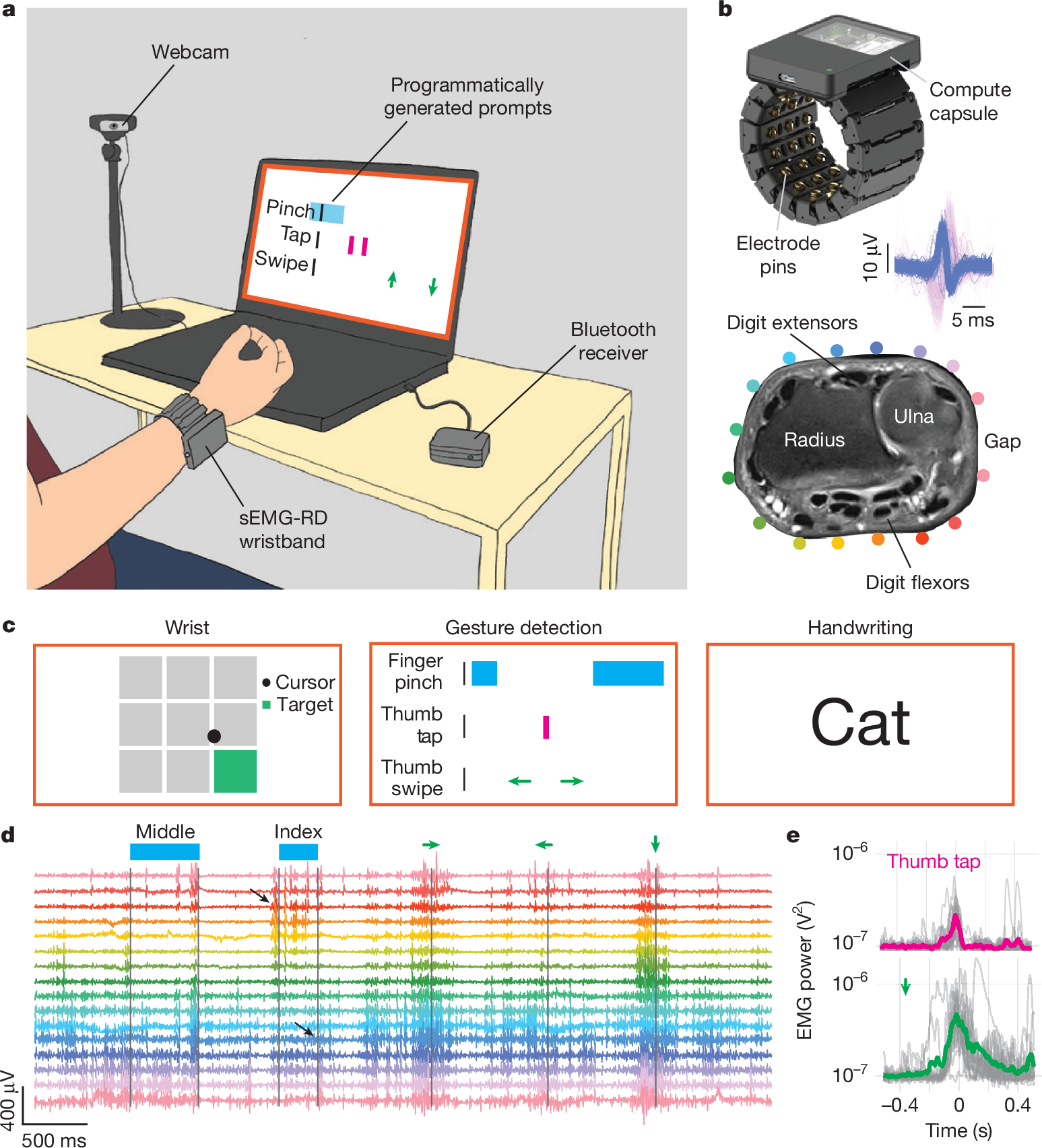
Generic neuromotor interface for handwriting from Meta (2025) [gni2025].
> **Architecture-only repository.** Documents the
> `braindecode.models.MetaNeuromotorHand` class. **No pretrained weights are
> distributed here.** Instantiate the model and train it on your own
> data.
## Quick start
```bash
pip install braindecode
```
```python
from braindecode.models import MetaNeuromotorHand
model = MetaNeuromotorHand(
n_chans=22,
sfreq=250,
input_window_seconds=4.0,
n_outputs=4,
)
```
The signal-shape arguments above are illustrative defaults — adjust to
match your recording.
## Documentation
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.MetaNeuromotorHand.html>
- Interactive browser (live instantiation, parameter counts):
<https://huggingface.co/spaces/braindecode/model-explorer>
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/meta_neuromotor.py#L34>
## Architecture
## Parameters
| Parameter | Type | Description |
|---|---|---|
| `n_outputs` | int | Vocabulary size for CTC. Defaults to `100` (handwriting charset). |
| `n_chans` | int | Number of EMG channels. Defaults to `16` (one armband). |
| `sfreq` | float | Sampling frequency in Hz. Defaults to `2000`. |
| `mpf_window_length` | int | MPF window length in samples. |
| `mpf_stride` | int | MPF frame stride in samples. |
| `mpf_n_fft` | int | STFT window / FFT size. |
| `mpf_fft_stride` | int | STFT hop size. Must divide `mpf_stride` and be `<= mpf_n_fft`. |
| `mpf_frequency_bins` | sequence of (float, float) or None | `(low, high)` Hz bands to average the cross-spectrum over. If `None`, all FFT frequency bins are used. |
| `mask_max_num_masks` | sequence of int | Max number of SpecAugment masks per dim (order matches `mask_dims`). |
| `mask_max_lengths` | sequence of int | Max mask length per dim (order matches `mask_dims`). |
| `mask_dims` | str | Axes to mask, among `"CFT"`. Defaults to `"TF"`. |
| `mask_value` | float | Filler value for masked regions. |
| `invariance_hidden_dims` | sequence of int | Hidden layer sizes of the per-rotation MLP. Output feature dim is `invariance_hidden_dims[-1]`. |
| `invariance_offsets` | sequence of int | Circular channel rotations to average over. |
| `num_adjacent_cov` | int | Number of adjacent off-diagonals of the cross-channel covariance matrix to keep. |
| `conformer_input_dim` | int | Conformer embedding dimension `D`. |
| `conformer_ffn_dim` | int | Feed-forward hidden dim inside each block. |
| `conformer_kernel_size` | int or sequence of int | Depthwise-conv kernel size per block. |
| `conformer_stride` | int or sequence of int | Depthwise-conv stride per block. As a scalar, applied only to the last block (entire encoder downsamples by `stride`); as a sequence of length `conformer_num_layers`, applied per block. Defaults to the paper's 15-layer schedule `(1, 1, 1, 1, 2) * 2 + (1,) * 5` (2x downsampling at blocks 5 and 10). When overriding `conformer_num_layers`, also pass a matching schedule or a scalar. |
| `conformer_num_heads` | int | Number of attention heads. |
| `conformer_attn_window_size` | int or sequence of int | Attention receptive field per block. Defaults to the paper's 15-layer schedule `(16,) * 10 + (8,) * 5`. When overriding `conformer_num_layers`, also pass a matching schedule or a scalar. |
| `conformer_num_layers` | int | Number of conformer blocks. |
| `drop_prob` | float | Dropout probability applied throughout the conformer (FFN, conv and attention blocks). |
| `time_reduction_stride` | int | Frame-stacking stride applied **before** the conformer. `1` disables it. |
| `log_softmax` | bool | If `True`, apply :func:`torch.nn.functional.log_softmax` to the emissions. Disabled by default (braindecode models return logits). |
| `activation` | type of nn.Module | Activation class used inside the conformer feed-forward and convolution blocks. Defaults to :class:`torch.nn.SiLU`. |
| `invariance_activation` | type of nn.Module | Activation class used inside the rotation-invariant MLP. Defaults to :class:`torch.nn.LeakyReLU`. |
## References
1. CTRL-labs at Reality Labs (Kaifosh, P., Reardon, T. R. et al.), 2025. A generic non-invasive neuromotor interface for human-computer interaction. Nature 645, 702-710. https://doi.org/10.1038/s41586-025-09255-w
2. Gulati, A. et al., 2020. Conformer: convolution-augmented transformer for speech recognition. Proc. Interspeech, 5036-5040.
3. Graves, A., Fernandez, S., Gomez, F., Schmidhuber, J., 2006. Connectionist temporal classification: labelling unsegmented sequence data with recurrent neural networks. Proc. ICML, 369-376.
4. Park, D. S. et al., 2019. SpecAugment: a simple data augmentation method for automatic speech recognition. Proc. Interspeech, 2613-2617.
5. Yu, J. et al., 2021. FastEmit: low-latency streaming ASR with sequence-level emission regularization. Proc. ICASSP.
6. Barachant, A., Barthelemy, Q., King, J.-R., Gramfort, A., Chevallier, S., Rodrigues, P. L. C., ... Aristimunha, B., 2026. pyRiemann (v0.10). Zenodo. https://doi.org/10.5281/zenodo.593816
## Citation
Cite the original architecture paper (see *References* above) and braindecode:
```bibtex
@article{aristimunha2025braindecode,
title = {Braindecode: a deep learning library for raw electrophysiological data},
author = {Aristimunha, Bruno and others},
journal = {Zenodo},
year = {2025},
doi = {10.5281/zenodo.17699192},
}
```
## License
BSD-3-Clause for the model code (matching braindecode).
Pretraining-derived weights, if you fine-tune from a checkpoint,
inherit the licence of that checkpoint and its training corpus.
|