Replace with clean markdown card
Browse files
README.md
CHANGED
|
@@ -9,19 +9,17 @@ tags:
|
|
| 9 |
- neuroscience
|
| 10 |
- braindecode
|
| 11 |
- foundation-model
|
| 12 |
-
- transformer
|
| 13 |
- sleep-staging
|
| 14 |
---
|
| 15 |
|
| 16 |
# BIOT
|
| 17 |
|
| 18 |
-
BIOT from Yang et al (2023)
|
| 19 |
|
| 20 |
-
> **Architecture-only repository.**
|
| 21 |
> `braindecode.models.BIOT` class. **No pretrained weights are
|
| 22 |
-
> distributed here**
|
| 23 |
-
> data
|
| 24 |
-
> separately.
|
| 25 |
|
| 26 |
## Quick start
|
| 27 |
|
|
@@ -40,184 +38,43 @@ model = BIOT(
|
|
| 40 |
)
|
| 41 |
```
|
| 42 |
|
| 43 |
-
The signal-shape arguments above are
|
| 44 |
-
|
| 45 |
|
| 46 |
## Documentation
|
| 47 |
-
|
| 48 |
-
-
|
| 49 |
-
<https://braindecode.org/stable/generated/braindecode.models.BIOT.html>
|
| 50 |
-
- Interactive browser with live instantiation:
|
| 51 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 52 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/biot.py#L56>
|
| 53 |
|
| 54 |
-
## Architecture description
|
| 55 |
-
|
| 56 |
-
The block below is the rendered class docstring (parameters,
|
| 57 |
-
references, architecture figure where available).
|
| 58 |
-
|
| 59 |
-
<div class='bd-doc'><main>
|
| 60 |
-
<p>BIOT from Yang et al (2023) [Yang2023]_</p>
|
| 61 |
-
<span style="display:inline-block;padding:2px 8px;border-radius:4px;background:#d9534f;color:white;font-size:11px;font-weight:600;margin-right:4px;">Foundation Model</span>
|
| 62 |
-
|
| 63 |
-
|
| 64 |
-
|
| 65 |
-
.. figure:: https://braindecode.org/dev/_static/model/biot.jpg
|
| 66 |
-
:align: center
|
| 67 |
-
:alt: BioT
|
| 68 |
-
|
| 69 |
-
BIOT: Cross-data Biosignal Learning in the Wild.
|
| 70 |
-
|
| 71 |
-
BIOT is a foundation model for biosignal classification. It is
|
| 72 |
-
a wrapper around the `BIOTEncoder` and `ClassificationHead` modules.
|
| 73 |
-
|
| 74 |
-
It is designed for N-dimensional biosignal data such as EEG, ECG, etc.
|
| 75 |
-
The method was proposed by Yang et al. [Yang2023]_ and the code is
|
| 76 |
-
available at [Code2023]_
|
| 77 |
-
|
| 78 |
-
The model is trained with a contrastive loss on large EEG datasets
|
| 79 |
-
TUH Abnormal EEG Corpus with 400K samples and Sleep Heart Health Study
|
| 80 |
-
5M. Here, we only provide the model architecture, not the pre-trained
|
| 81 |
-
weights or contrastive loss training.
|
| 82 |
-
|
| 83 |
-
The architecture is based on the `LinearAttentionTransformer` and
|
| 84 |
-
`PatchFrequencyEmbedding` modules.
|
| 85 |
-
The `BIOTEncoder` is a transformer that takes the input data and outputs
|
| 86 |
-
a fixed-size representation of the input data. More details are
|
| 87 |
-
present in the `BIOTEncoder` class.
|
| 88 |
-
|
| 89 |
-
The `ClassificationHead` is an ELU activation layer, followed by a simple
|
| 90 |
-
linear layer that takes the output of the `BIOTEncoder` and outputs
|
| 91 |
-
the classification probabilities.
|
| 92 |
-
|
| 93 |
-
.. important::
|
| 94 |
-
**Pre-trained Weights Available**
|
| 95 |
-
|
| 96 |
-
This model has pre-trained weights available on the Hugging Face Hub.
|
| 97 |
-
You can load them using:
|
| 98 |
-
|
| 99 |
-
.. code:: python
|
| 100 |
-
from braindecode.models import BIOT
|
| 101 |
-
|
| 102 |
-
# Load the original pre-trained model from Hugging Face Hub
|
| 103 |
-
# For 16-channel models:
|
| 104 |
-
model = BIOT.from_pretrained("braindecode/biot-pretrained-prest-16chs")
|
| 105 |
-
|
| 106 |
-
# For 18-channel models:
|
| 107 |
-
model = BIOT.from_pretrained("braindecode/biot-pretrained-shhs-prest-18chs")
|
| 108 |
-
model = BIOT.from_pretrained("braindecode/biot-pretrained-six-datasets-18chs")
|
| 109 |
-
|
| 110 |
-
To push your own trained model to the Hub:
|
| 111 |
-
|
| 112 |
-
.. code:: python
|
| 113 |
-
# After training your model
|
| 114 |
-
model.push_to_hub(
|
| 115 |
-
repo_id="username/my-biot-model", commit_message="Upload trained BIOT model"
|
| 116 |
-
)
|
| 117 |
-
|
| 118 |
-
Requires installing ``braindecode[hug]`` for Hub integration.
|
| 119 |
-
|
| 120 |
-
.. versionadded:: 0.9
|
| 121 |
-
|
| 122 |
-
Parameters
|
| 123 |
-
----------
|
| 124 |
-
embed_dim : int, optional
|
| 125 |
-
The size of the embedding layer, by default 256
|
| 126 |
-
num_heads : int, optional
|
| 127 |
-
The number of attention heads, by default 8
|
| 128 |
-
num_layers : int, optional
|
| 129 |
-
The number of transformer layers, by default 4
|
| 130 |
-
activation: nn.Module, default=nn.ELU
|
| 131 |
-
Activation function class to apply. Should be a PyTorch activation
|
| 132 |
-
module class like ``nn.ReLU`` or ``nn.ELU``. Default is ``nn.ELU``.
|
| 133 |
-
return_feature: bool, optional
|
| 134 |
-
Changing the output for the neural network. Default is single tensor
|
| 135 |
-
when return_feature is True, return embedding space too.
|
| 136 |
-
Default is False.
|
| 137 |
-
hop_length: int, optional
|
| 138 |
-
The hop length for the torch.stft transformation in the
|
| 139 |
-
encoder. The default is 100.
|
| 140 |
-
sfreq: int, optional
|
| 141 |
-
The sfreq parameter for the encoder. The default is 200
|
| 142 |
-
|
| 143 |
-
References
|
| 144 |
-
----------
|
| 145 |
-
.. [Yang2023] Yang, C., Westover, M.B. and Sun, J., 2023, November. BIOT:
|
| 146 |
-
Biosignal Transformer for Cross-data Learning in the Wild. In Thirty-seventh
|
| 147 |
-
Conference on Neural Information Processing Systems, NeurIPS.
|
| 148 |
-
.. [Code2023] Yang, C., Westover, M.B. and Sun, J., 2023. BIOT
|
| 149 |
-
Biosignal Transformer for Cross-data Learning in the Wild.
|
| 150 |
-
GitHub https://github.com/ycq091044/BIOT (accessed 2024-02-13)
|
| 151 |
-
|
| 152 |
-
.. rubric:: Hugging Face Hub integration
|
| 153 |
-
|
| 154 |
-
When the optional ``huggingface_hub`` package is installed, all models
|
| 155 |
-
automatically gain the ability to be pushed to and loaded from the
|
| 156 |
-
Hugging Face Hub. Install with::
|
| 157 |
-
|
| 158 |
-
pip install braindecode[hub]
|
| 159 |
-
|
| 160 |
-
**Pushing a model to the Hub:**
|
| 161 |
-
|
| 162 |
-
.. code::
|
| 163 |
-
from braindecode.models import BIOT
|
| 164 |
-
|
| 165 |
-
# Train your model
|
| 166 |
-
model = BIOT(n_chans=22, n_outputs=4, n_times=1000)
|
| 167 |
-
# ... training code ...
|
| 168 |
-
|
| 169 |
-
# Push to the Hub
|
| 170 |
-
model.push_to_hub(
|
| 171 |
-
repo_id="username/my-biot-model",
|
| 172 |
-
commit_message="Initial model upload",
|
| 173 |
-
)
|
| 174 |
-
|
| 175 |
-
**Loading a model from the Hub:**
|
| 176 |
-
|
| 177 |
-
.. code::
|
| 178 |
-
from braindecode.models import BIOT
|
| 179 |
-
|
| 180 |
-
# Load pretrained model
|
| 181 |
-
model = BIOT.from_pretrained("username/my-biot-model")
|
| 182 |
-
|
| 183 |
-
# Load with a different number of outputs (head is rebuilt automatically)
|
| 184 |
-
model = BIOT.from_pretrained("username/my-biot-model", n_outputs=4)
|
| 185 |
-
|
| 186 |
-
**Extracting features and replacing the head:**
|
| 187 |
|
| 188 |
-
|
| 189 |
-
import torch
|
| 190 |
|
| 191 |
-
|
| 192 |
-
# Extract encoder features (consistent dict across all models)
|
| 193 |
-
out = model(x, return_features=True)
|
| 194 |
-
features = out["features"]
|
| 195 |
|
| 196 |
-
# Replace the classification head
|
| 197 |
-
model.reset_head(n_outputs=10)
|
| 198 |
|
| 199 |
-
|
| 200 |
|
| 201 |
-
|
| 202 |
-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 203 |
|
| 204 |
-
config = model.get_config() # all __init__ params
|
| 205 |
-
with open("config.json", "w") as f:
|
| 206 |
-
json.dump(config, f)
|
| 207 |
|
| 208 |
-
|
| 209 |
|
| 210 |
-
|
| 211 |
-
|
| 212 |
-
saved to the Hub and restored when loading.
|
| 213 |
|
| 214 |
-
See :ref:`load-pretrained-models` for a complete tutorial.</main>
|
| 215 |
-
</div>
|
| 216 |
|
| 217 |
## Citation
|
| 218 |
|
| 219 |
-
|
| 220 |
-
*References* section above) and braindecode:
|
| 221 |
|
| 222 |
```bibtex
|
| 223 |
@article{aristimunha2025braindecode,
|
|
|
|
| 9 |
- neuroscience
|
| 10 |
- braindecode
|
| 11 |
- foundation-model
|
|
|
|
| 12 |
- sleep-staging
|
| 13 |
---
|
| 14 |
|
| 15 |
# BIOT
|
| 16 |
|
| 17 |
+
BIOT from Yang et al (2023) [Yang2023]
|
| 18 |
|
| 19 |
+
> **Architecture-only repository.** Documents the
|
| 20 |
> `braindecode.models.BIOT` class. **No pretrained weights are
|
| 21 |
+
> distributed here.** Instantiate the model and train it on your own
|
| 22 |
+
> data.
|
|
|
|
| 23 |
|
| 24 |
## Quick start
|
| 25 |
|
|
|
|
| 38 |
)
|
| 39 |
```
|
| 40 |
|
| 41 |
+
The signal-shape arguments above are illustrative defaults — adjust to
|
| 42 |
+
match your recording.
|
| 43 |
|
| 44 |
## Documentation
|
| 45 |
+
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.BIOT.html>
|
| 46 |
+
- Interactive browser (live instantiation, parameter counts):
|
|
|
|
|
|
|
| 47 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 48 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/biot.py#L56>
|
| 49 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 50 |
|
| 51 |
+
## Architecture
|
|
|
|
| 52 |
|
| 53 |
+
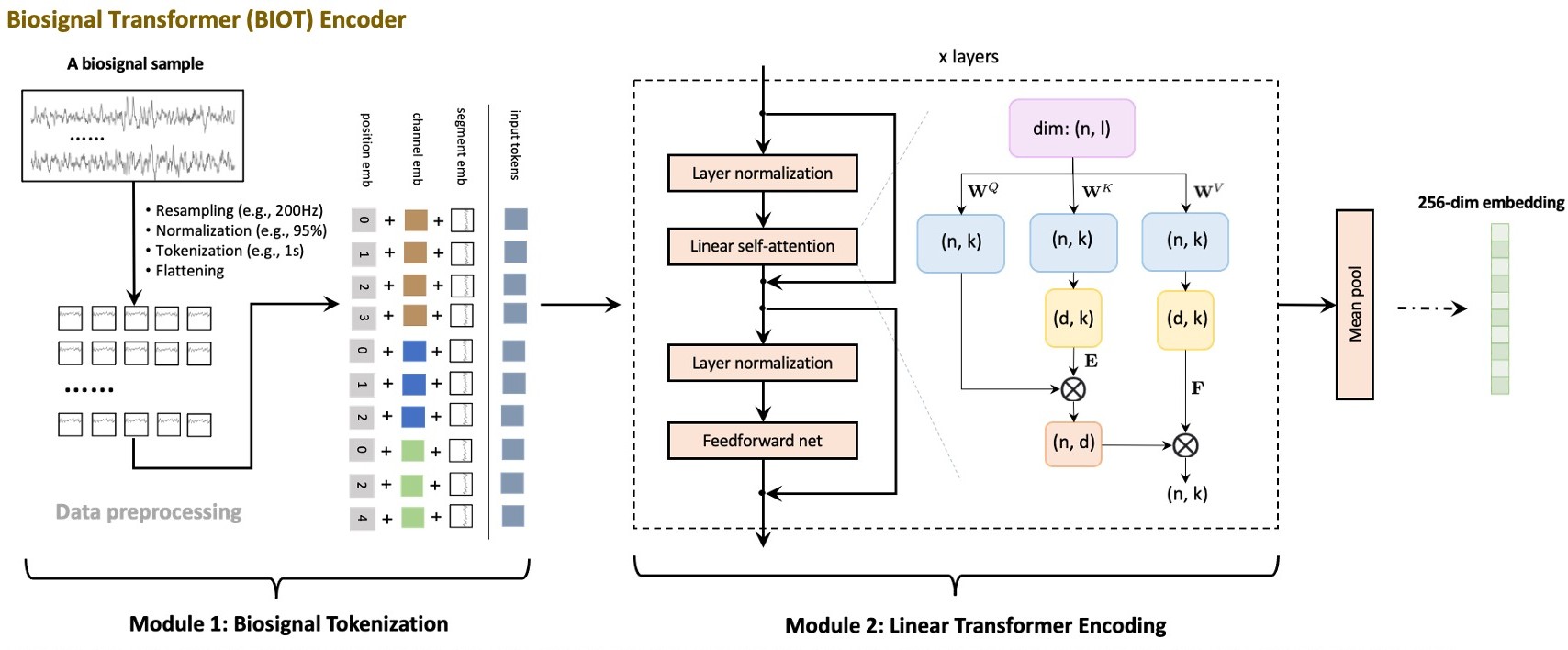
|
|
|
|
|
|
|
|
|
|
| 54 |
|
|
|
|
|
|
|
| 55 |
|
| 56 |
+
## Parameters
|
| 57 |
|
| 58 |
+
| Parameter | Type | Description |
|
| 59 |
+
|---|---|---|
|
| 60 |
+
| `embed_dim` | int, optional | The size of the embedding layer, by default 256 |
|
| 61 |
+
| `num_heads` | int, optional | The number of attention heads, by default 8 |
|
| 62 |
+
| `num_layers` | int, optional | The number of transformer layers, by default 4 |
|
| 63 |
+
| `activation: nn.Module, default=nn.ELU` | — | Activation function class to apply. Should be a PyTorch activation module class like `nn.ReLU` or `nn.ELU`. Default is `nn.ELU`. |
|
| 64 |
+
| `return_feature: bool, optional` | — | Changing the output for the neural network. Default is single tensor when return_feature is True, return embedding space too. Default is False. |
|
| 65 |
+
| `hop_length: int, optional` | — | The hop length for the torch.stft transformation in the encoder. The default is 100. |
|
| 66 |
+
| `sfreq: int, optional` | — | The sfreq parameter for the encoder. The default is 200 |
|
| 67 |
|
|
|
|
|
|
|
|
|
|
| 68 |
|
| 69 |
+
## References
|
| 70 |
|
| 71 |
+
1. Yang, C., Westover, M.B. and Sun, J., 2023, November. BIOT: Biosignal Transformer for Cross-data Learning in the Wild. In Thirty-seventh Conference on Neural Information Processing Systems, NeurIPS.
|
| 72 |
+
2. Yang, C., Westover, M.B. and Sun, J., 2023. BIOT Biosignal Transformer for Cross-data Learning in the Wild. GitHub https://github.com/ycq091044/BIOT (accessed 2024-02-13)
|
|
|
|
| 73 |
|
|
|
|
|
|
|
| 74 |
|
| 75 |
## Citation
|
| 76 |
|
| 77 |
+
Cite the original architecture paper (see *References* above) and braindecode:
|
|
|
|
| 78 |
|
| 79 |
```bibtex
|
| 80 |
@article{aristimunha2025braindecode,
|